Abstract
Traditional statistical approaches have advanced our understanding of the genetics of complex diseases, yet are limited to linear additive models. Here we applied machine learning (ML) to genome-wide data from 41,686 individuals in the largest European consortium on Alzheimer’s disease (AD) to investigate the effectiveness of various ML algorithms in replicating known findings, discovering novel loci, and predicting individuals at risk. We utilised Gradient Boosting Machines (GBMs), biological pathway-informed Neural Networks (NNs), and Model-based Multifactor Dimensionality Reduction (MB-MDR) models. ML approaches successfully captured all genome-wide significant genetic variants identified in the training set and 22% of associations from larger meta-analyses. They highlight 6 novel loci which replicate in an external dataset, including variants which map to ARHGAP25, LY6H, COG7, SOD1 and ZNF597. They further identify novel association in AP4E1, refining the genetic landscape of the known SPPL2A locus. Our results demonstrate that machine learning methods can achieve predictive performance comparable to classical approaches in genetic epidemiology and have the potential to uncover novel loci that remain undetected by traditional GWAS. These insights provide a complementary avenue for advancing the understanding of AD genetics.
Similar content being viewed by others
Introduction
Genome-wide association studies (GWAS) have enabled huge progress in identifying variants associated with the risk of developing Alzheimer’s disease (AD)1. Polygenic risk scores (PRS) based on these variants have greatly improved prediction of disease status2. However, inherent to GWAS and PRS are the assumptions that variants are independent predictors, linearly associated with the outcome, and therefore combine additively within and between loci3, with no interactions occurring between variants, or between genes and other risk factors. While such simplifying genetic assumptions have proved fruitful across a range of diseases and disorders4,5, they are at odds with biological evidence in AD that disease heterogeneity and responses from cells such as microglia are dependent on APOE status6,7,8,9. Further, there is genetic evidence suggesting that different variants are associated with the disease depending on APOE status10,11,12,13 and age at diagnosis or assessment14,15,16. As GWAS sample size increases and PRS approach limits on predictive performance, alternative modelling approaches are essential to maximise discoveries from existing data and enable a deeper understanding of AD genetics.
The confluence of increasingly large genetic data17, readily available computational resources, and mature methodologies presents a key opportunity for addressing this at scale by applying flexible data-driven machine learning (ML) models. Several studies have applied ML to the genetics of brain disorders and have been recently summarised18,19. Previous ML attempts have been impacted by high risk of bias20 and population stratification21, while AD studies in particular have been hampered by low sample size18, leaving a gap for comprehensive, large-scale studies that rigorously apply ML to genome-wide data. We conducted the largest genome-wide ML study in AD to date, marking a pivotal moment in the field. Our study presents a reproducible, bias-aware approach for ML model development, validation, and confounder adjustment. We trained three of the most prominent approaches in the field to compare predictive accuracy and uncover novel AD-associated risk loci which were not identified by traditional GWAS.
Results
Prediction
Gradient boosting, neural networks, MB-MDRC 1 d, and PRS models were compared for discovery and prediction (Fig. 1). Prediction of AD status between models was highly correlated in the test set, having pairwise correlations between r = 0.80 and r = 0.87 (Fig. 2). The highest correlations were observed for GBM-NN (r = 0.87) and GBM-MB-MDRC 1 d (r = 0.86). The weakest correlations were between NNs and PRS (r = 0.80). PRS was most strongly correlated with GBMs (r = 0.84). For discrimination between cases and controls, the highest AUC of 0.692 (95% CI: 0.683-0.701) was obtained with gradient boosting, and was not significantly different to an AUC of 0.689 (0.679-0.698) for PRS (Fig. 2, Supplementary Data 4). The AUCs remained within the 95% CI when we excluded any imputed variants from the data, indicating no risk of bias from their inclusion (GBMs: 0.683, NN: 0.674, and MBMDRC-1d: 0.667 without imputed variants). Predictions remained stable across repeats with different random train-test splits (Supplementary Data 4) and across the cohorts in the data (Supplementary Fig. 6). All models have a greater proportion of females in those predicted to be a case, reflecting the underling sex differences in the data (59% female, 62% female in cases, 57% in controls), except for GBMs which have a similar proportion in both cases and controls (Fig. 2i).
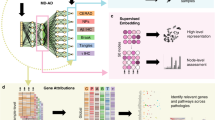
Data was separated into an initial balanced random split before model selection (cross-validation and hyperparameter tuning) in the training split (a). All models were subsequently evaluated for association (annotation, enrichment analysis, interaction testing and replication; b) and prediction (AUC and correlations; c). Interaction tests report p-values from the Wald test in logistic regression (two-sided) as standard, after correction for multiple testing The full pipeline was run four times per model to assess robustness. For prediction, AUC values, statistical tests, and correlation analyses are based on the initial train-test split. For association, variants were prioritised if they appeared in the top SNP selection in at least two repeats.
The top two most predictive approaches (GBM, PRS) were not significantly different by AUC, as measured by DeLong’s test, though prediction from MB-MDRC 1 d was significantly below other methods (a). Bars show the AUC from a single test split for each model, where whiskers are 95% CIs from the pROC package. Unadjusted p-values from tests (ns: not significant, ****: p < 0.0005) are annotated on panel a for DeLong’s two-sided test for correlated ROC curve; see supplementary Data 3 for exact values. Model predictions showed strong correlation, though correlation of the ranks is lower (b). Distributions of covariate-adjusted predictions for the most predictive approaches are similar but show a more distinct multimodal distribution for GBMs (c), NNs (d) and MB-MDRC 1d (f) compared to PRS (e), illustrating stronger influence of APOE risk alleles on predictions. Panel (g) shows the consistency across methods for individuals’ prediction scores, where the participants in the 5% extreme tails of GBM predictions are followed across predictions from NN, PRS and MB-MDRC 1 d. Classifications metrics are given in (h). Model predictions broken down by covariates are shown in (i) and (j), where dark blue indicates predicted cases, and light blue predicted controls. Box plots in (j) show the median (center line), the 25th and 75th percentiles (box limits), and the whiskers which extend to 1.5 times the interquartile range.
