Abstract
Profiling RNA modifications is essential to understand their functions and interactions. By taking the advantages of nanopore direct RNA sequencing, we present DirectRM, enabling simultaneous detection of six abundant modifications (N4-acetylcytidine, 1-methyladenosine, 5-methylcytidine, N7-methlguanosine, N6-methyladenosine, and pseudouridine) in native RNAs. Its two-stage pipeline identifies candidate modified kmers using binary classifier, then determines specific modifications and positions using an attention-based neural network. Trained with molecule-level features extracted from native RNA samples and validated on human cell lines and viral RNAs, DirectRM demonstrates high sensitivity, precision and robustness, outperforming existing tools. Crucially, we reveal the associations between modifications at both transcript and molecule-level. Modifications tend to proximate to each other on the transcript level, while at the molecule level, the presence of one modification is likely to reduce the occurrence of modifications at adjacent positions. DirectRM offers a powerful approach for studying epitranscriptome complexity and is expandable for future research.
Similar content being viewed by others
Introduction
While RNA was traditionally viewed as merely a mechanism for genetic information transfer, the fast-developing field of RNA modification or epitranscriptomics has revealed a complex and dynamic layer of regulation that influences various RNA-related processes1. The functional repertoire of RNA molecules can be diversified by modifications including methylation, pseudouridylation, and oxidation2,3,4,5,6,7. Those post-transcriptional modifications may function independently or collaborate with adjacent modifications to regulate various biological processes. They have been implicated in many molecular functions such as RNA stability, translation efficiency, splicing and degradation, and RNA-protein interactions. Biological processes thereby regulated include responses to environmental stimuli, cancer development, neurological disorders, and metabolic syndromes8,9,10,11,12,13,14,15,16,17,18,19.
Mapping and quantification of RNA modifications is crucial for understanding their functional implications and interactions in diverse biological contexts. In the past decades, Next Generation Sequencing (NGS) platforms, coupled to modification-specific treatments (e.g., antibody immunoprecipitation), have been commonly used for profiling and quantifying targeted modification types9,20,21,22,23,24,25,26,27,28,29. However, given that each type of modification requires a distinct enrichment method, it is unable to simultaneously profile the landscapes of multiple modification types. Also, independent experiments tend to capture inconsistent epitranscriptomic pictures, thus hindering the modeling of the RNA modification dynamics. Furthermore, different detection approaches may output quantification results in different formats (e.g., Log2Fold enrichment for MeRIP-seq versus Conversion rate for Bisulfite-seq). Ultimately, quantitative analysis results from different approaches cannot be directly compared and merged for systematic analysis. These limitations impede our ability to study the potential crosstalk between modifications.
A number of computational tools have been proposed, targeting the integration of epitranscriptomic data collected from different techniques or experiments for in silico detection of multiple types of RNA modifications with integrated prediction models, such as MultiRM30,31,32. These tools greatly advanced our understanding of the relationship between modifications33. However, it is important to note that these tools used primary sequence or genomic features for inference: this is obviously constant across cell lines or conditions, so that such methods cannot model the dynamic feature of RNA modifications30,31,32. Additionally, the isoform ambiguity issue of short-read NGS sequencing seriously restricts the ability of both experimental and relevant computational-based analysis of RNA modification to characterize isoform-specific modification landscapes34.
The direct RNA sequencing method, introduced by Oxford Nanopore Technologies (ONT), enables real-time, long-read, single-molecule, and direct sequencing of RNAs35. It sequences by deconvoluting the electric signals generated when individual molecules pass through the nanopore sensor36. Modifications along the molecules can be identified as distinct changes in the electric signals. Its ability to read native RNAs allows direct and simultaneous detection of all potential modifications along each molecule37.
To date, several tools have been implemented to infer RNA modifications from nanopore sequencing data. Examples include m6Anet, nanom6A, and DENA for m6A detection, nanoPsu and nanoMUD for pseudouridine (Ψ) detection, ELIGOS and Tombo for de novo detection, and TandemMod for multiple types of modification detection34,38,39,40,41,42,43,44,45,46,47,48. These pilot studies have demonstrated the feasibility of nanopore sequencing for modification detection with improved resolution (i.e., transcriptome-wide or molecule-level) and provided new insights into the epitranscriptome. However, there are still limitations. Firstly, existing tools are either focused on a single RNA modification type, mainly N6-methyladenosine (m6A) and Ψ, or only report non-canonical bases without specifying their modification types (e.g., modified adenosines), which constrains the study of the interplay between modifications. Additionally, some tools focused their analysis on certain sequence contexts. For instance, m6Anet and DENA only call m6A events on DRACH motifs, leaving a significant gap in the study of non-DRACH m6As43. In addition, these tools were trained with different groups of features (e.g., base call error features and signal features), algorithms (e.g., supervised and comparative methods), label sources, training samples (i.e., in vivo transcribed or in vitro transcribed (IVT) RNAs), and logics. The differences in detection methodologies lead to results that are highly specific to each method, making it difficult to directly compare or integrate findings across studies. This highlights the need for standardized approaches or integrative frameworks to enable more comprehensive analyses. Moreover, while the nanopore sensor can accommodate 5 or 9 nucleotides at a time, the presence of one canonical base can influence the signal of its neighbors. Since most existing tools only consider one modification type, their results tend to have false positives or false negatives caused by modified neighbors. TandemMod, which aimed to study multiple modifications, still suffers from the false positive issue since it applied independent models for each modification class46. Additionally, it was trained with IVT RNAs, which contain unnatural residues (e.g., consecutive m6As or m5Cs) and which may therefore distort the model.
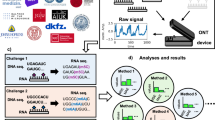
These challenges and limitations highlight a clear need for a more robust and versatile tool for the study of dynamic RNA modifications and their relationships. Here, we address these challenges by introducing DirectRM, which allows simultaneous detection of six of the most abundant RNA modifications, namely, N4-acetylcytidine (ac4C), 1-methyladenosine (m1A), 5-methylcytidine (m5C), N7-methylguanosine (m7G), m6A, and Ψ, through an integrated multi-label model. To capture the natural epitranscriptomic landscapes, we trained the model using molecule-level features extracted from nanopore sequencing data of natural human RNAs, paired with gold-standard labels derived from accurate NGS-based experiments. Notably, recognizing that RNA modifications are highly dynamic and exhibit significant variability across cell lines, we ensured that direct RNA sequencing and NGS-based experiments capture consistent, cell-specific modification profiles. After carefully evaluating alternative training strategies and model architectures, we implemented a two-stage detection pipeline: (1) de novo detection, where candidate modified kmers are identified with a binary classification network, and (2) modification position and class inference, where the specific modified base and its type within selected kmers are determined using an attention-based multi-label network. DirectRM was utilized to analyze modification patterns in two human cell lines, HepAD38 (parenchymal cell line derived from organs) and MT-4 (suspension cell line derived from blood), as well as viral RNAs from HTLV-1. This analysis provided initial insight into the dynamic epitranscriptome and revealed correlations between modifications across different biological contexts.
Results
Strict labeling enables high-accuracy neural de novo modification detection
Firstly, we performed de novo modification detection, where non-canonical kmers were identified regardless of their modification type. To precisely annotate or label kmers for model training (i.e., canonical or non-canonical), we tried two indicators. The Wilcoxon test was used to measure the deviation of mean current intensities from expected levels, and kmers with a p value smaller than 0.01 were considered as positive (Label A). The second indicator is the presence of more than one base call error, since previous studies have demonstrated that the modifications will increase the base calling errors (Label B). Base call errors refer to insertion, deletion, mismatch, and decreased base quality. Details for identifying the threshold for decreased base call qualities can be found in the Supplementary Note. We also tried a stricter criterion by taking the intersection of Labels A and B (Label C).
We visualized the signal feature distribution grouped by the three indicators with Principal Component Analysis (PCA) plot (Fig. 1a). It was found that the Wilcoxon test provided better clustering results, indicating better annotation of non-canonical kmers for de novo model training. The combination of the two indicators also showed satisfactory performance but provided many fewer positive instances. We further trained models based on the three label sets with HepAD38 samples, and Labels A and C derived from the Wilcoxon test achieved significantly better performance (Area Under Receiver Operating Characteristic (AUROC) = 0.954/0.949) than the base call error label (AUROC = 0.722) (Fig. 1b).
a PCA plot of features grouped by different labels or indicators. Label A refers to the p value from the two-sided Wilcoxon test of observed and expected mean signal intensities. Instances with a p value smaller than 0.01 are considered as positive. Exact p values for each instance can be found from the Source data file. Label B refers to the presence of base call errors. Instances with base call errors (i.e., insertion, deletion, mismatch, or decreased base call quality) are considered positive. Label C is the intersection of Label A and Label B. b ROC curves for the de novo model trained with Labels A, B, and C. c Precision, recall, and F1 scores under different coverage cutoffs. The results were derived from the IVET dataset. Source data are provided as a Source data file.
