Abstract
The biosynthesis of ribosomally synthesized and post-translationally modified peptides (RiPPs) involves binding of the N-terminal leader region of precursor peptides to peptide-modifying enzymes and subsequent modification of the C-terminal core. Canonical RiPP leaders are short and unstructured. However, atypical RiPP leaders called nitrile hydratase-like leader peptides (NHLPs) are longer and structured. The sequence and structural divergence dictate that the NHLPs engage with peptide modifying enzymes using binding modes that are distinct from that for the canonical short leader peptides. Here, we investigate the molecular basis of NHLP engagement with class II lanthionine synthetases (LanMs). Using genome mining, we identify a LanM that is selective for the NHLP while modifying diverse core peptides. The solution NMR structure of the NHLP reveals a rigid tertiary fold. Biochemically verified computational models support a bimodal mechanism for NHLP binding to LanMs. Findings developed here illuminate unique protein-protein interaction modalities that guide RiPP biosynthesis.
Similar content being viewed by others
Introduction
Covalent modification of proteins is one of nature’s methods of expanding and altering their structure, function, and reactivity. Numerous biochemical processes are devoted to introducing a myriad of post-translational modifications (PTMs) in proteins, and in responding to and modulating downstream effects of PTMs1. Protein modifications are often reversible, and a separate set of enzymes remove PTMs to turn off the altered functions of modified proteins2. Unlike PTMs affected upon proteins, modification of small peptides is often irreversible. The biosynthesis of most ribosomally synthesized and post-translationally modified peptides (RiPPs) involves molecular recognition and binding of an N-terminal region of the substrate peptide—referred to as the “leader” region—to the peptide modifying enzymes (Fig. 1a)3. This intermolecular peptide-protein interaction usually directs the modification of the C-terminal “core” region of the RiPP precursor peptide by the peptide modifying enzymes. The N-terminal leader region is not modified and is proteolytically removed to furnish mature RiPPs that are often endowed with pharmacologically favorable bioactivities4,5. The RiPP precursor peptides and the peptide modifying enzymes are encoded in genetic neighborhoods called biosynthetic gene clusters (BGCs). The bioinformatic detection of RiPP BGCs has established the genomic ubiquity and chemical diversity of RiPPs6,7.
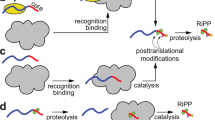
a Cartoon representation of RiPP biosynthesis in which the peptide modifying enzyme (green) binds to the leader region (blue) of the RiPP precursor peptide to install PTMs upon the core region (red). Proteolytic removal of the leader furnishes the mature RiPP. Lanthionine ring formation is shown as an example of a PTM. b Representative structures of short canonical leader peptides. Clockwise from top left: NMR structure of the cyanobactin PatE leader peptide (PDB: 2KYA) determined in 50% trifluoroethanol which promotes α-helix formation8, PatE leader peptide in complex with cyanobactin cyclodehydratase LynD dimer9, lasso peptide precursor TbiAα in complex with peptide binding protein TbiB111, and lanthipeptide precursor NisA in complex with dehydratase NisB dimer10. In complex with biosynthetic enzymes, the leader peptides (in yellow) can adopt isolated secondary structure elements, but no tertiary structures have been observed. c Cartoon schematic of the reaction catalyzed by LanM enzymes. The LanM dehydratase domain phosphorylates the Ser/Thr side chain hydroxyl followed by phosphate elimination to yield Dha/Dhb residues. Stereoselective Michael-type addition of the Cys side chain thiol at the Dha/Dhb-Cβ by the LanM cyclase domain leads to (methyl)lanthionine ring formation.
For most RiPP biosynthetic systems, the leader peptides are short (typically < 50 amino acids)3,4. While such canonical leader peptides can adopt isolated secondary structure elements upon binding to RiPP biosynthetic enzymes, the leader peptides are by themselves typically devoid of secondary and tertiary structure under physiological conditions (Fig. 1b)8,9,10,11. Haft’s bioinformatic description of nitrile hydratase-like leader peptides (NHLPs)—the mature products of which were subsequently named proteusins by Piel—upends this description12,13. The NHLPs are long, often exceeding 80 amino acids. Building upon these findings, we recently demonstrated that NHLPs adopt well-defined secondary and tertiary structures (vide infra)14. It was thus logical to hypothesize that the atypical length and structure of NHLPs would dictate their engagement with RiPP biosynthetic enzymes and would involve higher order protein-protein interactions spanning large molecular surfaces rather than peptide-protein interactions involving an isolated few amino acid residues of the leader peptide. However, contrary to this hypothesis, we previously found that the interactions of an NHLP with a peptide brominase involved only a short, transiently structured region at the C-terminus of the NHLP while the entirety of the structured NHLP nucleus was dispensable14. Whether the structured nucleus of the NHLP is likewise dispensable for binding to other RiPP biosynthetic enzymes is not apparent and requires evaluation of other NHLP/RiPP biosynthetic enzyme pairs.
Class II lanthionine synthetases (LanMs) are RiPP biosynthetic enzymes that install macrocyclizing thioether bonds in the core regions of lanthipeptide precursor peptides15,16. The LanM enzyme comprises of two domains: a kinase/dehydratase domain and a cyclase domain. The kinase/dehydratase domain (henceforth referred to as the dehydratase domain) phosphorylates Ser/Thr side chain hydroxyl(s) followed by phosphate elimination leading to dehydroalanine/dehydrobutyrine (Dha/Dhb) residue formation. This step is followed by stereoselective Michael-type addition of a Cys side chain thiol to the Dha/Dhb Cβ atom by the Zn2+-dependent cyclase domain leading to the macrocyclic thioether ring formation (Fig. 1c). The activity of LanM enzymes for canonical short and unstructured lanthipeptide precursor peptides, as well as for atypically long and structured NHLP-containing precursor peptides has been described16,17. While crystal structures of lanthipeptide modifying LanMs are available and the molecular basis for lanthipeptide precursor peptide binding to LanM enzymes has been investigated, no structural insights have been reported into NHLP binding to LanM enzymes18,19,20.
In this study, we investigated the molecular basis of NHLP engagement with LanM enzymes. Genome mining led to the discovery of a LanM enzyme that was found to be highly selective for the NHLP while modifying diverse core peptides. The solution NMR structure and relaxation measurements of the NHLP revealed a rigid tertiary fold with a flexible C-terminal tail. Computational models identified a bimodal mechanism for NHLP binding to RiPP biosynthetic enzymes, which was biochemically verified. This study brings to the fore the unique nature of NHLPs, and the corresponding changes adopted by NHLP-containing precursor peptide modifying enzymes that allow for intermolecular protein-protein interactions to guide catalysis.
Results and Discussion
Discovery of a NHLP-containing precursor peptide modifying LanM enzyme by genome mining
The activity of the LanM enzyme OspM, which modifies the precursor peptide OspA containing a NHLP, has been previously described by Piel17. We sought to discover additional LanM-NHLP pairs to study for two reasons: first, to identify new LanM substrates towards deciphering the types of natural products they generate and their substrate specificities, and second, to highlight conserved features across diverse NHLP-modifying LanMs. Using the OspM sequence as query, we generated a LanM sequence similarity network (SSN)21. The SSN demonstrated that OspM clustered separately from other class II lanthionine synthetases, such as GeoM, CinM, HalM2, CylM, and LchM that modify the canonical small lanthipeptide precursor peptides (Fig. 2a)22,23,24,25. The hyper promiscuous marine cyanobacterial LanMs—ProcM and SyncM—clustered separately from both OspM and GeoM-like enzymes26,27. We observed that OspM clustered with MppM, which we have recently described to modify a Nif11-like precursor peptide28. Akin to NHLPs, Nif11-like precursor peptides also possess unusually long leader sequences12,26.
a An SSN demonstrating OspM and MppM that respectively modify NHLP and Nif11-like precursor peptides with long leader sequences cluster separately from other LanMs that modify precursor peptides with short leader sequences. Note that the node corresponding to DprM (in red) clusters with OspM and MppM. b Organization of the dpr BGC. The ORFs encoding the DprE precursor peptides are colored yellow; those encoding RiPP biosynthetic enzymes and transporters are colored orange, and blue, respectively, and the dprM gene is colored red. The ORFs colored purple encode enzymes that are likely not involved in RiPP biosynthesis. c Alignment of the DprE1–E7 sequences. The −4, −7, and −12 residues relative to the leader/core boundary are marked. Note that the DprE NHLP sequences are highly conserved, but the core sequences are divergent. Residues predicted to be in involved in constructing the hydrophobic nucleus in the DprE NHLP structure (vide infra) are marked by blue triangles. d Methyllanthionine ring topologies generated by DprM. The Cys and Thr residues involved in methyllanthionine ring formation are highlighted in red. The Ser residues that are not modified by DprM are highlighted in blue. The ring size (number of atoms) is listed for each macrocycle and the N and the C termini for the core peptides are marked. Note that DprE2 and DprE5 have identical core sequences. The DprE6 core, which does not possess a Cys residue, is not shown since it is not modified by DprM. DprE8–E10 are non-natural (synthetic) precursor peptides designed to test the substrate scope of DprM. e Comparative efficiency of DprM for the formation of different ring sizes as determined by substrate conversion in in vitro assays using purified DprM and purified DprE peptides. Note that DprE8, DprE2, DprE1, and DprE10 precursor peptides yield methyllanthionine rings with 10, 16, 22, and 28 atoms, respectively. The mean and standard deviation from three independent experiments is plotted. Source data are provided as a Source Data file.
