Abstract
Nucleotide-binding leucine-rich repeat receptors (NLRs) function as core components of innate immunity in both plants and animals. In animals, NLR activation initiates caspase-mediated immune signaling. In contrast, plants lack caspases but instead contain metacaspases (MCAs/MCs), yet their role in antiviral immunity and whether they interface with NLR signaling remain largely unexplored. Here, we demonstrate that cleavage of the conserved immune regulator Bcl-2-associated athanogene 3 (BAG3) by metacaspase 4 (MCAIIa/MC4) induces cell death and activates antiviral immunity in plants. Upon Begomovirus infection, MC4 cleaves BAG3 to release its N-terminal functional domain (BAG3-N) from autoinhibition. BAG3-N assembles into oligomers and induces cell death, effectively inhibiting viral replication. This signaling also interfaces with NLR networks in certain plant species. Viral replication-associated proteins (Reps) counteract this defense response by binding to BAG3-N, highlighting an evolutionary arms race between plants and viruses. Evolutionary analyses reveal that a lysine substitution at position 50 of BAG3 confers its ability to induce cell death in angiosperms. These findings identify BAG3 as a conserved immune regulator linking metacaspase activation to antiviral defense, providing a mechanistic basis for engineering crops with enhanced resistance to insect-borne viruses.
Similar content being viewed by others
Introduction
Plants have evolved a complex immune system that employs cell-surface pattern recognition receptors (PRRs) and intracellular nucleotide-binding leucine-rich repeat receptors (NLRs) to combat pathogens1. PRRs detect pathogen-associated molecular patterns (PAMPs) and initiate PAMP-triggered immunity (PTI)2. In addition, plants detect insect herbivory through herbivore-associated molecular patterns (HAMPs) and damage-associated molecular patterns (DAMPs), initiating PTI-like immune responses3. However, pathogenic microbes and herbivores often deliver effectors into plant cells to suppress PTI and promote virulence4. To counteract such suppression, plants have evolved NLRs to trigger a more robust immune response known as effector-triggered immunity (ETI)1,5. NLRs can be functionally divided into sensor NLRs, which detect effectors, and helper NLRs, which transmit immune signals to trigger responses and typically induce programmed cell death (PCD) to restrict pathogen spread6.
In animals, caspases orchestrate PCD via cleavage of substrates such as gasdermins, leading to pyroptosis7,8. In contrast, plants lack canonical caspases and instead rely on metacaspases (MCAs/MCs), which play crucial roles in regulating PCD, particularly under stress conditions9. Arabidopsis thaliana encodes nine MCs, divided into type I (MC1-3/MCAIa-c) and type II (MC4-9/MCAIIa-f)10. Among these, MC4 cleaves the DAMP precursor PROPEP1 in a Ca²⁺-dependent manner, releasing the signaling peptide Pep1, which is recognized by the receptor PEPR1 to initiate defense responses upon mechanical damage10,11. Notably, PEPR1 is strongly upregulated upon whitefly (Bemisia tabaci) infestation12, suggesting that herbivory-induced stress may activate MC4 and enhance Pep1 signaling through PROPEP1 cleavage.
While MCs are primarily involved in early immune responses10,11, emerging evidence indicates that certain MCs also engage with intracellular NLRs13, pointing to a broader role in ETI. However, the downstream mechanisms linking metacaspase activity to immune execution remain poorly understood. Bcl-2-associated athanogene (BAG) family proteins are conserved regulators of PCD across eukaryotes14. In Arabidopsis, BAG6 is involved in autophagy-dependent cell death and defense against necrotrophic fungi15,16,17,18, while in rice, BAG4 contributes to defense against both bacterial and fungal pathogens19. Recent studies have identified BAGs as modulators of immune signaling, yet whether they couple metacaspase activity to immune execution remains unknown.
Geminiviruses are the largest family of plant viruses and comprise several genera, among which Begomoviruses are the most widespread and economically significant20. A notable member of this family is Tomato yellow leaf curl virus (TYLCV), a monopartite begomovirus transmitted by whitefly through a piercing-sucking feeding strategy. These single-stranded DNA viruses encode six proteins, with C1 serving as the replication-associated protein (Rep) essential for viral genome replication21. Like many viruses, Begomoviruses have evolved diverse strategies to suppress host immune responses22,23. However, whether they target metacaspase-dependent immune signaling to promote infection remains unknown.
Here, we identify BAG3 as a conserved immune regulator that connects metacaspase activity to antiviral immunity. Upon begomovirus infection, MC4 cleaves BAG3, releasing its N-terminal domain from autoinhibition. The released BAG3-N oligomerizes and triggers cell death, thereby restricting viral replication. Begomovirus Rep proteins counteract this response by binding BAG3-N and suppressing its activity, revealing a viral counter-defense strategy. These findings uncover a metacaspase-dependent antiviral pathway and highlight BAG3 as a potential target for engineering virus-resistant crops.
Results
MC4 is activated by herbivory and shifts substrate preference upon viral infection
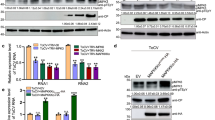
To investigate whether biotic stress, particularly the piercing-sucking feeding behavior of whiteflies, activates MC4, we examined the expression patterns of MCs. RNA sequencing revealed a marked upregulation of Arabidopsis MC4 expression following whitefly infestation (Fig. 1a). This upregulation was further confirmed by quantitative RT-PCR (qRT-PCR) (Fig. 1b). Under natural conditions, whiteflies transmit TYLCV. When plants were infected with TYLCV, MC4 expression was further elevated, whereas other MCs did not exhibit a similar increase (Fig. 1b). Western blot analysis revealed enhanced self-cleavage of MC4 in response to TYLCV infection (Fig. 1c and Supplementary Fig. 1a), indicating that MC4 undergoes auto-activation upon viral infection. These results suggest that MC4 plays an important role in plant defense against insects and viruses.
a Heat map depicting the expression profile of the Arabidopsis metacaspase genes (MCs) following 3 days of whitefly infestation. Expression levels are shown as log2(TPM + 1), averaged across three biological replicates. Color scale represents average intensity values in log2(TPM + 1). b Expression levels of Arabidopsis metacaspase genes were quantified by RT-qPCR after 3 days of whitefly infestation and TYLCV infection. Values are means ± standard deviation (SD) (n = 6). Data were analyzed by two-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. The experiment consisted of three biological replicates. c TYLCV infection promotes the activation of MC4. MC4-Flag and MC4C139A-Flag were transiently expressed in Mock- and TYLCV-treated N. benthamiana plants. Protein detections were performed using anti-Flag (α-Flag) and anti-CP (α-CP) antibodies. Coomassie brilliant blue (CBB) stained bands of the large subunit of Rubisco (rbcL) were used as a loading control. Three biological replicates were conducted. d Whitefly infestation promotes the processing of PROPEP1-YFP, and TYLCV infection disturbs this process. Protein detection was performed using an anti-GFP antibody. CBB-stained bands of the rbcL were used as a loading control. Three biological replicates were conducted. e The interaction between MC4 and BAG3 was assessed using a yeast two-hybrid (Y2H) system. The empty vectors pGBT9 and pGAD424 were used as negative controls. f Co-immunoprecipitation (Co-IP) analysis of MC4C139A-Flag and YFP-AtBAG3 in vivo. Total protein was extracted from N. benthamiana leaves transiently expressing 35S: GUS-Flag and 35S: MC4C139A-Flag together with 35S: YFP-AtBAG3. GFP-trap beads were used to precipitate the interaction complex. Anti-GFP (α-GFP) and anti-Flag (α-Flag) antibodies were used for detection. The experiment was repeated independently three times with similar results.
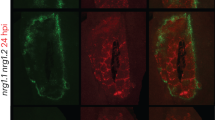
To examine whether whitefly infestation promotes Pep1 maturation, we treated PROPEP1-YFP and PROPEP1-YFP/mc4 Arabidopsis plants with whitefly infestation. Whitefly infestation significantly elevated the expression of PROPEP1 and promoted its cleavage (Fig. 1d and Supplementary Fig. 1b). However, under combined whitefly infestation and viral infection, although MC4 expression level was further induced, its ability to cleave PROPEP1 was markedly reduced. In the PROPEP1-YFP/mc4 plants, PROPEP1 processing was significantly diminished (Fig. 1d and Supplementary Fig. 1b). MC4 promotes PROPEP1 cleavage in response to whitefly infestation alone, but this cleavage is suppressed under combined whitefly and viral stress. These findings suggest that under the combined stress of herbivory and viral infection, MC4 may shift substrate preference to optimize the immune response.
