Abstract
Transcription and replication of the Nipah virus (NiV) are driven by the large protein (L) together with its essential co-factor phosphoprotein (P). L encodes all the viral enzymatic functions, including RNA-dependent RNA polymerase (RdRp) activity, while the tetrameric P is multi-modular. Here, we investigate the molecular mechanism of the NiV polymerase and build tools for anti-NiV drug discovery. We analyze and compare multiple cryo-EM structures of both full-length and truncated NiV polymerases from the Malaysia and Bangladesh strains. We identify two conserved loops in the polyribonucleotidyltransferase (PRNTase) domain of L and the binding between RdRp-PRNTase and CD domains. To further assess the mechanism of NiV polymerase activity, we establish a highly sensitive radioactive-labeled RNA synthesis assay and identify a back-priming activity in the NiV polymerase as well as a fluorescence and luminescent-based non-radioactive polymerase assay to enable high-throughput screening for L protein inhibitors. The combination of structural analysis and the development of both high-sensitive and high-throughput biochemical assays will enable the identification of new direct-acting antiviral candidates for treating highly pathogenic henipaviruses.
Similar content being viewed by others
Introduction
Nipah virus (NiV) has caused several outbreaks in Southeast Asia and the Western Pacific, with mortality rates ranging from 40 to over 90% since its initial emergence in Malaysia in 1998 and subsequent re-emergence in Bangladesh. NiV Bangladesh strain (NiV-B) is more infectious and pathogenic than the Malaysia strain (NiV-M)1,2,3 and the discovery of human-to-human transmission was a major development in understanding the virus’ spread and impact3,4,5. While monoclonal antibodies have shown promise in treating NiV infections6, there are currently no approved treatments.
As a member of the non-segmented negative-sense (NNS) RNA virus, NiV contains the largest genome (>18.2 kbp) within the family Paramyxoviridae7, which includes viruses responsible for various diseases, such as mumps (mumps virus), respiratory tract diseases in infants (human parainfluenza viruses, HPIV), and encephalitis (Measles). The enveloped NiV virion harbors a helical ribonucleoprotein (RNP) that is formed by the single-stranded genomic RNA (gRNA) and coated by nucleoprotein (N) protomers, with a ribonucleotide to N protein ratio of 6:18. Replication and transcription of gRNA require an RNA-dependent RNA Polymerase (RdRp) complex, which is formed by a large protein (L) and a homo-tetramer of phosphoprotein (P). The P protein consists of an N-terminal domain (NTD), a central oligomerization domain (OD), and a C-terminal domain (CTD) that contains an X domain (XD). With multiple modules binding to the N protein9,10, P is crucial for both releasing naked RNA and reassembling N-RNA of newly synthesized gRNA. NiV L protein consists of 2244 residues and five domains, which encode all enzymatic functions required for viral replication. During replication, a copy of the anti-genome (cRNA) is first synthesized, followed by the generation of full-length viral gRNA. The polymerase uses the Leader (Le) and Trailer complementary (TrC) RNA promoters at the 3’ terminus of the gRNA and cRNA to initiate replication. For transcription, L performs mRNA polymerization via RdRp and 5’ capping, cap methylation, and polyadenylation at the 3’ end of nascent viral mRNA11,12,13,14,15. Although L protein is the only viral protein with enzymatic activity, making it an attractive target for antiviral drug discovery, the requirement for BSL-4 containment makes high throughput screening on authentic NiV challenging.
In recent years, cryo-EM structures from multiple NNS RNA viruses have revealed five domains of L: the RNA-dependent RNA polymerase (RdRp) domain, polyribonucleotidyltransferase or capping (PRNTase or Cap) domain, connector domain (CD), methyltransferase domain (MT), and C-terminal domain (CTD)16,17,18,19. The binding mode between L and P tetramer was elucidated16,17,18,19,20,21,22,23,24,25,26, and L oligomerizations were observed for VSV and HPIV325,27,28.
Here we describe the structures and function of the NiV RdRp (L-P) complexes to help enable target-based drug discovery. We purified NiV RdRp from both Bangladesh and Malaysia strains and developed three independent in vitro assays: (1) a radio-labeled RNA polymerization assay, (2) a non-radioactive, fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA), and (3) a pyrophosphate-coupled luminescent assay. We validated these assays by demonstrating inhibition of the polymerase activity of NiV L-P with a tool compound. We determined and compared structures of L-P from two NiV strains in full-length and/or truncated forms and identified L truncation-related structural variations. In addition, we identified a NiV L-P structure containing CD and RdRp-PRNTase domains of L that suggests a novel polymerase conformation. Overall, the combination of high-throughput biochemical assays and structural biology facilitates the identification of new direct-acting antiviral candidates for treating highly pathogenic henipaviruses.
Results
Protein preparation and assay development for NiV L-P
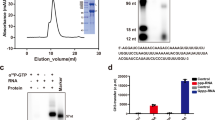
A baculovirus expression system was used to generate full-length (FL) NiV L-P from both M and B strains and an FL NiV-M with a mutation at the active site of the polymerase L(D832A)-P. In addition, we generated a truncated (TR) version of L-P containing only the RdRp and the PRNTase domain of each strain. Multiple chromatographic steps, including heparin column and size-exclusion chromatography (SEC), were included during purification, yielding well-separated SEC peaks that contain pure L and P proteins. Purified TR L-P from two strains were obtained in a dominant SEC peak with a Superose 6 Increase 10/300 GL column. However, the FL complexes of both M and B strains eluted in two separate SEC peaks at ~9.5 mL (peak #1) and ~11 mL (peak #2) (Figs. 1a and S1a).
a SDS-PAGE of the four purified NiV L-P constructs from NiV-M and NiV-B strains. From left to right: full-length (FL) NiV-M L-P, truncated (TR) NiV-M L(1-1463)-P, FL NiV-B L-P, and TR NiV-B L(2-1463)-P. Protein molecular weight markers are shown to the left of the gel images in kDa. b In vitro radioactive RdRp activity assay using the four NiV L-P constructs shown in (a) Le25 and TrC25 RNA serve as templates for synthesis; a reaction with no RNA is used as control. [α-32P] GTP is used as the radioactive signal tracer. RNA ladders (M) containing 6 bands are shown to the left of the gel images in nt. c In vitro radioactive RdRp activity assay for validation of the back priming activity of both FL and TR NiV L-P from both strains. Only [α-32P] ATP was used as the NTP. ~60 nM L-P was used. Sizes of the 3 major RNA products (26, 27, and 28 nt) were labeled. Predicted secondary structures of Le25 and TrC25 are shown on the right: a two-base pair and a free “U” at the 3’ terminus for Le25 and a 3’ three-base pair for TrC25 are formed. Back priming products with TrC25 were indicated by “AAA” in bold. The data shown above are representative results of at least two independent experiments. d Schematic of the fluorescence assay that monitors elongation activity. RdRp RNA-dependent RNA polymerase, BBT (2’-[2-benzothiazolyl]-6’-hydroxybenzothiazole) diphosphat, CIP calf intestinal phosphatase, 2pi pyrophosphate. e Fluorescent-based alkaline phosphatase-coupled assay (FAPA). Activity of varying concentrations (0–500 nM) of TR NiV-B/M L-P (left panel), means (±SD are plotted (n = 4). Here n = 4 refers to 4 technical replicates. Determination of Km for RNA with RdRp, TR NiV-B and NiV-M L-P exhibiting a Kmapp of ~5.6 µM and 3.3 µM, respectively (middle panel). Determination of Km for NTPs with RdRp, TR NiV-B, and NiV-M L-P exhibiting a Kmapp of ~22.79 µM and 20.9 µM, respectively (right panel). f Schematic of the luminescent coupled assay to monitor the RdRp activity. g Progress curves at varying concentrations (0-250 nM) of TR NiV-B L-P, displaying polymerase activity completion at ~30 min, as indicated by the dashed line. h Tool compound inhibition activity in the RdRp FAPA, coupled luminescence assay of TR NiV-B L-P (means ± SD are plotted (n = 2)). Here n refers to technical replicates. Source data are provided as a Source Data file.
