Abstract
Vault is a massive ribonucleoprotein complex found across Eukaryota. The major vault protein (MVP) oligomerizes into an ovular cage, which contains several minor vault components (MVCs) and is thought to transport transiently bound “cargo” molecules. Vertebrate vaults house a poly (ADP-ribose) polymerase (known as PARP4 in humans), which is the only MVC with known enzymatic activity. Despite being discovered decades ago, the molecular basis for PARP4’s interaction with MVP remains unclear. In this study, we determined the structure of the human vault cage in complex with PARP4 and its enzymatic substrate NAD+. The structures reveal atomic-level details of the protein-binding interface, as well as unexpected binding sites for NAD+ and related nucleotides within the interior of the vault cage. In addition, proteomics data show that human vaults purified from wild-type and PARP4-depleted cells interact with distinct subsets of proteins. Our results thereby support a model in which PARP4’s specific incorporation into the vault cage helps to regulate vault’s selection of cargo and its subcellular localization. Further, PARP4’s proximity to MVP’s NAD+-binding sites could support its enzymatic function within the vault.
Similar content being viewed by others
Introduction
The vault particle was identified more than three decades ago and has been found within most eukaryotic clades1, yet its molecular function is poorly understood. The particle’s primary structural element is known as the vault cage, an elliptical protein assembly measuring about 70 by 40 nm in dimension. In every species analyzed so far, the cage contains 78 copies of the major vault protein (MVP), which coalesce into two symmetrical, domed, 39mer halves2. Three constitutive minor vault components (MVCs) have been identified in the cage in vertebrates: poly (ADP-ribose) polymerase 4 (PARP4)3, telomerase component 1 (TEP1), and a class of small non-coding RNAs known as vault RNAs (vtRNAs)4. Vault activity has been implicated in the innate immune response5,6, tumor progression & chemotherapeutic drug resistance7,8, and metabolic regulation9,10. The vault cage structure has inspired hypotheses that the vault acts as a signaling scaffold, reaction crucible, or transport module for endogenous, transiently internalized molecules known as its “cargo”. Indeed, MVP has been implicated in the signaling activity and relocalization of specific macromolecules in response to different cellular perturbations6,9,11,12,13. However, its mechanism at the molecular level, including how it selects cargo molecules and their destinations, has not been elucidated. We hypothesized that some of these important regulatory roles might be performed by the MVCs, whose functions are not yet clear. In this study, we focus on PARP4, which is the only MVC with known catalytic activity14.
PARP4 is a member of the PARP superfamily of enzymes, which consume the critical metabolite NAD+ to deposit ADP-ribose (ADPr) modifications onto macromolecular substrates. The biological function of PARP proteins is driven in large part by their “targeting” domains, which determine their subcellular localization and, consequently, the molecules they modify15. PARP4’s specific docking to the interior of the vault cage via its MVP-Interaction or “MINT” domain (Fig. 1a) implies that it could regulate vault activity by modifying MVP, other minor components, or encapsulated vault cargo. To this end, PARP4 contains an unusual combination of domains involved in macromolecular interactions, including VWA, VIT, and BRCT (Fig. 1a), which could facilitate direct interactions between PARP4 and vault’s protein or nucleic acid cargo. In addition, there is broad interest in engineering a vault to deliver therapeutics into cells. Linking these molecules to the MINT domain has proven to be an effective way to package them into the vault cage16, generating interest in mutating the MINT domain to orchestrate optimal molecular loading and release17. These properties motivated us to investigate the structural basis for PARP4’s interaction with MVP, the impact of adding PARP4’s substrate NAD+ to the complex, and the effect of PARP4’s presence on the vault interactome. Here, we use single-particle cryo-EM to reveal the interaction interface between MVP and the MINT domain of PARP4, providing high-resolution structural insights. In addition, our structures reveal specific binding sites within the MVP cage for nucleotide ligands, including NAD+ and ADP. The presence of nucleotide-dependent enzymes in the vault interactome, along with the predicted location of PARP4’s catalytic domain near the NAD+ binding site, suggests that the vault may serve to store substrate for encapsulated enzymes. Finally, proteomic analysis shows that depletion of PARP4 alters the vault interactome, resulting in a reduced number of binding partners, which are more enriched in the nucleus and less associated with the cell membrane, suggesting that PARP4 may be involved in mediating vault cargo uptake.
a Domain diagrams of MVP (upper) and PARP4 (lower). b Dimensions and labeled regions of the vault cage, with one MVP monomer from the top half of the cage colored in pink and a medially symmetric monomer from the bottom in purple. c Magnified views of the vault medial interface, with atomic models of MVP fit to the cryo-EM map (gray mesh). Hydrophobic residues F6, I7, and I36, buttressed by two residues with complementary charges (D39 and R9), form the interface between two directly opposed MVP monomers (left panel). A network of charged (D39 and R9) and polar (N22, N40) side chains forms a dynamic interface between MVP monomers positioned diagonally across opposite halves (right panel). d Cross-sectional views of the apo-MVP (left) and MVP-PARP4 (right) cryo-EM maps, with potential corresponding to PARP4 colored in blue. Map view of a single copy of MVP3−5 bound to one copy of MINT (upper inset) and the corresponding model (lower inset).
Results
Structure of the human MVP-PARP4 complex
Full-length constructs of human MVP and PARP4 were cloned into baculoviral vectors for overexpression in insect cells, which do not produce endogenous MVP. An initial attempt to purify the MVP-PARP4 complex by expressing the proteins separately and mixing the cell lysates was unsuccessful, yielding only MVP cages without internalized PARP4 (Supplementary Fig. 1a). Therefore, we changed our expression strategy, simultaneously co-infecting cells with the MVP and PARP4 viruses and adding approximately 5–6 times more PARP4 than MVP virus. This allowed us to compensate for the more robust expression of MVP in insect cells and to saturate the PARP4 binding sites within the cage, maximizing the structural information we could obtain about the MVP-PARP4 interface. After the new expression strategy was executed, the cell lysate was subjected to exhaustive ultracentrifugation, gradient purification, and gel filtration, from which we recovered pure MVP-PARP4 complexes (Supplementary Figs. 1b and 2a).
Purified human MVP-PARP4 particles were then applied to carbon-coated grids and subjected to cryo-electron microscopy (cryo-EM) data collection and single-particle analysis. As a point of comparison, this process was repeated with apo-MVP particles (Supplementary Figs. 2 and 3). The resulting 3D maps allowed us to compare the human structures to previously published structures of the vault cage. Initially, we performed the reconstruction without enforcing any symmetry and found that the human vault cage adopts the same D39 symmetry as observed in previously published rat homolog structures2,18,19. Among several differences from these previous structures, however, is the prominent mass of poorly-resolved peptide potential jutting out from the tapered vault “caps”. Previous rat vault EM maps also demonstrate a slight protuberance at their ends, resulting from a C-terminal peptide that is blurred following symmetrization (Supplementary Fig. 4a). This feature, which we are calling the “knob” (Fig. 1b), is much more pronounced in the human vault particle, however (Supplementary Fig. 4a). This is likely the result of 30 additional residues at the human MVP C-terminus that are absent in the rat sequence (which is otherwise very similar, sharing 91% sequence identity with human MVP – Supplementary Fig. 4a). Our findings agree with previous work indicating that rat vaults engineered with a synthetic C-terminal tag exhibit a more prominent knob structure20. Because the knob is close to the symmetry axis and likely to deviate from perfect 39-fold symmetry, it could not be confidently modeled. However, the presence of the knob suggests that an intact human vault cage is not freely open at the ends (Supplementary Fig. 4b).
Our structural model also sheds light on interactions between the symmetrical halves of the human vault, which are thought to separate (either partially or completely) to engulf cargo into the cage18,21,22,23. Contrary to what was described in the original crystal structure of the rat vault cage, we did not observe salt bridges between E4 and R42 on MVP monomers that are directly opposite one another, despite the fact that the residues were conserved in both species (Supplementary Fig. 4c). This finding was in line with other rat vault cryo-EM structures that appeared after the publication of the crystal structure18,19, as well as a re-refinement of the original crystal structure24 (Supplementary Fig. 4d). In our structure, the previously observed hydrophobic interface18,19, which sits between medially symmetric copies of MVP, includes a phenylalanine in position 6 (Fig. 1c). Interestingly, the human F6 residue replaces an alanine at the same position in the rat MVP sequence (Supplementary Fig. 4c, d), indicating that the strength of the hydrophobic packing across the halves of the human vault cage may be stronger than that of the rat vault.
To either side of the hydrophobic interface, there are two conserved, charged residues, R9 and D39, the latter of which has previously been implicated in stabilizing inter-half associations18. Although the quality of the map at this relatively flexible region precludes the definitive assignment of their rotamer conformations, R9 and D39 appear to sit within range to form a salt bridge with one another and form, along with N22 and N40, a hydrogen bonding network of polar residues. These residues are located at the tight junctions between diagonally opposed copies of MVP. Their close proximity and apparent flexibility indicate that these four residues could dynamically interact with each of the others, depending on the local environment or the movement of the cage (Fig. 1c). In addition to contributing to the cumulative strength of the interaction between the cage halves, these polar residues could act as initial points of separation preceding the opening of the vault cage, depending on conditions such as changes in pH, salt concentration, or the introduction of other polar or charged molecules.
