Abstract
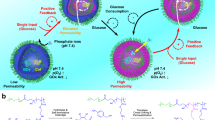
The plasma membrane (PM), a physical barrier separating cells from their environment, responds to fluctuating extracellular environment through receptor-mediated signaling. While these pathways have been extensively studied, the role of PM lipids remains poorly understood. Here, we show that phosphatidylinositol 4,5-bisphosphate (PIP2), a multifunctional phospholipid, translocates from the inner to the outer leaflet of the PM in response to extracellular acidification. A genome-wide screening identifies Transmembrane 9 superfamily 3 (TM9SF3) as a critical regulator for PIP2 translocation. During zebrafish gastrulation, when intracellular pH increases and extracellular interstitial fluid pH decreases, mutant anterior axial mesoderm lacking Tm9sf3 exhibits disorganized collective cell migration due to impaired PIP2-dependent cytoskeletal organization. Our results demonstrate that TM9SF3 mediates the PIP2 translocation when cells encounter a low pH for adapting the cells to their environment. Given that “pH-dependent PIP2 translocation” is evolutionarily conserved, cells may broadly employ lipid topology as a strategy to respond to extracellular stimuli.
Similar content being viewed by others
Introduction
The plasma membrane (PM) serves as a crucial interface between cells and their environments. Phospholipids, including glycerophospholipids and sphingolipids, and cholesterol constitute the lipid bilayer of the PM. The distribution of phospholipids between the inner and the outer leaflets of the PM is asymmetric. Phosphatidylcholine (PC) and sphingomyelin (SM) are predominantly localized to the outer leaflet, whereas phosphatidylserine (PS), phosphatidylethanolamine (PE), and phosphoinositides (PI) are enriched in the inner leaflet1,2,3. Cells can transiently scramble PM lipids to facilitate specific cellular functions. For example, PS and PE are actively translocated from the inner to the outer leaflet, a process essential for apoptotic cell clearance by macrophages4,5 and cytokinesis6,7, respectively. However, whether external stimuli can induce phospholipids translocation to trigger intracellular signaling, regulate cell migration, or modulate cellular morphology remains poorly understood.
Cells must continuously adapt to changing environments to maintain their integrity and function within the organism. While protein-based signal transduction cascades in the PM have been well characterized in response to extracellular biophysical stimuli such as mechanical force, temperature, and light8,9,10, the role of phospholipids as direct mediators of environmental adaptation is less explored. Developmental processes are regulated not only by genetic factors but also by environmental cues such as oxygen levels, temperature, and mechanical forces, which impact cell migration, proliferation, fate specification, and embryonic survival. pH is a fundamental physicochemical parameter affecting organisms across all domains of life11. Dysregulated pH homeostasis is associated with various diseases, including ischemia, cancer, and inflammation12. However, how cells sense and respond to changes in extracellular interstitial fluid pH (pHext) is not fully understood yet, particularly in the context of early embryonic development.
In this study, we demonstrate that embryonic cells respond to lower pHext (≤7.3) by altering the localization of PI between the inner and the outer leaflet of the PM during early development to regulate morphogenesis, even before the establishment of gas exchange and the urinary system, which are responsible for systemic pH regulation. Our findings highlight a previously unrecognized role of phospholipid dynamics in cellular responses to environmental pH during early development.
Results
Translocation of PIP2 to the outer leaflet of the PM in response to a lower pH
To investigate whether zebrafish cells change the asymmetric lipid distribution in the PM leaflets in response to varying pH, we examined the phospholipids located in the outer leaflet using membrane-impermeable probes. pH-dependent change in the localization of PS and PE was not observed, while PS was detected in the outer leaflet of the apoptotic cells4,13,14 (Fig. 1a–c). PIs are regulated by specific kinases and phosphatases and switch among distinct PI species15,16. Among them, PI(4)P and PI(4,5)P2 are the most abundant PIs in the PM. PI(3,4,5)P3 is produced from PI(4,5)P2 in response to extracellular stimulation. Hereafter, PI(4)P, PI(4,5)P2, and PI(3,4,5)P3 are referred to as PIP, PIP2, and PIP3. We utilized fluorescent protein probes: TagRFP-Phospholipase C (PLC) δ1-Pleckstrin homology (PH) (hereafter referred to as the PIP2 probe) for PIP2, TagRFP-P4M-SidM for PIP, and TagRFP-Akt-PH for PIP3, capable of detecting PIs located in the inner leaflet (Fig. 1d). Notably, we found that PIP2 was present at the outer leaflet when the cells were cultured at pH 7.2 but absent at pH 8.1 (Fig. 1a, c). Importantly, a mutant PIP2 probe deficient in PIP2 binding showed no fluorescence delineating the cell boundary under the same pH condition (Supplementary Fig. 1a). Additionally, the fluorescence intensity of the PIP2 probe under different pH conditions reflected the quantity of PIP2, rather than changes in the fluorescence of the PIP2 probe or the binding affinity between PIP2 and the probe (Supplementary Fig. 1b, c). These findings indicate that PIP2 can become enriched in the outer leaflet under a lower pHext environment.
a Confocal single-scanned cell images of isolated blastoderm cells at 4.5 hours post-fertilization (hpf), showing plasma membrane (PM) localization of various lipid-binding probes: TagRFP-PLCδ1-PH (PIP2; hereafter referred to as PIP2 probe), TagRFP-P4M-SidM (PIP), TagRFP-Akt-PH (PIP3), Annexin-V (PS), and FITC-SA-Ro (PE). Cells were incubated in Danieau’s buffer adjusted to pH 8.1 or pH 7.2, except for Annexin-V, which was used in a binding medium adjusted to pH 8.2 or pH 7.0. b Confocal single-scanned cell images of isolated blastoderm cells from Flag-Bax/H2A-BFP encoding mRNA-injected embryos at 4.5 hpf, incubated with Annexin-V in the binding medium adjusted to pH 8.2 or pH 7.0. c Quantification of fluorescence intensity at PM for each probe shown in (a). Individual cell values were normalized to the average intensity at pH 7.2 (set as 1). An identical methodology was employed for the quantitative analysis of lipid probes in this study. (PIP2, n = 288 (pH 8.1), n = 289 (pH 7.2); PE, n = 209 (pH 8.1), n = 168 (pH 7.2)). PM localization was not detected for PIP, PIP3, and PS. P-values were calculated using a two-tailed unpaired t-test for each lipid probe. The P values for PIP2 is <0.0001, and for PE is 0.0811. d Confocal single-scanned cell images of isolated blastoderm cells at 4.5 hpf embryos having TagRFP-P4M-SidM probe, PIP2 probe, and TagRFP-Akt-PH probe, injected at one-cell stage and imaged in Danieau’s buffer adjusted to pH 7.2. e Confocal single-scanned cell images of PIP2 probe on the PM of isolated blastoderm cells at 4.5 hpf, treated with DMSO or Ionomycin in L-15 medium. f Quantitative analysis of e. PM fluorescence intensity of TagRFP was measured in DMSO-treated (n = 121) and Ionomycin-treated (n = 160) cells. Individual cell values were normalized to the DMSO control (set as 1). P-values were calculated using a two-tailed unpaired t-test; **** indicates P < 0.0001. g Confocal single-scanned cell images of PIP2 probe of isolated blastoderm cells treated with DMSO or Nigericin/Valinomycin in Danieau’s buffer adjusted to pH 7.2. h Quantitative analysis of g. PM fluorescence intensity of TagRFP was measured for Nigericin/Valinomycin-treated (n = 141), FCCP-treated (n = 197), and control cells treated with DMSO (n = 140) or EtOH (n = 263). Values were normalized to their respective controls (set as 1). P-values were calculated using a two-tailed unpaired t-test; **** indicates P < 0.0001. All source data supporting the findings of this study are provided as a Source Data file. BF bright field, PS phosphatidylserine, PE phosphatidylethanolamine; PI, phosphatidylinositol, ND not detected, ns not significant, hpf hours post-fertilization. Scale bars, 20 µm.
PIP2 is a multifunctional lipid that regulates many cellular functions at the PM17. It acts as a substrate for second messenger involving PLC and PI 3-kinase (PI3K). PIP2 also activates actin-binding proteins that positively modulate the actin cytoskeleton, supporting processes from endocytosis to cell motility. Additionally, it regulates the activity of various ion channels and transporters. Given that PIP2 is typically present in the inner leaflet of the PM to perform these functions, we assumed that PIP2 detected in the outer leaflet had been translocated from the inner leaflet upon pH changes. Depletion of PIP2 at the inner leaflet by expressing the PIP2 phosphatase INPP5E18,19 (Supplementary Fig. 1d, e) or activating PLC activity with ionomycin20 (Fig. 1e, f) reduced PIP2 in the outer leaflet, indicating that PIP2 in the outer leaflet originates from the inner leaflet.
Next, we investigated whether mammalian cells also possess the pH-dependent PIP2 translocation machinery. In mammalian cells, PIP2, but not PIP and PIP3, was present in the outer leaflet of the PM when cultured in a low pH medium, as was found in the zebrafish cells (Supplementary Fig. 1f, g). Simultaneous visualization of PIP2 in both the inner and outer leaflets exhibited clear colocalization (Supplementary Fig. 1h), supporting the notion that PIP2 translocates from the inner to the outer leaflet. These results suggest that PIP2 translocation in response to a low pH environment is a conserved mechanism across vertebrates.
To investigate the correlation of PIP2 translocation with pHext and intracellular pH (pHint), we employed pH sensors for monitoring the pH dynamics inside and outside of the PM during PIP2 translocation. pH-sensitive SuperEcliptic (SE) pHluorin and the pH-insensitive mCherry were used to precisely monitor pH dynamics by ratiometry21. The probes for specifically monitoring pHext and pHint were tagged with GPI- (mCherry-SEpHluorin-GPI) or membrane- (Lyn-mCherry-SEpHluorin) anchoring sequences, respectively (Supplementary Fig. 2a). We found that pHint was unchanged when the cells were exposed to a low pH condition (Supplementary Fig. 2b, c), indicating that alterations in pHext do not affect pHint on the short time scale required for PIP2 translocation. These results suggest that cells sense pHext for PIP2 translocation. This finding suggests the presence of a membrane protein(s) capable of sensing pH fluctuation in the extracellular membrane surface, thereby promoting PIP2 translocation.
Subsequently, we investigated the significance of the pH gradient across the PM in the translocation of PIP2 to the outer leaflet. We eliminated the pH gradient across the PM by equilibrating pHext and pHint using ionophores or carbonyl cyanide 4-(trifluoromethoxy) phenylhydrazone (FCCP)21,22,23. These drug-treated cells showed a reduced amount of PIP2 in the outer leaflet in a medium adjusted to pH 7.2 compared to control cells (Fig. 1g, h). These results suggest that cells may perceive the pH gradient across the PM, rather than the absolute value of pHext, to facilitate the translocation of PIP2.
Identification of TM9SF3 as a guardian for asymmetric PIP2 distribution at the PM
We investigated the molecular mechanism of pH-dependent PIP2 translocation at the PM. The transverse movement (flop or flip) of PS is regulated by membrane proteins in an ATP- or Ca2+-dependent manner14. However, a decrease of neither ATP nor Ca2+ affected the translocation of PIP2 (Supplementary Fig. 2d, e), suggesting the presence of ATP- or Ca2+-independent PIP2 translocator in the cells.
To identify molecules involved in PIP2 translocation at the PM, we performed a comprehensive genome-wide transposon screening using a PiggyBac transposon-based system for floating 293F cells24,25. PIP2 was present in the outer leaflet of parental 293F cells under low pH (<pH 7.3) (Fig. 2a, b). A small subset of mutated cells with reduced affinity for the PIP2 probe was selectively sorted and expanded, and subsequently underwent an additional sorting step to acquire a cell population characterized by “low PIP2 exposure” (Fig. 2c, d). After the 10th round, additional sorting did not further increase the proportion of low PIP2 probe-binding cells, nor did it decrease the probe intensity per cell. The established mutant cells displayed reduced PIP2 translocation but normal PIP2 content within the cells (Supplementary Fig. 3a–c). Vector insertion sites were recovered using a splinkerette-PCR method and identified through conventional sequencing techniques26. Among the candidate molecules, we focused on membrane-associated molecules that might be potentially involved in the flopping of PIP2 and found that the transposon cassette was inserted in the Transmembrane 9 superfamily 3 gene (TM9SF3).
