Abstract
Cell migration is crucial in various biological processes, regulated by surrounding rigidity. Studies under static conditions suggest migration favors rigid substrates, as softer substrates (<4 kPa) do not provide sufficient traction forces. Here we show that mesenchymal stem cells (MSCs) can overcome this limitation when exposed to rapid cyclic changes in substrate rigidity. Under dynamic conditions, cell traction forces progressively rise, promoting a swift mechanical turnover of focal adhesions. This adaptation obviates the need for cell polarity and the mechanochemical turnover of focal adhesions typically required for traditional mesenchymal-type migration. The rapid migration speed together with the shape evolution during migration can be adequately predicted by our theoretical model that considers the force balance under dynamic conditions. Our findings underscore the innate capacity of cells to navigate through fluctuating mechanical cues, highlighting a versatile cellular response mechanism for understanding cell behaviors under dynamic physiological or pathological conditions.
Similar content being viewed by others
Introduction
Cell migration plays a crucial role across a wide range of physiological and pathological events, from tissue development and wound healing to cancer metastasis1,2,3. A wealth of research has uncovered the complex molecular mechanisms that drive cell migration, emphasizing the influence of both chemical signals and biophysical cues4,5,6. These cues include the density of cell adhesion ligands7,8,9, the morphologies10,11 and notably, the mechanical properties of the extracellular matrix (ECM)12,13,14. Among these, the rigidity of the substrate has been identified as a critical factor that governs cell migration. On rigid substrates, cells typically adopt a mesenchymal migration mode, characterized by an elongated morphology and the formation of stable focal adhesions (FAs)1,2,15,16,17. This behavior is integral to developmental processes18,19 and the spread of cancerous cells20,21,22.
The motor-clutch model, formulated by Chan and Odde23,24, provides a comprehensive framework to understand the impact of mechanical cues on cell migration, detailing the dynamics of adhesion and force transmission. According to this model, the polymerization of F-actin filaments drives the cell edge forward, counteracted by the retraction forces generated by ATP-dependent myosin II activity24,25. However, cells encounter difficulties migrating on substrates with rigidity below 4 kPa due to insufficient traction forces generated for propulsion13,26.
This conventional perspective, though informative, falls short of explaining cell migration in the naturally soft environments of various tissues and organs, such as the lungs, liver, and breast27,28. The strategies cells employ to migrate within these soft matrices remain poorly understood. Recent advancements highlight that both spatial gradient rigidity (durotaxis)20,26 and time-dependent rigidity (on viscoelastic substrates)14 significantly influence cell migration patterns. Notably, the identification of an optimal substrate viscoelasticity that promotes cell migration underscores the importance of traction force balance, as described in the molecular-clutch model. Our previous study indicates that rapid changes in substrate rigidity can result in the accumulation of mechanotransduction signaling proteins, thereby enhancing cell traction29. Building on these findings, we aim to investigate the impact of global temporally dynamic alterations in substrate rigidity on cell migration across soft substrates.
In this work, we investigate the effects of cyclic changes in substrate rigidity on the motility of cells on soft substrates. Remarkably, we find that rapid cyclic alterations in substrate rigidity, even with a modest amplitude of about 28%, can significantly increase the migration speed of human mesenchymal stem cells (hMSCs) by 36 folds. On these dynamic substrates, cells exhibit a unique cycle of morphological changes distinct from the common mesenchymal migration mode. The cells first elongate, increasing their aspect ratio, then swiftly revert to a more circular shape, and repeat this process in a random direction. This pattern indicates that cells can adapt their migration strategies in response to minor variations in mechanical cues, challenging the prevailing notion that cells are largely stationary on soft substrates. To further our understanding, we introduce a model that integrates mechanosensing with subsequent mechanotransduction pathways. Our findings suggest that a force imbalance induced by dynamic conditions acts as a powerful driver of rapid, random cell migration. This adaptive mechanism bypasses the conventional requirements of mesenchymal migration, which typically involves cell polarity, along with the turnover of adhesion and cytoskeleton molecules at both the leading and trailing edges. Our investigation into the dynamics of cell migration under variable mechanical conditions reveals a nuanced view of cell motility, offering significant insights into cellular behavior within both healthy and diseased states where substrate rigidity is not static.
Results
Fast cyclic changes in substrate rigidity increase cell migration speed
Cell migration speed correlates with substrate rigidity under static conditions. Cells demonstrate increased migration speeds with higher rigidity, or exhibit a biphasic response to rigidity13,26. Notably, slow migration occurred on the substrate with Young’s modulus below 4 kPa13. Our experiments have validated these trends with hMSCs. Specifically, hMSCs show minimal migration on soft substrates with a Young’s modulus of ~1.6 kPa, yet they migrate effectively on rigid substrates with a Young’s modulus of ~13.0 kPa (Fig. 1a, S1, 2, and Supplementary Movies 1 and 2). However, these observations do not entirely account for cell migration in the naturally soft and dynamic environments of organs like the lungs, liver, and breast, where the elastic moduli are often below 4 kPa. Given the dynamic nature of these biological environments, we proposed that dynamic mechanical conditions might similarly promote cell migration in these soft tissues. To test this hypothesis, we explored the effects of dynamically changing mechanical properties on cell migration behaviors.
a Cells exhibit sluggish movement on substrates possessing a Young’s modulus lower than 4 kPa. Particularly, hMSCs display limited migratory activity on soft substrates with a Young’s modulus around 1.6 kPa, while demonstrating efficient migration on rigid substrates with a Young’s modulus of ~13.0 kPa. The far-right panels show the trajectories of ~20 randomly selected migrating cells for each condition over 12 h. The far-left panels were created in BioRender. Z, Y. (2025) https://BioRender.com/ysxwnvq. b–e Time series of images of hMSC on the PYP hydrogels with different mechanical properties. The bottom-right panels show the trajectories of ~20 randomly selected migrating cells for each condition over 12 h. Times are indicated in hour:minute:second. Scale bar is 50 µm for all panels. f–h Average migration distance, average migration speed and mean square displacement (MSD) of hMSCs on the PYP hydrogels with different mechanical properties. The smooth, linear trends in the population-averaged plots for distance (f) and MSD (h) are emergent properties that result from averaging the asynchronous, intermittent movements of many individual cells. In f–h, n = 204, 202, 224, 207, 218, and 219 (1 min cycle, 5 min cycle, 10 min cycle, 2.2 kPa, 1.6 kPa, and 13.0 kPa) cells were examined, each from 3 independent experiments. In g, *** = p < 0.01, unpaired, two-tailed t-test. i, j Time series of images and average migration distance of hMSCs on the PYP hydrogels with fast cyclic rigidity change (transitioning between 2.2 kPa and 1.6 kPa) in the first 6 h and static rigidity (Young’s modulus of ~2.2 kPa) in the last 6 h. Times are indicated in hour:minute:second. Scale bar is 50 µm. In j, n = 176 cells were examined from 3 independent experiments. In f, h, and j, data are presented as mean values (central line) +/- standard deviation (shaded region). Source data are provided as a Source Data file.
a Time series of cell outlines from randomly selected hMSCs cultured on PYP hydrogels with different mechanical properties. Scale bar is 50 µm for all panels. b–e Cell area, circularity, roundness, and instantaneous migration speed of individual hMSCs on the PYP hydrogels with different mechanical properties. Representative figures for individual cells are shown in Fig. 2b–e, and additional representative statistics for multiple individual cells are presented in Fig. S8–11. f Quantitative analysis of directions (left) and direction autocorrelation (right) of cell migration on substrate with fast cyclic rigidity change (transitioning between 2.2 kPa and 1.6 kPa, 1 min on/off) and mesenchymal migration on static rigid substrates (Young’s modulus of ~13.0 kPa). In the right panel, data are presented as mean values (central line) +/- standard deviation (shaded region). In f, n = 204 and 219 (1 min cycle and 13.0 kPa) cells were examined, each from 3 independent experiments. g Representative point-scanning confocal images of hMSCs on substrate with fast cyclic rigidity change and static rigid substrates subject to indicated treatments immunostained with antibody against Myosin IIb and Golgi. Scale bar is 50 µm for all panels. h Quantification of Myosin IIb intensity polarity on the two sides and Golgi apparatus distribution angle from hMSCs on substrate with fast cyclic rigidity change and static rigid substrates. In h, n = 208 cells were examined from 3 independent experiments. In h, *** = p < 0.01, unpaired, two-tailed t-test. Source data are provided as a Source Data file.
We utilized photo-responsive hydrogels with dynamically and reversibly switchable rigidities for cell migration studies on substrates with fast cyclic rigidity changes. The fast and reversible conformational change in photo-active yellow protein (PYP) under light illumination30 enabled the hydrogels to rapidly switch between soft and rigid states for several hours (Fig. S3). Time-resolved rheological analyses and atomic force microscopy (AFM) nanoindentation measurements confirmed the complete reversibility of rigidity changes between the dark and light states under 1-min cycling intervals. The hydrogels maintained consistent, reversible behavior over 12 hours of cyclic testing, with negligible intracycle signal drift (Fig. S3). Throughout the rheological measurements, the loss modulus (G”) was found to be approximately one-sixth of the storage modulus (G’). This low loss tangent (tan δ « 1) indicates a predominantly elastic response, suggesting that time-dependent viscoelastic effects are minimal in this system. The near-complete overlap between approach and retract curves in AFM nanoindentation indicated minimal viscoelastic contributions during dynamic stiffness transitions (Fig. S4a, b). Complementary rheological characterization further demonstrated that PYP hydrogels exhibited negligible stress relaxation; when subjected to a controlled 10% strain, the initial resulting stress of approximately 2 kPa remained nearly constant over the measurement period (Fig. S4c). Together, these findings establish that the light-induced modulus changes in PYP hydrogels are predominantly elastic in nature, with no substantial alterations to time-dependent viscoelastic properties such as stress relaxation. Importantly, these modulus changes occurred without significant volumetric deformation of the hydrogels. Additionally, during hydrogel fabrication, an excess of RGD peptides (5 mM) was immobilized on the hydrogel surface, ensuring a constant ligand density throughout modulus modulation (Fig. S5). This effectively decouples the effects of ligand presentation from mechanical cues in modulating cellular responses. This unique switchable mechanical properties of these PYP hydrogels allowed us to investigate the impact of fast rigidity changes on cell migration.
