Abstract
Coagulation factor XII has been identified as a potential drug target that could prevent thrombosis without increasing the risk of bleeding. However, human data to support the development of factor XII-directed therapeutics are lacking. To assess the role of factor XII in venous thromboembolism, we examine genetic variation in the coding region of the F12 locus across 703,745 participants in the UK Biobank and NIH All of Us biorepositories. We find that heterozygous carriers of nonsense, frameshift, and essential splice site variants in F12 are protected against venous thromboembolism without an increased risk of bleeding or infection. We also show that F12 variant carriers generally experience a quantitative (type I) defect in circulating factor XII levels, though a subset of participants was also identified with possible qualitative (type II) deficiency. In vitro plasma-based thrombin generation is reduced at factor XII concentrations reflective of those seen in F12 variant carriers. We also show that F12 heterozygous mice are protected against venous thromboembolism and display an intermediate phenotype between wild-type and F12-null animals. We conclude that heterozygous loss of F12 represents a haploinsufficient state characterized by protection against venous thromboembolism and that therapeutically inhibiting factor XII is likely to be safe and effective.
Similar content being viewed by others
Introduction
Patients who experience venous thromboembolism (VTE) require treatment with anticoagulant medication, and current guidelines recommend lifelong therapy for patients with unprovoked VTE or persistent major risk factors1. However, all existing anticoagulant agents are associated with an increased risk of major and clinically-relevant non-major bleeding that ranges between 4 and 15% per year2,3,4,5,6. This means that a large proportion of treated patients will experience bleeding and many others cannot receive therapy at all due to pre-existing risk factors for this complication7,8,9.
Within this context, coagulation factor XII (FXII, F12) represents a potential drug target that could “decouple” hemostasis from therapeutic anti-thrombotic effect10,11,12,13. Severe congenital FXII deficiency in humans (activity <1%) does not cause a bleeding diathesis14,15,16, while FXII deletion or inhibition in preclinical models consistently protects against thrombosis without increasing bleeding17,18,19,20,21,22,23,24. These observations suggest that therapeutic FXII blockade could address a major unmet need by providing anti-thrombotic effect without a heightened risk of hemorrhage. However, despite the potential clinical importance of FXII, human data to support its role in VTE are limited25. To date, only two small epidemiologic studies have been performed to evaluate this question, one in the Netherlands26 and the other in the United States27. Neither revealed an association between circulating FXII levels and VTE. Both studies were limited by small sample size, the use of spot measurements of plasma FXII levels without germline genotyping data, inconsistent assay methods, ascertainment bias (including a reliance on individuals presenting to medical attention for VTE), and insufficiently matched controls. Because the human data remain discrepant with findings across animal studies consistently showing that loss of FXII is protective against VTE24,28, many have called for further research targeted at individuals with FXII deficiency29,30.
Here, we have leveraged population-scale multidimensional datasets to evaluate the clinical impacts of germline loss of function in F12. Our data show that heterozygous loss of F12 represents a haploinsufficient state associated with protection against a first VTE event.
Results
Study population and F12 variants
We included whole exome sequencing data from 414,670 participants in the UK Biobank (UKB) and whole genome sequencing data from 289,075 participants in the NIH All of Us (AoU) biorepository (Fig. 1a). Summary data for the two cohorts are shown in Table S1. For each moderately rare (MAF ≤ 1%) variant in the coding region of the F12 locus, 30 in silico prediction tools included in the dbNSFP database were used to assign a composite “functional impact score” (FIS) between 0 and 1, corresponding to the percentage of tools predicting a deleterious effect31. A higher FIS indicates that a variant is more likely to damage protein activity. High-confidence loss-of-function (HCLOF) variants, including those that result in frameshift, truncation (nonsense), and essential splice-site disruption, are considered the most likely to cause loss of protein function and were assigned an FIS of 1.0. Of 8231 individuals in the UKB and 7699 in AoU with at least one moderately rare nonsynonymous variant in F12, we identified 2253 carriers of 102 unique F12 variants with FIS = 1.0 (Data S1), 99.9% of whom were heterozygous. Within the coding region of F12, variants with FIS = 1.0 appeared to be relatively evenly distributed throughout the gene product (Fig. 1b). The ancestral breakdown of each cohort is shown in Fig. 1c.
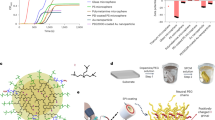
a Moderately rare (MAF ≤ 1%) variant filtering strategy, and b distribution and frequencies of high confidence loss-of-function (FIS = 1.0, MAF ≤ 1%) coding variants in the F12 locus identified in the UK Biobank (UKB) and NIH All of Us (AoU) datasets. There were 31 unique essential splice site variants that fell outside the exon boundaries and are not shown. Amino acids 1-19 (gray) comprise the FXII signal peptide. c Study breakdown by biobank and subpopulation as determined by principal components analysis of genetic ancestry. d For each moderately rare F12 variant, the in-cohort MAF was computed and plotted against the FIS value. All missense and HCLOF variants with in-cohort MAF ≤ 1% were included across the full range of FIS assignments. Variants are shown stratified by the dataset in which they were found (“UKB” or “AoU”), with variants shared across both datasets noted (“Both”). e Variant counts for F12 are shown according to FIS threshold (black circles). For comparison, the in-group median variant allele count (MAF ≤ 1%) at each FIS threshold is shown for two gene sets: vitamin K (VK)-dependent coagulation factors (F2, F7, F9, F10) (orange squares) and the larger group of essential humoral coagulation factors (F2, F5, F7, F8, F9, F10) (red triangles). Variant counts for F12 and each gene set were normalized to the number of variants at the FIS = 0 threshold. Abbreviations: FIS functional impact score, FN2 fibronectin type 2 domain, EGF EGF-like domain, FN1 fibronectin type 1 domain, KD kringle domain, PRR proline rich region, AFR African, AMR admixed American, EAS East Asian, EUR European, SAS South Asian, MID Middle Eastern, MAF minor allele frequency, VK vitamin K-dependent.
Despite the general expectation that physiologically deleterious variants are likely to be rarer, we found that in-cohort frequencies of F12 variants bore little relation to their FIS rating (Fig. 1d). Consistent with this finding, F12 variant allele counts failed to decline precipitously at the highest FIS cutoffs (Fig. 1e). For comparison, we evaluated the median allele counts at the same FIS thresholds for two groups of genes that are necessary for hemostasis: the vitamin K-dependent coagulation factors (F2, F7, F9, and F10) and a larger set of essential humoral coagulation factors (F2, F5, F7, F8, F9, and F10). For both gene sets, the median variant allele count at each FIS threshold demonstrated a marked decline as FIS rose, consistent with loss-of-function variants in these genes being under significant negative selection pressure (genetic constraint). Taken together, these data suggest that germline loss-of-function in F12 is well-tolerated.
