Abstract
During development, epithelia function as malleable sheets that undergo extensive remodeling to shape developing embryos. Optogenetic control of Rho signaling provides an avenue to investigate mechanisms of epithelial morphogenesis, but transgenic optogenetic tools can be limited by variability in expression levels and deleterious effects of transgenic overexpression on development. Here, we use CRISPR/Cas9 to tag Drosophila RhoGEF2 and Cysts/Dp114RhoGEF with components of the iLID/SspB optogenetic heterodimer, permitting light-dependent control over endogenous protein activities. Using quantitative optogenetic perturbations, we uncover a dose-dependence of tissue furrow depth and bending behavior on RhoGEF recruitment, revealing mechanisms by which developing embryos can shape tissues into particular morphologies. We show that at the onset of gastrulation, furrows formed by cell lateral contraction are oriented and size-constrained by basal actomyosin. Our findings demonstrate the use of quantitative, 3D-patterned perturbations of cell contractility to precisely shape tissue structures and interrogate developmental mechanics.
Similar content being viewed by others
Introduction
Embryonic development is an inherently biophysical process, requiring the integrated action of protein regulators and mechanical effectors to physically sculpt an embryo into a mature animal. Therefore, a proper understanding of embryogenesis requires dissecting both the biochemical pathways that direct morphogenesis and the mechanical behavior of the cells and tissues that enact the developmental program1. Across many animals, early development is defined by the sculpting of epithelia2. Although epithelial tissues are effectively 2D sheets, precise subcellular patterning of active stresses and mechanical properties can shape them into complex 3D morphologies3,4,5.
A well-studied mechanism to sculpt epithelial sheets is apical constriction, which directs tissue furrowing and invagination in a variety of contexts6,7,8. This process is characterized by a contractile apical actomyosin meshwork driving constriction of apical cell surfaces and expansion of basal cell surfaces9. However, recent theoretical and experimental work has additionally implicated lateral actomyosin contractility and cell apical-basal shortening in morphogenesis10,11. Epithelial furrowing driven by lateral cortical actomyosin plays roles in Drosophila cephalic furrow formation12,13 and wing disc morphogenesis14, as well as ascidian endoderm invagination15. More generally, lateral actomyosin contractility gives rise to a columnar-to-cuboidal transition during Drosophila pupal wing shaping16 and apoptosis-coupled tissue bending during Drosophila leg disc folding17. It is unknown why developing embryos employ different furrowing mechanisms in different contexts, or what functional consequences these may have. Additionally, while apical constriction provides a clear polarity-based cue guiding the direction of furrowing toward the interior of the embryo, it is unclear how lateral contraction translates cell shortening into a properly oriented furrow.
Although different morphogenetic processes are driven by distinct patterns of tissue stresses and mechanics, these are largely downstream of the widely conserved Rho signaling pathway18,19. In the Rho pathway, RhoGEFs and RhoGAPs cycle Rho GTPases between their active and inactive states to coordinate downstream actomyosin activity and direct the mechanical behavior of the epithelium. Different RhoGEFs and RhoGAPs are employed in a context-specific manner to initiate Rho signaling, but they converge on tuning the behavior of just a few classes of mechanically relevant molecules20. RhoGEF and RhoGAP activities must be balanced to carry out cell mechanical behaviors, and many are autoinhibited to prevent off-target activity21,22,23. Additionally, some RhoGEFs are known to engage in positive feedback interactions with active Rho124,25,26,27. Therefore, it is not clear to what extent tissues can modulate their shapes by controlling activity levels of a particular RhoGEF, or to what extent RhoGEFs act as on/off switches to enact tissue morphogenetic processes.
The Drosophila melanogaster embryo is a powerful model system for understanding how Rho signaling directs morphogenesis in a variety of contexts. RhoGEF2 and Cysts/Dp114RhoGEF, hereafter referred to as Cysts, are two RhoGEFs that play important roles in early fly development. During ventral furrow formation, RhoGEF2 directs apical actomyosin accumulation and apical constriction28,29. During cephalic furrow formation, RhoGEF2 is a major driver and Cysts is a minor driver of cell shortening30. During germband extension, RhoGEF2 mediates apical contractility while Cysts mediates junctional contractility31,32. Thus, different RhoGEFs have overlapping and distinct roles in embryogenesis, and it remains unclear to what extent a single RhoGEF can direct a range of morphogenetic behaviors on its own.
Optogenetics has emerged as a valuable tool to precisely manipulate spatiotemporal patterns of Rho signaling and contractile stresses33,34. Optogenetic tools perturbing the Rho pathway have produced significant insight into embryogenesis by facilitating perturbation of developmental events and reconstitution of some morphogenetic processes26,35,36. However, existing transgenic tools can be limited by variable expression levels, which hinder their quantitative use, as well as by potential artifacts of transgenic overexpression, which can unintentionally perturb development or compromise embryonic viability in some experimental conditions26.
Here, we develop two new optogenetic tools based on the tagging of Drosophila RhoGEF2 and Cysts at their endogenous loci with a component of the light-activated iLID/SspB heterodimer, producing viable embryos and healthy flies. We present an activation quantification strategy and uncover a dose-dependent relationship between RhoGEF recruitment and epithelial furrowing and bending. We find that at the onset of gastrulation, furrow asymmetry and furrow size constraints are influenced by a stiff basal actomyosin network, and that disrupting this basal network via scraps RNAi alters these mechanical behaviors. Finally, we demonstrate the extended applicability of the endogenous OptoRhoGEFs by manipulating distinct epithelial structures during multiple stages of embryonic development. Collectively, our results establish a framework for the precise optogenetic manipulation of tissue stresses and structures to interrogate biophysical mechanisms of morphogenesis.
Results
Development and characterization of endogenous OptoRhoGEFs
We engineered an optogenetic system into Drosophila melanogaster based on the LOV2-derived iLID/SspB molecular switch, in which light exposure causes an iLID conformational change that promotes its heterodimerization with SspB34. We localized the iLID component to the cell membrane and attached the SspB component to a RhoGEF, so that upon blue light exposure, the RhoGEF is targeted to the membrane, where it locally activates the Rho signaling pathway and induces cellular contractility (Fig. 1a). In contrast to existing transgenic optogenetic tools targeting Rho signaling26,35,36, here we used CRISPR/Cas9 to tag endogenous RhoGEF2 and Cysts loci with a SspB peptide, and optionally, a tagRFP-T sequence37, producing RhoGEF2-tgRFPt-SspB, RhoGEF2-SspB, SspB-tgRFPt-Cysts, and SspB-Cysts constructs38. The resulting flies were homozygous-viable. Co-expression of the SspB-tagged RhoGEFs with existing fluorescently-tagged RhoGEF2 or Cysts demonstrates that in the absence of optogenetic manipulation, the SspB-tagged RhoGEFs show canonical localization patterns, with RhoGEF2 enriched at furrow canals during cellularization (Supplementary Fig. 1a), and both RhoGEF2 and Cysts enriched at cell-cell junctions during germband extension (Supplementary Fig. 1b, c). We then co-expressed the SspB-tagged RhoGEFs in the blastoderm with a transgenic plasma membrane-localized iLID construct - either UASp>Venus-iLID-CaaX, UASp>mCherry-iLID-CaaX, or UASp>iLID-CaaX, depending on the fluorescent structures being visualized. When an SspB-bound GEF and an iLID-CaaX construct are co-expressed, they form the complete optogenetic tool, which we refer to here as OptoGEF2 and OptoCysts.
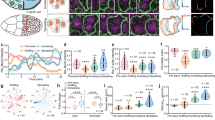
a Schematic: upon absorbing one 488-nm photon or two 927-nm photons, a membrane-bound iLID peptide undergoes a conformational change that promotes its binding to a GEF-bound SspB peptide, recruiting the GEF to the cell membrane. b Schematic: lateral RhoGEF recruitment is expected to induce apical-basal shortening and formation of a shallow furrow. c, d Two-photon lateral activation response of embryos co-expressing transgenic UASp>mCherry-iLID-CaaX (visualized) and 2 copies of endogenous RhoGEF2-tgRFPt-SspB (OptoGEF2, c) or transgenic UASp>mCherry-iLID-CaaX (visualized) and 2 copies of endogenous SspB-tgRFPt-Cysts (OptoCysts, d). White arrows in (c) highlight in-plane constriction behavior. e, f Two-photon lateral activation of embryos co-expressing UASp>Venus-iLID-CaaX, Sqh-mCherry (a mCherry-tagged Drosophila myosin II regulatory light chain, visualized), and 2 copies of endogenous RhoGEF2-SspB (OptoGEF2, e) or 2 copies of endogenous SspB-Cysts (OptoCysts, f). g, h Furrow depth-matched lateral activation experiments, where cells shorten by 10 µm, in OptoGEF2 (g) and OptoCysts (h) embryos (visualized with UASp>mCh-iLID-CaaX). i Schematic: Lateral activation results in persistent apical-basal shortening in both OptoGEF2 and OptoCysts embryos, but transient circumferential constriction only in OptoGEF2 embryos. For in-plane cross-sections, anterior is left and posterior is right. For transverse cross-sections, the apical tissue surface is oriented up and the basal surface is oriented down. For display in figure, brightness and contrast were adjusted between panels. Cyan outlines represent activation regions. In transverse views, yellow triangles mark slices shown in in-plane views. Scale bars are 10 µm.
OptoGEF2 and OptoCysts embryos are homozygous-viable. When cultured in the dark at 23 °C, the viability of OptoGEF2 embryos (0.87) was not significantly different from wild-type OreR flies (0.83; p = 0.18) and was significantly improved compared to an existing transgenic tool overexpressing RhoGEF2 catalytic DH & PH domains, hereafter referred to as RhoGEF2(DHPH)-Cry235 (0.38; p < 10–25) (Supplementary Fig. 2a). This improved viability represented a large increase in the fraction of embryos that reached cellularization (0.98 vs 0.50; p < 10–29) and a moderate increase in the fraction of cellularized embryos that ultimately hatched (0.89 vs 0.77; p = 0.003) (Supplementary Fig. 2b, c). OptoGEF2 embryos expressing mCh-iLID-CaaX and two copies of endogenously tagged RhoGEF2-tgRFPt-SspB successfully completed ventral furrow formation, cephalic furrow formation, and germband extension (Supplementary Movie 1). In the dark at 23 °C, OptoGEF2 embryos displayed myosin localization similar to wild-type (Supplementary Fig. 1j, k). This highlights that maintaining wild-type RhoGEF levels is important for experimentation in a healthy developmental background and demonstrates the utility of the endogenous OptoRhoGEFs in manipulating developmental mechanics. SspB-tagged RhoGEF2 and Cysts levels were highly consistent across embryos, displaying relative fluctuations of only 10–15%, in contrast to the UAS-driven iLID construct, which exhibited relative fluctuations of 45% (Supplementary Fig. 1g, h). These factors make endogenously tagged RhoGEFs ideal for quantitatively manipulating Rho signaling using optogenetics.