To uncover the potential substrates of MC4, we performed a yeast two-hybrid (Y2H) screen using MC4 as bait against an Arabidopsis cDNA library. Among the candidates, Arabidopsis BAG3 (AtBAG3) was notable given the proposed role of plant BAGs as positive regulators of cell death and immunity24. Phylogenetic analysis of BAG3 homologs across representative plant species revealed that BAG3 belongs to a well-conserved clade in angiosperms (Supplementary Fig. 2). The interaction between MC4 and AtBAG3 was first confirmed using Y2H assays (Fig. 1e). In co-immunoprecipitation (Co-IP) assays, the catalytically inactive mutant MC4C139A-Flag was co-immunoprecipitated by YFP-AtBAG3 (Fig. 1f). Moreover, bimolecular fluorescence complementation (BiFC) assays confirmed the interaction between MC4 and AtBAG3 in planta (Supplementary Fig. 3). Taken together, these results demonstrate that MC4 interacts with AtBAG3 in vitro and in vivo.
BAG3 modulates plant antiviral immunity through N-terminal domain-mediated cell death
To investigate the role of BAG3 in plant antiviral immunity, we generated Atbag3 knockout and YFP-AtBAG3 overexpression Arabidopsis lines (Supplementary Fig. 4a, b). Upon TYLCV infection, AtBAG3 overexpression plants exhibited severe necrosis at 21 days post inoculation (dpi) (Fig. 2a). Trypan blue staining confirmed extensive cell death in AtBAG3 overexpression plants (Fig. 2b). Quantitative PCR (qPCR) further revealed markedly reduced viral titers in AtBAG3 overexpression lines relative to Atbag3 mutants (Fig. 2c). Given that TYLCV primarily infects Solanaceae, with tomato (Solanum lycopersicum) as its principal host, we identified BAG3 homologs in tomato (Fig. 2d) and generated Slbag1/3/3-like (Slbag1/3/3 l) triple mutants (Supplementary Fig. 5a). Under normal conditions, no visible phenotypic differences were observed between Slbag1/3/3 l plants and wild-type (WT) plants (Supplementary Fig. 5b). Upon TYLCV infection, WT plants developed necrotic lesions and chlorosis, whereas Slbag1/3/3 l mutants lacked necrosis but exhibited pronounced leaf curling and stunted growth at 14 dpi (Fig. 2e). Necrosis and chlorosis typically indicate localized defense activation, whereas leaf curling and dwarfing are indicative of systemic infection and developmental disruption25,26. The observed symptoms in Slbag1/3/3 l plants suggest a weakened antiviral response. Consistently, viral titers were markedly elevated in Slbag1/3/3 l plants (Fig. 2f). Together, these findings establish BAG3 as a positive regulator of antiviral immunity in both Arabidopsis and tomato.
a Phenotypes of leaves from Arabidopsis Col-0, Atbag3 #9, and YFP-AtBAG3 #6 lines mock-treated or TYLCV-infected at 21 days post infiltration (dpi). The leaves are arranged from oldest (left) to youngest (right). The red arrows indicate the necrotic lesions. The experiment was repeated independently three times with similar results. b Trypan blue staining was used to assess cell death in leaves of Col-0, Atbag3 #9, and YFP-AtBAG3 #6 at 21 dpi with TYLCV infection. The experiment was repeated three times with similar results. c Quantification of viral genomic DNA by qPCR in Col-0, Atbag3 #9, and YFP-AtBAG3 #6 at 21 dpi post TYLCV infection. Values are means ± SD of six independent plants (n = 6) from one representative experiment. Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. d Schematic representation of the structure of the AtBAG3 and SlBAG1/3/3 l proteins. UbL, ubiquitin-like domain; BAG, BAG domain; Arginine, putative MC4 cleavage site. e Symptoms of TYLCV-infected WT and Slbag1/3/3 l #3 and #18 triple mutant tomato plants at 14 dpi. Scale bars = 1 cm. Three biological replicates were performed. The white arrows indicate the necrotic lesions. f Quantification of viral genomic DNA in systemic leaves of plants shown in Fig. 2e by qPCR. Values are means ± SD of eight independent plants (n = 8) from one representative experiment. P-values were calculated by an unpaired two-tailed Student’s t test. The experiment was repeated three times with similar results. g Cell death phenotypes induced by the empty vector (nEYFP), full-length AtBAG3, and domain truncations (AtBAG3-N and AtBAG3-C). The empty vector (nEYFP) was used as a negative control. Leaves were infiltrated in N. benthamiana and stained with trypan blue 2-3 days post-infiltration. Images under white and UV light are shown. Scale bars = 1 cm. The experiment was repeated three times with similar results. h Immunoblot detection of BAG3 and TYLCV coat protein (CP) in samples from Fig. 2e. Anti-BAG3 and anti-CP (α-CP) antibodies were used. Stained bands of rbcL were used as a loading control. The arrow indicates the putative cleaved product of SlBAG1/3/3 l. NS, nonspecific bands. The experiment was repeated three times with similar results. i Quantification of cell death induced by transient expression of AtBAG3, AtBAG3-N, and AtBAG3-C in N. benthamiana. Electrolyte leakage was measured from leaf disks at 2 dpi (n = 5, five biological independent samples from one representative experiment). Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. In box and whiskers, the center line indicates the median, the bounds of the box show the 25th and 75th percentiles, the whiskers indicate the minimum to maximum values. The experiments were performed three times with similar results. j Cell death phenotypes of AtBAG3-N truncations in N. benthamiana leaves. The empty vector (nEYFP) was used as a negative control. Infiltrated leaves were stained with trypan blue, and representative photographs were taken at 2-3 dpi. Images under white and UV light are shown. The experiments were performed three times with similar results. k Quantification of cell death induced by transient expression of AtBAG3-N truncations in N. benthamiana. Electrolyte leakage of leaf disks from N. benthamiana was measured at 2 dpi (n = 5, five biological independent samples from one representative experiment). Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. In box and whiskers, the center line indicates the median, the bounds of the box show the 25th and 75th percentiles, the whiskers indicate the minimum to maximum values. The experiments were performed three times with similar results.
The observed differences in necrosis between WT and bag mutant plants upon TYLCV infection prompted us to examine whether BAG3 directly mediates cell death. To verify the potential role of BAG3 in inducing cell death, we generated constructs encoding the full-length AtBAG3 protein. No cell death was observed in Nicotiana benthamiana (N. benthamiana) leaves transiently expressing AtBAG3 (Fig. 2g). Western blot analysis of SlBAG1/3/3 l expression in TYLCV-infected tomato plants (Fig. 2e) revealed the appearance of a distinct band at approximately 15 kDa beneath the major SlBAG1/3/3 l bands in TYLCV-infected samples (Fig. 2d, h). This band was consistently absent in mock-treated controls and Slbag1/3/3 l mutants. These findings suggest that SlBAG1, SlBAG3, and SlBAG3l may undergo proteolytic cleavage in response to viral infection, generating a smaller cleavage product.
Since BAG3 consists of an N-terminal ubiquitin-like (UbL) domain and a C-terminal BAG domain16, and both BAG3-N and BAG3-C fragments are approximately 15 kDa, we hypothesized that one of these fragments might induce cell death. To test this, we generated truncations encoding N-terminal (AtBAG3-N) and C-terminal (AtBAG3-C) domains. Notably, AtBAG3-N (1-122 amino acids, aa) was sufficient to induce cell death (Fig. 2g and Supplementary Fig. 6a). The increased ion conductivity indicated that AtBAG3-N expression exacerbated membrane damage (Fig. 2i). Given that the protein sequences of AtBAG3-N, SlBAG1-N, SlBAG3-N and SlBAG3l-N exhibit an 83.44% identity in their alignments (Supplementary Fig. 7a), we further explored the functional roles of their N-terminal domains by generating truncated versions of SlBAG1, SlBAG3 and SlBAG3l proteins. The results showed that the N-terminal domains of SlBAG1, SlBAG3 and SlBAG3l were able to induce cell death (Supplementary Fig. 7b–d), indicating that this domain acts as the key executor of BAG3-mediated cell death.
To identify the minimal functional domain of BAG3 required for cell death induction, we generated a series of truncation constructs and transiently expressed them in N. benthamiana leaves. C-terminal truncation analysis demonstrated that AtBAG3 retains its cell death-inducing activity until residue arginine (Arg, R) 122, beyond which the activity is completely abolished (Supplementary Fig. 8a, c). To further delineate the critical region, we systematically performed N-terminal truncations with single-residue resolution (Supplementary Fig. 8b, c), ultimately identifying the segment spanning R46 to R122 as the minimal functional domain essential for cell death induction (Fig. 2j, k and Supplementary Fig. 6b). These findings highlight BAG3-N as the core signaling domain responsible for BAG3-induced cell death, with R46-R122 identified as the minimal functional region essential for triggering cell death.