Identification of AD-associated loci
SNPs prioritized by ML approaches were required to appear in at least two random train-test splits, ensuring more robust associations; they include both known (Table 1) and novel variants (Table 2). Gradient boosting machines correctly distinguished the APOE haplotypes (Fig. 3a, showing distinct clusters). Figure 3b further highlights modelling of the APOE region across methods, wherein GBMs and NNs correctly identify causal SNPs. Known loci including CR1, BIN1, IDUA, OTULIN, RASA1, RASGEF1C, CLU, ABCA1, MS4A*, PICALM, ABCA7, APOE, and CASS4 (Table 1) are highlighted by ML models (Fig. 3c). In addition, several novel loci were identified with putative biological evidence for association with AD (ARHGAP25, COG7, LINC00924/LOC105369212, LY6H, SOD1 and ZNF597) which were replicated in Jansen et al.22 (Table 2). Association of an exonic missense variant was also highlighted in AP4E1, within 500 kb of the known SPPL2A locus (Table 2). Neural networks (NN) detected known loci in APOE, BIN1, CYP27C1, ABCA1 and ABCA7 (Fig. 3c) and an additional novel locus (SOD1), which also replicated in Jansen et al.22 (Table 2). MB-MDR 1 d identified SNPs in 24 genes, the majority of which map to the APOE region, for at least two train-test splits. Of these, 20 were identified by every possible split, indicating highly stable results. MB-MDR 1 d identified SNP-SNP pairs in the APOE region gene consistently through every train-test split. Single train-test splits also find genes outside of Chromosome 19 (Supplementary Fig. 7). The majority of candidate novel loci were identified by GBMs, which find multiple loci with evidence for association with AD-related traits such as cognition, pTau, AD age-at-onset and neurofibrillary tangles from previous GWAS (see Table 2 and Supplementary Data 5).
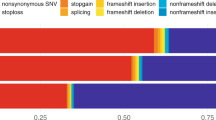
Uniform Manifold Approximation and Projection (UMAP) of raw (unscaled) SHAP values for GBM hits highlights that APOE alleles are identified and drive prediction (a). Neural networks and gradient boosting both rank the SNPs required to derive the e2 and e4 allele status for APOE as highest, unlike traditional GWAS (b). Values for neural networks and GBMs are not based on p-values, as described below, while p-values in (b,c) (GWAS) are from a logistic regression in the training split, using a logistic regression and p-values from a two-sided Wald test as standard. Manhattan plots are given for top hits only from gradient boosting (mean absolute SHAP values), neural networks (normalised network layer weights) and MB-MDRC 1 d (−log10 p-values), where hits from different random splits of the models are shown in different colours, and all variants from a single GWAS on the train split are shown in greyscale for comparison (right hand y-axis) (c). p-values for MB-MDRC 1 d in (b, c) are derived from a two-sided permutation-based test as implemented in MDMDR61,62. Hits from machine learning models (see Table 2) are enrichment for known Alzheimer’s disease processes (d).
Since ML may identify non-linear SNP-SNP interactions, pairwise interaction tests were performed for all top SNPs identified using machine learning models (Tables 1 and 2). Of 17,205 SNP-SNP pairwise interactions, 13 pairs were significant under a standard regression framework (encoded as a multiplicative interaction) after accounting for multiple testing and excluding pairs where both are within the APOE region. The two SNP-SNP pairs with the strongest evidence for association were between SNPs rs405509 and rs600550 (beta = 0.058, pFDR = 6.8 × 10−5), and SNPs rs405509 and rs12421663 (beta = −0.056, pFDR = 1.8 × 10−4), both of which involve SNPs in the APOE and MS4A* regions (Supplementary Data 6, Supplementary section 2.5).
Overlap of genes associated with disease risk across methodologies
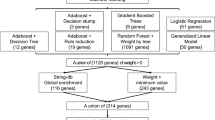
Known lead variants in APOE and BIN1 were the most important predictors in GBMs and NNs, with the SNPs used to derive APOE status (rs7412 and rs429358) identified and ranked as the most important SNPs by both GBMs and NNs but not by a GWAS in the training set (Fig. 3b). To compare the ML findings with GWAS results, we included SNPs not only identified in the training set in our study, but also all SNPs reported as genome-wide significant by larger meta-analyses (Supplementary Data 1) and applied the same gene annotation strategy as for the top variants from ML models (see “functional annotation and enrichment analysis”). In total, 130 genes were annotated, where more than one gene may be annotated to the same locus, and the same gene may be annotated to several independent loci. These genes correspond to 86 distinct loci reported in previous publications23,24,25. Of these, 19 loci were implicated by at least one ML methodology (Supplementary Fig. 8). APOE was found by all methods, seven loci (PICALM, IDUA, RASGEF1C, CLU, CR1, RTF2, RASA1) were found by GBMs and MB-MDRC 1 d, and two (ABCA7, BIN1) by MB-MDRC 1 d, GBMs and NNs. ABCA1 was only detected by NNs, and two (SPPL2A, ADAMTS1) only by GBMs (Table 1).
Motivated by published evidence suggesting that different variants are associated with the disease depending on APOE status, ML models were compared when trained with and without the APOE region in the same train-test split. Remarkably, GBMs without the APOE region found more known AD risk genes than when trained with APOE (see Supplementary section, 2.6), but with lower AUCs (Supplementary Fig. 9).
When the list of SNPs is limited to those which could be identified in the current data (using a GWAS in the corresponding training splits), all GWAS-significant SNPs were prioritised by at least one ML approach (Fig. 4). Furthermore, ~84% at the suggestive significance level (p ≤ 10−5) were retrieved by at least one ML approach.
a Genes mapped by the SNPs highlighted by each ML approach separately. Both ML approaches and GWAS significant SNPs were identified in the same training split. b Genes that are shared among at least two train-test splits. All GWAS genome-wide significant (p ≤ 5×10−8) SNPs in the train split were also identified by ML approaches. For simplicity, genes within 500 kb of a known locus or with at least one overlapping gene with the region were annotated only by the locus, including MS4A6A, CSTF1, EPHX2, CNN2, TOMM40/NECTIN2/CLPTM1/BCL3/BCAM/APOC1/APOC2/APOC4 and DGKQ/FAM53A, which were mapped to MS4A*, CASS4, CLU, ABCA7, APOE and IDUA regions, respectively. Subplots were created using ComplexUpset version 1.3.3.
Enrichment of ML findings in biologically relevant gene sets
The SNPs prioritised by ML (Tables 1 and 2) showed significant enrichment in microglial (p = 0.0024) and astrocytic regions (p = 0.0083), but not synaptic regions (p = 0.117). All 68 genes from 52 loci reported in Tables 1 and 2 were further analysed for protein-protein interaction scores using the online tool STRING (Supplementary Fig. 10). Pathway analyses demonstrated enrichment in various gene-ontology (GO) pathways (Fig. 3d, Supplementary Data 7). As expected, GO pathways yielded a number of biological processes of interest to Alzheimer’s disease, including regulation of amyloid beta formation (pFDR = 0.0014) and regulation of amyloid precursor protein catabolic process (pFDR = 0.00023, Fig. 3d).