One-photon (1 P) activation of OptoGEF2 or OptoCysts with 488-nm light illumination, either globally or locally, led to GEF relocalization to the plasma membrane (Fig. 1a, Supplementary Fig. 3b, c). However, to achieve subcellular recruitment, a two-photon (2 P) activation protocol is required35,39. To overcome the inefficient 2 P absorption of common optogenetic switches, such as iLID/SspB40, we developed a strategy in which strong optogenetic recruitment can be driven by an inefficient activation process if the light-sensitive iLID component is expressed in excess (Supplementary Fig. 3a). We direct the interested reader to the Supplementary Material to learn more about this approach and other considerations in designing these tools (Supplementary Fig. 3, Supplementary Note). This 2 P activation approach permitted OptoGEF2 z-specificity of 4.2 ± 0.3 µm, similar to RhoGEF2(DHPH)-Cry235 (3.9 ± 1.4 µm; p = 0.67) (Supplementary Fig. 2d–h).
Lateral RhoGEF recruitment causes tissue furrowing via cell shortening
Based on previous work employing the transgenic optogenetic tool RhoGEF2(DHPH)-Cry230,41, we hypothesized that 2 P activation of a lateral cell volume would recruit endogenous RhoGEF2 or Cysts to lateral cell interfaces, driving cell apical-basal shortening and producing a shallow furrow (Fig. 1b). Here, we define shortening as a reduction in the distance between the apical and basal cell surfaces, and furrowing as an overall movement of cells toward the basal surface of the epithelium, producing a pocket on the apical surface. This is distinct from invagination, which additionally requires cells to be internalized. We note that gastrulation in Drosophila and other animals requires furrowing and invagination toward the embryo interior.
We performed 927-nm lateral activation in embryos finishing cellularization, when cells were 32-35 µm long. We chose to experiment in embryos undergoing the blastula-to-gastrula transition because this is concomitant with the initiation of ventral and cephalic furrow formation, giving insight into gastrulation-stage epithelial mechanics. To avoid mechanical interference from the ventral furrow, we activated the dorsal side of the embryonic primary epithelium. In this tissue, RhoGEF2-tgRFPt-SspB and SspB-tgRFPt-Cysts both display cytoplasmic localization, with RhoGEF2-tgRFPt-SspB additionally showing some localization at the cellularization front (Supplementary Fig. 1d–f).
Activation resulted in cell shortening and tissue furrowing in both OptoGEF2 and OptoCysts embryos, with a stronger response in OptoGEF2 (Fig. 1c, d, Supplementary Movie 2). Activation of control embryos expressing only the iLID component of the optogenetic tool did not induce cell shape changes or myosin reorganization (Supplementary Fig. 4). Shortening was also induced by 800, 850, or 900-nm activation (Supplementary Fig. 5). During Drosophila development, selective employment of RhoGEF2 frequently requires both its release from microtubules (MTs) and its anchoring to a new subcellular membrane compartment27,29,42,43. Our results suggest that neither explicit release of RhoGEF2 from MTs nor relief of RhoGEF autoinhibition23 are required to elicit a mechanical response, and that providing membrane anchoring sites is sufficient to promote relocalization and contractile activity of either RhoGEF2 or Cysts.
Although activation of either OptoGEF2 or OptoCysts induced persistent basal-directed cell shortening, the two tools displayed different transient cell deformation states. At 1 min post-activation, OptoCysts embryos showed straight lateral interfaces, while OptoGEF2 embryos showed increased lateral curvature and unexpected circumferential constriction at a sub-nuclear plane, ~22 µm below the apical surface (Fig. 1c, d, lower middle panels). This temporarily produced cells with an hourglass-like lateral contour only in OptoGEF2 embryos (Fig. 1c, d, lower middle panels). Comparing tissue behaviors in OptoGEF2 and OptoCysts embryos that both formed a 10-µm deep furrow, circumferential constriction was only seen in OptoGEF2, suggesting that the phenotype was not due to the strength of the mechanical response (Fig. 1g, h, Supplementary Movie 2). Imaging nonmuscle myosin II (hereafter referred to as myosin), we found that Cysts recruitment resulted in accumulation of myosin at lateral cell-cell interfaces, as expected, while RhoGEF2 recruitment additionally organized myosin into ring-like structures at the basal ends of nuclei (Fig. 1e, f, upper middle panels). At two min post-activation, OptoCysts myosin recruitment is limited to lateral interfaces, while OptoGEF2 embryos have a distinct additional accumulation of myosin near the basal cell surface. (Fig. 1e, f, lower right panels).
To investigate the use of our endogenous optogenetic tools in alternative furrowing mechanisms, we also attempted to reconstitute apical constriction-based furrowing by selectively illuminating apical cell surfaces, as has been done with transgenic tools based on GEF catalytic domains26,30,35 (Supplementary Fig. 6a). In OptoGEF2 embryos, we found that apical activation caused localized accumulation of myosin at apical caps (Supplementary Fig. 6b), resulting in weak apical constriction and formation of a shallow furrow at the apical surface (Supplementary Fig. 6c). However, this was accompanied by cell shortening and lateral expansion rather than basal expansion, inconsistent with traditional pictures of apical constriction-driven furrowing.
Overall, lateral recruitment of either RhoGEF2 or Cysts leads to lateral actomyosin accumulation, cell shortening, and tissue furrowing. However, only RhoGEF2 recruitment induces transient circumferential constriction (Fig. 1i).
RhoGEF-driven cell shortening is dose-dependent
To dissect the mechanisms that specify the depth of an epithelial furrow during development, we asked whether we could precisely tune tissue furrowing by modulating the extent of OptoGEF2 or OptoCysts activation. To do this, we developed a pipeline to specify an input optogenetic dose and measure the resulting tissue furrowing response. After acquiring images of the unperturbed tissue (Fig. 2a), we scanned an IR laser (0–75 mW) for 40 s over a cylindrical lateral cell volume, 18 µm in diameter and extending from 5–25 µm below the apical cell surface (Fig. 2a’), activating the iLID in a 2 P process. We observed the tissue response for ~5 min (Fig. 2a”), up until the dorsal tissue started to be compressed by the invaginating posterior midgut. We then segmented the apical and basal surfaces of the tissue to extract relevant quantitative metrics of the furrowing response (Fig. 2a”’).
a Representative stills from each step of a lateral activation experiment, demonstrated in an OptoGEF2 embryo. After acquiring images of the unperturbed tissue (a), a 927-nm laser is scanned across a cylindrical cell volume measuring 18 µm in diameter and extending 5–25 µm beneath the apical tissue surface (a’). Tissue response to activation is recorded (a”) and quantified via segmentation of the induced furrow (a”’, blue region represents furrow segmentation). In all steps, UASp>mCh-iLID-CaaX is the fluorescent membrane marker and scale bars are 10 µm. b, b’, b” Representative traces of OptoGEF2 embryo responses to different amounts of iLID activation. Upper curve represents the average position of the apical tissue surface within the activation region; bottom curve represents the average position of the basal surface within the activation region. Cyan region represents the 40-s activation step. b Weak activation (activation magnitude = 0.15). b’ Moderate activation (activation magnitude = 0.46). b” Strong activation (activation magnitude = 1.29). c Region-averaged cell shortening as a function of activation magnitude. Activation magnitude was measured using the emitted fluorescence of the UASp>mCherry-iLID-CaaX fusion construct during 927-nm activation (shown in a’). d Maximum cell shortening at any point in the activation region versus the region-averaged cell shortening. e Maximum shortening of the basal surface versus maximum shortening of the apical surface. f Coefficient of determination of the linear fit to the combined OptoGEF2 & OptoCysts dataset as a function of the scaling factor applied to Cysts activation magnitudes (peak at scaling factor of 0.293). g OptoCysts/OptoGEF2 activity ratio (measured in f) and SspB-tgRFPt-Cysts/RhoGEF2-tgRFPt-SspB abundance ratio (measured in Supplementary Fig. 1g). Source data are provided as a Source Data file.
Based on the low variability in RhoGEF2 and Cysts expression levels (Supplementary Fig. 1h), we reasoned that the strength of the tissue response should be determined by the optogenetic dose we deliver—that is, by the number of iLID molecules that we photoconvert. This depends on two factors - the iLID protein level, which sets the base amount of inactive iLID molecules, and the photon flux passing through the illuminated cell volume, which depends on the activating laser power and determines what proportion of these iLID molecules are activated. For a 2 P activation process, we expect the activation dose to scale quadratically with the activating laser power and linearly with the iLID abundance. Both of these factors are captured quantitatively in the fluorescence emitted by the mCh-iLID-CaaX construct during our 40-s activation step, as mCherry is weakly excited by 927-nm light in a 2-photon process (Fig. 2a’). We thus use the relative iLID fluorescence from this activation step as a measure of activation magnitude, allowing us to compare tissue responses to different amounts of activation in different experiments and across embryos. This relies on the assumption that we are not saturating iLID activation, which holds here (Supplementary Fig. 3, Supplementary Note).
First, we examine the average positions of the apical and basal tissue surfaces within the tracked activation region over time (Fig. 2b-b”). Note that cells continue elongating by several microns until cellularization is fully completed, so that the final position of the basal surface is deeper than the initial position. Cellularization rates were the same between control, OptoGEF2, and OptoCysts embryos (Supplementary Fig. 1i), and were all consistent with wild-type cellularization rates44, indicating that differential cellularization rates do not play a role in tissue shape changes. We found that weak activation caused shallow, transient cell shortening (Fig. 2b), moderate activation elicited stronger, longer-lasting shortening (Fig. 2b’), and strong activation produced deep furrows that persisted through the end of the observation period (Fig. 2b”). This suggests that at sufficiently strong levels of activation, cell shortening can be converted into stable tissue furrowing.
Then, for each experiment, we extracted the maximum amount of cell shortening, calculated as the largest difference between instantaneous cell height and reference cell height based on embryo-specific cellularization rates. Applying this analysis, we found a striking grading of cell shortening response across a broad range of optogenetic doses for both OptoGEF2 and OptoCysts (Fig. 2c). We note that our OptoCysts lines expressed the iLID component at higher levels than our OptoGEF2 lines, allowing us to sample higher activation magnitudes with the Cysts tool. Previous studies modulating embryonic RhoGEF2 levels have shown it to be a dose-dependent regulator of myosin activity27,45,46, but RhoGEF2 overexpression tends to disrupt cell and tissue morphogenetic processes rather than enhance them27,45. Here, we report a dose-dependence of a tissue-scale, rather than molecular-scale, morphogenetic feature on levels of RhoGEF2 or Cysts activity. This dose-dependence suggests that although positive feedback likely amplifies RhoGEF activity in this context, it does not saturate it or cause switch-like behavior.