F12 variant carriers are protected against venous thromboembolism
Using Cox proportional hazards regression modeling with Firth’s penalized likelihood, we performed gene-level collapsing analyses of F12 in each dataset with a focus on FIS = 1.0 (HCLOF) variants (Table S2). Random-effects model meta-analysis of the two in-cohort results demonstrated that F12 variant carrier status was significantly associated with protection against a first VTE event (HR = 0.648, 95% CI: 0.496–0.846, P = 0.001) (Fig. 2a). When coding variants in the F12 locus across a range of FIS values were considered, point estimates for VTE risk declined steadily with higher FIS thresholds (Fig. 2b). By contrast, FIS stringency did not appear to be associated with bleeding risk.
a Cox proportional hazards regression with Firth’s penalized likelihood modeling was performed in the UK Biobank (UKB) and NIH All of Us (AoU) datasets, followed by random-effects model meta-analysis. All models were adjusted for sex, the first 10 principal components of genetic ancestry, and additional covariates as depicted in Table S3. (**) = value of ≤20 redacted to comply with NIH reporting regulations. b Cox proportional hazards regression for VTE followed by trans-cohort meta-analysis was repeated across a range of FIS thresholds with adjustment performed as in (a). Effect size estimates are nominally significant for the points displayed in red (P ≤ 0.024; significance threshold not adjusted for multiple comparisons). c A leave-one-variant-out (LOVO) analysis was performed using iterative Firth’s logistic regression modeling across all FIS = 1.0 variants in both cohorts. Outliers identified by the two-sided extreme studentized deviate (Grubbs) test are labeled. d Integrated Kaplan–Meier survival analysis across both UKB and AoU (N = 753,617) comparing incident VTE occurring after study enrollment between F12 variant carriers (blue) and non-carriers (black). Historical (prevalent) VTE events occurring prior to study enrollment were excluded. e For all F12 variant carriers (MAF ≤ 1%) in the UKB Pharma Proteomics Project (PPP) dataset with available plasma proteomics data (N = 626), we plotted variant FIS against the plasma FXII concentration in linearized NPX (L-NPX) units as determined by Olink®. The P-value for trend derived from the F-test is shown. f Mean (±SEM) circulating FXII levels as determined by Olink® (L-NPX) were compared between wild-type individuals (N = 41,041) and carriers of nonsense, frameshift, and insertion/deletion variants in F12 (FIS = 1.0) by unpaired two-sided t-test (N = 11). g Scatter plot showing the distribution of plasma FXII levels vs. the concentrations of GAPDH, a standard plasma housekeeping protein (N = 42,100). Vertical dotted lines represent the median plasma FXII value in L-NPX units for F12 variant carriers at FIS = 1.0 (blue) and the median plasma FXII concentration for the entire population (black). h Plasma samples from F12 variant carriers (FIS = 1.0, N = 29) and age- and sex-matched wild-type controls (N = 29) in the MGB Biobank were assayed for FXII concentration by enzyme-linked immunosorbent assay (ELISA) and compared by unpaired two-sided t-test. Carriers of essential splice site (ESS) variants were excluded from the analyses in (e–h).
We then performed a leave-one-variant-out (LOVO) sensitivity analysis using Firth’s logistic regression to identify any individual variants that may be driving the observed effect size estimates (Fig. 2c). No variant was found that, when excluded, appreciably weakened the association between F12 variant carrier status and protection against VTE, suggesting that our findings do not rely on any single influential variant. However, the exclusion of two essential splice site variants appeared to substantially strengthen the association between F12 variant carrier status and reduced VTE risk: chr5:177402460:C:T (“ESS1,” trans-cohort MAF: 0.031%) and chr5:177409044:A:C (“ESS2,” trans-cohort MAF: 0.16%). Whereas carriers of ESS1 had significantly reduced circulating FXII antigen levels according to the UKB Olink® plasma proteomics data, ESS2 did not appear to affect plasma FXII concentrations (Fig. S1A). Notably, excluding both ESS variants appeared to enhance the point estimate for protection against VTE in F12 variant carriers (HR = 0.454, 95% CI: 0.181–1.138, P = 0.092) (Fig. S1B). When evaluated separately by Cox regression, ESS1 carrier status (N = 393) was not associated with protection against VTE (HR = 1.02, 95% CI: 0.486–2.142, P = 0.96), whereas ESS2 carriers (N = 1447) experienced a significant reduction in disease risk (HR = 0.65, 95% CI: 0.45–0.89, P = 0.007) (Fig. S1B). These data could be consistent with ESS2 causing a type II (qualitative) deficiency in FXII. To further evaluate the biology of ESS1, we compared plasma samples of ESS1 variant carriers and participants with wild-type F12 in the MGB Biobank. As expected, ESS1 was associated with significantly decreased circulating FXII (12.74 µg/ml vs. 25.17 µg/ml, P = 0.0001). Western blotting for FXII showed that ESS1 variant carriers displayed a single band at approximately 50 kDa, consistent with the cleaved heavy chain of activated FXII (FXIIa) (Fig. S1C). By contrast, the majority of plasma FXII in wild-type individuals was in zymogen (uncleaved) form. Basal FXII activity in the plasma of ESS1 variant carriers (N = 6) greatly exceeded that of participants with wild-type F12 (N = 6) despite FXII being present at lower concentrations in ESS1 carriers (Fig. S1D). Taken together, these data are consistent with ESS1 being associated with a dysregulated form of FXII that is more easily activated at baseline.
We next investigated the potential interaction between F12 and common germline genetic risk factors for VTE (Table S3). As expected, carrier status for factor V Leiden (rs6025) and the prothrombin (F2) G20210A mutation (rs1799963) were strongly associated with VTE, as was the polygenic risk score (PRS) for VTE32. However, adjustment for these covariates and other established risk factors did not appreciably change the effect size estimates for F12 variant carrier status, suggesting that F12 functions independently of known environmental, acquired, and common genetic risk factors for VTE. Similarly, restricting our analysis to unrelated individuals (N = 565,807) did not significantly alter our findings (Table S4).
As an orthogonal approach and to assess the possibility of ongoing protection against VTE into later life, we conducted an integrated Kaplan–Meier analysis across both datasets (N = 753,617) after excluding all historical (prevalent) VTE events occurring prior to study enrollment (Fig. 2d). We found that F12 variant carriers (N = 2196) were at significantly lower risk of developing incident VTE compared to noncarriers (N = 751,421) (HR = 0.548, 95% CI: 0.384–0.777, log-rank P < 0.001) at a median (IQR) follow up of 10.7 (3.6) years.
Validation of F12 variant effect predictions
FXII plasma proteomics data from 44,464 individuals are available through the UKB Pharma Proteomics Project (PPP). After excluding ESS variant carriers, circulating FXII levels declined steadily with increasing F12 variant FIS (P < 0.0001 for trend) (Fig. 2e). The relationship between variant FIS and plasma FXII levels remained strong in a linear regression analysis adjusting for known determinants of circulating protein levels, including age, sex, ancestry, and estimated glomerular filtration rate (β = −0.407, SE = 0.04, P < 0.0001) (Table S5). Additionally, individuals with essential splice site, frameshift, and nonsense variants (FIS = 1.0) displayed mean plasma FXII levels that were significantly lower than wild-type participants (L-NPX 0.461 vs. 1.001, P < 0.0001) (Fig. 2f, g).
