Abstract
Recent lineage tracing based single-cell techniques (LT-scSeq), e.g., the Lineage And RNA RecoverY (LARRY) barcoding system, have enabled clonally resolved interpretation of differentiation trajectories. However, the heterogeneity of clone-specific kinetics remains understudied, both quantitatively and in terms of interpretability, thus limiting the power of barcoding systems to unravel how heterogeneous stem cell clones drive the overall cell population dynamics. Here, we present CLADES, a NeuralODE-based framework to faithfully estimate the clone and population-specific kinetics from both newly generated and publicly available LARRY LT-scSeq data. By incorporating a stochastic simulation algorithm (SSA) and differential expression gene (DEGs) analysis, CLADES yields the summary of cell division dynamics across differentiation time-courses and reconstructs the lineage tree of the progenitor cells in a quantitative way. Moreover, clone-level behaviors can be grouped into characteristic types by pooling individual clones into meta-clones for analyses at various resolutions. Finally, we show that meta-clone specific cellular behaviors identified by CLADES originate from hematopoietic stem and progenitor cells in distinct transcriptional states. In conclusion, we report a scalable approach to robustly quantify clone-specific differentiation kinetics of cellular populations for time-series systems with static barcoding designs.
Similar content being viewed by others
Introduction
One of the fundamental challenges in developmental biology is to understand the complicated cellular dynamics in a temporal order1,2. The interplay between cell proliferation (cellular expansion) and differentiation (phenotypic transition) plays an essential role in various biological processes, e.g., tissue development, regeneration, and the activation of innate immune response mechanisms3.
In recent years, the transcriptome-wide single-cell RNA sequencing (scRNA-seq) technique has been widely used as a scalable approach for studying cellular trajectories, either using a snapshot of transitioning cell populations or via a time-series design. Computational algorithms for dynamical analyses include pseudotime-based trajectory inference4,5,6,7,8, unspliced RNA-based RNA velocity methods for predicting temporal gene expression changes9,10,11 and metabolic labeling-based protocols for analyzing temporal dynamics, where nascent RNA is labeled with 4sU12,13,14.
However, scRNA-seq alone cannot offer fine-grained insights into the clone-level heterogeneity within cell clusters. Therefore, emerging lineage tracing techniques that utilize unique and inheritable DNA barcodes to track individual cells can offer a complementary approach to study cellular dynamics1,3,15.
Techniques of lineage-tracing coupled with single-cell sequencing (LT-scSeq) include retrospective analyses via endogenous genetic barcodes (e.g., mitochondrial variants and copy number variations16,17,18,19,20) and prospective designs with recently developed barcoding technologies via exogenous barcodes. Prospective designs can be broadly categorized into static barcoding with one-off induction to focus more on the clone-specific differentiation21,22, and dynamical barcoding with inductions at multiple time points to trace phylogenies23,24. In this study, we focus primarily on modeling static barcoding LT-scSeq data, where numerous progenitor cells are labeled with a unique barcode at an early point; then the barcode is propagated to all other populations (progeny from here on), hence facilitating the delineation of a high-resolution differentiation topology. One prominent example of this type of technology is the lentivirus-based system25,26 LARRY22, which has been recently employed, for example, to predict clonal fate bias in hematopoiesis22 and mouse brain formation27,28, unveiling new regulators/markers involved in reprogramming29, cell differentiation30, and identifying pathways relevant to cancer progression31,32.
Several computational algorithms have recently been proposed to analyze the population dynamics and LT-scSeq experiments. Depending on the space of the cell states, these methods can be grouped into different categories. First, the continuous models, such as Fischer et al.33, who developed Pseudodynamics, which models population distribution shifts to quantify developmental potentials for time-series data. Second, a common choice is to mimic a near-continuous space by employing a finite state mapping where each cell is treated as a state, e.g., LineageOT34, which maps cells from the same clone and adjacent time points using optimal transport; it was developed to recover lineage couplings in CRISPR-based lineage tracing datasets and outperforms the original OT method that operates without barcodes35. Another example is CoSpar36, which performs topology mapping under the constraints of sparsity and coherence. This method brought new insights to multi-clonal time series data analysis with respect to the identification of early cell fate bias at the cellular level. Third, discrete state space over cell types is also commonly used to ensure better interpretability, higher robustness, and computational efficiency37. Finally, other approaches have also significantly contributed to this field using various modalities, for example, by learning a smoothed transcription and regulatory dynamics38, analyzing the potency bias during HSC reactivation following platelet depletion39 or utilizing epimutation on DNA methylation for in vivo lineage tracing experiments40.
Despite various efforts to unveil more information from LT-scSeq data, several technical challenges persist; in addition to common issues such as the loss of barcodes or small clone size, scRNA-seq is destructive and only captures a fraction of the total cells, resulting in the possibility of tracking only the relative changes of cell state abundance36 rather than the actual kinetic rates that affect the overall population dynamics. This hampers the exploration of clone-specific dynamical patterns. Furthermore, having reliable kinetic rates enables the reconstruction of differentiation topologies and provides a quantitative estimation of division numbers39 as well as the likelihood of producing specific lineage outputs from a progenitor cell, which is useful under various scenarios, e.g., investigating the patterns related to differentiation, aging or disease.
Inspired by recent research efforts, and to address the aforementioned issues, we focused on the static barcoding system, (the LARRY, Fig. 1a) and developed a robust and generalizable algorithm to analyze LT-scSeq datasets, named as CLADES (Clonal Lineage Analysis with Differential Equations and Stochastic simulations), which comprises two key components: 1) a model estimator, based on NeuralODE41, to delineate clone-specific trajectories and state-dependent transition rates; 2) a data generator, via the Gillespie algorithm42, to simulate the differentiation topologies between progenitors and progenies. In general, for both components, CLADES uses the design of meta-clone-specific dynamics (see below) to handle many clones with few cell counts. CLADES also uses scaling factors and Poisson negative likelihood loss to efficiently handle the problems of barcode dropouts during the experiment (Methods).
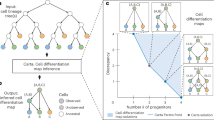
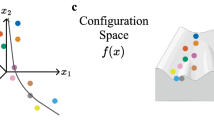
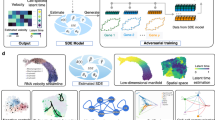
a General workflow of LT-scSeq experiment using static barcoding techniques with viral integration. DNA barcodes are induced at an early time point, then scRNA-seq data and clonal information are acquired at subsequent time points. b CLADES takes total cell counts and transition directions as input, then uses a neural net to estimate the transition rates between populations, and an ODE module to reconstruct cell counts at each time point. c CLADES is able to infer the dynamic changes of population size on various resolutions and the confidence interval of transition rates. d With the estimated kinetic rates of each clone, CLADES can further: 1) simulate detailed topologies of division and differentiation, and 2) infer the meta-clone level probability of lineage realization. e,f Given cell counts generated by either time-invariant or time-variant rates, the performance of both constant and dynamic mode are validated on test time points. e The constant mode (i) consistently performs better than the dynamic mode (ii) no matter how many time points were used to train the model; the robustness of the constant mode given time-invariant datasets was evaluated as well (iii). f the constant mode (i) performs worse than the dynamic mode (ii) except using only 2 training time points; the robustness of the dynamic mode given time-variant datasets was evaluated as well (iii). Mean_n is the mean value for all the test sets given n training time points. Based on the results of the above analysis, the minimum recommended time points for CLADES to have satisfactory performance is 3 (including the induction time point, which can be inferred from the data itself, Methods). Panel (a) and (d) are created in BioRender. Huang, Y. (2025) https://BioRender.com/ak99vwt.
Collectively, CLADES determines clone-specific dynamics and provides a quantitative description of the differentiation topology between a progenitor and its progeny.
Results
ODE function for (meta-) clone-specific dynamics
We model clone dynamics as a system of independent ordinary differential equations (ODEs) to infer the time-specific transition rates between cell states. The dynamics of each clone (or each meta-clone, namely a group of clones with similar dynamical profiles; see below) are described by the same equation but clone-specific parameters; this aligns with the assumption that only intra-clone transitions are allowed (cells keep the same clone identity throughout the experiment). Therefore, the estimation and inference processes are in parallel for each (meta-) clone. Without loss of generality, we describe the (multivariate) ODE function f for a specific clone c with parameters θ as shown in Eq.(1),
where ⋅ is the dot product of two matrices, ⊙ is element-wise multiplication, x(t) is a tensor of total counts for all cell states (interchangeably as cell populations), which has a dimension of (time-points, meta-clones, populations) and t refers to the real time of the biological system. K1(t) mimics the differentiation among populations and is based on the edges of the PAGA43 graph (denoted as L, Methods) with expert curation; it is non-negative and strictly upper triangular (see Supplementary Fig. S1a,b). K2(t) (the diagonal of the matrix L) is a one-dimensional vector, representing the net proliferation rate for each cell state. Generally, K1,2(t) can either be constant, which gives the classic population growth model with exponential change of population sizes, or a function of the real time t (Fig. 1b), which allows for more flexibility in our model44,45,46.
