Abstract
The creation of designer enzymes by incorporating genetically encoded non-canonical amino acids (ncAAs) could significantly expand the catalytic repertoire of the enzyme universe for abiological transformations. However, due to the limited availability of ncAAs with potential catalytic functional groups, progress in this field remains relatively slow. Herein, we present an efficient approach for enzyme design with organocatalytic ncAAs by integrating the biosynthesis and genetic incorporation of ncAAs harboring abiological catalytic residues into the protein scaffold. Based on this ncAAs in situ biosynthesis and incorporation system, our designer enzyme with an unnatural mercapto-aniline residue is created efficiently by feeding thiols to E. coli and shows significant catalytic activity for an enantioselective Friedel-Crafts alkylation reaction with excellent enantioselectivity and reactivity after directed evolution. This study provides a universal strategy for designing artificial enzymes with xenobiotic catalytic moieties with diverse biosynthesized ncAAs, thereby expanding the toolbox of biocatalysts for abiological transformations.
Similar content being viewed by others
Introduction
The potential of utilizing the catalytic abilities of enzyme catalysis for chemical synthesis has driven efforts to develop tailor-made biocatalysts capable of catalyzing non-biological transformations1,2,3,4,5,6. The introduction of abiological catalytic moiety into protein cavities is one of the most direct strategies to design enzymes with functions7,8,9,10,11,12. Genetic code expansion has emerged as a powerful technology for modifying protein structure and function13,14,15,16. The ability to encode non-canonical functionalities presents opportunities to generate a protein catalysts with non-natural catalytic functions17,18,19,20,21,22,23,24,25. Early examples include the use of genetically encoded para-azidophenylalanine as a precursor to a reactive aniline group for aldehyde functionalization26,27,28, the introduction of an unnatural reactive Me-histidine site for ester hydrolysis29,30, and the installation of a genetically encoded photosensitizer for photoenzymatic [2 + 2] cycloadditions31,32. Recently, a genetically encoded boronic-acid-containing designer enzyme for kinetic resolution of hydroxyketone was developed by the Roelfes group33 (Fig. 1a). Notably, to date, more than 300 non-canonical amino acids (ncAAs) have been successfully incorporated through stop codon suppression strategies, in contrast, utilizing ncAAs as a catalytic residues for artificial enzyme design has remained largely limited34,35,36.
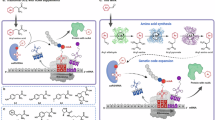
a Previous study on direct or indirect artificial design by the incorporation of exogenous supplemented non-canonical amino acids. b Our strategy for artificial enzyme design by the combination of endogenous S-functionalized cysteine biosynthesis and genetic incorporation. c The creation of designer enzyme SFC_pAPhC by feeding E. coli with 4-mercaptoaniline and catalyzed asymmetric Friedel-Crafts alkylation reaction.
The major barrier to the use of ncAAs in artificial enzyme design is that, in most cases, ncAAs must be provided exogenously during protein expression, which limits their application, especially when they are expensive, difficult to synthesize or have poor cell membrane penetration37,38. Moreover, the absence of effective orthogonal translation systems (OTS) that can incorporate ncAAs containing catalytic residues presents significant evolutionary hurdles39,40,41. In comparison to the extensive array of efficient chemical catalysts, only a limited number of ncAAs exhibit potential catalytic activities owing to their restricted structural diversity. Typically, the incorporated catalytic ncAAs exhibit low catalytic efficiency due to challenges in catalytic reactivity under aqueous phase, substrate binding, and steric hindrance, which may lower reaction rates. Therefore, the introduction of diverse synthetic non-natural amino acids with catalytic functions into proteins will be quite needed for enzyme design. In this regard, the development of biosynthetic pathways for ncAAs that can be integrated into the metabolism of host cells, coupled with a compatible orthogonal translation system, represents a promising approach to enhance the potential of using ncAA-containing artificial enzymes for new-to-nature biocatalysis42,43. This is particularly attractive for the directed evolution of designer enzymes and integration into existing biosynthetic pathways to create hybrid metabolic pathways44,45,46,47.
To explore the potential of unnatural amino acids as catalytic residues in designer enzymes, an expanded genetic code, combined with metabolic engineering for catalytic ncAA biosynthesis, would undoubtedly further diversify the chemical arsenal for creating enzymes with functions. However, only a small number of biosynthetic pathways for ncAAs have been reported to date, greatly restricting our ability to design enzymes with ncAA building blocks48,49,50,51. Based on this, we previously developed a biosynthesis system for S-arylcysteine, which has been demonstrated to produce ncAAs with a diverse array of structures from aromatic thiols in cells, some of these unnatural S-arylcysteines can be genetically encoded into GFP using the engineered Methanococcus jannaschii tRNATyr/TyrRS pair52. As such, we aim to explore the direct construction of a designer enzyme with in situ biosynthesized S-functionalized-cysteine in cells. As a proof of concept, the biosynthesis and genetic incorporation of S-(4-aminophenyl)-L-cysteine (pAPhC) are selected as a starting point, as it could act as a nucleophilic catalyst for activating aldehydes and exhibit similar catalytic properties to those of the previously reported artificial enzyme harboring an unnatural pAF residue26. Considering the impact of introducing a sulfur atom at the para position of aniline on the electron density and chain length, we hypothesize that the catalytic properties of unnatural aniline residue could be fine-tuned for enhanced reactivity and selectivity by introducing pAPhC into an appropriate protein (Fig. 1b).