MC4 cleaves BAG3 at R122
To investigate how BAG3 activity is regulated under resting conditions, we hypothesized that it adopts an auto-inhibited conformation via intra-molecular interactions between its N- and C-terminal domains. Co-expression of AtBAG3-N with AtBAG3-C in N. benthamiana revealed that AtBAG3-C suppressed AtBAG3-N-induced cell death (Fig. 3a), supporting an inhibitory role for the C-terminus. Notably, co-expression of AtBAG3-N with an empty vector still triggered cell death (Fig. 3a), indicating that endogenous N. benthamiana NbBAG3-C is insufficient to inhibit the activity of exogenous AtBAG3-N. Structural predictions using AlphaFold 3 indicated a direct interaction between AtBAG3-N and AtBAG3-C (Fig. 3b). Co-IP assays confirmed the inter-domain interaction of BAG3 (Fig. 3c). Moreover, BiFC assays further confirmed the interaction between BAG3-N and BAG3-C in planta (Supplementary Fig. 9). Together, these results demonstrate that AtBAG3 adopts an auto-inhibited conformation mediated by direct interaction between its N- and C-terminal regions under resting conditions.
a Cell death phenotypes of AtBAG3-N co-infiltrated with AtBAG3-C in N. benthamiana leaves and quantification of the cell death. nEYFP was used as a negative control. Representative images under UV light were taken at 2 dpi. Electrolyte leakage of leaf disks from N. benthamiana was measured at 2 dpi (n = 5). P-values were calculated by an unpaired two-tailed Student’s t test. The experiment was repeated independently three times with similar results. b The interaction model between AtBAG3-N (blue) and AtBAG3-C (pink) was predicted using AlphaFold 3. The stick model represents the potential interaction sites between AtBAG3-N and AtBAG3-C; this region is predicted with a high confidence score. c Co-IP analysis of AtBAG3-N-Flag and nEYFP-AtBAG3-C in vivo. Total proteins were extracted from N. benthamiana leaves transiently expressing 35S: nEYFP and 35S: nEYFP-AtBAG3-C together with 35S: AtBAG3-N-Flag. Flag-trap beads were used to precipitate the interaction complex, and anti-GFP (α-GFP) and anti-Flag (α-Flag) antibodies were used for detection. The experiments were performed three times with similar results. d Cell death phenotypes of MC4 co-infiltrated with AtBAG3 in N. benthamiana leaves. GUS-Flag and empty vector (nEYFP) were used as negative controls. Infiltrated leaves were stained with trypan blue, and representative photographs were taken at 2-3 dpi. Images under white and UV light are shown. The experiment was repeated independently three times with similar results. e Quantification of cell death induction of MC4 and AtBAG3 in N. benthamiana. Electrolyte leakage of leaf disks from N. benthamiana was measured at 2 dpi (n = 5). Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. In box and whiskers, the center line indicates the median, the bounds of the box show the 25th and 75th percentiles, the whiskers indicate the minimum to maximum values. The experiment was repeated independently three times with similar results. f Protein detection of Fig. 3d was performed using anti-Flag (α-Flag) and anti-GFP (α-GFP) antibodies. Stained bands of rbcL were used as a loading control. The experiment was repeated independently three times with similar results. g Only MC4, but not the catalytically inactive MC4C139A variant, can cleave AtBAG3 in N. benthamiana leaves. Replacement of AtBAG3 R122 with A or K impaired AtBAG3 cleavage in vivo. Stained bands of rbcL were used as a loading control. The experiment was repeated independently three times with similar results. h Only MC4 but not catalytically inactive MC4C139A variant can cleave AtBAG3 in vitro. MC4 cleaves AtBAG3 after residue R122. The experiment was repeated independently three times with similar results. i Top: cell death phenotypes of MC4 and the catalytically inactive MC4C139A variant co-infiltrated respectively with AtBAG3, AtBAG3R122A, and AtBAG3R122K in N. benthamiana leaves. Representative photographs were taken at 2-3 dpi. Images under UV light are shown. Bottom: quantification of cell death induction by MC4 and catalytically inactive MC4C139A variant co-infiltrated respectively with AtBAG3, AtBAG3R122A and AtBAG3R122K in N. benthamiana. Electrolyte leakage of leaf disks from N. benthamiana was measured at 2 dpi (n = 5). Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. In box and whiskers, the center line indicates the median, the bounds of the box show the 25th and 75th percentiles, the whiskers indicate the minimum to maximum values. The experiment was repeated independently three times with similar results.
Type II metacaspases, such as MC4, are known to cleave substrates after arginine (R) or lysine (K) residues27. The previously identified minimal cell death-inducing region of AtBAG3 spans residues R46-R122 (Fig. 2j, k). Considering the interaction between MC4 and AtBAG3, we hypothesized that MC4-mediated cleavage may release the active N-terminal domain of AtBAG3. Indeed, co-expression of AtBAG3 with MC4 induced cell death (Fig. 3d, e). Western blot analysis revealed that MC4 undergoes self-cleavage and cleaves AtBAG3, producing a product consistent with the size of AtBAG3-N (Fig. 3f).
To exclude potential interference from endogenous components, we identified N. benthamiana BAG3 homologs NbBAG1, NbBAG3, and NbBAG3-like (NbBAG3l) by sequence alignment and silenced them simultaneously using virus-induced gene silencing (VIGS) (Supplementary Fig. 10a). Silencing of NbBAG1/3/3l had no effect on AtBAG3/MC4-induced cell death (Supplementary Fig. 10b). In addition, co-expression of AtBAG3 or MC4 with empty vector did not induce cell death or AtBAG3 cleavage (Fig. 3d–f), confirming that endogenous NbBAG1/3/3 l do not interfere with AtBAG3/MC4-induced cell death under these conditions. In contrast, the catalytically inactive mutant MC4C139A failed to cleave AtBAG3 (Fig. 3g, h). Moreover, co-expression of AtBAG3 with MC4C139A failed to induce cell death (Fig. 3i), underscoring the requirement of MC4 enzymatic activity for AtBAG3 cleavage and subsequent cell death.
To pinpoint the cleavage site, we co-expressed MC4 with AtBAG3R122A (arginine to alanine) or the homotypic AtBAG3R122K (arginine to lysine) mutant individually. Neither AtBAG3R122A nor AtBAG3R122K triggered cell death when co-expressed with MC4 (Fig. 3i). The Mutated AtBAG3R122A cleavage site was not cleaved by MC4 in vivo. The homotypic AtBAG3R122K mutation was cleaved, albeit less efficiently (Fig. 3g). To further assess the direct dependence of AtBAG3 maturation on MC4, in vitro cleavage assays were performed using recombinant GST-MC4 and AtBAG3-His. Consistent with the in vivo results, mutated AtBAG3R122A-His cleavage sites were not cleaved by MC4 in vitro. Although the homotypic AtBAG3R122K mutation was cleaved, the efficiency was comparatively lower (Fig. 3h). Collectively, these results demonstrate that the precise cleavage at R122 by MC4 is essential for the release of the BAG3-N domain, which in turn induces cell death.
Oligomerization and plasma membrane localization are essential for BAG3-N-induced cell death
Self-interaction is a key mechanism underlying the activation of cell death regulators28,29,30,31,32. To determine whether BAG3-N-induced cell death requires self-interaction, we utilized AlphaFold 3 to predict the self-interaction of AtBAG3-N and identified several putative interaction sites (Fig. 4a). To validate these predictions functionally, we performed site-directed mutagenesis, substituting key residues with alanine (A) individually. Among these, substitution of lysine at position 50 with A (K50A) completely abolished the cell death phenotype, suggesting that K50 is essential for AtBAG3-N-induced cell death (Fig. 4b). Western blot analysis confirmed the correct expression of all proteins (Fig. 4c). Co-IP assays confirmed the self-interaction of AtBAG3-N (Fig. 4d), but revealed dramatically impaired self-interaction for the AtBAG3K50A-N mutant (Supplementary Fig. 11a). Consistently, BiFC assays demonstrated the self-interaction of AtBAG3-N, which was abolished in the AtBAG3K50A mutant (Supplementary Fig. 11b). Given that oligomerization underlies the activation of resistosome complexes such as ZAR1, we tested whether BAG3-N also forms oligomers33. Blue native PAGE (BN-PAGE) analysis showed that AtBAG3-N migrates as high molecular weight bands, potentially indicative of oligomeric assemblies, whereas the AtBAG3K50A mutant exhibited a marked reduction (Fig. 4e). These findings strongly indicate self-association as a prerequisite for BAG3-N oligomerization and subsequent cell death activity.