To truly estimate the expansion potential of different clones, relying on just the clusters’ relative proportion over time is not enough, as a cluster expanding in relative proportion over time could still result in a decreasing population, only with a slower shrinking rate compared to the other clusters, which would affect the accuracy of estimated rates. We solved this issue by measuring and then factoring in the culture’s expansion, following these steps at each time point: we measured the cells in the culture, then split a known fraction and sequenced the rest. We can thus obtain the real size of a clone (total cell counts from here on) after multiplying the relative sizes of the clusters by a scaling factor (which is a time-specific parameter) that also considers the number of cells lost during quality control (QC) in the sequencing protocol (Methods).
CLADES is designed for time-series lineage tracing experiments and mainly requires two types of input data: 1) the estimated total cell counts x(t) per time point ti, clone cj and population pk; 2) putative transition directions between populations, usually derived from the PAGA graph (Supplementary Note 1) with expert curation, which brings prior knowledge into the model (Fig. 1a, b). Under the common assumption that cellular divisions and differentiations between distinctive states are a stochastic process governed by a set of transition rates, we then interpolate the total cell counts on un-sequenced time points using NeuralODEs with biologically informed constraints (Fig. 1b, Methods). The underlying logic is solving an optimization problem between observations and model predictions with meaningful penalties.
CLADES takes the input data and feeds them into a multi-layer perceptron (MLP, Methods) with 2 layers, from which the model outputs the rate transition matrices (Supplementary Note 2) among populations and the predicted cell counts using an ODE solver. The rates can be either time-invariant or time-variant at the user’s discretion. After acquiring the rates, CLADES can reconstruct the dynamic changes of cell counts at both the population and the clone levels, and derive the associated confidence intervals (CI) of the kinetic rates (Fig. 1c) as well, providing a measure of uncertainty. By means of the Gillespie algorithm (Supplementary Note 3, Methods), CLADES can also provide a quantitative summary of division topologies and assess the probability of realizing the different fates of any progenitor cell type (Fig. 1d).
Performance and robustness of CLADES on synthetic datasets
In order to assess both the performance of the constant and the dynamic mode of CLADES, we used synthetic datasets as described in Supplementary Note 4.
We applied CLADES on a synthetic dataset governed by time-invariant rate ODE functions first, where the rates for each clone serve as the ground truth and cell counts are governed by Eq.(1). To test whether CLADES can recover the correct dynamics and provide guidance for appropriate usage with the LT-scSeq experiments, we conducted 4 independent trials with different sampling intervals as training sets and used the same 5 unobserved time points as testing sets, for both the constant and the dynamic modes of the model (Supplementary Table S1).
Generally, the performance of CLADES (Methods) improved along with the number of training time points, Fig. 1e (i, ii). Since the cell counts are the product of the transition rates, we observed similar trends in the absolute error of transition rates and the correlation of cell counts (Supplementary Fig. S2a). And the constant mode consistently performs better than the dynamic mode; this is reasonable as the synthetic data was generated based on the time-invariant rates, indicating the risk of overfitting for the dynamic mode when the underlying pattern of data is clean and simple (Fig. 1e). We also assessed the robustness of CLADES given datasets with different synthetic noise levels (Methods, Supplementary Note 4). Here we used trial index #3 (Supplementary Table S1) as an example, and we could observe that the constant mode performs fairly well if the noise level is smaller than 10, and gradually the performance deteriorates with a noise level higher than 10, Fig. 1e (iii).
With the time-variant scenario, we aim to demonstrate the limitations of the constant mode when facing complex data structures and highlight the flexibility and robustness of the dynamic mode. Specifically, similar synthetic settings were adopted (Supplementary Table S2), whilst this time the data was generated using time-variant rates. For both the constant and the dynamic modes, the performance increased when more training data were given, Fig. 1f (i, ii). However, the overall performance and additional evaluation metrics of the constant mode are always worse (Supplementary Fig. S2b). Interestingly, we also noticed that, apart from outperforming the constant mode when applied to its paired data (Fig. 1f (i, ii)), the dynamic mode generally yields better results even when it is applied to data generated via the other mode, Fig. 1e (ii) and Fig. 1f (i), suggesting that the dynamic mode is more robust to the ’unpaired patterns’. The performance of CLADES under different noise levels was assessed as well, (trial index #3, Supplementary Table S2). As shown in Fig. 1f (iii), the dynamic mode retains a recovery rate > 80% if the noise level is smaller than 15, and the performance gradually decreases after that.
In conclusion, for datasets with relatively simple patterns, the constant mode is suitable with tolerable errors, whilst for other biological systems that are more sophisticated, the dynamic mode of CLADES is preferred.
Characterizing the (meta-) clone-specific kinetic rates in human cord blood
We applied CLADES to a newly generated in vitro LARRY-based LT-scSeq data with three time points (24,885 barcoded out of 68,856 high-quality cells; see details in Supplementary Table S3 and Supplementary Note 5). The DNA barcodes were induced at Day 0 in CD34+ hematopoietic stem and progenitor cells (HSPCs) and sampled for sequencing at Day 3, 10 and 17 (Fig. 2a, Fig. 2b right). Upon culture, this pool of HSPCs differentiated into 12 distinct cell populations of progenitor or mature blood cells (Fig. 2b left), across the erythroid, myeloid, dendritic cell (DC) and mast cell (MC) lineages.
a Schematic of the data collection procedure for the LARRY human cord blood data. The total cell count is a 3-dimensional data. b UMAP illustration of the dataset, which contains 12 populations and 3 time points (HSC/MPP - Hematopoietic stem cell/Multipotent progenitor, DC- dendritic cell, MEMP - Megakaryocyte Erythroid and Mast cell Progenitors, NMP - Neutrophil monocyte progenitor). c Heatmap of the cell counts in log2 scale between populations and collected clone barcodes. d UMAP of the distributions of each meta-clone on the same landscape. NA represents non-barcoded cells. The meta-clones are formed using the population-time-specific kinetics, which is related to (c). e Upper panel: Proliferation and differentiation rates inferred by CLADES are generally less than 2, which aligns with the prior knowledge. Lower panel: the weighted mean of transition rates of all meta-clones combined resembles the rates of the whole biological system. f Model fitting results (dynamic mode) for meta-clones 2 and 7. The red line is the fitted curve, the gray line is the median, and the light blue region represents the 25–75% quantile. g Examples of inferred transition rates between populations of meta-clones 0, 2, and 7 at Day 3, which is the graphical illustration of kij in ODE functions (rows: starting cell states; columns: terminal cell states). h An example of the 95% confidence interval (CI) of the estimated transition rates, given by the bootstrapping approach. Blue dashed lines are CI, whilst the red dashed line is the original fitted value. i Left panel: Relative proportions of differentiated cell types of the terminal states at day 17 in meta-clones 2 and 7. Erythroid fate calculated as the sum of Early, Mid and Late Erythroid cells in each meta-clone. Right panel: Comparison of inferred transition rates for meta-clones 2 and 7 at different time points. Only values for which the difference between the two meta-clones is statistically significant are shown. Panel a is created in BioRender. Huang, Y. (2025) https://BioRender.com/wmlhrnj.
DNA barcodes with few cell counts make the analysis of individual clones infeasible (barcode matrix is shown in Fig. 2c and Supplementary Fig. S3); thus, to further reduce stochasticity and model complexity, clones were clustered into meta-clones (Supplementary Fig. S4) based on the similarity between time and population-specific counts of barcoded cells (Supplementary Fig. S5, Methods). The assumption is that hematopoiesis can be conceptualized as a clonal process, where clones have a finite number of differentiation behaviors; clones within a meta-clone will have similar kinetic rates and differentiation outcomes. As the initial barcoded HSPCs are heterogeneous, including both multi-potent and more lineage-restricted progenitors, we expect our approach to retrieve meta-clones with distinct kinetics of differentiation, being initiated by specific subsets of HSPCs. Indeed, we found 12 meta-clones with distinct outputs to four terminal fates (Mast cell, Late Erythroid, Monocyte and DC) (Fig. 2d, Supplementary Fig. S6 and Table S5). As an example, meta-clone 7 predominantly produces the Monocyte lineage, which originates from the most primitive HSC/MPP1 subpopulation, whilst meta-clone 0 predominantly produces cells of the mast cell and Erythroid lineage and correspondingly is initiated by Megakaryocyte Erythroid and Mast cell Progenitors (MEMP).
Given expert-curated putative transition directions derived from PAGA (Supplementary Fig. S1b), CLADES successfully predicted the total cell counts at the experimental time points (see Supplementary Table S4) and interpolated counts on unknown time points along the entire trajectory using both constant and dynamic modes while also providing associated estimation errors (Fig. 2f, Supplementary Fig. S7). Notably, the dynamic mode proved to have better performance provided that proper constraints are enforced to prevent model over-fitting.
As stated in equation (1), the population balance between cell states is governed by transition rates, which are the per capita output within a unit of time. Of note, the maximum inferred proliferation rate (2.5 per day) is consistent with the cell-cycle not lasting less than 10 h, which is biologically reasonable; differentiation frequency does not exceed 2.5 events per day (Methods, Fig. 2e, upper panel). Moreover, the weighted average behavior of all meta-clones should resemble the overall dynamics of the system (including both the barcoded and non-barcoded cells, background from here on, or BG for short, Fig. 2e, lower panel). CLADES calculates a transition rate matrix among cell states for any meta-clone at any time stamp (Fig. 2g).