Herein we report a designer enzyme construction system for the creation of S-Functionalized Cysteine dependent enzyme (SFC from here on) through the combination of ncAA in situ biosynthesis with genetic code expansion pathway in cells. Based on this system, a designer enzyme, SFC_pAPhC, with an unnatural amino residue is produced efficiently and shows significant catalytic activity for an enantioselective Friedel-Crafts alkylation reaction via the iminium pathway. Excellent enantioselectivity (up to 95% e.e.) and yields (up to 98%) are obtained after three rounds of directed evolution (Fig. 1c). Notably, our designed enzyme, SFC_V15pAPhC, exhibits opposite enantioselectivity for this Friedel-Crafts alkylation reaction comparing to the previously reported artificial enzyme, LmrR_pAF. Computational studies identify the key factors causing the distinct stereoselectivity of SFC_V15pAPhC and LmrR_V15pAF. This work introduces an affordable and efficient strategy for the design of artificial enzymes with diverse biosynthesized potential catalytic ncAAs, which not only significantly expands the toolbox of biocatalysts for reactions but also represents a promising strategy for creating hybrid metabolic pathways involving artificial enzymes.
Results
System construction for designer enzyme with in situ biosynthesized ncAAs
Our initial efforts involved the creation of designer enzyme with in situ biosynthesized ncAA by the integration of the biosynthesis system of S-arylcysteine, the compatible orthogonal translation system and the expression system of protein of interest. The biosynthetic method for genetically encoding S-arylcysteines into GFP protein by feeding cells with aromatic thiols was previously developed by our collaborator Liu52. In this regard, we sought to construct artificial enzyme based on the dimeric Lactococcal multidrug resistance regulatory (LmrR) protein, which is a small homodimeric protein that has been demonstrated to be a versatile scaffold for the design of artificial enzymes owing to its structure and evolvability28,53,54,55. The hybrid system was assembled by integrating three plasmids, which respectively encode engineered CysM (PBK_CysM-NtSat4), OTS (pUltra_PhSeRS), and LmrR variant (pET17b_LmrR_V15TAG) into Escherichia coli (Fig. 2a). After the successful fusion of the three plasmids into E. coli, the integrated ncAA biosysnthesis and genetic code expansion system was first tested by adding 1 mM PhSNa into the cell culture, after protein expression and purification, the corresponding protein LmrR containing a PhC residue was produced efficiently and confirmed by high-resolution MS (Fig. 2b, c). Based on the successfully constructed system, we sought to create a type of designer enzyme by introduce in situ biosynthesized S-(4-aminophenyl)-L-cysteine (pAPhC) into LmrR pocket, as the mercaptoaniline could act as a nucleophilic catalytic site. Position 15 was proved to be a superior site for catalytic ncAA incorporation owing to its surrounding environment in the hydrophobic pore of LmrR according to previous reports27, then was chosen to generate the designer enzyme SFC_V15pAPhC by adding 4-mercaptoaniline as precursor in the cell culture. However, the initial test shows low expression level of SFC_V15pAPhC, which is much lower than SFC_V15PhC and GFP_pAPhC. Further optimization of the cell growth conditions, such as medium, temperature, and precursor concentration, gave rise to an increased SFC_V15pAPhC expression level after affinity chromatography in 14 mg protein per liter culture (Fig. 2b).
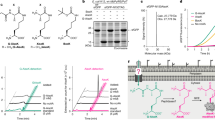
a Overview of the expression system for the production of SFC_V15pAPhC by the integration of three plasmids, PBK_CysM-NtSat4, pUltra_PhSeRS, and pET22b_LmrR_V15TAG with the adding of 4-Mercaptoaniline. b Outcome of the system optimization for the expression of SFC_V15pAPhC. c HRMS determination of LmrR_wt, SFC_V15PhC and designer enzyme SFC_V15pAPhC. d Crystal structure of the SFC_V15pAPhC homodimer (PDB: 8YRF). e Selected designer enzymes with mAPhC, oAPhC, and PyC residues were created in the system by feeding corresponding thiols into the engineered E. coli. All experiments are presented as mean ± standard deviation from three independent replicates. Source data are provided as a Source Data file.
To illustrate the advantages of in situ biosynthesis of ncAAs for the design of enzymes, the direct supplement of unnatural pAPhC, which was biosynthesized by engineered CysM, afforded almost half the protein yield compared to in situ biosynthesized pAPhC. This might be due to the low cell penetration efficiency of exogenous added pAPhC, resulting in a low intracellular concentration, which thereby limits the incorporation efficiency. High resolution mass spectrometry confirmed the successful incorporation of the unnatural pAPhC residue (Fig. 2c). The crystal structure of the designer enzyme SFC_V15pAPhC (PDB: 8YRF) provides more precise evidence for the success of our artificial enzyme design and offers structural support for subsequent study on the structure-activity relationship of the artificial enzymes (Fig. 2d). To verify the compatibility of this system and the ability to create diverse designer enzymes with distinguishing unnatural residues, several artificial enzymes with unnatural mAPhC, oAPhC and PyC were efficiently constructed by feeding corresponding thiols into the engineered E. coli (Figs. 2e and S5). Collectively, these results illustrate that our established system for in situ biosynthesis and incorporation of ncAAs constitutes an efficient and convenient approach for the construction of artificial enzymes, and exhibits significant potential for establishing a diverse range of artificial enzyme libraries.