Mining BGCs that encode LanM enzymes that cluster together with OspM and MppM led to identification of a Desulfotomaculum sp.-derived proteusin RiPP BGC (dpr BGC, Fig. 2b). The Desulfotomaculum spp. are anaerobic soil bacteria previously reported to harbor RiPP BGCs29. In addition to the putative lanthionine synthetase DprM, the dpr BGC encodes seven NHLP-containing precursor peptides (DprE1–E7, Table S1). The DprE peptides possess highly conserved leader sequences (Fig. 2c). All DprE leader sequences terminated with a Gly-Gly motif with hydrophobic amino acids occupying the −4, −7, and −12 positions relative to the leader/core boundary; the hydrophobic side chains of residues at these positions have been implicated in directing the activity of the C39 peptidase domain that is embedded at the N-terminus of the transporter DprT130. Unlike the conserved leader sequences, the DprE core sequences were variable with multiple Ser and Thr residues but contained at most one Cys residue that could conceivably be involved in (methyl)lanthionine ring formation (Fig. 1a). The DprE2 and DprE5 core sequences were identical, whereas the DprE6 core did not contain any Cys residues. While the DprE7 core was markedly divergent in sequence, each DprE core terminated in Tyr/Phe rich regions interspersed with Gly/Ser residues (Fig. 2c).
Other open reading frames (ORFs) in the dpr BGC encoded transporters (DprT1–T4), radical SAM enzymes (DprS1–S2), a RiPP recognition element (RRE) protein (DprR)31, and other putative biosynthetic enzymes (DprA–C). The dpr BGC is derived from metagenomic DNA and the RiPP(s) encoded herein were unknown. Nevertheless, detection of a LanM encoded in the vicinity of NHLP-containing precursor peptides presented an opportunity to query NHLP/LanM interactions. Furthermore, prior studies have established that RiPP biosynthetic enzymes encoded within BGCs or in bacterial genomes possessing multiple precursor peptides are broadly substrate tolerant leading us to query the biocatalytic potential of DprM for construction of (methyl)lanthionine macrocycles of various sizes26,27,32.
Activity and leader dependence for DprM
A synthetic gene codon-optimized for expression in Escherichia coli encoding DprM was co-expressed with synthetic genes encoding the DprE1–7 precursor peptides in E. coli using plasmid-based techniques that are well established for production of RiPPs33. The in vivo modified and thusly purified precursor peptides DprE1–5 and DprE7 demonstrated loss of 18 Da upon co-expression with dprM; treatment with iodoacetamide demonstrated that this mass loss corresponded to (methyl)lanthionine ring formation following Ser/Thr dehydration (Fig. S1–S14). Co-expression of the DprE6 peptide with DprM did not result in a − 18 Da mass loss. The DprE6 core sequence does not contain a Cys residue (Fig. 2c) but does possess Ser and Thr residues which were not dehydrated by DprM consistent with the observation that DprE1, DprE3, and DprE7 peptides underwent a solitary (methyl)lanthionine ring formation but no additional Ser/Thr side chain dehydrations. Discerning the methyllanthionine ring topology for DprE2, DprE4, and DprE5 was straightforward as there is a solitary Thr residue (and no Ser residues) in each of these three core peptides. Mass spectrometry fragmentation established that for DprE1, DprE3, and DprE7 DprM also installed only methyllanthionine rings; no lanthionine rings were generated (Fig. S15–S17). Degradation and derivatization experiments established that all six methyllanthionine rings were installed with the DL-configuration (Fig. S18)34. Taken together, these data allow for determination of the structures of the macrocyclic RiPPs generated by DprM (Fig. 2d). Furthermore, these data demonstrate the validity of using OspM as a query sequence to mine for additional lanthionine synthetases, such as DprM, that modify NHLP-containing precursor peptides.
It was evident that DprM was tolerant of different core sequences and could generate ring sizes ranging from 16 to 22 atoms. DprM preferred Thr residues for dehydration and macrocyclization. To test the substrate scope of DprM, three non-natural core sequences were appended to the DprE2leader to yield the DprE8–E10 precursor peptides (Table S1). Each of these peptides was modified by DprM to yield ring sizes ranging from 10 to 28 atoms (Fig. 2d, Fig. S19–24). The efficiency of different ring size formation was evaluated in vitro using purified DprM enzyme and purified DprE8 (ring size: 10 atoms), DprE2 (16 atoms), DprE1 (22 atoms), and DprE10 (28 atoms) peptides (Fig. S25–S28). Under identical reaction conditions with ~14-fold molar excess of the precursor peptides relative to DprM, maximal substrate turnover was observed for the natural DprE1 and DprE2 precursor peptides. The comparative efficiencies for generation of smaller (DprE8) and larger (DprE10) rings was reduced. However, despite the modestly compromised in vitro conversion, the substrate tolerance of DprM and its ability to construct a highly strained 10-atom macrocycle is noteworthy and potentially useful as a peptide rigidification modality35.
Next, we evaluated the leader peptide specificity for DprM. No modification of the DprE2 core peptide was observed in the absence of the leader peptide (Fig. S29). As mentioned above, the activity of OspM has been described, which modifies the NHLP-containing precursor peptide OspA17. To probe whether other NHLPs could support DprM activity, we generated a gene encoding a chimeric precursor peptide in which the DprE2core was appended to the OspAleader (Table S1). Upon in vitro treatment of OspAleader-DprE2core chimeric peptide with DprM, we observed that no modifications were affected upon the DprE2core (Fig. S30). Similarly, chimeric precursor peptides in which other NHLPs from PoyA, SrpE, and MprE precursor peptides were appended to the DprE2core were not modified by DprM (Table S1, Fig. S31–S33)13,32,36. These data allow us to posit that DprM is exquisitely selective for recognizing and binding to the DprE NHLPs.
Structure and dynamics of the NHLP DrpE2leader
We previously reported the solution NMR structure of the MprE NHLP, which demonstrated rigid secondary structure elements organized into a tertiary fold14. The MprE precursor peptides lead to the formation of linear azol(in)e containing RiPPs32. However, with a solitary structure in hand, it was unclear if other NHLPs would adopt similarly well-defined secondary and tertiary structures, or not. To gain insights into conservation within the NHLP structural fold which informs their interaction with the RiPP biosynthetic enzymes, we characterized the structure, stability, and dynamic properties of the NHLP DprE2leader. Far-UV circular dichroism (CD) spectroscopy demonstrated that the DprE2leader possessed stable secondary structural elements with 35.6% α-character, 19.3% β-character, and 45.1 % random coil content as predicted by the BeStSel server (Fig. 3a)37. The CD-derived melting temperature of DprE2leader was 67.1 °C consistent with a thermally stable and rigidly structured protein (Fig. 3b)14.
a Far-UV CD spectra for the DprE2leader acquired across 41–90 °C in 5 °C increments. b Temperature-dependent denaturation curve of DprE2leader as monitored by CD spectroscopy (at 222 nm). c 2D 1H-15N HSQC spectra of 192 μM DprE2leader peptide acquired at 25 °C at a 1H field of 800 MHz. Resonance assignments of NMR peaks corresponding to backbone amides are noted. d TALOS-N calculated RCI-S2 (black scatter, 0 and 1 signify disorder and rigid structure, respectively) and SSI (red = α helix; blue=β sheet) derived from backbone chemical shifts. A transiently structured region at the C-terminus of DprEleader is highlighted in an orange box in the RCI-S2 plot. e Cartoon representation for ten lowest energy solution structures of DprE2leader. The α-helices are colored red, the β-strands are colored blue, and the loops are colored gray. f Structure of DprE2leader in cartoon representation with the secondary structure elements labeled. g Previously reported structure of the NHLP-containing MprE precursor peptide in cartoon representation with the secondary structure elements labeled. Source data are provided as a Source Data file.
Uniformly 15N and 13C isotopically labeled DprE2leader peptide demonstrated a well-dispersed 2D 1H-15N HSQC spectrum, supporting a protein of stable secondary and tertiary structure (Fig. 3c). A majority (92%) of the chemical shifts corresponding to backbone HN, NH, Cα, Cβ, and CO atoms could be assigned using sequential assignment techniques based on three-dimensional triple-resonance NMR experiments. The secondary structure index (SSI) and random coil index order parameter (RCI-S2) were determined using TALOS-N38. The SSI indicated the presence of α-helices at the N-terminus of DprE2leader (residues 6–42) followed by a short span β-sheet (residues 51–65), and then a region lacking well-defined secondary structure (residues 66–86) (Fig. 3d). The RCI-S2 values provided evidence that the DprE2leader N-terminal region was well-ordered followed by increasing disorder towards the C-terminus, although a small region within the C-terminus exhibited a degree of order as highlighted by an increase in RCI-S2 value (residues 76–80, highlighted in orange in Fig. 3d). Next, we combined the chemical shifts, dihedral angle restraints from TALOS-N, and amide to amide NOE-based distance constraints to generate a set of energy minimized solution structure models for the DprE2leader peptide (Table S2).