The accuracy of the Olink® plasma proteomics platform is known to be analyte-dependent33. To orthogonally assess plasma FXII concentrations among F12 variant carriers, we obtained human plasma samples from participants in the MGB Biobank34,35 who are heterozygous for F12 variants with FIS = 1.0 (N = 29). Using enzyme-linked immunosorbent assay (ELISA), plasma FXII levels were compared between F12 variant carriers and age- and sex-matched individuals with wild-type F12 (N = 29) (Fig. 2h). Variant carriers had significantly lower mean ± SD plasma FXII levels (12.68 ± 4.01 µg/ml) compared to wild-type individuals (25.17 ± 11.76 µg/ml) (P < 0.001), consistent with the Olink® data.
F12 variant carrier status is not associated with bleeding or sepsis
Severe congenital FXII deficiency in humans (<5% activity) does not cause a bleeding diathesis despite markedly prolonged clinical clotting times36,37. Conversely, FXII is an important activator of the kinin-kallikrein system of innate immunity, and whether moderate FXII deficiency predisposes individuals to severe infection remains unknown29,38. We found that carrier status for F12 variants (FIS = 1.0) was not associated with an increased risk of coagulopathic bleeding (Fig. 3a and Table S6) or sepsis (Fig. 3a and Table S7) in a trans-cohort Cox regression analysis.
a Cox proportional hazards regression modeling followed by trans-cohort meta-analysis was performed to examine the associations between F12 variant carrier status (FIS = 1.0) and the occurrence of VTE, bleeding, and sepsis (blue) in the UKB and AoU datasets (N = 703,745). Using the same approach, separate effect size estimates (±95% CI) for each phenotype were generated using only synonymous variants in F12 (orange). Models were adjusted for sex and ancestry as well as the additional covariates listed in Tables S7 and S8. b, c In assessments restricted to UKB dataset (N = 414,670), we used Kaplan–Meier analysis to compare overall mortality between F12 variant carriers and non-carriers, followed by two-sample comparisons of markers of fertility. Two-sided t-test P values are shown; whiskers show 5th–95th percentile range, and individual values falling above the 95th percentile are not shown. (Live births: WT N = 244,079, variant N = 989, 424 outliers not shown; still births: WT N = 77,899, variant N = 449, 855 outliers not shown; children fathered: WT N = 204,370, variant N = 767, 796 outliers not shown). d Using serial Firth’s logistic regression analyses adjusting for age, sex, and ancestry, we evaluated the associations between 138 discrete infection phenotypes and F12 variant carrier status. The “any infection” category denotes positive status for any of the 138 phenotypes. Upward triangles represent directionally positive associations, whereas downward triangles represent directionally negative associations. The dashed line represents the Bonferroni-corrected statistical significance threshold, P < 3.6 × 10−4.
We next performed analyses using detailed data from UKB (N = 414,670) to identify potential adverse effects associated with F12 haploinsufficiency. Heterozygous loss of F12 was not associated with the higher baseline reticulocyte counts and lower baseline hemoglobin levels seen in participants with coagulopathic bleeding39 (Table S8). Moreover, F12 variant carriers did not experience significantly higher mortality (Fig. 3b) or diminished markers of fertility (Fig. 3c), except for a modest reduction in the number of children fathered (mean: 1.82 in non-carriers vs. 1.64 for carriers, P = 0.0003). Examining 138 discrete infection phenotypes (Fig. 3d and Data S2), we identified no significant associations with F12 variant carrier status. Further, variant carrier status was not associated with plasma levels of C-reactive protein (CRP), a common marker of systemic inflammation (Table S9).
Partial loss of FXII reduces thrombin generation in plasma
We next sought to determine whether partial FXII deficiency might afford protection against procoagulant challenge. FXII-deficient plasma spiked with 50% of normal FXII levels demonstrated a marked decrease in contact pathway (silica)-initiated thrombin generation that was not observed at higher FXII concentrations (Fig. 4a). As expected, extrinsic pathway (tissue factor)-initiated thrombin generation was not affected by plasma FXII concentration (Fig. 4b). These observations remained consistent across several metrics, including thrombin generation velocity (Fig. 4c), peak thrombin generation (Fig. 4d), and endogenous thrombin potential (Fig. 4e). The activated partial thromboplastin time (aPTT) declined sharply with increasing plasma FXII concentrations and was normalized at 50% (Fig. 4f), whereas the prothrombin time (PT) remained unaffected by plasma FXII concentration. Taken together, these data suggest that heterozygous loss of F12 is silent in clinical coagulation assays but nevertheless sufficient to reduce the procoagulant potential of platelet-poor plasma.
Representative thrombin generation curves for FXII-deficient plasma reconstituted with varying levels of FXII zymogen in a silica-initiated and b tissue factor (TF)-initiated assays. Quantitative summary data from calibrated automated thrombography assays are shown, including c thrombin generation velocity, d peak thrombin generation, and e endogenous thrombin potential (ETP) (n = 3 independent experiments, data presented as mean ± SD). Conditions marked by an asterisk (*) differ significantly (P < 0.05, by two-sided t test) from the condition with 100% FXII zymogen levels. f Activated partial thromboplastin time (aPTT) and prothrombin time (PT) assays performed on FXII-deficient plasma reconstituted with varying concentrations of FXII zymogen. Abbreviations: aPTT activated thromboplastin time, PT prothrombin time.
Heterozygous deletion of F12 is protective against VTE in vivo
Although biallelic deletion of F12 has consistently been shown to protect against thrombosis without contributing to bleeding in a range of animal models19,20,21,24,40, heterozygous loss of F12 has not been studied in detail. Given our finding that lifelong partial FXII deficiency among biobank participants is associated with a lower risk of VTE, we sought to evaluate whether heterozygous F12 knockout mice (F12+/-) are protected in an intravital electrolytic femoral vein injury model of thrombus formation. As expected, F12+/− mice demonstrated circulating FXII levels that were approximately 50% of normal when measured by ELISA (Fig. 5a) or western blot (Fig. 5b). Blood hemoglobin levels, white blood cell count, and platelet count were not significantly different in F12+/− mice compared to wild-type animals (F12+/+) and those with biallelic deletion of F12 (F12−/−) (Fig. S2). Real-time intravital microscopy showed a marked reduction in platelet and fibrin accumulation at sites of vascular injury in F12+/− animals relative to wild-type (F12+/+) (Fig. 5c and Video S1). Quantitative imaging analysis demonstrated that F12+/− mice displayed significantly lower platelet accumulation (P = 0.032, Fig. 5d, e) and fibrin formation (P = 0.025, Fig. 5f, g) in response to electrolytic femoral vein injury. F12+/− mice thus displayed an intermediate phenotype that fell between what was observed in F12−/− and F12+/+ animals.
