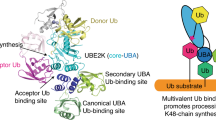
Abstract
Beyond the canonical K48-linked homotypic polyubiquitination for proteasome-targeted proteolysis, K11/K48-branched ubiquitin (Ub) chains are involved in fast-tracking protein turnover during cell cycle progression and proteotoxic stress. Here, we report cryo-EM structures of human 26S proteasome in a complex with a K11/K48-branched Ub chain. The structures revealed a multivalent substrate recognition mechanism involving a hitherto unknown K11-linked Ub binding site at the groove formed by RPN2 and RPN10 in addition to the canonical K48-linkage binding site formed by RPN10 and RPT4/5 coiled-coil. Additionally, RPN2 recognizes an alternating K11-K48-linkage through a conserved motif similar to the K48-specific T1 binding site of RPN1. The insights gleaned from these structures explain the molecular mechanism underlying the recognition of the K11/K48-branched Ub as a priority signal in the ubiquitin-mediated proteasomal degradation.
Similar content being viewed by others
Introduction
Ubiquitin (Ub) is implicated in a myriad of biological processes owing to its ability to form multiple types of polymeric conjugates through one (or more) of the seven lysine (K) residues or the N-terminal methionine (M1). In addition to the well-studied homotypic Ub chains, atypical Ub chain topologies have been identified. These include heterotypic mixed-linkage Ub chains, encompassing more than one type of isopeptide bond linkage along a linear chain, and complex heterotypic branched Ub chains, encompassing Ub molecules in which more than one lysine is involved in the chain formation resulting in multiple branching points within the same Ub chain1. Branched Ub chains account for 10–20% of Ub polymers2,3. Among the different types of branched Ub chains, K11-K48-linked Ub branching is the best characterized: it is preferentially recognized by the ubiquitin-proteasome system (UPS) for substrate degradation under specific cellular conditions, including cell cycle progression in early mitosis and proteotoxic stress that requires proteostasis maintenance4. The K11/K48-branched Ub chains mediate the timely degradation of mitotic regulators, misfolded nascent polypeptides, and pathological Huntingtin variants4,5,6. The K11/K48-branched Ub chains adopt different topologies in a cellular context-dependent manner4. However, the mechanism by which the proteasomal Ub receptors recognize the K11/K48-branched Ub chain and differentiate it from its homotypic counterparts remains unknown.
The 26S proteasome recognizes ubiquitinated substrates through three constitutive Ub/Ub-like (UBL) receptors – RPN1, RPN10, and RPN13 – located within the 19S regulatory particle (RP) subcomplex7,8,9,10,11. RPN10 binds to Ub through two α-helical ubiquitin interacting motifs (UIMs) tethered to the N-terminal Willebrand factor A (VWA) domain within the 19S RP7,12. RPN13 binds to Ub through its N-terminal pleckstrin receptor for ubiquitin (PRU) domain tethered to the flexible C-terminus of RPN2 while the C-terminal part of RPN13 forms a deubiquitinase adaptor (DEUBAD) domain to recruit ubiquitin C-terminal hydrolase L5 (UCHL5; also known as UCH37)8,9,13,14,15,16. The PRU and DEUBAD domains of RPN13 are connected by a disordered linker, providing accessibility to a large conformational space for substrate recognition17. RPN1 binds to Ub at the T1 site, formed by a three-helix bundle within its proteasome/cyclosome (PC) domain10 or at its N-terminal section11. The enhanced binding of K11/K48-branched Ub chains was reported for isolated RPN1 and RPN10, which could account for the accelerated proteasomal degradation of substrates marked with K11/K48-branched Ub chains6,18. Additionally, Ub binding of all three proteasomal Ub receptors involves auxiliary factors such as shuttling factors and deubiquitinating enzymes (DUBs) that further contribute to Ub chain specificity10,14,16,19,20. In particular, the RPN13-associated UCHL5 preferentially recognizes and removes K11/K48-branched Ub chains from proteasomal substrates21,22. Apart from UCHL5, the proteasome can reversibly bind to another auxiliary DUB, USP14. Several cryo-EM studies have revealed the molecular basis of USP14 (or its yeast homolog Ubp6) in the context of proteasome functions23,24,25,26,27,28. Unlike UCHL5, which exhibits K48-linkage-specific debranching activity, USP14 is proposed to be mainly K63-linkage specific29,30 or to catalyze the removal of supernumerary ubiquitin chains on a substrate en bloc31.
In addition to the canonical proteasomal Ub receptors, there is good biochemical and genetic evidence to suggest the presence of one or more hitherto unidentified cryptic Ub receptors within the 26S proteasome10,32. While Ub binding to RPT5 and Sem1/Dss1 may account for the cryptic Ub receptor, how they bind to Ub remains poorly understood33,34. Recently, two potential Ub binding sites have been suggested: the α5-subunit within the 20S core particle (CP) subcomplex and RPN2 within the 19S RP. The latter is a paralog of the RPN1 that is structurally homologous to the T1 Ub binding site of RPN135. UBL binding to RPN2 was observed by chemical cross-linking, but the molecular details remain elusive36. Importantly, whether the cryptic proteasomal Ub binding site is also implicated in recognizing branched Ub chains remains unknown. Despite the wealth of biochemical and structural understanding of the UPS, chiefly deduced from recent cryo-EM studies24,28,37,38,39,40,41,42,43,44,45,46,47, the molecular basis of how branched Ub chains are recognized and processed by the 26S proteasome remains poorly understood.
In this work, we report the cryo-EM structure of a tetra-ubiquitin with a K11/K48-branching point at the proximal Ub to form a well-defined tripartite binding interface with the 19S RP. The cryo-EM structures provide structural evidence to indicate RPN2 as a Ub receptor recognizing the K48-linkage extending from the K11-linked Ub, forming a unique alternating K11-K48-linkage stemming from the proximal Ub that helps position the K11-linked Ub branch into a groove formed by RPN2 and neighboring proteasomal subunits. The structural insights from this study illustrate the versatility of the 26S proteasome in decoding the complex Ub chain signaling needed to maintain proteostasis.
Results
Biochemical and structural characterization of a K11/K48-branched ubiquitin chain bound to the 26S proteasome complex
We reconstituted a functional complex of the human 26S proteasome in a complex with a polyubiquitinated substrate and the auxiliary proteins RPN13 and UCHL5. The substrate consisted of the intrinsically disordered residues 1-48 of S. cerevisiae Sic1 protein (hereafter Sic1PY)28 with a single lysine residue (K40) that serves as an anchoring point for ubiquitination by an engineered Rsp5 E3 ligase (hereafter Rsp5-HECTGML; Materials and Methods; Fig. 1a). While wild-type (WT) Rsp5 generates K63-linked Ub chains48, Rsp5-HECTGML has been engineered to generate K48-linked Ub chains, which was confirmed by Western blotting using Ub linkage-specific antibodies49. To exclude the possibility of generating a K63-linked Ub chain, a Ub variant (K63R) of which the lysine at position 63 is mutated into an arginine was used to produce the polyubiquitinated Sic1PY (Sic1PY-Ubn) (Supplementary Fig. 1). Dual fluorescence labelling for Sic1PY (Alexa647) and Ub (fluorescein) was introduced to enable simultaneous detection of the two, which helps distinguish substrate proteolysis from deubiquitination. Sic1PY-Ubn stably bound to the enzymatically active human 26S proteasome (Supplementary Fig. 2). The crude Sic1PY-Ubn reaction product was further fractionated by size-exclusion chromatography (SEC) to enrich the medium-length Ub chains (n = 4-8) to be efficiently processed by the 26S proteasome (Fig. 1b and Supplementary Fig. 2c). Contrary to the expectation of forming predominantly K48-linked homotypic Ub chains, however, Lbpro* Ub clipping2 and intact mass spectrometry (MS) analysis revealed the presence of doubly ubiquitinated (12.6%) and triply ubiquitinated (3.6%) Ub in addition to singly ubiquitinated Ub (41.8%), which is clear evidence of the formation of branched Ub chains (Fig. 1c). To identify the linkage types of the poly-Ub chains, we employed MS-based Ub absolute quantification (Ub-AQUA)50,51 to demonstrate that the SEC-enriched poly-Ub chains contained almost equal amounts of K11- and K48-linked Ub with a minor population of K33-linked Ub (Fig. 1d, Supplementary Table 1 and Supplementary Fig. 3).
a Construct design and labeling strategy of a polyubiquitinated Sic1PY. b 26S proteasome-mediated proteolysis of the polyubiquitinated Sic1PY detected by SDS-PAGE separation followed by fluorescence imaging of the exogenously labeled AlexaFluor647. c Evidence of Ub chain branching by Lbpro* digestion followed by intact MS analysis. The difference of 114 Da is indicative of a single ubiquitination site on the target Ub. The relative populations in percentage are derived from peak integration of the individual peaks. The experiments were carried in triplicate with the original data points shown in open circles and the error bars indicating the standard deviations. d MS Ub-AQUA/PRM quantification of the abundances of the individual Ub chain linkage types. K63 is omitted due to the use of the Ub(K63R) variant. The experiments were carried in triplicate with the original data points shown in open circles and the error bars indicating the standard deviations. Cryo-EM maps of the reconstituted proteasomal complex in the EB state (e), and substrate-processing ED state (f), Well-resolved cryo-EM map of the K11/K48-branched Ub4 chain can be identified in both states. g Segmented cryo-EM map of the proteasome-bound Ub4 in the ED state is superimposed with the atomic model to illustrate the clear definition of the K11/K48-branched Ub chain topology. The positions of the UbI, UbII, UbIII and UbIV are indicated in Roman numbers. The UbII density was lowpass filtered to enhance the readability.
UCHL5 is a proteasome-associated DUB that preferentially processes K11/K48-branched Ub chains21. The DUB activity of UCHL5 is activated upon binding to RPN138. The addition of UCHL5 to the reconstitute proteasomal complex may help capture K11/K48-branched Ub chains. However, UCHL5 could also disassemble the proteasome-bound Ub chains. To minimize the processing of the Sic1PY-Ubn by the endogenous UCHL5, we added an excess amount of preformed RPN13:UCHL5 complex with an alanine mutation for the catalytic cysteine of UCHL5(C88A). The presence of UCHL5, RPN13 and Sic1PY-Ubn in the reconstituted complex was confirmed by native gel electrophoresis combined with Western blotting and fluorescence imaging (Supplementary Fig. 2b). The ternary functional complex formation was additionally confirmed by negative staining electron microscopy (NSEM), which showed additional EM densities on the 19S RP of the reconstituted proteasomal complex when compared to the apo 26S proteasome (Supplementary Fig. 2d).
We successfully determined four cryo-EM structures of the reconstituted proteasomal complex after extensive classification and focused refinements (Supplementary Figs. 4, 5, and Supplementary Table 2 and Supplementary Movie 1). These structures resembled the previously reported substrate-free (apo) EA state, an Ub chain-bound EA, EB, and substrate-engaged ED state of human proteasome45. Informed by the Ub-AQUA data that indicated the most abundant linkage types (Fig. 1d), we generated the atomic models of a K11/K48-branched tetra-Ub (Ub4) chain bound to the EB and ED states (hereafter EB:Ub4 and ED:Ub4, respectively) (Fig. 1e, f), which showed good per-residue Q-scores52 for the individual Ubs (Supplementary Fig. 6). The individual Ubs adopted unique spatial distributions with respect to the neighboring Ub to enable the modeling of the most probable isopeptide bond linkages when considering the conformational landscape accessible to Ub53 (Supplementary Fig. 7). For the Ub chain-bound EA state, the most distal Ub could not be resolved in the cryo-EM map, but the core K11/K48-branched tri-Ub remained well-defined (hereafter EA:Ub3). Additionally, the ED:Ub4 state exhibited extra density extending from the proximal Ub into the base of the 19S RP, corresponding to the SicPY substrate poised for en bloc deubiquitination before being translocated into the 20S core particle (CP) for degradation (Fig. 1f, g).
The identification of substrate-free and Ub chain-bound 26S complexes in various conformational states allowed for the assessment of any preference for substrate binding by the RP2-CP or RP-CP complexes, as well as the examination of the cooperativity between opposing RPs within the RP2-CP. Classifications focusing on the 19S RP subcomplexes showed that in the majority of RP2-CP complexes (82% of those analyzed), at least one RP was found in an active (ED-like) substrate-processing conformation (Supplementary Fig. 8). On the other hand, only 19% of the analyzed RP-CP particles exhibited the 19S RP in an active conformation. Reciprocal analysis of classes representing the two 19S RPs in the RP2-CP complexes indicated that, generally, if one RP adopted the active conformation, the RP on the opposite side of the RP2-CP complex (RP’) adopted a resting (EA- or EB-like) conformation. Additionally, ~12% of the RP2-CP particles were identified in which both RPs were bound with the Ub chain and adopted an active conformation. It should be noted, however, that the stringent filtering of the particle image sets involved in the cryo-EM single particle reconstruction may affect the number of particles in each observed population; therefore, this analysis may not be the full representation of the true conformational landscape of the complex proteasome under native conditions.
