Abstract
Lipid nanoparticles (LNPs) are the most clinically relevant vehicles for mRNA vaccines. Despite the great successes, the toxicity caused by the high dose of lipid components still represents a great challenge. The suboptimal loading capacity of mRNA in LNPs not only compromises the vaccine’s efficacy but also heightens the risk of non-specific immune responses, accelerates clearance caused by anti-PEG IgG/IgM. These problems underscore the urgent need for improving mRNA loading capacity in LNPs to provide dose-sparing effects. Herein, we develop a metal ion mediated mRNA enrichment strategy to efficiently form a high-density mRNA core, and manganese ion (Mn2+) exhibits a unique capability to match the need. The prepared Mn-mRNA nanoparticle is subsequently coated with lipids to form the resulting nanosystem, L@Mn-mRNA, which achieved nearly twice the mRNA loading capacity compared to conventional mRNA vaccine formulations (LNP-mRNA). Remarkably, L@Mn-mRNA also demonstrates a 2-fold increase in cellular uptake efficiency compared to LNP-mRNA, attributed to the enhanced stiffness provided by the Mn-mRNA core. By combining improved mRNA loading with superior cellular uptake, L@Mn-mRNA achieves significantly enhanced antigen-specific immune responses and therapeutic efficacy as vaccines. We elucidate the mechanism behind Mn-mRNA construction and optimize the L@Mn-mRNA formulations, and this method is suitable for types of lipids and mRNAs. Moreover, L@Mn-mRNA also reduces the risk of anti-PEG IgG/IgM generation. Thus, this strategy holds significant potential as a platform for the next generation of lipid-based mRNA vaccines.
Similar content being viewed by others
Introduction
In the past infectious disease pandemic, the mRNA vaccine stepped onto the stage of combating diseases1,2,3,4. With the emergence of organ targeted mRNA delivery systems5,6,7,8,9,10 and mRNA tumor vaccines11,12,13, mRNA therapeutics are sprinting into a new era14,15. Despite the great successes, the mRNA loading capacity in the vaccines still represents a great challenge. For example, the mRNA component was less than 4% in weight in COVID-19 BNT162b2 vaccine developed by Pfizer Inc. and BioNTech SE, and less than 5% in mRNA-1273 COVID-19 vaccine developed by Moderna, Inc.16,17. In this case, to achieve the effective mRNA dose, the lipid dose will be relatively high. The toxicity and/or non-specific immune responses caused by the high dose of lipid components becomes a major concern of mRNA vaccines18,19,20,21, such as high rate of headache (62.8% of individuals aged 18–64) and fever (17.4% of individuals aged 18–64), according to FDA data on mRNA-1273 COVID-19 vaccine22. Thus, besides of screening novel phospholipids23,24,25,26 and optimizing the mRNA sequences27,28, how to efficiently improve the mRNA loading capacity in LNP systems is also crucial and challenging for mRNA vaccines (and other mRNA therapeutics)29,30.
Metal ions have been reported to assemble with short nucleic acids to form nanoparticles with high- nucleic acid loading content31,32,33, which inspired us to apply this strategy in enriching mRNA in vaccines. As the proposed procedures shown in Fig. 1A, we envision that, if the condensed metal ions-mRNA nanoparticle (M-mRNA) is coated with lipids, the resulted nanoparticle (L@M-mRNA) would have a high-density mRNA core, which would achieve higher mRNA loading than the conventional LNP-mRNA complexes. However, the mRNA activity maintenance and the assembly efficiency are the crucial techniques.
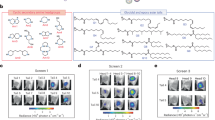
A Proposed procedures for increasing mRNA loading capacity through metal ion mediated mRNA enrichment. B Integrity evaluation of EGFP and Luciferase mRNA at 95 °C for different duration analyzed by agarose gel electrophoresis. C EGFP mRNA and Luciferase mRNA expression evaluation after being heated at 95 °C. Each group of mRNAs was transfected to DC 2.4 cells using LipofectamineTM 3000 Transfection Reagent. Relative EGFP expression was analyzed by flow cytometry. Relative luciferase expression was analyzed by microplate reader after incubated with luciferin. Data were presented as mean ± SD, n = 3 biologically independent samples. D Integrity evaluation of EGFP and Luciferase mRNA at 65 °C for different duration analyzed by agarose gel electrophoresis. E EGFP mRNA and Luciferase mRNA expression evaluation after being heated at 65 °C. Procedures were same with (C). Data were presented as mean ± SD, n = 3 biologically independent samples. F TEM images of Fe-mRNA, Cu-mRNA, Zn-mRNA, and Mn-mRNA heating at 65 °C for 5 min. The molar ratio of metal ions to bases (in mRNA) was 5:1. Scale bar, 100 nm. Luciferase mRNA (mLuc) was used in this experiment. G mRNA loading efficiency in Zn-mRNA and Mn-mRNA calculated by Quant-itTM RiboGreen RNA Assay Kit. Data were presented as mean ± SD, n = 4 technical replicates. Statistical significance of (C and E) was analyzed by one-way ANOVA with Tukey’s correction. Figure 1A was created in BioRender. Han, D. (2024) BioRender.com/AX272J1C1E. Source data are provided as a Source Data file.
Herein, we developed a highly efficient mRNA enrichment strategy to improve the mRNA loading capacity in lipid-based vaccine. In brief, we explored several commonly used metal ions (Fe2+, Cu2+, Zn2+, and Mn2+) to prepare the M-mRNA complexes, and found that Mn2+ could enrich mRNA (termed Mn-mRNA nanoparticles) in high efficiency without influencing the mRNA activity. Mn-mRNA was subsequently coated with lipids, leading to nanosystems (L@Mn-mRNA) with an almost twofold mRNA loading capacity compared to conventional mRNA vaccine formulations (LNP-mRNA). Remarkably, L@Mn-mRNA achieved two times increase in cellular uptake efficiency compared to LNP-mRNA, attributed to the enhanced stiffness provided by the Mn-mRNA core. Therefore, L@Mn-mRNA achieved significantly improved antigen-specific immune responses and therapeutic efficacy as vaccines. We elucidated the mechanism of Mn-mRNA construction and optimization of the L@Mn-mRNA formulations to establish a potential platform for the lipid-based mRNA vaccine.
Results and discussion
Design and screening of high-mRNA-loading nanoparticles
To increase the mRNA loading, we tried to construct a metal ion-mRNA (M-mRNA) nanoparticle as the core to enrich mRNA before lipids coating (Fig. 1A). The reported nanoscale complexes made of metal ions and short nucleic acids were typically prepared at 95 °C for over 30 min31,32,33. However, this harsh preparation condition may result in the degradation of mRNA. As shown in agarose gel electrophoresis results, EGFP mRNA (mEGFP), Luciferase mRNA (mLuc), and COVID-19 Spike protein mRNA (mSpike) degraded within 30 min at 95 °C (Fig. 1B and Supplementary Fig. 1), and the expression efficiency of mEGFP and mLuc in DC 2.4 cells (DCs) (transfected by LipofectamineTM 3000) was significantly reduced after heating (Fig. 1C). These results suggested that lower temperature and shorter incubation duration should be more favorable for enriching mRNA with metal ions. Therefore, we lowered the heating temperature to 65 °C, in which condition, the mRNA degradation and activity loss was significantly reduced, as demonstrated by the agarose gel (Fig. 1D and Supplementary Fig. 1) and DCs transfection experiments (Fig. 1E). Notably, over 95% of mRNAs (mEGFP, mLuc) maintained their activity within 5 min at 65 °C. Thus, we explored the potential for enriching mRNA under this condition.
Various metal ions were tested to assemble with mRNA at 65 °C. Among the Fe-mRNA, Cu-mRNA, Zn-mRNA, and Mn-mRNA complexes, only Mn2+ and Zn2+ groups could generate regular nanostructures (Fig. 1F). Then, we quantified the mRNA coordination ratio of Zn-mRNA and Mn-mRNA nanoparticles using a Quant-itTM RiboGreen RNA Assay Kit34,35. The results revealed that about 90% of mRNA formed nanoparticles in Mn-mRNA group within 5 min, whereas Zn-mRNA group required 15–30 min to reach similar efficiency (Fig. 1G). Considering the risk of mRNA degradation for prolonged treatment, we decided to use Mn2+ to enrich mRNAs at 65 °C for 5 min.
To investigate the mechanism of metal ions-mRNA complexes formation, we calculated the coordination bond lengths (l) and binding energy (ΔE) of metal ion with the adenine (A) and guanine (G) bases, which can in some extent reflect the stability of M-mRNA complex36,37. The selected sites were where transition metal ions most readily bind with bases N7/O6 (for G) and N7 (for A)36,37. As shown in Fig. 2A, Mn2+ and Zn2+ groups have longer bond lengths (no matter the l1, l2, and l3) than that of Fe2+ and Cu2+ groups, and Mn2+ have the lowest binding energy with bases compared with other metal ions. Shorter bond length and higher binding energy represents easier generation of stable M-mRNA complexes, which seems inconsistent with above results (that Fe2+ and Cu2+ groups did not form nanoparticles). We supposed that the formation of M-mRNA nanoparticles involved two steps: (1) binding, metal ions coordination with mRNA bases at room temperature, potentially forming complexes without regular morphologies; (2) rearrangement, the heating provided energy to break the metal ion-base coordination, leading to the reassembly of complexes and form nanoparticles (Fig. 2B). Thus, longer bond lengths (Mn2+ and Zn2+ with bases) and lower binding energy (Mn2+ with bases) indicated that lower temperature could realize the reassembly. To test our hypothesis, we monitored the heating process over 5 min and analyzed the spatial distribution of Mn element and P element (from mRNA) by using elemental analysis (EDS) mode of transmission electron microscopy (TEM). As shown in Fig. 2C, without heating, the reactants showed some aggregation (containing both Mn and P elements) and could not be collected by centrifugation, indicating that interactions between Mn and mRNA were already present. After 1 min of heating, no significant morphological change was observed, while particle morphology emerged at 3 min time point, though the structures was irregular. After 5 min heating, well-defined spherical nanoparticles were obtained. This observation supported our proposed mechanism of the M-mRNA formation (as shown in Fig. 2B), and the proposed coordination between the bases of mRNA and Mn2+ in the Mn-mRNA nanoparticles was shown in a 3D structure (Supplementary Fig. 2).