Progressing from the N-terminus, the NMR structure of the NHLP DprE2leader demonstrated a three α-helical bundle followed by a two-strand antiparallel β-sheet and a disordered C-terminal tail (Fig. 3e, f). These secondary structural elements buried a hydrophobic nucleus to which the three helices contributed most of the amino acid side chains (Trp4, Val12, Ile16, Phe26, Cys30, Pro34, and Val38) with the β2 strand contributing the solitary Val65 side chain (Fig. S34). These residues were highly conserved in all DprE sequences (Fig. 2c). Ionic interaction between the Glu8 and the Lys44 side chains, which lie at the N- and the C-termini respectively of the α1 and α3 helices, further rigidifies the helical bundle (Fig. S34). After the β2 strand, which terminates at the Pro67 residue, the remainder of the DprE2leader remains largely unstructured. Despite the low sequence similarity between the MprEleader and DprEleader sequences (31%), the structure of the DprE2leader is highly similar to the previously described MprEleader structure14. The α1–α3 and the β1 and β2 secondary structure elements are conserved, as is the loop region that extends from the β2 strand to the leader/core boundary (Fig. 3f, g, and Fig. S35)14. The MprEleader possesses an additional α4 helix; however, molecular dynamics simulations revealed that the MprE α4 helix was only transiently stable14. Further, the increase in RCI-S2 value of DprE2leader (residues 76–80) could also indicate the presence of a sparsely populated ordered state. The secondary structure elements and the tertiary structure demonstrate robust alignment between the DprE2leader and MprE unifying the structural description of the NHLPs. It is thus instructive to note that despite this overall structural conservation, the MprEleader could not substitute the DprEleader in supporting DprM activity (Fig. S33).
Next, we queried whether the presence of the core peptide influenced the NHLP structure, or not. Overlaid 1H-15N HSQC spectra of the full length DprE2 and DprE2leader demonstrated a good overall alignment with only modest chemical shift perturbations (Fig. S36). The full length DprE2 demonstrated similar CD spectra as DprE2leader with comparable melting temperatures (Fig. S37). Taken together, these observations allow us to posit that the core peptide has minimal effect on the NHLP structure. Furthermore, our previous description of the MprE solution structure contained the MprEcore sequence with excellent alignment still observed between the DprE2leader and MprEleader structures (vide supra)14.
With the overall structure of DprEleader thusly resolved, next, we queried the dynamic regions within the NHLP structure. Specifically, using solution NMR spectroscopy, we characterized psec to nsec timescale flexibility in the DprE2leader structure. We recorded 15N HSQC-based R1 and R2 relaxation rates in conjunction with 15N-{1H} heteronuclear NOE (HetNOE) values39. We found several regions of increased mobility within DprE2leader. The short, disordered loops connecting secondary structure elements as well as the C-terminal disordered region correlated with increased R1 values, and decreased R2 and 15N-{1H} HetNOE values indicative of motions on the psec to nsec timescale (Fig. 4). Taken together, the R1, R2, and HetNOE measurements of the DprE2leader are consistent with the calculated NMR structure and highlight the rigidity in the N-terminal region and flexibility within the C-terminal region. In addition, we observed a small increase in the 15N-{1H} HetNOE values for the DprE2leader residues 74–82 (shaded orange in Fig. 4c), consistent with increase in the RCI-S2 values for these residues (shaded orange in Fig. 3d). These observations are indicative of the possibility of a transiently populated ordered-state within the C-terminal tail of DprE2leader peptide.
Backbone dynamics of the DprE2leader using 15N probes revealed by measurement of a R1 (s−1), b R2 (s−1), and c 15N-{1H} HetNOE values recorded at 25 °C at a 1H field of 800 MHz. A DprE2leader secondary structure cartoon is provided for reference. Gray boxes highlight dynamic regions that are localized to loops connecting secondary structure elements as well as the C-terminal disordered region. A transiently structured region at the C-terminus of DprEleader is highlighted in an orange box in the hetNOE plot as in Fig. 3d. Error bars for the R1 and R2 values were determined using a statistical bootstrapping scheme in NMRFAM-Sparky. Errors bars for hetNOE values were determined from spectral noise in NMRFAM-Sparky. Source data are provided as a Source Data file.
Dissonance in NHLP/enzyme interactions
As alluded to before, there is scant understanding of the different modes of interactions between NHLPs and RiPP biosynthetic enzymes. Guided by the NMR solution structure of NHLP-containing MprE peptide, NMR chemical shift titrations, and biochemical assays, we previously described a model for interaction of the SrpE NHLP with the flavin-dependent halogenase SrpI14. The halogenase SrpI brominates a Trp side chain indole at the C-terminus of the SrpEcore36,40. Surprisingly, the highly structured N-terminal region of SrpE NHLP was completely dispensable for interactions with SrpI; instead, the L(X)4L motif that is positioned towards the C-terminus of the SrpEleader was the primary site for interaction with SrpI14. The L(X)4L motif is conserved in the MprEleader as well (Fig. 5a). Consequently, the MprEleader could substitute the SrpEleader in supporting SrpI activity41. Curiously, the L(X)4L motif is also conserved in the DprE2leader as well (Fig. 5a). Consequently, the chimeric peptide DprE2leader-SrpEcore was a viable substrate for SrpI (Table S1, Fig. S38). These data imply a fundamental disconnect in the mode of NHLP engagement by SrpI and by DprM wherein DprM was highly selective for the identity of the NHLP (MprEleader and SrpEleader did not support DprM activity), while SrpI demonstrated no such NHLP selectivity.
a Sequence alignment between NHLP-containing peptides MprE, SrpE, and DprE2. The secondary structure elements identified from the solution NMR structures of MprE and DprE2leader are denoted above, and below the sequence alignment, respectively. The L(X)4L motif which is conserved in all three sequences is identified, as are the sites of DprE2 truncation discussed in the text. The C-terminal Trp residue in the SrpEcore is brominated by SrpI. DprM generates the thioether linkage between the Thr and Cys side chains in the DprEcore. The resideus in boldface in the MprEcore are converted to azolines by YcaO cyclodehydratase MprC. b Comparison of the in vitro conversion of full length DprE2 to the macrocyclized product by DprM, and of the (Δ1–65)DprE2 and (Δ1–74)DprE2 peptides in which the NHLP has been truncated. Means and standard deviations from three independent experiments are plotted. Note that the similarly truncated SrpE peptides supported the in vitro activity of brominase SrpI14. c CD spectra of (Δ68–86)DprE2leader peptide acquired from 30 °C to 90 °C in 5 °C increments. d Temperature-dependent denaturation curves of (Δ68–86)DprE2leader and DprE2leader peptides as monitored by CD spectroscopy (at 222 nm). e 2D 1H-15N HSQC spectra of DprE2leader peptide (in red) superimposed with the 2D 1H-15N HSQC spectra of (Δ68–86)DprE2leader peptide (in blue). Spectra were acquired at 25 °C at a 1H field of 800 MHz. Resonance assignments of NMR peaks corresponding to backbone amides of the wild-type DprE2leader peptide are noted. The (Δ68–86)DprE2leader peptide could not be assigned due to limited stability. Source data are provided as a Source Data file.
The SrpEleader was truncated such that the entirety of the structured N-terminal region was removed but the L(X)4L motif was preserved. These truncated forms of the SrpEleader still supported SrpI activity14. In line with these observations, the DprE2leader in the DprE2leader-SrpEcore chimeric peptide was truncated, as illustrated in Fig. 5a. The truncated forms of the chimeric peptide—(Δ1–65)DprE2leader-SrpEcore and (Δ1–74)DprE2leader-SrpEcore— were also viable substrates for SrpI as evaluated using in vitro halogenation assays (Table S1, Fig. S39–S40). However, with similarly truncated leader sequences, the (Δ1–65)DprE2 and (Δ1–74)DprE2 peptides were not viable substrates for DprM (Fig. 5b, and Fig. S41–S43). These findings imply that the DprE2/DprM interactions are distinct from the SrpE/SrpI interactions.
Interactions of NHLPs with RiPP biosynthetic enzymes have been explored by scanning mutagenesis42. However, mutagenesis can impact the overall structure of the NHLP itself. To explore the effect of the truncation of the unstructured C-terminal tail on the N-terminal structured region of the NHLPs, various truncated forms of the DprE2leader were screened. Of these, the (Δ68–86)DprE2leader peptide—now bereft of the C-terminal unstructured tail—was expressed and purified (Fig. 5a, and Table S1). CD spectroscopy demonstrated that the truncated (Δ68–86)DprE2leader retained the requisite secondary structural elements (Fig. 5c). However, as compared to the full length DprE2leader, the truncated (Δ68–86)DprE2leader demonstrated a decrease in the melting temperature implying deletion of the C-terminal tail resulted in a disruption in the stability of the N-terminal structured region (Fig. 5d). This observation was corroborated by 2D 1H-15N HSQC spectra of the uniformly 15N labeled (Δ68–86)DprE2leader peptide that demonstrated significant chemical shift changes of NMR resonances corresponding to backbone amide groups as compared to the full length DprE2leader peptide (Fig. 5e).
Taken together, these observations illuminate a model wherein the structured N-terminal nucleus and the C-terminal unstructured tail work synergistically to stabilize the overall NHLP structure. Furthermore, NMR spectroscopy hints towards a transiently structured region with the C-terminal unstructured tail itself (Figs. 3d, and 4c). Overall, the NHLP structural complexity necessitates entirely different modalities for NHLP interactions with RiPP biosynthetic enzymes that are otherwise not explored by the much shorter leader peptides of other RiPP precursor peptides.