We cultured hMSCs on photo-responsive PYP hydrogels. Following a 24-h equilibration period post-plating, we subjected the hydrogels to various cyclic illumination protocols to assess the effects of cyclic rigidity changes on cell migration (Fig. 1a–e, S1–2, and Supplementary Movies 1–6). Cyclic illumination was continuously applied for 12 h to coincide with the cell migration observation window, with light intensity being rigorously monitored throughout the experiments to maintain consistent illumination conditions between experimental groups. To eliminate potential confounding effects from cell-cell interactions and specifically investigate cell-substrate interaction dynamics during migration, high-magnification video recordings were acquired in parallel with multicellular observations. The zoomed-in video datasets were analyzed to resolve individual cell movements, while separate multicellular recordings provided population-level context. Extended illumination had not impact on migration (Fig. S6, Supplementary Movie 7) or cell viability (Fig. S7) on photo-insensitive polyacrylamide hydrogels (PA gel, Young’s modulus of ~2.2 kPa). We used hMSCs cultured on PYP hydrogels under dark (Young’s modulus of ~2.2 kPa) and continuous illumination (Young’s modulus of ~1.6 kPa) as static state controls. Consistent with previous findings, cells showed limited migration on soft substrates after stable attachment on static hydrogels. To investigate the effects of dynamic conditions, we varied the cyclic illumination frequencies from 1 min to 30 min. Surprisingly, migration strongly correlated with switching frequencies, with higher frequencies resulting in greater cell movement after extended periods of rigidity switching (Fig. 1f–h). Peak migration occurred under 1-min cycles (1 min on/off), followed by 5-min cycles (5 min on/off). No noticeable changes in migration were observed with frequencies slower than 10 min (10 min on/off). Under 1-min cycle (transitioning between 2.2 kPa and 1.6 kPa) migration increased more than 36-fold compared to static soft substrates and over 2.5 times that of typical mesenchymal migration on static rigid substrates (Young’s modulus of ~13.0 kPa). Additionally, cell migration slowed down after the cessation of fast cyclic substrate rigidity changes (Fig. 1i, j, Supplementary Movie 8). This finding challenges the conventional notion that cells struggle to migrate on soft substrates and highlights the need for further investigation into the effects of fast cyclic rigidity changes. For convenience, we refer to the hydrogels that undergo cyclic rigidity changes between 2.2 kPa and 1.6 kPa at a frequency of 1 minute on/off as “dynamic soft substrates” hereafter.
Characterization of morphology of cells migrating on dynamic soft substrates
We then examined whether the cell migration on substrates with rapid cyclic changes in rigidity resembled the conventional mesenchymal migration observed on static, rigid substrates. As depicted in Supplementary Movies 2 and 3, the morphology of cells migrating on dynamic soft substrates differed significantly from that of mesenchymal migration on static substrates with a Young’s modulus of ~13.0 kPa.
Cells on dynamic soft substrates exhibited periodic morphological alterations (Fig. 2a), with cell area, circularity, round ratio, and instantaneous speed undergoing dramatic periodic changes (Fig. 2b–e and S8-11). The morphological changes involve an initial elongation phase, followed by a rapid snap-back, where the cell contracts along its long axis, resulting in a sharp decrease in the transverse-to-longitudinal ratio and a return to a rounded shape. Subsequently, this cycle of elongation and snap-back is repeated, leading to rapid migration. Interestingly, cell migration direction on the dynamic soft substrates is more random than mesenchymal migration on static rigid substrates. After the snap-back, the direction of cell re-elongation is random (Fig. 2f), in contrast to the persistence of the direction in mesenchymal migration. This observation is further substantiated by a MSD analysis, performed as described by Gorelik & Gautreau31. On a log-log plot, the MSD for cells on dynamic substrates increases linearly with time, yielding a persistence exponent (α)—defined as the slope of the log (MSD) vs. log (time) data—of approximately 1 (Fig. S12). An α value of ~1 is the hallmark of a classic random walk, providing strong quantitative evidence for non-persistent, diffusive-like motion. This contrasts with the mesenchymal migration on static rigid substrates, the shape of which is similar to the data presented by Gorelik & Gautreau31. For this static condition, the persistence exponent (α value) remains between 1 and 2, with an overall α value of approximately 1.2 obtained by performing a linear regression on the entire curve. Thus, both the MSD-derived α value of ~1 (Fig. S12) and direction autocorrelation analysis (Fig. 2f) confirm the non-persistent, random-walk-like nature of migration on dynamic substrates, which is characterized by a rapid loss of directional memory.
Previous research has demonstrated that cell polarization, arising from symmetry breaking, facilitates directed cell mesenchymal migration on static rigid substrates32. In contrast, the direction autocorrelation analysis of cell migration on dynamic soft substrates revealed that cell polarization does not drive this process, representing a migration mode characterized by maintained symmetry during movement (Fig. 2f). This finding reveals an unconventional framework for interpreting complex cell migratory behaviors.
To better understand the random migration direction of cells on dynamic soft substrates, as opposed to the directed migration seen in mesenchymal cells on static rigid substrates, we investigated the localization patterns of non-muscle Myosin IIb, which is essential for forming the rear of mesenchymal migrating cells on static substrates33. As shown in Fig. 2g, h, while Myosin IIb predominantly located at the rear of cells on static rigid substrates (Young’s modulus of ~13.0 kPa), in line with prior studies, this polarized localization pattern was absent in cells on dynamic soft substrates. Co-staining of the Golgi apparatus with Myosin IIb further revealed that hMSCs on dynamic soft substrates exhibited a broad perinuclear distribution of the Golgi apparatus (Fig. 2g, h), whereas those on static rigid substrates displayed a polarized distribution at one end relative to the nucleus. This observation further supports the absence of polarity in hMSCs on dynamic soft substrates. These findings indicate that, in contrast to mesenchymal migrating cells—which typically exhibit a distinct protruding front and a contractile rear—cells migrating on dynamic soft substrates lack polarity. This suggests that cell elongation is isotropic, occurring uniformly on both sides during the elongation phase, and that the detachment occurring during the rapid snap-back phase is random at either side.
Actin polymerization and adhesion dynamics during cell migration on dynamic soft substrates
After observing periodic morphological changes in cells migrating on dynamic soft substrates, we investigated the underlying cellular mechanisms. Since cell morphology is intricately tied to the cytoskeleton and cell adhesion, we focused on actin network activity and focal adhesion dynamics (Fig. 3a–d).
a–d Cell adhesion and morphology (a), relative F-actin intensity (b), quantification of F-actin intensity polarity (c) and FA area (d) of hMSCs on the PYP hydrogels with different mechanical properties. Scale bar is 50 µm for all panels. In b, c, and d, n = 210, 203, 214, and 212 (1 min cycle, 13.0 kPa, 2.2 kPa, and 1.6 kPa) cells were examined, each from 3 independent experiments. e–h Time series of images of hMSC migration (e, h), average cell migration speed (f) and probability of cell detachment (g) on PYP hydrogels with fast cyclic rigidity change with addition of NSC23766 (10 μM, –Rac, e), latrunculin A (100 nM, +Lat. A), and β1-integrin blocking (5 μg ml–1, –β1, h), or with different mechanical properties (g). In e and h, the bottom-right panel shows the trajectories of ~20 randomly selected migrating cells over 12 h. Times are indicated in hour:minute:second. Scale bar is 50 µm for all panels. In f, n = 204, 200, 202 and 210 (Ctrl, –Rac, +Lat. A, and –β1) cells were examined, each from 3 independent experiments. In h, n = 185 and 218 (1 min cycle and 2.2 kPa) cells were examined, each from 6 independent experiments. i–l Actin dynamics (i, j) and focal adhesion dynamics (k, l) of hMSCs on the PYP hydrogels with different mechanical properties. In i, representative kymographs are shown on the left side, scale bar is 20 µm for all panels. In j, n = 201, 198, 211 and 187 (2.2 kPa, 1.6 kPa, 13.0 kPa, and 1 min cycle) cells were examined, each from 3 independent experiments. In k, scale bar is 2 µm for all panels. In l, n = 185 and 186 (1 min cycle and 13.0 kPa) cells were examined, each from 3 independent experiments. Data in g and l are presented as mean values +/- standard deviation. In all graphs, *** = p < 0.01, n.s. = p > 0.1, unpaired, two-tailed t-test. Source data are provided as a Source Data file.
We assessed actin network activity using F-actin staining, which revealed an abundant and orderly alignment of actin stress fibers at the edges of hMSCs on dynamic soft substrates (Fig. 3a, b, and S13), indicating that actin polymerization plays a critical role in migration under these conditions. Interestingly, while F-actin typically localizes at the front of cells on static, rigid substrates, such polarization was absent on dynamic soft substrates, further highlighting the non-polar nature of migration in this environment (Fig. 3c).
To further investigate the role of the actin network, we used the inhibitor NSC23766 to block Rac134, which impedes actin nucleation (Fig. S14). This resulted in a significant reduction in cell migration on dynamic substrates (Fig. 3e, f, and Supplementary Movie 9). A similar cessation of migration was observed when actin polymerization was inhibited using latrunculin A (+Lat. A)35,36,37 (Fig. 3f and S15, Supplementary Movie 10), reinforcing the importance of enhanced actin dynamics for migration on dynamic soft substrates.
In addition to actin dynamics, we found that cell adhesion is strengthened on dynamic soft substrates, as evidenced by increased Paxillin accumulation (Fig. 3a, d) and elevated integrin recruitment (Fig. S16) in immunofluorescent staining. The size of focal adhesions was significantly larger on dynamic substrates than on static soft substrates (Young’s modulus ~2.2 or 1.6 kPa), suggesting that cell adhesion is enhanced during migration induced by cyclic changes in substrate rigidity.
Typically, the enlargement of adhesion plaques and the formation of platypodia are markers of stable cell adhesion. However, the rapid retraction observed following cell elongation suggests that these strengthened focal adhesions are more prone to rupture under dynamic conditions. Indeed, cells on dynamic soft substrates exhibited a higher detachment rate over 24 hours compared to those on static substrates (Fig. 3g).
To resolve this apparent contradiction and confirm the role of enlarged focal adhesions in promoting migration on dynamic soft substrates, we altered focal adhesion size by introducing β1-integrin blocking antibodies14 into the culture medium (Fig. S17). This treatment prevents unbound integrins from forming new integrin-ECM bonds, thereby shortening the lifespan of integrin-ECM interactions. Our results showed that β1-integrin blocking (–β1) significantly impaired cell migration on dynamic soft substrates (Fig. 3f, h, and Supplementary Movie 11), suggesting that the formation of enlarged focal adhesions is crucial for accelerating cell migration.
