Abstract
The vasopressin V2 receptor (V2R), a class A G protein-coupled receptor, is essential for regulating body water homeostasis. V2R antagonists have emerged as promising treatments for hyponatremia; however, the absence of structural information for antagonist-bound V2R hampers our understanding of antagonist recognition and the targeted design of V2R antagonists. In this study, we present two cryo-electron microscopy structures of inactive V2R bound to the clinically approved antagonists tolvaptan and conivaptan. Combined with functional analyses and molecular dynamic simulations, these structures reveal distinct binding poses: tolvaptan is deeply inserted within the binding pocket, whereas conivaptan is positioned at a shallower depth. Integrated analyses further define critical pharmacophoric features governing antagonist activity and unveil a TM7 helical conformation-dependent antagonism mechanism that is distinct from classical GPCR inactivation modes. Our findings deepen understanding of antagonist recognition and antagonism of V2R, providing a foundation for the development of V2R-targeted therapies.
Similar content being viewed by others
Introduction
The vasopressin V2 receptor (V2R), encoded by AVPR2, is a class A G protein-coupled receptor (GPCR) featuring the characteristic seven-transmembrane domain with an extracellular N-terminus and intracellular C-terminus. It is predominantly expressed in the kidney tubule, specifically in the distal convoluted tubule and collecting ducts1. V2R plays an essential role in the regulation of body water homeostasis by responding to the endogenous hormone arginine vasopressin (AVP), also known as antidiuretic hormone2,3. Upon AVP binding, activated V2R stimulates the Gαs subunit, which increases intracellular cAMP levels and activates protein kinase A (PKA). PKA then phosphorylates aquaporin-2 (AQP2) in renal collecting duct principal cells, inducing its translocation to the apical membrane to mediate water reabsorption2 (Fig. 1a). The importance of V2R is underscored by its involvement in various pathologies linked to water balance dysregulation4,5. For instance, X-linked nephrogenic diabetes insipidus (XNDI) is associated with loss-of-function pathogenic variants in AVPR2, while nephrogenic syndrome of inappropriate antidiuresis (NSIAD) is caused by gain-of-function mutations3, leading to impaired water excretion and subsequent hyponatremia (serum sodium concentration <135 mmol per liter)—the most common electrolyte derangement occurring in hospitalized patients6. Besides genetic disorders of V2R, the unsuppressed release of AVP or its enduring activation on V2R results in the syndrome of inappropriate antidiuretic hormone ADH release (SIADH), which is also closely associated with hyponatremia7. The pathophysiology of hyponatremia highlights the critical need for effective therapeutic strategies to regulate AVP signaling, further emphasizing the importance of developing V2R antagonists for clinical use.
a Schematic illustration of the regulatory function of V2R by endogenous hormone arginine vasopressin (AVP) and the antagonists tolvaptan and conivaptan. b Optimized construct for the structure determination of antagonist-bound V2R. c, d Orthogonal views of the density maps (left panel) and the models (right panel) of tolvaptan-V2R-Fab (c) and conivaptan-V2R-Fab complex (d). Tolvaptan-bound V2R is in green, tolvaptan is in pink, conivaptan-bound V2R is in purple, conivaptan is in orange, Fab-LC is in dark blue, Fab-HC is in cyan, BRIL is in gray. Components of V2R complexes are colored as indicated, with the EM density maps for tolvaptan and conivaptan displayed as transparent surfaces at contour levels of 0.40 and 0.26, respectively. LC light chain, HC heavy chain.
Currently, only two small-molecule V2R antagonists—tolvaptan and conivaptan—are approved for the treatment of hyponatremia8,9. Pharmacological profiling reveals that tolvaptan exhibits high selectivity for V2R, whereas conivaptan demonstrates dual antagonism of both V2R and V1aR10. However, these drugs pose significant safety concerns, such as hepatotoxicity risks associated with tolvaptan11. Clinical administration of both pharmacological agents necessitates close therapeutic monitoring, while their prolonged use is typically contraindicated due to potential adverse effects12. Therefore, there is an urgent need to develop new V2R antagonists with improved safety, efficacy, and selectivity. An accurate structural model of V2R can serve as a template for aiding the design of targeted antagonists.
Recent advancements in cryo-electron microscopy (cryo-EM) have enabled the elucidation of the AVP-V2R-Gs complex structures13,14,15. These structural insights enhance our understanding of AVP binding and V2R activation mechanisms. Nevertheless, detailed structural information regarding the interaction between antagonists and V2R remains unclear, hindering the rational design of V2R antagonists. In this study, we present the cryo-EM structures of V2R bound to two antagonists, tolvaptan and conivaptan. Combined with functional analysis, these structures uncover the mechanisms underlying antagonist recognition and antagonism of V2R, offering valuable templates for the design of targeted V2R antagonistic drugs.
Results
Cryo-EM structures of V2R bound to antagonists
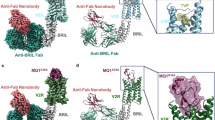
To facilitate structure determination of the antagonist-bound V2R, a thermostabilized apocytochrome b562RIL (BRIL) was inserted at the junction site between A2345.67 and A2646.28 within the intracellular loop 3 (ICL3) of the human V2R cDNA (Fig. 1b). The engineered V2R was expressed in sf9 cells and purified in the presence of tolvaptan and conivaptan. The samples were incubated with an anti-BRIL Fab16 to facilitate accurate image alignment for cryo-EM structure determination. Two cryo-EM structures of V2R bound to tolvaptan and conivaptan were determined at resolutions of 3.08 and 2.94 Å, respectively (Fig. 1c, d, Supplementary Figs. 1, 2, Supplementary Table 1). High-quality cryo-EM maps allow us to accurately model the antagonists, receptors, and anti-BRIL Fab (Supplementary Fig. 3). The overall structures of V2R consist of a transmembrane domain of seven transmembrane helices (TM1-TM7) linked by three intracellular loops (ICLs) and three extracellular loops (ECLs). The structures of antagonist-bound V2R are highly superimposable, with root-mean-square deviation (RMSD) value of 0.8 Å for the Cα atoms.