Despite the addition of an excess amount of UCHL5(C88A), we did not observe EM density that could be assigned to RPN13-bound UCHL5 nor the DEUBAD/PRU domains of RPN13 in any of the four functional states. This may be due to the intrinsic dynamics of RPN13 and UCHL5 on the proteasome17,54,55. Indeed, in the absence Sic1PY-Ubn, we identified by chemical cross-linking MS (XL-MS) analysis several contacts between UCHL5, RPN13 and proteasomal proteins that are spatially distant from each other (Supplementary Fig. 9a and Supplementary Data 1). In the presence of Sic1PY-Ubn, the XLs of UCHL5(C88A) and RPN13 were primarily linked to Ub, suggesting the displacement of UCHL5 by Sic1PY-Ubn (Supplementary Fig. 9b and Supplementary Data 2).
Structural mapping of the observed XLs onto the cryo-EM structures of the four different states was carried out using XMAS56 (Supplementary Fig. 10). The results showed multiple interactions between Ub and several proteasomal subunits and the long XLs (Cα-Cα distances exceeding 35 Å) were generally associated the Ubs. These long XL distances associated with Ubs were likely artifacts, due to the intrinsic difficulty to unambiguously distinguish between XL peptides with identical sequences but from different Ubs (Supplementary Fig. 10). Some XLs associated with the 19S RP subunits were found to be larger than 35 Å only found in the EA:Ub3 and EB:Ub4 states but not in the ED:Ub4 state, reflecting the conformational changes between these functional states (Supplementary Fig. 10a). When the XLs were mapped onto the crystal structures of UCHL5-RPN13-Ub57,58, all XLs were within 35 Å, indicating the robustness of our XL-MS analysis when applied to a simpler quaternary structure (Supplementary Fig. 11). The intrinsic dynamics of UCHL5, RPN10 UIM motifs (Fig. 1a), and RPN13 potentially allow the complex to target a wide range of ubiquitinated substrates. Nonetheless, such functional dynamics also makes it intrinsically challenging to define their precise locations on the proteasome59,60.
Structural basis of K11/K48-branched Ub chain bound to the 26S proteasome with multivalency
The proximal Ub in the EA:Ub3, EB:Ub4 and ED:Ub4 all adopted a conserved pose similar to several previously reported RPN11:Ub structures (Supplementary Fig. 12a). The K48 of the proximal Ub was linked to another Ub in contact with the RPT4/5 coiled-coil (CC) adjacent to RPN10 (Fig. 1e, f). The relative orientation of the RPT4/5 CC-bound K48-linked Ub was similar to previously reported cryo-EM densities of distal proteasome-bound Ub (Supplementary Fig. 12b)45,46, in line with the previously reported role of RPT5 in polyUb-binding by the proteasome33. The conformational variability of the distal K48-linked Ub observed in other Ub-receptor structures may explain the less resolved density of this Ub in our cryo-EM map of the complex (Supplementary Fig. 12c). Despite its dynamics, the density of the K48-linked Ub exhibited a clear Ub fold – a signature thumb and palm shape – enabling the modeling of the interactions between the K48-linked Ub and proteasomal subunits (Supplementary Fig. 6 and 7).
In addition to the canonical K48-linked Ub, we identified an additional Ub linked to the proximal Ub through a K11 linkage to form a K11/K48-branched Ub chain topology (Fig. 1f). The K11-linked Ub nicely tucked into a multi-subunit groove encompassing RPN2, RPN8, RPN9, RPN10, and RPN11. The ability of this Ub binding groove to bind K11-linked Ub was further confirmed by a control cryo-EM analysis of which Sic1PY was modified by a K11-only homotypic Ub chain. A well-resolved cryo-EM density was identified in the Ub binding groove but not in the RPT4/5 CC region (Supplementary Fig. 13). Compared to previously reported K11-linked di-Ub structures, proteasome-binding required substantial conformation rearrangements of the K11-linked Ub relative to the proximal Ub to dock into the identified Ub binding groove (Supplementary Fig. 12d)59,61. For simplicity, the proximal Ub, the K48-linked Ub, and the K11-linked Ub that formed the core K11/K48-branched Ub chain are hereafter defined as UbI, UbII, and UbIII, respectively (Fig. 1e–g).
In addition to the K11/K48-branched Ub3 core, we resolved the fourth Ub (defined as UbIV).) in the EB:Ub4 and ED:Ub4 states. UbIV was linked to UbIII through a K48-linkage, resulting in an alternating K11-K48-linkage stemming from UbI (Fig. 1e–g, and Supplementary Fig. 6 and 7). To the best of our knowledge, such an alternating linkage has not been documented in the literature. Collectively, our cryo-EM analysis revealed a multivalent binding mode between the human 26S proteasome and the K11/K48-branched Ub chain encompassing the canonical K48-link Ub binding site and the identified binding site for the K11-linked Ub chain.
Alternating K11-K48-linkage along the K11-linked branch of the Ub chain allows for an extensive interaction interface with the proteasomal 19S RP
Close examination of the molecular details of the interactions between the K11/K48-branched Ub chain and the proteasome revealed an extensive Ub interaction network involving RPN2, RPN8, RPN9, RPN10, RPN11, and the RPT4/5 CC in the EA:Ub3, EB:Ub4 and ED:Ub4 states. The multi-subunit binding interface sequestered a bulk part of the solvent-accessible surface area (∆SASA) of the K11-K48-linked Ub4 (Supplementary Table 3). The intermolecular hydrogen bonds involved C-terminal residues of UbI, particularly in the EA:Ub3 and EB:Ub4 states (Fig. 2a, b panel I and Supplementary Table 4). This is in line with the general functional role of RPN11 in removing the Ub chains en bloc from target substrates62. In both the EB:Ub4 and ED:Ub4 states, the I44 and I36 hydrophobic patches of UbI made extensive contacts with the hydrophobic lining of the active site of RPN11 (Fig. 2a, b panel I). In contrast, UbII made limited contacts with RPN10 and the RPT4 RPT5 CC primarily through hydrogen bonding. Notably, between the EB:Ub4 and ED:Ub4 states, the RPT4/5 CC underwent a pronounced lever motion over which UbII hopped over to contact opposite sides of the RPT4/5 CC: RPT5 was involved in hydrogen bonding with UbII in the EB:Ub4 state while RPN10 was involved in hydrogen bonding with UbII in the ED:Ub4 state (Fig. 2a, b panel II, Supplementary Tables 4 and 5).
Cryo-EM structures of the 19S RP in the EB:Ub4 state (a) and substrate-processing ED:Ub4 state (b) bound with the K11/K48-branched Ub4 with the same coloring scheme as in Fig. 1e. The expanded views of the individual Ubs in the two functional states are shown on the right panels with the individual Ub shown in cartoon representations and the proteasomal proteins in surface representations. The identities of the interacting residues in Ub are labeled in black. The identities of the proteasomal residues involved in forming salt bridges and/or hydrogen bonds with Ubs are indicated with matching colors according to the scheme shown on the left panels. Circled insets on the upper left corners of individual Ub panels illustrate the Ub binding interfaces outlined by black lines. The I44, I36 and D58 patches on the Ubs are colored hot pink, blue and brown, respectively. Proteasomal residues involved in contacting the I44, I36 and D58 patches are colored in the same way.
Regarding the identified K11-Ub chain binding groove, UbIII formed limited hydrogen bonds with RPN2 and RPN11 in all three functional states; UbIII made additional hydrogen bonds with RPN8 only in the ED:Ub4 state (Fig. 2a, b panel III and Supplementary Table 6). In contrast to the sparse interactions formed by UbII and UbIII, UbIV made extensive contacts with RPN2 and RPN9 (Fig. 2a, b panel IV, Fig. 3, Supplementary Tables 4 and 5). Among all the four Ubs, UbIV contributed to the second largest ∆SASA (after the proximal ubiquitin, UbI), accounting for 28% and 25% of the overall ∆SASA occupied by the entire K11-K48-linked Ub4 in the EB:Ub4 and ED:Ub4 complexes, respectively (Supplementary Table 3). Given the sparse contacts made by the K11-linked UbIII with RPN2, the primary role of UbIII appeared to restrict the conformational space available to UbIV for positioning it to the new Ub binding groove formed by RPN2 and RPN9. As a result, the unique alternating K11-K48-Ub linkage provided two anchoring points – UbI and UbIV – with optimal conformational complementarity with the 26S binding sites (Fig. 2). Collectively, the K11/K48-branched Ub4 contributed to a large ∆SASA of 3077 and 2938 Å2 within the 19 RP of the EB:Ub4 and ED:Ub4 states, respectively (Supplementary Table 3).
a Structural alignment of RPN1 (orchid) and RPN2 (fuchsia) focusing on the helical Ub binding motifs (highlighted in blue and hot pink, respectively). The alternating K11-K48-linked di-Ub is colored following the same scheme as in Fig. 1e. The structure of RPN1 is taken from the PDB entry 6J2N. The structure of the RPN2 in complex with the K11-K48-linked di-Ub is taken from the ED:Ub4 state in this study. b Expanded view of the K48-linked di-Ub in complex with RPN2. The conserved E538 and D564 are highlighted by red ovals. The inter-chain hydrogen bonds and salt bridges are shown in dashed white lines and dashed green lines, respectively. c LOGO representation of the sequence conservation of the K48-linkage binding motif shared by RPN2 and RPN1. The conserved acidic residues are indicated in red, and the other conserved hydrophobic residues are highlighted in black below the sequence alignment. They are numbered according to the sequence of RPN2.
RPN2 contains a conserved ubiquitin-binding motif
Our cryo-EM structures underscored the pivotal role of RPN2 in recognizing the K11-linked Ub branch with alternating K11-K48-linkages, namely UbIII-UbIV. Close examination of the structural motifs pertinent to Ub recognition revealed a conserved K48-linked di-Ub recognition motif within RPN2. The motif consists of three α-helices within the PC domain of RPN2, resembling the canonical T1 Ub receptor binding motif of RPN110 (Fig. 3a). Alignment of 23 RPN1 and 21 RPN2 sequences from different model organisms revealed two highly conserved negatively charged residues involved in Ub binding, corresponding to E538 and D564 in human RPN2 (Fig. 3 and Supplementary Fig. 14–15).
In the EB:Ub4 state, E538 of RPN2 formed a salt bridge with R72 of UbIV, while D564 formed a salt bridge with H68 of UbIV. The interaction was consolidated by R572 whose side-chain was hydrogen bonded to the backbone carbonyl of G47; concomitantly, Q537 made contacts with R72 and L73 of UbIV. The conserved hydrophobic surface encompassing A526, I529, V533, G534, Y535, A565, L566, and S569 made extensive contacts with L8, T9, I44, V70, L73, and R74 of UbIV (Fig. 3c). In the ED:Ub4 state, the interaction network between the RPN2 and UbIV underwent some rearrangement (Supplementary Tables 4 and 5). Specifically, E538 of RPN2 formed a salt bridge with R74 of UbIV, while D564 of RPN2 switched its salt bridge register to K6 of UbIV. An additional salt bridge was formed between H68 of UbIV and the conserved E568 of RPN2 in the ED:Ub4 complex, while the Q537 of RPN2 switched its hydrogen bonds to the backbone amide group of L73 instead of R72 in the EB:Ub4 state. The hydrophobic interactions between RPN2 and UbIV were generally conserved between the EB:Ub4 and ED:Ub4 states, with the latter exhibiting an additional contribution from the conserved L570 of RPN2.
In contrast to the extensive interactions between RPN2 and UbIV, RPN2 only made few interactions with UbIII. These included a hydrogen bond between Q540 of RPN2 and Q49 of UbIII, which existed in the EB:Ub4 state but not in ED:Ub4 state. Additionally, the hydrophobic interactions were formed mostly between the T539 of RPN2 and I44, A46, G47, and K48 of UbIII. Overall, the amphipathic nature of the di-Ub binding mode observed in RPN2 was similar but not identical to the one used by RPN1 (Supplementary Fig. 15). Importantly, the identified di-Ub binding motif of RPN2 could account for the previously proposed cryptic Ub receptor on the 19S RP in addition to RPN1, RPN10, and RPN13.