A The coordination bond lengths (l) and binding energy (ΔE) between metal ions and adenine (A(N7)) or guanine (G(N7/O6)). ΔE1 refers to the binding energy between the metal ions and the adenine, ΔE2 refers to the binding energy between the metal ions and the guanine. B Schematic illustration of the mechanism for Mn-mRNA nanoparticles formation after heating the system at 65 °C for 5 min. The heating provided energy for the rearrangement of Mn2+ and mRNA molecules. C The distribution of manganese and phosphorus elements of Mn-mRNA complexes before and after the heating at 65 °C for different time (1, 3, and 5 min) detected by Energy-Dispersive X-ray Spectroscopy (EDS) element analysis. Scale bar in first and second row, 500 nm; scale bar in third and fourth row, 100 nm. mLuc was used in this experiment. D Morphologies of Mn2+ and mRNA complexes with different molar ratios of Mn2+ to mRNA. Scale bar in 20:1 and 10:1 group, 500 nm, scale bar in other groups, 100 nm. mLuc was used in this experiment. E Mn-mRNA nanoparticles prepared at the same Mn2+ to mRNA molar ratio (5:1) at 65 °C for 5 min with different length mRNAs. Scale bar, 100 nm. Three times each experiment of (C–E) was repeated independently with similar results. Figure 2B was created in BioRender. Han, D. (2024) BioRender.com/ FL272J1BX9. Source data are provided as a Source Data file.
To investigate the relationship of Mn2+ and mRNA bases ratios in the Mn-mRNA formation, we tried to prepare the Mn-mRNA by using various molar ratios of these two components. Combining TEM images and dynamic light scattering (DLS) analysis, the system tended to generate uniform nanoparticles when the usage range of Mn2+ to mRNA base was controlled at 8:1 to 2:1 (Fig. 2D, Supplementary Fig. 3, and Supplementary Table 1), which was consistent with previous reports31,32. Neither increasing nor decreasing Mn2+ ratio (e.g., Mn2+ to bases molar ratio of 20:1 or 1:1) was benefit for the nanoparticle formation. Insufficient Mn2+ probably cannot drive coordination with mRNA, while excessive Mn2+ may lead to aggregations among particles (Supplementary Fig. 4). Considering that the 5:1 ratio group had lower PDI, we used this input ratio in the following investigations. In addition, the integrity of mRNA in Mn-mRNA was verified by agarose gel electrophoresis analysis, which demonstrated no detectable mRNA degradation (fragmentation) in the Mn-mOVA lane (Supplementary Fig. 5, as red dashed rectangle shown).
To evaluate the availability of Mn2+ and mRNA during the preparation of Mn-mRNA nanoparticles, we quantitatively analyzed the percentage of input Mn2+ in Mn-mRNA nanoparticles by using inductively coupled plasma mass spectrometry (ICP-MS). The result showed that 5% of the input Mn2+ involved the formation of Mn-mRNA. Notably, over 88% of the input mRNA had been involved in the formation of Mn-mRNA detected by Quant-itTM RiboGreen RNA Assay Kit, indicating a high efficiency in mRNA enrichment. Based on the above results, we calculated the weight percentage of Mn in Mn-mRNA nanoparticles was 4.4%, and the mRNA was 95.6% in weight (the detailed calculation process and results were presented on “Method” Section and Supplementary Table 2).
We respectively prepared Mn-mRNA with mSpike (4k nt), mLuc (1.9k nt), mOVA (1.6k nt), and mEGFP (1k nt) to verify the universality of the preparation conditions (65 °C heating for 5 min with 5:1 molar ratio of input Mn2+ to bases), though the particle size of Mn-mRNA had a decreasing trend with the mRNA length increasing. All groups could form regular nanoparticles (Fig. 2E and Supplementary Table 3), indicative of the universality of this approach.
Subsequently, we coated the Mn-mRNA with lipids (Fig. 3A) and explored the mass ratio of lipids (comprising four components) to the Mn-mRNA (mRNA mass instead of Mn-mRNA mass) ranging from 2.5:1 to 20:1. Upon lipids coating, the zeta potential of the particles increased and reached a positive value at a 10:1 ratio (15.5 ± 2.4 mV) (Fig. 3B). Further increasing the lipids ratio (e.g., 20:1) did not result in the further increase in zeta potential (13.1 ± 0.2 mV), while the polydispersity index (PDI) increased at ratios greater than 10:1 (Fig. 3B). In addition, the mRNA expression efficiency in DCs was enhanced with the increase of lipids usage till the ratio of lipids to Mn-mRNA reached 10:1 (Fig. 3C). Further increasing the lipids ratio did not significantly improve the transfection efficiency (relative mLuc expression: 1 ± 0.13 vs. 1 ± 0.09 for 10:1 vs. 20:1 of lipids to Mn-mRNA ratio). Therefore, we considered that the 10:1 ratio of lipids to Mn-mRNA could perform the sufficient coating and the mRNA encapsulation efficiency in this step was 98% (Supplementary Fig. 6). The mass ratio of Mn to mRNA in L@Mn-mRNA was consistent with that of Mn-mRNA nanoparticles (Supplementary Fig. 6). In which case, this formulation achieved a nearly 2-fold increase (9.1% vs. 4.7%) in mRNA loading capacity compared to current commercial products (Moderna mRNA-1273 COVID-19 vaccine) (Supplementary Table 4)38.
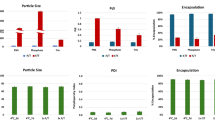
A Schematic illustration of L@Mn-mRNA preparation. B Size, PDI, and zeta potential of L@Mn-mRNA with different mass ratios of lipids to mRNA. Data were presented as mean ± SD, n = 3 technical replicates. C Relatively bioluminescence intensity evaluation in DC 2.4 cells after incubation with different groups (different mass ratios of lipids to mRNA) of L@Mn-mRNA for 24 h. Data were presented as mean ± SD, n = 4 biologically independent samples. P = 0.9999. Statistical significance was analyzed by two-tailed t-test. D Cryo-EM images of L@Mn-mRNA. mSpike, mLuc, mOVA, and mEGFP were respectively used for L@Mn-mRNA preparation. Scale bar, 50 nm. E Size, PDI, and zeta potential of L@Mn-mRNA nanoparticles detected by DLS. Data were presented as mean ± SD, n = 3 technical replicates. F Distribution of phosphorus (green) and manganese (red) elements of L@Mn-mRNA detected by Energy-Dispersive X-ray Spectroscopy (EDS) element analysis. There was no gap between two green (P) particles as shown by the single white dash line, and there was a gap between manganese regions labeled by two white dash lines. Scale bar, 10 nm. G The stability evaluation of L@Mn-mRNA detected by DLS. The L@Mn-mRNA storage temperature was 4 °C. Data were presented as mean ± SD, n = 3 technical replicates. H Cytotoxicity evaluation of L@Mn-mRNA and LNP-mRNA. L@Mn-mRNA and LNP-mRNA were respectively incubated with DC 2.4 Cells at equal mRNA concentration (left) (0, 0.1, 1, 2, 5, 10 μg/mL) or total lipid concentration (right) (0, 2, 20, 40, 100, 200 μg/mL) for 24 h. The cytotoxicity was evaluated by using CCK-8 assay. Data were presented as mean ± SD, n = 4 biologically independent samples. Statistical significance was analyzed by two-tailed t-test. mLuc was used in (B, C, F–H). SM-102 was used as ionizable lipid in the formulations in (B–H). Figure 3A was created in BioRender. Han, D. (2024) BioRender.com/ EU272J1C4F. Source data are provided as a Source Data file.
Cryo-EM images showed a typical core-shell structure of lipid-coated Mn-mRNA (L@Mn-mRNA), indicating that Mn-mRNA nanoparticles were successfully encapsulated by lipids (Fig. 3D). Four types of mRNA (mSpike, mLuc, mOVA, and mEGFP) and two types of commercialized ionizable lipids (SM-102 and Dlin-MC3-DMA) were used to investigate the universality of our method. Both SM-102 and Dlin-MC3-DMA could encapsulate these four types of mRNA (Fig. 3D, Supplementary Fig. 7). DLS results indicated an increase in the hydrodynamic diameter of the nanoparticles upon lipids coating, and the charge shifted from negative to positive in all groups (Fig. 3E and Supplementary Tables 3 and 5). To further confirm compositions of the core-shell structure, we employed elemental analysis to study the element distribution in L@Mn-mRNA nanoparticles. Distinct boundary could be observed in the Mn distribution, while the range of P was significantly larger than that of Mn (Fig. 3F). This result further demonstrated that the Mn-mRNA core had been coated by lipids.
The size, PDI, and zeta potential of L@Mn-mRNA did not have significant changes when incubated with PBS for three weeks (Fig. 3G), indicating a good stability. Meanwhile, we also prepared LNP-mRNA according to the commercial formulation as the control group for subsequent experiments38, and their morphology, size, zeta potential, and mRNA encapsulation efficiency were shown in Supplementary Fig. 8 and Supplementary Table 6. Besides, the cytotoxicity of L@Mn-mRNA and LNP-mRNA was evaluated on DCs (Fig. 3H). At the same mRNA dosage, the lipids content in L@Mn-mRNA is approximately half that of LNP-mRNA, in which test, the high concentration group (10 µg mRNA dose) of the L@Mn-mRNA had higher cell viability than that of LNP-mRNA, suggesting that L@Mn-mRNA with less lipids had the potential to reduce the cytotoxicity from formulation themselves when a higher therapeutic dosage was required. In another test, at equivalent lipids levels, L@Mn-mRNA showed no significant difference in cytotoxicity compared to LNP-mRNA at each concentration, indicating that the small amount of Mn2+ in L@Mn-mRNA did not induce cytotoxicity.