Binding mode of tolvaptan
Tolvaptan is a non-peptide V2R antagonist and has been approved for the treatment of hyponatremia and autosomal dominant polycystic kidney disease (ADPKD)17,18. It adopts a tilted insertion mode within the V2R orthosteric pocket (Fig. 2a). Its benzazepine group (designated as region A) and its neighboring methylphenyl group (region B) are oriented towards the extracellular surface, while the terminal methylphenyl group (region C) is positioned at the bottom of the pocket (Fig. 2a). The binding pose of tolvaptan closely aligns with the lowest-energy conformation observed in molecular dynamics (MD) simulations (Supplementary Fig. 4a, b), showing an RMSD of 1.25 Å (Supplementary Fig. 4b), indicating a stable binding mode of tolvaptan in V2R.
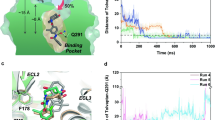
a Cross-section of the tolvaptan-binding pocket in V2R. Tolvaptan-bound V2R is in green, tolvaptan is in pink. b 2D presentation of interactions between tolvaptan and residues in the orthosteric binding pocket of V2R. The moieties of tolvaptan and their interacting residues are indicated by different colors for clarity. c, e Detailed interactions between tolvaptan and ligand-binding pocket residues in V2R. The hydrogen bond is highlighted with a cyan dashed line. d Effects of pocket residue mutations in V2R on the antagonistic activity of tolvaptan. Data are presented as means ± SEM from three independent experiments conducted in triplicate. One-way ANOVA followed by Dunnett’s multiple comparisons test was used to determine the P value compared with the response of WT. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001 were considered statistically significant. The exact P values are as follows: P = 0.9994 for V882.53A; P = 0.5906 for Q922.57A; P = 0.9475 for V932.58A; P = 0.9997 for M1203.33A; P = 0.0014 for I2095.42A; P = 0.5276 for W2846.48A; P = 0.0244 for Q2916.55A; P = 0.1013 for M3117.39A; P < 0.0001 for K1163.29A, M1233.36A, Q1744.60A, Y2055.38A, V2065.39A, F2876.51A, and F2886.52A.
At the upper orthosteric binding pocket, the benzazepine moiety in region A of tolvaptan is kinked and bends downward, creating intramolecular hydrophobic interactions with the neighboring methylphenyl group in region B. Both regions are surrounded by hydrophobic residues in TM3, TM5, and TM6, including M1203.33, M1233.36, V2065.39, I2095.42, F2876.51, F2886.52 and the aromatic ring of Y2055.38 (Fig. 2b, c). The majority of these residues make substantial contributions to the antagonistic activity of tolvaptan (Fig. 2d, Supplementary Fig. 5a–c). Notably, alanine substitutions of Y2055.38 and F2876.51 exhibit an over 200-fold decrease in the tolvaptan activity (Fig. 2d, Supplementary Table 2). These results reproduce previously reported mutagenesis data19, supporting the importance of these hydrophobic residues in the activity of tolvaptan. In addition to the identified hydrophobic residues, the hydrophilic residue Q2916.55 forms a hydrogen bond with the hydroxy group in region A, while the side chain of K1163.29 and Q1744.60 establishes a stable hydrogen bond with the carbonyl group in region B (Fig. 2b, c). These interactions are consistent with the MD simulation results, showing hydrogen bond occupancies of 45.38%, 16.63% and 94.93%, respectively (Supplementary Fig. 4c). Furthermore, the side chain of Q1744.60 also builds an intramolecular hydrogen bond with the hydroxyl group in the side chain of Y2055.38 (Fig. 2c, Supplementary Fig. 4b), stabilizing their conformation and enhancing the interaction between Q1744.60 and tolvaptan. Consequently, alanine mutations at Q2916.55, K1163.29, Q1744.60, and Y2055.38 remarkably attenuated tolvaptan-induced V2R antagonism, highlighting their critical roles in mediating the antagonistic activity of tolvaptan (Fig. 2c, d). These findings highlight the importance of region A and region B in the antagonistic activity of tolvaptan.
At the bottom of the binding pocket, Q922.57 engages in a hydrogen bond with the secondary amine group in region B. Analysis of MD simulations demonstrated a high hydrogen bond occupancy of 98.66% (Supplementary Fig. 4c). Region C of tolvaptan occupies a roughly hydrophobic cavity composed of V882.53, V932.58, W2846.48, and M3117.39 (Fig. 2e). However, these residues show negligible impacts on the antagonistic activity of tolvaptan (Fig. 2d), suggesting that region C makes a negligible contribution to the activity of tolvaptan. Consequently, these structural observations and mutational analyses support the contention that the physicochemical environment at the upper orthosteric binding pocket of V2R is the determinant of tolvaptan’s antagonistic activity by interacting with region A and region B. This conclusion is further supported by per-residue free energy decomposition analysis (Supplementary Fig. 4g).
Recognition of conivaptan by V2R
Compared to tolvaptan, another vaptan or V2R antagonist conivaptan presents a different binding pose. In contrast to tolvaptan, conivaptan is positioned in a relatively shallow binding pocket. Its benzoimidazoazepine moiety (region A) approaches the benzazepine group of tolvaptan, while regions B and C, which consist of a benzene group and a biphenyl group, are connected by an amide bond and oriented toward ECL1 (Fig. 3a, b).
a Cross-section of the conivaptan-binding pocket in V2R. Conivaptan-bound V2R is in purple, conivaptan is in orange. b Comparison of the binding pose of conivaptan and tolvaptan. The movement direction of ECL1 in conivaptan-V2R relative to that in tolvaptan is indicated by a red dashed arrow. c 2D presentation of interactions between conivaptan and the binding pocket residues in V2R. The moieties of conivaptan and their interacting residues are indicated by different colors for clarity. d, f Detailed interactions between conivaptan and ligand-binding pocket residues in V2R. The hydrogen bond is highlighted with a cyan dashed line. e Effects of pocket residue mutations in V2R on the antagonistic activity of conivaptan. Data are presented as means ± SEM from three independent experiments conducted in triplicate. One-way ANOVA followed by Dunnett’s multiple comparisons test was used to determine the P values compared with the response of WT. *P < 0.05; **P < 0.01; ***P < 0.001 and ****P < 0.0001 were considered statistically significant. The exact P values are as follows: P = 0.0041 for I2095.42A; P = 0.0002 for F2886.52A; P = 0.4160 for Q2916.55A; P = 0.1178 for F3077.35A; P = 0.4099 for M3117.39A; P < 0.0001 for F105ECL1A, M1203.33A, M1233.36A, F1784.64A, Y2055.38A, V2065.39A, and F2876.51A.