Interface between MVP and PARP4
The most striking feature in our MVP-PARP4 complex structure, however, was the prominent cryo-EM potential inside the vault cage. This additional potential, which corresponded to the PARP4 molecules occupying the cage, was clearly visible in the 3D reconstruction, 2D classes, and even in individual particles (Supplementary Fig. 2) when compared to the particles in the apo-MVP dataset (Supplementary Fig. 3). In a cross-sectional view, the C-terminus of PARP4 can be seen anchored proximal to the vault waist in between two beta-rich repeat domains of the MVP barrel. It then stretches out into the vault lumen, before the polypeptide chain moves towards the caps, hugging the walls of the cage (Fig. 1d). The long, disordered regions that bridge PARP4’s folded domains, their unconstrained movement in the vault lumen, and the variable occupancy of PARP4 per vault particle across the ensemble, rendered the majority of the enzyme too heterogeneous to be averaged to high resolution. Although processing the data with different symmetries did not measurably improve resolution for most of the protein structure (Supplementary Fig. 5a, b), imposing D39 symmetry ultimately allowed us to refine the MVP-PARP4 interface to near-atomic resolution (Supplementary Fig. 2). From there, we could dock and refine an AlphaFold model of PARP4’s MINT domain within the cryo-EM map (Fig. 1d), making relatively few alterations to the original coordinates (Supplementary Fig. 5c, d). The final refined model revealed the key structural details of PARP4’s binding within the MVP cage (Fig. 2a).
Prior to our structure, only inaccurate models of the MVP-MINT complex generated by predictive software, including GRAMM-X17 and AlphaFold, existed. Their inaccuracies may have been related to the complications presented by the curvature of the MVP cage with respect to MINT binding, since the alignment of bound MINT is out of register with the MVP cage symmetry (Supplementary Fig. 5e). Our structure shows that residues on MINT’s 4th and 5th, as well as its 7th and 8th, helices (when starting the count from the N-terminus of the domain) are crucial for MINT’s interaction with MVP (Fig. 2a). Surface analyses of the MINT-MVP interface reveal that it falls in a region of substantial electrostatic complementarity between MVP and PARP4. Specifically, an acidic stretch that runs along MVP’s third and fourth repeat domains aligns with a basic region between helices-4 and -5 of the MINT domain. Within this region, PARP4’s K1618 and MVP’s D214 residues forge a salt bridge, while several nearby residues engage in additional, specific electrostatic interactions. Notably, the side chains of PARP4 S1614 and R1689, as well as MVP R169 and Q164, appear to form hydrogen bonds with backbone amide groups of the opposite species. The side chains of PARP4 Q1682 and MVP Q172 are in range to hydrogen bond with one another as well (Fig. 2b). Having identified these residues as important drivers of complex formation, we sought to validate our structural results by mutating several of them to alanines and measuring the impact on binding.
a Atomic model (left) of the binding interface between the PARP4 MINT (blue) and MVP3-5 repeat domains (gray) from the FL PARP4-MVP structure (inset). MINT domain helices located at the PARP4-MVP interface are labeled with the letter h and associated helix number. Key residues from PARP4 and MVP are labeled in blue and gray, respectively. b The exposed interfaces of MINT (blue) and MVP (gray) are shown in cartoon (left) and surface (right) view representations. Critical interaction residues are labeled in black, with labels of additional residues that may support the interaction shown in gray. Surface views of the MINT and MVP3-5 models are colored by relative electrostatic potential (right), demonstrating electrostatic and steric complementarity at the interface. c Size exclusion chromatograms show individual traces of MVP3-5 (gray) and MINT constructs (blue), indicating their absorbance at 280 nm (upper). Individual traces are overlaid with complex traces following co-incubation of the WT (pink, left) and double mutant constructs (red, right). Corresponding SDS-PAGE gels (lower) from SEC binding assays showing co-elution of WT MVP3-5 and MINT (left) and the separate elutions of double mutant constructs (right) following their respective co-incubations and injections over a Superdex 75 increase column. Black bars denote the area over which fraction samples were collected, and numbers in the input lanes are shown in colors that correspond to the protein labels in the SEC traces above. The experiment was performed twice.
We designed PARP4 and MVP constructs that contained only the necessary interaction domains (MINT1562–1724 and MVP3-5102–274, respectively), both of which could be readily overexpressed in and purified from E. coli. Prior to injecting them over a gel filtration column, we incubated approximately equimolar concentrations of wild-type (WT) MINT and WT MVP3-5. The resulting peak eluted at a lower volume than the elution volumes of each individual species. We confirmed that they comigrated by SDS-PAGE gel (Fig. 2c). When we pooled fractions from the complex peak and re-injected the sample over the column, we observed a peak that eluted at the same volume, demonstrating the considerable stability of the complex (Supplementary Fig. 6a). After we confirmed that these domains could bind and remain stably intact over a size exclusion column, we mutated some of the previously identified interacting residues (the PARP4 residues S1614 and K1618 and MVP residues R169 and D214) to alanines. We dubbed the respective constructs the “SKA” and “RDA” mutants and used gel filtration to assay each construct’s ability to bind the corresponding species. Both mutant constructs (Fig. 2c), as well as combinations of each mutant and WT species (Supplementary Fig. 6b), were co-incubated prior to injection over a gel filtration column. In each case, we saw no evidence of complex formation between the truncation constructs, indicating that the interaction between PARP4 and MVP involves specific electrostatic interactions between the key residues. To verify that the mutations specifically altered the properties of the interaction site and not global protein conformation or stability, we used circular dichroism (CD) spectroscopy to confirm that the mutants exhibited the same secondary structure and thermal stability as their WT counterparts (Supplementary Fig. 7).
Structure of the NAD+-bound MVP cage
As a complement to our structure of an MVP bound to PARP4 alone, we sought to collect a cryo-EM dataset of the complex in the presence of NAD+ to determine whether active PARP4 would adopt an alternate conformation or whether its activity would influence the conformation or stability of the vault cage. Our previous observation that polar and charged residues stabilized inter-half MVP interactions led us to hypothesize that PARP4 could modify these residues with bulky ADP-ribose groups, consequently triggering an initial separation between the halves that could destabilize the cage and permit cargo entry. To test whether active PARP4 would destabilize the cage, we incubated the MVP-PARP4 complex with 1 mM of NAD+ for several hours, dialyzed excess NAD+ out overnight, and froze grids with the complex solution for data collection. Contrary to our expectations, introducing NAD+ did not appear to decrease the proportion of intact vault cages on the grid (Supplementary Fig. 8). However, after collecting and processing a full dataset of these MVP-PARP4 complexes in the presence of NAD+, we noticed some additional, previously unobserved cryo-EM potential within the interior of the “shoulder” region of the vault cage. We soon found that we could dock an NAD+ molecule into this potential (Fig. 3a, b). Notably, we also observed changes in MVP’s structure near the binding site, providing additional evidence for the notion that MVP binds NAD+ directly. Namely, when NAD+ was present, several residues in the binding site (including R437, L517, and R511) adopted different rotamer conformations, and a loop appeared to be displaced from its previous position directly in front of the pocket (Fig. 3b).
a Cross-section of the upper half of the MVP cryo-EM map, with adjacent MVP monomers (black, cartoon view) fit into the potential. b Magnified top view (left) of the ligand signal in the MVP-PARP4-NAD+ map (pink) compared to the corresponding vacant site in the MVP-PARP4 map (gray). An atomic model of NAD+ is docked into the site in the MVP-PARP4-NAD+ map (upper panels). Coordinates for each structure are shown in cartoon view, with zoned NAD+ potential in the MVP-PARP4-NAD+ map shown as black mesh. Residues that adopt different rotamers in the presence and absence of NAD+ are labeled in gray and pink, respectively, and the “keyhole loop” is boxed with a black dashed line. Front-facing views of the same maps (right). c Adjacent NAD+-bound MVP monomers MVP1 and MVP2 in cartoon view (colored blue and pink respectively, left) and in surface view (colored by electrostatic potential, right). Magnified views of MVP1 and MVP2 SPFH domains, with the keyhole loop shown in a darker hue (inset, left). Surface view of the same model (inset, right). Bound NAD+ is shown in purple. d Atomic model of MVP’s NAD+-binding site. Side chains (stick view) of residues from MVP1 and MVP2 (blue and pink, respectively) that comprise the boundaries of the site are shown in cartoon view and labeled (upper left). Side chains of residues from MVP2 only, including those that neutralize the diphosphate’s positive charge (lower left). Stick view of the same residues within the protein surface, which is colored by electrostatic potential (upper and lower right).
