Abstract
Light-driven water splitting by plants, algae and cyanobacteria is pivotal for global bioenergetics and biomass formation. A manganese cluster bound to the photosystem II proteins catalyzes the complex reaction at high rate, but the rate-determining factors are insufficiently understood. Here we trace the oxygen-evolution transition by time-resolved polarography and infrared spectroscopy for cyanobacterial photosystems genetically modified at two strategic sites, complemented by computational chemistry. Our results highlight three rate-determining roles of the protein environment of the metal cluster: acceleration of proton-coupled electron transfer, acceleration of substrate-water insertion after O2-formation, and balancing of rate-determining enthalpic and entropic contributions. Whereas in general the substrate-water insertion step may be unresolvable in time-resolved experiments, here it likely becomes traceable because of deceleration by genetic modification. Our results may stimulate new time-resolved experiments on substrate-water insertion in photosynthesis, clarification of enthalpy-entropy compensation in enzyme catalysis, and knowledge-guided development of inorganic catalyst materials.
Similar content being viewed by others
Introduction
Plants, algae, and cyanobacteria harvest solar energy and convert it into chemical energy by producing carbohydrates from water and carbon dioxide, coupled with the release of molecular oxygen (O2) into the atmosphere. This process of oxygenic photosynthesis facilitates primary biomass formation and provides the chemically stored energy that “fuels” life on Earth. A protein-cofactor complex called photosystem II (PSII) plays a central role by facilitating the light-driven oxidation of two “substrate water” molecules, thereby generating O2 as well as four energized electrons (reducing equivalents) and four protons1,2,3,4,5. Here, we address the clearly crucial but still insufficiently understood roles of H-bonded protein-bound water clusters in photosynthetic oxygen evolution. Progress comes from the informative combination of problem-adapted methodology: site-directed mutagenesis verified by protein mass spectroscopy, infrared (IR) difference spectroscopy in various frequency ranges, activation energy analyses based on time-resolved O2-polarography, tracking of reaction sequences by time-resolved IR spectroscopy, and large-scale molecular dynamics (MD) simulation with a force-field specifically parameterized for the PSII state prior to onset of O2-formation.
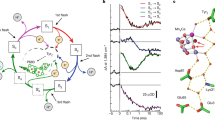
The absorption of four light quanta by the PSII pigments is required to complete one turnover of the water-oxidation cycle, each initiating the primary charge separation and resulting in the formation of a chlorophyll cation radical (P680+) at the electron donor side of PSII. The chlorophyll radical oxidizes a redox-active tyrosine residue (YZ), followed by oxidation of the protein-bound Mn4CaO5-6 cluster that facilitates the water oxidation chemistry in conjunction with its protein-water environment1,2,3,4,5. Figure 1 shows the sequence of events in water oxidation and its atomistic key players. The basic model proposed by Kok already in 1970 comprises four semi-stable Si-states (S0–S3, with S1 being the dark-stable S-state) and the transiently formed S4-state, where the respective subscript indicates the number of accumulated oxidizing equivalents (Fig. 1a, inner circle)6. Whereas in the three S-state transition leading from the S0 to S3, oxidizing equivalents are accumulated by oxidation of Mn ions1,7,8,9, the S4-state is likely rather reached by oxidation of an oxygen (O6), resulting in a Mn-bound oxyl radical10.
a Water oxidation cycle: Starting in the dark-stable S1 state, four consecutive light flashes drive PSII through the five semi-stable S-states of Kok’s classical S-state cycle (inner circle of Si-states6). By alternating oxidation (e─) and proton removal (H+) steps, the Mn4CaOx cluster accumulates up to 4 oxidation equivalents, while avoiding excessive charge accumulation (outer cycle of Sin/+ states11). The electrons are removed by transfer to the previously oxidized tyrosine residue (YZ); the protons move to the periphery of PSII; two water molecules are inserted3,87; O2 is formed and released. Approximate room temperature values of the respective time constants are indicated13. b Active-site structure: Water molecules and selected residues in the vicinity of the Mn4CaO5-6 cluster based on the 2.07 Å two-flash (S3-rich) crystallographic model (PDB ID: 6DHO, PSII from Thermosynechococcus vestitus BP-1; also used for Mn4CaO6 cluster structure in panel a)16. Residues from the D1, D2, and CP43 subunits are depicted in green, yellow, and pink, respectively; Mn in purple, Ca in yellow, Cl in green. Water molecules and six μ-oxo bridges (O1, O2,…, O6) are depicted as red or orange spheres. The pentagon of H-bonded water molecules, consisting of W26 to W30, has been termed “water wheel” and brought into play as an element of substrate water delivery16,17,19,46. Numbering of water molecules is based on ref. 16. Approximate boundaries of the Cl1, O4, and O1 channels near the Mn4CaO5-6 cluster are depicted in orange (Cl1), blue (O4), and yellow (O1). The proposed directionalities in proton-coupled electron transfer and water insertion in the oxygen-evolving S3→ S4→ S0 transition are indicated, as are the two key residues investigated here by genetic modification (D1-D61 and D1-N298).
Water oxidation in PSII cannot be understood without considering the four protons that need to be removed from two water molecules prior to O2 formation. Therefore, Kok’s reaction cycle model has been extended to include four proton-removal steps (Fig. 1a, outer circle)11, which mostly can be followed in time-resolved experiments (see time constants indicated in Fig. 1a)12,13. The binding of the two water molecules that represent the substrates of the water oxidation chemistry is also indicated in the extended S-state cycle of Fig. 1a, as established primarily by advanced H2O16/H2O18 water exchange experiments3,14. Here we focus on the oxygen-evolving S3→S4→S0 transition, for which we have previously inferred the following sequence of events from experimental and computational results10:
-
(1)
The light-induced YZox oxidation promotes proton release from the proton-sharing D2-E312—D1-E65 dyad within about 300 µs, YZox S3+ YZox S3n + H+.
-
(2)
Electron transfer from the Mn4CaO6-cluster to YZox is coupled to proton transfer (PT) to D1-D61 and results in oxyl-radical formation within about 3 ms, YZox S3n → YZ S4+.
-
(3)
Not rate-limiting and thus kinetically invisible (i) peroxide formation, (ii) O2-formation, and (iii) O2-release result in a Mn4CaO4-cluster with vacant water binding sites, S4+ → S0+ + O2.
-
(4)
Deprotonation of a water molecule and hydroxide insertion in the Mn4CaO4− cluster results in the stable S0-state of the Mn4CaO5-cluster, S0+ + H2O → S0n + H+, which seems to be a kinetically invisible step in the wild-type PSII15 but possibly resolvable in the here investigated D1-N298A variant.
The Mn4CaO4-6 cluster is connected to the thylakoid lumen by extensive networks of H-bonded water molecules and amino-acid sidechains (Fig. 1b), which have been termed “channels”. The three major channels (“Cl1”, “O4”, and “O1”) likely facilitate proton egress and/or substrate water access16,17,18,19,20,21,22,23,24. The “Cl1” (or “broad”) channel leads from YZ and the Mn4CaO5-6 cluster’s Ca ion past aspartate-61 of the D1 subunit (D1-D61) and the Cl‒ ion that is coordinated by lysine-317 of the D2 subunit (D2-K317). This pathway likely facilitates proton egress during the S2→S325,26,27,28 and S3→S010,18,25,26,27,29,30,31 transitions. The “O1” or “large” channel leads from YZ and the Mn4CaO5-6 cluster’s Ca ion past O1. Recent studies have focused on this channel as the primary entry pathway of substrate water16,17,19,20,21,22,23,24,28,32. Asparagine-298 (D1-N298) provides a link between YZ (via its proton acceptor, D1-H190) and the water wheel (waters W26, W27, W28, W29, and W30) that has been proposed to serve as a water delivery site via the O1 channel17,19; tautomerization to its imidic acid form might support a proton translocating role of D1-N298A33. The critical importance of D1-N298 in the water oxidation reactions of PSII has been demonstrated by mutagenesis and IR studies. First, mutations of D1-N298 substantially decrease steady-state rates of O2 evolution detected under high-light (saturating) continuous illumination34,35, but the mechanistic origin of the decreased rate could not be identified. Second, from static FTIR studies it was concluded that the N298A mutation perturbed the network of H-bonds that surround the Mn4CaO5-6 cluster and substantially diminished the efficiencies of the S2→S3 and S3→S0 transitions34. Third, time resolved FTIR studies show that the N298A mutation decreases the rates of the S1→S2 and S2→S3 transitions 3-7-fold, but the oxygen-evolving S3→4→S0 transition was not investigated28. Here, we show that the N298A PSII variant can undergo the oxygen-evolving S3→S4→S0 transition equally fast and efficiently as the wild-type PSII. A paradox—between the severely reduced O₂-evolution rate under continuous illumination and the rapid O₂ formation detected by time-resolved polarography—is identified and resolved by a scenario supported by time-resolved IR experiments: a severe slowdown of substrate-water insertion after O2 formation by the sidechain exchange in the N298A variant.
We also conducted time-resolved IR studies on the D1-D61A variant of PSII, for comparison with wild-type PSII and the N298A variant. Comparison with D61A is especially informative because of its clearly pivotal role in PSII water oxidation. It facilitates Mn4CaO5-6 cluster oxidations10,36,37,38,39 and mutations of D61 substantially decrease steady-state rates of O2 evolution27,29,40, perturb the H-bond networks surrounding the Mn4CaO5-6 cluster25,30, and diminish the efficiencies25,30 as well as rates29,41,42 of the individual S-state transitions, all this likely relating to its role as an acceptor of protons removed from the substrate water molecules10,37,43,44. (In the following the mutations D61A and N298A of the D1 protein subunit will be simply referred to as D61A and N298A).