In detail, region A and B of conivaptan are surrounded by hydrophobic residues, including M1203.33, M1233.36, F1784.64, Y2055.38, V2065.39, I2095.42, F2876.51, F2886.52, and M3117.39 (Fig. 3c, d). Intriguingly, in contrast to tolvaptan-bound V2R, the side chain of F1784.64 undergoes a significant displacement to form hydrophobic interactions with three aromatic rings in regions A, B, and C (Fig. 3b). The structural observation is supported by the mutational analysis that substituting the majority of these residues with alanine dramatically impairs the antagonistic activity of conivaptan (Fig. 3e, Supplementary Fig. 5d–f).
Additionally, structural comparison of V2R bound to the two vaptans reveals distinct interaction patterns involving region C: tolvaptan’s region C extends deeper into the binding pocket, whereas conivaptan’s region C directly engages F105ECL1 (Fig. 3b). This represents the largest structural discrepancy between the tolvaptan- and conivaptan-bound V2R structures. The biphenyl moiety in region C of conivaptan forms π-π interactions with F105ECL1 and F3077.35 through its two benzene rings, respectively (Fig. 3f). Replacing F105ECL1 with alanine results in an 80-fold decrease in conivaptan activity, indicative of the importance of region C on conivaptan activity (Fig. 3e, Supplementary Table 3). The only hydrogen bond interaction identified in the structure occurs between Q2916.55 and the carbonyl group in region B (Fig. 3c, d). Nonetheless, mutation analysis indicates that replacing Q2916.55 with alanine exerts a negligible effect on the activity of conivaptan (Fig. 3e).
Unlike the stable binding mode of tolvaptan, the MD simulation analysis reveals a more dynamic binding characteristic of conivaptan within V2R. Regions B and C of conivaptan move toward the core of the helix bundle, with an RMSD of 4.06 Å between the representative simulated conformation and the cryo-EM structure (Supplementary Fig. 4e). Globally, binding free energy decomposition analysis showed that MD simulations largely confirmed cryo-EM identified residues (Supplementary Fig. 4h). A notable exception was Q922.57, which contributed significantly to conivaptan binding by forming a hydrogen bond with its amide group (occupancy: 39.93%, Supplementary Fig. 4f), a feature that was not apparent in static structure analysis. The functional significance of Q922.57 was confirmed by mutagenesis studies, with Q922.57A mutation causing a 31-fold loss in conivaptan’s antagonistic activity (Supplementary Table 3). These MD simulation results offer a plausible explanation for the relatively weak density observed in region C, emphasizing the flexibility of its interaction with V2R, and also suggest an alternative binding conformation of conivaptan.
Inactive conformation of V2R
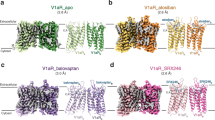
A pairwise structural comparison of the antagonist-bound with the AVP-bound V2R (PDB: 7DW9) confirms that the former is in an inactive state. Upon binding to antagonists, the extreme cytoplasmic end of TM6 of V2R undergoes a pronounced inward displacement of 8.6-8.9 Å, as measured at the Cα atom of V2666.30, relative to the AVP-bound V2R. This displacement represents the hallmark conformational change associated with the inactivation of class A GPCRs. Additionally, the intracellular portion of TM7 exhibits an outward shift compared to AVP-bound V2R (Fig. 4a, b, Supplementary Fig. 6a, b). These conformational changes close the cytoplasmic cavity, preventing its opening and the coupling of the G protein heterotrimer, which aligns with the classic conformational alterations associated with class A GPCR antagonism (Supplementary Fig. 7a). In addition, structure superposition reveals notable conformational changes in the extracellular region of V2R. Upon binding of tolvaptan, the extracellular end of TM2 undergoes an unwinding of one helical turn, accompanied by a half-helical downward movement of TM3, relative to AVP-bound V2R (Supplementary Fig. 7b). Meanwhile, outward movements of the extracellular ends of TM1, TM6, and TM7 are also observed in the tolvaptan-bound V2R compared to the AVP-bound V2R (Supplementary Fig. 7c). These structural observations demonstrate the characteristics of the inactive V2R in comparison to the active receptor.
a Comparison of the overall structures of tolvaptan-V2R and AVP-V2R complexes in inactive and active conformations, respectively. AVP-bound V2R is in blue, AVP is in light blue, tolvaptan-bound V2R is in green, tolvaptan is in pink. The binding depths of tolvaptan and AVP are marked by corresponding colored dashed lines. b Conformational changes in TM6 and TM7 between inactive and active V2R. The movement direction of TM6 in tolvaptan-V2R relative to AVP-V2R complexes is indicated by a red dashed line. c Structural superposition of tolvaptan and AVP in the V2R pocket. d, e Conformational changes of V2R during the conformational transition from the active to inactive state. The hydrogen bond between Tyr2 of AVP and L3127.40 in TM7 is indicated by a cyan dashed line. The movements of the residues are marked by red dashed arrows.
Antagonism mechanism of V2R
The structural comparison of inactive and active V2R provides insights into the mechanism of antagonism at V2R. The overall binding pose of tolvaptan resembles that of the AVP segment Tyr2-Phe3 but is inserted deeper into the helix core of V2R (Fig. 4c). Its methylphenyl moiety sterically approaches the side chain of Tyr2 of AVP, which is crucial for AVP activity13, both of which are positioned at the bottom of the binding pocket (Fig. 4c). It was reported that the hydrogen bond between Tyr2 of AVP and the backbone carbonyl group of L3127.40 in V2R pulls and distorts the helix of the TM7 segment between L3127.40 and S3157.43 13. However, the corresponding interaction is absent due to the lack of a hydrogen bond donor in the methylphenyl moiety of tolvaptan. Accordingly, the helical continuity of TM7 is maintained upon tolvaptan binding (Fig. 4d). Meanwhile, in contrast to AVP-bound V2R, the TM7 segment at the bottom of the binding pocket is positioned notably outward upon tolvaptan binding. These conformational changes on TM7 may propagate antagonistic effects to the cytoplasmic end of TM6, which adopts a conformation incompatible with G protein coupling.
