Abstract
Methane hydroxylation presents a promising approach to produce high-energy methanol from potent greenhouse gases, thereby contributing to a more sustainable future. Despite its environmental importance, current research on this process remains challenging due to the harsh operating conditions for the activation of inert C-H bond in methane. In nature, methane monooxygenase converts methane by activating the C-H bonds through its hydroxylase, under ambient conditions, receiving electrons from NADH via a reductase. Beyond the traditional biological approach, the development of NADH-independent biocatalytic systems could open new avenues for cost-effective and sustainable methane conversion. Herein, we report an NADH-free biosolar platform that activates hydroxylase for eco-friendly methanol production. The xanthene-based light harvester spontaneously associates with hydroxylase and directly transfers its photoexcited electrons to the diiron active site, eliminating the need for a cofactor or reductase. Halogenation of xanthene accelerates direct electron transfer to the active site by increasing the polarizability and spin-orbit coupling of the light harvesters. Accordingly, the direct photobiocatalytic platform achieved a methanol time yield of 7.52 mmol gcat−1 h−1. This work provides the design concept of solar-driven biocatalytic methane hydroxylation under ambient conditions, suggesting a promising approach for implementing methanol biomanufacturing.
Similar content being viewed by others
Introduction
Methane is a major greenhouse gas, with a global warming potential approximately 30 times higher than that of carbon dioxide1,2. Significant efforts have been devoted to the valorization of methane gas, including decarbonization3,4, syngas production5,6, and chemical synthesis7,8,9. While methane liquefaction into methanol is the most promising option due to its utility as a liquid fuel and basic building block10, the process requires high energy and cost due to the harsh operating conditions (T > 800 °C; P > 20 bar) to activate the inert C–H bonds in methane11. Recently, photocatalytic conversion has gained attention as it utilizes photon energy rather than thermal energy. Leveraging photon energy, the photocatalyst activates methane via oxygen-containing radicals generated from its photogenerated holes, leading to methanol production at room temperature8,12. However, oxygen-containing radicals can also overoxidize methanol to produce undesired carbon dioxide13,14. Although a few photocatalysts have achieved high methanol selectivity (>90%) by controlling the active radicals, their catalytic productivities remain unsatisfactory7,14,15. Therefore, innovative catalytic platforms are required to achieve both high selectivity and productivity.
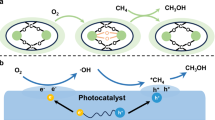
Methane monooxygenase (MMO) in methanotrophs catalyzes O2-dependent methane oxidation with high methanol selectivity under ambient conditions16,17,18,19,20,21. In the soluble form of the enzyme (sMMO), methane conversion is achieved using three separate protein components: hydroxylase (MMOH), reductase (MMOR), and regulatory protein (MMOB). MMOH activates methane via a glutamate/histidine-coordinated dinuclear iron center22,23, while MMOR transfers two electrons from nicotinamide adenine dinucleotide cofactors to MMOH via its electron transfer domain (flavin adenine dinucleotide; [2Fe-2S], ferredoxin) from nicotinamide adenine dinucleotide cofactors (NADH), priming it to react with O2 to form methanol24. MMOB enhances the activity of MMOH, facilitating substrate access to diiron active sites (Fig. 1a)18,25. The overall sMMO-driven reaction is as follows:
a Natural, cofactor-dependent MMOH reduction assisted by MMOR and MMOB. Upon the electron transfer by NADH cofactor and MMOR to diiron active site, MMOH catalyzes the methane hydroxylation using O2 as a green oxidant. MMOB facilitates substrate access to active site. b Biosolar MMOH reduction using XD as a light harvester. The XD transfers its photoinduced electrons directly to the active site, gaining electrons from sacrificial donors (TEOA and water), and then MMOH drives the methane hydroxylation under solar illumination. FL fluorescein (X = H, Y = H), EY eosin Y (X = Br, Y = H), ErB erythrosine B (X = I, Y = H), RB rose bengal (X = I, Y = Cl), MMOHox resting diferric MMOH, MMOHred diferrous MMOH, kET electron transfer rate.
Despite its effectiveness, reliance on NADH during the catalytic process can pose challenges for the widespread application of MMO systems26,27.
Herein, we present an NADH-free, direct biosolar platform for the conversion of methane to methanol, featuring high catalytic productivity using xanthene dyes (XDs) as light harvesters. Photobiocatalysis, which integrates light-driven processes with biocatalysts, presents a sustainable approach to renewable energy conversion28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43. By utilizing light energy to drive selective redox reactions, it allows for efficient and environmentally friendly chemical transformations. Within this regime, XDs are promising photosensitizers that directly activate redox biocatalysts, harvesting visible light with a high quantum yield44, and have been hybridized with the MMO system for photobiocatalytic methane conversion in this study. Eosin Y (EY) and its derivatives are complexed with MMOH and transfer their photoinduced electrons to the diiron active center of MMOH for high-yield methanol production (Fig. 1b). Furthermore, the photoinduced electron transfer from XDs to MMOH is facilitated by the strong spin-orbit coupling induced by the heavy halogen atoms in xanthene. Among the XDs, rose bengal (RB) exhibited the fastest rate of electron transfer (kET = 1.46 × 1011 s−1) due to the heavy halogen atoms, and consequently achieved a methanol time yield (7.52 mmol gcat−1 h−1) under ambient conditions.
Results
Direct photoreduction of MMOH
We first examined the possibility of complex formation between the XD and MMO for solar-hybrid biocatalyst development to achieve methane conversion. To this end, we used MMOH as a model enzyme and EY as a model light-harvesting dye because MMOH has a supreme oxygenation activity among the sMMO components (MMOH, MMOR, and MMOB) and the EY is a most common light harvester to reduce redox enzymes (Fig. 2 and Supplementary Figs. 1–5)28,45. In absorption spectra, the EY revealed a red shift of absorption maximum with increasing concentrations of MMOH, suggesting the association of EY with MMOH (Fig. 2a and Supplementary Fig. 4). EY-MMOH coupling was confirmed using 1H nuclear magnetic resonance (NMR) spectroscopy (Fig. 2b). The addition of MMOH to the EY solution resulted in an upfield shift in the outer rim protons of xanthene (Ha and Hb) and benzoate (Hc, Hd, He, and Hf) due to the shielding effect of the functional groups (e.g., carboxylate, amide, and imidazole) in MMOH. Surface-enhanced Raman spectroscopy was used to further investigate the specific interactions between EY and MMOH (Fig. 2c). In the Raman spectra, the characteristic peaks of aromatic ring stretch (~645 cm−1)/deformation (~712 cm−1), C–O–C ring stretch (~1180 cm−1), and asymmetric stretch of carboxylate (~1560 cm−1) in EY46 disappeared after the coupling with MMOH due to the noncovalent interactions (e.g., π–π stacking, hydrogen bonding, and ion-dipole interaction) between the dye molecules and proteins. The reduced infrared peak intensities of the C–O (~1090 cm−1), C=C (~1605 cm−1), and C=O (~1730 cm−1) stretches47 in the EY/MMOH complex corroborated the self-association of EY with MMOH through noncovalent interactions (Supplementary Fig. 5). We investigated the possibility of sMMO reduction by photoexcited EY molecules using photoluminescence (PL) spectroscopy. Oxidative fluorescence quenching of EY was observed with MMOH and MMOR, but not with MMOB (Fig. 2d and Supplementary Fig. 6). MMOB without redox moieties cannot receive photoinduced electrons from EY. The Stern–Volmer constant of MMOH (1.95 ×104 M−1) was two times higher than that of MMOR. This result was ascribed to the facile binding of EY to MMOH and the rapid photoinduced electron transfer to the diiron active center. To elucidate the detailed electron transfer mechanism in MMOH, we measured the electron paramagnetic resonance (EPR) of the protein (Supplementary Fig. 7). The EPR signal of MMOH in a diferrous state (FeIIFeII) at g = 16 appeared upon illumination in the presence of EY, while other ones at g = 4.3 [ferric ion (FeIII)] and g = 1.83 [mixed-valent form (FeIIFeIII)] vanished48. The result indicates the EY-sensitized reduction of the diiron active site in MMOH.