An in vitro RNA synthesis assay was performed using leader (Le) and trailer complementary (TrC) promoter sequences of NiV as templates and [α-32P]-GTP or [α-32P]-ATP as the signal tracer. Purified FL proteins from two SEC peaks were both active, and peak #2 was used in all the subsequent assays (Fig. S1a, b). RNA synthesis was detected with Le25 and TrC25 for all four wildtype constructs, FL, and TR (Fig. 1b). The mutation D832A in NiV-M L abolished RNA synthesis (Fig. S1c). The TR proteins produced RNA products that are longer than the template (25 nt) and reminiscent of the back-priming (or 3’ extension) products of the NNS virus RdRp29,30,31,32, while the FL showed both the back-priming products (>25 nts) and multiple de novo products (≤25 nts) (Fig. 1b).
To further validate the back-priming activity observed in Fig. 1b, we repeated the assay but supplied only [α-32P]-ATP as the NTP for the reaction (Fig. 1c). The scheme illustrates the predicted hairpins of Le25 and TrC25. We hypothesize that the addition of ATP is sufficient for the generation of RNA products of 26–28 nts in length when combined with the TrC25 template, as shown on the gel. The mismatch at the 3’ end of Le25 suggests that RNA synthesis through back-priming is not feasible, and hence no bands were observed on the gel for these reactions (Fig. 1c). Screening of various lengths of Le and TrC RNA with FL proteins consistently showed that TrC but not Le RNA generated back-priming products (Fig. S1d). These results indicate that the FL and TR L-P of both strains can synthesize products longer than the given template via a back-priming mechanism. However, only the FL protein can produce de novo RNA products (Fig. 1b).
To aid high-throughput drug discovery, we developed both fluorescent and luminescent-based assays to measure the polymerase activity of the TR NiV-B/M L-P. The fluorescence-based alkaline phosphatase-coupled assay (FAPA) was optimized for use in 384-well plates (Fig. 1d). Using TrC25 as the template for the polymerase reaction, a 3.5-fold signal window was observed at 125 nM of TR NiV-B L-P and a 4-fold signal window was observed at 62 nM of TR NiV-M L-P (Fig. 1e). Km values were obtained by plotting the initial velocities of the observed RNA synthesis activity as a function of either RNA or nucleotide to delineate the enzyme affinities to RNA and nucleotide (Fig. S1e). The NiV-B apparent Km for RNA and NTP were determined as ~5.6 µM and ~22.79 µM, respectively. The NiV-M apparent Km for RNA and NTP were ~3.3 µM and 21 µM, respectively (Fig. 1e). As a complementary method for measuring elongation activity, the luminescent-based assay was developed as a continuous assay that detects pyrophosphate production in real-time (Fig. 1f). Pyrophosphate formation depends on NiV RdRp concentration, and the luminescent signal was only observed in the presence of the enzyme, NTPs, and TrC25 RNA. A 6-fold signal window was detected at 250 nM of TR NiV-B L-P. Here, an n-fold signal window refers to the ratio of the fluorescent or luminescent signal at the concentration of enzyme tested (in the absence of inhibitor) to the fluorescent or luminescent signal in the absence of protein (Fig. 1e, g). To further validate the FAPA and luminescence assays, a tool compound that inhibits the polymerase was employed. The IC50 value for this compound was 0.8 µM and showed inhibition in the two independent assay formats (Fig. 1h). Both non-radioactive polymerase assays were robust, as indicated by a Z’ > 0.5.
Overall structures of FL/TR NiV L-P from different strains
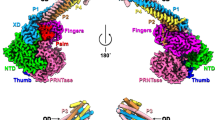
Single particle cryo-EM was applied for FL NiV L-P and TR L-P from both M and B strains. Sample quality, including purity, molecular weight, and sample homogeneity, was confirmed by SDS-PAGE and SEC (Figs. 1a, 2a and S1a). We determined the structure of the FL NiV-M L-P polymerase to 2.92 Å resolution. Similar to our in vitro radioactive RNA synthesis assay, the peak #2 sample of FL NiV-M L-P was used (Figs. S1a and S2). In the atomic model, the tetrameric P, with each monomer at various lengths, and the RdRp (residue 7-970) and PRNTase (residue 971-1446) domains of L were modeled. For P, four P-OD (residue 535-582, 532-589, 532-579, 538-578, respectively), one P-CTD fragment (residue 594-610), one P-XD (residue 657-706) and its preceding loop (residue 636-656) were resolved. Some regions of P, including the P-NTD, were not resolved, likely due to their lack of binding to L. Four fragments, including residues 586-711, 1140-1153, 1267-1289, and 1338-1361 in L, were not modeled due to poor density (Fig. 2b). No density was observed for the CD, MT, and CTD domains in L, suggesting high flexibility of these domains. This is also observed in other Paramyxoviruses L reconstructions such as HPIV5 and Newcastle disease virus (NDV)20,24 (Fig. 2a, b).
a Domain organization of NiV L and P proteins. L contains the following five domains: RdRp domain (marine), PRNTase domain (lime green), CD (gray), MT domain (gray), and CTD (gray). Residues in the active site of the RdRp (GDN) and the PRNTase domain (GXXT and HR) are indicated. Locations of six conserved regions (CR-I to CR-VI) are highlighted. P contains an N-terminal Domain (NTD, in gray), an oligomerization domain (OD, in orange, brown, light pink, or dark pink), and a C-terminal Domain (CTD, in dark pink). At the C terminus, P has an X domain (XD, in dark pink). The NiV proteins P interacts with are indicated on the top. Predicted phosphorylation sites of P are shown below. Cryo-EM density map fitted with the model for FL NiV-M L-P (b), TR NiV-M L-P (c), and TR NiV-B L-P (d) and another view of the models with a 180° turn. In b, c, d, individual domains are colored as depicted in (a), and the right panels are views with the NTP entry channel facing outward, defined as the front view in this study.
TR L-P structures from NiV-M and NiV-B were determined at 2.99 Å and 3.43 Å resolution, respectively, displaying similar assembly and architectures to FL NiV-M L-P (Figs. 2c, d and S3). In TR NiV-B L-P, we obtained the density to model an extra double a-helix turn structure on the N terminus of each P protomer, which was similarly observed in the crystal structures of NiV and Sendai virus P-ODs33,34,35. In addition, we resolved extra density for L alone for FL NiV-M L-P and TR NiV-B L-P (Figs. S2 and S3), suggesting that L and L-P are in equilibrium in the purified complex samples.
Overall, our structures show that L has a structurally conserved RdRp domain and a PRNTase domain that forms a ring shape. The catalytic residues 831-GDN are located in the center of the RdRp-PRNTase ring. The antenna-like P-OD is docked to and extends from the RdRp domain, which separates the PRNTase domain and the P protein. Between P-OD and P-XD, a long loop attaches to the edge of the NTP entry (Fig. 2).
Structural comparison for L from FL NiV-M L-P and TR NiV-M & B L-P
Consisting of 970 residues, the NiV L RdRp domain shares a “fingers-palm-thumb” right-hand fold. The catalytic residues (831-GDN) are held by seven conserved motifs that belong to the palm subdomain (motif A-E) or the fingers subdomain (motif F-G). The palm subdomain is at the junction of the other two subdomains, while the N-terminal region (NTR) structurally supports the active site (Figs. 3a, S4 and S5a). In contrast, the PRNTase domain is much smaller than the RdRp domain. The alpha helix-rich part of the PRNTase domain forms the putative RNA template exit channel, and the loop-rich region accommodates the priming and intrusion loops (Fig. 3b). In NNS virus polymerases, these two loops are functional. Located over the RdRp catalytic cavity, the priming loop facilitates the formation of an initial dinucleotide in RNA synthesis36. The HR (histidine-arginine) motif in the intrusion loop catalyzes mRNA capping37 (Figs. 3b and S5b).
a Cartoon representation of the RdRp domain from FL NiV-M L-P. Conserved RdRp motifs A-G are highlighted in rainbow colors within the “fingers-palm-thumb” subdomains, and the N-terminal region (NTR) is shown in gray. G831 in the RdRp catalytic site (GDN) is shown as magenta spheres. The unresolved part between the fingers and motif A in the palm (named preA) is shown as a dashed red line. b Architecture of the RdRp-PRNTase domain from FL NiV-M L-P. Cartoon representation of PRNTase domain on the surface of RdRp domain. RdRp and PRNTase domains are shown in color as depicted in Fig. 2. G831 of GDN is shown as magenta spheres. The loop-rich region is indicated and the residues at the boundaries of the priming and intrusion loops are shown as orange and blue spheres, respectively. RNA template entry, NTP entry, and template exit channels are labeled in yellow and shown as spheres (calculated by CAVER Web)57. c Superposition of TR NiV-M and NiV-B L protein RdRp domain (left panel) and PRNTase domain (right panel). Residues that differ are highlighted as orange spheres. Each variation is labeled to show the change from the Malaysia to the Bangladesh strain. d Superposition of the FL and TR NiV-M L-P structures and highlights on their differences by numbers 1–4 around the RdRp catalytic center, priming loop, and intrusion loop. e Detailed views of the structural differences in RdRp and PRNTase domains observed between FL and TR NiV-M L-P as indicated in (d).