To further investigate how dynamic rigidity variations influence actin dynamics, we implemented live-cell imaging to quantify retrograde flow patterns (Supplementary Movies 12–14). Through systematic analysis of complete morphological cycles combined with multi-timepoint velocity measurements over 12-h periods under dynamic mechanical conditions, we observed distinct flow characteristics (Fig. 3i, j). Quantitative comparisons demonstrated that dynamic substrates (transitioning between 2.2 kPa and 1.6 kPa, 1 min on/off) exhibited significantly faster actin retrograde flow velocities compared to static controls across all rigidity levels (soft: 2.2 or 1.6 kPa; rigid: 13.0 kPa). These direct measurements confirm that dynamic rigidity changes enhance actin activity.
To further evaluate the impact of dynamic rigidity variations on focal adhesion dynamics, we performed live-cell fluorescence recovery after photobleaching (FRAP) experiments using Talin-GFP to directly compare adhesion turnover between dynamic and static substrates (Supplementary Movies 15-18). Quantitative time-resolved analysis demonstrated significantly faster Talin fluorescence recovery in dynamically modulated rigidity environments compared to static controls across all rigidity levels (soft: 2.2 or 1.6 kPa; rigid: 13.0 kPa), confirming enhanced focal adhesion remodeling through accelerated assembly-disassembly kinetics (Fig. 3k, l, and S18).
Force imbalance drives the detachment of adhesions and the “snap-back” process
We hypothesized that the migration rate depends not solely on actin network activity or focal adhesions independently, but on their interplay. Our previous study suggests that rapid rigidity changes lead to an accumulation of mechanosignaling proteins, such as phosphorylated FAK (pFAK), due to the slower dephosphorylation rate compared to the frequency of cyclic changes29. This accumulation enhances actin polymerization and increases the phosphorylation level of Myosin IIa, resulting in higher endogenous forces generated by the cells. As a consequence, the increased traction forces promote integrin engagement at focal adhesions, leading to the formation of platypodia.
However, there is a time lag between the rise in endogenous forces from enhanced actin activity and the corresponding increase in adhesion strength due to focal adhesion growth. This lag creates an imbalance, where the internal forces cannot be fully countered by the external adhesion strength. Once this imbalance reaches a critical threshold, a collapse of focal adhesions at their weakest points occurs, initiating the ‘snap-back’ process observed during migration on substrates undergoing fast cyclic rigidity changes.
To validate this hypothesis, we compared the internal cellular traction forces with the external adhesion forces between the cells and hydrogel substrates. Adhesion forces were directly measured using traction force microscopy. Since directly measuring endogenous traction forces from actin network activity is challenging, we used intracellular phosphorylated Myosin IIa (pMyosin IIa) levels as an indicator of internal traction forces. These measurements were conducted on dynamic soft substrates (PYP hydrogels with fast cyclic rigidity changes of 1 min on/off) and on static rigid substrates (PYP hydrogels with a Young’s modulus of ~13.0 kPa) as controls.
The results showed that cell adhesion forces on dynamic soft substrates were comparable to those on static rigid substrates (Fig. 4a, b and S19). Consistent with previous observations of morphological symmetry, cell adhesion forces on static rigid substrates exhibited the characteristic front-rear asymmetry of classical mesenchymal migration, while those on dynamic substrates with fast cyclic rigidity changes demonstrated increased symmetry, further emphasizing the non-polar migration behavior in this mechanical environment (Fig. S20). However, intracellular pMyosin IIa levels were significantly higher on dynamic soft substrates (Fig. 4c, d). Notably, spatial analysis demonstrated significantly elevated pMyosin IIa levels in both cytoplasmic compartments and F-actin-enriched regions, with enhanced co-localization between pMyosin IIa and F-actin (Fig. S21), substantiating the reinforcement of cell-endogenous contractile forces. Western Blot analysis and immunofluorescence (Figs. S22, 23) further demonstrated that, in addition to pMyosin IIa, the phosphorylation levels of other signaling proteins, such as pFAK and pMLC, were also elevated on dynamic soft substrates. Assuming that internal forces are balanced with cell-ECM adhesions on static rigid substrates (Young’s modulus of ~13.0 kPa), these results indicate that the intracellular forces generated by pMyosin IIa significantly exceed the extracellular adhesion forces on fast dynamic soft substrates (transitioning between 2.2 kPa and 1.6 kPa, 1 min on/off).
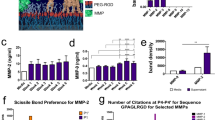
a, b Adhesion forces of hMSCs on static soft (2.2 kPa) and rigid (13.0 kPa) PYP hydrogels and on PYP hydrogel with fast cyclic rigidity change (1 min cycle) over 12 h. To better represent the average cell force, the results are presented as traction forces per unit area (N m-2), also known as traction stress (Pa). Scale bar is 50 µm for all panels. In b, n = 215, 212 and 204 (2.2 kPa, 1 min cycle, and 13.0 kPa) cells were examined, each from 3 independent experiments. c, d Mechnosignalling protein pMyosin IIa level of hMSCs on static soft (2.2 kPa) and rigid (13.0 kPa) PYP hydrogels and on PYP hydrogel with fast cyclic rigidity change (1 min cycle) over 12 h. Scale bar is 50 µm for all panels. In d, n = 213, 205 and 214 (2.2 kPa, 1 min cycle, and 13.0 kPa) cells were examined, each from 3 independent experiments. e Time series of images of hMSCs migration on PYP hydrogels with fast cyclic rigidity change with (OE Talin) overexpressing Talin. The far-right panel shows the trajectories of ~20 randomly selected migrating cells under the condition over 12 h. Times are indicated in hour:minute: second. Scale bar is 50 µm for all panels. f Time series of images of hMSCs migration on PYP hydrogels with fast cyclic rigidity change with the addition of non-phototoxic Blebbistatin inhibitors (25 μM, –pMyosin). The far-right panel shows the trajectories of ~20 randomly selected migrating cells under the condition over 12 h. Times are indicated in hour:minute:second. Scale bar is 50 µm for all panels. g Average cell migration speed on PYP hydrogels with fast cyclic rigidity change with and without (Ctrl) overexpressing Talin (OE Talin) and addition of non-phototoxic Blebbistatin inhibitors (–pMyosin). In g, n = 204, 202, and 222 (Ctrl, OE Talin, and –pMyosin) cells were examined, each from 3 independent experiments. In all graphs, *** = p < 0.01, n.s. = p > 0.1, unpaired, two-tailed t-test. Source data are provided as a Source Data file.
To further test whether the imbalance between internal and external forces drives fast cell migration, we conducted additional control experiments manipulating inner and outer forces. We transfected cells with plasmids overexpressing Talin (OE Talin)38 to enhance cell adhesion and regulate migration (Fig. S24). The results showed that cells overexpressing Talin on dynamic soft substrates shifted to a mesenchymal migration mode, resulting in a considerable decrease in migration speed and a transition in migration direction from random movement to more persistent orientation (Fig. 4e, g, Fig. S25, Supplementary Movie 19). Talin overexpression strengthens the connection between stress fibers and integrins, stabilizing the molecular clutch and reinforcing cell-substrate adhesion. Even though endogenous forces were enhanced, they were insufficient to break cell-substrate connections, forcing cells to rely on the chemical turnover of focal adhesions for migration, similar to mesenchymal migration observed on static rigid substrates. Under dynamic mechanical conditions, Talin overexpression promoted focal adhesion assembly compared to non-overexpressing controls, effectively resolving intracellular/extracellular force imbalance through reinforced adhesion-mediated force transmission. This restoration of mechanical homeostasis converted the dynamic substrate-induced rapid migration mode observed in non-overexpressing cells into conventional mesenchymal migration patterns, aligning with the adhesion-stabilized force equilibrium paradigm.
Conversely, when endogenous cellular forces were reduced by inhibiting Myosin IIa phosphorylation (–pMyosin) with a non-phototoxic blebbistatin derivative39, cells were also unable to migrate rapidly on dynamic soft substrates (Fig. 4f, g, Supplementary Movie 20). Under dynamic mechanical conditions, inhibition of Myosin IIa phosphorylation attenuated intracellular contractility compared to non-inhibited controls, effectively resolving intracellular/extracellular force imbalance and suppressing the cyclic cytoskeletal remodeling essential for increasing migration speed in phosphorylation-competent cells. This transition to a low-tension homeostatic state impaired the mechanochemical feedback loop necessary for sustained traction force generation, culminating in complete migratory cessation under dynamic substrate conditions. This confirms that the accumulation of sufficient intracellular forces, exceeding the adhesion strength, is crucial for fast cell migration on dynamic soft substrates.
Quantitative model
To quantitatively understand the rapid cell migration on dynamic soft substrates, we propose a refined physical model. In our previous work, we developed a model that accounted for the rapid dynamic changes in substrate rigidity due to the accumulation of cell forces. This model integrated both mechanosensing and mechanosignaling dynamics by combining the molecular clutch model24 with a rigidity-dependent FAK phosphorylation mechanism40. In the current study, we further advance this model by incorporating the cell spreading module proposed by Shenoy and coworkers41, enabling us to predict the dynamic changes in cell morphology and migration.