a Spectroscopic analysis of interaction between EY (10 µM) and MMOH (2–10 µM) in an aqueous solution. b 1H NMR (400 MHz, 298 K) spectra of EY and EY/MMOH complex solution. Sample: deuterium oxide solution containing EY (100 µM) and MMOH (2–10 µM). The characters of a–f indicate the hydrogen of EY depicted in the inset. c Surface-enhanced Raman spectra of EY and EY/MMOH complex. Substrate: hydrophilic Ag nanoparticles. Excitation: 633 nm. Aqueous solution (5 µL) containing EY (10 µM) with or without MMOH (10 µM) was dropcasted on the Ag substrate, where the solution dried for 24 h for Raman experiments. d Stern–Volmer plots for the oxidative quenching of EY (1 µM) with proteins (MMOH, MMOR, and MMOB) in phosphate buffer (100 mM, pH 7). Excitation: 500 nm. Ksv indicates a Stern–Volmer constant.
Electron transfer from EY to MMOH
In our computational workflow (Fig. 3), we highlight the initial electron acceptor on MMOH surface, providing insights into its role in the electron transfer from EY to MMOH. Blind docking simulations revealed several binding clusters (Supplementary Fig. 8). Among these, the cluster 2 was found to the most probable binding site based on its optimal binding energy (−6.83 kcal/mol) and proximity to the diiron cluster (EY–Fe distance: ~15 Å) (Fig. 4a, b and Supplementary Fig. 8). To refine this binding pose, we performed energy minimization followed by 100 ns of molecular dynamics (MD) simulations under isothermal-isobaric ensemble (NPT) conditions. The cluster 2 remained stable throughout the 100 ns simulation, maintaining the ~16 Å EY–Fe distance, while other clusters showed significant fluctuations with EY–Fe separations (Supplementary Fig. 9). The convergence of the protein root-mean-square deviation (RMSD) across three independent 100 ns MD runs underscores the robustness and reliability of our simulations (Supplementary Fig. 10). Throughout the simulations, K323 consistently maintained a ~4 Å distance from EY and formed frequent hydrogen bonds, whereas R320 exhibited larger fluctuations (~6 Å) (Supplementary Fig. 11). The relative binding energy (ΔGbind) of EY to the surface of MMOH was calculated using molecular mechanics-generalized born surface area (MM-GBSA). The calculated binding free energy (∆∆GMM-GBSA) was −3.51 kcal/mol related to suitable noncovalent bond between the dye and MMOH (Supplementary Table 1). The docked EY molecule was encompassed by the residues K323, R320, W317, M247, Y251, Y324, and V255 (Fig. 4c). To identify the primary electron acceptor, we assessed the spin densities and electron transfer rates (kET). Based on our analysis, K323 exhibited the highest spin density of 0.78 e/Å3, followed by R320, W317, M247, Y251, Y324, and V255 (Fig. 4d). The kET from EY to K323 (kET = 1.61 × 106 s−1) is the highest, significantly surpassing the rates to R320 (1.68 × 105 s−1), W317 (1.46 × 101 s−1), M247 (3.83 × 102 s−1), Y251 (3.01 × 104 s−1), Y324 (1.64 × 104 s−1), and V255 (2.80 × 104 s−1) (Table 1). The low reorganization energy (λ = 2.84 eV) facilitates the high kET, implying that less molecular structural adjustment is required for the electron transfer. Furthermore, K323 has negative free energy of reaction (∆G° = −0.40 eV), indicating that the electron transfer process is spontaneously favorable. In addition, the shortest distances between the electron-emitting oxide ion of EY on the tricyclic ring and the residues were measured to ascertain their spatial layouts. Computational analyses revealed a complex electron transfer mechanism from EY to MMOH, particularly on the MMOH surface. Our computational analyses have identified lysine residue K323 as the primary electron acceptor, which is substantiated by several critical factors, including the high kET, minimal reorganization energy, negative free energy suggesting spontaneous transfer, lowest activation energy barrier for swift electron transfer, and close proximity to EY. Experimental results, such as the decrease in binding affinity and methanol productivity of the K323A (lysine at K323 changed to alanine), R320A (arginine at R320 changed to alanine), and R320A/K323A mutants compared with MMOH wild-type (WT) supported the computational insights, which highlights the indispensable role of the lysine residue K323 in maintaining the functional integrity of MMOH (Supplementary Figs. 12 and 13).
a Blind docking is performed to determine the binding site of EY (dye) on the MMOH protein surface. b The result of blind docking reveals clusters of docked EY on the MMOH surface. The largest cluster, representing the binding site, is highlighted in the inset. c Potential first electron acceptor residues selected based on their proximity to the best binding pose of EY. d Marcus curve analysis utilized to confirm the first electron acceptor residue, considering EY as the donor and the identified acceptor residue.
a Blind docking result of EY on the entire surface of the MMOH protein. The largest EY cluster with the best binding energy is analyzed, considering the diiron core’s location in the α subunit of MMOH. b The statistical study identified the largest cluster based on a 2.0 RMSD cutoff. The inset shows the cluster with the largest number of conformations (seven) among the top five clusters and the lowest average binding energy. c The resulting binding pose of EY on the MMOH structure. The pose highlights the proximity of EY to neighboring residues as potential candidates for the first electron acceptor. d Spin-density values of the residues on the MMOH surface. K323 exhibits the highest spin density. Error bars represent the means ± SD (n = 3).