The present investigation addresses three roles of the protein-water environment of the catalytic metal cluster in the oxygen-evolving S3→S4→S0 transition, clearly demonstrating that the H-bond protein-water clusters surrounding the Mn4CaO4-6 cluster are not merely a passive protein-water matrix but also an essential part of the evolutionary fine-tuned active site. (a) Confirming previous experimental studies29,45, we now find by time-resolved IR spectroscopy in the D61A variant a drastic slowdown of the rate-limiting step of S4-formation, explainable by the crucial role of the D61 in the proton transport path for removing a proton from a Mn-bound substrate hydroxide concomitantly with oxyl radical formation10. (b) For the N298A variant, the combination of time-resolved O2-polarography and IR spectroscopy provides evidence that the insertion of a “fresh” substrate water after O2-release is optimized by the protein-water network. We discover that the N298A modification likely causes a pronounced slowdown of the deprotonation-coupled water insertion step, which is also investigated by molecular mechanics (MD) simulations for ca. 106 atoms over a total simulation period of 5 µs per PSII variant. (c) Unexpectedly, we encounter a third role of the protein-water network by analysis of the temperature dependence of the O2-formation step. The enthalpic and entropic contribution to the free energy of activation are strongly affected by the exchange of a single residue (N298A), with an almost unvaried rate constant value at room temperature. This interesting case of entropy-enthalpy compensation is also investigated by extensive MD simulations. Finally, we discuss possible implications for water oxidation in inorganic catalyst materials.
Results
Targeted protein variations, oxygen-evolving activities, and verification of site mutations
Two PSII point mutants of Synechocystis sp. PCC 6803 were investigated for which previous investigation have suggested that they could affect the processes in the oxygen-evolving S3→S0 transition (see Fig. 1a) in distinctly different ways. A specific aspartate (D61 of the D1 protein) has been frequently suggested to act as a pivotal proton acceptor in several of the S-state transitions, transiently accepting protons removed from substrate water molecules25,27,29,30,31. Recently, a very specific proposal on its role in the rate-determining step(s) of the oxygen-evolving transition has been presented10. Briefly, the rate-determining step is the formation of a reactive oxyl radical, which is facilitated by coupling the transfer of an electron to the previously light-oxidized tyrosine sidechain (YZ in Fig. 1b) to PT from a metal-bound hydroxide to D61, the latter involving three further metal-bound water species. This S3→S4 transition is preceded by deprotonation of a glutamate pair (D1-E65—D2-E312, not shown in Fig. 1b) in the S3+→S3N transition that precedes the proton-coupled electron transfer and likely “prepares” the D61 by structurally mediated increase in its proton affinity10,44, in line with recent results obtained by time-resolved crystallography46. Thus, the exchange of the protonatable carboxylate sidechain of the D61 against an inert methyl group (D61A mutant) is predicted to profoundly affect light-driven oxygen evolution10,37,44,47. Indeed, a strong decrease in the rate of oxygen evolution is observed for the D61A variant (Table 1), in good quantitative agreement with results previously obtained for the same genetic variant31,45.
The second here selected protein variation is the asparagine-alanine exchange at position 298 of the D1-protein (N298A mutant). It is more remote from the site of oxyl radical formation and O-O bond formation, and thus it is not expected to be directly involved in these reactions. An indirect, possibly energetic effect on these processes might result from H-bonding of the asparagine to the histidine (H190) that accepts a proton upon YZ oxidation and shuttles it back in the S3→S4 transition. Yet of particular interest regarding the role of this residue is its association with the “water wheel” comprising W29, W28, W26, W27, and W30, which has been suggested, based on crystallographic data16,17,19,46, to be crucial in water delivery to the active-site metal cluster.
Even though the N298 residue likely is not related to the rate-determining step of oxyl radical formation in the S3→S4 transition, the light-saturated steady-state rates of O2 evolution of the N298A variant were at least equally strongly diminished as found for the D61A residue exchange, see Table 1. The reduction in the O2 evolution in comparison to unmodified wild-type (WT) protein is similar to those reported previously in Synechocystis sp. PCC 680334 and Chlamydomonas reinhardtii35 for the N298A and D61A variants.
In the D61A variant, the decreases in steady-state oxygen evolution corresponds to a reaction time constant that is increased by clearly more than one order of magnitude. (Note that the influence of the sidechain exchange on the time-resolved O2 polarography data is expected to be greater than that on the corresponding steady-state rate; this is expected because the latter is determined by a co-limitation of acceptor and donor side processes). In the N298A variant, however, the severely reduced steady-state rate of oxygen evolution is not matched by a deceleration of O2-evolution time constant, but rather a slight acceleration from 3.0 to 1.9 ms is observed for O2-evolution at 10 °C. To explain this divergence, we propose that in the N298A variant the reaction steps of oxyl radical formation, O-O bond formation and O2 release are largely unaffected by the residue exchange, whereas subsequent steps of insertion of a hydroxide in the void left by O2-release are slowed down significantly.
Recently the posttranslational modification or even restoration of genetic variations obtained by site-directed at the donor side of PSII has been described48. To verify the presence of the D1-A61 or D1-A298 residues in mutation-bearing PSII core complexes48,49, the amino acid sequences of chymotryptic or chymotryptic/tryptic peptides of isolated D1 subunits were analyzed by mass spectrometry. In N298A, at least 99% of the residue at the mutation position was the desired alanine (see Supplementary Figs. 1 and 2). In D61A, approx. 98.5% of the residue at the mutation position was the desired alanine (see Supplementary Figs. 3 and 4). These results are in marked contrast to the post-translational restoration of the wild-type amino acid residue recently observed in the mutants D1-D170H48,49, D1-E189Q48, and D1-D342N48.
Enthalpy and entropy of activation from time-resolved O2-polarography
Oxygen release transients were measured at temperatures ranging from 0 to 35 °C, for wild-type and N298A thylakoid membranes (Fig. 2a, b). The O2 transients were simulated with a diffusion model, as described in the Methods section, to obtain a time constant for O2 evolution (τO2) and a time constant for a preceding reaction step (τpre)29,45. These time constants likely are assignable to proton removal in the YZoxS3+→YZoxS3n transition (τpre) and the overall rate-limiting oxyl-radical formation of the YZoxS3n→YZS4 transition (τO2)13,50,51. The Arrhenius activation-energy analysis was applied to both sets of time constants (Fig. 2c). Here, the rate constant of oxygen evolution is of special interest. Applying Eyring’s transition state theory52,53,54, the enthalpic and entropic contribution to the free energy of activation were calculated for the rate constant of oxygen evolution (kO2 = 1/τO2) as described previously10,45; the applicability of transition state theory is discussed elsewhere55. The remarkable result from this analysis is that the N298A residue exchange increases the entropy of activation significantly, but also decreases the activation enthalpy such that the rate constants in the room temperature range remain similar to that of the wild-type PSII.
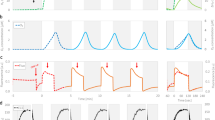
Normalized O2-evolution transients of a wild-type and b N298A PSII thylakoids at eight different temperatures (0–35 °C). c Arrhenius plot of time constants extracted from (a, b) using a diffusion model. The time constant τO2 reflects the oxygen evolution rate (τO2 = 1/kO2); τpre corresponds to proton removal preceding O2 evolution. The slope of the Arrhenius plot reveals activation energies for the preceding reactions of 269 ± 25 meV (WT) and 405 ± 24 meV (N298A), and for the oxygen-evolution step of 421 ± 22 meV (WT) and 321 ± 18 meV (N298A). For the oxygen evolution step, the Arrhenius parameters and the corresponding enthalpic and entropic contributions to the free energy of activation are shown in Table 2, for the preceding reaction in Supplementary Table 1. An overview of all fit parameters involved in simulation of the O2-evolution transients is provided in Supplementary Fig. 6. At each temperature, at least three independent data sets were obtained and averaged, before determining the time constants by simulation and fit with the diffusion model described in ref. 29; the error bars in panel c correspond to parameter uncertainties derived from the covariance matrix (calculated by the least-squares minimization algorithm). For a direct comparison of O2-evolution transients of wild-type and N298A at 20 °C, see Supplementary Fig. 5.
Static FTIR difference spectra
The static FTIR S-state difference spectra of Supplementary Fig. 7 inform on changes in the protein-water environment of the Mn4Ca-oxo cluster that are associated with the respective S-state transition and detectable after its completion. Minor spectral changes are detectable in the mid-frequency S2-minus-S1 spectrum (Supplementary Fig. 7a), which resemble alterations produced by the mutations of many other residues in the protein-water environment near the Mn4Ca-oxo cluster (see Table 1 in ref. 56). This suggests that the N298A mutation also perturbs the protein-water environment that couples to the S-state transitions, as was concluded previously34 and discussed for D61A elsewhere30. Perturbation of the protein-water environment in both PSII variants, N298A and D61A, is shown more directly in the weakly H-bonded O-H stretching region of the IR spectra (Supplementary Fig. 7b).
Comparison of the static difference spectra does not provide evidence for major modification of the S-state transitions in either N298A or D61A. The extent of spectral modifications in N298A is especially minor, also in the structure-sensitive amide I and amide II regions, rendering conformational changes of the protein environment in the vicinity of the Mn4Ca-oxo cluster unlikely.
Time-resolved IR reveals biphasicity in the O2-evolution transition of D1-N298A
PSII was driven through its reaction cycle with its four semi-stable S-states (Fig. 1a) by application of a sequence of 10 laser flashes (532 nm, ca. 5 ns halfwidth) with a 1 s interval between flashes. Flash-induced IR transients were obtained at selected wavenumbers for wild-type, N298A, and D61A; see Supplementary Figs. 13 and 14 for flash-induced transients of all four S-state transitions. To reach a noise level below 5 µOD, signal averaging of data collected for 500–5000 flash sequences applied to dark-adapted PSII core complexes was required at each wavenumber selected for investigation (Supplementary Table 2). The tuneable quantum cascade laser (QCL) laser system was able to cover the spectral range from 1650 to 1300 cm−1, but to match the noise level requirements with the available amount of PSII core complexes, gapless coverage of this spectral range was out of the question and a selection of a small number of informative wavenumbers was approached.