Enhanced cellular uptake of L@Mn-mRNA by dendritic cells
Then we investigated the cellular uptake of L@Mn-mRNA (first labeled the mRNA (mLuc) with Cy5). Under equivalent mRNA concentration (dose), the L@Mn-Cy5-mRNA group (half lipid content of the LNP-mRNA group) exhibited enhanced Cy5 uptake both at 4 (1.68 ± 0.10 fold) and 10 (1.80 ± 0.29 fold) hours (Fig. 4A and Supplementary Figs. 9A and 10A) imaged by using confocal laser scanning microscopy (CLSM). Flow cytometry results indicated that the Cy5 signal in L@Mn-Cy5-mRNA treated DCs was nearly twofold of that in LNP-Cy5-mRNA treated group (Fig. 4B, C and Supplementary Fig. 11A, B). In another experiment, we labeled DMG-PEG2K with Cy5 to prepare the LNP-mRNA and L@Mn-mRNA (named as Cy5-LNP-mRNA and Cy5-L@Mn-mRNA), and DCs were incubated with these two types of nanoparticles at the same mRNA dose (Cy5-labeling lipids in Cy5-L@Mn-mRNA was half dose of that in Cy5-LNP-mRNA) for 4 h and 10 h, respectively. Both CLSM and flow cytometry showed similar Cy5 signal intensity in each group (relative intensity at 4 h: 1 ± 0.16 vs. 0.98 ± 0.14; at 10 h: 1 ± 0.09 vs. 1.08 ± 0.05) (Fig. 4D–F and Supplementary Figs. 9B, 10B, and 11C, D). These results suggested that the cellular uptake efficiency of L@Mn-mRNA was about twofold of LNP-mRNA at the same mRNA dosage.
A Cellular uptake of LNP-Cy5-mRNA and L@Mn-Cy5-mRNA at 4 h imaged by CLSM. Same Cy5-mRNA dose (0.5 μg/mL) was used to incubate with DC 2.4 cells. Red, Cy5 labeled mRNA; green, Lysotracker green; blue, Hoechst. Scale bar, 10 µm. B Cellular uptake of LNP-Cy5-mRNA and L@Mn-Cy5-mRNA at 4 h time point detected by flow cytometry. C Quantitative analysis of cellular uptake from flow cytometry. Mean fluorescence intensity of L@Mn-Cy5-mRNA was twofold of that in LNP-Cy5-mRNA. Data were presented as mean ± SD, n = 5 biologically independent samples. P < 0.0001. D Cellular uptake of Cy5-LNP-mRNA and Cy5-L@Mn-mRNA at 4 h time point imaged by CLSM. Cy5 labeled lipid (Cy5-DMG-PEG2K) was used in this experiment. Same mRNA dose (0.5 μg/mL) was used, which meant the Cy5-DMG-PEG2K in Cy5-L@Mn-mRNA was about 50% of that in Cy5-LNP-mRNA. Red, Cy5 labeled DMG-PEG2K; green, Lysotracker green; blue, Hoechst. Scale bar, 10 µm. E Cellular uptake of Cy5-LNP-mRNA and Cy5-L@Mn-mRNA at 4 h time point detected by flow cytometry. F Quantitative analysis of cellular uptake from flow cytometry. Mean fluorescence intensity of Cy5-L@Mn-mRNA was similar to that of Cy5-LNP-mRNA. Data were presented as mean ± SD, n = 5 biologically independent samples. G, H Stiffness of L@Mn-mRNA and LNP-mRNA measured by AFM. Scale bar in AFM images, 200 nm. The Young’s modulus of L@Mn-mRNA was 1.3-fold of that in LNP-mRNA. Twenty nanoparticles of each group were randomly collected to calculate the stiffness. Data were presented as mean ± SD, n = 20 technical replicates. P < 0.0001. SM-102 and mLuc were used. Statistical significance of (C, F, and H) was analyzed by two-tailed t-test. Figure 4G was created in BioRender. Han, D. (2024) BioRender.com/ HX272J1C5X. Source data are provided as a Source Data file.
We deduced that the enhanced cellular uptake was attributed to the enhanced stiffness provided by the Mn-mRNA core39,40. Therefore, we used atomic force microscopy (AFM) in mechanical mode to measure the Young’s modulus of LNP-mRNA and L@Mn-mRNA (Fig. 4G). After randomly selecting twenty particles, we found that the Young’s modulus of L@Mn-mRNA was about 1.3 times higher than that of LNP-mRNA (Fig. 4H). This result supported our hypothesis that increased stiffness enhanced the cellular uptake.
Additionally, confocal microscopy results showed that the colocalization of Cy5 with lysosomes (lysotracker green) decreased after 10 h of incubation with the cells (Supplementary Fig. 9A, B), indicating that L@Mn-mRNA could achieve lysosomal escape, and the Pearson correlation coefficient analysis showed that L@Mn-mRNA had a superior efficiency of lysosomal escape than that of LNP-mRNA (Supplementary Fig. 12). This phenomenon is probably attribute to the increased stiffness of L@Mn-mRNA (compared to LNP-mRNA), which facilitated enhanced lysosomal escape efficiency41. Notably, since the Mn-mRNA core could dissociate under physiological conditions (at PBS or FBS) (Supplementary Fig. 13) due to the presence of abundant ions and proteins, the mRNA expression would not be affected after the lysosomal escape.
L@Mn-mRNA promoted maturation and antigen presentation of dendritic cells in vitro and in vivo
Before we investigated immune effect of L@Mn-mRNA vaccine platform, we used mLuc as the reporter mRNA to compare the in vitro and in vivo expression efficiency of L@Mn-mRNA and LNP-mRNA. We incubated these two types of nanoparticles with bone marrow-derived dendritic cells (BMDCs) for 12 and 24 h, and quantified bioluminescence intensity at each time point using a plate reader. The result showed that L@Mn-mLuc had nearly two times higher in vitro expression efficiency than LNP-mLuc (Supplementary Fig. 14). We subcutaneously injected L@Mn-mLuc and LNP-mLuc into the tail bases of C57BL/6 mice, followed by bioluminescence imaging using the in vivo imaging system (IVIS) at 6 and 24 h post-injection. The images indicated that L@Mn-mLuc group had 4.1 times higher expression efficiency in vivo than LNP-mLuc at 6 h and twofold of LNP-mLuc at 24 h (Fig. 5A and Supplementary Fig. 15), probably due to its superior capability entering cells. After 3-week storage, the mRNA expression efficiency in L@Mn-mLuc reduced to 75 %, while the LNP-mLuc group reduced to 69 % (Supplementary Fig. 16). This phenomenon further demonstrated a comparable stability between L@Mn-mLuc and LNP-mLuc. We also do the subcutaneous injections on both sides of the groin, and the bioluminescence signal (Supplementary Fig. 17A) had a similar trend with Supplementary Fig. 15. The tissues were collected post the 24 h imaging, and both groups exhibited heightened expression exclusively in the inguinal lymph nodes (iLN) proximal to the injection site (Supplementary Fig. 17B). Overall, due to the improved cellular uptake, the L@Mn-mRNA significantly enhanced the mRNA expression efficiency both in vitro and in vivo, validating its potential to be an efficient mRNA delivery platform. In addition, the incorporation of Mn2+ did not alter the stability of the L@Mn-mRNA.
A In vivo Luciferase expression of LNP-mLuc and L@Mn-mLuc at 6 h after subcutaneous injection to mice imaged by IVIS (5 μg mLuc per mouse). The bioluminescence signal in L@Mn-mLuc group was about 4.1-fold stronger than that of LNP-mLuc group. Data were presented as mean ± SD, n = 3 biologically independent samples. P = 0.0282. Statistical significance was analyzed by two-tailed t-test. Minimum: 3 × 105 p/s/cm2/sr; Maximum: 8× 106 p/s/cm2/sr. B–D Flow cytometry examination of BMDCs maturation percentage (CD80+, CD86+, CD40+ cells percentages) 24 h after incubation with PBS, LNP-mOVA (mOVA dose: 1 μg/mL), L@Mn-mOVA (mOVA dose: 0.5 μg/mL), and L@Mn-mOVA (mOVA dose: 1 μg/mL). Data were presented as mean ± SD, n = 4 biologically independent samples. E, F IL-6 (E) and TNF-α (F) secretion from BMDCs after treatment with each formulation for 24 h detected by ELISA Kit. Data were presented as mean ± SD, n = 4 biologically independent samples. G Flow cytometry analysis of SIINFEKL-H2Kb+ presentation on BMDCs after treatment with each formulation for 24 h. Data were presented as mean ± SD, n = 4 biologically independent samples. H–J Flow cytometry analysis of matured dendritic cells (CD86+, CD80+, CD40+) in LNs after administration with PBS, LNP-mOVA (mOVA dose: 10 μg per mouse), L@Mn-mOVA (half dosage, mOVA dose: 5 μg per mouse), and L@Mn-mOVA (mOVA dose: 10 μg per mouse) for 24 h Data were presented as mean ± SD, n = 4 biologically independent samples. K Flow cytometry analysis of SIINFEKL-H2Kb+ presentation on matured dendritic cells in lymph nodes with each formulation for 24 h. mOVA dose was the same with that of (H–J). Data were presented as mean ± SD, n = 4 biologically independent samples. SM-102 was used in this part. Statistical significance of (B–K) was analyzed by one-way ANOVA with Tukey’s correction. Source data are provided as a Source Data file.