Discussion
In leveraging the largest genotyped case-control AD dataset, this study demonstrates that the current scale of data has reached a threshold where ML can achieve similar predictive accuracy to classical methods and uncover novel genetic insights into AD. Gradient boosting, neural networks, and multifactor dimensionality reduction were selected due to their prominence in the field, demonstrated performance, and complementary strengths. These methods offer distinct advantages: highly performant tree ensembles (GBMs), flexible networks incorporating prior knowledge (NNs), and detection of SNP-SNP interactions (MB-MDR)26. Here they were applied to identify novel genes not detectable via GWAS, and to compare ML prediction accuracies with PRS.
ML methods correctly identified the lead SNPs used to calculate the APOE haplotype, rs7412 and rs429358, as having the greatest impact on the model, in contrast to classical univariable linear models (traditional GWAS) which do not distinguish between top variants by p-value, and rank rs7412 lower (Fig. 3c). PRS for AD risk prediction perform best when modelled with two predictors: PRS calculated without the APOE region and a separately coded weighting of the APOE e2 and e4 alleles27. Here, we demonstrate that ML approaches can accurately identify APOE clusters and achieve comparable prediction accuracy without including the derived APOE variable as a second predictor. ML approaches further detected the lead SNPs identified in large meta-analyses GWAS for several key genes, including BIN1 (rs6733839), PICALM (rs3851179), ABCA7 (rs3752246, a coding variant), CASS4 (rs6069736) and CR1 (rs4844610). More broadly, ML highlighted SNPs in around 22% of genes identified by larger meta-analysis GWAS23,24,25, while comparison to a GWAS undertaken on the same (EADB-core) split that was used to train machine learning models, showed that 100% of genome-wide significant SNPs (p ≤ 5 × 10−8) were retrieved by ML approaches. This demonstrates that the majority of findings expected under a linear additive GWAS paradigm can be prioritised using flexible machine learning models. Though strides have been made in ML-based prediction of AD from genetic data, for example28, to our knowledge we present the first well-powered, genome-wide ML-based gene discovery study detecting nearly a quarter of known genes found in larger GWAS meta-analyses from the literature which contain around twenty times the sample size, while also identifying multiple putative regions with credible associations to AD biology, marking an important benchmark for these methods.
Putative novel genes which replicate in an independent GWAS consist of ARHGAP25, COG7, LINC00924/LOC105369212, LY6H, SOD1 and ZNF597, which have potential relevance to AD. ARHGAP25 encodes the Arhgap25 protein which is expressed in macrophages where it affects phagocytosis29 through modulation of the actin cytoskeleton30. Wu et al.31 demonstrated that Ly6h is among the proteins competing to bind to α7 subunits of nicotinic acetylcholine receptors (nAChRs), which are expressed throughout the brain and enable fast cholinergic transmission at synapses. These proteins function collectively to maintain the optimal α7 assembly required for neuronal function and viability. This delicate balance is disrupted during Alzheimer’s disease due to Aβ-driven reduction in Ly6h. Notably, an increase in Ly6h in human cerebrospinal fluid correlates with elevated Alzheimer’s disease severity31.
COG7 (Component of Oligomeric Golgi Complex 7) encodes a protein integral to Golgi apparatus function, which is essential for protein glycosylation and trafficking. Disruptions in Golgi function have been implicated in various neurodegenerative disorders, including Alzheimer’s disease. Mutations in COG7 are associated with Congenital Disorders of Glycosylation (CDG)32, which frequently manifest with neurological impairments. Abnormal glycosylation processes have been implicated in Alzheimer’s disease pathology, particularly affecting tau protein processing and amyloid precursor protein (APP) metabolism33.
ML hits which replicate also include the missense variant rs2306331 (AP4E1). This is within the known locus of SPPL2A (maximum r2 0.32, minimum distance 177 kb with rs2306331 and GWAS catalogue hits for SPPL2A), but may be an independent signal with the region. The AP4E1 gene encodes the ε subunit of adaptor protein complex-4 (AP-4), which facilitates the transport of amyloid precursor protein (APP) from the trans-Golgi network to endosomes. Disruption of the APP-AP-4 interaction enhances γ-secretase-catalysed cleavage of APP to amyloid-β peptide, suggesting that AP-4 deficiency may constitute a potential risk factor for Alzheimer’s disease34.
We also note that ML-derived hits in the know loci IDUA and DGKQ (rs4690324, linked via sQTLs in GTEX to splicing of IDUA in the brain35) are also related to heparan sulfate metabolism36, in addition to multiple brain diseases37, traits38, and lipid metabolism39. Neural networks also highlight a new potential AD-relevant locus. All GO-derived pathways, independent of association to AD, were used in the neural network analysis as hidden layers. This approach highlighted SNPs mapping to SOD1 (cytogenic band 21q22.11), found in the region of APP (21q21.3), with lead SNPs from NNs around 5 Mb away from the closest NN-based hits in APP. SOD1 has been widely investigated for its role in antioxidant defense system, showing impaired expression in AD patients40,41, but is currently primarily implicated in ALS.
In our study, we adopted an MB-MDR approach to detect both SNP main effects and pair-wise SNP-SNP interactions. The method is an improvement over classical pair-wise statistical interaction approaches as it can detect more complex interactions. However, it implements a somewhat conservative strategy for variant discovery, as interaction studies are hampered by several factors that can increase the number of false positives including, but not limited to, LD and interference of major loci, which may contribute to phantom epistasis. To address this limitation, we opted for a model-based (MB) form of MDR, though the results did not highlight novel loci.
Comparing individual scores, ML predictions achieved correlations of 0.8-0.84 with PRS, indicating ML models give predictions which are broadly consistent with well-established approaches. It also shows that identification of novel signals through flexible modelling of complex effects will introduce deviations from the predictions of a simple linear additive model. Disease risk prediction from ML, as assessed by AUC, was higher (but not significantly) than PRS. Similar results have been reported in psychiatry21 and coronary artery disease42. This is likely to be due to several reasons. First, SNPs in general are (at best) only correlated with the causal variants, making it particularly difficult to detect nonlinear effects and interactions, which are the main potential advantages of ML over PRS. Second, genetic predictors are weak as compared to some other predictors (e.g. biomarkers43), and the upper bound for AUC in complex trait genetics in practice falls substantially below 144. Weak predictor-response relationships are an inherent challenge to finding patterns with flexible models, and complex models may at-present still be under-powered to achieve a clear improvement in AUC. Third, large GWAS discover and replicate SNPs, which in the resulting summary statistics show small association effect sizes. The effect sizes may however be higher in more homogenous samples. For example, the OR for APOE is around 3.4 in samples of mean age 72−73 years45 but is reduced in samples over 90 years46. In pathologically-confirmed samples, which are also generally older than clinical samples, some of the GWAS-derived SNP effect sizes are higher than reported in clinically-assessed AD GWAS47. In more homogenous samples in terms of age, population, and cognitive scores (such as The Alzheimer’s Disease Neuroimaging Initiative (ADNI)48), the AD PRS AUC are higher than that in clinical samples49,50. Thus, summary statistics from large GWAS meta-analyses enable PRS which predict with moderate AUC in any data set, but which do not achieve high accuracy as effect sizes are averaged across many studies with slightly different features such as recruitment criteria and outcome definitions, and therefore genetic architectures, rather than being specific to a particular one.