To further evaluate the performance of indicators, we tested the model performance with In Vitro Epitranscriptome (IVET) datasets (i.e., m6A, m1A, m5C paired with an unmodified control). Since the IVET dataset was sequenced with the RNA002 chemistry, we used the de novo model implemented for the RNA002 for comparison. Table 1 and Supplementary Table 1 summarize the F1-score, precision, and recall of model performance on the three datasets. While the Label A model showed a slightly better AUROC than Label C on the validation dataset, its performance on the IVET datasets was notably worse, averaging an F1-score of 0.639 compared to Label C’s 0.7. Thus, we decided to use the stricter indicator (i.e., p value of the Wilcoxon test smaller than 0.01 and the presence of base call errors) for de novo model training.
The read-level results provided by the de novo model can be pooled to estimate the site-level modification probability. We evaluated the influence of coverage cutoff on the recall and precision of detection with the IVET datasets. As shown in Fig. 1c, increasing the coverage cutoff led to a notable trade-off between recall and precision. Higher coverage improved recall by enabling more comprehensive detection of modification sites, as insufficient coverage at lower levels may leave modified reads undetected. However, this improvement in recall was accompanied by a slight decline in precision, likely due to the accumulation of false positives at higher coverage cutoffs. To optimize the precision-recall balance, the F1-score was calculated across coverage cutoffs. F1-score increased sharply from 1× to 10× coverage, followed by a performance plateau beyond 10×, suggesting this threshold represents an inflection point where additional reads do not necessarily improve the performance. Thus, we suggest using 10× coverage when estimating the site-level modification probability and modification level, which enables sufficient detection performance while accommodating detection on low-abundance transcripts.
After optimizing the de novo detection model, we performed benchmark comparison with published de novo detection models, namely Tombo and ELIGOS, on the IVET dataset. It is worth noting that Tombo and ELIGOS utilize statistical tests to identify modified kmers, where they compare the observed kmer features with expected ones. Statistical methods, as employed by these two methods, rely heavily on pre-existing knowledge of nanopore signals. Tombo and ELIGOS were primarily implemented for RNA002 chemistry, which only has 45 possible kmers. The relatively small number of possibilities makes it feasible for statistical methods to handle. However, in the scenario of the RNA004 chemistry, the number of possible kmers exponentially increases to 49 possibilities, which presents a significant challenge for statistical methods. In contrast, our model leverages the power of deep learning algorithms to perform de novo modification detection. Deep learning models are inherently designed to handle high-dimensional data and can effectively learn from and adapt to more complex scenarios. According to the results (Table 1 and Supplementary Table 1), our deep learning-based model outperformed the two statistical methods on the benchmark datasets sequenced with the RNA002 chemistry and shows promise as a more effective approach for handling the increased complexity of the RNA004 samples.
In order to demonstrate the feasibility of the de novo model on natural samples, we collected 570 and 509 modification sites (single-base resolution) for HepAD38 and MT-4, respectively, and 46 modification peaks for HTLV-1 samples. The kmer level and read-level results were pooled to estimate the modification probabilities for candidate sites. According to Table 2, the de novo model successfully identified most candidate sites in the three samples, demonstrating its detection ability when applied to native RNA samples.
Modification inference model training with NGS-derived native modification labels
After identifying non-canonical kmers, the next step is to determine the modification type(s) and the exact positions of modification(s) within kmers. While prior methodologies frequently employed IVT data for molecule-level labeling purity, this approach introduces significant biological artifacts. IVT systems generate non-physiological modification patterns, exemplified by consecutive m6A residues, since it has complete nucleotide replacement with modifications34,44. Quantitative analysis revealed that approximately 90% of residues in standard IVT 9-mers exhibit modification frequencies exceeding natural observations provided by NGS techniques (Fig. 2a). Furthermore, IVT dataset are inherently limited to single modification types, failing to capture the biologically critical phenomenon of modification co-occurrence (Fig. 2b). Since nanopore sequencer operates on a scale of 5 or 9 nucleotides (5 nt for RNA002 kit and 9 nt for RNA004 kit), kmers with multiple modifications are expected to have distinguishable signals from kmers with single modifications. For example, the event of kmers having both m5C and ac4C sites is significantly different from those of kmers having only m5C or ac4C modifications (Fig. 2b). These results confirmed the frequent co-occurrence of modification events and distinct combinatorial modification signal profiles, necessitating models that account for modification interactions.
a Pie chart showing the percentage of natural and unnatural IVT 9-mers. Natural 9-mers are defined as those with a modification count consistent with NGS observations. Unnatural 9-mers are categorized based on the number of additional modifications compared to NGS observations: low aberrant (1 additional modification), moderate aberrant (2 additional modifications), high aberrant (3–4 additional modifications), and severe aberrant (>5 additional modifications). b Upset plot showing the co-occurrence of different modifications within 9-mer windows. c Graphical illustrations of label generation from NGS techniques. d ROC plots illustrating the performance of multi-label models implemented for RNA002 and RNA004 chemistry. e Transcriptomic distribution of identified m6A sites. f Motif analysis of identified m6A sites. g Bar charts showing AUROC scores for the independent and integrated multi-label models. h Pie charts showing the percentage of sites with different numbers of modified neighbors. i Bar charts showing the error rate of independent and integrated multi-label on different IVET datasets. Source data are provided as a Source data file.
To preserve authentic epitranscriptomic complexity, we therefore utilized gold-standard NGS-derived modification sites for training label generation. Figure 2c demonstrates the label generation strategies. Cell-specific modification coordinates were obtained by overlapping collected NGS sites from public sources with our cell-specific peaks. Modified kmers that overlapped with collected sites were considered as positive instances for the relevant class.
In order to determine the modification identity and class, while capturing the interactions between modifications, we planned to use an attention-based multiple instances and multi-label learning framework. We primarily proposed 8 model structures and performed systematic comparisons from the computational and biological perspectives (Supplementary Tables 2–4, Supplementary Figs. 1–3). The detailed model optimization process can be found in the Supplementary Note. The final proposed structure is composed of six parallel sub-models, each dedicated to one modification class, and each featuring an attention mechanism that determines the modified position(s) within the kmer bag, an LTSM-based feature extractor for key feature extraction, and a fully connected classifier that provides the modification probability for the target class. The model was optimized in an integrated manner with multi-label learning algorithms to capture the cross-modification dependencies and influences. Evaluation on RNA002 and RNA004 datasets demonstrated state-of-the-art performance: the model achieved excellent binary AUROC for each class (>0.95) and multi-label Area Under Precision-Recall Curve (PRAUC) of 0.92 (Fig. 2d). Beyond machine learning metrics, the model successfully recapitulated fundamental biological patterns of RNA modifications. Detected m6A sites exhibited pronounced enrichment near the end of the coding sequence (CDS) region and precisely identified the consensus DRACH motifs (Fig. 2e, f). These patterns align with previous m6A studies and confirm our model’s capacity to extract meaningful biological signatures.
Integrated model allows better modification detection than independent models
To comprehensively validate the robustness of our integrated multi-label model, we further conducted systematic comparisons against independently optimized binary classifiers, where each modification-specific sub-model was trained separately using binary classification algorithms. Our integrated model demonstrated significant performance enhancements at the molecule level, improving AUROC scores by 0.06–0.13 across all modification classes compared to independent binary classifiers (Fig. 2g).
At the site-level evaluation, we quantified the distribution of modification sites by counting occurrences in each group defined by the number of modified neighbors within 9-mer windows. This operation is based on a critical consideration that the nanopore protein can accommodate multiple nucleotides. Thus, the presence of modified neighbors may warn of potential false positives. According to Fig. 2h, the integrated model generated modification profiles showing substantially stronger concordance with NGS ground truth, indicating better capacity to mitigate false positives induced by adjacent modifications and better capacity to recapitulate the epitranscriptomic patterns. In contrast, within the independently identified datasets, a large proportion of sites (43%) were found to have other modified sites within 9 bp up or downstream, which is greatly higher than in the collected NGS sets (20%). This observation can be more likely explained by the influence of nearby non-canonical signals rather than each of these loci being a true positive. The possible explanation for the improvement can be that the integrated model leverages the multi-label learning algorithm, which allows it to capture dependencies and correlations between different modification types. For example, certain modifications may co-occur or mutually exclude each other. It is noticeable that multi-label learning faces the imbalanced positive-to-negative issue, making the model prioritize the majority class. In many scenarios, such an issue could introduce undesired bias. However, in our case, the dataset is composed of approximately a 1:5 positive-to-negative ratio, which may make the model aware of the sparse nature of modifications. Also, to mitigate the potential bias caused by this class imbalance issue, we applied the Asymmetric Loss, which ensures that the model pays sufficient attention to the positive minority class during training. Moreover, since the independent model was optimized exclusively for a single modification type, when confronted with co-occurring modifications or complex signal patterns (e.g., overlapped modification signals), it may exhibit systematic failures. According to Fig. 2b, we have confirmed the frequent co-occurrence modification events and demonstrated that they have distinct modification profiles. To this end, the significant inconsistency between the independent-model results and NGS profiles is attributable to the fact that the independent model failed to deconvolute those overlapped signals. In contrast, the integrated multi-label model can resolve these limitations through interconnected optimization that explicitly learns those complex modification signal patterns. This holistic view enables our integrated multi-label model to make more informed analyses.