A model for two-state recognition of the DprE2 NHLP by DprM
Guided by the structure of the MprE NHLP, we have previously described a model for the intermolecular interaction between SrpEleader and the brominase SrpI14. Despite exhaustive efforts, the limited stability of DprM precluded biophysical characterization of the DprE2leader/DprM interaction. Thus, we turned to computational modeling of the DprE2leader/DprM complex supported by biochemical interrogation of the model. A DprE2leader/DprM complex was modeled using AlphaFold3 in 1:1 stoichiometry in the presence of ATP/Mg2+ and Zn2+ 43. Our previous report of the crystal structure of the cytolysin synthetase CylM and the availability of the crystal structure of lichenicidin synthetase LicM2 allowed us to validate the appropriateness of the AlphaFold3-generated model18,19. Like CylM and LicM2, DprM was modeled by AlphaFold3 to possess N-terminal dehydratase and C-terminal cyclase domains (Fig. S44). Included in the model were ATP/Mg2+ and a Zn2+ cation, which are required for activity of the dehydratase and cyclase domains, respectively. We observed that the Zn2+ ion was modeled to be bound by three Cys side chains in the DprM cyclase domain at a site analogous to the Zn2+ binding sites in CylM and LicM2 (Fig. S45). The ATP binding site in the dehydratase domain of DprM was similarly analogous to the dehydratase domains of CylM and LicM2 with Mg2+ coordinated to ATP (Fig. S46). Thus, the requisite two-domain architecture of LanMs and small molecule cofactors/substrates were likely modeled correctly by AlphaFold3.
The trihelical bundle and the buried hydrophobic nucleus of the DprE2leader NHLP was modeled by AlphaFold3 to be in excellent agreement with the experimentally determined solution NMR structure. This trihelical bundle remained intact upon interaction with DprM, as did the salt bridge interaction between the DprE2 Glu8 and Lys44 side chains. As compared to the CylM and LicM2 dehydratase domains, the dehydratase domain of DprM was modeled to possess an additional short helix positioned within a long loop that is not present in the CylM and LicM2 dehydratase domains (Fig. S46); this short DprM helix contacted the N-terminal trihelical bundle of the DprE2 leader peptide (Fig. 6a). The DprM helix inserted two Trp side chain indoles—Trp203 and Trp207—into the DprE2 trihelical hydrophobic nucleus (Fig. 6b). This interaction model, which is predicated upon hydrophobic interactions between DprE2leader and DprM, satisfies the previously described biochemical data showing that the truncation of the DprE2 to (Δ1–65)DprE2 and (Δ1–74)DprE2 peptides such that the N-terminal hydrophobic nucleus was removed led to loss in DprM activity (Fig. 5b). To further test the hydrophobic interaction locus predicted by AlphaFold3, the DprM Trp203 and Trp207 residues were mutated to Phe, Leu, and Ala and the activity of the mutant DprM enzymes was compared to the wild type DprM activity. Note that both these residues are distant from the dehydratase and the cyclase active sites (Fig. 6a). In support of the interaction model, mutation of Trp203 and Trp207 led to reduction in DprM activity (Fig. 6c, and Fig. S47–S54). At the other end of the DprE2leader, side chain carboxylates of two Glu residues, DprE2 Glu74 and DprE2 Glu77 were predicted to be coordinated by a constellation of DprM Lys and Arg side chains (Fig. 6d). Mutation of the DprE2 Glu74 and Glu77 residues to Ala, individually and together, also led to loss in DprM activity (Fig. 6e, and Fig. S55–S57).
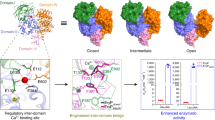
a Two views of the AlphaFold3-predicted DprE2leader/DprM model. DprM is shown as a transparent surface in brown; a long loop region absent in the CylM and LicM2 dehydratase domains is shown in cartoon representation. The DprE2leader is shown in blue. The DprM dehydratase and cyclase domains are modeled with the ATP/Mg2+ and Zn2+ bound, respectively. The bimodal binding model involves the interaction of the DprM Trp203 and Trp207 side chains with the trihelical bundle at the N-terminus of DprE2leader (see panel b) and by the interaction of DprM basic residues labeled ‘K’ and ‘R’ (see panel d) with the unstructured C-terminal region of the DprE2leader. b Two views for the engagement of the DprM Trp203 and Trp207 side chains into the hydrophobic nucleus of the DprE2leader. DprE2leader side chains that constitute the hydrophobic nucleus are shown in stick-ball representation, as are the DprE2leader Glu8 and Lys44 side chains. c In vitro activity of the wild type and mutant DprM enzymes. The mean and the standard deviation of substrate conversion in three independent reactions is plotted. d Two views for the binding of DprE2 Glu74 and Glu77 side chains with a constellation of DprM residues with basic side chains. This site of interaction involves the transiently structured region of DprE2leader highlighted in brown in Figs. 3d and 4c that is modeled by AlphaFold3 to adopt a short α-helix upon binding to DprM. e In vitro activity of the wild type DprM with wild type DrpE2, and with the DprE2 Glu74Ala, Glu77Ala, and the Glu74Ala/Glu77Ala mutant peptides. The mean and the standard deviation of substrate conversion in three independent reactions is plotted. f–j Comparison of loop regions in the crystal structures of f CylM and g LicM2, and the AlphaFold3-predicted models of h DprE2leader/DprM, i MppEleader/MppM, and j OspAleader/OspM. The loop regions are colored magenta. The residues at the loop peripheries are numbered. The hydrophobic amino acid side chains that are inserted by the LanM into the hydrophobic nuclei of the leader peptides are shown in stick-ball representation. Source data are provided as a Source data file.
Towards further exploration of how the abovementioned mutations could disrupt the DprE2/DprM interaction, we performed computational modeling within Rosetta’s REF2015 forcefield together with the DprE2leader/DprM AlphaFold3 model to calculate the ∆∆G of mutations44. The DprE2leader/DprM(Trp203Ala/Trp207Ala) mutant complex showed an unfavorable ∆∆G of +25.24 kcal/mol relative to the DprE2leader/DprM(wild-type). Analysis of the contribution of each of Rosetta’s energy terms to the total energy difference revealed major changes to Lennard-Jones attractive potential between residues (fa_atr), highlighting how the double Trp to Ala mutations disrupted local van der Waals forces at the DprE2/DprM interface (Fig. S58). Likewise, the DprE2leader(Glu74Ala/Glu77Ala)/DprM mutant complex demonstrated an unfavorable ∆∆G of +2.51 kcal/mol relative to the wild-type. Analysis of the contribution of each of Rosetta’s energy terms to the total energy difference revealed major changes to Lennard-Jones attractive potential between residues (fa_atr) and the Coulombic electrostatic potential (fa_elec), highlighting how the double Glu to Ala mutation altered both van der Waals and electrostatic forces at the DprM/DprE2 interface (Fig. S58).
We next performed in silico site saturation mutagenesis by scanning over DprM residues 201–209 or DprE2 residues that were modeled to be proximal to DprM using Rosetta to query all possible point mutations (except for cysteine). The predicted change in energy was determined by comparing the total Rosetta score of the mutant versus wild-type DprE2leader/DprM complex. From the view of DprM, we found that mutations in DprM residues 201–209 were associated with deleterious changes in binding energies, suggesting the DprE2leader/DprM interaction is highly specialized and optimized (Fig. S59). Results from the experimental and in silico DprM mutations at the Trp203 and Trp207 positions were highly consistent. From the view of DprE2leader, we found several sites that were not associated with deleterious changes in binding energies (these mainly face away from DprM) but many that were not (Fig. S60). Results from the experimental and in silico DprE2 mutations at the Glu74 and Glu77 sites were also highly consistent.
The applicability of the interaction model described above in which the structured N-terminal region of the NHLP makes hydrophobic interaction with the LanM is not restricted to DprE2/DprM only but likely extends to other LanMs as well. As mentioned previously, we have recently described the activity of the cyanobacterial MppM that constructs a lanthionine ring in the MppE core peptide28. The MppE leader peptide belongs to the Nif11-like family, and akin to NHLPs, is atypically long and structured12,26. The AlphaFold3 predicted model of MppE interaction with MppM demonstrated analogous insertion of four Leu side chains by MppM into the hydrophobic nucleus of MppE (Fig. S61). Simultaneous mutagenesis of these MppM Leu residues to Ala and in vitro comparison to the wild type MppM enzyme demonstrated loss in activity of the mutant MppM enzyme (Fig. S62–S63). The dehydratase domain of OspM also shows similar hydrophobic interactions with the OspA NHLP (Fig. S64)17.
Taken together, these data allow us to posit that the interactions of the NHLP DprE2leader with DprM are bimodal in nature such that the N-terminal structured region of the DprE2leader makes hydrophobic contact with DprM and the C-terminal unstructured region of the DprE2leader is involved in salt bridge interactions with DprM. The AlphaFold3 models also showed that the C-terminal region of the DprE2leader (residues 74–82) exhibited helical characteristics in complex with DprM, consistent with experimental observations of transiently sampled structural features within the apo DprE2leader (Figs. 3d, and 4c). The DprE2leader/DprM intermolecular interactions are different from the SrpEleader/SrpI interactions wherein the Leu side chains comprising the L(X)4L motif in SrpEleader were principally involved in hydrophobic interactions with SrpI akin to lanthipeptide leader peptide interactions with the C39 proteases30. As mentioned previously, elimination of the entire N-terminal structured region of SrpE had no effect on SrpI activity14. Thus, the structured nucleus of NHLPs is dispensable for binding to some RiPP biosynthetic enzymes but not others.
