Abstract
47% of hereditable diseases are caused by single C•G-to-T•A base conversions, which means efficient A-to-G base editing tools (ABEs) have great potential for the treatment of these diseases. However, the existing efficient ABEs, while catalyzing targeted A-to-G conversion, cause high A or C bystander editing and off-target events, which poses safety concerns for their clinical applications. To overcome this shortcoming, we have developed ABE8e-YA (ABE8e with TadA-8e A48E) for efficient and accurate editing of As in YA motifs with YAY > YAR (Y = T or C, R = A or G) hierarchy through structure-oriented rational design. Compared with ABE3.1, which is currently the only ABE version with a YAC motif preference, ABE8e-YA exhibits an average A-to-G editing efficiency improvement of an up to 3.1-fold increase in the indicated YA motif while maintaining reduced bystander C editing and minimized DNA or RNA off-targets. Additionally, we demonstrate that ABE8e-YA efficiently and precisely corrects pathogenic mutations in human cells, suggesting its high suitability for addressing 9.3% of pathogenic point mutations, higher than that of ABE8e and ABE9. Moreover, by using ABE8e-YA, we efficiently and precisely generate hypocholesterolemia and tail-loss mouse models mimicking human-associated disease, as well as performed in vivo mouse proprotein convertase subtilisin/kexin type 9 (Pcsk9) base editing for hypercholesterolemia gene therapy. Together these data indicate its great potential in broad applications for disease modeling and gene therapy.
Similar content being viewed by others
Introduction
About 60% of hereditable diseases are caused by single base mutations. 47% of these are caused by single C•G-to-T•A base conversions, which urgently requires effective T-to-C or A-to-G (the complementary DNA strand) correction tools to break through the limitations of the low efficiency of traditional CRISPR/Cas9-mediated homologous recombination1.
The emergence of the new generation of gene-editing tools, specifically adenine base editors, which possess inherent advantages such as high efficiency and safety, has opened up the possibility of achieving a complete cure for genetic diseases caused by C•G-to-T•A mutations2. ABEs have been successfully applied in various in vivo gene therapy studies in mice, including in the treatment of type I tyrosinemia3, progeria4, Duchenne muscular dystrophy5, and hereditary deafness6. A clinical trial initiated by Verve Therapeutics is currently underway to treat hypercholesterolemia using ABEs7.
However, currently, the most efficient version of an ABE features an editing window of 3-9 nucleotides (nt), which inevitably leads to bystander editing while targeting the intended base, posing safety risks for clinical disease treatment. Among the 47% pathogenic point mutations that can be potentially corrected by ABEs, the proportion that can be targeted by ABE8e is 18.0% (corresponding proportion is 39.8% when PAM is NG). However, 12.3% of those (corresponding proportion is 24.4% when the PAM is expanded to NG) targeted by ABE8e will cause bystander mutations (Supplementary Fig. 1). The development of ABEs with motif sequence preferences can effectively minimize the occurrence of such potential safety risks. In recent years, scientists around the world have made significant progress on the accuracy and efficiency of base editing. For C-to-T base editing, a series of improved cytosine base editors (CBEs) have been developed, such as through structure-oriented design of cytosine deaminases to construct motif-specific CBEs (TC, CC and WRC, W = A or T, R = A or G)8,9,10,11, engineering cytosine base editors with only 1-2nt editing windows12, re-evolving TadA variants transformed ABEs into more accurate and less off-target CBEs13,14,15, and more. For ABEs, some progress has also been made, such as rational design of accurate ABE9 for only editing A5 or A6 positions16. However, limitations of ABEs are still present. For example, ABE3.1, the only ABE with a YAC motif preference, has low efficiency and induces more C bystanders, as well as generated greater off-targets. Therefore, ABEs that could efficiently catalyze A-to-G conversion in specific motifs without introducing more bystanders and off-targets are still in urgent need.
Here, we use structure-oriented rational design to generate ABE8e-YA, a mutant with enhanced preference for YA motif sequences. Compared to ABE8e, ABE8e-YA can precisely edit YA sequences while minimizing bystanders. More importantly, compared with ABE3.1, ABE8e-YA exhibits an average 3.1-fold increased A-to-G editing efficiency within the indicated motif while maintaining reduced bystander C editing and minimized DNA or RNA off-targets. We also demonstrate that ABE8e-YA can effectively correct disease-causing point mutations while minimizing detrimental mutations in human cells. Moreover, by injecting ABE8e-YA mRNA and single-guide RNAs (sgRNAs) targeting the splice donor of mouse Pcsk9 and Tbxt into zygotes, a PCSK9 loss-of-function mouse model of hypocholesterolemia and a short-tailed mouse model for simulating mouse evolution are efficiently generated, respectively. Finally, we deliver lipid nanoparticle-packaged ABE8e-YA mRNA and a sgRNA targeting the GT splice donor site of Pcsk9 into mice. High base editing efficiency for Pcsk9 is observed, and a substantial reduction in protein expression of PCSK9 and low-density lipoprotein cholesterol (LDL-C) in plasma is noted.
Results
Structure-based evolution of TadA-8e for motif sequences preference
The earlier reported ABE with a YAC motif preference was ABE3.12, but its A-to-G editing efficiency was low (1.2–31.1%) and it induced relatively high C bystander editing (Supplementary Fig. 2). Efficiency-wise, ABE8e is currently considered the more effective version of ABE. However, ABE8e not only has higher editing activity and a wider editing window causing it to tend to induce more bystander edits and off-targets, but this also causes it to lose its sequence preference17. To address this limitation, we attempted to develop ABEs that restore the inherent motif sequence specificity through rational design according to the crystal structure of ABE8e (PDB:6VPC)18. Based on our hypothesis that modifying residues around the DNA substrate-interaction region of TadA-8e can modulate DNA binding or DNA selectivity. We selected the following amino acids in TadA-8e that interact with the DNA Non-target Strand (NTS): R26-E27-V28-P29, N46-A48-I49, M61, V82, R107, G115, Y123, A142-L145-D147-Y149-Q154-A158. These amino acids were modified to alter their polarity (e.g., N46A/L, V82T, Y123A, L145T, Y149A, A158S), electrical properties (e.g., E27R, R107C, D147R, Q154H/R), both polarity and electrical properties simultaneously (e.g., R26A/G, E27A, A48D, I49K, R107A, A142E), or side chain size (e.g., V28A/G, P29A, I49A, M61I, G115M).