This newly discovered ligand-binding site sits at the interface between two MVP monomers (“MVP1” and “MVP2”). Interestingly, it is found within their SPFH (Stomatin/Flotillin/Prohibitin/Hflk) domain, an oligomeric protein motif enriched in lipid microdomains throughout the tree of life25. In addition to the canonical α-helical and β-sheet sub-domains characteristic of SPFH family proteins26, MVP’s SPFH domain contains an unusual loop we have dubbed the “keyhole loop”. This loop extends into the vault lumen, creating the small pocket that holds NAD+ (Fig. 3b, black dashed boxes). To verify that the potential represented bound NAD+ and not covalently attached ADP-ribose (ADPr), we employed two experimental strategies. First, we obtained the structure of MVP alone (without the ADP-ribosyltransferase PARP4) in the presence of NAD+, and then we conducted a mass spectrometry-based analysis to detect ADP-ribosylation on the vault cage. In our MVP-NAD+ cryo-EM map, we observed the same ligand potential in the SPFH domain pocket, indicating that MVP specifically binds NAD+ and that PARP4 is not necessary for the interaction (Supplementary Fig. 9). Further, after incubating NAD+ with both the MVP-PARP4 complex and MVP alone and submitting both samples for LC–MS/MS, we were unable to identify an ADPr modification near the binding pocket by LC–MS/MS in either sample. This provided additional evidence that the cryo-EM potential corresponded to non-covalently bound NAD+. Whereas the majority of the vault’s interior surface is relatively smooth and acidic, the binding site of NAD+ is a recessed pocket lined by basic residues (Fig. 3c), which we determined to be crucial for recruiting and stabilizing NAD+.
Biochemical basis for the MVP-nucleotide interaction
Analysis of the amino acid side chains lining the vault’s ligand-binding pocket revealed that MVP1 and MVP2 play distinct roles in stabilizing bound NAD+. Residues on MVP1 provide important steric constraints, with residues P508, H509, and P565 defining the left-side boundary of the pocket. In addition, H509’s side chain appears to hydrogen bond with the OH groups of NAD+’s distal ribose. MVP1’s R507 and K573 residues may serve to help neutralize the negative charge introduced by NAD+’s phosphates (Fig. 3d). However, the basic residues on MVP2’s keyhole loop (R504, K506, and R511) likely do so to a much greater degree, with R511 and K506 in range to form salt bridges with the diphosphate group. Additional MVP2 residues, including R437, D521, and W545, appear to form hydrogen bonds with the ribose moieties of NAD+, strengthening the electrostatic attraction between MVP2 and its ligand. Finally, hydrophobic residues from MVP2 (L517 and F522) form a border along the top and bottom of the pocket (Fig. 3d).
In all, our structural data depict a well-defined, distinct NAD+-binding fold between the SPFH domains of two adjacent MVP monomers, whose electrostatic properties are ideal for stabilizing NAD+’s diphosphate and ribose groups at the site. We therefore hypothesized that this pocket could also bind small molecules with structural motifs similar to NAD+. To investigate the potential versatility of the pocket, we determined the cryo-EM structure of human MVP in the presence of ADP (Supplementary Fig. 10). Similar to the NAD+-bound structure, the cryo-EM map exhibited additional potential behind the keyhole loop. However, this potential was not as distinct or as prominent as that found in the MVP-NAD+ map (Supplementary Figs. 11a and 12). We also conducted isothermal titration calorimetry experiments to determine MVP’s affinity for the ligands and found that both NAD+ and ADP bound in the tens of micromolar range (Supplementary Fig. 11b), suggesting that the combination of a purine base, ribose sugar, and diphosphate group are adequate for the ligands’ association to the vault cage.
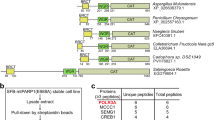
Identifying the nucleotide-binding site prompted us to ask if there was a functional link between the vault’s resident PARP enzyme and MVP’s ability to harbor its enzymatic substrate (given the importance of NAD+ availability in PARP regulation27). Namely, we wondered whether MVP’s nucleotide-binding site could provide an NAD+ reservoir for vault-bound PARP4. Although we were unable to identify its modification targets, enzyme assays indicated that vault-bound PARP4 is able to consume NAD+ substrate (Supplementary Fig. 13), suggesting that it retains its catalytic activity in the vault. Intriguingly, when we attempted a rigid body fit of the previously published crystal structure of PARP4’s catalytic module (comprising its HD and ART domains), we found that the volume of PARP4's potential adjacent to MVP’s SPFH domain was wide enough to accommodate this region (Fig. 4a, b). To further probe the relationship between MVP’s NAD+- and PARP4-binding sites, we analyzed the conservation of key residues at those sites by performing a multiple sequence alignment (MSA) of MVP from several representative vertebrate and invertebrate species. We found that the conservation patterns of MVP residues involved in PARP4 binding (present only in the vertebrate species) differed from those responsible for the NAD+ interaction (which had variable conservation of residues between the two groups) (Fig. 4c). This could reflect a model in which PARP4 began to opportunistically occupy the vault in higher order organisms to take advantage of stored NAD+ as an enzymatic substrate.
a Cross-sectional views of the MVP-PARP4-NAD+ (ternary complex) cryo-EM maps, with color coding by molecular species (left) and local resolution (center), as well as a transparent view of the map with the MVP-MINT-NAD+ molecular model docked into the high-resolution region (right). b Domain diagram of PARP4 (upper) with the HD-ART and VIT-VWA regions highlighted by dashed line boxes. Cross-sectional view of the ternary complex map with AlphaFold2 models and crystal structure coordinates (PDB 8SX174) of the boxed PARP4 domains docked into it, along with models of MINT and NAD+ for reference (left). View of the docked models turned 90° to show their alignment in the map (right). Magnified views of the docked models (insets, lower). Label colors correspond to the color of each model. c Multiple sequence alignment (MSA) of the PARP4 and NAD+-binding regions of MVP in a representative sample of vertebrate (blue) and invertebrate (black) species. Residues important for human MVP’s interaction with PARP4 and NAD+ are outlined in gray and pink, respectively.
Vault coimmunoprecipitation (co-IP) from WT and PARP4-depleted cells
In order to find native vault interaction proteins, we developed an anti-human MVP synthetic antigen binding reagent (sAB) that could be biotinylated and subsequently used for proteomic analysis (Fig. 5a). We chose to use A549 cells (derived from human lung epithelia) as our native vault source due to their high constitutive expression of vault proteins and vault’s documented, medically relevant activity in lung tissue5,6,28. We engineered a PARP4-depleted cell line using CRISPR/Cas9 so that we could determine PARP4’s influence on the vault interactome. We verified protein-level PARP4 depletion via Western blot (Fig. 5b), though spectral counts in our MS data suggested that a very small amount of PARP4 peptides remained. Significant hits from our co-IPs (with log2(Fold Change) > 1.5 and −log10(p-value) > 1.3) included known MVCs (black data points, Fig. 5c, d) and COP1 (an E3 ubiquitin ligase previously observed to interact with vault29), which validated our approach. We identified a total of 52 hits from the WT A549 cells (Fig. 5c) and subjected them to gene ontology (GO) network analysis (Fig. 5e) to evaluate their biological roles and cellular compartmentalization. We then compared the co-IP hits from the WT and PARP4-depleted cell lines, sorting them into three groups. 34 “WT hits” were either exclusively identified or significantly enriched in WT cells, while 16 “KO hits” were uniquely present in our PARP4-depleted cells. 18 “shared hits” were identified in both datasets without significant differential enrichment in either (Fig. 5f).
a Cryo-EM image of an MVP-MINT particle with anti-MVP sABs (indicated by red arrows) bound to each cap (right panel), next to an MVP-MINT particle without bound sABs (left panel). Schematic of anti-MVP coimmunoprecipitation (co-IP) experiment (right). b Proof-of-concept anti-MVP and -PARP4 western blot showing vault proteins pulled down from WT and PARP4 KO A549 cells by the anti-MVP sAB. The blot depicts chemiluminescent signal corresponding to MVP, PARP4, and α-tubulin (as loading control) in co-IP samples from WT, MVP KO, and PARP4 KO cells. PARP4 depletion in the KO cell line was verified by Western blot at least four times. c Volcano plot showing differential enrichment of species co-IP’ed with anti-MVP (right) and anti-GFP control (left) sABs from WT A549 cells. Enrichment (log2(fold change) of peptide abundance from the experimental/control IP condition >1.5) and significance (−log10(p-value), calculated via two-tailed homoscedastic t-test, across triplicate experiments >1.3) shown as solid and dashed lines, respectively. Representative enriched hits are labeled, with vault components written in black text and genes associated with gene regulation, nucleotide metabolism, and cell migration and invasion written in pink, gold, and teal, respectively. d Volcano plot showing differential enrichment of species co-IP’ed with the anti-MVP sAB from WT A549 (right) and PARP4 KO A549 cells (left). e Network of hits co-IP’ed with MVP from WT A549 cells. Connecting lines represent previously established interactions (left). Key showing the proportion of proteins tagged with significant gene ontology (GO) biological process terms and their false discovery rates (FDR) (right). f Venn diagram showing hits unique to or significantly enriched in the WT (versus PARP4 KO) co-IP dataset (blue, left), hits unique to the PARP4 KO cell co-IP dataset (pink, right), or hits present in both datasets without significant differential enrichment in either (purple, center). g Bar graphs depicting the proportion of proteins in each dataset tagged with specific GO biological process (upper) and cellular component terms (center), as well as the proportion of those previously observed to either interact with (left) or be modified with (right) ADP-ribose (lower).