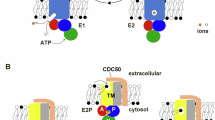
a Confocal single-scanned cell images of 293F cells incubated with PIP2 probe in Ringer’s solution adjusted to indicated pH. b Quantification of TagRFP signal at the PM in (a). Values were normalized to the average intensity at pH 6.0 (set as 1). (n = 104 (pH 8.3); 113 (pH 7.8); 104 (pH 7.3); 101 (pH 6.8); 105 (pH 6.3); 105 (pH 6.0)). P-values were calculated using one-way ANOVA followed by Tukey’s multiple comparisons test. P = 0.4783 and P < 0.0001 for comparisons between pH 8.3 vs pH 7.8, and pH 7.8 vs pH 7.3, respectively. c Schematic diagram of the PiggyBac-based screening method. d Generation of a “low PIP2 exposure” 293F cell population through FACS sorting. e Schematic representation of the TM9SF3 protein structure. f Western blot analysis confirming the knockout of TM9SF3 in 293F cells. g Confocal single-scanned cell images of TM9SF3-/− 293F cells incubated with PIP2 probe in Ringer’s solution adjusted to pH 7.3. h Quantification of TagRFP signal at the PM in (g). Values were normalized to the WT average (set as 1). (WT: n = 122; TM9SF3−/− #1: n = 105; TM9SF3−/− #2: n = 112). P-values were calculated using one-way ANOVA followed by Tukey’s multiple comparisons test. **** indicates P < 0.0001. The P value for the comparisons between TM9SF3−/−#1 and #2 is 0.8495. i Mass spectrometric analysis of PIP2 levels in WT and TM9SF3−/− #1 293F cells after 10 minutes incubation in Ringer’s solution adjusted to indicated pH (n = 3). P-values were calculated using 2-way ANOVA followed by Tukey’s multiple comparisons test. All P-values exceed 0.990. j Cell surface biotinylation assay in DLD-1 cells. Biotinylated cell surface proteins appear in the “PM” lane; unbiotinylated cytosolic proteins appear in the “Cyto” lane. k Quantitative analysis of TM9SF3 localization from (j). The relative expression levels were normalized to the cytoplasmic signal (set as 1; n = 3 biological replicates). P-values were calculated using a two-tailed unpaired t-test. P-value is 0.0824. l Schematic illustration of TM9SF3-mediated PIP2 translocation in response to acidic extracellular pH. m Confocal images of Dictyostelium AX2 (WT), phg1a-, and phg1b- cells incubated with the PIP2 probe in a culture medium adjusted to pH 6.5 or pH 7.0. n Quantification of TagRFP signal at the PM in (m). Values were normalized to the average of WT cells at pH 7.0 (set as 1) (WT, n = 106 (pH 6.5), n = 57 (pH 7.0); phg1a-, n = 167 (pH 6.5), n = 113 (pH 7.0); phg1b-, n = 127 (pH 6.5), n = 119 (pH 7.0)). P-values were calculated using a two-tailed unpaired t-test for each genetically distinct Dictyostelium strain. **** indicates P < 0.0001. The P value for the comparison between phg1b- at pH 7.0 and 6.5 is 0.3144. b, h, i, k, n Data are presented as mean values ± SD. All source data are provided as a Source Data file. PBase PiggyBac transposase, PB PiggyBac, SB Sleeping Beauty, SA splice acceptor, pA bidirectional SV40 polyadenylation signal, CAG cytomegalovirus enhancer and chicken beta-actin promoter, SD Foxf2-exon-1 splice donor, MW molecular weight, PM plasma membrane, Cyto Cytosol. Scale bars, 20 µm.
Although TM9SF3 is upregulated in gastric cancer27, the function of TM9SF3 is still unclear. TM9SF3 is predicted to be a nine membrane-spanning molecule with a large extracellular domain at the N-terminus, based on the sequence of amino acids28 (Fig. 2e). To assess the potential involvement of TM9SF3 in the pH-dependent PIP2 translocation machinery, we established two independent 293F cell clones harboring biallelic mutations in TM9SF3 (TM9SF3−/− #1 and #2) by CRISPR/Cas9 system (Fig. 2f). Both clones completely lost the ability to translocate PIP2 in pH 7.3 condition (Fig. 2g, h). This observation was further corroborated by TM9SF3 knockdown cells (Supplementary Fig. 3d–f). Neither did DLD-1 cells lacking TM9SF3 expose PIP2 on the outer leaflet of the PM (Supplementary Fig. 3g, h), indicating that TM9SF3 is essential for PIP2 translocation regardless of cell type. The requirement for PIP2 translocation was exclusive to TM9SF3 in our screening. Knockdown of other candidate transmembrane proteins, such as TMEM253 and KCNJ3, did not affect PIP2 translocation (Supplementary Fig. 3i).
We further confirmed that the depletion of TM9SF3 did not affect the total amount or the diversity in the molecular species of PIP2 (Fig. 2i and Supplementary Fig. 4a–c). It is thus unlikely that TM9SF3 regulates PIP2 synthesis. A biotinylation assay for cell surface proteins showed that endogenous TM9SF3 is localized both at the PM and within intracellular compartments (Fig. 2j, k, and Supplementary Fig. 4d, e). Furthermore, TM9SF3 at the PM in vivo was confirmed by the fractional analysis of the mice intestine and liver (Supplementary Fig. 4f, g). Collectively, these findings indicate that TM9SF3, localized at the PM, may mediate extracellular acidification-dependent PIP2 translocation, although the underlying mechanism remains unknown (Fig. 2l).
To address the evolutionary origin of the pH-dependent PIP2 translocation, we examined Dictyostelium discoideum (D. discoideum) amoeba, the most primitive organism known to possess PIs, including PIP229. We found that D. discoideum exhibited PIP2 translocation in a pH-dependent manner (Fig. 2m, n). The mutant lacking Phg1b, the ortholog of TM9SF1/330, exhibited the lack of PIP2 translocation under low pH conditions, while Phg1a, the ortholog of TM9SF2/4, did not (Fig. 2m, n). These results suggest that the function of TM9SF3 as a pH transducer is evolutionarily conserved from invertebrates to vertebrates.
Requirement of TM9SF3 for the translocation of PIP2 in vivo under low pH
To explore the biological relevance of pH-dependent TM9SF3-mediated PIP2 translocation in vivo, we used zebrafish embryos. We initially examined the dynamics of pH during zebrafish gastrulation, as it involves dynamic cell migration that consumes high energy. A pH sensor reporter line, Tg(ef1:Lyn-mCherry-SEpHluorin) allowed us to visualize pHint and revealed that pHint became alkalinized from 6 to 14 hours post-fertilization (hpf) (Fig. 3a, b, and Supplementary Movie 1). To monitor the pHext, we observed the embryo injected with mCherry-SEpHluorin-GPI mRNA enabling us to visualize the pHext and found that the pHext gradually became acidic, although a ratiometry from mCherry-EGFP-GPI, a pH-insensitive control, did not change (Fig. 3a, c, and Supplementary Fig. 5a and Supplementary Movie 2). These results suggest that embryonic cells are exposed to a low pH environment during gastrulation. Furthermore, disruption of pH gradient across the PM by ionophores resulted in failure of gastrulation (Supplementary Fig. 5b), suggesting that the response to varying pH in cells might be important for morphogenesis during gastrulation.
a Schematic illustration of zebrafish embryogenesis from 6 to 14 hpf. b Ratiometric images of SEpHluorin/mCherry fluorescence intensity in a Tg(EF:Lyn-mCherry-SEpHluorin) embryo from 6 to 14 hpf. The color scale represents relative pH, with blue to purple indicating acidic conditions and yellow to red indicating alkaline. This color scheme is used consistently for all pH analyses in the study. Lateral view; anterior is up. c Ratiometric images of SEpHluorin/mCherry fluorescence intensity in an embryo injected with mCherry-SEpHluorin-GPI encoding mRNA from 6 to 14 hpf. Lateral view; anterior is up. d Whole-mount in situ hybridization showing tm9sf3 mRNA expression in zebrafish embryos at 4 hpf. e Western blot analysis comparing protein expression in WT and tm9sf3−/− zebrafish embryos at 4 hpf. f Immunofluorescence images of Tm9sf3 in WT and tm9sf3−/− zebrafish embryos at 4 hpf. Insets from the upper panel (yellow outline) are enlarged below. Animal pole view. g Confocal single-scanned cell images of isolated blastoderm cells from WT, tm9sf3−/−, and tm9sf3 mRNA-injected tm9sf3−/− zebrafish embryos at 4 hpf incubated with PIP2 probe in Danieau’s buffer adjusted to pH 7.2. h Quantitative analysis of (f). The mean fluorescence intensity of TagRFP on the PM was normalized to the average value in WT cells (set as 1) (WT, n = 181; tm9sf3−/−, n = 153; tm9sf3−/− + Tm9sf3, n = 137). P-values were calculated using one-way ANOVA followed by Tukey’s multiple comparisons test. **** indicates P < 0.0001. The P value for the comparison between WT and tm9sf3−/− + Tm9sf3 mRNA is 0.1522. i Confocal scanned z-stack images of WT and tm9sf3−/− embryos at 7 hpf, injected with mRNA encoding membrane-anchored EGFP and a secretable PIP2 probe (see Methods) at 7 hpf. Enlarged views of EVL and blastoderm cells are shown. Animal pole view. j Images depicting littermate WT, tm9sf3+/-, and tm9sf3−/− zebrafish at 1 month post-fertilization (mpf). Dashed lines indicate the body length used for quantification. Lateral view. k Quantitative analysis of body length from (j) (WT, n = 54; tm9sf3+/-, n = 96; tm9sf3−/−, n = 7). P-values were calculated using one-way ANOVA followed by Tukey’s multiple comparisons test. The P values for the comparison are as follows: WT vs tm9sf3+/-, 0.6064; WT vs tm9sf3−/−, 0.0011; and tm9sf3+/- vs tm9sf3−/−, 0.0274. l Survival rate of zebrafish shown in (j). h, k Data are presented as mean values ± SD. All source data are provided as a Source Data file. SE superecliptic, EVL enveloping layer, WT wild-type. Scale bars, 100 µm in (b, c), 200 µm (d), 50 µm (f), 20 µm (g, i) and 1 mm (j).
To investigate the role of TM9SF3 in the developing embryos, we generated the loss-of-function alleles, tm9sf3ncv135 (Supplementary Fig. 5c). Sequencing of homozygous mutant fish showed 14 bases deletion in tm9sf3 (hereafter referred to as tm9sf3−/−) (Supplementary Fig. 5c, d). Whole-mount in situ hybridization (WISH) unveiled the ubiquitous expression of tm9sf3 mRNA from 2 to 48 hpf (Fig. 3d and Supplementary Fig. 5e). Western blot analysis of proteins extracted from maternal-zygotic tm9sf3−/− embryos revealed a depletion of Tm9sf3 (Fig. 3e). A very weak band was still observed in the mutant lane. However, a second, independently generated tm9sf3 mutant line also showed the same weak band (Supplementary Fig. 5f, g). Therefore, we believe that this band does not represent residual TM9SF3, but rather cross-reactivity with other members of the TM9SF family - such as TM9SF1, TM9SF2, or TM9SF4 - which share 33 %, 26 %, and 27 % identity with the antigen sequence, respectively. Nonsense-mediated mRNA decay was not observed in tm9sf3−/− mutant at 6 and 10 hpf (Supplementary Fig. 5h, i). Furthermore, the mRNA expression of other TM9SF family members, tm9sf1, 2, and 4, remained unchanged, suggesting that the genetic compensation is unlikely to occur in tm9sf3−/−31 (Supplementary Fig. 5h, i). Given that tm9sf3 mRNA is maternally provided, maternal-zygotic tm9sf3−/− embryos were hereafter employed as knockout embryos in this study unless otherwise specified.
Tm9sf3 was not detectable in the PM of both enveloping layer (EVL) and blastoderm cells in tm9sf3−/− embryos (Fig. 3f and Supplementary Fig. 5j). Blastoderm cells from WT but not those from the tm9sf3−/− embryos showed localization of PIP2 in the outer leaflet when cells were cultured under pH 7.2 (Fig. 3g, h). Furthermore, we investigated whether TM9SF3-dependent PIP2 translocation occurs in embryos as well as isolated blastoderm cells. Another PIP2 probe, designed to be secreted and containing tandem PIP2 binding domains to enhance affinity for PIP2, secreted from the cells bound to the PM of EVL and blastoderm cell in WT embryos, but not in tm9sf3−/− embryos (Fig. 3i).