As the transition rates are point estimates, CLADES uses bootstrapping (Methods) to estimate the 95% CI of the transition rates (two examples in Fig. 2h and complete results in Supplementary Fig. S8) and statistical tests to assess the significance of dynamical rates among meta-clones (Methods, Supplementary Fig. S9). Moreover, we only consider rates inferred for populations that have at least four cells at the respective time point. Interestingly, distinct meta-clones exhibit differences in rates at specific stages of progenitor/precursor maturation. As an example, here we compare two meta-clones, meta-clones 2 and 7, both originating in HSC/MPP 1, but which produce strikingly different differentiated output at day 17 (see pie charts of Fig. 2i, left panel). The differentiated output of meta-clone 2 consists of approximately 50% DCs and 40% monocytes. In contrast, the differentiated output of meta-clone 7 comprises less than 1% DCs and 85% monocytes. Importantly, CLADES estimates transition rates between distinct subpopulations that are consistent with the differentiation behaviors of meta-clones 2 and 7 (Fig. 2i, right panel), with, for example, a significantly higher transition rate from HSC/MPP 2 to DC progenitors on day 10 and day 17 in meta-clone 2 than in meta-clone 7 and conversely, significantly higher transition rates within the Monocytic compartment for meta-clone 7 than meta-clone 2 on Day 10 and 17 (Fig. 2i, right panel).
Resolving cell division history and fate realization of progenitor cells
After estimating transition rates between cell states using CLADES, we then employed the modified Gillespie algorithm (a stochastic simulation algorithm originally used to depict chemical reaction processes, algorithm 1) to further delineate the behavior of the meta-clones describing our system. A common workflow of a Gillespie simulation trial starts with a single progenitor (e.g., HSC/MPP 1). For each step, the time interval until the next reaction (proliferation, differentiation, or apoptosis) is extracted from an exponential distribution (where the parameter λ is a state-rate-dependent value); then, a reaction is picked to occur based on the currently available cell states and the previously estimated transition rate matrices. The simulation continues and updates cell states until certain stopping criteria are met (Fig. 3a).
a Examples of lineage trees resulting from the Gillespie algorithm; each simulation trial starts with one progenitor cell (not necessarily a HSC/MPP 1). Nodes represent cell ID, that is, its order of generation. The edge label ti is the order of occurred reactions. b Examples of the number of division events needed to produce the first progeny starting from one HSC/MPP 1 for meta-clones 2 and 7, respectively. c Examples of the number of division events needed to produce the first Mast cell or Early Erythroid starting from one HSC/MPP 1 for meta-clones 0 and background, respectively. The mean value is reported as well. d Bar plot shows the number of trials producing each progeny starting from HSC/MPP 1 among all meta-clones. E.g., meta-clones 0, 2, 6, and 7 are multi-potent clones whilst meta-clones 4 and 5 are more likely to be uni-potent clones. The y axis indicates number of simulated trials a certain progeny has been generated out of all simulations (1000 in our analysis).
We ran Gillespie on all meta-clones and, for each meta-clone, we inferred several properties, including: 1) the number of cell division events that occurred between the original HSC/MPP 1 (or other progenitor cell of interest) and the first cell produced for each progeny; 2) the number of trials where a certain population was produced out of all possible simulation trials (1000 in this case). Our simulations, therefore, yielded a quantitative summary of differentiation topology and terminal state realization for each meta-clone. After combining all the simulation results together, we could compare the differences in fold change among all meta-clones or with respect to the background cells (Supplementary Fig. S10a for different initial conditions and S10b for the comprehensive comparison between meta-clones). Comparing meta-clones against each other, we could identify correlations between division events of a particular lineage and the potency of a meta-clone. For example, for meta-clone 2, which produces large numbers of DCs, the Gillespie simulation estimated approximately five divisions from HSC/MPP 1 to the production of the first DC. In contrast, for meta-clone 7, which does not produce DC effectively, more than 12 divisions are predicted (Fig. 3b). For the myeloid lineage, for instance, it takes meta-clone 0 around 5 division events on average to produce the first Mast cell, and ten division events for the early erythroid respectively, which is similar with that of the background (Fig. 3c). This indicates that meta-clone 0 shows similar behavior with others in the myeloid lineage.
Next, we scored the capability of a meta-clone to produce progeny. To this aim, we needed to define when to consider that a cell type has been produced. We opted for setting the threshold of at least 40 cells to exist in the compartment at any time point. Meta-clones have distinct differentiation behaviors (Supplementary Fig. S6). Some of them are multi-potent; for instance, meta-clones 2 and 7 have 4 terminal states, while meta-clone 0 and 6 have 3. Others are either bi-potent (meta-clone 5) or uni-potent, being committed to a specific lineage such as Monocytes (meta-clone 4), see Fig. 3d. We also noticed that meta-clones with a similar fate realization, but different total regenerative outputs, can also bear differential kinetic rates (Fig. 3b, Supplementary Table S5). Generally, we saw that lineage output is highly meta-clone specific, and that the probability of producing each progeny (including both intermediate and terminal states) given a progenitor can be inferred probabilistically using Gillespie simulations. Indeed, the number of simulation trials in which a specific progeny is produced directly correlates with the variety in lineage realization (Fig. 3d).
CLADES recapitulates the cellular dynamics of murine hematopoiesis
We applied CLADES to a publicly available mouse hematopoietic dataset that was introduced by Weinreb et al.22. This dataset contains around 130,000 sequenced cells (Supplementary Table S6), allowing us to explore the gene signatures corresponding to differential outputs. Compared with the dataset described in the previous section (4 terminal fates out of 12 cell states), there are more potential terminal fates (10 out of 22 cell states) in this dataset. While the number of time points is the same (3 sequenced time points, days 2, 4 and 6, Fig. 4a), cells are followed over only 6 days, and therefore the extent of differentiation is lower than that of the cord blood dataset.
a UMAP illustration of the mouse hematopoietic dataset, which contains three time points and 22 manually defined populations (Baso - Basophil, DC - Bendritic cell, Eos - Eosinophil, Ery - Erythroid, Ly - Lymphoid, Meg - Megakaryocyte, Mono- Monocyte, Neu - Neutrophil, prog - progenitor, pDC - Plasmacytoid dendritic cells). b We defined 13 meta-clones based on the time-state-dependent number of barcoded cells. c The putative transition directions were given by the PAGA graph with expert curation, which acted as a guide for the model. d Left panel: the overview of the estimated proliferation and differentiation rates across all evaluation time points. Right panel: the weighted rates among meta-clones resemble the patterns of the whole dataset. e (i) Lineage realization for each meta-clone given a prog_2, (ii) lineage realization for each meta-clone given a prog_3.
This dataset has 5859 unique clones in total, with some of them being multi-potent whilst others being uni-potent. Of note, only 1989 clones (also with limited number of cells) appeared in the terminal states. This result indicates the low capture rate of barcodes and justifies the necessity to merge individual clones into the meta-clones. We then followed the same preprocessing pipeline as the cord blood dataset and constructed 13 meta-clones using again the time and state-dependent number of cells in each clone as features (Fig. 4b for combined UMAP, Supplementary Fig. S11 for used features and S12 for UMAP of each meta-clone). Distinct behaviors can be seen in different meta-clones (Supplementary Table S8). The number of multi-potent meta-clones is higher than the CB dataset, with several giving rise to seven or more terminal states (meta-clones 1, 2, 3, 4). Of interest two meta-clones, 5 and 8 produce very few cells from any terminal state with the progeny of early time-points retained within the most primitive progenitor space (Supplementary Fig. S12). As this dataset does not contain flow cytometry-based counting, we used an estimated fold expansion of hematopoietic stem and progenitor cells in culture as an alternative to the total cell counts and scaling factors (Methods). We applied both constant and dynamic modes of CLADES using the PAGA graph with expert curation as the guided transition directions (Fig. 4c). Interestingly, we found that both modes performed similarly, suggesting that, in this dataset, the kinetics of the in vitro system do not change much during the time span of the experiment (Supplementary Table S7).
The distribution of estimated proliferation as well as differentiation rates provided by CLADES overall falls within a reasonable range (Fig. 4d, left panel), with the weighted average of meta-clones across all evaluated time points resembling the rates of the whole system with a Pearson correlation score of 0.819 (Fig. 4d, right panel). Using our method, the likelihood of each meta-clone to produce differentiated progeny can be calculated, thereby simulating which progenitor would be potentially responsible for the lineage output per meta-clone. For example, if this simulation is begun from prog_2, the expected potency can be seen for multiple meta-clones, including meta-clones 1, 2, 3 and 4 (Fig. 4e (i)). Of note, this is not the case for meta-clones 0 and 7, yet if these simulations are begun from prog_3, the expected lineage realization is seen (Fig. 4e (ii)), indicating that prog_3 is the main contributor to the lineage output of these clones. This further highlights the utility of the model to resolve the contribution of each progenitor to the global differentiated output.