To investigate the cellular role of RPN2, we performed CRISPR genome editing to modify the PSMD1 gene, which encodes RPN2 (Supplementary Fig. 16a). We first disrupted PSMD1 in HEK293T cells by electroporating Cas9 ribonucleoproteins, achieving an initial percentage of insertion or deletion (% indel) and knockout (KO) score of 72% and 59% respectively on day 3 post editing, indicating effective gene disruption. However, % indel and KO score declined to 39% and 12% on day 21, suggesting that unedited or heterozygous cells outcompeted homozygous KO cells (Supplementary Fig. 16b). This observation indicates that PSMD1 is essential for cell viability and that complete loss of RPN2 impairs proteostasis and is not tolerated. To further examine this, we performed single-cell sorting to isolate PSMD1 KO clones. DNA sequencing and immunoblotting confirmed that all viable clones were heterozygous KOs (Supplementary Figs. 16b and 16c). Real-time cell proliferation (RTCA) analysis showed that the heterozygous KO clone proliferated more slowly than parental cells, supporting the essential role of RPN2 in promoting cell growth (Supplementary Fig. 16h).
In addition to gene KO, we introduced targeted point mutations into the endogenous PSMD1 locus using CRISPR knock-in (KI). We designed homology-directed repair templates (HDRT) encoding a fragment of PSMD1 cDNA (exon 13 to 15), the double-mutations E538K and D564K, an egfp reporter gene and a synthetic poly-adenosine sequence (Supplementary Fig. 16d). Successful KI enabled targeted integration of the HDRT at PSMD1 exon 13, bypassing the endogenous transcription. Co-expression of EGFP allowed enrichment of KI cells and isolation of homozygous mutant clones as validated by genomic PCR, Sanger sequencing and flow cytometry (Supplementary Fig. 16e–g). RTCA revealed that the double-mutant exhibited reduced growth rates compared to wild-type KI cells (Supplementary Fig. 16i).
To assess the functional impact of these mutations, we treated the mutant cells with nocodazole (for cell cycle arrest) without and with the proteasome inhibitor MG132 and analyzed the accumulation of proteasome substrates by immunoblotting. Cyclin A, cyclin B1, and Nek2 have previously been reported to carry K11/K48-branched ubiquitin chains during cell cycle arrest6. Without MG132 treatment, a slightly increased accumulation of cyclin A and cyclin B1 was observed for 2X KI with respect to WT KI; the accumulation was less prominent for Nek2. With MG132 treatment, more pronounced accumulation was observed for cyclin A, cyclin B1 and Nek2 for 2X KI with respect to KI WT (Supplementary Fig. 17a–c). The same trend was observed for the overall ubiquitination pool (Supplementary Fig. 17d). The effects were much less obvious in the heterozygous KO relative to untreated control cells, suggesting that a single copy of WT RPN2 was sufficient to maintain the proteostasis under the experimental conditions. Together, our gene-editing results demonstrate that RPN2 is essential for cell proliferation (Supplementary Fig. 16). Substitution of key residues in the RPN2 groove disrupts recognition and processing of ubiquitinated substrates, resulting in imbalanced proteostasis.
Substrate binding triggers a cascade of conformational changes in the 19S RP
The 19S RP exists in many functional states with distinct conformational arrangements of the individual subunits. Our cryo-EM data revealed four distinct functional states that resemble the previously reported conformational states of the human 26S proteasome, namely the EA, EB, and ED states45 (Supplementary Fig. 18). These functional states bound to Ub3 or Ub4 to form the EA:Ub3, EB:Ub4, and ED:Ub4 state (Fig. 4 and Supplementary Fig. 19). The initial substrate binding to the apo EA state did not induce discernable conformational changes: the overall conformation of the 19S RP did not change from the apo EA state to the EA:Ub3 state. The EB:Ub4 state underwent subtle but significant conformational changes relative to the EA:Ub3 state, especially for RPN1, RPN2 and the RPT4/5 CC (Fig. 4a). These subunit motions were accompanied by an overall tilting motion of the 19S RP lid, resulting in a pronounced displacement of the center of mass (COM) of RPN2 by 11 Å (Supplementary Fig. 19a). Such a motion opened up the space between the 19S lid and the oligosaccharide/oligonucleotide binding (OB) ring of the ATPase RPT subunits, thereby facilitating substrate access to the AAA+ ATPase entry pore. Finally, the ED:Ub4 state underwent the most substantial conformational changes in the entire 19S RP lid, with the most notable being the lever motion of the RPT4/5 CC leading to a switch of UbII binding mode (Fig. 4b). Furthermore, the CP gate opened up in this state upon the appropriate alignment of the AAA+ ring, the OB ring, and the α-subunit ring of the 20S CP, thereby allowing substrate passage into the proteolytic chamber for degradation (Fig. 4c and Supplementary Fig. 19b), which is the hallmark of the ED state45.
a Structural comparison of the 19S RP in the apo EA state, EA:Ub3 state, EB:Ub4 state, and ED:Ub4 state following the same rendering scheme as in Fig. 1. The structures of the individual domains are purposely low-pass filtered to a resolution of 15 Å by ChimeraX according to their atomic models to highlight the large domain motions and CP gate alignment without the interferences of the finer details. For the Ub-bound states, the conformations of RPN1, RPN2 and RPT4/5 in the preceding states are outlined in dashed lines to highlight the rigid-body motions associated with the functional state transitions. The missing UbIV in the EA:Ub3 state is indicated by an orange dashed line. b All 19S RP subunits but RPT1/2, RPT3/6 and RPT4/5 are shown in transparent surfaces to highlight the lever motions of the individual CCs, especially for the transition of the RPT4/5 CC from the EB:Ub4 state to the ED:Ub4 state. c CP gate opening for substrate passage into the proteolytic chamber in the ED:Ub4 state. The seven subunits of the α-ring are color-ramped from white to dark indigo. Expanded views of the latching loop of RPN2 (residues 811-875; left panel) and the RPN11 Ins-1 loop (residues 76-88; right panel) in the EA:Ub3 state (d), EB:Ub4 state (e), and ED:Ub4 state (f). These intrinsically disordered loops are sufficiently rigidified and visible to cryo-EM analysis only in the ED:Ub4 state.
In addition to the rigid-body motions of the individual subunits, the EB:Ub4 state exhibited partially resolved EM density of the intrinsically disordered residues 811-875 of RPN2 as a result of its interaction with the RPT1/2 CC. This loop became resolved in the ED:Ub4 state. The structure of the ED:Ub4 state illustrated how the intrinsically disordered loop latched onto RPN1 near the RPT1/2 CC to help maintain the inter-subunit geometrical arrangements (Fig. 4d–f and Supplementary Fig. 19c). Additionally, the Ins-1 loop of RPN11, pertinent to the activation of DUB activity of RPN1163, also became resolved when transiting from the EA:Ub3 state to the ED:Ub4 state, facilitated by the loop hairpin (residues 54–74) of RPT5 in the EB:Ub4 state (Fig. 4d–f and Supplementary Fig. 19d). Collectively, the structural insights gleaned from the four distinct functional states underscored the dynamic interplay and crosstalk between the individual subunits within the 19S RP and 20S CP.
Discussion
In this work, we have generated a K11/K48-branched Ub chain-modified Sic1PY using an engineered chimeric Rsp5 Ub E3 ligase, Rsp5-HECTGML49. The substrate could be efficiently recognized and processed by the human 26S proteasome (Fig. 1 and Supplementary Fig. 2). We determined the cryo-EM structures of the K11/K48-branched Ub chains with the branching point located at the proximal Ub, based on which we revealed how the K48-linked UbIV extended from the K11-linked UbIII to form a hitherto unseen alternating K11-K48-linked Ub chain stemming from the proximal UbI (Fig. 1e-g). RPN2, RPN8, RPN9, RPN10, and RPN11 collectively form a multi-subunit groove to accommodate the K11-linked Ub chain. The K11-linkage specificity of this multi-subunit binding groove was corroborated by the cryo-EM analysis of the K11-only homotypic Ub chain (Supplementary Fig. 13). The K11/K48-branched Ub chain made extensive hydrogen bonds and hydrophobic interactions with the 19S RP through the canonical I36 and I44 hydrophobic patches within the individual Ubs (Fig. 2). Specifically, the K11-K48-linked di-Ub was recognized by conserved Ub binding motif within RPN2, reminiscent of T1 Ub binding site on RPN1 (Fig. 3). Unlike RPN1, which binds to K48-linked di-Ub using the T1 helical bundle alone10, the recognition of the K11-linked Ub chain involves several proteasomal subunits in addition to RPN2: UbIII formed several salt bridges with RPN8 and RPN11; UbIV formed several salt bridges with RPN9 (Fig. 2). RPN2 alone may not be sufficient to bind to the alternating K11-K48-linked Ub chain stably. In other words, the multivalent nature of this unique Ub chain binding groove may explain why previous studies have failed to identify this cryptic Ub receptor because this binding groove consists of a collection of proteasomal subunits10,32.
Despite the intrinsic heterogeneity of the Ub chain length and linkages of the Sic1PY-Ubn used in this study, the ability to resolve the cryo-EM maps of the branched Ub4 is a testament that the unique multivalent binding mode provides superior avidity over homotypic Ub chains and other forms of Ub linkages, facilitating in silico enrichment of particle images to yield the well-defined cryo-EM maps of the K11/K48-branched Ub chains (Supplementary Fig. 6). Superposition of previously reported Ub chain structures extending from the structure of UbIV showed steric clashes between the extended Ub moieties and RPN9 that line the bottom of the K11-Ub binding groove (Supplementary Fig. 20). It is plausible that UCHL5, known to edit K11/K48-branched Ub chains21, may function as a molecular ruler to trim additional Ub moieties extending from UbIV to avoid these steric clashes. However, despite the addition of the catalytically inactive UCHL5, we could not map its position on the substrate-bound complex. Indeed, Walters and co-workers concluded in their recent cryo-EM study that the Ub binding sites on RPN10 and RPN13, which tethers the UCHL5 to the proteasome, are too dynamic to be underpinned by cryo-EM46.
The structural snapshots of the free and substrate-bound 26S proteasome revealed abundant and long-range conformational changes associated with the Ub-receptor binding and the catalytic cycle of the 26S proteasome. Between the EB:Ub4 state and ED:Ub4 state, the RPT4/5 CC underwent a significant lever motion to expose UbII. The lever motion was accompanied by the CP gate opening accomplished by the appropriate alignment of the basal ring and the OB ring (Fig. 4 and Supplementary Fig. 19). The conformational changes within the ATPase assembly were reminiscent of the previously reported activation mechanism of archaeal PAN ATPase, and the activation mechanism of the ATPase unfoldase activity upon Ub binding to the proteasomal Ub receptors47,64. The extensive tripartite binding interface formed by the K11/K48-branched Ub chain may generate a favorable driving force to trigger these allosterically coupled conformational changes together with the ordering of several intrinsically disordered loops (Fig. 4d–f and Supplementary Fig. 19d). Our structural findings thus suggested a pivotal role of the ATPase base of the 19S RP as a hub to sense and transmit allosteric regulatory signals triggered by substrate binding40,43,65,66.
Recently, Haselbach and colleagues reported the cryo-EM structure of a K11/K48-branched tri-Ub that shows essentially the same binding mode as reported in our current study67. Mutations in RPN2 and RPN11 result in accumulation of securin, a model substrate used in their study to be K11/K48-ubiquitinated through the concerted work of APC/C, UBE2C and UBE2S. Meanwhile, Haselbach and colleagues also reported in a separate study the cryo-EM structure of the 26S proteasome in complex with PITHD1 that functions as an proteasome inhibitor in the context of cellular dormancy68. PITHD1 inhibits the proteasome in a triple lock mechanism by targeting the same K11-Ub chain binding groove as we reported herein through a folded PITH domain and the insertion of an extended tail into the ATPase gate that prevent substrate entrance into the 20S CP. The overlapping binding mode between the PITH domain and the K11-linked Ub chain may suggest the interference of the recognition of K11/K48-branched Ub chains by PITHD1 as a mechanism to inhibit the proteasome during mitosis.