Plasma levels of FXII antigen were determined in F12+/+ (N = 6), F12+/− (N = 6), and F12−/− (N = 7) mice by a ELISA (median ± IQR, P value computed using one-way ANOVA) and b western blotting. c Venous thrombus formation was evaluated in F12+/+, F12+/− and F12−/− mice using the femoral vein electrolytic injury model with representative images of platelet accumulation (green) and fibrin formation (red) after vascular injury. Scale bar = 200 µm. d Quantification of platelet fluorescence intensity (in relative fluorescence units, RFU; data presented as mean ± SEM) over time and e the integrated platelet fluorescence intensity expressed as area under the curve (AUC ± SD) values, according to F12 genotype (F12+/+, N = 10; F12+/−, N = 10; F12−/−, N = 8; P value computed using one-way ANOVA). Similarly, fibrin fluorescence intensity over time (f) and the integrated fibrin fluorescence intensity (g) were quantified by genotype. Abbreviations: RFU relative fluorescence units, AUC area under curve.
Discussion
We have employed multidimensional data from over 700,000 individuals to conduct what to our knowledge is the largest study to date of FXII deficiency. We show that heterozygous loss of function in F12 constitutes a haploinsufficient state characterized by lower circulating FXII levels, reduced plasma thrombin generation, and protection against VTE in vivo.
We found that F12 variant carrier status is associated with protection against a first VTE event (HR = 0.648) similar to that observed among individuals with group O blood41,42 or those receiving prophylactic therapy with aspirin43,44,45. Notably, we demonstrate that plasma-based thrombin generation is significantly diminished at FXII concentrations ~50% of normal, corresponding to levels observed in heterozygous F12 variant carriers. We corroborated these findings by showing that F12+/− mice are significantly protected against venous thrombosis. Our data are at variance with prior results in a murine mesenteric arteriole thrombosis model suggesting that the F12+/− genotype is not associated with protection; this discrepancy may be due to the different vascular beds and model systems evaluated in that study20. Although the precise molecular mechanism of FXII activation in vivo remains unclear, our data fit a model in which FXII is needed to achieve sufficient thrombin generation under conditions of thrombosis but not physiologic hemostasis. Moreover, these results suggest that FXII remains an important contributor to VTE pathogenesis independent of effects mediated by tissue factor and/or thrombin-mediated back activation of FXI46.
Consistent with preclinical data17,18,19,20,21,22,23,24 and decades of observation, FXII deficiency in our analysis was not associated with an increased risk of bleeding or infection. We also show that loss-of-function variants in F12 appear to be under low negative selection pressure according to indirect metrics of genetic constraint. At minimum, these data support the hypothesis that physiologic hemostasis is decoupled from pathologic thrombosis at the level of FXII and strongly suggest that lifelong moderate FXII deficiency is unlikely to reduce reproductive fitness. To date, no other coagulation factor has been found to have these properties.
Our data also challenge the canonical view that humoral coagulation factor deficiencies are inherited solely as recessive, dominant, and X-linked bleeding disorders. Rather, partial (heterozygous) loss of function in some coagulation factor genes may represent haploinsufficient states that result in unexpected phenotypes. It is likely that established Mendelian definitions of inheritance fail to capture the full complexity of relationships between genotype and clinical presentation among individuals with coagulation factor gene defects. Within this context, well-annotated, large-scale germline genomic datasets offer an unparalleled opportunity to identify conditions that exist on a spectrum of recessive to dominant inheritance and uncover biological effects that do not require total genetic loss of function47,48,49.
This work included a number of important limitations. First, despite the size of the cohorts used, we were only able to analyze heterozygous F12 loss of function. While we expect protection against VTE to be more pronounced in individuals with biallelic loss of F12, we cannot rule out the possibility that this genotype is also associated with previously unrecognized adverse effects (e.g. increased infection risk). Second, we remain limited in our ability to resolve the effects of individual F12 variants; we await the advent of larger datasets to help address this issue. Third, our study relied on in silico predictions of variant effect. To mitigate this concern, we focused our primary analysis solely on HCLOF variants that are highly likely to disrupt FXII activity in vivo and directly confirmed that carriers had significantly lower plasma FXII levels.
Strengths of our approach include the use of a population genomics analysis that helps overcome many key limitations affecting prior studies. Ascertainment bias is of particular concern in population-based FXII research in light of our data showing that FXII haploinsufficiency is likely silent on standard clinical coagulation assays and does not lead carriers to present to medical attention. This study also featured plasma proteomics and gold-standard ELISA validation of predicted germline variant effects. Perhaps most importantly, we were able to validate our findings in an animal model and demonstrate a protective phenotype for venous thrombosis in F12 heterozygous mice.
In summary, we have shown that genetically-defined FXII deficiency is protective against a first VTE event in a large human population. These findings help resolve longstanding uncertainty surrounding the role of FXII in thrombotic disease and suggest that targeting FXII is likely to be a safe and effective therapeutic strategy.
Methods
Ethics
Study design and conduct complied with all relevant regulations regarding the use of human study participants and was conducted in accordance with criteria set forth by the Declaration of Helsinki. The UK Biobank (UKB) resource was approved by the UK Biobank Research Ethics Committee. Use of UKB data was conducted under application number 17488 and was approved by the Mass General Brigham (MGB) Institutional Review Board. Use of the All of Us resource was approved by the All of Us Institutional Review Board, and the work performed in this study was approved under a data use agreement between the Massachusetts General Hospital and the All of Us program. Use of the MGB Biobank was approved by the MGB Institutional Review Board (protocols 2009P002312 and 2023P001908). Participants in all three biobanks provided written informed consent.
Mice used in this study were male and female 8–12 week old F12+/+, F12+/−, and F12−/− littermates on a C57Bl6/J background. All animal studies were approved by the UNC Chapel Hill Institutional Animal Care and Use Committee (IACUC) under protocol number 23-196. Mice were group housed in environmentally enriched individually ventilated cages on a standard light/dark cycle with food and water provided ad libitum. Animal welfare was monitored by trained technicians with access to a certified veterinarian. Mice were euthanized under terminal anesthesia by exsanguination and cervical dislocation.
Multiomic datasets
The UK Biobank
The UK Biobank (UKB) is a national biorepository program containing data from approximately 500,000 participants enrolled between 2006 and 2010 in the United Kingdom50. The UKB includes individual-level whole exome sequencing and array genotyping data as well as extensive clinical data, including laboratory results, diagnosis codes, and procedure codes. Additionally, Olink® Explore 3072 data are available for about 50,000 participants.
