Abstract
Pre- and post-transcriptional mechanisms, including alternative promoters, termination signals, and splicing, play essential roles in diversifying protein output by generating distinct RNA and protein isoforms. Two major challenges in characterizing the cellular function of alternative isoforms are the lack of experimental methods to specifically and efficiently modulate isoform expression and computational tools for complex experimental design and analysis. To address these gaps, we develop and methodically test an isoform-specific knockdown strategy which pairs the RNA-targeting CRISPR/Cas13d system with guide RNAs that span exon-exon junctions. In parallel, we provide computational tools for experimental design and analysis. In this study, we demonstrate that junction-targeting achieves robust and isoform-specific RNA knockdown across diverse alternative isoform events, genes, and cell types.
Similar content being viewed by others
Introduction
The majority of human genes generate alternative RNA and protein isoforms through mechanisms such as alternative splicing (AS), alternative transcription initiation, and alternative polyadenylation1,2,3. In some cases, alternative isoforms regulate gene dosage through nonsense-mediated decay, translation efficiency, and stability4,5. In others, RNA isoforms are translated into distinct proteins that can vary in localization, protein-protein interactions, and nucleic acid binding6,7,8. Alternative transcriptional and splicing events are dynamic throughout development and differentiation, particularly in brain and muscle tissues1,2,9,10.
Genetic variants that affect AS can have significant consequences. Cis-acting variants within a single gene can be sufficient to drive disease, while trans-acting variants in RNA binding proteins can disrupt entire splicing networks11,12,13. Splicing quantitative trait locus analysis has demonstrated that a substantial proportion of complex disease heritability is mediated through genetic effects on AS14,15,16,17,18. Widespread splicing alterations are also observed across cancers, with ~30% more AS events in cancer cells relative to normal cells3,19,20.
Recent advances in long read RNA-sequencing have dramatically expanded the catalog of annotated transcripts. Over the past five years alone, GENCODE added ~55,000 transcripts, bringing the total to ~250,00021. Despite this, the functional roles of most isoforms remain unknown–highlighting the need for both experimental and computational methods to interrogate isoform-specific functions.
While gene-level perturbation is routine, isoform-level functional studies remain challenging. CRISPR-based DNA-targeting systems have been used to perturb alternative transcription and splicing events, including cassette exon deletion22,23, promoter inhibition24, and splice site editing25. RNA-based methods such as RNAi26,27 and antisense oligonucleotides28 primarily target cassette exons. Most recently, catalytically inactive RNA-targeting CRISPRs have been directed to splicing regulatory regions to alter pre-mRNA processing29,30,31,32. However, these strategies typically focus on a single event type, often cassette exons, and require significant computational work to confidently identify expressed isoforms and to link a transcriptional or splicing event with its corresponding isoform(s).
Here, to address these limitations, we introduce and systematically evaluate a versatile platform for isoform-specific knockdown that pairs the CRISPR/Cas13d RNA-targeting system33,34,35,36,37,38 with guide RNAs (gRNAs) that target mature RNA exon-exon junctions (EEJs). We show that EEJs are broadly targetable by Cas13, and that existing models, including our TIGER deep learning model36, accurately predict EEJ gRNA efficacy, suggesting similar rules govern targeting of EEJs and exons. The critical innovation of our strategy is its ability to achieve isoform-specific knockdown using gRNAs targeting unique EEJs across diverse transcriptional and splicing event types. We validate EEJ-specific knockdown by reverse-transcription quantitative PCR (RT-qPCR) across twelve EEJs in four genes and isoform-specific knockdown by long-read RNA-sequencing (LRS) of six RBFOX2 EEJs. For RBFOX2 isoforms with alternative first exons, we also assess protein-level knockdown and measure the downstream effects on splicing. Finally, we provide TIGER efficacy predictions for ~2.2 million EEJ-targeting gRNAs, the Isoviz R package for experimental design, and practical recommendations for isoform-level perturbation using our EEJ-targeting strategy.
Results
Cas13d essentiality screen shows broad applicability of targeting common EEJs for RNA knockdown
Our strategy is, to our knowledge, the first to propose explicitly targeting EEJs in mature RNA for isoform-specific knockdown. EEJs, as opposed to exons, can be selectively targeted across all types of alternative transcriptional and splicing events, expanding the number of potentially targetable isoforms in complex transcriptomes. To evaluate this potential, we categorized EEJs based on their isoform specificity: (1) common EEJs are present in all isoforms of a gene, (2) fully unique EEJs are exclusive to a single isoform, and (3) partially unique EEJs are shared amongst a subset of isoforms (Fig. 1a). We applied this classification to GENCODE v41 Basic Annotations21 and identified 53,890 protein-coding and 23,502 lncRNA EEJs that are uniquely targetable across 36,604 and 11,053 isoforms, respectively (Fully Unique in Fig. 1b). Given that isoform expression is often tissue-specific, some partially unique EEJs may become fully unique in specific contexts. By combining fully and partially unique categories, we estimate that our EEJ strategy has the potential to target upwards of 97% of protein-coding and lncRNA isoforms from multi-isoform genes.
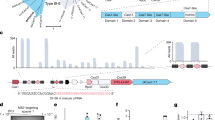
a Schematic of EEJ categories: common (gray), fully unique (spanning pink-purple exons), and partially unique (spanning pink-yellow exons). Eight 23 bp gRNAs are tiled per EEJ, starting from 15 to 8 bp upstream. b Counts of EEJs and isoforms in GENCODE v41 (Basic), categorized as in (a), across 18,568 protein-coding and 14,849 lncRNA genes. c Library design summary: 50,310 gRNAs targeting 6932 EEJs across 942 genes from Gold Standard40,41, DepMap Common42, and DepMap A37543 essential gene sets. Controls included 1972 non-essential and 963 non-targeting (NT) gRNAs. d Schematic of the essentiality screen: Cells expressing gRNAs that knockdown essential genes are depleted over the 21 day time course. e Cumulative density function (CDF) plot of Day 21/Day 0 log2 fold changes (LFCs) for gRNAs targeting essential genes in categories from (c). Dashed lines indicate 30.3% active gRNAs for Gold Standard genes (defined by LFC < 0.01 quantile of NT gRNA normal distribution). f Barplot of active gRNA counts per EEJ (max 8/8) for Gold Standard genes. EEJs with 0/8 active gRNAs (28%) are excluded. g Percentage of targetable EEJs (≥1 active gRNA) per Gold Standard gene. Genes with ≥6 EEJs are on the x-axis, ordered by RNAi-based essentiality. Source data are provided on GSE242107 and Zenodo. Panels a and d were created in BioRender. Schertzer, M. (2025) https://BioRender.com/81f7cfw.
EEJ accessibility, and thus targetability, has not been systematically evaluated for any RNA-targeting technology. Therefore, the first step in testing our strategy was to evaluate whether gRNAs spanning common EEJs could effectively knockdown RNA, independent of isoform specificity. We considered several factors that might limit junction accessibility–constraints that are less relevant for targeting exons. First, Cas13d complex-target formation could be sterically hindered by the exon-junction complex39 or by splicing factor binding. Additionally, EEJs have a shorter nuclear lifespan than exons, as they are only accessible to nuclear Cas13d in the brief window after splicing but before export. While a previous Cas13d essentiality screen in HEK293FT cells provided some supporting evidence that gRNAs targeting EEJs could knockdown RNA, the number of EEJ gRNAs tested was limited (n = 1559) across just 16 essential genes (Supplementary Fig. 1A)36.
To assess whether Cas13d-gRNA complexes can efficiently access EEJs at scale, we conducted an essentiality screen exclusively targeting common EEJs in RNA. This screen–designed without gRNA efficacy predictions–tested 50,310 gRNAs across 6,932 EEJs in 942 genes. Previous Cas13d essentiality screens targeted far fewer genes (e.g., 16 or 55) and primarily focused on exons36,38. Genes are categorized here into three overlapping essentiality lists: (1) Gold Standard40,41, (2) DepMap Common42, and (3) DepMap A37543 (Fig. 1c). As this screen aimed to evaluate EEJ accessibility rather than isoform specificity, we prioritized common EEJs expressed in A375 cells to achieve total gene knockdown44 (‘Guide library design‘ in Methods). Controls included 1972 non-essential (NE) gene gRNAs and 963 non-targeting (NT) gRNAs (Fig. 1c). Because Cas13d does not require a protospacer adjacent motif (PAM), we tiled eight 23 bp gRNAs per EEJ from −15bp to +15 bp, expecting a mix of active and inactive gRNAs. The gRNA pool was transduced in two biological replicates into monoclonal Cas13d-expressing A375 cells, and gRNA depletion was quantified by log2 fold change (LFC) at Days 7, 14, and 21 post-induction. We used gRNA depletion as a proxy for RNA knockdown, i.e., a gRNA with a significant negative LFC corresponds to an active gRNA that effectively knocks down its essential gene target (Fig. 1d).
To evaluate screen quality, we observed concordance between biological replicates at Day 21 (R = 0.69 for gRNAs targeting Gold Standard essential genes; Supplementary Fig. 1B), and a positive correlation with shared gRNAs from a prior HEK293FT screen (R = 0.67, Supplementary Fig. 1c)36. We also examined potential Cas13d collateral activity45,46,47 by analyzing NE gene control gRNAs targeting 409 genes ranging from 10 to 492 transcripts per million (TPM). As expected, the knockdown of these non-essential genes by Cas13d did not lead to cell death and thus, gRNA depletion in our screen (Supplementary Fig. 1D).
We next evaluated gRNA efficiency and EEJ accessibility across essential genes. We anticipated that gRNA depletion would vary widely across EEJs and genes, driven primarily by two factors: gRNA efficiency and the degree of gene essentiality. As a first step, we classified gRNAs as active or inactive based on a −1.96 LFC cutoff (corresponding to the 0.01 quantile of the NT control gRNAs’ normal distribution). Among Gold Standard essential genes, a highly curated set that are almost universally essential40,41, 30.3% of EEJ gRNAs were active (Fig. 1e, Supplementary Fig. 1E), consistent with prior estimates (31.5% and 32.3% in exons and EEJs36; Supplementary Fig. 1A). This high-confidence Gold Standard gene set was used for subsequent analyses. For 840 common EEJs with all 8 tiling gRNAs (116 Gold Standard genes), 72% were targetable (≥1 active gRNA; Fig. 1f). EEJ targetability varied across genes–for example, all EEJs in CCT8, EIF3B, and PSMD1 were targetable while <50% were targetable for EXOSC10, DDX49, and FTSJ3 (Fig. 1g). This variation tracked with RNAi-based estimates of gene essentiality, with less essential genes showing fewer active gRNAs (Fig. 1g). This suggests that gRNA effectiveness may be underestimated in the screen. Overall, these results demonstrate that EEJs are broadly accessible to the RNA-guided Cas13d nuclease and targetable for RNA knockdown.
Deep learning uncovers similar rules for gRNAs targeting EEJs and exons
Our screen results (Fig. 1) indicated that 30.3% of randomly selected EEJ gRNAs are effective. The TIGER deep learning model can predict gRNA efficiency and enrich for active gRNAs beyond random selection36, but it and similar models were trained primarily on exon-targeting gRNAs35,36,37,38. Compared to exons, EEJs are bound by distinct RNA-binding proteins and may show differential accessibility to nuclear-localized Cas13/gRNA complexes due to splicing efficiency, transcript position, and time to export. These factors may influence gRNA efficacy and design. It remains unclear whether a model trained on EEJ-targeting data would learn additional, or different, features. As our screen includes more EEJ-targeting gRNAs than any previous Cas13 screen, we are well-positioned to investigate this question.
To train TIGER model adaptations, we first developed SEABASS, a Bayesian linear mixed model that integrates LFCs across timepoints and replicates (Methods). SEABASS outputs two values per gRNA: a slope of LFC over time and a standard error to quantify uncertainty (Fig. 2a). Unlike simple averages, SEABASS captures variance of de-noised data for model training. A more negative slope indicates a more active gRNA, while slopes near 0 indicate inactive gRNAs (Fig. 2a).
a Schematic of the SEABASS linear mixed model. Input: LFCs for all timepoints and replicates for EEJ gRNAs. Output: slope, where a more negative slope corresponds to a more active gRNA, and standard error. b Summary of TIGER model adaptations. ‘Cell non-specific set’ is defined below the table. MFE, Minimum free energy. c Cross-validated area under the precision-recall curve (AUPRC) values for each model across gene essentiality thresholds. Random gRNA selection baseline shown for comparison. Dotted line marks the 25% essentiality threshold used in (d). d Pearson correlations between predicted and observed gRNA activity at the 25% essentiality cutoff. Observed values are either LFCs or SEABASS slopes. *p < 0.0001 (two-sided). e SHapley Additive exPlanations (SHAP) plots comparing positional nucleotide contributions for TIGER (solid line) and TIGERjunction (dashed line). Colors denote nucleotide identity (e.g. A is blue). SHAP values indicate predicted impact on LFC when that position has the specific nucleotide. f SHAP value profiles comparing TIGERsite (solid) and TIGERjunction (dashed lines) across EEJs. TIGERsite predicts the mean LFC of all eight gRNAs tiled across an EEJ using a 101 bp window (−50/+50). TIGERjunction predicts the LFC of individual gRNAs using only the 23 bp target sequence. SHAP values from TIGERjunction were tiled across the EEJ at the same eight positions used in the screen (tiles; dotted lines), and then positionally averaged (tile mean; dashed lines). This average closely matches the TIGERsite SHAP profile within the tiling window (gray bars). X-axis indicates gRNA target position centered at the splice site. Source data are provided as a Source Data file.