To examine whether TM9SF3 contributes to zebrafish development, we compared the growth of tm9sf3−/− embryos with WT embryos. The maternal-zygotic tm9sf3−/− embryos grew normally up to 24 hpf (Supplementary Fig. 5k). To enable comparisons among littermates, we crossed heterozygous mutants to obtain zygotic tm9sf3−/− mutant embryos and WT embryos. From larva to juvenile stage, the zygotic tm9sf3−/− fish exhibited growth retardation and increased mortality during 7 dpf to 1 month post-fertilization (mpf) (Fig. 3j–l and Supplementary Fig. 5l–n). These results suggest that Tm9sf3, involved in the flopping of PIP2 in response to fluctuating pHext, is essential for development.
TM9SF3 functions under low pH during embryogenesis
We were intrigued by the mechanisms through which cells respond to fluctuating pHext during the morphogenesis of gastrulation (Fig. 3a–c). PIP2 is crucial for coordinating a functional cytoskeleton with the PM17, thereby regulating cell motility. Thus, we assumed that cell motility might be affected in the tm9sf3−/− embryos. We examined the migration of prechordal plate (ppl) cells in the anterior axial mesoderm using the reporter line Tg(gsc:EGFP-CAAX)32 (Fig. 4a). The pHext surrounding ppl became gradually acidic between 4 to 10 hpf (Supplementary Fig. 6a). In WT embryos, ppl showed cohesive and well-coordinated collective cell migration during 6.5 to 7 hpf, whereas tm9sf3−/− embryos showed reduced cohesion within ppl (Fig. 4b and Supplementary Movie 3). In the mutant embryos, ppl cells tended to migrate individually toward the animal pole, exhibiting spiky membrane protrusions. Disorganized migration of ppl was quantified by tracking cells marked by introduced nuclear TagBFP within ppl (Fig. 4c and Supplementary Fig. 6b and Supplementary Movie 4, 5). Ppl cells in tm9sf3−/− embryo exhibited a loss of coordinated cell migration, characterized by random and rapid movement compared to WT cells (Fig. 4d, e). Consequently, the displacement length of the tracked path was greater in tm9sf3−/− embryo than in WT (Supplementary Fig. 6c, d). These results suggested that ppl cells adapt to pH fluctuation to control their migratory behavior through TM9SF3-mediated PIP2 translocation.
a Schematic illustration of Tg(gsc:EGFP-CAAX) embryos from dorsal view during 6.5 to 7 hpf. EGFP marks gsc-positive cells. b Representative images of Tg(gsc:EGFP-CAAX) in WT and tm9sf3−/− embryos at 6.5 and 7 hpf. Yellow arrows indicate individual cells detached from the collective migration. Areas outlined in red are enlarged and displayed to the right. c Nuclei tracking of ppl cells labeled with H2A-BFP in WT and tm9sf3−/− embryos from 6.5 to 7 hpf. “Spot” colors represent displacement length, while “track” colors represent elapsed time. d, e Quantitative analysis of (c). 3D trajectories of individual cell migration from 6.5 to 7 hpf, with each cell’s initial position (X, Y, Z) at 6.5 hpf set to (0, 0, 0) (d). Colors indicate elapsed time. 2D plot of displacement lengths for each cell from 6.5 to 7 hpf (e). For each time point, WT (n = 108-123 cells) and tm9sf3−/− (n = 113–130 cells) were analyzed. Data are presented as mean values ± SD. f Time-lapse images of ppl in tm9sf3−/−; Tg(gsc:EGFP-CAAX) embryo injected with Lifeact-RFP mRNA from 6.5 to 7 hpf. Time indicates the elapsed time. Yellow asterisk marks the cell isolated from the collective ppl, and white arrows indicate membrane protrusions. g Schematic illustration of cell migration shown in (f), indicated by an asterisk. All source data are provided as a Source Data file. Scale bars, 50 µm (b), 10 µm (enlarged images in b), 200 µm (c), and 20 µm (f).
PIP2 binds to actin regulators controlling membrane blebs, lamellipodia, and filopodia. Ppl cells exhibit predominant blebs and lamellipodia and less filopodia during collective migration33. To visualize the actin cytoskeletal change in the ppl cells of tm9sf3−/− embryos, we introduced Lifeact-RFP and observed alterations in membrane shape. Ppl cells of tm9sf3−/− embryos showed filopodia-like protrusion and consequently detached from the ppl by decreased strength of cell-cell adhesions (Fig. 4f, g and Supplementary Fig. 6e and Supplementary Movies 6, 7). These results suggested that the cells were not damaged; instead, they actively altered their migration properties from blebs/lamellipodia to filopodial protrusion to move individually in tm9sf3−/− embryo.
TM9SF3 regulates ppl migration in response to pHext by modulating cytoskeletal dynamics
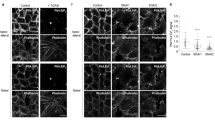
We investigated the molecular mechanisms by which Tm9sf3 regulates the migration of ppl cells in response to varying pHext during gastrulation. We hypothesized that cell functions, such as adhesion and migration, which are essential for coordinated migratory behavior, might be disorganized in tm9sf3−/− embryos. To investigate the cellular properties underlying these behaviors, we analyzed non-muscle myosin II (myosin) activity - which regulates cell-cell adhesion by concentrating cadherins at contacts34 - as well as actin polymerization activity. We performed immunostaining for phosphorylated myosin and filamentous actin (F-actin), respectively. During gastrulation, from 7 to 8 hpf, ppl in WT embryos exhibited higher levels of phosphorylated myosin and F-actin compared to ppl in tm9sf3−/− embryos at 8 hpf (Fig. 5a–c). Spatial analysis revealed that phosphorylated myosin levels were relatively higher at the front of the ppl compared to the rear in WT embryos; however, this spatial distribution was absent in tm9sf3−/− embryos at 8 hpf (Fig. 5d). Conversely, in WT embryos, F-actin levels were relatively higher at the rear of the ppl compared to the front (Fig. 5d). This inverse relationship, indicative of an anti-correlation between phosphorylated myosin and F-actin, was absent in tm9sf3−/− embryos (Fig. 5e). The uncoordinated activity of myosin and actin polymerization in the ppl of tm9sf3−/− embryos resulted in cells at the leading edge exhibiting an increased number of filopodia (Fig. 5f). These unbalanced adhesive and migratory properties likely reduce cohesiveness and enhance the migratory rate, ultimately disrupting collective cell behavior (Fig. 4). To analyze how PIP2 levels in the inner memebrane influences cellular activities, we introduced the PIP2-producing enzyme PI(4)P 5-kinase I-gamma (PIP5K) into WT embryos to promote PIP2 production in the PM19. PIP5K-injected embryos exhibited reduced levels of phosphorylated myosin and polymerized actin (Supplementary Fig. 7a, b), resembling the phenotype observed in the ppl of tm9sf3−/− embryos. These findings suggest that decreased PIP2 levels in the inner membrane in response to a low pH environment reduce actin polymerization while enhancing myosin activity, both of which are essential for spatially regulated collective cell migration.
a Immunofluorescence images showing phosphorylated myosin light chain (pMLC), filamentous actin (F-actin), and nuclei in Tg(gsc:EGFP-CAAX) at 8 hpf in WT and tm9sf3−/− backgrounds. Quantitative analysis of pMLC (b) and F-actin (c) levels in ppl at 7 and 8 hpf, corresponding to (a). Mean fluorescence intensity per embryo is plotted, with WT at 7 hpf set as 1 (WT: n = 9 at 7 and 8 hpf; tm9sf3−/−; n = 9 at 7 and 8 hpf). Data are presented as mean values ± SD. d Digital section images from (a), showing the distribution of pMLC and F-actin across ppl cells. The color spectrum indicates relative signal intensity. Regions corresponding to the front and rear of ppl are highlighted with yellow and blue boxes, respectively, and shown at higher magnification below. e Quantitative comparison of the pMLC to F-actin ratio from the front to the rear of ppl based on (d). f Live images of ppl cells at the leading edge of Tg(gsc:EGFP-CAAX) in WT and tm9sf3−/− backgrounds expressing injected Lifeact-RFP mRNA at 7 hpf. White arrows indicate filopodia formation at the leading edge of ppl. WT and tm9sf3−/− embryos correspond to those in Supplementary Fig. 6, (e) and Fig. 4, (f), respectively. Results are representative of two independent embryos. g Fluorescence images of mixed ppl cells from WT and tm9sf3−/− embryos incubated in the medium adjusted to pH 7 or 8 for 1, 4, and 8 hours. Results are representative of two independent experiments. h Schematic model summarizing cytoskeletal dynamics in WT and tm9sf3−/− ppl cells under low pH environment. All source data are provided as a Source Data file. Scale bars, 50 µm (a, d), 10 µm (enlarged panels in d), 20 µm (f, g).
PIP2 functions as a signaling substrate; however, no defects in cell fate specification were observed, as indicated by the normal expression level of mesodermal markers (goosecoid (gsc), no tail (ntl)) and endodermal markers (sox17 and casanova (cas)) (Supplementary Fig. 7c). Further analysis of cell mortality and proliferation in ppl at 7 hpf was performed using immunostaining for cleaved caspase-3 and phospho-histone H3, respectively. The number of phospho-histone H3-positive cells was comparable between groups, suggesting consistent levels of proliferation (Supplementary Fig. 7d and e). Similarly, cleaved caspase-3-positive cells were not detected in ppl, indicating an absence of apoptosis-mediated cell death in both WT and tm9sf3−/− embryos (Supplementary Fig. 7f). These results indicate that WT and tm9sf3−/− embryos exhibit similar cell populations and comparable cellular conditions. Collectively, these results suggest that the lack of coordinated cell behavior in ppl of tm9sf3−/− embryos is attributable to the disruption of the spatial coordination of cytoskeletal components.
To examine how cytoskeletal changes in response to low pH affect cell behavior in a cell-autonomous manner, we analyzed ppl cells from WT and tm9sf3−/− embryos using an in vitro system. Ppl cells were induced via cyclops mRNA injection35 and confirmed by goosecoid promoter-driven GFP expression (Supplementary Fig. 7g). WT ppl cells formed clusters more rapidly than those from tm9sf3−/− embryos, indicating greater cohesion at pH 7 (Supplementary Fig. 7h). In culture medium conditioned to pH 7, but not pH 8, WT ppl cells exhibited greater cohesiveness than tm9sf3−/− ppl cells, which underwent cell sorting after 8 hours of incubation (Fig. 5g and Supplementary Fig. 7i–l and Supplementary Movie 8, 9). These results suggest that low pH, sufficient to translocate PIP2, promotes cell clustering, potentially via upregulated myosin activity, which is essential for collective cell migration.
Discussion
In this study, we demonstrated that extracellular acidic pH triggers the flopping of PIP2. Under physiological conditions, pHext is maintained in a slightly alkaline environment (pH 7.3-7.4) compared to pHint (pH 7.2)36. The threshold of PIP2 translocation from the inner to the outer leaflet, approximately pH 7.3, suggests the presence of a finely tuned pH response system in the PM, enabling cells to adapt to fluctuating environments.
All embryonic cells are surrounded by extracellular fluid, whose physical and chemical properties may influence organismal development. However, how variations in these fluid properties affect cellular behavior during early development remain largely unknown. Here, we show that during early zebrafish development, pHext gradually decreases after the onset of gastrulation. In Tm9sf3-deficient embryos, impaired adaptation to pH leads to distinct ppl migration phenotypes due to disorganized cytoskeletal dynamics, where migration overrides adhesion. Our findings indicate that a decrease in pHext promotes ppl cell clustering and enhances tissue cohesiveness - processes essential for morphogenesis - highlighting pHext as a critical regulator of early development.