Photobiocatalytic methane hydroxylation
Protein film voltammetry was used to investigate the roles of MMOH, MMOR, and MMOB in methane hydroxylation. In cyclic voltammograms, the cathodic current between 0 and −0.2 V [vs. reversible hydrogen electrode (RHE)] increased with the addition of MMOH and methane, suggesting electron transfer from a glassy carbon electrode (GCE) to MMOH for methane activation (Supplementary Fig. 14a). The cathodic current further increased when MMOH was coupled with MMOB in a methane-saturated buffer (Supplementary Fig. 14b), indicating that MMOH activates methane by utilizing electrons from the GCE, while MMOB facilitates methane access to MMOH18,25. In contrast, the addition of MMOR to MMOH decreased the cathodic current of MMOH due to electron dissipation by MMOR. Furthermore, we measured the transient photocurrent responses of EY-stained proteins to examine the possible electron transfer between EY and proteins under visible light. Figure 5a depicts a photocurrent in the presence of MMOH, which further increases with the addition of methane and MMOB. The photocurrent responses indicated that EY transferred its photoinduced electrons to MMOH for methane activation, whereas MMOB expedited the catalytic turnover of methane. In contrast, MMOR reduced the photocurrent of the EY/MMOH complex by dissipating the photoexcited electrons that MMOH could use for methane activation. This hypothesis is supported by the PL quenching of EY by MMOR and the photocurrent of EY/MMOR, which are evidence of the EY-sensitized reduction of MMOR (Figs. 2d and 5a and Supplementary Figs. 15 and 16). The charge transfer resistance between EY and the MMOH/MMOB complex exhibited 1.75-fold decrease after light illumination (Fig. 5b and Supplementary Fig. 17), indicating that protein reduction by photoexcited EY occurred much faster than that by GCE49,50,51,52.
a Transient photocurrent response of EY with proteins (MMOH, MMOR, and MMOB) at −0.09 V (vs. RHE). b Nyquist plots of EY with proteins (MMOH and MMOB) at −0.09 V (vs. RHE). Inset: equivalent circuit for the plots. Red line represents the fitted data. Electrolyte of (a, b): methane-saturated phosphate buffer (100 mM, pH 7). Working electrode of (a, b): glassy carbon electrode coated with EY and protein mixture solution. c Control experiments on a methanol time yield of the photobiocatalytic methane conversion platform. Electrolyte: methane-saturated TEOA buffer (100 mM, pH 7.5) containing MMOH (2.5 µM), MMOR (5 µM), and MMOB (5 µM) with EY (20 µM). Methanol time yield was determined at 1-h reaction. d Methanol time yield and quantum yield of EY/MMOH/MMOB in terms of photon flux. Light source of (a–d): xenon lamp, I = 0.58 µE cm−2 s−1, λ > 400 nm. Error bars represent the means ± SD (n = 3). e Energy diagram for a conventional methane hydroxylation by multi-protein components (MMOH, MMOR, and MMOB). f Energy diagram for a photobiocatalytic methane hydroxylation by EY. The oxidation potentials of ground-state and excited-state EY were obtained from voltammetric/spectroscopic analyses. The redox potentials of NADH and redox proteins (MMOH, MMOR, and protein complexes) were based on the literature25,27,55.
We assessed the photobiocatalytic performances of EY-protein complex under ambient conditions (T = 25 °C; P = 1 atm) with visible light irradiation (I = 0.58 µE cm−2 s−1, λ > 400 nm). According to our results (Supplementary Figs. 18 and 19), a triethanolamine (TEOA) functions as an optimal electron donor to sustain photobiocatalytic methane hydroxylation. Control experiments involving each reaction component (TEOA, light harvester, sMMO, methane, and light) produced negligible methanol, confirming that the photobioreduction by EY is a crucial step in methane hydroxylation (Fig. 5c). Methane isotopic tests using 13C methane disclosed that the generated methanol originated from a photobiocatalytic methane hydroxylation (Supplementary Fig. 20)53. When coupled with either MMOB or MMOR, the EY/MMOH complex demonstrated increased (1.36 times) and decreased (1.22 times) methanol time yields, respectively. MMOB accelerated methane activation in MMOH, while MMOR hindered the activation by consuming photoexcited electrons from EY. As a result, EY/MMOH/MMOB exhibited the highest methanol time yield (4.13 mmol gcat−1 h−1) and methanol productivity (22.59 mmol gcat−1; 6 h reaction) among EY-protein complexes (Fig. 5c and Supplementary Fig. 21). Isothermal titration calorimetry (ITC) results indicate the binding interactions between ligands (EY and MMOR) and MMOH are non-competitive (Supplementary Fig. 22 and Supplementary Table 2). We compared methanol production from various MMO configurations differing in enzyme source (native or recombinant) and reducing equivalents (NADH or EY) (Supplementary Fig. 23). While the recombinant MMO showed lower activity—possibly due to misfolding or incomplete complex assembly—the EY-driven native system exhibited a 1.9-fold higher activity than the native NADH system. We speculate that the intact configuration of key residues in the native enzyme (e.g., K323, R320) facilitates efficient photoinduced electron transfer to MMOH, thereby contributing to the enhanced photocatalytic performance. Increasing the photon flux further boosted the initial production rate, showing a high quantum yield of 0.13 (Fig. 5d). Guided by voltammetric and spectroscopic analyses, we constructed an energy diagram for the direct activation pathway of MMOH, in which EY exhibits a sufficiently high driving force (excited-state oxidation potential: −0.58 V vs. RHE) to reduce MMOH (Fig. 5e, f, Supplementary Fig. 24 and Supplementary Table 3)27,54,55.
Biosolar methane hydroxylation by xanthene derivatives
Next, we investigated the structural effects of XD molecules on the photobiocatalytic methane hydroxylation. We selected nonhalogenated [fluorescein (FL)] and halogenated [EY, Erythrosine B (ErB), and RB] dye molecules for our study (Fig. 1b and Supplementary Figs. 25 and 26) due to the influence of halogen atoms on the physicochemical and photophysical properties of XD56. The binding affinity of each XD for MMOH using ITC (Fig. 6a, Supplementary Fig. 27, and Supplementary Table 4) and was confirmed to be driven enthalpically. RB exhibited the highest binding affinity to MMOH (with a lowest Kd of 9.23 ± 1.3 μM) compared to ErB, EY, and FL. The high polarizability of RB due to heavy halogen atoms increases the hydrophobic interactions between halogenated xanthene molecules and proteins57, leading to its high binding affinity to MMOH. This favorable binding of RB to MMOH is desirable for photobiocatalytic methane hydroxylation. The XD-sensitized reduction of MMOH was further investigated using PL spectroscopy. All XDs exhibited oxidative fluorescence quenching in the presence of MMOH, indicating the photoreduction of MMOH by the XDs (Supplementary Figs. 28 and 29). Moreover, the Stern–Volmer constants increased as XD halogenation increased. With heavier halogen atoms, the average PL lifetime decreases in the following order: RB (0.83 ns) < ErB (1.30 ns) < EY (2.01 ns) < FL (4.33 ns) (Fig. 6b). The heavy halogen atoms in xanthene expedite the intersystem crossing of electrons by increasing the spin-orbit coupling between the singlet-triplet states56. Accordingly, the fluorescence lifetime and PL intensity from the singlet states decreased as the triplet formation was enhanced by strong spin-orbit coupling (Fig. 6c). Overall, RB with heavy iodine and chlorine atoms exhibited the fastest decay lifetime (0.83 ns) and the highest Stern–Volmer constant (4.15 × 104 M−1) among the XDs, indicating a rapid photoinduced electron transfer from RB to MMOH. We further evaluated thermodynamic indices, such as the oxidation potentials of the ground and excited states of XDs, for electron transfer involved in MMOH reduction by XDs to compare the driving forces of the redox reactions. Upon excitation of XD by visible light, the photoinduced electrons on the excited-state of XD exhibit sufficient potential (FL: −0.55 V vs. RHE, EY: −0.58 V, ErB: −0.62 V, and RB: −0.69 V) to drive MMOH reduction (reduction potential of 0.33 V vs. RHE)55, and TEOA oxidation (oxidation potential of 1.46 V vs. RHE)27 stabilizes XD to sustain the photocatalytic turnover (Supplementary Figs. 30–32 and Supplementary Table 5). The excited RB molecules exhibit the highest oxidation potential (−0.69 V vs. RHE) with the most negative Gibbs energy (−1.02 eV) to reduce MMOH, indicating that RB can act as the most effective light harvester for MMOH reduction among xanthene derivatives (Supplementary Tables 3 and 5).