We retrained several TIGER model adaptations (Fig. 2b): (1) the original TIGER model trained on exon LFCs, (2) TIGERjunction trained on EEJ LFCs, (3) TIGERbass trained on EEJ SEABASS slopes, and (4) TIGERsite trained on EEJ LFCs with additional sequence context. All models except TIGERsite included cell non-specific features–such as RNA structure–that were previously shown to modestly boost TIGER performance36.
To evaluate how well TIGER, TIGERjunction, and TIGERbass classify active versus inactive EEJ-targeting gRNAs for essential genes, we generated cross-validated area under precision-recall curve (AUPRC) values using observed screen data as labels. Because gene essentiality varies, we assessed performance across increasingly stringent cutoffs (5–45% active gRNAs per gene), including random gRNA selection as a baseline (Fig. 2c). TIGERbass performs best at a lower cutoff, when the data is noisier, but all three models converge to an AUPRC of 0.88 at the most stringent cutoff. Notably, all TIGER-based models outperform random selection by ~twofold (Fig. 2c). At a moderate cutoff of 25%, where we have more data, we observed similar trends using Pearson correlation (Fig. 2d). TIGERbass not only outperforms the other models when tested against its predictive modality of SEABASS slopes but also modestly outperforms TIGER and TIGERjunction at their predictive modality of LFC, confirming the benefit of SEABASS’s de-noising (R = 0.63 for TIGERbass versus R = 0.61 and 0.55 for TIGERjunction and TIGER, respectively) (Fig. 2d).
Following previous work36,38, we used SHapley Additive exPlanations (SHAP)48 to assess features used by the model to predict gRNA efficacy. First, we focused on nucleotide preferences for active gRNAs by comparing SHAP values across the 23 bp target RNA sequence for TIGER, which is trained on exon LFC data alone versus TIGERjunction, which is trained on EEJ LFC data alone (dashed line; Fig. 2e). Pearson correlations of positional nucleotides and observed LFC data produced similar plots. (Supplementary Fig. 2A), indicating that sequence determinants of gRNA efficiency are indistinguishable for targeting EEJs and exons.
Next, we asked whether there was broader sequence context surrounding EEJs that influence gRNA efficiency, for example as a result of secondary structure or sequence-specific splicing factor binding. We compared two adaptations of TIGER: 1) TIGERjunction that is trained using 23 bp of target RNA sequence and 2) TIGERsite that considers −50/+50 bp around each EEJ site in the mature RNA (Fig. 2b). We trained TIGERjunction to predict a gRNA’s LFC and trained TIGERsite to predict the mean of the eight gRNAs’ LFCs that target each EEJ. When we tiled TIGERjunction SHAP values across the EEJ as we tiled gRNAs across the EEJ in our screen (TIGERjunction tiles) and positionally averaged TIGERjunction SHAP values (TIGERjunction tile mean), we recovered almost exactly the TIGERsite SHAP profile within the tiling window (between the gray dotted lines; Fig. 2f). This suggests that TIGERsite simply learns to average the sequence preferences of the eight possible gRNA alignments despite having the opportunity to learn additional novel EEJ sequence preferences. We conclude from this analysis that the predictive signal is almost entirely derived from the local sequence of the gRNA/target, with minimal if any contribution from broader sequence context–in line with findings from Wessels et al.36.
TIGERbass input included 23 bp of target sequence and a set of cell non-specific features relating to RNA structure (Fig. 2b). From this, the model achieved a Pearson correlation of 0.59 between predicted and observed slopes (5% gene essentiality cutoff; Supplementary Fig. S2B). To explain discrepancies between the predicted and observed results, we calculated residuals for each gRNA and investigated their association with a set of non-sequence features, not included in TIGERbass training (Supplementary Figs. S2C). We considered RNA half-life estimates, gene expression, and nuclear RNA localization in A375 (Cas13d is localized to the nucleus), gene length, intron length, relative EEJ position in a gene, and gRNA tiling position. We fit a linear regression to evaluate feature associations with residuals, but found negligible relationships across all non-sequence features tested (Supplementary Fig. 2D, E). We conclude that gRNA sequence, and thus its target RNA sequence, are the primary features predictive of Cas13d gRNA EEJ targeting efficiency. We hypothesize that this will hold true for future RNA-targeting CRISPR systems. Practically, this means that one can use existing prediction tools for EEJ gRNA design such as TIGER or other models35,38.
Isoviz R package automates experimental design for Cas13d-based isoform-centric studies
The next step is to link EEJ targetability with isoform specificity, but mammalian transcriptomes are complex, making both experimental design and interpretation challenging. First, there is uncertainty regarding which isoforms are expressed and at what levels in a given cell type. Furthermore, some events, especially alternative 5’ and 3’ splice sites (SS), are difficult to visualize from a typical transcript diagram but could have substantial consequences, especially if they introduce a frameshift. Finally, selecting efficient gRNAs that span specific EEJs can be time-consuming. The individual resources and tools currently available to address these challenges are disjointed and require extensive manual, error-prone labor.
To aid in designing isoform-centric experiments, especially CRISPR-based ones, we developed Isoviz, an R package that integrates transcript-level annotations from GENCODE or long read analyses, spliced read counts for each EEJ, and TIGER gRNA predictions (Fig. 3a). The result of this integration shows a clear link between each EEJ, its expression within a cell type, and its corresponding isoform(s). Based on this, a user can quickly design gRNAs to EEJs that map uniquely or partially uniquely to knockdown an isoform in their cell type of interest.
a Overview of Isoviz inputs and functions. Required inputs (red boxes) vary by function; default input files are available on GitHub but can be replaced by user-defined data. Pre-processing steps (yellow) reformat input for downstream functions. Main visualization and output functions (purple) operate per user-specified gene. b Isoviz output for COP1 and SPAG9, showing isoform structures and two selected EEJs per gene plotted as introns. EEJs are labeled with read counts and corresponding isoforms. Users can select all or a subset of EEJs for plotting. c Isoviz output table showing gRNAs targeting EEJs in (b) and their TIGER scores. d RT-qPCR quantification of COP1 and SPAG9 RNA knockdown using EEJ-targeting gRNAs from (c) at 24 hours of doxycycline. Knockdown is relative to cells transfected with an empty gRNA cassette (n = 4 gRNAs per EEJ). Each boxplot represents three RT-qPCR measurements from two biological replicates. Box plots were made using ggplot and show the median (center line), the 25th and 75th percentiles (box bounds), and the range of values within 1.5× the interquartile range (whiskers). Outliers beyond this range are shown as individual points. Source data are provided as a Source Data file.
We illustrate the utility of Isoviz in the design of EEJ-based RNA knockdown experiments for two genes with relatively simple transcript structures, COP1 and SPAG9. Isoviz allows for the visualization of all EEJs and their corresponding isoforms, followed by a table with the top TIGER scoring gRNAs for each EEJ, with the option to filter for a subset of EEJs (Fig. 3b, c). Figure 3b identifies that the COP1⋅20769 EEJ is specific to an exon inclusion event in the COP1-204 isoform, and the SPAG9⋅117144 EEJ belongs to an exon exclusion event that is partially unique and maps to all isoforms except SPAG9-201 (EEJ IDs are assigned in Isoviz for ease of comparison). To select gRNAs that target these EEJs, Fig. 3c shows four of the eight possible gRNA options per EEJ and their TIGER score. Both visualization and table outputs save the user significant time and reduce the possibility of error.
RT-qPCR confirms EEJ-specific RNA knockdown using EEJ gRNAs
To directly measure the EEJ-specificity and magnitude of RNA knockdown for COP1 and SPAG9 gRNAs designed with Isoviz (Fig. 3c), we created a non-viral, piggyBac-Cas13d vector system. In brief, we cloned a doxycycline (dox)-inducible Cas13d (NLS-RfxCas13d-NLS) and a gRNA cassette (hU6-DR-BsmBI)37 into two separate piggyBac transposon backbones49, creating PB-Cas13 and PB-rtTA-gRNA, respectively50,51,52. We generated stable HEK293 cell lines for the 16 COP1 and SPAG9 gRNAs (Fig. 3c) and measured RNA knockdown, with junction-specific primers, via RT-qPCR after 24 hrs of dox induction of Cas13d. All COP1 and SPAG9 gRNAs produced robust and consistent knockdown across two biological replicates, ranging from 39 to 83% reduction in RNA expression relative to control (Fig. 3d, Supplementary 3A). We further show that RNA knockdown for all 8 COP1 gRNAs was similar in hESCs and across different timepoints of Cas13 dox-induction (Supplementary 3B). Importantly, robust knockdown (>80%) can be observed as early as 12 h post dox-induction and is maintained at similar levels through the longest tested timepoint of 72hrs. In line with a previous study53, we found that RT-qPCR primers flanking or overlapping the gRNA cutsite showed the most RNA knockdown, and we designed all RT-qPCR primers for future experiments accordingly (Supplementary Fig. 3C).
To provide further evidence of the specificity, efficiency, and versatility of our EEJ strategy, we used Isoviz to visualize annotated transcript structure and design gRNAs for diverse splicing and transcriptional event types in the MKNK2 and RBFOX2 genes in hESCs and HEK293 cell lines (Supplementary Fig. 4A, B, respectively). In MKNK2, we targeted two unique EEJs of alternative last exons in HEK293 cells54. By RT-qPCR, we measured RNA knockdown of 78% at the 128207 EEJ, which maps to MKNK2-202 and MKNK2-211, and 87% at the 128208 EEJ of MKNK2-201 (Supplementary Fig. 4C). Importantly, the alternate MKNK2 EEJ in each experiment was minimally affected.
In RBFOX2, the transcript structure is more complex, where each annotated transcript contains combinations of 4 different event types. This includes four alternative TSS (AltTSS), two separate alternative 3’ SS events (Alt3SS.1 and Alt3SS.2), and a cassette exon (CassetteEx; Supplementary Fig. 4b). Importantly, the subtle base pair differences at Alt3SS.1 (3 bp) and Alt3SS.2 (12 bp) EEJs cannot be distinguished by eye, exemplifying how Isoviz visualization is necessary for accurately identifying small splicing changes and mapping them to their corresponding isoform(s). We did not test EEJ gRNAs at the cassette exon event due to low expression of the exclusion product present in RBFOX2-205 and RBFOX2-207. For all other EEJs represented in Supplementary Fig. 4B, we again use RT-qPCR to measure robust and EEJ-specific RNA knockdown in both hESC and HEK293 cell lines (Supplementary Fig. 4D, E). Together, the COP1, SPAG9, MKNK2 and RBFOX2 experiments emphasize the versatility of our EEJ strategy to target a broad range of alternative transcriptional and splicing events.
PacBio long read RNA-sequencing confirms RBFOX2 isoform-specific knockdown using EEJ gRNAs
We have shown that EEJ gRNAs achieve consistent and robust RNA knockdown across diverse event types, cell types, replicates, and timepoints. A crucial next step is to determine how EEJ-centric RNA knockdown quantification relates to desired isoform-specific knockdown. To accurately identify which isoforms are expressed in hESCs and quantify isoform-level differences upon knockdown, we performed PacBio long read RNA-sequencing (LRS55) on hESC Empty (Control) and the six RBFOX2 EEJ RNA samples from Supplementary Fig. 4D. We chose RBFOX2 here because of its high isoform complexity, where each EEJ gRNA is expected to target a subset of multiple isoforms. In fact, LRS analysis identified 12 RBFOX2 isoforms expressed in the unperturbed hESC control, six of which have not been previously annotated in GENCODE (Fig. 4a, top panel; isoforms >0.6 TPM or 10 full length reads in the control). The bottom panel of Fig. 4a shows Isoviz integration of RBFOX2 EEJ-level information with LRS transcript-level structure for both annotated and novel isoforms.