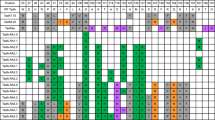
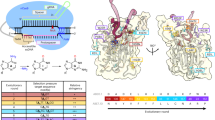
Based on the above principles, we designed a series of mutants of ABE8e and screened for ABEs that restore its YAC motif sequence preference (Fig. 1a). A human endogenous target site (HEK2) with A3CA5CA7A8 sequence was selected for testing. Among these mutations, ABEs that could edit A5 while reducing or having no editing efficiency for A3, A7 and A8 were selected as candidates (Fig. 1b). After co-transfecting ABE8e mutants and HEK2 sgRNA in HEK293T cells for 72 h, we collected genomic DNA from the cells for amplicon high-throughput sequencing (HTS). After analyzing the A-to-G or C-to-D (D = A,G or T) base editing efficiency in the FASTQ file from HTS using BE-Analyzer19, we found that almost all of the ABE8e mutants can edit both A5 and A7, while only ABE8e A48D exhibited reduced A7 and C6 base editing activity without sacrificing on-target A5 activity (Fig. 1c). This indicated that A48 may be the key amino acid of TadA-8e that influences its motif sequence preference. In order to further identify the TadA-8e mutant capable of minimizing bystander editing, we saturated the mutant A48 amino acid and further tested it with the same target in HEK293T cells. HTS data showed that ABE8e A48E maintains comparable editing activity at A5 while minimizing bystander editing activity at A7 (a 3.0-fold decrease) compared to ABE8e A48D (Fig. 1d). Structural modeling using PyMOL revealed that the A48E substitution introduces a glutamate side chain (-CH2-CH2-COO⁻) generating electrostatic repulsion with the DNA phosphate backbone. This interaction displaces the substrate toward the opposite face of the deamination pocket, compressing the van der Waals gap within the active site (Fig. 1e). The resulting steric constraints preferentially accommodate smaller pyrimidines (C/T), thereby enhancing YA motif sequences preference. We named the ABE8e A48E variant “ABE8e-YA” and continued to characterize its activity.
a Schematic illustration of the conceptual design to create an efficient YA motif-preferred ABE. b Overview of the interaction of TadA-8e (yellow) with the single-stranded DNA substrate (orange) (Protein Data Bank (PDB): 6VPC). Cas9n is shown in green, sgRNA in cyan, and double-stranded DNA in orange. Amino acids spatially contacting or adjacent to the substrate DNA are labeled on the enlarged image. c The A-to-G base editing efficiency of ABE8e or ABE8e variants at an endogenous genomic loci containing YAC motif and multiple As with non-YAC motif (HEK site2) in HEK293T cells. Data are mean (n = 3 independent experiments). d The A-to-G base editing efficiency of ABE8e or ABE8e variants of A48 saturation mutagenesis at HEK site2 in HEK293T cells. Data are mean (n = 3 independent experiments). e The A48E mutation in TadA8e alters ABE’ s sequence preference by modifying molecular interactions. Red dashed lines indicate steric clashes between the protein and ligand, yellow dashed represent favorable van der Waals interactions. Source data are provided as a Source Data file.
Characterization of ABE8e-YA
To unbiasedly profile the characteristics of ABE8e-YA, 23 endogenous targets were tested in HEK293T cells using ABE8e, ABE9 and ABE3.1 as controls. Among these targets, they almost cover NAN (N = A, T, C and G) and some targets contained NCN (N = A, T, C and G) within its editing window. After analyzing the A-to-G editing efficiency of the most highly-edited As in each target across all 23 target sites, we found that the efficiencies of ABE8e were 43.8-90.9%, the maximum of which was higher than all other ABE treated groups (Fig. 2a). The efficiencies of ABE8e-YA were 1.4–90%, which was more efficient than that of ABE9 (0.7-78.7%) and of ABE3.1 (0–75.3%) (Fig. 2a). After standardization based on ABE3.1, the fold-changes of ABE8e-YA were significantly lower than that of ABE8e, but higher that of ABE9 and ABE3.1 (Fig. 2a, b). Through analyzing the A-to-G editing efficiency of each protospacer of all tested targets, we found ABE8e can efficiently edit almost all As without motif preference at major editing window (A3-A8) and even edit As outside of the major window (A2 or A10-A13) with considerable efficiency (Fig. 2a, c). ABE9, ABE3.1 and ABE8e-YA have a similar editing window that was narrower than that of ABE8e, suggesting they are all accurate versions of an ABE suitable for precision gene therapy (Fig. 2a, c). We further analyzed the motif preference of these base editors and no motif preference for ABE8e and ABE9 was observed (Fig. 2d). It is noteworthy that ABE3.1 is capable of effective editing of both YA (including YAC) and displays a slight preference for AA sequences. In contrast, ABE8e-YA exhibits a preference for YA motifs, with a YAY > YAR hierarchy (where Y = T or C, and R = A or G) (Fig. 2a, d, e). Moreover, the average A-to-G efficiencies of ABE8e-YA at A4, A5, A6 and A7 were 39.2%, 57.1%, 36.7% and 15.1%, respectively, far higher than the corresponding efficiencies of ABE3.1(Fig. 2a, c). Since ABE8e has been reported to have cytosine deaminase activity20, we further analyzed the C editing of ABE8e-YA at 12 targets. The results showed that ABE8e-YA induced lower or near background C-to-D (D-A, G or T) editing activity compared to ABE8e. At FANCF-sg5, ABE8e-YA decreased the editing efficiency of C induced by ABE8e by up to 10.4 times, which was comparable to ABE9 (Fig. 2f). ABE8e-YA also induced no more indels than that of ABE8e and ABE3.1 (Fig. 2g). Together, ABE8e-YA is highly efficient at YA motif preference with significantly reduced adenine and cytosine bystander editing effects.