For exome sequencing, the revised version of the IDT xGen Exome Research Panel V.1.0 was used on Illumina NovaSeq 6000 instruments, achieving over 20X coverage at 95% of sites. The procedures for sequencing, alignment, variant calling, and joint genotyping have been previously described51,52. This study used the OQFE exome call set, adhering closely to a previously described quality control pipeline31. Briefly, low-quality genotypes were set to “missing,” and variants with a < 90% call rate, failed Hardy-Weinberg equilibrium test result (P < 1 × 10−15), or presence in a low-complexity region were removed. Sample-level quality control included the removal of samples that were duplicates, had mismatches between exome sequencing and genotyping array data, had mismatches between genetically-inferred and self-reported sex, had low call rates, or were outliers (outside 8 standard deviations from the mean) for several additional metrics. Samples from participants who had withdrawn consent were also removed. Quality control of individual level data was performed using Hail version 0.2 (hail.is) and PLINK version 2.0.a (www.cog-genomics.org/plink/2.0/).
NIH All of Us
The National Institutes of Health All of Us (AoU) Research Program dataset contains approximately 400,000 participants with whole-genome sequencing and linked electronic health record data. The ultimate goal of the program is to recruit one million adult participants (age ≥ 18 years) across a diverse cross-section of the US population53. Sequencing, variant calling, and quality control were performed as previously described54. Briefly, sequencing was conducted on Illumina NovaSeq 6000 instruments following manufacturer-defined best practices. Variant calling was performed using Illumina’s DRAGEN pipeline (version 3.4.12), harmonized between different AoU Genome Centers. A stringent central QC procedure was applied, as described elsewhere55.
Mass General Brigham Biobank
The Mass General Brigham (MGB) Biobank is currently enrolling participants across the Mass General Brigham health system in and around Boston, MA, and contains approximately 53,000 participants with whole-exome sequencing and linked electronic health record data, as well as blood samples available upon request for a subset of participants. Samples were sequenced on Illumina NovaSeq devices using a custom exome panel (TWIST Human Core Exome), with a target depth of at least 20X coverage at >85% of sites. Sequence alignment, processing, and variant joint-calling were performed using the Genome Analysis ToolKit (GATK v4.1), following GATK best practices, after which a stringent QC pipeline was applied as described above (“UK Biobank”).
Variant annotation and functional impact score (FIS)
Variants were assigned a minor allele frequency (MAF) based on the highest ancestry-associated minor allele frequency (MAF) value in the Genome Aggregation Database (gnomAD) v2.1.1, restricted to major outbred continental superpopulations: European, African, South Asian, East Asian, and Admixed-American56,57. The Loss-of-Function Transcript Effect Estimator (LOFTEE)56 plug-in implemented in Variant Effect Predictor (VEP) v.10558 was used to identify high-confidence loss-of-function (HCLOF) variants that are the most likely to damage protein activity, including those that result in truncation (nonsense), frameshift, and essential splice site disruption. We removed any HCLOF variants flagged by LOFTEE as questionable, e.g. variants in poorly conserved exons or those in tandem acceptor (alternative splicing) or noncanonical splice sites.
For analyses including variant “functional impact scores” (FIS), the protein-level consequence of each variant was determined using dbNSFP v4.3a59 For missense variants, 30 in silico prediction tools included in the dbNSFP database were used to assign each variant a composite FIS between 0 and 1 corresponding to the percentage of tools that predicted a deleterious effect31. A higher FIS indicates that a variant is more likely to damage protein activity, with HCLOF variants assigned an FIS of 1.0. Variants were depicted in lollipop format using the trackviewer and lolliplot packages implemented in R version 4.2.3.
Phenotype definitions
For the UKB and AoU datasets, curated disease phenotypes for venous thromboembolism (VTE), bleeding, and sepsis were crafted from ICD-9 and -10 codes (Data S3)39. Participants with at least one code corresponding to the inclusion criteria for each phenotype were considered cases; those who met inclusion criteria but also met exclusion criteria at an earlier date were excluded.
Ancestry and relatedness definitions
Genetically-defined ancestry was determined through principal component analysis (PCA) for all modeling analyses, with the first 10 principal components (PCs) of ancestry included as covariates. Ancestry categorizations presented in the summary data (European, East Asian, South Asian, African, Admixed American, Middle Eastern, and Other) were defined using ADMIXTURE as previously described60. Briefly, ADMIXTURE models were trained using 87,398 variants identified in 2504 samples from the 1000 Genomes Project, which included whole genome sequencing data and known global ancestries. Likelihood estimations based on these variants were performed on UKB and AoU samples using genotyping array data. For sensitivity analyses restricted to unrelated individuals, we excluded third-degree or closer relatives. To identify the unrelated subset of participants, the KING-robust algorithm was used to compute pairwise kinship estimates for all participants and pairs with kinship coefficients ≥0.0442 were considered related. Individuals related to multiple others were iteratively removed until no related pairs remained, and in each remaining pair one participant was removed at random, leaving 543,262 participants after accounting for missing data31.
Genetic association analyses
We tested for associations between the burden of rare, germline variants in F12 (FIS = 1.0, HCLOF) and the occurrence of phenotypes of interest. Rare variants were defined as those with a global minor allele frequency (MAF) of ≤1% in the Broad Institute’s Genome Aggregation Database (gnomAD)57. Rare variants classified as HCLOF were used in our analyses unless otherwise noted. Except where otherwise indicated, the associations between variant burden (i.e., number of qualifying variants in F12) and phenotypes of interest were assessed using Cox regressions with Firth’s penalized likelihood correction, an approach that accounts for case-control imbalance, using the coxphf package in R version 4.0 (UKB) or 4.4 (AoU). The presence of qualifying variants in F12 was collapsed into a single variable, and our regression models adjusted at minimum for age at enrollment, sex, sequencing batch, and principal components 1–10 of genetic ancestry, with additional covariates included as indicated. Models included only participants with data available for all tested covariates. Meta-analyses were performed using summary statistics derived from sub-analyses and incorporated into a random-effects model using the meta package in R version 4.0.
A leave-one-variant-out analysis to determine the impact of individual variants was performed by iteratively removing carriers of each unique F12 HCLOF variant and performing Firth’s logistic regressions (logistf package in R 4.0 in the UKB and 4.4 in AoU) comparing the risk of VTE among carriers of the remaining variants vs. non-carriers. These models adjusted for age, sex, and the first 4 principal components of genetic ancestry. Outliers were identified using the two-sided extreme studentized deviate (Grubbs) test in Prism version 10.4.1.
For sensitivity analyses including the factor V Leiden (rs6025) and prothrombin G20210A (rs1799963) variants in the UKB, carrier status was obtained using directly sequenced (rs6025) or imputed data (rs1799963) from the UKB Affymetrix Axiom® genotyping array as described elsewhere50; carrier status was obtained from WGS data in the AoU dataset. For sensitivity analyses using polygenic risk scores (PRS) for VTE, the standard PRS for venous thromboembolic disease was used in the UKB (UKB field 26289). A custom VTE PRS was generated in AoU using PLINK 2.0 with variant weights obtained from Polygenic Score Catalog ID PGS00179661.