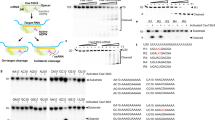
a Isoviz visualization of annotated and novel RBFOX2 isoforms detected by PacBio LRS in hESCs (≥10 full-length reads in control). TPM values are listed in parentheses next to each isoform. The lower panel shows six selected EEJs plotted as introns from three events–an alternative TSS (AltTSS), and two alternative 3’ splice sites (Alt3SS.1: 3 bp, Alt3SS.2: 12 bp). EEJs are labeled with short-read coverage, relative abundance (%), and corresponding isoforms. RBFOX2⋅178147 EEJ defines N-short isoforms; RBFOX2⋅178149 defines N-long. LRS was performed as a single replicate. b Proportion of isoforms containing RBFOX2⋅178147 or RBFOX2⋅178149 EEJs in LRS data. Each box represents one isoform, colored as in (a), AltTSS. c Comparison of RBFOX2 isoform TPMs between control and AltTSS knockdowns in LRS data. Each point is an isoform colored by the whether it contains the EEJ being targeted with that knockdown. RBFOX2-208 is the most abundant RNA isoform detected with LRS and is marked for reference. d Diagram of RBFOX2 protein isoforms highlights N-terminal amino acid (AA) differences (N-short MEK: 8 AA; N-long MAE: 78 AA). Western blot shows total RBFOX2 protein expression in control and knockdowns; quantification normalized to GAPDH. Band sizes are consistent with antibody documentation (expected bands: ~40kDa N-short and ~47 kDa for N-long). e Fragment Ion Intensities for two shared RBFOX2 peptides measured by targeted mass spectrometry, serving as a complementary method to measure total RBFOX2 knockdown. n = 3 injection replicates. Box plots were made in ggplot and show the median (center line), the 25th and 75th percentiles (box bounds), and the range of values within 1.5× the interquartile range (whiskers). f Splicing changes in skipped exon events at 96 hr timepoint. Each point represents an event; deviation from the diagonal indicates differential splicing relative to control. Total knockdown gRNA (targets EEJ present in all RBFOX2 isoforms) used as a positive control. g Validation of two skipped exon events from (f): GOLIM4 and CLSTN1. Percent spliced in (PSI) is calculated across timepoints from RNA-seq data (n = 1 for each timepoint) and for 72 h timepoint from RT-PCR gels (top band = inclusion, bottom = skipping). PSI calculated from band intensities (ImageJ). Source data are provided as a Source Data file and in Supplementary Data 2, 3, and GSE242107.
To quantify isoforms in the LRS data, we first examined isoform expression per EEJ for the AltTSS event in the unperturbed hESC Control. Seven isoforms containing the RBFOX2⋅178147 EEJ (termed N-short) had a combined expression of 70.4 TPM (1,182 full length reads), while four isoforms containing the RBFOX2⋅178149 EEJ (termed N-long) were expressed at 21.7 TPM (364 full length reads) accounting for 76% and 24%, respectively, of total RBFOX2 RNA expression in the Control (Fig. 4b). Knockdown of the more abundant N-short EEJ reduced total RBFOX2 RNA expression from 92 TPM to 35 TPM whereas knockdown of the less abundant N-long EEJ had minimal impact on total expression (92 TPM to 89 TPM in Fig. 4b). This limited effect was due to an increase in N-short RNA isoform expression (70 TPM to 85 TPM), despite the expected decrease in N-long RNA (21 TPM to 3.1 TPM), suggesting a compensatory feedback mechanism.
Next, we compared full-length TPM for each of the RBFOX2 isoforms in control relative to EEJ knockdown of the two AltTSS events (Fig. 4c). As expected, isoforms containing the targeted EEJ (colored points as in a) showed decreased TPM, while isoforms without the targeted EEJ (gray points) did not (Fig. 4c, Supplementary Fig. 4F, G). As an example, the point for the most abundant RBFOX2 RNA isoform in the hESC control, RBFOX2-208 at 28.1 TPM (472 full length reads), is marked in each plot. RBFOX2-208 contains EEJs 178147, 178145, and 178135 and decreases significantly in each knockdown from 28.1 to 3.31, 5.75, and 15.6, respectively (Fig. 4c, Supplementary Fig. 4F, G). Conversely, RBFOX2-208 does not contain EEJs 178149, 178146, or 178136 and does not decrease in these knockdowns. Similar plots for the Alt3SS⋅1 and Alt3SS⋅2 events are in Supplementary Fig. 4F, G. These results confirm that targeting unique or partially unique EEJs is highly specific in knocking down its intended isoform target(s). Remarkably, this remains true for EEJs that differ by only three nucleotides, as in Alt3SS⋅1 (Supplementary Fig. 4F).
Functional impact of RBFOX2 isoform-specific knockdown in hESCs
The ultimate goal of developing an isoform-specific targeting system is to advance our understanding of isoform biology. For most isoforms of interest, the functional unit is the protein, so evaluating protein-level knockdown is essential for interpretation of downstream effects. We focused on the RBFOX2 AltTSS event, which had the greatest impact on the protein sequence, with N-short and N-long isoforms containing 8 and 78 unique amino acids at their respective N-termini (Fig. 4d). Surprisingly, the knockdown of N-short isoforms, which are most abundant at the RNA level (shown by short and long read RNA-seq in Figs. 4a, b, respectively), showed a minimal decrease in RBFOX2 total protein expression relative to control via western blot (27% reduction; Fig. 4d). Conversely, N-long isoform knockdown resulted in an 83% reduction in total RBFOX2 protein. We confirmed these total protein-level changes by quantifying the reduction in fragment ion intensity of RBFOX2 peptides common to all isoforms using targeted mass spectrometry (Fig. 4e, Supplementary Fig. 5A, B, Supplementary Data 1, 2). These findings show that the lesser abundant RNA isoforms, N-long, are the predominant RBFOX2 protein isoforms in hESCs and further suggest that they are the functionally relevant isoforms.
RBFOX2 is a well-studied splicing factor that binds RNA through a highly conserved RNA Recognition Motif (RRM) and interacts with a specific set of splicing regulators via its C-terminal protein domain56,57. Importantly, both N-short and N-long isoforms share the RRM and C-terminal domain, but the biological relevance of RBFOX2’s diverse N-termini remains largely unexplored. To investigate downstream splicing changes following N-short versus N-long isoform-specific knockdown, we performed RNA-sequencing followed by rMATS analysis across three RBFOX2 knockdown timepoints: 24, 48, and 96 hrs. In the RBFOX2 total knockdown–using a gRNA targeting an EEJ shared across all isoforms–we identified 93 skipped exon (SE) events that were consistently differentially spliced across all timepoints (Fisher’s combined test, adjusted for multiple testing using Benjamini-Hochberg correction, FDR < 0.05; Fig. 4f). Of these 93 SE events, 0 and 35 were significant across all timepoints in N-short and N-long knockdown, respectively (Fig. 4f; Supplementary Data 3). We highlight the effects on two SE events in GOLIM4 and CSTN1 (Fig. 4g left) and validate using PCR (Fig. 4g right), demonstrating concordance with RNA-seq rMATS quantification. These functional results are consistent with our observation that RBFOX2 protein-level changes are specific to N-long knockdown.
Collectively, these data show that, despite their low RNA abundance, N-long isoforms execute the splicing regulatory function of RBFOX2 in hESCs. The discrepancy between RNA and protein isoform expression may stem from factors such as translation initiation, translation efficiency, or protein stability. Our findings highlight the need to consider these factors more broadly as potential mechanisms by which alternative isoforms impact phenotypes. Notably, this discovery was only possible with a system that targets the endogenous locus and is capable of targeting alternative first exons, not just cassette exons, demonstrating the value of our strategy for understanding the functional impact of isoform diversity.
Practical applicability of our unified EEJ-targeting strategy and computational tools
Finally, we integrate our experimental and computational frameworks to address key practical considerations for isoform-specific knockdown studies. Specifically, we assess: (1) how TIGER-based gRNA selection improves screen performance, (2) how well TIGER scores predict RNA knockdown levels, and (3) the likelihood of identifying an effective gRNA for a given EEJ or isoform.
To address point 1, we re-evaluated screen performance in Fig. 1 by generating receiver operating characteristic (ROC) and precision-recall (PR) curves using gold standard essential and non-essential genes as positive and negative labels, respectively (Supplementary Fig. 5C, D). We compared screen performance with and without TIGER-based gRNA filtering and observed improvements in classification as TIGER score thresholds increased. AUROC improved from 0.79 (all gRNAs) to 0.86 (TIGER score >0.2, Q3) and 0.89 (TIGER score >0.55, Q4), while AUPRC increased from 0.58 to 0.74 and 0.81, respectively–confirming TIGER’s utility for screen design. We also provide Cas13-specific gene essentiality scores, calculated as the median LFC per gene considering gRNAs with TIGER score >0.2 (Supplementary Data 4).
To address point 2–how well TIGER scores predict the level of RNA knockdown–we plot correlations between observed screen LFC and TIGER scores for the ten most essential genes with greater than 50 predicted active gRNAs (R = −0.46 to −0.72; Fig. 5a). Classifying gRNAs with TIGER score >0.2 as effective yields high precision across these genes, ranging from 0.89 (SF3B1; 64/72 gRNAs) to 0.62 (DYNC1H1; 137/221 gRNAs). Experimenters can adjust the TIGER score threshold to prioritize precision: raising the threshold to the top Q4 quartile (score >0.55) increases precision to 0.97 (SF3B1) and 0.80 (DYNC1H1). For screen design, we recommend prioritizing high precision to ensure confidence that selected gRNAs will be effective. Conversely, for low-throughput experiments targeting a few genes, we suggest selecting the top two predicted gRNAs per EEJ, regardless of score, and testing both for RNA knockdown (see additional guidance in Methods: Practical use of TIGER predictions for gRNA design).
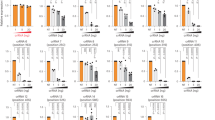
a TIGER scores versus observed log2 fold changes (LFC) for gRNAs targeting the 10 most essential genes in our Cas13 screen (>50 predicted active gRNAs/gene). Each point is a gRNA colored by quartiles of TIGER scores (Q4 = predicted most active). Vertical and horizontal gray lines mark TIGER score of 0.2 and LFC of −1.96 (0.01 quantile of NT control gRNAs’ normal distribution), respectively. Precision (Pr) and recall (Re) are shown for TIGER score >0.2. Linear regression lines (black) and 95% confidence intervals (gray shading) are shown for each plot. Pearson correlations are displayed per gene. p < 0.001 (two-sided) for all comparisons. b Pearson correlations between RT-qPCR measured RNA knockdown and predictions from TIGER and two published Cas13 models. Data aggregated from Figs. 3 and 4 (HEK293). Each point represents an average of ≥2 biological replicates and ≥2 technical replicates. The gray vertical line on the TIGER plot marks a score of 0.2. Linear regression lines (black) and 95% confidence intervals (gray shading) are shown. Pearson correlations are displayed. p < 0.001 (two-sided) for all comparisons. c TIGER scores for gRNAs tiling GENCODE v41 EEJs (8 gRNAs/EEJ; −15/+8 bp to −8/+15 bp relative to splice site). gRNAs with homopolymers or multi-mapping were excluded. Colored ticks at the top correspond to gRNA in (b). Dashed line marks recommended threshold (score >0.2). d Total EEJ gRNAs from GENCODE (green) and those predicted active (score >0.2, purple) for protein-coding and lncRNA genes. e GENCODE-annotated EEJs, isoforms, and genes (green; n above each bar) versus those likely targetable with predicted active EEJ gRNAs (purple; score >0.2). f Isoform category counts (Fig. 1b), adjusted to exclude isoforms with no predicted active EEJ gRNAs (gray). Single-transcript genes were also excluded (n = 5195 protein-coding and n = 11,187 lncRNA). Source data are provided as a Source Data file.
While our essentiality screen conveniently allowed for the testing of thousands of gRNAs at once, it provided only a proxy for the level of RNA knockdown. To assess how TIGER predictions translate to actual RNA knockdown, we aggregated all of the HEK293 RT-qPCR data generated in this paper (34 gRNAs across 14 junctions and 4 genes) and compared observed RNA knockdown to predictions from three publicly available deep learning models for Cas13 gRNA efficacy prediction: TIGER36, Wei et al. and Cheng et al. TIGER performs the best (R = −0.86), and all gRNAs with TIGER score >0.2 were effective to knockdown RNA > 40% (Fig. 5b). Additionally, Wei et al. achieved a good correlation of −0.74 and Cheng et al. shows a −0.56 correlation. All models, except Cheng et al., agreed on which gRNAs would have very low activity (Fig. 5b). Indeed, these gRNAs did not substantially knockdown RNA.
To address point 3, we estimated the likelihood of identifying an effective gRNA for targeting EEJs and isoforms genome-wide. We used TIGER to score eight tiling gRNAs per EEJ across all GENCODE human basic annotation transcripts (1.8 million gRNAs after filtering out gRNAs with homopolymers and non-unique sequence; Fig. 5c). Based on our assessment of TIGER performance and gRNA effectiveness (Fig. 5a, b), we set a TIGER score threshold of 0.2 to identify 899,673 predicted active EEJ-spanning gRNAs (Fig. 5c, d). These active gRNAs target 201,905 EEJs across 80,440 isoforms, belonging to 17,752 protein-coding and 12,831 lncRNA genes (Fig. 5e). Remarkably, if we reconsider the number of isoforms from Fig. 1b that have unique or partially unique EEJs that can be targeted with a predicted active EEJ gRNA, we conclude that up to 89% of isoforms from multi-isoform genes (excluding the 5195 protein-coding and 11,187 lncRNA single transcript genes) are uniquely targetable (Fig. 5f). These targetable isoforms harbor almost all types of splicing and transcriptional events, including cassette exons, alternative 5’ and 3’ SS, and alternative first and last exons.