The shortening response was not uniform over the whole activation region, resulting in the maximum shortening within the region being slightly greater than the average shortening (Fig. 2d). Consistent with our qualitative observations in Fig. 1c, d, we found that shortening from the apical surface dominated the response, with shortening from the basal surface playing only a minor role (Fig. 2e). Using our dose-response curves, we calculated an activity ratio of OptoCysts compared to OptoGEF2 by finding the scaling factor which, when applied to the OptoCysts activation magnitudes, minimized the residuals to the joint OptoGEF2-OptoCysts shortening vs. activation magnitude linear fit (Fig. 2f). This activity ratio represents the relative dose-response sensitivity of the two tools and depends on the expression levels of the two RhoGEFs, their relative catalytic activities, and any potentially differential steric hindrance imposed by the optogenetic tags. We calculated an activity ratio of 0.29 (Fig. 2g), which is comparable to our measured Cysts/RhoGEF2 protein level ratio of 0.40 (Fig. 2g, Supplementary Fig. 1g), and to a Cysts/RhoGEF2 abundance ratio of 0.37 extracted from an embryo proteomics dataset47, suggesting that differences in tool activity can largely be explained by differences in cellular levels of the two GEFs.
Our dose quantification approach is validated by three additional observations. First, at constant activating laser power, shortening scales with iLID levels (Supplementary Fig. 7a). Second, shortening behavior is much better accounted for by our activation magnitude parameter, which combines information about activating laser power and iLID levels, than by activating laser power alone (Supplementary Fig. 7b, c). Finally, when we calculate our dose using an alternative measure, defined as the laser power squared times the relative iLID protein level, we also find dose-dependent shortening behavior (Supplementary Fig. 7d) and linear scaling with our activation magnitude parameter (Supplementary Fig. 7e). In alternative experimental setups where iLID fluorescence during activation can’t be measured, this alternative approach may prove useful.
Overall, our endogenous optogenetic tools allowed us to uncover a dose-dependent relationship between RhoGEF recruitment and furrow depth. This suggests that during development, furrow morphologies can be specified by modulating the amount of RhoGEF membrane anchoring sites. This is true of both RhoGEF2 and Cysts, suggesting that these two RhoGEFs overlap at least partially in the tissue deformations they are able to drive.
Activation dose modulates the amount of RhoGEF recruited to the membrane
Although we found that furrow depth scales with our administered optogenetic dose, it is not obvious how this dose translates to actual changes in RhoGEF activity or localization. An unintuitive feature of a non-equimolar optogenetic switch in which the light-sensitive component is in excess (here, iLID) is that the timescale of SspB unbinding becomes decoupled from the timescale of iLID dark-state reversion48. This is apparent via both an equilibrium argument, in which additional activated iLID beyond the amount required for near-full SspB translocation acts to increase the duration of full SspB recruitment, as well as a kinetic argument, in which excess unbound iLID at the membrane can trap SspB after it unbinds from an iLID molecule but before it diffuses into the cytoplasm48. In OptoGEF2 and OptoCysts embryos, total iLID levels are in excess of RhoGEF levels, as described above. However, given the inefficiency of the 2 P activation process40 (Supplementary Fig. 3), it’s uncertain if activated iLID is also in excess. Thus, depending on whether the RhoGEF or the active iLID is in excess, an increase in optogenetic dose will lead to either more RhoGEF molecules being recruited, or a roughly constant number of RhoGEF molecules being recruited for an increased duration (Fig. 3a).
a Schematic: depending on the relative abundance of active iLID and SspB-bound RhoGEF, an increase in the number of activated iLID molecules can result in either increased magnitude or increased duration of GEF recruitment. b Shortening vs. activation magnitude in response to lateral activation of embryos laid by mothers expressing UASp>mCh-iLID-CaaX and either heterozygous or homozygous for RhoGEF2-tgRFPt-SspB. c, d Furrowing timescale as a function of either activation strength (c) or final furrow depth (d). e, f Recovery timescale as a function of either activation strength (e) or furrow depth (f). Furrowing and recovery timescales are defined as the time at which the tissue reaches 1/e = 37% of its final shortened or recovered height. P values in (e, f) are calculated using a two-tailed t test against the null hypothesis that the relationship between recovery time and activation magnitude (e) or recovery time and furrow depth (f) is the same for OptoGEF2 and OptoCysts embryos. Source data are provided as a Source Data file.
To dissect this, we measured the timescales of tissue furrowing and tissue recovery, the latter of which we could only quantify in furrows that relaxed. Surprisingly, we found that furrowing time was independent of the GEF activated, the activation magnitude, or the ultimate furrow depth, taking an average of 70 s across all experiments (Fig. 3c, d). The recovery time, on the other hand, did scale with activation magnitude and with furrow depth (Fig. 3e, f). Notably, the OptoGEF2 and OptoCysts recovery times only collapsed onto a single curve when plotted versus furrow depth, suggesting that the timescale of recovery is better explained by the final mechanical state of the furrow than by iLID reversion kinetics.
We argue that the optogenetic dose modulates the amount of RhoGEF at the membrane based on the following observations. First, if the behavior of the tissue is elastic-like, with a given amount of membrane-bound GEF corresponding to a given cell height strain, then we would only observe the dose-dependent furrow depth seen in Fig. 2c if our dose modulates the amount of GEF. Second, if the behavior of the tissue is viscous-like, with a given amount of membrane-bound GEF driving a given cell height strain rate, then we would only observe the constant furrow time seen in Fig. 3c if our dose changed the magnitude, rather than the duration, of GEF recruitment. Thus, in either case, our quantitative measurements are more consistent with the dose modulating the amount of membrane-bound GEF. This interpretation is supported by the furrow recovery timescales in Fig. 3f being mechanically determined and is consistent with, though not proven by, the observation that embryos heterozygous for RhoGEF2-tgRFPt-SspB showed a weaker dose-dependent shortening response than their homozygous counterparts (Fig. 3b). Although this is the most parsimonious interpretation, we cannot rule out potential complexities arising from biochemical feedback or endogenous protein localization patterns in determining the timescale or strength of the response. In summary, our optogenetic dose tunes furrow depth by changing the amount of RhoGEF2 recruited to the cell membrane, and furrow recovery dynamics scale with the depth of the induced furrow.
Large regions of lateral Rho activation are associated with tissue bending
Comparing the wide, apical constriction-driven ventral furrow to the narrow, lateral contraction-driven cephalic furrow, we wondered how lateral contraction scaled as a furrowing mechanism in the primary epithelium. To explore this, we varied activation region diameters up to 50 µm. Strikingly, we found that while 18-µm regions underwent coherent shortening toward the basal surface, eventually forming a furrow (Fig. 4a), 50-µm regions instead produced highly curved tissues that predominantly shortened toward the apical surface and failed to form a productive furrow (Fig. 4c, Supplementary Movie 3). This phenotype was also observed in OptoCysts embryos (Supplementary Fig. 8a), although bending was not as strong. At intermediate region sizes, some tissues showed strong bending, but were still able to undergo net basal-oriented shortening to produce a furrow (Fig. 4b). Activation of a single lateral interface resulted in shortening of the two cells sharing the interface and tissue furrowing toward the basal surface (Supplementary Fig. 8b), similar to previous work30. Cellular myosin patterns in response to OptoGEF2 activation were the same in 50-µm activation regions as in 18-µm activation regions, characterized by initial lateral myosin accumulation, transient subnuclear contractile rings, and persistent basal myosin (Fig. 1e, Supplementary Fig. 8c). Bent tissues showed a unique behavior in apical slices, with a central tissue plug surrounded by acellular perivitelline space (Fig. 4d). We used this behavior to classify embryos as either bending or non-bending. However, we did observe a continuous range of tissue morphologies from flat to curved, so bending behavior was not binary (Fig. 4e). Plotting bending phenotype as a function of activation region size and activation magnitude, we found that larger activation regions required smaller amounts of activation to exhibit bending (Fig. 4f).
a–c Lateral activation response of an 18-µm region (a), a 37-µm region (b), and a 50-µm region (c). Activation magnitudes are 1.29 (a), 1.39 (b), 1.02 (c). d In-plane view of the bending response of the embryo shown in (c). e Apical slice of an activated tissue exhibiting an intermediate bending phenotype (50 µm activation region diameter, activation magnitude = 0.36). f Tissue bending phenotype as a function of the size of the activation region and the strength of activation. Bending is classified using the behavior shown in (d). g Schematic: tissue curvature can arise from lateral contraction via either cell-intrinsic basal stiffness (left) or cell-extrinsic compressive forces (right). h Shortening vs activation magnitude for small and large activation regions (p = 0.73, two-tailed t test against the null hypothesis that the relationship between cell shortening and activation magnitude is the same for 18-µm and 50-µm activation regions). i, j An 840-nm laser was used to activate a 52-µm cylindrical region and then perforate a neighboring portion of the vitelline membrane, relieving hydrostatic pressure and tissue compression via outward-driven tissue flow. Resulting tissue response is shown in in-plane (i) and transverse (j) slices. Yellow dashed line in (i) marks orientation of orthogonal slice shown in (j). k, l Activation of a 45 × 15 µm elliptical prism, 5 to 25 µm below the apical surface, oriented either along the AP (k) or DV (l) axis. In all images, UASp>mCh-iLID-CaaX is the membrane marker. For display in figure, brightness and contrast were adjusted between panels. Cyan outlines represent activation regions. In transverse views, yellow triangles mark slices shown in in-plane views. Scale bars in (a–e, j, k, l) are 10 µm. Scale bars in (i) are 20 µm. Source data are provided as a Source Data file.
We considered two mechanisms by which lateral contractility could produce curved tissues (Fig. 4g), both relying on volume conservation-based coupling between cell shortening and cell area expansion49. In one model, tissue-intrinsic basal stiffness induces bending by restricting basal area expansion, producing wedge-shaped cells and net tissue curvature50. In another model, tissue area expansion pushes against surrounding non-activated cells and the resulting external compressive force bends the activated tissue via a buckling or wrinkling instability51,52. The former model predicts curvature from tissue-intrinsic forces, while the latter model predicts curvature from tissue-extrinsic forces.