Unlike tolvaptan, conivaptan overlaps with the Phe3-Cys6 segment of AVP (Supplementary Fig. 8a). Notably, structural analysis reveals that conivaptan adopts a unique spatial orientation distal from the characteristic TM7 distortion region (L3127.40-S3157.43), exhibiting no direct contact with this structural motif. Instead, our findings suggest that conivaptan may exert its antagonistic effect by occupying the orthosteric binding pocket, thereby sterically hindering the critical interaction between Tyr2 in AVP and the distortion region of TM7 (Supplementary Fig. 8b).
The side chain rotamer of the toggle switch residue W6.48 has been proposed to play a toggling role in class A GPCR activation20. Compared to the AVP-V2R structure, the side chain of W2846.48 in tolvaptan-bound V2R structures only exhibits a slight upward movement without a significant rotamer switch (Fig. 4d, e). Furthermore, W2846.48 is not critical to the agonistic activity of AVP or the antagonistic effect of tolvaptan (Supplementary Fig. 9a, b). This contrasts with previous reports that mutating W6.48 to alanine eliminates basal activity and significantly impairs agonist-induced GPCR activation21. Additionally, the distance between conivaptan and W2846.48 in conivaptan-bound V2R is excessively large to support any direct interaction (Supplementary Fig. 9c). These findings suggest a toggle switch non-dependent regulatory mechanism of V2R, thereby enhancing our understanding of the regulatory mechanisms underlying GPCR activity.
Discussion
In this study, we resolved two cryo-EM structures of inactive V2R bound to two clinical drugs tolvaptan and conivaptan. The tolvaptan-V2R model was refined using PDB 9HAP as a reference22. Our model is largely consistent with 9HAP in both overall architecture and ligand binding conformation. Additional electron density was observed, enabling the modeling of previously unresolved residues (e.g., F196ECL2, R2025.35 and W2936.57). Combined with mutational analysis, these structures provide insights into the diversity in antagonist recognition and reveal the underlying mechanism of V2R antagonism, which plays a critical role in regulating water homeostasis and cardiovascular health.
Structural analysis reveals that the peptide class A GPCRs have nearly three-fold larger binding cavities than non-peptide GPCRs23. As a member of the peptide class of GPCRs, V2R features a large binding pocket to accommodate the endogenous “spoon-like” cyclic peptide, AVP. This pocket also provides ample open space, allowing for diverse binding poses of small molecule ligands. Indeed, two non-peptide antagonists adopt different binding poses within the orthosteric binding pocket of V2R. Tolvaptan is deeply inserted into the binding pocket, while conivaptan presents a shallower binding pose and extends toward ECL1.
The two antagonists share similar benzazepine-derived pharmacophores, defined as region A. From a structural aspect, despite their markedly different binding poses, region A groups of both antagonists are spatially proximate and crucial for their antagonist activity. In addition, both antagonists exhibit similarities to specific segments of AVP, although the overlapping portions between tolvaptan and AVP differ from those between conivaptan and AVP (Supplementary Fig. 10). MD simulation results demonstrate that the binding pose of tolvaptan remains stable, whereas regions B and C of conivaptan shifted toward the core of the helix bundle, underscoring the dynamic binding behavior of conivaptan. Interestingly, the MD-simulated structure of conivaptan retains partial similarity to AVP, with the overlapping region extending from Tyr2 to Phe3 (Supplementary Fig. 8c), which more closely resembles the binding mode of tolvaptan (Supplementary Fig. 8d). This not only explains the relatively weak density of region C in the cryo-EM conivaptan-bound V2R structure but also implies a dynamic binding mode for conivaptan. These structural insights can be utilized for the rational design of V2R-targeted antagonists.
In addition, region B of tolvaptan appears to contribute to the receptor subtype selectivity. Previous studies demonstrated that introducing substituents such as methyl, chloro, or fluoro groups into the region B substantially enhanced their selectivity for V2R over V1AR (Supplementary Fig. 11)24,25,26. Notably, tolvaptan shows approximately 193-fold greater V2R selectivity compared to its des-methyl analog24. These findings underscore the importance of region B modifications in guiding rational design toward subtype-selective V2R antagonists.
Furthermore, the resolved inactive V2R configurations address a critical gap in understanding V2R antagonism mechanisms. Structural analyses reveal that antagonist binding induces structural reorganization of TM7, restoring helical continuity at the bottom of the binding pocket that becomes distorted upon AVP engagement (Supplementary Fig. 10). This conformational change appears to prevent the necessary conformational changes acquired for G protein coupling. This TM7 helical conformation-dependent antagonism suggests a distinct inactivation mechanism for class A GPCRs, differing from that observed in receptors like the β2-adrenergic receptor and endothelin receptor type A (Supplementary Fig. 12). In summary, the structures of V2R bound to the two antagonists, tolvaptan and conivaptan, together with MD simulation and functional analyses, not only deepen our understanding of V2R pharmacology but also facilitate the development of more effective therapeutic agents.
Limitations of this study
It should be noted that the density corresponding to the biphenyl moiety of conivaptan is relatively weaker compared to other ligand segments. This observation aligns with the dynamic behavior of conivaptan observed in MD simulations, which suggests conformational flexibility in this region. Nevertheless, the overall binding pose of conivaptan is supported by our mutagenesis data, indicating that the resolved structure represents a dominant conformation under the experimental conditions. The alternative binding mode for conivaptan proposed by MD simulation needs further experimental validation.
Methods
Construct
Wild-type human V2R (residues 1–234 and 264–371) was cloned into a pFastBac vector (Invitrogen) using homologous recombination (CloneExpress One Step Cloning Kit, Vazyme) and modified to contain an N-terminal FLAG and C-terminal His8 tags. The strategies of cytochrome b562 RIL (BRIL) fusion and the incorporation of anti-BRIL Fabs were employed to introduce a rigid mass, addressing the molecular weight barrier of GPCRs for cryo-EM structure determination27,28. BRIL was inserted into the ICL3, with the insertion sites optimized to ensure continuous connectivity. Two short linkers derived from A2A adenosine receptor (ARRQL between residue 234 and the N-terminus of BRIL, and ERARSTL between the C-terminus of BRIL and residue 264) were used to further improve the rigid attachment of BRIL to V2R. The anti-BRIL Fab16 was cloned into a pFastBac vector (Invitrogen) with N-terminal His8 tags.