a Dissociation constants (Kd) of XDs to MMOH calculated from ITC experiments. Error bars represent the means ± SD (n = 3). b Fluorescence decay spectra of XDs (1 µM) with MMOH (6 µM) in phosphate buffer (100 mM, pH 7). Excitation: 467 nm. Emission: 518 nm (FL), 538 nm (EY), 555 nm (ErB), and 580 nm (RB). The lifetime was estimated by fitting the spectra with a triple-exponential function. c Energy diagram of XDs for the direct photoreduction of MMOH. Substitution of atoms to heavier halogens in XDs increases the intersystem crossing (ISC)-induced triplet formation by the strong spin-orbit coupling (SOC), thereby enhancing the efficiency of photoinduced electron transfer (PET) from XDs to MMOH. d Methanol time yield and quantum yield of XD/MMOH. Electrolyte: methane-saturated TEOA buffer (100 mM, pH 7.5) containing XD (20 µM) and MMOH (2.5 µM). Light source: xenon lamp, I = 0.58 µE cm−2 s−1, λ > 400 nm. Error bars represent the means ± SD (n = 3). e Comparison of catalytic performances for methane hydroxylation over RB/MMOH/MMOB complexes with other catalysts12,13,19,53,62,63,64,65,66,67,68. The dashed line represents catalysis that occurs under ambient conditions (T = 25 °C; P = 1 atm).
We conducted blind docking simulations for the XD/MMOH complexes to investigate the binding of XD to MMOH and the subsequent electron transfer between them. RB exhibited the lowest binding free energy (−7.65 kcal/mol) among the light harvesters, indicating the highest binding affinity to the MMOH surface (Supplementary Table 1). The MM-GBSA simulations provided a more accurate binding free energy by considering the MM interactions and solvation effects58,59,60,61. Following docking analysis, the MM-GBSA results further supported the higher binding interaction of RB with the MMOH surface; RB exhibited the lowest binding free energy (−5.26 kcal/mol) in the molecular mechanics Poisson-Boltzmann surface area (MMPBSA) simulations, indicating a highly favorable and stable binding interaction compared to other XDs. In addition, RB (kET = 1.46 × 1011 s−1) exhibits a significantly higher rate of electron transfer to the first acceptor residue K323 than the other dyes, ErB (kET = 1.91 × 108 s−1), EY (kET = 1.61 × 106 s−1), and FL (kET = 2.77 × 104 s−1) (Supplementary Figs. 33 and 34, and Table 2). This is mainly due to the lowest reorganization energy of RB (λ = 1.57 eV), indicating minimal structural rearrangement during the electron transfer. Lower reorganization energy reduces the energy barrier (∆G‡) to transfer electrons, accelerating kET. Furthermore, RB has the highest driving force (∆G° = −0.53 eV), and shortest donor–acceptor distance (R = 3.1 Å) for a foster proficient electron transfer among all the XDs, leading to the highest kET (Table 2). The results are supported by a strong correlation between the kET, reorganization energy, and distance between the donor and acceptor (Supplementary Fig. 35). We estimated and compared the catalytic productivities of various XD-sensitized MMOH under visible light. The RB/MMOH complexes exhibited the highest photobiocatalytic performance, with a methanol time yield of 4.93 mmol gcat−1 h−1, a quantum yield of 0.15, and an inactivation rate of 8.73% h−1 compared to ErB (3.93 mmol gcat−1 h−1, 0.12, and 9.9% h−1), EY (3.04 mmol gcat−1 h−1, 0.09, and 10% h−1), and FL (1.20 mmol gcat−1 h−1, 0.04, and 9.49% h−1) (Fig. 6d and Supplementary Fig. 36). The XD’s photobleaching can be responsible for the inactivation (Supplementary Fig. 37). RB is the most potent light harvester for activating MMOH due to favorable binding to the protein and strong redox power to reduce the diiron active site. To demonstrate full-fledged biosolar methane hydroxylation under ambient conditions (T = 25 °C; P = 1 atm) and solar light (at an air mass 1.5 global intensity, 100 mW cm−2), we evaluated the catalytic performance of MMOH using RB as a light harvester and MMOB as a catalytic turnover enhancer. The RB/MMOH/MMOB complexes produced 39.8 mmol gcat−1 of methanol over 6 h with an initial rate of 7.52 mmol gcat−1 h−1 (Supplementary Fig. 38). Furthermore, the catalytic activity of RB/MMOH/MMOB for methane hydroxylation to methanol was confirmed by methane isotopic experiments using 13C methane (Supplementary Fig. 39)53.
The RB-MMO platform outperformed cutting-edge thermal, photo, and biocatalysts for methane hydroxylation (Fig. 6e and Supplementary Table 6). The reported thermal catalysts, such as metal-organic frameworks13,62 and zeolites63,64,65, require harsh operating conditions (T > 200 °C; P > 6 atm) to achieve meaningful catalytic turnover (0.001–3.8 mmol gcat−1 h−1), which is highly energy-intensive. Photoactive metal oxides12,53,66,67 and methane-oxidizing biocatalysts19,68 have enabled methane-to-methanol conversion under mild conditions, but their catalytic productivity remains unsatisfactory (<0.4 mmol gcat−1 h−1). In contrast, our biosolar platform achieved benchmark productivity (7.52 mmol gcat−1 h−1), owing to the direct photoreduction of MMOH by a powerful RB light harvester, ranking among the state-of-the-art catalysts for methane conversion.