The RdRp-PRNTase domains in TR NiV-M/B L-P were structurally similar, with a root-mean-square deviation (RMSD) value of 0.358 Å over 1274 atoms (Fig. 3c and S5). Nine residue differences between NiV-M to NiV-B were mapped to the RdRp and three to the PRNTase domain (Fig. 3c). In the RdRp domain, six mutations occurred in the NTR (R36K, D71N, I77V, I94T, K113R, V252I), two (E534D and K783E) were located to the finger and one (I890V) to the palm subdomain (Fig. 3c). K1157R, R1181K, and R1262K occurred in the PRNTase domain. Structure-based sequence alignment identified eight more mutations mapped to an area between the finger and the motif A of the palm subdomain that is unique to NiV and not present in other NNS viruses (Fig. S4). However, the EM density in this region (K580-K711, named preA in this study) was not resolved.
Next, the superposition of FL and TR NiV-M L-P structures showed that more than 20 residues near the active sites were resolved in the FL but not the TR, particularly at five locations: V829-G831 next to the RdRp catalytic sites (831-GDN); K544–T546 in the loop of motif F of the RdRp domain (E543-R551); K581-V585 at the beginning of the preA region; G1232-L1234 and H1264-S1266 before the priming loop (S1266 - P1290), and G1383-K1385 after the intrusion loop (P1338–V1362). In the TR NiV-B L-P, similar structural disorders were also observed, especially at loop I1230-T1244, before the priming loop, and loop D1375-I1391, after the intrusion loop, which were named pre-priming loop and post-intrusion loop, respectively (Figs. 3d, e and S6)24. Taken together, our results show that the FL has a more stable structure on the RdRp-PRNTase domains compared to the TR protein.
Interactions between L and P in FL NiV-M L-P
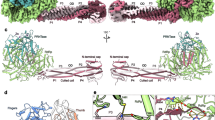
In FL NiV-M L-P, a conserved antiparallel beta-sheet is formed between the RdRp and one P protomer24,25, herein named P1. We name the other P protomers P2, P3, and P4, from a counterclockwise direction using a top view from the N terminus of the P-OD (Fig. 4a). The four P protomers are nearly identical in P-OD but vary in the extending arm (Fig. 4a). Six L-P interfaces were identified using the PDBePISA online server38 (Fig. 4a). Residues M575-I578 of P1 and K385-E388 of L form the antiparallel beta-sheet at interface 1, and K795, L382, and Y389 of L likely form hydrogen bonds with G582, K581, and M574 of P1, respectively (Figs. 1 and 4b). In interface 2, not only are multiple hydrogen bonds presumably present, including one formed by S565 of P2 and H423 of L, but the L residues Y389, M393, and I737 also have extensive hydrophobic interactions with residues of the two proximal protomers of P, including M574 and I576 of P2 and L571 of P1 (Figs. 2 and 4b). The likely hydrogen bond formed by M574 of P1 and Y389 of L bridges interfaces 1 and 2 (Figs. 1, 2 and 4b). P2 itself extends close to the template entry tunnel of RdRp to form interface 3, where K583 and R586 of P2 contact E740 and E744 of L through electrostatic interactions, and E585 and K589 of P2 likely form hydrogen bonds with Q454, D456, and C457 of L, respectively (Figs. 3 and 4b). Interfaces 4 and 5 involve fewer residues than other interfaces and are maintained similarly by potential hydrogen bonds and electrostatic interactions (Figs. 4, 4b and 5). At interface 6, residues from a single a-helix of P-XD interact with three a-helices of RdRp via multiple suggested hydrogen bonds and salt bridges, making a relatively stable L-P binding site (Fig. 4b). The interface area of L and P is 2870 Å2 and involves the RNA template entry side and the NTP entry side.
a A front view of the FL NiV-M L-P is shown in the cartoon and colored as depicted in Fig. 2. Four P protomers are labeled by P1, P2, P3, and P4. The P protein fragments extend from the C-terminal of P-OD and are highlighted by a thicker loop radius. G831 in the GDN is shown as magenta spheres. In the dashed line box on the top, four P protomers are shown side by side and in superposition. The P-OD part is nearly identical, whereas the structures downstream vary depending on their binding sites on the L protein. P-CTD is shown, which is assigned to P4. Two side views are shown below to highlight the locations of the six L-P binding interfaces. The NTP entry and template entry are indicated. b Magnified view of the interactions between L and P proteins at the six interfaces. Structural elements, including the RdRp and P protomers and residues that participate in the interactions, are indicated. The potential hydrogen bonds and electrostatic interactions are depicted as dashed lines, with oxygen and nitrogen atoms shown in red and blue, respectively.
Structure of FL NiV-M L-P-CD
Single-particle cryo-EM analysis was also applied to the peak #1 sample of FL NiV-M protein from another batch of purification with buffers at higher pH, which contains pure L and P and shows RdRp activity, allowing the determination of the structure to 2.99 Å resolution (Figs. S1a, b, S2 and S7a). Compared to FL NiV-M L-P, a significant amount of extra density next to the RdRp-PRNTase domains was observed upon map reconstruction (Figs. 5 and S2). A similar density was assigned to the CD domain from a 2nd L (CD’) binding to a L-P complex in the partial HPIV3 L-P dimer structure (PDB ID: 8KDB)25. An AlphaFold 2 (AF)39 model of NiV-M L CD domain (residues 1475-1757) was generated to aid model building, with the mean predicted local distance difference test (pLDDT) and the uncertainty to be 89.31% and 4.98%, respectively. The AF model of CD fits the extra density, and the atomic models of five fragments with 17–38 residues in length were successfully built (Figs. 5b and S6). However, other parts of the CD could not be built due to the poor density. Modeling of the MT and CTD domains was not consistent with the size and shape of the density observed and could, therefore, be ruled out from the model (Fig. S7b). These results suggest this newly discovered density is the CD domain of NiV L protein; therefore, we named this structure FL NiV-M L-P-CD.
a Domain organization of NiV L and P proteins. The color code used in Fig. 2a is followed except for the CD domain shown in yellow. b Model building for the FL NiV-M L-P-CD structure determined using the peak #1 sample. The fitting between the RdRp-PRNTase and the map is shown on the left, with the extra density boxed. Fitting for the AF model of CD domain and the modeled CD domain with the extra density are shown on the right. c Superposition of the structures of FL NiV-M L-P and FL NiV-M L-P-CD in this study. The RMSD is indicated. d Partial interface between CD and RdRp-PRNTase in FL NiV-M L-P-CD. The residues (K191, E1540, L973, and D1641) involved in polar interactions identified using the PDBePISA online server are shown in stick and indicated. Individual domains are colored as depicted in (a).
Superposition of FL NiV-M L-P-CD and FL NiV-M L-P in this study showed that they are nearly identical with a RMSD value of 0.321 Å over 1304 atoms (Fig. 5c), suggesting the high stability of the RdRp-PRNTase core of the polymerase. Interface analysis between the core and CD domain revealed that E1540 and D1641 in CD form salt bridge and hydrogen bond with K191 in the RdRp domain and L973 in the PRNTase domain, respectively (Fig. 5d). In HPIV3, corresponding residues L1483 and N1584 were also found to have interactions with F195 in RdRp and Q914 in PRNTase in the partial L-P dimer structure25. Nearby residues, especially L971-N982 and D1619-Q1652 of NiV L, are of the same length and highly conserved in the closely related Paramyxoviruses, but not in other NNS viruses, including VSV and RSV (Fig. S4). Our results support that a conserved interface forms between the RdRp-PRNTase and CD domains of the L protein in Paramyxoviruses polymerase.