Clonal kinetics and characteristics can be inferred from early coordinated gene signatures
We showed earlier that the difference in outputs in terms of time-scales and fate realization of early progenitors unveils differences in transition rates. We thus sought to connect such differences to the possible heterogeneity in the transcriptomic signatures, arguing that the latter may coordinate the transition rates towards each lineage and terminal fates.
Initial comparisons of differentially expressed genes (DEGs) within the earliest progenitor population highlighted the recapitulated lineage priming reported by Weinreb et al.22. For instance, meta-clone 0 expresses genes associated with monocytes (Rbms1 and Sirpa) whereas meta-clone 4 expresses Podxl, Pbx1 and Igals9, which are associated with the megakaryocytic lineage (Supplementary Fig. S14).
The 13 meta-clones identified in this dataset have distinct behaviors (Fig. 5a, Supplementary Fig. S14 and S15). Even meta-clones with similar differentiation outputs, for example meta-clones 0 and 1, which both give rise to late progenitors in the monocyte and neutrophil lineages, differ in the final cell output of the terminal states at day 6 (predominance of monocyte progenitors in meta-clone 0 and of neutrophil progenitors in meta-clone 1; Fig. 5b). To further investigate the gene expression programs at play within the progenitor populations of these meta-clones, we analyzed the molecular signatures in the prog_1 population from the mouse hematopoietic dataset, as prog_1 is the most primitive progenitor population represented in this dataset (Fig. 4a, c). We calculated DEGs between meta-clone 1 and 0 within this population (Fig. 5c). We found meta-clone-specific genes that are characteristic of specific hematopoietic lineages and suggest lineage priming even in the most primitive progenitors (Fig. 5d). For example, up-regulation of genes important in neutrophils (Cd48, Chek1, Cited2, Pcna and Thy1) can be seen in meta-clone 1 whilst genes specifically related to monocytes can be seen in meta-clone 0 (Ccl24, Ccr2 and Zfp36) (Fig.5e). Whilst it was previously demonstrated that the potential potency of these meta-clones is generated from different progenitor populations, the subsequent progenitor population (prog_2) exhibits further lineage specific gene signatures (Fig. 5f–h), including lineage-specific transcription factors such as Gfi1 and Cebpe for neutrophilic lineages (meta-clone 1) and cell surface markers associated with monocytes (Cd33 and Cd52, meta-clone 0).
a Stacked distribution of cell counts in percentage view where the x axis shows the proportion of cell counts for each population, the y axis shows the meta-clones, and the number at the side of each bar is the total counts for each meta-clone. b Distribution of late progenitors at day 6 for meta-clones 0 and 1. c Left: only cells from the earliest progenitor populations were used for DEGs analysis, which is prog_1, as shown with brown color in the UMAP landscape for all cells. Right, examples of meta-clone-specific UMAPs. d Volcano plot of DEGs between prog_1 cells within meta-clone 1 and meta-clone 0. Identified genes associated with specific terminal states (Chek1, Cited2, Pcna), Thy1, Cd48). e Examples of gene expression violin plots for prog_1 cells for Chek1, Pcna, F2r and Nfix, illustrating early lineage expression in the most primitive progenitor population. f Left: only cells from the progenitor 2 populations were used for DEGs analysis, as shown with pink color in the UMAP landscape for all cells. Right, examples of meta-clone-specific UMAPs. g Volcano plot of DEGs between prog_2 cells within meta-clone 1 and meta-clone 0. Lineage specific gene expression can be seen, Gfi1, Alox5, Cebpe, Cd200r and S1pr4. h Examples of gene expression violin plots for prog_2 cells (Irf8, Csf1r, Gfi1 and Cebpe). i Volcano plots of DEGs between prog_1 cells within meta-clone 5 vs rest and meta-clone 8 vs rest. Upregulated genes are mostly associated with the stemness of a cell (e.g., B2m, Hmga2). d,g,i Wilcoxon rank-sum two-sided test was used to calculate the DEGs between same identity of cells from different meta-clones.
As mentioned previously, both meta-clones 5 and 8 are retained within the primitive progenitor space during the 6-day incubation period with extremely low differentiation rates to terminal states, with most of the daughter cells remaining in prog_1 and prog_2 cell state and a few in prog_4 (see Fig. 5a). To further analyze this unique behavior, we computed the DEGs between meta-clone 5 and all other meta-clones within prog_1 cell state (Fig. 5i). This showed significantly upregulated expression of genes associated with proliferation and self-renewal (Tgfbr3, Evi5, S1pr1, Selp and Yes1) whilst genes associated with specific hematopoietic lineages were downregulated (Gzmb, Elane, Mpo, Irf1). Interestingly, differential gene expression of meta-clone 8 and all other meta-clones within prog_1 population highlighted high expression of markers associated with HSCs (Fgd5) and HSPC entry into cell cycle (Ccne1), and downregulation of genes associated with lineage specification (Gfi1, Ccl3 and Cxcl10), potentially explaining the relative lack of differentiation observed in these meta-clones.
Nevertheless, we saw with our analysis that meta-clones with similar fate realizations can produce progeny in different proportions. For example, meta-clones 2 & 7 of the human cord blood data are both multi-potent, but have different offspring sizes (Supplementary Table S8). Meta-clones 0~4 of the mouse hematopoietic dataset can generate most of the terminal states, but the contribution to each population differs, as shown in Fig. 5a and Supplementary Table S8. This analysis suggests that we can further identify sub-states of progenitor cells based on the magnitude and the rates at which their progeny are produced.
Similarly, the DEGs of meta-clones 1~4 compared to meta-clone 0 (Supplementary Fig. S16 & S17) illustrate that, for multi-potent clones, the transcriptomic heterogeneity at early stages can possibly be linked to differences in fate output and population size of the offspring.
In conclusion, meta-clones analysis allows for a fine-grained definition of specific differentiation behaviors, including both lineage preference as well as size of the generated progenies. In addition, transcriptomic signatures can be assigned to these behaviors, providing a molecular basis for such behaviors.
Discussion
In this paper, we presented CLADES, a NeuralODE-based method to estimate both the proliferation and differentiation rates along the time course data. With the Gillespie algorithm, CLADES delineates detailed division topology and quantitatively summarizes the lineage output of progenitor cells. Importantly, the cellular behaviors identified by CLADES through the meta-clone approach can be associated with specific transcriptional states of the cells initiating these behaviors at the earliest time point.
Many mathematical models for hematopoiesis have assumed that adult physiological hematopoiesis is in perfectly homeostatic conditions (referred to as “steady state”), where kinetic rates are constant values. On the other hand, more recent models37,47 have begun incorporating the idea that, given that the relative abundance of the different populations, including HSCs, varies with aging, dependent on the context, there isn’t necessarily a steady state. CLADES can partially address this problem thanks to the constant vs dynamic mode option. Indeed, if the constant mode outperforms the dynamic one, we can conclude that the kinetic rates are nearly constant, which may lead to a stationary growth or a steady state, whilst if the dynamical mode performs better, then the rates are time-dependent and it is more likely that the system is not in a steady state.
Though CLADES has offered new perspectives to explore the LT-scSeq data, there are still a few challenges and limitations that exist. Firstly, CLADES requires a multi-shot lineage tracing design, limiting its use to experimental designs with at least 2, ideally 3, sequencing time points.
Secondly, CLADES is formulated based on the static barcoding with viral integration, and for now, it cannot be applied to analyze data from cumulative barcoding techniques (like CRISPR-Cas9 DNA editing24,48, which allows multi-time barcoding, and hence provides a fine-grained structure of sub-clones) or in vivo experiments40. However, for retrospective barcoding (often with endogenous genetic variants, for instance, MAESTER49), we expect that CLADES can be applied with minimal extension.
Thirdly, although CLADES aims to quantitatively summarize the differentiation kinetics and lineage output of each clone as the primary goal, while examining the regulation and determination from the cellular transcriptome in a separate step, the current model only uses cells with barcodes and, therefore, lacks analysis of cells without a barcode. Future work includes combining both barcodes and transcription data to infer the kinetic rates and lineage output, and mapping cells without barcoding information to meta-clones, since limited barcoding efficiency can affect the performance of the algorithm and DEG analysis as well.
Finally, although we focused on the hematopoietic system in both humans and mice to evaluate our model, CLADES is, in principle, broadly applicable to analyze other developmental systems. Moreover, it may be further applied to study cancer progression, where different clones may have distinct phenotypic properties, e.g., cancer plasticity50.
Methods
Ethical Statement
Human cord blood biological samples were sourced ethically, and the research was conducted in accordance with the terms of the informed consents under an institutional review board/research ethics committee-approved protocol as specified below. One umbilical cord blood (CB) sample from one male newborn (as assigned as birth) was obtained with informed consent from a healthy donor by the Cambridge Blood and Stem Cell Biobank (CBSB) in accordance with regulated procedures approved by the relevant Research and Ethics Committees (18/EE/0199 and 24/EE/0116 Research Studies). No participant compensation was provided.