A similar situation affects variant discovery from flexible models which can detect interactions: nuanced relationships between predictors are notoriously difficult to replicate51, a situation further complicated by varying LD between tagged and causal variants. Such SNPs may nonetheless represent important loci which impact disease risk in specific contexts without being consistently associated across enough studies and circumstances to reach genome-wide significance under linear models in meta-analyses. While machine learning models here do not show signs of overfitting, and top SNP rankings by SHAP values are consistent in both the train and test splits, many SNPs identified do not replicate in an external dataset, which is also the case for standard GWAS. In particular, in PRS approaches using different priors (for Bayesian models) or LD pruning parameters (in clumping and thresholding approaches), the resulting set of SNPs and their estimated effects will differ27,52. Similarly, when effects of SNPs are jointly estimated in sparse, high-dimensional ML models, either simultaneously or in a stage-wise manner (as in gradient boosting), different top associations and predictions are expected. Despite these caveats, we show that the novel findings suggest SNPs which are biologically relevant to AD.
This study has a number of limitations. First, our study attempted to run and compare reasonably diverse methods for predicting AD risk, with the advantage of implementing them in a unified dataset. As a result, we found that the PRS and ML prediction showed a similar prediction performance based upon ranking individuals according by their prediction scores. The predictions from ML and PRS, however, were highly correlated, explaining the similarity of the prediction accuracies and reducing the likelihood that an ensemble of the different methods would improve prediction. Second, the methods used for selection of top SNPs in interpreting model results reflect inherent differences in how the ML approaches work, and no standard thresholds exist at the moment for identifying important features across these distinct frameworks. This methodological variability, which is a broader challenge in the field, likely contributes to the incomplete overlap between the top SNPs identified by different ML approaches, making replication in external summary statistics a critical step to ensure robust findings. Third, the influence of APOE status on model outcomes is a key component of all models in the study. While APOE SNPs were included as predictors, we did not conduct analyses stratified by APOE carrier status, especially ε3/ε3, ε3/ε4, or ε4/ε4 carriers. Future work should explore whether predictive accuracy and associations differ meaningfully across these strata. Finally, although our post-hoc analyses indicate that cohort or genotyping centre effects do not drive associations or predictions, alternative machine learning approaches such as federated learning (FL) may improve modelling where underlying predictor distributions diverge53, while also allowing for models which include data from more cohorts without diminishing privacy. Indeed, within a central learning paradigm, current best practices for data harmonisation involve merging datasets based on shared high-quality SNPs that have undergone rigorous quality control, ensuring high imputation accuracy, reasonably large minor allele frequencies (MAF), and other metrics in each separate study. However, these QC steps alone do not guarantee that differences arising from ancestry, clinical assessment criteria, and genotyping chips or batches used to generate the data are fully accounted for. Applying flexible ML within an FL framework leverages the advantages of client-specific data, which is often more homogeneous as it is typically generated locally, within the same population, using similar clinical assessments and the same genotyping chip. Results from each “client” can then inform analyses for other clients by tuning parameters (e.g., increasing or decreasing neural network weights for variants in specific genomic regions), creating a more flexible and adaptive analysis framework.
In conclusion, this study demonstrates that machine learning can uncover both known and novel genetic loci for AD, providing a powerful complement to traditional GWAS while still achieving competitive predictive performance. Though replication challenges remain, ML successfully prioritised established risk loci and identified biologically relevant novel associations, including variants in ARHGAP25, COG7, LY6H, and SOD1. These findings highlight ML’s potential to refine our understanding of AD beyond additive genetic effects and expand the toolkit available for maximising discovery from available data. With expanding datasets and computational advances, ML could further enhance risk prediction and gene discovery, particularly through federated learning and multi-omic integration. This study marks a key step toward leveraging ML for deeper insights into AD genetics.
Methods
Data
Data were obtained from the European Alzheimer & Dementia Biobank (EADB) consortium which combines genetic and clinical data from 16 countries and has been described previously23. All study protocols were reviewed and approved by the respective institutional review boards overseeing the cohorts (see supplementary information for details). Individuals were genotyped at three centres. Data were accessed after quality control procedures and data harmonisation were applied to give the EADB-core sample. All participants are unrelated individuals of European ancestry, encompassing 20,013 clinically defined AD cases and 21,673 control, after excluding participants present in Kunkle et al.45. 59% of the sample is female, split as 62% in cases and 57% in controls, with a median age at baseline of 73 (IQR = 14). Data and splitting procedures are described further in Supplementary section 1.1. Informed consent was obtained in writing from all study participants. For individuals with significant cognitive impairment, consent was secured from a caregiver, legal guardian, or other authorized proxy.
Quality control (QC)
Analyses used directly genotyped (non-imputed) variants to ensure high data quality. To avoid excluding key known AD loci not covered in the genotyped data, we also included 67 imputed variants in previously reported genome-wide significant loci23,24,25 (Supplementary Fig. 1) which were not already present in the genotyped data, converting the imputed dosages to the most probable (probability ≥ 0.9) genotypes, and applying further quality control as described previously23. Data were further filtered for a minor allele frequency of 5%, to ensure variants were common enough to reliably observe interactions. SNPs were clumped (R2 = 0.75 following Joiret et al.54, window = 1 Mb) using stage 1 summary statistics from Kunkle et al.45, after removing individuals common between the summary statistics and the genotyped data. Out of 81 SNPs previously reported as genome-wide significant, 52 survived minor allele frequency (MAF) and clumping procedures (Supplementary Fig. 1; Supplementary Data 1), giving a combined 215,193 SNPs (Supplementary section 1.1). Analyses were also run without inclusion of the previously reported 67 imputed variants to confirm that their inclusion did not artificially inflate performance estimates.