Discussion
We show that EEJs are broadly accessible to the Cas13d complex–and therefore targetable–for RNA knockdown. This expands the diversity of RNAs that can be specifically targeted, as EEJs often contain sequences unique to individual isoforms. Our EEJ-targeting strategy offers a versatile approach for isoform-specific RNA knockdown–one that is robust and applicable across diverse alternative isoform events, genes, and cell types.
Initially, we considered that targeting EEJs might be challenging due to their transient nature in the nucleus, or the Cas13d complex being blocked by the disassembling spliceosome or the exon junction complex (EJC). Based on this, we examined a list of EEJ-relevant features for TIGER model training (Fig. 2, Supplementary Fig. S2). We also explored explicitly including predicted or measured RBP binding as features58, but observed no improvement in predictive accuracy (results not shown). We caution against over-interpreting this finding given the limited cross cell-type accuracy of existing RBP binding predictors and the lack of RBP binding profiles in A375. However, to offer further evidence, we did not detect any effect of broader EEJ sequence on gRNA efficacy beyond the 23 bp gRNA/target sequence itself (Fig. 2f), implying that RBP binding does not play a substantial role in determining gRNA efficacy. Thus, as with exon-targeting gRNAs, EEJ-targeting gRNA efficiency is primarily dependent on gRNA and target RNA sequence, and these sequence determinants are indistinguishable between the two. Based on this, we expect that a gRNA will work similarly across different cell types when its target is expressed.
Although not tested here, we speculate that, with minimal adaptations, the strategy of targeting EEJs could be extended to other species that undergo splicing59, and that RNA-targeting systems beyond Cas13d could also be effectively utilized for EEJ-based isoform specific knockdown. These findings have additional implications for RNA-based therapeutics including Antisense Oligonucleotides and siRNAs60,61,62.
A major challenge in interpreting isoform biology is distinguishing isoform-level from gene-level effects, especially in knockdown/knockout as opposed to isoform-switching systems. We describe three possible scenarios in which isoform-specific function can be inferred in a high throughput screening context. First, a greater phenotypic effect is measured upon knockdown of a lower expressed RNA isoform. Our RBFOX2 case study provides a compelling example of this: knocking down the relatively lowly expressed N-long RNA isoforms have a greater effect on downstream splicing than knocking down the more abundant N-short RNA isoforms. Barring a non-monotonic relationship between total RBFOX2 expression and splicing, this strongly indicates isoform-specific function–likely driven by enhanced translation or protein stability in the N-long isoforms. Second, knockdown of two isoforms have opposing effects. For example, one isoform might reduce proliferation while another promotes it. Third, there is a stronger phenotypic effect upon knockdown of the higher expressed isoform. This is the most challenging scenario to interpret because the observed phenotype could be due to isoform-specific function or simply from gene-level loss of expression. If one assumes a linear relationship between total gene expression and phenotype, these more subtle differences in isoform-specific differences can be detected. Importantly, in a screening context, SEABASS supports testing for isoform differences for all three scenarios.
Although we do not perform an isoform-specific functional screen here, we propose that our EEJ targeting strategy can be applied in a high throughput setting. However, selecting an appropriate screening output is critical when designing isoform-specific screens. An interesting future direction is using Perturb-seq to generate high-dimensional single-cell RNA-seq phenotypic data for the detection of isoform-specific effects. CRISPRi knockdown typically produces a median of 45 differentially expressed genes (DEGs) for non-essential genes and 500 DEGs for essential genes63. This suggests that knocking down two isoforms of a gene with meaningfully different function will likely result in detectably different expression responses, even for genes not directly involved in gene regulation. Ultimately, as the number of detectable isoforms continues to grow across diverse cell types and disease contexts, isoform-level screening platforms will be valuable for prioritizing isoforms of interest for further experimental follow-up.
We were particularly interested in the knockdown of the two RBFOX2 alternative TSS events, as previous studies have primarily focused on C-terminal differences56,57. While we were expecting to identify isoforms with distinct protein functions, our findings instead highlight an interesting and potentially widespread role of alternative transcription and splicing: regulating gene dosage. This echoes recent work demonstrating that many genetic effects on splicing likely impact phenotypes through nonsense-mediated decay (and consequently, dosage) rather than altering protein function64. We show here that additional mechanisms–such as translation efficiency and protein stability–must also be considered. These mechanisms are particularly relevant for isoforms that vary at their N-termini, where 5’UTRs directly contribute to translation initiation and the N-terminal amino acid affects protein stability65,66.
It is now clear that isoform diversity plays a pivotal role in development, cell-type specific regulatory programs, evolution, and disease–from Mendelian conditions to cancer and complex traits such as neuropsychiatric disorders. Despite this knowledge, it has remained challenging to study the functions of individual alternative isoforms in an endogenous cellular context with existing experimental and computational methods. In this paper, we have made significant progress to address this challenge.
Methods
Ethical statement
HUES66 was derived at Harvard University, where ethics approval was obtained prior to its registration with NIH under NIH Human Embryonic Stem Cell Registry; NIHhESC-10-0057. The New York Genome Center cannot speak to the consent process or conditions surrounding donation except to note that all NIH-registered human embryonic stem cell lines are in compliance with federal requirements. At the New York Genome Center, all research involving human embryonic stem cells is overseen by its Institutional Stem Cell Research Oversight Committee (ISCRO). This project was originally approved by the NYGC ISCRO on July 11, 2019 under protocol # ISCRO004.
Cas13d essentiality screen
Monoclonal cell line generation for screen
A375 cells were acquired from American Type Culture Collection (ATCC Cat #CRL-1619). A375 cells were maintained at 37 °C with 5% CO2 in DMEM media (ThermoFisher Cat #11965118) supplemented with 10% serum (Serum Plus II, Sigma-Aldrich, Cat #14009C).
To generate the doxycycline-inducible Cas13d A375 cell line used in the screen, we transduced cells with lentivirus carrying the previously published plasmid, TRE_NLS-RfxCas13d-NLS-HA (Addgene #138149) at a low multiplicity of infection (MOI < 1). Cells underwent drug selection for one week with 5ug/ml of blasticidin (ThermoFisher, Cat #A1113903). Colonies were picked after sparse plating, expanded, dox induced, and screened for Cas13d expression using western blotting and immunofluorescence methods with an anti-HA antibody (Cell Signaling Cat #2367). Cas13 knockdown activity was confirmed by FACS using CD46 as a positive control (Cell Signaling Cat #13241).
Guide library design
To design the Cas13d gRNA library targeting exon-exon junctions (EEJs) in A375 melanoma cells, we identified EEJs that have evidence of expression. First, we downloaded three publicly available RNA-seq datasets from A375 cells (SRR6515912, SRR6515913, and SRR6515914 from GSE10973144;. Next, we processed sequencing reads according to LeafCutter recommendations15: reads were aligned to hg38 (GENCODE v32 gtf annotations) using STAR v2.7.1 with parameters ‘--twopassMode’ and ‘--outSAMstrandField intronMotif’67. Sequencing reads that overlap EEJs were counted using ‘regtools junction extract’ v0.5.2 with parameters ‘-a 8 -m 50 -M 500000 -s 1’68. Next, to define intron clusters using LeafCutter, we ran the ‘leafcutter_cluster_regtools.py’ python script with ‘-m 30 -p 0.01 -l 500000 parameters’15.
To select genes to target in our essentiality screen, we employed a multi-step filtering approach. First, we considered 1000 candidate genes that had the lowest LFC (i.e. genes that, when knocked out, reduced cell proliferation) from the GeCKO essentiality screen using CRISPR/Cas9 in A375 cells 43DEPMAP43. Next, we overlapped this gene list with the LeafCutter A375 EEJ counts discussed above, and we filtered out EEJs that had <10 junction counts. We used Leafcutter clusters to assess if the EEJs were common or unique in A375 cells, requiring that unique EEJs have at least 15% usage across all three replicates. Next, we designed eight 23 bp gRNAs overlapping each EEJ, starting at −15bp and +8 bp and sliding by 1 bp to −8bp and +15 bp relative to the EEJ. We chose this tiling window of up to 15 bp overlapping a single exon based on previous work showing loss of RNA knockdown efficiency using gRNAs less than 20 bp37. We chose this conservative 15 bp cutoff, but others could test 16/7, 17/6, 18/5, and 19/4 splits to see if they can achieve isoform-specific RNA knockdown. We used ‘bedtools fasta’ to get the sequence.
To obtain the final set of gRNAs, we used Bowtie69 to align reads to the transcriptome69 and filtered out gRNAs that aligned to more than one gene locus when allowing up to 2 mismatches (-v 2). Additionally, we eliminated gRNAs with homopolymers with a length 5 or more for A’s, G’s, and C’s and 4 or more for T’s. After filtering, our final essential gene screen library contained 50,310 gRNAs targeting 6932 EEJs in 942 essential genes. As controls, we included 963 non-targeting guides (random 23 bp sequences with no matches in the transcriptome) and 1972 common EEJ gRNAs targeting 409 non-essential genes.
gRNA library synthesis, cloning, and amplification
Pooled gRNA libraries were synthesized as single-stranded oligonucleotides (Twist Biosciences) and resuspended to a concentration of 10 ng/µl in TE. Cloning and sequencing were performed the same as by Wessels and colleagues36.
Briefly, overhangs for Gibson cloning were added to the oligonucleotide library by PCR using the oligo_amp_FW and oligo_amp_REV primers (Supplementary Data 5). Libraries were PCR amplified in 8x 50ul reactions per 10,000 gRNAs (0.5 µl Q5 polymerase (NEB Cat #M0493), 10 µl 5× reaction buffer, 2 µl oligo pool (1 ng/µl), 2.5 µl of each forward and reverse primer (10 µM), 2.5 µl dNTPs (10 mM) and 30 µl water). PCR conditions were 98 °C/30 s, 10× (98 °C/10 s, 63° C/10 s and 72 °C/15 s) and 72 °C/3 min. The PCR amplified library was gel-purified, quantified, and cloned into BsmBI-digested pLentiRfxGuide-Puro (Addgene #138151) via Gibson Assembly. Eight Gibson reactions were performed with a 20-µl reaction volume each time (500 ng digested plasmid (0.088 pmol), 123.15 ng purified oligo pool (1.3245 pmol, 15:1 molar ratio), 10 µl 2× Gibson Assembly Master Mix (NEB)), incubated for 1 h at 50 °C. Next, to expand the gRNA library, the assembled library plasmid pool was electroporated into Endura cells (Lucigen, Cat #60242-2) at 50–100 ng/µl. After electroporation, cells were recovered in LB medium for 1 h and plated on LB agar carbenicillin at 37 °C for 12–14 h. To achieve good library representation, we aimed to get a coverage of >200 colonies per gRNA. The library plasmid pool was extracted from harvested bacterial cells with the IBI Maxiprep Kit (Cat #IB47125). Complete library representation with minimal bias was verified by Illumina sequencing (MiSeq, Cat #MS-103-1002).
gRNA library screening and sequencing
Lentivirus was produced via transfection of library plasmid pool and appropriate packaging plasmids (psPAX2, Addgene #12260 and pMD2.G, Addgene #12259) using linear polyethylenimine MW25000 (Polysciences, Cat #23966). We seeded ten million A375 cells per 10 cm dish and transfected with 60 µl polyethylenimine, 9.2 µg plasmid pool, 6.4 µg psPAX2 and 4.4 µg pMD2.G. At 3d post-transfection, viral supernatant was collected and passed through a 0.45-µm filter and stored at −80 °C until further use.
Doxycycline-inducible RfxCas13d-NLS A375 cells were transduced with the pooled library lentivirus in separate two infection replicates, ensuring at least 1000× guide representation in the selected cell pool per infection replicate using spinfection. After 24 h, cells were selected with 1 µg ml−1 puromycin (ThermoFisher, Cat #A1113803), resulting in ~30% cell survival. Puromycin selection was performed 72 h after the addition of puromycin. Assuming independent infection events (Poisson), we determined that ~83% of surviving cells received a single sgRNA construct70. After completed puromycin selection, input sample was collected (Day 0), and RfxCas13d expression was induced by the addition of 1 µg/ml doxycycline (Sigma-Aldrich, Cat #D9891). Cells were passed every 2–3 d (maintaining full representation) and supplemented with fresh doxycycline. We collected genomic DNA (gDNA; at least 1,000 cells per construct representation) from each sample on Day 0, Day 7, Day 14, and Day 21.