Parameter definitions and data structures
Denote the number of time points available as T, the number of meta-clones as C and the number of populations as P. We define the following terms for each individual meta-clone c,
-
\({x}_{t,p}\in {{\mathbb{R}}}^{T * P}\): original real number of cells in the dish for population p and time t;
-
\({y}_{t,p}\in {{\mathbb{R}}}^{T * P}\): number of cells sequenced from the dish for population p and time t;
-
\({K}_{1}\in {{\mathbb{R}}}^{P * P}\): transition matrix of cell differentiation rates between populations. Its values are constrained to be non-negative and strictly upper triangular (to avoid the reversed differentiation process);
-
\({K}_{2}\in {{\mathbb{R}}}^{1 * P}\): rates of the overall effects of proliferation and apoptosis processes combined within a population, diagonal of the topology graph L. These rates can be negative or positive, depending on whether apoptosis exceeds proliferation or not;
-
L ∈ {0, 1}P*P: topology graph of cell states, binary version of K1 + K2, derived from PAGA with expert curation; after creating the graph edges, we inspected the biological plausibility of each edge and removed transitions that can be confidently ruled out based on previous knowledge;
-
Papop ∈ {0, 1}1*P: vector of fully differentiated populations (terminal fates) with limited proliferation ability;
-
Pprol ∈ {0, 1}1*P: vector of progenitor populations (e.g., HSCs) with strong proliferation ability;
-
\({\mu }_{t}\in {{\mathbb{R}}}^{+}\): scaling factor between xt,p and yt,p for time t;
-
tcut: stabilization term used in the modified Gillespie algorithm, a trade-off between simulation accuracy and time complexity, default is 1e−4;
-
li: penalty terms used to regularize parameters K1,2, where \(i\in {\mathbb{Z}}\);
-
λ ∈ (0, 1]: adjustable parameter controlling the magnitude of each penalty term in the loss function.
Annotation of the human cord blood dataset
To annotate the cord blood dataset, we first transferred the labels from the fetal liver atlas published by Popescu et al.51 by means of the Seurat label transfer algorithm (functions FindTransferAnchors and TransferData). We then clustered our landscape by means of the Leiden algorithm in the scanpy package and assigned to each cluster the cell type of the most commonly transferred label. Finally, for the two clusters labeled as HSC/MPP from the label transfer, we manually inspected the expression of genes known to be highly expressed in the most immature HSC/MPPs and also compared genes differentially expressed between these 2 clusters. From this inspection, the cluster with the most immature features was labeled HSC/MPP1, while the one with differentiative features was labeled HSC/MPP2.
NeuralODE-based architecture
Given a population balance model, the per capita growth/transition rates (Eq. (2)) can be treated as either time-invariant (constant value) or time-variant (they assume a different value at each time point). For a time-invariant scenario, the K1,2 themselves are the trainable parameters, whilst for the time-variant scenario, the ODE block is built upon a 2-layer multi-layer perceptron (MLP, the number of hidden dimensions is dependent on the number of populations, default is 32) with xt,p as input and K1,2 as output. Softplus activation function was used since it has a unique gradient, which is theoretically better than other non-smooth non-linear activation functions such as ReLU and LeakyReLU, given the inner characteristics of NeuralODE41. K1 is further masked by the topology graph L to confine the empirically infeasible direction of transitions (e.g., backward transitions or transitions from Late Erythroids to Monocytes). Squaring ensures the inferred rates to be non-negative in K1 and the overall transition matrix π(t) can be inferred when combining the estimated rates; the diag function is used to transform a vector into a zero-like matrix where the diagonal is that vector. In summary:
where the parameters w1, w2 are the weight matrix and the weight vector, respectively. MLP with 2 dense layers and a relatively small hidden dimension was used because the model is run at the meta-clone rather than at the individual clone scale, which significantly reduces the number of rates to be inferred, and more hidden layers (or larger layer dimensions) would inevitably lead to the risk of over-fitting.
Scaling factor for experimental cell counts
As scRNA-seq is destructive by nature, cells sequenced at later time points cannot quantitatively reflect the accurate dynamics of cell counts during this period. Since our model is based on the real number of cells at each time point, sequenced cell counts were scaled back to total cell counts in the culture environment based on additional information (e.g., either manual counts or data from fluorescence-activated cell sorting, FACS52), before being fed to the model.
In order to calculate the scaling factor between sequenced counts and real total counts of the human cord blood data, the number of cells in the dish was measured at each sequencing time (Supplementary Table S3, Supplementary Table S5). Then the estimated total number of cells at each time point is computed in a chained cascading way:
where FAC1,2,3, y1,2,3, and c1,2,3 are the numbers of cells sorted in the dish, sequenced in the experiment, and with clonal information at different time points, respectively, and x1,2,3 is a cell count tensor with a shape of (time, meta-clone, population). Of note, for downstream analysis, we only consider rates inferred for populations that have at least 4 cells (before scaling) at the respective time point.
For the cord blood dataset, we introduced some restrictions based on biological knowledge, by forcing: 1) all Ery populations (Early Ery, Mid Ery, Late Ery) to have zero counts at day 0 and day 3, 2) all DC populations (DC and DC precursors) to have zero counts at day 0 and day 3, 3) all HSC/MPP1 populations to have zero counts at day 17.
The scaling factors for the mouse hematopoiesis data are calculated similarly, except that we used the fold change of cell counts provided by the authors, as the exact number of cells cultured is not accessible.
Initial conditions of the NeuralODE framework
Solving an ODE system is essentially an initial value problem (usually the initial value is the first available time point). Static barcoding techniques like LARRY offer a retrospective ground truth, that if a barcode is seen at a later time point, one cell must have been barcoded with it at the induction time point. The LARRY system produces a vast variability of combinations of barcodes, making the possibility of having two cells with the same barcode extremely low. Therefore, this experimental protocol of LARRY allows inferring the initial condition (unobserved data at Day 0), which can be used by CLADES.
Specifically, apart from the sequenced data, we manually added an extra time point, Day 0, to the dataset, in which the number of initially labeled cells equals the number of unique barcodes, e.g., for each meta-clone. However, we do not know which populations were initially barcoded. Assuming that the distribution of barcoded cells does not change much between Day 0 and Day 3, we took the Day 3 distribution and scaled it to the expected number of initially labeled cells. Note that, in doing so, we only relied on the clones for which we have cells at all time points. Consequentially, some meta-clones were removed from our analysis, since none of their clones satisfied our requirement.
As a model whose number of parameters is larger than the available data points tends to have a large solution space, an extra data point should make it more constrained (less flexible), and have a smaller estimation error after the optimization process.
Parameter inference
Given a LARRY-based LT-scSeq dataset with noise due to detection or loss of barcodes, we formulate the cell counts at each time point as sampled from a Poisson distribution with means given by the neural network (we found it to be more robust than the commonly used GaussianNLL loss). We minimize the negative log-likelihood loss for each meta-clone separately as follows,
Whilst the inputs and outputs of the NeuralODE algorithm are based on real cell counts xt,p to interpolate and mimic the natural development process, our reconstruction loss is based on the original sequenced cell counts yt,p to avoid making a stiff ODE that is difficult to solve and to speed up the back-propagation process53, as the number of cell counts can easily scale to millions. Besides reconstruction loss, the model also incorporates the penalty terms shown in Eq.(5), where the default values are λ0, λ5 = 1.0, λ1, λ2, λ3 = 0.5, and λ4 = 0.1,
Specifically, the rationale behind these penalties is:
-
the transition rates in topology L should not be too large to violate the biological prior knowledge. We used 6 per day here because cell cycle cannot last too quickly (6 per day means 4 h duration on average). However, this is an adjustable parameter at the user’s discretion.
-
fully differentiated populations should have limited proliferation ability (e.g., Late Erythroids).
-
theoretically, populations with 0 cell counts at a certain time point should not have the ability to either proliferate or differentiate; however, to account for the possibility of barcode dropouts or sequencing error which mistakenly assigned 0 cell count to a population, a softer penalty is applied to tackle this issue (besides the major recon loss).
-
the weighted mean of the estimated rates for each meta-clone should mimic the dynamics of background cells. Here, “background” stands for all available cells, independent of the barcode presence/quality.
-
we hypothesize that the apoptosis process should not be too quick in a homeostatic environment.
-
early progenitor populations should have an overall positive net growth rate (e.g., HSCs).
We used L1 norm for most penalties due to its ability of selecting non-zero parameters, except for l2 and l4, where L2 norm was applied to make the penalty less stringent. For l2, considering an ideal scenario where cell counts sequenced at each population and time point are 100% accurate, then the population with 0 counts should have neither proliferation nor differentiation ability. However, in reality, loss of barcodes or sequencing error at each time point could introduce extra dropouts to cell counts, especially for small-sized clones, making the data harder to analyze (e.g., in the cord blood dataset, meta-clone 1 does not have any HSC/MPP 1 or HSC/MPP 2, whilst most of the progenies exist). Using the aforementioned technique, CLADES has the ability to counteract this negative effect and automatically interpolate cell counts to recover a smoothed trajectory. For l4, our intention was to introduce moderate constraints to the model in terms of suppressing the negative transition rates, which would increase the model’s flexibility when facing complicated systems.
The overall cost is the sum of both penalty terms and reconstruction loss at each time point t, clone c, population p, respectively
From DNA barcodes to meta-clones
According to the experimental protocol, the inheritable DNA barcodes are induced in undifferentiated populations and their immediate progenies at Day 0. Due to the typical challenges in clonal analysis, e.g., cell dropout, barcode homoplasy or loss of barcodes36, although hundreds of unique clones are captured, a large proportion only contains few cells or appears in limited time points and populations. Therefore, analyzing the behavior of individual clones is infeasible.