Discussion
Our study identifies structural similarities and differences between the NiV B and M strains and the full-length and truncated L protein. We showed that the RdRp-PRNTase of L is more stable in the full-length NiV L-P than in the truncated. We developed a radioactive assay and two high-throughput, non-radioactive biochemical assays to demonstrate the functionality of the proteins used in this study and accelerate anti-NiV drug discovery.
De novo RNA synthesis and back-priming have been previously described for RSV viruses29; however, the regulation of this mechanism and its role during viral replication remain poorly understood. In this study, we demonstrate that the L protein of NiV is capable of synthesizing RNA de novo and catalyzing the 3’ extension of the RNA template via back-priming. Notably, the full-length L protein can perform RNA synthesis through both mechanisms, whereas the truncated form cannot generate the de novo products. Kleiner et al. demonstrated that the truncated L of RSV (RdRp-PRNTase) is unable to generate the de novo products that the full-length L protein can generate40. They also discovered that the RdRp-PRNTase of RSV L can synthesize a 2 nt product necessary for initiating RNA synthesis. Our data did not provide evidence of the 2 nt product, which may be due to the low resolution of the gel. These observations, combined with our finding that the NiV RdRp-PRNTase is more stable in the full-length L, suggest that the three C-terminal domains of L are essential in the elongation activity of RNA replication. Additionally, we show that the back-priming mechanism depends on the RNA template and the predicted secondary structure it can form. We generated an AF3 model using the RdRp-PRNTase of NiV L and its TrC19 RNA which produces the back-priming product (Fig. S1d). The AF prediction indicates that the predicted 3-base pairings in the TrC RNA can be stabilized by the L protein and positioned towards the 831-GDN site for RNA 3’ extension (Fig. S8), thereby validating our in vitro assays. More studies are needed to dissect and fully understand this mechanism’s relevance during viral replication.
While the 32P assay directly measures RdRp activity through visualization of newly synthesized RNA products, this gel-based assay is unsuitable for screening. Here, we report the development of two non-radioactive assay formats, a fluorescence-based assay, and a continuous luminescent-based assay. Of note is that both assays were developed with the truncated L protein containing the RdRp and PRNTase domains. The Z’ values for both assays were >0.5, indicating robust assay performance. Previously, using a radiometric de novo RdRp activity assay, it has been reported that the NiV polymerase exhibits a Km of ~79 µM and 18 µM for CTP and ATP, respectively41, while the RNA Km has not yet been described in the literature. Here, for the first time, we were able to develop biochemical plate-based assays that allowed us to characterize NiV RdRp kinetically. Developing high-throughput NiV RdRp assays is critical in enabling target-based drug discovery against NiV.
Our structural comparison of TR NiV-M/B L-P reveals that I890V mutation occurs in the motif E of the palm subdomain (Fig. 3). In the FL NiV-M L-P, the distance between I890 and G831 in the RdRp catalytic center 831-GDN is ~7.6 Å, and the side chain of I890 protrudes towards the NTP entry. This structural proximity suggests that shortening the hydrophobic side chain in the I890V mutation likely impacts the RdRp activity of the NiV polymerase.
Comparison of TR and FL NiV RdRp in this study suggests that the presence of the three C-terminal domains (CD, MT, and CTD) could potentially stabilize the pre-priming and post-intrusion loops in RdRp-PRNTase (Fig. 3d, e). Similar to our study of NiV L-P, an NDV L-P structure (PDB ID: 7YOT) without the three flexible C-terminal domains of L was obtained using full-length L and P proteins, and the pre-priming and post-intrusion loops were nearly completely resolved, suggesting higher stability24 (Fig. S9a). A complete NDV L-P structure containing all five domains of L was also determined, in which S1154 in the pre-priming loop (L1151-E1165) and D1296 in the post-intrusion loop (D1296-V1312) likely form a hydrogen bond and a salt bridge with the MT domain24 (Fig. S9b). Structure-based alignment showed that these two residues correspond to S1233 and D1375 in NiV and are also highly conserved in the Paramyxoviridae family (Fig. S4). In VSV L-P (PDB ID: 6U1X), corresponding residues E1117 (pre-priming loop) and D1255 (post-intrusion loop) interact with R1740 and S1706 in the MT domain21. Based on our results and the literature, we propose that the MT domain of L could interact and stabilize the pre-priming and post-intrusion loops in the PRNTase domain.
We also determined the structure of FL NiV-M L-P-CD containing parts of the CD domain bound to the RdRp-PRNTase domains. Three 2D class averages obtained using the selected 68,791 particles matched three different projections of our FL NiV-M L-P-CD structure superposed with the AF CD domain model (Fig. S7c). Our FL NiV-M L-P-CD structure raised the question of whether it represents a monomer or dimer state of the polymerase. Despite the pH variation (7.6-8.0) of buffers used in L-P purification, peaks #1 and #2 were repeatedly obtained, suggesting dimerization occurs in the earlier eluted peak #1 (Fig. S7a). Mass photometry results confirmed the formation of L-P dimer in peak #1 but also showed that the majority is a monomer (Fig. S7d). Recently, cryo-EM structures of the full-length NiV polymerase were reported by several groups42,43,44,45,46,47, however, the purified L-P complex corresponding to peak #1 in this study has not been used. Meanwhile, the crystal structure of the L CD domain was reported at 1.85 Å resolution45, which we superimposed on our FL NiV-M L-P-CD structure to check if a monomeric polymerase model is possible. The distance (70.8 Å) between the end residues (P1446 at the C terminus of the RdRp-PRNTase in FL NiV-M L-P-CD and L1481 at the N terminus of CD crystal structure) could allow the linker (34 residues) to connect them in the same polymerase molecule (Fig. S7e), thus not excluding a monomer model. Our results suggest that the CD domain in FL NiV-M L-P-CD could come from the same L-P complex or a second one.
Superposing our FL NiV-M L-P-CD structure with the three C-terminal domains of L in the HPIV5 L-P (PDB ID: 6V85) reveals a possible conformation for the five domains of L protein (Fig. S10), further supporting that the RdRp-PRNTase-CD interface found in this study is true in nature. By modeling the Le10 RNA bound to the RSV L-P (PDB ID: 8SNX) into this superposition, we found that the model describes a state where the MT domain blocks the RNA template entry (Fig. S10). Therefore, the binding between RdRp-PRNTase and CD domains could regulate the RNA entry for RNA synthesis in the NiV.
In NNS virus polymerase, multi-modular P protein interacts with the L protein as a tetramer containing protomer P1-P4. However, how the four P protomers are arranged on the surface of L is poorly understood. By applying our P1-P4 nomenclature, we compared the L-P binding modes of NiV in this study and other NNS viruses from the literature. We found that one hydrophobic residue in L (Y389 in NiV of this study) acts as a wedge in between P1 and P2, likely stabilizing the P protomers with hydrophobic interactions. Using the same box size for particle image extraction, the P3 and P4 are unresolvable in one MuV L-P structure (PDB ID: 8YXM), but all four protomers are resolved in another MuV L-P structure using the same sample (PDB ID: 8IZL)26. Additionally, P1, P2, and P4 can stretch to either the template or NTP entry side, but P3 has only been shown to bind the template entry side on L. For example, P2 in our NiV L-P structure and the EBOV L-VP35 extend towards two opposite directions (Fig. S11). This suggests the interactions between L and specific P protomers are interchangeable, and P monomers may constantly bind and disassociate from the polymerase complex during the different stages of RNA synthesis.
Previously published structures of the NiV polymerase with resolutions ranging from 2.3 to 3.19 Å detail the interactions between the L and P proteins, align with and enhance our findings42,43,44,45,46,47. Recently, Sala et al. reported the NiV polymerase-RNA complex in an early elongation state48, in which the template RNA superposes well with the TrC19 RNA used in our AF3 model for the RdRp-PRNTase-RNA complex (Fig. S8). However, the L protein back-priming capabilities and structural analysis between two NiV strains presented here provide unique structural and functional insights.
In summary, our biochemical and structural analysis of NiV-M and NiV-B polymerases reveals essential aspects of the mechanisms of NiV RNA synthesis, including how the important loops in the RdRp-PRNTase domain could be stabilized in the polymerase and how the binding between the RdRp-PRNTase and CD domains could influence the RNA template entry. These findings will be essential for developing structure-based drug discovery efforts targeting NiV RdRp.