Then the capability of L@Mn-mOVA to activate the dendritic cells maturation and antigen presentation was evaluated in vitro and in vivo. BMDCs were respectively treated with PBS, LNP-mOVA, L@Mn-mOVA (half dosage) and L@Mn-mOVA. After 24-h incubation, the dendritic cells maturation markers, such as CD80, CD86, CD40, were detected by using flow cytometry (Fig. 5B–D), and the secretion of pro-inflammatory cytokines IL-6 and TNF-α in the cell culture medium was tested by ELISA kits (Fig. 5E, F). The results showed that the L@Mn-mOVA group was 1.2-fold expression of CD80, 1.2-fold expression of CD86, and 1.7-fold expression of CD40 compared to the LNP-mOVA (Fig. 5B–D). The IL-6 and TNF-α secretion was both approximately threefold of that in the LNP-mOVA group (Fig. 5E, F). The antigen presentation was characterized by measuring the frequency of OVA (SIINFEKL)-H2Kb complexes on the surface of BMDCs, and the L@Mn-mOVA group exhibited a nearly 1.7 times higher antigen presentation compared to LNP-mOVA group (Fig. 5G). Notably, even the L@Mn-mOVA (half dosage) group showed significant increases in these results compared to the LNP-mOVA group.
We also explored if the immune response enhancement caused by Mn2+, because Mn2+ can be act as an agonist of cGAS-STING pathway42,43,44,45. However, we did not observe significant STING activation after incubating L@Mn-mOVA with RAW-Lucia ISG reporter cells (for STING activation detection46,47) compared to the MnCl2 treated groups (100 μM & 200 μM, as previous reported48,49) (Supplementary Fig. 18), probably because the Mn2+ concentration in L@Mn-mOVA was much lower than that of MnCl2 groups. For example, Mn2+ concentration was only 8.4 µM even the mRNA dose reached 10 µg/mL in L@Mn-mOVA formulation. In addition, no differences were detected between L@Mn-mOVA and LNP-mOVA groups in STING activation (Supplementary Fig. 18). Therefore, the enhanced immune response in the L@Mn-mOVA group was more likely attributed to high mRNA loading and improved cellular uptake.
To evaluate in vivo responses, each formulation (PBS, LNP-mOVA, L@Mn-mOVA with different dosages) was subcutaneously injected into the inguinal region of C57BL/6 mice. iLNs were harvested for flow cytometric analysis at the 24 h time point after administration. with the results showing that L@Mn-mOVA group exhibited 2.5-fold expression of CD80, 1.7-fold expression of CD86, and 4.1-fold expression of CD40 compared to the group treated with LNP-mOVA (Fig. 5H–J). As for the in vivo antigen presentation, the L@Mn-mOVA group achieved 1.9-fold SIINFEKL-H-2Kb-positive cells compared to the LNP-mOVA group (Fig. 5K). In addition, the level of these markers in the half dosage mOVA in L@Mn-mOVA group was comparable to LNP-mOVA group, and some makers (such as CD40+ and SIINFEKL-H-2Kb-positive cells) were higher than that of LNP-mOVA. The gating strategy of DCs maturation and antigen presentation are presented in Supplementary Fig. 19. To further verify whether Mn2+ exerts an adjuvant function in vivo, we subcutaneously administered formulations into the inguinal region of mice on Day 0, 3, and 6, and detected the expression of p-IRF-3 (cGAS-STING pathway’s downstream marker) in the iLNs on Day 13. No significant difference in p-IRF-3 expression was observed between the LNP-mOVA group and the L@Mn-mOVA group, indicating that the L@Mn-mOVA did not lead to additional activation of the STING-related pathway compared to LNP-mOVA treatment (Supplementary Fig. 20).
Therapeutic efficacy of L@Mn-mRNA vaccine in B16-OVA melanoma tumor model
To evaluate the anti-tumor efficacy of L@Mn-mRNA vaccine, the B16-OVA melanoma tumor model was established on C57BL/6 mice, and the mice were subcutaneously administered in the inguinal regions with PBS, LNP-mOVA (10 μg, representing 10 μg mOVA per mouse), L@Mn-mOVA (5 μg, representing half dosage 5 μg mOVA per mouse) and L@Mn-mOVA (10 μg, representing 10 μg mOVA per mouse) three times (on Day 4, 7, and 10) (Fig. 6A). The L@Mn-mOVA group exhibited higher efficiency on tumor-suppression than that of the LNP-mOVA group (Fig. 6B and Supplementary Fig. 21A), with improving median survival rate by 26% (46–58 days) (Supplementary Fig. 21B). The half dosage L@Mn-mOVA still exhibited a certain degree of superior anti-tumor effect compared to the LNP-mOVA group, extending median survival by 13% (46–52 days) (Fig. 6B and Supplementary Fig. 21B).
A Time line of therapeutic procedures. B Tumor growth curves of each mouse after treatment with each formulation. mOVA dose in LNP-mOVA and L@Mn-mOVA were 10 μg per mouse, mOVA dose in L@Mn-mOVA half dose was 5 μg per mouse. n = 5 biologically independent samples of each group. C SIINFEKL tetramer+ in CD8+ cells in the whole blood after the treatment (at Day 17) analyzed by using flow cytometry. Data were presented as mean ± SD, n = 5 biologically independent samples. D Flow cytometry analysis of SIINFEKL tetramer+ in CD8+ cells in the spleen after the treatment (at Day 17). Data were presented as mean ± SD, n = 5 biologically independent samples. E, F The IFN-γ secretion by splenocytes after restimulation with OVA peptide was determined by the ELISpot assay. Data were presented as mean ± SD, n = 5 biologically independent samples. G–J Serum levels of IL-6 (G), TNF-α (H), IFN-β (I), and IFN-α (J) were evaluated by using ELISA kits. Data were presented as mean ± SD, n = 4 biologically independent samples. K–M The infiltrating immune cells in tumors of each group after treatment (at Day 17) detected by using flow cytometry, including CD3+CD4+ T cells (K), CD3+CD8+ T cells (L), and CD11c+ DCs (M). Data were presented as mean ± SD, n = 4 biologically independent samples. N Time line of immune memory measurement. O, P Immune memory evaluation on Day 70 after treatment by flow cytometry. The proportions of central memory T cells (CD3+ , CD8+ , CD44high and CD62Lhigh) (O) and effector memory T cells (CD3+, CD8+, CD44high, CD62Llow) (P) in the splenocytes detected by using flow cytometry. Data were presented as mean ± SD, n = 5 biologically independent samples. SM-102 was used in this part. Statistical significance of (C, D, F–M, and O, P) were analyzed by one-way ANOVA with Tukey’s correction. Source data are provided as a Source Data file.
Then we evaluated the antigen-specific immune response in blood and splenocytes. The L@Mn-mOVA (10 μg) group showed eightfold increase in the proportion of SIINFEKL tetramer+ cells within CD8+ T cells in the blood and 1.8-fold increase in the spleen compared to the LNP-mOVA group (Fig. 6C, D). Although the difference between the L@Mn-mOVA (5 μg) group and the LNP-mOVA group in the blood was negligible, the L@Mn-mOVA (5 μg) group showed a significant increase in SIINFEKL tetramer+ cells within CD8+ T cells compared to the LNP-mOVA group in the spleen (Fig. 6C, D). The corresponding gating strategy was presented in Supplementary Fig. 22. Further, splenocytes isolated from mice subjected to various treatment groups were analyzed through the IFN-ELISpot assay. The half-dose L@Mn-mOVA (5 μg) group and the L@Mn-mOVA group (10 μg) exhibited threefold and 3.5-fold increases in the number of spots compared to the LNP-mOVA group (Fig. 6E, F), respectively. There results demonstrated that the L@Mn-mOVA groups (including the half-dose group) effectively activated antigen-specific immune responses.
For the serum pro-inflammatory cytokines (IL-6, TNF-α, IFN-α, and IFN-β) detection, the L@Mn-mRNA group had a significant increase compared to the LNP-mOVA group (Fig. 6G–J), and the TNF-α and IFN-β in L@Mn-mOVA (5 μg) group were also higher than that of the LNP-mOVA group, indicating an enhanced immunogenic effectiveness.
Then the immune infiltration at the tumor site was assessed by flow cytometry. The L@Mn-mOVA (10 μg) group exhibited twofold ratio of CD3+CD8+ T cells and 1.8-fold ratio of CD3+CD4+ T cells compared to that of LNP-mOVA group (Fig. 6K, L). The L@Mn-mOVA (10 μg) group also exhibited 1.5-fold number of CD11c+ cells (antigen-presenting cells) (Fig. 6M). The L@Mn-mOVA (5 μg) group showed a significant increase in the number of CD3+CD8+ T cells, CD3+CD4+ T cells and similar number of CD11c+ cells compared to the LNP-mOVA group. Thus, compared to the LNP-mOVA group, the L@Mn-mOVA exhibited a more robust stimulation of the immune system in combating tumors. Additionally, the half-dose group, L@Mn-mOVA (5 μg) showed comparable levels of immune activation to LNP-mOVA group. The corresponding gating strategy was presented in Supplementary Fig. 23.
To evaluate the capability of L@Mn-mRNA on inducing long-term immune memory, we subcutaneously administered the formulations on Day 0, Day 3, and Day 6 to C57BL/6 mice without tumor inoculation, and the mice were sacrificed for analysis on Day 70 (Fig. 6N). At the same dosage, the L@Mn-mOVA group had a higher proportion of CD44highCD62Lhigh T cells (central memory T cells) and CD44highCD62Llow T cells (effector memory T cells) in the spleen compared to the LNP-mOVA group, indicating stronger immune memory (Fig. 6O, P). The gating strategy of central memory T cells and effector memory T cells was presented in Supplementary Fig. 24. Based on these results, the L@Mn-mOVA exhibited superior anti-tumor efficacy compared to LNP-mOVA at the same dosage. Even the L@Mn-mOVA (5 μg) group, which meant the lipid usage was a quarter of that in LNP-mOVA, achieved effective anti-tumor immune response.