PIP2 is a well-established positive regulator of the actin cytoskeleton37. Potentially elevated inner leaflet PIP2 levels in Tm9sf3-deficient ppl cells likely contribute to their increased migratory activity. This is evident from the highly dynamic, actin-rich protrusions at the leading edge of Tm9sf3-deficient ppl, which may facilitate cell dissociation and enhance overall cell movement. These effects may be mediated by PIP2-dependent cytoskeletal modulators such as Neuronal Wiskott-Aldrich syndrome protein (N-WASP), formins, and Ezrin/Radixin/Moesin (ERM) proteins, which play key roles in linking the cytoskeleton to the PM17,33. The PDGF-PI3K pathway has been reported to be crucial for ppl cell migration during gastrulation38. PI3K utilizes PIP2 as a substrate to produce PIP3, an instructive signal that drives actin polymerization and promotes cell migration. This signaling pathway may be upregulated in Tm9sf3-deficient cells, as indicated by their strong protrusive activity. Despite the observed reduction in F-actin levels, as indicated by phalloidin staining, Tm9sf3-deficient cells exhibit increased actin dynamics. This may suggest that actin assembly, turnover, and spatiotemporal regulation are enhanced to support cell migration. The interplay between PIP2 and cytoskeletal remodeling mechanisms likely enables these cells to maintain their motility despite a lower overall F-actin content. Previous studies have reported that PIP2 binding to vinculin promotes focal adhesion (FA) dynamics39,40. Thus, one possible scenario is that increased PIP2 levels at the inner leaflet of the PM, due to the loss of tm9sf3, may enhance FA turnover and thereby promote cell migration, as demonstrated in Fig. 4e. Further investigation is required to elucidate the precise molecular pathways by which Tm9sf3 deficiency alters actin dynamics and facilitates cell migration.
In WT embryos, myosin activity at the cell periphery in ppl cells increased during gastrulation, with particularly elevated activity at the leading edge of the ppl. However, in Tm9sf3-deficient embryos, these spatiotemporal patterns of myosin activity were not observed. This is likely due to a reduced pH response caused by impaired TM9SF3-mediated PIP2 translocation, as an increased PIP2 level, induced by PIP5K, has also been shown to decrease myosin activity. Since myosin activity regulates the formation and stability of the cell-cell junctions, myosin-driven contractility is crucial for contact formation during zebrafish gastrulation41. Consequently, reduced myosin activity in Tm9sf3-deficient embryos likely leads to decreased cell cohesion, as seen in the more random movement of ppl cells. When induced ppl cells from WT and Tm9sf3-deficient embryos were mixed, WT cells exhibited higher cohesiveness than Tm9sf3-deficient cells at pH 7, but not at pH 8. These results suggest that extracellular pH influences cell cohesion via Tm9sf3, though how decreased inner PIP2 or increased outer PIP2 regulates cell cohesion remains unclear. So far, no specific molecules have been identified that regulate myosin activity in ppl cells. Further analysis is needed to determine how pH influences cell behavior through specific signaling pathways.
PIP2 is a key regulator of various ion channels and transporters17,42. Recent studies have identified that a proton-activated chloride channel (PAC) is inhibited by PIP2 binding to its extracellular domain43,44,45,46. Interestingly, the pH thresholds for PIP2 translocation (<pH 7.3) and PAC activation (<pH 5.0) are distinct, suggesting that TM9SF3 may play a role in preventing excessive PAC activation under acidic environments. This implies that the pH-dependent regulatory function of PIP2 extends beyond the inner leaflet to the outer leaflet of the PM. Further investigation is required to understand how PIP2 level alterations across both leaflets impact cellular function and how cells detect and respond to pH gradient at the PM, particularly through the role of TM9SF3 in PIP2 translocation. While we have demonstrated that TM9SF3 is essential for pH-dependent PIP2 translocation, detailed mechanistic insights are still required to understand how cells sense pH gradients across the PM and how TM9SF3 contributes to PIP2 translocation.
Elevated TM9SF3 expression levels have been reported in peripheral blood T cells of T-cell leukemia patients47 and in tumor tissues of gastric cancer patients, where it correlates positively with malignancy27. Functionally, TM9SF3 has been shown to promote cancer cell invasion and proliferation, highlighting its potential role in carcinogenesis. A defining feature of the tumor microenvironment is acidosis, driven by the accumulation of acidic metabolic byproducts such as lactic acid due to heightened metabolic activity and poor perfusion48. Cancer cells with increased TM9SF3 expression may exhibit enhanced adaptation to these heterogeneous acidic conditions, promoting collective migration and facilitating metastasis49. Thus, targeting TM9SF3 could be a promising therapeutic strategy to disrupt cancer cell adaptation to acidosis.
Our findings suggest that PIP2 undergoes functional modifications not only through conventional mechanisms - such as enzymatic regulation of its expression and localization17 - but also through topological distribution changes at the PM. This introduces a distinct regulatory mechanism for multifunctional signal lipids. The PM is characterized by an asymmetric phospholipid distribution, with PIs, PS, and PE predominantly localized in the inner leaflet. Our results indicate that vertical lipid translocation, triggered by external stimuli, occurs more frequently than previously recognized and plays a critical role in lipid function at the PM. These findings suggest that lipids within the bilayer may acquire other functional roles by modulating their topology in response to environmental cues. We anticipate that future research will uncover previously unknown lipid functions in cellular signaling and adaptation.
Methods
Zebrafish (Danio rerio) strains, transgenic lines, and mutant lines
The experiments using zebrafish were approved by the animal experimentation committee of the institutional review board of the National Cerebral and Cardiovascular Center (permit number: 23041), and were performed according to the guidelines of the Institute. Embryos were raised in E3 medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2・2H2O, 0.33 mM MgSO4・7H2O) and incubated at 28.5 °C50. Developmental staging was referred to chronological and morphological criteria51. We used the AB strain as wild-type (WT). The following zebrafish lines, Tg(ef1:Lifeact-EGFP) ncv518, Tg(ef1:Lyn-mCherry-SEpHluorin) ncv520, and tm9sf3ncv135 used in this study were generated as described in the experimental procedures. Tg(gsc:EGFP-CAAX)32 were kindly provided by C-P. Heisenberg (Institute of Science and Technology, Austria).
Generation of transgenic zebrafish lines
To generate Tg(ef1:Lifeact-EGFP)ncv518 and Tg(ef1:Lyn-mCherry-SEpHluorin) ncv520 zebrafish lines, the corresponding Tol2-based plasmid DNA (25 pg) was microinjected along with Tol2 transposase mRNA (25 pg)52 into one-cell stage embryos of AB. Tol2 transposase mRNA was in vitro transcribed from NotI-linearized expression plasmids with SP6 RNA polymerase using a mMessage mMachine mRNA kit (AM1340, Thermo Fisher Scientific). The embryos were raised to adulthood and crossed with AB to identify germline-transmitting founder fishes. Throughout the text, all Tg lines used in this study are simply described without their line numbers. For example, Tg(ef1:Lifeact-EGFP) ncv518 is abbreviated to Tg(ef1:Lifeact-EGFP).
Generation of knockout zebrafish line by transcription activator-like effector nuclease (TALEN)
TALEN targeting tm9sf3 was designed using TAL Effector Nucleotide Targeter 2.0 (https://tale-nt.cac.cornell.edu) and was assembled via the Golden Gate method53. TALEN repeat carriable di-residues (RVDs) were cloned into an RCIscript-Goldy TALEN vector (Addgene #38142). tm9sf3 TALEN mRNA was in vitro transcribed from SacI-linearized expression plasmids with T3 RNA polymerase using a mMessage mMachine mRNA kit (AM1348, Thermo Fisher Scientific). Embryos, injected with 15 pg of the TALEN mRNA at the one-cell stage, were raised to adulthood and crossed with AB to identify germline-mutated founders. Screening for founders was conducted by genomic PCR and subsequent sequencing using the following primer sets: 5’- CAACCACTGTCATAGAACCAGGA -3’ and 5’- TTCTCGGCCCGCAGTTTATC -3’. By sequencing the PCR product, we identified the tm9sf3ncv135 allele harboring a 14 nucleotide deletion and another tm9sf3 allele harboring an 8 nucleotide deletion, both in the first exon (Supplementary Fig. 5c, d, f, g).
Plasmids
cDNA fragment encoding zebrafish Tm9sf3 was amplified by PCR using a cDNA library derived from zebrafish embryos and subcloned into pCR4 blunt TOPO vector (Invitrogen) to generate pCR4-TOPO_Tm9sf3. Tm9sf3 was amplified by PCR and subsequently subcloned into a pCS2+ vector using BamHI and XhoI sites to generate pCS2+_Tm9sf3. Lyn-INPP5E19 was subcloned into pEGFP-N3 using EcoRI and BamHI sites to generate pEGFP_Lyn-INPP5E, following Lyn-INPP5E-EGFP was subcloned into pCS2+ vector using EcoRI and XbaI sites to generate pCS2+_Lyn-INPP5E-EGFP. Tandem PIP2 binding PH domain of mouse PLC δ154 tagged with TagRFP was subcloned into a pCS2+ vector. Subsequently, the N-terminal signal sequence of a GPI-anchored T-cadherin (MQPRTPLTLCVLLSQVLLVTSAV)55 was fused to the N-terminus of TagRFP-mPLCδ1(x2) for facilitating secretion, using inverse PCR method.
The PH domain of PLCδ156 and Akt57, and the P4M domain of SidM58 were amplified by PCR and subsequently subcloned into pGEX_TagRFP to generate pGEX_TagRFP-PLCδ1-PH, pGEX_TagRFP-Akt-PH, and pGEX_TagRFP-P4M-SidM vectors, respectively. K30L, K32L, and R40L mutations, designed to generate PIP2 non-binding probe20, were introduced into pGEX_TagRFP-PLCδ1-PH using PCR-based mutagenesis with the primer sets: 5’- GGCAGCCAGCTTCTGTTGGTGTTGTCCAGCTCGTGGCGTAGGGAA-3’ and 5’- CTGTAGCTTGTAGAAGAGTTCCCTACGCCACGAGCTGGACAACAC-3’. TagRFP-PLCδ1-PH was amplified by PCR and subsequently subcloned into pCS2+ vector to generate pCS2+_TagRFP-PLCδ1-PH, following TagRFP was replaced with EGFP amplified by PCR using EcoRI and Bglll sites to generate pCS2+_EGFP-PLCδ1-PH. EGFP- PLCδ1-PH was further subcloned into a pPBbsr2 vector using EcoRI and XhoI sites.
Not I site in pEGFP-Lyn-tailed mCherry-SEpHluorin (Addgene#32002) was replaced with XbaI site. Lyn-tailed mCherry-SEpHluorin was subcloned into pCS2+ vector using XhoI and XbaI sites to generate pCS2+_Lyn-tailed mCherry-SEpHluorin. N-terminal fusion of membrane translocation signal sequence from mouse acrosin (MVEMLPTVAVLVLAVSVVAKDNTT) and linker sequence (LQEFAT), and C-terminal fusion of linker sequence (LE) and mouse Thy-1 GPI anchor signal sequence (with a cysteine-to-asparagine mutation at the ω site of mouse Thy-1 GPI anchoring sequence indicated as an underline) (NGGISLLVQNTSWMLLLLLSLSLLQALDFISL) were generated by PCR59, followed subcloned into pCS2+ vector to produce pCS2+_mCherry-SEpHluorin-GPI. SEpHluorin was replaced with EGFP using an In-Fusion HD cloning kit (639650, TAKARA Bio) to produce pCS2+_mCherry-EGFP-GPI.
The Tol2 vector systems were kindly provided by K. Kawakami60,61 (National Institute of Genetics, Japan) and P. Pantazis62 (ETH Zurich, Switzerland). Lyn-tailed mCherry-SEpHluorin and Lifeact-EGFP were subcloned into pTol2 vector, which has a Xenopus EF1α enhancer-promoter60,61.
Mouse PIP5KIC63519 and tagBFP were subcloned into the pCS2+ vector using In-Fusion cloning to construct pCS2 + _PIP5K-tagBFP. Additionally, the PM-targeting signal (MGCIKSKRK, a myristoylation motif derived from Lyn tyrosine kinase) was replaced with PIP5KIC635 to generate pCS2 + _Lyn-tagBFP.