Methods
Protein preparation of the NiV polymerase
To obtain active (NiV, Malaysia and Bangladesh strain) L-P complexes, codon-optimized coding sequences of L (GenBank: AAK29089.1 for M and GenBank AAY43917.1.1 for B) and P (GenBank: AAF73378.1 for M and GenBank: AAY43912.1 for B) were synthesized to have an N-terminal 10XHis tag and 3XStrep tag, respectively, and cloned into pFastBac-Dual vectors (Synbio Technologies) for expression in Spodoptera frugiperda 9/21 (SF9/21, ThermoFisher, Cat No. 1496015 and 11497013) cells. A second open reading frame for mCherry or GFP was included in the Malaysian strain L or P plasmid vector, respectively, to guide the baculovirus generation in insect cells. Truncated L genes containing the RdRp and PRNTase domains were subcloned into the pFastBac-Dual vector. The vector for NiV-M L D832A was generated by the Q5 Site-Directed Mutagenesis Kit (NEB). All the self-cloned plasmids were confirmed by Sanger sequencing.
For the full-length Malaysia strain L-P complex, the insect cells were harvested by centrifugation (3500 × g, 4 °C, 30 min) at 48 h post-infection and lysed by dounce homogenizer in buffer A containing 30 mM Tris-HCl, pH 7.6–8.0, 300 mM NaCl, 10% glycerol, 1 mM Tris(2-carboxyethyl) phosphine (TCEP), 1 mM Phenylmethanesulfonyl fluoride (PMSF). The cell lysate was centrifugated (104,000 g, 4 °C, 1 h), and then the supernatant was filtered (0.45 μm filter) and loaded through a Strep-tactin Sepharose gravity column (IBA), which was pre-equilibrated with buffer B containing 30 mM Tris-HCl, pH 7.6-8.0, 300 mM NaCl, 10% glycerol, 1 mM TCEP. The bound protein was eluted using buffer B supplemented with 1–10 mM D-disthiobiotin. The eluted proteins were slowly diluted with an equal volume of buffer C containing 30 mM Tris-HCl, pH 7.6–8.0, 10% glycerol, 1 mM TCEP and ran by a Heparin column (Cytiva) with a gradient of buffer D containing 30 mM Tris-HCl, pH 7.6–8.0, 1.0 M NaCl, 10% glycerol, 1 mM TCEP. The eluted L-P peak was pooled and further purified by the Ni-NTA affinity column (Thermal Fisher). The L-P complex was eluted using buffer B supplemented with 100–200 mM imidazole. The protein was then concentrated with a 100 kDa cutoff Millipore Ultra centrifugal filter and purified by SEC using a Superose 6 10/300 column (Cytiva) pre-equilibrated with buffer E containing 30 mM Tris-HCl, pH 7.6–8.0, 0.3 M NaCl, 1 mM TCEP. Similar procedures were used for the truncated protein of the Malaysia strain L-P, except the pH of all buffers was kept at 7.6. Similar procedures were used for the full-length and truncated L-P of the Bangladesh strain, except that the Ni-NTA column was omitted. After Strep-tactin column and Heparin column purification, the L-P complex was concentrated with a 100 kDa cutoff Millipore Ultra centrifugal filter and purified by SEC using a Superose 6 10/300 column (Cytiva) pre-equilibrated with buffer F containing 20 mM HEPES, pH 7.5, 300 mM NaCl, 5% glycerol, 6 mM MgCl2, 1 mM TCEP. Size exclusion chromatography peak fractions containing L-P were pooled, concentrated, flash-frozen in aliquots, and stored at −80 °C before use.
Radioactivity-based polymerase NiV RdRp activity assays
Standard desalting purified RNA oligoribonucleotides representing residues 7, 10, 13, 16, 19, 22, and 25 of the Leader (Le) or trailer complementary (TrC) sequences of NiV-M were purchased (IDT) and used for in vitro RNA dependent RNA polymerization assays. NTPs were purchased from NEB. Radioactive isotope-labeled nucleotides [α-32P] GTP and [α-32P] ATP were purchased from Perkin Elmer (800 Ci/mmol, 10 mCi/ml, 250 μCi). The RNA ladder was generated by incubating [ɣ-32P] ATP and various lengths of RNA templates with T4 PNK (NEB). Purified full-length and truncated NiV Malaysia strain L-P complex samples were supplemented with 6 mM MgCl2 and incubated on ice for 30 min before the assay. Purified NiV Bangladesh strain L-P samples were directly used in the assay. The reaction mixtures were composed of 2 μM RNA template, L-P complexes, NTPs (50 uM [α-32P] GTP/ATP and 1.25 mM for the other three NTPs or as specified), and reaction buffer (50 mM Tris-HCl, pH 7.4, 8 mM MgCl2, 5 mM DTT, 10% glycerol) in a final volume of 10 μl. The reactions were started by adding the L-P samples and incubating at 30 °C for 3 h. Reactions were terminated by adding stop buffer (90% formamide, 20 mM EDTA, 0.02% bromophenol blue) and heated at 90 °C for 5 min. 3 ul of the final sample was resolved by denaturing urea PAGE (20% acrylamide-bis acrylamide 37.5:1, 1XTBE, 7 M urea). The RNA sequencing gel was exposed to a phosphor screen overnight and imaged by phosphor imaging with a Typhoon biomolecular imager.
Fluorescence-based alkaline phosphatase-coupled polymerase assay (FAPA) with truncated NiV L-P
A fluorometric assay that monitors elongation activity was developed and validated. Here, the RdRp enzyme incorporates an adenosine monophosphate residue into the RNA while simultaneously releasing a BBT (2’-[2-benzothiazoyl]-6’-hydroxybenzothiazole) diphosphate. Subsequently, Calf Intestinal Alkaline phosphatase cleaves the BBT-PPi by-product and liberates a free BBT anion fluorescent molecule. Free BBT can be detected at excitation max 422 nm and emission max 566 nm. The RNA template consists of 25 nucleotides derived from the trailer complementary sequence of the NiV (Malaysia strain). Time course experiments were performed on the 25 nt Trailer RNA template using either 125 nM or 62 nM of truncated NiV-B or NiV-M L-P, respectively, and 100 µM Atto-ATP, UTP, CTP, and GTP. RNA was titrated from 0 to 30 µM to determine the Kmapp. The converse set of experiments determined the Kmapp for NTP, using 125 nM or 62 nM of NiV-B or NiV-M enzyme and 20 µM of RNA. Nucleotides were titrated from 0 to 250 µM. To identify compounds that are competitive with nucleotides, the assay will be performed under pseudo-first-order kinetics, whereby nucleotide concentrations will be added below Km. In contrast, RNA concentrations will be added at saturating concentrations (>5× Km). The assay quality was assessed by determining the Z’ value. The Z’ values are calculated by evaluating the signal dynamic range and the signal variation of the active control (minimum) and the total signal (uninhibited reaction, total). Here, 80 nL of 100% DMSO was used as the neutral control, and 80 nL of tool compound was used for the minimum active control (final concentration 100 µM). The assay was performed by adding 4 µl of the enzyme to the well and allowing the enzyme to pre-incubate with the compound for 15 min at 30 °C. The reaction was then initiated by the addition of 4 µL of RNA and nucleotide solution. The Z’ factor was calculated using the formula:
The assay performs well with a Z’ factor above 0.5 and a %CV for neutral and active controls below 10%.
Luminescence coupled assay to monitor NiV RdRp activity
In this luminescence-coupled assay, the initial step is the enzymatic reaction between the L-P (RdRp), the RNA template, and the incoming nucleotides. As the nucleotides get incorporated into the RNA, the resulting byproduct is pyrophosphate. We utilized the PPiLight Assay Kit (Lonza PPiLight_Catalog #: LT07-610), which detects inorganic pyrophosphate (PPi). The detection reagent catalyzes the AMP conversion to ATP in the presence of PPi, which is subsequently consumed by luciferin in the presence of luciferase and produces light.