For the assessment of discrete infection phenotypes within the UKB dataset, associations between F12 variant carrier status and 138 infection-related phecodes (https://phewascatalog.org/phewas/#phex) were evaluated by serial Firth’s logistic regression models with adjustment for age, sex, and the first 4 principal components of genetic ancestry.
Multiple linear regression modeling was performed using the glm package implemented in R version 4.0 with adjustments as indicated.
Kaplan–Meier analysis
For Kaplan–Meier analyses, case/control status was determined at the time of last follow-up. Age at disease onset was defined as the earliest of either (1) the first appearance of a qualifying billing code in the electronic medical record or (2) age at second or subsequent visit if the condition was identified during a UKB visit. Individuals with phenotypes identified on the date of the first (baseline) UKB visit were excluded. For our VTE analysis, we included only incident disease and excluded prevalent (historical) events occurring prior to the date of enrollment. For our mortality analysis, analysis began with participant date of birth. Survival curves were drawn using GraphPad Prism version 10.1.2, with P values computed by the log-rank method. Hazard ratios were computed using a univariate (unadjusted) Cox proportional hazards model with the survival R package (VTE analysis) or the Mantel-Haenszel method implemented in Graphpad Prism v.10.4.1 (mortality analysis).
Human plasma measurements
EDTA-anticoagulated plasma from F12 variant carriers and age- and sex-matched controls were obtained from the MGB Biobank. Plasma levels of FXII antigen were determined by ELISA (IHUFXIIKTT, Innovative Research, Novi, MI) in technical duplicates. In separate experiments, congenital FXII deficient platelet poor plasma (George King Biomedical, Overland Park, KS) was reconstituted with varying levels of FXII zymogen (Prolytix, Essex Junction, VT). Contact pathway-initiated thrombin generation was evaluated using calibrated automated thrombography as previously described62. In brief, in technical duplicates per condition, 10 μL of a trigger solution containing silica (1:24000 final, Kontact, Pacific Hemostasis, Waltham, MA) or tissue factor (5 pM final, Dade Innovin, Siemens, Munich, Germany) and phospholipid (4 μM final concentration, Synapse, Maastricht, Netherlands) or calibrator (Stago, Parsippany, NJ) was added to 40 μL of plasma followed by addition of 10 μL of a FluCa substrate solution (Stago, Parsippany, NJ). Fluorogenic substrate cleavage was monitored using a microplate reader (FluoroskanAscent, Thermo Fisher Scientific, Waltham, MA) with data recorded and analyzed using Thrombinoscope software (v5, Thrombinoscope, Maastricht, The Netherlands). Tissue factor-initiated reactions were conducted in the presence of corn trypsin inhibitor (50 μg/mL final, Prolytix, Essex Junction, VT). Data were evaluated using ANOVA followed by Dunnett’s post-test for comparisons against the 100% FXII condition.
FXIIa activity assay
EDTA-anticoagulated plasma from ESS1 variant carriers and F12 wild-type controls were diluted 1:10 in Hepes buffered saline, pH 7.4 (HBS). Separately, a stock consisting of S-2302 substrate (681 µM, Diapharma), soybean trypsin inhibitor (1.43 µM, Sigma), apxiban (21.76 µM, Cayman Chemical), hirudin (2 units/ml, Aniara), and EDTA (25 mM) was made. Each diluted plasma sample (50 µl) was added to a clear 96-well non-treated polypropylene plate (Greiner Bio-one, cat#: 655201), together with 50 µl of substrate stock (100 µl per well total). Each sample was run in triplicate. FXIIa activity was then recorded as the change in absorbance at 405 nm over one hour at 37 °C, averaged over 3 wells. A standard curve was made by diluting 1, 3, 10, 30, and 100 nM of purified FXIIa (Enzyme Research Laboratories) in 100 µl of HBS, with substrate cleavage measured in the same fashion. Absorbance slopes from the known concentrations of FXIIa were then used to determine the concentrations of FXIIa present in the plasma samples.
Coagulation assays
Prothrombin (PT) and activated partial thromboplastin (aPTT) clotting times were performed on a Stago Start 4 hemostasis analyzer in accordance with the manufacturer’s instructions. The PT and aPTT assays utilized the Neoplastine® CI Plus and PTT-A reagents, respectively.
Murine blood and plasma measurements
Whole blood was collected from male and female 8–12 week old F12+/+, F12+/- and F12−/− littermates by injection of 200 μL of 3.8% sodium citrate (Ricca Chemical Co, Arlington, TX) into the inferior vena cava followed by collection of 600 μL of whole blood. Complete blood counts were determined using an automated analyzer (Element HT5, Heska, Loveland, CO). Platelet poor plasma was generated by centrifugation of whole blood at 4500 × g for 15 min at room temperature. Plasma FXII antigen levels were determined by ELISA (IMSFXIIKTT, Innovative Research, Novi, MI) and by western blotting (anti-FXII primary antibody, Affinity Biologicals).
Mouse femoral vein electrolytic injury model
Male 8–12 week old F12+/+, F12+/−, and F12−/− littermates were subjected to a femoral vein electrolytic injury model of venous thrombosis24. Male mice were used exclusively to ensure consistency in vessel diameter. Anesthetized mice were administered Alexa488-labeled anti-GPIX antibody (4 μg/mouse, Emfret Analytics, Germany) and Alexa647-labeled anti-fibrin antibody (2 μg/mouse, clone 59D8, in-house) intravenously to label platelets and fibrin respectively. A 100 μm diameter stainless steel wire was used to apply a 1.5 V direct current at 0.02 A generated by a linear power supply (DP832, Rigol, Beavertown, OR) to the ventral surface of the femoral vein for 30 s to induce an electrolytic injury. Accumulation of fluorescently labeled platelets and fibrin was monitored by intravital videomicroscopy using a stereo microscope (SMZ25, Nikon, Tokyo, Japan) coupled to a digital camera (ORCA Flash 4.0, Hamamatsu, Japan). Data were analyzed using NIS-Elements software (Nikon, Tokyo, Japan). The operator (D.S.P.) was blinded to animal genotype.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Access to individual level genetic and phenotypic UK Biobank data is available to approved researchers through application on the UK Biobank website [www.ukbiobank.ac.uk] and the final release of the UK Biobank exome sequencing dataset is available on the DNAnexus Research Analysis Platform [www.ukbiobank.ac.uk/enable-your-research/research-analysis-platform]. More information about registration for data access can be found at www.ukbiobank.ac.uk/register-apply/. Use of UK Biobank Data was performed under application number 17488. Access to genetic and phenotypic data from All of Us is available to United States-based researchers through the All of Us Researcher Workbench [www.researchallofus.org/register/]. A publicly available data browser can be found at databrowser.researchallofus.org. Our work with the All of Us dataset was approved under a data use agreement between the Massachusetts General Hospital and the All of Us program. Access to individual-level data from the Mass General Brigham Biobank is not currently freely available to researchers outside of the Mass General Brigham healthcare system. Other datasets used in this manuscript include the dbNSFP database [sites.google.com/site/jpopgen/dbNSFP], v.4.2a and v.4.3a, gnomAD exomes v.2.1 [gnomad.broadinstitute.org/downloads], and Ensembl release 105 [www.ensembl.org/info/data/index.html]. A publicly available repository of UK Biobank genotype/phenotype associations can be found at https://hugeamp.org:8000/research.html?pageid=600_traits_app_home. Source Data are provided as a Source Data file. Source data are provided with this paper.