Reagents
The following primary antibodies were used in this study: anti-TM9SF3 (HPA039609, Atlas antibodies); anti-CD29 (Integrin β1) (610467, BD Bioscience); anti-β-actin (A5441, Sigma-Aldrich); anti-phospho-Myosin Light Chain 2 (3671, Cell Signaling); anti-ZO1 (33-9100, Invitrogen); anti-E-cadherin (61081, BD Bioscience); anti-MTCO1 (PA5-79701, Invitrogen); anti-Tubulin (DM1A, Sigma), anti-Na+, K+-ATPase (a kind gift from Dr. C. Toyoshima, The University of Tokyo, Japan), anti-Cleaved Caspase-3 (#9661, Cell Signaling), anti-Phospho Histone H3 Alexa Fluor 647 conjugate (#3458, Cell Signaling). The antibody against zebrafish TM9SF3 used for immunohistochemistry was generated in this study (BiCell). The peptide used as antigen was designed (YHNRQETYKYFSLPF) and synthesized, and immunized in rabbits. The secondary antibodies used for immunohistochemistry were Alexa 488-conjugated anti-rabbit IgG and anti-rat IgG (A11034 and A11006, Thermo Fisher Scientific), used for western blot were peroxidase-linked anti-mouse, rat, and rabbit IgG (NA931, NA934, NA935, Amersham), and used for whole-mount in situ hybridization was anti-DIG antibody (11093274910, Roche).
The following drugs were used in this study: Valinomycin and nigericin (P35379, Thermo Fisher Scientific); Oligomycin A (11342, Cayman Chemicals); BAPTA (B-6769, Molecular Probes); Ionomycin (407950, Calbiochem); FCCP (C2920, Sigma-Aldrich); DMSO (041-29351, FUJIFILM Wako).
Synthesis and injection of mRNA
mRNA was synthesized using SP6 mMessage mMachine Kit and injected into one-cell stage embryos. The following templates were used in this study: pCS2+_mCherry-SEpHluorin-GPI; pCS2+_mCherry-EGFP-GPI; pCS2+_Tm9sf3; pCS2+_h2afva-tagBFP63; pCS2+_Lyn-INPP5E-EGFP; pCS_zT2TP52; pCS2+_zBax13,64; pCS2+_membrane anchored-EGFP; pCS2+_Lifeact-RFP65; pCS2 + _Lyn-tagBFP; pCS2 + _PIP5K-tagBFP; pCS2 + _cyclops66.
Cell culture
Mouse embryonic fibroblast cells (MEF) (CBA-310, Cell Biolabs, Inc.), DLD-1 (JCRB9094, National Institutes of Biomedical Innovation, Health and Nutrition), and 293F (R79007, Invitrogen) cells were used in this study. MEF in high glucose contained Dulbecco’s Modified Eagle Medium (DMEM) (043-30085, FUJIFILM Wako) and DLD-1 in RPMI1640 (11875093, Thermo Fisher Scientific) supplemented with 10% fetal bovine serum were cultured in a 37 °C incubator containing a humidified atmosphere of 5% CO2 in the air. 293F in FreeStyle 293 expression medium (12338018, Thermo Fisher Scientific) was cultured in a 37 °C incubator containing a humidified atmosphere of 8% CO2 in the air on an orbital shaker platform rotating at 135 rpm.
Image acquisition and analysis
Stereomicroscopic images were acquired with an SZX16 stereomicroscope with a DP2-BSW camera regulated with CellSens imaging software (Olympus); an SZX7-ILST-C stereomicroscope equipped with a DP21 camera (Olympus). Confocal images were acquired with upright FV1200 (UMPLFLN 10x/0.30 NA; XLUMPLFLN20XW 20x/1.00 NA, Olympus); upright FV3000 (UMPLFLN 10x/0.30 NA; XLUMPLFLN20XW 20x/1.00 NA, Olympus); inverted FV3000 (UPLXAPO 40x/0.95 NA; UPSAPO60XO 60x/1.35 NA, Olympus); inverted FV4000 (UPLSAPO 40x/0.95 NA); and inverted LSM880 (Plan Apochromat 63x/1.4 NA, ZEISS) confocal microscopes. For the fast time-lapse analysis, images were acquired with Andor spinning disc confocal Dragonfly200 (Andor Technology Ltd) based on ECLIPSE FN1 upright microscope (Nikon), equipped with water-immersion LWD 25x/1.1 NA, 16x/0.80 NA (Nikon), Zyla4.2 PLUS USB 3.0 sCMOS cameras (Andor Technology Ltd) and P-725.4 PIFOC piezo nano-positioner (Physik Instrumente) regulated with Fusion software (Andor Technology Ltd). For the super-resolution analysis, images were acquired with LSM880 with AiryScan. All images were acquired sequentially to avoid cross-detection of the fluorescent signals. Confocal images were processed and analyzed with Imaris software (ver 9.9 and 10.1) (Oxford Instruments); NIS-Elements (Nikon) to create 3D ratio images of SEpHluorin expressing embryos; and Fiji software67.
Preparation of recombinant protein biosensors
Recombinant protein probes for detecting lipids were prepared by a bacterial expression system. Respective constructs of TagRFP-PLCδ1-PH, TagRFP-Akt-PH, and TagRFP-P4M-SidM ligated into the pGEX bacterial expression vector were introduced into the BL21 (DE3) pLysS strain of Escherichia coli. Protein expression was induced by the addition of isopropyl β-D-1-thiogalactopyranoside (IPTG) at a final concentration of 0.5 mM, following incubation for overnight at 16 °C. Bacteria were harvested and lysed in phosphate-buffered saline (PBS) containing 1% Triton X-100 supplemented with 1 mM PMSF. After freeze-thawing followed by a brief sonication on ice, a crude extract was obtained in the supernatant after centrifugation at 12,000 x g for 30 min at 4 °C. The crude extract was incubated with glutathione sepharose (GSH) (GE Healthcare) for 1 h at 4 °C. GSH-Sepharose was washed with ice-cold PBS-Triton and subsequently with PBS to remove detergents using a column (Sarstedt). 1 ml of GSH buffer adjusted to pH8-9 with NaOH containing 50 mM Tris, 50 mM GSH, and 200 mM NaCl was added, and incubated on ice for 5 min. The obtained elution fraction was dialyzed in PBS for subsequent assays.
To measure the fluorescent intensity of TagRFP-PLCδ1-PH under different pH conditions, 2 μg of the probe was added to 1 ml of distilled water (DW) and dispensed 100 μl each into a 96-well plate. Following pH was adjusted using 20 mM HEPES to the pH range from pH 6.0 to 8.3. The fluorescence excited by a 550 nm wavelength was measured on a 96-well plate reader, POLARstarOmega-6 (BMG Labtech). After subtracting the measured fluorescent intensity of DW as background, the respective intensity was compared with the intensity at pH 7.3, serving as a reference point.
Lipid probes binding assay
Zebrafish embryos around 4 to 5 hpf were dechorionated in E3 medium and transferred to 200 μl DMEM without sodium bicarbonate (12100061, GIBCO) adjusted to the indicated pH with 20 mM HEPES (Fig. 1a, d, g and Fig. 3g and Supplementary Fig. 1a); CO2-independent L-15 medium (pH7.3) (Fig. 1e and Supplementary Fig. 1d); Annexin V binding buffer (K103-100, BioVision) (Fig. 1a, b). Embryos were mechanically dissociated into single cells by mild pipetting, following yolk proteins were removed by 2 successive wash steps with 200 μl same fresh medium, followed by centrifugation at 300 x g for 2 min. After 2 wash steps, dissociated cells were added with 100 μl of fresh medium and transferred to a 35-mm glass-bottom dish (Asahi Techno Glass), followed by the addition of 2 μg of indicated lipid binding probes, 0.25 μg of FITC-SA-Ro7, or Annexin V-Cy5. FCCP (1 µM), Valinomycin and nigericin (10 µM), EtOH, and DMSO were added from the washing step.
293F cells were added with 2 μg of lipid binding probes in 100 μl Ringer’s solution without NaHCO3 (128.3 mM NaCl, 1.13 mM CaCl2, and 1.34 mM KCl) adjusted to the indicated pH with 20 mM HEPES (Fig. 2a, g) or FreeStyle 293 expression medium (Supplementary Fig. 3a, e, i). MEF and DLD-1 cultured on a collagen-coated glass bottom dish were replaced with Ringer’s solution adjusted to the indicated pH with 20 mM HEPES containing the lipid probe (Supplementary Fig. 1f, g and 3h) or Live Cell Imaging Solution (pH 7.3) (A59688DJ, Thermo Fisher Scientific) (Supplementary Fig. 1h).
Images were acquired using LSM880 or FV3000 inverted confocal microscopes. The fluorescence intensity of the probes on the PM was quantified using Fiji software.
Binding assay of PIP2 and TagRFP-PLCδ1-PH probe
Silica beads-supported lipid bilayers were prepared using previously described protocols with modifications68. Stock solutions of PIP2 and phosphatidylcholine (PC) were individually aliquoted into polypropylene centrifuge tubes, and dried in vacuo (VC-36R, TAITEC). Lipids were hydrated in an HNa buffer (150 mM NaCl, 20 mM HEPES, pH 7.5) with vortexing, and were subjected to two freeze-thaw cycles using liquid nitrogen before sonication to generate liposomes using a Branson Sonifier 250 (Branson Ultrasonic). 5 ×106 silica beads (5 μm diameter, Bangs Laboratories, Inc.) in 100 μL of 400 mM NaCl, 20 mM HEPES (pH 7.5) were added with PIP2 liposomes (final concentration 1 μM) and incubated at 25 °C with intermittent vortexing. After 1 hour, PC liposomes (final concentration 20 μM) were added and further incubated for 15 minutes at 25 °C. The beads were washed once and resuspended in 100 μL of HNa buffers at different pH levels: pH 7.3 and pH 7.9.
The resultant lipid-coated beads were incubated with 2 μg of purified recombinant TagRFP-PLCδ1-PH protein at 25 °C for 15 minutes with continuous agitation. After centrifugation (10,000 x g for 30 seconds), the supernatants were transferred to new tubes, and the sedimented beads were added to HNa buffer containing 1% NP-40 for extraction. Concentrations of the recombinant protein in the supernatant (Cs) and bead fractions (Cb) were determined by fluorometric measurement with a Fluoroscan fluorometer at excitation 544 nm/emission 590 nm using ScanLT software (Thermo Fisher Scientific). Data are expressed as CB/(CS + CB).
Chemical treatments of zebrafish embryos
Embryos were treated with valinomycin and nigericin (10 µM each) to equilibrate extracellular and intracellular pH. As a control, the embryos were incubated in the buffer containing the same amount of DMSO.
Ratiometric pH measurement in zebrafish embryos
To monitor intracellular and extracellular pH (pHint and pHext) dynamics during development, Tg(ef1:Lyn-mCherry-SEpHluorin) embryos and mCherry-SEpHluorin (EGFP)-GPI mRNA-injected embryos were used to analyze the pHint and pHext, respectively. Embryos were dechorionated and mounted in 0.5% low-melting-point agarose (Invitrogen) on a 35-mm diameter glass-bottom dish and covered with E3 medium. pH was measured by calculating the SEpHluorin/mCherry fluorescence emission ratio at the PM using a 488 nm laser for SEpHluorin and a 561 nm laser for mCherry. Three-dimensional ratiometric images were generated using NIS-Elements software.
Ratiometric pH measurement in cell culture
For the ratiometric pH measurement, the fluorescence reporter encoding pCS2+-Lyn-mCherry-SEpHluorin and pCS2+-mCherry-SEpHluorin-GPI were transfected into DLD-1 for measuring the pHint and pHext, respectively. Plasmid transfections were performed using ViaFect (E4981, Promega) according to the manufacturer’s protocol. SEpHluorin and mCherry were excited at 488 and 561 nm lasers, respectively. The emitted fluorescence was acquired, followed by use to calculate the 488/561 ratio21. The signal of these probes expressed on the PM was measured and further analyzed for measuring pH. pH calibration standard was obtained by using Nigericin/Valinomycin-containing calibration buffers adjusted to the indicated pH (P35379, Thermo Fisher Scientific). The pH calibration data was applied to convert the ratio image to pH. The alterations of pHint and pHext in response to the varied extracellular fluid pH were calculated for every single cell and averaged, using Fiji software for the image analysis.
Generation of the stable cell line for EGFP-PLCδ1-PH by piggyBac transposon system
To establish a stable cell line, a piggyBac transposon-based system was applied. The fluorescence reporter vector pPBbsr2-EGFP-PLCδ1-PH was co-transfected with the transposase vector pCMV-mPBase (neo-)69, using viafect. One day after transfection, cells were added with 10 ug/ml blasticidin S (029-18701, FUJIFILM Wako), and selected for the stable cell line.