The RNA template used in the assay comprises 25 nucleotides and is the same as the template used in the FAPA assay. A series of titrations were performed to evaluate the activity of truncated NiV L-P across a range of 0–250 nM, using 5 µM of the RNA template, 10 µM of NTPs (UTP, GTP, CTP), and 100 µM of α-S-ATP. Here, in addition to the standard, unmodified nucleotides, we used adenosine-5’-O-(1-thiotriphosphate) (α-S-ATP). This ATP analog is a poor substrate for luciferase but can be used as a substrate for RdRp nucleotide incorporation into the RNA template. To monitor the protein activity, 5 μl of the enzyme was titrated with buffer, which consists of 50 mM Tris-HCl (pH 7.4), 8 mM MgCl2, 5 mM DTT, 10% glycerol, 0.1% Triton-X-100, 40 units/ml RNAsin, and 0.01% BSA, followed by 5 ul of the substrate (RNA and nucleotide mix), resulting in a total reaction volume of 10 ul. 5 ul of converting reagent and 5 μl of detection reagent were added immediately. Luminescence activity was measured at a 0.1-s integration reading, with the temperature set at 37 °C. The reaction progress curve was obtained at different time points, and the optimal window size was observed at 35 min. Assay quality was similarly evaluated by determining the Z’ value (as described above).
Cryo-EM screen and dataset collection for NiV-M L-P
We glow-discharged UltrAuFoil R 1.2/1.3 holey gold supports 300-mesh gold grids (Quantifoil Micro Tools) at 20 mA for 30 s. To vitrify the sample, we applied 3.0 μl of FL/TR NiV-M L-P complexes (~0.7/0.9 mg/mL) onto each grid and plunge-froze the sample in liquid ethane using a Vitrobot Mark ΙV (Thermo Fisher Scientific, TFS) after a 3 s blotting with no force and with filter papers pre-saturated for 30 min under 100% humidity. We screened the homogeneity of the sample with a Talos 120 KV TEM (TFS) and a Talos Arctica 200 kV TEM (TFS) at the IEMC of Emory University. For the peak #2 FL sample, images were collected at the National Center for Cryo-EM Access and Training (NCCAT) at the New York Structural Biology Center with Leginon 3.5 on TFS Titan Krios microscopes operated at an acceleration voltage of 300 kV with a Gatan K3 camera with a 0.809 Å pixel size. The energy filter slit width was 20 eV. The defocus range was set from −0.6 to −2 μm. The total accumulated dose was about 55 electrons per square ångström. 13,158 micrographs were collected. For the peak #1 FL sample, images were collected at the Midwest Consortium at Purdue University with a Titan Krios microscope (TFS) operated at an acceleration voltage of 300 kV with a Gatan K3 camera. The pixel size was obtained using the gold diffraction data to 0.874 Å initially, and calibrated and validated to 0.84 Å during dataset processing. The energy filter slit width was 20 eV. The defocus range was set from −0.5 to −2 μm. The total accumulated dose was 52 electrons per square ångström. 9782 micrographs were collected. For the TR NiV-M L-P, images were collected at NCCAT with Leginon 3.5 on TFS Titan Krios microscopes operated at an acceleration voltage of 300 kV with a TFS Selectris-Falcon4i camera with a 0.927 Å pixel size. The energy filter slit width was 20 eV. The defocus range was set from −0.6 to −2 μm. The total accumulated dose was about 50.35 electrons per square ångström. A tilted stage (30–45° at each square) was set, and 13,418 micrographs were collected.
Cryo-EM screen and dataset collection for NiV-B L-P
The model was generated from data collected on six different grids due to preferred orientation issues. Briefly, 3 µL of the sample were applied to the following: Quantifoil 1.2/1.3 holey carbon 300 mesh copper grid (one of the six total) with graphene oxide support glow-discharged at 5 mA for 10 s, UltrAufoil 1.2/1.3 holey 300 mesh gold grids (five of the six total) glow-discharged at 20 mA for either 10 s or 20 s. The graphene oxide-containing grid was blotted for 10 s and the gold grids for 6 s. All grids were plunge frozen in liquid ethane using a Vitrobot Mark III (Thermo Fisher Scientific) set to 4 °C, 100% relative humidity, and positive 20 blot force. The truncated L-P was diluted to around 1 μM for the graphene oxide-containing grid and 4-2 μM for the gold grids. The protein was diluted for all grids in 20 mM Tris pH 7.5, 300 mM NaCl, 6 mM MgCl2, 2.5% glycerol, and 1 mM TCEP. However, 0.03% w/v CHAPS detergent was added to two gold grids to induce new views of the protein.
Grids were stored in liquid nitrogen before data collection or immediately put on the microscope. Data was collected using EPU (Thermo Fisher Scientific) on a Glacios (Thermo Fisher Scientific) operating at 200 kV. Movies were recorded using Falcon 3 (Thermo Fisher Scientific) in counting mode at a corrected magnification of ×120,000, corresponding to a physical pixel size of 0.86 Å and a dose rate of 1.21 e-/pixel/frame. Each movie was recorded as a stack of 29 subframes, each accumulated for 29 s, totaling ~35 electrons per Å2. Defocus values ranged between −0.8 and −2.0 μm. For two (gold) of the six grids, the stage was tilted to either 20 degrees or 30° to expand the view angles of the protein. 11,285 movies were collected.
Cryo-EM data processing
The FL NiV-M L-P (peak #2) dataset was processed using cryoSPARC V4.2.149. Pre-processing was performed using patch motion correction and patch-CTF estimation with default settings. For particle picking, based on its diameter and the ring shape formed by RdRp and PRNTase domain, ~3000 potential NiV L-P particles were manually picked from 200 micrographs. After 2D classification, good-looking classes were selected and then used to train the topaz model with the same 200 micrographs50,51. A final topaz pick model was obtained after multiple cycles of topaz pick and 2D classification and then used to pick particles from the whole dataset (1.4 million particles). To remove duplicates, particles with an origin closer than 100 Å from each other were discarded. These particles were 2D classified, and the selected particles (885 K) were used to generate 4 new ab initio models that were then refined using heterogeneous refinement, local motion correction, homogeneous refinement, and local refinement with recentering, leading to a final reconstruction of 2.92 Å resolution. The TR NiV-M L-P dataset was processed using a similar procedure. After particle picking and 2D classification, 581 K particles were subjected to 3D ab initio and refinements, including global and local CTF refinement, finally yielding a 2.99 Å resolution map.
The TR NiV-B L-P dataset: movie frames were aligned and summed into a single micrograph using Relion v452 motion correction and then imported into cryoSPARC v.4. CTF parameters were determined using patch CTF estimation in cryoSPARC. Particles were picked via template picker in cryoSPARC using previous 2D class averages from the same data as templates. Initially, 1,634,076 particles for the truncated L/P protein complex were identified after multiple rounds of 2D classification in cryoSPARC. Particles were run through two steps of heterogeneous refinement (first with 3 classes and second with 6), ending at a final particle count of 183,266. Running cryoSPARC non-uniform refinement on this pool of particles led to a final map at a global resolution of 3.62 Å (EM map 1). After the final round of 3D refinement, the density of the P protein in EM map 1 was very weak, so we re-centered on the P protein and re-extracted an intermediate particle stack consisting of particles of the truncated L-P complex. After multiple rounds of 2D classification to remove poor recentered particles, we had a final particle count of 346,748. Running cryoSPARC homogeneous refinement on these particles led to a reconstructed map at a global resolution of 3.43 Å (EM map 2) with a better-resolved P protein (EM map 2).