a The self-interaction model of AtBAG3-N were predicted using AlphaFold 3. AtBAG3-N monomers are represented in pink and purple. The stick model represents the potential self-interaction sites; this region is predicted with a high confidence score. b Cell death phenotypes of AtBAG3-N point mutants transiently expressed in N. benthamiana leaves. Representative photographs were taken at 2-3 dpi. Images under white and UV light are shown. The experiment was repeated independently three times with similar results. c Protein detection of Fig. 4b was performed using an anti-GFP (α-GFP) antibody. Stained bands of rbcL were used as a loading control. d Co-IP analysis of AtBAG3-N-Flag and nEYFP-AtBAG3-N in vivo. Total protein was extracted from N. benthamiana leaves at 2 dpi after transient co-expression of 35S: nEYFP-AtBAG3-N and 35S: AtBAG3-N-Flag. Flag-trap beads were used to precipitate the interaction complex, and anti-GFP (α-GFP) and anti-Flag (α-Flag) antibodies were used for detection. e AtBAG3-N oligomerization state, as detected by BN-PAGE. Square brackets point to oligomers. NS, nonspecific bands (upper image). The protein expression level and loading control were detected by SDS-PAGE and rbcL, respectively (bottom image). f Subcellular localization of YFP-HA, YFP-AtBAG3 and YFP-AtBAG3-N was observed in N. benthamiana at 2 dpi by confocal microscopy. Scale bars = 25 μm. The experiment was repeated three times with similar results. g YFP fluorescence was observed after cell plasmolysis in N. benthamiana leaves. Leaves were treated with 5 % NaCl for 5–10 mins to trigger cell plasmolysis. The cell wall is marked by dotted lines. Scale bars =25 μm. The experiment was repeated three times with similar results. h Total protein (T) was separated into soluble (S) and plasma membrane (PM) fractions. H+-ATPase was used as a plasma membrane marker. Antibodies against GFP and H+-ATPase were used. The experiment was repeated independently three times with similar results.
Proper subcellular localization is crucial for the function of proteins that induce cell death32,34,35. To determine the localization of full-length AtBAG3 and AtBAG3-N, we performed confocal microscopy, which revealed that YFP-AtBAG3-N fluorescence completely overlapped with the plasma membrane (PM) marker CD3-1007-RFP36 (Fig. 4f). In addition, plasmolysis analysis further supported the possible plasma membrane localization of YFP-AtBAG3-N (Fig. 4g). To confirm this, we performed cellular compartment fractionation assays in N. benthamiana leaves expressing YFP-HA, YFP-AtBAG3, YFP-AtBAG3-N and YFP-AtBAG3K50A-N. While full-length AtBAG3 was predominantly detected in soluble fractions, AtBAG3-N was exclusively detected in the plasma membrane fraction, further supporting the plasma membrane localization of AtBAG3-N (Fig. 4h). Moreover, the AtBAG3K50A-N mutation not only impaired oligomerization but also disrupted the plasma membrane localization of AtBAG3-N (Fig. 4e, g, h). Although AtBAG3-N lacks canonical lipid modifications or transmembrane domains, it is localized at the plasma membrane (Fig. 4f–h), suggesting that its recruitment may occur through protein-protein interactions. These findings suggest that both oligomerization and PM localization are essential for BAG3-N-induced cell death.
BAG3-N induces conserved cell death across angiosperms
BAG proteins are a conserved family across eukaryotes14. To assess whether BAG3-N-induced cell death is evolutionarily conserved, we performed sequence alignments to identify homologs of AtBAG3-N across a diverse range of plant species. These included monocots such as Oryza sativa (Os-) and Zea mays (Zm-); dicots such as Solanum lycopersicum (Sl-), Davidia involucrata (Di-), Weddellina squamulosa (Ws-) and Terniopsis brevis (Tb-); gymnosperms such as Picea sitchensis (Ps-); pteridophytes such as Selaginella moellendorffii (Sm-); and bryophytes including the liverwort Marchantia polymorpha (Mp-) and the moss Physcomitrium patens (Pp-). Phylogenetic analysis was conducted for these species (Fig. 5a). To further investigate whether BAG3-N-induced cell death is conserved in monocots, we expressed the N-terminal domains of OsBAG3 and ZmBAG3 in N. benthamiana leaf cells. Both OsBAG3-N and ZmBAG3-N induced cell death (Supplementary Fig 12a–c). In contrast, the N-terminal domains of MpBAG3 and PpBAG3 (MpBAG3-N and PpBAG3-N), derived from bryophytes, failed to trigger cell death (Supplementary Fig. 12d–f). Sequence alignment revealed that a conserved lysine (K) at position 50 in angiosperms is mutated to threonine (T) / serine (S) in gymnosperms and lower plant lineages (Fig. 5a). Mutant variants BAG3K50T-N and BAG3K50S-N failed to induce cell death (Supplementary Fig. 13a, b), indicating that K50 is critical for the function of BAG3-N. Taken together, our findings suggest that the BAG3-N-induced cell death response is conserved across angiosperms and may contribute to broad-spectrum antiviral defense.
a The maximum likelihood phylogenetic tree was generated in MEGA using BAG3-N domain sequences of 13 BAG3 homologs identified from Oryza sativa (Os-), Zea mays (Zm-), Arabidopsis thaliana (At-), Davidia involucrata (Di-), Solanum lycopersicum (Sl-), Weddellina squamulosa (Ws-), Terniopsis brevis (Tb-), Picea sitchensis (Ps-), Selaginella moellendorffii (Sm-), Marchantia polymorpha (Mp-) and Physcomitrium patens (Pp-). The red frame highlights the amino acid at position 50 of BAG3 across different species. b Cell death phenotypes induced by the empty vector (nEYFP) and AtBAG3-N infiltrated in the leaves of WT, Nbeds1, Nbnrc2/3, Nbnrc4 and Nbnrc2/3/4 mutants. Representative photographs were taken at 2-3 dpi. Images under white and UV light are shown. The experiment was repeated independently three times with similar results. c GUS, NbNRC2 or NbNRC3 was co-expressed with AtBAG3-N in WT and Nbnrc2/3 mutant lines. GUS and empty vector (nEYFP) were used as a negative control. Representative photographs were taken at 2-3 dpi. Images under white and UV light are shown. The experiment was repeated independently three times with similar results. d BN-PAGE assay testing the oligomeric status of AtBAG3-N co-expressed with NbNRC2 in the leaves of WT and Nbnrc2/3 (upper image). Square brackets point to oligomers. NS, nonspecific bands (upper image). The protein expression level and loading control were detected by SDS-PAGE and rbcL, respectively (bottom image). The experiment was repeated independently three times with similar results. e Imaging of AtBAG3-N-induced cell death in Arabidopsis protoplasts. Protoplasts expressing AtBAG3-N were stained with PI before imaging. Bright-field image and autofluorescence were collected to show the morphology of the protoplasts. PI staining images were collected to show the dead cells. Scale bars = 25 μm. The experiment was repeated independently three times with similar results. f Quantification of cell death induced by AtBAG3-N expression in Arabidopsis protoplasts. The number of viable cells in culture based on the quantitation of ATP, which indicates the presence of metabolically active cells (n = 24). P-values were calculated by an unpaired two-tailed Student’s t test. In box and whiskers, the center line indicates the median, the bounds of the box show the 25th and 75th percentiles, the whiskers indicate the minimum to maximum values. The experiment was repeated independently three times with similar results.
Based on the observation that BAG3-N-induced cell death is a conserved response, we sought to investigate whether this response occurs independently through BAG3-N alone or requires the assistance of other proteins. Specifically, we aimed to determine whether the induction of cell death by BAG3-N in plants relies on EDS1, a family of proteins essential for the function of all tested Toll/interleukin-1 receptor NLRs (TNLs)6, or on helper NLRs, such as NRC2/3/4, which are known to play pivotal roles in the immune responses of various coiled-coil NLRs (CNLs) in solanaceous plants37. To verify whether the cell death triggered by BAG3-N requires these immune signaling components, we expressed AtBAG3-N in leaves of Nbeds1, Nbnrc2/3, Nbnrc4 and Nbnrc2/3/4 mutants and observed the cell death phenotype. AtBAG3-N was able to induce cell death in Nbeds1 and Nbnrc4 mutants, whereas it failed to induce cell death in Nbnrc2/3 and Nbnrc2/3/4 mutants (Fig. 5b). Western blot analysis revealed markedly reduced AtBAG3-N protein levels in Nbnrc2/3 and nrc2/3/4 plants (Supplementary Fig. 14a), suggesting that the lack of cell death may be due to insufficient BAG3-N expression. Co-expression of NbNRC2 or NbNRC3 with AtBAG3-N in Nbnrc2/3 mutants rescued the cell death phenotype (Fig. 5c and Supplementary Fig. 14b). Notably, the resting state of NRC2 forms homodimers or tetramers38, while its activated form adopts a hexameric structure31. BN-PAGE revealed that co-expression of AtBAG3-N and NbNRC2 led to the accumulation of dimeric and tetrameric NRC2 complexes (Fig. 5d), suggesting that AtBAG3-N functions via inactive NRC2 oligomers. Given that AtBAG3-N expression was reduced in Nbnrc2/3 and Nbnrc2/3/4 mutants (Supplementary Fig. 14a), we cannot rule out the possibility that loss of cell death is partially due to protein instability.