The wavenumbers of 1384, 1400, 1514, and 1544 cm−1 were selected for the following reasons: Based on the FTIR difference spectra of the S3→S0 transition of Supplementary Fig. 7 and results described in ref. 10, at 1384, 1514 and 1544 cm−1, comparably large contributions of the millisecond phase of oxygen evolution were expected (and indeed found); the 1400 cm−1 wavenumber was chosen to facilitate direct comparison to results obtained in refs. 28,57,58. The 1384 cm−1 and the 1400 cm−1 wavenumbers are predominantly associated with the stretching vibrations of carboxylate residues ligating the metal ions of the Mn4Ca-oxo cluster59,60,61. At 1514 cm−1 a strong contribution of the CO stretching vibrations of the redox-active tyrosine radical has been predicted62 and therefore this wavenumber was selected. However, we find that the IR transients at 1514 cm−1 do not match the expected oxidation and reduction kinetics of YZox in the various S-state transitions suggesting that prominent signals of other origin(s) overlap. At 1544 cm−1 the amide II vibrations become visible61,63, which may inform on conformational changes of the protein backbone63. Moreover, we collected IR transients at 1478 cm−1 (Supplementary Fig. 17), which reflect the oxidation state changes of the quinones at the acceptor side of PSII.
Figure 3 shows that the IR difference absorption at 700-800 ms after the excitation flash exhibits a clear flash-number dependence in all data sets. Wild-type PSII and N298A show similar period-of-four oscillating patterns, indicating that both samples are going through the S-state cycle with similar turn-over efficiency. Also, the absolute ΔA values are mostly similar, implying an undiminished amount of O2-evolving PSII in N298A. In D61A, the oscillating pattern is significantly more dampened, but still clearly discernible, indicating that the sample is cycling with a greatly reduced transition efficiency.
Data of wild-type PSII (blue), as well as PSII with point mutations D61A (green) and N298A (red) of the D1 protein subunit is shown for a 1384 cm−1, b 1400 cm−1, c 1514 cm−1, and d 1544 cm−1. Each data point was obtained by averaging the IR difference absorption between 700 and 800 ms following the nth excitation flash (n = 1,…, 10). The exciting laser flashes were spaced by 1 s.
In the following, we analyze the IR transients of the oxygen-evolution transition, S3→S4→S0, induced predominantly by the third flash applied to dark-adapted PSII (for data on other S-state transitions see Supplementary Fig. 11). The IR transients of the S3→S0 transition of wild-type PSII show a well-resolved kinetic phase with a time constant of 4.7 ms at all four probed wavenumbers (Fig. 4a–d, blue), which concurs with the O2 evolution rate determined by O2 polarography at 10 °C (see Table 1). In the D61A variant this phase is strongly retarded in the IR data (360 ms phase, Fig. 4a–d, green), suggesting that the O2-evolution rate is strongly affected by the mutation. The N298A variant, however, displays a more complex behavior. In first experiments, we collected data at 1400 cm−1, which at first glance suggested that the O2 evolution in the N298A variant is significantly slowed (Fig. 4b). This, however, would be at odds with the O2-polarography results. Further data sets at 1384, 1514, and 1544 cm−1 then revealed biphasic behavior with phases of 6.5 and 75 ms. The amplitudes of these two phases show a clear wavenumber-dependent behavior (Fig. 4e–h); at 1384 cm−1 and 1514 cm−1 the slower phase dominates, while at 1544 cm−1, the faster phase is more prominent; at 1400 cm−1 the faster phase is essentially absent. A 60 ms phase is well resolved in the WT and likely assignable to acceptor side processes, whereas the origin of the 15 ms phase in D61A remains unclear; both phases are not specific to the S3→S4→S0 transition; see Supplementary Figs. 15 and 16 and their discussion in the Supplementary Information. Simulation of the transients moreover reveal time constants of 480, 790, and 990 µs with the similar negative amplitudes for the WT, D61A, and N298A variants, which are likely assignable to the PT preceding oxygen evolution12,50,51,58 and also visible as τpre in the O2-transients (Fig. 2). Noteworthily, the PT phase is largely unaffected by both residue exchanges, aside from a moderate slowing down (1.6–2.1-fold), which previously has been seen for the D61A variant also in time-resolved O2-polarography experiments45.
a–d A deconvolution algorithm was applied to the wild-type and N298A transients of 10 consecutive excitation flashes, resulting in four transients corresponding to the “pure” S-state transitions. These are similar to the transients recorded after the 3rd saturating excitation flash (compare Supplementary Fig. 12). For the D61A variant, the original 3rd flash data is shown without deconvolution, because of uncertainties regarding the likely highly S-state dependent miss factor. Wild-type data is shown in blue, D61A in green, and N298A in red; the black lines were obtained by curve-fitting with the parameters shown in Supplementary Table 4. e–h Amplitudes corresponding to the time constants obtained from fitting the data on the left to a sum of exponentials. The bars filled in white are assigned to processes that are not specific for the oxygen-evolving S3→S0 transitions, see Supplementary Figs. 15 and 16. The time constants were determined globally by a joint fit of transients collected at several wavenumbers with the same set of time constants. Uncertainty ranges were estimated by performing fits of various subsets of the four transients, for which the mean values of the time constants and amplitude are shown here, with the error bars indicating the respective standard deviations of the amplitudes (Supplementary Figs. 8–10 and Tables 3, 4). The time constant values are indicated by arrows in (a–d). The estimated uncertainty ranges of the time constants are 4.7 ± 0.3 ms and 60 ± 35 ms (wild-type), 15 ± 8 ms and 357 ± 58 ms (D61A), 6.5 ± 0.8 ms and 75 ± 9 ms (N298A). Further details on the S-state deconvolution, curve-fitting, and estimation of average time constants and their uncertainty ranges are presented in the Methods section and in Section 4 of the Supplementary Materials. For a closer look at the millisecond kinetics at 1384 and 1544 cm−1, see also Supplementary Fig. 18.
Population of intermediate states in the S3→S4→S0 transition—a possible scenario
The time-resolved IR experiments suggest that in the S3→S4→S0 transition, instead of a single millisecond phase there are two kinetic phases—and thus two processes—resolvable in IR transients of the N298A variant. Since only the faster process coincides with appearance of O2 as detected by time-resolved O2-polarography, the slower time constant relates to a process in the oxygen-evolving transition that occurs after O2-release. In the WT-PSII, it is faster than the O2-evolution step and thus not resolvable in time-resolved experiments. In the N298A variant, however, this process is slowed down by minimally one order of magnitude and thereby becomes detectable as a separate kinetic phase. The existence of this slow process in the N298A variant also resolves the incongruity between fast O2-formation detected by time-resolved polarography and severely reduced rates of O2-evolution detected for continuous illumination at high photon fluence rates (Table 1). Since the slow process in the N298A variant occurs after O2-release, we assign it to the water deprotonation and hydroxide insertion needed to refill the vacant O5 site, finally resulting in Mn4CaO5 cluster in its S0 state. Which of these steps, water deprotonation and insertion, represents the rate-limiting bottleneck remains open as does their sequence; likely these processes proceed in a tightly coupled way.
The relation of the experimentally determined rate constants to the populations of the hypothetical intermediates is shown in Fig. 5 for the WT, D61A and N298A variant; the proposed assignment of the B→C and C→D transitions to atomistic processes is schematically shown in Fig. 6.
The populations were calculated based on the reaction scheme of (a) and the rate constants experimentally determined in the time-resolved IR experiment (k = 1/τ) for b WT PSII, c the D61A variant, and d the N298A variant; the respective IR time constants are indicated in (b–d). Each reaction step is assumed to be associated with a significant free energy change so that the backreaction can be neglected. For the WT and the D61A variant, the C intermediate likely does not accumulate to a resolvable extent because kCD is significantly greater (faster) than kBC; therefore, kCD values were arbitrarily chosen to equal 20 times the respective kBC values. The dashed violet lines indicate the time courses of produced dioxygen.
a Oxygen atom O6 is oxidized by transferring an electron to YZ; the proton bound to O6 is concomitantly transferred to D61 in a Grotthus-type mechanism. Subsequently, O6 and O5 form an O=O bond while reducing Mn4, Mn3, and Mn1 from +IV to +III. This step takes place within about 5 ms in wild-type PSII as well as in PSII containing the N298A mutation, but is severely slowed down in the D61A mutant. Here a specific mode of O-O bond formation is indicated10,43,88,89; alternatives have been discussed1,90. b A new water molecule is inserted into the Mn4CaOx cluster, and a proton is simultaneously released to the bulk, leading to the formation of the S0 state (shown in c). This step is faster than O2 release and thus “invisible” in wild-type PSII as well as in PSII containing the D61A mutation. We propose that in the N298A mutant, this step is slowed down roughly ten-fold and thus becomes visible for spectroscopic methods. The given time constants are approximate values at 10 °C. The arrangements of the non-hydrogen atoms schematically shown in (a, c) correspond to crystallographically resolved structures16,46. The structures in panel b, the location of H-atoms, and all the particle movements indicated by arrows have been deduced by structure-based computational chemistry previously10 and thus are more hypothetical than the arrangement of non-hydrogen atoms shown in (a, c).
Molecular dynamics simulations rationalize the influence of the asparagine-alanine exchange
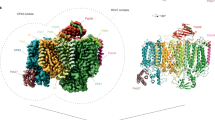
MD simulations were performed on the cryo-EM structure from the mesophilic cyanobacterium, Synechocystis sp. PCC 680364. To model the S3YZox state, the coordinates of Mn4CaO6 cluster and the C-terminus of D1 (D1-Ala 344) were taken from the crystallographically characterized PSII of Thermosynechococcus vestitus BP-146. Force-field parameters for the MnCa-oxo cluster were derived as described in the Supplementary Information. For the complete PSII dimer model immersed in a lipid-bilayer–water environment (about 106 atoms), we calculated five trajectories of 500 ns for wild-type PSII and the N298A variant. Due to the homodimeric organization of PSII, this corresponds to a total of 10 trajectories of 500 ns duration. Neglecting the first 100 ns of each trajectory, a total simulation time of 4 µs was analyzed, with selected results shown in Figs. 7 and 8. The overall structural stability of the simulated systems during the unrestrained MD simulations was proven by calculating the time evolution of different properties (see Supplementary Fig. 19).