The FL NiV-M L-P (peak #1) dataset was processed using cryoSPARC V4.2.1. Pre-processing was performed using patch motion correction and patch-CTF estimation with default settings. Blob picking was first tried for particle picking but failed. Then, based on its diameter and the ring shape formed by RdRp and PRNTase domain, 2,530 potential NiV L-P particles were manually picked from 300 micrographs. After 2D classification, good-looking classes were selected and then used to train the topaz model with the same 300 micrographs. A final topaz pick model was obtained after multiple cycles of topaz pick and 2D classification and then used to pick particles from the whole dataset (1.27 million particles). To remove duplicates, particles with an origin closer than 100 Å from each other were discarded. These particles were 2D classified, and the selected particles (1.02 million) were used to generate 3 ab initio models that were then subjected to 3D refinement. A map (502 K particles) containing a significant extra blob compared to the peak #2 sample was obtained using heterogeneous refinement. Then, particles for this novel 3D class were re-extracted and used for one round of Topaz pick from 600 micrographs. More particles were picked using the Topaz model. After 3D classification, 69,991 particles were selected for homogeneous refinement and non-uniform refinement, leading to a map at 3.64 Å resolution that contains the extra density for the CD domain. Model building on this map revealed an expansion of the RdRp-PRNTase core that distorted the RdRp catalytic site. Since the data-collection microscope was retired, we estimated the pixel size to be 0.84 Å by inspecting the fitting between the reported Nipah polymerase structures (PDB IDs: 9BDQ and 9FUX) that have same conformations into our map. Then the 69,991 particles were re-extracted and used for 3D reconstruction and refinement, after reference-based motion correction on the re-extracted particles with the refined 3D model, non-uniform refinement and local refinement using a mask created on the RdRp core plus the extra density, a map at a global resolution of 2.99 Å was obtained with 68,791 particles. We further validated the applied pixel size by using the “fit spherical aberration” function in global CTF refinement by obtaining a spherical aberration value at 2.7153.
Model building and refinement
For FL NiV-M L-P, the 2.92 Å resolution map was first fitted with the RdRp domain of the HPIV5 L-P complex cryo-EM structure (PDB ID: 6V85) using UCSF Chimera X53. Then, the coordinates of a NiV L RdRp-PRNTase (residues 1–1463) model predicted by AlphaFold 239 (https://neurosnap.ai/) were fitted into the map by superimposition with the HPIV5 L RdRp domain. The published crystal structure of the OD domain of NiV P protein (PDB ID: 6EB9) was positioned to fit the density for P protein. The structure model for the missing parts was manually built by WinCOOT54. Multiple cycles of manual adjustment in WinCoot and real-space refinement in PHENIX_refine55 were used to improve model geometry using secondary structure restraints compared with the HPIV5 L-P model. Models of TR NiV-M L-P and FL NiV-M L-P-CD were built using FL NiV-M L-P as the template. For TR NiV-B L-P, the molecular models for the truncated NiV-B L-P complex were built using an AlphaFold 2 starting model and consecutive rounds of manual building in Coot and real space refinement in Phenix. The structures of the CD, MT, and CTD domains of L were predicted by AlphaFold 2 (https://neurosnap.ai/) with residues 1475-1757, 1758-2083, and 2084-2244 of the NiV L protein (UniProt ID: Q997F0) as the input. The mean predicted local distance difference test (pLDDT) and the uncertainty are 89.31 and 4.98%, 81.62 and 6.7%, 88.83 and 1.35% for the three domains, respectively. Model and map statistics are summarized in Tables S1 and S2. The structure of NiV L RdRp-PRNTase complexed with the Tr19 RNA was predicted using AlphaFold 356 (https://neurosnap.ai/). Residues 1–1463 of the NiV L protein and the Tr19 sequence (5’-UUCUUUACCCUUGUUCGGU-3’) were used as the input.
Mass photometry
A Refeyn TwoMP mass photometer (Refeyn LTD, Oxford, UK) was used for the mass photometry analysis of the peak #1 sample of FL NiV-M L-P at room temperature. Glass coverslips and gaskets were cleaned with HPLC-grade water and isopropanol. NiV L-P was diluted to 100 nM in SEC buffer. Nineteen microliters of buffer were used to find the camera focus prior to loading 1 μl of the sample onto the gasket. The acquisition camera image size was set to medium. Data was collected as a 1-min movie and then processed using ratiometric imaging. The Refeyn TwoMP instrument was calibrated using the MassFerence P1 standard containing a protein with several oligomeric forms ranging from 86 to 344 kDa to correlate ratiometric contrast to mass. Data were analyzed using DiscoverMP version 2.3 software (Refeyn LTD, Oxford, UK).
Figure preparation
Protein sequence alignment was performed with T-coffee (https://tcoffee.crg.eu/apps/tcoffee/do:expresso) combined with secondary structures, and the alignment results were displayed with ESPript (https://espript.ibcp.fr/). Buried interface areas were calculated by PDBePISA (https://www.ebi.ac.uk/pdbe/pisa/). UCSF Chimera X and Pymol (https://pymol.org/2/) were used for structure visualization and figure generation. Cavities in the protein complex were calculated using CAVERweb57.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM density maps and atomic coordinates generated in this study have been deposited to the Electron Microscopy Data Bank (EMDB; https://www.ebi.ac.uk/emdb/) and the PDB (https://www.rcsb.org), respectively, with the following accession codes or numbers: PDB ID: 9COK and EMD-45782 for the FL NiV-M L-P; PDB ID: 9MUW and EMD-48649 for the TR NiV-M L-P; PDB ID: 9CWO and EMD-45968 for the TR NiV-B L-P; PDB ID: 9MZH and EMD-45896 for the FL NiV-M L-P-CD. Source data are provided with this paper.
References
Chua, K. B. Nipah virus outbreak in Malaysia. J. Clin. Virol. 26, 265–275 (2003).
Hossain, M. J. et al. Clinical presentation of Nipah virus infection in Bangladesh. Clin. Infect. Dis. 46, 977–984 (2008).
Mire, C. E. et al. Pathogenic Differences between Nipah virus Bangladesh and Malaysia strains in primates: implications for antibody therapy. Sci. Rep. 6, 30916 (2016).
Chadha, M. S. et al. Nipah virus-associated encephalitis outbreak, Siliguri, India. Emerg. Infect. Dis. 12, 235–240 (2006).
Banerjee, S. et al. Nipah virus disease: a rare and intractable disease. Intractable Rare Dis. Res. 8, 1–8 (2019).
Tit-Oon, P. et al. Prediction of the binding interface between monoclonal antibody m102.4 and Nipah attachment glycoprotein using structure-guided alanine scanning and computational docking. Sci. Rep. 10, 18256 (2020).
Lo, M. K. et al. Determination of the henipavirus phosphoprotein gene mRNA editing frequencies and detection of the C, V and W proteins of Nipah virus in virus-infected cells. J. Gen. Virol. 90, 398–404 (2009).
Ker, D. S., Jenkins, H. T., Greive, S. J. & Antson, A. A. CryoEM structure of the Nipah virus nucleocapsid assembly. PLoS Pathog. 17, e1009740 (2021).
Omi-Furutani, M., Yoneda, M., Fujita, K., Ikeda, F. & Kai, C. Novel phosphoprotein-interacting region in Nipah virus nucleocapsid protein and its involvement in viral replication. J. Virol. 84, 9793–9799 (2010).
Shu, Y. et al. Plasticity in structural and functional interactions between the phosphoprotein and nucleoprotein of measles virus. J. Biol. Chem. 287, 11951–11967 (2012).
Grosfeld, H., Hill, M. G. & Collins, P. L. RNA replication by respiratory syncytial virus (RSV) is directed by the N, P, and L proteins; transcription also occurs under these conditions but requires RSV superinfection for efficient synthesis of full-length mRNA. J. Virol. 69, 5677–5686 (1995).
Sleat, D. E. & Banerjee, A. K. Transcriptional activity and mutational analysis of recombinant vesicular stomatitis virus RNA polymerase. J. Virol. 67, 1334–1339 (1993).
Grdzelishvili, V. Z. et al. A single amino acid change in the L-polymerase protein of vesicular stomatitis virus completely abolishes viral mRNA cap methylation. J. Virol. 79, 7327–7337 (2005).
Ogino, T. & Banerjee, A. K. Unconventional mechanism of mRNA capping by the RNA-dependent RNA polymerase of vesicular stomatitis virus. Mol. Cell 25, 85–97 (2007).
Liang, B. Structures of the mononegavirales polymerases. J. Virol. 94 https://doi.org/10.1128/JVI.00175-20 (2020).
Liang, B. et al. Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 162, 314–327 (2015).
Gilman, M. S. A. et al. Structure of the respiratory syncytial virus polymerase complex. Cell 179, 193–204 e114 (2019).
Cao, D. et al. Cryo-EM structure of the respiratory syncytial virus RNA polymerase. Nat. Commun. 11, 368 (2020).
Pan, J. et al. Structure of the human metapneumovirus polymerase phosphoprotein complex. Nature 577, 275–279 (2020).