Protein expression and purification
The V2R-BRIL fusion and anti-BRIL Fab were co-expressed in sf9 insect cells using the Bac-to-Bac baculovirus expression system (Thermo Fisher Scientific). Cells were grown in SIM SF expression medium (Sino Biological) at a density of 3 × 106 cells/mL before co-infection with V2R-BRIL and anti-BRIL Fab viruses at a ratio of 1:1. The cells were harvested by centrifugation at 750 g for 20 min after 48 h infection. The cell pellets were resuspended and lysed in lysis buffer containing 20 mM HEPES, pH 7.4, 100 mM NaCl, 5 mM MgCl2, 5 mM CaCl2, and 10% (v/v) glycerol supplemented with EDTA-free protease inhibitor cocktail (TargetMol). The suspension was subsequently incubated with 10 μM V2R antagonist for 30 min. Then the membrane was solubilized by the addition of 0.5% (w/v) lauryl maltose neopentyl glycol (LMNG, Anatrace) and 0.1% (w/v) cholesterol hemisuccinate (CHS, Anatrace) for 2 h at 4 °C. The supernatant was collected by centrifugation at 140,000 × g for 35 min and then incubated with Anti-DYKDDDDK G1 Affinity Beads (GenScript) for 2 h at 4 °C. The resin was separated by centrifugation at 600 × g for 3 min, loaded into a gravity flow column, and washed with 20 column volumes of 20 mM HEPES, pH 7.4, 100 mM NaCl, 1 μM V2R antagonist, 0.1% (w/v) LMNG and 0.02%(w/v) CHS. The resin was further washed with 20 column volumes of 20 mM HEPES, pH 7.4, 100 mM NaCl, 1 μM V2R antagonist, 0.05% (w/v) LMNG and 0.01%(w/v) CHS and finally eluted with 20 mM HEPES, pH 7.4, 100 mM NaCl, 1 μM V2R antagonist, 0.01% (w/v) LMNG, 0.002%(w/v) CHS, and 0.2 mg/mL FLAG peptide. The protein was concentrated using an Amicon Ultra centrifugal filter (molecular weight cut-off of 100 kDa, Millipore) and injected onto a Superdex 200 Increase 10/300 GL column (Cytiva) equilibrated in the buffer containing 20 mM HEPES, pH 7.4, 100 mM NaCl, 1 μM V2R antagonist, 0.00075% (w/v) LMNG, 0.00025% (w/v) glyco-diosgenin (GDN, Anatrace) and 0.00015% (w/v) CHS. The complex fractions were collected and concentrated for electron microscopy experiments.
Cryo-EM grid preparation and data collection
For the preparation of cryo-EM grids, 3 μL of the purified tolvaptan-V2R-Fab and conivaptan-V2R-Fab complexes at the concentrations of 23 mg/mL, 27 mg/mL, and 28 mg/mL, respectively, were applied onto glow-discharged holey carbon grids (Au300, R1.2/1.3, Quantifoil). Excess samples were blotted for 4 s with standard vitrobot filter paper (Ted Pella) under 100% humidity at 4 °C. Then the samples were vitrified by plunging into liquid ethane using a Vitrobot Mark IV (Thermo Fisher Scientific). Cryo-EM data collection was performed on a Titan Krios G4 at 300 kV accelerating voltage in the Advanced Center for Electron Microscopy at Shanghai Institute of Materia Medica, Chinese Academy of Sciences (Shanghai, China). A total of 10,878 movies for the tolvaptan-V2R-Fab complex and 5867 movies for the conivaptan-V2R-Fab complex were collected by a Gatan K3 Summit direct electron detector at a pixel size of 0.824 Å using the EPU software (FEI, Eindhoven, the Netherlands). Movies were obtained at a total dose of 50 e− Å−2 over 3 s exposure with a defocus ranging from −1.0 to 3.0 μm.
Cryo-EM data processing
All dose-fractionated image stacks were processed for beam-induced motion correction using MotionCor229 within RELION 4.030. Subsequent data processing was performed using CryoSPARC 4.4.1.
For the conivaptan-V2R-Fab complex, 1955 micrographs were initially selected for analysis. After blob picking, 1,668,261 particles were extracted. Following three rounds of 2D classification, 747,127 particles were retained for ab-initio reconstruction to generate four reference models. 2D templates were then used for particle picking across all micrographs, resulting in the extraction of 3,702,735 particles, which were subjected to several rounds of hetero refinement. Notably, the map of the solved inactive A2AR complex (EMDB: EMD-25648), along with the four previously generated reference models, were employed as references. A total of 189,727 particles were selected and used to reconstruct a model, achieving a global resolution of 3.31 Å. Following local refinement, the final map exhibited a global resolution of 2.94 Å at an FSC of 0.143.
For the tolvaptan-V2R-Fab complex, 1700 micrographs were initially selected for analysis. After blob picking, 1,440,079 particles were extracted. Following three rounds of 2D classification, 341,192 particles were retained. Using the inactive A2AR map and four previously generated reference models from the conivaptan-V2R-Fab complex dataset, we identified a specific class, which was then used for Topaz training. The Topaz-trained model was subsequently applied for particle picking across all micrographs, leading to the extraction of 6,721,794 particles, which underwent several rounds of hetero refinement. A total of 326,036 particles were selected and used to reconstruct a model with a global resolution of 3.30 Å. After local refinement, the final map achieved a global resolution of 3.08 Å at an FSC of 0.143.
Model building and refinement
The V2R structure from Alphafold2 prediction was used as the initial reference model for receptor building31. The structures of the light and heavy chains of anti-BRIL Fab were derived from PDB entry 6CBV16 and rigid-body fitted into the density using UCSF Chimera32. The model was then modified using COOT33 and ISOLDE34, followed by refinement in PHENIX35. Final model statistics were validated using Comprehensive Validation (cryo-EM) in PHENIX35, and are provided in Supplementary Information, Table S1.
Radioligand binding assay
[3H]-vasopressin (specific activity 89.2 Ci·mmol−1) was obtained from PerkinElmer (Boston, MA, USA). Tolvaptan, Conivaptan and unlabeled vasopressin were purchased from MCE (MedChemExpress, NJ, USA). All other reagents were of analytical grade and obtained from standard commercial sources.