a Water distributions and MD conformations around residue N298 (wild-type, blue) and A298 (N298A mutant, red) are represented by extracting 40 snapshots by each of the 10 independent 400 ns-long trajectories. Water wheel molecules were selected within a 3.8 Å-radius sphere centered on the center of mass of the Cα atoms of residues V185, H190, L343, A344, CP43-A386, and CP43-I398. Red and blue dots and lines represent water molecules and residues. A less frequently sampled cluster in the upper left identifies a third, alternative water binding site on the calcium, consistent with recent structural data for T. vestitus24 (see Supplementary Fig. 20). Yellow spheres indicate the five waters of the pentagon, and W3 and W4 from a cryo-EM structure (PDB ID: 7N8O, Synechocystis sp. PCC 6803)64. MD snapshots were fitted to this structure using the backbone atoms of D1-protein residues 165–210, 303–344, and CP43-protein residues 366–418. b The cavity boundaries for wild-type (blue) and N398A (red) were defined using a central 6 Å sphere centered on the water wheel, and an outer 4.5 Å-radius sphere centered on the Mn4Ca cluster; see Supplementary Information for details. c Histograms of the water occupancy for the “water wheel” within the 3.8 Å-radius spherical region (detailed in panel a) for wild-type (blue) and N298A (red). d The Mn4CaO5-6 cluster and its surrounding environment, including waters W3, W4 (calcium-bound), W26–W30 (water wheel), and selected residues. We analyzed the water exchange between the cyan 3.0 Å-radius sphere centered on the Ca and the red 3.8 Å-radius sphere centered on the water wheel cluster. Table reports the probability of water transitions from the red to the cyan sphere, estimated from transition events within consecutive 10 ps intervals.
a Wild-type (WT) PSII simulations show a strong H-bond of the Yz-–H190–N298 triad. In the N298A simulation, the backbone carbonyl of L297 partly substitutes the asparagine in the H-bond network. b Fluctuations of the H-bonds involving the epsilon nitrogen (Nε) and delta nitrogen (Nδ) of H190 in one representative monomer trajectory (i.e., M9) for both the wild-type (blue) and the N298A mutant (red); more details are reported in Supplementary Fig. 21. c Representation of key geometrical features around H190: the angle Θ between the planes formed by the rings of YZ and H190 and the distance d of the H-bond of the (Nδ) nitrogen. d Distribution of distances between the hydrogen attached to the delta nitrogen (Nδ) of H190 and N298 (wild-type, blue) or L297 (N298A mutant, red). e Distribution of the angle, Θ, between Yz and H190 in wild-type (blue) and the N298A mutant (red), reflecting the relative orientation of planes defined in panel c. f Key residues likely involved in the oxyl-radical formation step. g Distribution of E189-Ca distances. h Illustration of a hypothetical energy landscape for wild-type (blue) and N298A mutant (red), including a pre-transition state (Pre-TS) and transition state (TS). The softer profile in the N298A mutant may explain an increased entropy and decreased enthalpy of activation (H0act).
Focusing on the water molecules shown in the MD snapshots of Fig. 7a, the MD simulations reproduce most of the various water positions observed by cryo-EM64. These results are also consistent with those obtained for T. vestitus (see Supplementary Fig. 20). The latter are compared to X-ray crystallography46 and fit especially well with a recent cryo-EM structure24, which has added several transiently or partially occupied water positions (W4a, W4b, W26a). The main effect of the N298A modification may be described as a less constrained conformational space in the vicinity of this residue, corresponding to about 20% increase in local volume compared to the wild-type (Fig. 7b). By exchange of the asparagine sidechain against the smaller alanine sidechain, the water wheel (W26–W30) gains space and expands into the gained space (Fig. 7a), which is not coupled to an increase in the average number of contributing water molecules but rather associated with a moderate decrease from about 5.0 down to 4.5 (Fig. 7c and Supplementary Table 5). Thus, the wild-type water wheel may be seen as more constrained when compared to the N298A variant, possibly associated with a higher free energy of the water-wheel structure and thereby favoring the water insertion step in the wild-type PSII by a reduced free energy difference between the initial state and the transition state. This aligns with our finding that water molecules are nearly three times less likely to transition from the water wheel to the Mn4Ca-oxo cluster in the N298A mutant (Fig. 7d).
A sizeable entropic contribution to the free energy of activation of the rate-determining step of the oxygen-evolution transition (−T0ΔS‡ in Table 2) has been assigned to a high number of conformations of the H-bonded protein-water network in the surrounding of the Mn4Ca-oxo cluster and YZ, with only a specific subset of these acting as an active conformation that allows the rate-limiting step to proceed10,55. Our MD simulations illustrate and support this entropic contribution by showing pronounced fluctuations (see also Supplementary Movie 1). These include H-bond distances and protein conformations that relate directly to the reaction coordinate(s) and the likely transition state structure, as shown in Fig. 8 and summarized as follows:
-
1.
The H-bond to the second nitrogen (Nδ) of His190 to the asparagine sidechain in the WT is substituted in the N298A variant by rarer (11% probability) H-bonding to the backbone carbonyl of L297 in competition with water molecules (Supplementary Figs. 21 and 22), thereby likely increasing the conformational variability of the H190 sidechain (Supplementary Fig. 23).
-
2.
For the YZ-H190 pair, the MD simulations do not reveal changes in H-bonding itself (Supplementary Fig. 24) but rather highlight pronounced fluctuations in the relative orientation of the planes formed by their rings (Fig. 8e). We note that the relative YZ—H190 position changes upon YZ, oxidation/reduction, as observed experimentally in crystallographic snapshot data23,46 and computationally10, indicating a close relation to the respective reaction coordinate. The YZ—H190 motions are influenced by the two conformers associated with the second nitrogen of His190 (Fig. 8a and Supplementary Fig. 25), which also appear to modulate the number of water molecules populating the water wheel as defined in Fig. 7, possibly perturbing its structure and dynamics (Supplementary Fig. 26).
-
3.
The MD simulations reveal a pronounced influence on the position of the Glu189 sidechain (Fig. 8f, g). Here, the active conformation in oxyl radical and subsequent O-O bond formation likely corresponds to the occupancy peak of the WT at 3.5 Å55; this occupancy peak is not only strongly diminished but also highly “blurred” in N298A, resulting a broad plateau level, which indicates a severe destabilization of the active conformation compared to WT PSII.
Currently, it is not possible to quantify the entropic and enthalpic contribution to ΔG‡ of the rate-determining step, but the results of our MD simulations render an increased entropic and concomitantly decreased enthalpic contribution plausible. Denoting the active conformation of the protein-water network reached before the onset of the rate-limiting step in O2 formation as the pre-transition state (pre-TS), a softer energy landscape for the N298A mutant may result in a less stabilized pre-TS, thereby increasing the effective entropy of activation and decreasing the effective enthalpy of activation (Fig. 8h).
Discussion
In photosynthetic water splitting, the protein determines structure and the spatial arrangement of the catalytic metal cluster with its inorganic Mn4CaO5/6 core and YZ, a nearby redox-active tyrosine sidechain. Aside from the basic structural role of the protein, the adjacent and more remote networks of H-bonded protein-water clusters facilitate efficient water oxidation by “orchestrating” the coupling between electron transfer steps and proton movements and likely also the insertion of the substrate-water molecules. Our study addresses three types of orchestrating protein roles, illustrated by analyses of the PSII variants with residue exchanges that affect the rate constants (reciprocal time constants) of the rate-limiting step of the oxygen-evolving S3→S4→S0 transition.
Coupling of electron and proton transfer
In the rate-limiting S3→S4 transition, the electron transfer likely is coupled to Grotthus-type PT from the substrate-hydroxide bound at Mn1 to the D61 carboxylate sidechain10 (Fig. 1). Accordingly, a clear effect of the aspartate-alanine exchange in the D61A variant is expected and has previously been detected by time-resolved O2-polarography45, for the D61N variant also by time-resolved UV-vis experiments29. We can now confirm the drastic slowdown of the O2-evolution step by infrared spectroscopy. In light of the recent proposal of close coupling between S4-formation and PT to D61 in the rate-determining step of the oxygen-evolution transition (ref. 10, see also refs. 43,44), it is surprising that O2-evolution is slowed down by about two orders of magnitude but is not fully inhibited. Seemingly, the PT via the D61 sidechain is not obligatory but can be replaced by one or more alternative routes of coupling the YZ-reducing ET step to proton relocation; these are by about two orders of magnitude slower, but still allow for oxygen evolution. However, the oxygen evolution step proceeds at severely reduced transition efficiency (high miss factor, low amplitude of corresponding IR signals), explainable by competing recombination reactions during the strongly extended YZox lifetime. This illustrates the role of the protein in acceleration of the O2-formation step to a level that facilitates an average quantum efficiency of the reactions in photosynthetic water oxidation well above 80% (miss factors well below 20%). Aside from providing relevant support to previous findings25,30,45, the time-resolved IR data of the D61A variant is of importance in the present study because it facilitates comparison with the N298A variant.
Orchestrating substrate-water binding
In the N298A variant, we encountered a paradox of severely reduced O2-evolution under continuous illumination of saturating intensity but slightly accelerated O2-release for excitation with short flashes of visible light. The time-resolved infrared spectroscopy supports resolution of this paradox by a plausible scenario.
The rate-determining proton-coupled ET step in the S3→S4 formation as well as the subsequent processes of fast peroxide formation and O2-release largely proceed in the N298A variant as they do in the wild-type, aside from the shift between entropy and enthalpy of activation discussed further below. These processes create a vacancy in the Mn4Ca-oxo cluster that needs to be refilled to reach the S0-state, by binding of a hydroxide to Ca and two five-coordinate MnIII ions (Ca, Mn3 and Mn4 in Fig. 6b)16,46,65,66. This is associated with the release of a proton11,67,68 (Fig. 1a) stemming from coupled H2O deprotonation and hydroxide insertion into the vacant metal-cluster site. Despite its complexity, the Mn4Ca-cluster reconstruction by water deprotonation and hydroxide insertion is faster than the rate limiting S3→S4 transition in the wild-type PSII but not so in the N298A variant, where it is slowed down to about 75 ms (time-constant value). Consequently, only in the IR transients of the N298A variant the metal cluster reconstruction by hydroxide insertion is reflected as a separate kinetic phase. The time constant of 75 ms for completion of the S3→S0 transition is in line with the strongly diminished continuous-illumination O2-evolution in the N298A variant.