LanMs are class defining lanthipeptide synthesizing enzymes16. Our previous investigation into the mode of leader peptide binding to a non-NHLP lanthipeptide biosynthesizing HalM2 had revealed that the dehydratase domain was the principal site of interaction with the precursor leader peptide20. These findings were consistent with prior observations that the LanM dehydratase domains could be separated from the cyclase domains to yield functional leader-dependent dehydratases18,45,46. The models proposed in this study fall in line with these observations wherein AlphaFold3 models the dehydratase domains of DprM, MppM, and OspM to be involved in interactions with the corresponding NHLPs. The long loop region in DprM which inserts the Trp203 and Trp207 side chains into the hydrophobic nucleus of DprE2leader trihelical bundle is positioned at the interface of the structural elements termed as the N-lobe and the capping helices in our previous description on the crystal structure of CylM (Fig. 6f)18. Sixteen CylM residues, 162–177, which would correspond to the key DprM loop were not modeled into the CylM crystal structure which preclude definitive comparison with the corresponding 34 residue region of DprM. The crystal structure of LicM2 (PDB: 6ST5) demonstrates a small well-ordered seven residue loop in this region which is distinctly different in conformation from the model of the DprE2leader/DprM binding interface (Fig. 6g, h). These findings can be extended to the interaction models for MppEleader/MppM as well as OspAleader/OspM binding as well (Fig. 6i, j).
An enduring question is whether the AlphaFold3-predicted models are representative of molecular truth, or not? While the models presented above are all in line with biochemical data, a doubt still lingers47. Could there be an alternate way to test the validity of AlphaFold3 predictions? While the experimentally determined solution NMR structures of DprE and MprE leader peptides are remarkably similar, the MprEleader does not support the DprM activity. It is thus likely that the MprEleader does not engage with DprM in the same productive fashion that DprEleader peptides do. While computational predictions based on co-evolution can be limited in their utility to recover non-native states of conformationally flexible proteins48,49, the AlphaFold3-predicted structure of DprM in complex with the MprEleader demonstrates that the trihelical bundle of the MprEleader is disengaged with the Trp203 and Trp207 side chains (Fig. S65). The different conformations of the NHLP trihelical bundles relative to the DprM interaction loop observed here are likely representative of the leader peptide specificity for DprM. Taken together, it is tantalizing to posit that LanMs that interact with NHLPs and Nif11-like peptides have evolved a structural element to coordinate to these atypically long and structured leader peptides which could form the basis of their separate clustering in SSNs (Fig. 2a). Mining the LanM SSN together with AlphaFold3 modeling demonstrates that numerous other LanMs that cluster with DprM preserve the bimodal architecture for binding with NHLPs and Nif11-like leader peptides (Fig. S66–S70). It is likely that other enzymes have similarly evolved distinct intermolecular protein-protein interaction modalities with long protein-like leader peptides, understanding of which will enrich the intricate structural biology underpinning RiPP biosynthesis. The biotechnology potential of DprM could be perceived as high due to its ability to construct various methyllanthionine ring sizes but this assertion is juxtaposed against the compromised atom economy imposed by the requirement of a long leader peptide, as well as strict leader peptide specificity which could compromise combinatorial modifications by other RiPP biosynthetic enzymes.
Methods
General materials and methods
All chemicals, solvents, and media components were obtained commercially from Sigma-Aldrich, Fisher Scientific, and VWR, and used without further purification. Phusion high-fidelity DNA polymerase and Gibson assembly Master Mix were purchased from New England Biolabs. PrimeSTAR DNA polymerase Master Mix was purchased from Takaro Bio. Synthetic DNA fragments were ordered from Twist Biosciences. Isotopically labeled U-15NH4Cl and U-13C6-labeled glucose were purchased from Cambridge Isotope Laboratories. Mass spectra were recorded on Agilent 6530 C time of flight (ToF) mass spectrometer equipped with an electrospray ionization (ESI) source coupled to an Agilent 1260 liquid chromatography system equipped with a diode array detector. CD spectra were acquired using a Jasco J-815 spectropolarimeter equipped with a Neslab RTE-111 (Thermo Scientific) circulating water bath.
Cloning and mutagenesis
Standard molecular biology techniques were used to construct plasmids for recombinant protein production in E. coli. Target DNA fragments were amplified using either Phusion or PrimeSTAR high-fidelity DNA polymerases. Gibson assembly was used to subclone genes encoding proteins of interest into the target vectors. The plasmid sequences were confirmed by Sanger sequencing or by Nanopore sequencing. Genes encoding the wild type, chimeric, and mutant forms of the precursor peptides DprE and MppE were cloned in pET28(+) vector. For in vivo co-expression expression, the dprM and mppM genes were cloned into the multiple cloning site-2 (MCS-2) of the pCDF-Duet vector. For recombinant protein production, dprM, dprM mutants, mppM, mppM mutant(s), and srpI were also cloned into the pET28(+) vector. Gene encoding the DprE-truncated peptides were cloned into pET28-MBP vector. Synthetic genes optimized for expression in E. coli were used as templates for amplification and subcloning of amplicons in plasmid vectors. Typical PCR reactions (20 μL) contained 20 ng template DNA, 1 μM each of reverse and forward primers, along with DNA polymerases. PCR amplicons were subcloned into plasmid vectors using standard Gibson assembly protocols. All the mutations were installed using site directed mutagenesis protocols.
Generation of the LanM sequence similarity network (SSN)
An SSN for LanMs illustrated in Fig. 2a was created using the Enzyme Function Initiative- Enzyme Similarity Tool (EFI-EST)21. The sequence for OspM was used as the query sequence and the SSN was generated using an E-value of 5 and maximum blast sequence length of 1000 amino acids17. Using these parameters, 1000 LanM sequences were retrieved (969 unique) to generate the SSN using an alignment score cutoff of 100. The SSN was visualized in Cytoscape 3.8.2 using a 90% identity cut-off for grouping sequences into representative nodes50.
Co-expression of dprM with dprE1–E10 and peptide purification
For co-expression of dprE1–10 with dprM, genes encoding DprE1–10 were cloned in pET28 plasmids such that the DprE1–10 peptides were appended with an N-terminal His6 tag and co-transformed in E. coli BL21(DE3) host together with pCDF-Duet plasmid vector carrying dprM or empty pCDF-Duet vector as negative control. Colonies were grown under appropriate antibiotic kanamycin (50 μg/mL, final concentration) and streptomycin (50 μg/mL, final concentration) selection on Luria-Bertani (LB) agar media for 16 h. A single colony was picked and inoculated in 10 mL of terrific broth (TB) supplemented with appropriate antibiotics for 16 h at 37 °C. This inoculum was used to initiate 1 L TB cultures supplemented with antibiotics. Cultures were incubated with shaking at 37 °C until optical density measured at 600 nm (OD600) reached 0.6. Cultures were cooled at 18 °C for 1 h before protein expression was inducted by addition of 0.3 mM (final concentration) isopropyl-β-d-1-thiogalactopyranoside (IPTG). The cultures were further incubated at 18 °C, 180 rpm shaking for 12–16 h before harvesting the cells by centrifugation at 2000 × g for 30 min.
All steps for protein purification were performed at 4 °C or on ice. Cell pellets were resuspended in lysis buffer A (20 mM Tris-Cl (pH 7.5), 1 M NaCl) and cells were lysed by sonication or homogenization. The lysate was clarified by centrifugation at 36,000 × g for 45 min. The supernatant was loaded onto a 5 mL His-Trap Ni-NTA column. The column was washed extensively with wash buffer B (20 mM Tris-Cl (pH 7.5), 30 mM imidazole, 1 M NaCl,) until a stable UV-absorbance base line was observed, and bound proteins were eluted using a linear gradient from 0% to 100 % across 25 min of elution buffer (20 mM Tris-HCl (pH 7.5), 500 mM imidazole, 1 M NaCl) using ÄKTAprime plus FPLC system. Purity of eluent protein fractions were analyzed by SDS-PAGE, and fractions containing protein of interest were pooled and dialyzed overnight in the buffer containing 20 mM Tris-Cl (pH 7.5), 500 mM NaCl before storage in small aliquots at −80 °C.
Digestion of purified peptides with LahT150 peptidase and iodoacetamide (IAA) derivatization
The broad substrate tolerance of the peptidase LahT150 has been described previously30. LahT150-catalyzed leader cleavage reactions were performed in a total reaction volume of 200 μL containing 50 mM HEPES-Na (pH 7.5), 5 mM DTT, 26 μM LahT150 protease, 169 μM unmodified or DprM-modified peptides DprE1–10. All components were added and incubated at 30 °C for 6 h. 180 μL of this reaction mixture was withdrawn and 50 mM IAA was added for acetamide labeling; IAA labeling was allowed to proceed for 16 h in dark at 25 °C. An equal volume of MeOH + 1% (v/v) trifluoroacetic acid (TFA) was added to quench the reactions. The precipitates were removed by centrifugation at 16,000 × g for 30 min. The supernatants were withdrawn and analyzed by liquid chromatography/mass spectrometry (LC/MS).