Affinity at the V2R-BRIL fusion or the V2R were determined on membranes from HEK293T cells transiently transfected with the V2R-BRIL fusion or the wild type V2R. The collected cells were homogenized in a chilled (4 °C) assay buffer (100 mM NaCl, 10 mM MgCl2, 0.1 mg/ml bacitracin, 1 mg/ml BSA, 20 mM Tris-HCl, pH 7.4). Afterward, membranes and the cytosolic fraction were separated by centrifugation at 48,000 g at 4 °C for 25 min. The pellet was resuspended in 3 mL of the assay buffer, and the homogenization and centrifugation step were repeated. Subsequently, membrane aliquots containing 5 μg of protein were incubated in a total volume of 250 μL assay buffer at 25 °C for 90 min with 2.3 nM [3H]-vasopressin to detect the displacements of radioligands in the presence of different concentrations of Tolvaptan or Conivaptan. Nonspecific binding was determined in the presence of 10 μM unlabeled vasopressin and represented less than 10% of the total binding. Incubations were terminated by rapid vacuum filtration to separate the bound and free radioligand through 96-well GF/C filter plates (UniFilter-96 GF/C, PerkinElmer) using a PerkinElmer Filtermate-harvester. The radioactivity was determined by a β scintillation counter (MicroBeta2 Microplate Counters).
Site-directed mutagenesis
The results of radioligand binding experiments showed that BRIL fusion has no impact on the binding affinity of tolvaptan and conivaptan to V2R (Supplementary Fig. 13). Therefore, we performed mutagenesis on the WT without BRIL. Site-directed mutants were generated using polymerase chain reaction with pcDNA3.1(+)-human V2R-His as the template. This study included a total of 18 mutations: V882.53A, Q922.57A, V932.58A, W992.64A, F105ECL1A, Q1744.60A, F1784.64A, Q1193.32A, M1203.33A, M1233.36A, Y2055.38A, V2065.39A, I2095.42A, W2846.48A, F2876.51A, F2886.52A, Q2916.55A, M3117.39A. The plasma membrane expression levels of both wild-type and mutant V2R proteins were quantified using Western blot analysis (Supplementary Fig. 14). Cell lysates were prepared from transfected cells using cell lysis buffer (Beyotime). Proteins were separated by SDS-PAGE and transferred onto nitrocellulose membranes (Pall Corporation). Immunoblotting was performed using anti-Tubulin (Proteintech) and anti-His-Tag (Proteintech) as primary antibodies. Horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (H + L) (Beyotime) and goat anti-mouse IgG (H + L) (Beyotime) were used as secondary antibodies.
Cell culture and transfection
HEK293T cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with stable glutamine, 10% newborn calf serum, streptomycin, and penicillin at 37 °C in a humidified 5% CO2 atmosphere. When the cells reached 60%–70% confluency, they were transfected with plasmid DNA using GP-transfect-Mate (GenePharma, Shanghai, China), according to the manufacturer’s instructions. After 48 h, the cells were collected for the cAMP assays.
cAMP assay
Experiments were performed using the cAMP kit (PerkinElmer, Shanghai, China). Vasopressin, the endogenous V2R agonist, was diluted with cAMP reaction buffer to create a gradient concentration. These diluted solutions were subsequently mixed with the HEK293T cells transiently transfected with wild-type and mutant V2 receptors at 37 °C for 30 min. Cellular cAMP level was subsequently determined according to the manufacturer’s instructions. cAMP production in response to the maximum vasopressin concentration was set as 100%, and all other cAMP levels were normalized for comparative analysis. Concentration-response curves and EC50 values for vasopressin were determined for each V2R mutation. The individually determined EC50 values were used to stimulate basal cAMP production, allowing assessment of tolvaptan’s and conivaptan’s ability to inhibit cAMP accumulation at the mutant V2 receptors. To generate concentration-dependent curves for tolvaptan and conivaptan, cells (at a concentration of 2500 cells/well) were exposed to a mixture of tolvaptan/conivaptan and vasopressin for 30 min at 37 °C. The incubation was terminated by consecutively adding 5 μL of dye-labeled cAMP and 5 μL of cryptate-labeled anti-cAMP antibody in lysis buffer, followed by an additional one-hour incubation. Fluorescence signals were recorded at 665 nm and 620 nm using Thermo Scientific Varioskan LUX (Vantaa, Finland)36.
System preparation and molecular dynamics simulations
In this study, classical molecular dynamics simulations were employed to complement or validate the cryo-EM structural data of V2R in complex with tolvaptan and conivaptan. All simulations were performed using Amber20 software37. The resolved cryo-EM structures of V2R bound to tolvaptan and conivaptan were used to build the simulation systems. The missing residues in ECL2 and ICL2 were modeled using the Robetta server38. The anti-BRIL Fabs were excluded, and residues in ICL3 were not modelled. The open N- and C- termini, as well as the terminal residues at the intracellular ends of TM5 and TM6, were capped with neutral acetyl and methylamide groups. All residues and ligands were maintained in their standard protonation states, determined based on pKa values at pH 7.4 ± 0.05. The protonation state of the methylimidazole moiety in conivaptan was predicted using Schrödinger’s Epik module (Schrödinger Release 2022-3), which indicated that 3H-methylimidazole tautomer is the predominant form under physiological conditions. Therefore, this tautomer was chosen for subsequent MD simulations. The V2R structure was aligned in the Orientations of Proteins in Membranes (OPM) database39 and inserted into an 85 Å × 85 Å 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) bilayer using the CHARMM-GUI webserver. Following published works on GPCR MD simulations40,41, the LIPID17 force field was employed to parametrize the lipid bilayer, while the ff19SB force field42 was applied to the protein. The water model was chosen as TIP3P43, and the systems were set to a salt concentration of 0.15 M. The restrained electrostatic potential (RESP)44 of tolvaptan and conivaptan was characterized in Gaussian09 software45 with HF/6-31 G* basis. The ligands were parametrized using the general Amber force field (GAFF)46. A comprehensive summary of the system setup is provided in Supplementary Table 4.