To distinguish between these potential mechanisms, we first note that curvature was not the result of a wrinkling instability, as activation regions measuring from 25-75 µm in diameter always resulted in only a single tissue crest, indicating that the bending pattern was not spatially periodic (Fig. 4f, Supplementary Fig. 8d). Moreover, cell heights were equal in tissue grooves and crests, inconsistent with the groove-crest symmetry-breaking predicted during a wrinkling instability52 (Supplementary Fig. 8e, f). In a buckling-type process, we expect a buildup of in-plane compressive stress during cell shortening that abruptly transitions to out-of-plane curvature once bending becomes energetically favorable. Here, we found that the scaling of maximum cell shortening with activation magnitude was equivalent in 18-µm and 50-µm regions (Fig. 4h) and that tissues displayed a continuous range of bending behaviors (Fig. 4e), both of which are inconsistent with a simple buckling process. Finally, relieving compressive stress and hydrostatic pressure via IR laser vitelline membrane micro-perforation30 did not relax tissue curvature, further suggesting that bending is a tissue-intrinsic process (Fig. 4i, j, Supplementary Movie 4).
A prediction of outward bending being driven by a tissue-intrinsic mechanism is that lateral activation of an elliptical region would result in tissue bending along the long (major) axis of the ellipse. On the other hand, if bending in large regions arises from tissue-extrinsic compression, the elliptical region would tend to furrow normally, as compression along the major axis could be relieved by expansion along the minor axis. Thus, we activated a 45 × 15 µm ellipse, oriented either along the embryo AP or DV axis, noting that the dorsal epithelium has higher intrinsic curvature along its DV, compared to its AP, axis (Fig. 4k, l). Consistent with tissue-intrinsic bending, AP-oriented ellipses resulted in an outward bending phenotype similar to that observed in 50-µm circular activation regions (Fig. 4c, k). Interestingly, DV-oriented ellipses instead showed a phenotype that appears more similar to the furrowing observed in 18-µm circular activation regions (Fig. 4a, l). Upon investigation of the shape of the activated tissue, we saw that the DV-oriented ellipses resulted in activated regions that are curved outward along the DV axis (similar to the tissue-intrinsic prediction), but the curvature of these activated regions tracked the intrinsic curvature of the embryo, so that it did not induce obvious outward bulging. Thus, the behavior of the DV-oriented activation ellipses suggests that innate embryo curvature may allow for large groups of cells to furrow without strong outward bending, as long as the region of cells is oriented along an axis of high embryo curvature.
Overall, we found that large regions of lateral Rho activation displayed disrupted furrowing behavior, and instead exhibited a bending phenotype. Our observations rule out wrinkling and buckling as drivers of tissue curvature, and are consistent with bending being tissue-intrinsic. This prompted us to consider that basal tissue stiffness may be the source of tissue bending in our experiments.
Perturbing basal mechanics alters tissue furrowing and bending
During these studies, we identified multiple tissue behaviors that seemed consistent with the presence of a mechanically stiff basal layer in the primary epithelium - that is, a basal layer that resists deformation. These included preferential cell shortening toward the basal surface for small activation regions (Fig. 2e) and tissue bending for large activation regions (Fig. 4c). Both behaviors persisted when the activation depth was changed from 5–25 µm to 5–10 µm below the apical surface (Supplementary Fig. 8g–j), ruling out off-target basal activation via two-photon axial spreading53 or iLID-CaaX diffusion48 as their source. Given that Drosophila cellularization is associated with a stiff, actomyosin-rich basal front54,55, with a recent cantilever-based study reporting a large stiffness differential between apical and basal cell surfaces at mid-cellularization56, we hypothesized that mechanical perturbation of the basal tissue surface would alter activation-induced tissue shape changes.
To test this, we repeated these experiments in a scraps/anillin (scra) RNAi background, in which cellularization is perturbed, characterized by destabilized yolk stalks, improper actomyosin organization and reduced myosin levels at basal furrow canals, vesiculation of lateral membranes, and eventual disintegration of the epithelium after initial gastrulation movements57,58,59. By the onset of gastrulation, rather than forming a planar sheet with narrow, circular yolk stalks, basal myosin II instead accumulates into segmented, bar-like structures at cell-cell interfaces57 (Fig. 5a). This is thought to arise from a loss of Scraps/Anillin-mediated recruitment of septins to furrow canals, resulting in a failure of the basal actin network to organize into circular, tightly packed filaments58. Laser ablation of the basal tissue surface in OptoGEF2 and OptoGEF2 + scra RNAi embryos revealed that the presence of scra RNAi reduced the initial retraction distance (velocity) by 62%, indicating perturbed basal mechanics and suggesting decreased basal tension in the tissue (Supplementary Fig. 9a–d). This disrupted basal actomyosin organization makes scra RNAi embryos a useful system for exploring the role of basal tissue mechanical properties in directing morphogenetic behaviors. Thus, we predicted that furrowing direction bias and bending behavior would both be reverted by scra RNAi treatment.
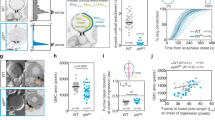
a Basal myosin II at the onset of gastrulation in unactivated OptoGEF2 + scra RNAi embryos (left) and unactivated OptoGEF2 control embryos (right). b Schematic: we hypothesize that a stiff basal layer is required to properly orient cephalic furrow formation and to translate cell shortening into productive tissue furrowing. c, d Two-photon lateral activation of an 18-µm diameter region in embryos expressing OptoGEF2 + scra RNAi (c) or OptoGEF2 alone (d). e, f Two-photon lateral activation of a 50-µm region in embryos expressing OptoGEF2 + scra RNAi (e) or OptoGEF2 alone (f). g, h Cephalic furrow formation in embryos expressing OptoGEF2 + scra RNAi, with myosin (g) or membrane (h) visualized. i Cephalic furrow formation in unactivated control embryos expressing OptoGEF2 alone. In (c–f, h, i), UASp>mCh-iLID-Caax is the membrane marker. In (a, g), myosin is visualized with Sqh-mCh. For display in figure, brightness and contrast were adjusted between panels. Cyan outlines represent activation regions. White and cyan arrows are guides to the eye. In transverse views, yellow triangles mark slices shown in in-plane views. Scale bars are 10 µm.
Consistent with our predictions, small lateral activation regions in OptoGEF2 + scra RNAi embryos caused shortening from the cell basal surface instead of from the apical surface, which was seen in embryos expressing OptoGEF2 alone (Fig. 5c, d). Large activation regions in OptoGEF2 + scra RNAi embryos caused shortening from the basal surface, but no overall tissue curvature, as demonstrated by the absence of the apical indentations that characterized bending in OptoGEF2 embryos (Fig. 5e, f). This suggests that a basal actomyosin layer is required for productive cell furrowing in response to lateral activation (Fig. 5b) and is the source of the bending phenotypes observed in large activation regions (Fig. 4g, left).
To understand how the disrupted basal layer in scra RNAi embryos affects lateral contraction-driven morphogenesis, we imaged cephalic furrow dynamics in OptoGEF2 + scra RNAi embryos without any activation. Basal myosin in cephalic furrow initiator cells showed the same segmented structure as in the rest of the embryo, suggesting that basal mechanical properties are also perturbed in these cells (Supplementary Fig. 9e). Notably, while lateral actomyosin still accumulated in cephalic furrow initiator cells (Fig. 5g), it now frequently caused these initiator cells to shorten from their basal surfaces rather than their apical surfaces (Fig. 5h, i, Supplementary Movie 5). This was not a result of broadly disrupted epithelial morphogenesis, as ventral furrow formation still occurs normally in scra RNAi embryos59, suggesting that the perturbed phenotype is particular to mechanisms driving cephalic furrow formation. Nonetheless, we cannot rule out potential contributions of other effects of scra RNAi, particularly the disruption of lateral cell membranes, toward these phenotypes.
Overall, in both our optogenetic perturbations and endogenous cephalic furrow formation, productive lateral contraction-driven tissue furrowing towards the interior of the embryo requires a stiff basal layer. Although lateral contractility is the active force driving furrowing, passive basal stiffness seems to provide the symmetry-breaking mechanical cue that orients it. However, these basal mechanical properties have the additional effect of imposing a size constraint on lateral contraction-driven furrows, beyond which tissues bend instead of coherently furrowing. Here, bending obstructs productive furrowing in a size-dependent manner because large activation regions cause tissues to form an arc that subtends a larger angle, deviating more from an approximately flat morphology. Forcing the activated tissue to adopt this larger arc shape while still maintaining continuity with non-activated tissue along its peripheral basal surface causes the center of the activated region to bulge toward the embryo exterior instead of furrowing toward the embryo interior. We found that this bending behavior disrupts the ability of large regions of lateral contraction to form a coherent furrow, imposing an effective size constraint on furrows formed by this mechanism. However, embryo curvature can modulate the impact of this size constraint by allowing large, curved furrows to track the intrinsic curvature of the embryo.
Endogenous OptoRhoGEFs in other developmental contexts
We wanted to evaluate the utility of the endogenous OptoRhoGEFs for studying a range of developmental processes during Drosophila embryonic development, beyond reconstituting furrowing in the primary epithelium during the blastula-to-gastrula transition at late stage 5/early stage 6. To do this, we performed one-photon optogenetic perturbations in various tissues of OptoGEF2 embryos at developmental stages from cellularization to dorsal closure.
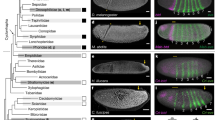
First, we performed optogenetic perturbations to the forming primary epithelium during early cellularization (early stage 5). Consistent with previous studies60, local activation of the cellularizing primary epithelium, when cells were 5 µm tall, resulted in constriction of activated cells, which partially relaxed upon cessation of photoactivation (Fig. 6a, b, Supplementary Movie 6). Second, we performed optogenetic perturbations of the amnioserosa later in development during dorsal closure (stage 14). Activation of the entire amnioserosa resulted in rapid amnioserosa contraction, decreased dorsal opening width, and loss of cell area oscillations, all of which were reverted after photoactivation (Fig. 6c, d, Supplementary Movie 7). Third, we performed optogenetic perturbations of the lateral epidermis during late dorsal closure (stage 15). Activation of the left and right lateral epidermis resulted in stretching of amnioserosa cells and increased dorsal opening width, which reverted after activation (Fig. 6e, f, Supplementary Movie 8). Fourth, we tested single-cell perturbations in the amnioserosa during dorsal closure. Activation of a single cell resulted in reduced cell area and contraction of the tissue toward that cell (Fig. 6g, Supplementary Movie 9), consistent with previous studies61. Finally, we demonstrate arbitrary, rapid, reversible deformations of the embryo during dorsal closure in response to a 5-step activation protocol that targets multiple embryo regions (Supplementary Movie 10).
a 1 P photoactivation of a 30-µm circular region in the cellularizing primary epithelium, when cells were 5 µm tall. Images are 5-µm sub-apical slices, taken just before photoactivation (left), just after 7 min of continuous photoactivation (center), and after an additional 7 min of recovery (right). b Area change of the cyan region shown in (a, left) over time. c 1 P photoactivation of the entire amnioserosa during early dorsal closure. Images are 4-µm apical maximum intensity projections, taken just before photoactivation (left), just after 12 min of continuous photoactivation (center), and after an additional 5 min of recovery (right). d Change in amnioserosa width along the DV axis over time for the experiment shown in (c). Distinct cyan shades represent different photoactivation areas, which were changed over the course of the experiment to track the boundaries of the amnioserosa as it contracted. e 1 P photoactivation of the left & right lateral epidermis during late dorsal closure. Images are 6-µm sub-apical slices, taken just before photoactivation (left), just after 5 min of continuous photoactivation (center), and after an additional 7 min of recovery (right). f Change in amnioserosa width along the DV axis for the experiment shown in (e) over time, measured via PIV. g 1 P photoactivation of a single cell (8-µm diameter circle) in the amnioserosa during early dorsal closure. Images are 3-µm apical maximum intensity projections, taken just before photoactivation (left), just after 3 min of photoactivation (center), and just after 3 min of photoactivation, overlaid with the PIV-derived velocity field (right). All experiments shown are in OptoGEF2 embryos. In (a, c, e, g), cyan outlines represent regions of photoactivation. In (b, d, f), t = 0 min corresponds to the initiation of photoactivation and shaded cyan regions represent the duration of photoactivation. For display in figure, brightness and contrast were adjusted between panels. Scale bars in (a, c, e, g) are 20 µm. Source data are provided as a Source Data file.
