Abstract
Asthma is a heterogeneous condition that is often comorbid with obesity and influenced by diverse risk factors. Elucidating the association of gut microbial characteristics with asthma could improve our understanding of the pathophysiology. Here, we investigate relationships of host genetics and stool microbiota characteristics with asthma among US Hispanic/Latino adults, while considering the influence of obesity status, using host whole genome sequencing and stool shotgun metagenomic microbiota data from participants of the Hispanic Community Health Study/Study of Latinos. We evaluate cross-sectional associations of microbiota characteristics with asthma and analyse whether they are modified by obesity status (body mass index≥30 kg/m2). We assess differences in alpha diversity, beta diversity, and taxonomic abundance with asthma, independent of obesity, and interactions between asthma and obesity using covariate-adjusted regression-based methods. We generate an asthma polygenic risk score (PRS) and compare the classification accuracy of genetic and microbial factors for asthma status. We report that asthma is associated with differences in overall taxonomic composition (beta diversity; p = 0.001), which is not dependent on obesity status (p = 0.31). Asthma is not associated with alpha diversity metrics (p > 0.17), though obesity is associated with lower alpha diversity (p < 0.01). We identify multiple taxa that are associated with asthma, including decreased abundance of Lactobacillus and Enterococcus species, and some taxonomic associations vary by obesity status. Compared to models including baseline risk factors and an asthma PRS, microbial information improves classification accuracy of asthma (p = 0.04). Our results support that there are microbiota characteristics associated with asthma in Hispanic/Latino adults independent of obesity.
Similar content being viewed by others
Introduction
Asthma is a heterogeneous condition influenced by demographic factors, environmental stimuli, and genetic predisposition1. The gut microbiota may influence disease etiology and severity, possibly through immune regulation, which plays a key role in asthma2. Microbiota and microbiota-derived metabolites can induce or dampen inflammatory processes2, and a fecal inoculant of bacteria protective against childhood asthma improved airway inflammation in a murine model3. Nasal, oral, and lung microbiota have been linked to asthma4,5, but few large-scale studies have examined the gut microbiota in asthma, particularly in adults. Understanding the role of the gut microbiota in the pathophysiology of asthma could inform preventive efforts, more accurate endotyping, or microbiota-based therapies2,6,7.
Obesity is often comorbid with asthma and associated with increased asthma symptoms and reduced lung function8. Gut microbiota has been studied extensively in relation to obesity and may contribute to weight gain and metabolic dysfunction9. A prior small study of ~200 individuals reported changes in the gut microbiota composition associated with asthma and with obesity, and enhanced changes when the conditions were comorbid10. Genetics also contribute to both conditions with a highly polygenic architecture11,12, though asthma genetic risk scores have somewhat limited accuracy11. Augmenting these risk scores with microbial information—which reflects environmental factors and non-genetic pathways—could advance personalized medicine, particularly if the combined information could accurately predict disease incidence or disease course.
Both obesity and asthma prevalence vary significantly across racial and ethnic groups. Hispanic/Latino adults are 20% more likely to be obese than non-Hispanic Whites, and children are 80% more likely13. Some Hispanic/Latino subpopulations have extremely high asthma prevalence, such as Puerto Ricans, whose lifetime prevalence is almost 2.5 times that of non-Hispanic Whites14. Furthermore, Hispanic/Latino individuals have higher complication rates from asthma compared to non-Hispanic Whites15. Thus, it is important to investigate asthma, and specifically obese asthma, in diverse communities that carry a high burden of these conditions.
In this study, we investigate the contribution of gut microbiota characteristics to asthma in a population of over 2400 Hispanic/Latino adults in the United States, while accounting for and evaluating the influence of obesity. The work is motivated by the hypothesis that asthma is associated with distinct gut microbiome characteristics relative to non-asthmatics, with differences by obesity status. We evaluate whether microbial information improves classification accuracy for asthma phenotypes independent of individual risk factors, including an asthma polygenic risk score (PRS). We also apply machine learning methods to determine the utility of gut microbiota for asthma subtyping.
Results
This study included 2404 Hispanics/Latino adults, 191 (7.9%) of whom had a current asthma diagnosis (86 non-obese asthmatics; 105 obese asthmatics; Table 1). Our initial sample had 3035 individuals, and we excluded 348 individuals with a history of asthma but no current asthma diagnosis, 238 individuals with unknown asthma status, and 45 with <100,000 sequencing reads in the microbiota data for the final sample size 2404. Females comprised 63.4% of the study cohort and 77.5% of those with asthma (69.8% of those with non-obese asthma; 83.8% of those with obese asthma). Almost one third (30.9%) of asthma cases were childhood onset (≤12 years). Average dietary quality was highest among non-obese non-asthmatic individuals (AHEI = 51.7, standard deviation (SD): 7.8) and lowest among those with asthma (non-obese asthma: 47.5 (7.5); obese asthma: 47.1 (7.7)). The cohort was predominantly individuals who self-reported national backgrounds of Mexico (41.1%), Puerto Rico (17.7%), Cuba (12.8%), and the Dominican Republic (10.3%). Asthma is known to vary greatly by Hispanic/Latino background, with particularly high prevalence in Puerto Ricans16, which was the case in this cohort. Obesity prevalence was high overall (42.6%) and even higher among those with asthma (55.0%), particularly Dominicans (75.9%).
Overall taxonomic composition associated with asthma and obesity
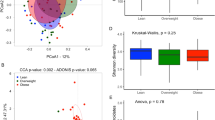
Gut microbiota taxonomic composition, summarized by Aitchison distance metric (beta diversity), differed significantly with current asthma status vs. no history of asthma (R2 = 0.11%; p = 0.001), adjusting for covariates, including obesity, which was likewise significant (R2 = 0.12%; p = 0.001) (Table S1 and Fig. 1a–d). The interaction between asthma and obesity was not significant (R2 = 0.04%; p = 0.30).