Designer enzyme catalysis
Having established the efficient system for pAPhC biosynthesis and adjacent incorporation into position 15 of LmrR, we selected three additional positions (19, 89, and 92), which are demonstrated to be superior sites for ncAAs incorporation26, and situated in the hydrophobic cavity of the protein dimer interface for pAPhC incorporation (Fig. 3a). To explore the catalytic potential of the designed unnatural mercapto-aniline residue in these artificial enzymes, we choose to investigate the enantioselective Friedel-Crafts alkylation reaction of indole to α,β unsaturated aldehyde by iminium catalysis56. The initial test was performed with 1 mM of 2-methylindole 1a and 5 mM trans-2-pentenal 2a in the presence of SFC_V15pAPhC. The desired addition product 3a was obtained with desirable activity in 74% e.e. and 39% yield, and with R configuration according to the reported study57. Interestingly, we found that SFC_V15pAPhC showed the opposite stereoselectivity compared to LmrR_V15pAF with (S)−3a in this reaction. These results suggest the introduction of sulfur atoms led to discernible variations in the reactivity and stereoselectivity exhibited by SFC_V15pAPhC and LmrR_V15pAF (Fig. 3b). No catalytic activity was observed when using wildtype LmrR or the precursor for pAPhC biosynthesis, 4-mercaptoaniline in the reaction. Notably, the free amino acid pAPhC or LmrR in which the pAPhC residue was substituted with PhC (without amino catalytic group), showed limited catalytic activity and no stereoselectivity, demonstrating a strong synergistic combination of the pAPhC side chain in the LmrR scaffold for catalysis. Besides, LmrR with pAPhC incorporated at position 19, 89, and 92 catalyzed the reaction with different activity and enantioselectivity, indicating a distinct chemical environment to promote catalysis. In addition to SFC_pAPhC, we investigated a similar variant SFC_V15mAPhC which the amino group is on the meta position of sulfur, generated form the same three plasmids system with the precursor 3-mercaptoaniline, also showed considerable activity (Fig. 3c). These results suggest our designed pAPhC site of the artificial enzyme act as the catalytic center, to form an electrophilic iminium intermediate, which then was attacked by nucleophilic indole to give the final product.
a Overview of the homo-dimeric structure of SFC_V15pAPhC (PDB: 8YRF) and enhanced view of the hydrophobic pocket showing selected positions for pAPhC catalytic residue incorporation. b Asymmetric Friedel-Crafts alkylation of enal 1a and indole 2a to form alkylation product (R)−3a by SFC_V15pAPhC and (S)−3a by LmrR_V15pAF. c Reaction yields (blue) and e.e. (yellow) values of product 3a with different SFC_pAPhC variants as well as negative controls. Reaction conditions: Enzyme varaints (5.0 mol%), 5 mM 1a and 1 mM 2a in 50 mM NaH2PO4, 150 mM NaCl (pH 6.5) buffer with DMF (4.0% v/v). Reactions conducted for 10 h at 4 °C and analyzed by normal-phase chiral column with e.e. and yield, unless noted otherwise. Yields are determined by HPLC analysis using a calibration curve of pure product 3a. Error bars represent ± standard deviation of the mean over 3 independent replicates. Absolute configuration of 3a assigned by comparison of order of elution on chiral normal-phase HPLC with enantioenriched reference compound and the literature57,67. Source data are provided as a Source Data file.
Designer enzyme directed evolution
With the good starting enantioselectivity and activity achieved by SFC_V15pAPhC (SFC1.0), we sought to identify important residues near the catalytic site by alanine scanning to further improve the reaction enantioselectivity and efficiency. Guided by the crystal structure of SFC1.0 (PDB: 8YRF), which is obtained by us, we selected 10 positions which were considered important for catalysis by LmrR-based artificial enzymes based on previous studies, and mutated these positions to alanine. During the alanine scanning with those purified mutants, eight of ten variants showed positive or negative effects on the enantioselectivity and yield obtained in catalysis, indicating potential influence for catalytic activity. In order to rapidly identify beneficial mutations, we created site-specific codon libraries which encodes 12 amino acids (NDT codon, N = A/T/C/G, D = A/G/T, T = T) with a good representation of the functional diversity of all 20 amino acids at the eight positions (L18, N19, K22, N88, M89, A92, F93, S95) based on SFC1.0 (Fig. 4a, b). After second round of screening, we found an improved mutants SFC_V15pAPhC_F93I, which afford higher yield and enantioselectivity in production of 3a compared to SFC1.0. In a similar manner, we continued with a third round of library screening based on SFC_V15pAPhC_F93I (hereafter, SFC2.0), selecting three positions (A92, M8, K22) for NDT codon construction. This third round of screening identified three further improved mutations, as A92S (SFC3.1), A92Y(SFC3.2), and M8R (SFC3.3). Then in the fourth round of evolution, we created three site-specific NDT libraries based on these three evolved mutants, as SFC3.1_M8NDT, SFC3.2_M8NDT, and SFC3.3_A92NDT for further screening. Gratifyingly, further evolved mutant SFC_V15pAPhC_F93I_M8R_A92D (SFC4.0) was identified, displayed both improved stereoselectivity and reactivity in comparison with preceding mutants. Further investigations on reaction parameters, including buffer pH, temperature, substrates ratio and enzyme loadings, resulted in 95% e.e., and 98% yield for the generation of 3a. The enzyme loading could be lowered to 1.0 mol% and 2.0 mol% with retentive excellent enantioselectivity and activity (Fig. 4c, d).