References
Ortel, T. L. et al. American Society of Hematology 2020 guidelines for management of venous thromboembolism: treatment of deep vein thrombosis and pulmonary embolism. Blood Adv. 4, 4693–4738 (2020).
Agnelli, G. et al. Apixaban for the treatment of venous thromboembolism associated with cancer. N. Engl. J. Med. 382, 1599–1607 (2020).
Granger, C. B. et al. Apixaban versus warfarin in patients with atrial fibrillation. N. Engl. J. Med. 365, 981–992 (2011).
Connolly, S. J. et al. Apixaban in patients with atrial fibrillation. N. Engl. J. Med. 364, 806–817 (2011).
Investigators, E.-P. et al. Oral rivaroxaban for the treatment of symptomatic pulmonary embolism. N. Engl. J. Med. 366, 1287–1297 (2012).
Patel, M. R. et al. Rivaroxaban versus warfarin in nonvalvular atrial fibrillation. N. Engl. J. Med. 365, 883–891 (2011).
Iyer, G. S., Tesfaye, H., Khan, N. F., Zakoul, H. & Bykov, K. Trends in the Use of Oral Anticoagulants for adults with venous thromboembolism in the US, 2010–2020. JAMA Netw. Open 6, e234059 (2023).
Carney, B. J. et al. Anticoagulation in cancer-associated thromboembolism with thrombocytopenia: a prospective, multicenter cohort study. Blood Adv. 5, 5546–5553 (2021).
Brunton, N. E. et al. Delayed anticoagulation in venous thromboembolism: Reasons and associated outcomes. Res Pract. Thromb. Haemost. 5, e12500 (2021).
Srivastava, P. & Gailani, D. The rebirth of the contact pathway: a new therapeutic target. Curr. Opin. Hematol. 27, 311–319 (2020).
Nickel, K. F., Long, A. T., Fuchs, T. A., Butler, L. M. & Renne, T. Factor XII as a therapeutic target in thromboembolic and inflammatory diseases. Arterioscler. Thrombosis. Vasc. Biol. 37, 13–20 (2017).
Davoine, C., Bouckaert, C., Fillet, M. & Pochet, L. Factor XII/XIIa inhibitors: tTheir discovery, development, and potential indications. Eur. J. Med. Chem. 208, 112753 (2020).
Tillman, B. & Gailani, D. Inhibition of factors XI and XII for prevention of thrombosis induced by artificial surfaces. Semin. Thrombosis Hemost. 44, 60–69 (2018).
Loscalzo, J. Thrombosis and Hemorrhage, (Lippincott Williams and Wilkins, 2003).
Zeerleder, S. et al. Reevaluation of the incidence of thromboembolic complications in congenital factor XII deficiency-a study on 73 subjects from 14 Swiss families. Thromb. Haemost. 82, 1240–1246 (1999).
Girolami, A. et al. Comparable levels of activity and antigen in factor XII deficiency: a study of 21 homozygotes and 58 heterozygotes. Clin. Appl. Thromb. Hemost. 11, 335–338 (2005).
Renné, T., Schmaier, A. H., Nickel, K. F., Blombäck, M. & Maas, C. In vivo roles of factor XII. Blood 120, 4296–4303 (2012).
Stavrou, E. & Schmaier, A. H. Factor XII: what does it contribute to our understanding of the physiology and pathophysiology of hemostasis & thrombosis. Thrombosis Res. 125, 210–215 (2010).
Chaudhry, S. A. et al. Cationic zinc is required for factor XII recruitment and activation by stimulated platelets and for thrombus formation in vivo. J. Thromb. Haemost. 18, 2318–2328 (2020).
Renne, T. et al. Defective thrombus formation in mice lacking coagulation factor XII. J. Exp. Med. 202, 271–281 (2005).
Kleinschnitz, C. et al. Targeting coagulation factor XII provides protection from pathological thrombosis in cerebral ischemia without interfering with hemostasis. J. Exp. Med. 203, 513–518 (2006).
Larsson, M. et al. A factor XIIa inhibitory antibody provides thromboprotection in extracorporeal circulation without increasing bleeding risk. Sci. Transl. Med. 6, 222ra217 (2014).
Matafonov, A. et al. Factor XII inhibition reduces thrombus formation in a primate thrombosis model. Blood 123, 1739–1746 (2014).
Grover, S. P., Olson, T. M., Cooley, B. C. & Mackman, N. Model-dependent contributions of FXII and FXI to venous thrombosis in mice. J. Thromb. Haemost. 18, 2899–2909 (2020).
Key, N. S. Epidemiologic and clinical data linking factors XI and XII to thrombosis. Hematol. Am. Soc.Hematol. Educ. Prog. 2014, 66–70 (2014).
Koster, T., Rosendaal, F. R., Briet, E. & Vandenbroucke, J. P. John Hageman’s factor and deep-vein thrombosis: Leiden thrombophilia Study. Br. J. Haematol. 87, 422–424 (1994).
Cushman, M., O’Meara, E. S., Folsom, A. R. & Heckbert, S. R. Coagulation factors IX through XIII and the risk of future venous thrombosis: the Longitudinal Investigation of Thromboembolism Etiology. Blood 114, 2878–2883 (2009).
von Bruhl, M. L. et al. Monocytes, neutrophils, and platelets cooperate to initiate and propagate venous thrombosis in mice in vivo. J. Exp. Med. 209, 819–835 (2012).
Maas, C. & Renne, T. Coagulation factor XII in thrombosis and inflammation. Blood 131, 1903–1909 (2018).
Kenne, E. et al. Factor XII: a novel target for safe prevention of thrombosis and inflammation. J. Intern. Med. 278, 571–585 (2015).
Jurgens, S. J. et al. Analysis of rare genetic variation underlying cardiometabolic diseases and traits among 200,000 individuals in the UK Biobank. Nat. Genet. 54, 240–250 (2022).
Thompson, D. J. et al. UK Biobank release and systematic evaluation of optimised polygenic risk scores for 53 diseases and quantitative traits. medRxiv, https://doi.org/10.1101/2022.06.16.22276246 (2022).
Eldjarn, G. H. et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 622, 348–358 (2023).
Karlson, E. W., Boutin, N. T., Hoffnagle, A. G. & Allen, N. L. Building the partners HealthCare Biobank at partners personalized medicine: informed consent, return of research results, recruitment lessons and operational considerations. J. Pers. Med. 6, 2 (2016).