Discussion
In summary, we designed a biosolar platform for conversion of methane to methanol by coupling XD light harvesters with MMOH. In the photobiocatalytic process, XD generates photoinduced electrons and subsequently reduces MMOH, thereby facilitating methane activation. Our spectroscopic and computational investigations revealed that the rate of photoinduced electron transfer from XD to MMOH was enhanced by the strong spin-orbit coupling induced by the heavy halogen atoms in XD. Thus, RB with heavy halogen atoms exhibited the highest rate of electron transfer (kET = 1.46 × 1011 s−1) to MMOH compared to those of other dyes (kET < 1.90 × 108). Consequently, the RB-MMO system achieved the solar bioconversion of methane to methanol at a production rate of 7.52 mmol gcat−1 h−1. Our direct biosolar platform successfully demonstrates (1) eco-friendly chemical synthesis using a potent greenhouse gas (methane), (2) facile methane activation by XD-MMO complexes under ambient conditions, and (3) high catalytic turnover of MMOH with high selectivity. This study presents a pioneering example of biosolar methane conversion to methanol using light as a clean energy source under ambient conditions.
Methods
Spectroscopic characterizations
The absorption spectra of XD, with and without proteins (MMOH, MMOR, and MMOB), were acquired using a V-650 spectrophotometer (JASCO). 1H NMR spectra were obtained using a 400 MHz NMR spectrometer (Bruker) with a water suppression technique to enhance the signal-to-noise ratio. Quantitative 1H NMR was performed to quantify the amount of methanol using a maleic acid as a standard. To this end, we mixed the reaction sample (540 μL) with a standard solution [maleic acid (10 mM) in deuterium oxide, 60 μL]. We calculated the methanol quantity using Eq. (2).
where Canalyte and Cstandard are the molar concentrations of analyte and standard, Nanalyte and Nstandard are the proton numbers corresponding to the peaks of analyte and standard, Ianalyte and Istandard are the integrals of the peaks of analyte and standard. Mass spectra were recorded on an XEVO G2-XS QTof (Waters Corporation). Raman spectra (excitation wavelength: 633 nm) were collected using an ARAMIS (Horiba Jobin Yvon) instrument with hydrophilic Ag nanoparticles to improve the signal-to-noise ratio. The infrared peak of the functional group on the XD with or without proteins was confirmed using a Nicolet iS50 (Thermo Fisher Scientific Instrument). PL spectra (excitation: 470 nm (FL), 500 nm (EY), 520 nm (ErB), or 530 nm (RB)) were recorded on an RF-5301PC spectrofluorophotometer (Shimadzu). Time-resolved PL decay profiles (excitation: 467 nm) were obtained using a Fluorolog3 spectrofluorometer (HORIBA). Fluorescence decay curves were obtained using a time-correlated single-photon counting approach with a time resolution of 10 ps. The fluorescence lifetimes were estimated using the best fit to a triple-exponential function (Eq. 3):
where α and τ are the amplitude and decay time, respectively, and I(t) represents the fluorescent intensity at a given time, t. The average lifetimes of τ (τavg) were evaluated using Eq. (4).
Details for simulations of charge transfer
The computational workflow depicted in Fig. 3 reveals the electron transfer mechanism between the light harvester and MMOH. Starting with blind docking (Fig. 4a), the workflow locates the binding site of the EY dye on the MMOH surface, yielding multiple clusters of docked molecules. The inset in Fig. 4b shows the largest cluster, indicating the most probable binding sites. Next, the potential first electron acceptor residues were singled out (Fig. 4c), which were close proximal to the optimal binding mode of EY and potentially engaged in the initial electron transfer. Subsequently, Marcus’s curve analysis was used to validate the first electron-acceptor residue. This computational workflow highlights the complex electron transfer from the light harvester to MMOH, including binding-site identification and electron-acceptor selection69,70.
Blind docking analysis
The crystal structure data for sMMO (PDB ID:6VK5) were obtained and analyzed by selecting chains including the D and H chains (MMOB) for further computation. Although only chain A was necessary for the calculation, the selection of additional chains prevented blindly docked EY from binding to the chain-binding region of MMO, thereby ensuring a more accurate analysis. The EY structure was retrieved from the PubChem database (PubChem ID: 11048)71. AutoDock Tools 1.5.772 was used for all the EY-to-MMO molecular docking experiments. All residues in the active site were included in the blind grid computation, with a 0.5 Å spacing between atoms, and the box was fixed at 126 × 126 × 126 Å. One thousand Lamarckian Genetic Algorithm (LGA) runs were performed, and the computation was discontinued after a maximum of 25 × 106 energy assessments. The crossover rate of 0.8 was used, and the population size was restricted to 150 based on the Lamarckian GA58. The conformation with the lowest binding energy from the largest cluster was selected and examined using MD simulations. Maestro (Schrodinger 2022 suite)73 was used to visualize the hydrophobic interactions and hydrogen bonds between ligands and proteins.
Molecular dynamics simulation and molecular mechanics-generalized born surface area analysis
MD simulations for interaction analysis and evaluation of binding free energy were performed using Desmond (Schrodinger 2022 suite)74. All systems were arranged using the “System Builder” in Maestro. The selected docking results were subjected to energy minimization and placed in the orthorhombic box with a buffer distance of 10 Å to create a hydration model. The SCP water model75 was used to develop the hydration model. The cutoff radii for the van der Waals force, time step, initial temperature, and system pressure were set to 9 Å, 2.0 fs, 300 K, and 1.01 bar, respectively. The electrostatic force was evaluated by dividing the near and far terms, and the boundary between the near and far terms was determined to be 9 Å. The sampling interval during the simulation was set to 2 ps. The MD simulations were performed using the NPT ensemble for 100 ns using Desmond60. The stability of surface binding was further validated using MMPBSA calculations, employing 250 evenly spaced structures derived from the stable portion of a 100 ns simulation performed in Desmond with the optimized potentials for liquid simulations (OPLS) force field76,77. These structures were converted to PDB format78 and analyzed using the “MMPBSA.py” script79. The binding free energy was assessed by considering van der Waals, electrostatic, desolvation, and internal strain energies, along with free energy decomposition to determine key residue contributions. The calculations utilized default parameters recommended by AMBER, as outlined in prior publications59,60,61.
Quantum mechanics/molecular mechanics analysis
For quantum mechanics/molecular mechanics (QM/MM) minimizations, an 8 Å layer of water molecules surrounding the proteins and the active site region was optimized in oxidized form for iron (Fe3+) with QSite (Schrodinger 2022 suite)80 using unrestricted density functional theory (DFT) calculations. Furthermore, using QM/MM-based spin-density calculations in MMOH, we located the transfer region and identified the first electron-acceptor residue. This residue serves as the initial recipient of an electron from the dye located on the protein’s surface81,82. In our QM/MM calculations, the transfer region was included in the QM region, and the remaining protein was treated as the MM region, where electron donors/acceptors were frozen (previously optimized QM region). An extra electron was injected into the QM region and the doublet state. Single-point calculations using the QM/MM method were performed with QSITE using unrestricted DFT combined with the def2-TZVP basis set (def2-TZVP for heavy atoms and 6-311++G** for lighter atoms)83,84,85 and residues, and regions containing extra spin density were identified. This method was used to identify the residue with the highest spin density for selecting the first electron acceptor from the dye.