In our model, myosin motors pull the actin filaments toward the cell center, creating a retrograde flow (Fig. 5a). The molecular clutches connect F-actin to the substrate and can randomly disengage or engage, with their dissociation or association rates defined by \({r}_{{off}}\) and \({r}_{{on}}\), respectively. When engaged, these clutches impede the retrograde flow, allowing the polymerization of the leading edge to advance the cell membrane, thereby facilitating cell spreading and elongation7,42,43,44,45. The clutches are modeled as springs with a specific stiffness \({k}_{c}\), and the RGD-coated hydrogel substrate is simplified as another spring with its own stiffness \({k}_{{ECM}}\). We simulate dynamic changes in rigidity by varying \({k}_{{ECM}}\). When the substrate softens (decreasing from\(\,{k}_{{ECM}}^{{stiff}}\) to \({k}_{{ECM}}^{{soft}}\)), external adhesion forces reduce instantly following Hooke’s law. This necessitates a rapid adjustment in the molecular clutch chains to maintain force equilibrium at the adhesion sites. Conversely, as the substrate rigidifies, new clutches form, incorporating both cytoplasmic pFAK and its inactive forms within focal adhesions. Supplementary Table S1 contains the model parameters (see Supplementary Information for more details).
a Schematic of the molecular clutch model of a cell attached to a substrate. b–d Average migration distance, MSD and FA area of hMSCs on substrates with different mechanical properties over 12 h in the experiments and simulations. In b-d, n = 204, 224 and 214 (1 min cycle, 10 min cycle, and 2.2 kPa) cells were examined, each from 3 independent experiments. In d, *** = p < 0.01, unpaired, two-tailed t-test. (e-f) Average migration distance and MSD of hMSCs on substrates with fast cyclic rigidity changes over 12 h, slowing the actin polymerization rate (–Rac) and lowering the number of motors (–pMyosin) in the experiments and simulations. In e, f, n = 204, 200 and 222 (Ctrl, –Rac, and –pMyosin) cells were examined, each from 3 independent experiments. In b, c, e, and f, the experimental data are presented as mean values (points) +/- standard deviation (shaded region); the simulation results are shown as mean values (solid line). g Bond lifetime (time elapsed between initial attachment of a single bond to its breakage) versus interval of cyclic change. Data are presented as mean values +/- standard deviation (error bars), which reflects the predicted variability. h Fast cyclic substrate rigidity changes trigger rapid cell migration. I) → II) Fast cyclic substrate rigidity changes periodically accumulate signalling proteins because the slow dephosphorylation rate of pFAK during the softening phase and the rapid phosphorylation rate during the rigidifying phase. The accumulated signalling proteins further catalyse the mechanotransduction pathways to promote actin polymerization, resulting in cell elongation. II) → III) The continuous accumulation of mechanosignalling protein pMyosin IIa makes the pMyosin IIa-generated intracellular force continue to exceed the adhesion force between cell and ECM. As a result, the entire focal adhesion on one side of the cell collapses and the cell snap-backs to the other side, which leads to rapid cell migration. In all panels, the simulation results are calculated from 200 independent simulation runs. Source data are provided as a Source Data file.
As an initial step, we verified that our simulation replicated key experimental findings. We simulated the effects of varying cyclic frequencies of rigidity changes on cell migration speed and mean square displacement (MSD). The simulations predicted that faster cyclic changes in substrate rigidity would increase both migration distance and MSD (Fig. 5b, c), with the average migration distance and MSD following a linear relationship with time, consistent with experimental results. This emergent linearity at the population level can be attributed to the underlying single-cell dynamics. The timing of ‘snap-back’ events, which are triggered when intracellular contractile forces (pMyosin IIa) exceed extracellular adhesion forces, varies significantly between individual cells (as shown in Supplementary Movie 3). This temporal heterogeneity is fundamentally driven by the stochastic nature of integrin-ECM binding dynamics. Consequently, the dispersion of these asynchronous snap-back events averages out the discontinuities of single-cell trajectories, giving rise to the apparent linear increase in average migration distance over time at the ensemble level. Furthermore, the simulations accurately reflected the experimental impact of fast cyclic rigidity changes on focal adhesion size (Fig. 5d). The simulations also demonstrated a reduction in migration speed when actin network activity was inhibited by the Rac1 inhibitor NSC23766 (–Rac) or latrunculin A (+Lat. A), or when Myosin IIa phosphorylation or integrin activity was suppressed (Fig. 5e, f and S26, 27). Additionally, our simulations predict that cell–matrix bond lifetime decreases with faster cyclic rigidity changes, as the higher force loading rate shortens the lifetime of bonds before rupture (Fig. 5g). This prediction is mechanistically corroborated by FRAP analyses demonstrating significantly accelerated adhesion turnover kinetics under dynamic rigidity modulation compared to static substrate conditions across all tested rigidity levels (Young’s modulus of ~2.2, 1.6 and 13.0 kPa).
These simulations offer insights into the molecular mechanisms underlying how fast cyclic changes in substrate rigidity influence cell migration (Fig. 5h). As substrate rigidity decreases, components of the molecular clutch disassemble, releasing pFAK into the cytoplasm. However, pFAK does not immediately dephosphorylate within 5 minutes. As the substrate rigidifies again, the molecular clutch reassembles, capturing pFAK from the cytoplasm and generating newly phosphorylated FAK in focal adhesions (FAK has a half-life of ~10 seconds in adhesions before being phosphorylated to pFAK)46,47,48, leading to pFAK accumulation. With prolonged cyclic changes in substrate rigidity, these fast mechanical stimuli periodically accumulate signaling proteins, further amplifying mechanotransduction pathways (e.g., pFAK/Src-RhoA/ROCK-pMyosin IIa), enhancing pMyosin IIa-generated intracellular forces, actin polymerization, and cell adhesion (refer to the Supplementary Document for further details).
Critically, the accumulation of mechanosignaling proteins under dynamic conditions drives intracellular force amplification through two competing mechanisms: (1) Force-enhanced actomyosin contractility promotes FA assembly by mechanically recruiting plaque components. (2) Pre-activated mechanosignaling proteins (e.g., pMyosin IIa) within newly assembled molecular clutches generate elevated intrinsic forces. Rather than distributing forces across clutches, this pre-activation renders FAs prone to force overload.
Our experimental evidence (Fig. 4c, d and S20) demonstrates significant increases in pMyosin IIa levels both in cytoplasmic compartments and F-actin-enriched regions, with enhanced colocalization between pMyosin IIa and F-actin. These findings confirm that newly assembled molecular clutches contain elevated levels of pre-activated mechanosignaling proteins such as pMyosin IIa, which directly contributes to FA instability and reduced lifetime.
The continuous accumulation of mechanosignaling proteins like pMyosin IIa results in intracellular traction forces consistently exceeding adhesion forces with the ECM, which accelerates actin retrograde flow (Fig. S28). This acceleration in actin polymerization facilitates ongoing cell elongation, increasing cytoskeletal resistance forces and ultimately amplifying the forces exerted by the molecular clutches within focal adhesions. When these forces accumulate to a critical level, focal adhesions on one side of the cell collapse, propelling the cell to snap back in the opposite direction. Under rapid cyclic rigidity changes, the cell re-extends in a random direction, repeating the process, resulting in stochastic migration patterns. Conversely, when cyclic stimulation ceases, pMyosin IIa accumulation stops, and intracellular traction forces no longer surpass adhesion forces. This reduction in force disrupts actin retrograde flow and cytoskeletal resistance, inhibiting further actin polymerization and cell elongation, thus slowing cell migration. Therefore, the dynamic accumulation of mechanosignaling proteins, driven by rapid cyclic changes in substrate rigidity, is crucial in facilitating rapid cell migration on dynamically changing substrates.
Quantitative model predicts cell morphodynamics and migration speeds
After confirming that our quantitative model accurately reflects the effects of fast cyclic changes in substrate rigidity on cell migration, we sought to experimentally validate the model’s predictions concerning signaling protein accumulation and its impact on cell morphodynamics, elongation duration, and migration speed. Specifically, we modulated the softening amplitude from the rigid state to 28% and 40% of the rigid state to control the rate of signaling protein accumulation (Fig. 6a, Supplementary Movie 21). The larger softening/stiffening amplitude leads to greater disassembly of molecular clutches, releasing more pFAK into the cytoplasm where it becomes available for re-recruitment upon re-stiffening, resulting in a net increase in the signaling protein accumulation rate as demonstrated in our previous work29. The model predicted that increasing the softening amplitude would accelerate the rate of cell elongation due to more rapid actin polymerization (Fig. 6b). However, this faster elongation rate also caused a quicker increase in cytoskeletal resistance forces. As a result, the combination of faster cytoskeletal resistance buildup and the collapse of a higher number of molecular clutches during each softening event led to a more rapid collapse of focal adhesions and a shortened elongation duration (Fig. 6c). While the reduced elongation duration resulted in smaller maximum cell elongation (Fig. 6d), the faster elongation rate contributed to an overall increase in migration speed (Fig. 6e and Fig. S29).
a Time series of images of hMSC migration (above) and outlines (below) on PYP hydrogels with fast cyclic rigidity change (1 min on/off) with larger softening amplitude from the rigid state (40% of the rigid state, transitioning between 2.2 kPa and 1.3 kPa,). The far-right panel shows the trajectories of ~20 randomly selected migrating cells under the condition over 12 h. Scale bar is 50 µm for all panels. b–d Cell elongation characteristics of hMSCs on substrates with fast cyclic rigidity changes with different softening amplitude from the rigid state (28% and 40% of the rigid state) predicted by our simulation and validated by experiments. In b–f, n = 204 and 213 (28% and 40%) cells were examined, each from 3 independent experiments. In b, the simulation results are shown as mean values (central line) +/- standard deviation (shaded region), which reflects the predicted variability. The experimental data are presented as mean values (points) +/- standard deviation (error bars). e–g Average migration distance, MSD and FA area of hMSCs on substrates with fast cyclic rigidity changes with different softening amplitude from the rigid state (28% and 40% of the rigid state) over 12 h, predicted by our simulation and validated by experiments. In g, n = 210 and 216 (28% and 40%) cells were examined, each from 3 independent experiments. In e and f, the experimental data are presented as mean values (points) +/- standard deviation (shaded region); the simulation results are shown as the mean values (solid line). In all graphs, *** = p < 0.01, unpaired, two-tailed t-test. Source data are provided as a Source Data file.
Despite the faster migration speed, the shorter elongation duration, which reflects a higher snap-back frequency, combined with the random migration direction, led to a decrease in the mean square displacement (MSD) of cell migration (Fig. 6f). This apparent paradox can be resolved by noting the distinction between scalar speed, derived from total path length, and vector-based MSD. The more rapid rate of elongation at 40% amplitude increases the total path length traveled per unit time, thus leading to a higher calculated migration speed. In contrast, the MSD is a vector-dependent quantity that measures net displacement. The shortened elongation duration in this condition—reflecting a higher snap-back frequency—reduces the effective “step length” of each migration cycle. Furthermore, a quantitative analysis of migration directions confirms that the cell re-orients randomly after each snap-back event in both cases (Fig. 2f and S30a). Consistent with this, the analysis of log-log MSD plots yields a persistence exponent (α-value) of approximately 1 for both conditions—the hallmark of a classic random migration (Fig. S30b). The more frequent directional resets from the faster snapbacks in the 40% amplitude condition cause a slightly faster decay in the direction autocorrelation function (Fig. S30c). Taken together, these factors—a shorter effective step length combined with a migration mode that, while fundamentally random (α ≈ 1), exhibits a more rapid loss of directional memory—synergistically lead to a less efficient accumulation of net displacement and, consequently, a lower overall MSD. Additionally, the faster accumulation of signaling proteins increased focal adhesion area (Fig. 6g). Experimentally, we confirmed all of these predictions, validating the model’s accuracy. Notably, the ability of our computational model to integrate these competing effects of speed and step length, and to correctly predict this non-intuitive outcome, underscores its predictive fidelity. For more detailed information, please refer to the Supplementary Information.