In summary, our study provides insight into how the human 26S proteasome recognizes a K11/K48-branched Ub chain through multivalent interactions involving multiple proteasomal subunits, namely RPN2, RPN8, RPN9, RPN10, RPN11 and the RPT4/5 CC. The newly identified K11-Ub binding groove on the 19S RP is particularly complex. While we have identified a conserved K48-linked Ub binding motif on RPN2, which is structurally similar to that of the T1 motif of RPN1, RPN2 alone is not sufficient to form a stable complex with the unique alternating K11-K48-linked Ub chain. Additional contacts made by RPN8, RPN9, and RPN10 may be necessary to generate sufficient multivalency (Fig. 2). This multi-subunit Ub binding groove helps account for the previously speculated cryptic Ub receptor10,32. Collectively, the K11/K48-branched Ub chain forms a large interaction surface with the proteasome that affords avidity, providing expedited turnover of K11/K48-branched Ub conjugated substrates at specific biological checkpoints for maintaining proteostasis in a timely fashion4,6.
Methods
Plasmids, genes, and site-directed mutagenesis
The plasmids pRSFDuet_His-TEV-gsggs-Ub(WT), pRSFDuet_His-TEV-gsggs-Ub(K0), pET21d_His-hE1, pETDuet_His-SUMO-gg-UBCH7, pGEX_TEV-SENP2, pMBP_TEV-Rsp5-WW3-HECT, and pMBP_TEV-Rsp5 WW3-HECTGML were synthesized by GenScript, USA. pET19b_RPN13 was a gift from Joan Conaway & Ronald Conaway (Addgene plasmid #19423; (http://n2t.net/addgene:19423); RRID: Addgene_19423)16. pOPINB-AMSH* was a gift from David Komander (Addgene plasmid #66712; (http://n2t.net/addgene:66712) RRID: Addgene_66712)69. Plasmid encoding the N-terminal hexahistidine (His6)-UCHL5(WT) gene was taken from the in-house plasmid collection. Plasmids encoding His6-TEV-gsggs-Ub(K63R), -Ub(K48R), -Ub(K11-only), -Ub(K48-only) and His6-UCHL5(C88A) were constructed using site-directed mutagenesis. K11-only or K48-only referred to the Ub variants of which all lysines but K11 or K48 were mutated into arginines, respectively. For each of the mutations, a pair of complementary mutagenic primers were designed with which the entire plasmid was amplified in a thermocycling reaction using a KOD (Invitrogen, USA) high-fidelity DNA polymerase and the plasmids encoding His6-TEV-gsggs-Ub(WT), -Ub(K0), or His6-UCHL5(WT) respectively as a template. The synthetic gene encoding the substrate polypeptide (later called Sic1PY) exhibiting an efficient degradation initiation region for the proteasome engagement was derived from the C-terminal sequence of S. cerevisiae Sic1 protein (residues 1-48). The sequence was modified to contain an additional PPxY motif and single lysine for ubiquitination by the Rsp5-E3 Ub ligase, a single cysteine for site-specific labeling, and His6-tag at its C-terminus. The gene cloned in pET-9a plasmid by NdeI/BamHI was purchased from GenScript, USA.
Human 26S proteasome
The human 26S proteasome complex used in the study was purchased from UBPBio, USA (Catalog No.: A1100 and A1101). The complex was expressed in human HEK 293 and affinity purified following the procedure described by Wang et al.70 with manufacture modifications. The proteasome sample was custom-made to be stored in HEPES buffer as frozen aliquots, shipped on dry ice and stored at −80 °C before use. The aliquots were thawed immediately before use. The purity of the sample was confirmed by native and SDS-PAGE; the activity of the sample was confirmed using a Suc-LLVY-AMC substrate followed by spectrometric analysis.
Recombinant protein expression and purification
The recombinant proteins used in the study, except human His6-RPN13, were expressed in E. coli BL21-CodonPlus (DE3)-RIL competent cells (Agilent Technologies). Proteins were expressed and purified following the protocol described below unless otherwise stated. During the purification procedures, protein solutions were kept on ice where possible, except for size-exclusion chromatography (SEC), during which the columns were kept at room temperature. A HiLoad 16/600 Superdex 200 pg column (GE Healthcare, USA) connected to an FPLC system (ÄKTA Purifier UPC 10 or ÄKTA Pure 25 L, GE Healthcare, USA) was used for SEC unless otherwise specified. Human ubiquitin-activating enzyme His6-Uba1 (E1) was purified at 4 °C at all times, including the SEC steps.
After harvesting by centrifugation at 4 °C, 6000 g for 15 min using a JLA 8.1 rotor in Avanti -26 XP Series centrifuges. (Beckman Coulter, USA), cells were lysed using NanoLyzer N2 homogenizer (Gogene Corporation, Taiwan) operating at 18–20 kpsi, unless otherwise stated. After lysis, cell debris, and aggregates were removed by 30 min centrifugation at 4 °C, 20,000 g using a JA 25.5 rotor in Avanti -26 XP Series centrifuges. (Beckman Coulter, USA). All the His-tag-based affinity purifications were done using the Ni-NTA resin (cOmplete™ His-Tag Purification Resin; Roche, Switzerland) for immobilized metal affinity chromatography (IMAC). The resin was incubated with protein at 4 °C for 2 h with gentle mixing and then packed into empty PD10 open columns (Cytiva, USA) for subsequent wash and elution. The wash and elution buffer were supplemented with 3 mM and 250 mM imidazole, respectively. After each purification step, the eluents were checked by SDS-PAGE to identify the target proteins and evaluate their purity. When applicable, the affinity purification tag was removed by incubation with TEV protease at a 1:50 (v/v) ratio overnight at 4 °C with simultaneous dialysis to remove residual imidazole. After the final SEC step, individual fractions containing the target proteins were pooled, concentrated by Amicon Ultra centrifugal ultrafiltration filters (Merck Millipore, USA), and checked by a NanoPhotometer N60 (IMPLEN, Germany) to determine their concentrations by their absorbance at 280 nm, aliquoted, snap-frozen and stored at −80 °C until further use.
Ub(WT), Ub(K63R), Ub(K43R), and Ub(K11-only) were expressed at 20 °C for 16–20 h after inducing the bacteria cells with 0.6 mM IPTG. The buffer used for cell lysis and subsequent IMAC purification is composed of 30 mM Tris-HCl, pH 7.5, 100 mM NaCl, and 5 mM β-mercaptoethanol (βME). The N-terminal His6-tag was removed from the protein by TEV protease followed by IMAC. The remaining impurities and cleaved His-tag were removed in buffer A (30 mM HEPES, pH 7.5, 100 mM NaCl, and 5 mM βME).
The recombinant E1 and the ubiquitin-conjugating enzyme UBCH7 (E2) were expressed at 16 °C for 20 h after inducing the bacteria cells with 0.6 mM IPTG. The harvested cells were disrupted by sonication (Qsonica, USA) in lysis buffer (50 mM Tris-HCl, pH 7.6, 200 mM NaCl, and 5 mM βME).
After IMAC purification, fractions containing the E1 protein were diluted with 50 mM Tris-HCl, pH 7.6, and 2 mM dithiothreitol (DTT) to reduce NaCl concentration to 40 mM and were further separated by anion-exchange chromatography using Q Sepharose Fast Flow resin (Cytiva, USA) packed into an empty PD10 open-column (Cytiva, USA). The E1 protein was eluted with a NaCl step gradient ranging from 0 to 400 mM with a 40 mM increment. The purification was completed by SEC using a Superdex 200 Increase 10/300 column (Cytiva, USA) in buffer B (30 mM HEPES, pH 7.5, 200 mM NaCl, and 0.5 mM TCEP).
The E2 protein construct used for the study contained an N-terminal His6-SUMO tag. To remove the SUMO tag, IMAC-purified protein was treated with in-house made His6-SENP2 protease at a 1:50 (v/v) ratio. After reducing the imidazole concentration by overnight dialysis at 4 °C, the SUMO tag was removed by Ni-NTA, and the flow-through fractions containing the E2 protein were collected. In the final step, the E2 protein was purified by SEC in buffer B.
The WW3-HECT domain of yeast ubiquitin ligase Rsp5 (E3(WT)) and its mutated version with C-terminal 3 amino acid insertion, namely Rsp5 WW3-HECTGML (E3), were expressed at 16 °C for 16 h after inducing the bacteria cells with 0.5 mM IPTG. Cells were lysed in 30 mM Tris-HCl, pH 8.0, 200 mM NaCl, 10% glycerol, 2 mM βME, and 0.5 mM PMSF. The same buffer was also used for washing and elution during subsequent IMAC purification. Both constructs contained an N-terminal His6-tag followed by a TEV cleavage site. The target proteins were purified by SEC in buffer C (20 mM HEPES, pH 7.5, 200 mM NaCl, 10% glycerol, and 0.5 mM TCEP).
UCHL5(WT) and UCHL5(C88A) were N-terminally His6-tagged. They were over-expressed at 16 °C for 20 h after induction with 0.4 mM IPTG. Cells were lysed in buffer C (3 mM DTT was used instead of 0.5 mM TCEP). The same buffer was used for the His-tag affinity purification and was supplemented with 3 mM and 200 mM imidazole for wash and elution, respectively. The His6-tag was not removed from the expressed protein. They were purified by SEC in buffer C.
RPN13 was N-terminal His-tagged. It was over-expressed in E. coli BL21 (DE3) (Yeastern Biotech, Taiwan) strain after induction with 0.4 mM IPTG. Cells were lysed in a buffer containing 20 mM Tris-HCl, pH 7.5, 150 mM NaCl, 5 mM DTT, and 1 mM PMSF. IMAC was carried out in buffer D (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, and 2 mM DTT) supplemented with 5 mM or 250 mM imidazole for wash and elution, respectively. SEC was performed using a HiLoad 16/600 Superdex 75 pg column (Cytiva, USA) in buffer C.
The RPN13-UCHL5 complexes were formed by mixing RPN13 with UCHL5(WT) or UCHL5(C88A) mutant at a molar ratio of 1:1.3. Mixtures were incubated on ice for 2 h, and excess free protein was removed by SEC in buffer C. The deubiquitinase (DUB) activity of RPN13-UCHL5(WT) was confirmed by comparing the DUB activity of free UCHL5(WT) using a model fluorogenic substrate Ub-AMC (UBPBio, USA)71.
Recombinant Sic1PY was overexpressed at 16 °C overnight after inducing the bacteria with 0.1 mM IPTG. The harvested cells were disrupted by sonication (amplitude 10, 10 s/10 s on/off, the total processing time of 20 min) in lysis buffer D supplemented with cOmplete™, EDTA-free Protease Inhibitor Cocktail (Roche, Switzerland). Sic1PY was purified by IMAC aided by its C-terminal His-tag. IMAC was performed in 20 mM Tris-HCl, pH 8.0, 200 mM NaCl, 10% glycerol, and 2 mM DTT.
His-tagged Lbpro* (29–184 aa, UniProt ID: P05161) was synthesized and subcloned to pRSFDuet expression vector by GenScript (Piscataway, NJ). Lbpro*(L102W) was generated by site-directed mutagenesis to cleave ubiquitin2. Over-expression of Lbpro* was induced by 0.6 mM IPTG. Lbpro* was purified by IMAC, followed by SEC using a HiLoad 16/600 Superdex 200 pg column (Cytiva, USA). Fractions containing Lbpro* were collected, concentrated, and flash-frozen by liquid nitrogen.
Site-specific fluorescent labeling of ubiquitin and Sic1PY
To facilitate the substrate detection in electrophoretic gels and distinguish deubiquitination of the substrate from Sic1PY proteolysis, we fluorescently labeled Ub with Fluorescein-5-Maleimide (AnaSpec, USA) and Sic1PY with Alexa Fluor 647 C2 Maleimide (Thermo Fisher Scientific, USA). A cysteine was engineered at the C-terminus of Sic1PY; a cysteine was introduced to the N-terminus of a Ub variant. Both proteins were fluorescently labeled through maleimide-mediated covalent modification. Fluorescein-5-Maleimide and Alexa Fluor 647 C2 Maleimide were dissolved according to manufacturer instructions at 0.1 M concentration. Before adding the dye, the protein was reduced by DTT at a final concentration of 30 mM on ice for 30 min. The reducing agent was removed by desalting using a disposable PD 10 desalting column (Cytiva, USA). The reduced protein in 25 mM HEPES, pH 7.0, and 150 mM NaCl was incubated with a five-fold excess of the dye at 4 °C overnight. The reaction was stopped by adding a quenching buffer to a final concentration of 50 mM Tris and 20 mM DTT. A second desalting step subsequently removed the unreacted fluorescent dye. The fluorescently labeled proteins were resolved by 4-20% SurePAGE (GeneScript, USA), and the protein concentration was estimated based on the measured dye-specific absorbance.