The N298 residue might affect the reactions in the oxygen-evolving S3→S4→S0 transitions in different ways. First, its sidechain H-bonds the protonated Nδ-H of the pivotal histidine-190 (H190). The complete loss of this H-bond likely would affect the proton affinity (pKa) of the H190-Nε severely, concomitantly the redox potential for YZ oxidation, and consequently the rate constant of the rate-determining proton-coupled ET in the S3→S4 transition. Experimentally, this is not observed, which our MD simulations can explain by the formation of an alternative H-bond to the peptide carbonyl of the neighboring residue (L297), which is present throughout a small fraction of the MD simulation time.
Second, the N298 sidechain lines the cavity of the so-called water wheel, a cluster of about 5 water molecules, which based on crystallographic data have been suggested to provide the H2O molecule for insertion into the Mn4Ca-oxo cluster, in the S2→S3 transition and likely also in the S4→S0 transition17,19,46. Our findings suggest a drastic slowdown of the water insertion step in the S4→S0 transition of the N298A variant, thereby providing experimental support for a crucial role of the water-wheel cluster.
Since extensive water movements proceed easily in the picosecond time domain (see Supplementary Movie 1), time constants in the millisecond domain do not relate to the water mobility itself, but rather to the Gibbs free energies of the initial state and the relevant transition state. Upon replacement of asparagine with a short alanine sidechain in the N298A variant, the water wheel gains space and expands towards the A298 sidechain (Fig. 7a). We hypothesize that, thereby, the free energy of the initial state of the water insertion step is lowered and consequently the activation energy increased, resulting in a major slow-down. However, also more complex explanations may apply.
The here-reported mutagenetic deceleration of later steps in the oxygen-evolving transition paves the road for more detailed spectroscopic and structural studies of these later steps, which previously were hidden behind the rate-determining YZoxS3→YZS4 transition. Further experimental studies on these kinetically hidden transitions will be highly valuable, or even indispensable, on the way towards a complete understanding of all aspects of photosynthetic water oxidation. Mutagenic deceleration may be an especially promising strategy but also in wild-type PSII, later steps might be resolvable46.
Balancing entropy and enthalpy of activation
The absolute rate constant value of the rate-determining step on photosynthetic oxygen evolution cannot be understood without considering a pronounced “entropic slow-down” likely assignable to multiple conformations of the H-bond protein-water network surrounding YZ and the Mn4Ca-oxo cluster of PSII10,55. We now detect a significantly increased entropic and reduced enthalpic contribution to the free energy of activation in the N298A variant that is caused by the exchange of a single residue only. The here-reported MD simulations with a force field developed for cyanobacterial PSII in the YZoxS3 state provide evidence for an increased flexibility in key protein sidechains in the N298A variant and reduced probability to reach the transition state conformation, in line with an increased entropic contribution to the free-energy of activation of the YZoxS3→YZS4 transition. The increased flexibility suggests less stringent energetic constraints regarding nuclear movements, including the ones required to reach the transition state, which may explain the reduced enthalpy of activation.
Relevance for inorganic (electro)catalysts
On first glance, it may appear far-fetched to compare the processes at the protein-bound Mn4Ca complex of PSII to inorganic catalyst materials. However, also the active phase of electrocatalyst materials based on first row transition metals typically comprises redox-active transition metals interconnected by bridging oxides and hydroxides69,70,71, likely with extensive H-bonding of bridging and terminal HxO to nearby water molecules72; for Mn oxide catalysts, also a beneficial role of Ca ions has been demonstrated73,74. These materials are often discussed and addressed in computational studies by assuming simultaneous oxidation and deprotonation with focus on adsorption energies in various oxidation/protonation states, without considering the reaction kinetics (activation energies) of proton-coupled electron transfer, O2-release, and substrate-water insertion (adsorption) in more detail. Previously, we showed that the slowest step in the reaction cascade of PSII water oxidation likely is the oxyl-radical formation by coupling the oxidation of a metal-bound hydroxide with its deprotonation in a Grotthus-type PT with an interesting relay functionality of a second metal ion10. In spite of its evolutionary optimization, this step is still associated with a sizeable free-energy of activation of about 580 meV (time constant around 2 ms). Now we provide evidence that in a genetic PSII variant, also the deprotonation-coupled binding (adsorption) of a substrate hydroxide may be associated with a sizeable free-energy of activation (about 670 meV, time constant around 70 ms). The magnitude of these two activation energies points to the need to consider increasingly more kinetic limitations (entropy and enthalpy of activation) in electrocatalyst materials, which also in these materials could relate to multiple conformations and fluctuations of H-bond networks in vicinity of the catalytic site.
The rate-limiting processes can be addressed especially well for the Mn4Ca-oxo cluster of PSII and its surrounding H-bonded water clusters because (i) experimentally, the transition between four semi-stable intermediates can be traced individually in time-resolved experiments and (ii) the well-defined active-site structure and adjacent water clusters favor detailed computational studies. Therefore, we propose to see the biological water oxidation as a model system that can inspire and direct the investigation of possibly analogous rate-limiting processes in electrocatalyst materials, and thereby the knowledge-guided development of new materials with diminished kinetic limitations.
Methods
Construction and propagation of mutants
The D1-D61A mutation has been described previously40. The D1-N298A mutation was created by changing the wild-type AAC codon of N298 to GCC in the psbA-2 gene of Synechocystis sp. PCC 680375. The mutation-bearing plasmid was transformed into a Synechocystis strain that lacks all three psbA genes and has a hexa-histidine tag (His-tag) at the C-terminus of CP4776. Cultures were maintained and propagated as described previously30. To verify the integrity of the mutant cultures that were harvested for the purification of thylakoid membranes and PSII core complexes, the DNA sequence of the relevant portion of the psbA-2 gene was sequenced after PCR amplification of genomic DNA purified from an aliquot of each harvested culture. Purification of genomic DNA and PCR amplification of a 1380-bp fragment containing the entire psbA-2 gene was performed as described previously75. No trace of the wild-type codon was detected in any of the mutant cultures.
Purification of thylakoid membranes
Harvested cells were suspended in 1.2 M betaine, 10% (v/v) glycerol, 50 mM MES-NaOH (pH 6.0), 5 mM CaCl2, 5 mM MgCl2, 1 mM benzamidine, 1 mM ε-amino-n-caproic acid, 1 mM phenymethylsulfonyl fluoride, and 0.05 mg/mL DNAse I, then broken in a glass bead homogenizer30,77. After removal of debris and unbroken cells by low-speed centrifugation, the resulting thylakoid membranes were concentrated by ultracentrifugation (20 min at 186,000 × g in a Beckman Ti45 rotor) and suspended in TM buffer [1.2 M betaine, 10% (v/v) glycerol, 50 mM MES-NaOH (pH 6.0), 20 mM CaCl2, 5 mM MgCl2] to a concentration of 1.0–1.5 mg of Chl/mL. The resulting membranes were either flash-frozen in liquid nitrogen or used immediately for the purification of PSII core complexes.
Purification of PSII core complexes
Freshly prepared thylakoid membranes were extracted with n-dodecyl β-D-maltoside (at concentrations of 1 mg of Chl/mL and 1% (w/v) detergent) with gentle stirring in darkness for 10 min30,77. After unextracted material was removed by centrifugation (12 min at 35,300 × g in a Beckman JA-17 rotor), the supernatant was loaded onto a 40 mL, 5 cm diameter column of Ni-NTA superflow affinity resin (Qiagen, Inc., Valencia, CA) that had been equilibrated with PSII buffer [1.2 M betaine, 10% (v/v) glycerol, 50 mM MES-NaOH (pH 6.0), 20 mM CaCl2, 5 mM MgCl2, 0.03% (w/v) n-dodecyl β-D-maltoside]. After the column was washed with four bed volumes of PSII buffer at 5 mL/min, it was eluted with four bed volumes of PSII buffer containing 50 mM histidine at 5 mL/min. The eluted PSII core complexes were brought to 1 mM EDTA and concentrated by ultrafiltration. The purified PSII core complexes [in 1.2 M betaine, 10% (v/v) glycerol, 50 mM MES-NaOH (pH 6.0), 20 mM CaCl2, 5 mM MgCl2, 50 mM histidine, 1 mM EDTA, and 0.03% (w/v) n-dodecyl β-D-maltoside] were aliquoted, frozen in liquid N2, and stored at 80 K.
Mass spectrometry
PSII core complexes were extracted of lipids and pigments with methanol and ethyl acetate, then denatured with 5.2% lithium lauryl sulfate, 172 mM Tris-HCl (pH 8.0), and 40 mM dithiothreitol78. Denatured PSII subunits were separated by SDS-PAGE in gels containing 20% acrylamide and 6 M urea26,78. Excised D1 subunits were further denatured with urea (8 M), reduced with DTT (20 mM), alkylated with indoleacetic acid (50 mM), and buffer-exchanged into ammonium bicarbonate (50 mM). Subunits containing D1-D61A were digested with chymotrypsin for 16 h at 37 °C. Subunits containing D1-N298A were digested with trypsin for 16 h at 37 °C followed by partial digestion with chymotrypsin (2 h at 37 °C). Digested solutions were desalted with C18 Zip Tips (Waters Corporation), dried, and resuspended in 20 µL 0.1% formic acid. The resulting peptides were separated by nano-HPLC chromatography (EASY-nLC 1200, Thermo Scientific). Peptide samples (5 µL) were loaded onto a trap column (Thermo Scientific Acclaim PepMap C18, 75 μm × 2 cm) with a flow rate of 10 μl/min for 3 min, then separated on an analytical column (Acclaim PepMap C18, 75 μm × 25 cm) with a linear gradient of 3–37% buffer B (80% acetonitrile, 0.1% formic acid) in buffer A (0.1% formic acid) at a flow rate of 300 nL/min over 180 min. The column temperature was maintained at 45 °C. The nano-HPLC was coupled online with an electrospray tandem mass spectrometer (Orbitrap Fusion Tribrid Mass Spectrometer that was equipped with an EASY-Spray ion source). The electrospray voltage was 2.2 kV. The mass spectrometer was operated in data-dependent mode to switch automatically between MS and MS/MS acquisitions. MS spectra (m/z 375–1500) were acquired with a mass resolution of 60 K. High energy collisional dissociation (HCD) fragmentation was employed for recording MS/MS spectra. Raw data were processed and analyzed with MaxQuant (Version 2.1.4.0) with an in-house database containing the D1 protein and its mutants. Data from two preparations each of D1-D61A and D1-N298A core complexes were analyzed, each in comparison with wild-type PSII.