Genetic screening assay
The pATP1-CAGGS-Puro vector carrying cytomegalovirus enhancer, chicken β-actin promoter (CAG), and bifunctional activating/inactivating transposons24,25 was co-transfected with the transposase encoding pCMV-mPBase (neo-) vector69 into 293F cells, using 293 fectin (12347, Thermo Fisher Scientific) according to the manufacturer’s protocol. One day after transfection, cells were selected with 2 μg/ml Puromycin for the next 14 days (Sigma-Aldrich).
The cells that lost the ability to translocate PIP2 across the plasma membrane were enriched by repeating the FACS sorting. 1×106 mutagenized 293F cells in 1 ml of FreeStyle 293 expression medium were incubated with TagRFP-PLCδ1-PH probe at room temperature (RT) for 5 min. The cells were washed once with fresh medium and collected by centrifugation, and suspended in 1 ml of FreeStyle medium containing SYTOX Blue (Thermo Fisher Scientific) to remove the non-binding probe. The cells were subjected to flow cytometry using FACS Aria III (BD Bioscience), and a few percent of the cells, which exhibited less binding of the probe, were collected. The sorted cells were grown for 1 week in FreeStyle 293 expression medium containing Penicillin-Streptomycin, and subjected to a next round of sorting as described above. The resultant cells exhibiting less binding of the probe were designated as the mutant population.
Mapping of gene-trap insertion sites
The Splinkelette PCR method was performed to identify the transposon insertion site26. Genomic DNA was isolated from 7.5×106 293F cells using 750 μl cell lysis buffer containing 50 mM Tris-HCl (pH 8.0), 1 % SDS, 0.1 M EDTA (pH 8.0), and 100 mM NaCl. After adding 100 μg proteinase K, samples were reacted overnight at 55 °C. 250 μl 5 M NaCl was added, followed by centrifugation at 17,700 x g for 15 min at 4 °C. Supernatant was transferred to a new tube and added with 500 μl 2-propanol, followed by centrifugation at 17,700 x g for 15 min at 4 °C. The pellet was washed with 1 ml EtOH and dried, following dissolving into 50 μl DW.
10 μg isolated genomic DNA was digested with Sau3AI (R0169, New England Biolabs). The oligonucleotides, “Long-strand adaptor” (5’- CGAAGAGTAACCGTTGCTAGGAGAGACCGTGGCTGAATGAGACTGGTGTCGACACTAGTGG -3’) and “Short-strand adaptor” (5’- GATCCCACTAGTGTCGACACCAGTCTCTAATTTTTTTTTTCAAAAAAA -3’) were resuspended in 5x NEB buffer 2 to a concentration of 50 µM, and mixed at a final concentration of 25 µM each26. This adaptor mix was denatured and annealed by heating it to 95 °C for 5 min and then cooling it to RT at the rate of 1 °C every 15 sec. Sau3AI-digested genomic DNA was heated at 65 °C for 20 min and placed on ice, followed reacted with an adaptor mix and T4 DNA ligase (2011, TAKARA Bio) in T4 DNA ligase buffer overnight at 4 °C. Reaction was terminated by heat inactivation of T4 DNA ligase at 65 °C for20 min. Splinkelette adaptor-ligated DNA was digested with EcoRV (R0195, New England Biolabs) at 37 °C for 6 h. The reaction was terminated by heat inactivation of T4 DNA ligase at 65 °C for 20 min. The ligated DNA was purified by the QIAquick Gel Extraction Kit (28704, QIAGEN) according to the manufacturer’s instructions. The DNA purified as above was subjected to a nested PCR using the 1st PCR primer sets (5’- CGAAGAGTAACCGTTGCTAGGAGAGACC -3’ and 5’-GCGTCATTTTGACTCACGCGGTCGTTATAG -3’) and 2nd PCR primer sets (5’-GTGGCTGAATGAGACTGGTGTCGAC -3’ and 5’-AAATCAGTGACACTTACCGCATTG -3’). Amplified PCR products were cloned into the pCR4 blunt TOPO vector and sequenced.
Generation of knockout cell lines by CRISPR/Cas9 system
The 20-nt-long CRISPR RNA (crRNA) was predesigned and synthesized by Integrated DNA Technologies (IDT). The sequence of crRNA for targeting the TM9SF3 gene was 5’- GTTTCTTGACGATTATGGTA -3’. To assemble the CRISPR Ribonucleoprotein (RNP) complex, duplex RNA made of a crRNA and a trans-activating crRNA (tracrRNA) was prepared by annealing crRNA to tracrRNA. 1 µM each of crRNA and tracrRNA in Nuclease-free Duplex buffer was mixed and annealed in a heat block at 95 °C for 5 min, following cooling to RT. To form the RNP complex, 60 nM each of duplex RNA and Cas9 protein were incubated at RT for 5 min. The RNP complex was delivered into cells using RNAiMAX (13778, Thermo Fisher Scientific). To establish the cell lines from a single cell, transfected cells were diluted and plated onto 96 well plates. For single-cell cloning of 293F, a chemically defined HE150 medium (HE150-0005, GMEP) was used for culturing the cells and incubated in static conditions. Genomic DNAs were isolated from the cells using QuickExtract DNA Extraction Solution (QE09050, Lucigen), followed by PCRs to screen for the colonies carrying homozygous knockout of TM9SF3. Primers used for PCR are 5’- TGCCTGAGACTACAATCAGAGG -3’ and 5’- AACTGGAAGGTGCTACTGGC -3’. Amplified PCR products were analyzed using the MultiNa microchip electrophoresis system (MCE-202, Shimadzu) to preselect potential mutant cell populations. PCR amplicons were subsequently cloned into the pCR4 blunt TOPO vector and sequenced. Sequencing analysis identified two independent homozygous TM9SF3 knockout 293F cell lines, designated TM9SF3−/−#1 and #2, harboring 5- and 56- nucleotide deletions, respectively, in the second exon (Fig. 2f, g). Additionally, a compound heterozygous TM9SF3−/− DLD-1 cell line was identified, harboring 1- and 2- nucleotide deletions (Supplementary Fig. 3g, h).
siRNA-mediated gene knockdown
For knockdown experiments, Dharmacon SMART pool-ON-TARGETplus siRNAs (a mixture of four different siRNAs; Thermo Fisher Scientific) against human genes were used: TM9SF3 (L-010220-02); TMEM263(L-015480-02); KCNJ3(L-006248-00). ON-TARGETplus Non-targeting siRNA Pool (D-001810-10) was used as a control siRNA. RNAs (20 nM) were transfected into cells with Lipofectamine RNAi MAX. Analyses were performed 48 hours after transfection. We confirmed the efficiency of knockdown by quantitative PCR (qPCR) for TM9SF3. qPCR was performed using 20 ng of extracted total RNA by Trizol (15596026, Invitrogen) and Quantifast Probe RT-PCR kit (204256, QIAGEN). Primers used for qPCR are; 5’- GCTTCATGATGCTGGTGCTG -3’ and 5’- TTGTCCATTGCCACCGGTAA -3’ for human TM9SF3; 5’- CTGGCATTGCCGACAGGAT -3’ and 5’- GCAATGATCTTGATCTTCATTGTGC -3’ for human β-actin.
Western blotting analysis
Cultured 293F and DLD-1 cells were washed with ice-cold PBS, lysed at 4 °C in RIPA buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl2, 0.5 % NaDoc, 1 % NP-40, and 0.1 % SDS, added with a protease inhibitor cocktail (11697498001, Roche), followed by centrifugation at 17,700 x g for 15 min. The supernatants were mixed with 1x SDS sample buffer and subjected to western blot analysis. The samples were separated by SDS-polyacrylamide gel electrophoresis using Mini-PROTEAN TGX Gels (4561096, BIO-RAD), followed by transfer to PVDF membrane, and immunoblotted with the indicated antibodies. Chemiluminescence signals were detected using the ChemiDoc Imaging system (1708370, BIO-RAD) and quantified using Image Lab software (BIO-RAD).
After removing the chorion and yolk using forceps at 4 hpf, AB or maternal-zygotic tm9sf3−/− mutant embryos were directly lysed in 1x SDS sample buffer and subjected to a western blot analysis with anti-TM9SF3 antibody and anti-β-actin antibody.
Mass spectrometric lipid analysis
Cellular PIP2 was measured as described previously70. Briefly, acidic phospholipids extracted from parental and TM9SF3−/− 293F cells were derivatized by a methylation reaction with (trimethylsilyl) diazomethane (T1146, Tokyo Chemical Industry), followed by LC-MS/MS analysis. LC-MS/MS was performed using a triple quadrupole mass spectrometer QTRAP6500 (ABSciex) and a Nexera X2 HPLC system (Shimadzu) combined with a PAL HTC-xt (CTC Analytics, Switzerland) autosampler. The obtained chromatographic data were analyzed by MultiQuant 3.0.2 software (ABSciex). Concentration of each molecular species was calculated by dividing the peak area of each species by that of C37:4 PIP2 added to the cell pellets as an internal standard before lipid extraction. The relative PIP2 concentrations to phosphatidylserine in each sample are presented.
Cell surface biotinylation assay
DLD-1 cells were washed twice with ice-cold PBS and then labeled with Sulfo-NHS-SS-biotin (89881, Thermo Fisher Scientific), dissolved in ice-cold PBS, for 30 min at 4 °C with gentle agitation. The reaction was terminated by quenching solution, and the labeled cells were collected into the conical tubes. After washing twice with TBS, a lysis buffer containing a protease inhibitor cocktail was added to the cells. The lysate was transferred to 1.5 ml tubes and incubated for 30 min on ice, mixing vigorously every 5 min for 5 sec. After centrifugation at 10,000 x g for 2 min at 4 °C, the supernatant was transferred to the column, which contains avidin agarose in a collection tube. The supernatant and avidin agarose were mixed in the column for 60 min at RT. After centrifugation of the mixture at 1000 x g for 1 min, elution was used as a non-biotinylated “cytosol” protein. Then the avidin beads were washed twice with wash buffer and bound proteins were eluted by SDS-PAGE sample buffer with 50 mM DTT for 60 min at RT. After centrifugation of the column at 1000 x g for 2 min, elution was used as a biotinylated “plasma membrane” protein.
Fractional analysis of mouse tissues
Small intestine and thigh muscles from 3-month-old C57BL6J mice were homogenized in 3 volumes of solution A (20 mM Tris-HCl, pH 7.6, 0.32 M sucrose, 3 mM MgCl2, and protease inhibitor cocktail) using a motor-driven Potter-Elehjem homogenizer (HK-1, AS ONE). Muscles were cut into small pieces using scissors prior to homogenization. Homogenates were filtered through a nylon mesh, and 0.6 volume of solution B (0.25 M sucrose) was added. The mixture was then layered over an equal volume of solution A in a centrifuge tube and centrifuged at 700 × g for 10 min at 4 °C to remove the nuclear fraction. The resulting supernatant was then centrifuged at 7000 g for 10 min at 4 °C. The obtained organelle-enriched pellet was resuspended in 1 ml of ice-cold solution B, and the supernatant was kept. This supernatant was ultracentrifuged at 105,000 g (CP80WX, Himac) for 100 min at 4 °C. The resulting pellet was resuspended in 100 μl of solution B as a “microsome fraction” (MS) containing plasma membrane. This supernatant was kept as a “cytosolic fraction” (Cytosol). The resuspended mitochondria-enriched pellet was subjected to sequential centrifugation at 12,000 g for 10 min and 24,000 g for 10 min at 4 °C. The obtained pellet was resuspended in 1 ml of ice-cold solution B as “mitochondria fraction” (MT). Each fraction was mixed with SDS sample buffer and subjected to western blot analysis.
Dictyostelium cell strains and culture condition
Dictyostelium discoideum strains used in this study are AX2 (WT), phg1a- (NBRP Nenkin: S91332), and phg1b-. phg1b- mutant was generated by the CRISPR-Cas9 system using all-in-one CRISPR/Cas9 vector, pTM1285 (NBRP Nenkin: G90426)71. DNA sequencing identified that the 80th amino acid residue of the phg1b gene in the deficient strain was replaced by a stop codon. Cells were grown axenically in HL5 medium, including glucose (Formedium) in 90-mm culture dishes at 21 °C.