To further validate our hypothesis, we performed validation on high-purity IVET datasets and revealed a fourfold reduction in the error rate: the integrated multi-label model achieved an approximately 5% error rate versus a 20% error rate for the independent binary classifiers (Fig. 2i). This evidence collectively establishes that independent optimization introduces systematic detection artifacts, while our end-to-end multi-label model maintains biological fidelity across molecular, site, and experimental validation levels.
DirectRM: simultaneous identification of multiple RNA modifications
The above experiments, comparisons, and optimizations resulted in the final DirectRM framework, which allows simultaneous detection of multiple RNA modifications from direct RNA sequencing data. Figure 3 illustrates the overall design of DirectRM. Firstly, molecule-level kmer features, including base call error features and various signal features, were extracted from natural RNA samples. Then, the binary de novo detection model was trained to identify potential modified kmers. To infer the modification class and position along those candidate modification kmers, an integrated multi-label model was trained. The overall model is composed of six independent models for each modification class. Each model is composed of a feature extractor, which extracts key features from the vector, an attention mechanism, which provides the attention score for each kmer position, and fully connected layers, which provide the modification probability for the target class. The overall model was trained in an integrated manner with cell-specific gold-standard labels derived from paired NGS-based experiments. The optimized model will output the attention score of each kmer position and the modification probability of each class. Classes with a modification probability larger than 0.5 will be considered as present in the kmer region, and the exact modification position will be assigned to relevant nucleotides with the highest attention score. Conflicting estimations will be weighted averaged to provide the results. Optionally, molecule-level results can be collated to estimate the transcript-level or gene-level modification rate. We implemented DirectRM for both the established RNA002 sample and the latest RNA004 chemistry. In addition, DirectRM allows adaptive selection of the re-squiggle algorithm: both the ONT remora and the latest Uncalled4 are supported49. Detailed benchmark comparison on the re-squiggle algorithm was performed and can be found in the Supplementary Table 5 and Supplementary Note. DirectRM is publicly available at: https://github.com/yuxinPenny/DirectRM50.
Overall design of DirectRM.
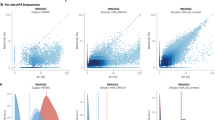
To rigorously evaluate DirectRM’s performance, we conducted systematic benchmarking against published modification detection tools trained independently for specific modification classes, mainly m6A and Ψ. Samples of five different human cell lines and one mouse cell line were collected for comparison. For m6A detection, three representative tools, namely m6Anet, DENA, and nanom6A, were evaluated. Cell-relevant m6A sites were collected and used to validate the results of the tools. Since different tools report varied numbers of m6A sites, to ensure a fair comparison, we performed 10 rounds of random sampling, selecting 3000 sites from each tool’s results in each round. For each round, we calculated the percentage of sites validated by cell-relevant NGS sites. As systematically benchmarked in Table 3 and Supplementary Data 1, DirectRM demonstrated superior accuracy in predicting m6A modifications across multiple cell lines. In A549 samples, up to 46.18% of DirectRM-identified sites were validated by NGS techniques, significantly outperforming nanom6A (24.94%), m6Anet (18.97%), and DENA (14.03%). This performance advantage was consistently observed in all tested cell lines, where DirectRM consistently achieved the highest validation rates among comparative tools. Notably, the absolute percentage of validated sites across different cell lines (e.g., 1%–46%) is attributable to the data availability (e.g., MeRIP-seq samples) for cell lines. Specifically, the number of confidently detectable cell-specific sites ranged from 126,220 (A549) to 12184 (K562), resulting in proportional fluctuations in validation rates. Collectively, these results establish that DirectRM robustly maintains performance advantages despite cell-specific epitranscriptomic complexity. Critically, DirectRM uniquely enables detection at non-DRACH motifs, a capability absent in comparative tools. Due to the limited availability of cell-specific Ψ datasets across diverse cell lines, our benchmarking approach for Ψ detection tools required methodological adaptation. To maintain rigorous evaluation standards despite this constraint, we implemented a benchmarking strategy using all confidently detected single-base resolution sites aggregated from available datasets. DirectRM was benchmarked against nanoPsu and nanoMUD using 5282 and 4291 experimentally validated human and mouse Ψ sites, respectively. DirectRM again demonstrated significant advantages, as its predictions are more accurate and reliable. In contrast, both NanoMUD and NanoPsu, which were trained with IVT data, exhibit an extremely high number of false positives in their results. This comprehensive validation establishes DirectRM as a versatile framework that simultaneously maintains the state-of-the-art accuracy while extending detection capacity to previously unaddressable contexts.
Generalization capability of DirectRM demonstrated on the human MT-4 cell line
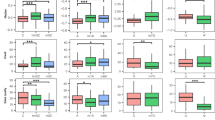
To demonstrate that our optimized framework can be generalized to different cell lines, we replicated the HepAD38-related experiments using the MT-4 cell line sequenced with RNA004 chemistry (i.e., two replicates of ONT sequencing sample and three replicates of NGS-based modification detection for the independent test). Following the training data preparation strategy on the HepAD38 training sample, we prepared independent testing datasets with two MT-4 replicates. Read-level signal features, base call error features, and sequence features were extracted, and MT-4 relevant sites were used as labels for testing. We visualized the molecule-level performance with Receiver Operating Characteristics (ROC) curves for each class (Fig. 4a). The results suggested an excellent generalizing ability at the molecule level, with all AUROC scores larger than 0.85. Moreover, sites for two additional modification types, namely A-to-I and hm5C, were collected. Firstly, the sensitivity of the de novo detection model was assessed, and it was found that modification sites can be accurately identified (Fig. 4b). Then, we calculated the probabilities for A-to-I and hm5C sites of being one of the six targeted modifications, and the results were summarized in Fig. 4c. The low probabilities provided for the two non-training modifications (Median probabilities around 0.25) indicate excellent specificity of the multi-label model.
a ROC curves for molecule-level evaluation of the DirectRM model on the two replicates of the MT-4 datasets. b Percentage of A-to-I and hm5C sites that have been successfully identified by the de novo model. c Distribution of the site-level modification probabilities for A-to-I and hm5C sites of being ac4C, m1A, m5C, m7G, and Ψ provided by the DirectRM model for the two MT-4 biological replicates. Data are presented as boxplots showing the median (center line), the 25th and 75th percentiles (box limits), and the whiskers extending to the most extreme data points within 1.5 × interquartile range (IQR) from the box. Individual data points beyond the whiskers are plotted as outliers (dots). n = 1000 for each group. d Percentage of DirectRM-identified MT-4 sites validated by the NGS technique on mRNAs. e Percentage of sites identified on non-coding RNAs. The “others” category includes miscRNA, snoRNA, etc. f Upset plot showing the intersection of HepAD38 modification profiles and the two replicates of MT-4 samples. g Dot plot showing the site modification rate under HepAD38 cell lines, and two replicates of MT-4 samples. Differentially modified sites under different cell conditions or replicates were defined as: FC < 1/3 or FC > 3. The dashed line depicts the predicted mean values of the Y-axis for given values of X. The surrounding shaded area represents the 95% confidence interval for the regression line. Source data are provided as a Source data file.
We then validated the collected profiles with gold-standard sites collected from the NGS technique. According to Fig. 4d, within the DirectRM collected sites, a proportion of sites can be validated by accurate NGS techniques. For example, more than half of the identified m6A and Ψ sites were also reported by NGS-based techniques. In addition to those NGS-reported sites, DirectRM also reported numerous sites that were ignored by previous NGS-based experiments. It is worth noting that since NGS-based techniques are likely to enrich polyA-tailed RNAs, the detection of modifications was very limited to certain RNA types, especially mRNAs. In our study, by taking the advantages of nanopore sequencing, 9012 sites on diverse types of modifications, including lncRNA, rRNA, and snoRNA, were identified (Fig. 4e). Some of the sites were also reported by other studies. For instance, a Ψ site (18S:406) on rRNA was also reported by Milenkovic et al.51. This site was identified as a tissue-specific rRNA modification site and thus can be used for tissue and cell development stage prediction. More detailed information on modifications to non-coding RNAs, as well as evidence that supports those modification sites (258 sites in total), can be found in Supplementary Data 2.