Prior to production simulations, the systems underwent three rounds of energy minimization, each comprising 5000 steps (1000 steps of steepest descent optimization and 4000 steps of conjugate gradient optimization) to eliminate unnatural collisions. Subsequently, the systems were gradually heated from 0 to 300 K within the canonical (NVT) ensemble for 1 ns, with all heavy atoms constrained with 10.0 kcal·mol−1 Å−2 constraint. Following the heating phase, the systems were equilibrated in the isothermal isobaric (NPT) ensemble for 60 ns, during which harmonic restraints on protein backbone and ligand atoms were gradually reduced from 10.0 to 0 kcal·mol−1 Å−2. The SHAKE algorithm47 was used to constrain bond lengths involving hydrogen atoms. The particle mesh Ewald (PME) method48 was employed to calculate long-range electrostatic interactions, with a nonbonded cutoff set as 10 Å. The productive simulations were performed with a 2 fs time step. For each system, three independent 1 μs production runs were performed with different initial velocities.
Analysis of molecular dynamics trajectories
The analysis of MD simulation data was carried out using the CPPTRAJ49 module of Amber20. The root-mean-square deviation (RMSD) of ligands and protein backbone residues was calculated relative to the first frame of the production MD trajectories. The RMSD analysis showed that all simulation systems have reached convergence (Supplementary Fig. 4i, j). Hydrogen bond was defined based on geometric criteria: a hydrogen bond was considered formed if the distance between the acceptor heavy atom (A) and the donor heavy atom (D) was less than 3.5 Å, and the angle between the acceptor atom, donor hydrogen atom (H), and donor atom (A-H-D) was greater than 120°. To identify the key residues involved in the binding of ligands to the V2R, per-residue free energy decomposition analyses were conducted using the molecular mechanics/generalized Born surface area (MM/GBSA) method with the MMPBSA.py program50. A total of 1000 frames were extracted from the last 100 ns of trajectory for energy calculations. To extract representative conformations, principal component analysis (PCA) was performed on the final 100 ns of the trajectory. The eigenvectors of principal components were derived from the diagonalized covariance matrix of the RMSDs for the heavy atoms of ligands and critical residues involved in hydrogen bonding or contributing less than −1.5 kcal/mol to the binding energy. The first two principal components were subsequently used as reaction coordinates to construct the free energy landscape, with the lowest-energy conformation identified from the landscape serving as the representative conformation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The density maps and structure coordinates have been deposited to the EMDB and the PDB under accession numbers EMD-63949 and 9U81 for the tolvaptan-bound V2R complex; EMD-63948 and 9U80 for the conivaptan-bound V2R complex. EMD-25648, 6CBV, and 9HAP were used for cryo-EM data processing and model building. 7DW9, 6TPK, 3SN6, 3NY8, 8XVI, and 8XVJ were used for structure analysis. The initial coordinates, simulation input files, and molecular dynamics trajectories generated in this study have been deposited in the Zenodo repository (https://zenodo.org/records/16275945, https://doi.org/10.5281/zenodo.16275945) and are publicly accessible without restrictions. Source data are provided with this paper.
Change history
23 March 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41467-026-68448-7
References
Glavas, M. et al. Vasopressin and its analogues: from natural hormones to multitasking peptides. Int J. Mol. Sci. 23, 3068 (2022).
Perisic, M., Woolcock, K., Hering, A., Mendel, H. & Muttenthaler, M. Oxytocin and vasopressin signaling in health and disease. Trends Biochem. Sci. 49, 361–377 (2024).
Erdelyi, L. S., Hunyady, L. & Balla, A. V2 vasopressin receptor mutations: future personalized therapy based on individual molecular biology. Front. Endocrinol. 14, 1173601 (2023).
Girault-Sotias, P. E., Gerbier, R., Flahault, A., de Mota, N. & Llorens-Cortes, C. Apelin and vasopressin: the Yin and Yang of Water Balance. Front Endocrinol. 12, 735515 (2021).
Gilles, N. Natural peptide toxins as an option for renewed treatment of type 2 vasopressin receptor-related diseases. Biology 12, 544 (2023).
Kim, S., Jo, C. H. & Kim, G. H. The role of vasopressin v2 receptor in drug-induced hyponatremia. Front. Physiol. 12, 797039 (2021).
Schrier, R. W. et al. Tolvaptan, a selective oral vasopressin V2-receptor antagonist, for hyponatremia. N. Engl. J. Med. 355, 2099–2112 (2006).
Torres, V. E. et al. Tolvaptan in patients with autosomal dominant polycystic kidney disease. N. Engl. J. Med. 367, 2407–2418 (2012).
Ghali, J. K. et al. Efficacy and safety of oral conivaptan: a V1A/V2 vasopressin receptor antagonist, assessed in a randomized, placebo-controlled trial in patients with euvolemic or hypervolemic hyponatremia. J. Clin. Endocrinol. Metab. 91, 2145–2152 (2006).
Crombie, A. L. et al. Synthesis and evaluation of azabicyclo[3.2.1]octane derivatives as potent mixed vasopressin antagonists. Bioorg. Med. Chem. Lett. 20, 3742–3745 (2010).
Berl, T. et al. Oral tolvaptan is safe and effective in chronic hyponatremia. J. Am. Soc. Nephrol. 21, 705–712 (2010).
Warren, A. M., Grossmann, M., Christ-Crain, M. & Russell, N. Syndrome of inappropriate antidiuresis: from pathophysiology to management. Endocr. Rev. 44, 819–861 (2023).
Zhou, F. et al. Molecular basis of ligand recognition and activation of human V2 vasopressin receptor. Cell Res 31, 929–931 (2021).
Wang, L. et al. Cryo-EM structure of the AVP-vasopressin receptor 2-Gs signaling complex. Cell Res. 31, 932–934 (2021).
Bous, J. et al. Cryo-electron microscopy structure of the antidiuretic hormone arginine-vasopressin V2 receptor signaling complex. Sci. Adv. 7, eabg5628 (2021).
Mukherjee, S. et al. Synthetic antibodies against BRIL as universal fiducial marks for single-particle cryoEM structure determination of membrane proteins. Nat. Commun. 11, 1598 (2020).
Rondon-Berrios, H. & Berl, T. Vasopressin receptor antagonists: Characteristics and clinical role. Best. Pr. Res. Clin. Endocrinol. Metab. 30, 289–303 (2016).