Many of the most overrepresented genes in all three groups were active in pro-survival and pro-proliferative pathways, including a number of cytosolic enzymes involved in nucleotide metabolism (gold data points, Fig. 5c, d), and proteins that operate at the gene regulation level (pink data points, Fig. 5c, d). WT hits in the “gold set” included ACAD11, AKR1C3, and AKR1B1, all nucleotide-binding dehydrogenase/reductases that promote cell growth and survival in the wake of stress30,31, while those in the “pink set” included RUVBL232, HNRNPDL33, and DARS2. Although it is canonically known as a tRNA synthetase, DARS2 has recently been implicated in regulating metabolism by regulating splicing34 and transcription35. A third set of highly abundant genes in our pulldowns regulate cell migration or invasion (teal data points, Fig. 5c, d), including TRO36, one of the most abundant hits, as well as RACK137, WDR1138, MICAL239, and — the only gene in this group to inhibit cell migration — TCAF140. Finally, SLC25A1, a solute carrier, emerged as an interesting hit for its homology with another solute carrier that transports NAD+ into the mitochondria41. Broadly speaking, most WT hits were associated with either the “metabolic process” or “organelle organization” GO biological process terms. Their “cellular component” GO terms (derived from the Ensembl database) place them throughout the cell, with a little over a quarter able to localize to the plasma membrane and a little over half present in the nucleus.
By contrast, according to their cellular component GO terms, none of the KO hits localize to the plasma membrane, and over three-quarters spend time in the nucleus. Further, fewer KO hits were associated with organelle organization than WT hits, though a similar proportion were involved in metabolic processes (Fig. 5g, Supplementary Table 1). Like the WT hits, many of the KO hits belonged in the “pink set” for their ability to regulate cell growth and survival at the gene expression level. Among them were the KRAB-type zinc fingers ZNF420, a known negative regulator of apoptosis42, and ZNF45, whose expression has been observed to be upregulated in some tumor types43. Three functionally related RNA-binding proteins also emerged as prominent hits, all of which have been shown to regulate gene expression following interactions with the 3′UTRs of various mRNAs44,45. The most abundant of these, DAZAP1, has also been implicated in splicing events that promote cell growth46, while the others (PUM1 and PUM2) have been associated with cancer cell proliferation44,47.
The shared hits exhibited GO functions and subcellular localization patterns broadly similar to those of the WT hits (Fig. 5g). “Gold set” pro-growth/-survival metabolic enzymes include LMTK248 and ALDH1A149 (which uses an NAD cofactor), while “pink set” hits that regulate gene expression were represented by components of the SMC5/6 complex. This assembly is a cohesin/condensin-like structure that promotes cell growth and protects the cell from genotoxic stress (and even viral infection) by interacting with DNA and mediating homologous recombination (HR)50,51. The complex’s ability to regulate HR at telomeres also links SMC5 and SMC6 to organelle organization and “telomere organization” functions in the GO network analysis, a function shared by several other hits, including the MVC TEP1. Hits with the “organelle organization” designation also included KPNB1, which, along with IPO7, regulates nucleocytoplasmic shuttling52, a previously hypothesized function for vault. Finally, GMPPA, which mediates protein glycosylation as a regulatory subunit in the GDP-mannose biosynthetic enzyme complex, has been linked to tumor growth53 and chemotherapeutic drug resistance54, pathologies previously linked to MVP upregulation7,8.
Discussion
Our structure of the MVP-PARP4 complex resolves a longstanding question in the field regarding the biochemical basis for the MINT domain’s association with the vault cage. Along with shedding light on the vault’s native function, this structure could inform efforts to develop vault-based biomolecule delivery systems that use MINT or mutants thereof as a fusion domain17. Interestingly, when we constructed full-length PARP4 and MVP mutants with SKA and RDA mutations, respectively, we found that the point mutations reduced, but did not eliminate PARP4 incorporation into the MVP cage (Supplementary Fig. 6c). This may be due to the elevated concentration of each protein in our recombinant system or could indicate the presence of additional interaction interfaces between MVP and PARP4 aside from the one we resolved between the MVP3-5 and MINT domains. In support of the latter possibility, we identified some non-resolvable potential along the interior wall of the vault cage cap that appears to be exclusive to complexes with PARP4 (Supplementary Fig. 6d) and could conceivably represent an additional point of contact between the two vault components. Going forward, characterization of this and any other PARP4-MVP interactions will further our understanding of PARP4 recruitment and stabilization in the vault cage.
In addition, our discovery of MVP’s structurally distinct nucleotide-binding pocket introduces several new questions to the field. Provided that NAD+ is MVP’s preferred ligand, its association with the vault cage could have a number of functional consequences. By analogy to the functions of other NAD+-binding proteins55, MVP’s interaction with NAD+ could potentially stabilize a particular conformational state56,57 (thereby altering MVP’s binding properties58) or influence NAD+ compartmentalization and availability for enzymes that require it41. Both functions appear possible. With respect to the former, bound NAD+ displaces a loop (which was too flexible to be completely modeled) just beneath MVP’s SPFH domain, appearing to push it further into the lumen (Fig. 3b) where it could potentially interact with vault cargo or MVCs more readily. As to the latter function, NAD+’s proximity to PARP4, as well as our discovery of vault cargo molecules that bind NAD+ (Supplementary Table 1), could signal vault’s ability to act as a repository for enzymes that utilize NAD+ as a cofactor or substrate. While we did not detect a PARP4 modification target or very robust NAD+-consumption activity in our system, it is possible that, in vivo, PARP4 interacts with additional factors that promote its catalytic activity.
The identification of additional ligands that may occupy the binding site raises further questions about the site’s function. Although the ligand potential in our MVP-ADP cryo-EM map appeared weaker than that in our MVP-NAD+ map (possibly due to its smaller size), ADP’s affinity for the MVP cage was on the same order of magnitude as NAD+ in ITC experiments (Supplementary Fig. 8d). The cellular concentration of ADP is generally much lower than that of NAD+, but it can increase rapidly in response to the introduction of metabolic stress59, suggesting that different ligands could compete to occupy the binding site under different conditions. Finally, given ADPr’s structural similarity to NAD+, the SPFH domain could conceivably serve to anchor ADP-ribosylated substrates in the vault lumen. Such interactions, however, would likely require substantial conformational changes within the SPFH domain to accommodate the additional bulk of the modified protein or nucleic acid attached to the ADPr moiety. Although further work is needed to define its precise function, MVP’s nucleotide-binding capacity indicates that vault may be able to adapt its behavior in response to changes in cellular metabolite concentrations, offering one potential explanation for its functional versatility in different cell types and conditions.
PARP4’s role in the vault complex still requires investigation. However, our proteomics data indicate that vaults from WT and PARP4-depleted cells exhibit distinct proteins in their interactomes, suggesting that PARP4 may play a role in regulating vault’s cargo selection or subcellular localization. “KO hits” from PARP4-depleted cells were more likely to be found in the nucleus. Accordingly, they were often associated with gene regulation and nucleic acid binding. By contrast, many “WT” hits localized to the plasma membrane and regulated cell migration and invasion functions. PARP4’s VWA domain (a protein-protein interaction domain present in cell surface proteins that regulate cell migration60) could conceivably facilitate vault’s recruitment of certain cargo proteins. We have not yet determined whether PARP4’s catalytic activity influences vault cargo recruitment or if the cargo molecules themselves are being modified. Our analysis of WT and KO hits indicates that a similar proportion of genes in each dataset have previously been observed to bind ADP-ribose61,62,63. However, a greater proportion of WT hits than KO hits have been observed to accept ADP-ribose modifications64. This subset of WT hits could constitute some of vault-bound PARP4’s modification targets (Fig. 5g, Supplementary Table 1). Further exploration of these molecules could thus shed light on the vault’s roles in cell signaling. More broadly, our identification of native vault-binding partners involved in cell growth, proliferation, chemoresistance, and migration can begin to explain MVP’s previously observed roles in cancer progression7,65 and drug resistance8, as well as metabolic regulation10, a function now further cemented by our discovery of its association with the critical metabolite NAD+.
Methods
Plasmid construction and baculovirus generation
Human MVP and PARP4 genes were synthesized by GenScript and subcloned into the pVL1393 baculovirus transfer vector. pVL1393-MVP and pVL1393-PARP4 plasmids were incubated with BestBac2.0 linearized baculoviral DNA (Expression Systems, 91-002) and transfected into adherent Sf9 insect cells using Cellfectin II (Invitrogen 10362-100). After 7 days, the cells were pelleted by centrifugation, and P0 baculovirus was harvested from the supernatant fraction. P0 virus was amplified in Sf9 suspension cells for an additional week to generate P1 baculovirus stocks. P1 viruses were, in turn, amplified in Sf9 cells to generate P2 stocks. P1 and P2 stocks were used to infect Hi5 cells for protein expression. Constructs for E. coli expression of WT and mutant MVP3-5 and MINT domains were cloned into the pET47b(+) vector, which introduced an N-terminal His-tag and HRV 3C cut sites. Alanine mutations in the MVP and PARP4 truncation constructs and full-length sequences were imposed by site-directed mutagenesis and confirmed by Sanger sequencing. All primers used for this study are listed in Supplementary Table 2.