Boutin, N. T. et al. The evolution of a large biobank at mass General Brigham. J. Pers. Med. 12, 1323 (2022).
Schmaier, A. H. Antithrombotic potential of the contact activation pathway. Curr. Opin. Hematol. 23, 445–452 (2016).
Muller, F., Gailani, D. & Renne, T. Factor XI and XII as antithrombotic targets. Curr. Opin. Hematol. 18, 349–355 (2011).
Renné, T. & Stavrou, E. X. Roles of factor XII in innate immunity. Front. Immunol. 10, 2011 (2019).
Haj, A. K. et al. Loss of Function in Protein Z (PROZ) is associated with increased risk of ischemic stroke in the UK Biobank. J. Thromb. Haemost. 23, 171–180 (2024).
Beck, S. et al. Generation of a humanized FXII knock-in mouse-A powerful model system to test novel anti-thrombotic agents. J. Thromb. Haemost. 19, 2835–2840 (2021).
Dentali, F. et al. Non-O blood type is the commonest genetic risk factor for VTE: results from a meta-analysis of the literature. Semin. Thromb. Hemost. 38, 535–548 (2012).
Ohira, T. et al. ABO blood group, other risk factors and incidence of venous thromboembolism: the Longitudinal Investigation of Thromboembolism Etiology (LITE). J. Thromb. Haemost. 5, 1455–1461 (2007).
Prevention of pulmonary embolism and deep vein thrombosis with low dose aspirin: Pulmonary Embolism Prevention (PEP) trial. Lancet 355, 1295–1302 (2000).
Brighton, T. A. et al. Low-dose aspirin for preventing recurrent venous thromboembolism. N. Engl. J. Med. 367, 1979–1987 (2012).
Becattini, C. et al. Aspirin for preventing the recurrence of venous thromboembolism. N. Engl. J. Med. 366, 1959–1967 (2012).
Gailani, D. & Broze, G. J. Jr. Factor XI activation in a revised model of blood coagulation. Science 253, 909–912 (1991).
Lipov, A. et al. Exploring the complex spectrum of dominance and recessiveness in genetic cardiomyopathies. Nat. Cardiovasc. Res. 2, 1078–1094 (2023).
Heyne, H. O. et al. Mono- and biallelic variant effects on disease at biobank scale. Nature 613, 519–525 (2023).
Barton, A. R., Hujoel, M. L. A., Mukamel, R. E., Sherman, M. A. & Loh, P. R. A spectrum of recessiveness among Mendelian disease variants in UK Biobank. Am. J. Hum. Genet 109, 1298–1307 (2022).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
UK Biobank – Final Exome Data Release FAQs/July. https://www.ukbiobank.ac.uk/media/najcnoaz/access_064-uk-biobank-exome-release-faq_v11-1_final-002.pdf. accessed 1 March 2022.
U. K. Biobank WES Protocol / September 2021. https://biobank.ndph.ox.ac.uk/ukb/ukb/docs/UKB_WES_Protocol.pdf.
All of Us Research Program, I. et al. The “All of Us” Research Program. N. Engl. J. Med. 381, 668–676 (2019).
All of Us Beta Release Genomic Quality Report (ARCHIVED C2021Q3R6 CDR CT Dataset v5). https://support.researchallofus.org/hc/en-us/articles/7002036503700-All-of-Us-Beta-Release-Genomic-Quality-Report-ARCHIVED-C2021Q3R6-CDR-CT-Dataset-v5. Accessed 6 May 2024.
All of Us Genomic Quality Report. https://support.researchallofus.org/hc/en-us/articles/4617899955092-All-of-Us-Genomic-Quality-Report. accessed 6 May 2024.
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100 (2024).
McLaren, W. et al. The ensembl variant effect predictor. Genome. Biol. 17, 122 (2016).
Liu, X., Wu, C., Li, C. & Boerwinkle, E. dbNSFP v3.0: a one-stop database of functional predictions and annotations for human nonsynonymous and splice-site SNVs. Hum. Mutat. 37, 235–241 (2016).
Jurgens, S. J. et al. Adjusting for common variant polygenic scores improves yield in rare variant association analyses. Nat. Genet. 55, 544–548 (2023).
Wang, Y. et al. Global Biobank analyses provide lessons for developing polygenic risk scores across diverse cohorts. Cell Genom. 3, 100241 (2023).
Grover, S. P. et al. C1 inhibitor deficiency enhances contact pathway-mediated activation of coagulation and venous thrombosis. Blood 141, 2390–2401 (2023).
Acknowledgements
We thank the participants of the UK Biobank, NIH All of Us, and Mass General Brigham Biobank programs; this work would have been impossible without their generous contributions. We are grateful to Dr. John-Bjarne Hansen for his assistance with the project. We are also indebted to Drs. George Vassiliou and Sruthi Cheloor Kovilakam for their advice and assistance with the figures. This work was supported by funding from the Fondation Leducq (14CVD01), National Institutes of Health (R01 HL092577, R01 HL139731, R01 HL157635), American Heart Association (18SFRN34110082), and European Union (MAESTRIA 965286) awarded to P.T.E.; National Institutes of Health (1R01HL166246 and R03 HL162761) and the CSL Behring Heimburger Award to P.K.B.; National Institutes of Health (R01HL171042), and American Society of Hematology (Scholar Award) to S.P.G.; National Institutes of Health (R35HL144976) to W.B.; an Amsterdam UMC doctoral fellowship and the Junior Clinical Scientist Fellowship (03-007-2022-0035) from the Dutch Heart Foundation awarded to S.J.J.; and the BioData Ecosystem fellowship awarded to S.H.C.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.K.H., P.K.B., S.P.G., A.B. and P.T.E. Methodology: A.K.H., J.T.R., S.J.J., X.W., S.H.C., L-C.W., X.W. and S.K. Multiomic Investigations: A.K.H., S.J.J., A.R.E, S.C. and P.K.B. Animal studies: D.S.P., S.P.G., W.B., and N.M. In vitro studies: H.E., K.B., L.F., J.F., S.P.G., W.B., N.M. and P.K.B. Formal analysis: A.K.H., P.K.B., S.J.J., X.W., S.C., J.R., A.R.E. and S.H.C. Visualization: P.K.B. and A.K.H. Funding acquisition: P.T.E. and P.K.B. Supervision: P.K.B. and P.T.E. Writing – original draft: P.K.B., S.P.G. and A.K.H. Writing – review and editing: S.J.J., S.P.G., A.K.H., P.K.B., and P.T.E.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sarah Beck, Jeffrey Weitz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Haj, A.K., Paul, D.S., Jurgens, S.J. et al. Coagulation factor XII haploinsufficiency is protective against venous thromboembolism in a population-scale multidimensional analysis. Nat Commun 16, 8176 (2025). https://doi.org/10.1038/s41467-025-62789-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62789-5