Taxonomy of study samples by participant asthma status (N = 2404: 191 asthmatics, 2213 no history of asthma) at the A phylum, B genus, and C species level. D Principal component plot of the gut microbiota composition summarized by the Aitchison distance metric, with colors and ellipses according to asthma status. In covariate-adjusted regression-based models evaluating the association between asthma status and the overall gut microbiota (beta diversity), there are highly significant (p = 0.001; Table S1) differences in gut microbiota composition by current asthma status (versus no history of asthma) but not a significant interaction between asthma and obesity (p = 0.31). E Plots of the relationship between current asthma status (versus no history of asthma) and obesity status (N = 2404: 1025 obese, 1379 non-obese) and standardized metrics of alpha diversity estimated by linear regression models. The x-axes show the effect estimate for asthma (left panel) and obesity (right panel) and 95% confidence intervals per standard deviation change in the alpha diversity metric (y-axis). The models control for: age, sex, and current smoking status, income, education level, Hispanic/Latino background (Mexican, Puerto Rican, Cuban, Dominican, or Other), generation in the mainland US, relocation age to the mainland US, and the Alternate Healthy Eating Index, a summary measure of dietary quality. The asthma model also controls for obesity status and vice versa. Source data for these figures are provided as Source Data files.
Lower alpha diversity associated with higher odds of obesity but not asthma
We assessed microbial richness, evenness, and phylogenetic relatedness via four alpha diversity metrics (Fig. 1e), none of which were significantly associated with asthma in adjusted models (p ≥ 0.12), controlling for obesity. Obesity was associated with lower evenness and Shannon diversity (β = −0.14; 95% confidence interval (CI): −0.22, −0.06; p = 0.0005); (β = −0.12; 95% CI: −0.20, −0.04; p = 0.003, respectively). There was no evidence of a statistical interaction between asthma and obesity for any of the alpha diversity metrics (p > 0.78).
Taxonomic and functional differences with asthma status
We split the data into training (N = 1353) and testing (N = 1051) sets to identify gut microbiota taxa and functions that differ by asthma status in the training dataset, and then examine the significant taxa in the testing dataset. In the training data, 58 taxa associated with asthma, independent of obesity (Supplementary Data 1); 29 of these showed significant associations with asthma in the testing data, ten of which remained significant after adjusting for multiple comparisons (Fig. 2a and Supplementary Data 2; prevalence and abundance shown in Fig. S1). Four of the taxa had higher relative abundance among asthmatics (Tissierellia bacterium S7-1-4, Porphyromonas asaccharolytica, Siccibacter turicensis, Alloscardovia criceti); the others had lower abundance (Enterococcus casseliflavus, Lactobacillus crispatus, Oxalobacter formigenes, Lactobacillus amylovorus, Lactobacillus reuteri, Enterococcus durans). For the analysis, a log transformation was applied to the taxonomic counts with a pseudo-count added when abundance was zero. We performed a sensitivity analysis to assess whether our results are robust to the choice of pseudo-count. The four taxa that passed this sensitivity analysis are indicated with an asterisk in the figure (T. bacterium S7-1-4, O. formigenes, L. reuteri, E. durans); those failing the sensitivity analysis could be false positives. We also identified 15 taxa that associated with the interaction between asthma and obesity in the training data (Supplementary Data 3); three of the taxa were significant in the testing data after adjusting for multiple comparisons (Bifidobacterium minimum, Enterococcus italicus, Fusobacterium mortiferum; Fig. 2b and Supplementary Data 3–5; prevalence and abundance shown in Fig. S2). However, based on the results of the pseudo-count sensitivity analyses, these significant differences in abundance by asthma/obesity status that we observed could be due to false positives.
These plots show results of differential abundance analyses using ANCOM-BC (Analysis of Compositions of Microbiomes with Bias Correction). A The taxa shown on the y-axis were significantly (q < 0.05) associated with current asthma status (versus no history of asthma) in both the training (N = 1353: 96 asthmatics, 1257 non-asthmatics) and testing datasets (N = 1051: 95 asthmatics, 956 non-asthmatics)). The estimated log-fold change (± standard error) in the taxa abundance in the testing dataset is shown on the x-axis. Taxa significantly increased with asthma are shown in red and those decreased with asthma are shown in blue. B For the taxa on the y-axis, the association with asthma was dependent on obesity status; the interaction between asthma and obesity status was significant in both the training and testing datasets (q < 0.05). In order to interpret the interaction effects, the figures depict the change in log-fold abundance (± standard error) in the testing dataset for non-obese asthma (N = 42), obese asthma (N = 53), and non-asthmatic obesity (N = 406) versus non-obese non-asthmatic (N = 550). The comparisons that were significant are shown in red (increased abundance) or blue (decreased abundance); and non-significant results are in gray. For both A and B, the statistical models involve adding a pseudo-count when taxonomic abundance is zero; the asterisks (*) indicate that the associations were robust to the choice of pseudo-count in sensitivity analyses. All models controlled for age, sex, current smoking status, income, education level, Hispanic/Latino background, generation in the mainland US, relocation age to the mainland US, the Alternate Healthy Eating Index, and obesity status in models of current asthma status (A). Source data for these figures are provided as Source Data files.
There were ten functional modules associated with asthma independent of obesity status (Supplementary Data 6), of which seven were significant in the testing dataset, all related to genetic information processing (Fig. S3a and Supplementary Data 7). Given the similarity of the effect estimates, we evaluated the correlations among these functions, and they were high (95%), indicating that these functions are largely redundant. All of these modules failed the pseudo-count sensitivity analyses and could be false positives. There were six functional modules with statistically significant interactions between asthma and obesity status in the training data (Supplementary Data 8); three of these were significant in the testing dataset (Fig. S3b and Supplementary Data 9, 10). These functions are related to carbohydrate and lipid metabolism and environmental information processing, but the only significant finding that was robust to false positives in the sensitivity analysis was decreased carbohydrate metabolism related to 2-oxoisovalerate:ferredoxin oxidoreductase with non-asthmatic obesity.
Assessment of causal relationships of taxa with asthma
We examined the taxa that were significantly associated with asthma in the testing dataset (Fig. 2) for causal effects on asthma using two-sample Mendelian Randomization (MR). Some of the taxa in our results did not have genetic instruments at the species level based on prior literature, in which case we used instruments at the genus level when available; in some cases, neither were available. Furthermore, most of the taxa did not have strong genetic instruments (i.e., genetic instruments highly predictive of taxonomic abundance, usually an association p < 10−8); thus, we assessed genetic instruments at a range of p-value thresholds (from p < 10−6 to 10−8; Table S2). We found no evidence of significant causal relationships between any of these taxa and asthma (Fig. S3 and Supplementary Data 11).