Discussion
In this study, we discovered that fast cyclic changes in substrate rigidity play a crucial role in regulating adherent cell migration on soft substrates. Our findings reveal that as the frequency of cyclic rigidity changes increases, so does the migration speed. Migration on these substrates is characterized by a continuous cycle of cell elongation and snap-back, with the direction of migration being random and independent of the initial cell spreading. However, it is possible that different types of adherent cells may exhibit distinct migration behaviors due to variations in the dephosphorylation rates of mechanosignaling proteins49,50,51 or the strength of cell-ECM interactions13,16,20,22.
While our findings elucidate how dynamic rigidity changes regulate mesenchymal motility, it is essential to emphasize that these conclusions are derived exclusively from 2D models. Three-dimensional migration involves distinct mechanobiological constraints, including matrix remodeling and spatial confinement, which may elicit divergent responses to rigidity dynamics. Future work employing 3D dynamically tunable matrices will be critical to extend these principles to physiologically relevant microenvironments.
Our investigation into cell migration on soft substrates is motivated by the recognition that many soft tissues exhibit dynamic, periodic mechanical properties. During tissue development and disease progression27,28, the mechanical microenvironment undergoes dynamic changes over time. Even under normal conditions, tissues and organs experience rhythmic mechanical fluctuations due to natural functions such as heartbeats52, breathing53, blood vessel contractions54, and digestive movements55. These cyclical mechanical forces are essential for proper physiological function, adaptation to varying demands, and maintaining homeostasis. For example, the cyclic biophysical forces generated by normal respiration are vital for maintaining alveolar epithelial cell fate56. Understanding how cells respond dynamically to changes in the mechanical environment is key to regulating cell behavior and fate.
This study introduces temporal rigidity dynamics as a regulatory mechanism for cell migration in soft microenvironments. While prior work established rigidity gradients as spatial guidance cues16,20,26, we demonstrate that time-varying mechanical signals (transitioning between 2.2 kPa and 1.6 kPa) in uniform substrates fundamentally alter migration mechanics. The key innovation lies in revealing how rapid (1-min cycle) rigidity changes reprogram mechanoadaptation: cells transition from stable polarization on static substrates to a dynamic migration mode characterized by cyclic morphological reshaping and transient adhesion remodeling. This mechanical plasticity enables sustained motility without directional bias – a behavior distinct from both durotaxis and conventional mesenchymal migration.
Crucially, our phototunable hydrogel platform achieves dynamic rigidity modulation within physiological softness ranges (<4 kPa), recapitulating the temporal mechanical variations found in developing or remodeling tissues14,27. The discovery that non-directional migration speed can be increased through periodic rigidity changes – rather than spatial gradients – redefines our understanding of mechanoregulation, showing cells prioritize temporal mechanical dynamics over spatial cues in compliant environments. By decoupling temporal and spatial mechanical variables, we establish substrate dynamics as an independent control parameter for migration, a paradigm shift from static or gradient-based models.
Rapid actin polymerization, marked by the formation of actin-rich stress fibers, is often associated with cell migration on rigid, elastic substrates, where it drives cellular movement2,16,17. Similarly, cell migration on soft substrates with fast cyclic rigidity changes is frequently linked to lamellipodia-mediated mesenchymal migration, as both processes involve rapid actin polymerization and enhanced focal adhesion at the cell edge8,15,16,17. On rigid substrates, mesenchymal migration is stabilized by a dynamic balance between the intracellular forces generated by pMyosin IIa and the adhesion forces at focal adhesions. Additionally, microtubule detyrosination induces symmetry breaking, leading to cell polarization and causing mesenchymal migration to follow a straight path, aligned with the initial direction of movement32. In contrast, cell migration on soft substrates with fast cyclic rigidity changes exhibits random re-elongation directions, resulting in migration paths that are independent of the initial orientation.
Although the snap-back process observed in our study resembles amoeboid-mode migration in three-dimensional environments, where cells exhibit significant morphological changes, the underlying mechanisms are distinct. Amoeboid-mode migrating cells are rounded, exhibit weak adhesion to their surroundings, and are characterized by blebbing driven by forced nuclear movement57,58,59. Our findings demonstrate that cells can achieve rapid migration and undergo dramatic morphological changes while maintaining strong adhesion and mechanotransduction, facilitated by the dynamic cyclic mechanical properties of the ECM.
While cell migration in static mechanical environments has been extensively studied, migration in dynamic mechanical environments remains less understood. Our research provides insights into how cells adapt their migration strategies in response to dynamic, periodic mechanical cues. We found that cells can achieve rapid migration in high-frequency dynamic environments, significantly enhancing their adhesion until they reach a more stable, static state. This enhanced adhesion facilitates rapid establishment of ECM connections in relatively static environments, improving the cells’ ability to find stable environments within dynamic settings and contributing to overall environmental homeostasis. Thus, this mode of migration highlights the importance of dynamic substrate rigidity changes in maintaining cellular homeostasis.
The functional significance of this distinct migratory behavior likely depends on the specific biological context, with potentially contrasting roles in physiology and disease. Our findings suggest this behavior could function as a “search-and-settle” mechanism, enabling cells to efficiently escape local, transiently unstable microdomains, such as those from micro-injury or remodeling, and subsequently arrest upon finding a stable niche. Conversely, this same mechanism could be co-opted in pathological conditions like cancer or fibrosis, where the tissue microenvironment often loses mechanical homeostasis. A potential implication is that the constantly fluctuating matrix stiffness in such a chaotic environment could perpetually trigger high-speed motility, thereby increasing the probability of cancer cell invasion. These potential roles are speculative and highlight a clear need for future research. In vivo investigations will be crucial to fully understand the complex interplay between dynamic mechanical environments and cell behavior in both maintaining health and driving disease.
Methods
Statistics & reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. Sample sizes were determined based on the variation observed in pilot experiments to ensure that the magnitude and consistency of measurable differences between groups were reliable. All statistical analyses were conducted using Origin (v2025b). Two-tailed unpaired Student’s t-tests were used for comparisons between two groups. In this work, *** represents p < 0.01, ** represents p < 0.05, * represents p < 0.1, n.s. represents p > 0.1. All graphs and data are representative of more than three independent experiments. Values are presented as mean values +/- standard deviation (s.d.). The experiments were randomized; all samples were randomly allocated to experimental groups. The investigators were blinded to group allocation during experiments and outcome assessment, including data collection and analysis.
Protein expression and purification
PYP 36/128 genes were cloned into pQE80L vectors. ApoPYP was expressed in Escherichia coli (BL21) and dissolved in 4 mol/L urea phosphate-buffered saline (PBS) buffer (10 mmol/L, pH 7.4). P-hydroxycinnamic anhydride (100 mg for apoPYP isolated from a 1 L culture) was added to the solution, and the reaction mixture was stirred at 4 °C for 4 hours. The precipitate was removed by centrifugation (6708 × g, 60 min). The supernatant was taken and dialyzed against PBS buffer (10 mmol/L, pH 7.4) at 4 °C, and the PBS buffer was changed 3-5 times. Again, the precipitate was removed by centrifugation (6708 × g, 60 min). The supernatant was collected, and then Co2+-NTA protein resin (GE healthcare, Shanghai, China) was used to purify the proteins. Finally, the purified PYP mutants were dialyzed against PBS buffer (10 mmol/L, pH 7.4) at 4 °C.
Preparation of hydrogels
We introduced cysteine (Cys) mutations at a pair of surface-exposed residues, 36 and 128 (PYP 36/128), to facilitate crosslinking of the protein with an 8-armed PEG-Mal (Molecular Weight: 20 kDa) for hydrogel construction (PYP hydrogel). In this study, PYP hydrogels were formed at a poly(ethylene glycol) (PEG)-to-PYP mass ratio of 2:3. Additionally, RGD was incorporated at a final concentration of 5 mM to ensure cell adhesion to the hydrogel surface.
Photo-tuning the mechanical properties of PYP hydrogels
For the AFM nano-indentation and cyclic substrate rigidity experiments, we utilized an optical fiber-guided LED source with tunable light intensities and maximum wavelengths (PE-4000, CoolLED, USA). An optical power meter (818-ST-UV/DB-840C, Newport, USA) was employed to measure the light intensity. AFM-based nano-indentation measurements were undertaken with an atomic force microscope (JPK, Nanowizard IV) at 37 °C to match the temperature of cell culture environments. Silicon nitride cantilevers (MLCT-A; products of Bruker, the half angle of the quadratic pyramidal face θ was 20°) were used in all experiments. Under all experimental conditions, the heat effects were negligible.
Rheometry analysis of PYP hydrogels
The rheological properties of the hydrogels, including storage modulus, loss modulus, and stress relaxation, were characterized following previously described methods60. Briefly, PYP hydrogels were formed into cylindrical discs (8 mm diameter, 3 mm height). Parallel plate shear rheometry was conducted using a HAAKE RheoStress 6000 rheometer at 37 °C. Dynamic time sweep tests to measure storage (G’) and loss (G”) moduli were conducted at a constant strain of 2% and an oscillatory frequency of 1 rad s⁻¹. Stress relaxation measurements were carried out by applying a single step strain of 10%. Under all experimental conditions, the heat effects were negligible.
Cell lines
Human mesenchymal stem cells (hMSCs) were obtained from Hunan Maybio Bio-Pharmaceutical Co., Ltd. (Cat# R7788115), donated from a female.
HMSCs cell culture
HMSCs were cultured in αMEM (Gibco, Thermo Fisher Scientific) with penicillin/streptomycin (1%, Gibco, Thermo Fisher Scientific) and fetal bovine serum (10%, Gibco, Thermo Fisher Scientific) in a 5% CO2 atmosphere at 37 °C. The culture medium was changed every other one to two days until the cells were passaged at approximately 90% confluency. After trypsinization, hMSCs were seeded (1×106 cells/mL) on PYP gels (the PYP concentration was 50 mg mL-1, and the Young’s modulus was 2.2 kPa when not specified).