Safety evaluation and bioeffects of L@Mn-mRNA vaccine
It is crucial to evaluate the safety of the L@Mn-mRNA vaccine platform. During the therapeutic period, no fluctuations were observed in the body weight of the mice (Supplementary Fig. 25), and we found no significant organic disease in the major organs (heart, liver, spleen, lung, and kidney) after treatment compared to that of healthy mice (Supplementary Fig. 26). In addition, blood biochemical criterion (alanine aminotransferase (ALT), aspartate aminotransferase (AST), total protein (TP), albumin (ALB), globulin (GLB), albumin/globulin ratio (A/G) and alkaline phosphatase (ALP) for evaluating liver functions; blood urea nitrogen (BUN) and creatinine (CR) for evaluating kidney function; triglycerides (TG) for cardiovascular health) did not had significant differences in all groups (Supplementary Fig. 27), which indicated the formulations did not affect the functions of major organs.
Manganese accumulation in organs is another safety concern of the L@Mn-mRNA vaccine platform, since high manganese accumulation in brain may cause neurotoxic50. After subcutaneously administering L@Mn-mRNA at Day 0, 3, and 6, we measured the manganese content in major organs (heart, liver, spleen, lung, kidney, brain) and blood of mice on Day 13. Although there was a certain level increase of manganese content in the liver (1.18 ± 0.07 fold) and kidney (1.25 ± 0.17 fold) for the L@Mn-mRNA group compared to the control group (Supplementary Fig. 28), the manganese concentration remained within the normal range for mice (because manganese is a type of microelement)51,52,53,54. For other organs, including the brain, no significant manganese accumulation was observed after the treatment of L@Mn-mOVA (Supplementary Fig. 28). These results demonstrated that the Mn2+ component in L@Mn-mOVA vaccine would not generate toxicity.
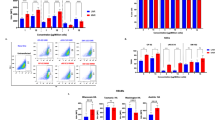
Meanwhile, we conducted antibody tests for anti-PEG on blood samples from each group of mice before and after treatment. The timeline of the anti-PEG IgM/IgG antibody titers measurement was presented in Fig. 7A. In the PBS group, anti-PEG IgM levels showed no significant change before and after administration. In the LNP-mRNA (10 μg) group, 60% of the mice exhibited at least a 20-fold increase in anti-PEG IgM levels. In comparison, 60% of the mice showed a tenfold increase in anti-PEG IgM level in L@Mn-mRNA (10 μg) group and only 40% of the mice had a fivefold increase in L@Mn-mRNA (5 μg) group. Similarly, as for the measurement of anti-PEG IgG, in the LNP-mRNA (10 μg) group, 80% of the mice exhibited a substantial increase (ranging from a minimum of sixfold to a maximum of ninefold). In the L@Mn-mRNA (10 μg) group, only 20% of the mice exhibited a notable elevation in antibody levels (ninefold), with the remaining mice displaying marginal increments. In the L@Mn-mRNA (5 μg) group, all mice had no significant increase (Fig. 7B). These results revealed that a reduced lipids input (in the L@Mn-mRNA formulation) would be benefit to avoid a higher production of anti-drug antibodies, which potentially offered an effective strategy to avoid the accelerated blood clearance.
A The timeline of the anti-PEG IgM/IgG titers. B Anti-PEG IgM/IgG measurement detected by ELISA Kit. Anti-PEG IgM was detected on Day −1 and Day 13, and anti-PEG IgG was detected on Day −1 and Day 20. Day −1 represented the day before performing the injection of each formulation. n = 5 biologically independent samples. SM-102 was used in this experiment. C Time line of administrating and imaging. D In vivo Luciferase expression of LNP-mLuc and L@Mn-mLuc at 6 h after subcutaneous injection to mice imaged by IVIS (10 μg mLuc per mouse) at Day 0, 13, and 20. Data were presented as mean ± SD, n = 5 biologically independent samples. Minimum: 4 × 105 p/s/cm2/sr; Maximum: 4 × 107 p/s/cm2/sr. Source data are provided as a Source Data file.
To further demonstrate that the L@Mn-mRNA vaccine platform could overcome the mRNA expression decrease caused by anti-PEG IgG/IgM mediated clearance, we subcutaneously injected LNP-mLuc and L@Mn-mLuc (10 μg mLuc per mouse) into the tail bases of C57BL/6 mice on Day 0, 3, and 6, and we measured mRNA expression efficiency on Day 13 and 20 (Fig. 7C, D). The L@Mn-mLuc group exhibited around 93% and 35 % mLuc expression (compared to the first injection) on Day 13 and Day 20, respectively. However, the LNP-mLuc group only had less than 15% mLuc expression (compared to the first injection) on Day 13 and Day 20. These results demonstrated that the L@Mn-mRNA could maintain the mRNA expression efficiency in a certain degree, which was probably because of the reduced risk of anti-PEG IgG/IgM generation.
In summary, we have developed a platform technique for improving mRNA loading in lipid-based nanoparticles. Mn2+ could efficiently enrich mRNA and form a high-density mRNA core before the lipid encapsulation. The developed platform (L@Mn-mRNA) integrated advantages on increased mRNA loading, enhanced cellular uptake and decreased the risk of lipid or PEG induced side-effects or rapid clearance. The platform technique could be suitable for types of novel ionizable lipids and mRNA. Future studies may focus on the scale-up preparation and the universality to diseases (for mRNA-based therapy). Thus, this work holds the potential to provide references for the next generation of lipid-based mRNA therapeutics.
Methods
Materials
All mRNAs were purchased from APE×BIO or TriLink Biotechnologies. Manganese (Ⅱ) chloride tetrahydrate was obtained from Alfa Aesar (China) Chemicals Co., Ltd. DMG-PEG2K (R-PEG-0021), Cy5-DMG-PEG2K (R-PF-0170), and DSPC (LP-R4-076) obtained from Xi’an Ruixi Biological Technology Co., Ltd. DLin-MC3-DMA was purchased from AVT (Shanghai) Pharmaceutical Tech Co., Ltd. Cholesterol was purchased from Solarbio Biotech Co., Ltd. SM-102 was purchased from Xiamen Sinopeg Biotech Co., Ltd. Hoechst 33342 (62249), LysoTracker® Green DND-26 (L7526) and Lipofectamine 3000 (L3000075) were purchased from Invitrogen Trading (Shanghai) Co., Ltd. FITC anti-mouse CD11c antibody (117305, 1:100), PE-Cy7 anti-mouse CD86 antibody (105013, 1:100), APC anti-mouse CD80 antibody (104713, 1:100), APC anti-mouse CD40 antibody (124611, 1:100), APC anti-mouse H-2Kb bound to SIINFEKL antibody (141605, 1:100), PE anti-mouse H-2Kb bound to SIINFEKL antibody (141603, 1:100), PE anti-mouse CD40 antibody (124609, 1:100), FITC anti-mouse CD3 antibody (100203, 1:100), PE-Cy7 anti-mouse CD8 antibody (100721, 1:100), APC anti-mouse CD4 antibody (100515, 1:100), APC anti-mouse CD11c antibody (117309, 1:100), APC anti-mouse CD44 antibody (103011, 1:100) and PE anti-mouse CD62L antibody (104407, 1:100) were purchased from Biolegend. T-Select H-2kb OVA Tetramer-SIINFEKL-PE mouse (TS-5001-1C) were purchased from MBL Life Science. IFN-1α (MM-47294M1) and IFN-1β (MM-46462M1) ELISA kit assay were obtained from Meimian Industrial Co., Ltd. Mouse Anti-PEG IgM ELISA (#PEGM-1) and Mouse Anti-PEG IgG ELISA (#PEGG-1) were purchased from Life Diagnostics, Inc. The mouse IL-6 (1210602), TNF-α (1217202), IFN-γ pre-coated ELISpot kit (2210005) were obtained from Dakewe Biotech Company. OVA257–264 (SIINFEKL) were obtained from Solarbio Biotech Co., Ltd. Phenylmethanesulfonyl fluoride was purchased from Beijing Lablead. D-Luciferin (D1007) was purchased from Solarbio Biotech Co., Ltd. Phospho-IRF-3 (Ser396) Rabbit mAb #29047 and Anti-rabbit IgG, HRP-linked Antibody 7074 (Goat) were purchased from Cell Signaling Technology. RPMI 1640 medium and Opti-MEM reduced serum medium were obtained from Thermo Fisher Scientific Inc.
Mn-mRNA and Lipid@Mn-mRNA (L@Mn-mRNA) preparation
For a reaction volume of 200 μL, we selected a 600 μL EP tube. First, 50 μL of 6 mM Mn2+ solution (manganese chloride tetrahydrate aqueous solution) was mixed with 130 μL of RNzyme-free water. Then, 20 μL of 1 mg/mL mRNA was combined with the former mixture. It was then vortexed for 6 min. Afterwards, the sample was heated in the metal bath at 65 °C for 5 min. After completion of the reaction, the sample was cooled to the room temperature. The Mn-mRNA nanoparticles were collected through centrifugation at 60,000 × g for 15 min, and the Mn-mRNA nanoparticles are dispersed in 200 μL of enzyme-free water. The mRNA and Mn2+ that were not included in Mn-mRNA nanoparticles could be removed though the centrifugation. SM-102 (or Dlin-MC3-DMA), cholesterol, DSPC, and DMG-PEG2000 were mixed according to the proportions specified in the formulation (molar ratio of 50:38.5:10:1.5) using anhydrous ethanol as the solvent. A volume ratio of anhydrous ethanol to aqueous phase (1:3) and a total lipid to mRNA mass ratio (10:1) was maintained. The aqueous phase was then rapidly injected into the ethanol phase with a pipette, followed by rapid pipetting for 150 times. It was allowed to stand at room temperature for 1 h. Finally, excess ethanol from the system was removed using a dialysis cup with a cut-off molecular weight of 10 kDa to obtain the L@Mn-mRNA.