Preparation of Sic1PY-Ubn
In vitro ubiquitination assay was performed in a 10 mL reaction mixture containing 50 mM HEPES, pH 7.0, 250 mM NaCl, 10 mM MgCl2, 7.5 mM ATP, 0.05 mg/mL BSA, 250 nM UBE1 (E1), 750 nM UbcH7 (E2), 1.2 µM Rsp5-HECTGML (E3), together with 7.15 µM Ub (10% fluorescein-labeled) and 1.25 µM Sic1PY (10% AlexaFluor-647-labelled) at 37 °C for 30 min. The reaction mixture was incubated at 4 °C for 2 h with 3 mL of cOmplete Ni-NTA resin (Roche, Switzerland) equilibrated in buffer E (50 mM Tris-HCl, pH 7.5, 500 mM NaCl, 0.005% NP40). Polyubiquitinated Sic1PY and E1 (both His-tagged) were eluted by buffer E supplemented with 250 mM imidazole. The product was fractionated and E1 removed by SEC using a Superdex 75 increase 10/300 column (Cytiva, USA) with buffer F (30 mM HEPES, pH 7.5, 100 mM NaCl, 5% glycerol, 0.5 mM TCEP, and 0.005% NP-40). The fractions containing the substrate with Ub chains of intermediate length (4-10 Ubs) were collected and used for further experiments.
Native-gel separation of 26S proteasome
To resolve the proteasomal complexes, 3–8% Tris-acetate gels (Invitrogen, USA) were used for native electrophoresis. The reconstituted samples were mixed with 4× loading buffer (100 mM Tris-HCl, pH 8.0, 20% glycerol, 2 mM ATP, 0.002% bromophenol blue) immediately before loading. 1–2 ug of proteasomal complex was loaded per well. The separation was carried out at 4 °C and 150 V for 4 h, using a TBE running buffer supplemented with 0.5 mM DTT, 0.5 mM ATP, and 2 mM MgCl2 in a XCell SureLock Mini-Cell (Invitrogen, USA).
In-gel 26S proteasome activity
Native gels containing 26S proteasomes were incubated with reaction buffer (50 mM Tris-HCl, pH 7.5, 10 mM MgCl2, 1 mM ATP, 1 mM DTT, 50 μM Suc-LLVY-AMC) for 30 min at 37 °C. The fluorescence signal of the product generated by 26S proteasomes around the complex position in the gel was visualized and quantified by ChemiDoc XRS+ Imaging Systems with supplied Image Lab Software (BioRad, USA).
Immunodetection of native 26S proteasome components
For immunoblotting, the native gels containing the 26S proteasomes were transferred to Amersham Hybond LFP, 0.2 PVDF membranes (Cytiva, USA) using Bio-Rad mini-protean 3 transfer system and buffer containing 25 mM Tris base, 192 mM glycine, 0.1% SDS (sodium dodecyl sulfate) for 4 h at 70 V (current limit: 350 mA) in cold room. The membranes were incubated with blocking buffer containing 5% non-fat dry milk in TBST (Tris-buffered saline with 0.1% Tween-20) at room temperature for 1 h and then incubated with primary antibodies diluted in blocking buffer at 4 °C overnight. Next, the membranes were washed five times for 5 min each with TBST and then incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies ×10,000-fold diluted in blocking buffer at room temperature for 1 h. After five times 5 min wash with TBST, the membrane was covered with Clarity Western ECL Blotting Substrate (BioRad, USA) and incubated for 5 min. Finally, the developed HRP signal was recorded by iBright Imaging System 5.3.0 (Invitrogen, USA).
For re-probing with different antibodies, the membranes were stripped by incubating the membrane five times for 10 min each in a solution containing 1.5% glycine, pH 2.2, 0.1% SDS, and 1% Tween20. The membrane was subsequently washed twice for 10 min each with PBS and twice for 10 min each with TBST. The membrane was then again blocked, and proteins were detected in the same way as described above.
Western blot analysis of ubiquitinated substrates
The ubiquitinated substrates were resolved by a 4-20% SurePAGE (GeneScript, USA) using Tris-MOPS running buffer supplied with the gels. Before loading on the gel, samples were mixed with 4× loading buffer (200 mM Tris–HCl (pH 6.8), 8% SDS, 0.2% bromophenol blue, 40% glycerol, 20% βME) and heated at 70 °C for 10 min. The electrophoretic separation was carried out at room temperature in an XCell SureLock Mini-Cell (Invitrogen, USA) at 200 V for 30 min. Gels containing the resolved samples were transferred to nitrocellulose membranes using a Bio-Rad mini-protean 3 transfer system and buffer containing 25 mM Tris base, 190 mM glycine, 20% Methanol for 1 h at 100 V (current limit: 350 mA) at room temperature. Alternatively, the proteins were transferred to a PVDF membrane using iBlot Dry Blotting System and dedicated transfer stacks (Invitrogen, USA). The immunodetection procedure was performed the same way described above for analysis of the native 26S proteasome complexes.
Negative stain electron microscopy (NSEM)
Four microliters of 30× diluted human 26S proteasome (1 mg/mL, UBPBio, USA) alone or mixed with SEC-purified ubiquitinated substrate were placed on carbon-coated grids that were glow-discharged at 25 mA for 30 s. After staining with 2% uranyl formate, the grids were blotted and dried in the air for 1 day. Images were collected using a FEI Tecnai G2-F20 electron microscope at 200 keV (FEI, USA). A magnification of 29,000× was used, corresponding to a pixel size of 2.9 Å. The micrographs were converted to mrc format using EMAN2 and imported to RELION-3.072 for particle picking. Picked particle stacks were exported to CryoSPARC v2.1473,74,75, where further data processing steps were conducted, including 2D classification, ab-initio 3D reconstruction, 3D classification, and final 3D refinement. The resulting 3D structures were visualized using UCSF-ChimeraX76,77.
Mass spectrometry analysis of Ub chain compositions
Sic1PY-Ubn was treated with Lbpro* to determine the polyubiquitin chain architecture. A batch of the ubiquitinated substrate was prepared following the above ubiquitination protocol without fluorescently labeled Sic1PY and Ub. The purified Sic1PY-Ubn was treated with 10 µM Lbpro* in 25 mM Tris-HCl, pH 7.5, 100 mM NaCl, and 10 uM EDTA and incubated at 37 °C for 20 h. To determine Ub linkage types, the Lbpro*-treated sample was separated by 4-20% SurePAGE (GeneScript, USA) using Tris-MOPS running buffer supplied with the gels and stained with One-Step Blue® (Biotium, USA). The protein bands were excised from the gel, diced, and destained with 25% (v/v) acetonitrile (ACN) and 5 % Trifluoroacetic acid (TFA). The diced gel was reduced by 50 mM Dithioerythritol (DTE) in 25 mM ammonium bicarbonate (ABC) at 37 °C for 1 h, and subsequently alkylated by 100 mM iodoacetamide (IAM) at room temperature in the dark for 1 h. The diced gel was washed with 25% (v/v) ACN in 25 mM ABC, pH 8.5 for 5 min once, dehydrated with 25% ACN in 25 mM ABC for 5 min, and dried under vacuum. The gel pieces were rehydrated with a 1:20 protease to protein ratio with a Mass Spectrometry Grade Lys-C (Fujifilm-Wako, USA) in 25 mM ABC digested at 37 °C for 3 h. Following Lys-C digestion, the same amount of sequencing grade Trypsin (Promega, USA) in 25 mM ABC was added for digestion at 37 °C for 16 h. Tryptic peptides were extracted twice with 50% (v/v) ACN containing 5% (v/v) TFA for 3 min each time with moderate sonication. The extracted solutions were pooled and dried by speed vacuum. The peptide mixture was aliquoted and desalted by a C18-ZipTip (Merck Millipore, USA) and eluted with 50% ACN in 0.1% (v/v) formic acid (FA). 200 fmol of ubiquitin-absolute quantification (Ub-AQUA) peptide, which is a gift from Dr. Yasushi Saeki in Tokyo Metropolitan Institute of Medical Science, Japan, was added into the extracted peptide as ideal internal standards and subsequently concentrated peptide mixture to a final volume of less than 2 µL using by speed vacuum followed by H2O2 treatment by diluting the peptide mixture to 20 μL of 0.1 % TFA and 0.05 % H2O2 at 4 °C overnight to oxidize methionine. Finally, the ubiquitin-absolute quantification/ parallel reaction monitoring (Ub-AQUA/ PRM) assay was used to estimate the abundance of each Ub-chain type linkage of signature peptides with di-glycine (GG) modification at a specific lysine against the total number of the modified peptides detected in the sample.
LC-MS-based Ub-AQUA/PRM analysis was performed on a Thermo UltiMate 3000 RSLCnano system connected to a Thermo Orbitrap Fusion mass spectrometer (Thermo Fisher Scientific, Germany) equipped with a nanospray interface (New Objective, USA). Peptide mixtures were loaded onto a 75 μm ID, 25 cm length C18 BEH column (Waters, USA) packed with 1.7 μm particles with a pore of 130 Å and were separated using a segmented gradient in 30 min from 5% to 35% solvent B (0.1% (v/v) FA in ACN) at a flow rate of 300 nL/min. Solvent A was 0.1% (v/v) FA in water. The mass spectrometer was operated in the PRM mode. Briefly, the MS1 scans of peptide precursors from 350 to 1600 m/z were performed at 120 K resolution with a 2 × 105 ion count target. The PRM scans were performed by isolation window at 1 Da with the quadrupole, HCD fragmentation with normalized collision energy of 30, and analysis at 30 K resolution in the orbitrap. The PRM scans ion count target was 1 × 106, and the max injection time was 54 ms. The PRM scans were triggered by time-scheduled targeting precursor ions selected for target peptides (without heavy isotope labeling, Supplementary Table 1) in ±2 min elution windows.
The PRM data were processed using Skyline 22.2.0.255 (MacCoss Lab Software, USA) to generate XIC and perform peak integration. All peaks were manually inspected to confirm correct detection and peak boundaries. Peak integration and calculation of the ratios between light endogenous and the heavy-labeled peptide (L/H) were done in Skyline, and result reports were exported from the software. Light-to-heavy peptide peak area ratios were used for the target peptide quantitation analysis.
For determining the branching of the Ub chains, an aliquot of the Lbpro*-treated substrate sample corresponding to two micrograms of the pure protein was first diluted with 50% (v/v) Methanol followed by 50% (v/v) ACN and 1% (v/v) FA. An aliquot corresponding to one pmol of the pure protein was subsequently injected via the LockSpray Exact Mass Ionization Source (Waters, USA) with a syringe pump (Harvard Apparatus, USA) and held a flow rate of 3 uL/min throughout the analysis to the mass spectrometer. The mass of intact proteins was determined using Waters Synapt G2 HDMS mass spectrometer (Waters, USA). The acquired spectra were deconvoluted to zero-charge spectrum using MaxEnt1 algorithm of the MassLynx 4.1 software (Waters, USA).
Cryo-EM sample preparation
To prepare 26S-UCHL5(C88A)-Sic1PY-Ubn(K63R) complexes, 40 μL of 1 mg/mL purified human 26S proteasome (UBPBio, USA) was first incubated at 37 °C for 10 min and then supplemented with preformed RPN13:UCHL5(C88A) to a final concentration of 1 μM and incubated for another 10 min at room temperature. To prevent substrate en bloc deubiquitination by RPN11, the sample was treated with 3 mM o-phenanthroline (oPA) and incubated for an extra 5-10 min. Next, 40 μL of 2-3 µM ubiquitinated Sic1PY substrate, supplemented with 2 mM ATP and 5 mM MgCl2, was added to the mixture and incubated for another 5 min at room temperature. The sample was then diluted with 350 μL of cold buffer composed of 20 mM HEPES, pH 7.6, 40 mM NaCl, 5 mM MgCl2, 2 mM βME, 0.5 mM oPA, and concentrated to 40 μL using 100 kDa MWCO Amicon Ultra-0.5 mL Centrifugal Filter (MerckMillipore, USA) to reduce glycerol content. The sample was incubated at room temperature immediately before vitrification. Three microliters of the reconstituted sample were applied onto 300-mesh Quantifoil R1.2/1.3 holey carbon grids, which were glow-charged at 20 mA for 30 s before use. After incubation for 3 s, the grids were blotted for 2.5 s at 4 °C and 100% humidity and vitrified using a Vitrobot Mark IV. (Thermo Fisher Scientific, USA).