Notably, there is currently no evidence indicating the presence of NRC2/3 in Arabidopsis. To better understand BAG3-N-induced cell death, we stained Arabidopsis protoplasts with propidium iodide (PI) to assess plasma membrane integrity32. At 12 h post-transfection, protoplasts expressing AtBAG3-N exhibited PI uptake, suggesting a loss of plasma membrane integrity (Fig. 5e). We then evaluated AtBAG3-N-induced cell death in protoplasts using the Cell Titer-Glo Luminescent Cell Viability Assay, which quantifies cellular ATP levels33. AtBAG3-N expression led to a significant reduction in cell viability (Fig. 5f). These results indicate that BAG3-N can induce cell death in Arabidopsis protoplasts. Together, these results suggest that BAG3-N-induced cell death is conserved across angiosperms but engages divergent downstream signaling modules depending on species context.
Viral Rep protein suppresses BAG3-N-induced cell death
Plant viruses have evolved various strategies to subvert host defenses and facilitate infection3,39,40. To investigate whether TYLCV-encoded proteins suppress BAG3-N-induced cell death, we co-expressed AtBAG3-N with each of the six canonical TYLCV proteins in N. benthamiana and monitored the cell death phenotypes. Among the viral proteins tested, only Rep markedly suppressed AtBAG3-N-induced cell death (Fig. 6a). Electrolyte leakage assays showed reduced membrane damage upon co-expression of Rep with AtBAG3-N (Fig. 6b). Western blot analysis confirmed the correct expression of all proteins (Supplementary Fig. 15).
a Cell death phenotypes of AtBAG3-N co-infiltrated with the six canonical TYLCV-encoded viral proteins (Rep, C2, C3, C4, V1 and V2), respectively, in N. benthamiana leaf cells. GUS and empty vector (nEYFP) were used as a negative control. Representative photographs were taken at 2 dpi. Images under white and UV light are shown. The experiment was repeated independently three times with similar results. b Quantification of cell death induction of AtBAG3-N co-expressed with TYLCV-encoded viral proteins in N. benthamiana. Electrolyte leakage of leaf disks from N. benthamiana was measured at 2 dpi. Values are means ± SD of five biological independent samples (n = 5) from one representative experiment. Data were analyzed by one-way ANOVA followed by Tukey’s test for multiple comparisons. Different lowercase letters indicated statistically significant differences (P < 0.05). The corresponding P-values can be found in the Source Data. The experiment was repeated independently three times with similar results. c Co-IP analysis of nEYFP-AtBAG3-N and TYLCV-encoded viral proteins in vivo. Total protein was extracted from N. benthamiana leaves transiently expressing 35S: nEYFP or 35S: nEYFP-AtBAG3-N together with TYLCV-encoded viral proteins. Flag-trap beads were used to precipitate the interaction complex, and anti-GFP (α-GFP) and anti-Flag (α-Flag) antibodies were used for detection. The experiment was repeated independently three times with similar results. d Co-IP analysis of Rep-Flag with nEYFP, nEYFP-AtBAG3, and nEYFP-AtBAG3-N. Total protein was extracted from N. benthamiana leaves transiently expressing 35S: nEYFP, 35S: nEYFP-AtBAG3 and 35S: nEYFP-AtBAG3-N together with 35S: Rep-Flag. Flag-trap beads were used to precipitate the interaction complex, anti-GFP(α-GFP) and anti-Flag (α-Flag) antibodies were used for detection. The experiment was repeated independently three times with similar results. e BiFC assay verifies the interactions between Rep-cEYFP and nEYFP-AtBAG3, and between Rep-cEYFP and nEYFP-AtBAG3-N in planta. Scale bars = 25 μm. The experiment was repeated independently three times with similar results. f The interaction model between AtBAG3-N (blue) and Rep (purple) was predicted using AlphaFold 3. The stick model represents the potential interaction sites between AtBAG3-N and TYLCV Rep; this region is predicted with a high confidence score.
To elucidate the mechanism by which Rep suppresses BAG3-N-induced cell death, we examined whether Rep interacts with AtBAG3-N or the full-length AtBAG3. Co-IP assays revealed that, among all TYLCV-encoded proteins, only Rep was specifically associated with AtBAG3-N (Fig. 6c). Notably, Rep did not associate with full-length AtBAG3, suggesting that it selectively targets the N-terminal domain of AtBAG3 (Fig. 6d). In addition, BiFC assays further confirmed the Rep and AtBAG3-N interaction in planta (Fig. 6e). Structure predictions using AlphaFold 3 suggested that Rep mimics the interaction pattern of AtBAG3-C with AtBAG3-N (Fig. 6f). Collectively, these results indicate that Rep suppresses BAG3-N-induced cell death by directly associating with BAG3-N.
Notably, while the Rep protein of TYLCV is sufficient to suppress BAG3-N-induced cell death, the ability to inhibit this response varies among Sri Lankan cassava mosaic virus (SLCMV) strains. Specifically, the recently emerged SLCMVHN7 strain effectively suppresses AtBAG3-N-induced cell death, whereas the SLCMVCol strain failed to do so (Supplementary Fig. 16a, b), suggesting strain-specific differences in immune evasion capacity.
Discussion
The plant immune system relies on a sophisticated interplay between cell surface-localized PRRs and intracellular NLRs, constituting a multilayered defense against diverse pathogens1. While plant metacaspases have been implicated in stress and immune signaling27, their downstream substrates have remained largely unidentified. Previous studies have primarily addressed their roles in peptide maturation10,11 and general regulation of cell death41,42,43,44,45, with little evidence connecting them to antiviral immunity pathways. Our findings identify BAG3 as a proteolytic substrate of the type II metacaspase MC4, providing a direct molecular connection between metacaspase activity and antiviral immunity. Upon begomovirus infection, MC4 cleaves BAG3, releasing its N-terminal domain from autoinhibition. BAG3-N triggers cell death, acting downstream of MC4. This cleavage-dependent activation resembles the mechanism of pyroptosis in animals, wherein caspase-mediated cleavage of gasdermins initiates inflammatory cell death7,8. Despite the absence of caspases and gasdermins in plants9,46, the proteolytic activation of BAG3-N may reflect a convergent evolution of immune signaling strategies across kingdoms.
BAG3-N exhibits hallmark features of an immune execution module: it self-assembles into oligomeric complexes, localizes to cellular membranes, and induces cell death28,29,30,32. These properties, together with its β-sheet-rich structural propensity, suggest that BAG3-N may assemble into a supramolecular complex with structural similarities to resistosomes. However, unlike canonical resistosomes, direct evidence for membrane pore formation of BAG3-N is lacking, and functional equivalence remains speculative. Alternatively, BAG3-N may represent a novel class of oligomeric signaling module distinct from canonical NLR resistosomes. It may also act as a component of a higher-order immune signaling assembly. Structural characterization of BAG3-N and its interaction partners will be critical to elucidating its precise role in plant immune signaling.
Plant viruses have evolved sophisticated strategies to suppress host immune responses23. In line with the co-evolutionary arms race between viruses and plant hosts, we demonstrate that geminiviral Rep proteins interact with BAG3-N to suppress its cell death-inducing activity. Notably, TYLCV Rep is capable of suppressing BAG3-N-induced cell death, but the suppression efficiency differs among SLCMV strains. The recently emerged SLCMVHN7 strain exhibits enhanced suppression capacity, correlated with a C-terminal truncation in its Rep protein47. This structural modification likely reflects adaptive evolution, potentially allowing the virus to more effectively attenuate BAG3-N-mediated defense without compromising its replication or systemic movement. Such viral diversification mirrors allelic variation in bacterial effectors, such as Pseudomonas syringae HopZ variants that differentially interact with ZED1-related kinases to evade immune surveillance48.