To extract gDNA, screen cells were lysed as in ref. 70 with 12 ml of NK lysis buffer for 100 million cells (50 mM Tris, 50 mM ethylenediaminetetraacetic acid, 1% SDS and pH 8). Once cells were resuspended, 60 µl of 20 mg/ml Proteinase K (Qiagen) was added and the sample was incubated at 55 °C overnight. The next day, 60 µl of 20 mg/ml RNase A (Qiagen) was added and mixed, and samples were incubated at 37 °C for 30 min. Then, 4 ml of prechilled 7.5 M ammonium acetate was added, and samples were vortexed and spun at 4000 × g for 10 min. The supernatant was placed in a new tube, mixed well with 12 ml isopropanol and spun at 4000 × g for 10 min. DNA pellets were washed with 12 ml of 70% ethanol, spun and dried, and pellets were resuspended with 0.2× TE buffer (Sigma-Aldrich). In addition, we also generated linearized plasmid library input and diluted it down to mimic similar copy number conditions as the gDNA samples.
To amplify the gRNA cassette from gDNA and add indexing for Illumina sequencing, we used a two-step PCR protocol, PCR1 and PCR2, respectively. For the PCR1 reaction, we used 960 µg gDNA for each sample. We performed 96× 100 µl PCR1 reactions per sample (10 µl 10× Taq buffer, 0.02 U/µl Taq-B enzyme (Enzymatics, P7250L), 0.2 mM dNTPs, 0.2 µM forward and reverse primers and 100 ng gDNA per µl). Thermocycler conditions were 94 °C/30 s, 20× (94 °C/10 s, 55 °C/30 s, 68 °C/45 s) and 68 °C/3 min. For each sample, all PCR1 products were pooled and mixed. For the PCR2 reaction, we performed 2 reactions per sample (20 µl 5× NEB Q5 buffer, 0.01 U/µl Q5 enzyme, 20 µl PCR1 product, 0.2 mM dNTPs and 0.4 µM forward and reverse PCR2 primers in 100 µl). Thermocycling conditions for PCR2 were 98 °C/30 s, 7× (98 °C/10 s, 63 °C/30 s and 72 °C/45 s) and 72 °C/5 min. For each sample, PCR2 products were pooled, followed by normalization (gel-based band densitometry quantification), before combining equal amounts of uniquely barcoded samples.
The pooled product was then purified using SPRI beads. First, we performed a 0.6× vol/vol SPRI to remove gDNA carryover, followed by the addition of a 0.3× vol/vol SPRI (0.6 + 0.3 = 0.9× final) to the supernatant to purify the ~260 bp PCR product. The final amplicons were sequenced on Illumina NextSeq 500-II HighOutput 1 × 150 v2.5 (Cat #20024907).
Oligonucleotides can be found in Supplementary Data 5.
Cas13d essentiality screen analysis
Data pre-processing
Cutadapt v3.571 was used to demultiplex reads based on barcode sequences in the forward primer of PCR2 during screen library prep (Barcodes in Supplementary Data 6). We included the 8 bp barcode plus 8 bp of the U6 sequence (“TCTTGTGG”) to ensure the proper position of the barcode match within the read. We allowed for 1 mismatch within the 16 bp sequence (-e 1 -O 16, –action=none). Cutadapt was also used to trim 5’ (54 bp) and 3’ (16 bp) sequences upstream and downstream of the gRNA sequence (-e 0.1). Bowtie v1.1.269 index was built from the screen library fasta file and reads were subsequently aligned to the library index with strict parameters of -v 0 that do not allow for mismatches and -m 1 that only allow for a single alignment (--norc --best --strata). We used an in house R script that inputs the sam alignment file and outputs a raw count matrix of gRNA read counts per sample.
Technical replicates, re-sequencing of the same library for greater read depth, were combined for each sample and all future analyses were performed on this pool. Then, we added 1 to all values to prevent division by 0 for entirely depleted gRNAs. Following DESeq72 we divided counts \({K}_{{gt}}\) for each guide g by its geometric mean \({K}_{g}^{R}={({\Pi }_{t}{K}_{{gt}})}^{-1}\) across time-points t to obtain \({D}_{{gt}}={K}_{{gt}}/{K}_{g}^{R}\). We then normalize by the median-of-ratios \(s={media}{n}_{g\in {nt}}({D}_{{gt}})\) corresponding to the non-targeting (nt) gRNAs to give normalized counts \({Q}_{{gt}}={D}_{{gt}}/s\). We visualized these normalized counts and removed gRNAs that had low or high detection in the Day 0 sample (cutoffs are 0.01 and 0.99 quartiles of normalized gRNA count distribution). Finally, we calculated log2 fold changes (LFCs) by dividing normalized counts across each time point by the corresponding replicate at Day 0 and taking their log2.
Essential gene classification lists for screen evaluation
We considered three essential gene lists to evaluate screen performance across our 942 essential genes: (1) Gold Standard (n = 217 genes40,41), (2) DEPMAP Common Essential genes list (CRISPRInferredCommonEssentials.csv, n = 1,552 genes42), and (3) DEPMAP GeCKO A375 data43.
Defining active versus inactive gRNAs in Cas13d essentiality screen
To label gRNAs as active vs inactive, we employ the same methodology as36. We used maximum likelihood estimation to fit a normal distribution to the mean LFC of non-targeting gRNA replicates. An active gRNA was defined as one whose LFC falls below the 1st percentile of this distribution (LFC < −1.96 in our screen).
Filtering of essential gene list for TIGER model training
Different screening platforms and cell types are known to return non-overlapping sets of essential genes, largely due to variability in gene essentiality41,73. Some genes require complete knockout to impact cell viability or proliferation, while others show significant effects with only partial knockdown. Previous Cas13 RNA knockdown essentiality screens targeted a limited number of genes (16 and 55 in refs. 36,38), so that there is no comprehensive essential gene list specific for the Cas13 platform. Gene-level variability could bias gRNA efficacy estimation if a gene’s gRNAs are mis-classified as inactive only because the knockdown was not sufficient to reduce cell proliferation. To avoid this bias and ensure accurate labels for model training, we filtered out genes with less than 5% active gRNAs, i.e. fewer than 1 gRNA depleted out of 20 (LFC < −1.96, corresponding to the 0.01 quantile of the NT gRNA’s normal distribution). This removed 356 genes. This filtered list is only used for TIGER model training.
Comparison to previous RNAi essentiality screens
A375 RNAi gene essentiality scores were downloaded from DepMap (file: D2_combined_gene_dep_scores.csv). A single score indicating LFC was provided for every gene.
SEABASS linear mixed model
Existing approaches for analyzing essentiality screen data do not handle multiple time-points and replicates. We therefore developed a probabilistic, hierarchical linear mixed modeling approach which we call SEABASS (Screen Efficacy Analysis with BAyesian StatisticS). We model log2 fold changes \({y}_{{gtr}}\) for guide \(g\) at time-point (week) \(t\) and replicate \(r\) using the linear mixed model,
where \({\beta }_{g}\) is a per guide slope, \({\epsilon }_{{gtr}}\sim T(0,{\sigma }_{1},{\tau }_{1})\) is noise, \({\alpha }_{{gr}}\sim T(0,{\sigma }_{2},{\tau }_{2})\) are random slope (i.e. random effect) terms, and \(T(m,\sigma,\tau )\) is a Student-t distribution with mean \(m\), scale \(\sigma\), degrees of freedom \(\tau\) and therefore variance \(\tau {\sigma }^{2}/(\tau -2)\) (for \(\tau > 2\)). The hyperparameters \({\{{\sigma }_{1}},{{\tau }_{1}},{{\sigma }_{2}},{{\tau }_{2}}\}\) are shared across all guides (and genes). By fitting \({\tau }_{1}\) and \({\tau }_{2}\) we can control how heavy-tailed the noise distribution is. For \(\tau=1\) the Student-t corresponds to a Cauchy distribution (extremely heavy tails), and for \(\tau \to \infty\) a Gaussian (light tails). The noise distribution parameters \(\{{\sigma }_{1},{\tau }_{1},{\sigma }_{2},{\tau }_{2}\}\) are learnt on the non-targeting (NT) guides only, where we fix \({\beta }_{g}=0\). We additionally explored using Laplace or explicit Gaussian or Cauchy distributions for the noise and random slope terms but found these provided a worse fit to the NT data according to the evidence lower bound (ELBO). Our use of the Student-t distribution endows SEABASS with natural robustness to outliers.
We put a gene-dependent prior on the per-guide slopes \({\beta }_{g}\sim D(0,{s}_{\gamma })\) where \(D(m,s)\) is a location-scale distribution with mean \(0\) and scale \(s\), and \(\gamma\) is the gene targeted by guide \(g\). The per gene scales \({s}_{\gamma }\) capture differences in gene essentiality. We explored \(D\) being Gaussian, Cauchy, Student-t or Laplace, and choose Laplace since it gave the lowest estimated false positive rate (assuming all significantly positive \({\beta }_{g}\) are false positives). We put a log-normal prior on \({s}_{\gamma }\) i.e. \(\log ({s}_{\gamma })\sim N({\mu }_{s},{\sigma }_{s}^{2})\).
We use stochastic variational inference (SVI) in pyro to fit the model jointly across all guides (and all genes). We use a structured variational posterior where \(\{{\beta }_{g},{\alpha }_{g1},\cdots,{\alpha }_{{gR}}\}\) for a guide are drawn from a multivariate normal (where \(R\) is the number of replicates). This is to account for the strong posterior dependencies we expect between these variables. For \({s}_{\gamma }\) we use a (diagonal/mean-field) normal variational posterior on each \(\log ({s}_{\gamma })\). We optimize \(\{{\sigma }_{1},{\tau }_{1},{\sigma }_{2},{\tau }_{2},{\mu }_{s},{\sigma }_{s}\}\) by placing Dirac delta variational “posteriors” on these parameters.
We found that SVI using just one Monte Carlo (MC) sample for gradient estimation (the default in pyro) and the Adam optimizer did not fully converge. To address this, we developed a strategy where we monitored the ELBO for optimization having stalled, assessed by the ELBO for the last 10 epochs not showing a statistically significant (p > 0.05) downward slope. We then double the number of MC samples used for gradient estimation and resume optimization. We go up to a maximum of 32 MC samples. This resulted in improved ELBOs and agreement between parameter estimates across random initializations.
TIGER model architecture
We closely mimicked the published TIGER architecture36: two convolutional layers with 32 filters, each of length 4, and rectified linear unit (ReLU) activations. This is followed by a max pooling layer with a pool size of 2. The data is then flattened into a vector and subjected to dropout with a rate of 0.25 for regularization. Subsequently, the features are concatenated with non-sequential features and fed into a dense layer with 128 sigmoid outputs. Another dropout layer with a rate of 0.1 is applied here. This is succeeded by a dense layer with 32 sigmoid outputs and another dropout layer with the same 0.1 rate. The final layer is a linear output layer, producing a scalar LFC prediction. We additionally score these predictions (i.e. map them to the unit interval) using a sigmoid function similar to35,36,38. In particular, we map the 0.1 quantile (very active LFCs) of all gencode LFC predictions to 0.9 (a high score) and the 0.9 quantile (inactive LFCs) of all gencode LFC predictions to 0.1 (a low score).
In generating TIGER’s, TIGERjunctions’s, and TIGERbass’s predictions for our screen, all gencode EEJs, and our RT-qPCR tested gRNAs, we performed 10-fold cross validation, ensuring that any gencode or RT-qPCR gRNA never appears in the training pool. This procedure results in a single prediction for gRNAs that appear in the training set TIGER uses36. TIGERjunction uses our screen data, and TIGERbass uses SEABASS slope estimates from our screen data–and an ensemble of 10 predictions for gRNAs that did not, which we then average. TIGERjunction and TIGERbass only consider transcript sequence since guide sequence is redundant in our setting of perfectly matched gRNAs. For TIGER guide models (Fig. 2, Supplementary Fig. 2), we use +/−1 nucleotide of additional transcript sequence context like the published TIGER architecture36. However, we do not normalize for gene essentiality as TIGER does, since this requires a significant number of guides targeting a single gene, which our screen does not have.
In the EEJ-targeting setting, Wessels et al.’s additional scalar features related to gRNA position along a transcript, distance to nearby EEJs, and transcript length are ill-defined. Their results suggest these features have minimal impact on predictions (via SHAP analysis) and on predictive performances (via feature holdouts). Therefore, we only consider the following features for our training of TIGER, TIGERjunction, and TIGERbass (Fig. 2, Supplementary Fig. S2):
-
Target accessibility
-
log unpaired
-
log unpaired at position 11
-
log unpaired at position 19
-
log unpaired at position 25
-
-
Hybridization minimum free energy (MFE)
-
positions 1 and 23
-
positions 3 and 12
-
positions 15 and 9
-
-
Guide minimum free energy (MFE)
-
Guide secondary structure
-
presence of direct repeat stem loop (a Boolean variable)
-
presence of a g-quadruplex (a Boolean variable)
-
For TIGERsite (Fig. 2), we provide 100 nt of transcript sequence only and do not consider these additional scalar features.