Electron transfer rate calculations
Based on the spin-density-based method, residues were obtained from QM/MM calculations, which provide a high density when a free electron is introduced into the system. Calculations using the QM/MM approach yielded residues that were produced using a spin-density-based method. These calculations yield a high spin density when free electrons are injected into the system. The next step in the computation involved transferring the residue with the highest spin density to the MM column and recalculating the spin density using the parameters as described above86. Here, we computed the driving force (∆G°), and reorganization energy (λ), and obtained activation energy (∆G‡), which is derived from the driving force and reorganization energy, using the complete Marcus equation as follows (Eq. 5):
where HDA denotes the electronic coupling between the donor and acceptor. Assuming that the electronic coupling remains constant and that the solvent is not involved in the reaction, the Moser–Dutton ruler can be used to predict the degree of confidence, and the donor–acceptor distance (R) can be calculated as follows87,88 (Eq. 6):
The Marcus curve is a useful tool for determining the relative efficiencies of different competitive electron-transfer routes. The most favorable first acceptor residue in terms of energy efficiency was determined by plotting the rate of electron transfer as a function of the free energy difference. In addition, the curve can be used to determine the activation energy required for electron transfer, which may provide additional information on the reaction mechanism89,90. In the simulations, the sample frames were obtained from structures with the lowest amount of energy. The top 50 lowest-energy structures were selected for the computation. Marcus curves for the electron exchange between EY and K323 were plotted by fitting polynomial curves to the reactant state, where the electron-containing the donor, dye, and acceptor are all in a neutral state, and the product state, where the acceptor has gained an electron. Both states were fitted using polynomial curves to generate a diabatic free-energy curve.
Photobiocatalytic reactions and analyses
For a photobiocatalytic conversion of methane to methanol, we prepared a TEOA buffer (100 mM, pH 7.5, 1.5 mL) containing MMOH (2.5 µM), MMOR (5 µM), and MMOB (5 µM) with XD (20 µM). The electrolyte was bubbled with CH4 and O2 gases for 20 min, and then added to a septa-sealed borosilicate glass bottle using a magnetic stirrer. The system was stirred at 500 rpm under visible light (I = 0.58 µE cm−2 s−1, λ > 400 nm) with a supply of CH4 (2 mL) and O2 (2 mL) every 1 h. We quantified methanol using a 7890 A gas chromatograph (Agilent Technologies) equipped with a DB-1 column (length: 60 m, diameter: 0.320 µm) and flame ionization detector. The oven temperature programs are listed in Supplementary Table 7. A predetermined calibration graph was used for methanol quantification with ethanol as an internal standard. The methanol reaction time and quantum yields were determined using Eqs. (7) and (8).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data underlying this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Dummer, N. F. et al. Methane oxidation to methanol. Chem. Rev. 123, 6359–6411 (2023).
Song, H. & Ye, J. Direct photocatalytic conversion of methane to value-added chemicals. Trends Chem. 4, 1094–1105 (2022).
Marzeion, B., Kaser, G., Maussion, F. & Champollion, N. Limited influence of climate change mitigation on short-term glacier mass loss. Nat. Clim. Change 8, 305–308 (2018).
Alves, L., Pereira, V., Lagarteira, T. & Mendes, A. Catalytic methane decomposition to boost the energy transition: scientific and technological advancements. Renew. Sustain. Energy Rev. 137, 110465 (2021).
Olivos-Suarez, A. I. et al. Strategies for the direct catalytic valorization of methane using heterogeneous catalysis: challenges and opportunities. ACS Catal. 6, 2965–2981 (2016).
Aasberg-Petersen, K. et al. Natural gas to synthesis gas—catalysts and catalytic processes. J. Nat. Gas Sci. Eng. 3, 423–459 (2011).
Thyssen, V. V., Vilela, V. B., de Florio, D. Z., Ferlauto, A. S. & Fonseca, F. C. Direct conversion of methane to C2 hydrocarbons in solid-state membrane reactors at high temperatures. Chem. Rev. 122, 3966–3995 (2022).
Song, H., Meng, X., Wang, Z. -j, Liu, H. & Ye, J. Solar-energy-mediated methane conversion. Joule 3, 1606–1636 (2019).
Bai, S. et al. High-efficiency direct methane conversion to oxygenates on a cerium dioxide nanowires supported rhodium single-atom catalyst. Nat. Commun. 11, 954 (2020).
Ravi, M., Ranocchiari, M. & van Bokhoven, J. A. The direct catalytic oxidation of methane to methanol—a critical assessment. Angew. Chem. Int. Ed. 56, 16464–16483 (2017).
Sushkevich, V. L., Palagin, D., Ranocchiari, M. & van Bokhoven, J. A. Selective anaerobic oxidation of methane enables direct synthesis of methanol. Science 356, 523–527 (2017).
Villa, K., Murcia-López, S., Morante, J. R. & Andreu, T. An insight on the role of La in mesoporous WO3 for the photocatalytic conversion of methane into methanol. Appl. Catal. B 187, 30–36 (2016).
Cui, X. et al. Room-temperature methane conversion by graphene-confined single iron atoms. Chem 4, 1902–1910 (2018).
Fan, Y. et al. Selective photocatalytic oxidation of methane by quantum-sized bismuth vanadate. Nat. Sustain. 4, 509–515 (2021).
Xie, J. et al. Highly selective oxidation of methane to methanol at ambient conditions by titanium dioxide-supported iron species. Nat. Catal. 1, 889–896 (2018).
Shu, L. et al. An Fe2IVO2 diamond core structure for the key intermediate Q of methane monooxygenase. Science 275, 515–518 (1997).
Banerjee, R., Proshlyakov, Y., Lipscomb, J. D. & Proshlyakov, D. A. Structure of the key species in the enzymatic oxidation of methane to methanol. Nature 518, 431–434 (2015).
Schulz, C. E., Castillo, R. G., Pantazis, D. A., DeBeer, S. & Neese, F. Structure–spectroscopy correlations for intermediate Q of soluble methane monooxygenase: insights from QM/MM calculations. J. Am. Chem. Soc. 143, 6560–6577 (2021).
Kim, H. J. et al. Biological conversion of methane to methanol through genetic reassembly of native catalytic domains. Nat. Catal. 2, 342–353 (2019).
Hwang, I. Y., Kalyuzhnaya, M. G. & Lee, E. Y. Quantitative assessment of methane bioconversion based on kinetics and bioenergetics. Bioresour. Technol. 410, 131269 (2024).
Fox, B. G., Froland, W. A., Dege, J. E. & Lipscomb, J. D. Methane monooxygenase from Methylosinus trichosporium OB3b: purification and properties of a three-component system with high specific activity from a type II methanotroph. J. Biol. Chem. 264, 10023–10033 (1989).
Kal, S. & Que, L. Jr Activation of a non-heme FeIII-OOH by a second FeIII to hydroxylate strong C−H bonds: possible implications for soluble methane monooxygenase. Angew. Chem. Int. Ed. 58, 8484–8488 (2019).