Together, our findings reveal BAG3 as a molecular bridge linking metacaspase activity to NLR-mediated immune activation. In the resting state, BAG3 adopts an autoinhibited conformation maintained by intramolecular interactions between its N-terminal (BAG3-N) and C-terminal (BAG3-C) domains. This self-inhibitory configuration prevents the activation of downstream immune responses, thereby allowing normal plant growth. When plants are infected by TYLCV, MC4 is activated, leading to the cleavage of BAG3, which releases the N-terminal functional domain, BAG3-N. The released BAG3-N induces cell death in plant leaves, thereby restricting viral infection, and this signaling interfaces with NLR networks in certain plant species. However, to overcome this plant defense mechanism, the viral replication-associated proteins Reps interact with BAG3-N, inhibiting its cell death-inducing function (Supplementary Fig. 17). This work extends the current paradigm of plant cell death regulation and uncovers a viral target within this regulatory axis. Future studies should dissect the structural basis of BAG3-N. Given the evolutionary conservation of BAG3-N across plant lineages, synthetic deployment of this immune module may provide a promising avenue for engineering resistance to insect-borne viruses.
Methods
Plant materials and growth conditions
Arabidopsis thaliana (A. thaliana) plants used in this study include wild-type (Col-0), Atbag3 #9, 35S: YFP-AtBAG3 #6, mc4, PROPEP1-YFP and PROPEP1-YFP/mc4 lines. mc4, PROPEP1-YFP and PROPEP1-YFP/mc4 Arabidopsis lines were described previously11. We generated Atbag3 #9 and YFP-AtBAG3 #6 Arabidopsis lines. The 35S: YFP-AtBAG3-fused plasmid was used for stable plant transformation. Arabidopsis transformations were performed using the floral-dip method49. A guide RNA (gRNA) targeting AtBAG3 was designed using CRISPR-P 2.0. Atbag3 mutants were identified by sequencing PCR amplicons of the target loci. Sterilized seeds were incubated on Murashige and Skoog medium at 4 °C for 7 days before being transferred to a growth chamber (22 °C with 10 h of light / 14 h of dark).
Nicotiana benthamiana (N. benthamiana) plants used in this study include WT, Nbeds1, Nbnrc2/3, Nbnrc4 and Nbnrc2/3/4. Nbeds1, Nbnrc2/3, Nbnrc4 and Nbnrc2/3/4 were described previously50,51,52,53. N. benthamiana were grown in an insect-free growth chamber at 25 °C, with 12 h of light / 12 h of dark cycle.
Solanum lycopersicum (S. lycopersicum) cv. Micro-Tom was used to generate stable CRISPR/Cas9-edited tomato plants. Tomato plants were grown in plant growth rooms at 25 °C with 16 h of light / 8 h of dark cycle. The crispr-Slbag1/3/3 l transgenic S. lycopersicum plants were generated by transformation with the binary vector PV58KN-SlBAG1/3/3 l. Six guide RNAs targeting the open reading frame of SlBAG1/3/3 l were designed using CRISPR-P 2.0. The edited target sites were characterized by PCR sequencing to select the homozygous transgenic lines.
Whitefly and virus inoculation
Whiteflies were collected in the field in Chaoyang District, Beijing, China and identified as Bemisia tabaci MEAM1, B biotype (mtCOI, GenBank accession number MF579701). The whitefly population was maintained in a growth chamber at 25 °C and 65 % RH on cotton, under a 12 h of light / 12 h of dark cycle.
For Tomato yellow leaf curl virus (TYLCV) infection, 20–25 non-viruliferous adult whiteflies were confined in clip-cages on TYLCV-infected tomato plants for a three-day acquisition access period and then transferred to four-week-old healthy A. thaliana or tomato plants for a 3-day inoculation access period. Inoculations were performed on wild-type (WT), YFP-AtBAG3 #6 and Atbag3 #9 plants, as well as on WT and Slbag1/3/3 l plants, with six replicates for each plant type54,55. Total DNA was extracted from systemic infected leaves at 14 dpi and analyzed by qPCR with virus-specific primers.
Plasmid construction and generation of transgenic plants
For yeast two-hybrid experiments, DNA fragment encoding AtBAG3 was cloned into the pGADT7 vector to generate AD construct. Full-length MC4 was cloned into the pGBKT7 vector to generate BD constructs.
For BiFC, the DNA fragments of MC4 and AtBAG3 were cloned into the pENTR-3C entry vector and subsequently transferred into pBA3032 and pBA3036 destination vectors through attL× attR (LR) recombination reactions. This generated fusion genes with cEYFP or nEYFP at the C or N terminus, controlled by the CaMV 35S promoter.
For the Co-IP assay in vivo, full-length AtBAG3 was cloned into a pH7WGY2 empty vector to generate YFP fusion constructs. Full-length MC4C139A was cloned into the pMDC32 empty vector to generate the MC4C139A-Flag fusion construct.
Truncations and site-directed mutants of AtBAG3, SlBAG1/3/3 l, OsBAG3, ZmBAG3, MpBAG3 and PpBAG3 were cloned into pENTR-3C entry vector and then transferred into pBA3036 destination vector through LR recombination reactions to generate fusion genes with nEYFP at the N terminus under the control of the CaMV 35S promoter.
All plasmids were confirmed by DNA sequencing. Primers and plasmids used in this study are listed in Supplementary Data 1.
Antibody preparation
The DNA fragment of AtBAG3 was cloned into pET-30a vector to generate His-tagged constructs. His-tagged protein was purified using Ni-NTA agarose (Qiagen, R90101) according to the manufacturer’s instructions. The purified protein was used as an antigen to immunize rabbits for the production of polyclonal antibodies by BGI (Beijing, China).
RNA-sequencing analysis
Thirty adult whiteflies were transferred to two-week-old A. thaliana plants for three days. Mock plants were treated identically without insects. Leaves were collected for RNA extraction, with three independent biological replicates per group. RNA libraries were prepared and sequenced by BGI (Beijing, China). Reads were mapped to the Arabidopsis reference genome (TAIR10), and gene expression levels were quantified and normalized using TPM values.
Yeast two-hybrid analysis
Y2H assays were performed as previously described56. The library of Arabidopsis Mate and Plate was used to screen with the Matchmaker Gold Yeast Two-Hybrid System according to the manufacturer’s protocol (Clontech). All constructs were transformed into the yeast strain AH109 through the method of modified lithium acetate. Yeast cotransformants were screened on selective dropout (SD) medium lacking Leu, Trp, and His (SD-Leu-Trp-His) supplemented with 2 mM 3-AT. Interaction between MC4 and AtBAG3 proteins was confirmed according to the manufacturer’s protocol. The yeast strain AH109 was co-transformed with AD-AtBAG3 and BD-MC4. Yeast co-transformants were first plated on SD-Leu-Trp medium and then transferred to SD-Leu-Trp-His plates with 2 mM 3-AT for further screening.
Bimolecular fluorescence complementation (BiFC)
BiFC assays were performed as previously described57. The nEYFP-AtBAG3 and MC4-cEYFP constructs, as well as the nEYFP-AtBAG3-N and Rep-cEYFP constructs, were transformed into A. tumefaciens strain EHA105 and co-infiltrated into N. benthamiana leaves. Unfused cEYFP was used as a negative control. Fluorescence was observed at 2 days post-infiltration (dpi) using confocal microscopy.
Co-immunoprecipitation (Co-IP)
Approximately 1 g N. benthamiana leaf tissue was collected and ground into powder in liquid nitrogen. Proteins were extracted in a cold extraction buffer (50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 2 mM MgCl2, 0.5 mM EDTA, 0.1% Triton, 0.5% NP-40, 10% glycerol, 1 mM phenylmethylsulfonyl fluoride (PMSF) and one protease inhibitor cocktail/100 mL (Sigma-Aldrich, USA)). The protein extracts were then incubated with 25 μL GFP-trap or Flag-trap beads for 4 h at 4 °C. After incubation, the beads were washed three times with extraction buffer and resuspended in SDS-PAGE loading buffer before being used for immunoblot analysis. α-GFP antibody (Transgen, HT801-01; 1:5000 dilution) and α-FLAG antibody (Transgen, HT201-01; 1:10,000 dilution) were used as primary antibodies for immunoblotting, followed by Goat anti-mouse lgG-HRP (Transgen, HS002-01; 1:10,000 dilution) antibodies.