Taken together, our results demonstrate the utility of the endogenous OptoRhoGEFs in manipulating tissues across several stages of embryogenesis.
Discussion
Here, we developed two optogenetic tools based on the endogenous tagging of different RhoGEFs with a component of a light-activated heterodimer. We demonstrate the efficacy of these tools for the precise reconstitution and manipulation of diverse actomyosin-driven processes during Drosophila embryonic development. They add to a growing body of optogenetic tools permitting light-directed control of endogenous protein activity62,63,64.
The endogenous nature of these tools allowed us to accurately quantify our optogenetic input by accounting for expression variability of only the iLID component. In theory, a dose could be precisely quantified in any optogenetic system by measuring both component levels48, but this requires two dedicated fluorescence channels and is prohibitively difficult for optogenetics applications. Using this approach, we uncovered a dose-dependent relationship between RhoGEF recruitment and induced furrow depth (Fig. 2c). The roughly linear dose-response curve is surprising, given the potentially nonlinear kinetic and mechanical processes involved in the activation response.
Through our quantitative studies, we uncovered several key principles of lateral contraction-driven tissue morphogenesis. We found that furrows formed by this mechanism can have their morphologies specified by grading levels of RhoGEF membrane anchors, which acts by modulating the amount of membrane-bound RhoGEF (Figs. 2c, 3). However, patterning of tissue apical-basal mechanical properties plays a crucial role in determining the resulting tissue deformations. A stiff basal actomyosin layer, present at the onset of gastrulation, not only provides a symmetry-breaking cue to orient tissue furrowing (Fig. 5c, d), but also constrains the size of productive furrows (Fig. 5e, f). This stiff basal network is characteristic of cellularization56, but is a transient state in gastrulation and persists only for the first several minutes of ventral and cephalic furrow formation. Nonetheless, apical-basal asymmetry is a defining property of epithelia, and the mechanics described here are potentially representative of many epithelia with mature basement membranes. In other contexts, a stiff apical surface could direct furrowing behaviors, or buckling and wrinkling processes may emerge in the absence of an explicit symmetry-breaking cue.
OptoGEF2 produced unexpected cellular behaviors that were not predicted from a simple model of Rho activation. First, we observed that OptoGEF2, but not OptoCysts or RhoGEF2(DHPH)-Cry230, produced unexpected myosin localization patterns - namely, subnuclear circumferentially constricted rings (Fig. 1e, upper middle) and persistent basal myosin (Fig. 1e, lower right). This difference suggests that RhoGEF2-specific interactions, beyond those of its DH & PH domains, drive these behaviors. For example, persistent basal myosin may arise from PDZ domain-dependent RhoGEF2 binding to Slam, an anchor directing RhoGEF2 localization at the cellularization front65 (Supplementary Fig. 1a). The absence of circumferential constriction following lateral OptoCysts (Fig. 1f) or RhoGEF2(DHPH)-Cry230 recruitment suggests that it is not necessary for furrowing. Future studies using RhoGEF2-based optogenetic constructs with mutated PDZ, RH, and SxIP domains will be useful in determining the origin of the circumferential constriction phenotype.
Second, the weak apical constriction response induced by OptoGEF2 (Supplementary Fig. 6) is surprising given that RhoGEF2(DHPH)-Cry2 is able to elicit a strong apical constriction response35. This weak apical constriction in OptoGEF2 is reminiscent of Concertina mutants, in which RhoGEF2 is not released from MTs preceding ventral furrow formation, but still elicits a moderate residual level of apical constriction due to T48 anchoring at apical membranes29,42. In this context, active iLID plays a similar role to T48 - both redirect a non-sequestered subpopulation of RhoGEF2 to the apical surface, where they drive a small amount of apical constriction that is insufficient for robust tissue invagination. These observations suggest that OptoGEF2 may have some limitations associated with endogenous RhoGEF2 localization, full-length RhoGEF2 interacting partners, or RhoGEF2 autoinhibition. Even so, both OptoRhoGEFs were potent tools for manipulating cell and tissue-level contractile behaviors throughout embryonic development. Future studies with OptoRhoGEFs will help to elucidate regulatory principles of full-length RhoGEFs and their contribution to actomyosin patterning during development.
The ventral and cephalic furrows employ similar strategies to permit cell internalization past a stiff basal surface. Both the presumptive mesoderm and cephalic furrow initiator cells undergo precocious loss of basal actomyosin in order to couple cellular constriction to basal expansion and invagination12,30,66,67. Ectopic local stiffening of the basal tissue surface via optogenetics prevents both ventral and cephalic furrow internalization30,68. Our results suggest that preceding relaxation, cephalic furrow formation takes advantage of the stiff tissue basal surface to orient cell shortening toward the embryo interior (Fig. 5g, h). This may prime cells to invaginate once basal myosin is lost, either by stabilizing the shortened cells via plastic deformation69 or by overcoming an energy barrier associated with cell movement70,71. Alternatively, given that basal relaxation is sufficient to cause cell ingression in the primary epithelium30, that cell pressure differentials play major roles in determining cephalic furrow cell shapes72, and that hydrostatic pressure excess can enhance tissue folding past a compliant surface73, the role of the basal sheet may instead be to increase hydrostatic pressure in response to cell contraction.
Surprisingly, we find that lateral contraction does not scale well as a furrowing mechanism in the primary epithelium. When a large group of cells shortens at once, restricted basal area expansion causes tissues to adopt curved shapes, forcing central cells to move toward the vitelline membrane instead of toward the embryonic interior (Fig. 4c, d). Notably, in anisotropic regions with high lateral contractility, productive furrowing can still occur when the long axis of the group of cells is oriented along an axis of high curvature of the embryo (Fig. 4k, l). This suggests that embryo-wide curvature and geometric constraints may play a role in cephalic furrow formation. Although reminiscent of buckling processes in other epithelia and developing systems74,75,76,77, the bending process described here is distinct because it requires no external forces and exhibits no sudden transition from a non-buckled to a buckled state (Fig. 4e, h). This continuous change in curvature as a function of lateral contraction has been described theoretically50, but to our knowledge, has not been demonstrated experimentally. It will be interesting to re-examine wedged cell shapes and curved epithelial morphologies in embryos to see if this curvature mechanism is at play during development.
Endogenous optogenetic tools, combined with the dose quantification strategies we present here, provide a powerful toolbox to interrogate epithelial morphogenesis. We build upon the spatial and temporal control inherent to optogenetic systems by additionally permitting quantitative induction of tool activation and biomechanical responses. Our work lays a foundation for the precise modulation of contractile forces from the subcellular to tissue scales, allowing for careful interrogation of developmental mechanics and control of 3D shapes of engineered living systems and materials.
Methods
Cloning
To make pBI-UASp-mCherry-iLID-CaaX, pBI-UASp-Venus-iLID-CaaX, and pBI-UASp-iLID-Caax, Venus and iLID-CaaX were amplified from pLL7.0: Venus-iLID-CAAX (Addgene #60411, gift from Brian Kuhlman) and mCherry was amplified from pBI-UASp-mCherry-Cry2phr-RhoGAP71E36. Fragments were cloned into a pENTR/D-Topo vector (Life Technologies, Carlsbad, CA) using the NEBuilder HiFi DNA Assembly Kit (New England BioLabs, Ipswich, MA) and recombined into a UASp-attB destination vector (gift of F. Wirtz-Peitz) using the Gateway cloning system (Life Technologies, Carlsbad, CA).
To make pHD-RhoGEF2-tgRFPt-SspB-DsRed, pHD-RhoGEF2-SspB-DsRed, pHD-SspB-tgRFPt-DsRed-Cysts, and pHD-SspB-DsRed-Cysts, SspB and tgRFPt were amplified from pLL7.0: tgRFPt-SSPB WT (Addgene #60415, gift from Brian Kuhlman), DsRed was amplified from pScarlessHD-sfGFP-DsRed (Addgene #80811, gift from Kate O’Connor-Giles), and homology arms were amplified from genomic DNA of vas>Cas9 (III) (BDSC #51324). We used FlyBase (release FB2020_06) to obtain RhoGEF2 and Cysts homology sequences. Fragments were cloned into a pScarlessHD-sfGFP-DsRed donor vector (Addgene #80811, gift from Kate O’Connor-Giles) using the NEBuilder HiFi DNA Assembly Kit (New England BioLabs, Ipswich, MA).
Donor plasmid PAM sites were mutated using the QuikChange II XL Site-Directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA). The gRNA-encoding plasmids pU6-BbsI-chiRNA-RhoGEF2 and pU6-BbsI-chiRNA-Cysts were made by insertion of gRNA oligos into pU6-BbsI-chiRNA (Addgene #45946, gift from Melissa Harrison and Kate O’Connor-Giles and Jill Wildonger) using the Q5 Site-Directed Mutagenesis Kit (New England BioLabs, Ipswich, MA) or T4 DNA ligase (New England BioLabs, Ipswich, MA). The primers used to make the CRISPR constructs are listed in Supplementary Data 1.