Expression and purification of vault cage complexes
Hi5 cells were grown to one liter at a density of about 2 × 106 cells/ml and infected with 1 ml MVP P1 baculovirus. Cells grown to produce the MVP-PARP4 complex were simultaneously infected with 5–6 ml hPARP4 P2 baculovirus. Infected cells were rotated on an orbital shaker at 120 rpm at 27 °C. After 65 h, they were harvested by centrifugation at 900 × g for 15 min. Cell pellets were washed with 20 ml of phosphate-buffered saline (PBS) per 0.5 L of cells, transferred to 50 ml Falcon tubes, and centrifuged for an additional 15 min at 900 × g. Cells were decanted, and aliquots were flash frozen in liquid nitrogen prior to long-term storage at −80 °C. For purification, 0.5 L cell aliquots were thawed in room temperature water for 10 min, then resuspended in 120 ml Buffer A (50 mM Tris pH 7.4, 75 mM NaCl, 1.5 mM MgCl2, & 1 mM DTT), with 1% Triton X-100 and 1 mM phenylmethylsulfonyl fluoride (PMSF). Pellets were disrupted with 30–40 strokes of a Type A Dounce Homogenizer (VWR, cat. No. KT885301-0100). Lysate was transferred to centrifuge tubes and vortexed twice during a 20-min incubation on ice. Lysate was cleared by centrifugation at 20,000 × g for 15 min, and the supernatant was harvested for additional purification steps. High-molecular-weight complexes were pelleted by centrifugation at 150,000 × g for one hour. Pellets were resuspended in 6 ml of Buffer A with 7% Ficoll (GE Healthcare 45-002-020) and sucrose added. The resuspension was centrifuged for 45 min at 43,000 × g in order to pellet microsomal contaminants. The supernatant fraction was diluted in 21 ml of Buffer A and pelleted by ultracentrifugation for 3 h at 200,000 × g. Pellets were resuspended in 0.8 ml of Buffer A and incubated with 500 µg RNase A as well as 50 U of RNase T1 for 15 min. Denatured ribosomal proteins were pelleted by centrifugation at 30,000 × g for 20 min. The supernatant was then transferred to a new 1.5 ml tube for an additional 15-min spin. The remaining supernatant was taken up to 1 ml with Buffer A and gently pipetted over a 20%–60% sucrose gradient. Gradients were centrifuged at 78,000 × g for 16 h. Fractions from the 45% and upper half of the 50% gradient fractions were pooled and dialyzed in ADP-ribosylation buffer (50 mM HEPES pH 8, 5 mM MgCl2, 5 mM CaCl2, 0.25 mM DTT) overnight. Dialysate was concentrated to 500 µl and injected over a Superose 6 increase 10/300 GL column (Cytiva). Fractions were aliquoted and flash frozen with liquid nitrogen prior to storage at −80 °C.
To assess full-length mutant versus WT PARP4 incorporation into mutant and WT vault cages, full-length SKA PARP4 + RDA MVP and WT PARP4 + MVP were co-expressed and purified over a sucrose gradient as described above. The ratio of the adjusted volumes of MVP and PARP4 gel bands in each corresponding lane was calculated. Bands were detected using ImageLab 6.0.1, with “custom” sensitivity set to 50%. Adjusted volumes (int.) of bands corresponding to MVP and PARP4 were determined (with default background subtraction settings) to calculate relative ratios of MVP:PARP4 band intensities.
Expression and purification of MVP3-5 and MINT truncation constructs
pET47b-MVP3-5 and pET47b-MINT WT and mutant constructs were transformed into Rosetta 2(DE3)pLysS Singles E. coli competent cells. Protein production was induced by the addition of 0.5 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) at OD600 0.6–0.75, and cells were kept at 18 °C and rotated at 220 rpm overnight. For purification, cells were pelleted by centrifugation at 7000 × g for 20 min, and each pellet was resuspended in 50 ml Ni-NTA buffer (20 mM Na2HPO4/NaH2PO4 pH 7.4, 300 mM NaCl, 1 mM PMSF) with 10 mM of imidazole. Resuspended cell solutions were subjected to sonication for 3 min at the 75% amplitude setting, with a pulse sequence of 3 s on/5 s off (QSonica). Lysate was cleared by centrifugation at 37,400 × g for 45 min, and the supernatant was flowed over 6 ml of washed Ni-NTA resin (ThermoFisher, cat no. PI88223) in a gravity column. Resin beads were washed with 150–200 ml Ni-NTA buffer with 20 mM Imidazole. Proteins were eluted from the column following 5-min incubations in 15–25 ml Ni-NTA buffer with 250 mM Imidazole. Protein eluate was dialyzed into 2 L SEC buffer (50 mM Tris pH 8, 150 mM NaCl, and 0.5 mM tris(2-carboxyethyl)phosphine (TCEP-HCl)) and subjected to His-tag cleavage with PreScission Protease overnight. Dialyzed protein was concentrated, filtered, and injected over a Superdex 75 increase 10/300 GL column (Cytiva). Peak fractions were pooled and used immediately for gel filtration peak shift assays.
Cryo-electron microscopy sample preparation
Vault cage peak fractions were concentrated and incubated at 4 °C for 3.5–4 h in the presence of 1 mM NAD+, 1 mM ADP, or no ligand. Samples were dialyzed in ADPr reaction buffer overnight to remove excess ligand. Vault proteins were then concentrated and subjected to 5 min of centrifugation at 16,000 × g to clear aggregates. Quantifoil R1.2/1.3, 200 mesh copper grids with 2 nanometers of continuous carbon film were glow discharged using a 15 s, 5-watt program on the Solarus 950 Plasma Cleaner System (Gatan). Three microliters of sample (~1.5–1.8 mg ml−1) were applied to the carbon-coated grid surface. The climate chamber of the Vitrobot Mark IV (ThermoFisher) was set to 8 °C and 100% humidity. Grids were blotted for 1–2 s at blot force 0 with standard Vitrobot filter paper (Ted Pella, 47000-100), then plunge-frozen into liquid ethane.
Cryo-electron microscopy data collection
Data were collected at one of two facilities, both of which used a 300 kV Titan Krios transmission electron microscope (ThermoFisher). At the Advanced Electron Microscopy Facility at the University of Chicago, movies were recorded on a K3 direct detector camera (Gatan) at a nominal magnification of 64,000× (translating to a pixel size of 0.672 Å at the detector), in super-resolution counting mode by image shift. EPU software was set to automated acquisition mode and collected (depending on the dataset) between 4951 and 6968 image stacks, each with 40 frames, subject to a total dose of 45 e−/Å2. The defocus range was set to −1.0 to −2.5 μm. At the HHMI Janelia Farms Cryo-Electron Microscopy Facility, movies were recorded on a K3 Bioquantum camera (Gatan) at a nominal magnification of 53,000× (translating to a pixel size of 0.66 Å at the detector), in super-resolution counting mode by image shift. SerialEM software was used to collect data using a strategy of 3 by 3 + 1 shots per hole, acquiring (depending on the dataset) between 3551 and 7386 image stacks. The defocus range was set to −0.7 to −1.7 μm. See Table 1 for additional details.
Cryo-electron microscopy image processing
Stack images were exported to cryoSPARC live (v3.3.1 or v4.4.0), where they underwent motion correction and CTF determination. Particles were picked using 2D class averages generated from initial templates, then exported to cryoSPARC66 for additional processing. Contaminants, broken vaults, and poorly aligned classes were eliminated following initial 2D classification. Depending on the dataset, between 174,624 and 670,709 particles were used to generate the initial set of 2D classes or ab initio models (which were typically of low quality but useful as templates to sort particles during heterogeneous refinement). For the first homogeneous refinement, a low-pass filtered MVP cage map from an earlier dataset served as the initial model, with D39 symmetry imposed. Additional 2D classification or heterogeneous refinement jobs were used to further sort vault cage particles. The resulting particles were typically used as inputs for subsequent rounds of either homogeneous or non-uniform refinement, in which the program was instructed to optimize the per-particle defocus and per-exposure-group CTF parameters, fit the anisotropic beam magnification, and apply Ewald sphere correction using the correct curvature sign, as determined by running homogeneous reconstruction with both. See Supplementary Figs. 2, 3, 8, 9, and 10 for more details.