Rosenzweig, A. C., Frederick, C. A., Lippard, S. J. & Nordlund, P. Crystal structure of a bacterial non-haem iron hydroxylase that catalyses the biological oxidation of methane. Nature 366, 537–543 (1993).
Kim, H. et al. MMOD-induced structural changes of hydroxylase in soluble methane monooxygenase. Sci. Adv. 5, eaax0059 (2019).
Merkx, M. et al. Dioxygen activation and methane hydroxylation by soluble methane monooxygenase: a tale of two irons and three proteins. Angew. Chem. Int. Ed. 40, 2782–2807 (2001).
Kim, J. & Park, C. B. Collaborative catalysis for solar biosynthesis. Trends Chem. 5, 133–146 (2023).
Lee, S. H., Choi, D. S., Kuk, S. K. & Park, C. B. Photobiocatalysis: activating redox enzymes by direct or indirect transfer of photoinduced electrons. Angew. Chem. Int. Ed. 57, 7958–7985 (2018).
Park, J. H. et al. Cofactor-free light-driven whole-cell cytochrome P450 catalysis. Angew. Chem. Int. Ed. 54, 969–973 (2015).
Jang, J., Aurangzeb Khan, W., Hollmann, F., Won, K. & Park, C. B. Biosolar oxygenation of hydrocarbons by photocatalytic artificial wood. ACS Sustain. Chem. Eng. 12, 8950–8957 (2024).
Kim, C. H., Kim, J., Hollmann, F. & Park, C. B. Photoelectrocatalytic N2 fixation and C-H oxyfunctionalization driven by H2O oxidation. Appl. Catal. B Environ. 336, 122925 (2023).
Lee, C. H., Kim, J. & Park, C. B. Z-schematic artificial leaf structure for biosolar oxyfunctionalization of hydrocarbons. ACS Energy Lett. 8, 2513–2521 (2023).
Yun, C.-H., Kim, J., Hollmann, F. & Park, C. B. Light-driven biocatalytic oxidation. Chem. Sci. 13, 12260–12279 (2022).
Kim, J., Jang, J., Hilberath, T., Hollmann, F. & Park, C. B. Photoelectrocatalytic biosynthesis fuelled by microplastics. Nat. Synth. 1, 776–786 (2022).
Shin, J., Andreas Hutomo, C., Kim, J., Jang, J. & Park, C. B. Natural pollen exine-templated synthesis of photocatalytic metal oxides with high surface area and oxygen vacancies. Appl. Surf. Sci. 599, 154064 (2022).
Kim, J. et al. Unbiased photoelectrode interfaces for solar coupling of lignin oxidation with biocatalytic C═C bond hydrogenation. ACS Appl. Mater. Interfaces 14, 11465–11473 (2022).
Wang, D. et al. Lignin-fueled photoelectrochemical platform for light-driven redox biotransformation. Green. Chem. 22, 5151–5160 (2020).
Choi, D. S., Kim, J., Hollmann, F. & Park, C. B. Solar-assisted eBiorefinery: photoelectrochemical pairing of oxyfunctionalization and hydrogenation reactions. Angew. Chem. Int. Ed. 59, 15886–15890 (2020).
Choi, D. S. et al. Bias-free in situ H2O2 generation in a photovoltaic-photoelectrochemical tandem cell for biocatalytic oxyfunctionalization. ACS Catal. 9, 10562–10566 (2019).
Kim, J. et al. Nicotinamide adenine dinucleotide as a photocatalyst. Sci. Adv. 5, eaax0501 (2019).
Willot, S. J. P. et al. Expanding the spectrum of light-driven peroxygenase reactions. ACS Catal. 9, 890–894 (2019).
Lee, Y. W. et al. Unbiased biocatalytic solar-to-chemical conversion by FeOOH/BiVO4/perovskite tandem structure. Nat. Commun. 9, 4208 (2018).
Kim, J. et al. Biocatalytic C=C bond reduction through carbon nanodot-sensitized regeneration of NADH analogues. Angew. Chem. Int. Ed. 57, 13825–13828 (2018).
Son, E. J. et al. Carbon nanotube–graphitic carbon nitride hybrid films for flavoenzyme-catalyzed photoelectrochemical cells. Adv. Funct. Mater. 28, 1705232 (2018).
Lee, S. H. et al. Cofactor-free, direct photoactivation of enoate reductases for the asymmetric reduction of C=C bonds. Angew. Chem. Int. Ed. 56, 8681–8685 (2017).
Wang, D., Kim, J. & Park, C. B. Lignin-induced CaCO3 vaterite structure for biocatalytic artificial photosynthesis. ACS Appl. Mater. Interfaces 13, 58522–58531 (2021).
Greeneltch, N. G. et al. Near-infrared surface-enhanced Raman spectroscopy (NIR-SERS) for the identification of eosin Y: theoretical calculations and evaluation of two different nanoplasmonic substrates. J. Phys. Chem. A 116, 11863–11869 (2012).
Huang, X.-Y., Bin, J.-P., Bu, H.-T., Jiang, G.-B. & Zeng, M.-H. Removal of anionic dye eosin Y from aqueous solution using ethylenediamine modified chitosan. Carbohydr. Polym. 84, 1350–1356 (2011).
Fox, B. G. et al. Moessbauer, EPR, and ENDOR studies of the hydroxylase and reductase components of methane monooxygenase from Methylosinus trichosporium OB3b. J. Am. Chem. Soc. 115, 3688–3701 (1993).
Kim, J., Nguyen, T. V. T., Kim, Y. H., Hollmann, F. & Park, C. B. Lignin as a multifunctional photocatalyst for solar-powered biocatalytic oxyfunctionalization of C–H bonds. Nat. Synth. 1, 217–226 (2022).
Jang, J. et al. Self-assembled protective layer by symmetric ionic liquid for long-cycling lithium–metal batteries. Adv. Energy Mater. 12, 2103955 (2022).
Kuk, S. K. et al. CO2-reductive, copper oxide-based photobiocathode for Z-scheme semi-artificial leaf structure. ChemSusChem 13, 2940–2944 (2020).
Jang, J. et al. Miniature Li+ solvation by symmetric molecular design for practical and safe Li-metal batteries. Nat. Energy 10, 502–512 (2025).
Zheng, K. et al. Room-temperature photooxidation of CH4 to CH3OH with nearly 100% selectivity over hetero-ZnO/Fe2O3 porous nanosheets. J. Am. Chem. Soc. 144, 12357–12366 (2022).
Kim, J. & Park, C. B. Shedding light on biocatalysis: photoelectrochemical platforms for solar-driven biotransformation. Curr. Opin. Biotechnol. 49, 122–129 (2019).
Paulsen, K. E. et al. Oxidation-reduction potentials of the methane monooxygenase hydroxylase component from Methylosinus trichosporium OB3b. Biochemistry 33, 713–722 (1994).
Jhonsi, M. A., Srinivasan, V. & Kathiravan, A. Light induced behavior of xanthene dyes with benzyl viologen. Synth. Met. 196, 131–138 (2014).