For the preparation of Fe-mRNA, Cu-mRNA, and Zn-mRNA, 6 mM M2+ solutions were respectively prepared using ferric chloride tetrahydrate, anhydrous copper (II) sulfate, and zinc nitrate enneahydrate. The ratios of M2+ to bases were referred from the literature31,32. The remaining steps were identical to the preparation method of Mn-mRNA. When preparing Mn-mRNA with different Mn2+ to nucleotide molar ratios, the amount of manganese ions was adjusted accordingly. When preparing L@Mn-mRNA with different total lipid-to-mRNA mass ratios, the amount of lipids was adjusted accordingly.
Mn-mRNA, L@Mn-mRNA, and LNP-mRNA characterization
The morphological characterization of the particles was imaged using transmission electron microscopy (Hitachi HT7700) and cryo-electron microscopy (Thermo Fisher Titan Krios), respectively. Distribution of phosphorus (green) and manganese (red) elements of particles was detected by Energy-Dispersive X-ray Spectroscopy (EDS) element analysis from field-emission transmission electron microscope (JEOL JEM-F200). The size, polydispersity index (PDI) and zeta potentials of Mn-mRNA, L@Mn-mRNA, and LNP-mRNA were measured by using DLS (Malvern Panalytical Mastersizer). Each set of data was tested three times. Diameters are reported as the number mean peak average. The characterization of particle size and zeta potential for all particles was conducted by diluting the particles in water at a ratio of 1:50. To calculate mRNA reaction efficiency, Zn-mRNA or Mn-mRNA nanoparticles and the corresponding supernatant were dissolved by mixing them with a hydrochloric acid solution at pH 334. The mRNA content in both particles and supernatant was measured separately using the Quant-itTM RiboGreen RNA Assay Kit (n = 3 in each group). The Mn2+ content was measured using ICP-MS (Perkinelmer NexION 300X). The mRNA encapsulation efficiency of LNP-mRNA was calculated by Quant-it™ RiboGreen RNA Assay Kit according to the instructions.
The quantitative calculation of the Mn-mRNA composition was showed below:
-
1.
Mn ratio involved in the reaction (by ICP-MS) equals to “Mn in the precipitate (Mn-mRNA) (μg)”/“Mn in the precipitate (Mn-mRNA) (μg) + Mn in the supernatant solution (μg)”
-
2.
Mn amount involved in the reaction equals to “Mn Input × Mn ratio involved in the reaction”
-
3.
mRNA amount involved in the reaction equals to “mRNA Input” × “mRNA encapsulation efficiency”
-
4.
The proportion of mRNA in Mn-mRNA equals to “mRNA involved (μg)” / “Mn involved (μg) + mRNA involved (μg)”
-
5.
The proportion of Mn in Mn-mRNA equals to “Mn involved (μg)”/“Mn involved (μg) + mRNA involved (μg)”
L@Mn-mRNA was mixed with a solution containing 2% Triton ×-100 and hydrochloric acid (pH 3) to disrupt the lipid membrane and dissociate Mn-mRNA complexes. The mRNA content was then measured using the RiboGreen assay kit. The Mn content was determined by using ICP-MS, as previously described.
The mRNA capacity of L@Mn-mRNA was determined using the following formula:
mRNA capacity equals to “mRNA content/(Mn content + mRNA content + lipid content)”
The mRNA capacity of LNP-mRNA was determined using the following formula:
mRNA capacity equals to “mRNA content/(mRNA content + lipid content)”
Agarose gel electrophoresis
Agarose gel electrophoresis was run on a 1% agarose gel using 1× TAE as electrophoresis buffer at 100 V for 30 min. The mRNA sample amount in each well was 0.5 μg. Subsequently, we utilized ultraviolet light for imaging.
Simulated calculation
The geometric and electronic properties of metal ions-mRNA have been computed using the Gaussian 09 W program package at B3LYP/6-31 G(d,p) level55. In order to ensure the feasibility of the calculation, we have adjusted the number of self-selected electrons in systems. After optimization, the length of bonds was measured by Mutifwn56. Utilizing the iefpcm implicit solvent model, single-point energy calculations for the structures were performed employing the wb97xd57 functional with the def2TZVP58 basis set. Frequency calculations were carried out using the 6-311Gd59 basis set to determine the binding energies of the complexes at 298 K for different components.
LNP-mRNA and L@Mn-mRNA Young’s moduli test from AFM
Initially, a circular mica sheet with a diameter of 1 cm was affixed securely onto an iron plate. Subsequently, 30 μL of LNP-mRNA and L@Mn-mRNA were incubated respectively on the mica sheet for 20 min. After gently washing the mica sheet with PBS, the sheet was then covered with 20 μL of PBS. AFM imaging and force measurements were carried out in aqueous medium at 26 ± 1 °C by using a Bruker Multimode-8 with NanoScope Analysis software (Version 1.9). The probe used for measurements was SCANASYST-FLUID + . AFM images were recorded with the following scanning parameters: scan rate, 0.977 Hz; peak force amplitude, 40 nm; peak force frequency, 2 KHz and spring constant, 0.7 N/m. The force between the tip and the sample was carefully maintained so as not to collapse the nanoparticles. The resolution was 256 × 256 pixels per AFM. Twenty data points were selected for quantitative analysis in each group.
Cells and animals
Female C57BL/6 (6–8 weeks old) were purchased from SPF Biotechnology. This study was approved by the ethics committee of the National Center for Nanoscience and Technology of China (approval number NCNST21-2103-0401). The animals were maintained in a controlled environment with a 12-h light/dark cycle. The ambient temperature was maintained at 22–24 °C, and the humidity level was kept at 40–60%. The B16-OVA cell line was purchased from Meisen Chinese Tissue Culture Collections (CTCC; cell line number CTCC-003-0259). The DC 2.4 cell line was obtained from the American Type Culture Collection (Manassas, VA, USA). The B16-OVA cells and DC 2.4 cells were cultured in RPMI 1640 containing 10% fetal bovine serum (FBS), Pen-Strep (100 U/mL and 100 μg/mL, respectively) solution at 37 °C with 5% CO2. Raw-Lucia ISG cells (Raw-Lucia) were purchased from InvivoGen. BMDCs were obtained through the following steps. After euthanizing C57BL/6 mice (6–8 weeks), they were placed in 50 mL centrifuge tubes filled with 75% ethanol for 10 min for sterilization. Subsequently, the leg bones were removed, muscles were stripped, and bones on both sides were clipped. Bone marrow was then flushed into new culture dishes using a needle. Red blood cell lysis buffer was added afterwards, and the lysis was terminated after 3 min. Finally, cells were collected by centrifugation and resuspended in culture medium containing 20 ng/mL GM-CSF (Peprotech) and 20 ng/mL IL-4 (Peprotech), with changing medium every two days. On Day 6, the nonadherent cells were aspirated and incubated in 6-well plates with fresh medium for further investigation.
In vitro transfection
DC 2.4 cells were seeded in a 96-well plate at a density of 104 cells per well. Different kinds of mRNAs were heated for different durations in a metal bath, then mixed with Lipofectamine™ 3000 Transfection Reagent according to the instructions to form complexes (n = 3 in each group). These complexes were then transfected into cells at a dose of 100 ng mRNA per well. After 24 h, the EGFP expression level was measured using flow cytometry (ACEA NovoExpress) with 10000 live cells in FITC channel. The luciferase mRNA expression level was detected by mixing an equal volume of cell suspension and the Bright-Lite Plus Luciferase Assay System (Vazyme), and then results were measured by a plate reader.
Cellular uptake and lysosomal escape
L@Mn-Cy5-Mrna (L@Mn-Cm) and LNP-Cy5-mRNA (LNP-Cm) were synthesized using Cy5-labeled mRNA. Cy5-L@Mn-mRNA (CL@Mn-m) and Cy5-LNP-mRNA (CLNP) were synthesized using Cy5-labeled DMG-PEG2K. The particles were separately incubated with DC 2.4 cells for 4 and 10 h. DC 2.4 cells were stained with Hoechst 33342 (1:500) for 5 min and washed with Opti-MEM medium three times. After that, DC 2.4 cells were stained with LysoTracker® Green DND-26 (1:10000) for 30 min and washed with Opti-MEM medium three times. The cellular uptake (Hoechst 33342/405 nm; Cy5/640 nm) and lysosome escape events (DND-26/488 nm) were captured using a CLSM (Olympus FV3000). The endosomal escape ratio was analyzed using an image analysis algorithm (ImageJ).
DC 2.4 cells were seeded in a 12-well plate at a density of 105 cells per well. Subsequently, LNP-mRNA and L@Mn-mRNA nanoparticles were separately incubated with DC 2.4 cells at an equal mRNA dosage for varying durations (mRNA dose: 0.5 μg/mL). Finally, cell uptake was quantitatively analyzed using flow cytometry (ACEA NovoExpress) with 10,000 live cells in APC channel (n = 5 in each group).
In vitro and in vivo bioluminescence
L@Mn-mLuc and LNP-mLuc were respectively incubated in BMDCs for 12 and 24 h. Subsequently, an equal volume of cell suspension and the Bright-Lite Plus Luciferase Assay System (Vazyme) were mixed. Finally, the bioluminescence intensity was measured using a plate reader.
At 6 and 24 h after the injection of L@Mn-mLuc and LNP-mLuc (mRNA dose: 5 μg per mouse), mice were injected intraperitoneally with 0.2 mL d-luciferin (15 mg/mL in DPBS). Ten minutes later, the mice were imaged with an in vivo imaging system (PerkinElmer IVIS). Bioluminescence was quantified using the Living Image software (PerkinElmer).