Image acquisition and analysis in Dictyostelium cells
Dictyostelium cells were starved in KK2 buffer (20 mM potassium phosphate buffer, pH 6.2) for 3 hours. Starved cells were washed three times with 20 mM HEPES buffer at pH 6.5 or 7.0 and transferred to 35-mm glass-bottom dishes (Iwaki) containing 20 mM HEPES buffer at pH 6.5 or 7.0, respectively. After 15 min of incubation at 21 °C to allow cells to attach to the bottom, TagRFP-PLCδ1-PH probe was added and incubated for 5 min at 21 °C. DIC and fluorescence images of Dictyostelium cells were acquired using an inverted fluorescence microscope IX83 (Olympus) equipped with a confocal unit CSU-W1 (Yokogawa), an sCMOS camera (Prime 95B, Photometrics), and an objective (UPLXAPO 40×/0.95 NA, Olympus). Confocal images were processed and analyzed with Fiji software67 and Prism 10 software (GraphPad Software).
Immunohistochemistry
Embryos were fixed in 4% paraformaldehyde (PFA) overnight at 4 °C. After fixation, embryos were dechorionated and washed in PBST (PBS with 0.1% Tween20) 3 times. Embryos were blocked in blocking solution (PBS with 5% goat serum and 0.3% Triton) for 1 h at RT. Detergent was not used for visualizing TM9SF3 on the PM (Fig. 3f and Supplementary Fig. 5j). Subsequently, embryos were incubated with primary antibody overnight at 4 °C. Protein reacting with antibody was visualized with species-matched Alexa Fluor-labeled secondary antibody (Thermo Fisher Scientific), along with Phalloidin (Invitrogen) and DAPI for counterstaining F-actin and nuclei, respectively.
Whole-mount in situ hybridization (WISH)
The sense and antisense tm9sf3 RNA probes labeled with digoxigenin (DIG) were prepared by using an RNA labeling kit (11175033910, Roche). pCR4-TOPO_Tm9sf3 was used as a template vector. WISH was performed according to the standard protocol as described previously72. Briefly, embryos were fixed in 4% PFA in PBS, dehydrated in MeOH, and gradually rehydrated in PBS containing 0.1% Tween 20 (PBS-T). Embryos were hybridized with RNA probes at 65°C overnight in hybridization buffer (5xSSC, 50% formamide, 5 mM EDTA, 0.1% Tween 20, 50 ug/ml heparin, and 1 mg/ml torula RNA). After hybridization, embryos were incubated with anti-DIG antibody conjugated with alkaline phosphate in blocking buffer at 4°C overnight. The colorimetric reaction was carried out using BM purple (11442074001, Roche). Images of the embryos were recorded with an SZX16 Stereo microscope.
Quantitative PCR for embryos
Embryos were collected at the indicated stage and dissolved in Trizol (Invitrogen) to extract total RNAs. RNAs were transcribed into cDNA by PrimeScript FAST RT reagent kit with gDNA Eraser (RR092, TAKARA), following the manufacturer’s instructions. Quantitative PCR was done with TB Green Premix Ex Taq2 FAST qPCR (RR830, TAKARA) using a Real-time PCR detection system (CFX connect, Bio-Rad). The β-actin gene was used as a reference. Primer sequences used for gsc, ntl, sox17, and cas were designed from the previous paper73. Primers used for zebrafish tm9sfs are; 5’-GACCGTATGGCTGAATCTCTTT-3’ and 5’-GCAACTGATCCACCTCTTTCT-3’ for tm9sf1, 5’-CACCATTCTCCTCTGCTACTTC-3’ and 5’-GGATGGCATAGACCAGGAAATAA-3’ for tm9sf2, 5’-GTGTGCTGCATCTGCTTATTT-3’ and 5’-GCTGGCTCCATAAACCATTTC-3’ for tm9sf3, 5’-TTACCCAGCTGTCGTCTTTG-3’ and 5’-GAGCCAGCATCGTAGTGAAA-3’ for tm9sf4.
Body axis measurement
Heterozygous mutant tm9sf3+/- male and female fish were paired for mating, and the laid eggs were raised to the indicated time points. Tricaine-treated anesthetized 7 dpf embryos and 1 mpf juvenile fish were used for measuring the body axis. Stereomicroscopic images were acquired with an SZX16 stereomicroscope equipped with a DP2-BSW camera for 7 dpf embryos from the top view and SZX7-ILST-C stereomicroscope equipped with a DP21 camera for 1 mpf fish from the lateral view. The measurement was conducted for the distance from the tip of the mouth to the tip of the caudal fin for 7 dpf and from the tip of the mouth to the root of the caudal fin for 1 mpf. After the capture of the images, the genotype was determined by PCR.
Image acquisition and quantitative analysis of gsc-positive axial mesoderm cells
Tg(gsc:EGFP-CAAX) in WT and tm9sf3−/− backgrounds were embedded in agarose under the stereomicroscope at approximately 6 hpf and imaged from 6.5 to 7 hpf using Dragonfly200 with 30 seconds intervals for 30 minutes (total 60 time points). To visualize nuclear or actin dynamics, 100 pg of mRNA encoding h2afva-tagBFP or 200 pg of mRNA encoding Lifeact-RFP was injected at one-cell stage embryos, respectively. 3D visualization and quantitative analyses were performed using Imaris 10.1 software. For the analysis of cell migratory properties, gsc-positive cells expressing EGFP-CAAX were selected using machine learning-assisted segmentation, and their nuclei were subsequently labeled using the “Mask” function. Nuclei were then selected with the “spot” function and automatically tracked, followed by manual tracking. Cells that were successfully tracked for at least 25 minutes (50 time points) were included in subsequent analysis. Displacement length and displacement change (Δlength), were calculated using Imaris and plotted on the 2D graphs with Prism 7 software. For spatial analysis, each cell’s position (X, Y, Z) at the initial time point was defined as the origin (0, 0, 0), and subsequent positional changes were plotted in 3D using Imaris. Unique color codes were assigned to each cell track to facilitate visual distinction in the representation of migratory parameters.
Quantitative analysis of immunostained ppl cells
For quantitative analysis of ppl cells, Tg(gsc:EGFP-CAAX) embryos in WT and tm9sf3−/− backgrounds were fixed at indicated time points and subjected to immunostaining. Quantitative image analysis was performed on 3D images using Imaris software (versions 9.9 and 10.1). gsc-positive cells expressing EGFP-CAAX were identified using machine learning-assisted segmentation and used to generate masks for other image channels. Segmented images of ppl were reconstructed and analyzed. The mean signal intensity within the ppl region was quantified for each embryo and used for statistical analysis. For phospho-MLC and F-actin quantification, the mean signal intensity within the ppl was normalized to the average WT value at 7 hpf and used for comparative analysis. For spatial analysis (Fig. 5e), line scan analysis was performed along the midline of the ppl from the leading edge to the rear. The phospho-myosin signal was divided by the phalloidin signal, and the resulting ratio was plotted. For phospho-histone H3 staining, the number of positive cells within the segmented ppl was counted and analyzed.
In vitro, ppl progenitor cell culture
To culture ppl progenitor cells, WT and tm9sf3−/− embryos injected with 100 pg of cyc mRNA were manually dechorionated in E3 buffer at 6 hpf. For the cell sorting experiments, ppl progenitor cells were prepared from Tg(gsc:EGFP-CAAX) embryos injected with 100 pg of cyc mRNA and 30 pg of h2afva-tagBFP mRNA, and from tm9sf3−/− embryos co-injected with 100 pg of cyc mRNA, 30 pg of h2afva-tagBFP mRNA, and 50 pg of Lifeact-RFP mRNA. Embryos from each group were mixed prior to cell preparation. Groups of 5-10 embryos were transferred to DMEM/F-12 culture medium (Gibco, 12500-062) adjusted to pH 7 or 8 using 20 mM Hepes. Embryos were mechanically dissociated by gentle pipetting, followed by centrifugation at 300 g for 2 minutes. This step was repeated twice to ensure efficient cell separation and removal of yolk granules. For in vitro assays, glass-bottom dishes (IWAKI) were coated with fibronectin. Each dish was prepared by adding 150 μl of 20 μg/ml bovine fibronectin (PMC, SFN01) and incubating overnight. Dishes were then washed twice with distilled water and coated with 50 mg/ml BSA (SIGMA, A2153) for 10 minutes prior to the experiment. Images were acquired using an inverted confocal microscope. For the quantification of cell sorting, the position of each cells was analyzed using Imaris software. EGFP and RFP signals were used to segment WT and tm9sf3−/− cells, respectively, and the position of nuclei in each cell was used for further analysis. The centroid of all cells was manually calculated using Excel software (Microsoft), and the distance of each cell from the centroid was measured and plotted. Two-dimensional graphs representing these distances with gradient color coding, as well as local cell density maps, were generated and analyzed using OriginPro software (ver 2025) (LightStone). The region surrounding the centroid is shown in the panel in Fig. 5g.
Statistics and Reproducibility
All experiments were repeated at least twice, and similar results were acquired. No statistical method was used to predetermine sample size. Data distribution was assumed to be normal, but was not formally tested. Experiments were not randomized, and investigators were not blinded during data collection or analysis. No data were excluded from the analysis.
Statistical analysis was performed using Prism 7 and 10 (GraphPad software). All data are presented as mean ± SD. Statistical significance between multiple groups was assessed using one-way ANOVA with Tukey’s test. Comparisons between two groups were analyzed using a two-tailed Student’s t-test. Data was considered statistically significant if the p-value < 0.05 (*), <0.01 (**), <0.001 (***), <0.0001 (****).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study, as presented in the main text and supplementary materials, are available. Plasmids and zebrafish lines generated in this study, except for the tm9sf3 8 bp deletion mutant (Supplementary Fig. 5f, g), are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Bretscher, M. S. Asymmetrical lipid bilayer structure for biological membranes. Nat. N. Biol. 236, 11–12 (1972).
Verkleij, A. J. et al. The asymmetric distribution of phospholipids in the human red cell membrane. A combined study using phospholipases and freeze-etch electron microscopy. Biochim Biophys. Acta 323, 178–193 (1973).
Devaux, P. F. Static and dynamic lipid asymmetry in cell membranes. Biochemistry 30, 1163–1173 (1991).
Nagata, S., Suzuki, J., Segawa, K. & Fujii, T. Exposure of phosphatidylserine on the cell surface. Cell Death Differ. 23, 952–961 (2016).
Fadok, V. A. et al. Exposure of phosphatidylserine on the surface of apoptotic lymphocytes triggers specific recognition and removal by macrophages. J. Immunol. 148, 2207–2216 (1992).
Emoto, K. & Umeda, M. An essential role for a membrane lipid in cytokinesis. Regulation of contractile ring disassembly by redistribution of phosphatidylethanolamine. J. Cell Biol. 149, 1215–1224 (2000).
Emoto, K. et al. Redistribution of phosphatidylethanolamine at the cleavage furrow of dividing cells during cytokinesis. Proc. Natl. Acad. Sci. USA 93, 12867–12872 (1996).
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010).
Caterina, M. J. et al. The capsaicin receptor: a heat-activated ion channel in the pain pathway. Nature 389, 816–824 (1997).
Nagel, G. et al. Channelrhodopsin-1: a light-gated proton channel in green algae. Science 296, 2395–2398 (2002).
Tsai, H. H. & Schmidt, W. The enigma of environmental pH sensing in plants. Nat. Plants 7, 106–115 (2021).
Quade, B. N., Parker, M. D. & Occhipinti, R. The therapeutic importance of acid-base balance. Biochem. Pharm. 183, 114278 (2021).
Hoijman, E. et al. Cooperative epithelial phagocytosis enables error correction in the early embryo. Nature 590, 618–623 (2021).
Sakuragi, T. & Nagata, S. Regulation of phospholipid distribution in the lipid bilayer by flippases and scramblases. Nat. Rev. Mol. Cell Biol. 24, 1–21 (2023).
Hammond, G. R. & Balla, T. Polyphosphoinositide binding domains: Key to inositol lipid biology. Biochim. Biophys. Acta 1851, 746–758 (2015).
Posor, Y., Jang, W. & Haucke, V. Phosphoinositides as membrane organizers. Nat. Rev. Mol. Cell Biol. 23, 797–816 (2022).
Wills, R. C. & Hammond, G. R. V. PI(4,5)P2: signaling the plasma membrane. Biochem. J. 479, 2311–2325 (2022).