An asthma polygenic risk score and gut microbiota characteristics independently improved models of current asthma and obese asthma
The asthma PRS was elevated among individuals with current asthma versus those with no history of asthma (Fig. 3a). While the interaction between the PRS and obesity was not significant (p = 0.31), the PRS was higher among those with obese asthma compared to non-obese asthma in covariate-adjusted regression models (β = 0.41; p = 0.03). The area under the curve (AUC) for the asthma PRS was 0.62 for current asthma, 0.60 for non-obese asthma, and 0.66 for obese asthma (Fig. 3b).
A The distribution of asthma PRS values for individuals in the subset of individuals in this study with genetic data (N = 1089: 133 current asthma, 956 no history of asthma), and specifically for those with non-obese asthma (N = 65) and obese asthma (N = 68). The violin plots show the distributions of PRS, with box plots showing the median, first and third quartiles, whiskers that extend from the hinge to the largest value no further than 1.5 * interquartile range, and outliers. B Area under the curve (AUC) values and 95% confidence intervals from logistic regression models of asthma outcomes as a function of the asthma PRS in the subset of individuals in this study with genetic data (N = 1089): current asthma (N = 133) versus and no history of asthma (N = 956); non-obese asthma (N = 65) and obese asthma (N = 68) versus non-obese/non-asthmatic (N = 550). C The AUC of sequentially adjusted logistic regression models in the testing data (N = 1051) of: (1) current asthma (N = 95) versus and no history of asthma (N = 956); and (2) non-obese asthma (N = 42) and obese asthma (N = 53) versus non-obese/non-asthmatic (N = 550). The models sequentially adjust for: (1) baseline covariates (age, sex, current smoking status, income, education level, Hispanic/Latino background, generation in the mainland US, relocation age to the mainland US, the Alternate Healthy Eating Index, and the first five genetic principal components); (2) the asthma PRS; and (3) a summary measure of microbiota species. The p value of each model compared to the previous model is shown with the level of statistical significance indicated by asterisks. Source data for these figures are provided as Source Data files. AUC area under the curve, CI confidence interval, PRS polygenic risk score.
We sequentially evaluated (1) baseline covariates (described above) plus five genetic principal components, (2) the asthma PRS, and (3) microbiota features (Table S3) as predictors of asthma outcomes using a stepwise approach. Compared to baseline risk factors alone, the PRS significantly improved models of current asthma versus no history of asthma and obese asthma (but not non-obese asthma) versus non-obese/non-asthmatics (Fig. 3c). The addition of microbiota information further improved the models of current asthma and obese asthma. The improvements in AUC—and overall AUC values—were highest for obese asthma (0.814 in the baseline model, 0.844 with the addition of the PRS, and 0.860 in the final model with microbiome information; compared to 0.766, 0.779, and 0.783 for current asthma, and 0.808, 0.810, and 0.811 for non-obese asthma). Functional microbiota information did not improve models of any asthma outcomes (Fig. S3c and Table S4). We also examined interactions between genetic factors (the PRS and top variants) and the taxa associated with asthma; none were significant after adjusting for multiple comparisons (q ≥ 0.90), though many were nominally significant (Supplementary Data 12).
Clinical asthma subtypes do not correlate well with gut microbiota characteristics
There are multiple clinically observed asthma subtypes, and microbiome information may help to understand differences in the physiology of the subtypes. Thus, we clustered all individuals in HCHS/SOL with current asthma based on the available clinical characteristics, such as demographics, childhood asthma onset, BMI, and lung function, and we identified two asthma subtypes (Fig. 4a, b). The subtypes correlated strongly with asthma onset: 96.6% of individuals had childhood-onset asthma in subtype 1 versus 25.6% in subtype 2. Subtype 2 had higher obesity prevalence and proportion female, and poorer lung function. We sought to differentiate these two asthma subtypes using microbiota species (Fig. 4c), which showed low accuracy AUC = 0.52 (0.42–0.62). Since childhood- versus adult-onset asthma was the primary defining feature of these subtypes, we additionally applied the same approach to try to differentiate asthmatics by onset using species information, which showed greater accuracy (Fig. 4c, d; AUC = 0.59 (0.51–0.67)).
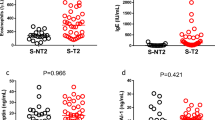
A Using clustering techniques applied to clinical information, we identified two asthma subtypes among all HCHS/SOL asthmatics (N = 2027: 955 Subtype 1 (purple), 1072 Subtype 2 (green)). The subset of N = 191 asthmatics in the microbiome study are shown in darker colors. B The table shows characteristics of the individuals in each subtype. Subtype 1 was predominantly individuals with child-onset asthma (96.6%), whereas subtype 2 was generally characterized by adult-onset asthma (74.4%) and female sex (83.3%). The statistical comparisons are based on chi-squared tests for categorical variables and ANOVA for continuous variables. C Receiver operating characteristic (ROC) curves for machine learning models (Lasso with logistic regression) using species-level taxa in the subset of individuals with microbiome data (N = 191) to differentiate: (1) the asthma subtypes (purple); and (2) childhood versus adult-onset asthma (orange), which was the primary differentiating characteristic of the subtypes. An area under the curve (AUC) value of 0.5 is equivalent to random guessing, whereas values above 0.5 have greater accuracy. The AUC values indicate that microbiota information cannot accurately differentiate between the asthma subtypes but can differentiate childhood versus adult-onset asthma. D The topmost important taxa from the machine learning model of childhood versus adult-onset asthma are shown with their importance scores; the direction of effect and colors indicate whether the taxa are enriched with childhood (purple) or adult (teal) onset asthma. Source data for these figures are provided as Source Data files. AUC Area under the curve.
Discussion
Asthma is a complex condition that is frequently comorbid with obesity. Here, we examined the relationships among gut microbiota, genetic risk factors, and asthma phenotypes in Hispanic/Latino adults. We observed differences in overall taxonomic structure (beta diversity) with current asthma status versus those with no history of asthma. We report microbiota taxonomic and functional associations with asthma, some of which were modified by obesity status. While many of the identified taxa were not major components of the overall composition, low-abundance taxa can have potent biological effects17. We investigated the use of MR to assess the causal effects of these taxa, but most of the taxa did not have adequately strong genetic instrumental variables to justify this type of causal inference. Gut microbiota information enhanced models of asthma beyond baseline risk factors and an asthma PRS.