Immunofluorescence
Cultured hMSCs were first fixed in 4% paraformaldehyde (Sigma, Germany) and then treated with 0.25% (v/v) Triton-X 100 (Sigma, Germany) in PBS buffer. Nonspecific antibody binding was blocked by incubating the samples with 1% (w/v) bovine serum albumin (BSA) in PBST (0.1% v/v Triton-X 100 in PBS buffer) before incubation with the primary antibody (1:250) diluted in PBST with 1% BSA. Samples were then incubated with the secondary antibody (1:500), Alexa 488 conjugated phalloidin (1:500, Thermo Fisher Scientific) to detect F-actin, and 4′,6-diamidino-2-phenylindole (DAPI, 1:500, KeyGen) to visualize the nucleus diluted in PBST with 1% BSA.
Analysis of polarity
For analysis of polarity, cells were divided into two parts by a line through the center of the nucleus perpendicular. The polarity was calculated from sum-projected images as the integrated density in one part of the cell divided by the integrated density in the other part of the cell. The data is presented as absolute values in a log2 scale (\(\left|{\log }_{2}\left({si}{de}1/{side}2\right)\right|\)).
Quantification of the Golgi apparatus distribution angle
For quantification of the Golgi apparatus distribution angle, the nuclear centroid was set as the origin of a polar coordinate system. The angular distribution was measured by calculating the angle occupied by Golgi apparatus-associated signals. The distribution angle was defined as the absolute angular difference between the maximum and minimum angles of the Golgi apparatus-positive regions relative to the nucleus origin, with 360° representing complete perinuclear encircling.
Antibodies
The primary antibodies used were Phospho-FAK/PTK2 pTyr397 Antibody (Novex, 700255, WB 1:1000, IF 1:200), Phospho-Myosin IIa (Ser1943) (D7Z7T) Rabbit mAb (Cell signaling Technology, 14611S, WB 1:1000, IF 1:200), Paxillin Mouse Polyclonal Antibody (BD bioscience, 610052, IF 1:200), Rabbit Monoclonal anti-Phospho-Myosin Light Chain 2 (Thr18/Ser19) (Cell Signaling Technology, 95777, WB 1:1000, IF 1:200), Talin-1 Polyclonal antibody (ProteinTech, 14168-1-AP, WB 1:1000, IF 1:200), Mouse Monoclonal anti-Integrin beta 1 (Abcam, ab24693, IF 1:200), Anti-Giantin antibody (Abcam, ab80864, IF 1:200), Rabbit Polyclonal anti-Myosin IIb (Cell Signaling Technology, 3404S, IF 1:200), Mouse Polyclonal anti-RAC1 (ProteinTech, 24072-1-AP, IF 1:200), Anti-beta Actin antibody (Abcam, ab8227, WB 1:1000). The secondary antibodies used were Alexa Fluor 568 anti-rabbit (Thermo Fisher Scientific, A-11011, IF 1:200), Alexa Fluor 647 anti-mouse (Thermo Fisher Scientific, A-32728, IF 1:200) and Goat Anti-Rabbit IgG H&L (HRP) (Abcam, ab205718, WB 1:2000).
Inhibitors
Rac1 Inhibitor NSC23766 was procured from Cayman (Cat# 13196). Latrunculin A was procured from Tocris Bioscience (Cat# 76343-93-6). Para-aminoblebbistatin was procured from MCE (Cat# HY-111474).
OE Talin cDNA transfection
Human OE Talin cDNA was procured from YouBio Biotech Corporation (Hunan, China). Cell transfections were conducted using Lipofectamine 3000 reagent from Invitrogen (Carlsbad, CA, USA).
Western blot
After hMSCs were cultured under different condition, cells were washed with PBS buffer and then lysed using lysis buffer (3 mM MgCl2, 100 mM Hepes-NaOH, 300 mM NaCl, 1 mM EDTA, 20% glycerol, 0.2% SDS, 1% sodium deoxycholate, 2% TX-100 (w/v), pH 7.5) containing phenylmethylsulfonyl fluoride (PMSF, 1 mM, Thermo Fisher Scientific). The remains were collected and frozen on ice for 30 minutes. The precipitate was removed by centrifugation (12,219 × g, 15 min). The supernatant was taken, and the total protein concentration of each sample was determined using the Bicinchoninic Acid (BCA) protein concentration determination kit (Beyotime). Proteins in each sample were separated by SDS-PAGE and then electrotransferred to a poly(1,1-difluoroethylene) (PVDF) membrane in transfer buffer (0.025 M Tris, 0.192 M glycine, 0.037% SDS, 20% methanol). Next, the PVDF membrane was blocked overnight at 4 °C with 5% nonfat dry milk (w/v) in TBST buffer (20 mM Tris, pH 7.6, 137 mM NaCl2, and 0.1% Tween-20 (v/v) and then incubated overnight at 4 °C with the primary antibodies (1:10000). After being washed 5 times with TBST buffer (10 min each), the blots were incubated with HRP-conjugated secondary antibodies (1:10000) for 1 h at 37 °C. Again, the blots were washed 5 times with TBST buffer (10 min each), and then enhanced chemiluminescence (ECL) Star (Beyotime) was used to visualize the protein bands. Finally, blots were exposed and photographed in a CLIN X chemical exposure apparatus. Uncropped blots with molecular weight markers are shown in Fig. S31.
Measurement of cell speed and MSD
Cell migration trajectories were systematically tracked through time-lapse imaging analysis using ImageJ software. Subsequently, the distance of cell movement over time was calculated following the kinematic definition:
where \(x\left(t\right)\) represents the position vector of the cell at time \(t\). Finally, the cell migration speed was determined by applying a linear regression fit to the displacement (\(s\left(t\right)\)) versus time (\(t\)), where the slope of the regression line corresponds to the cell’s average speed. Mean squared displacement was computed through ensemble averaging across \(N\) cell trajectories using the statistical mechanical framework:
where \({x}^{\left(i\right)}\left(t\right)\) represents the position vector of the \(i\)-th cell at time \(t\).
Measurement of cell actin fibers alignment
All imaging was performed on a Leica STELLARIS 5 (Germany), and examined under a 20× objective. The images were processed using the ImageJ plugin FibrilTool, which utilizes the nematic tensor concept to measure fiber alignment. This technique primarily relies on first-order derivatives, making it less sensitive to noise. To obtain quantitative data, circular statistics were applied to directional data to analyze the properties of tangent directions within the region of interest (ROI). The circular average of these tangent directions defines the average fiber orientation in the region, while the circular variance provides a score indicating the degree of fiber order, equivalent to computing the nematic tensor. In this scoring system, 0 represents disorder and 1 signifies complete order. For more details, please refer to the work of Arezki Boudaoud et al. (2014)61.
Measuring probability of cell detachment of hMSC
The probability of cell detachment was calculated using live-cell imaging. Cells were imaged every 10 minutes over a 24-hour period to monitor detachment events. The proportion of detached cells was determined by counting the number of cells that had detached from the substrate relative to the total number of cells observed at each time point. This process was repeated across multiple experimental replicates to ensure statistical reliability. The probability of cell detachment was then computed as the ratio of detached cells to the total number of cells over the 24-hour observation period.
Actin flow experiments
Live-cell imaging of actin dynamics was performed using SPY555-FastAct (Spirochrome), a cell-permeable F-actin probe. Cells were incubated with SPY555-FastAct in culture medium for 2 h at 37 °C following the manufacturer’s protocol (dilution 1:1000 in culture medium), followed by three washes with PBS to remove excess probe. Time-lapse imaging was conducted at 10-second intervals using a confocal laser scanning microscope equipped with a 100× oil immersion objective and environmental chamber (37 °C, 5% CO₂).
For actin flow quantification, image sequences were imported into ImageJ 1.54p (NIH). Kymographs were generated using the Reslice function along manually selected straight-line ROIs positioned in regions exhibiting actin flow. The resulting kymographs displayed spatial information (x-axis) versus temporal progression (y-axis, 10 s/line). Actin flow speed was calculated from the slope of linear streaks in kymographs by measuring the angle of deflection relative to the vertical axis.
Focal adhesion dynamics
Focal adhesion turnover was assessed via Talin-GFP fluorescence recovery after photobleaching (FRAP). Cells were transiently transfected with Talin-GFP plasmid (gift from Qiang Wei) using Lipofectamine 3000 (Thermo Fisher) and imaged 24 hours post-transfection. A defined region of interest within individual focal adhesions was photobleached using a 488 nm laser at 100% intensity. Post-bleach recovery was monitored using a 100× oil immersion objective on a confocal microscope. Recovery curves were normalized to pre-bleach intensity and fitted to a single-exponential model \(F\left(t\right)={F}_{0}\,+\,{F}_{1}\,\left(1-{e}^{-{kt}}\right)\) in software ImageJ to calculate half-recovery times \(({t}_{\frac{1}{2}}=\,{\mathrm{ln}}\,2/k)\) and mobile fractions \(({F}_{M}={F}_{1}/\,({F}_{0}\,+\,{F}_{1}))\).
Simulation
The study employs a motor-clutch model24 to investigate cellular spreading and migration dynamics. In this framework, myosin motors exert forces on actin filament bundles, inducing retrograde flow of actin towards the cell center. Consistent with the stochastic nature of integrin-ligand binding, clutches within the model exhibit stochastic binding and unbinding behaviors, characterized by association and dissociation rates denoted as \({r}_{{on}}\) and \({r}_{{off},i}\), respectively. The probability of a clutch \(i\) remaining engaged \({P}_{b,i}\) is expressed as \({P}_{b,i}=\frac{{r}_{{on}}}{{r}_{{on}}+{r}_{{off},i}}\). The lifetime of each integrin-ligand bond is defined as the time elapsed between clutch-substrate binding and unbinding. As F-actin undergoes retrograde motion, each bound clutch (treated as a linear spring with stiffness \({k}_{c}\)) experiences stretching, generating a resisting force \({F}_{c,i}\). This force can be represented as \({F}_{c,i}={k}_{c}({x}_{c,i}-{x}_{s})\), where \({x}_{c,i}\) and \({x}_{s}\) denote displacements of the filament end and substrate end of the clutch, respectively. Utilizing a Bell model62, the dissociation rate \({r}_{{off},i}\) is anticipated to exponentially increase with \({F}_{c,i}\), described by \({r}_{{off},i}={r}_{{off}}^{0}{e}^{\frac{{F}_{c,i}}{{F}_{b}}}\), where \({F}_{b}\) represents a characteristic force, and \({r}_{{off}}^{0}\) signifies the clutch breaking rate in the absence of any force. Recent experimental evidence highlights the growth of FAs on stiff substrates through integrin recruitment to adhesion complexes via talin unfolding25,63. Based on these findings and the results obtained in this study, the model predicts a progressive increase in the number of integrins \({n}_{c}\) over time with a speed of \({k}_{e}\). Building upon the model proposed by Elosegui-Artola et al.63 for FA reinforcement, the association rate \({r}_{{on}}\) is modified to \({r}_{{on}}^{0}(1+\alpha ({F}_{a}-{F}_{{cr}}))\) once the average clutch force \({F}_{a}\) surpasses a force threshold \({F}_{{cr}}\). A value of 0.2 per piconewton is selected for \(\alpha\) based on previous study41. The substrate force \({F}_{s}\) can be related to the actin flow speed, \({V}_{f}\), through the Hill relation2:
where \({N}_{M}\) is the number of pMyosin IIa motors; \({v}_{\max }\) is the maximum actin flow speed; and \({F}_{M}\) is the characteristic stalling force of a single pMyosin IIa motor. Consequently, the force loading rate and actin flow speed are enhanced by pMyosin IIa accumulation.