It is straightforward to use the pooled information, under the assumption that clones with similar kinetics should in turn produce similar cell counts at a specific time point t for a specific population p. Using cell counts at (ti, pi) as features, clones with alike characteristics were then clustered together to form meta-clones using the Leiden clustering method from the scanpy package54.
Here we give a detailed step-by-step guideline on how the meta-clones are yielded in our experiments,
-
1.
The LARRY system induced a few thousand clones (3940 for the cord blood dataset and 5859 for the mouse hematopoiesis dataset), of which clones with cell numbers less than or equal to 2 are filtered out, which leads to 606 clones and 4214 clones respectively;
-
2.
calculate the principle components of the filtered time-population-dependent barcode matrix using default values
$${\mathtt{sc.tl.pca()}}$$ -
3.
get the neighbours’ information
$${\mathtt{sc.pp.neighbors()}}$$ -
4.
get the reduced dimension visualization of the matrix, as an indication of the relationships of each clone
$${\mathtt{sc.tl.umap()}}$$ -
5.
meta-clones are based on the aggregation of each individual clone; here we used Leiden clustering as the example
$${\mathtt{sc.tl.leiden()}}$$
Whilst the resolution of Leiden clustering is arbitrary, we generally follow the rules that having at least 15 clones per meta-clone (after QC). Given the sparsity nature of the barcoding data, fewer clones in a meta-clone may translate into noisier estimations. The number of meta-clones is a hyper-parameter that can be adjusted to explore the data at different levels of resolutions; in our analysis, the default way is to set the parameter resolution in Leiden clustering equal to 1 (Supplementary Note 5).
Of note, an extra meta-clone named as background was included (BG, including both the barcoded and non-barcoded cells). Meta-clone BG represents the average behavior of the entire system, and the weighted average of the kinetics of all the meta-clones should resemble the dynamic profiles of the meta-clone BG.
Model initialization and training strategies
For the constant mode of CLADES, the Kaiming Uniform55 was used as the default to initialize the rates (K1,2) of the (meta-) clone-specific ODE function,
where fin is the dimension of the input layer (number of defined cell populations). For the dynamic mode, since the estimated rates for each meta-clone are given by linear layers with a small input, the weights would be too large if the parameters were initialized by the aforementioned approach because of the inversely proportional property. Therefore, to avoid problems like gradient exploding or numerical overflow which might make the adaptive solver unable to solve, we used standard normal distribution N ~ (0, 0.01) to initialize the weights in MLP as an alternative.
The training strategy also varies for both synthetic and real-world datasets. For the synthetic dataset, the number of training time points used varies for different simulation trials, whilst they all use the same testing set for evaluation, which allows a direct comparison between each trial (Supplementary Table S1 & S2). However, for real-world datasets such as the cord blood or mouse hematopoietic data, the available time points are limited. Therefore, splitting them into training and test sets is not feasible, because that would undermine the model’s ability to learn sufficient patterns, and we used all 3 available time points as the training set.
Nevertheless, the performance of CLADES can still be guaranteed, due to 1) the analysis of synthetic data proved that 3~4 input time points are sufficient to get a fairly accurate predictions; 2) and we have multiple biologically informed penalties to prevent the possibility of over-fitting.
As for the configurations used to train the model: the default epochs are 1500 and for the ODE block of the dynamic mode, softplus activation function and a hidden dimension size of 32 were used. AdamW optimizer with default settings was adopted and the learning rate can be adjusted using either multi-step learning rate decay every 200 epochs with decay rate γ = 0.5, or an automatic way based on the general losses. The initial learning rate for constant mode and dynamic mode are 5e−2 and 1e−3, respectively.
Bootstrapping for model confidence intervals
As described in the parameter inference section, PoissonNLL was used instead of GaussianNLL because it is more robust to sparse data with lineage relationships. However, we could not directly compute the estimation error or 95% CI based on this loss function.
To generate such data and get a comprehensive analysis of the rates, we adopted the bootstrapping strategy, which randomly samples the observation data with replacement M times (the initial time point t0 is not sampled). Based on the central limit theorem, we get the percentile CI for each transition rate and the quantile estimation error for the total cell counts after ranking the fitted parameters \({\hat{\theta }}_{0},{\hat{\theta }}_{1},...,{\hat{\theta }}_{M-1},{\hat{\theta }}_{M}\) of all bootstrapping trials.
For statistical analysis and the comparison of rates within different meta-clones, we mainly used the student t test and Mann Whitney U rank test based on whether the distribution of bootstrapped values follows a standard normal distribution or not. The p-values of multiple tests were corrected using the Benjamini-Hochberg procedure (false discovery rate, FDR-BH) with 0.05 as the threshold.
It is worth noting that, when performing the tests, we took precautions to avoid abnormal significance in the bootstrapping results, especially for populations with zero cell counts. Therefore, we introduced the mean absolute change of parameters as an additional metric to assess the significance between a pair of meta-clones (see Model evaluation metrics section). Furthermore, we decided to deem as non-significant also the rates estimated for cell-populations that have less than 4 cells at the corresponding time point.
The bootstrapping approach might over-estimate the CI given limited available experimental time points; alternative approaches include increasing sequencing times or profiling the likelihoods56,57 after fitting the model, at the cost of a complicated experimental design or a linearly increased computational complexity.
Model evaluation metrics
The following evaluation metrics were used to help assess the performance and usage of CLADES, for both synthetic and real-world datasets.
Average recovery rate
We used this to demonstrate the model’s ability to interpolate cell counts on unknown time points (for synthetic datasets) and to assess the model’s performance in reconstructing the original cell counts (for real-world datasets) with the following formula,
where xt,c,p and \({\hat{x}}_{t,c,p}\) are the observed and predicted cell counts at any given time points, meta-clones, and populations. Since cell counts from different time points usually have distinctive magnitudes (shown in the scaling factors), this metric is often evaluated on a per-time-point basis to ensure the comparability between different trials.
Correlation of cell counts
Whilst the average recovery rate focused on the error of each individual data point, this metric is more coarse-grained, which models the general trends of cell counts given a time stamp and a meta-clone
as shown in the equation, the correlation is based on the cell counts of available populations. Thus, it is reasonable if the correlation is extremely high, as the number of populations is usually not a large number.
Absolute error of rates
Both evaluation metrics above are focused on the accuracy of predicted cell counts, for each (meta-) clone, the number of descendant cell counts is the product of the orchestrated transition rates between upstream and downstream cell states.
Therefore, it is also useful to investigate the model’s performance in recovering interim parameters.
where K is the transition rate matrix, including both proliferation and differentiation rates.
The absolute error of rates is a more direct indicator of model’s performance, as sometimes the error accumulated in transition rates could not be reflected by the predicted cell counts.
Mean absolute difference of parameters
This is mainly used in the post-processing part of the bootstrapping results, together with statistical tests. As in some cases, the variance of distribution of rates given by bootstrapping is small, and that leads to a significant statistical difference between meta-clones, which might lead to wrong interpretation when the total cell counts are 0.
Suppose we need to compare two rates and from M bootstrapping trials, we can get \({K}_{{\theta }_{i},{c}_{1},{p}_{1},{p}_{2}}\) and \({K}_{{\theta }_{i},{c}_{2},{p}_{1},{p}_{2}}\). Then the mean absolute difference of rates is defined as,
and the default value for this metric is 0.1, so that any difference lower than this threshold would be considered as not significant.
Using Gillespie to simulate differentiation landscapes
We adopted and modified the original Gillespie algorithm to achieve a balance between accuracy and time complexity (Supplementary Note 3, algorithm 1). As tcut was introduced in the algorithm design to guarantee convergence (e.g., the time increment between two simulation steps Δt = max(Δt, tcut)), the cell counts generated by the stochastic simulation algorithm do not perfectly resemble the observed counts, whilst this does not affect the division statistics of a progenitor cell to produce a certain progeny and the likelihood of producing a lineage.
The division summary is delineated in the following way: starting from an early progenitor cell at the initial time point (e.g., HSC/MPP 1 or prog_1); the number of proliferation events was counted until the first progeny was produced. This progeny could either be a later progenitor or a specific cell fate (e.g., prog_2 or Erythroids). In some realizations, there were only differentiation events; thus, the number of divisions was 0.
In order to explore the variety of fates produced by different progenitors, we run the simulation 1000 times, starting each time with just a single progenitor cell. The starting progenitor can be any, but we focused on the cell types that were initially barcoded to recapitulate the observation and get more insight about which cell type is producing the different outputs during the time span of the experiment. Note that we consider as “produced” a lineage if there are at least 40 cells simultaneously in that lineage at any point of the simulation.
The original Gillespie algorithm sometimes falls into an infinite loop of choosing the same reaction, due to the extremely small time increments. Though the introduction of tcut does not affect the analysis mentioned above, we lose other information that the Gillespie algorithm could provide, e.g., analysis of cell count dynamics and statistics of reactions.