Prior work of asthma and obesity concluded that the combination of these two conditions has additive effects on systemic inflammation and the gut microbiome10. We did not examine immunological phenotypes, but our results do not indicate a strong additive effect of asthma and obesity on the gut microbiome. For example, we did not see significant interactions between asthma and obesity in relation to alpha or beta diversity, and only a few taxa showed an association with asthma that was dependent on obesity status. These interaction effects were not driven by “enhanced” or “additive” effects in obese asthma, i.e., effects that were present with non-obese asthma or non-asthmatic obesity that were exacerbated when the conditions were both present. Our sample size of asthmatics was almost twice as large as the prior study, but even larger samples may be necessary to fully clarify the additive effects of these two conditions on the microbiome.
Chronic inflammation is characteristic of both asthma and obesity. Lactobacillus is a well-known anti-inflammatory microbial genus, and many of the species that were protectively associated with asthma in our results are in this genus, including L. reuteri, L. crispatus, and L. amylovorus. Probiotic supplements often contain Lactobacillus species, particularly L. reuteri. In human trials, L. reuteri probiotics improved symptoms in asthmatics and those with allergic rhinitis and reduced Th2-associated pro-inflammatory cytokines18. In mouse models, L. reuteri strains alleviated airway inflammation, promoted the proliferation of Tregs, and modulated gut microbiota composition overall, enriching the prevalence of Lactobacillus and Enterococcus species and enhancing the generation of butyrate, which is a strongly anti-inflammatory short-chain fatty acid metabolite19. Our results likewise show protective associations of some species within the Enterococcus genus, including E. durans, E. casseliflavus, and E. italicus. We performed an MR analysis to assess whether the taxa identified in our analyses were causally related to asthma, but we were unable to identify strong genetic instruments for most of the taxa (with the exception of Bifidobacterium), and we found no evidence of causal links. The association between functional lactase gene variants and Bifidobacterium is one of the most robust genetic-taxonomic links across prior literature; however, this association is subject to strong gene-diet interactions with dairy intake20,21,22. Unfortunately, the generally weak associations between genetic variants and microbial taxa are a major impediment to the utility of MR studies of the microbiome. While many recent studies have employed this approach, a recent review of MR studies concluded that they were typically of poor quality with inappropriate application of the method23.
Our results suggest that lower gut alpha diversity is associated with increased odds of obesity. This relationship was not observed with asthma. Prior studies examining the association between gut microbiota diversity and obesity, childhood asthma, and adult asthma have been inconsistent24,25,26, as have studies of respiratory alpha diversity and asthma27. This inconsistency could reflect the heterogeneity of these conditions across individuals, heterogeneity in microbiota-disease relationships across diverse populations, or data quality issues28.
In addition to microbiota information, we examined an asthma PRS, which had slightly better performance compared to previously reported asthma PRS in adults (0.62 compared to AUCs of 0.51–0.57)11. There is evidence that genetics and environment interact in ways that contribute to asthma pathogenesis and that some of those effects are at least partially mediated through the microbiome1. In this study, we did not see statistically significant interactions between genetic risk factors for asthma (PRS or top variants) and asthma-associated taxa, or between the PRS and obesity status in relation to asthma. However, in our results, the PRS and microbiota information independently improved the accuracy of models of current asthma status (versus those with no history of asthma) and obese asthma (versus non-obese/non-asthmatic), which suggests that these different types of information capture different aspects of risk. This aligns with prior evidence that genetic and microbiota information additively improve models for numerous traits, including incident disease29,30. We observed that the asthma PRS was higher on average among those with obese asthma than with non-obese asthma, and it did not significantly improve models of non-obese asthma beyond baseline risk factors. Since these conditions are so commonly comorbid, prior asthma genome-wide association studies may more accurately reflect obese asthma than non-obese asthma. Many prior studies have also noted a genetic correlation between asthma and BMI, particularly for later onset asthma—with estimates as high as 0.2531,32,33. Genetic predisposition for obesity is associated with asthma in the HCHS/SOL cohort, particularly childhood-onset asthma34. Thus, genetic factors involved in obesity may also play a role in asthma (or vice versa), and our results suggest a similar trend for some microbial factors. An important area for future work is to combine genetic and microbiome signatures to try to predict asthma incidence or disease course.
Asthma encompasses a heterogeneous group of clinical symptoms that may more closely reflect a syndrome with multiple phenotypes than a single disease35. Numerous efforts have sought to define asthma subtypes based on clinical characteristics36,37,38,39, hoping that this may lead to more effective targeted therapies40. Two of the subtypes seen frequently across studies are characterized by: (1) early-onset asthma, usually with high prevalence of allergic/atopic disease; and (2) later-onset disease with poorer lung function, predominantly occurring in women and individuals with obesity36. We identified asthma subtypes consistent with this prior work. Since the link between clinical subtypes and pathophysiology remains unclear40, molecular data may improve our understanding of the underlying biology of asthma subtypes37. Thus, we used a machine learning approach to try to identify microbial signatures of the asthma subtypes in this study. While this was unsuccessful, one of the primary defining characteristics of the subtypes was childhood- vs. adult-onset asthma, which we were able to differentiate with moderate accuracy using microbiota species information. It is thought that adult-onset disease is more environmentally driven (which may be reflected in microbial signatures), whereas childhood-onset is more genetically driven, specifically by dysregulated allergy and epithelial barrier function genes41. Prevotella was among the microbial taxa in our findings that was enriched in adult-onset versus childhood-onset asthma. Prior work on obese asthma has linked Prevotella-dominated microbiota composition with increased chemokines, regardless of obesity or asthma status10; thus, our result could reflect the increased inflammation that is characteristic of adult-onset asthma37. This is an important area for further research in larger cohorts of asthmatics, particularly combining genetic, microbiota, and other molecular information, such as metabolomics or transcriptomics.