Assessing model feature importance
To determine TIGER’s learned gRNA design rules, we performed ten-fold cross validation, collecting Shapley additive explanations (SHAP)48 values for each element in the fold such that we have a SHAP value for every dataset element. Averaging these values conditioned on positional nucleotide identity, we observed TIGER learns known gRNA design rules both when trained on exon data36 and when trained on our EEJ screen data. We similarly collected SHAP values for the junction-sequence TIGER model (Fig. 2d) that predicts the average of an EEJ’s tiled gRNAs’ LFCs from 50 nt up- and down-stream of the EEJ.
Comparison to other publicly available models
To obtain Cheng et al. predictions for all EEJ-spanning gRNAs, we re-trained the DeepCas13 model using LFCs for the 5726 gRNAs provided in their paper35(https://bitbucket.org/weililab/deepcas13/src/master/). We then used this model to predict gRNA efficiency for all 2.2 million GENCODE EEJ-spanning gRNAs. DeepCas13 outputs slightly different but highly correlated (r > 0.94) predictions each time it is trained. Therefore, we repeated this process 5 times and averaged them to get a single prediction score per gRNA.
To obtain Wei et al. predictions for all EEJ-spanning gRNAs, we downloaded the already generated predictions for Human RefSeq coding genes [refseq_coding_guides_prediction_sorted.csv] from https://www.rnatargeting.org/38 and merged the 7.4 million gRNA predictions with our GENCODE EEJ gRNA list. Their model uses gRNAs that are 30 bp long, so we extracted the first 23 bp. This results in gRNA predictions for 1,617,364 of the 2.2 million GENCODE gRNAs.
Analysis of non-sequence feature association with gRNA efficacy residuals
In addition to sequence and the ‘Cell non-specific set’ of features (Fig. 2b and Methods: TIGER Model Architecture) that are known to be important from ref. 36, we considered certain non-sequence features that might be more important to EEJ targetability. We hypothesized that these features may help explain discrepancies in gRNA efficacy predictions beyond what is captured by TIGERbass. Guide-level residuals were calculated as observed slopes minus predicted slopes, and EEJ residuals were calculated by averaging all available gRNAs per EEJ. The number of guides, EEJs and genes evaluated herein included 27,804, 3822 and 480 respectively.
For non-sequence features, some datasets were generated in-house, while others were pulled from additional sources.
A375 gene expression, percent gene nuclear, and percent EEJ nuclear were calculated from A375 RNA-seq data generated in this paper (see ‘Methods: Illumina short read sequencing analysis’ and Supplementary Data 6 for additional information). Briefly, gene counts were obtained using featureCounts74 and were normalized to RPKM (Reads Per Kilobase Million) and junction counts were obtained using RegTools68. Percent nuclear values were obtained by dividing the nuclear read counts by the sum of the nuclear and cytoplasmic read counts for a gene or EEJ. We recognize that these are not absolutely quantifications of nuclear localization but are relative quantifications for comparison between genes or EEJs.
Gene length and intron length were obtained from Hg38 coordinates from the ‘annotables’ package (https://github.com/stephenturner/annotables). Intron lengths were calculated using observed junction lengths. gRNA tiling position and relative EEJ position were obtained from GENCODE v41 annotations. gRNA tiling position indicates positions 1–8 along an EEJ where 1 is −15bp/+8 bp and 8 is −8bp/+15 bp. Relative EEJ position was calculated based on a EEJs’ basepair genomic distance to the gene TSS.
To obtain mean RNA half lives, we downloaded a matrix of genes and RNA half lives across 33 studies75. Half lives were averaged for all available studies for each gene.
All available features were centered and scaled to ensure that all predictor variables were on a similar scale. We then fit a multiple linear regression to assess each feature’s regression coefficient, standard error and p-value.
Practical use of Isoviz and TIGER for gRNA design
We provide four ways to access TIGER predictions for gRNA design.
-
1)
tiger.nygenome.org webtool: Ideal for single gene experiments, exon or EEJ based
-
Convenient (no coding involved)
-
Can predict on any sequence
-
Option to find off targets
-
Low throughput
-
Model version does not include secondary structure as features (this has minimal effect on TIGER performance)
-
-
2)
https://github.com/daklab/tiger: Ideal for screen design, exon or EEJ based
-
High throughput
-
Requires some coding
-
-
3)
Isoviz R package: Ideal for EEJ based single gene
-
Only contains prediction for annotated EEJs
-
gRNAs tiling EEJ from -15 to +15
-
Requires some coding
-
-
4)
Zenodo with DOI 10.5281/zenodo.14968148: Ideal for EEJ based screen design.
-
Only contains prediction for annotated EEJs
-
gRNAs tiling EEJ from −15 to +15
-
Convenient
-
At various TIGER score thresholds, we consider the trade off between Precision (for gRNAs predicted to work, how many will actually work?) and Recall (for all gRNAs that work, how many do you select?) in Fig. 5. We propose a score >0.2 as a reasonable threshold for most experiments. However, in a screen design, we recommend increasing the threshold to prioritize Precision, at the cost of decreasing Recall (so you will not include some good gRNAs in your screen). However, if you have a single EEJ or gene that you want to target in a low-throughput experiment, we recommend selecting the top two predicted gRNAs for that EEJ, independent of score, and testing them for RNA knockdown.
One additional consideration is the tiling window around an EEJ. We chose a conservative cutoff for our screen and subsequent experiments, requiring at most 15 bp overlapping a single exon (15/8 split for a 23 bp gRNA). However, Wessels et al. showed loss of RNA knockdown efficiency using gRNAs less than 20 bps. This suggests that gRNAs outside the windows tested here–such as 16/7, 17/6, 18/5, and 19/4–could also target isoforms specifically, but you would need to verify that for yourself. This would be worth trying if you can’t find a good gRNA within the −15 to +15 window.
In Fig. 5, we exclude GENCODE EEJ gRNAs with homopolymers and non-unique sequences. However, in Isoviz and in the table on Zenodo (DOI 10.5281/zenodo.14968148), we include these gRNAs, but mark in a column why we recommend filtering it out. Importantly, the Isoviz table and Zenodo download do not include off target predictions–this is a feature only available with the webtool. Instead, we provide a column that includes the broader target sequence at that EEJ. You can copy and paste this sequence into the TIGER webtool (run it in off target mode) and get all off target loci with the predicted knockdown effect of those off targets (see Supplementary Fig. 6 for demo).
Finally, when designing gRNAs for an isoform-centric experiment, an initial challenge is confidently identifying which isoforms are expressed in the cell type of interest. In our experience, LRS results are more accurate to determine the presence or absence of specific isoforms. In our PacBio data, we detect a significant number of unannotated isoforms, consistent with other studies (Reese et al.). For example, we were surprised to find that the predominant RBFOX2 isoform containing the most upstream alternative start EEJ (RBFOX2.178149) was an unannotated isoform and not RBFOX2-220 (ENST00000695854.1), the canonical isoform, or RBFOX2-209 (ENST00000438146.7). However, we find EEJ counts from short read RNA-sequencing data to be more accurate for event level comparison within a sample (comparing the same transcript across samples is ok with LRS). Importantly, Isoviz can incorporate both long and short read data, using long reads to identify isoform structures and short reads to give additional quantitative support at the EEJ-level.
piggyBac-Cas13 knockdown experiments
Cloning of Cas13d and gRNA constructs into piggyBac transposon system
To create the doxycycline-inducible Cas13d piggyBac vector, the TRE_NLS-RfxCas13d-NLS-HA was cloned from Addgene plasmid # 13814937 into the piggyBac backbone from Addgene plasmid # 12602949 by digestion with SpeI and EcoRI followed by Gibson Assembly (NEB). As an aside, we initially tested four versions of Cas13d for targeting EEJs: NLS-Cas13-NLS, NLS-Cas13-NES, NES-Cas13-NLS, and NES-Cas13-NES. We decided to use NLS-Cas13-NLS for all experiments in this paper.
To create the rtTA-sgRNA expressing piggyBac vector, the hU6_RfxCas13d-DR_BsmBI was cloned from Addgene plasmid # 13815137 in the piggyBac backbone from Addgene plasmid # 12602849 by digestion with SfiI and BglII followed by Gibson Assembly (NEB). Oligonucleotides used for cloning are in Supplementary Data 5. Full plasmid sequences were verified using plasmidsaurus long read sequencing (https://www.plasmidsaurus.com/).
To clone individual gRNAs, PB_hU6_RfxCas13d-DR_BsmBI was digested with BsmBI (NEB R0739), dephosphorylated with Alkaline Phosphatase (CIP; NEB M0525S), and gel extracted. Oligonucleotides were synthesized for each top and bottom gRNA as 5’-AAAC [23 bp Guide sequence]-3‘ and 5’-AAAA [23 bp Target Sequence]-3’, respectively. Note that Guide Sequence and Target Sequence are outputs of TIGER webtool and are reverse complements of one another. Oligos were annealed using the standard protocol76. Briefly, 1 µl each of 100 µM top and bottom gRNA oligos were mixed with 0.5 µl T4 PNK enzyme (NEB M0201S) and 1 µl 10x T4 ligation buffer (NEB B0202S) in 10 µl total volume and gRNAs were annealed in a thermocycler using the protocol: 37 °C for 30 min, 95 °C for 5 min, ramp down to 25 °C at 5 °C/min. To ligate the annealed oligos into the digested plasmid, 2 µl of 1:100 annealed oligos were mixed with 25 ng digested plasmid, 0.25 µl T7 ligase, and 5 µl 2x T7 Stick together buffer (NEB M0318L). The ligation reaction was incubated at 25 °C for 20 min and transformed into competent cells. Guide RNA sequences cloned and tested for RNA knockdown in this study are in Supplementary Data 7.
We plan to submit both plasmids to Addgene after publication: (1) PB_TRE_NLS-RfxCas13d-NLS-HA and (2) PB_hU6_RfxCas13d-DR_BsmBI.
To allow propagation of the piggyBAC transposase from System Biosciences on ampicillin plates, the transposase was cloned into SmaI and HindIII sites into pUC19 (NEB) as in ref. 77.
Cell culture
HEK293 cells were acquired from ATCC (Cat #CRL-1573). HEK293 cells were maintained at 37 °C with 5% CO2 in DMEM media (ThermoFisher Cat #11965118) supplemented with 10% serum (Serum Plus II, Sigma-Aldrich, Cat #14009C).
HUES66 human embryonic stem cells (hESCs) were obtained from Harvard University. Cells were maintained at 37 °C with 5% CO2 in StemFlex media (ThermoFisher Cat #A3349401) and grown on Geltrex-coated plates (ThermoFisher Cat #A1413302). For passaging, Accutase (ThermoFisher Cat #A1110501) was used to dissociate cells and 10 µM ROCK inhibitor Y-27632 dihydrochloride (Tocris Cat #1254) was added to the media for plating.
Stable transfections of piggyBac Cas13 system
With piggyBac, the number of integrations of cargo plasmids can be tightly controlled by changing the ratio of cargo plasmids to PB transposase plasmid49,78,79. We initially tested two gRNAs at five different gRNA:Cas13:transposase ratios in human embryonic stem cells (hESCs). We perform all experiments in this paper at a 2:2:1 ratio.
To generate stable Cas13/gRNA-expressing HEK293 cell lines, 2 ×105 cells were plated in a single well of a 12-well plate and transfected the following day using Lipofectamine 3000 (Invitrogen #L3000001) according to the manufacturer’s protocol. A 2:2:1 ratio of piggyBac cargo vectors (Cas13d and gRNA) and pUC19-piggyBac transposase, totaling 1.25 µg total of plasmid DNA was mixed with Mix #1 (50 µl OptiMEM +2.5 µl P3000) per well. Mix #2 (50 µl OptiMEM +1.88 µl Lipofectamine 3000 reagent) was added and incubated for 10 min before adding to individual wells. On the day after transfection, cells were selected with 100 µg/mL Hygromycin (ThermoFisher #10687010) and 400 µg/mL G418 (ThermoFisher #10131035) for 7–9 days. After selection, Cas13d expression was induced by adding 1 µg/mL doxycycline (Sigma #D3447-500MG) to the media for 24 h before cells were harvested.
RNA extraction and RT-qPCR analysis
RNA was extracted using TRIzol (Invitrogen #15596018) according to the manufacturer’s protocol. The Qubit RNA HS Assay Kit (Invitrogen #Q32855) was used to quantify RNA and the High Capacity cDNA Reverse Transcription Kit (Invitrogen #4368813) was used to RT 500 ng–1 µg of RNA. qPCR was performed using iTaq Universal SYBR Green (Bio-Rad #1725122) and primers listed in Supplementary Data 5.