Fly stocks
Drosophila melanogaster was cultured at 23 °C using standard techniques. Crosses were maintained in the dark at 23 °C. Imaging was done in the dark at room temperature (~21 °C). Crosses and embryos were handled in dark rooms with all light sources covered by a Red 25 Wratten Filter (Kodak, Rochester, NY).
iLID constructs under a UASp promoter were expressed with a maternal α-tubulin matα-tub15 Gal4-VP16 driver (mat15, gift from D. St Johnston). CRISPR/Cas9 knock-ins at RhoGEF2 and Cysts loci were done following the flyCRISPR method38 (https://flycrispr.org/). Guide RNAs were designed using the flyCRISPR target finder (http://targetfinder.flycrispr.neuro.brown.edu/). Homology donor and gRNA plasmids were co-injected into vas>Cas9 (III) (BDSC #51324) (BestGene Inc, Chino Hills, CA). After homology-mediated cassette insertion, the 3xP3-DsRed selectable marker was flipped out by crossing to nos-PBac (III)78. Transgenic iLID lines were generated by PhiC31-directed integration into an attP2 docking site (BDSC #8622) (BestGene Inc, Chino Hills, CA).
The fly stocks generated for this study were RhoGEF2-tgRFPt-SspB, RhoGEF2-SspB, SspB-tgRFPt-Cysts, SspB-Cysts, UASp>iLID-CaaX (III), UASp>Venus-iLID-CaaX (III), UASp>mCherry-iLID-CaaX (III). Other stocks used in this study were RhoGEF2>RhoGEF2-GFP (III)79, Cysts-GFP79, mat15 (III) (gift from D. St Johnston), mat67 (II) (gift from D. St Johnston), osk>Gal4 (III) (BDSC #44242), sqh>Sqh-mCh (II) (BDSC #99923), sqh>Sqh-mCherry (III) (BDSC #59024), UASp>CIBN-pmGFP (II)35, UASp>RhoGEF2(DHPH)-Cry2 (III)35, UASp>RhoGEF2(DHPH)-Cry2-mCherry (III)35, UASp>OptoSOS (II) (BDSC #95111), UASp>scra(RNAi) (III) (BDSC #41841), vas>Cas9 (III) (BDSC #51324), Oregon-R, sqh>gap43-mCherry (III) (gift from Adam Martin80). The fly genotypes used in each experiment are listed in Supplementary Table 1.
Live imaging and photomanipulation
Embryos were collected in apple agar plates in the dark, dechorionated for 2 min in 50% bleach, washed with water for 1 min, selected under Halocarbon oil 27 (Sigma-Aldrich, St. Louis, MO), and mounted in a 50/50 mixture of Halocarbon oil 27 and Halocarbon oil 700 (Sigma-Aldrich, St. Louis, MO) between a glass coverslip (VWR International, Radnor, PA) and an oxygen-permeable membrane (YSI, Yellow Springs, OH). For most experiments, embryos were imaged and activated on a Zeiss LSM 880 NLO upright two photon confocal microscope, equipped with a diode laser for 561 nm excitation, an argon laser for 488 nm excitation, a Mai Tai DeepSee 2 photon laser for 690-1040 nm excitation, an Airyscan detector with a FAST module, and a Plan-Apochromat 40x/1.0 NA water objective. For cephalic furrow imaging (Fig. 5g–i), cellularization/dorsal closure imaging (Fig. 6), and imaging of RhoGEF2 & Cysts localization (Supplementary Fig. 1a–f), embryos were imaged on a Zeiss LSM 880 inverted microscope, equipped with a diode laser for 561 nm excitation, an Airyscan detector with a FAST module, and a C-Apochromat 40x/1.2 NA water objective (Figs. 5g–i, 6) or a Plan-Apochromat 63x/1.40 NA oil objective (Supplementary Fig. 1a–f). Laser power measurements were performed using a S170C Microscope Slide Power Sensor (Thorlabs, Newton, NJ) and a PM100D Power and Energy Meter Console (Thorlabs, Newton, NJ), with the sensor positioned at the sample plane.
For lateral activation experiments, embryos were activated on their dorsal surfaces, at the midpoint between the anterior and posterior poles. Embryos were activated at the onset of gastrulation, when cells were between 32 and 35 µm long. During lateral activation experiments, embryos were activated by scanning a 840-nm (Fig. 4i, j), 900-nm (Fig. 1e, f, Supplementary Fig. 4c, 4d, 8c, Supplementary Movie 4) or 927-nm (all other figures) Mai Tai laser in a circular region through a cylindrical stack extending from 5 to 25 µm below the apical cell surface, sampled every 1 µm, with a pixel dwell of 1.85 µs, a pixel spacing of 0.12 µm, and a laser power between 0 and 70% (0–75 mW at sample plane), for 5 total iterations. For single time point apical activation experiments (Supplementary Fig. 2d, 2f), a Mai Tai laser was scanned in a 24-µm circular region at a single apical slice for a single time point, with a pixel dwell of 1.85 µs and a pixel spacing of 0.12 µm. For OptoGEF2 (Supplementary Fig. 2d), the laser was set to 900 nm at 25% power (33 mW at sample plane) and scanned for 7-15 iterations. For RhoGEF2(DHPH)-Cry2 (Supplementary Fig. 2f), the laser was set to 925 nm at 30% power (31 mW at sample plane) and scanned for 5–10 iterations. For sustained apical activation experiments (Supplementary Fig. 6), a 927-nm Mai Tai laser was scanned in a 18-µm circular region at a single apical slice, with a pixel dwell of 1.85 µs, a pixel spacing of 0.12 µm, a laser power of 45% (47 mW at sample plane), for 3 iterations per frame, repeated every frame (every 22 s). Movies tracking membrane markers and myosin were taken as 44-µm stacks with 2-µm slice spacing, taking 22 s/frame. For 1 P activation during early cellularization and dorsal closure (Fig. 6), embryos were activated with a 488-nm argon laser focused on the apical surface, with a pixel dwell of 1.4 µs, a pixel spacing of 0.15 µm, and a laser transmission set to 20% for 3 total iterations every frame (every ~20 s).
Vitelline membrane micro-perforation (Fig. 4i, j, Supplementary Movie 4) was performed by scanning a 840-nm laser at 100% power (nominally 2600 mW at the laser source) over a 5 µm x 10 µm patch of the vitelline membrane, sampled every 0.12 µm in xy and every 0.5 µm in z, for 1 iteration30. Laser ablation (Supplementary Fig. 9a–d) was performed by scanning a 800-nm laser at 100% power (nominally 2620 mW at the laser source) over a 1-pixel thick, 47-µm line spanning the cellularization front when cells were 30 µm long, sampled every 0.12 µm, for 25 iterations. Following ablation, tissue behavior was observed for 15 s at a 0.35-s acquisition interval.
Furrow segmentation and tracking
Optogenetically induced furrows were quantified by segmenting tissue apical and basal boundaries. Tissue apical surfaces were segmented using an ilastik autocontext workflow trained on manually annotated data with all color/intensity, edge, and texture features selected81. Output pixel probabilities were fed into an ilastik object classification workflow with 2 x 2 smoothing, hysteresis thresholding (core probability 0.5, edge probability 0.35), and were classified using the following features: convexity, number of defects, convex hull center, size in pixels, mean intensity, and histogram of intensity in neighborhood (10 x 10). Segmentations and classifications were manually confirmed and corrected when necessary. Due to poor basal membrane signal, basal tissue boundaries were segmented manually in ImageJ. Tissue movement was tracked and corrected for using PIVlab82 (FFT window deformation, 128px first pass, 64px second pass, 32px third pass).
Furrow quantification
Using the segmented tissue boundaries, cell height at each point in the tissue was calculated as the difference between the z-position of the apical tissue boundary and that of the basal tissue boundary. To avoid artifacts from embryo curvature, this was only done at coverslip-adjacent portions of the embryo. A reference cell height curve, representing what cell height would be in the absence of activation, was constructed by using non-activated portions of the embryo to calculate an embryo-specific cellularization rate, which was used to estimate expected cell lengthening through the end of the experiment. The shortening at each point of the tissue was then calculated as the difference between the instantaneous cell height and the reference cell height curve. Average shortening is measured as the largest cell shortening at any time point in the experiment, averaged across all points within the activation region. Maximum shortening is measured as the largest cell shortening at any time point in the experiment, measured at any point within the activation region. The movement of the activation region was tracked and accounted for via PIV, as described above. Furrowing and recovery timescales were calculated by finding the time point at which the maximum cell height reached 1/e = 37% of its final shortened or recovered height, interpolating between adjacent time points when necessary. Furrowing and recovery timescales were only measured in experiments where cells shortened by more than 1 µm.
Activation quantification
iLID activation levels were quantified via the fluorescence signal emitted during the first activation time point (20-µm stack, 5–25 µm below the embryo apical surface, sampled every µm, 21 slices total, shown in Fig. 2a’). A standard fluorescence curve was calculated as the average fluorescence intensity at each slice, averaged across all experiments. For each experiment, the normalized fluorescence at each slice was calculated as that experiment’s fluorescence at that slice divided by the standard curve’s fluorescence at that slice. The final normalized activation level for each experiment was calculated as the average of the experiment’s normalized fluorescence across all slices. This was used as the activation magnitude for the experiments in the main text. For the alternative activation magnitude calculation (Supplementary Fig. 7d, e), the same procedure was instead applied to the mCh-iLID-CaaX fluorescence signal at the final pre-activation time point (Fig. 2a) to get an iLID protein level. This protein level was converted into a proxy for activation magnitude by multiplying it with the square of the laser power, calculated as the product of the user-input laser power (% max) with the maximum laser power (mW) measured at the sample plane.
To quantify z-spreading of OptoGEF2 and RhoGEF2(DHPH)-Cry2 (Supplementary Fig. 2h), raw myosin intensity signal was quantified within a 12-µm circle centered at the activation region, at 0-12 µm below the apical surface, sampled every 2 µm. The xy-movement of the activation region was corrected via PIV, as described above. As cells moved away from the vitelline membrane following activation, the slice of peak myosin intensity, subject to a 3.7-µm Gaussian blur filter, was tracked and labeled as z = 0 µm below the apical surface. A Gaussian was fit to the myosin z-intensity profiles at the time point of peak apical myosin signal—this time point is reported in Supplementary Fig. 2j. The half width at half maximum (HWHM) of the Gaussian fit is reported as a proxy for z-spreading in Supplementary Fig. 2h.
Viability studies
Cages were kept at 23 °C in 24-h dark. Embryos were collected into apple agar plates for 2 h. After collection, the same number of embryos of each genotype (OptoGEF2, RhoGEF2(DHPH)-Cry2, OreR) were arranged in a grid on an apple agar plate coated with Halocarbon oil 27 (Sigma-Aldrich, St. Louis, MO). Plates were imaged on a Leica M165 FC stereomicroscope equipped with a Plan APO 1.0x M-series objective and a DFC7000T camera (Leica Microsystems, Chicago, IL) for 36 hours, with an image taken every 10 min. Then, the number of embryos that were collected, that cellularized, and that hatched were counted to calculate viability (number hatched divided by number collected), cellularization fraction (number cellularized divided by number collected), and hatch fraction (number hatched divided by number cellularized). For RhoGEF2(DHPH)-Cry2, a significant fraction of embryos that were laid were misshapen or aberrantly small. As these embryos were extremely unlikely to hatch, only embryos of proper morphology were counted. Thus, the viability and cellularization fractions reported for RhoGEF2(DHPH)-Cry2 represent an upper bound on the true RhoGEF2(DHPH)-Cry2 viability and cellularization fractions.