Statistics and reproducibility
No statistical method was used to predetermine sample size, as there are no standardised calculations for machine learning. For consistency, all implemented approaches were evaluated using the same QC and random train-test splits of the data (Fig. 1) which were well balanced for case-control status, age-at-baseline or assessment, sex, genotyping centre, and the distribution of all principal components (Supplementary Fig. 2). Participants were randomly separated into 70–30% train-test splits, with the same split applied to all algorithms, resulting in 29,180 individuals in the training set (14,006 cases; 15,174 controls) and 12,506 for testing (6007 cases; 6499 controls), each with 215,193 predictors after quality control procedures. ML models were built with and without SNPs from the APOE region (Chr19:44.4–46.5 Mb). In training ML and PRS models, analyses were adjusted for covariates comprising genetic sex, 20 principal component (PCs), and genotyping centre. The adjustment method was altered to be appropriate for each modelling approach: covariates were included in the final layer for NNs, with predictions and importance scores taken from non-covariate nodes (see Supplementary Fig. 3); for GBMs and MB-MDR, covariates were z-transformed and then regressed-off from the data before modelling. All reported area under the receiver operator characteristic curve (AUC) values are calculated on the predicted probabilities from models (without thresholding) and adjusted again for confounders in the test split. We utilise penalisation and cross-validated random (GBMs) or grid-based (NNs and MB-MDR) hyperparameter search to reduce the likelihood of overfitting. Details for training and covariate adjustment are given in Supplementary sections 1.2-1.4. To ensure robust results, stability was assessed by re-running all models on three additional random 70–30% train-test splits of the data.
Gradient boosting machines (GBMs)
GBMs were trained using version 1.7.6 of the XGBoost package55, an efficient implementation of regularised gradient boosting, and the dask package56, version 2023.1.1, which allows for distributed training of ML models across multiple nodes in a high-performance computing (HPC) cluster. Hyperparameters for learning rate, tree depth, and column sampling fraction were tuned on a random subsample of the training set using random search (Supplementary section 1.2). Importance of all predictors in GBMs was assessed using SHapley Additive exPlanatory (SHAP) values57,58.
Neural networks (NNs)
NN models were built with GenNet59, which uses a biologically-driven configuration in which the connections between the input layer, representing SNPs, and the first hidden layer, representing genes, are defined using knowledge available in annotation databases. NN architectures were built with nine hidden layers, including an initial SNP-to-gene layer annotating all SNPs to the nearest gene using annotations from ANNOVAR60, followed by layers defined from the hierarchy of terms from the Gene Ontology consortium (GO terms), where connections between layers progress from local pathways to more general ones as they move deeper through the network, using all available pathway annotations (Supplementary Fig. 3, Supplementary Data 2). Model hyperparameters for batch size, learning rate (LR) and L1 (default) penalization were tuned during training (Supplementary section 1.3).
Model based multifactor dimensionality reduction (MB-MDR)
A multi-dimensional reduction strategy was implemented using the MB-MDR methodology61,62 with MBMDR 4.4.1 software. An approximation routine was used to accelerate permutation-based significance assessment and multiple testing63, when searching for disease-susceptibility multi-locus genotypes, adjusted for confounders. Single and interacting variants under various pair-wise and higher order epistasis models were combined to create multi-locus risk scores (MB-MDRC) and estimate an individual’s susceptibility to a trait with the MBMDRClassifieR package available in R64. SNPs and SNP-SNP interactions were included in the MB-MDR risk score if they passed the permutation-based multiple testing corrected threshold that had the best performance in the train split (Supplementary section 1.4).
Polygenic risk scores (PRS)
PRS were calculated with LDAK-Bolt-Predict from the LDAK package, the most predictive heritability and linkage disequilibrium-informed PRS when individual genotypes are available65. LDAK-Bolt-Predict reweights variants using Gaussian priors informed by heritability models, shrinking effect sizes without the need for p-value thresholding65. To give the same information to both PRS and all machine learning models, PRS were derived using summary statistics which were generated in the training set by running a genome-wide association study (GWAS), adjusting for the same confounders in PLINK v2.00a3.3LM66 (Supplementary section 1.5).
Selection of top predictors
The top SNPs identified by machine learning were selected by the permutation-based p-value threshold defined above for MB-MDR (padj < 1), empirically by taking the extreme tail of the distribution for SHAP values (μ | SHAP | > 0.0005) in gradient boosting (Supplementary Fig. 4), summed weights (padj < 0.05) for neural networks (Supplementary Fig. 5), and by applying the Boruta algorithm67 (gradient boosting models only). Only SNPs which were present in top SNPs for at least two train-test random data splits of a given method were prioritised. Details on predictor selection are given in Supplementary sections 1.6 and 2.2-2.4.
Replication of top predictors
The prioritised loci were reported as novel if they were identified with more than one split of a machine learning model, replicated at p ≤ 0.05 significance level in an external independent dataset (Jansen et al.22) and had no genome-wide hits from AD summary statistics in the GWAS catalogue. The direction of effect for ML approaches is not available as different subgroups may have varying associations with the outcome for a given SNP, and consequently it is not directly reportable as in standard GWAS.
Functional annotation and enrichment analysis
Publicly available tools were used for further annotation: SNPs were annotated with dbSNP build 15668. SNPs were initially positionally mapped to genes using ANNOVAR60 version 2020-06-07 and Gencode v40, and then reannotated using functional evidence from the Open Targets Genetics portal where available69. Genome coordinates use build GRCh38.p14. Pathway analysis and protein-protein interaction from the consensus annotated genes were determined using STRING v12.070 (Supplementary section 1.7).
The list of top SNPs within each ML analysis was tested for enrichment in the list of genes expressed in microglia (n = 761)71, astrocytes (n = 757)72, and synapses (n = 1535)73, with a window size of 35 kilobase (kb) upstream and 10 kb downstream of regions to include regulatory elements. To account for non-independent SNPs in the genomic regions, enrichment p-values were derived using a bootstrap approach (Supplementary section 1.8) and presented without correction for multiple testing for the number of cell types tested.
Statistical interactions
The ML methods used may include interactions but not explicitly test for them. The top SNPs from all ML approaches (Tables 1 and 2) were therefore formally tested for pair-wise interactions in the whole dataset under a regression framework, assuming a multiplicative relationship between SNPs, i.e. logit(y) = β0 + β1SNP1 + β2SNP2 + β3SNP1*SNP2, where SNPs are coded additively (0, 1, 2) with zero-count compensation, and covariates are included in the model. Pairs were assessed by the p-value for the interaction term after adjusting for multiple comparisons using a false discovery rate (FDR) Benjamini-Hochberg (p = 0.05) threshold for significance (Supplementary section 1.9). Putative SNP interactions were visualized in python using shap 0.4158, statsmodels 0.13.574, matplotlib 3.7.175, and seaborn 0.12.276. The overall workflow is presented in Fig. 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data are protected and are not publically available due to data privacy laws. They can be accessed by application to the EADB consortium. The data generated in this study are provided in the Supplementary Information file.