Cryo-electron microscopy model building, refinement, and validation
Model building was completed in COOT67 with starting models including the rat homolog of MVP from a previous crystal structure of the vault cage (PDB accession no. 4V60) and PARP4’s MINT domain predicted by AlphaFold268. Small-molecule coordinates were retrieved from the REFMAC monomer library in COOT. Full-length PARP4 was present in the complex, but only a model of the MINT domain could be confidently built into the potential. The potential at the interaction interface of the MVP-PARP4 maps initially seemed to suggest a 1:1 MVP-PARP4 ratio, because the most prominent features of the MINT domain (large, rigid helices) appeared 78 times throughout the cage. However, when we began to build the model, it became clear that space constraints within the cage would not permit this stoichiometry. Ultimately, we had to delete two regions on either side of the MINT domain (residues 1587–1593 on one side of the domain and 1646–1650 and 1655–1661 on the other) for the map to accommodate adjacent MVP-MINT coordinates. Because we observed this phenomenon with or without the imposition of symmetry, we believe that the overestimation of bound MINT domains was an artifact inherent to aligning a symmetrical cage with different configurations of an internally bound protein during data analysis. The final model, containing two copies of MVP related by a 2-fold axis, was refined in real space and validated using PHENIX69. See Table 1 for the details of model statistics.
Isothermal titration calorimetry (ITC)
MVP cage protein was purified mostly as described above, with the exception that the buffer was DTT-free (since it tends to oxidize during injection and distort the ITC baseline). The sample was concentrated to 320 µl and subjected to a 10-min spin at 16,000 × g, at which point the supernatant was transferred to a new tube. The MVP monomer concentration was calculated to be approximately 50 µM by protein absorbance measurement at 280 nm. 5 mM solutions of NAD+, ADP, and adenine were prepared in the buffer used for MVP SEC. ITC binding experiments were conducted using a MicroCal PEAQ-ITC instrument (Malvern Panalytical). Following an initial 0.4 µl ligand injection, 2 μl ligand solutions were titrated into buffer (as a control) or MVP 18 times at a rate of 2 s μl−1 at 150 s time intervals. Stirring speed and experimental temperature were kept at 750 rpm and 25 °C, respectively. Estimated binding parameters were derived from the line fit. For the fit, the number of binding sites (N) was held at one (based on our structural data), plots underwent baseline adjustments, and ligand-only isotherms were assigned as controls and subtracted via the line method in the Malvern PEAQ-ITC software (version 1.41).
Gel filtration peak shift assays
Wild-type and mutant MINT (100 µM) and MVP3-5 (90 µM) constructs were individually injected onto a Superdex 75 increase 10/300 GL column (Cytiva) to establish the peak height and elution volume of each construct. To assess complex formation, wild-type MINT (100 µM) was incubated with wild-type MVP3-5 (90 µM) overnight at 4 °C. The mixture was then injected onto the Superdex 75 increase column. This process was repeated with a mixture of the mutant MINT and MVP3-5 constructs, as well as with combinations of each WT and mutant construct. 2× Laemmli loading dye was added to peak fraction samples, which were then boiled and loaded onto 15% SDS-PAGE gels. Bands were resolved with Coomassie stain.
A549 cell culture and PARP4 KO cell line generation
A549 cells were cultured in DMEM (MilliporeSigma) supplemented with 10% fetal bovine calf serum (Cytiva), 100 U ml–1 penicillin, and 100 µg ml–1 streptomycin in a humidified incubator (with 95% air, 5% CO2) at 37 °C. Mutant cells were generated via CRISPR/Cas9 transfection. A guide RNA targeting PARP4 was annealed into the pSpCas9(BB)-2A-GFP plasmid (PX458, addgene) (Supplementary Table 2). The guide RNA-Cas9 plasmid was transfected into A549 cells using Lipofectamine 3000 (ThermoFisher) and a cell line-specific protocol provided by ThermoFisher (58). Cells were viewed under a fluorescence microscope to verify that transfection had occurred, then trypsinized and pooled for fluorescence-assisted cell sorting (FACS) at the University of Chicago Flow Cytometry Core. Each cell that exhibited low-level fluorescence was selected and transferred to an individual well of a 96-well plate. Plates were transferred to a 37 °C incubator and kept there overnight, after which the cells were supplemented with fresh media. Single clone cell lines were expanded to larger plates as needed. The genomic DNA of the cell lines that persisted was isolated and genotyped to identify mutants. Western blot assays were performed to verify protein-level depletion (Fig. 5b).
anti-MVP sAB selection and purification
Human MVP was expressed and purified as described above, with the exception that MVP was purified in primary amine-free buffer (150 mM NaCl, 50 mM HEPES pH 7.4, 1.5 mM MgCl2, 0.1 mM TCEP-HCl) prior to sAB selection using the phage display library as described previously70. MVP was biotinylated and immobilized on streptavidin beads, at which point the beads were washed with selection buffer and subjected to four rounds of phage display panning at increasingly lower concentrations of MVP (200–10 nM). The nineteen sAB candidates that emerged from the final round of clones were subjected to a phage ELISA assay, and the twelve that displayed the greatest affinity and specificity for MVP were subcloned into the RH2.2 expression vector for subsequent purification as documented previously71.
Vault immunoprecipitation and mass spectrometry
Three anti-MVP and control IP biological replicates were conducted per cell line (WT versus PARP4 KO) for a total of twelve samples. WT and PARP4 KO A549 cells were seeded (at 7.5E5 cells per 15-cm plate) three days prior to the experiment. Two 15-cm plates were seeded per condition replicate (for a total of 24 plates). On the day of the experiment, cells were washed with 8 and then 4 ml ice cold PBS, then subjected to lysis with 1.7 ml fresh NETN buffer: 150 mM NaCl, 50 mM Tris pH 7.5, 5 mM EDTA, 0.5% NP-40 (MilliporeSigma), 1 mM DTT, 1 mM PMSF, 1 complete protease inhibitor tablet (Roche) per 20 ml buffer, and 250 nM ADP-HPD PARG (MilliporeSigma) inhibitor. Cells were incubated with NETN buffer at 4 °C for 6 min, at which point the lysate was collected with a cell scraper and transferred to a 2 ml centrifugation tube. Tubes were vortexed for 30 s and kept on ice for an additional 15 min. Cell debris and nuclei were pelleted for 8 min at 16,000 × g, after which the lysate supernatant was transferred to a fresh 2 ml tube and placed on ice. Following a Bradford (Bio-Rad) assay to quantify total lysate protein material per replicate, volumes containing 6 mg lysate protein were used as starting material per coimmunoprecipitation replicate. A total of 55 mg (per replicate) of biotinylated anti-MVP or anti-GFP (control) sAB were incubated with magnetic streptavidin beads (Promega, Cat. No. Z5482) at 4 °C. After being washed with NETN buffer three times, the sAB-bead mixtures were added to the harvested lysate and incubated in an overhead rotator at 4 °C for two hours in order to limit the binding of false positives. Samples were then subjected to a stringent series of eight washes with NETN buffers with varying NaCl concentrations. The wash cycle started with the highest (2× at 500 mM) NaCl concentration and progressed to lower (3× at 300 mM), then lowest (3× at 150 mM) NaCl concentration buffers. All washes were conducted on an overhead rotator at 4 °C. Finally, immunoprecipitated proteins were briefly washed with 900 µl ddH2O to eliminate buffer contaminants, then eluted from the streptavidin beads with 82 µl 0.1 M acetic acid, pH 4.5, and incubated at 37 °C. Eluates were immediately stored at −80 °C for subsequent MS analysis.
Prior to digestion, samples were lysed using 1× S-Trap lysis buffer (5% SDS, 50 mM TEAB, pH adjusted to 7.55 using 12% phosphoric acid) and bead-beating to shear the DNA. The samples were reduced with TCEP (20 mM final) for 30 min at 65 °C and alkylated with 80 mM iodoacetamide (IAA) in the dark for 30 min at room temperature. Samples were acidified using 12% phosphoric acid (final concentration 1.2%) and diluted using a 6x volume of S-Trap binding buffer (methanol containing 100 mM TEAB, pH adjusted to 7.2 using 12% phosphoric acid). Diluted samples were loaded onto an S-Trap micro column. Finally, SDS was completely removed from the sample by washing the filter 3 times with 160 μL of S-Trap binding buffer. Pure protein samples on the filter were then digested using 2 μg of trypsin/Lys-C (#V5073, Promega; 1:20 weight-to-weight ratio) in 50 mM ammonium bicarbonate, 0.5 mM CaCl2 at 37 °C overnight. Digested peptides were collected by washing the filter in three steps with 40 μL of 50 mM ammonium bicarbonate, pH 8.5, 40 μL of 0.15% formic acid, and 40 μL of 0.15% formic acid in 60% acetonitrile. Eluate was dried in vacuo and stored at −80 °C prior to analysis. The analysis of the samples was performed using an Exploris 480 mass spectrometer, in conjunction with an UltiMate 3000 liquid chromatography system (Thermo Scientific). A MonoCap column from GL Sciences, measuring 50 cm in length and 0.75 mm in inner diameter (Cat. No. 5020-10006), was employed. The flow rate was maintained at 500 nL/min, and the temperature was held constant at 25 °C.