RNA modifications exhibit highly dynamic features, with their presence and patterns varying across different cell lines, tissues, and biological replications, highlighting their context-dependent regulatory roles. To this end, we systematically compared RNA modification profiles across distinct cell lines (HepAD38 vs MT-4) and biological replicates (MT-4 rep 1 vs MT-4 rep 2). As shown in Fig. 4f, analysis revealed 16,848 HepAD38-specific sites, while the remaining sites were conserved between HepAD38 and MT-4 cell lines. Notably, dynamic modification patterns were also observed across replicates: 7106 and 3.576 sites were uniquely identified in the two MT-4 replicates. To quantify modification dynamics at shared sites (present in both cell lines and replicates), we compared their modification rate under respective cellular conditions. Differentially modified sites were rigorously defined as those exhibiting >3-fold change (FC > 3 or FC < 1/3) in modification rates (Fig. 4g). Strikingly, pairwise comparison between cell lines demonstrated significant divergence in modification intensities (Spearman correlation ρ = 0.62), with a substantial fraction of sites classified as differentially modified sites. This pronounced heterogeneity underscores the cell-type-dependent plasticity of the epitranscriptome. Conversely, site modification rates remained relatively static across different MT-4 replicates, suggesting consistent enzymatic activities or regulations.
Site-directed mutagenesis verifies the modifications identified by DirectRM on HTLV-1 viral RNAs
In addition to human cell lines, DirectRM was also applied to study the RNA modification landscapes on HTLV-1 RNAs. To evaluate the feasibility of our approach on viral RNAs, we performed similar tests to those with MT-4 samples. Firstly, we tested the sensitivity of the de novo model and the specificity of the multi-label model with candidate A-to-I and hm5C sites collected from the NGS technique. The results illustrated that DirectRM achieved equally good performance on viral RNAs as with human cells. All the A-to-I and hm5C sites were successfully screened by the de novo detection model and can be distinguished from the six target modifications by the multi-label model (Fig. 5a). Candidate modifications were then obtained from MeRIP-seq experiments and used to validate the profiles provided by DirectRM. A large proportion of sites were also validated by NGS techniques, especially m6A and Ψ, which illustrated the applicability of DirectRM for viral samples (Fig. 5b).
a Distribution of site-level modification probabilities for A-to-I and hm5C sites of being ac4C, m1A, m5C, m7G, and Ψ provided by the multi-label model for the two HTLV-1 biological replicates. Data are presented as boxplots showing the median (center line), the 25th and 75th percentiles (box limits), and the whiskers extending to the most extreme data points within 1.5× interquartile range (IQR) from the box. Individual data points beyond the whiskers are plotted as outliers (dots). n = 1000 for each group. b Percentage of DirectRM-identified sites validated by the NGS technique. c The circos plot shows the quantitative modification profiles on HTLV-1 isoforms. d Relative modification levels for wildtype and mutant strains. Data are presented as mean values ± SD for each group (n = 3). All individual data points are overlaid as jittered points. Significance levels are: ***p < 0.001, ****p < 0.0001 based on one-way ANOVA tests with three biological replicates. Exact p values are provided in the Source data file. Source data are provided as a Source data file.
HTLV-1 is a single-stranded RNA virus that can cause adult T-cell leukemia. Its genome contains four genes and can express ten possible genes. After evaluating the performance of DirectRM on viral samples, we applied it to unveil the modification landscapes of these genes. In our study, we sequenced RNAs for seven of the ten genes and found that modifications were primarily enriched in two genes, namely Tax and Env, and were most enriched in the Tax gene (Fig. 5c and Supplementary Fig. 4). Among the six target modifications, four of them, namely m1A, m5C, m6A, and m7G, were confidently identified (modification rate > 0.1). To further validate the reliability of our detection, we performed experimental verification on six high-confidence sites on the Tax gene. Firstly, according to Supplementary Fig. 5, the MeRIP-qPCR analysis confirmed the presence of the four modifications on the Tax gene. We then constructed mutant plasmids containing single-nucleotide substitutions for modification sites (e.g., A to C mutation for the m6A site) and then transfected them into HEK293 cells. Subsequent MeRIP-qPCR analysis revealed a significant reduction in modification enrichment signals at mutant sites compared to their wild-type counterparts (Fig. 5d). This strong concordance between DirectRM detection and experimental validation underscores the reliability of our framework, particularly for the epitranscriptomic studies in viral RNAs.
Molecule-level investigation revealed potential associations between modifications
Previous studies have observed that the genomic distance between two modifications is significantly shorter than random, indicating potential clustering or positive associations between modifications30. In this study, by taking advantage of nanopore long reads, we obtained isoform-specific epitranscriptomic profiles with DirectRM on HepAD38 and MT-4 cells and evaluated the transcriptomic distance between modifications. Firstly, for all possible pairs of modifications (n = 30), we calculated the distance from testing modification sites to their nearest reference modification sites (denoted as query and subject here, respectively) and applied the median of distances to measure the distance between two modifications. The six modification profiles were permutated, and distances were calculated as above as a random control. As in the genome, a strong clustering feature between modifications can also be observed in different splicing isoforms. As shown in Fig. 6a and Supplementary Fig. 6a, the distance between most modifications is significantly closer than random (p value = 0.015 and 0.038). Then, we visualized the detailed distance information between modifications with a heatmap (Supplementary Fig. 6b). The color intensity represents the magnitude of the difference between the observed and permutated distances between two modifications. In general, the distance between two arbitrary modifications is closer than random, while in certain cases (e.g., from m6A to ac4C), their distance can be longer than random.
a Transcriptomic distance between two modifications in HepAD38. Data are presented as boxplots showing the median (center line), the 25th and 75th percentiles (box limits), and the whiskers extending to the most extreme data points within 1.5 × interquartile range (IQR) from the box. All individual data points are overlaid as jittered points. n = 30 (6 × 5) for each group. Significance level is p = 0.015 based on the one-sided t-test. b Modification rates for sites with and without modified neighbors in HepAD38. Sites with other modifications within a 100 bp region were considered as having modified neighbors. NA indicates that the site was the only site detected on that transcript. Data are presented as boxplots showing the median (center line), the 25th and 75th percentiles (box limits), and the whiskers extending to the most extreme data points within 1.5× interquartile range (IQR) from the box. n = 1000 for each group. Outliers are not shown in the figure. Significance levels are ****p < 0.0001 based on two-sided t-tests. Exact p values are provided in the Source data file. c Calculation of probability and conditional probability for the presence of modifications at adjacent positions. The pie chart shows the proportion of modification pairs that increase or reduce the presence of each other. d Read-level investigations of adjacent modification sites located in protein-binding regions. Three examples, including (1) m6A (ENST00000273968.5:576) and m5C (ENST00000273968.5:572) sites located in the binding region of YTHDF2 protein, (2) m7G (ENST00000313601.11:1830) and m6A (ENST00000313601.11:1857) site located in the binding region of IGF2BP3 protein, and (3) m6A (ENST00000375114.7:1779) and m5C (ENST00000375114.7:1759) site located in the co-binding region of YBX1 and IGF2BP protein family. The heatmap shows the read-level modification status for target sites. 30 reads were sampled for visualization. e Probabilities and conditional probabilities for the presence of modification at the above example sites. f SHAP waterfall plot shows the contribution of modifications to the transcript level. Source data are provided as a Source data file.
Given the co-occurrence feature of modifications, further analysis was conducted to explore the potential interaction between modifications. Firstly, we classified modifications into three groups: (1) modifications sites with modified neighbors (i.e., distance <100), (2) sites without modified neighbors, and (3) sites which were the only one detected on that transcript, denoted as NA (Fig. 6b and Supplementary Fig. 6c). Interestingly, we found that sites with modified neighbors have significantly lower modification rate than the other two groups. We hypothesized that different modifications may play similar roles in certain biological processes. When other modifications are present nearby, responsibilities can be shared, thus reducing the need for a high modification level to activate the function. Also, modifications can play opposing functions (e.g., stabilize RNA or lead to RNA degradation). In this case, the presence of one modification can influence and potentially reduce the presence of the other, resulting in a decrease in the level of modification. Additionally, we evaluated the relationship between distance to their nearest modification sites and site modification rate. According to Supplementary Fig. 6d, within 100 bp, the longer the distance to other modifications, the higher the modification rate, indicating the less the impact on the modification level by other modifications. When the distance is greater than 100, the modification levels remain stable with the increase of distance, indicating that the modification rate is not likely to be influenced by modification sites beyond a 100 bp region.