Mass spectrometric detection and characterization of modified and unmodified DprE core peptides
An Agilent Poroshell 120 2.7 μm C18 reversed-phase column (100×4.6 mm) was coupled to an Agilent 6530 C ToF mass spectrometer equipped with an ESI source. This LC/MS setup was used for detection of modified product and the unmodified substrate peptides after digestion with LahT150. A flow rate of 0.3 mL/min was used with solvent A: water + 0.1% (v/v) formic acid and solvent B: MeCN + 0.1% (v/v) formic acid. Three different collision induced disassociation (CID) energies were used to acquire peptide fragmentation spectra; these were 30 eV, 50 eV, and 70 eV. All mass spectra were acquired in the positive ionization mode with MS1 range from m/z 50–3000 Da and MS2 range from m/z 50–3000. Unmodified substrate and modified product peptides were eluted from the stationary phase column using the following solvent gradient: 5% solvent B from 0 to 5 min, linear gradient from 5% solvent B to 100% solvent B from 5 to 18 min, 100% solvent B from 18 to 22 min, linear gradient from 100% solvent B to 5% solvent B from 22 to 24 min, and 5% solvent B from 24 to 30 min. Mass spectrometry data were acquired between 5 min and 24 min. Area under the extracted ion chromatograms (EICs) for [M + 2H]2+ ions was used to determine the relative amount of substrate and product peptides in the reaction mixtures. The stereochemistry of the methyllanthionine rings was established using degradation and derivatization experiments as described previously28,34.
Expression of dprE and mppE (and mutants thereof) and peptide purification
For general overexpression of N-His6-DprE1–10, N-His6-DprE2leader, mutants of N-His6-DprE2, N-His6-MppE, truncated peptides of N-His6-DprE2, chimeric peptides of N-His6-DprE2 in pET28 and LahT150 protease, 20 ng plasmid DNA was transformed in E. coli BL21(DE3). Colonies were grown under kanamycin (50 μg/mL, final concentration) selection on LB agar media for 16 h at 37 °C. A single colony was picked and inoculated in 10 mL of TB media supplemented with kanamycin for 16 h at 37 °C. This inoculum was used to initiate 1 L TB cultures. Cultures were incubated with shaking at 37 °C until OD600 reached 0.6. Cultures were cooled at 18 °C for 1 h before protein expression was inducted by addition of 0.3 mM IPTG (final concentration). The cultures were further incubated at 18 °C, 180 rpm for 12–16 h before harvesting the cells by centrifugation at 2000 × g for 30 min.
All steps for protein purification were performed at 4 °C or on ice. Cell pellets were resuspended in lysis buffer A (20 mM Tris-Cl (pH 7.5), 1 M NaCl) and cells were lysed by sonication or homogenization. The lysate was clarified by centrifugation at 36,000 × g for 45 min at 4 °C. The supernatant was loaded onto a 5 mL His-Trap Ni-NTA column at 4 °C. The column was washed extensively with wash buffer B (20 mM Tris-Cl (pH 7.5), 30 mM imidazole, 1 M NaCl) till a stable absorbance base line was observed, and protein was eluted with a linear gradient from 0% to 100 % in 25 min of elution buffer (20 mM Tris-HCl (pH 7.5), 500 mM imidazole, 1 M NaCl) using ÄKTAprime plus FPLC system. Purity of eluent protein fractions were analyzed by SDS-PAGE, and fractions containing protein of interest were pooled and dialyzed overnight in the buffer containing 20 mM Tris-Cl (pH 7.5), 500 mM NaCl before storage.
Expression of dprM, mppM and srpI (and mutants thereof) and protein purification
The pET28 plasmid vectors containing genes encoding DprM, MppM, and SrpI and chaperone protein encoding plasmids pGKJE8 or pGro7 (Takara Biosciences) were co-transformed in E. coli BL21(DE3). Colonies were grown under kanamycin (50 μg/mL, final concentration) and chloramphenicol (34 μg/mL, final concentration) selection on LB agar media for 16 h at 37 °C. A single colony was picked and inoculated in 10 mL of TB media for 16 h at 37 °C. This inoculum was used to initiate 1 L TB cultures. Cultures were incubated with shaking at 37 °C until OD600 reached 0.6. Cultures were cooled at 18 °C for 1 h before protein expression was inducted by addition of 0.5 mM (final concentration) IPTG and 250 mg L-arabinose (when pGro7 chaperone plasmid was used) and 500 mg of L-arabinose and 0.5 ng of tetracycline (when pGKJE8 chaperone plasmid was used). Cultures were further incubated at 18 °C, 180 rpm for 12–16 h before harvesting by centrifugation 2000 × g for 30 min at 4 °C.
All steps for protein purification were performed at 4 °C or on ice. Cell pellets were resuspended in lysis buffer A (20 mM Tris-Cl (pH 7.5), 1 M NaCl) and cells were lysed by sonication or homogenization. The lysate was clarified by centrifugation at 36,000 × g for 45 min at 4 °C. The supernatant was loaded onto a 5 mL His-Trap Ni-NTA column at 4 °C. The column was washed extensively with wash buffer B (20 mM Tris-Cl (pH 7.5), 30 mM imidazole, 1 M NaCl) at 2 mL/min flow rate till a stable UV-absorbance base line was observed. Proteins were eluted using a linear gradient from 0% to 100 % across 40 min of elution buffer (20 mM Tris-HCl (pH 7.5), 500 mM imidazole, 1 M NaCl) using ÄKTAprime plus FPLC system. The purity of eluent protein fractions was analyzed by SDS-PAGE, and fractions containing protein of interest were pooled and desalted in 50 mM HEPES-Na (pH 7.5), 500 mM NaCl, 10% (v/v) glycerol buffer using PD-10 columns before storage at −80 °C in small aliquots.
In vitro macrocyclization assays with purified DprM and MppM, and mutants thereof
To monitor the cyclization of different substrate peptides by DprM and MppM, all experiments were performed in triplicate. Assays were performed in a total volume of 200 μL comprising of 50 mM HEPES-Na (pH 7.5), 1 mM TCEP, 5 mM MgCl2, 5 mM ATP, 16.6 μM substrate DprE and MppE peptides, and 1.2 μM DprM and MppM. Reactions were incubated at 30 °C for 1 h. To this was added 26.3 μM LahT150 protease. For the DprE2 (E74A/E77A) mutant peptide was added 0.25 μg GluC, followed by incubation at 30 °C for another 1 h. To the reaction tube was then added equal volume of MeOH + 1% (v/v) TFA. Precipitated proteins were removed by centrifugation at 16,000 × g for 30 min at room temperature. The supernatants were withdrawn to HPLC vials. Two sets of negative control reactions were performed, omitting ATP, and omitting DprM/MppM.
Expression and purification of isotopically labeled DprE2leader and full length DprE2
Plasmid encoding NHis6-DprE2leader and NHis6-DprE2 were transformed into E. coli BL21(DE3). Using procedures described previously14, 1 L minimal media was inoculated with 5 mL overnight inoculum, supplemented with kanamycin, 3 g of 13C-labeled glucose, and 15N-labeled ammonium chloride. The cultures were incubated at 37 °C with shaking until the OD600 reached 0.9. Protein expression was induced by adding 0.8 mM IPTG after allowing the culture to cool down at 18 °C for 1 h. Induced cultures were further incubated at 18 °C, 180 rpm shaking for 18 h. The cultures were harvested by centrifugation at 5000 × g for 30 min at 4 °C and the cell pellet was resuspended in lysis buffer A (20 mM Tris (pH 7.5), 1 M NaCl). The resuspended cell pellet was lysed by sonication and lysate was removed by centrifugation at 35,000 × g for 45 min. The supernatant was loaded on to 5 mL His-Trap Ni-NTA column and the protein purification was performed using an ÄKTA go chromatography instrument. The column was then washed with wash buffer B (20 mM Tris (pH 7.5), 1 M NaCl, 30 mM imidazole) and, eluted using a linear gradient from 0% to 100% across 30 min with elution buffer B (20 mM Tris (pH 7.5), 1 M NaCl, 1 M imidazole). The protein was dialyzed overnight at 4 °C with IEX buffer A (20 mM Tris (pH 8.9), 100 mM NaCl). The dialyzed protein was then loaded on to the His-Trap Q column and eluted with a linear gradient of IEX buffer B (20 mM Tris (pH 7.5), 500 mM NaCl). Eluted protein fractions were further purified by size exclusion chromatography (SEC) using a Superdex 75 16/200 column with SEC buffer (20 mM Na-phosphate (pH 7.0), 100 mM NaCl). Using 3 kDa Amicon centrifugal filter at 4 °C, the pure fraction with the highest protein concentration was concentrated. Bradford assay was used to determine the protein concentration, and the concentrated protein was used to acquire NMR and CD spectroscopy data.
Circular dichroism spectroscopy
CD spectra were acquired using purified 50 μM DprE2leader peptide and 20 μM full length DprE2 peptide. Thermal denaturation experiments were performed by increasing the temperature from 41.28 to 90 °C for DprE2leader peptide and 10 to 90 °C for full length DprE2 peptide in 1 °C/min increments using a circulating water bath. CD spectra acquisition wavelength ranged from 200 to 300 nm. A 0.1 cm path length cuvette was used. Three replicate spectra, scanned at rate of 500 nm/min, were averaged. Each averaged spectrum was background-corrected and converted to molar circular dichroism [Δ\({{{\rm{\varepsilon }}}}\)]. CD data at 41.28 °C and 10 °C from 200–300 nm was deconvoluted into component spectra using the BeStSel server37. Melting temperature was determined using Boltzmann sigmoid model.