During the analysis, mobile phase A (0.15% formic acid in water) and mobile phase B (0.15% formic acid in 100% acetonitrile) were utilized. A gradient spanning 130 min was employed: 5% B for 5 min, followed by a transition from 5% to 17% B over 100 min, a transition from 17% to 30% B over 19 min, and a rapid transition from 30% to 95% B in 1 min. The composition was maintained at 95% B for 5 min. A full-scan MS spectrum ranging from 350 to 1650 m/z was collected at a resolution of 60,000 at m/z 200. The maximum injection time was set to 50 ms, with an AGC target value of 3e6. The cycle time for data acquisition was set to 3 s, and the intensity threshold was set at 5e4. For MS/MS scans, a resolution of 15,000 was employed, with a maximum acquisition time of 40 ms and an AGC target value of 4e4. The isolation window at the Orbitrap cell was set to 1.6 m/z, and the first mass was set to 110 m/z. The collision energy for HCD was set to 32. A dynamic exclusion duration of 10 s was implemented. Charge states 1–8 were included, while unassigned charge states were ignored. The heated capillary temperature was set to 300 °C. Raw MS data were processed and searched with Proteome Discoverer (version 3.0.0.757; Thermo Fisher Scientific)72 using the Sequest HT search engine. The precursor mass tolerance of 10 ppm and fragment mass tolerance of 0.02 Da were used. The human database was downloaded from Uniprot. A 1% FDR cutoff was applied to filter the data, which was estimated by Percolator. Trypsin (full) was set as the enzyme in the search. The maximum mass cleavages were set to 2, and the peptide lengths were set from 6 to 144. Carbamidomethyl (+57.021 Da on C) was selected as a fixed modification, whereas oxidation (+15.995 Da on M), protein N-terminal Met-loss (−131.040 Da), Acetyl (+42.011 Da on N-terminus), and Protein N-terminal Met-loss+Acetyl (−89.030 Da) were dynamic modifications.
To assess enrichment of species that were present in the experimental but not control IPs, imputation values were calculated based on normal distribution using Perseus 2.0.1173. Additional analysis and visualization were performed with Microsoft Excel and R Studio. To identify co-IP hits, we looked for proteins with differential abundance values in the MVP IP versus control IP samples. Protein species with an average differential abundance value of log2(Fold Change) > 1.5, averaged over three replicates, were considered enriched. The significance of these enriched proteins was evaluated via a 2-tailed, homoscedastic t-test. Enriched species with −log10(p-value) > 1.3 (indicating a p-value of <0.05) were designated “hits”.
PARP activity assays
Changes in NAD+ concentration in the presence of FL MVP-PARP4, FL MVP, and the PARP1 catalytic module were measured using the NAD/NADH-Glo assay (Promega, Cat. No. G9071). Serial dilutions of proteins were incubated with 20 μM NAD+ in a white, 384-well plate for 3 h. Reactions were diluted with 25 μl of 2× luciferase detection reagent (prepared according to the “extended range” formulation recommended for PARP assays) for 20 min. Luminescence (corresponding to unconsumed NAD+ in the reaction) of triplicate PARP reactions was read on the FlexStation3 using the default Spectra iD5 luminescence endpoint detection settings. Average luminescence values for each concentration were plotted as relative light units (RLU).
Circular dichroism (CD) spectroscopy
CD measurements for each construct were recorded using a Jasco J-715 spectropolarimeter. MVP3-5 and MINT constructs were concentrated to 6.8 and 10 μM, respectively, and 180 μl were loaded into a quartz cuvette with a 1-mm optical path. Scans to identify wavelengths associated with ellipticity minima or maxima were performed at 4 °C between either 200- or 205-nm and 250-nm, using a step of 1-nm. Temperature scans were recorded from 4 to 90 °C in 1 °C intervals at a constant wavelength of 229-nm for MVP3-5 and 222-nm for MINT constructs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM maps of the human vault cage on its own, bound to NAD+, bound to PARP4, bound to PARP4 and NAD+, and bound to ADP have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-44953 (apo-MVP map), 44954 (MVP-NAD+ map), 44955 (MVP-PARP4 map), 44957 (MVP-PARP4-NAD+ map), and 44959 (MVP-ADP map), respectively. Atomic coordinates for the above complexes have been deposited in the Protein Data Bank with accession codes 9BW5 (apo-MVP structure), 9MXH (MVP-NAD+ structure), 9BW6 (MVP-PARP4 structure), 9BW7 (MVP-PARP4-NAD+ structure), and 9MXV (MVP-ADP structure), respectively. The raw protein mass spectrometry data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD052802. Source data are supplied in accompanying source data files. Source data are provided with this paper.
References
Daly, T. K., Sutherland-Smith, A. J. & Penny, D. In silico resurrection of the major vault protein suggests it is ancestral in modern eukaryotes. Genome Biol. Evol. 5, 1567–1583 (2013).
Tanaka, H. et al. The structure of rat liver vault at 3.5 angstrom resolution. Science 323, 384–388 (2009).
Kickhoefer, V. A. et al. The 193-Kd vault protein, VPARP, is a novel poly(ADP-Ribose) polymerase. J. Cell Biol. 146, 917–928 (1999).
Kickhoefer, V. A. et al. The telomerase/vault-associated protein Tep1 Is required for vault RNA stability and its association with the vault particle. J. Cell Biol. 152, 157–164 (2001).
Kowalski, M. P. et al. Host resistance to lung infection mediated by major vault protein in epithelial cells. Science 317, 130–132 (2007).
Peng, N. et al. Inducible major vault protein plays a pivotal role in double-stranded RNA– or virus-induced proinflammatory response. J. Immunol. 196, 2753–2766 (2016).
Lötsch, D. et al. Major vault protein supports glioblastoma survival and migration by upregulating the EGFR/PI3K signalling axis. Oncotarget 4, 1904–1918 (2013).
Scheffer, G. L. et al. The drug resistance-related protein LRP is the human major vault protein. Nat. Med. 1, 578–582 (1995).
Zhang, W. et al. Feedback regulation on PTEN/AKT pathway by the ER stress kinase PERK mediated by interaction with the Vault complex. Cell. Signal 27, 436–442 (2015).
Ben, J. et al. Major vault protein suppresses obesity and atherosclerosis through inhibiting IKK–NF-κB signaling mediated inflammation. Nat. Commun. 10, 1801 (2019).
Liu, S. et al. Human hepatitis B virus surface and e antigens inhibit major vault protein signaling in interferon induction pathways. J. Hepatol. 62, 10151023 (2015).
Teng, Y. et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 8, 14448 (2017).
Wakatsuki, S. et al. Small noncoding vault RNA modulates synapse formation by amplifying MAPK signaling. J. Cell Biol. 220, e201911078 (2021).
Vyas, S. et al. Family-wide analysis of poly(ADP-ribose) polymerase activity. Nat. Commun. 5, 4426 (2014).
Cohen, M. S. & Chang, P. Insights into the biogenesis, function, and regulation of ADP-ribosylation. Nat. Chem. Biol. 14, 236–243 (2018).
Kickhoefer, V. A. et al. Engineering of vault nanocapsules with enzymatic and fluorescent properties. Proc. Natl. Acad. Sci. USA 102, 4348–4352 (2005).
Yu, K. et al. Modulation of the Vault Protein-Protein Interaction for Tuning of Molecular Release. Sci. Rep. 7, 14816 (2017).
Guerra, P. et al. Symmetry disruption commits vault particles to disassembly. Sci. Adv. 8, eabj7795 (2022).
Ding, K. et al. Solution structures of engineered vault particles. Structure 26, 619–626.e3 (2018).
Kickhoefer, V. A. et al. Targeting vault nanoparticles to specific cell surface receptors. ACS Nano 3, 27–36 (2009).
Poderycki, M. J. et al. The vault exterior shell is a dynamic structure that allows incorporation of vault-associated proteins into its interior. Biochemistry 45, 12184–12193 (2006).
Woodward, C. L., Mendonça, L. M. & Jensen, G. J. Direct visualization of vaults within intact cells by electron cryo-tomography. Cell. Mol. Life Sci. 72, 3401–3409 (2015).
Kedersha, N. L. et al. Vaults. III. Vault ribonucleoprotein particles open into flower-like structures with octagonal symmetry. J. Cell Biol. 112, 225–235 (1991).
Casanas, A. et al. New features of vault architecture and dynamics revealed by novel refinement using the deformable elastic network approach. Acta Crystallogr. D Biol. Crystallogr. 69, 1054–1061 (2013).
Daumke, O. & Lewin, G. R. SPFH protein cage — one ring to rule them all. Cell Res. 32, 117–118 (2022).
Kataoka, K. et al. A cryptic phosphate-binding pocket on the SPFH domain of human stomatin that regulates a novel fibril-like self-assembly. Curr. Res. Struct. Biol. 4, 158–166 (2022).
Hopp, A.-K. et al. Mitochondrial NAD+ controls nuclear ARTD1-induced ADP-ribosylation. Mol. Cell 81, 340–354.e5 (2021).
Rayo, J. et al. Immunoediting role for major vault protein in apoptotic signaling induced by bacterial N-acyl homoserine lactones. Proc. Natl. Acad. Sci. USA 118, e2012529118 (2021).
Yi, C. et al. Major vault protein, in concert with constitutively photomorphogenic 1, negatively regulates c-Jun–mediated activator protein 1 transcription in mammalian cells. Cancer Res. 65, 5835–5840 (2005).