a The A-to-G editing efficiency of ABE8e, ABE9, ABE3.1 and ABE8e-YA were tested at 23 endogenous target sites in HEK293T cells. Data are mean (n = 3 independent experiments). b The editing efficiency at highly-edited adenines induced by ABE8e, ABE9, ABE3.1 and ABE8e-YA was tested at 23 endogenous target sites in HEK293T cells. Each data point represents the highest editing efficiency from A to G in the editing window of each target site, which is standardized by ABE3.1 and calculated from three independent experiments. Data are median with range (n = 23 target sites). P-values were determined by two-sided paired Wilcoxon rank-sum test. ABE8e vs ABE9: P < 0.0001, ABE8e vs ABE3.1: P < 0.0001, ABE8e vs ABE8e-YA: P < 0.001, ABE9 vs ABE8e-YA: P = 0.0010, ABE3.1 vs ABE8e-YA: P < 0.0001. c Average A-to-G editing efficiency of ABE8e, ABE9, ABE3.1 and ABE8e-YA was tested at 23 endogenous target sites in HEK293T cells. Data are means ± SD. d The motif preference analysis of ABE8e, ABE9, ABE3.1 and ABE8e-YA at 23 endogenous target sites in HEK293T cells. Each data point represents average A-to-G editing efficiency at all motifs within the editing window of each target site, calculated from 3 independent experiments. The thick centre line within the violin plot shows the median, while the thinner horizontal lines show the first and third quartiles. e A sequence motif logo reflecting the context preferences of ABE8e, ABE9, ABE3.1 and ABE8e-YA at 23 endogenous target sites in HEK293T cells. f Average C-to-D(A/C/G) editing efficiency of ABE8e, ABE9, ABE3.1 and ABE8e-YA was tested at 12 endogenous target sites in HEK293T cells. Data are means ± SD (n = 3 independent experiments). g The indels induced by ABE8e, ABE9, ABE3.1 and ABE8e-YA at 23 endogenous target sites in HEK293T cells. Each data point represents average indels frequency at each target site calculated from 3 independent experiments. Data are median with range (n = 23 target sites), and P-values were determined by two-sided paired Wilcoxon rank-sum test. ABE8e vs ABE8e-YA: P = 0.4144; ABE9 vs ABE8e-YA: P = 0.0135; ABE3.1 vs ABE8e-YA: P = 0.6010. Source data are provided as a Source Data file.
Off-target evaluation of ABE8e-YA
Next, we performed off-target assessment of ABE8e-YA in three ways: sgRNA-dependent DNA off-target, sgRNA-independent DNA off-target and whole-transcriptomic RNA off-target. 17 off-targets from in silico prediction or Cas9 known off-targets were used to evaluate the sgRNA-dependent DNA off-target. The results showed that 4/17 off-targets were observed for ABE8e and no sgRNA-dependent DNA off-targets were observed for both ABE8e-YA and ABE9 (Fig. 3a). A modified R-loop assay was used to evaluate the sgRNA-independent DNA off-target effects21. HTS data showed that ABE8e-YA, similar to ABE9, induced no sgRNA-independent DNA off-targets, which were far fewer than that of ABE8e (Fig. 3b and Supplementary Fig. 3). RNA-Seq was used to evaluate whole-transcriptomic RNA off-target effects22,23.The results showed that ABE8e-YA induced 5 times lower the off-target rate than that of ABE8e, though slightly higher than that of ABE9 (Fig. 3c). In summary, ABE8e-YA is a highly efficient base editing tool with high specificity.
a Cas9-dependent DNA on- and off-target analysis of the indicated targets (PD-1-sg4 and HEK3) by ABE8e, ABE9, ABE3.1 and ABE8e-YA in HEK293T cells. Lowercase protospacer sequences represent mismatched bases compared to their corresponding on-target sequences. Data are means ± SD (n = 3 independent experiments), and P-values were determined by two-tailed Student’ s t-test. ABE8e vs ABE8e-YA: PD-1-sg4, OT5: P = 0.0012, OT7: P = 0.0047; HEK site3 OT3: P < 0.0001, OT4: P = 0.0029. b Cas9-independent DNA off-target analysis of ABE8e, ABE9, ABE3.1 and ABE8e-YA using the modified orthogonal R-loop assay at each R-loop site with nSaCas9-sgRNA. Data are means ± SD (n = 3 independent experiments). P-values were determined by two-tailed Student’ s t-test. ABE8e vs ABE8e-YA at R-loop1: P = 0.0002; R-loop2: P < 0.0001; R-loop3: P = 0.0075; R-loop4: P = 0.0010; R-loop5: P = 0.0473; R-loop6: P = 0.0030. c RNA off-target editing activity by ABE8e, ABE9, ABE3.1 and ABE8e-YA using RNA-seq. Each biological replicate is listed on the bottom. Significance was tested by two-tailed Student’s t test (a, b). d ABE8e, ABE9 and ABE8e-YA were used to edit HEK293T stable cell lines carrying pathogenic GALT c.413 C > T (galactosemia) and OTC c.533 C > T (ornithine transcarbamylase deficiency) mutations. Representative sequencing reads show A-to-G and C-to-D conversions (percentages indicate corresponding reads/total reads). e The A-to-G or C-to-D (%) base editing of by ABE8e, ABE9 and ABE8e-YA at the HEK293T stable cells. Data are mean (n = 3 independent experiments). Source data are provided as a Source Data file.