Kisseleva, M. V., Wilson, M. P. & Majerus, P. W. The isolation and characterization of a cDNA encoding phospholipid-specific inositol polyphosphate 5-phosphatase. J. Biol. Chem. 275, 20110–20116 (2000).
Kanemaru, K. et al. Plasma membrane phosphatidylinositol (4,5)-bisphosphate is critical for the determination of epithelial characteristics. Nat. Commun. 13, 2347 (2022).
Varnai, P. & Balla, T. Visualization of phosphoinositides that bind pleckstrin homology domains: calcium- and agonist-induced dynamic changes and relationship to myo-[3H]inositol-labeled phosphoinositide pools. J. Cell Biol. 143, 501–510 (1998).
Koivusalo, M. et al. Amiloride inhibits macropinocytosis by lowering submembranous pH and preventing Rac1 and Cdc42 signaling. J. Cell Biol. 188, 547–563 (2010).
Lauger, P. Carrier-mediated ion transport. Science 178, 24–30 (1972).
Benz, R. & McLaughlin, S. The molecular mechanism of action of the proton ionophore FCCP (carbonylcyanide p-trifluoromethoxyphenylhydrazone). Biophys. J. 41, 381–398 (1983).
Rad, R. et al. PiggyBac transposon mutagenesis: a tool for cancer gene discovery in mice. Science 330, 1104–1107 (2010).
Friedrich, M. J. et al. Genome-wide transposon screening and quantitative insertion site sequencing for cancer gene discovery in mice. Nat. Protoc. 12, 289–309 (2017).
Uren, A. G. et al. A high-throughput splinkerette-PCR method for the isolation and sequencing of retroviral insertion sites. Nat. Protoc. 4, 789–798 (2009).
Oo, H. Z. et al. Identification of novel transmembrane proteins in scirrhous-type gastric cancer by the Escherichia coli ampicillin secretion trap (CAST) method: TM9SF3 participates in tumor invasion and serves as a prognostic factor. Pathobiology 81, 138–148 (2014).
Pruvot, B. et al. Comparative analysis of nonaspanin protein sequences and expression studies in zebrafish. Immunogenetics 62, 681–699 (2010).
Michell, R. H. Inositol derivatives: evolution and functions. Nat. Rev. Mol. Cell Biol. 9, 151–161 (2008).
Benghezal, M. et al. Synergistic control of cellular adhesion by transmembrane 9 proteins. Mol. Biol. Cell 14, 2890–2899 (2003).
El-Brolosy, M. A. et al. Genetic compensation triggered by mutant mRNA degradation. Nature 568, 193–197 (2019).
Smutny, M. et al. Friction forces position the neural anlage. Nat. Cell Biol. 19, 306–317 (2017).
Diz-Munoz, A. et al. Control of directed cell migration in vivo by membrane-to-cortex attachment. PLoS Biol. 8, e1000544 (2010).
Shewan, A. M. et al. Myosin 2 is a key Rho kinase target necessary for the local concentration of E-cadherin at cell-cell contacts. Mol. Biol. Cell 16, 4531–4542 (2005).
Krieg, M. et al. Tensile forces govern germ-layer organization in zebrafish. Nat. Cell Biol. 10, 429–436 (2008).
Casey, J. R., Grinstein, S. & Orlowski, J. Sensors and regulators of intracellular pH. Nat. Rev. Mol. Cell Biol. 11, 50–61 (2010).
Saarikangas, J., Zhao, H. & Lappalainen, P. Regulation of the actin cytoskeleton-plasma membrane interplay by phosphoinositides. Physiol. Rev. 90, 259–289 (2010).
Montero, J. A., Kilian, B., Chan, J., Bayliss, P. E. & Heisenberg, C. P. Phosphoinositide 3-kinase is required for process outgrowth and cell polarization of gastrulating mesendodermal cells. Curr. Biol. 13, 1279–1289 (2003).
Chandrasekar, I. et al. Vinculin acts as a sensor in lipid regulation of adhesion-site turnover. J. Cell Sci. 118, 1461–1472 (2005).
Saunders, R. M. et al. Role of vinculin in regulating focal adhesion turnover. Eur. J. Cell Biol. 85, 487–500 (2006).
Maitre, J. L. et al. Adhesion functions in cell sorting by mechanically coupling the cortices of adhering cells. Science 338, 253–256 (2012).
Suh, B. C. & Hille, B. PIP2 is a necessary cofactor for ion channel function: how and why? Annu. Rev. Biophys. 37, 175–195 (2008).
Yang, J. et al. PAC, an evolutionarily conserved membrane protein, is a proton-activated chloride channel. Science 364, 395–399 (2019).
Osei-Owusu, J. et al. Molecular determinants of pH sensing in the proton-activated chloride channel. Proc. Natl. Acad. Sci. USA 119, e2200727119 (2022).
Mihaljevic, L., Ruan, Z., Osei-Owusu, J., Lu, W. & Qiu, Z. Inhibition of the proton-activated chloride channel PAC by PIP(2). Elife 12, https://doi.org/10.7554/eLife.83935 (2023).
Ullrich, F. et al. Identification of TMEM206 proteins as pore of PAORAC/ASOR acid-sensitive chloride channels. Elife 8, https://doi.org/10.7554/eLife.49187 (2019).
Shen, L., Du, X., Ma, H. & Mei, S. miR-1193 Suppresses the Proliferation and Invasion of Human T-Cell Leukemia Cells Through Directly Targeting the Transmembrane 9 Superfamily 3 (TM9SF3). Oncol. Res. 25, 1643–1651 (2017).
Boedtkjer, E. & Pedersen, S. F. The Acidic Tumor Microenvironment As A Driver Of Cancer. Annu. Rev. Physiol. 82, 103–126 (2020).
Yamamoto, A., Doak, A. E. & Cheung, K. J. Orchestration of collective migration and metastasis by tumor cell clusters. Annu Rev. Pathol. 18, 231–256 (2023).
Westerfield, M. The zebrafish book. A guide for the laboratory use of zebrafish (Danio rerio). 4th ed., Univ. of Oregon Press, Eugene. (2000).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Suster, M. L., Abe, G., Schouw, A. & Kawakami, K. Transposon-mediated BAC transgenesis in zebrafish. Nat. Protoc. 6, 1998–2021 (2011).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Oikawa, T. et al. Tks5-dependent formation of circumferential podosomes/invadopodia mediates cell-cell fusion. J. Cell Biol. 197, 553–568 (2012).
Duellman, T., Burnett, J. & Yang, J. Quantitation of secreted proteins using mCherry fusion constructs and a fluorescent microplate reader. Anal. Biochem. 473, 34–40 (2015).
Idevall-Hagren, O., Dickson, E. J., Hille, B., Toomre, D. K. & De Camilli, P. Optogenetic control of phosphoinositide metabolism. Proc. Natl. Acad. Sci. USA 109, E2316–E2323 (2012).
Furutani, M., Tsujita, K., Itoh, T., Ijuin, T. & Takenawa, T. Application of phosphoinositide-binding domains for the detection and quantification of specific phosphoinositides. Anal. Biochem. 355, 8–18 (2006).
Hammond, G. R., Machner, M. P. & Balla, T. A novel probe for phosphatidylinositol 4-phosphate reveals multiple pools beyond the Golgi. J. Cell Biol. 205, 113–126 (2014).
Kondoh, G. Development of glycosylphosphatidylinositol-anchored enhanced green fluorescent protein. One-step visualization of GPI fate in global tissues and ubiquitous cell surface marking. Methods Mol. Biol. 183, 215–224 (2002).
Kawakami, K. et al. A transposon-mediated gene trap approach identifies developmentally regulated genes in zebrafish. Dev. Cell 7, 133–144 (2004).
Urasaki, A., Morvan, G. & Kawakami, K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics 174, 639–649 (2006).
Dempsey, W. P., Fraser, S. E. & Pantazis, P. PhOTO zebrafish: a transgenic resource for in vivo lineage tracing during development and regeneration. PLoS One 7, e32888 (2012).
Compagnon, J. et al. The notochord breaks bilateral symmetry by controlling cell shapes in the zebrafish laterality organ. Dev. Cell 31, 774–783 (2014).
Popgeorgiev, N. et al. The apoptotic regulator Nrz controls cytoskeletal dynamics via the regulation of Ca2+ trafficking in the zebrafish blastula. Dev. Cell 20, 663–676 (2011).
Riedl, J. et al. Lifeact: a versatile marker to visualize F-actin. Nat. Methods 5, 605–607 (2008).
Rebagliati, M. R., Toyama, R., Haffter, P. & Dawid, I. B. cyclops encodes a nodal-related factor involved in midline signaling. Proc. Natl. Acad. Sci. USA 95, 9932–9937 (1998).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Pucadyil, T. J. & Schmid, S. L. Real-time visualization of dynamin-catalyzed membrane fission and vesicle release. Cell 135, 1263–1275 (2008).
Yusa, K., Rad, R., Takeda, J. & Bradley, A. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat. Methods 6, 363–369 (2009).
Morioka, S. et al. A mass spectrometric method for in-depth profiling of phosphoinositide regioisomers and their disease-associated regulation. Nat. Commun. 13, 83 (2022).
Sekine, R., Kawata, T. & Muramoto, T. CRISPR/Cas9 mediated targeting of multiple genes in Dictyostelium. Sci. Rep. 8, 8471 (2018).
Chiba, A. et al. Osteocrin, a peptide secreted from the heart and other tissues, contributes to cranial osteogenesis and chondrogenesis in zebrafish. Development 144, 334–344 (2017).
Sako, K. et al. Optogenetic Control of Nodal Signaling Reveals a Temporal Pattern of Nodal Signaling Regulating Cell Fate Specification during Gastrulation. Cell Rep. 16, 866–877 (2016).
Acknowledgements
We thank C. Toyoshima, Y. Harada, and R. Verena for their thoughtful discussion, Y. Okamura, K. Segawa, M. Oginuma, and K. Tsujita for critical reading of the manuscript, R. Rad (pATP1-CAGGS-Puro vector), C. Toyoshima (anti-Na+, K+-ATPase antibody), C-P. Heisenberg (Tg(gsc:EGFP-CAAX)), and K. Emoto (FITC-SA-Ro) for sharing the material, M. Sone, T. Sato, K. Hiratomi, S. Toyoshima, T. Babazono, and E. Hanimura for technical assistance. RCIscript-GoldyTALEN was a gift from Daniel Carlson & Stephen Ekker (Addgene plasmid#38142); Lyn-tailed mCherry-SEpHluorin was a gift from Sergio Grinstein (Addgene plasmid #32002); iRFP-PH-PLCδ1 was a gift from Pietro De Camilli (Addgene#66841). K.S., Y.V.M., and N.M. acknowledge funding from JSPS KAKENHI (21K05983 and 24K09294 to K.S. and 19H01022 to N.M.), JST PRESTO Grant (JPMJPR204B to Y.V.M.), JST grant (JPMJPF2018 and JPMJMS2024-3 to N.M.), and The Takeda Science Foundation (K.S. and N.M.) This work was supported by AMED-CREST (JP25gm1710007h0003), Nanken-Kyoten, the Multilayered Stress Diseases program, and the Initiative for High-Depth Omics, IST; by COI-NEXT Support Unit for Imaging Science and Evident (FV3000) at Kento; by the Joint Usage/Research Center for Proteo-Interactome (PRiME), the Proteo-Science Center, Ehime University; by JSPS KAKENHI Grant Number JP22H04926, Grant-in-Aid for Transformative Research Areas ― Platforms for Advanced Technologies and Research Resources “Advanced Bioimaging Support”.
Author information
Authors and Affiliations
Contributions
K.S. conceptualized the project, performed experiments and analyzed the data. Y.V.M. contributed to Dictyostelium cell experiments. S.M., J.H., J.S. and T.S. contributed to lipid analysis. H.N. and M.F. contributed to zebrafish experiments. Y.N. and Y.S. contributed to protein purification. K.K. and N.M. supervised the project. K.S., Y.V.M., and N.M. acquired funding. K.S. wrote the original draft with inputs from all authors, and K.S. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks De-Li Shi, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sako, K., Morimoto, Y.V., Morioka, S. et al. Cells adapt to extracellular acidic pH through TM9SF3-mediated PI(4,5)P2 flop. Nat Commun 16, 8852 (2025). https://doi.org/10.1038/s41467-025-63524-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63524-w