Cryo-EM data collection
Cryo-EM data acquisition was performed on a 300 keV Titan Krios transmission electron microscope (Thermo Fisher Scientific, USA) equipped with a Gatan K3 direct detector (Gatan, USA) in a super-resolution mode using the EPU software (Thermo Fisher Scientific, USA). Movies were collected with a defocus range of -1.2 to -1.8 μm at a magnification of 64,000×, corresponding to a super-resolution pixel size of 0.668 Å. A total of 48–50 e-/Å2 was distributed over 50 frames with an exposure time of 1.8 s. The datasets were energy-filtered with a slit width of 15–20 eV, and the dose rates were adjusted to 8–10 e-/pixel/s.
Cryo-EM data processing and model building
A total of 15,242 micrographs were recorded, then aligned and two-fold binned (resulting in a pixel size of 1.4 Å) using MotionCor2 (UCSF)78 and CTF corrected by CTFFIND4.179. The particle images of the single- (RP-CP) and double-capped (RP2-CP) proteasome particles were manually picked and used as a training set for automated particle picking by Topaz convolutional neural network, which resulted in an initial dataset of 2,695,911 particles80. Further data processing, including 2D and 3D classifications, was performed in RELION-3.181, while the final nonuniform refinement and variability analysis was performed in CryoSPARC v3.1 (Structura Biotech, Canada)73,74,75. The initial dataset of picked particles was extracted, and four-fold binned (resulting in a pixel size of 5.6 Å, box size of 150 pixels) was cleaned by 2D and global 3D classification, resulting in a dataset containing 527,610 RP-CP and 575,778 RP2-CP proteasome particles. The particles representing RP2-CP due to their pseudo symmetry were refined, imposing the C2 symmetry restrain, then symmetry-expanded (doubling the number of the RP2-CP particles), and again refined with C1 symmetry with mask focused on only one RP of the complex. The symmetry expansion procedure was used to allow the merging of the RP2-CP particles with the RP-CP complexes while simultaneously reducing the heterogeneity of the dataset. After combining, all the particles were re-centered on one RP (RP of interest, in the case of the symmetry-expanded RP2-CP the refined RP) and re-extracted with two-fold binning (resulting pixel size of 2.8 Å, box size of 300 pixels). After 3D classification focused on the RP of interest, the particles’ signal was partially subtracted, leaving only the signal of the RP of interest and the first ring of α subunits composing the CP subcomplex. The classification yielded two conformations of the RP, one with 532,818 particles resembling ED active state of the human 26S complex and the second with 736,915 RPs resembling the resting EA state of the human 26S proteasome. After global refinement, particles in each subset were re-extracted at full resolution (resulting in a pixel size of 1.4 Å, box clipped to 400 pixels). Further classifications of the EA-like particles focused on the ubiquitin-binding region filtered out 153,646 particles without any obvious proteasome-bound ubiquitin densities. The subset produced 3.4 Å resolution map of “empty” proteasomes after nonuniform refinement in CryoSPARC v3.1. Heterogeneity among the remaining 202,823 EA-like particles was resolved by 3D Variability Analysis within CryoSPARC v3.1 (Structura Biotech, Canada)73,74,75, which revealed the existence to EA-like proteasomes with three (EA:Ub3) or EB-like proteasomes with four (EB:Ub4) ubiquitin bound at the RP, which accounted for 54,794 and 52,216 particles, respectively. The particles representing both states were then refined by nonuniform refinement in CryoSPARC v3.1, yielding a map of the EA:Ub3 and EB:Ub4 with an overall resolution of 3.6 Å and 3.8 Å, respectively.
The subset of ED particles was also classified with a mask focused on the ubiquitin-binding region. Traces of proteasome-bound Ub were found in all of the obtained classes; however, for further processing, only 212,581 particles with the best and unambiguously defined Ub densities were selected. Based on subsequent classification aimed at resolving heterogeneity in the AAA+ ATPase, we extracted two subsets of ED:Ub4 particles with slightly different conformations of the ATPase subcomplex accounting for 84,476 and 49,828 particles, which, after nonuniform refinement in CryoSPARC v3.1 produced maps with an overall resolution of 3.8 Å and 4 Å, respectively. The reconstructed ED:Ub4 maps showed no significant differences in the region of interest (the Ub and Ub-binding region). The 3.8 Å map provided better detail of the AAA+ ATPase, so we used it to build the atomic model of the ED:Ub4 state. All the reconstructed cryo-EM maps of the Ub-bound complexes were further focus-refined with masks around RPN3:RPN7 region, Ub and neighboring interfaces, and AAA+ ATPase. The obtained focused maps were merged in Phenix using Combine Focused Maps procedure82. The resulting combined cryo-EM maps were used for final atomic model refinement. The atomic models of the empty complex, EA:Ub3, EB:Ub4, and ED:Ub4 states of the complex were built using Phenix and COOT82,83 based on PDB 6MSB (EA state) and 6MSK (ED state) structures as references45. Atomic details and interactions of the proteasome-bound K11/K48-branched Ub chain and the neighboring subunits were refined using ISOLDE84 within UCSF ChimeraX76,77. The latter was also used to visualize the cryo-EM maps and atomic models. All cryo-EM data collection and refinement statistics are reported in Supplementary Table 2.
Quantification of the functional states of the 19S RP in the doubly capped (RP2-CP) and singly capped (RP-CP) 26 proteasome
To quantify the relative populations of the 19S RP in the active and resting states in the RP2-CP and RP-CP, we used the particle stacks from the early steps of data processing, representing 575,778 RP2-CP and 527,610 RP-CP particles. We refined each subset with C1 symmetry and subsequently ran focused 3D classification (without particle alignment and with the regularization parameter T = 64) on each of the RPs (RP and RP’) in the RP2-CP complex and conducted a similar classification focused on the RP of the RP-CP particles. We divided the particles based on class assignments and calculated class occupancies. We then traced which RPs originated from the same RP2-CP particle and plotted the connection between the RPs representing different conformations of the opposing RPs in the same RP2-CP complexes, using pySankey (https://github.com/anazalea/pysankey).
Substrate degradation assay
The rate and efficiency of proteasome-mediated degradation of Sic1PY ubiquitinated with branched Ub chains (generated using Ub(K63R) mutant), or with homotypic K11-, or K48-linked chains (generated using the Ub(K11-only) or Ub(K48-only) variants, respectively) was estimated based on quantification of the degradation products separated in SDS-PAGE. The degradation reaction contained 60 nM human 26S proteasome in 30 mM HEPES, pH 7.4, 50 mM NaCl, 10% glycerol, 0.5 mM TCEP, 0.5 mg/mL BSA, ATP regeneration system (50 mM ATP, 100 mM MgCl2, 3.2 mg/mL creatine kinase, 160 mM creatine phosphate), and 10 nM of Sic1PY-Ubn. The concentrations of the substrates labeled with branched-, K11-, and K48-linked Ub chains used for the degradation assay were normalized based on the AlexaFluor-647 signal of each of the substrate preparations. To check the effect of UCHL5 on the proteasome-mediated degradation of the substrates, the reactions were additionally supplemented with preformed SEC-purified complexes of RPN13-UCHL5(WT), catalytically inactive RPN13-UCHL5(C88A), or RPN13 alone as a control. The reactions were carried at 37 °C for 30 min, after which they were stopped by adding SDS-PAGE loading dye into the sample and 10 min heating at 37 °C. Degradation products and remaining undegraded substrates were resolved by a 4-20% SurePAGE (GeneScript, USA) using Tris-MOPS running buffer supplied with the gels. The amount of degraded substrate was estimated by quantifying the AlexaFluor-647 signal of the small molecular weight degradation products appearing in the very bottom of the gel in the samples treated with the proteasome. The degradation efficiency of each of the tested Ub chains (expressed as a faction of degraded substrate labeled with the particular Ub chain) was calculated by comparing the signal of the degradation products with the total signal of the degradable substrate, i.e., the sum of remaining substrate labeled with Ub chains of at least four Ub (corresponding to signals of molecular weight of ≥50 kDa) and the degraded substrate (which equals to the signal of degradation product). The statistical significance of the observed differences was calculated using two-way ANOVA performed using GraphPad Prism 8 (GraphPad Software, USA).
Sample preparation for XL-MS
The functional 26S proteasome complex samples for the XL-MS experiment were prepared the same way as for the cryo-EM experiment, except that one of the samples was prepared without the substrate. The formed complex samples, with and without the substrate (n = 1), each containing 50 μg of the human 26S proteasome (UBPBio, USA) in a total volume of 100 μL were then individually cross-linked using disuccinimidyl sulfoxide (DSSO) at a final concentration of 80 μM for 1 h following previously described protocols85. The amount of DSSO equates to a 300-fold excess of crosslinker to the 26S proteasome. The reactions were stopped by adding ABC to a final concentration of 5 mM.
Cross-linked samples were then buffer-exchanged to 50 mM triethylammonium bicarbonate buffer using 100 kDa MWCO Amicon Ultra-0.5 mL Centrifugal Filter (Merck Millipore, USA). Samples were reduced and alkylated by 10.7 mM TCEP and 21.4 mM chloroacetamide at 95 °C for 10 min. Proteins were proteolyzed with Lys-C (1:50 w/w) and trypsin (1:50 w/w) for 4 h at 37 °C. The peptide digests were desalted using Sep-Pak C8 cartridges (Waters, USA), dried under vacuum, and reconstituted in 0.05% TFA and 1% (v/v) ACN for subsequent LC/MS measurements.
Samples were separated by reverse phase LC using a Thermo Scientific™ UltiMate™ 3000 UHPLC system connected to a Thermo Scientific™ EASY-Spray™ column, 50 cm × 75 µm over a 130 min with 4-25% gradient (A: water, 0.1% FA; B: ACN, 0.1% FA) at a flow rate of 300 nL/min. The cross-linked samples were analyzed on a Thermo Scientific™ Orbitrap Eclipse™ Tribrid™ mass spectrometer with FAIMS Pro™ device (Thermo Fisher Scientific, USA) and Instrument Control Software version 3.4 using a stepped collision energy (SCE) MS2 HCD of 21-26-31% or MS2CID-MS3CID acquisition strategies86. MS2 sequencing events were triggered for precursors of charge states +3 to +8. Mass-difference-dependent CID-MS3 acquisitions were triggered if a unique mass difference (Δ = 31.9721 Da) was observed in the CID-MS2 spectrum. MS1 and MS2 scans were acquired in the Orbitrap with a mass resolution of 60,000 and 30,000 correspondently, whereas MS3 scans were acquired in the ion trap. Precursor isolation windows were set to 1.6 m/z for MS2 scans and 2.5 m/z for MS3 scans.
XL identification and quantification
The raw data were analyzed using Proteome Discoverer 3.2 software (Thermo Fisher Scientific, USA) with XlinkX node 3.287 for cross-linked peptides and a SEQUEST HT search engine for unmodified peptides, looplinks and monolinks. The following parameters were applied for Proteome Discoverer: MS1 ion mass tolerance, 5 ppm; MS2 ion mass tolerance, 20 ppm; MS3 ion mass tolerance, 0.6 Da; maximal number of missed cleavages, 2; minimum peptide length, 6.
Carbamidomethylation ( + 57.021 Da) was used as a static modification for cysteine. Different DSSO cross-linked mass modifications were used as variable modifications for lysine in addition to methionine oxidation ( + 15.996 Da). Data were searched against a database containing Uniprot/SwissProt 64 entries of human SwissProt database based on protein identifications from the proteomics experiment of 26S proteasome complex. The false discovery rate (FDR) was set to 1% at CSM and cross-linkers levels. Additionally, cross-links were filtered with an identification score of ≥40 and Δ-score of ≥4. Results visualization was performed using xiNET88 and the XMAS plug-in for ChimeraX56.