Abdella, R., Aggarwal, M., Okura, T., Lamb, R. A. & He, Y. Structure of a paramyxovirus polymerase complex reveals a unique methyltransferase-CTD conformation. Proc. Natl. Acad. Sci. USA 117, 4931–4941 (2020).
Jenni, S. et al. Structure of the vesicular stomatitis virus L protein in complex with its phosphoprotein cofactor. Cell Rep. 30, 53–60 e55 (2020).
Horwitz, J. A., Jenni, S., Harrison, S. C. & Whelan, S. P. J. Structure of a rabies virus polymerase complex from electron cryo-microscopy. Proc. Natl. Acad. Sci. USA 117, 2099–2107 (2020).
Yuan, B. et al. Structure of the Ebola virus polymerase complex. Nature 610, 394–401 (2022).
Cong, J. et al. Structure of the newcastle disease virus L protein in complex with tetrameric phosphoprotein. Nat. Commun. 14, 1324 (2023).
Xie, J. et al. Structural basis for dimerization of a paramyxovirus polymerase complex. Nat. Commun. 15, 3163 (2024).
Li, T. et al. Structures of the mumps virus polymerase complex via cryo-electron microscopy. Nat. Commun. 15, 4189 (2024).
Rahmeh, A. A. et al. Critical phosphoprotein elements that regulate polymerase architecture and function in vesicular stomatitis virus. Proc. Natl. Acad. Sci. USA 109, 14628–14633 (2012).
Rahmeh, A. A. et al. Molecular architecture of the vesicular stomatitis virus RNA polymerase. Proc. Natl. Acad. Sci. USA 107, 20075–20080 (2010).
Noton, S. L., Aljabr, W., Hiscox, J. A., Matthews, D. A. & Fearns, R. Factors affecting de novo RNA synthesis and back-priming by the respiratory syncytial virus polymerase. Virology 462-463, 318–327 (2014).
Cao, D. et al. In vitro primer-based RNA elongation and promoter fine mapping of the respiratory syncytial virus. J. Virol. 95 https://doi.org/10.1128/JVI.01897-20 (2020).
Shareef, A. M., Ludeke, B., Jordan, P., Deval, J. & Fearns, R. Comparison of RNA synthesis initiation properties of non-segmented negative strand RNA virus polymerases. PLoS Pathog. 17, e1010151 (2021).
Peng, Q. et al. Molecular mechanism of de novo replication by the Ebola virus polymerase. Nature 622, 603–610 (2023).
Bruhn, J. F. et al. Crystal structure of the Nipah virus phosphoprotein tetramerization domain. J. Virol. 88, 758–762 (2014).
Tarbouriech, N., Curran, J., Ruigrok, R. W. & Burmeister, W. P. Tetrameric coiled coil domain of Sendai virus phosphoprotein. Nat. Struct. Biol. 7, 777–781 (2000).
Jensen, M. R. et al. Structural description of the Nipah virus phosphoprotein and its interaction with STAT1. Biophys. J. 118, 2470–2488 (2020).
Cressey, T. N. et al. Distinctive features of the respiratory syncytial virus priming loop compared to other non-segmented negative strand RNA viruses. PLoS Pathog. 18, e1010451 (2022).
Ogino, T. & Banerjee, A. K. The HR motif in the RNA-dependent RNA polymerase L protein of Chandipura virus is required for unconventional mRNA-capping activity. J. Gen. Virol. 91, 1311–1314 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Kleiner, V. A. et al. Conserved allosteric inhibitory site on the respiratory syncytial virus and human metapneumovirus RNA-dependent RNA polymerases. Commun. Biol. 6, 649 (2023).
Jordan, P. C. et al. Initiation, extension, and termination of RNA synthesis by a paramyxovirus polymerase. PLoS Pathog. 14, e1006889 (2018).
Yang, G., Wang, D. & Liu, B. Structure of the Nipah virus polymerase phosphoprotein complex. Nat. Commun. 15, 8673 (2024).
Peng, Q. et al. Cryo-EM structure of Nipah virus L-P polymerase complex. Nat. Commun. 15, 10524 (2024).
Wang, Y. et al. Cryo-EM structure of Nipah virus RNA polymerase complex. Sci. Adv. 10, eadr7116 (2024).
Balikci, E. et al. Structure of the Nipah virus polymerase complex. EMBO J. 44, 563–586 (2025).
Hu, S. et al. Structural and functional analysis of the Nipah virus polymerase complex. Cell https://doi.org/10.1016/j.cell.2024.12.021 (2025).
Xue, L. et al. Cryo-EM structures of Nipah virus polymerase complex reveal highly varied interactions between L and P proteins among paramyxoviruses. Protein Cell https://doi.org/10.1093/procel/pwaf014 (2025).
Sala, F. A., Ditter, K., Dybkov, O., Urlaub, H. & Hillen, H. S. Structural basis of Nipah virus RNA synthesis. Nat. Commun. 16, 2261 (2025).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Res. Comput. Mol. Biol. 10812, 245–247 (2018).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D Struct. Biol. 73, 496–502 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Stourac, J. et al. Caver Web 1.0: identification of tunnels and channels in proteins and analysis of ligand transport. Nucleic Acids Res. 47, W414–W422 (2019).
Acknowledgements
We thank the members of the laboratory of B.L. for their support and critical discussions. We also acknowledge the assistance and services provided at the Robert P. Apkarian Integrated Electron Microscopy Core (IEMC) at Emory University. We thank Drs. Eric Sundberg, and Maros Huliciak for assistance in the Mass Photometry analysis at the Department of Biochemistry at Emory University. We would like to express our gratitude to Thierry Diagana for leading the University of Texas Medical Branch (UTMB)-Novartis Alliance for Pandemic Preparedness (UNAPP), a collaboration between Global Health, Biomedical Research at Novartis and UTMB as one of the NIH Antiviral Drug Discovery (AViDD) Centers for Pathogens of Pandemic Concern. We also thank Pei-Yong Shi (currently at GlaxoSmithKline LLC) for his initial leadership of UNAPP during his tenure at UTMB and Vineet Menachery for his continued support and leadership. We thank the Global Health Project Management (Kit Chan), Alliance Management and Partnering (Thomas Krucker, Elianna Amin), Legal (Daniel Raymond, Marcus Hall), and Finance team (Marco Tulio Ramirez, Tracey Heinrich, and Jean Claude Poilevey) for their operational and administrative support. We also thank Gwynn Pardee for baculovirus generation and protein expression and Cathy Luu, Mikias Woldegiorgis, and Heidi Carias Martinez for additional expression support. Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number U19AI171413. Cryo-EM data collections were carried out at the Midwest Consortium at Purdue University (NIH U24 GM116789) and the National Center for Cryo-EM Access and Training (NCCAT) at the New York Structural Biology Center, supported by the NIH Common Fund Transformative High-Resolution Cryo-Electron Microscopy program (U24 GM129539). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceptualization B.L. J.S., M.C.C., N.J.; Methodology B.L., Z.C., J.Q.D., C.D., C.Z.B., T.T., L.X., A.R., D.S., D.P.; Validation B.L., Z.C.; Formal analysis Z.C., B.L., A.R., D.C.; Investigation B.L., Z.C., C.D., C.Z.B.; Resources B.L., C.D., C.Z.B., M.D., G.S.; Data curation Z.C., B.L., C.D., C.Z.B., A.R. Writing—original draft Z.C., B.L.; Writing—review and editing Z.C., B.L., J.H., L.W., C.Z.B., J.Q.D., J.S., A.R.; Visualization Z.C., B.L., J.S.; Supervision B.L., J.S., N.J., M.C.C., M.D., G.S.; Project administration B.L., J.S.; Funding acquisition B.L., N.J.
Corresponding authors
Ethics declarations
Competing interests
The Authors declare the following competing interests: J.Q.D., C.D., C.Z.B., D.P., A.R., D.S., T.T., L.X., M.D., G.S., M.C.C., N.J., and J.S. are employees of Novartis Pharma AG and some own shares in Novartis Pharma AG. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Loïc Carrique, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Z., Quirit Dudley, J., Deniston, C. et al. Cryo-EM structures of Nipah virus polymerases and high-throughput RdRp assay development enable anti-NiV drug discovery. Nat Commun 16, 6655 (2025). https://doi.org/10.1038/s41467-025-61764-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61764-4