In vitro protease assay
The plasmids pET-30a-AtBAG3, pGST-MC4 and pGST-MC4C139A were transformed into E. coli (BL21) DE3 cells. The recombinant GST- and His- tagged proteins were expressed overnight at 18 °C with 1 mM IPTG induction and were separately purified using glutathione Sepharose and Ni-NTA agarose beads according to the manufacturer’s instructions. The in vitro protease assay was performed as described previously58. Briefly, 1 μg of purified recombinant BAG3 protein was incubated with 1 µg of purified recombinant MC4 or MC4C139A proteins in a reaction buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 10% (w/v) glycerol and 50 µM CaCl2) at 30 °C for 20 min. The reaction was stopped by the SDS-PAGE loading buffer supplemented with 50 mM EGTA. α-GST antibody (Transgen, HT601-01; 1:5000 dilution) and α-His antibody (Transgen, HT501-01; 1:5000 dilution) were used for immunoblotting.
qRT-PCR
Total RNA was extracted using the Plant RNA Purification Reagent (Invitrogen), and 2 μg of total RNA from each sample was reverse transcribed using TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix (Transgen, AT311-03). qPCR was performed on the CFX 96 system (Bio-Rad) using Thunderbird SYBR qPCR mix (Toyobo, QPS-201). The primers for the target genes are listed in Supplementary Data 1. Data were normalized to NbEF1α, AtACTIN or SlACTIN expression using the 2−ΔΔCt method. qRT-PCR experiments were biologically replicated three times, and the results were consistent across all replicates.
Blue native-polyacrylamide gel electrophoresis (BN-PAGE)
BN-PAGE was performed using the NativePAGE™ Novex® Bis-Tris Gel System (Thermo Fisher Scientific, USA) according to the manufacturer’s protocol. nEYFP-AtBAG3-N and the empty vector were transiently expressed in N. benthamiana leaves individually using A. tumefaciens. Plant samples were collected 36 h post-inoculation, and proteins were extracted for subsequent native PAGE analysis59. α-GFP antibody (Abclonal, AE011; 1:5000 dilution) and α-FLAG antibody (Transgen, HT201-01; 1:10,000 dilution) were used as primary antibodies for immunoblotting.
Trypan blue staining
The trypan blue staining solution was prepared by dissolving 0.015 g of trypan blue in a mixture of 10 ml of deionized water, 10 ml of glycerol, 10 ml of lactic acid, 10ml of water-saturated phenol and 40 ml of absolute ethanol. Leaves were immersed in the staining buffer, boiled for 10 min and then incubated for 5 h at room temperature. Afterward, the leaves were destained using 2.5 mg/mL chloral hydrate.
Membrane fractionation
Plasma membrane fractionation was performed using the MinuteTM Plasma Membrane Protein Isolation Kit, according to the manufacturer’s protocol (Invent, SM-005-P, USA). The plasma membrane marker protein, H+-ATPase, was detected using an anti-H+-ATPase monoclonal antibody (Agrisera, AS07 260; 1:5000 dilution). The uncropped blots of immunoblotting images are provided in the Source Data files.
Ion leakage assay
The ion leakage of treated leaf samples was measured according to a previous report, with some modifications60. Briefly, five leaf disks, each with a diameter of 5 mm, were collected and floated on 5 ml of double-distilled water at 25 °C for 3 h. The sample conductivity was then measured and recorded as value A. The leaf disks were subsequently treated at 95 °C for 25 min, and the conductivity of the cooled solution (25 °C) was recorded as value B. Ion leakage was calculated as a percentage using the formula: ion leakage = (value A / value B) × 100.
Phylogenetic analysis
To analyze the phylogenetic relationships of BAG3 and BAG3-N homologs across plant species, AtBAG3 and AtBAG3-N protein sequences from Arabidopsis thaliana were used as query sequences to perform BLASTP searches against public databases. Homologs were identified from a broad range of taxa. The BAG3 and BAG3-N domain sequences were extracted, and multiple sequence alignment was performed using MUSCLE (default parameters). The maximum likelihood phylogenetic trees were constructed using MEGA (v10.2.6), applying the JTT substitution model and a bootstrap test with 1000 replicates to assess the reliability of the inferred relationships. The resulting phylogenetic tree was visualized using iTOL to highlight taxonomic clustering and conserved sequence features.
Protoplast-based cell death analyses by confocal microscopy
Arabidopsis thaliana Protoplasts were isolated and transfected using the Arabidopsis Protoplasts Preparation and Transformation Kit, according to the manufacturer’s instructions (Coolaber, PPT101-10T, China). The nEYFP and nEYFP-AtBAG3-N constructs were transfected into A. thaliana protoplasts and incubated for 12 h post-transfection. To assess protoplast morphology and viability, cells were stained with 100 ng/mL propidium iodide (Sigma, P4170, USA) for 30 min at room temperature. The protoplasts in culture medium were then imaged by the Leica TCS SP8 STED confocal microscope. The autofluorescence in protoplasts was excited by the 488 nm laser, and the signal was collected to indicate the morphology of chloroplasts as well as the whole protoplasts.
Protoplast viability analysis
Protoplast viability was measured using the Cell Titer-Glo Luminescent Cell Viability Assay, according to the manufacturer’s instructions (Promega, G7570, USA). Luminescence signals were recorded using a microplate reader. The maximum luminescence value obtained from the control group was defined as 100% viability, and the survival rates for the experimental groups were calculated as the ratio of their respective luminescence values to the control maximum.
Statistical analyses
A significance level of 0.05 was used for statistical analysis. One-way or two-way analysis of variance (ANOVA), followed by Tukey’s multiple range test, was used for data analysis involving more than two groups. For comparisons between two groups, an unpaired two-tailed Student’s t test was performed. All statistical analyses were carried out using GraphPad Prism. The corresponding P-values can be found in the Source Data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Our RNA-seq raw data have been uploaded to the National Center for Biotechnology Information (NCBI, www.ncbi.nlm.nih.gov). RNA-seq data sets are accessible under accession number PRJNA1214420. The original data points in graphs and uncropped blots of immunoblotting images are provided in the Source Data files. Source data are provided in this paper.
References
Jones, J. D., Staskawicz, B. J. & Dangl, J. L. The plant immune system: from discovery to deployment. Cell 187, 2095–2116 (2024).
Dodds, P. N., Chen, J. & Outram, M. A. Pathogen perception and signaling in plant immunity. Plant Cell 36, 1465–1481 (2024).
Ye, J., Zhang, L., Zhang, X., Wu, X. & Fang, R. Plant defense networks against insect-borne pathogens. Trends Plant Sci. 26, 272–287 (2021).
Feng, F. & Zhou, J. Plant–bacterial pathogen interactions mediated by type III effectors. Curr. Opin. Plant Biol. 15, 469–476 (2012).
Dodds, P. N. & Rathjen, J. P. Plant immunity: towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 11, 539–548 (2010).
Wang, J., Song, W. & Chai, J. Structure, biochemical function, and signaling mechanism of plant NLRs. Mol. Plant 16, 75–95 (2023).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Minina, E. A. et al. Classification and nomenclature of metacaspases and paracaspases: no more confusion with caspases. Mol. Cell 77, 927–929 (2020).
Shen, W., Liu, J. & Li, J. Type-II metacaspases mediate the processing of plant elicitor peptides in Arabidopsis. Mol. Plant 12, 1524–1533 (2019).
Hander, T. et al. Damage on plants activates Ca²⁺-dependent metacaspases for release of immunomodulatory peptides. Science 363, eaar7486 (2019).
Wang, N. et al. A whitefly effector Bsp9 targets host immunity regulator WRKY33 to promote performance. Philos. Trans. R. Soc. B 374, 20180313 (2019).
Salguero-Linares, J. et al. Lack of At MC1 catalytic activity triggers autoimmunity dependent on NLR stability. EMBO Rep. 26, 2378–2412 (2025).
Thanthrige, N. et al. Centrality of BAGs in plant PCD, stress responses, and host defense. Trends Plant Sci. 25, 1131–1140 (2020).
Li, Y., Kabbage, M., Liu, W. & Dickman, M. B. Aspartyl protease-mediated cleavage of BAG6 is necessary for autophagy and fungal resistance in plants. Plant Cell 28, 233–247 (2016).
Fang, S. et al. Structural insight into plant programmed cell death mediated by BAG proteins in Arabidopsis thaliana. Biol. Crystallogr. 69, 934–945 (2013).
Li, Y. & Dickman, M. Processing of AtBAG6 triggers autophagy and fungal resistance. Plant Signal. Behav. 11, e1183086 (2016).
Kang, C. et al. AtBAG6, a novel calmodulin-binding protein, induces programmed cell death in yeast and plants. Cell Death Differ. 13, 84–95 (2006).
You, Q. et al. An E3 ubiquitin ligase-BAG protein module controls plant innate immunity and broad-spectrum disease resistance. Cell Host Microbe 20, 758–769 (2016).
Zhao, P. et al. Red-light is an environmental effector for mutualism between begomovirus and its vector whitefly. PLoS Pathog. 17, e1008770 (2021).
Wang, N. et al. Diverse begomoviruses evolutionarily hijack plant terpenoid-based defense to promote whitefly performance. Cells 12, 149 (2022).