To contextualize this analysis within cell spreading phenomena, it is pertinent to consider lamellipodia, flat sheet-like structures propelled by actin polymerization-induced protrusive forces against the cell membrane. During spreading, actin monomers migrate from the cell center to the periphery to sustain continuous polymerization, countering the retrograde flow of F-actin. Consequently, the cell spreading speed \({V}_{s}\) can be expressed as \({V}_{s}={V}_{p}-{V}_{f}\), where \({V}_{p}\) denotes polymerization velocity and \({V}_{f}\) signifies retrograde flow velocity. As cell spreading progresses, the deformed membrane and cytoskeleton resist further deformation, exerting increasing forces on binding integrins. Treating the cytoskeleton/membrane as a viscoelastic material with height \(h\), the resistance force \({f}_{r}\) can be calculated as \({f}_{r}=h({k}_{m}{\epsilon }_{r}+{\eta }_{m}\frac{d{\varepsilon }_{r}}{{dt}})\)24,64, where \({k}_{m}\) and \({\eta }_{m}\) represent the effective stiffness and viscosity of the cytoskeleton, respectively. Here, \({\epsilon }_{r}=\frac{R\left(t\right)-{R}_{0}}{{R}_{0}}\) denotes the mean radial strain in the deformed cytoskeleton, with \(R\left(t\right)\) representing the spreading radius at time \(t\). As the force between integrins and the substrate \({f}_{s}={f}_{r}+\sum {F}_{c,i}\) increases, integrins on one side of the cell detach from the substrate, prompting cell migration towards the opposite side.
Cell migration is influenced by two key factors. On one hand, it is determined by the interplay between the polymerization speed of actin and the retrograde flow speed, which together affect the rate of cell protrusion. On the other hand, the probability of cell translocation due to adhesion site rupture is calculated based on the balance of internal and external forces acting on integrins at each time point. Both aspects collectively contribute to the overall migration behavior.
To gauge the robustness of the model to changes in parameters, one-at-a-time parameter variations were performed when holding every other parameter fixed at their assumed values. Parameters were varied over 0.1- and 10-fold of their assumed values, and the model was found to be robust and tolerant of significant changes. For some parameters, the allowable range did not allow for extreme fold changes, so in these cases, the maximum or minimum fold change of the parameter was used to calculate the sensitivity instead.
The predictions of our computational model are inherently stochastic, with variability arising primarily from the probabilistic nature of the simulated ECM-integrin connections. In the main text, for figures where simulation results are plotted alongside experimental data, we have in some cases presented only the mean of the simulation. This was a deliberate choice to avoid visual clutter that would arise from overlapping error bands (i.e., the standard deviation from the experiment and the standard deviation from the simulation), which can hinder the readability and interpretation of the data. To provide a full and transparent characterization of the model’s predicted variability for these specific cases, we present the unabridged data here in Fig. S32. For each relevant simulation, the data is shown as the ensemble average (solid line) surrounded by a shaded region representing one standard deviation (mean values +/- SD).
Traction force microscopy
Traction force microscopy experiments were performed as described previously29. The images of fluorescent beads were analyzed using a previously reported program based on Python 3.8. The images with and without cells were first aligned using the phase_cross_correlation function provided by the scikit-image module. After alignment, the overlapping regions were cropped, and the deformation fields were calculated using an iterative cross-correlation-based particle image velocimetry (PIV) method provided by the OpenPIV module. In this method, the images were divided into smaller regions, known as interrogation windows, and the cross-correlation between the corresponding windows was used to measure deformation. The calculation was repeated while progressively decreasing the size of the interrogation windows to achieve better PIV resolution.
Once the deformation field was obtained, outliers were filtered and replaced using a weighted average. The problem of calculating the traction field from the deformation field was then converted into a simple matrix equation using a Green’s tensor \(M\):
where \({{\rm{u}}}\) represents the deformation field and \({{\rm{f}}}\) represents the traction field. Since solving the matrix \(M\) in real space is generally challenging, a widely adopted method is to solve it in Fourier space:
where \(\widetilde{u}\), \(\widetilde{M}\), and \(\widetilde{f}\) are the Fourier transforms of the deformation field, the Green’s tensor, and the traction field, respectively. This equation can be solved analytically, allowing for the calculation of traction fields directly in Fourier space. The inverse Fourier transform is then applied to obtain the traction fields in real space.
The TFM output is presented as force per unit area in Pascals (Pa), offering a more accurate depiction of the spatial distribution of cellular traction forces. This approach is known as the Fourier Transform Traction Cytometry (FTTC) method, which was employed in our study.
In our study, different Young’s modulus values were used for the dark and light conditions to calculate the traction forces, as determined from prior experiments (Fig. S3). Additionally, to ensure the reliability of the experimental data, bead images obtained under the respective conditions (without cells) were used as controls during the calculation of traction forces.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data used to generate the main and Supplementary figures are provided in the Source Data file. Any additional information required to reanalyze the data, including the original high-resolution immunofluorescence images, have been uploaded to GitHub for anonymous access (https://github.com/yuyousama/cellMigrationData), and are publicly available as of the date of publication. A permanent reference65 (https://doi.org/10.5281/zenodo.16742360) for downloading the data has been provided. Source data are provided with this paper.
Code availability
The original code for the simulation model has been uploaded to GitHub for anonymous access (https://github.com/yuyousama/cellMigrationModel) and is publicly available as of the publication date. We have provided a permanent reference66 (https://doi.org/10.5281/zenodo.16742356) for the specific code version used in this study.
References
Ridley, A. J. et al. Cell migration: integrating signals from front to back. Science 302, 1704–1709 (2003).
Lauffenburger, D. A. & Horwitz, A. F. Cell migration: A physically integrated molecular process. Cell 84, 359–369 (1996).
Trepat, X., Chen, Z. & Jacobson, K. Cell migration. Compr. Physiol. 2, 2369–2392 (2012).
van Helvert, S., Storm, C. & Friedl, P. Mechanoreciprocity in cell migration. Nat. Cell Biol. 20, 8–20 (2018).
Charras, G. & Sahai, E. Physical influences of the extracellular environment on cell migration. Nat. Rev. Mol. Cell Biol. 15, 813–824 (2014).
Schwarzbauer, J. E. Cell migration: May the force be with you. Curr. Biol. 7, R292–R294 (1997).
Kechagia, J. Z., Ivaska, J. & Roca-Cusachs, P. Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 20, 457–473 (2019).
Kim, D. H. & Wirtz, D. Predicting how cells spread and migrate: focal adhesion size does matter. Cell Adh Migr. 7, 293–296 (2013).
Seetharaman, S. & Etienne-Manneville, S. Integrin diversity brings specificity in mechanotransduction. Biol. Cell 110, 49–64 (2018).
Reversat, A. et al. Cellular locomotion using environmental topography. Nature 582, 582–585 (2020).
Wang, W. Y., Davidson, C. D., Lin, D. & Baker, B. M. Actomyosin contractility-dependent matrix stretch and recoil induces rapid cell migration. Nat. Commun. 10, 1186 (2019).
Wisdom, K. M. et al. Matrix mechanical plasticity regulates cancer cell migration through confining microenvironments. Nat. Commun. 9, 4144 (2018).
Bangasser, B. L. et al. Shifting the optimal stiffness for cell migration. Nat. Commun. 8, 15313 (2017).
Adebowale, K. et al. Enhanced substrate stress relaxation promotes filopodia-mediated cell migration. Nat. Mater. 20, 1290–1299 (2021).
Oakes, P. W. et al. Lamellipodium is a myosin-independent mechanosensor. Proc. Natl Acad. Sci. USA 115, 2646–2651 (2018).
Lo, C. M., Wang, H. B., Dembo, M. & Wang, Y. L. Cell movement is guided by the rigidity of the substrate. Biophys. J. 79, 144–152 (2000).
Shafqat-Abbasi, H. et al. An analysis toolbox to explore mesenchymal migration heterogeneity reveals adaptive switching between distinct modes. Elife 5, e11384 (2016).
Barriga, E. H., Franze, K., Charras, G. & Mayor, R. Tissue stiffening coordinates morphogenesis by triggering collective cell migration in vivo. Nature 554, 523–527 (2018).
Zhu, M. et al. Spatial mapping of tissue properties in vivo reveals a 3D stiffness gradient in the mouse limb bud. Proc. Natl Acad. Sci. USA 117, 4781–4791 (2020).
DuChez, B., Doyle, A., Dimitriadis, E. & Yamada, K. Durotaxis by human cancer cells. Biophys. J. 116. (2019) https://doi.org/10.1016/j.bpj.2019.01.009.
McKenzie, A. J. et al. The mechanical microenvironment regulates ovarian cancer cell morphology, migration, and spheroid disaggregation. Sci. Rep. 8, 7228 (2018).
Yeoman, B. et al. Adhesion strength and contractility enable metastatic cells to become adurotactic. Cell Rep. 34, 108816 (2021).
Bangasser, B. L., Rosenfeld, S. S. & Odde, D. J. Determinants of maximal force transmission in a motor-clutch model of cell traction in a compliant microenvironment. Biophys. J. 105, 581–592 (2013).