Experimental procedures for the LARRY human cord blood dataset
1) LARRY barcoding plasmid propagation and analysis of barcode library diversity using 10X. The LARRY Barcode Version 1 library was a gift from Fernando Camargo (Addgene, #140024). The library was amplified and the lentiviral vector produced according to the associated published protocol22 with some minor adaptations. Briefly, plasmids were introduced into ElectroMAX Stbl4 competent cells (Life Technologies) using a MicroPulser Electroporator (Biorad) and incubated for 1 h at 37 °C before spreading over 24 large Agar + Ampicillin plates. After 24 h at 32 °C, colonies were harvested through scraping using pre-warmed LB medium containing Ampicillin. The resulting culture (approx. 1.5L) was incubated at 37 °C for 2 h before isolating plasmid DNA using Megaprep kits (Machery-Nagel).
A reference library was made through sequencing of PCR-amplified barcodes from the LARRY plasmid library. 10ng of LARRY plasmids was taken as input for a two-step PCR; the first step adds Illumina Read1 and Read2 sequences (5’ACACTCTTTCCCTACACG ACGCTCTTCCGATCTTGTGACGTCACAGGTCGACACCAGTCTCATT3’ and 5’GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCGAGTAACCGTTGCTAGGAGAGACCATA3’). The second step adds the P5 and P7 flow cell attachment sequences and 7bp sample indices (P5 5’AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT3’ and P7 5’CAAGCAGAAGACGGCATACGAGANNNNNNNGTGACTGGAGTTCAGACGTGCTCTTCCGATC3’). 8ng of PCR1 product was taken as input for PCR2. The PCR program used was as follows: 98 °C 2 min, eight cycles of 98 °C 10 s, 58 °C (PCR1), 62.5 °C (PCR2) 20 s, 72 °C 30 s, followed by final elongation 72 °C 5 min and 4 °C indefinitely. In between PCR1 and PCR2, PCR purification was performed using the QIAquick PCR purification kit (Qiagen). Purification of PCR2 product was carried out using Ampure XP beads (Beckman Colter) before sequencing on a Novaseq instrument at the Cancer Research UK Cambridge Institute Genomics facility. The resulting list of LARRY barcodes was used as a reference list in downstream analysis.
2) LARRY library lentivirus preparation. Lentiviral vector was produced by transforming the amplified LARRY and packaging plasmids (psPAX2 and pMD2.G) into HEK293T cells using the TransIT-LT1 transfection reagent (Mirus) and incubated at 37 °C. Twenty-four hours after transfection, 500μl of 0.5 mM Sodium Butyrate (Sigma-Aldrich) was added to the cells and the culture continued at 37 °C incubation. LARRY lentivirus was harvested 48 h after transfection, filtered through a 0.45 μm PES filter (Whatman) and concentrated 100-fold by centrifugation in a Beckman Colter Optima XPM-80 ultracentrifuge at 20,000 x g for 2 h at 4 °C. The lentivirus was aliquoted and stored at −80 °C. Titration of the lentiviral barcoding library was performed on HEK293T cells, with a read out obtained by flow cytometry 3 days after infection.
3) Cord blood samples. Umbilical CB samples were obtained with informed consent from healthy donors by Cambridge Blood and Stem Cell Biobank (CBSB) in accordance with regulated procedures approved by the relevant Research and Ethics Committees (18/EE/0199 and 24/EE/0116 Research Studies). MNCs were obtained using Pancoll density gradient centrifugation of diluted (1:1 with PBS) CB. Red cells were lysed before positive selection for CD34+ cells using the micro beads CD34+ selection kit and AutoMACS cell separation technology (Miltenyi Biotech). CB CD34+ cells were then stored at −150 °C until use in experiments.
4) FACS. To sort CD34+ cells for experiments from CB samples, the cells were thawed by dropwise addition of pre-warmed Rich Thawing Medium containing Iscove’s Modified Dulbecco’s Medium (IMDM, Life Technologies), 0.1 mg/ml DNase (Lorne Laboratories) and 50% Fetal Bovine Serum (FBS, Life Technologies) before centrifugation (500 x g 5 min) and resuspension in PBS + 3% FBS. The samples were stained with antibodies (Table 1).
Cells were incubated for 20 min at room temperature and washed with PBS + 3% FBS. DAPI (Biolegend) was added 1 in 100 (final concentration 137 μg/ml) to identify viable cells. Unstained cells and compensation beads (Invitrogen) were used for compensation and as controls to set appropriate gates. Lineage− CD11c− CD34+ cells were sorted into Eppendorf tubes using a BD Influx approved for CL2-sorting.
5) In vitro barcoding, culture and sampling for 10X. The cord blood CD34+ cells were cultured following the reference conditions58 (Tables 2 & 3), that promote the differentiation of cells towards Megakaryocyte, Erythroid and Myeloid lineages (MEM) with reduced levels of EPO cytokine to avoid biasing the culture towards the erythroid lineage.
For barcode labeling, 75,000 sorted Lineage− CD11c− CD34+ cells were seeded at 5000 cells per well in a 96-well round-bottom plate. LARRY lentivirus was added directly to the culture at an MOI of 60. After 24 h, the virus was diluted out and cells transferred to a 96-well flat-bottom plate. At 3 days post-transduction, GFP+ and GFP− cells were FACS-sorted. Two thirds of the GFP+ fraction (8856 cells) was sent for scRNA-seq analysis (10X genomics) and the remaining third (4428 cells) was re-plated in 500 μl reduced EPO MEM culture conditions in a 24-well plate. GFP− cells were also analyzed by scRNA-seq. At day 10 post-transduction cells were stained for GlyA (PE, 1:1000 dilution, BD) using the protocol above. GFP+ GlyA− cells were FACS-sorted, with 40,000 cells processed for scRNA-seq and the remainder replated, as previously. At day 17 post-transduction, 40,000 GFPhi GlyA− and 40,000 GFPmid GlyA− cells were FACS-sorted and processed for scRNA-seq.
6) Sample preparation and LARRY cDNA enrichment for scRNAseq (10X Genomics). Up to 20,000 live cells of interest were sorted into 300μl MEM media and kept on ice before centrifugation. Cells were resuspended in PBS + 0.04% BSA (miltenyi) and further processed for single cell sequencing using the Chromium Single Cell 3’ Library & Gel Bead Kit v3 (10X Genomics) following manufacturer’s protocols. LARRY molecules constitute a small fraction of the total cDNA library; thus, we adapted previously published protocols22,59 to enrich the LARRY barcodes and ensure a sufficient number of reads per barcode. Using a portion of each sample’s cDNA, LARRY sequences were PCR-amplified while simultaneously adding Illumina primers and indices necessary for sequencing and identifying the enriched samples in downstream analysis (Supplementary Table S9). Purification of the PCR product was carried out using Ampure XP beads (Beckman Colter) before sequencing alongside the full sample cDNA library on a NovaSeq 6000 at the Cancer Research UK Cambridge Institute Genomics facility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
CLADES used two public datasets to demonstrate its applicability and the downstream functions: 1) a newly generated LARRY-based human cord blood LT-scSeq data, and the raw data is deposited under GEO with accession number GSE276896. The scripts for pre-processing the data can be found here. Processed adata, and files for model inputs are stored and can be accessed via https://figshare.com/articles/dataset/CLADES/27908142 with DOI number https://doi.org/10.6084/m9.figshare.27908142; 2) the mouse hematopoietic system is a publicly available dataset22,36, and we used the same data as described in the literatures, however with a more fine-grained annotation of cell populations. Relevant data can be accessed via the same figshare repository as well. We have deposited all the synthetic datasets used in this project into https://figshare.com/articles/dataset/CLADES/27908142 as well. The scripts for generating Supplementary Figs. can be found in several different notebooks within the GitHub repository https://github.com/StatBiomed/clonaltrans, covering both the data preprocessing and downstream analysis parts.
Code availability
CLADES is a Python package and is publicly available at https://github.com/StatBiomed/clonaltrans with both constant and dynamic modes implemented. It also contains documentation, pipelines, and Jupyter Notebooks to reproduce figures and results mentioned in this paper. The specific version used for this manuscript is v1.2.0, with Zenodo DOI identifier https://doi.org/10.5281/zenodo.1581140860.
References
Kester, L. & Van Oudenaarden, A. Single-cell transcriptomics meets lineage tracing. Cell Stem Cell 23, 166–179 (2018).
Woodworth, M. B., Girskis, K. M. & Walsh, C. A. Building a lineage from single cells: genetic techniques for cell lineage tracking. Nat. Rev. Genet. 18, 230–244 (2017).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Setty, M. et al. Characterization of cell fate probabilities in single-cell data with Palantir. Nat. Biotechnol. 37, 451–460 (2019).
Qiu, X. et al. Mapping transcriptomic vector fields of single cells. Cell 185, 690–711 (2022).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
La Manno, G. et al. Rna velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Soldatov, R. A., Kharchenko, P. V. & Theis, F. J. RNA velocity-"current challenges and future perspectives. Mol. Syst. Biol. 17, e10282 (2021).
Gao, M., Qiao, C. & Huang, Y. Unitvelo: temporally unified RNA velocity reinforces single-cell trajectory inference. Nat. Commun. 13, 6586 (2022).
Erhard, F. et al. Time-resolved single-cell RNA-seq using metabolic RNA labelling. Nat. Rev. Methods Prim. 2, 77 (2022).