a Asymmetric Friedel-Crafts alkylation of enal 1a and indole 2a to form alkylation product (R)−3a by SFC_V15pAPhC mutants. b Enhanced view of the hydrophobic pocket showing selected positions for enzyme directed evolution. c The iterative-site-specific mutagenesis strategy for the direct evolution of SFC_V15pAPhC. d Reaction outcomes of e.e. (yellow) and yield (blue) afforded by different evolved SFC_V15pAPhC variants. Reaction conditions: Enzyme varaints (5.0 mol% dimer concentration), 5 mM 1a and 1 mM 2a in 50 mM NaH2PO4, 150 mM NaCl (pH 6.5) buffer with DMF (4.0% v/v). Reactions conducted for 10 h at 4 °C and analyzed by normal-phase chiral column with e.e. and yield, unless noted otherwise. Yields are determined by HPLC analysis using a calibration curve of pure product 3a. Error bars represent ± standard deviation of the mean over 3 independent replicates. Source data are provided as a Source Data file.
Mechanistic experiment and kinetic characterization
In our proposed catalytic cycle, unnatural pAPhC residue accelerate the Friedel-Crafts alkylation reactions through the formation of an iminium ion upon condensation with 2-pentenal 2a. In order to verify this catalytic mechanism, we conducted a mechanistic experiment to trap this intermediate by incubating SFC4.0 (Fig. 5a) with 2-pentenal. When subjecting these samples to high-resolution mass spectrometry, SFC4.0 displayed a dominant peak corresponding to the single modified iminium ion intermediate (Figs. S12 and S13). This result suggests that the iminium ion intermediate between pAPhC and 2-pentenal may be involved in catalysis process (Fig. 5b). To evaluate the catalytic performance of the evolved mutant SFC4.0, the kinetics of the reaction was measured. The initial rates were fitted to the Michaelis-Menten equation, giving an apparent catalytic efficiency (kcat/KM(2-pentenal)) of 0.64 ± 0.22 M−1 s−1 for SFC4.0, and (kcat/KM(2-pentenal)) of 0.12 ± 0.04 M−1 s−1 for SFC1.0. The catalytic efficiency results showed 5.5-fold-improvement by our engineered variant (Fig. 5c). Directed evolution of this enzyme for the Friedel−Crafts reaction gave an apparent increase in catalytic efficiency.
a Overview of the homodimeric structure of SFC4.0 (PDB: 9IIJ) and enhanced view of the hydrophobic pocket showing the evolved mutations. b Capture of the iminium ion intermediate by the condensation of SFC4.0 with enal 1a. c Kinetic characterization of SFC1.0 and evolved mutant SFC4.0 for the reaction. Data are presented as mean values ± SD as appropriate. Error bars represent ± standard deviation of the mean over three independent replicates. d Substrate scope of the enantioselective Friedel−Crafts alkylation reaction with SFC4.0 and SFC1.0. The absolute configuration of 3f assigned according to the reported compound. e Scale-up reaction and cascade with transaminase to generate indolethylamine. Source data are provided as a Source Data file.
Substrate scope investigations
With the optimal reaction conditions and evolved SFC4.0 mutant in hand, we turned to investigate the substrate scope of the enantioselective Friedel−Crafts alkylation reaction by variation of the substituted enals and indoles with SFC4.0 and SFC1.0 respectively (Fig. 5d). A series of substituted enals underwent stereoselective alkylation smoothly, with different length alkyl chain (3a–3d) and phenyl (3e) being well tolerated. Alkylation products were obtained with up to 95% e.e. and yields were good to excellent (up to 98%), highlighting the compatibility of artificial enzymes with enals. Notably, in the case of cinnamaldehyde, SFC1.0 showed much better stereoselectivity and slightly higher yield than evolved mutant SFC4.0, indicating distinct mutants exhibit specific selectivity towards various substrates. Variations in the substitutions of indoles influence the reaction outcomes, especially for reaction yield (3f–3m). Indole substrates without the 2-methyl group (3f–3i) generally exhibit good enantioselectivity but suffered from low reactivity, mainly owing to the decreasing nucleophilicity of the indole without 2-methyl. Notably, product 3 f was obtained with an enantiomeric excess (e.e.) of 72%, and its absolute configuration was determined to be R. This assignment aligns with the reported compound57 but exhibits the opposite configuration compared to the enantioselectivity previously reported by the artificial enzyme LmrR_V15pAF. Then, a variety of substituted 2-methyl indoles bearing either electron-donating or electron-withdrawing groups on the indole ring and N-protection were well tolerated (3j–3m), with good to excellent stereoselectivities and good yields. Furthermore, the reaction can be scaled up with equal efficiency and enantioselectivity (3a), even to the 1 mmol scale as evidenced by the synthesis of (R)−3a. A biocatalytic cascade was achieved by the combination of artificial enzyme SFC4.0 and natural transaminase for the production of unnatural indolethylamine 4 with excellent enantioselectivity and moderate conversion, which emphasize the potential application value of artificial enzymes in the biosynthesis of non-natural products58,59,60 (Fig. 5e).