Code availability
Code for running biologically-informed neural networks via GenNet and Tensorflow is already published and available (https://github.com/ArnovanHilten/GenNet). Code used for running distributed gradient boosting using dask and XGBoost (DAXOS v0.1.0) with covariate adjustment, preprocessing of genotypes, and applying distributed Boruta-SHAP feature selection has been made publicly available (https://github.com/seafloor/daxos)77. Pathway annotation and enrichment tests are also available on GitHub (https://github.com/seafloor/escott-price-lab-pipelines/tree/main/workflows/pathways). We use MBMDR version 4.4.1 (http://bio3.giga.ulg.ac.be/index.php/software/mb-mdr).
Change history
04 September 2025
In this article the affiliation details for Barbara Borroni were incorrectly given as ‘Molecular Marker Laboratory, IRCCS Istituto Centro San Giovanni di Dio Fatebenefratelli, Brescia, Italy’, ‘Department of Clinical and Experimental Sciences, University of Brescia, Brescia, Italy’ and ‘Cognitive and Behavioural Neurology, Department of Continuity of Care and Frailty, ASST Spedali Civili Brescia, Brescia, Italy’ but should have been ‘Molecular Marker Laboratory, IRCCS Istituto Centro San Giovanni di Dio Fatebenefratelli, Brescia, Italy’, ‘Department of Clinical and Experimental Sciences, University of Brescia, Brescia, Italy’. The original article has been corrected.
References
Lambert J. C., Ramirez A., Grenier-Boley B., Bellenguez C. Step by step: towards a better understanding of the genetic architecture of Alzheimer’s disease. Mol. Psychiatry 1–12 (2023).
Baker, E. & Escott-Price, V. Polygenic Risk Scores in Alzheimer’s Disease: Current Applications and Future Directions. Front Digit Health 2, 556191 (2020).
Nelson, R. M., Pettersson, M. E. & Carlborg, Ö A century after Fisher: time for a new paradigm in quantitative genetics. Trends Genet. 29, 669–676 (2013).
Lewis C. M., Vassos E. Polygenic risk scores: from research tools to clinical instruments. Genome Med. 12 (2020)
Visscher P. M., et al. 10 Years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101 (2017).
Liu, C. C. et al. Cell-autonomous effects of APOE4 in restricting microglial response in brain homeostasis and Alzheimer’s disease. Nat. Immunol. 24, 1854–1866 (2023). 24:11. 2023 Oct 19.
Yin, Z. et al. APOE4 impairs the microglial response in Alzheimer’s disease by inducing TGFβ-mediated checkpoints. Nat. Immunol. 24, 1839–1853 (2023).
Lin, Y. T. et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1141–1154.e7 (2018).
Emrani, S., Arain, H. A., DeMarshall, C. & Nuriel, T. APOE4 is associated with cognitive and pathological heterogeneity in patients with Alzheimer’s disease: a systematic review. Alzheimer’s Res. Ther. 12, 1–19 (2020).
Ma, Y. et al. Analysis of whole-exome sequencing data for Alzheimer disease stratified by APOE genotype. JAMA Neurol. 76, 1099–1108 (2019).
Bracher-Smith, M. et al. Whole genome analysis in APOE4 homozygotes identifies the DAB1-RELN pathway in Alzheimer’s disease pathogenesis. Neurobiol. Aging 119, 67–76 (2022).
Jun, G. et al. A novel Alzheimer disease locus located near the gene encoding tau protein. Mol. Psychiatry 21, 108–117 (2016).
Park, J. H. et al. Novel Alzheimer’s disease risk variants identified based on whole-genome sequencing of APOE ε4 carriers. Transl. Psychiatry 11, 1–12 (2021).
Escott-Price, V. & Schmidt, K. M. Pitfalls of predicting age-related traits by polygenic risk scores. Ann. Hum. Genet 87, 203–209 (2023).
Ishioka, Y. L. et al. Effects of the APOE epsilon 4 allele and education on cognitive function in Japanese centenarians. Age 38, 495–503 (2016).
Hayden, K. M. et al. Cognitive resilience among APOE ε4 carriers in the oldest old. Int. J. Geriatr. Psychiatry 34, 1833 (2019).
Abdellaoui, A., Yengo, L., Verweij, K. J. H. & Visscher, P. M. 15 years of GWAS discovery: realizing the promise. Am. J. Hum. Genet. 110, 179–194 (2023).
Rowe T. W., et al. Machine learning for the life-time risk prediction of Alzheimer’s disease: a systematic review. Brain Commun. 3 (2021)
Alatrany, A. S., Hussain, A. J., Mustafina, J. & Al-Jumeily, D. Machine learning approaches and applications in genome wide association study for Alzheimer’s disease: a systematic review. IEEE Access 10, 62831–62847 (2022).
Christodoulou, E. et al. A systematic review shows no performance benefit of machine learning over logistic regression for clinical prediction models. J. Clin. Epidemiol. 110, 12–22 (2019).
Bracher-Smith, M., Crawford, K. & Escott-Price, V. Machine learning for genetic prediction of psychiatric disorders: a systematic review. Mol. Psychiatry 26, 70–79 (2020).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Bellenguez, C. et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat. Genet. 54, 412–436 https://www.nature.com/articles/s41588-022-01024-z54:4 [Internet]. [cited 24 Jan 2023].
Andrews, S. J., Fulton-Howard, B. & Goate, A. Interpretation of risk loci from genome-wide association studies of Alzheimer’s disease. Lancet Neurol. 19, 326–335 (2020).
Wightman, D. P. et al. A genome-wide association study with 1,126,563 individuals identifies new risk loci for Alzheimer’s disease. Nat. Genet. 53, 1276–1282 https://www.nature.com/articles/s41588-021-00921-z53:9 [Internet]. [cited 25 Nov 2021].
Van Steen K. Travelling the world of gene–gene interactions. Brief Bioinform [Internet]. 2012 Jan 1 [cited 28 Feb 2025];13:1–19. Available from: https://doi.org/10.1093/bib/bbr012.
Leonenko, G. et al. Identifying individuals with high risk of Alzheimer’s disease using polygenic risk scores. Nat. Commun. 12, 1–10 (2021). https://www.nature.com/articles/s41467-021-24082-z12:1 [Internet]. [cited 6 Sep 2021].