Introduction of therapeutic mutation with high precision using ABE8e-YA in human cells
To assess ABE8e-YA’s suitability for precision gene therapy, we generated stable HEK293T cell lines carrying two pathogenic mutations: GALT c.413 C > T24 (Transferase Deficiency Galactosemia) and OTC c.533 C > T25 (Ornithine Transcarbamylase Deficiency) with bystander editing identified as risk mutation in the ClinVar database. The results showed that ABE8e-YA has higher desired base editing efficiency at A6 in GALT (45.6%) and A6 in OTC (71.35%) than that of ABE9, though its efficiency slightly lower than that of ABE8e at these sites (Fig. 3d, e). However, ABE8e also induced undesired base mutations with high efficiency, such as A3 in GALT and A3 in OTC which potentially bring the risk of additional disease26. In contrast, ABE8e-YA exhibited dramatically reduced or undetectable activity at these undesired loci (Fig. 3d, e).
We further counted these pathogenic mutations targeted by the efficient version of ABE (taking ABE8e as an example here) but caused bystander editing categorized as possibly presenting a risk of disease in the ClinVar database. Among all pathogenic point mutations suitable for A-to-G correction, 5.7% (5413), 2% (1841), and 9.3% (8783) were suitable for correction without introducing risk mutations when using ABE8e, ABE9, and ABE8e-YA, respectively (Supplementary Fig. 5 and Supplementary Data 2). When PAM was further expanded from NGG to NG, the proportion of precise disease-treatment events increased to 15.4% (14,472) for ABE8e, 6.4% (6033) for ABE9 and 22.9% (21,622) for ABE8e-YA, respectively (Supplementary Fig. 5 and Supplementary Data 2). These data suggest that ABE8e-YA is superior to existing ABE tools in achieving precise A-to-G base correction for future clinical gene therapy applications.
Generation of disease associated mouse models with high precision using ABE8e-YA
Base editors have been applied for generating animal disease models that are essential for basic medical research27. When base editors are used to generate animal models of disease, they induce not only the target base conversions but also bystander base editing which potentially perturbs analysis of the relationship between the mouse disease phenotype and base mutations. Therefore, we used ABE8e-YA to generate mouse models to minimize the occurrence of bystander events. We first targeted the Pcsk9 to generate hypocholesterolemia mouse models by the zygote microinjection of ABE8e-YA mRNA and sgRNA targeting the splice site of Pcsk9 with ABE8e and ABE9 as controls (Fig. 4a and Supplementary Fig. 6)28. A-to-G editing efficiency was quantified using CRISPResso2 to analyze the HTS of tail biopsy-derived genomic DNA29. The results showed that ABE8e-YA, ABE9 and ABE8e induced efficient base editing in pups, with 17/18 (ABE8e-YA), 16/16 (ABE9) and 11/11 (ABE8e), successfully edited, respectively (Fig. 4b, c and Supplementary Fig. 6). However, only ABE8e-YA induced precise and efficient A-to-G conversion of the opposite DNA strand of GT splice site with minimized bystander editing in pups. 15/17 base-edited mice carried homozygous target base edits in ABE8e-YA treated group. In ABE8e treated group, no mice harboring homozygous target base edits in 11 F0 base-edited mice and, in the remaining mice, not only the target base A6, but also bystander A4 were edited. However, no homozygous mice were generated in the ABE9 treated group (Fig. 4c and Supplementary Fig. 6). The phenotype of the pups carried precise target base editing by ABE8e-YA were further confirmed by the detection of PCSK9 and LDL-C (Fig. 4d, e). These data suggest that ABEs, especially our highly efficient version of ABE, enabled efficient base editing in embryos to produce homozygous mice with the disease phenotype.
a The workflow for generating mouse models by ABEs, the sampling process and sequencing steps. b The target sequence of ABE8e-YA or ABE8e in the proprotein convertase subtilisin/kexin type 9 (Pcsk9) exon 1 locus. The PAM sequence and sgRNA target sequence are shown in orange and black underline, respectively. The desired A-to-G conversion to disrupt the opposite DNA strand of the GT splice site is shown in red, and the GT splice site in blue. c Genotyping of representative F0 pups of Pcsk9 base editing by ABE8e, ABE9 and ABE8e-YA. The frequencies of wild-type (WT) and mutant alleles were determined by analyzing HTS using CRISPResso2. The percentage values on the right represent the frequencies of the indicated mutant alleles. The red position indicates the base where the mutation occurs. d, e The expression of PCSK9 and LDL-C in plasma of F0 pups from 8-week-old wild-type and founder (#CP03, #CP14 and #CP15) mice. Data are means ± SD (n = 3 independent experiments). f The target sequence of ABE8e-YA or ABE8e Tbxt exon 6 locus. The PAM sequence and sgRNA target sequence are shown in orange and black underline, respectively. The desired A-to-G conversion is shown in red, and the GT splice site in blue. g Genotyping of representative F0 pups of Tbxt base editing by ABE8e-YA. The frequencies of wild-type (WT) and mutant alleles were determined by analyzing HTS using CRISPResso2. The percentage values on the right represent the frequencies of the indicated mutant alleles. h The tail-loss phenotype of Tbxt base editing F0 pups from 8-week-old wild-type and founder (#CT02, #CT03 and #CT04) mice. Source data are provided as a Source Data file.