We perform standard RT-qPCR analysis to measure the relative RNA knockdown at each target EEJ, comparing measurements taken using the same qPCR primers across samples. To control for sample to sample variation in RNA quantity and RT efficiency, we normalize each target Ct value to the Ct value of a housekeeping control gene within the same sample. Below each figure, we list the number of technical replicates (measurements from the same cDNA sample) and biological replicates (measurements from different transfections) performed in each assay.
PCR validation of skipped exon splicing events
PCR reactions were performed with 2 µl cDNA from HUES66 RBFOX2 knockdown cells treated with 1 µg/mL doxycycline for 3 days. Phusion DNA Polymerase with HF buffer (Thermo F530L) and primers flanking the GOLIM4 and CLSTN1 skipped exon events were used. PCR was carried out for 30 cycles, and products were run on a 2% agarose gel. Exon inclusion and exclusion bands were quantified using ImageJ. Primers are listed in Supplementary Data 5.
RNA fractionation
To isolate RNA from cytosolic and nuclear fractions, cells were washed twice with 1 mL cold PBS and scraped in 1 mL PBS + 1 mM PMSF + 1:100 protease inhibitor cocktail (PIC, Sigma P8340). 200 uL was removed at this step and added to 1 mL Trizol (total RNA). The remaining cells were centrifuged at 1500 × g. for 5 min, and resuspended in 250 uL low salt solution (10 mM KCl, 1.5 mM MgCl2, 20 mM Tris-HCl pH 7.5) supplemented with 1 mM PMSF, 1 mM DTT, and 1x PIC. Triton X-100 was added to a final concentration of 0.1% and cells were rotated for 10 min at 4 °C, then centrifuged for 5 min at 1500 × g. 200 uL of supernatant was removed and added to 1 mL Trizol (cytosolic fraction). The remaining supernatant was discarded and the nuclear pellet was washed by rotating for 2 min at 4 °C in low salt solution without Triton X-100 and centrifuged at 1300 × g for 10 min. Nuclear pellet was resuspended in 1 mL Trizol (nuclear fraction). Isolation of RNA from Trizol was performed according to the manufacturer protocol.
Proteomics experiments and analysis
Protein extraction
RBFOX2 knockdown HUES66 cells (3 days after induction with 1 µg/mL doxycycline) were washed with PBS and then lysed with 300ul ice cold RIPA (Thermo #89900) supplemented with 1 mM PMSF (Thermo #36978) and 1x protease inhibitor cocktail (Sigma #P8340) in a 12-well plate on ice for 5 min. Cells were scraped, transferred to 1.7 ml microcentrifuge tubes and sonicated at 20% intensity for 1 × 30 s pulse (Qsonica Q125 Sonicator with 1/8” diameter probe). Samples were centrifuged at ~14,000 × g for 15 min at 4 °C and the supernatant was transferred to a new tube.
Western blot
Prior to western blotting, protein levels were quantified using the BCA Protein Assay Kit (Thermo #23227). 15 µg total protein was prepared in 1x SDS sample buffer (Thermo #LC2676) with 50 mM DTT and boiled for 5 minutes at 100 °C. Proteins were transferred onto a nitrocellulose membrane, blocked in 5% milk for 1 h at room temperature, and incubated in primary antibody overnight at 4 °C. Primary antibodies used were GAPDH (Cell Signaling, #2118; 1:1000) and RBFOX2 (Bethyl Laboratories, A300-864A; 1:1,000). Membranes were washed 3x for 10 minutes each in 1x TBST and incubated in secondary antibody for 1 h at RT (Thermo, #31460; 1:10,000). Membranes were washed 2 × 10 min in TBST and 1 × 10 min in TBS followed by 5 min ECL incubation (SuperSignal West Pico Chemiluminescent Substrate, Thermo, #34577) and imaging using the C-Digit Blot Scanner (Licor).
Protein digestion
Whole cell lysate pellets were prepared for each of three conditions: Control, N-short Knockdown, or N-long Knockdown. Approximately 10% of each lysate was used as input for each of two filter-aided sample preparations (FASP) adapted from ref. 80: one using Trypsin (Promega, V5280) and the other using Asp-N (Promega, VA1160), 1 µg. Each digestion yielded ~20 µg of peptide as assayed by NanoDrop A280 (ThermoFisher Scientific). Peptides were desalted using Pierce Peptide Desalting Spin Columns (Pierce, ThermoFisher Scientific). The manufacturer’s protocol was used with a substitution of 0.1% trifluoroacetic acid (TFA) with 0.1% formic acid (Optima LC/MS grade, Thermo Fisher Scientific).
LC-MS targeting of RBFOX2 peptides
Desalted samples were analyzed in triplicate by nanoLC-MS/MS using a Dionex Ultimate 3000 (Thermo Fisher Scientific, Bremen, Germany) coupled to an Orbitrap Eclipse Tribrid mass spectrometer (Thermo Fisher Scientific, Bremen, Germany). Each injection of ~1 μg peptides was loaded onto an Acclaim PepMap 100 trap column (300 μm × 5 mm × 5 μm C18) and gradient-eluted from an Aurora Ultimate TS analytical column (75 μm × 25 cm, 1.7 μm C18) equilibrated in 100% solvent A (0.1% formic acid in water) and 0% solvent B (80% acetonitrile in 0.1% formic acid). The peptides were eluted into the mass spectrometer at 400 nL/min up to 100% B over a period of 90 min. MS2 spectra were acquired using the tMS2 (otherwise known as PRM) targeting method. Using this method, only the m/z corresponding to peptides of interest were selected for MS2 acquisition. Separate lists were used for Trypsin-generated peptides and AspN-generated peptides (see Supplementary Data 1 for list of peptides). The following instrument settings were used: positive ion mode was used with 1.7 kV at the spray source, RF lens at 30% with advanced peak determination and XCalibur version 4.5.445.18. Full MS scans were acquired in the Orbitrap from 300 to 2000 m/z with 120,000 resolution. In parallel, the tMS2 targeting was performed on a list of m/z corresponding to either 11 tryptic or 18 AspN-derived peptides of interest. Ions were isolated in the quadrupole, HCD was used for fragmentation (30% normalized collision energy). The resulting fragments were detected in the Orbitrap at 15,000 resolution with standard AGC target and dynamic maximum injection time mode.
Data analysis using Skyline
Target peptide detection was evaluated using the Skyline software suite81 (Skyline (64-bit) 24.1.0.199 (6a0775ef83)). Two tryptic peptides shared among all RBFOX2 isoforms, GFGFVTFENSADADR and ILDVEIIFNER, met dot product spectral correlation criterion of at least 0.5. For each common peptide, the fragment ion intensity from each triplicate injection was normalized relative to the average fragment ion intensity of the Control to generate the boxplots in Fig. 4e (see Supplementary Data 2 for replicate values). All Skyline and mass spec raw file data was uploaded to the Panorama Web repository.
High throughput sequencing and additional analyses
Analysis of GENCODE basic annotations
The R code used to generate the GENCODE summary barplot in Fig. 1b (human) are added to the Isoviz package (isoviz_junction_to_transcript_summary.R). The code can be run in two ways: 1) using GENCODE annotations or 2) long read sequencing generated annotations specific to a cell type. GENCODE annotations for additional species beyond human may also be used. The table includes all EEJs present in the input file with two classification columns: ‘junction_category’ indicates whether the EEJ is classified as common, fully unique, partially unique, or single transcript and ‘Isoform_targetable’ indicates whether the isoform can be targeted uniquely. When comparing classifications of EEJs and isoforms using long read data annotations, outputs are expected to vary based on cell type. It is recommended to consider EEJ and transcript classifications specific to your cell type of interest when designing experiments.
Illumina short read sequencing
Per RNA sample, we combined 1 µg of total RNA + 2 µl 1:100 ERCC RNA Spike-in Mix #1 (Invitrogen #4456740). Libraries were prepared using the Kapa RNA HyperPrep Kit with RiboErase (Roche #08098131702), pooled, and sequenced using the Nextseq 500/550 High Output Kit with 150 cycles (Illumina #20024907).
Illumina short read sequencing analysis
RNA-seq reads were aligned to hg38 genome using STAR v2.7.1 and ‘--twopassMode Basic’ and ‘--outSAMstrandField intronMotif’ parameters (Index built with GENCODE v41 basic annotations gtf67). Samtools v1.9 was used to filter for reads with a mapping quality greater than 20, sorted, and indexed82.
To determine the gene-level RPKM and TPM counts in A375 total, nuclear, and cytoplasmic RNA-seq data, alignment bam files were overlaid with the GENCODE v41 basic annotation gtf file using featureCounts (subread v2.0.4)74.
HEK293, hESC, and A375 junction counts (provided in Isoviz and used for non-sequence features in Supplementary Fig. S2) were generated using Regtools extract v0.5.268 with ‘-a 8 -m 50 -M 500000 -s 0’ parameters.
rMATS differential splicing analysis
Differential splicing analysis was performed using rMATS v4.1.183 and GENCODE v41 Basic Annotations gtf. First, we ran rMATS –task prep for each aligned bam across knockdown conditions and timepoints (--b1 all_bams.txt, -t single --libType fr-firststrand --readLength 158 --variable-read-length --allow-clipping). Next, we combined the prep results in a single post step (--task post). This generates a summary of all data across a single set of events with the same event IDs. Finally, at each timepoint, we statistically compared events in the control to each of the RBFOX2 knockdowns using prepare_stat_inputs.py for each comparison followed by rMATS with –task stat for the final output.
Because we lacked biological replicates at each timepoint for statistical analysis, we used Fisher’s method to integrate the three rMATS p-values for each event across time points. Specifically, we calculate the test statistic,

where i indexes time-points, and then calculated a combined p-value as \(p=1-{F}_{6}(T)\) where\(\,{F}_{6}\) is the CDF of a chi-squared distribution with 6 degrees of freedom. This combined p-value approach enabled us to identify splicing events that consistently responded to RBFOX2 knockdown.
For analysis in Fig. 4, we only considered the 93 skipped exon events (SE) with false discovery rate <0.05 in the total knockdown sample (Fisher’s combined test across timepoints, adjusted for false discovery using the Benjamini-Hochberg correction). We compare these SE events across total, N-short, and N-long RBFOX2 knockdowns. A summary of these events is in Supplementary Data 3.
PacBio long read sequencing
Prior to sending RNA samples to PacBio for long read sequencing, the RNA integrity number (RIN) was calculated using the RNA 6000 Nano Kit (Agilent #5067-1511). RIN scores ranged between 8.7 and 9.7, indicating high quality, intact RNA. Per RNA sample, we combined 1 µg of total RNA + 2 µl 1:100 ERCC RNA Spike-in Mix #1 (Invitrogen #4456740).
MAS-Seq bulk Iso-Seq libraries were generated in collaboration with PacBio using a pre-commercial protocol that implements the MAS-Seq concatenation method55. Briefly, cDNA molecules were concatenated into an ordered array, in this case, an 8-fold array, and sequenced on the Revio system. The concatenated array is sequenced as a HiFi read and then bioinformatically de-concatenated into segmented reads (S-reads) which represent the original cDNA molecules.
PacBio long read sequencing analysis
A pre-commercial version of the SMRT LINK analysis pipeline was used to generate HiFi reads, S-reads, and subsequent full-length reads and high-quality isoform sequences. After the isoseq3 refine step where full-length non-chimeric reads were obtained in bam file format, we continue with our own analysis pipeline.
Minimap2 v.2.1784 was used to align reads to GRCh38 human genome reference with -ax splice:hq -uf parameters. Secondary alignments were removed, and aligned reads were filtered for mapping quality greater than or equal to 60. The resulting bam files were converted to BED12 format and PSL format with FLAIR’s helper scripts85; https://flair.readthedocs.io/en/latest/cite.html). Then, additional filtering was applied to select for full length reads based on overlap with annotated transcription start sites (TSS) and transcription end sites (TES). Our custom script required overlap between −10 bp to +50 bp of TSS and −50 bp to +10 bp of TES (using human GENCODE v41 Basic annotation). Reads that shared the same junction chain were grouped together and collapsed, using a custom script adapted from FLAIR collapse85. Importantly, our custom collapse script ignores start and end coordinates of the reads, allowing reads with the same junction chain, but slight variations in TSS or TES, to be grouped together. Unspliced reads (i.e. mono-exonic reads) were excluded. Resulting full length, collapsed reads were matched with transcripts in GENCODE v41 basic annotations, and any transcripts without a match were still overlapped with the proper gene and was assigned a unique transcript ID through our custom scripts. Finally, the number of reads grouped together per transcript were tallied and output to a separate file per sample. The “million” in TPM was calculated based on the total number of full length reads after filtering.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data generated in this study has been deposited to NCBI Gene Expression Omnibus (GEO) under accession number GSE242107. All mass spectrometry data generated in this study has been deposited to Panorama Web repository under accession number PXD060902. TIGER gRNA predictions across all GENCODE v41 EEJs can be downloaded from Zenodo with DOI 10.5281/zenodo.1496814886. Source data are provided with this paper.