This is one of the largest studies of the gut microbiota and asthma in adults; however, there are some limitations, and our results may not be generalizable to non-Hispanic/Latino populations. The cross-sectional nature of the data limits our ability to understand the direction of effects, and observational data cannot be used to determine causality. Prior work in this and other cohorts has established associations of Hispanic/Latino background, age at relocation to the mainland US, sex42, and diet with asthma43,44, obesity45, and gut microbiota46. We controlled for these and other relevant factors, but there remains the possibility of residual confounding due to imperfect measurement and unmeasured confounding (e.g., environmental exposures, urbanicity, atopy), which could impact our findings. Furthermore, we were unable to investigate all the possible complex interactions among the measured confounders, given the number of asthmatics in this study and the inter-individual heterogeneity of asthma and the microbiome. Similarly, we had limited power for our analyses of interaction effects. These are important areas for further research. Many measures relevant to asthma were unavailable or limited to a subset of participants, hindering our ability to perform in-depth analyses of inflammation, comorbid allergies, asthma severity, or exacerbations. Furthermore, there is likely some misclassification of asthma status as our definition was based on self-report, and this misclassification could be differential based on obesity status. Despite these limitations, we were able to leverage the substantial strengths of the cohort, including the large sample size, extensive data collection, and unique population, to help elucidate the relationships of host genetics and stool microbiota characteristics with asthma status. Our results highlight taxa that might offer new avenues for asthma prevention or treatment strategies with further research, such as supporting beneficial taxa like strains of Lactobacillus and Enterococcus.
In conclusion, this study detected numerous associations between gut microbiota characteristics and asthma. Asthma is often comorbid with obesity, and we identified some taxonomic associations with asthma that were dependent on obesity status. However, we did not find strong overall evidence of “additive” effects of these two conditions on the microbiome. We found differences in the taxonomic signatures of adult- vs. child-onset asthma. Thus, our results support that there is a gut microbial signature of asthma that may vary to some extent with obesity and age at onset.
Methods
This study complies with complies with relevant ethical regulations and was approved by the Albert Einstein College of Medicine institutional review board.
Study population
Hispanic Community Health Study/Study of Latinos (HCHS/SOL) is a population-based cohort study of 16,415 adults, aged 18–74 years, self-identifying as Hispanic/Latino recruited through four field centers in Miami, San Diego, Chicago and the Bronx area of New York. HCHS/SOL has had three in-person study visits and an annual telephone interview between visits. Our analysis focused on the subset of participants in the Gut Origins of Latino Diabetes (GOLD) ancillary study that coincided approximately with the second study visit46. Study design, participant measures collection, and microbiota data generation are summarized in the Supplementary Methods45,46,47,48. HCHS/SOL was approved by institutional review boards at each center and the data coordinating center (University of North Carolina); all participants gave written informed consent.
Outcomes
The primary outcome of interest in this study was current asthma status, defined based on self-report of physician-diagnosed asthma and asthma medication in the four weeks preceding the second study visit. We compared cases to those without asthma reported at the first two study visits or in annual follow-up, and we excluded individuals without current asthma but who had previously reported an asthma diagnosis to avoid potential misclassification due to temporal symptom variation. We were interested in the influence of obesity status on the relationships of gut microbiota and genetic characteristics with asthma. BMI was calculated based on measured weight and height, and obesity status was defined as BMI ≥ 30 kg/m2. For many of our analyses, we modeled asthma and obesity as predictors (rather than outcome variables) to test their statistical interaction. We also performed some analyses using categorical asthma/obesity status: non-obese asthma (current asthma, body mass index (BMI) < 30), obese asthma (current asthma, BMI ≥ 30), non-asthmatic obesity (BMI ≥ 30, no asthma history), and the referent group of non-obese/non-asthmatic (BMI < 30, no asthma history).
Covariates
Information on demographics (including self-reported sex, age, Hispanic background and relocation history), behaviors, health status, family and medical histories, and medication use was collected using structured questionnaires, and anthropometric traits were measured at in-person visits47,48. Two 24-h dietary recalls were collected during the HCHS/SOL baseline period. The first recall was collected in person at the clinic, and the second recall was collected by telephone within 3 months of the first recall. The Alternate Healthy Eating Index (AHEI) was calculated to represent adherence to overall healthy dietary pattern recommendations49.
Fecal sample collection and processing
Participants enrolled in the GOLD study self-collected two fecal samples, one via Whatman Flinders Technology Associates (FTA) card (GE Healthcare, Chicago, IL) and another placed in a tube of RNAlater (Invitrogen, Carlsbad, CA)45,46. DNA extracted from the FTA card underwent shotgun metagenomic sequencing via a shallow-coverage method using Illumina NovaSeq platforms50. The bioinformatics processing the sequencing data is described in the Supplementary Methods. The alpha diversity indices (observed features, Pielou’s evenness, Shannon diversity index, and Faith’s phylogenetic distance51,52) were calculated using QIIME version 2021.453, and beta diversity Aitchison distances were calculated using the R vegan package54,55. Functional profiles were obtained using SHOGUN56 via sequence alignment to a nucleotide gene database derived from NCBI RefSeq (release 82) and the Kyoto Encyclopedia of Genes and Genomes database57 (release 94.0) and summarized at the module level, which correspond to functional units of gene sets. Both taxonomic and functional abundances were centered log-ratio transformed prior to analyses, with a pseudo-count (half of the minimum of the relative abundance) added to zeros before taking logs58.
Whole genome sequencing data
The genetic data were generated by the Trans-Omics for Precision Medicine (TOPMed) program using whole genome sequencing to a median depth of 30X using Illumina HiSeq X technology (study id: phs001395; more details provided in the Supplementary Methods).
Statistical analysis
We compared participant characteristics by asthma/obesity status using chi-squared tests for categorical variables and ANOVA for continuous variables.
Microbiota diversity, taxonomy, and functions
We assessed the association of asthma with the overall taxonomic composition (beta diversity) using permutational ANOVA of the Aitchison distance metric54 using the R package GUniFrac v1.859. We used linear regressions to evaluate the association of asthma status with four scaled measures of alpha diversity (observed features, Pielou’s evenness, Shannon Diversity Index, and Faith’s phylogenetic diversity) to capture the richness, evenness, and phylogenetic relatedness of microbiota within each sample. We included asthma and obesity status in the models, as well as the interaction between them, which we removed in cases where it was not statistically significant. We additionally controlled for the following covariates throughout our analyses: age, sex, smoking status, income, education level, Hispanic/Latino background (Mexican, Puerto Rican, Cuban, Dominican or Other), generation in the US, relocation age to the US mainland, and the AHEI. In models of alpha diversity, we additionally controlled for total sequencing depth.