However, for some other disease-causing genes that are lethal with homozygous mutations, the most efficient version of ABE is not always suitable. For example, in Tbxt, a gene that influences the evolution of the mouse tail, a single exon 6-skipped transcript is sufficient to induce a tail-loss phenotype (Fig. 4f)30. We designed a sgRNA targeting the splice site of the Tbxt such that its disruption may possibly cause exon 6 skipping. By co-microinjection of ABE8e-YA with sgRNA into mouse zygotes, 8 (8/8) pups were edited in the ABE8e-YA treated group (Fig. 4g and Supplementary Fig. 7). Heterozygous mice (#CT03, #CT04) displayed almost no tail phenotype and mice with reduced editing efficiency (#CT02) showed a short-tailed phenotype (Fig. 4h). Co-microinjection of ABE8e and the sgRNA was used as a control, however, we found no pups were born in this group. The probable cause is that homozygous editing by ABE8e in mice leads to complete exon 6 skipping and thus death in utero. These data suggest that ABE8e-YA has significant potential for generating disease models. More importantly, ABE8e-YA shows superior advantages in animal models of diseases targeting homozygous lethal genes compared with other high efficiency version of ABEs.
In vivo adenine base editing Pcsk9 in mouse using ABE8e-YA
To further validate ABE8e-YA’s potential for gene therapy in vivo, we used lipid nanoparticle (LNP)–packaged delivery of mRNA encoding an ABE8e-YA and a sgRNA targeting the GT splice donor site of Pcsk9 in mice. ABE8e with the same guide served as the control (Fig. 5a). Mice were injected with LNP (designated as Day 0), and blood samples of the mice were collected from the tail vein on Days −2, 7, 14, and 21 for serum analysis. At day 21 post-injection, mice were euthanized and their livers and other organs were harvested for HTS, followed by analysis using CRISPResso2. Higher base editing efficiencies for ABE8e (43.9%) and ABE8e-YA (45%) were observed exclusively in the liver in the group receiving high-dose LNP delivery (2 mg/kg) (Fig. 5b). Notably, ABE8e-YA precisely induced the desired base mutation (A6) while minimizing bystander editing (A4) (Fig. 5b, Supplementary Fig. 8a). Low-dose delivery of ABEs (0.5 mg/kg and 0.125 mg/kg) resulted in a loss of in vivo base editing efficiency (Fig. 5b). No DNA off-target effects were detected (Supplementary Fig. 8b). During this time, we also monitored the expression levels of PCSK9 and LDL-C in plasma. The results also showed substantial reduction in both of these proteins in both the ABE8e and the ABE8e-YA treated groups (Fig. 5c, d). Meanwhile, LNP-packaged ABE8e-YA delivery did not induce liver injury as observed through the detection of serum alanine aminotransferase (ALT) and aspartate transaminase (AST) (Supplementary Fig. 8c, d). These data suggested that ABE8e-YA has great potential in clinical gene therapy in vivo.
a The workflow for the mouse Pcsk9 base editing in vivo by the delivery of LNP-packaged ABE8e-YA. The target sequence of ABE8e-YA in the Pcsk9 exon 1 locus. The PAM sequence and sgRNA target sequence are shown in green and orange underline, respectively. The desired A-to-G conversion, designed to disrupt the GT splice site on the opposite DNA strand, is shown in red, and the GT splice site in blue. b The A-to-G editing efficiency of Pcsk9 in adult mice 3-weeks after the delivery of LNP-packaged ABE8e-YA and sgRNA. Data are means ± SD (n = 3 individual mice) c, d Expression of PCSK9 and LDL-C in plasma of adult mice before and after the delivery of LNP-packaged ABE8e-YA and sgRNA. Data are means ± SD (n = 3 for the control group, n = 3 for the ABE8e-YA group). Source data are provided as a Source Data file.
Discussion
In this study, we have developed a YA motif-preferred adenine base editor by structure-oriented rational design of TadA-8e and named it ABE8e-YA. Compared to the most efficient ABE version currently available, ABE8e, ABE8e-YA has YA motif preference and a narrower editing window, while reducing bystander A and C editing and DNA/RNA off-targets. ABE3.1, currently the only sequence-specific ABE version, has unpredictable C editing and RNA off-targets. ABE8e-YA not only effectively alleviates the limitations of previous ABEs, but also has the advantages of high efficiency and small size. Compared with the current most accurate ABE, ABE9, ABE8e-YA not only has the same editing efficiency at A5 and A6, but also a higher editing efficiency at A4 and A7 sites, where ABE9 cannot edit. Therefore, ABE8e-YA is an elegant adenine base editor with high efficiency, precision, and a small size. However, although ABE8e-YA has high accuracy, it also sacrifices on-target editing efficiency. Therefore, it is necessary to further engineer deaminase in the future to improve its efficiency while maintaining its accuracy. For cytosine base editors, TC, CC, and WRC motif preference editors have been reported, while similar ABE tools are still lacking. Thus, further exploration is needed to discover new ABEs with different motif performance, such as RA (R = A or G), to meet the need for clinical disease treatment.
Furthermore, we have also demonstrated that ABE8e-YA can be effectively used to correct disease-causing point mutations in human cells, especially in editing windows where there is an A mutation with a safety risk. According to statistics in the ClinVar database, 9.3% (8,783) of disease treatment events could be treated using ABE8e-YA rather than other ABEs by precisely targeting YA motif. When the PAM was further expanded from NGG to NG, the ratio of precise disease treatment events was further expanded to 22.9% (21,622). The correction outcomes of the two pathogenic mutations (GALT c.413 C > T, Transferase Deficiency Galactosemia; OTC c.533 C > T, Ornithine transcarbamylase deficiency) further validated the precision of ABE8e-YA in addressing pathogenic point mutations. Finally, we demonstrated its effectiveness and precision for disease modeling and gene therapy in vivo, suggesting that ABE8e-YA has great potential in precisely targeting specific bases for future clinical gene therapy.
In summary, ABE8e-YA is a highly efficient and specific base editing tool, which has great potential in animal model generation and disease gene therapy.