Qiu, Q. et al. Massively parallel and time-resolved RNA sequencing in single cells with SCNT-seq. Nat. Methods 17, 991–1001 (2020).
Maizels, R. J., Snell, D. M. & Briscoe, J. Reconstructing developmental trajectories using latent dynamical systems and time-resolved transcriptomics. Cell Syst. 15, 411–424 (2024).
Chen, C., Liao, Y. & Peng, G. Connecting past and present: single-cell lineage tracing. Protein Cell 13, 790–807 (2022).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339 (2019).
Kwok, A. W. C. et al. Mquad enables clonal substructure discovery using single cell mitochondrial variants. Nat. Commun. 13, 1205 (2022).
Huang, R. et al. Robust analysis of allele-specific copy number alterations from scRNA-seq data with XClone. bioRxiv 15, 6684 (2023).
Xue, Y., Su, Z., Lin, X., Ho, M. K. & Yu, K. H. Single-cell lineage tracing with endogenous markers. Biophys. Rev. 16, 125–139 (2024).
Chung, H. M. & Huang, Y. Interpretable variational encoding of genotypes identifies comprehensive clonality and lineages in single cells geometrically. bioRxiv 2024–07 (2024).
Guo, C. et al. Celltag indexing: genetic barcode-based sample multiplexing for single-cell genomics. Genome Biol. 20, 1–13 (2019).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
Sashittal, P., Schmidt, H., Chan, M. & Raphael, B. J. Startle: a star homoplasy approach for CRISPR-Cas9 lineage tracing. Cell Syst. 14, 1113–1121 (2023).
Spanjaard, B. et al. Simultaneous lineage tracing and cell-type identification using CRISPR–Cas9–induced genetic scars. Nat. Biotechnol. 36, 469–473 (2018).
Lu, R., Neff, N. F., Quake, S. R. & Weissman, I. L. Tracking single hematopoietic stem cells in vivo using high-throughput sequencing in conjunction with viral genetic barcoding. Nat. Biotechnol. 29, 928–933 (2011).
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013).
Bandler, R. C. et al. Single-cell delineation of lineage and genetic identity in the mouse brain. Nature 601, 404–409 (2022).
Ratz, M. et al. Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics. Nat. Neurosci. 25, 285–294 (2022).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Rodriguez-Fraticelli, A. E. et al. Single-cell lineage tracing unveils a role for tcf15 in haematopoiesis. Nature 583, 585–589 (2020).
Merino, D. et al. Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer. Nat. Commun. 10, 766 (2019).
Rodriguez-Meira, A. et al. Single-cell multi-omics identifies chronic inflammation as a driver of tp53-mutant leukemic evolution. Nat. Genet. 55, 1531–1541 (2023).
Fischer, D. S. et al. Inferring population dynamics from single-cell RNA-sequencing time series data. Nat. Biotechnol. 37, 461–468 (2019).
Forrow, A. & Schiebinger, G. Lineageot is a unified framework for lineage tracing and trajectory inference. Nat. Commun. 12, 4940 (2021).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943 (2019).
Wang, S.-W., Herriges, M. J., Hurley, K., Kotton, D. N. & Klein, A. M. Cospar identifies early cell fate biases from single-cell transcriptomic and lineage information. Nat. Biotechnol. 40, 1066–1074 (2022).
Kucinski, I. et al. A time-and single-cell-resolved model of murine bone marrow hematopoiesis. Cell Stem Cell 31, 244–259 (2024).
Tong, A. et al. Learning transcriptional and regulatory dynamics driving cancer cell plasticity using neural ODE-based optimal transport. bioRxiv 2023-03 (2023).
Wojtowicz, E. E. et al. Panhematopoietic RNA barcoding enables kinetic measurements of nucleate and anucleate lineages and the activation of myeloid clones following acute platelet depletion. Genome Biol. 24, 152 (2023).
Wang, S.-W. Methyltree: exploring epimutations for accurate and non-invasive lineage tracing. Nat. Methods 22, 463–464 (2025).
Chen, R. T., Rubanova, Y., Bettencourt, J. & Duvenaud, D. K. Neural ordinary differential equations. Adv. Neural Inf. Process. Syst. 31, 1–13 (2018).
Lu, T., Volfson, D., Tsimring, L. & Hasty, J. Cellular growth and division in the Gillespie algorithm. Syst. Biol. 1, 121–128 (2004).
Wolf, F. A. et al. Paga: graph abstraction reconciles clustering with trajectory inference through a topology-preserving map of single cells. Genome Biol. 20, 1–9 (2019).
Waldherr, S. Estimation methods for heterogeneous cell population models in systems biology. J. R. Soc. Interface 15, 20180530 (2018).
Charlebois, D. A. & Balázsi, G. Modeling cell population dynamics. Silico Biol. 13, 21–39 (2019).
Stumpf, P. S. et al. Stem cell differentiation as a non-Markov stochastic process. Cell Syst. 5, 268–282 (2017).
Takahashi, M. et al. Reconciling flux experiments for quantitative modeling of normal and malignant hematopoietic stem/progenitor dynamics. Stem Cell Rep. 16, 741–753 (2021).
Bowling, S. et al. An engineered CRISPR-Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422 (2020).
Miller, T. E. et al. Mitochondrial variant enrichment from high-throughput single-cell RNA sequencing resolves clonal populations. Nat. Biotechnol. 40, 1030–1034 (2022).
Marjanovic, N. D. et al. Emergence of a high-plasticity cell state during lung cancer evolution. Cancer Cell 38, 229–246 (2020).
Popescu, D.-M. et al. Decoding human fetal liver haematopoiesis. Nature 574, 365–371 (2019).
Orfao, A. & Ruiz-Argüelles, A. General concepts about cell sorting techniques. Clin. Biochem. 29, 5–9 (1996).
Kim, S., Ji, W., Deng, S., Ma, Y. & Rackauckas, C. Stiff neural ordinary differential equations. Chaos Interdiscip. J. Nonlinear Sci. 31, 093122 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. Scanpy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
He, K., Zhang, X., Ren, S. & Sun, J. Delving deep into rectifiers: surpassing human-level performance on imagenet classification. In Proc. IEEE International Conference on Computer Vision, 1026–1034 (IEEE, 2015).
Meeker, W. Q. & Escobar, L. A. Teaching about approximate confidence regions based on maximum likelihood estimation. Am. Stat. 49, 48–53 (1995).
Raue, A. et al. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics 25, 1923–1929 (2009).
Belluschi, S. et al. Myelo-lymphoid lineage restriction occurs in the human haematopoietic stem cell compartment before lymphoid-primed multipotent progenitors. Nat. Commun. 9, 4100 (2018).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Gao, M. et al. A hybrid neuralode-gillespie approach for unveiling clonal cell fate and differentiation dynamics. StatBiomed clonaltrans https://doi.org/10.5281/zenodo.15811408 (2025).
Acknowledgements
We kindly thank cord blood donors for their tissue donation, Dr. Joanna Baxter and the team of the Cambridge Blood and Stem Cell Biobank for consenting and collecting cord blood samples. We also thank Reiner Schulte and Gabriela Grondys-Kotarba at the Cambridge Institute for Medical Research Flow Cytometry Core Facility for flow cytometry cell sorting, and the CRUK Cambridge Institute Genomics Center for sequencing. This project is supported by the National Natural Science Foundation of China (No. 62222217), the University of Hong Kong through a startup fund and a seed fund (Y.H.), and the InnoHK initiative of the Innovation and Technology Commission of the Hong Kong Special Administrative Region Government. This project is also funded by Wellcome (215116/Z/18/Z; E.L. and B.G.) and previous core support grants from Wellcome and Medical Research Council (MRC) to the Wellcome-MRC Cambridge Stem Cell Institute (203151/Z/16/Z). E.L. was supported by a Wellcome - Royal Society Sir Henry Dale Fellowship 107630/Z/15/Z. M.H. is supported by a Sir Henry Wellcome Postdoctoral Fellowship (224055/Z/21/Z). M.B. is funded in part also by the Wellcome Trust [226795/Z/22/Z] This research was funded in whole, or in part, by the Wellcome Trust. For the purpose of open access, the author has applied a Creative Commons Attribution (CC BY) license to any Author Accepted Manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
B.G., M.B., E.L., and Y.H. conceived the project. M.G. developed the algorithm, implemented the package CLADES, and analyzed the data with Y.H., M.B., and W.Z.’s support. S.C., M.H., and E.C. generated the cord blood dataset. M.B. processed the data with S.C.’s and Y.C.’s support. M.G., M.B., and Y.H. wrote the manuscript with inputs from all authors. N.W. and E.L. edited the manuscript and contributed to the biological interpretation of the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
: Nature Communications thanks Shou-Wen Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gao, M., Barile, M., Chabra, S. et al. CLADES: a hybrid NeuralODE-Gillespie approach for unveiling clonal cell fate and differentiation dynamics. Nat Commun 16, 8174 (2025). https://doi.org/10.1038/s41467-025-63150-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63150-6
This article is cited by
-
TemporalVAE: atlas-assisted temporal mapping of time-series single-cell transcriptomes during embryogenesis
Nature Cell Biology (2025)