To validate the above hypothesis, we investigated the associations between modifications at the molecule level. For each pair of neighboring modification sites (i.e., distance smaller than 100), denoted as query site of modification A and subject site of modification B here, we calculated the probability for the presence of modification A on query position, denoted as P(query = A), and the condition probability for the presence of modification A given that modification B occurred on subject position on the same molecule, denoted as P(query = A|subject = B) here (Fig. 6c). According to the pie chart, for the majority of pairs of adjacent modification sites, the presence of one modification will reduce the presence of the other modification on the same molecule, with P(query = A) > P(query = A|subject = B). This can explain the earlier observation that sites with modified neighbors show significantly lower modification occupancy than sites without modified neighbors. Supplementary Fig. 6e, f shows the detailed numbers of site pairs between two given modifications that negatively or positively influence the presence of each other, respectively.
To infer the functional implications of those associated modifications, we analyzed their interactions with RNA-binding proteins and quantified their influence on the transcript level with SHAP analysis. Two examples of negatively associated neighboring modifications in HepAD38 cell lines illustrate the approach: (1) one m6A (ENST00000273968.5:576) and one m5C (ENST00000273968.5:572) site located in the binding region of YTHDF2 protein, and (2) m7G (ENST00000313601.11:1830) and m6A (ENST00000313601.11:1857) sites located in the binding region of IGF2BP3 protein (Fig. 6d, e). SHAP analysis was used to evaluate the contribution of each modification to the transcript level (gene expression level). Figure 6f shows the contribution of m6A and m5C within the YTHDF2 binding region on transcript ENST00000273968.5, and m7G and m6A within the IGF2BP3 binding region on transcript ENST00000313601.11, to the transcript expression level. In the first case, m5C and m6A modifications contribute oppositely to the transcript expression level. According to previous research, YTHDF2 protein can bind to m6A to promote the degradation of transcripts, while binding to m5C will stabilize the RNA structure and protect it from degradation33. Since m6A and m5C interact with the YTHDF2 protein to play opposing functions, these two modifications thus reduce the presence of each other in that region. Regarding the second case, m6A can bind to IGF2BP3 to protect transcript ENST00000313601.11 from degradation, while conversely, IGF2BP3 can promote mRNA degradation by interacting with m7G modification. This result is also consistent with the finding of Liu et al.52. In addition to the two negatively associated examples, we also presented one positively associated example in the co-binding region of YBX1 and the IGF2BP protein family. The likelihood of the presence of m6A or m5C increased significantly when the other modification occurred on the same reads. Since both m6A and m5C can promote the transcript expression level by interacting with YBX1 and IGF2BP proteins, we hypothesized that these two modifications may need to collaborate to exert function, so that the presence of one modification will increase the presence of the other. In addition to these examples, in HepAD38 cell lines, we identified 705 negatively and 155 positively associated cases in the binding region of 56 RNA-binding proteins on 216 isoforms (see Supplementary Data 3 for details). In MT-4 cell lines, we identified 489 negatively and 123 positively associated cases in the binding region of 59 RNA-binding proteins on 216 isoforms (see Supplementary Data 4 for details).
Discussion
Integrated analysis of different RNA modifications can provide valuable insight into their crosstalk and provide a better understanding of the complex epitranscriptome. With NGS platforms, different types of modifications are treated differently, introducing various technical biases and difficulties to the analysis. As a promising alternative to NGS platforms, the nanopore direct RNA sequencing technique offers the potential to detect all modifications present in each individual molecule. However, published tools were designed to identify certain modification types and were trained based on distinct fashions. In this project, we proposed the first integrated detection framework, DirectRM, for six abundant modifications, namely ac4C, m1A, m5C, m6A, m7G, and Ψ, from natural nanopore sequencing data.
De novo modification detection was first conducted to identify candidate non-canonical kmers. Three types of indicators, including the difference between observed and expected signal means, the presence of base call errors, and the combination of the two, were evaluated, with the combined indicator showing the best performance.
Subsequently, to determine the exact modification types and positions within modified kmers, an attention-based multi-instance multi-label learning framework was proposed. The architecture consists of six sub-models (one for each modification class), each equipped with an attention mechanism to determine the most likely modification position (based on the highest attention score) and a classifier to predict the modification probability for that class. The model was optimized with NGS labels derived from native RNAs, which better represent the biological complexity of RNA modifications, and an integrated multi-label learning framework, which enables the model to effectively capture the modification dependencies and to deconvolute complex modification signals. Systematic comparisons between our integrated multi-label model and independent binary models, along with benchmarking against existing ONT-based detection tools, demonstrated the robustness and practicality of our framework.
We also demonstrated the applicability of DirectRM on different human cell lines and viral RNAs. The results showed a high level of generalizing ability, with high AUROC scores of read-level results, high reproducibility, sensitivity, and specificity, further supported by experimental validation. Additionally, we observed the dynamic nature of RNA modifications, revealing distinct modification landscapes and levels across different cell lines, highlighting the biological variability and complexity of the epitranscriptome.
By taking advantage of nanopore long-read and single-molecule sequencing, we characterized the modification landscape on splicing isoforms and individual molecules with DirectRM. The association between modifications was investigated and analyzed from the transcriptome level to the read level. It was found that the presence of adjacent modifications (distance smaller than 100) on the same read will reduce (in most cases) or increase the presence of testing modifications and thus influence the modification rate and further RNA fate or levels. We also identified a number of associated modification site pairs i that are associated with RNA-binding proteins n HepAD38 and MT-4 cell lines, which indicate possible functional implications for associated modifications.
DirectRM enables integrated detection of multiple types of modifications with high accuracy, sensitivity, specificity, reproducibility, and generalizing ability. It has provided valuable insights into the dynamic and complex epitranscriptome and crosstalk between modifications, both in terms of position and occupancy. However, there are still some limitations to address. For instance, the current analysis focuses on only six of the most well-studied modifications, leaving room for future research to include additional modification types. Carefully designed synthetic datasets incorporating various modification types and better simulating real epitranscriptomic landscapes could be leveraged to enhance the model’s performance and capabilities. As our integrated detection model achieved a better balance between precision and recall compared to independent models, as evidenced by its significantly lower false positive rate. We still recognize the need for further evaluation to fully address the possibility of missed true positives. Additionally, the analysis of modification crosstalk remains constrained. For example, when assessing the distance between modifications, only linear distances were considered. Future studies could incorporate structural context, such as RNA secondary structure or circular RNA formation, to provide a more comprehensive understanding. Furthermore, when linking modification pairs to RNA-binding proteins, incorporating protein-binding preferences and additional experimental data could enhance the authenticity and depth of the analysis.
Methods
Data collection
To train and validate DirectRM, we used replicate samples from the human HepAD38 and MT-4 cell lines.
Cell culture
The HepAD38 (ATCC, CRL-3561) samples were provided by Prof. Quan Yuan’s laboratory from Xiamen University, Fujian, China. The procedures for sample collection, storage, and processing were carried out according to the protocols recommended by the providing laboratory. HepAD38 cells were stored in Dulbecco’s modified Eagle’s medium (DMEM) solution, which consists of Fetal Bovine Serum (FBS) at 10% concentration, tetracycline at 3 μg/mL concentration (HY-B0474, MedChemExpress, Shanghai, China), and G418 at 400 μg/mL concentration (Catalog no. 345810, Sigma-Aldrich, St. Louis, MO, USA). HEK293T cell line was obtained from the American Type Culture Collection (ATCC, CRL-3216) and cultured in DMEM. The HTLV-1-transformed T cells MT-4 were purchased from Changzhou Haotiangi Biotechnology Co., Ltd(HTQ-XB-0733) and cultured in RPMI 1640 medium. All cell lines were cultured in the recommended growth medium augmented with 10% FBS (ST30-3302, PAN Seratech) and incubated at 37 °C under 5% CO2. All samples were tested for mycoplasma contamination.
Immunoprecipitation-based methods for RNA modification detection
To obtain the dynamic RNA modification profiles for different cell lines, we performed independent modification detection experiments on HepAD38, MT-4, and HTLV-1 RNAs. The antibody-immunoprecipitation-based methods for ac4C, m1A, m5C, hm5C (for MT-4 samples), m6A, and m7G detection were performed by SeqHealth Technology Co., Ltd (Wuhan, China), namely ac4C-RIP-seq, m1A-MeRIP-seq, m5C-MeRIP-seq, hm5C-MeRIP-seq, m6A-MeRIP-seq, and m7G-MeRIP-seq. Briefly, RNA was extracted from HepAD38 or MT-4 samples using Trizol reagent (Invitrogen, CA, USA), and poly(A) RNA selection was applied to remove ribosomal RNA. Purified RNAs were fragmented, denatured, and sheared into approximately 100nt fragments. Then, RNA fragments were split into the Input and the Immunoprecipitation (IP) sample. IP samples contain modified fragments enriched by modification-specific antibodies. The IP and Input samples were reverse transcribed to a cDNA library and finally subjected to paired-end sequencing (PE150) on the Illumina NovaseqTM 6000 platform (SeqHealth Technology Co., Ltd). All experiments were replicated three times.