NMR assignments and structure determination
Unless stated otherwise, all solution nuclear magnetic resonance (NMR) experiments were conducted at 25 °C in 3 mm NMR tubes in a TCI cryoprobe equipped Bruker Avance III HD 800 MHz spectrometer at the Georgia Tech NMR Center. All data were collected on TopSpin v3.5pl5, processed in NMRPipe v11.5 and analyzed in NMRFAM-SPARKY v3.1951,52.
Chemical shift assignments
Uniformly double-labeled 15N/13C His6-DprE2leader peptide (residues 1–86) was prepared in NMR buffer (20 mM Na-phosphate (pH 6.5), 100 mM NaCl) at concentrations of 192 μM. Backbone resonances were assigned using sequential assignment strategies with standard triple-resonance experiments acquired with non-uniform sampling with 18 to 20% Poisson Gap sampling schedule and reconstructed with istHMS10: 3D HNCO (Bruker pulse sequence hncogp3d), 3D HNCA (Bruker pulse sequence hncagp3d), 3D HNCACB (Bruker pulse sequence hncacbgp3d), 3D CBCACONH (Bruker pulse sequence cbcaconhgp3d), and 3D SOFAST NOESY (pulse sequence sfnoesyNhmqc3d.HnHa-NHn)53. For 3D experiments, the acquisition times were set to 92 msec in 1H, 11 msec in 15N, 23 msec in 13CO and 10.5/5 msec in 13Cα,β. Recycle delay (d1) was set to 1 sec in all experiments except the NOESY where 0.45 sec was used. 3D amide-amide SOFAST NOESY experiments (pulse sequence sfnoesyNhmqc3d.HnHa-NHn) were collected with 112 scans, 0.45 sec recycle delay (d1), and 350 msec mixing time (d8) and NOE cross-peaks were assigned manually in NMRFAM-SPARKY53.
Structure calculations in CS-Rosetta
NMR structure calculations for DprE2leader were performed using automated Python scripts with the CS-Rosetta3 toolkit v3.354. Assigned chemical shifts of DprE2leader were fed into TALOS-N to determine secondary structure index (SSI), random coil index order parameter (RCI-S2), and dihedral angle restraints. DprE2leader chemical shifts were used in CS-Rosetta to pick fragments of amino acid lengths three and nine38. Protein sequence, 3-mer/9-mer fragments, backbone chemical shifts, and NOE restraints obtained from 3D amide-amide SOFAST NOESY experiments were used as input for the abrelax CS-Rosetta protocol. The 10 lowest energy models were selected from the 10,000 models calculated, that fit the experimental NOE data and dihedral angle restraints. Final structures were validated with MolProbity55.
Relaxation measurements of DprE2 leader peptide
Uniformly double-labeled 15N/13C His6-DprE2leader peptide (residues 1–86) was prepared in NMR buffer (20 mM Na-phosphate (pH 6.5), 100 mM NaCl, 10% D2O) at concentrations of 328 μM. Backbone 15N amide relaxation rates were determined with longitudinal (T1) (Bruker pulse sequence hsqct1etf3gpsi3d) and transverse (T2) (Bruker pulse sequence hsqct2etf3gpsi3d) relaxation data were acquired at 25 °C as a series of 2D1 H-15N HSQC spectra with the 15N-13C labeled DprE2leader peptide39. The relaxation delay times for T1 increments were selected within the range of 10 to 2400 msec (10, 20, 60, 100, 200, 400, 600, 800, 1200, 1600, 2000, 2400, 60, 60), while the values for T2 increments were selected between 16.96 to 237.44 msec (16.96, 33.92, 67.84, 101.76, 135.68, 169.6, 203.53, 237.44, 33.92, 33.92). Recycle delay (d1) was set to 3 sec for T1 and 1.5 sec for T2. The heteronuclear 15N{1H}-NOE (Bruker pulse sequence hsqcnoef3gpsi3d) was measured by acquiring two 1H-15N HSQC type spectra with and without proton saturation times of 3 sec. The heteronuclear NOE values were determined from the ratio of intensities of peaks in the two spectra (saturated/unsaturated). Recycle delay (d1) was set to 3 sec for the hetNOE experiment. The relaxation rates (R1 = 1/T1 and R2 = 1/T2) were calculated by fitting the signal decay rates to an exponential decay model in NMRFAM-SPARKY. Error bars for the R1 and R2 values were determined using a statistical bootstrapping scheme in NMRFAM-Sparky39. Errors bars for hetNOE values were determined from spectral noise in NMRFAM-Sparky39. Source data are provided as a Source Data file.
HSQC measurements of full length DprE2 peptide
Uniformly 15N-labeled full length His6-DprE2 peptide (residues 1–99) was prepared in NMR buffer (20 mM Na-phosphate (pH 6.5), 100 mM NaCl, 10% D2O) at concentrations of 782 μM. Backbone 1H-15N HSQC (Bruker pulse sequence hsqcetfpf3gpsi) data were acquired with 8 scans, 1 s recycle delay, and 30 ms in the indirect 15N dimension at 25 °C in 5 mm NMR tubes in a TCI cryoprobe equipped Bruker Avance III HD 700 MHz spectrometer. Data were collected on TopSpin v3.5pl5, processed in NMRPipe v11.5 and analyzed in NMRFAM-SPARKY v3.19. Chemical shift perturbations (CSPs, p.p.m.) were determined from weighted average of amide 1H and 15N chemical shift changes for full length DprE2 versus DprE2leader using the equation ΔδNH = [1/2 (ΔδH2 + ΔδN2/25)]½.
AlphaFold3 modeling
The ternary complexes between LanM enzymes (monomer) and the corresponding proteusin/Nif-11 like leader peptides (also monomers) were predicted in the presence of one each of Zn2+, Mg2+, and ATP using the online AlphaFold3 server (https://alphafoldserver.com). The following ipTM and pTM values were obtained:
Computational modeling in Rosetta
The AlphaFold3 model of the DprM/DprE2leader complex was used as an input for Rosetta. Towards in silico ∆∆G calculations for double mutant complexes (DprM W203A,W207A/DprE2 and DprM/DprE2 E74A,E77A), the REF2015 forcefield of Rosetta v2021.16 was used to minimize bond angles/geometry and side-chain rotamer conformations using Cartesian-space refinement with Idealize and Relax applications, respectively44,56,57,58. The Cartesian version of Rosetta’s ∆∆G protocol was used to generate, pack, and score mutations relative to the wild-type structure. The ∆∆G was then calculated as the difference between the energy of the mutated complex and relative to the wild-type complex. The code employed is provided in the Supplementary Information document.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
NMR resonance assignments for DprE2 leader peptide have been deposited in the Biological Magnetic Resonance Data Bank (https://bmrb.io/) under accession number 31187. Coordinates for the DprE2 structure have been deposited to the Protein Data Bank (PDB, https://www.rcsb.org/) with the accession code 9CJX. All data supporting the findings of this study are available from the corresponding authors upon request. Source data are provided with this paper.
Code availability
Rosetta code for computational ∆∆G of mutation and site saturation mutagenesis of the DprE2/DprM complex AlphaFold3 model are provided at Github: (https://github.com/mcshanlab/Vidya_et-al_DprM-DprE2_Rosetta). The AlphaFold3 model of the DprE2leader/DprM complex is also provided therein.
References
Lin, H. & Carroll, K.S. Introduction: Posttranslational protein modification. Chem. Rev. 118, 887–888 (2018).
Suskiewicz, M. J. The logic of protein post-translational modifications (PTMs): Chemistry, mechanisms and evolution of protein regulation through covalent attachments. BioEssays 46, 2300178 (2024).
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Montalbán-López, M. et al. New developments in RiPP discovery, enzymology and engineering. Nat. Prod. Rep. 38, 130–239 (2021).
Li, Y. & Rebuffat, S. The manifold roles of microbial ribosomal peptide–based natural products in physiology and ecology. J. Biol. Chem. 295, 34–54 (2020).
Kloosterman, A. M., Medema, M. H. & van Wezel, G. P. Omics-based strategies to discover novel classes of RiPP natural products. Curr. Opin. Biotechnol. 69, 60–67 (2021).
Russell, A. H. & Truman, A. W. Genome mining strategies for ribosomally synthesised and post-translationally modified peptides. Comput. Struct. Biotechnol. J. 18, 1838–1851 (2020).
Houssen, W. E. et al. Solution structure of the leader sequence of the patellamide precursor peptide, PatE1–34. ChemBioChem 11, 1867–1873 (2010).
Koehnke, J. et al. Structural analysis of leader peptide binding enables leader-free cyanobactin processing. Nat. Chem. Biol. 11, 558–563 (2015).
Ortega, M. A. et al. Structure and mechanism of the tRNA-dependent lantibiotic dehydratase NisB. Nature 517, 509–512 (2015).
Chekan Jonathan, R., Ongpipattanakul, C. & Nair Satish, K. Steric complementarity directs sequence promiscuous leader binding in RiPP biosynthesis. Proc. Natl. Acad. Sci. 116, 24049–24055 (2019).
Haft, D. H., Basu, M. K. & Mitchell, D. A. Expansion of ribosomally produced natural products: a nitrile hydratase- and Nif11-related precursor family. BMC Biol. 8, 70 (2010).