Egbaria, K. & Friedman, M. Adsorption of fluorescein dyes on albumin microspheres. Pharm. Res. 9, 629–635 (1992).
Sirin, S. et al. A computational approach to enzyme design: predicting ω-aminotransferase catalytic activity using docking and MM-GBSA scoring. J. Chem. Inf. Model. 54, 2334–2346 (2014).
Nicastro, R. et al. Malonyl-CoA is a conserved endogenous ATP-competitive mTORC1 inhibitor. Nat. Cell Biol. 25, 1303–1318 (2023).
Chen, W. et al. Structural insights into drug transport by an aquaglyceroporin. Nat. Commun. 15, 3985 (2024).
Ri, K. et al. Molecular mechanism of choline and ethanolamine transport in humans. Nature 630, 501–508 (2024).
Osadchii, D. Y. et al. Isolated Fe sites in metal organic frameworks catalyze the direct conversion of methane to methanol. ACS Catal. 8, 5542–5548 (2018).
Qi, G. et al. Au-ZSM-5 catalyses the selective oxidation of CH4 to CH3OH and CH3COOH using O2. Nat. Catal. 5, 45–54 (2022).
Tomkins, P. et al. Isothermal cyclic conversion of methane into methanol over copper-exchanged zeolite at low temperature. Angew. Chem. Int. Ed. 55, 5467–5471 (2016).
Ab Rahim, M. H. et al. Oxidation of methane to methanol with hydrogen peroxide using supported gold–palladium alloy nanoparticles. Angew. Chem. Int. Ed. 52, 1280–1284 (2013).
Feng, N. et al. Efficient and selective photocatalytic CH4 conversion to CH3OH with O2 by controlling overoxidation on TiO2. Nat. Commun. 12, 4652 (2021).
Zhu, W. et al. Facet-dependent enhancement in the activity of bismuth vanadate microcrystals for the photocatalytic conversion of methane to methanol. ACS Appl. Nano Mater. 1, 6683–6691 (2018).
Koo, C. W., Tucci, F. J., He, Y. & Rosenzweig, A. C. Recovery of particulate methane monooxygenase structure and activity in a lipid bilayer. Science 375, 1287–1291 (2022).
de la Lande, A., Babcock, N. S., Řezáč, J., Sanders, B. C. & Salahub, D. R. Surface residues dynamically organize water bridges to enhance electron transfer between proteins. Proc. Natl Acad. Sci. USA 107, 11799–11804 (2010).
Nilsen-Moe, A. et al. Proton-coupled electron transfer from tyrosine in the interior of a de novo protein: mechanisms and primary proton acceptor. J. Am. Chem. Soc. 142, 11550–11559 (2020).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F. & Hermans, J. Interaction Models for Water in Relation to Protein Hydration. In: Intermolecular Forces: Proceedings of the Fourteenth Jerusalem Symposium on Quantum Chemistry and Biochemistry, Jerusalem, (ed. Pullman, B.) 331–342 (D. Reidel Publishing Company, Dordrecht, 1981).
Shaw, D. E. et al. Anton 2: Raising the Bar for Performance and Programmability in a Special-Purpose Molecular Dynamics Supercomputer. In SC ’14: Proceedings of the International Conference for High Performance Computing 41–53 (Networking, Storage and Analysis. IEEE Press, 2014).
Li, J. et al. The VSGB 2.0 model: a next generation energy model for high resolution protein structure modeling. Proteins 79, 2794–2812 (2011).
Madhavi Sastry, G., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Shivakumar, D. et al. Prediction of absolute solvation free energies using molecular dynamics free energy perturbation and the OPLS force field. J. Chem. Theory Comput. 6, 1509–1519 (2010).
Wu, X. et al. Computational design of α-conotoxins to target specific nicotinic acetylcholine receptor subtypes. Chem. Eur. J. 30, e202302909 (2024).
Xue, J., Huang, X. & Zhu, Y. Using molecular dynamics simulations to evaluate active designs of cephradine hydrolase by molecular mechanics/Poisson–Boltzmann surface area and molecular mechanics/generalized Born surface area methods. RSC Adv. 9, 13868–13877 (2019).
Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236 (1996).
Murphy, R. B., Philipp, D. M. & Friesner, R. A. A mixed quantum mechanics/molecular mechanics (QM/MM) method for large-scale modeling of chemistry in protein environments. J. Comput. Chem. 21, 1442–1457 (2000).
Philipp, D. M. & Friesner, R. A. Mixed ab initio QM/MM modeling using frozen orbitals and tests with alanine dipeptide and tetrapeptide. J. Comput. Chem. 20, 1468–1494 (1999).
Melo, M. C. R. et al. NAMD goes quantum: an integrative suite for hybrid simulations. Nat. Methods 15, 351–354 (2018).
Zhang, A. et al. Catalytic cycle of formate dehydrogenase captured by single-molecule conductance. Nat. Catal. 6, 266–275 (2023).
Csizi, K.-S., Steiner, M. & Reiher, M. Nanoscale chemical reaction exploration with a quantum magnifying glass. Nat. Commun. 15, 5320 (2024).
Lucas, M. F., Rousseau, D. L. & Guallar, V. Electron transfer pathways in cytochrome c oxidase. Biochim. Biophys. Acta Bioenerg. 1807, 1305–1313 (2011).
Guallar, V. & Wallrapp, F. H. QM/MM methods: looking inside heme proteins biochemistry. Biophys. Chem. 149, 1–11 (2010).
Blumberger, J. Free energies for biological electron transfer from QM/MM calculation: method, application and critical assessment. Phys. Chem. Chem. Phys. 10, 5651–5667 (2008).
Blumberger, J. Recent advances in the theory and molecular simulation of biological electron transfer reactions. Chem. Rev. 115, 11191–11238 (2015).
Darcy, J. W., Kolmar, S. S. & Mayer, J. M. Transition state asymmetry in C–H bond cleavage by proton-coupled electron transfer. J. Am. Chem. Soc. 141, 10777–10787 (2019).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grants funded by the Ministry of Science and ICT (Grant No. RS-2023-00222078, RS-2024-00440681, RS-2024-00460425, RS-2024-00351665). C.B.P. acknowledges support from the NRF of Korea (Grant No. RS-2023-00222078, RS-2024-00440681, RS-2024-00460425). J.-K.L. acknowledges support from the NRF of Korea (Grant No. RS-2023-00222078, RS-2024-00440681, RS-2024-00351665).
Author information
Authors and Affiliations
Contributions
C.B.P. and J.-K.L. conceptualized the project. J.J. designed and conducted experiments. R.L. prepared MMOH, MMOR, and MMOB samples and performed ITC analysis. D.D. carried out computational simulation works. J.J., R.L., and D.D. prepared the original draft. C.B.P. and J.-K.L. reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sandy Schmidt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jang, J., Das, D., Lestari, R. et al. Solar-hybrid biocatalyst for methane hydroxylation to methanol. Nat Commun 16, 9310 (2025). https://doi.org/10.1038/s41467-025-64362-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64362-6