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE79 partner repository with the dataset identifier PXD068038.
Steady-state O2 evolution measurements
Initial light-saturated rates of O2 evolution were measured at 25.0 °C in a solution of 1 M sucrose, 50 mM MES-NaOH (pH 6.5), 25 mM CaCl2, and 10 mM NaCl that was held in a water-jacketed cell that was outfitted with a Clark-type oxygen electrode. For measurements, concentrated PSII core complexes (5 μg Chl) were diluted into 1.6 mL of assay buffer containing 1 mM potassium ferricyanide and 0.4 mM 2,6-dichloro-p-benzoquinone (the latter was purified by sublimation and diluted into the assay buffer immediately before the PSII core complexes were added). Red-filtered illumination was provided by two Dolan-Jenner (Woburn, MA) Model 180 fiber optics illuminators equipped with light guides. Error ranges were obtained as a standard deviation of 4–8 experiments.
Time-resolved O2 polarography measurements
An aliquot of thylakoid membranes (equivalent to 10 µg of chlorophyll) was suspended in buffer (150 mM NaCl, 25 mM MES, 1 M betaine, 5 mM MgCl2 and 5 mM CaCl2, adjusted to pH 6.2 with NaOH) and placed onto a bare platinum and silver-ring electrode29, which was then centrifuged in a swing-out rotor at 10,000 × g for 10 min (at 4 °C). The polarization voltage of –0.95 V required for O2-reduction at the platinum electrode was provided by a custom-built potentiostat and was switched on 15 s prior to the first excitation flash. For each of 80 flashes spaced by 900 ms, the current signal was recorded for a total of 500 ms (20 ms before and 480 ms after the flash). Slow drift contributions to the current signals were suppressed by a first-order high-pass filter (time constant of 100 ms); its influence was considered in simulation (fit) of the transients. Sample excitation was achieved by saturating flashes (613 nm, 40 µs flash duration) from a red high-performance light-emitting diode (LED) operated at a maximal current density of around 150 A. The sample temperature was monitored by a miniature sensor within the sample buffer and controlled by Peltier elements.
A numerical diffusion model, inter alia, taking into account O2 diffusion through the sample as well as the rate of O2 consumption at the electrode, was used to accurately determine the rate constant of O2 formation, as detailed elsewhere29.
O2 polarography experiments were performed at 8 distinct temperatures between 0 and 35 °C for wild-type PSII and for the N298A variant. At each temperature, at least three independent data sets were obtained and averaged before determining the time constants with the diffusion model; the given error is the one estimated by the fitting algorithm. The time constants were subjected to Arrhenius analysis (Fig. 2c), allowing for the determination of activation energy (Ea) and pre-exponential factor (A) by linear regression. The enthalpy and entropy of activation (ΔH‡ and ΔS‡) were calculated from Ea and A by applying Eyring’s (and Polanyi’s) transition state theory52,53,54, using the equations provided as Supplementray Equations. The error estimations for Ea and A shown in Table 2 were obtained from the least-squares optimization process; the provided range of uncertainty for Ea corresponds to the 1σ confidence interval of the slope value of the linear regression. As ΔH‡ is calculated by subtracting a constant number from Ea, the error of ΔH‡ is equal to the error of Ea. The 1σ confidence interval of ΔG‡ was determined from the error range of the time constant τO2 at 20 °C. The error of TΔS‡ is the sum of the errors of ΔG‡ and ΔH‡.
Static FTIR difference spectroscopy
PSII core complexes were exchanged into a buffer containing 40 mM sucrose, 10 mM MES-NaOH (pH 6.0), 5 mM CaCl2, 5 mM NaCl, and 0.06% (w/v) n-dodecyl β-D-maltoside, concentrated to approx. 3 mg of Chl/mL, mixed with 1/10 volume of fresh 100 mM potassium ferricyanide, spread into the center 13 mm of a 25 mm BaF2 window, then dried lightly with N2 gas30,80. The lightly dried samples were rehydrated to 95% relative humidity by placing six 1 μL drops of a solution of 40% (v/v) glycerol around the periphery of the sample81. A second window was placed over the first with a thin O-ring spacer in between. Sealed samples were equilibrated in the FTIR sample compartment at 0 °C in darkness for 1.5 h, illuminated with 6 pre-flashes, then dark-adapted for 30 min30,80. For each sample, the absorbance at 1657 cm−1 (amide I band) was 0.6–1.1. Spectra were recorded with a Bruker Vertex 70 spectrometer (Bruker Optics, Billerica, MA) that held a pre-amplified, midrange D317 photovoltaic MCT detector (Kolmar Technologies, Inc., Newburyport, MA). Actinic flashes (∼20 mJ/flash, ∼7 ns fwhm) were provided by a frequency-doubled Q-switched Nd:YAG laser (BRIO, Quantel USA, Bozeman, MT). After dark adaptation, samples were given six flashes at 13 s intervals. Two transmission spectra were recorded before the first flash and one transmission spectrum was recorded starting 0.33 s after the first and subsequent flashes (each transmission spectrum consisted of 100 scans). The 0.33 s delay was included to allow the oxidation of QA•─ by the ferricyanide. Difference spectra of the successive S-state transitions (e.g., Sn+1–minus–Sn difference spectra), were obtained by dividing the transmission spectrum obtained after the nth flash by the transmission spectrum obtained before the nth flash, then converting the ratio to units of absorption. The background noise level and the stability of the baseline were obtained by dividing the second pre-flash transmission spectrum by the first and converting the ratio to units of absorption (these spectra are labeled dark-dark in each figure—these are control difference spectra obtained in the presence of the sample but without a flash being given). The sample was then dark-adapted for 30 min and the cycle was repeated. For each sample, the illumination cycle was repeated 19 times. The spectra of 12–26 samples were averaged.
Time-resolved single-frequency IR difference spectroscopy
PSII core complexes were left in the storage buffer but were concentrated to approximately 3 mg of Chl/ml and mixed with a 1/10 volume of fresh 100 mM potassium ferricyanide. About 10 µl of the sample solution was pipetted onto each of five 25 mm CaF2 plates. The CaF2 plates were prepared beforehand with a thin ring of vacuum grease and a 15 µm PTFE spacer. A second CaF2 plate was placed on top of the first, sealing the sample in between. The samples were mounted onto an x-y-movable sample holder in a sample compartment, which was continuously flushed with dry air and kept at 10 °C. Per sample plate 39 individual spots were measured, resulting in a total of 195 sample spots. Prior to the first measurement the sample was illuminated with 2 pre-flashes and subsequently given an hour for dark adaptation. After being subjected to a flash sequence (6 s without excitation, followed by 10 sequential saturating flashes with 1 s flash spacing), each spot was given an hour to dark adapt before being measured again. The 10th excitation flash was about three times stronger than the first nine flashes, which allows for the estimation and correction of a flash-induced heat artifact as described previously10,57. The excitation flashes were provided by a frequency-doubled Q-switched Nd:YAG laser (Minilite II, Amplitude Laser, Inc., Milpitas, CA). The IR source was a QCL covering the wavenumber range of 1650 – 1300 cm−1 (MIRcat, Daylight Solutions, Inc., San Diego, CA). The transmitted IR signal was measured with a 10 MHz pre-amplified MCT detector (Vigo Systems, Poland) and a second, identical detector was used as reference to improve the overall signal-to-noise (S/N). The intensity of the excitation flashes was measured by a power meter and a photodiode was used to determine the timing of the flashes. Data processing and analysis was performed in Python 3.7. An artifact signal of unknown origin was found in the millisecond region of the transients; as the same artifact was also recorded by the reference detector, we were able to correct for this artifact.
For analysis of the S3→S0 transition of wild-type PSII and the N298A variant, the flash-induced IR transients were deconvolved into transients corresponding to the individual S-state transitions, in a similar manner as e.g., described in refs. 10,82. For this an initial starting population of 90% S1 and 10% S0 were assumed. The miss factor of each dataset was determined by simulating the flash-dependency of the IR difference signals and was estimated to be 10–15% for wild-type PSII and 12–18% for N298A. A deconvolution of the D61A data was not (satisfactorily) possible due to the strongly retarded behavior of this variant.
To analyze the kinetics, the transients were fit to a sum of exponentials:
For each PSII variant, the time constants (τi) were determined globally over all four wavenumbers, while the amplitudes (Ai) and the offset (y0) were left to vary freely. The parameters were optimized by a least-squares algorithm. The error of the parameters was estimated by performing fits of multiple subsets of the four transients and calculating the standard deviation. For more details see Section 4 in the Supporting Information.
Classical molecular dynamics simulations
As initial structure for our simulations we have considered the cryo-EM structure of Synechocystis photosystem II resolved at 1.93 Å resolution (Protein Data Bank ID 7N8O)64.
MD simulations were performed using the GROMACS package83. The dimeric complex was solvated in a water box with dimensions 31.3 × 20.9 × 15.3 nm and embedded into a POPC lipid bilayer, following the general scheme described in refs. 84,85. All simulated systems were composed of approximately 1,000,000 atoms.
In total, ten classical MD simulations were performed to investigate the wild-type protein and the N298A variant. Five unrestrained 500 ns simulations were run for each system in the NPT ensemble with different starting velocities (referred to as WT-1, WT-2, WT-3, WT-4, WT-5, N298A-1, N298A-2, N298A-3, N298A-4, and N298A-5). Additional details on the methodology, including system setup, parameterization, and simulation protocols, are available in the Supplementary Information (Molecular Dynamics section).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The time-resolved IR data obtained for 10 consecutive excitation flashes (applied to wild-type PSII, N298A and D61A) are available at Zenodo86 [https://doi.org/10.5281/zenodo.15789664]. The mass spectrometry data are available via ProteomeXchange with identifier PXD068038. Source data are provided with this paper.
Code availability
Python code (Python 3.7.16) was used to correct the time-resolved IR data for S-state starting populations and cycle inefficiency (miss factor) and to fit the data globally to a sum of exponentials. This code is analogous to code deposited previously (https://doi.org/10.5281/zenodo.7682034). Standard GROMACS packages were used for the MD simulations, which are generally available—see also ref. 83.