Western blot analysis
GAPDH and p-IRF-3 protein expression in iLNs were analyzed by western blot. C57BL/6 mice were administered multiple subcutaneous injections of PBS, LNP-mOVA (10 μg), L@Mn-mOVA (5 μg), and L@Mn-mOVA (10 μg) into the inguinal region of mice on Day 0, 3, 6, and detected the expression of p-IRF-3 in the iLNs on Day 13. Specifically, on Day 13, the iLN of the mice were removed, homogenized, and proteins were extracted using lysis buffer. The lysate was then centrifuged in a pre-chilled centrifuge. Centrifugation was conducted at 4 °C and 11,000 × g for 20 min. After centrifugation, the supernatants were harvested for subsequent protein concentration measurement. Protein levels were determined using a BCA assay kit (Beyotime), following the protocol provided by the manufacturer. Equal amounts of protein (15 μg per sample) were separated via SDS-polyacrylamide gel electrophoresis and then transferred onto PVDF membranes. Membranes were blocked using 5% bovine serum albumin to prevent non-specific binding, and subsequently incubated with primary antibodies. After washing, membranes were treated with the appropriate secondary antibodies. Protein bands were detected using enhanced chemiluminescence reagents (Bio-Rad), and densitometric analysis of the bands was performed using ImageJ software. Each experimental group included three independent samples (n = 3).
Enzyme-linked immunosorbent assay (ELISA)
PBS, LNP-mOVA (2 μg/well), L@Mn-mOVA (1 μg/well), and L@Mn-mOVA (2 μg/well) were separately incubated with BMDCs in a 6-well plate (3 × 105 cells per well) for 24 h. The supernatant was collected for the detection of IL-6 and TNF-α secretion from BMDCs. After diluting the supernatant fivefold, 50 µL of the test sample was added to the pre-coated plates. Subsequent steps were carried out according to the manufacturer’s instructions. Finally, the results were read using a plate reader.
For the measurement of cytokines in serum, blood samples were collected from each group of mice at the endpoint of treatment (Day 17). After preparing serum from the blood samples, the serum was diluted fivefold. Then 50 µL of the test sample was added to pre-coated plates. Subsequent steps were carried out as described above (n = 4 in each group).
L@Mn-mRNA promoted maturation and antigen presentation of DCs in vitro and in vivo
BMDCs were seeded into a 12-well plate (1 × 105 cells per well) and then incubated with PBS, LNP-mOVA (mOVA dose: 1 μg/mL), L@Mn-mOVA (mOVA dose: 0.5 μg/mL), and L@Mn-mOVA (mOVA dose: 1 μg/mL) for 24 h. The medium from each group was collected for ELISA of TNF-α, IL-6 according to the manufacturer’s instructions. Meanwhile, cells were collected and stained with FITC anti-mouse CD11c antibody (1:100), PE-Cy7 anti-mouse CD86 antibody (1:100), APC anti-mouse CD80 antibody (1:100), APC anti-mouse CD40 antibody (1:100), and PE anti-mouse SIINFEKL-H-2kb antibody (1:100) for 30 min at 4 °C before the detection of flow cytometry (ACEA NovoExpress).
iLN from each mouse was harvested at 24 h post-injection of PBS, LNP-mOVA (10 μg mOVA per mouse), L@Mn-mOVA (5 μg mOVA per mouse), and L@Mn-mOVA (10 μg mOVA per mouse) at the inguinal regions and were gently mechanically disrupted using sterile pestles in RPMI 1640 medium in a 1.5 mL tube. The resulting cell suspensions were collected and stained with FITC anti-mouse CD11c antibody (1:100), PE-Cy7 anti-mouse CD86 antibody (1:100), APC anti-mouse CD80 antibody (1:100), PE anti-mouse CD40 antibody (1:100), and APC anti-mouse SIINFEKL-H-2kb antibody (1:100) for 30 min at 4 °C before being analyzed by flow cytometry (ACEA NovoExpress).
Anti-tumor effects in a subcutaneous B16-OVA cancer model
To investigate the anti-tumor efficacy of the L@Mn-mRNA vaccine in a murine melanoma model, C57BL/6 mice were subcutaneously injected with 1 × 106 B16-OVA cells into the right flank on Day 0. Mice were randomly divided into four groups and subcutaneously administered with PBS, LNP-mOVA (10 μg mOVA per mouse), L@Mn-mOVA (5 μg mOVA per mouse), or L@Mn-mOVA (10 μg mOVA per mouse) at the bilateral inguinal regions on Days 4, 7, and 10. The PBS group served as a negative control. Tumor dimensions were measured every other day using vernier calipers, and tumor volumes were calculated as length × width2 × 0.5. On Day 17, mice were sacrificed, and tumors were harvested, and digested into single-cell suspensions, and analyzed by flow cytometry to characterize immune cell infiltration. Cells were stained for 30 min at 4 °C with FITC anti-mouse CD3 (1:100), PE-Cy7 anti-mouse CD8a (1:100), APC anti-mouse CD4 (1:100), and APC anti-mouse CD11c (1:100), and then analyzed using a flow cytometer (ACEA NovoExpress).
Blood and spleens were collected to assess antigen-specific T cell responses. Single-cell suspensions were stained with T-Select H-2Kb OVA Tetramer-SIINFEKL-PE (1:100) and PE-Cy7 anti-mouse CD8a (1:100), followed by flow cytometric analysis (ACEA NovoExpress). Spleens were further processed for IFN-γ ELISpot assays. Briefly, splenocytes (1 × 105 cells per well) were seeded into 96-well plates pre-coated with anti-mouse IFN-γ antibodies and stimulated with SIINFEKL peptide for 20 h. After incubation, plates were sequentially treated with a biotinylated detection antibody and streptavidin-conjugated ALP. Spots were developed using a substrate solution and quantified using an automated ELISpot reader (Dakewe Biotech).
To evaluate the biosafety of the L@Mn-mRNA vaccines, blood and major organs (heart, liver, spleen, lung, and kidney) were collected and subjected to H&E staining for examination.
Long-term immune memory test
The C57BL/6 mice (6–8 weeks, female) were subcutaneously administered with PBS, LNP-mOVA (10 μg mOVA per mouse), L@Mn-mOVA (5 μg mOVA per mouse), and L@Mn-mOVA (10 μg mOVA per mouse) at the right bilateral inguinal regions on Day 0, 3, and 6. To analyze memory T cells, splenocytes were collected at Day 70 (n = 5) and stained with FITC anti-mouse CD3 (1:100), PE-Cy7 anti-mouse CD8 (1:100), APC anti-mouse CD44 (1:100), and PE anti-mouse CD62L (1:100) antibodies. The central memory T cells (CD3+CD8+CD44highCD62Lhigh) and effector memory T cells (CD3+CD8+CD44highCD62Llow) were analyzed by flow cytometry (ACEA NovoExpress).
Anti-PEG IgG and IgM measurement
The C57BL/6 mice (6–8 weeks, female) were subcutaneously administered with PBS, LNP-mOVA (10 μg mOVA per mouse), L@Mn-mOVA (5 μg mOVA per mouse), and L@Mn-mOVA (10 μg mOVA per mouse) at the right bilateral inguinal regions on Day 0, 3, and 6. To detect Anti-peg IgM and Anti-peg IgG, blood of each group was respectively collected at Day −1, 13, and Day 20. After allowing the whole blood to stand and then centrifuging it to obtain plasma, the plasma was diluted 500-fold. ELISA assay kits are utilized to separately measure the antibody titers of Anti-peg IgM and Anti-peg IgG. Briefly, 100 μL diluted samples was dispensed into the coated wells. Then, 100 μL diluted HRP conjugate was added into each well. After dispensing 100 μL TMB reagent into each well, the plate was gently mix for 20 min. Finally, 100 μL stop solution was added, and the results were collected at a plate reader.
Statistical analysis
All results are presented as the mean ± S.D. from at least three independent experiments. Statistical differences between two groups were determined by two-tailed t-test, and the differences among three or more groups were determined by one-way ANOVA with the Tukey multiple comparison post-test. P values of P < 0.05 were regarded as significant. Animal survival rates were compared with the log-rank test using GraphPad Prism 9.0 (GraphPad Software, San Diego, CA, USA).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the conclusions of this study are included in the manuscript and supplementary materials. Source data are provided with this paper. Source data is available for Figs. 1–7 and Supplementary Figs. 1–28 in the associated source data file Source data are provided with this paper.
References
Fang, E. et al. Advances in COVID-19 mRNA vaccine development. Signal Transduct. Target. Ther. 7, 94 (2022).
Chaudhary, N., Weissman, D. & Whitehead, K. A. mRNA vaccines for infectious diseases: principles, delivery and clinical translation. Nat. Rev. Drug Discov. 20, 817–838 (2021).
Szabó, G. T., Mahiny, A. J. & Vlatkovic, I. COVID-19 mRNA vaccines: platforms and current developments. Mol. Ther. 30, 1850–1868 (2022).
Ye, Z. et al. The mRNA vaccine revolution: COVID-19 has launched the future of vaccinology. ACS Nano 17, 15231–15253 (2023).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Wang, X. et al. Preparation of selective organ-targeting (SORT) lipid nanoparticles (LNPs) using multiple technical methods for tissue-specific mRNA delivery. Nat. Protoc. 18, 265–291 (2023).
Xue, L. et al. High-throughput barcoding of nanoparticles identifies cationic, degradable lipid-like materials for mRNA delivery to the lungs in female preclinical models. Nat. Commun. 15, 1884 (2024).
Lokugamage, M. P. et al. Optimization of lipid nanoparticles for the delivery of nebulized therapeutic mRNA to the lungs. Nat. Biomed. Eng. 5, 1059–1068 (2021).
Lian, X. et al. Bone-marrow-homing lipid nanoparticles for genome editing in diseased and malignant haematopoietic stem cells. Nat. Nanotechnol. 19, 1409–1417 (2024).
Suberi, A. et al. Polymer nanoparticles deliver mRNA to the lung for mucosal vaccination. Sci. Transl. Med. 15, eabq0603 (2023).
Zong, Y., Lin, Y., Wei, T. & Cheng, Q. Lipid Nanoparticle (LNP) enables mRNA delivery for cancer therapy. Adv. Mater. 35, 2303261 (2023).
Miao, L. et al. Delivery of mRNA vaccines with heterocyclic lipids increases anti-tumor efficacy by STING-mediated immune cell activation. Nat. Biotechnol. 37, 1174–1185 (2019).