Methods
Plasmid construction
The DNA sequences used in this research can be found in Supplementary Data 3. The ABE8e (#138489) and lentiCRISPR v2 (#52961) plasmids were obtained from Addgene. Polymerase chain reaction (PCR) was performed using KOD-Plus-Neo DNA Polymerase (TOYOBO, Code: KOD-401). Base editor (BE) plasmids developed in this study were assembled using ClonExpress MultiS One Step, as described in the Supplementary Data 3. sgRNA expression plasmids were constructed according to established protocols27. Briefly, oligonucleotides listed in Supplementary Data 1 were denatured at 95 °C for 5 min followed by slow cooling to room temperature. Annealed oligonucleotides were ligated into BbsI-linearized U6-sgRNA(sp)-EF1α-GFP for sgRNA expression (Thermo Fisher Scientific).
Human cell culture
HEK293T cell lines (ATCC CRL-3216) were cultured in Dulbecco’ s Modified Eagle’ s Medium (DMEM; Gibco) containing 10% fetal bovine serum (FBS; Gibco) and 1% penicillin-streptomycin (Gibco). The cell lines were tested negative for mycoplasma contamination by PCR, and were cultivated at 37 °C, 5% CO2 in the incubator.
Stable cell line generation
The HEK293T stable cell line was established using a lentiviral vector incorporating a puromycin resistance marker. We cloned into this vector a 200-bp DNA fragment that spanned a disease-associated mutation and included roughly 100 bp of flanking sequence on both sides. In brief, we used a combination of 12 μg of the specific lentiviral transfer plasmids (Lenti-GALT-sg1 and Lenti-OTC-sg1), along with 6 μg of pMD2.G and 9 μg of psPAX2, for co-transfection into HEK293T cells that were nearly 85% confluent in a 10 cm dish. At 72 h post-transfection, virus-containing supernatant was harvested. After centrifugation at 1699 × g (10 min, 4 °C), the supernatant was cleared of cell debris. The clarified solution was subsequently passed through a 0.45 µm membrane filter with minimal protein adsorption properties (Millipore). Then it was serially diluted and added to a 24-well plate, each well containing 5 × 104 HEK293T cells. We transferred the transduced cells to new plates after 24 h and added puromycin (2 µg ml−1) for selection. Post-puromycin selection (1 week), cells were plated at clonal density in 96-well plates. Isolated clones were expanded for subsequent transfection.
Cell transfection and fluorescence-activated cell sorting (FACS)
In DNA base editing experiments, HEK293T cells were plated in 24-well plates and transfected when they reached about 80% confluency. Next, cells were transfected with pre-formed DNA-PEI complexes containing 1 μg total plasmid DNA (750 ng ABE plasmid/250 ng sgRNA plasmid) and 3 μl PEI (Polysciences) in serum-free medium, with 15 min room temperature incubation prior to addition. After three days, transfected cells were digested with 0.25% trypsin (Gibco) for fluorescence-activated cell sorting (FACS). Genomic DNA was isolated using the QuickExtract™ DNA Extraction Solution (QE09050, Epicentre) according to the manufacturer’s instructions. For RNA off-target analysis, HEK293T cells were seeded into 10-cm dishes and transfected with 30 μg of Cas9n-P2A-GFP, ABE8e-P2A-GFP, ABE9-P2A-GFP, ABE8e-YA-P2A-GFP and ABE3.1-P2A-GFP using PEI at approximately 80% confluency. Three days after transfection, cells were washed with phosphate-buffered saline (PBS) and detached using 0.25% trypsin (Gibco). FACS was performed on a BD FACSAriaIII system (BD Biosciences) controlled by FACSDiva software (v8.0.2). Following doublet exclusion through forward/side scatter gating (Supplementary Fig. 9), the top 15% GFP-positive cells (300,000–400,000 cells) were collected. Total RNA was subsequently isolated from sorted cells using RNAiso Plus reagent (Takara).
Enhanced orthogonal R-loop assay
In this study, a modified orthogonal R-loop assay was employed for Cas9-independent DNA off-target analysis, which replaced dSaCas9-sgRNA plasmid with nSaCas9-sgRNA plasmid at each R-loop site. For the transfection process, a mixture of 3 μl polyethyleneimine (PEI, Polysciences), 850 ng plasmid DNA (250 ng SpCas9 sgRNA plasmid, 300 ng base editor plasmid (ABE8e, ABE9, ABE3.1, ABE8e-YA) and 300 ng nSaCas9 containing sgRNA plasmid were added to the cells. After three days, transfected cells were treated with 0.25% trypsin (Gibco) for sorting and then genomic DNA was extracted using the QuickExtract™ DNA Extraction Solution (QE09050, Epicentre), adhering to the manufacturer’s instructions.
RNA sequencing (RNA-Seq) experiments
RNA sequencing was performed as previously described in ref. 27. A total of 3 μg RNA per sample was used as input material for sample preparations. Sequencing libraries were prepared using an NEB Next Ultra RNA Library Prep Kit for Illumina (NEB) following the manufacturer’s recommendations. After assessing library quality with the Agilent Bioanalyzer 2100, index codes were added to assign sequences to individual samples. The index-coded samples were clustered on a cBot Cluster Generation System using the TruSeq PE Cluster Kit v3-cBot-HS (Illumina) according to the manufacturer’s guidelines. Following cluster generation, the libraries were sequenced on an Illumina HiSeq platform, generating 125-bp or 150-bp paired-end reads.
RNA sequence variant calling and quality control
RNA sequence variant calling and quality control were performed as previously described in ref. 23. Briefly, raw fastq format data were initially processed through in-house Perl Scripts. Adapter sequences were removed, and low-quality bases were trimmed to produce clean data using Trimmomatic. Concurrently, the Q20, Q30, and GC content of the clean data were calculated. All further analyses were conducted using this high-quality clean data. The reference genome index was built using HISAT2 version 2.0.5 and paired-end clean reads were aligned to the reference genome (Ensemble GRCh38) with the same HISAT2 version. Single-nucleotide polymorphism calling was carried out using GATK version 4.0. Variant loci in the base editor overexpression groups were filtered to eliminate sites without high-confidence reference genotype calls in the control group.