PSMD1 knockout by Cas9 ribonucleoprotein electroporation
PSMD1 knockout (KO) was conducted in HEK293T cell line by electroporation of Cas9 ribonucleoproteins (RNP) as illustrated in Supplementary Fig. 16a. HEK293T cells were obtained from American Type Culture Collection and maintained in Complete DMEM medium containing DMEM with high glucose (HyClone) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 25 mM HEPES, 1X GlutaMAX, 1X penicillin-streptomycin. Recombinant Cas9 protein was expressed and purified from Escherichia coli, and single-guide RNAs (sgRNAs) were synthesized by in vitro transcription as previously described89. The sgRNA targeting PSMD1 (5’-CCTTAGATCGTTAGACACGG-3’) was designed using the Benchling CRISPR design tool (www.benchling.com). For Cas9 RNP formation, 48 pmol of sgRNA was incubated with 40 pmol of Cas9 protein at 37 °C for 25 min. Cas9 RNP was then mixed with 2 × 10⁵ HEK293T cells in 20 μL of SF buffer (Lonza) and electroporated using a 16-well nucleocuvette and the 4D-Nucleofector X Unit (Lonza) with pulse code DS150. After electroporation, 100 μL of pre-warmed complete DMEM was added, and cells were incubated at 37 °C for 15 min for recovery. The cells were transferred to a 24-well plate containing 1 mL of medium and maintained under standard culture conditions. On day 3 post electroporation, a fraction of cells was harvested and analyzed for KO efficiency. On day 5, single cells were isolated by fluorescence-activated cell sorting (FACS) using FACSAria III (BD Biosciences) at the flow cytometry core facility of the Institute of Biomedical Sciences, Academia Sinica. to establish single clones. One cell was isolated into each well of a flat-bottom 96-well plate containing 200 μL complete DMEM for expansion. On day 18, the edited cell population and viable single clones were analyzed again for KO efficiency.
For KO analysis, cells were harvested by centrifugation at 300 × g for 5 min, washed once with DPBS, and lysed in QuickExtract (Lucigen, USA) by sequential incubation at 65 °C for 15 min, 98 °C for 5 min, and 12 °C for 5 min. Genomic DNA (100 ng) was used for PCR amplification using KAPA HiFi HotStart PCR Kit (Roche, Switzerland) with forward primer (5’-GCTGAGTTTATCACTGACCTTTTGATTAT-3’) and reverse primer (5’-CTCTTCCATCCTCCCATACATTACTAAA-3’). Amplicons were purified by QIAquick PCR Purification Kit (Qiagen, Germany), sequenced at the Sanger sequencing facility at the Institute of Biomedical Sciences in Academia Sinica, and analyzed for indel percentage using the Inference of CRISPR Edits tool (https://ice.synthego.com) with default settings.
PSMD1 mutations knock-in (KI)
CRISPR-mediated homology-directed repair (HDR) was performed to insert double mutant PSMD1 sequence (E538K/D564K) at the endogenous PSMD1 exon 13 locus (2X KI). Also, a synthetic wild-type (WT KI) sequence was inserted as an editing control. HDR templates consisted of a 3’ PSMD1 sequence fused to a T2A self-cleaving sequence and egfp reporter gene, flanked by ~550-nt homology arms (Supplementary Fig. 16b). HDR templates were synthesized as plasmids by GenScript, and the sequences are listed in Supplementary Information. Forty pmol of Cas9 RNP was co-electroporated with 1 μg of HDR plasmid into 2 × 10⁵ HEK293T cells using the same electroporation condition as in gene KO. Successful knock-in produced EGFP+ cells, which were isolated by FACS using FACSAria III on day 7 post electroporation. To establish single clones, one cell was seeded into each well on a flat-bottom 96-well plate containing 200 μL Complete DMEM for expansion. On day 14, each clone was validated by genomic PCR, junction PCR, Sanger sequencing and flow cytometry. The confirmed mutant clones were cultured for two more weeks before proliferation assay and drug treatment.
For genomic PCR, cells were lysed using QuickExtract (Lucigen, USA) as described above. PCR amplification was performed using KAPA HiFi HotStart DNA Polymerase kit (KAPA Biosystems), forward primer (5’-TGATTATCACATCCTTTGATCATTCTTACT-3’) and reverse primer (5’-TTTTAAAGTCAGTAAAACATCCAACTGAAA-3’). Junction PCR used forward primer (5’-TGATTATCACATCCTTTGATCATTCTTACT-3’) and reverse primer (5’-CATCATCATTAACATCACTTACAGCAACA-3’). Amplicons were resolved by DNA gel electrophoresis in TAE buffer and stained with SYBR Safe (Thermo Fischer Scientific) for visualization. Amplicons were also purified by QIAquick PCR Purification Kit (Qiagen, Germany) for Sanger sequencing.
To assess EGFP expression, the knock-in cells were harvested, washed once with 1 mL of DPBS (Corning), and resuspended in ice-cold flow buffer (DPBS supplemented with 2% FBS, 25 mM HEPES, and 0.5 mM EDTA) containing DAPI (BioLegend). The experiments were performed in CytoFLEX (Beckman Coulter) and the data were analyzed with FlowJo (BD Biosciences).
Cell proliferation assay
Cell proliferation was assessed using the xCELLigence Real-Time Cell Analysis (RTCA) system (ACEA Biosciences). Parental HEK293T, PSMD1-KO and 2X mutant cells were seeded at 2 × 10⁴ per well in 200 μL complete DMEM into 16-well E-plates with microelectrodes (ACEA Biosciences) and incubated at 37 °C with 5% CO₂ for cell proliferation. Cell density was measured by electrical impedance and recorded as cell index every 15 min. All cell lines were routinely tested for mycoplasma contamination using the EZ-PCR detection assay kit (Biological Industries).
Impact of PSMD1 KO and KI by immunoblotting
Untreated control cells, KO, WT KI and 2X KI cells were seeded at 8 × 105 per well in 2 mL complete DMEM into 6-well plate, treated with 100 ng/mL nocodazole (Sigma, USA) for 16 h and 15 μM MG132 for last 3 h of synchronization. Cells were harvested and washed once with 1 mL DPBS. Cells were resuspended and lysed in 50 mM Tris pH 8.0, 250 mM NaCl, 5 mM EDTA, 1% NP-40. After centrifugation at 13,000 rpm, 4 °C for 30 min, supernatants were mixed with 4X loading dye and heated at 95 °C for 5 min. 10 µL Samples were loaded into 12% SDS-PAGE and run at 180 V for 45 min. All the proteins were transferred from gel to PVDF membrane by using Trans-Blot Turbo Blotting System (BIO-RAD, USA) and Trans-Blot Turbo RTA Transfer Kit, PVDF (BIO-RAD, USA). PVDF membranes were blocked by 5% skim milk (Becton Dickinson, USA) dissolved in TBST buffer at room temperature for 1 h. Primary antibodies (anti-GAPDH, batch# 3600003301, AC033, Abclonal, USA; anti-PSMD1, Lot# J2219, SC-166038, Santa Cruz, USA; anti-ubiquitin, Lot# C2124, SC-8017, Santa Cruz, USA; anti-cyclinA, Lot# L1724, SC-271682, Santa Cruz, USA; anti-cyclinB1, Lot# J2324, SC-245, Santa Cruz, USA; anti-Nek2, Lot# D1224, SC-55601, Santa Cruz, USA) were diluted 1000X with TBST buffer and incubated on a shaking platform at 4 °C overnight. Next day, membranes were washed with TBST buffer for 5 min three times before the secondary antibody. Secondary antibody (anti-mouse, Lot# 38, 7076S, Cell Signaling, USA) was performed 5000X dilution in TBST buffer and incubated at room temperature for 1 h, then washed with TBST buffer for 5 min four times. Protein signals were activated by Clarity Western ECL Substrate (BIO-RAD, USA), then detected by iBright FL1500 Image System (Thermo Fisher Scientific, USA) and analyzed by iBright Analysis Software 5.3.0 (Thermo Fisher Scientific, USA).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates of the apo EA state generated in this study have been deposited in the Protein Data Bank (PDB) under the accession code 8JRI (EA state). The atomic coordinates of the 26S proteasome complexed with substrates in different states generated in this study have been deposited in the Protein Data Bank (PDB) under the accession codes 8JRT (EA:Ub3 state), 8JTI (EB:Ub4 state) and 8K0G (ED:Ub4 state). The cryo-EM map of the apo EA state has been deposited in the Electron Microscopy Data Bank (EMDB) under the accession codes EMD-36598 (consensus). The cryo-EM maps of the EA:Ub3 state have been deposited in the EMDB under the accession codes EMD-36605 (composite), EMD-37269 (consensus), EMD-37273 (local refinement focused on Ub binding region), EMD-37276 (local refinement focused on Rpn3/Rpn7 region), and EMD-37277 (local refinement focused on AAA+ ATPase subcomplex). The cryo-EM maps of the EB:Ub4 state have been deposited in the EMDB under the accession codes EMD-36645 (composite), EMD-37317 (consensus), EMD-37319 (local refinement focused on Ub binding region), EMD-37327 (local refinement focused on Rpn3/Rpn7 region), and EMD-37328 (local refinement focused on AAA+ ATPase subcomplex). The cryo-EM maps of the ED:Ub4 state have been deposited in the EMDB under the accession codes EMD-36764 (composite), EMD-37334 (consensus), EMD-37335 (local refinement focused on Ub binding region), EMD-37341 (local refinement focused on the RP lid), and EMD-37344 (local refinement focused on AAA+ ATPase subcomplex). The RAW files of XL-MS analysis have been deposited to PRIDE - PRoteomics IDEntifications Database under the accession codes PXD064931 (Cross-link MS analysis of the human 26S proteasome in complex with RPN13:UCHL5 without Sic1PY-Ubn), PXD064932 (Cross-link MS analysis of human 26S proteasome in complex with RPN13:UCHL5 with Sic1PY-Ubn). The RAW files of the Ub-AQUA/ PRM and Intact MS analysis were deposited to MassIVE database under accession code MSV000092883 and MSV000092884, respectively. Source data are provided with this paper.
References
French, M. E., Koehler, C. F. & Hunter, T. Emerging functions of branched ubiquitin chains. Cell Discov. 7, 6 (2021).
Swatek, K. N. et al. Insights into ubiquitin chain architecture using Ub-clipping. Nature 572, 533–537 (2019).
Haakonsen, D. L. & Rape, M. Branching out: Improved signaling by heterotypic ubiquitin chains. Trends Cell Biol. 29, 704–716 (2019).
Yau, R. G. et al. Assembly and function of heterotypic ubiquitin chains in cell-cycle and protein quality control. Cell 171, 918–933 e920 (2017).
Thrower, J. S., Hoffman, L., Rechsteiner, M. & Pickart, C. M. Recognition of the polyubiquitin proteolytic signal. EMBO J. 19, 94–102 (2000).
Meyer, H. J. & Rape, M. Enhanced protein degradation by branched ubiquitin chains. Cell 157, 910–921 (2014).
Deveraux, Q., Ustrell, V., Pickart, C. & Rechsteiner, M. A 26 S protease subunit that binds ubiquitin conjugates. J. Biol. Chem. 269, 7059–7061 (1994).
Husnjak, K. et al. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 453, 481–488 (2008).
Schreiner, P. et al. Ubiquitin docking at the proteasome through a novel pleckstrin-homology domain interaction. Nature 453, 548–552 (2008).
Shi, Y. et al. Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome. Science 351, 6275 (2016).
Boughton, A. J., Liu, L., Lavy, T., Kleifeld, O. & Fushman, D. A novel recognition site for polyubiquitin and ubiquitin-like signals in an unexpected region of proteasomal subunit Rpn1. J. Biol. Chem. 297, 101052 (2021).
Young, P., Deveraux, Q., Beal, R. E., Pickart, C. M. & Rechsteiner, M. Characterization of two polyubiquitin binding sites in the 26 S protease subunit 5a. J. Biol. Chem. 273, 5461–5467 (1998).
VanderLinden, R. T., Hemmis, C. W., Yao, T., Robinson, H. & Hill, C. P. Structure and energetics of pairwise interactions between proteasome subunits RPN2, RPN13, and ubiquitin clarify a substrate recruitment mechanism. J. Biol. Chem. 292, 9493–9504 (2017).
Hamazaki, J. et al. A novel proteasome interacting protein recruits the deubiquitinating enzyme UCH37 to 26S proteasomes. EMBO J. 25, 4524–4536 (2006).
Qiu, X. B. et al. hRpn13/ADRM1/GP110 is a novel proteasome subunit that binds the deubiquitinating enzyme, UCH37. EMBO J. 25, 5742–5753 (2006).
Yao, T. et al. Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nat. Cell Biol. 8, 994–1002 (2006).
Chen, X., Lee, B. H., Finley, D. & Walters, K. J. Structure of proteasome ubiquitin receptor hRpn13 and its activation by the scaffolding protein hRpn2. Mol. Cell 38, 404–415 (2010).
Boughton, A. J., Krueger, S. & Fushman, D. Branching via K11 and K48 bestows ubiquitin chains with a unique interdomain interface and enhanced affinity for proteasomal subunit Rpn1. Structure 28, 29–43.e26 (2020).
Chen, X. et al. Structure of hRpn10 bound to UBQLN2 UBL illustrates basis for complementarity between shuttle factors and substrates at the proteasome. J. Mol. Biol. 431, 939–955 (2019).