Chan, C. E. & Odde, D. J. Traction dynamics of filopodia on compliant substrates. Science 322, 1687–1691 (2008).
Elosegui-Artola, A. et al. Mechanical regulation of a molecular clutch defines force transmission and transduction in response to matrix rigidity. Nat. Cell Biol. 18, 540–548 (2016).
Pathak, A. & Kumar, S. Independent regulation of tumor cell migration by matrix stiffness and confinement. Proc. Natl Acad. Sci. USA 109, 10334–10339 (2012).
Chaudhuri, O., Cooper-White, J., Janmey, P. A., Mooney, D. J. & Shenoy, V. B. Effects of extracellular matrix viscoelasticity on cellular behaviour. Nature 584, 535–546 (2020).
Liu, K., Wiendels, M., Yuan, H., Ruan, C. & Kouwer, P. H. J. Cell-matrix reciprocity in 3D culture models with nonlinear elasticity. Bioact. Mater. 9, 316–331 (2022).
Yang, J. et al. Photo-tunable hydrogels reveal cellular sensing of rapid rigidity changes through the accumulation of mechanical signaling molecules. Cell Stem Cell https://doi.org/10.1016/j.stem.2024.09.016.
Zhao, J. M. et al. Single-molecule detection of structural changes during Per-Arnt-Sim (PAS) domain activation. Proc. Natl Acad. Sci. Usa. 103, 11561–11566 (2006).
Gorelik, R. & Gautreau, A. Quantitative and unbiased analysis of directional persistence in cell migration. Nat. Protoc. 9, 1931–1943 (2014).
Lavrsen, K. et al. Microtubule detyrosination drives symmetry breaking to polarize cells for directed cell migration. Proc. Natl Acad. Sci. USA 120, e2300322120 (2023).
Vicente-Manzanares, M., Koach, M. A., Whitmore, L., Lamers, M. L. & Horwitz, A. F. Segregation and activation of myosin IIB creates a rear in migrating cells. J. Cell Biol. 183, 543–554 (2008).
Chung, C. Y., Lee, S., Briscoe, C., Ellsworth, C. & Firtel, R. A. Role of Rac in controlling the actin cytoskeleton and chemotaxis in motile cells. Proc. Natl Acad. Sci. USA 97, 5225–5230 (2000).
Fujiwara, I., Zweifel, M. E., Courtemanche, N. & Pollard, T. D. Latrunculin A accelerates actin filament depolymerization in addition to sequestering actin monomers. Curr. Biol. 28, 3183–3192.e3182 (2018).
Ayscough, K. Use of latrunculin-A, an actin monomer-binding drug. Methods Enzymol. 298, 18–25 (1998).
Morita, R., Shigeta, Y. & Harada, R. Latrunculin resistance mechanism of non-conventional actin NAP1 uncovered by molecular dynamics simulations. Cytoskeleton (Hoboken) 81, 143–150 (2024).
Liu, J. et al. Talin determines the nanoscale architecture of focal adhesions. Proc. Natl Acad. Sci. USA 112, E4864–E4873 (2015).
Várkuti, B. H. et al. A highly soluble, non-phototoxic, non-fluorescent blebbistatin derivative. Sci. Rep. 6, 26141 (2016).
Cheng, B. et al. Nanoscale integrin cluster dynamics controls cellular mechanosensing via FAKY397 phosphorylation. Sci. Adv. 6, eaax1909 (2020).
Gong, Z. et al. Matching material and cellular timescales maximizes cell spreading on viscoelastic substrates. Proc. Natl Acad. Sci. USA 115, E2686–e2695 (2018).
Zhang, Z. et al. Programmable and reversible integrin-mediated cell adhesion reveals hysteresis in actin kinetics that alters subsequent mechanotransduction. Adv. Sci. (Weinh.) 10, e2302421 (2023).
Panciera, T., Azzolin, L., Cordenonsi, M. & Piccolo, S. Mechanobiology of YAP and TAZ in physiology and disease. Nat. Rev. Mol. Cell Biol. 18, 758–770 (2017).
Dasgupta, I. & McCollum, D. Control of cellular responses to mechanical cues through YAP/TAZ regulation. J. Biol. Chem. 294, 17693–17706 (2019).
Ladoux, B. & Mège, R. M. Mechanobiology of collective cell behaviours. Nat. Rev. Mol. Cell Biol. 18, 743–757 (2017).
Stutchbury, B., Atherton, P., Tsang, R., Wang, D. Y. & Ballestrem, C. Distinct focal adhesion protein modules control different aspects of mechanotransduction. J. Cell Sci. 130, 1612–1624 (2017).
Le Coq, J., Acebrón, I., Rodrigo Martin, B., López Navajas, P. & Lietha, D. New insights into FAK structure and function in focal adhesions. J. Cell Sci. 135 https://doi.org/10.1242/jcs.259089 (2022).
Seong, J. et al. Detection of focal adhesion kinase activation at membrane microdomains by fluorescence resonance energy transfer. Nat. Commun. 2, 406 (2011).
Alfieri, R., Giovannetti, E., Bonelli, M. & Cavazzoni, A. New treatment opportunities in phosphatase and tensin homolog (PTEN)-deficient tumors: Focus on PTEN/focal adhesion kinase pathway. Front. Oncol. 7, 8 (2017).
Chetram, M. A. & Hinton, C. V. PTEN regulation of ERK1/2 signaling in cancer. J. Receptors Signal Transduct. 32, 190–195 (2012).
Zheng, Y. et al. FAK Phosphorylation by ERK Primes Ras-induced tyrosine dephosphorylation of FAK mediated by PIN1 and PTP-PEST. Mol. Cell 35, 11–25 (2009).
Crnko, S., Du Pré, B. C., Sluijter, J. P. G. & Van Laake, L. W. Circadian rhythms and the molecular clock in cardiovascular biology and disease. Nat. Rev. Cardiol. 16, 437–447 (2019).
Smith, J. C. Respiratory rhythm and pattern generation: Brainstem cellular and circuit mechanisms. Handb. Clin. Neurol. 188, 1–35 (2022).
Ahmadi, A., Dabidi Roshan, V. & Jalali, A. Coronary vasomotion and exercise-induced adaptations in coronary artery disease patients: A systematic review and meta-analysis. J. Res Med Sci. 25, 76 (2020).
Barbara, G. et al. The intestinal microenvironment and functional gastrointestinal disorders. Gastroenterology 150, 1305–1318.e1308 (2016).
Shiraishi, K. et al. Biophysical forces mediated by respiration maintain lung alveolar epithelial cell fate. Cell 186, 1478–1492.e1415 (2023).
Ruprecht, V. et al. Cortical contractility triggers a stochastic switch to fast amoeboid cell motility. Cell 160, 673–685 (2015).
Venturini, V. et al. The nucleus measures shape changes for cellular proprioception to control dynamic cell behavior. Science 370 https://doi.org/10.1126/science.aba2644 (2020).
Lomakin, A. J. et al. The nucleus acts as a ruler tailoring cell responses to spatial constraints. Science 370 https://doi.org/10.1126/science.aba2894 (2020).
Wu, X. et al. Reversible hydrogels with tunable mechanical properties for optically controlling cell migration. Nano Res. 11, 5556–5565 (2018).
Boudaoud, A. et al. FibrilTool, an ImageJ plug-in to quantify fibrillar structures in raw microscopy images. Nat. Protoc. 9, 457–463 (2014).
Bell, G. I. Models for the specific adhesion of cells to cells. Science 200, 618–627 (1978).
Elosegui-Artola, A. et al. Rigidity sensing and adaptation through regulation of integrin types. Nat. Mater. 13, 631–637 (2014).
Bangasser, B. L. & Odde, D. J. Master equation-based analysis of a motor-clutch model for cell traction force. Cell Mol. Bioeng. 6, 449–459 (2013).
Yu, Z. & Jiapeng, Y. Dynamic rigidity changes enable rapid cell migration on soft substrates: Cell Migration data. https://doi.org/10.5281/zenodo.16742360 (2025).
Yu, Z. J., Yang. Dynamic Rigidity changes enable rapid cell migration on soft substrates: Cell migration model (v1.1). https://doi.org/10.5281/zenodo.16742356 (2025).
Acknowledgements
This work is supported mainly by the National Key R&D Program of China (Grant No. 2024YFA0919300 to Yi.C.), the National Natural Science Foundation of China (No. T2225016 to Yi.C., T2222020 to Q.W., T2322010 to B.X. and 11934008 to Yi.C.), Shandong Provincial Natural Science Foundation (grant Nos. ZR2024QC217 to J.Y.), Shandong Provincial Laboratory Project (grant Nos. SYS202202 to C.J.), Research Project of Jinan Microecological Biomedicine Shandong Laboratory (grant JNL-2023004Q to J.Y., 2025008B to X.G., 2025009B to C.J. and 2025010B to Yi.C.), and the Natural Science Foundation of Jiangsu Province (No. BK20220120 to B.X.).
Author information
Authors and Affiliations
Contributions
Investigation: J.Y., Y.Z., S.W., L.D., Yi.C., Conceptualization: J.Y., Y.Z., Q.W., Yi.C., Methodology: J.Y., Y.Z., S.W., Q.W., Yi.C., Formal Analysis: J.Y., Y.Z., S.W., Software: J.Y., Y.Z., Writing – original draft: J.Y., Writing – review and editing: Y.Z., S.W., P.W., L.D., L.L., Yu.C., X.H., B.X., W.W., C.J., X.G., Q.W., Yi.C., Data curation: J.Y., Y.Z., S.W., Visualization: J.Y., Y.Z., S.W., L.L., Funding acquisition: J.Y., B.X., C.J., X.G., Q.W., Yi.C., Project administration: C.J., Q.W., Yi.C. Supervision: B.X., W.W., C.J., X.G., Q.W., Yi.C., Resources: B.X., W.W., C.J., X.G., Q.W., Yi.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Manuel Salmeron-Sanchez, who co-reviewed with Giuseppe Ciccone, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, J., Zhang, Y., Wang, S. et al. Dynamic rigidity changes enable rapid cell migration on soft substrates. Nat Commun 16, 8793 (2025). https://doi.org/10.1038/s41467-025-63854-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63854-9
This article is cited by
-
A Phenotype-Structured PDE Framework for Investigating the Role of Hypoxic Memory on Tumor Invasion under Cyclic Hypoxia
Bulletin of Mathematical Biology (2026)