References
Cox, N., Pantazis, D. A. & Lubitz, W. Current understanding of the mechanism of water oxidation in photosystem II and its relation to XFEL data. Annu. Rev. Biochem. 89, 795–820 (2020).
Junge, W. Oxygenic photosynthesis: history, status and perspective. Q. Rev. Biol. 52, E1 (2019).
Shevela, D., Kern, J. F., Govindjee, G. & Messinger, J. Solar energy conversion by photosystem II: principles and structures. Photosynth. Res. 156, 279–307 (2023).
Vinyard, D. J. & Brudvig, G. W. Progress toward a molecular mechanism of water oxidation in photosystem II. Annu. Rev. Phys. Chem. 68, 101–116 (2017).
Dau, H. & Zaharieva, I. Principles, efficiency, and blueprint character of solar-energy conversion in photosynthetic water oxidation. Acc. Chem. Res. 42, 1861–1870 (2009).
Kok, B., Forbush, B. & McGloin, M. Cooperation of charges in photosynthetic O2 evolution - I. A linear four-step mechanism. Photochem. Photobiol. 11, 457–475 (1970).
Luzzolino, L., Dittmer, J., Dörner, W., Meyer-Klaucke, W. & Dau, H. X-ray absorption spectroscopy on layered photosystem II membrane particles suggests manganese-centered oxidation of the oxygen-evolving complex for the S0-S1, S1-S2, and S2-S3 transitions of the water oxidation cycle. Biochemistry 37, 17112–17119 (1998).
Dau, H., Liebisch, P. & Haumann, M. X-ray absorption spectroscopy to analyze nuclear geometry and electronic structure of biological metal centers—potential and questions examined with special focus on the tetra-nuclear manganese complex of oxygenic photosynthesis. Anal. Bioanal. Chem. 376, 562–583 (2003).
Pantazis, D. A. The S3 state of the oxygen-evolving complex: overview of spectroscopy and XFEL crystallography with a critical evaluation of early-onset models for O–O bond formation. Inorganics 7, 55 (2019).
Greife, P. et al. The electron–proton bottleneck of photosynthetic oxygen evolution. Nature 617, 623–628 (2023).
Dau, H. & Haumann, M. Eight steps preceding O-O bond formation in oxygenic photosynthesis—a basic reaction cycle of the photosystem II manganese complex. Biochim. Biophys. Acta 1767, 472–483 (2007).
Haumann, M. et al. Photosynthetic O2 formation tracked by time-resolved x-ray experiments. Science 310, 1019–1021 (2005).
Klauss, A., Haumann, M. & Dau, H. Alternating electron and proton transfer steps in photosynthetic water oxidation. Proc. Natl Acad. Sci. USA 109, 16035–16040 (2012).
Chernev, P., Aydin, A. O. & Messinger, J. On the simulation and interpretation of substrate-water exchange experiments in photosynthetic water oxidation. Photosynth. Res. 162, 413–426 (2024).
Boussac, A., Sellés, J. & Sugiura, M. Kinetics of reformation of the S0 state capable of progressing to the S1 state after the O2 release by photosystem II. Photosynth. Res. 163, 5 (2025).
Kern, J. et al. Structures of the intermediates of Kok’s photosynthetic water oxidation clock. Nature 563, 421–425 (2018).
Ibrahim, M. et al. Untangling the sequence of events during the S2→ S3 transition in photosystem II and implications for the water oxidation mechanism. Proc. Natl Acad. Sci. USA 117, 12624–12635 (2020).
Kaur, D. et al. Proton exit pathways surrounding the oxygen evolving complex of photosystem II. Biochim. Biophys. Acta 1862, 148446 (2021).
Hussein, R. et al. Structural dynamics in the water and proton channels of photosystem II during the S2 to S3 transition. Nat. Commun. 12, 6531 (2021).
Sirohiwal, A. & Pantazis, D. A. Functional water networks in fully hydrated photosystem II. J. Am. Chem. Soc. 144, 22035–22050 (2022).
Doyle, M. D. et al. Water networks in photosystem II using crystalline molecular dynamics simulations and room-temperature XFEL serial crystallography. J. Am. Chem. Soc. 145, 14621–14635 (2023).
Hussein, R. et al. Evolutionary diversity of proton and water channels on the oxidizing side of photosystem II and their relevance to function. Photosynth. Res. 158, 91–107 (2023).
Li, H. et al. Oxygen-evolving photosystem II structures during S1–S2–S3 transitions. Nature 626, 670–677 (2024).
Hussein, R. et al. Cryo–electron microscopy reveals hydrogen positions and water networks in photosystem II. Science 384, 1349–1355 (2024).
Service, R. J., Hillier, W. & Debus, R. J. Evidence from FTIR difference spectroscopy of an extensive network of hydrogen bonds near the oxygen-evolving Mn4Ca cluster of photosystem II involving D1-Glu65, D2-Glu312, and D1-Glu329. Biochemistry 49, 6655–6669 (2010).
Pokhrel, R., Service, R. J., Debus, R. J. & Brudvig, G. W. Mutation of lysine 317 in the D2 subunit of photosystem II alters chloride binding and proton transport. Biochemistry 52, 4758–4773 (2013).
Kuroda, H. et al. Proton transfer pathway from the oxygen-evolving complex in photosystem II substantiated by extensive mutagenesis. Biochim. Biophys. Acta 1862, 148329 (2021).
Okamoto, Y., Shimada, Y., Nagao, R. & Noguchi, T. Proton and water transfer pathways in the S2→ S3 transition of the water-oxidizing complex in photosystem II: time-resolved infrared analysis of the effects of D1-N298A mutation and NO3– substitution. J. Phys. Chem. B 125, 6864–6873 (2021).
Dilbeck, P. L. et al. The mutation D1-D61N in Synechocystis sp. PCC 6803 allows the observation of pH-sensitive intermediates in the formation and release of O2 from photosystem II. Biochemistry 51, 1079–1091 (2012).
Debus, R. J. Evidence from FTIR difference spectroscopy that D1-Asp61 influences the water reactions of the oxygen-evolving Mn4CaO5 cluster of photosystem II. Biochemistry 53, 2941–2955 (2014).
Ghosh, I. et al. Insights into proton-transfer pathways during water oxidation in photosystem II. J. Phys. Chem. B 123, 8195–8202 (2019).
Aydin, A. O. et al. Probing substrate water access through the O1 channel of Photosystem II by single site mutations and membrane inlet mass spectrometry. Photosynth. Res. 163, 28 (2025).
Chrysina, M., de Mendonça Silva, J. C., Zahariou, G., Pantazis, D. A. & Ioannidis, N. Proton translocation via tautomerization of Asn298 during the S2–S3 state transition in the oxygen-evolving complex of photosystem II. J. Phys. Chem. B 123, 3068–3078 (2019).
Nagao, R., Ueoka-Nakanishi, H. & Noguchi, T. D1-Asn-298 in photosystem II is involved in a hydrogen-bond network near the redox-active tyrosine YZ for proton exit during water oxidation. J. Biol. Chem. 292, 20046–20057 (2017).
Kuroda, H., Kodama, N., Sun, X.-Y., Ozawa, S. -i & Takahashi, Y. Requirement for Asn298 on D1 protein for oxygen evolution: analyses by exhaustive amino acid substitution in the green alga Chlamydomonas reinhardtii. Plant Cell Physiol. 55, 1266–1275 (2014).
Retegan, M. et al. A five-coordinate Mn(IV) intermediate in biological water oxidation: spectroscopic signature and a pivot mechanism for water binding. Chem. Sci. 7, 72–84 (2016).
Narzi, D., Bovi, D. & Guidoni, L. Pathway for Mn-cluster oxidation by tyrosine-Z in the S2 state of photosystem II. Proc. Natl Acad. Sci. USA 111, 8723–8728 (2014).
Amin, M. et al. Proton-coupled electron transfer during the S-state transitions of the oxygen-evolving complex of photosystem II. J. Phys. Chem. B 119, 7366–7377 (2015).
Wang, J., Askerka, M., Brudvig, G. W. & Batista, V. S. Crystallographic data support the carousel mechanism of water supply to the oxygen-evolving complex of photosystem II. ACS Energy Lett. 2, 2299–2306 (2017).
Chu, H.-A., Nguyen, A. P. & Debus, R. J. Amino acid residues that influence the binding of manganese or calcium to Photosystem II. 1. The lumenal interhelical domains of the D1 polypeptide. Biochemistry 34, 5839–5858 (1995).
Hundelt, M., Hays, A. M., Debus, R. J. & Junge, W. Oxygenic photosystem II: the mutation D1-D61N in synechocystis sp. PCC 6803 retards S-state transitions without affecting electron transfer from YZ to P680+. Biochemistry 37, 14450–14456 (1998).
Hundelt, M., Hays, A.-M. A., Debus, R. J. & Junge, W. in Photosynthesis: Mechanisms and Effects (ed. Garab, G.) 1387–1390 (Springer, 1998).
Siegbahn, P. E. M. Water oxidation mechanism in photosystem II, including oxidations, proton release pathways, O―O bond formation and O2 release. Biochim. Biophys. Acta 1827, 1003–1019 (2013).
Allgöwer, F., Gamiz-Hernandez, A. P., Rutherford, A. W. & Kaila, V. R. Molecular principles of redox-coupled protonation dynamics in photosystem II. J. Am. Chem. Soc. 144, 7171–7180 (2022).
Bao, H. & Burnap, R. L. Structural rearrangements preceding dioxygen formation by the water oxidation complex of photosystem II. Proc. Natl Acad. Sci. U. S. A 112, E6139–E6147 (2015).
Bhowmick, A. et al. Structural evidence for intermediates during O2 formation in photosystem II. Nature 617, 629–636 (2023).
Siegbahn, P. E. Mechanisms for proton release during water oxidation in the S2 to S3 and S3 to S4 transitions in photosystem II. Phys. Chem. Chem. Phys. 14, 4849–4856 (2012).
Shimada, Y. et al. Post-translational amino acid conversion in photosystem II as a possible origin of photosynthetic oxygen evolution. Nat. Commun. 13, 4211 (2022).