Quantification of cellular protein levels
To measure cellular levels of RhoGEF2 and Cysts, embryos homozygous for RhoGEF2-tgRFPt-SspB, SspB-tgRFPt-Cysts, or vas-Cas9 (III) were mounted as described. Using bright-field transmitted light, embryos were selected that were in late stage 5/early stage 6 and had their dorsal surfaces facing the coverslip. A single fluorescent image was taken 5 µm below the vitelline membrane over a 15-s scan period using the 561-nm laser line. A fluorescence signal was calculated by measuring the average fluorescence intensity across the tissue, leaving a 5-µm buffer from the vitelline membrane to exclude vitelline membrane autofluorescence. A background fluorescence level was calculated as the average fluorescence across 5 measured vas-Cas9 (III) embryos. The abundance of RhoGEF2-tgRFPt-SspB and SspB-tgRFPt-Cysts in each embryo was calculated as the average fluorescence of that embryo minus the background vas-Cas9 (III) fluorescence value.
To compare levels of cellular RhoGEF2-tgRFPt-SspB and UASp>mCh-iLID-CaaX, the same procedure as above was performed using either embryos homozygous for RhoGEF2-tgRFPt-SspB or embryos only expressing UASp>mCh-iLID-CaaX with a mat15 driver. Acquisition settings were tuned to account for the difference in fluorescence intensity levels between RhoGEF2 & iLID. We note that this introduces additional noise to the low-signal RhoGEF2 fluorescence intensity measurements. To account for the use of different fluorophores in the two constructs, measured fluorescence intensities were scaled by the relative excitation of each fluorophore at 561 nm as well as the theoretical brightness of each fluorophore37,83, to yield a final measurement of relative abundance.
Ablation analysis
Post-ablation retraction distances (Supplementary Fig. 9c, d) were calculated via PIV analysis using PIVlab82 (FFT window deformation, 1st pass 102 px), resulting in a 9 x 9 field of displacement vectors. Left & right retraction distances at each time step were calculated by averaging the x-displacements of the middle 3 vectors in the columns immediately to the left and to the right of the cut, neglecting the central vector column centered at the cut. Total retraction distance at each time step was calculated by subtracting the displacement of the left side of the cut from the displacement of the right side of the cut. Cumulative retraction distances were calculated by integrating these displacements over each time step.
Figure preparation
All figures were prepared in Figma. Graphs were made in Python using Matplotlib84. Microscopy images and movies were prepared in ImageJ85. For images of membrane markers displayed in figures, brightness and contrast were adjusted between panels to maximize image contrast and permit membrane visualization in photobleached tissues, as noted in figure captions. In transverse slices of membrane markers, fluorescence signal was normalized at all z-slices to account for decreased fluorescence signal at higher depths. When present, z-drift was manually corrected. Images were rotated and reflected so that in dorsal views, anterior is left and posterior is right, and in lateral views, anterior is left, posterior is right, dorsal is up, and ventral is down. In transverse views, apical is up and basal is down.
Statistics and reproducibility
Statistical analyses were done in Python using SciPy86. ANOVAs, chi-squared tests, and t-tests were performed as noted. Tests for coincidence of two lines were done as described in Glantz 2012, Ch. 887. All measurements were taken from distinct samples.
The images in Fig. 1c are representative of 16/17 OptoGEF2 embryos; 1/17 embryos displayed no visible shortening response (see Fig. 2c). The images in Fig. 1d are representative of 9/11 OptoCysts embryos; 2/11 embryos displayed no visible shortening response (see Fig. 2c). The images in Fig. 1e are representative of 4/4 OptoGEF2 embryos. The images in Fig. 1f are representative of 5/5 OptoCysts embryos. The images in Fig. 1g are representative of 14/17 OptoGEF2 embryos; 3/17 embryos did not display circumferential constriction. The images in Fig. 1h are representative of 11/11 OptoCysts embryos. The images in Fig. 4i, j are representative of 3/3 embryos. The images in Fig. 4k, l are representative of 3/3 embryos (AP-oriented ellipse) and 3/3 embryos (DV-oriented ellipse). The images in Fig. 5a are representative of 12/12 OptoGEF2 embryos and 10/10 OptoGEF2 + scra RNAi embryos. The images of perturbed furrowing polarity in Fig. 5c are representative of 4/6 OptoGEF2 + scra RNAi embryos (2/6 displayed apical-directed shortening and 2/6 displayed symmetric shortening); 2/6 embryos displayed basal-directed shortening. The images of basal-directed shortening in Fig. 5d are representative of 16/17 OptoGEF2 embryos; 1/17 embryos displayed apical-directed shortening. The images of apical-directed shortening without bending in Fig. 5e are representative of 3/4 OptoGEF2 + scra RNAi embryos; 1/4 embryos displayed bending. The images of apical-directed shortening with bending in Fig. 5f are representative of 5/8 OptoGEF2 embryos; 3/8 embryos displayed shortening without bending. The images of partially inverted cephalic furrow formation in Fig. 5g, h are representative of 4/4 OptoGEF2 + scra RNAi embryos. The images of cephalic furrow formation in Fig. 5i are representative of 3/3 OptoGEF2 embryos. The in-plane constriction behavior in cellularizing OptoGEF2 embryos, shown in Fig. 6a, b, is representative of 3/3 embryos. The amnioserosa constriction experiments shown in Fig. 6c, d are representative of 3/3 OptoGEF2 embryos. The lateral epidermis constriction experiments shown in Fig. 6e, f are representative of 3/3 OptoGEF2 embryos. The single-cell constriction experiments shown in Fig. 6g are representative of 3/3 OptoGEF2 embryos.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The primary microscopy data used in this study are too large to deposit in a public repository, but are available from the corresponding author upon request. Source data are provided with this paper.
Code availability
No custom code was generated.
References
Collinet, C. & Lecuit, T. Programmed and self-organized flow of information during morphogenesis. Nat. Rev. Mol. Cell Biol. 22, 245–265 (2021).
Gilmour, D., Rembold, M. & Leptin, M. From morphogen to morphogenesis and back. Nature 541, 311–320 (2017).
Pilot, F. & Lecuit, T. Compartmentalized morphogenesis in epithelia: From cell to tissue shape. Dev. Dynam. 232, 685–694 (2005).
Zartman, J. J. & Shvartsman, S. Y. Unit operations of tissue development: Epithelial folding. Annu. Rev. Chem. Biomol. Eng. 1, 231–246 (2010).
Tozluoǧlu, M. & Mao, Y. On folding morphogenesis, a mechanical problem. Philos. Trans. R. Soc. B 375, 20190564 (2020).
Baker, P. C. & Schroeder, T. E. Cytoplasmic filaments and morphogenetic movement in the amphibian neural tube. Dev. Biol. 15, 432–450 (1967).
Martin, A. C., Kaschube, M. & Wieschaus, E. F. Pulsed contractions of an actin–myosin network drive apical constriction. Nature 457, 495–499 (2009).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Martin, A. C. & Goldstein, B. Apical constriction: themes and variations on a cellular mechanism driving morphogenesis. Development 141, 1987–1998 (2014).
Kondo, T. & Hayashi, S. Mechanisms of cell height changes that mediate epithelial invagination. Dev. Growth Differ. 57, 313–323 (2015).
Ioannou, F., Dawi, M. A., Tetley, R. J., Mao, Y. & Muñoz, J. J. Development of a new 3D hybrid model for epithelia morphogenesis. Front. Bioeng. Biotechnol. 8, 405 (2020).
Spencer, A. K., Siddiqui, B. A. & Thomas, J. H. Cell shape change and invagination of the cephalic furrow involves reorganization of F-actin. Dev. Biol. 402, 192–207 (2015).
Eritano, A. S. et al. Tissue-scale mechanical coupling reduces morphogenetic noise to ensure precision during epithelial folding. Dev. Cell 53, 212–228.e12 (2020).
Sui, L. et al. Differential lateral and basal tension drive folding of Drosophila wing discs through two distinct mechanisms. Nat. Commun. 9, 4620 (2018).
Sherrard, K., Robin, F., Lemaire, P. & Munro, E. Sequential activation of apical and basolateral contractility drives ascidian endoderm invagination. Curr. Biol. 20, 1499–1510 (2010).
Athilingam, T., Tiwari, P., Toyama, Y. & Saunders, T. E. Mechanics of epidermal morphogenesis in the Drosophila pupa. Semin. Cell Dev. Biol. 120, 171–180 (2021).
Monier, B. et al. Apico-basal forces exerted by apoptotic cells drive epithelium folding. Nature 518, 245–248 (2015).
Etienne-Manneville, S. & Hall, A. Rho GTPases in cell biology. Nature 420, 629–635 (2002).
Simões, S. et al. Compartmentalisation of Rho regulators directs cell invagination during tissue morphogenesis. Development 133, 4257–4267 (2006).
Denk-Lobnig, M. & Martin, A. C. Modular regulation of Rho family GTPases in development. Small GTPases 10, 122–129 (2019).
Mikawa, M., Su, L. & Parsons, S. J. Opposing roles of p190RhoGAP and Ect2 RhoGEF in regulating cytokinesis. Cell Cycle 7, 2003–2012 (2008).
Mason, F. M., Xie, S., Vasquez, C. G., Tworoger, M. & Martin, A. C. RhoA GTPase inhibition organizes contraction during epithelial morphogenesis. J. Cell Biol. 214, 603–617 (2016).
Müller, P. M. et al. Systems analysis of RhoGEF and RhoGAP regulatory proteins reveals spatially organized RAC1 signalling from integrin adhesions. Nat. Cell. Biol. 22, 498–511 (2020).
Chen, Z. et al. Activated RhoA binds to the pleckstrin homology (PH) domain of PDZ-RhoGEF, a potential site for autoregulation. J. Biol. Chem. 285, 21070–21081 (2010).
Medina, F. et al. Activated RhoA is a positive feedback regulator of the Lbc family of Rho guanine nucleotide exchange factor proteins. J. Biol. Chem. 288, 11325–11333 (2013).
Rich, A., Fehon, R. G. & Glotzer, M. Rho1 activation recapitulates early gastrulation events in the ventral, but not dorsal, epithelium of Drosophila embryos. eLife 9, e56893 (2020).
Lin, B., Luo, J. & Lehmann, R. An AMPK phosphoregulated RhoGEF feedback loop tunes cortical flow–driven amoeboid migration in vivo. Sci. Adv. 8, eabo0323 (2022).