We split the data into training and testing sets to identify specific microbiota taxa and functional pathways robustly associated with asthma (see Supplementary Methods). We applied the analysis of compositions of microbiomes with bias correction (R package ANCOMBC v2.8.1)60 in the training set to the species-level taxa and microbiota functions (summarized as modules, i.e., functional units of gene sets) with adjustment for covariates. We used the training data to identify features that were significantly associated with asthma status (in models controlling for obesity status) or with the interaction between asthma and obesity (q < 0.05). We then evaluated these taxa in the testing data. To understand the interaction effects for taxa showing significant asthma/obesity interactions in the testing data (q < 0.05), we ran stratified models comparing: (1) non-obese asthma; (2) obese asthma; and (3) non-asthmatic obesity to non-obese non-asthmatics. In the training and testing data, we used the Holm method to account for multiple comparisons, as recommended by the authors of ANCOM-BC61. We also implemented the sensitivity analysis in the ANCOMBC package that evaluates whether the results were robust to the choice of pseudo-count on the log-transformed data (and thus unlikely to be false positives) with an alpha value of 0.0561. Among the 2733 species and 445 functional modules detected across the samples, we included features that were detectible in at least 90% of participants (N = 599 species; 388 functional modules).
Estimating causal effects of identified taxa
MR62 is a statistical method to estimate causality of exposures on outcomes using genetic instrumental variables to circumvent some of the usual biases present in observational epidemiologic studies. We applied two-sample MR to assess causal effects of taxa on asthma, focusing on the subset of taxa that were significantly associated with asthma in our testing dataset. Genome wide summary statistics for the species were obtained from previously published manuscripts from the Dutch Microbiome Project and the FINRISK study when available63,64, and genus-level summary statistics from the MiBioGen consortium65 otherwise. Summary statistics for asthma were retrieved from the Global Biobank Meta-Analysis Initiative (GBMI)66. See the Supplementary Methods for more detail. We adjusted the p-values using the Benjamini–Hochberg67 false discovery rate (FDR) to account for multiple comparisons.
Polygenic risk scores (PRS)
We generated an asthma PRS based on prior work by Wang et al.68 using summary statistics from GBMI11 with weights that are available on the Polygenic Score Catalogue69 (PGS001787). We standardized the PRS to have mean zero and standard deviation one. We evaluated the PRS based on odds ratios and the receiver operating characteristic AUC values from logistic models of asthma (current asthma versus non-asthmatic; non-obese asthma versus non-obese non-asthmatic; and obese asthma versus non-obese non-asthmatic). We also tested the interaction between the asthma PRS and obesity in relation to current asthma.
Parsimonious models of asthma/obesity outcomes
We fit logistic regression models of asthma outcomes (current asthma versus those with no history of asthma; non-obese asthma versus non-obese non-asthmatics; obese asthma versus non-obese non-asthmatics) in the testing data sequentially adjusting for the following: (1) baseline covariates (those described above) and the first five genetic principal components to control for global ancestry; (2) asthma PRS; and (3) a summary microbiota measure (see Supplementary Methods). We assessed sequential model improvement using a likelihood ratio test and AUC.
Interactions between genetic factors and taxa
We examined interactions between genetic factors (including the asthma PRS and the 100 genetic variants from the PRS most strongly associated with asthma) and the taxa significantly associated with asthma in the testing dataset using logistic regressions and controlling for the same covariates described above. We used FDR to adjust the interaction p values for multiple comparisons.
Asthma subtyping
Using the full HCHS/SOL dataset with known asthma status, we used using partitioning around medoids to cluster the subset of individuals with current asthma (N = 2027) based on demographic and clinical characteristics (e.g., age, sex, asthma onset, BMI, clinical characteristics). We used the R package mice to impute missing values among these clinical variables. We used Lasso logistic regression with 5-fold cross validation and 50 repeats to characterize the predominant microbiota species associated with the subtypes in the subsample of asthmatic individuals with microbiome data. We estimated the AUC in the validation loops using the R package MLeval v0.370. A primary difference between the subtypes was the age at asthma onset (childhood or adult). Thus, we also applied a similar Lasso approach to try to differentiate childhood versus adult-onset asthma based on species-level taxa. See the Supplementary Methods for more details.
We conducted analyses in the NHLBI BioData Catalyst71 environment powered by Terra and R v4.4.272. Our statistical significance cutoff was 0.05 after adjustment for multiple comparisons where appropriate, as noted.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
HCHS/SOL data are on the National Institutes of Health repositories dbGap (accession number phs000810.v1.p1; phs001395.v2.p1) and accessible through BioData Catalyst71. Gut microbiota sequence data are deposited in QIITA (ID 11666) and EMBL-EBI ENA (ERP117287)73. HCHS/SOL has an established process for researchers to apply for access to participant data and materials with a review process by the project’s Steering Committee. These policies are described at https://sites.cscc.unc.edu/hchs/ (accessed August 28, 2025). The senior author will accept reasonable requests for data and specimen access, which will be referred to the Steering Committee of the HCHS/SOL project and addressed in a timely manner. All other data are available in the article and its Supplementary files or from the corresponding author upon request. Source data are provided with this paper.
Code availability
The code relevant to this project is at https://github.com/Stanislabski/AsthmaObesity.
References
Tang, H. H. F., Teo, S. M., Sly, P. D., Holt, P. G. & Inouye, M. The intersect of genetics, environment, and microbiota in asthma—perspectives and challenges. J. Allergy Clin. Immunol. 147, 781–793 (2021).
Sokolowska, M., Frei, R., Lunjani, N., Akdis, C. A. & O’Mahony, L. Microbiome and asthma. Asthma Res. Pract. 4, 1 (2018).
Arrieta, M.-C. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152–307ra152 (2015).
Miller, R. L., Grayson, M. H. & Strothman, K. Advances in asthma: new understandings of asthma’s natural history, risk factors, underlying mechanisms, and clinical management. J. Allergy Clin. Immunol. 148, 1430–1441 (2021).
Hufnagl, K., Pali-Schöll, I., Roth-Walter, F. & Jensen-Jarolim, E. Dysbiosis of the gut and lung microbiome has a role in asthma. Semin. Immunopathol. 42, 75–93 (2020).