High-throughput DNA sequencing and data analysis
On-target and off-target regions were amplified by PCR using high-throughput sequencing primer pairs flanking these sites (listed in the Supplementary Data 1). PCR amplification was carried out with KOD-Plus-Neo DNA Polymerase according to the manufacturer’s instructions with 100–150 ng of genomic DNA as a template. For the preparation of high-throughput sequencing (HTS) libraries, site-specific primers containing an adaptor sequence (forward 5′-GGAGTGAGTACGGTGTGC-3′; backward 5′-GAGTTGGATGCTGGATGG-3′) at the 5′ end were employed in the PCR. Subsequently, the products were subjected to a second-round of PCR using primers with different barcode sequences. Then, barcoded PCR amplicons were pooled and sequenced on Illumina HiSeq (AnnoroadGene Technology sequenced Co., Ltd). FASTQ files were aligned to full-length as previously described29, with editing efficiencies calculated using BE-Analyzer19 or CRISPResso229.
Preparation of mRNAs and sgRNAs and Microinjection in mice
Phosphorothioate-modified and 2’-O-methyl-modified sgRNAs were synthesized by GenScript (Nanjing, China) (Supplementary Data 3). mRNA was prepared following established methods27. Briefly, for each ABE variant (ABE8e, ABE8e-YA, and ABE9), the template was separately PCR-amplified with primers T7-ABEs-mRNA-F/R (Supplementary Data 1), followed by in vitro transcription (mMESSAGE mMACHINE T7 Kit) and purification (MEGAclear Kit)31.
Wild-type female C57BL/6 J mice (4–6 weeks old) and ICR mice (8–12 weeks old), bred in-house, were used as embryo donors and foster mothers, respectively. Mice were housed in a specific pathogen-free (SPF) facility with a 12 h light/dark cycle, ad libitum access to food and water, ambient temperature maintained at 20–22 °C, and relative humidity of 40–60%. A 2 nl mixture containing either ABE8e, ABE8e-YA or ABE9 mRNA (200 ng μl−1) and sgRNA (100 ng μl−1) was co-injected into wild-type one-cell stage embryos. Injected zygotes were transferred into pseudopregnant female mice immediately using an Eppendorf TransferMan NK2 micromanipulator. Approximately 20 days post-transplantation, F0 pups were born. Genomic DNA was extracted from 1-week-old postnatal tail biopsies using the QuickExtract™ DNA Extraction Solution (QE09050, Epicenter) following the manufacturer’s protocol.
LNP treatment and serum analysis
All mouse cohorts were maintained at specific pathogen-free (SPF) facilities in Suzhou Institute of Systems Medicine and approved by Institutional Animal Care and Use Committee. Female C57BL/6 J mice (6–8 weeks old received intravenous tail vein injections of LNP-formulated ABE8e-YA or ABE8e mRNA complexed with Pcsk9-targeting sgRNA at a 2:1 ratio (Starna Therapeutics, Suzhou). Three dose levels were evaluated: 2 mg/kg, 0.5 mg/kg, and 0.125 mg/kg. Control mice were administered an equal volume of sterile saline. To assess the serum PCSK9 and low-density lipoprotein cholesterol (LDL-C) levels, mice were fasted overnight for 12 hours before blood collection via tail tip sampling. Blood samples clotted at room temperature. Serum samples from each time point were concurrently collected and was separated by centrifugation (3000 × g, 10 min). The Proteintech ELISA kit (#KE10050) was employed for PCSK9 quantification, while Solarbio kits #BC5335, #BC1555 and #BC1565 measured LDL-C, ALT, and AST levels respectively. Manufacturer-specified protocols guided all procedures. For terminal procedures, mice were euthanized by CO₂ inhalation. DNA was extracted from the excised median liver lobe to assess on-target and off-target site editing efficiency.
Statistics & reproducibility
Unless stated otherwise, data are presented as means ± SD from three biologically independent experiments performed on separate days. All bar plots and figures were generated using Prism 9.3 (GraphPad). The statistical significance of differences between two groups was assessed using an unpaired two-tailed Student’s t-test or two-sided paired Wilcoxon rank-sum test within Prism 10 (GraphPad). A P-value of less than 0.05 was considered to indicate significance. RNA-seq data was analyzed using Trim Galore (version 0.6.6), STAR (version 2.7.1a), SAMtools (version 1.14), Picard MarkDuplicates module (version 2.23.9) software. For the analysis of FACS data, FlowJo v.10 was employed. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Molecular visualization and comparative analysis
Structural models of the TadA8e complex and its mutant derivatives (TadA8e-A48E + RA and TadA8e-A48E + YA) were analyzed using PyMOL (v3.1.0a0 Open-Source). Initial coordinates for the wild-type complex were derived from the experimentally resolved structure (PDB ID: 6VPC). Site-directed mutagenesis (A48E) and ligand substitution (RA → YA) were modeled in PyMOL using the Mutagenesis and Ligand Docking modules, followed by energy minimization (AMBER force field) to optimize side-chain conformations.
Ethical statement
This study strictly adhered to all applicable ethical guidelines. All mouse-related procedures were carefully designed to comply with institutional standards and were approved by the Institutional Animal Care and Use Committee (IACUC) at the Suzhou Institute of Systems Medicine, Chinese Academy of Medical Sciences & Peking Union Medical College, Suzhou, China.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw high-throughput sequencing data and RNA-seq sequencing data generated in this study have been deposited to the NCBI Sequence Read Archive (SRA) under accession code PRJNA1193511. Plasmids encoding ABE8e-YA are available. Source data are provided with this paper.
Code availability
The code for calculating pathogenic point mutations suitable for ABE8e-YA in the ClinVar database is available at Code Ocean [https://codeocean.com/capsule/2242462/tree] without any access restrictions or usage limitations.