Cao, X. et al. Benzodiazepine derivatives as potent vasopressin V2 receptor antagonists for the treatment of autosomal dominant kidney disease. J. Med. Chem. 65, 9295–9311 (2022).
Liu, H. L. et al. Structural basis of tolvaptan binding to the vasopressin V2 receptor. Acta Pharm. Sin. 45, 2441–2449 (2024).
Shi, L. et al. Beta2 adrenergic receptor activation. Modulation of the proline kink in transmembrane 6 by a rotamer toggle switch. J. Biol. Chem. 277, 40989–40996 (2002).
Holst, B. et al. A conserved aromatic lock for the tryptophan rotameric switch in TM-VI of seven-transmembrane receptors. J. Biol. Chem. 285, 3973–3985 (2010).
Fouillen, A. et al. Inactive structures of the vasopressin V2 receptor reveal distinct binding modes for Tolvaptan and Mambaquaretin toxin. Nat Commun. 16, 3899 (2025).
Foster, S. R. et al. Discovery of human signaling systems: pairing peptides to G protein-coupled receptors. Cell 179, 895–908 e21 (2019).
Kondo, K. et al. 7-Chloro-5-hydroxy-1-[2-methyl-4-(2-methylbenzoyl-amino)benzoyl]-2,3,4,5-tetrahydro-1H-1-benzazepine (OPC-41061): a potent, orally active nonpeptide arginine vasopressin V2 receptor antagonist. Bioorg. Med. Chem. 7, 1743–1754 (1999).
Albright, J. D. et al. 5-Fluoro-2-methyl-N-[4-(5H-pyrrolo[2,1-c]-[1, 4]benzodiazepin-10(11H)-ylcarbonyl)-3-chlorophenyl]benzamide (VPA-985): an orally active arginine vasopressin antagonist with selectivity for V2 receptors. J. Med. Chem. 41, 2442–2444 (1998).
Matthews, J. M. et al. Potent nonpeptide vasopressin receptor antagonists based on oxazino- and thiazinobenzodiazepine templates. Bioorg. Med. Chem. Lett. 14, 2747–2752 (2004).
Chen, H., Huang, W. & Li, X. Structures of oxysterol sensor EBI2/GPR183, a key regulator of the immune response. Structure 30, 1016–1024.e5 (2022).
Tsutsumi, N. et al. Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling. Elife 9, e58464 (2020).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem J. 478, 4169–4185 (2021).
Tunyasuvunakool, K. et al. Highly accurate protein structure prediction for the human proteome. Nature 596, 590–596 (2021).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D. Struct. Biol. 74, 519–530 (2018).
Adams, P. D. et al. Recent developments in the PHENIX software for automated crystallographic structure determination. J. Synchrotron Radiat. 11, 53–55 (2004).
Yun, Y. et al. Long residence time adenosine A1 receptor agonists produce sustained wash-resistant antilipolytic effect in rat adipocytes. Biochem. Pharm. 164, 45–52 (2019).
Case, D. A. et al. AMBER 2020. University of California, San Francisco. (2020).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Zhang, M. et al. Cryo-EM structure of an activated GPCR-G protein complex in lipid nanodiscs. Nat. Struct. Mol. Biol. 28, 258–267 (2021).
Liu, H. et al. Structural insights into ligand recognition and activation of the medium-chain fatty acid-sensing receptor GPR84. Nat. Commun. 14, 3271 (2023).
Tian, C. et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput 16, 528–552 (2020).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Frisch, M. J. et al. Gaussian 09, Revision A.02. (Gaussian, Inc., 2009).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Essmann, U. et al. A smooth particle mesh Ewald method. The Journal of Chemical Physics 103 (1995).
Roe, D. R. & Cheatham, T. E. III PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Miller, B. R. III et al. MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Acknowledgements
The cryo-EM data were collected at the Advanced Center for Electron Microscopy, Shanghai Institute of Materia Medica. We thank all staff at the institution for their assistance with cryo-EM data collection. This work was partially supported by the National Key Research and Development Program of China (2023YFC3605500 to W.Y.); the National Natural Science Foundation (32171187 to Y.J.; 82121005 to Y.J. and H.E.X.; 22077110 to D.G.; 32130022 to H.E.X.; 22107090 to H.L); the National Key R&D Program of China (2022YFC2703105 to H.E.X.); Program of Shanghai Academic/Technology Research Leader (22XD1425200 to Y.J.); Shanghai Oriental Talents Program (Y.J.); the Jiangsu Outstanding Youth Fund (BK20240051 to D.G.); the CAS Strategic Priority Research Program (XDB37030103 to H.E.X.); the Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to H.E.X.); China Postdoctoral Science Foundation (2023M731487 to Y.L.Y.); Shanghai Post-doctoral Excellence Program (2023018 to Y.L.Y. and 2022232 to B.P.); Shanghai Sailing Program (23YF1460700 to Y.L.Y. and 22YF1461200 to B.P.).
Author information
Authors and Affiliations
Contributions
T.Z. designed the expression constructs, expressed and purified the V2R samples, and prepared figures and the initial manuscript. H.L. conducted molecular dynamics simulations and prepared corresponding figures. C.Y. and Y.X. calculated cryo-EM data, built and refined structural models. Y.T. participated in the refinement of structural models. Y.Z., Y.C., and H.S. conducted site-directed mutagenesis experiment and cAMP assay. B.P. and C.W. prepared cryo-EM grids. K.W. conducted cryo-EM data collection. Y.L.Y., S.J., Y.S., and W.Y. participated in structural and functional analysis. H.E.X. supervised the structure determination. D.G. supervised the molecular dynamics simulations, mutagenesis, and cAMP assay. Y.J. and D.G. conceived and designed the experiments. Y.J. initiated collaborations with D.G. and H.E.X., supervised the project, and wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Giuseppe Deganutti, Jagannath Maharana and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, T., Liu, H., You, C. et al. Structural insights into antagonist recognition by the vasopressin V2 receptor. Nat Commun 16, 9734 (2025). https://doi.org/10.1038/s41467-025-64735-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64735-x
This article is cited by
-
Molecular basis of antagonism of the dimeric human arginine vasopressin receptor 1A
Nature Communications (2026)