Lanisnik Rizner, T. Enzymes of the AKR1B and AKR1C subfamilies and uterine diseases. Front. Pharmacol. 3, 34 (2012).
Jiang, D. et al. Analysis of p53 transactivation domain mutants reveals Acad11 as a metabolic target important for p53 pro-survival function. Cell Rep. 10, 1096–1109 (2015).
Wang, H. et al. The transcriptional coactivator RUVBL2 regulates Pol II clustering with diverse transcription factors. Nat. Commun. 13, 5703 (2022).
Li, R. Z. et al. hnRNPDL extensively regulates transcription and alternative splicing. Gene 687, 125–134 (2019).
Gu, X. et al. Aspartyl-tRNA synthetase 2 orchestrates iron-sulfur metabolism in hematopoietic stem cells via fine-tuning alternative RNA splicing. Cell Rep. 42, 113264 (2023).
Guang, S. et al. Mutations in DARS2 result in global dysregulation of mRNA metabolism and splicing. Sci. Rep. 13, 13042 (2023).
Chen, K. Y. et al. Identification of trophinin as an enhancer for cell invasion and a prognostic factor for early stage lung cancer. Eur. J. Cancer 43, 782–790 (2007).
Adams, D. R., Ron, D. & Kiely, P. A. RACK1, a multifaceted scaffolding protein: structure and function. Cell Commun. Signal. 9, 22 (2011).
Lee, J. et al. Coordination of canonical and noncanonical Hedgehog signalling pathways mediated by WDR11 during primordial germ cell development. Sci. Rep. 13, 12309 (2023).
Mariotti, S. et al. MICAL2 is a novel human cancer gene controlling mesenchymal to epithelial transition involved in cancer growth and invasion. Oncotarget 7, 1808–1825 (2016).
Gkika, D. et al. TRP channel-associated factors are a novel protein family that regulates TRPM8 trafficking and activity. J. Cell Biol. 208, 89–107 (2015).
Luongo, T. S. et al. SLC25A51 is a mammalian mitochondrial NAD(+) transporter. Nature 588, 174–179 (2020).
Tian, C. et al. KRAB-type zinc-finger protein Apak specifically regulates p53-dependent apoptosis. Nat. Cell Biol. 11, 580–591 (2009).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Smialek, M. J. et al. Role of PUM RNA-binding proteins in cancer. Cancers https://doi.org/10.3390/cancers13010129 (2021).
Wang, Q. et al. RNA binding protein DAZAP1 promotes HCC progression and regulates ferroptosis by interacting with SLC7A11 mRNA. Exp. Cell Res. 399, 112453 (2021).
Choudhury, R. et al. The splicing activator DAZAP1 integrates splicing control into MEK/Erk-regulated cell proliferation and migration. Nat. Commun. 5, 3078 (2014).
Gong, Y. et al. PUMILIO proteins promote colorectal cancer growth via suppressing p21. Nat. Commun. 13, 1627 (2022).
Zhang, R. et al. Lemur tyrosine kinase 2 acts as a positive regulator of NF-κB activation and colon cancer cell proliferation. Cancer Lett. 454, 70–77 (2019).
Huang, Y.-K. et al. The role of ALDH1A1 in glioblastoma proliferation and invasion. Chem. Biol. Interact. 402, 111202 (2024).
Piccoli, G. D., Torres-Rosell, J. & Aragón, L. The unnamed complex: what do we know about Smc5-Smc6?. Chromosome Res. 17, 251–263 (2009).
Peng, X. P. & Zhao, X. The multi-functional Smc5/6 complex in genome protection and disease. Nat. Struct. Mol. Biol. 30, 724–734 (2023).
Ivic, N. et al. Fuzzy interactions form and shape the histone transport complex. Mol. Cell 73, 1191–1203.e6 (2019).
Ding, J.-H. et al. Guanosine diphosphate–mannose suppresses homologous recombination repair and potentiates antitumor immunity in triple-negative breast cancer. Sci. Transl. Med. 16, eadg7740 (2024).
Liu, D. et al. Transcriptomic and metabolomic profiling in helicobacter pylori–induced gastric cancer identified prognosis- and immunotherapy-relevant gene signatures. Front. Cell Dev. Biol. 9, 769409 (2021).
Duarte-Pereira, S. et al. Study of NAD-interacting proteins highlights the extent of NAD regulatory roles in the cell and its potential as a therapeutic target. J. Integr. Bioinform. 20, 20220049 (2023).
Kumar, V. et al. Transcription corepressor CtBP is an NAD(+)-regulated dehydrogenase. Mol. Cell 10, 857–869 (2002).
Park, Y. W., Jang, Y. Y., Joo, H. K. & Lee, J. Y. Structural Analysis of Redox-sensing Transcriptional Repressor Rex from Thermotoga maritima. Sci. Rep. 8, 13244 (2018).
Li, J. et al. A conserved NAD+ binding pocket that regulates protein-protein interactions during aging. Science 355, 1312–1317 (2017).
Innocenti, B., Pozzan, T. & Fasolato, C. Intracellular ADP modulates the Ca release-activated Ca current in a temperature- and Ca-dependent way. J. Biol. Chem. 271, 8582–8587 (1996).
Pinon, P. & Wehrle-Haller, B. Integrins: versatile receptors controlling melanocyte adhesion, migration and proliferation. Pigment Cell Melanoma Res. 24, 282–294 (2011).
Gagné, J.-P. et al. Quantitative proteomics profiling of the poly(ADP-ribose)-related response to genotoxic stress. Nucleic Acids Res. 40, 7788–7805 (2012).
Wright, R. H. et al. ADP-ribose-derived nuclear ATP synthesis by NUDIX5 is required for chromatin remodeling. Science 352, 1221–1225 (2016).
Dasovich, M. et al. Identifying poly(ADP-ribose)-binding proteins with photoaffinity-based proteomics. J. Am. Chem. Soc. 143, 3037–3042 (2021).
Ayyappan, V. et al. ADPriboDB 2.0: an updated database of ADP-ribosylated proteins. Nucleic Acids Res. 4, D261–d265 (2021).
Yu, H. et al. Major vault protein promotes hepatocellular carcinoma through targeting interferon regulatory factor 2 and decreasing p53 activity. Hepatology 72, 518–534 (2020).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 12141221 (2020).
Emsley, P. et al. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Miller, K. R. et al. T cell receptor-like recognition of tumor in vivo by synthetic antibody fragment. PLoS ONE 7, e43746 (2012).
Paduch, M. et al. Generating conformation-specific synthetic antibodies to trap proteins in selected functional states. Methods 60, 3–14 (2013).
Orsburn, B. C. Proteome discoverer-a community enhanced data processing suite for protein informatics. Proteomes 9, 15 (2021).
Rudolph, J. D. & Cox, J. A network module for the perseus software for computational proteomics facilitates proteome interaction graph analysis. J. Proteome Res. 18, 2052–2064 (2019).
Frigon, L. & Pascal, J. M. Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP. Nucleic Acids Res. 51, 12492–12507 (2023).
Acknowledgements
We thank Jotham Austin, Tera Lavoie, and James Fuller at the University of Chicago Advanced Electron Microscopy Core, as well as Rui Yan, Xiaowei Zhao, and Zhiheng Yu at the HHMI Janelia Research Campus, for their assistance with cryo-EM data collection. We also thank Virender Kumar, Varun Sharma, and Himanshi Yadav for maintenance of and assistance with the University of Chicago Research Computing Cluster resources and the Beagle3 GPU cluster, which is supported by the National Institute of Health (NIH) under the High-End Instrumentation (HEI) grant program award S10OD028655. Isothermal titration calorimetry data were collected with the help of Robel D. Demissie and Hyun Lee at the University of Chicago-Illinois Biophysics Core. ADP-ribosylation identification mass spectrometry data were collected and analyzed with the help of Lasanthi Jayathilaka at the University of Illinois-Chicago Mass Spectrometry Core, and proteomics data were collected and analyzed with the help of Allen Huff and Samuel Weng at the University of Chicago Proteomics Platform. We are grateful to the Sosnick lab and Michael Baxa at the University of Chicago for the use of their instrument and expertise to collect CD spectra. This work was supported by the NIH under grant number R35GM143052 to M.Z. and by a training grant T32GM144290 to J.E.L.
Author information
Authors and Affiliations
Contributions
J.E.L. and M.Z. designed the research. J.E.L. performed all laboratory experiments and prepared specimens for cryo-EM data collection. J.E.L. and M.Z. collected and processed cryo-EM data and built and refined atomic models. J.E.L. and M.Z. wrote the manuscript with input from the other authors. R.S. and S.A. assisted with revisions. S.M. and A.A.K. performed and directed synthetic antigen binding reagent selection and validation assays. Y.X. provided plasmids and reagents for protein expression and optimized the pull-down protocol for mass spectrometry, and K.R. assisted with protein purification.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lodwick, J.E., Shen, R., Erramilli, S. et al. Structural insights into the roles of PARP4 and NAD+ binding in the human vault cage. Nat Commun 16, 6724 (2025). https://doi.org/10.1038/s41467-025-61981-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61981-x