Kitajima-Ihara, T. et al. Fourier transform infrared and mass spectrometry analyses of a site-directed mutant of D1-Asp170 as a ligand to the water-oxidizing Mn4CaO5 cluster in photosystem II. Biochim. Biophys. Acta 1861, 148086 (2020).
Gerencser, L. & Dau, H. Water oxidation by photosystem II: H2O−D2O Exchange and the influence of pH support formation of an intermediate by removal of a proton before dioxygen creation. Biochemistry 49, 10098–10106 (2010).
Zaharieva, I., Dau, H. & Haumann, M. Sequential and coupled proton and electron transfer events in the S2 → S3 transition of photosynthetic water oxidation revealed by time-resolved X-ray absorption spectroscopy. Biochemistry 55, 6996-–7004 (2016).
Eyring, H. The activated complex in chemical reactions. J. Chem. Phys. 3, 107–115 (1935).
Evans, M. G. & Polanyi, M. Some applications of the transition state method to the calculation of reaction velocities, especially in solution. Trans. Faraday Soc. 31, 875–894 (1935).
Mortimer, R. G. & Eyring, H. Elementary transition-state theory of the Soret and Dufour effects. Proc. Natl Acad. Sci. USA 77, 1728–1731 (1980).
Dau, H. & Greife, P. Applicability of transition state theory to the (proton-coupled) electron transfer in photosynthetic water oxidation with emphasis on the entropy of activation. Inorganics 11, 389 (2023).
Debus, R. J. Alteration of the O2-producing Mn4Ca cluster in photosystem II by the mutation of a metal ligand. Biochemistry 60, 3841–3855 (2021).
Mäusle, S. M. et al. Activation energies for two steps in the S2 → S3 transition of photosynthetic water oxidation from time-resolved single-frequency infrared spectroscopy. J. Chem. Phys. 153, 215101 (2020).
Sugie, K., Kato, Y., Nagao, R. & Noguchi, T. Mechanism of the oxygen-evolving process in the water-oxidizing complex of photosystem II, as revealed by time-resolved infrared spectroscopy. J. Phys. Chem. B 129, 6172–6187(2025).
Debus, R. J. FTIR studies of metal ligands, networks of hydrogen bonds, and water molecules near the active site Mn4CaO5 cluster in Photosystem II. Biochim. Biophys. Acta 1847, 19–34 (2015).
Nakamura, S. & Noguchi, T. Quantum mechanics/molecular mechanics simulation of the ligand vibrations of the water-oxidizing Mn4CaO5 cluster in photosystem II. Proc. Natl Acad. Sci. U. S. A 113, 12727–12732 (2016).
Noguchi, T. & Sugiura, M. Analysis of flash-induced FTIR difference spectra of the S-state cycle in the photosynthetic water-oxidizing complex by uniform 15N and 13C isotope labeling. Biochemistry 42, 6035–6042 (2003).
Berthomieu, C., Hienerwadel, R., Boussac, A., Breton, J. & Diner, B. A. Hydrogen bonding of redox-active tyrosine Z of photosystem II probed by FTIR difference spectroscopy. Biochemistry 37, 10547–10554 (1998).
Barth, A. Infrared spectroscopy of proteins. Biochim. Biophys. Acta 1767, 1073–1101 (2007).
Gisriel, C. J. et al. High-resolution cryo-electron microscopy structure of photosystem II from the mesophilic cyanobacterium, Synechocystis sp. PCC 6803. Proc. Natl Acad. Sci. USA 119, e2116765118 (2022).
Pal, R. et al. S0-state model of the oxygen-evolving complex of photosystem II. Biochemistry 52, 7703–7706 (2013).
Krewald, V. et al. Metal oxidation states in biological water splitting. Chem. Sci. 6, 1676–1695 (2015).
Junge, W., Haumann, M., Ahlbrink, R., Mulkidjanian, A. & Clausen, J. Electrostatics and proton transfer in photosynthetic water oxidation. Philos. Trans. R. Soc. B 357, 1407–1418 (2002).
Dau, H. & Haumann, M. The manganese complex of photosystem II in its reaction cycle—basic framework and possible realization at the atomic level. Coord. Chem. Rev. 252, 273–295 (2008).
Dau, H. et al. The mechanism of water oxidation: from electrolysis via homogeneous to biological catalysis. ChemCatChem 2, 724–761 (2010).
Bergmann, A. et al. Unified structural motifs of the catalytically active state of Co(oxyhydr)oxides during the electrochemical oxygen evolution reaction. Nat. Catal. 1, 711–719 (2018).
Song, F. et al. Transition metal oxides as electrocatalysts for the oxygen evolution reaction in alkaline solutions: an application-inspired renaissance. J. Am. Chem. Soc. 140, 7748–7759 (2018).
Mattioli, G., Giannozzi, P., Amore Bonapasta, A. & Guidoni, L. Reaction pathways for oxygen evolution promoted by cobalt catalyst. J. Am. Chem. Soc. 135, 15353–15363 (2013).
Wiechen, M., Zaharieva, I., Dau, H. & Kurz, P. Layered manganese oxides for water-oxidation: alkaline earth cations influence catalytic activity in a photosystem II-like fashion. Chem. Sci. 3, 2330–2339 (2012).
González-Flores, D. et al. Electrosynthesis of biomimetic manganese-calcium oxides for water oxidation catalysis - atomic structure and functionality. ChemSusChem 9, 379–387 (2016).
Chu, H.-A., Nguyen, A. P. & Debus, R. J. Site-directed photosystem II mutants with perturbed oxygen-evolving properties. 1. Instability or inefficient assembly of the manganese cluster in vivo. Biochemistry 33, 6137–6149 (1994).
Debus, R. J. et al. Does histidine 332 of the D1 polypeptide ligate the manganese cluster in photosystem II? An electron spin echo envelope modulation study. Biochemistry 40, 3690–3699 (2001).
Tang, X.-S. & Diner, B. A. Biochemical and spectroscopic characterization of a new oxygen-evolving photosystem II core complex from the cyanobacterium synechocystis PCC 6803. Biochemistry 33, 4594–4603 (1994).
Kashino, Y., Koike, H. & Satoh, K. An improved sodium dodecyl sulfate-polyacrylamide gel electrophoresis system for the analysis of membrane protein complexes. Electrophoresis 22, 1004–1007 (2001).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2024).
Kim, C. J. & Debus, R. J. Evidence from FTIR difference spectroscopy that a substrate H2O molecule for O2 formation in photosystem II is provided by the Ca ion of the catalytic Mn4CaO5 cluster. Biochemistry 56, 2558–2570 (2017).
Noguchi, T. & Sugiura, M. Flash-induced FTIR difference spectra of the water oxidizing complex in moderately hydrated photosystem II core films: Effect of hydration extent on S-state transitions. Biochemistry 41, 2322–2330 (2002).
Isgandarova, S., Renger, G. & Messinger, J. Functional differences of photosystem II from synechococcus elongatus and spinach characterized by flash induced oxygen evolution patterns. Biochemistry 42, 8929–8938 (2003).
Van der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Narzi, D., Coccia, E., Manzoli, M. & Guidoni, L. Impact of molecular flexibility on the site energy shift of chlorophylls in Photosystem II. Biophys. Chem. 229, 93–98 (2017).
Narzi, D. & Guidoni, L. Structural and dynamic insights into Mn4Ca cluster-depleted Photosystem II. Phys. Chem. Chem. Phys. 23, 27428–27436 (2021).
Mäusle, S. M., Debus, R. J. & Dau, H. Supplementary data for “Three rate-determining protein roles in photosynthetic O2-evolution addressed by time-resolved experiments on genetically modified photosystems”. Zenodo https://doi.org/10.5281/zenodo.15789664 (2025).
de Lichtenberg, C., Kim, C. J., Chernev, P., Debus, R. J. & Messinger, J. The exchange of the fast substrate water in the S2 state of photosystem II is limited by diffusion of bulk water through channels – implications for the water oxidation mechanism. Chem. Sci. 12, 12763–12775 (2021).
Siegbahn, P. E. O.-O. bond formation in the S(4) state of the oxygen-evolving complex in photosystem II. Chemistry 12, 9217–9227 (2006).
Narzi, D., Capone, M., Bovi, D. & Guidoni, L. Evolution from S3 to S4 state of the oxygen evolving complex in Photosystem II monitored by QM/MM dynamics. Chem. Eur. J. 24, 10820–10828 (2018).
Lubitz, W., Pantazis, D. A. & Cox, N. Water oxidation in oxygenic photosynthesis studied by magnetic resonance techniques. FEBS Lett. 597, 6–29 (2023).
Acknowledgements
S.M.M., R.A., P.S.S., and H.D. gratefully acknowledge financial support by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) provided to the collaborative research center on Protonation Dynamics in Protein Function (SFB 1078, project A4/Dau) and under Germany’s Excellence Strategy—EXC 2008—390540038-UniSysCat. G.P., C.D.S., D.N. and L.G. acknowledge the CINECA award under the ISCRA initiative (project IsB29_PSIICMD, D.N.), for the availability of high-performance computing resources and support, and would like to especially thank Paola Alberigo for her valuable assistance in facilitating and extending the computational resources needed for this work. This work was supported by the Department of Energy, Office of Basic Energy Sciences, and Division of Chemical Sciences Grant DE-SC0005291 to R.J.D. We thank Dr. Quanqing Zhang, IIGB Proteomics Manager at UC Riverside, for the protein mass spectrometry analyses.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
S.M.M. collected, analyzed, and conceptualized the time-resolved IR data; preliminary data was collected by P.S.S. G.P., C.D.S. and D.N. performed the computational research. R.A. collected and analyzed the time-resolved O2 polarography data. R.J.D. constructed the site-directed PSII mutants, purified PSII complexes, collected and analyzed the FTIR data. H.D., R.J.D. and L.G. designed and conceptualized the experimental and computational research. H.D., S.M.M., R.J.D., G.P., D.N. and L.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maria Drosou, Dimitrios Pantazis, David Vinyard and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mäusle, S.M., Parisse, G., Assunção, R. et al. Three rate-determining protein roles in photosynthetic O2-evolution addressed by time-resolved experiments on genetically modified photosystems. Nat Commun 16, 9515 (2025). https://doi.org/10.1038/s41467-025-64513-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64513-9