Sayour, E. J., Boczkowski, D., Mitchell, D. A. & Nair, S. K. Cancer mRNA vaccines: clinical advances and future opportunities. Nat. Rev. Clin. Oncol. 21, 489–500 (2024).
Barbier, A. J., Jiang, A. Y., Zhang, P., Wooster, R. & Anderson, D. G. The clinical progress of mRNA vaccines and immunotherapies. Nat. Biotechnol. 40, 840–854 (2022).
Wang, C., Zhang, Y. & Dong, Y. Lipid Nanoparticle–mRNA formulations for therapeutic applications. Acc. Chem. Res. 54, 4283–4293 (2021).
Chen, K. et al. mRNA vaccines against SARS-CoV-2 variants delivered by lipid nanoparticles based on novel ionizable lipids. Adv. Funct. Mater. 32, 2204692 (2022).
Escalona-Rayo, O. et al. In vitro and in vivo evaluation of clinically-approved ionizable cationic lipids shows divergent results between mRNA transfection and vaccine efficacy. Biomed. Pharmacother. 165, 115065 (2023).
Szebeni, J. et al. Applying lessons learned from nanomedicines to understand rare hypersensitivity reactions to mRNA-based SARS-CoV-2 vaccines. Nat. Nanotechnol. 17, 337–346 (2022).
Bitounis, D., Jacquinet, E., Rogers, M. A. & Amiji, M. M. Strategies to reduce the risks of mRNA drug and vaccine toxicity. Nat. Rev. Drug Discov. 23, 281–300 (2024).
Trougakos, I. P. et al. COVID-19 mRNA vaccine-induced adverse effects: unwinding the unknowns. Trends Mol. Med. 28, 800–802 (2022).
Ndeupen, S. et al. The mRNA-LNP platform’s lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. iScience 24, 103479 (2021).
FACT SHEET FOR HEALTHCARE PROVIDERS ADMINISTERING VACCINE (VACCINATION PROVIDERS) EMERGENCY USE AUTHORIZATION (EUA) OF THE MODERNA COVID-19 VACCINE TO PREVENT CORONAVIRUS DISEASE 2019 (COVID-19). https://www.fda.gov/media/144637/download (2022).
Li, B. et al. Accelerating ionizable lipid discovery for mRNA delivery using machine learning and combinatorial chemistry. Nat. Mater. 23, 1002–1008 (2024).
Riley, R. S. et al. Ioni zable lipid nanoparticles for in utero mRNA delivery. Sci. Adv. 7, eaba1028 (2021).
Zhu, Y. et al. Screening for lipid nanoparticles that modulate the immune activity of helper T cells towards enhanced antitumor activity. Nat. Biomed. Eng. 8, 544–560 (2024).
Sun, Y. et al. In vivo editing of lung stem cells for durable gene correction in mice. Science 384, 1196–1202 (2024).
Metkar, M., Pepin, C. S. & Moore, M. J. Tailor made: the art of therapeutic mRNA design. Nat. Rev. Drug Discov. 23, 67–83 (2024).
Bicknell, A. A. Attenuating ribosome load improves protein output from mRNA by limiting translation-dependent mRNA decay. Cell Reports 43, 114098 (2024).
Hajj, K. A. & Whitehead, K. A. Tools for translation: non-viral materials for therapeutic mRNA delivery. Nat. Rev. Mater. 2, 17056 (2017).
Chen, S. et al. Nanotechnology-based mRNA vaccines. Nat. Rev. Methods Prim. 3, 63 (2023).
Li, M. et al. Engineering multifunctional DNA hybrid nanospheres through coordination-driven self-assembly. Angew. Chem. Int. Ed. 58, 1350–1354 (2019).
Liu, C. et al. Self-assembly of copper–DNAzyme nanohybrids for dual-catalytic tumor therapy. Angew. Chem. 133, 14445–14449 (2021).
Zou, Z. et al. Zn2+-coordination-driven RNA assembly with retained integrity and biological functions. Angew. Chem. 133, 23152–23158 (2021).
Lim, K. S., Lee, D. Y., Valencia, G. M., Won, Y.-W. & Bull, D. A. Nano-self-assembly of nucleic acids capable of transfection without a gene carrier. Adv. Funct. Mater. 25, 5445–5451 (2015).
Muntean, C. M., Nalpantidis, K., Feldmann, I. & Deckert, V. Zn2+–DNA interactions in aqueous systems: a Raman spectroscopic study. Spectroscopy 23, 155–163 (2009).
Martin, R. B. Nucleoside sites for transition metal ion binding. Acc. Chem. Res. 18, 32–38 (1985).
Berti, L. & Burley, G. A. Nucleic acid and nucleotide-mediated synthesis of inorganic nanoparticles. Nat. Nanotechnol. 3, 81–87 (2008).
Schoenmaker, L. et al. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int. J. Pharm. 601, 120586 (2021).
Hui, Y. et al. Understanding the effects of nanocapsular mechanical property on passive and active tumor targeting. ACS Nano 12, 2846–2857 (2018).
Anselmo, A. C. & Mitragotri, S. Impact of particle elasticity on particle-based drug delivery systems. Adv. Drug Deliv. Rev. 108, 51–67 (2017).
Wang, S. & Lv, Y. Silica-coated liquid metal nanoparticles with different stiffness for cellular uptake-enhanced tumor photothermal therapy. Biomater. Adv. 161, 213872 (2024).
Song, Y. et al. Manganese enhances the antitumor function of CD8+ T cells by inducing type I interferon production. Cell. Mol. Immunol. 18, 1571–1574 (2021).
Zhang, R. et al. Manganese salts function as potent adjuvants. Cell. Mol. Immunol. 18, 1222–1234 (2021).
Zhou, M. et al. Manganese-enriched Zinc peroxide functional nanoparticles for potentiating cancer immunotherapy. Nano Lett. 23, 10350–10359 (2023).
Sun, X. et al. Amplifying STING activation by cyclic dinucleotide–manganese particles for local and systemic cancer metalloimmunotherapy. Nat. Nanotechnol. 16, 1260–1270 (2021).
Dosta, P. et al. Investigation of the enhanced antitumor potency of STING agonist after conjugation to polymer nanoparticles. Nat. Nanotechnol. 18, 1351–1363 (2023).
Kuchitsu, Y. et al. STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nat. Cell Biol. 25, 453–466 (2023).
Lv, M. et al. Manganese is critical for antitumor immune responses via cGAS-STING and improves the efficacy of clinical immunotherapy. Cell Res. 30, 966–979 (2020).
Wang, C. et al. Manganese increases the sensitivity of the cGAS-STING pathway for double-stranded DNA and is required for the host defense against DNA viruses. Immunity 48, 675–687.e677 (2018).
Miah, M. R. et al. The effects of manganese overexposure on brain health. Neurochem. Int. 135, 104688 (2020).
Tsuji, T. et al. Elemental concentration of manganese and potassium in brain and other organs of fetal, sucking and developmental mice. J. Radioanal. Nucl. Chem. 258, 49–53 (2003).
Rucker, R. B., Fascetti, A. J. & Keen, C. L. Chapter 22 - Trace minerals. In Clinical Biochemistry of Domestic Animals (Sixth Edition) (eds Kaneko, J. J., et al.) 663–693 (Academic Press, San Diego, 2008).
Seo, Y. A., Elkhader, J. A. & Wessling-Resnick, M. Distribution of manganese and other biometals in flatiron mice. BioMetals 29, 147–155 (2016).
Singh, M. Effect of fluoride on tissue manganese levels in the mouse. Sci. Total Environ. 22, 285–288 (1982).
Frisch, M. et al. Gaussian. http://www.gaussian.com/ (2009).
Lu, T. & Chen, F. Multiwfn: a multifunctional wavefunction analyzer. J. Comput. Chem. 33, 580–592 (2012).
Chai, J.-D. & Head-Gordon, M. Long-range corrected hybrid density functionals with damped atom–atom dispersion corrections. Phys. Chem. Chem. Phys. 10, 6615–6620 (2008).
Weigend, F. & Ahlrichs, R. Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: design and assessment of accuracy. Phys. Chem. Chem. Phys. 7, 3297–3305 (2005).
Petersson, A. et al. A complete basis set model chemistry. I. The total energies of closed-shell atoms and hydrides of the first-row elements. J. Chem. Phys. 89, 2193–2218 (1988).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFA0909900), the Beijing Natural Science Foundation (Z220022), and the National Natural Science Foundation of China (32271391). We sincerely acknowledge the valuable discussions with Ms. Yike Wu, Dr. Na Chen, Dr. Shuai Liu, Dr. Guangna Liu, and Dr. Keman Cheng. Elements of Figs. 1–4 and Supplementary Figs. 4 and 6 were created with BioRender.com and had got the publication license.
Author information
Authors and Affiliations
Contributions
X.M., T.J., and G.N. conceived the idea, analyzed the data, and wrote the manuscript. X.M., S.L., A.L., S.Z., Zongran Liu, M.Z., Zhongxian Li, J.Z., Junxi Li, Q.C., W.L. performed experiments. H.Q., Jiaqi Lin, and X.L. provided advice on in vivo experiments. X.S. and H.W. supervised and conducted the computational simulation portion. X.Z. and Junxi Li guided and implemented the cryo-electron microscopy section. All authors contributed to the discussion and editing of manuscript. Z.G., H.L., and L.L. provided suggestions for the overall work.
Corresponding authors
Ethics declarations
Competing interests
X.M. and T.J. have filed the patent application on this research. The other authors declare no conflict of interest.
Peer review
Peer review information
Nature Communications thanks Hao Song and Ronghua Yang for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, X., Liu, S., Zhang, S. et al. Engineering of mRNA vaccine platform with reduced lipids and enhanced efficacy. Nat Commun 16, 8913 (2025). https://doi.org/10.1038/s41467-025-63965-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63965-3