Freeman, M. F. et al. Metagenome mining reveals polytheonamides as posttranslationally modified ribosomal peptides. Science 338, 387–390 (2012).
Nguyen, N. A. et al. Disordered regions in proteusin peptides guide post-translational modification by a flavin-dependent RiPP brominase. Nat. Commun. 15, 1265 (2024).
Xie, L. et al. Lacticin 481: in vitro reconstitution of lantibiotic synthetase activity. Science 303, 679–681 (2004).
Repka, L. M., Chekan, J. R., Nair, S. K. & van der Donk, W. A. Mechanistic understanding of lanthipeptide biosynthetic enzymes. Chem. Rev. 117, 5457–5520 (2017).
Bösch, N. M. et al. Landornamides: antiviral ornithine-containing ribosomal peptides discovered through genome mining. Angew. Chem. Int. Ed. 59, 11763–11768 (2020).
Dong, S.-H. et al. The enterococcal cytolysin synthetase has an unanticipated lipid kinase fold. eLife 4, e07607 (2015).
Gonsior, M., Martins, B.M. Crystal structure of LicM2. https://www.rcsb.org/structure/6ST5 (2022).
Rahman, I. R. et al. Substrate recognition by the class II lanthipeptide synthetase HalM2. ACS Chem. Biol. 15, 1473–1486 (2020).
Gerlt, J. A. et al. Enzyme function initiative-enzyme similarity tool (EFI-EST): a web tool for generating protein sequence similarity networks. Biochim Biophys. Acta 1854, 1019–1037. (2015).
McClerren, A. L. et al. Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic. Proc. Natl. Acad. Sci. 103, 17243–17248 (2006).
Shenkarev, Z. O. et al. Isolation, structure elucidation, and synergistic antibacterial activity of a novel two-component lantibiotic lichenicidin from Bacillus licheniformis VK21. Biochemistry 49, 6462–6472 (2010).
Ökesli, A., Cooper, L. E., Fogle, E. J. & van der Donk, W. A. Nine post-translational modifications during the biosynthesis of cinnamycin. J. Am. Chem. Soc. 133, 13753–13760 (2011).
Garg, N., Tang, W., Goto, Y., Nair, S. K. & van der Donk, W. A. Lantibiotics from Geobacillus thermodenitrificans. Proc. Natl. Acad. Sci. USA 109, 5241–5246. (2012).
Li, B. et al. Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proc. Natl. Acad. Sci. 107, 10430–10435 (2010).
Arias-Orozco, P., Inklaar, M., Lanooij, J., Cebrián, R. & Kuipers, O. P. Functional expression and characterization of the highly promiscuous lanthipeptide synthetase SyncM, enabling the production of lanthipeptides with a broad range of ring topologies. ACS Synth. Biol. 10, 2579–2591 (2021).
Saha, N., Vidya, F. N. U., Luo, Y., van der Donk, W. A. & Agarwal, V. Transformation-Guided Genome Mining Provides Access to Brominated Lanthipeptides. Org. Lett. 27, 984–988 (2025).
Letzel, A.-C., Pidot, S. J. & Hertweck, C. Genome mining for ribosomally synthesized and post-translationally modified peptides (RiPPs) in anaerobic bacteria. BMC Genomics 15, 983 (2014).
Bobeica, S. C. et al. Insights into AMS/PCAT transporters from biochemical and structural characterization of a double Glycine motif protease. eLife 8, e42305 (2019).
Burkhart, B. J., Hudson, G. A., Dunbar, K. L. & Mitchell, D. A. A prevalent peptide-binding domain guides ribosomal natural product biosynthesis. Nat. Chem. Biol. 11, 564–570 (2015).
Nguyen, N. A. et al. A silent biosynthetic gene cluster from a methanotrophic bacterium potentiates discovery of a substrate promiscuous proteusin cyclodehydratase. ACS Chem. Biol. 17, 1577–1585 (2022).
Shi, Y., Yang, X., Garg, N. & van der Donk, W. A. Production of lantipeptides in Escherichia coli. J. Am. Chem. Soc. 133, 2338–2341 (2011).
Luo, Y., Xu, S., Frerk, A. M. & van der Donk, W. A. Facile method for determining lanthipeptide stereochemistry. Anal. Chem. 96, 1767–1773 (2024).
Viel, J. H. & Kuipers, O. P. Modular use of the uniquely small ring a of mersacidin generates the smallest ribosomally produced lanthipeptide. ACS Synth. Biol. 11, 3078–3087 (2022).
Nguyen, N. A. et al. An obligate peptidyl brominase underlies the discovery of highly distributed biosynthetic gene clusters in marine sponge microbiomes. J. Am. Chem. Soc. 143, 10221–10231 (2021).
Micsonai, A. et al. BeStSel: webserver for secondary structure and fold prediction for protein CD spectroscopy. Nucleic Acids Res. 50, W90–W98 (2022).
Shen, Y. & Bax, A., Protein structural information derived from NMR chemical shift with the neural network program TALOS-N. In Artificial Neural Networks, Cartwright, H., Ed. Springer New York (2015).
Kawale, A. A. & Burmann, B. M. Characterization of backbone dynamics using solution NMR spectroscopy to discern the functional plasticity of structurally analogous proteins. STAR Protoc. 2, 100919 (2021).
Saha, N., Vidya, F. N. U., Xie, R. & Agarwal, V. Halogenase-Assisted Alkyne/Aryl Bromide Sonogashira Coupling for Ribosomally Synthesized Peptides. J. Am. Chem. Soc. 146, 30009–30013 (2024).
Nguyen, N. A. & Agarwal, V. A leader-guided substrate tolerant RiPP brominase allows Suzuki–Miyaura cross-coupling reactions for peptides and proteins. Biochemistry 62, 1838–1843 (2023).
Fuchs, S. W. et al. A lanthipeptide-like N-terminal leader region guides peptide epimerization by radical SAM epimerases: implications for RiPP evolution. Angew. Chem. Int. Ed. 55, 12330–12333 (2016).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Alford, R. F. et al. The rosetta all-atom energy function for macromolecular modeling and design. J. Chem. Theory Comput. 13, 3031–3048 (2017).
Shimafuji, C. et al. In vitro catalytic activity of N-terminal and C-terminal domains in NukM, the post-translational modification enzyme of nukacin ISK-1. J. Biosci. Bioeng. 120, 624–629 (2015).
Thibodeaux, C. J., Wagoner, J., Yu, Y. & van der Donk, W. A. Leader peptide establishes dehydration order, promotes efficiency, and ensures fidelity during lacticin 481 biosynthesis. J. Am. Chem. Soc. 138, 6436–6444 (2016).
Agarwal, V. & McShan, A. C. The power and pitfalls of AlphaFold2 for structure prediction beyond rigid globular proteins. Nat. Chem. Biol. 20, 950–959 (2024).
Buel, G. R. & Walters, K. J. Can AlphaFold2 predict the impact of missense mutations on structure?. Nat. Struct. Mol. Biol. 29, 1–2 (2022).
Pak, M. A. et al. Using AlphaFold to predict the impact of single mutations on protein stability and function. PLOS ONE 18, e0282689 (2023).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomolecular NMR 6, 277–293 (1995).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Frueh, D. P. Practical aspects of NMR signal assignment in larger and challenging proteins. Prog. Nucl. Magn. Reson. Spectrosc. 78, 47–75 (2014).
Shen, Y. et al. Consistent blind protein structure generation from NMR chemical shift data. Proc. Natl. Acad. Sci. 105, 4685–4690 (2008).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Leman, J. K. et al. Macromolecular modeling and design in Rosetta: recent methods and frameworks. Nat. Methods 17, 665–680 (2020).
Nivón, L. G., Moretti, R. & Baker, D. A pareto-optimal refinement method for protein design scaffolds. PLoS One 8, e59004 (2013).
Froning, K. et al. Computational stabilization of T cell receptors allows pairing with antibodies to form bispecifics. Nat. Commun. 11, 2330 (2020).
Acknowledgements
The authors are thankful to the National Institutes of Health (1R35GM142882 to V.A. and 1R37GM058822 to W.A.V.) for support. A.C.M. acknowledges start-up funding from the Georgia Institute of Technology. This research was supported in part through research cyberinfrastructure resources and services provided by the Partnership for an Advanced Computing Environment (PACE) at the Georgia Institute of Technology, Atlanta, Georgia, USA.
Author information
Authors and Affiliations
Contributions
F.N.U.V., W.A.V., A.C.M., and V.A. designed research. F.N.U.V. prepared recombinant protein samples. FNU V., A.C.M., and H.W. performed NMR experiments, resonance assignments, and structure calculations. F.N.U.V. performed circular dichroism experiments. A.C.M. and V.A. performed molecular modeling. F.N.U.V. performed enzyme activity assays. Y.L. performed methyllanthionine configuration determination experiments and analyzed data with W.A.V. F.N.U.V., W.A.V., A.C.M., and V.A. wrote the paper with feedback from all authors. A.C.M. and V.A. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yong-Xin Li, A. Link, and George P. Lisi for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vidya, F., Luo, Y., Wu, H. et al. Large protein-like leader peptides engage differently with RiPP halogenases and lanthionine synthetases. Nat Commun 16, 9273 (2025). https://doi.org/10.1038/s41467-025-64365-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64365-3