The raw data from the above antibody-immunoprecipitation-based methods were collected and processed with stringent bioinformatic pipelines to obtain the modification peaks for each modification. First, TrimGalore (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) was used to remove adapter sequences and reads of low quality. The cleaned reads were then aligned to the human reference genome GRCh38 (UCSC genome browser) using Hisat253. Then, peak calling was performed with exomePeak2 based on the IP and the Input sample. To identify HTLV-1 viral RNAs, unmapped reads from the human genome were further aligned to the HTLV-1 genome (GenBank: AB513134.1) using Bowtie54. Those mapped reads were considered as HTLV-1 RNAs and were subsequently subjected to exomePeak2 (https://github.com/ZW-xjtlu/exomePeak2) to identify modification peaks on the viral genome. Additionally, the input sample for m6A-MeRIP-seq was analyzed to identify A-to-I sites55. The RED-ML tool was used here. The single-nucleotide polymorphism, simple repeat region, and Alu regions as required by the RED-ML tool were downloaded from the UCSC genome browser. The REDItools “Denovo” function was used to detect A-to-I sites on HTLV-1 viral RNA.
PA-Ψ-seq
To obtain Ψ profiles from HepAD38, MT-4, and PA-Ψ-seq experiments were performed. Cells were cultured in 10 cm dishes with a cell density of 6 × 106 cells/dish and then pulsed with 100 µM of 4-Thiouridine (Selleck Cat # E1292) for a further 24 h. RNA was extracted using Trizol (Invitrogen, cat # 15596026). Immunoprecipitation was performed using antipseudouridine (anti-Ψ) antibody (MBL # 347-3). The antibody-RNA mixture was then crosslinked with UV 365 nm 2500 × 100 µJ/cm2 twice in a UV Stratalinker, and then treated with RNase T1 (0.1 U/µL) for 15 min at 22 °C. The immunoprecipitated RNAs were processed with QIAseq® miRNA UDI Library Kit (Qiagen, 331505) to generate the cDNA library for sequencing. The cDNA library was quantified and finally sequenced on the Illumina NovaseqTM 6000 platform using PE150 mode.
Similarly, the raw data from PA-Ψ-seq were first aligned to the human reference genome GRCh38 using Bowtie, and in MT-4 samples, unmapped reads were further aligned to the HTLV-1 genome reference to identify HTLV-1-specific reads. The enriched regions (i.e., modification peaks) of Ψ were identified with the MACS2 peak caller and then annotated by Chipseeker.
Single-base modification sites collection
It is important to remember that the above antibody-immunoprecipitation-based methods and PA-Ψ-seq have not achieved single-base resolution for modification detection: they report modification peaks instead. To locate modification sites within obtained peaks, we collected modification sites from public sources. We collected 9645 ac4C, 4320 m1A, 4008 m5C, 427760 m6A, 8487 m7G and 5282 Ψ sites from m6A-atlas2, m7GHub, and other sources56,57,58,59,60,61,62. To ensure consistency between modification profiles captured by different experiments, for each cell line, only sites located within cell-specific peaks were considered. After filtration, 136 ac4C, 710 m1A, 68 m5C, 115866 m6A, 478 m7G and 708 Ψ were kept for HepAD38 cell line and 199 ac4C, 532 m1A, 50 m5C, 81188 m6A, 744 m7G and 744 Ψ for MT-4 cell line. Since very few ac4C, m1A, and m5C sites were kept after filtration, an insufficient number for model training, we randomly selected 1000 additional sites for each in further analysis.
Nanopore direct RNA sequencing
Samples were sequenced with RNA004 chemistry, which is newly introduced as an eventual replacement for RNA002 chemistry. Since the RNA002 kit is currently the most used, we also performed direct RNA sequencing with this for the training sample (i.e., HepAD38). Two replicates for each cell line were acquired.
Samples sequenced with RNA004 kits were acquired in Pod5 format. The raw Pod5 data were base-called and mapped to the reference sequencing by Dorado v0.6.2 with the RNA004_130bps_hac@v3.0.1 model. The alignment file SAM file was compressed, sorted, and indexed with Samtools. Then, we performed signal refinement (i.e., re-squiggle) with ONT Remora and Uncalled4, which are compatible with the RNA002 and RNA004 kits49. A systematic benchmarking of alignment algorithms was conducted, with Remora ultimately selected in our downstream analysis. For data in fast5 format (i.e., sequenced with RNA002 kit), it should be converted to Pod5 with the pod5 convert_to_pod5 function before launching Dorado. The GRCh38 genome and transcriptome sequences downloaded from UCSC were used as references for HepAD38 and MT-4 reads, while the AB513134.1 genome sequence downloaded from NCBI was used as a reference to identify HTLV-1 reads.
Machine learning approaches
Feature extraction
For each qualifying read, its related BAM file and Pod5 file were loaded and combined by the Remora Python API to form an object that contains sequence, mapping, signal, and signal refinement information. We extracted features for targeted regions with a 9-mer sliding window (i.e., the sequencing unit) and a step size of 5nt, which creates positional overlaps between consecutive instances, thus ensuring critical motifs can be fully captured even at the end of the window. Two groups of features, namely the base call error features and signal features, as well as the kmer sequence, were extracted. In terms of signal features, we calculated the mean, median, minimum, maximum, 1st and 3rd quantiles, standard deviations, and length of the signal event for each position within the kmer. Base call error features include insertion, deletion, mismatch, and base qualities. The kmer sequence was one-hot encoded.
De novo modification detection
De novo detection refers to identifying non-canonical kmers with an unknown modification type. To optimize the de novo detection model, we benchmarked three types of indicators of modified and unmodified kmers: (1) difference between observed means of current intensities and expected levels (provided by Remora), kmers with Wilcoxon test p value smaller than 0.01 were considered as positive; (2) base call errors, kmers having more than one insertion, deletion, mismatch, or base with decreased qualities, were considered as positive; and (3) kmers in the intersection of (1) and (2) were considered as positive.
An equal number of modified and unmodified negative instances extracted from HepAD38 samples (training sample) were combined and partitioned 70:30 into a training:validation dataset. A deep network composed of a bidirectional LTSM unit, a fully connected network, and a SoftMax function was used. Binary cross-entropy loss was used for model parameter tuning, and an Adam optimizer with an initial learning rate of 0.01 was used for model updates. The StepLR learning rate scheduler with decay of 0.1 and an early stop mechanism was used to avoid overfitting. Model performance was evaluated by commonly used binary classification metrics, including AUROC, PRAUC, accuracy, and F1-score.
The optimized de novo detection model was then applied to samples to select candidate modified kmers for subsequent analysis.
Modification type and exact location inference
To determine the modification type (i.e., ac4C, m1A, m5C, m6A, m7G, and Ψ) and exact location within selected kmers, for each modification class, we applied an attention-based multi-instance multi-label learning architecture. The overall model is composed of six sub-models, and each focuses on one modification type. The sub-model contains three components: (1) an attention mechanism that determines the position of modification within the kmer bag, (2) a feature extractor that captures the key feature patterns, and (3) a fully connected network that provides the modification probabilities for the target class. We primarily proposed 8 model architectures and performed systematic benchmarking from both the computational and biological perspectives to select the optimal model structure.
The training dataset comprises candidate modified kmers identified from the HepAD38 samples by the de novo detection model. The candidate label sets were generated from the gold-standard NGS techniques. Modified kmers overlapped with HepAD38-related modification sites were considered positive for the relevant class. For example, if a kmer overlapped with HepAD38-relevant m6A and m7G sites, its candidate label sets will be [0,0,0,1,1,0]. The dataset was randomly divided into 70:30 training: validation datasets. It is worth noting that all negative (i.e., [0,0,0,0,0,0]) instances were removed from the training dataset.
The multi-label learning algorithm was used to optimize the modification inference model. Given the positive-negative imbalance issues of multi-label learning, we carefully selected the Asymmetric Loss function for model parameter tuning. Similarly to de novo model training, an Adam optimizer with a learning rate of 0.01 was used for model optimization. StepLR with a learning rate decay of 0.1 and early stopping was used to avoid overfitting. Model performance was evaluated with multi-label accuracy, multi-label AUPRC, as well as the binary AUROC, precision, and recall for each class.
The optimized multi-label was used to infer the kmer modification status in new samples. Classes with a modification probability larger than 0.5 will be considered present in the kmer region, and the exact modification position will be assigned to the relevant nucleotide with the highest attention score. Given that the choice of different window sizes and step sizes may lead to redundant estimation of certain positions, conflicting estimations will be weighted averaged to provide the final results.