References
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Song, C. Q. et al. Adenine base editing in an adult mouse model of tyrosinaemia. Nat. Biomed. Eng. 4, 125–130 (2020).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson-Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Cui, C. et al. A base editor for the long-term restoration of auditory function in mice with recessive profound deafness. Nat. Biomed. Eng. 9, 40–56 (2025).
Verve Therapeutics. A Phase 1b Single Ascending Dose Study to Evaluate the Safety of VERVE-201 in Patients with Refractory Hyperlipidemia. ClinicalTrials. gov Identifier: NCT06451770 (2024).
Gehrke, J. M. et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 36, 977–982 (2018).
Lee, S. et al. Single C-to-T substitution using engineered APOBEC3G-nCas9 base editors with minimum genome- and transcriptome-wide off-target effects. Sci. Adv. 6, eaba1773 (2020).
Liu, L. D. et al. Intrinsic Nucleotide Preference of Diversifying Base Editors Guides Antibody Ex Vivo Affinity Maturation. Cell Rep. 25, 884–892.e883 (2018).
Liu, Z. et al. Improved base editor for efficient editing in GC contexts in rabbits with an optimized AID-Cas9 fusion. FASEB J. 33, 9210–9219 (2019).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Chen, L. et al. Re-engineering the adenine deaminase TadA-8e for efficient and specific CRISPR-based cytosine base editing. Nat. Biotechnol. 41, 663–672 (2022).
Lam, D. K. et al. Improved cytosine base editors generated from TadA variants. Nat. Biotechnol. 41, 686–697 (2023).
Neugebauer, M. E. et al. Evolution of an adenine base editor into a small, efficient cytosine base editor with low off-target activity. Nat. Biotechnol. 41, 673–685 (2022).
Chen, L. et al. Engineering a precise adenine base editor with minimal bystander editing. Nat. Chem. Biol. 19, 101–110 (2022).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Lapinaite, A. et al. DNA capture by a CRISPR-Cas9-guided adenine base editor. Science 369, 566–571 (2020).
Hwang, G. H. et al. Web-based design and analysis tools for CRISPR base editing. BMC Bioinform 19, 542 (2018).
Jeong, Y. K. et al. Adenine base editor engineering reduces editing of bystander cytosines. Nat. Biotechnol. 39, 1426–1433 (2021).
Doman, J. L. et al. Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors. Nat. Biotechnol. 38, 620–628 (2020).
Grünewald, J. et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors. Nature 569, 433–437 (2019).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Yang, Y. P. et al. Molecular analysis in newborns from Texas affected with galactosemia. Hum. Mutat. 19, 82–83 (2002).
Oppliger Leibundgut, E. O. et al. Ornithine transcarbamylase deficiency: new sites with increased probability of mutation. Hum. Genet 95, 191–196 (1995).
Yamaguchi, S. et al. Mutations and polymorphisms in the human ornithine transcarbamylase (OTC) gene. Hum. Genet 27, 626–632 (2006).
Zhang, X. et al. Increasing the efficiency and targeting range of cytidine base editors through fusion of a single-stranded DNA-binding protein domain. Nat. Cell Biol. 22, 740–750 (2020).
Fitzgerald, K. et al. Effect of an RNA interference drug on the synthesis of proprotein convertase subtilisin/kexin type 9 (PCSK9) and the concentration of serum LDL cholesterol in healthy volunteers: a randomised, single-blind, placebo-controlled, phase 1 trial. Lancet 383, 60–68 (2014).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Xia, B. et al. On the genetic basis of tail-loss evolution in humans and apes. Nature 626, 1042–1048 (2024).
Yang, L. et al. Increasing targeting scope of adenosine base editors in mouse and rat embryos through fusion of TadA deaminase with Cas9 variants. Protein Cell 9, 814–819 (2018).
Acknowledgements
We thank Professor Dali Li from East China Normal University for generously providing a variety of base editor plasmids, which greatly accelerated the progress of this project during the establishment of our research group. We thank Yuanqing He and Naiyun Ma from the Laboratory Animal Science Center of Suzhou Institute of Systems Medicine for their help in generating base-edited mice. We thank Chunyi Hu from the Faculty of Science of National University of Singapore for crucial guidance in elucidating the working mechanism of ABE8e-YA from a structural biology perspective. This work was partially supported by grants from National Key R&D Program of China (No.2022YFC3400200 to X.Z. and No.2022YFA1103400 to X.Z.), the National Natural Science Foundation of China (No.82522046 to X.Z. and No.32201223 to X.Z.), the NCTIB Fund for R&D Platform for Cell and Gene Therapy, the CAMS Innovation Fund for Medical Sciences (No.2022-I2M-1-024 to X.Z., No.2022-I2M-2-004 to X.Z. and No. 2023-I2M-2-005 to X.Z.), the Suzhou Municipal Key Laboratory (No.SZS2022005), the Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (No.2022-RC180-08 to X.Z.) and the Special Research Fund for Central Universities, Peking Union Medical College (No.3332024087 to Y.Z.).
Author information
Authors and Affiliations
Contributions
X.Z. designed and supervised the project. M.S., Y.L., and J.R. performed most of the experiments for base editing in HEK293T cells and mouse embryos with the help of Y.Z. and J.W.; Q.C. performed the analysis for base editors to correct pathogenic point mutations in the ClinVar database and the experiments for base editing in vivo in mice. X.Z. wrote the manuscript with the input from all the authors and R.V.G. polished it.
Corresponding author
Ethics declarations
Competing interests
X.Z., H.L., J.R., Y.L., and Y.H. have submitted patent applications (application no. 202311786372.7, under review) based on the results reported in this study. This patent mainly relates to ABE8e-YA in this paper. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shang, M., Li, Y., Cao, Q. et al. A motif preferred adenine base editor with minimal bystander and off-targets editing. Nat Commun 16, 9153 (2025). https://doi.org/10.1038/s41467-025-64203-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64203-6