Computational study on enantioselectivity
Given that our designed enzyme SFC_V15pAPhC exhibits opposite enantioselectivity in the Friedel-Crafts alkylation reaction compared to the previously reported artificial enzyme LmrR_pAF, we aim to identify the factors controlling the stereoselectivity of both SFC_V15pAPhC and LmrR_V15pAF using molecular dynamics (MD) simulations. As the pathways producing different enantiomers of the product diverge at the iminium intermediate, which is formed by the aldimine condensation between the non-canonical residue and trans-2-hexenal (Fig. S14), the simulations focused on this intermediate. Since the (E)-isomer is energetically more favorable than the (Z)-isomer (Fig. S18), only the (E)-configuration was considered for the iminium intermediate in the simulations. Therefore, the relative positioning of the methyl indole to the iminium group determines whether the Re- or Si-face of the C=N double bond will be attacked, ultimately leading to the formation of different products.
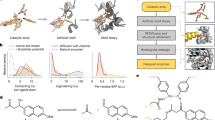
For LmrR_V15pAF, MD simulations without the binding of methyl indole reveal two representative conformations. One conformation is characterized by the hydrogen bond interactions between the iminium group and D100, while the other is defined by the π–π interaction between the C=C double bond and F93. Based on these structures, the methyl indole was then docked into the binding pocket and the resulting complexes were subjected to further MD simulations. Interestingly, these simulations converged to a very similar binding mode of the methyl indole (Fig. 6a, b), in which the iminium group forms a hydrogen bond with D100, and the 2-methyl indole is positioned on its Re-face. This finally leads to the formation of the (S)-product. Analysis of the structures shows that the preferred binding location of 2-methyl indole can be attributed to the beneficial interactions with the residues bearing aromatic rings in the Re-face, which are absent in the Si-face.
The simulations start from the initial structures in which the methyl indole was docked into (a) the Re-face and (b) the Si-face of the iminium group in LmrR_V15pAF, and (c) the Re-face and (d) the Si-face of the iminium group in SFC_V15pAPhC. Source data are provided as a Source Data file.
In the case of SFC_V15pAPhC, MD simulations without 2-methyl indole binding reveal a stable hydrogen bond interaction between the iminium group and the N88 residue. Further MD simulations with 2-methyl indole binding to different faces of the iminium group show that the 2-methyl indole can be stably accommodated in the Si-face but not in the Re-face (Fig. 6c, d). When bound to the Re-face, methyl indole quickly dissociates from the reaction site during the simulation. The reaction is thus predicted to be R-selective, consistent with the experimental outcome. For the evolved mutant SFC4.0, the simulations indicate that the iminium group forms a stable hydrogen bond with the newly-introduced D92 side chain, which enlarges the binding pocket in the Si-face (Fig. S21). This change in the hydrogen bond pattern accounts for the increased enantioselectivity for the mutant compared to the wide-type.
According to the MD simulations, the residues at positions 88 and 92 play crucial roles in governing the activity and enantioselectivity of SFC. The catalytic performance of certain mutants during the directed evolution process provided experimental validation for this conclusion. In the first round of evolution, a mutation library targeting position N88 was constructed, and mutations at this site to other amino acids consistently resulted in reduced activity. In the third round of evolution, mutations at position 92 to leucine (L), serine (S), and histidine (H) exhibited lower reactivity and enantioselectivity compared to the A92D mutant. As for LmrR_V15pAF, the crucial role of D100 drawn from MD simulations is consistent with the previous experimental study56, in which the D100A mutation significantly reduced the enantioselectivity of LmrR_V15pAF.
Finally, the binding free energies of the indole substrate to the enzymes with the iminium intermediate were calculated, with values of −13.5, −16.5, and −10.3 kcal/mol for LmrR_V15pAF, SFC1.0, and SFC4.0, respectively. These negative values indicate that indole readily binds to the active site with high affinity in all three enzymes. Interestingly, the observed trend in binding affinities is opposite to the experimentally measured yield discussed above, suggesting that reaction outcomes are influenced by factors beyond binding affinity alone.