Chen, X. et al. Structures of Rpn1 T1:Rad23 and hRpn13:hPLIC2 reveal distinct binding mechanisms between substrate receptors and shuttle factors of the proteasome. Structure 24, 1257–1270 (2016).
Deol, K. K. et al. Proteasome-bound UCH37/UCHL5 debranches ubiquitin chains to promote degradation. Mol. Cell 80, 796–809.e799 (2020).
Song, A. et al. Branched ubiquitin chain binding and deubiquitination by UCH37 facilitate proteasome clearance of stress-induced inclusions. Elife 10, e72798 (2021).
Hung, K. Y. S. et al. Allosteric control of Ubp6 and the proteasome via a bidirectional switch. Nat. Commun. 13, 838 (2022).
Wehmer, M. et al. Structural insights into the functional cycle of the ATPase module of the 26S proteasome. Proc. Natl Acad. Sci. USA 114, 1305–1310 (2017).
Huang, X., Luan, B., Wu, J. & Shi, Y. An atomic structure of the human 26S proteasome. Nat. Struct. Mol. Biol. 23, 778–785 (2016).
Bashore, C. et al. Ubp6 deubiquitinase controls conformational dynamics and substrate degradation of the 26S proteasome. Nat. Struct. Mol. Biol. 22, 712–719 (2015).
Aufderheide, A. et al. Structural characterization of the interaction of Ubp6 with the 26S proteasome. Proc. Natl Acad. Sci. USA 112, 8626–8631 (2015).
Zhang, S. et al. USP14-regulated allostery of the human proteasome by time-resolved cryo-EM. Nature 605, 567–574 (2022).
Mansour, W. et al. Disassembly of Lys11 and mixed linkage polyubiquitin conjugates provides insights into function of proteasomal deubiquitinases Rpn11 and Ubp6. J. Biol. Chem. 290, 4688–4704 (2015).
Vaden, J. H. et al. Ubiquitin-specific protease 14 regulates c-Jun N-terminal kinase signaling at the neuromuscular junction. Mol. Neurodegener. 10, 3 (2015).
Lee, B. H. et al. USP14 deubiquitinates proteasome-bound substrates that are ubiquitinated at multiple sites. Nature 532, 398–401 (2016).
Martinez-Fonts, K. et al. The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates. Nat. Commun. 11, 477 (2020).
Lam, Y. A., Lawson, T. G., Velayutham, M., Zweier, J. L. & Pickart, C. M. A proteasomal ATPase subunit recognizes the polyubiquitin degradation signal. Nature 416, 763–767 (2002).
Paraskevopoulos, K. et al. Dss1 is a 26S proteasome ubiquitin receptor. Mol. Cell 56, 453–461 (2014).
Zhaolong, W. et al. Hidden dynamics of proteasome autoregulation discovered by cryo-EM data-driven deep learning. Preprint at bioRxiv https://doi.org/10.1101/2020.12.22.423932 (2022).
Saeki, Y., Sone, T., Toh-e, A. & Yokosawa, H. Identification of ubiquitin-like protein-binding subunits of the 26S proteasome. Biochem Biophys. Res Commun. 296, 813–819 (2002).
Matyskiela, M. E., Lander, G. C. & Martin, A. Conformational switching of the 26S proteasome enables substrate degradation. Nat. Struct. Mol. Biol. 20, 781–788 (2013).
Sledz, P. et al. Structure of the 26S proteasome with ATP-gammaS bound provides insights into the mechanism of nucleotide-dependent substrate translocation. Proc. Natl Acad. Sci. USA 110, 7264–7269 (2013).
Unverdorben, P. et al. Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome. Proc. Natl Acad. Sci. USA 111, 5544–5549 (2014).
Chen, S. et al. Structural basis for dynamic regulation of the human 26S proteasome. Proc. Natl Acad. Sci. USA 113, 12991–12996 (2016).
Ding, Z. et al. High-resolution cryo-EM structure of the proteasome in complex with ADP-AlFx. Cell Res 27, 373–385 (2017).
Zhu, Y. et al. Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome. Nat. Commun. 9, 1360 (2018).
Eisele, M. R. et al. Expanded coverage of the 26S proteasome conformational landscape reveals mechanisms of peptidase gating. Cell Rep. 24, 1301–1315.e1305 (2018).
de la Pena, A. H., Goodall, E. A., Gates, S. N., Lander, G. C. & Martin, A. substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis-driven translocation. Science 362, eaav0725 (2018).
Dong, Y. et al. Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565, 49–55 (2019).
Chen, X. et al. Cryo-EM reveals unanchored M1-ubiquitin chain binding at hRpn11 of the 26S proteasome. Structure 28, 1206–1217 e1204 (2020).
Ding, Z. et al. Structural snapshots of 26S proteasome reveal tetraubiquitin-induced conformations. Mol. Cell 73, 1150–1161 e1156 (2019).
Galan, J. M. & Haguenauer-Tsapis, R. Ubiquitin lys63 is involved in ubiquitination of a yeast plasma membrane protein. EMBO J. 16, 5847–5854 (1997).
Kim, H. C. & Huibregtse, J. M. Polyubiquitination by HECT E3s and the determinants of chain type specificity. Mol. Cell Biol. 29, 3307–3318 (2009).
Ohtake, F., Tsuchiya, H., Tanaka, K. & Saeki, Y. Methods to measure ubiquitin chain length and linkage. Methods Enzymol. 618, 105–133 (2019).
Tsuchiya, H., Tanaka, K. & Saeki, Y. The parallel reaction monitoring method contributes to a highly sensitive polyubiquitin chain quantification. Biochem Biophys. Res Commun. 436, 223–229 (2013).
Pintilie, G. et al. Measurement of atom resolvability in cryo-EM maps with Q-scores. Nat. methods 17, 328–334 (2020).
Lange, O. F. et al. Recognition dynamics up to microseconds revealed from an RDC-derived ubiquitin ensemble in solution. Science 320, 1471–1475 (2008).
Gadhave, K., Kumar, P., Kapuganti, S. K., Uversky, V. N. & Giri, R. Unstructured biology of proteins from ubiquitin-proteasome system: Roles in cancer and neurodegenerative diseases. Biomolecules 10, 796 (2020).
Lu, X. et al. Structure of the Rpn13-Rpn2 complex provides insights for Rpn13 and Uch37 as anticancer targets. Nat. Commun. 8, 15540 (2017).
Lagerwaard, I. M., Albanese, P., Jankevics, A. & Scheltema, R. A. Xlink mapping and analySis (XMAS) - smooth integrative modeling in chimeraX. Preprint at bioRxiv. https://doi.org/10.1101/2022.04.21.489026 (2022).
Vander Linden, R. T. et al. Structural basis for the activation and inhibition of the UCH37 deubiquitylase. Mol. Cell 57, 901–911 (2015).
Sahtoe, D. D. et al. Mechanism of UCH-L5 activation and inhibition by DEUBAD domains in RPN13 and INO80G. Mol. Cell 57, 887–900 (2015).
Lu, X., Ebelle, D. L., Matsuo, H. & Walters, K. J. An extended conformation for K48 ubiquitin chains revealed by the hRpn2:Rpn13:K48-diubiquitin structure. Structure 28, 495–506 e493 (2020).
Osei-Amponsa, V. & Walters, K. J. Proteasome substrate receptors and their therapeutic potential. Trends Biochem Sci. 47, 950–964 (2022).
Zhang, N. et al. Structure of the s5a:k48-linked diubiquitin complex and its interactions with rpn13. Mol. Cell 35, 280–290 (2009).
Yao, T. & Cohen, R. E. A cryptic protease couples deubiquitination and degradation by the proteasome. Nature 419, 403–407 (2002).
Worden, E. J., Dong, K. C. & Martin, A. An AAA motor-driven mechanical switch in Rpn11 controls deubiquitination at the 26S proteasome. Mol. Cell 67, 799–811.e798 (2017).
Cundiff, M. D. et al. Ubiquitin receptors are required for substrate-mediated activation of the proteasome’s unfolding ability. Sci. Rep. 9, 14506 (2019).
Inobe, T., Fishbain, S., Prakash, S. & Matouschek, A. Defining the geometry of the two-component proteasome degron. Nat. Chem. Biol. 7, 161–167 (2011).
Smith, D. M. et al. Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s alpha ring opens the gate for substrate entry. Mol. Cell 27, 731–744 (2007).
Amann, S. J. et al. Structural basis for the ubiquitin chain recognition of the human 26S proteasome. Preprint at bioRxiv. https://doi.org/10.1101/2025.04.07.647569 (2025).
Amann, S. J. et al. PITHD1: An endogenous inhibitor of the 26S proteasome during cellular dormancy. Preprint at bioRxiv https://doi.org/10.1101/2024.12.04.626795 (2024).
Michel, M. A. et al. Assembly and specific recognition of k29- and k33-linked polyubiquitin. Mol. Cell 58, 95–109 (2015).
Wang, X. et al. Mass spectrometric characterization of the affinity-purified human 26S proteasome complex. Biochemistry 46, 3553–3565 (2007).
Wilkinson, K. D., Gan-Erdene, T. & Kolli, N. Derivitization of the C-terminus of ubiquitin and ubiquitin-like proteins using intein chemistry: methods and uses. Methods Enzymol. 399, 37–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife https://doi.org/10.7554/eLife.42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Punjani, A. & Fleet, D. J. 3D variability analysis: Resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr D. Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D. Biol. Crystallogr 66, 486–501 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr D. Struct. Biol. 74, 519–530 (2018).
Wang, X. et al. The proteasome-interacting Ecm29 protein disassembles the 26S proteasome in response to oxidative stress. J. Biol. Chem. 292, 16310–16320 (2017).
Schnirch, L. et al. Expanding the depth and sensitivity of cross-link identification by differential ion mobility using high-field asymmetric waveform ion mobility spectrometry. Anal. Chem. 92, 10495–10503 (2020).
Heissel, S. et al. Fast and accurate disulfide bridge detection. Mol. Cell Proteom. 23, 100759 (2024).
Combe, C. W., Fischer, L. & Rappsilber, J. xiNET: cross-link network maps with residue resolution. Mol. Cell Proteom. 14, 1137–1147 (2015).
Lin, S. W., Nguyen, V. Q. & Lin, S. Preparation of Cas9 ribonucleoproteins for genome editing. Bio Protoc. 12, e4420 (2022).
Acknowledgements
We thank Dr. Yasushi Saeki at the Tokyo Metropolitan Institute of Medical Science, Japan, for sharing the Ub-AQUA peptides. We thank Academia Sinica Common Mass Spectrometry Facilities (AS-CFII-111-209) and Academia Sinica Cryo-EM Center (AS-CFII-111-210) for data collection; both are funded by the Academia Sinica Core Facility and Innovative Instrument Project. Taiwan Protein Project (AS-KPQ-109-TPP2) is acknowledged for supporting the cryo-EM facility. We thank the protein core facility, biophysics core facility and imaging core facility of the Institute of Biological Chemistry, Academia Sinica, for supporting the sample preparation and analyses. This work was supported by Academia Sinica intramural fund to SL and STDH, an Academia Sinica Career Development Award to STDH (AS-CDA-109-L08), an Academia Sinica Career Development Award to KPW (AS-CDA-110-L03), an Academia Sinica Investigator Award to STDH (AS-IV-114-L04), and funding from the National Science and Technology Council (NSTC), Taiwan (110-2311-B-001-013-MY3 and 113-2311-B-001-017-MY3 to STDH; 112-2811-B-001-006, 113-2811-B-001-019 and 114-2811-B-001-041 to TC).
Author information
Authors and Affiliations
Contributions
Conceptualization: P.D., S.-T.D.H., Investigation: P.D., S.N.C., T.C., Y.S.W., H.A.S., J.Y.C.H., M.C.T., S.L., R.V., S.-T.D.H., Data visualization: P.D., T.C., Y.S.W., S.-T.D.H., Methodology: P.D., S.Y.L., S.L., R.V., Y.C.C., K.P.W., S.-T.D.H., Project administration: P.D., S.-T.D.H., Supervision of the study: P.D., S.-T.D.H., Funding acquisition: S.L., K.P.W., S.-T.D.H., Writing–original draft: P.D., S.-T.D.H., Writing–review/editing: P.D., T.C., Y.S.W., S.L., R.V., S.-T.D.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kitaik Lee, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Draczkowski, P., Chen, SN., Chen, T. et al. Structural basis of K11/K48-branched ubiquitin chain recognition by the human 26S proteasome. Nat Commun 16, 9094 (2025). https://doi.org/10.1038/s41467-025-64719-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64719-x