Barcik, W., Boutin, R. C. T., Sokolowska, M. & Finlay, B. B. The role of lung and gut microbiota in the pathology of asthma. Immunity 52, 241–255 (2020).
Alashkar Alhamwe, B. et al. Intranasal administration of Acinetobacter lwoffii in a murine model of asthma induces IL-6-mediated protection associated with cecal microbiota changes. Allergy 78, 1245–1257 (2023).
Peters, U., Dixon, A. E. & Forno, E. Obesity and asthma. J. Allergy Clin. Immunol. 141, 1169–1179 (2018).
Aron-Wisnewsky, J., Warmbrunn, M. V., Nieuwdorp, M. & Clément, K. Metabolism and metabolic disorders and the microbiome: the intestinal microbiota associated with obesity, lipid metabolism, and metabolic health-pathophysiology and therapeutic strategies. Gastroenterology 160, 573–599 (2021).
Michalovich, D. et al. Obesity and disease severity magnify disturbed microbiome-immune interactions in asthma patients. Nat. Commun. 10, 5711 (2019).
Tsuo, K. et al. Multi-ancestry meta-analysis of asthma identifies novel associations and highlights the value of increased power and diversity. Cell Genom. 2, 100212 (2022).
Khera, A. V. et al. Polygenic prediction of weight and obesity trajectories from birth to adulthood. Cell 177, 587–596.e9 (2019).
Obesity and Hispanic Americans-The office of minority health. https://minorityhealth.hhs.gov/omh/browse.aspx?lvl=4&lvlid=70.
González Burchard, E. & Borrell, L. N. Need for racial and ethnic diversity in asthma precision medicine. N. Engl. J. Med. 385, 2297–2298 (2021).
Asthma and Hispanic Americans - The office of minority health. https://www.minorityhealth.hhs.gov/omh/browse.aspx?lvl=4&lvlid=60.
Barr, R. G. et al. Pulmonary disease and age at immigration among Hispanics. Results from the Hispanic community health study/study of Latinos. Am. J. Respir. Crit. Care Med. 193, 386–395 (2016).
Han, G., Luong, H. & Vaishnava, S. Low abundance members of the gut microbiome exhibit high immunogenicity. Gut Microbes 14, 2104086 (2022).
Li, L. et al. Efficacy and safety of Lactobacillus reuteri CCFM1040 in allergic rhinitis and asthma: a randomized, placebo-controlled trial. Front. Nutr. 9, 862934 (2022).
Li, L. et al. Lactobacillus reuteri attenuated allergic inflammation induced by HDM in the mouse and modulated gut microbes. PLoS ONE 15, e0231865 (2020).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Wang, J. et al. Meta-analysis of human genome-microbiome association studies: the MiBioGen consortium initiative. Microbiome 6, 101 (2018).
Goodrich, J. K., Davenport, E. R., Clark, A. G. & Ley, R. E. The relationship between the human genome and microbiome comes into view. Annu. Rev. Genet. 51, 413–433 (2017).
Hatcher, C. et al. A systematic review of the applications of Mendelian randomization assessing the causal relevance of the gut microbiome in human health and disease. Preprint at https://doi.org/10.1101/2025.06.03.25328787 (2025).
Zimmermann, P., Messina, N., Mohn, W. W., Finlay, B. B. & Curtis, N. Association between the intestinal microbiota and allergic sensitization, eczema, and asthma: a systematic review. J. Allergy Clin. Immunol. 143, 467–485 (2019).
Wang, Q. et al. A metagenome-wide association study of gut microbiota in asthma in UK adults. BMC Microbiol. 18, 114 (2018).
Legatzki, A., Rösler, B. & von Mutius, E. Microbiome diversity and asthma and allergy risk. Curr. Allergy Asthma Rep. 14, 466 (2014).
Avalos-Fernandez, M. et al. The respiratory microbiota alpha-diversity in chronic lung diseases: first systematic review and meta-analysis. Respir. Res. 23, 214 (2022).
Stanislawski, M. A., Dabelea, D., Lange, L. A., Wagner, B. D. & Lozupone, C. A. Gut microbiota phenotypes of obesity. NPJ Biofilms Microbiomes 5, 18 (2019).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Liu, Y. et al. Integration of polygenic and gut metagenomic risk prediction for common diseases. Nat. Aging 4, 584–594 (2024).
Zhu, Z. et al. Shared genetic and experimental links between obesity-related traits and asthma subtypes in UK Biobank. J. Allergy Clin. Immunol. 145, 537–549 (2020).
Han, X. et al. Obesity-related biomarkers underlie a shared genetic architecture between childhood body mass index and childhood asthma. Commun. Biol. 5, 1–10 (2022).
Thomsen, S. F., Duffy, D. L., Kyvik, K. O., Skytthe, A. & Backer, V. Risk of asthma in adult twins with type 2 diabetes and increased body mass index. Allergy 66, 562–568 (2011).
Guo, Y. et al. Genetic predisposition to obesity is associated with asthma in US Hispanics/Latinos: results from the Hispanic community health study/study of Latinos. Allergy 73, 1547–1550 (2018).
Kleinert, S. & Horton, R. After asthma: airways diseases need a new name and a revolution. Lancet 391, 292–294 (2018).
Moore, W. C. et al. Identification of asthma phenotypes using cluster analysis in the severe asthma research program. Am. J. Respir. Crit. Care Med. 181, 315–323 (2010).
Wenzel, S. E. Asthma phenotypes: the evolution from clinical to molecular approaches. Nat. Med. 18, 716–725 (2012).
Lefaudeux, D. et al. U-BIOPRED clinical adult asthma clusters linked to a subset of sputum omics. J. Allergy Clin. Immunol. 139, 1797–1807 (2017).
Deliu, M., Sperrin, M., Belgrave, D. & Custovic, A. Identification of asthma subtypes using clustering methodologies. Pulm. Ther. 2, 19–41 (2016).
Fajt, M. L. & Wenzel, S. E. Asthma phenotypes and the use of biologic medications in asthma and allergic disease: the next steps toward personalized care. J. Allergy Clin. Immunol. 135, 299–310 (2015).