Discussion
In summary, we propose a concept of integration of ncAAs biosynthesis and incorporation firmly into the realm of enzyme design and established an efficient strategy for the construction of artificial enzymes with unnatural catalytic residues through the creation of a cell factory that integrates the biosynthesis and genetic incorporation of organocatalytic ncAAs into the protein of interest. Through this approach, we developed an artificial enzyme harboring an unnatural catalytic pAPhC residue and demonstrated its potential as an efficient catalyst for enantioselective Friedel-Crafts alkylation reactions. The biocatalytic enantioselective Friedel-Crafts alkylation of enals and indoles was achieved using the designer enzyme SFC_pAPhC. After rounds of directed evolution based on an optimized platform, diverse chiral indole alcohol products were produced with satisfactory yield and enantioselectivity (up to 95% e.e., and up to 98% yield). Computational studies revealed that the (R)-enantiopreference of SFC_V15pAPhC can be attributed to the favorable binding of the substrate to the Si face of the key iminium intermediate, ultimately leading to the formation of (R) enantiomer. Our study not only demonstrates a artificial enzyme for enantioselective Friedel-Crafts alkylation but also provides a universal strategy to create diverse, evolvable artificial enzymes with xenobiotic catalytic functions, enabling the catalysis of reactions through in situ ncAAs biosynthesis and incorporation.
Methods
General procedure for artificial enzyme catalyzed reactions
Reactions were set up with purified enzymes and reactants in 300 μL total volume in a 2 mL microcentrifuge tube. Stock solutions of protein in pH 6.5 PBS buffer (50 mM NaCl, 150 mM NaH2PO4) to give the specified final concentration. Stock solutions of indoles (25 mM in DMF, 12 μL added, final concentration 1 mM) and enals (125 mM, 12 μL added to give final concentrations of 5 mM) substrates were added. The microcentrifuge tubes were then mixed by continuous inversion in a cold room at 4 °C for the specified reaction time. After the reaction time had elapsed, the solution of 2-phenylquinoline as internal standard (12 μL, 5 mM in DMF) were added. The reaction products and internal standard were then extracted by vortex mixing with EtOAc (1 mL) and the organic extract was dried over Na2SO4, filtered and evaporated to dryness. The residue thus obtained was redissolved by vortex mixing with HPLC grade solvent (heptane: isopropanol 4:1, 150 μL) and analysed by normal-phase chiral HPLC to determine yield and enantioselectivity.
Computational method
The crystal structures of LmrR_V15pAF (PDB ID: 6I8N), SFC_V15pAPhC (PDB ID: 8YRF), and SFC_V15pAPhC_IRD (PDB ID: 9IIJ) were used as starting structures for the molecular dynamics (MD) simulations. These structures were pretreated by removing water and ligand molecules in PyMOL, adding missing residues using MODELLER 10.461. The pKa values of the ionizable residues in the system were calculated using PROPKA362. The protonation states of these residues were determined by comparing the predicted pKa and the experimental pH values, which were further confirmed through careful visual inspection of the hydrogen-bonding networks. All missing hydrogen atoms were added by the LEaP program of AMBER 20 package63. The non-canonical amino acid in three enzymes was replaced by the iminium intermediate generated by the aldimine condensation with trans-2-hexenal. MD simulations were performed using the AMBER 20 software. The standard AMBER ff14SB force field64 was used for the simulation of the protein. For the iminium intermediate and 2-methylindole, force field parameters were generated using the general AMBER force field (GAFF2)65. The binding free energies of the indole substrate were obtained from the molecular mechanics/Poisson-Boltzmann surface area (MM-PBSA) calculations66. Additional details and parameters for the MD simulations are provided in the Supporting Information.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Authors declare that all data supporting the findings of this study are available within the paper and its supplementary information files. The PDB structures used in this work include 8YRF and 9IIJ. Source data are provided with this paper.
References
Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. 3, 203–213 (2020).
Lovelock, S. L. et al. The road to fully programmable protein catalysis. Nature 606, 49–58 (2022).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2017).
Huang, P. S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Renata, H., Wang, J. Z. & Arnold, F. H. Expanding the enzyme universe: accessing non-natural reactions by mechanism-guided directed evolution. Angew. Chem. Int. Ed. 54, 3351–3367 (2015).
Leveson-Gower, R. B., Mayer, C. & Roelfes, G. The importance of catalytic promiscuity for enzyme design and evolution. Nat. Rev. Chem. 3, 687–705 (2019).
Okeley, N. M. & Donk, W. A. V. D. Novel cofactors via post-translational modifications of enzyme active sites. Chem. Biol. 7, 159–171 (2000).
Appel, M. J. & Bertozzi, C. R. Formylglycine, a post-translationally generated residue with unique catalytic capabilities and biotechnology applications. ACS Chem. Biol. 10, 72–84 (2015).
Lewis, J. C. Artificial metalloenzymes and metallopeptide catalysts for organic synthesis. Cheminform 45, 2954–2975 (2013).
Bos, J. & Roelfes, G. Artificial metalloenzymes for enantioselective catalysis. Curr. Opin. Chem. Biol. 19, 135–143 (2014).
Zhu, Z., Hu, Q., Fu, Y., Tong, Y. & Zhou, Z. Design and evolution of an enzyme for the asymmetric Michael addition of cyclic ketones to nitroolefins by enamine catalysis. Angew. Chem. Int. Ed. 63, e202404312 (2024).
Guo, J. et al. Chemogenetic evolution of diversified photoenzymes for enantioselective [2+2] cycloadditions in whole cells. J. Am. Chem. Soc. 146, 19030–19041 (2024).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Young, D. D. & Schultz, P. G. Playing with the molecules of life. ACS Chem. Biol. 13, 854–870 (2018).