Code availability
Code is publicly available for SEABASS on github [https://github.com/daklab/seabass] and pypi [https://pypi.org/project/seabass/0.0.5/], Isoviz R package on github [https://github.com/daklab/isoviz], and TIGER on github https://github.com/daklab/tiger and with a user-friendly website for gRNA prediction at [tiger.nygenome.org].
References
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Stanley, R. F. & Abdel-Wahab, O. Dysregulation and therapeutic targeting of RNA splicing in cancer. Nat. Cancer 3, 536–546 (2022).
Lewis, B. P., Green, R. E. & Brenner, S. E. Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc. Natl Acad. Sci. Usa. 100, 189–192 (2003).
Yan, Q. et al. Systematic discovery of regulated and conserved alternative exons in the mammalian brain reveals NMD modulating chromatin regulators. Proc. Natl Acad. Sci. 112, 3445–3450 (2015).
Rogalska, M. E., Vivori, C. & Valcárcel, J. Regulation of pre-mRNA splicing: roles in physiology and disease, and therapeutic prospects. Nat. Rev. Genet. 24, 251–269 (2023).
Kelemen, O. et al. Function of alternative splicing. Gene 514, 1–30 (2013).
Nilsen, T. W. & Graveley, B. R. Expansion of the eukaryotic proteome by alternative splicing. Nature 463, 457–463 (2010).
Vuong, C. K., Black, D. L. & Zheng, S. The neurogenetics of alternative splicing. Nat. Rev. Neurosci. 17, 265–281 (2016).
Baralle, F. E. & Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 18, 437–451 (2017).
Scotti, M. M. & Swanson, M. S. RNA mis-splicing in disease. Nat. Rev. Genet. 17, 19–32 (2015).
Chen, S., Benbarche, S. & Abdel-Wahab, O. Splicing factor mutations in hematologic malignancies. Blood 138, 599–612 (2021).
Seiler, M. et al. Somatic mutational landscape of splicing factor genes and their functional consequences across 33 cancer types. Cell Rep. 23, 282–296.e4 (2018).
Raj, T. et al. Integrative transcriptome analyses of the aging brain implicate altered splicing in Alzheimer’s disease susceptibility. Nat. Genet. 50, 1584–1592 (2018).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using LeafCutter. Nat. Genet. 50, 151–158 (2017).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Kahles, A. et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224.e6 (2018).
Danan-Gotthold, M. et al. Identification of recurrent regulated alternative splicing events across human solid tumors. Nucleic Acids Res. 43, 5130–5144 (2015).
Frankish, A. et al. GENCODE 2021. Nucleic Acids Res. 49, D916–D923 (2021).
Thomas, J. D. et al. RNA isoform screens uncover the essentiality and tumor-suppressor activity of ultraconserved poison exons. Nat. Genet. 52, 84–94 (2020).
Gonatopoulos-Pournatzis, T. et al. Genetic interaction mapping and exon-resolution functional genomics with a hybrid Cas9-Cas12a platform. Nat. Biotechnol. 38, 638–648 (2020).
Davies, R. et al. CRISPRi enables isoform-specific loss-of-function screens and identification of gastric cancer-specific isoform dependencies. Genome Biol. 22, 47 (2021).
Gapinske, M. et al. CRISPR-SKIP: programmable gene splicing with single base editors. Genome Biol. 19, 107 (2018).
Celotto, A. M. & Graveley, B. R. Exon-specific RNAi: a tool for dissecting the functional relevance of alternative splicing. RNA 8, 718–724 (2002).
Prinos, P. et al. Alternative splicing of SYK regulates mitosis and cell survival. Nat. Struct. Mol. Biol. 18, 673–679 (2011).
Villemaire, J., Dion, I., Elela, S. A. & Chabot, B. Reprogramming alternative pre-messenger RNA splicing through the use of protein-binding antisense oligonucleotides. J. Biol. Chem. 278, 50031–50039 (2003).
Du, M., Jillette, N., Zhu, J. J., Li, S. & Cheng, A. W. CRISPR artificial splicing factors. Nat. Commun. 11, 2973 (2020).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676.e14 (2018).
Recinos, Y. et al. CRISPR-dCas13d-based deep screening of proximal and distal splicing-regulatory elements. Nat. Commun. 15, 3839 (2024).
Li, J. D., Taipale, M. & Blencowe, B. J. Efficient, specific, and combinatorial control of endogenous exon splicing with dCasRx-RBM25. Mol. Cell 84, 2573–2589.e5 (2024).
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Cheng, X. et al. Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches. Nat. Commun. 14, 1–14 (2023).
Wessels, H.-H. et al. Prediction of on-target and off-target activity of CRISPR-Cas13d guide RNAs using deep learning. Nat. Biotechnol. 42, 628-637 (2024).
Wessels, H.-H. et al. Massively parallel Cas13 screens reveal principles for guide RNA design. Nat. Biotechnol. 38, 722–727 (2020).
Wei, J. et al. Deep learning and CRISPR-Cas13d ortholog discovery for optimized RNA targeting. Cell Syst. 14, 1087–1102.e13 (2023).
Hir, H. L., Saulière, J. & Wang, Z. The exon junction complex as a node of post-transcriptional networks. Nat. Rev. Mol. Cell Biol. 17, 41–54 (2015).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733 (2014).
Morgens, D. W., Deans, R. M., Li, A. & Bassik, M. C. Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes. Nat. Biotechnol. 34, 634–636 (2016).
Tsherniak, A. et al. Defining a Cancer Dependency Map. Cell 170, 564–576.e16 (2017).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Regan-Fendt, K. E. et al. Synergy from gene expression and network mining (SynGeNet) method predicts synergistic drug combinations for diverse melanoma genomic subtypes. NPJ Syst. Biol. Appl 5, 6 (2019).
Wang, Q. et al. The CRISPR-Cas13a Gene-Editing System Induces Collateral Cleavage of RNA in Glioma Cells. Adv. Sci. 6, 1901299 (2019).
Ai, Y., Liang, D. & Wilusz, J. E. CRISPR/Cas13 effectors have differing extents of off-target effects that limit their utility in eukaryotic cells. Nucleic Acids Res. 50, e65 (2022).
Shi, P. et al. Collateral activity of the CRISPR/RfxCas13d system in human cells. Commun. Biol. 6, 1–8 (2023).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 30, 4768–4777 (2017).
Schertzer, M. D. et al. A piggyBac-based toolkit for inducible genome editing in mammalian cells. RNA 25, 1047–1058 (2019).
Ding, S. et al. Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 122, 473–483 (2005).
Wang, G. et al. Efficient, footprint-free human iPSC genome editing by consolidation of Cas9/CRISPR and piggyBac technologies. Nat. Protoc. 12, 88–103 (2017).
Li, S., Zhang, A., Xue, H., Li, D. & Liu, Y. One-Step piggyBac Transposon-Based CRISPR/Cas9 Activation of Multiple Genes. Mol. Ther. Nucleic Acids 8, 64–76 (2017).
Burris, B. J. D., Molina Vargas, A. M., Park, B. J. & O’Connell, M. R. Optimization of specific RNA knockdown in mammalian cells with CRISPR-Cas13. Methods 206, 58–68 (2022).
Maimon, A. et al. Mnk2 alternative splicing modulates the p38-MAPK pathway and impacts Ras-induced transformation. Cell Rep. 7, 501–513 (2014).
Al’Khafaji, A. M. et al. High-throughput RNA isoform sequencing using programmed cDNA concatenation. Nat. Biotechnol. 42, 582–586 (2023).
Damianov, A. et al. Rbfox proteins regulate splicing as part of a large multiprotein complex LASR. Cell 165, 606–619 (2016).
Peyda, P., Lin, C.-H., Onwuzurike, K. & Black, D. L. The Rbfox1/LASR complex controls alternative pre-mRNA splicing by recognition of multipart RNA regulatory modules. Genes Dev. https://doi.org/10.1101/gad.352105.124 (2025).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Kushawah, G. et al. CRISPR-Cas13d INDUCES Efficient mRNA knockdown in animal embryos. Dev. Cell 54, 805–817.e7 (2020).
Urbanski, L. M., Leclair, N. & Anczuków, O. Alternative-splicing defects in cancer: splicing regulators and their downstream targets, guiding the way to novel cancer therapeutics. Wiley Interdiscip. Rev. RNA 9, e1476 (2018).
Cartegni, L. & Krainer, A. R. Correction of disease-associated exon skipping by synthetic exon-specific activators. Nat. Struct. Biol. 10, 120–125 (2003).
Finkel, R. S. et al. Treatment of infantile-onset spinal muscular atrophy with nusinersen: a phase 2, open-label, dose-escalation study. Lancet 388, 3017–3026 (2016).
Nadig, A. et al. Transcriptome-wide characterization of genetic perturbations. bioRxivorg, https://doi.org/10.1101/2024.07.03.601903 (2024).
Fair, B. et al. Global impact of unproductive splicing on human gene expression. Nat. Genet. 56, 1851–1861 (2024).
Bachmair, A., Finley, D. & Varshavsky, A. In vivo half-life of a protein is a function of its amino-terminal residue. Science 234, 179–186 (1986).
Varshavsky, A. The N-end rule pathway and regulation by proteolysis: The N-End Rule Pathway. Protein Sci. 20, 1298–1345 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Cotto, K. C. et al. Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer. Nat. Commun. 14, 1589 (2023).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Sanson, K. R. et al. Optimized libraries for CRISPR-Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 1–15 (2018).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2013).
Agarwal, V. & Kelley, D. R. The genetic and biochemical determinants of mRNA degradation rates in mammals. Genome Biol. 23, 245 (2022).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Kirk, J. M. et al. Functional classification of long non-coding RNAs by k-mer content. Nat. Genet. 50, 1474–1482 (2018).
Cadiñanos, J. & Bradley, A. Generation of an inducible and optimized piggyBac transposon system. Nucleic Acids Res. 35, e87 (2007).
Wilson, M. H., Coates, C. J. & George, A. L. Jr. PiggyBac transposon-mediated gene transfer in human cells. Mol. Ther. 15, 139–145 (2007).
Wiśniewski, J. R. Filter-aided sample preparation for proteome analysis. Methods Mol. Biol. 1841, 3–10 (2018).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl Acad. Sci. USA. 111, E5593–E5601 (2014).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Tang, A. D. et al. Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns. Nat. Commun. 11, 1–12 (2020).
Schertzer, M. & Stirn, A. TIGER predictions for all GENCODE v41 Exon-exon Junction guide RNAs. Zenodo, https://doi.org/10.5281/ZENODO.14968148 (2025).
Acknowledgements
We thank PacBio for partnering with us to generate data for this paper using a pre-commercial product. M.D.S. was supported by NIH/NIGMS (F32GM142213). G.M.S. is supported by NIH/NIGMS (R35GM142647). N.E.S. is supported by NIH/NHGRI (DP2HG010099, R01HG012790), NIH/NCI (R01CA218668, R01CA279135), the NIH/NIAID (R01AI176601), the NIH/NHLBI (R01HL168247), the Simons Foundation for Autism Research (Genomics of ASD 896724), the Chan Zuckerberg Initiative (2024-338515), and NYU and NYGC funds. D.A.K. is supported by Columbia University and NYGC startup funds, and NIH/NCI (R21CA272345). D.A.K. and N.E.S. are supported by the MacMillan Center for the Study of the Non-Coding Cancer Genome. A.S. and D.A.K. were supported by NSF CAREER DBI2146398. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.D.S., L.P., and D.A.K. conceived the study. L.P. and H.W. performed the CRISPR screen. M.D.S., S.H.P., A.R., E.D.J., and C.H. performed all additional experiments. M.D.S., A.S., K.I., S.H.P., A.D., E.D.J., and D.A.K performed analyses. M.D.S. and K.I. developed Isoviz. A.D. and D.A.K developed SEABASS. A.S. developed the variants of TIGER. H.W., G.M.S., and N.E.S. provided reagents and advice. M.D.S and D.A.K. wrote the paper with input from co-authors. M.D.S. and D.A.K. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
G.M.S. is on the scientific advisory board of Quantum-Si Incorporated and holds stock in Quantum-Si Incorporated. N.E.S. is an adviser to Qiagen and a co-founder and adviser of TruEdit Bio and OverT Bio. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Schertzer, M.D., Stirn, A., Isaev, K. et al. Cas13d-mediated isoform-specific RNA knockdown with a unified computational and experimental toolbox. Nat Commun 16, 6948 (2025). https://doi.org/10.1038/s41467-025-62066-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62066-5
This article is cited by
-
Proteoform medicine: characterizing and targeting protein forms in human disease
Nature Reviews Genetics (2026)
-
RNA structure modulates Cas13 activity and enables mismatch detection
Nature Biotechnology (2025)
-
OY-TES-1 Splice Variant V5a in Glioma: A Driver of Malignancy and Potential Therapeutic Target
Current Medical Science (2025)