Zhou, X. et al. Deep learning-based polygenic risk analysis for Alzheimer’s disease prediction. Commun. Med. ;3, 1–20 (2023). 3:1. 2023 Apr 6.
Schlam, D. et al. Phosphoinositide 3-kinase enables phagocytosis of large particles by terminating actin assembly through Rac/Cdc42 GTPase-activating proteins. Nature. Nat. Commun. 6, 1–12 (2015). https://www.nature.com/articles/ncomms96236:1 [Internet] [cited 28 Feb 2025].
Csépányi-Kömi, R., Sirokmány, G., Geiszt, M. & Ligeti, E. ARHGAP25, a novel Rac GTPase-activating protein, regulates phagocytosis in human neutrophilic granulocytes. Blood [Internet]. 119, 573–582 (2012). [cited 28 Feb 2025].
Wu, M., Liu, C. Z., Barrall, E. A., Rissman, R. A. & Joiner, W. J. Unbalanced regulation of α7 nAChRs by Ly6h and NACHO contributes to neurotoxicity in Alzheimer’s disease. J. Neurosci. [Internet]. 41, 8461 (2021). https://pmc.ncbi.nlm.nih.gov/articles/PMC8513707/ Oct 13 [cited 28 Feb 2025].
Wu, X. et al. Mutation of the COG complex subunit gene COG7 causes a lethal congenital disorder. Nat. Med. 10, 518–523 (2004). https://www.nature.com/articles/nm104110:5 [Internet]. 2004 Apr 25 [cited 28 Feb 2025].
Haukedal H., Freude K. K. Implications of glycosylation in Alzheimer’s disease. Front. Neurosci. [Internet]. Jan 13 [cited 28 Feb 2025];14. Available from: https://pubmed.ncbi.nlm.nih.gov/33519371/ (2021).
Burgos P. V., et al. Sorting of the Alzheimer’s disease amyloid precursor protein mediated by the AP-4 complex. Dev Cell [Internet]. 2010 Mar 16 [cited 28 Feb 2025];18:425. Available from: https://pmc.ncbi.nlm.nih.gov/articles/PMC2841041/.
GTEX. rs4690324 in DGKQ linked to IDUA through sQTLs [Internet]. [cited 2024 Apr 18]. Available from: https://www.gtexportal.org/home/snp/rs4690324.
Fabregat A., et al. Reactome. [cited 16 Apr 2024]. IDUA [lysosomal lumen]. Available from: https://reactome.org/content/detail/R-HSA-1678661 (2017).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014). https://www.nature.com/articles/ng.304346:9 [Internet]. 2014 Sep 1 [cited 28 Feb 2025].
Grasby K. L., et al. The genetic architecture of the human cerebral cortex. Science [Internet]. Mar 20 [cited 2025 Feb 28];367. Available from: https://pubmed.ncbi.nlm.nih.gov/32193296/ (2020).
Lind L. Genetic determinants of clustering of cardiometabolic risk factors in U.K. biobank. Metab. Syndr. Relat. Disord. [Internet]. 2020 [cited 28 Feb 2025];18:121–127. Available from: https://pubmed.ncbi.nlm.nih.gov/31928498/.
Li, Y., Ma, C. & Wei, Y. Relationship between superoxide dismutase 1 and patients with Alzheimer’s disease. Int J. Clin. Exp. Pathol. 10, 3517–3522 (2017).
Spisak, K. et al. rs2070424 of the SOD1 gene is associated with risk of Alzheimer’s disease. Neurol. Neurochir. Pol. 48, 342–345 (2014).
Gola, D., Erdmann, J., Müller-Myhsok, B., Schunkert, H. & König, I. R. Polygenic risk scores outperform machine learning methods in predicting coronary artery disease status. Genet Epidemiol. 44, 125–138 (2020).
Blanco, K. et al. Systematic review: fluid biomarkers and machine learning methods to improve the diagnosis from mild cognitive impairment to Alzheimer’s disease. Alzheimers Res. Ther. 15, 1–16 (2023).
Wray, N. R., Yang, J., Goddard, M. E. & Visscher, P. M. The genetic interpretation of area under the roc curve in genomic profiling. PLoS Genet. 6, e1000864 (2010).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019). https://www.nature.com/articles/s41588-019-0358-251:3 [Internet]. 2019 Feb 28 [cited 26 Jul 2021].
Farrer, L. A. et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease: a meta-analysis. JAMA 278, 1349–1356 (1997).
Escott-Price, V., Myers, A. J., Huentelman, M. & Hardy, J. Polygenic risk score analysis of pathologically confirmed Alzheimer’s disease. Ann. Neurol. 82, 311 (2017).
Weber, C. J. et al. The worldwide Alzheimer’s disease neuroimaging initiative: ADNI-3 updates and global perspectives. Alzheimer’s Dement. 7, e12226 (2021).
Leonenko, G. et al. Genetic risk for alzheimer disease is distinct from genetic risk for amyloid deposition. Ann. Neurol. 86, 427–435 (2019).
Bellou, E. et al. Age-dependent effect of APOE and polygenic component on Alzheimer’s disease. Neurobiol. Aging 93, 69–77 (2020).
Frankel, W. N. & Schork, N. J. Who’s afraid of epistasis? Nat. Genet 14, 371–373 (1996).
Ding, Y. et al. Large uncertainty in individual polygenic risk score estimation impacts PRS-based risk stratification. Nat. Genet. 54, 30–39 (2021). https://www.nature.com/articles/s41588-021-00961-554:1 [Internet]. 2021 Dec 20 [cited 22 Jan 2024].
Zhu, H., Xu, J., Liu, S. & Jin, Y. Federated learning on non-IID data: a survey. Neurocomputing 465, 371–390 (2021).
Joiret, M., Mahachie John, J. M., Gusareva, E. S. & Van Steen, K. Confounding of linkage disequilibrium patterns in large scale DNA based gene-gene interaction studies. BioData Min. 12, 1–23 (2019).
Chen T., Guestrin C. XGBoost. In: Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining - KDD ’16. New York, New York, USA. p. 785–794. (ACM Press, 2016).
Rocklin M. Dask: parallel computation with blocked algorithms and task scheduling. In: Proc. 14th Python in Science Conference (2015);
Lundberg S. M., Allen P. G., Lee S. I. A Unified approach to interpreting model predictions. Adv Neural Inf Process Syst. 30 (2017)
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
van Hilten, A. et al. GenNet framework: interpretable deep learning for predicting phenotypes from genetic data. Commun. Biol. 4, 1–9 (2021). 4:1. 2021 Sep 17.
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164–e164 (2010).