Brouwer, B. et al. Noncanonical amino acids: bringing new-to-nature functionalities to biocatalysis. Chem. Rev. 124, 10877–10923 (2024).
Agostini, F. et al. Biocatalysis with unnatural amino acids: enzymology meets xenobiology. Angew. Chem. Int. Ed. 56, 9680–9703 (2017).
Drienovská, I. & Roelfes, G. Expanding the enzyme universe with genetically encoded unnatural amino acids. Nat. Catal. 3, 193–202 (2020).
Neumann-Staubitz, P. & Neumann, H. The use of unnatural amino acids to study and engineer protein function. Curr. Opin. Struct. Biol. 38, 119–128 (2016).
Drienovská, I., Rioz-Martínez, A., Draksharapu, A. & Roelfes, G. Novel artificial metalloenzymes by in vivo incorporation of metal-binding unnatural amino acids. Chem. Sci. 6, 770–776 (2015).
Ugwumba, I. N. et al. Improving a natural enzyme activity through incorporation of unnatural amino acids. J. Am. Chem. Soc. 133, 326–333 (2011).
Hu, C., Chan, S. I., Sawyer, E. B., Yu, Y. & Wang, J. Metalloprotein design using genetic code expansion. Chem. Soc. Rev. 43, 6498–6510 (2014).
Xiao, H. et al. Exploring the potential impact of an expanded genetic code on protein function. Proc. Natl. Acad. Sci. USA 112, 6961–6966 (2015).
Yu, M. Z. et al. An artificial enzyme for asymmetric nitrocyclopropanation of α, β-unsaturated aldehydes—design and evolution. Angew. Chem. Int. Ed. 63, e202401635 (2024).
Williams, T. L. et al. Secondary amine catalysis in enzyme design: broadening protein template diversity through genetic code expansion. Angew. Chem. Int. Ed. 63, e202403098 (2024).
Drienovska, I., Mayer, C., Dulson, C. & Roelfes, G. A designer enzyme for hydrazone and oxime formation featuring an unnatural catalytic aniline residue. Nat. Chem. 10, 946–952 (2018).
Mayer, C., Dulson, C., Reddem, E., Thunnissen, A.-M. W. H. & Roelfes, G. Directed evolution of a designer enzyme featuring an unnatural catalytic amino acid. Angew. Chem. Int. Ed. 58, 2083–2087 (2019).
Zhou, Z. & Roelfes, G. Synergistic catalysis in an artificial enzyme by simultaneous action of two abiological catalytic sites. Nat. Catal. 3, 289–294 (2020).
Burke, A. J. et al. Design and evolution of an enzyme with a non-canonical organocatalytic mechanism. Nature 570, 219–223 (2019).
Hutton, A. E. et al. A non-canonical nucleophile unlocks a new mechanistic pathway in a designed enzyme. Nat. Commun. 15, 1956 (2024).
Trimble, J. S. et al. A designed photoenzyme for enantioselective [2+ 2] cycloadditions. Nature 611, 709–714 (2022).
Sun, N. et al. Enantioselective [2+2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022).
Longwitz, L., Leveson-Gower, R. B., Rozeboom, H. J., Thunnissen, A.-M. W. & Roelfes, G. Boron catalysis in a designer enzyme. Nature 629, 824–829 (2024).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Li, J. C., Liu, T., Wang, Y., Mehta, A. P. & Schultz, P. G. Enhancing protein stability with genetically encoded noncanonical amino acids. J. Am. Chem. Soc. 140, 15997–16000 (2018).
Yu, Y., Hu, C., Xia, L. & Wang, J. Artificial metalloenzyme design with unnatural amino acids and non-native cofactors. ACS Catal. 8, 1851–1863 (2018).
Wang, Y.-S., Fang, X., Wallace, A. L., Wu, B. & Liu, W. R. A rationally designed pyrrolysyl-tRNA synthetase mutant with a broad substrate spectrum. J. Am. Chem. Soc. 134, 2950–2953 (2012).
Brustad, E., Bushey, M. L., Brock, A., Chittuluru, J. & Schultz, P. G. A promiscuous aminoacyl-tRNA synthetase that incorporates cysteine, methionine, and alanine homologs into proteins. Bioorg. Med. Chem. Lett. 18, 6004–6006 (2008).
Young, D. D. et al. An evolved aminoacyl-tRNA synthetase with atypical polysubstrate specificity. Biochemistry 50, 1894–1900 (2011).
Takahito, M. et al. Codon reassignment in the Escherichia coli genetic code. Nucleic Acids Res. 38, 8188–8195 (2010).
Johnson, D. B. F., Xu, J., Shen, Z., Takimoto, J. K. & Wang, L. RF1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nat. Chem. Biol. 7, 779–786 (2011).
Almhjell, P. J., Boville, C. E. & Arnold, F. H. Engineering enzymes for noncanonical amino acid synthesis. Chem. Soc. Rev. 47, 8980–8997 (2018.
Mehl, R. A. et al. Generation of a bacterium with a 21 amino acid genetic code. J. Am. Chem. Soc. 125, 935–939 (2003).
Jeschek, M., Panke, S. & Ward, T. R. Artificial metalloenzymes on the verge of new-to-nature metabolism. Trends Biotechnol. 36, 60–72 (2018).