Benchmark comparison of de novo detection models
We compared our de novo detection model with two published tools that allow de novo detection, Tombo and ELIGOS. The IVET datasets from plant cDNA libraries were downloaded from the GEO database under accession number GSE227087 and used as benchmark datasets63. The IVET datasets are composed of three modified datasets (m6A, m1A, and m5C), which were used as positive, and one unmodified negative control. On each dataset, Tombo, ELIGOS, and DirectRM models were run following the provided guidance to get relevant results. It is noticeable that IVET samples were sequenced with RNA002 chemistry; the DirectRM de novo detection model implemented for RNA002 chemistry was used for comparison. F1-score, Precision, and Recall were calculated based on the results to evaluate the performance of models.
Benchmark comparison of modification detection models
We compared our DirectRM framework with published modification detection tools, including m6Anet, nanom6A, and DENA for m6A detection, and NanoMUD, NanoPsu for Ψ detection. We collected direct RNA sequencing samples for five human cell lines, namely HeLa (PRJNA872027), HEK293 (PRJEB40872), K562 (PRJNA931346), A549 (PRJEB44348), HepG2 (PRJEB44348), as well as one mouse mESC (PRJEB39819) sample from the NCBI SRA database. On each sample, these tools were run following the provided guidance to get relevant results. For benchmarking of m6A detection tools, for each cell line, we collected m6A-MeRIP-seq data from m6A-atlas2 and obtained cell-relevant m6A sites by overlapping the collected sites with peaks. Results derived from different ONT tools were validated by cell-relevant NGS sites. Since different tools report varied numbers of m6A sites, to ensure a fair comparison, we performed 10 rounds of random sampling, selecting 3000 sites from each tool’s results in each round. For each round, we calculated the percentage of sites validated by cell-relevant NGS sites. In terms of Ψ modification, due to the limited availability of cell-specific Ψ datasets across diverse cell lines, we adapted to use confidently detected single-base resolution sites aggregated from available datasets. Similarly, an equal number of sites were sampled from the results of different tools, and the percentage of sites being validated was calculated. Notably, since existing tools were implemented for RNA002 chemistry, these collected samples were sequenced with the RNA002 kit, and the DirectRM model implemented for the RNA002 kit was used for comparison.
Validation on the human MT-4 cell line
To demonstrate that the multi-label model trained with HepAD38 can be generalized to different cell lines, we prepared an independent testing dataset with MT-4 samples. Similarly, multi-label accuracy, multi-label AUPRC, and binary AUROC, precision, and recall for each class were calculated to evaluate the performance. Then, quantitative modification profiles were obtained with DirectRM. The ONT-derived results were compared and validated using gold-standard NGS-derived sites. To evaluate the feasibility of the overall detection pipeline, in addition to the six target modification classes, we collected sites for A-to-I and hm5C modification. We assessed whether those sites can be accurately identified with the de novo detection model and subsequently be distinguished from the six testing classes with the multi-label model.
Validation of HTLV-1 viral RNAs
Plasmid constructions
The gene sequence of Tax was designed based on the reference sequence obtained from the NCBI database (GenBank Accession AB513134). The wild-type DNA sequence of the target genes was synthesized by Fuzhou Sunya Biotechnology Co., LTD. The DNA CDS was cloned into the XbaI-NotI(NEB) sites of the pCDH-CMV-MCS-EF1a-mCherry-T2A-Puro vector, resulting in constructs named pCDH-Tax-wt. Mutant versions of the Tax CDS were also generated, including m6A (A7380C, A7402C), m1A(A7553C, A7666C), m5C (C7373A), and m7G (G7978T, G7576T), and denoted as pCDH-Tax-A7380C, pCDH-Tax-A7402C, pCDH-Tax-A7553C, pCDH-Tax-A7666C, pCDH-Tax-C7373A, pCDH-Tax-G7978T, and pCDH-Tax-G7576T, respectively. Plasmid transfections were performed using Lipofectamine 3000 transfection reagent (L3000008; Invitrogen) according to the manufacturer’s protocol.
MeRIP-qPCR
The primary antibodies used in this study included: rabbit monoclonal antibody against m6A (SAB5600251, Sigma), mouse monoclonal antibody against m5C (68301-1-Ig, Proteintech), mouse monoclonal antibody against m7G (68302-1-Ig, Proteintech), mouse monoclonal antibody against m1A (68636-1-Ig, Proteintech), and normal rabbit/mouse IgG antibody (sc-2027/sc-2025, Santa Cruz Biotechnology). Total RNA was extracted from HEK293T cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). For immunoprecipitation, 50 μg of RNA was resuspended in immunoprecipitation (IP) buffer (150 mmol/L NaCl, 0.1% NP-40, 10 mmol/L Tris-HCl, pH 7.4) and incubated with 50 μL of magnetic beads (Selleck) pre-coated with the antibody mixture for 2 h at room temperature. The RNA-antibody mixture was then incubated overnight at 4 °C with either modification-specific antibodies or rabbit/mouse IgG antibody (Santa Cruz Biotechnology). Beads were washed six times with IP buffer and then incubated with 300 μL of elution buffer (5 mmol/L Tris-HCl, pH 7.5, 1 mmol/L EDTA, pH 8.0, 0.05% SDS, 4.2 μL 20 mg/mL proteinase K) at 50 °C for 1.5 h. The eluted RNA was purified using TRIZOL (Accurate Biology, Changsha, Hunan, China). One mg of total RNA was reverse transcribed into first-strand cDNA using the Evo M-MLV RT kit (AG11707; Accurate Biology) following the manufacturer’s instructions. RT-qPCR was performed with the SYBR Green Pro Taq HS qPCR Kit (AG11701; Accurate Biology). The primer sequences for the Tax gene were forward, 5′- CACTCAACCCTCACCACTCC-3′; reverse, 5′- TGGGGCTCATGGTCATTGTC-3′. Each qPCR analysis included at least three biological replicates, and three independent experiments were conducted.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The original contributions presented in the study are publicly available. The RIP-seq and PA-Ψ-seq data for MT-4 samples have been deposited in the National Center for Biotechnology Information (NCBI) database under accession code GSE287582. The RIP-seq data and PA-Ψ-seq data from HepAD38 samples have been deposited in the NCBI database under accession codes GSE286110 and GSE288115. The Direct RNA Sequencing (DRS) data generated in this study, including two replicates of HTLV-1 infected MT-4 samples sequenced with the SQK-RNA004 kit and two HepAD38 samples sequenced with the SQK-RNA002 and SQK-RNA004 kit, have been deposited in the NCBI database under accession code PRJNA1273605. The In Vitro Epitranscriptome (IVET) DRS datasets were obtained from the NCBI database under accession code GSE227087. The DRS data for human HeLa, HEK293, K562, A549, HepG2, and mouse mESC samples were obtained from the NCBI database under accession codes PRJNA872027, PRJEB40872, PRJEB39819, PRJNA931346, PRJEB44348. Source data used to generate the figures are provided with this paper. Source data are provided with this paper.
Code availability
The code used to develop the model, perform the analyses, and generate results in this study is publicly available and has been deposited in https://github.com/yuxinPenny/DirectRM at GitHub repository. The specific version of the code associated with this publication is archived in Zenodo and is accessible via https://doi.org/10.5281/zenodo.1693847850. DirectRM is licensed under the terms of the MIT license.
References
Brenner, S., Jacob, F. & Meselson, M. An unstable intermediate carrying information from genes to ribosomes for protein synthesis. Nature 190, 576–581 (1961).
Netzband, R. & Pager, C. T. Epitranscriptomic marks: emerging modulators of RNA virus gene expression. Wiley Interdiscip. Rev. RNA 11, e1576 (2020).
Kennedy, E. M. et al. Posttranscriptional m(6)A editing of HIV-1 mRNAs enhances viral gene expression. Cell Host Microbe 19, 675–685 (2016).
Cohn, W. E. Pseudouridine, a carbon-carbon linked ribonucleoside in ribonucleic acids: isolation, structure, and chemical characteristics. J. Biol. Chem. 235, 1488–1498 (1960).
Shi, H., Chai, P., Jia, R. & Fan, X. Novel insight into the regulatory roles of diverse RNA modifications: re-defining the bridge between transcription and translation. Mol. Cancer 19, 78 (2020).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Lee, S. Y., Kim, J. J. & Miller, K. M. Emerging roles of RNA modifications in genome integrity. Brief. Funct. Genom. 20, 106–112 (2021).
Li, X., Ma, S. & Yi, C. Pseudouridine: the fifth RNA nucleotide with renewed interests. Curr. Opin. Chem. Biol. 33, 108–116 (2016).
Carlile, T. M. et al. Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells. Nature 515, 143–146 (2014).
Anderson, B. R. et al. Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation. Nucleic Acids Res. 38, 5884–5892 (2010).
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013).
Torres, A. G., Batlle, E. & Ribas de Pouplana, L. Role of tRNA modifications in human diseases. Trends Mol. Med. 20, 306–314 (2014).
Batista, P. J. et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Haussmann, I. U. et al. m(6)A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature 540, 301–304 (2016).