Cattaert, T. et al. Model-based multifactor dimensionality reduction for detecting epistasis in case–control data in the presence of noise. Ann. Hum. Genet. 75, 78–89 (2011).
Mahachie John, J. M., Van Lishout, F. & Van Steen, K. Model-based multifactor dimensionality reduction to detect epistasis for quantitative traits in the presence of error-free and noisy data. Eur. J. Hum. Genet. 19, 696–703 (2011).
Lishout F. et al. K Van. gammaMAXT: a fast multiple-testing correction algorithm. BioData Min. 8 (2015)
Gola, D. & König, I. R. Empowering individual trait prediction using interactions for precision medicine. BMC Bioinforma. 22, 74 (2021).
Verhaeghe J., Van Der Donckt J., Ongenae F., Van Hoecke S. Powershap: a power-full shapley feature selection method. 71–87 (2022).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Kursa, M. B. & Rudnicki, W. R. Feature selection with the boruta package. J. Stat. Softw. 36, 1–13 (2010).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Mountjoy, E. et al. An open approach to systematically prioritize causal variants and genes at all published human GWAS trait-associated loci. Nat. Genet. 53, 1527–1533 (2021). 53:11. 2021 Oct 28.
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, 1248–1259 (1979). 2017.
Endo F., et al. Molecular basis of astrocyte diversity and morphology across the CNS in health and disease. Science. 2022;378 (1979)
Koopmans, F. et al. SynGO: an evidence-based, expert-curated knowledge base for the synapse. Neuron 103, 217–234.e4 (2019).
Seabold S., Perktold J. Statsmodels: econometric and statistical modeling with Python. SciPy. 92–96 (2010).
Hunter, J. D. Matplotlib. Comput Sci. Eng. 9, 90–95 (2007).
Waskom, M. L. seaborn: statistical data visualization. J. Open Source Softw. 6, 3021 (2021).
Bracher-Smith M. Machine learning in Alzheimer’s disease genetics (2025).
Acknowledgements
We thank the University of Lille’s Intensive Scientific Computing Mesocentre. M.B.S. and V.E.P. are supported by the UK Dementia Research Institute [UK DRI-3206] through UK DRI Ltd, principally funded by the Medical Research Council and by the UK Medical Research Council: MR/T04604X/1, MR/P005748/1. K.V.S. has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreements No 813533 (MLFPM) and No 860895 (TranSYS), and FNRS convention PDR T.0294.24 “Expanded PRS embracing pathways and interactions for increased clinical utility”. F.M. (Federico Melograna) was also funded by the Marie Sklodowska-Curie grant agreement 860895 (TranSYS). J.W. (Julie Williams) is supported by Moondance Dementia Research Laboratory. B.M.T. and G.R. (Gennady Roshchupkin) are funded by ZonMw VIDI (#09150171910068) and ZonMw Veni grant (1936320) respectively. N.A. is funded by GSK. RG and AD were supported by the Italian Ministry of Health (Ricerca Corrente).
Author information
Authors and Affiliations
Consortia
Contributions
M.B.S., F.M. (Federico Melograna) and B.U. contributed equally to the study conception and design, analysis and interpretation of data, creation of code for analysis and primary drafting and revision of the manuscript. C.B., B.G.B., D.D., A.J.N., P.H., B.M.T., M.H. (Marc Hulsman), I.D.R., R.C.M., A.R. (Agustín Ruiz), R.F.S., N.A., G.R. (Gennady Roshchupkin) and J.C. contributed to the study design, interpretation of data or substantial revisions. K.V.S., C.V.D. and V.E.P. contributed equally to the study conception, interpretation of data, manuscript drafting and substantial revisions. All remaining authors—S.V.D.L., A.C., F.K., O.P., A.S. (Anja Schneider), M.D., D.R., N.S. (Norbert Scherbaum), J.D., S.R.H., L.H., L.M.P., E.D., T.G., J.W. (Jens Wiltfang), S.H.H., S.M. (Susanne Moebus), T.T., N.S. (Nikolaos Scarmeas), O.D.I., F.M. (Fermin Moreno), J.P.T., M.J.B., P.P., R.S.V., V.A., M.B., P.G.G., R.P., P.M. (Pablo Mir), L.M.R., G.P.R., J.M.G.A., E.R.R., H.S., S.H., A.D.M., S.M. (Shima Mehrabian), L.T. (Latchezar Traykov), J.H., M.V., N.S. (Nicolai Sandau), J.Q.T., Y.A.P., H.H., J.V.S., I.R. (Inez Ramakers), F.V., P.S., C.G., G.P., V.G., J.W. (Julie Williams), P.A., A.B., J.F.D., G.N., C.D., F.P. (Florence Pasquier), O.H., S.D., E.G., J.P., R.G., D.G., B.A., P.M. (Patrizia Mecocci), V.S., L.P., A.S. (Alessio Squassina), L.T. (Lucio Tremolizzo), B.B., M.W., B.N., M.S., D.S., I.R. (Innocenzo Rainero), A.D., F.P. (Fabrizio Piras), C.M., G.R., F.J., P.K., T.M., P.S.J., K.S., M.I., M.H. (Mikko Hiltunen), R.S., W.V.D.F., O.A.A., A.R. (Alfredo Ramirez), and those in the EADB umbrella group (see Supplementary)—contributed to data acquisition and/or minor revisions.
Corresponding authors
Ethics declarations
Competing interests
Outside the submitted work, T.G. received consulting fees from AbbVie, Alector, Anavex, Biogen, BMS; Cogthera, Eli Lilly, Functional Neuromodulation, Grifols, Iqvia, Janssen, Noselab, Novo Nordisk, NuiCare, Orphanzyme, Roche Diagnostics, Roche Pharma, UCB, and Vivoryon; lecture fees from Biogen, Eisai, Grifols, Medical Tribune, Novo Nordisk, Roche Pharma, Schwabe, and Synlab; and has received grants to his institution from Biogen, Eisai, and Roche Diagnostics. N.A. received funding from GSK. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Payam Barnaghi, who co-reviewed with Antigone Fogel, Mike Nalls, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bracher-Smith, M., Melograna, F., Ulm, B. et al. Machine learning in Alzheimer’s disease genetics. Nat Commun 16, 6726 (2025). https://doi.org/10.1038/s41467-025-61650-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61650-z
This article is cited by
-
Immunometabolism Reframes Alzheimer’s Disease: From Systemic Dysmetabolism to Glial Rewiring
Cellular and Molecular Neurobiology (2026)
-
Unsupervised machine learning integrates genomic variants and EMR to unravel mechanisms of brain hemorrhage and epilepsy as early indicators of Alzheimer’s in down syndrome
Alzheimer's Research & Therapy (2025)