Huang, J. et al. Unnatural biosynthesis by an engineered microorganism with heterologously expressed natural enzymes and an artificial metalloenzyme. Nat. Chem. 13, 1186–1191 (2021).
Gu, Y. et al. Directed evolution of artificial metalloenzymes in whole cells. Angew. Chem. Int. Ed. 61, e202110519 (2022).
Bloomer, B. J., Clark, D. S. & Hartwig, J. F. Progress, challenges, and opportunities with artificial metalloenzymes in biosynthesis. Biochemistry 62, 221–228 (2023).
Chen, L. et al. Advances in biosynthesis of non-canonical amino acids (ncAAs) and the methods of ncAAs incorporation into proteins. Molecules 28, 6745 (2023).
Chen, Y. et al. Biosynthesis and genetic incorporation of 3, 4-dihydroxy-l-phenylalanine into proteins in Escherichia Coli. J. Mol. Biol. 434, 167412 (2022).
Chen, Y. et al. Unleashing the potential of noncanonical amino acid biosynthesis to create cells with precision tyrosine sulfation. Nat. Commun. 13, 5434 (2022).
Yang, Y. et al. Directed evolution of the fluorescent protein CGP with in situ biosynthesized noncanonical amino acids. Appl. Environ. Microbiol. 90, e0186323 (2024).
Wang, Y. et al. Expanding the structural diversity of protein building blocks with noncanonical amino acids biosynthesized from aromatic thiols. Angew. Chem. Int. Ed. 60, 10040–10048 (2021).
Deng, L. et al. Transfer hydrogenation of CO 2 into formaldehyde from aqueous glycerol heterogeneously catalyzed by Ru bound to LDH. Chem. Commun. 57, 5167–5170 (2021).
Zhou, Z. & Roelfes, G. Synergistic catalysis of tandem Michael addition/enantioselective protonation reactions by an artificial enzyme. ACS Catal. 11, 9366–9369 (2021).
Atta, L. O., Zhou, Z. & Roelfes, G. In vivo biocatalytic cascades featuring an artificial-enzyme-catalysed new-to-nature reaction. Angew. Chem. Int. Ed. 62, e202214191 (2023).
Leveson-Gower, R. B., Zhou, Z., Drienovska, I. & Roelfes, G. Unlocking iminium catalysis in artificial enzymes to create a Friedel-Crafts alkylase. ACS Catal. 11, 6763–6770 (2021).
Austin, J. F. & MacMillan, D. W. Enantioselective organocatalytic indole alkylations. Design of a new and highly effective chiral amine for iminium catalysis. J. Am. Chem. Soc. 124, 1172–1173 (2002).
Ishikura, M., Abe, T., Choshi, T. & Hibino, S. Simple indole alkaloids and those with a nonrearranged monoterpenoid unit. Nat. Prod. Rep. 32, 1389–1471 (2015).
Yang, L., Hou, A., Jiang, Q., Cheng, M. & Liu, Y. Methodological development and applications of tryptamine-ynamide cyclizations in synthesizing core skeletons of indole alkaloids. J. Org. Chem. 88, 11377–11391 (2023).
Zeeli, S. et al. Synthesis and biological evaluation of derivatives of indoline as highly potent antioxidant and anti-inflammatory agents. J. Med. Chem. 61, 4004–4019 (2018).
Webb, B. & Sali, A. Protein structure modeling with MODELLER. In Structural Genomics: General Applications (eds Chen, Y. W. & Yiu, C.-P. B.) 239–255 (Springer US, 2021).
Søndergaard, C. R., Olsson, M. H., Rostkowski, M. & Jensen, J. H. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Case, D. A. et al. Amber 2020 (University of California, San Francisco, 2020).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 25, 247–260 (2006).
Miller, B. R. III et al. MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Liang, X., Li, S. & Su, W. Highly stereoselective imidazolethiones mediated Friedel–Crafts alkylation of indole derivatives. Tetrahedron Lett. 53, 289–291 (2012).
Acknowledgements
We thank G. Roelfes (University of Groningen) and Y.-C. Chen (Sichuan University) for insightful discussions. We appreciate the financial support from the National Key Research and Development Program of China (2021YFA0911500 to Z.Z. and X.S.), the National Natural Science Foundation of China (22207043 to Z.Z.), the Natural Science Foundation of Jiangsu Province (BK20221092 to Z.Z.), and the Tianjin Synthetic Biotechnology Innovation Capacity Improvement Project (TSBICIP-CXRC-026 to X.S.).
Author information
Authors and Affiliations
Contributions
Z.Z. coordinated and conceived the project. W.H. performed the experimental work, analyzed the data, and performed crystal-growing experiments. Y.W., Y.B., and Z.Z. (Zhixi Zhu) assisted in biocatalytic system construction experiments. D.Y. collected and analyzed the X-ray data of proteins. S.W. performed computational modeling under the supervision of X.S. T.L. developed the ncAAs biosynthesis and incorporation system. Z.Z. and X.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, W., Wang, S., Wei, Y. et al. Design and evolution of artificial enzyme with in-situ biosynthesized non-canonical amino acid. Nat Commun 16, 8698 (2025). https://doi.org/10.1038/s41467-025-63733-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63733-3