Chen, J. et al. m(6)A Regulates Neurogenesis and Neuronal Development by Modulating Histone Methyltransferase Ezh2. Genom. Proteom. Bioinform. 17, 154–168 (2019).
Engel, M. et al. The Role of m(6)A/m-RNA methylation in stress response regulation. Neuron 99, 389–403 e389 (2018).
Kmietczyk, V. et al. m(6)A-mRNA methylation regulates cardiac gene expression and cellular growth. Life Sci Alliance 2, e201800233 (2019).
Liu, B. et al. A potentially abundant junctional RNA motif stabilized by m(6)A and Mg(2). Nat. Commun. 9, 2761 (2018).
Nombela, P., Miguel-López, B. & Blanco, S. The role of m(6)A, m(5)C and Ψ RNA modifications in cancer: novel therapeutic opportunities. Mol. Cancer 20, 18 (2021).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Chen, K. et al. High-resolution N(6)-methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angew. Chem. Int. Ed. 54, 1587–1590 (2015).
Ke, S. et al. A majority of m6A residues are in the last exons, allowing the potential for 3’ UTR regulation. Genes Dev. 29, 2037–2053 (2015).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Molinie, B. et al. m(6)A-LAIC-seq reveals the census and complexity of the m(6)A epitranscriptome. Nat. Methods 13, 692–698 (2016).
Koh, C. W. Q., Goh, Y. T. & Goh, W. S. S. Atlas of quantitative single-base-resolution N(6)-methyl-adenine methylomes. Nat. Commun. 10, 5636 (2019).
Garcia-Campos, M. A. et al. Deciphering the “m(6)A Code” via antibody-independent quantitative profiling. Cell 178, 731–747 e716 (2019).
Zhang, Z. et al. Single-base mapping of m(6)A by an antibody-independent method. Sci. Adv. 5, eaax0250 (2019).
Meyer, K. D. DART-seq: an antibody-free method for global m(6)A detection. Nat. Methods 16, 1275–1280 (2019).
Marchand, V. et al. AlkAniline-Seq: profiling of m(7) G and m(3) C RNA modifications at single nucleotide resolution. Angew. Chem. Int. Ed. 57, 16785–16790 (2018).
Song, Z. et al. Attention-based multi-label neural networks for integrated prediction and interpretation of twelve widely occurring RNA modifications. Nat. Commun. 12, 4011 (2021).
Liu, K. & Chen, W. iMRM: a platform for simultaneously identifying multiple kinds of RNA modifications. Bioinformatics 36, 3336–3342 (2020).
Salekin, S. et al. Predicting sites of epitranscriptome modifications using unsupervised representation learning based on generative adversarial networks. Front. Phys. 8, 196 (2020).
Chen, Z. et al. YTHDF2 promotes ATP synthesis and immune evasion in B cell malignancies. Cell 188, 331–351 (2025).
Jenjaroenpun, P. et al. Decoding the epitranscriptional landscape from native RNA sequences. Nucleic Acids Res. 49, e7 (2021).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Lu, H., Giordano, F. & Ning, Z. Oxford nanopore MinION sequencing and genome assembly. Genom. Proteom.Bioinform. 14, 265–279 (2016).
McIntyre, A. B. R. et al. Single-molecule sequencing detection of N6-methyladenine in microbial reference materials. Nat. Commun. 10, 579 (2019).
Liu, H. et al. Accurate detection of m(6)A RNA modifications in native RNA sequences. Nat. Commun. 10, 4079 (2019).
Price, A. M. et al. Direct RNA sequencing reveals m(6)A modifications on adenovirus RNA are necessary for efficient splicing. Nat. Commun. 11, 6016 (2020).
Pratanwanich, P. N. et al. Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore. Nat. Biotechnol. 39, 1394–1402 (2021).
Gao, Y. et al. Quantitative profiling of N(6)-methyladenosine at single-base resolution in stem-differentiating xylem of Populus trichocarpa using Nanopore direct RNA sequencing. Genome Biol. 22, 22 (2021).
Huang, S. et al. Interferon inducible pseudouridine modification in human mRNA by quantitative nanopore profiling. Genome Biol. 22, 330 (2021).
Qin, H. et al. DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N(6)-methyladenosine on RNA. Genome Biol. 23, 25 (2022).
Zhang, Y. et al. NanoMUD: profiling of pseudouridine and N1-methylpseudouridine using Oxford Nanopore direct RNA sequencing. Int. J. Biol. Macromol. 270, 132433 (2024).
Hendra, C. et al. Detection of m6A from direct RNA sequencing using a multiple instance learning framework. Nat. Methods 19, 1590–1598 (2022).
Li, J. et al. YTHDF1 promotes mRNA degradation via YTHDF1-AGO2 interaction and phase separation. Cell Prolif. 55, e13157 (2022).
Chen, S., Meng, J. & Zhang, Y. Quantitative profiling N1-methyladenosine (m1A) RNA methylation from Oxford nanopore direct RNA sequencing data. Methods 228, 30–37 (2024).
Li, J. et al. Detection and Quantification of 5moU RNA Modification from Direct RNA Sequencing Data. Curr. Genom. 25, 212–225 (2024).
Kovaka, S. et al. Uncalled4 improves nanopore DNA and RNA modification detection via fast and accurate signal alignment. Nat. Methods 22, 681–691 (2025).
Zhang. Y. et al. DirectRM: integrated detection of landscape and crosstalk between multiple RNA modifications using direct RNA sequencing. DirectRM. https://doi.org/10.5281/zenodo.16938478 (2025).
Milenkovic, I. et al. Epitranscriptomic rRNA fingerprinting reveals tissue-of-origin and tumor-specific signatures. Mol. Cell 85, 177–190.e177 (2025).
Liu, C. et al. IGF2BP3 promotes mRNA degradation through internal m(7)G modification. Nat. Commun. 15, 7421 (2024).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Picardi, E. & Pesole, G. REDItools: high-throughput RNA editing detection made easy. Bioinformatics 29, 1813–1814 (2013).
Liang, Z. et al. m6A-Atlas v2.0: updated resources for unraveling the N6-methyladenosine (m6A) epitranscriptome among multiple species. Nucleic Acids Res. 52, D194–d202 (2024).
Zhang, Y. et al. DirectRMDB: a database of post-transcriptional RNA modifications unveiled from direct RNA sequencing technology. Nucleic Acids Res. 51, D106–d116 (2023).
Wang, X. et al. m7GHub V2.0: an updated database for decoding the N7-methylguanosine (m7G) epitranscriptome. Nucleic Acids Res. 52, D203–d212 (2024).
Chen, K. et al. RMDisease: a database of genetic variants that affect RNA modifications, with implications for epitranscriptome pathogenesis. Nucleic Acids Res. 49, D1396–d1404 (2021).
Song, B. et al. RMDisease V2.0: an updated database of genetic variants that affect RNA modifications with disease and trait implication. Nucleic Acids Res. 51, D1388–d1396 (2023).
Lu, L. et al. Base-resolution m(5)C profiling across the mammalian transcriptome by bisulfite-free enzyme-assisted chemical labeling approach. Mol. Cell 84, 2984–3000.e2988 (2024).
Dai, Q. et al. Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA. Nat. Biotechnol. 42, 1559–1570 (2024).
Wu, Y. et al. Transfer learning enables identification of multiple types of RNA modifications using nanopore direct RNA sequencing. Nat. Commun. 15, 4049 (2024).
Acknowledgements
This work was funded by grants from The National Natural Science Foundation of China (Grant No. 32300552 to D.H.), Joint Funds for the Innovation of Science and Technology in Fujian Province (Grant No. 2023Y9004 to K.C.), Fujian Medical University High-level Talent Research Startup Funding Project (Grant No. XRCZX2021019 to K.C.), Fujian Research and Training Grants for Young and Middle-aged Leaders in Healthcare (Grant No. 2023WJWZQN02 to K.C.), Jiangsu Science and Technology Program (Grant No. BK20230279 to D.H.). Additionally, the authors thank Hang Li and Mengqian Lin for technical support and Ruimei Chen for assistance with the manuscript preparation.
Author information
Authors and Affiliations
Contributions
K.C., D.H., and J.M. (Jia Meng) conceived the idea and supervised the project. Y.Z. performed computational analysis and wrote the manuscript. Y.W. (Yuecheng Wu) prepared the cell samples and performed the wet-lab validation on HTLV-1. J.M. (Jiongming Ma) and H.W. help with the computational analysis. Y.W. (Yiyu Wu) and L.L. help with the wet-lab experiments. G.J. and D.R. co-supervised the project. All authors read and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Y., Wu, Y., Ma, J. et al. DirectRM: integrated detection of landscape and crosstalk between multiple RNA modifications using direct RNA sequencing. Nat Commun 16, 9450 (2025). https://doi.org/10.1038/s41467-025-64495-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64495-8