Häcker, U. & Perrimon, N. DRhoGEF2 encodes a member of the Dbl family of oncogenes and controls cell shape changes during gastrulation in Drosophila. Genes Dev. 12, 274–284 (1998).
Kölsch, V., Seher, T., Fernandez-Ballester, G. J., Serrano, L. & Leptin, M. Control of Drosophila gastrulation by apical localization of adherens junctions and RhoGEF2. Science 315, 384–386 (2007).
Popkova, A. et al. A mechanical wave travels along a genetic guide to drive the formation of an epithelial furrow during Drosophila gastrulation. Dev. Cell 59, 400–414.e5 (2024).
Garcia De Las Bayonas, A., Philippe, J.-M., Lellouch, A. C. & Lecuit, T. Distinct RhoGEFs activate apical and junctional contractility under control of G proteins during epithelial morphogenesis. Curr. Biol. 29, 3370–3385.e7 (2019).
Silver, J. T. et al. Apical polarity proteins recruit the RhoGEF Cysts to promote junctional myosin assembly. J. Cell Biol. 218, 3397–3414 (2019).
Kennedy, M. J. et al. Rapid blue-light–mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl. Acad. Sci. Usa. 112, 112–117 (2015).
Izquierdo, E., Quinkler, T. & De Renzis, S. Guided morphogenesis through optogenetic activation of Rho signalling during early Drosophila embryogenesis. Nat. Commun. 9, 2366 (2018).
Herrera-Perez, R. M., Cupo, C., Allan, C., Lin, A. & Kasza, K. E. Using optogenetics to link myosin patterns to contractile cell behaviors during convergent extension. Biophys. J. 120, 4214–4229 (2021).
Shaner, N. C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Gratz, S. J. et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 196, 961–971 (2014).
Krueger, D. et al. Principles and applications of optogenetics in developmental biology. Development 146, dev175067 (2019).
Kinjo, T. et al. FRET-assisted photoactivation of flavoproteins for in vivo two-photon optogenetics. Nat. Methods 16, 1029–1036 (2019).
Sui, L. & Dahmann, C. Increased lateral tension is sufficient for epithelial folding in Drosophila. Development 147, dev194316 (2020).
Parks, S. & Wieschaus, E. The Drosophila gastrulation gene concertina encodes a Gα-like protein. Cell 64, 447–458 (1991).
Rogers, S. L., Wiedemann, U., Häcker, U., Turck, C. & Vale, R. D. Drosophila RhoGEF2 associates with microtubule plus ends in an EB1-dependent manner. Curr. Biol. 14, 1827–1833 (2004).
Lecuit, T. & Wieschaus, E. Polarized insertion of new membrane from a cytoplasmic reservoir during cleavage of the Drosophila embryo. J. Cell Biol. 150, 849–860 (2000).
Azevedo, D. et al. DRhoGEF2 regulates cellular tension and cell pulsations in the amnioserosa during Drosophila dorsal closure. PLoS ONE 6, e23964 (2011).
Kerridge, S. et al. Modular activation of Rho1 by GPCR signalling imparts polarized myosin II activation during morphogenesis. Nat. Cell Biol. 18, 261–270 (2016).
Casas-Vila, N. et al. The developmental proteome of Drosophila melanogaster. Genome Res. 27, 1273–1285 (2017).
Natwick, D. E. & Collins, S. R. Optimized iLID membrane anchors for local optogenetic protein recruitment. ACS Synth. Biol. 10, 1009–1023 (2021).
Gelbart, M. A. et al. Volume conservation principle involved in cell lengthening and nucleus movement during tissue morphogenesis. Proc. Natl. Acad. Sci. Usa. 109, 19298–19303 (2012).
Guerra Santillán, K. Y., Dahmann, C. & Fischer-Friedrich, E. Elastic contractile stress in the basement membrane generates basal tension in epithelia. PRX Life 2, 013004 (2024).
Hannezo, E., Prost, J. & Joanny, J.-F. Theory of epithelial sheet morphology in three dimensions. Proc. Natl. Acad. Sci. Usa. 111, 27–32 (2014).
Andrenšek, U., Ziherl, P. & Krajnc, M. Wrinkling instability in unsupported epithelial sheets. Phys. Rev. Lett. 130, 198401 (2023).
Dong, C.-Y., Koenig, K. & So, P. Characterizing point spread functions of two-photon fluorescence microscopy in turbid medium. J. Biomed. Opt. 8, 450 (2003).
Warn, R. & Magrath, R. F-actin distribution during the cellularization of the embryo visualized with FL-phalloidin. Exp. Cell Res. 143, 103–114 (1983).
Warn, R. M. & Robert-Nicoud, M. F-actin organization during the cellularization of the Drosophila embryo as revealed with a confocal laser scanning microscope. J. Cell Sci. 96, 35–42 (1990).
Cheikh, M. I. et al. A comprehensive model of Drosophila epithelium reveals the role of embryo geometry and cell topology in mechanical responses. eLife 12, e85569 (2023).
Thomas, J. H. & Wieschaus, E. src64 and tec29 are required for microfilament contraction during Drosophila cellularization. Development 131, 863–871 (2004).
Field, C. M., Coughlin, M., Doberstein, S., Marty, T. & Sullivan, W. Characterization of anillin mutants reveals essential roles in septin localization and plasma membrane integrity. Development 132, 2849–2860 (2005).
Goldner, A. N., Cheikh, M. I., Osterfield, M. & Doubrovinski, K. Viscous shear is a key force in Drosophila ventral furrow morphogenesis. Development 151, dev202892 (2024).
Krueger, D., Quinkler, T., Mortensen, S. A., Sachse, C. & De Renzis, S. Cross-linker–mediated regulation of actin network organization controls tissue morphogenesis. J. Cell Biol. 218, 2743–2761 (2019).
Lv, Z., Zhang, N., Zhang, X., Großhans, J. & Kong, D. The lateral epidermis actively counteracts pulling by the amnioserosa during dorsal closure. Front. Cell Dev. Biol. 10, 865397 (2022).
Redchuk, T. A. et al. Optogenetic regulation of endogenous proteins. Nat. Commun. 11, 605 (2020).
Manoilov, K. Y. U., Verkhusha, V. V. & Shcherbakova, D. M. A guide to the optogenetic regulation of endogenous molecules. Nat. Methods 18, 1027–1037 (2021).
Kook, Y. H. et al. AAV-compatible optogenetic tools for activating endogenous calcium channels in vivo. Mol. Brain 16, 73 (2023).
Wenzl, C., Yan, S., Laupsien, P. & Großhans, J. Localization of RhoGEF2 during Drosophila cellularization is developmentally controlled by slam. Mech. Dev. 127, 371–384 (2010).
Dawes-Hoang, R. E. et al. Folded gastrulation, cell shape change and the control of myosin localization. Development 132, 4165–4178 (2005).
Polyakov, O. et al. Passive mechanical forces control cell-shape change during Drosophila ventral furrow formation. Biophys. J. 107, 998–1010 (2014).
Krueger, D., Tardivo, P., Nguyen, C. & De Renzis, S. Downregulation of basal myosin-II is required for cell shape changes and tissue invagination. EMBO J. 37, e100170 (2018).
Molnar, K. & Labouesse, M. The plastic cell: mechanical deformation of cells and tissues. Open Biol. 11, 210006 (2021).
Bi, D., Lopez, J. H., Schwarz, J. M. & Manning, M. L. Energy barriers and cell migration in densely packed tissues. Soft Matter 10, 1885 (2014).
Takeda, H., Kameo, Y., Inoue, Y. & Adachi, T. An energy landscape approach to understanding variety and robustness in tissue morphogenesis. Biomech. Model. Mechan. 19, 471–479 (2020).
Niloy, R. A., Holcomb, M. C., Thomas, J. H. & Blawzdziewicz, J. The mechanics of cephalic furrow formation in the Drosophila embryo. Biophys. J. 122, 3843–3859 (2023).
Santillán, K. Y. G., Jantzen, C., Dahmann, C. & Fischer-Friedrich, E. Epithelial folding through local degradation of an elastic basement membrane plate. Adv. Phys. Res. 3, 2400062 (2024).
Savin, T. et al. On the growth and form of the gut. Nature 476, 57–62 (2011).
Nelson, C. M. On buckling morphogenesis. J. Biomech. Eng. 138, 021005 (2016).
Trushko, A. et al. Buckling of an epithelium growing under spherical confinement. Dev. Cell 54, 655–668.e6 (2020).
Wang, H., Liu, Y.-Q., Hang, J.-T., Xu, G.-K. & Feng, X.-Q. Dynamic high-order buckling and spontaneous recovery of active epithelial tissues. J. Mech. Phys. Solids 183, 105496 (2024).
Keenan, S. E. et al. Rapid dynamics of signal-dependent transcriptional repression by Capicua. Dev. Cell 52, 794–801.e4 (2020).
Di Pietro, F. et al. Systematic analysis of RhoGEF/GAP localizations uncovers regulators of mechanosensing and junction formation during epithelial cell division. Curr. Biol. 33, 858–874.e7 (2023).
Martin, A. C., Gelbart, M., Fernandez-Gonzalez, R., Kaschube, M. & Wieschaus, E. F. Integration of contractile forces during tissue invagination. J. Cell Biol. 188, 735–749 (2010).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Thielicke, W. & Stamhuis, E. J. PIVlab – Towards user-friendly, affordable and accurate digital particle image velocimetry in MATLAB. J. Open Res. Softw. 2 (2014).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Glantz, S. A. Primer of biostatistics (McGraw-Hill, New York, 2012).
Acknowledgements
We thank Stas Shvartsman and Liz Gavis for their contributions to conceptualization of the endogenous optogenetic tools and for helpful discussions. We thank Bex Pendrak, Sameer Thukral, and Kasza Lab members for helpful discussions. Stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) were used in this study. This work was performed in part at the Live Imaging and Bioenergetics Facility at the Advanced Science Research Center at The Graduate Center of the City University of New York. This work was supported by NIH Grant R35GM138380 to KEK and NIH Grant 1F31HD118793-01 to ADC; KEK holds an NSF CAREER Award, Packard Fellowship, and Sloan Research Fellowship in Physics.
Author information
Authors and Affiliations
Contributions
The tools were conceptualized and designed by A.D.C., C.A.D., R.M.H., and K.E.K. Cloning and Drosophila transgenesis were done by A.D.C., C.A.D., and R.M.H. Experimental design was done by A.D.C. and K.E.K. Experimentation and data analysis were performed by A.D.C. The manuscript was written by A.D.C. and K.E.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Countryman, A.D., Doherty, C.A., Herrera-Perez, R.M. et al. Endogenous OptoRhoGEFs reveal biophysical principles of epithelial tissue furrowing. Nat Commun 16, 7665 (2025). https://doi.org/10.1038/s41467-025-62483-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62483-6