Jiang, X. et al. Plant viruses convergently target NPR1 with various strategies to suppress salicylic acid-mediated antiviral immunity. J. Integr. Plant Biol. 67, 1395–1412 (2025).
Ge, L., Zhou, X. & Li, F. Plant–virus arms race beyond RNA interference. Trends Plant Sci. 29, 16–19 (2024).
Zhou, Y. et al. Double-faced role of Bcl-2-associated athanogene 7 in plant–Phytophthora interaction. J. Exp. Bot. 72, 5751–5765 (2021).
Moshe, A., Gorovits, R., Liu, Y. & Czosnek, H. Tomato plant cell death induced by inhibition of HSP90 is alleviated by Tomato yellow leaf curl virus infection. Mol. Plant Pathol. 17, 247–260 (2016).
Cao, X., Huang, M., Wang, S., Li, T. & Huang, Y. Tomato yellow leaf curl virus: Characteristics, influence, and regulation mechanism. Plant Physiol. Biochem. 213, 108812 (2024).
Basak, S. & Kundu, P. Plant metacaspases: Decoding their dynamics in development and disease. Plant Physiol. Biochem. 180, 50–63 (2022).
Liu, F. et al. Activation of the helper NRC4 immune receptor forms a hexameric resistosome. Cell 187, 4877–4889 (2024).
Wang, W. et al. WeiTsing, a pericycle-expressed ion channel, safeguards the stele to confer clubroot resistance. Cell 186, 2656–2671 (2023).
Jacob, P. et al. Plant “helper” immune receptors are Ca²⁺-permeable nonselective cation channels. Science 373, 420–425 (2021).
Madhuprakash, J. et al. A disease resistance protein triggers oligomerization of its NLR helper into a hexameric resistosome to mediate innate immunity. Sci. Adv. 10, eadr2594 (2024).
Bi, G. et al. The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling. Cell 184, 3528–3541 (2021).
Hu, M., Qi, J., Bi, G. & Zhou, J. Bacterial effectors induce oligomerization of immune receptor ZAR1 in vivo. Mol. Plant 13, 793–801 (2020).
Wang, J. et al. Reconstitution and structure of a plant NLR resistosome conferring immunity. Science 364, eaav5870 (2019).
Wu, X., Zhang, X., Wang, H., Fang, R. -x & Ye, J. Structure–function analyses of coiled-coil immune receptors define a hydrophobic module for improving plant virus resistance. J. Exp. Bot. 74, 1372–1388 (2023).
Nelson, B. K., Cai, X. & Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Wu, C. et al. NLR network mediates immunity to diverse plant pathogens. Proc. Natl. Acad. Sci. USA 114, 8113–8118 (2017).
Ma, S. et al. Oligomerization-mediated autoinhibition and cofactor binding of a plant NLR. Nature 632, 869–876 (2024).
Wu, X. et al. The Orthotospovirus nonstructural protein NSs suppresses plant MYC-regulated jasmonate signaling leading to enhanced vector attraction and performance. PLoS Pathog. 15, e1007897 (2019).
Liu, B. et al. Plant defence negates pathogen manipulation of vector behaviour. Funct. Ecol. 31, 1574–1581 (2017).
Coll, N. S. et al. The plant metacaspase AtMC1 in pathogen-triggered programmed cell death and aging: functional linkage with autophagy. Cell Death Differ. 21, 1399–1408 (2014).
Tsiatsiani, L. et al. Metacaspases. Cell Death Differ. 18, 1279–1288 (2011).
Minina, E. A. et al. Autophagy and metacaspase determine the mode of cell death in plants. J. Cell Biol. 203, 917–927 (2013).
Coll, N. S. et al. Arabidopsis type I metacaspases control cell death. Science 330, 1393–1397 (2010).
Watanabe, N. & Lam, E. Arabidopsis metacaspase 2d is a positive mediator of cell death induced during biotic and abiotic stresses. Plant J. 66, 969–982 (2011).
Devant, P. & Kagan, J. C. Molecular mechanisms of gasdermin D pore-forming activity. Nat. Immunol. 24, 1064–1075 (2023).
Wang, D. et al. A 7-amino-acid motif of Rep protein essential for virulence is critical for triggering host defense against Sri Lankan cassava mosaic virus. Mol. Plant Microbe Interact. 33, 78–86 (2020).
Lewis, J. D. et al. The Arabidopsis ZED1 pseudokinase is required for ZAR1-mediated immunity induced by the Pseudomonas syringae type III effector HopZ1a. Proc. Natl. Acad. Sci. USA 110, 18722–18727 (2013).
Zhang, X., Henriques, R., Lin, S., Niu, Q. & Chua, N. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 1, 641–646 (2006).
Wu, C. et al. NRC4 gene cluster is not essential for bacterial flagellin-triggered immunity. Plant Physiol. 182, 455–459 (2020).
Adachi, H. et al. An N-terminal motif in NLR immune receptors is functionally conserved across distantly related plant species. Elife 8, e49956 (2019).
Witek, K. et al. A complex resistance locus in Solanum americanum recognizes a conserved Phytophthora effector. Nat. Plants 7, 198–208 (2021).
Schultink, A., Qi, T., Lee, A., Steinbrenner, A. D. & Staskawicz, B. Roq1 mediates recognition of the Xanthomonas and Pseudomonas effector proteins XopQ and HopQ1. Plant J. 92, 787–795 (2017).
Cui, X., Tao, X., Xie, Y., Fauquet, C. M. & Zhou, X. A DNAβ associated with Tomato yellow leaf curl China virus is required for symptom induction. J. Virol. 78, 13966–13974 (2004).
Yang, J. et al. βC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev. 22, 2564–2577 (2008).
Zhang, X. et al. Near-infrared light and PIF4 promote plant antiviral defense by enhancing RNA interference. Plant Commun. 5, 100665 (2024).
Zhao, P. et al. Viruses mobilize plant immunity to deter nonvector insect herbivores. Sci. Adv. 5, eaav9801 (2019).
Minina, E. A., Stael, S., Van Breusegem, F. & Bozhkov, P. V. Plant metacaspase activation and activity. Methods Mol. Biol. 1133, 237–253 (2014).
Na Ayutthaya, P. P., Lundberg, D., Weigel, D. & Li, L. Blue native polyacrylamide gel electrophoresis (BN-PAGE) for the analysis of protein oligomers in plants. Curr. Protoc. Plant Biol. 5, e20107 (2020).
Zhu, M. et al. The intracellular immune receptor Sw-5b confers broad-spectrum resistance to tospoviruses through recognition of a conserved 21-amino acid viral effector epitope. Plant Cell 29, 2214–2232 (2017).
Acknowledgements
We thank Dr. Brian Staskawicz (University of California, USA) for kindly providing the Nbeds1 N. benthamiana mutant seeds. We thank Dr. Sophien Kamoun (University of East Anglia, UK) for kindly providing the Nbnrc2/3, Nbnrc4 and Nbnrc2/3/4 N. benthamiana mutants seeds. We thank Dr. Simon Stael (The Swedish University, Sweden) for kindly providing the mc4, PROPEP1-YFP/mc4 Arabidopsis seeds. We thank Dr. Jianfeng Li (Sun Yat-sen University, China) for kindly providing the mc4/5/6/7, PROPEP1 promoter: PROPEP1-GFP-HA, PROPEP1-GFP-HA/mc4/5/6/7 Arabidopsis seeds. We thank Dr. Dawei Li (China Agricultural University, China) for kindly providing the Nbmc4 N. benthamiana mutants seeds. We thank Dr. Hailong Guo (China Agricultural University, China) for kindly providing the NbNRC2, NbNRC3 and NbNRC4 plasmids. The study was supported by the National Natural Science Foundation of China (32125032), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0810000, XDA0450000), the China National Key Research and Development Program (2023YFC2606800, 2024YFC3406000), the CAS Projects for Young Scientists in Basic Research (YSBR-080), the Inner Mongolia Science and Technology Program (2023KJHZ0018) and the project of State Key Laboratory of Discovery and Utilization of Functional Components in Traditional Chinese Medicine.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.Y.; Investigation, L.L., Y.J., H.W., X.C., and W.Z. Writing-Original Draft, J.Y. and L.L.; Writing-Review & Editing, J.Y., L.L., Y.J., X.L., P.Z., and Y.S.; Funding Acquisition, J.Y.; Supervision, J.Y., R.F. and X.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, L., Jiang, Y., Zhao, P. et al. Cleavage of Bcl-2-associated athanogene by metacaspase determines plant antiviral immunity. Nat Commun 16, 9102 (2025). https://doi.org/10.1038/s41467-025-64021-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64021-w