Pividori, M., Schoettler, N., Nicolae, D. L., Ober, C. & Im, H. K. Shared and distinct genetic risk factors for childhood-onset and adult-onset asthma: genome-wide and transcriptome-wide studies. Lancet Respir. Med. 7, 509–522 (2019).
Chowdhury, N. U., Guntur, V. P., Newcomb, D. C. & Wechsler, M. E. Sex and gender in asthma. Eur. Respir. Rev. 30, 210067 (2021).
Jerschow, E. et al. Effect of relocation to the U.S. on asthma risk among Hispanics. Am. J. Prev. Med. 52, 579–588 (2017).
Han, Y.-Y. et al. Dietary patterns, asthma, and lung function in the Hispanic community health study/study of Latinos. Ann. Am. Thorac. Soc. 17, 293–301 (2020).
Wang, Z. et al. Microbial co-occurrence complicates associations of gut microbiome with US immigration, dietary intake and obesity. Genome Biol. 22, 336 (2021).
Kaplan, R. C. et al. Gut microbiome composition in the Hispanic community health study/study of Latinos is shaped by geographic relocation, environmental factors, and obesity. Genome Biol. 20, 219 (2019).
LaVange, L. M. et al. Sample design and cohort selection in the Hispanic community health study/study of Latinos. Ann. Epidemiol. 20, 642–649 (2010).
Sorlie, P. D. et al. Design and implementation of the Hispanic community health study/study of Latinos. Ann. Epidemiol. 20, 629–641 (2010).
Estrella, M. L. et al. Alternate healthy eating index is positively associated with cognitive function among middle-aged and older Hispanics/Latinos in the HCHS/SOL. J. Nutr. 150, 1478–1487 (2020).
Costello, M. et al. Characterization and remediation of sample index swaps by non-redundant dual indexing on massively parallel sequencing platforms. BMC Genom. 19, 332 (2018).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Pielou, E. C. The measurement of diversity in different types of biological collections. J. Theor. Biol. 13, 131–144 (1966).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Aitchison, J. The statistical analysis of compositional data. J. R. Stat. Soc. 44, 139–160 (1982).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.6-10, https://github.com/vegandevs/vegan (2025).
Hillmann, B. et al. Evaluating the information content of shallow shotgun metagenomics. mSystems 3, https://doi.org/10.1128/msystems.00069-18 (2018).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Lahti, L. & Sudarshan, S. microbiome R package. http://microbiome.github.io (2012).
Jun Chen, Xianyang Zhang, Lu Yang, Lujun Zhang. GUniFrac: Generalized UniFrac Distances, distance-based multivariate methods and feature-based univariate methods for microbiome data analysis. 1.8 https://doi.org/10.32614/CRAN.package.GUniFrac (2012).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979).
Pierce, B. L. & Burgess, S. Efficient design for mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am. J. Epidemiol. 178, 1177–1184 (2013).
Lopera-Maya, E. A. et al. Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch Microbiome Project. Nat. Genet. 54, 143–151 (2022).
Qin, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. https://doi.org/10.1101/2020.09.12.20193045 (2020).
Kurilshikov, A. et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat. Genet. 53, 156–165 (2021).
Global Biobank Meta-analysis Initiative Resources. Global Biobank Meta-analysis Initiative https://www.globalbiobankmeta.org/resources.
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Wang, Y. et al. Global Biobank analyses provide lessons for developing polygenic risk scores across diverse cohorts. Cell Genom. 3, 100241 (2023).
Lambert, S. A. et al. The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat. Genet. 53, 420–425 (2021).
John, C. R. MLeval: machine learning model evaluation. R package version 0.3. Comprehensive R Archive Network (CRAN) (2020).
National Heart, L., and Blood Institute, National Institutes of Health, U. S. Department of Health and Human Services. The NHLBI BioData Catalyst. Preprint at https://zenodo.org/record/3822858 (2020).
R. Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing.. https://www.R-project.org/ (Vienna, Austria, 2024).
R. C. Kaplan et al. EMBL-EBI European Nucleotide Archive. Burk_SOL GOLD. ERP117287. https://www.ebi.ac.uk/ena/browser/view/PRJEB34394.
Acknowledgements
HCHS/SOL is a collaborative study supported by contracts from the National Heart, Lung, and Blood Institute (NHLBI) to the University of North Carolina (HHSN268201300001I/N01-HC-65233), University of Miami (HHSN268201300004I/N01-HC-65234), Albert Einstein College of Medicine (HHSN268201300002I/N01-HC-65235), University of Illinois at Chicago (HHSN268201300003I/N01-HC-65236 Northwestern University), and San Diego State University (HHSN268201300005I/N01-HC-65237). The following have contributed to the HCHS/SOL through a transfer of funds to the NHLBI: National Institute on Minority Health and Health Disparities, National Institute on Deafness and Other Communication Disorders, National Institute of Dental and Craniofacial Research, National Institute of Diabetes and Digestive and Kidney Disease, National Institute of Neurological Disorders and Stroke, NIH Institution-Office of Dietary Supplements. Additional funding was provided to RK and RB: HHSN268200625235C, R01HL105756, R01HL136266, R01MD011389, R01HL140976, R01DK119268, R01HL060712, R01DK134672, R01DK120870, R01AG085320, and the New York Regional Center for Diabetes Translation Research (P30DK111022); to M.A.S.: BioData Catalyst Fellows Program (1OT3HL147154) and K01HL157658; and to FH: R01HL157069 and R01AI152504. None of the funding agencies had a role in the design, conduct, interpretation, or reporting of this study. The authors wish to acknowledge the contributions of the consortium working on the development of the NHLBI BioData Catalyst® (BDC) ecosystem.
Author information
Authors and Affiliations
Contributions
M.A.S., R.D.B., and R.C.K. designed the study. M.A.S. and N.G. reviewed relevant literature to inform the design. M.A.S. and E.L. curated the data and performed the analyses. C.H.A., E.M.L., and L.A.L. provided guidance on coding and analyses. K.L. performed the Mendelian Randomization analyses. MAS generated the figures and data visualizations. M.A.S., E.L., K.L., L.A.L., E.M.L., F.H., R.D.B., and R.C.K. were involved in the interpretation of results. K.B., C.L.A., and F.H. provided expertise related to asthma, the HCHS/SOL data, and/or the study population. MAS wrote the original draft with input from E.L.; C.A.H., K.L., L.A.L., E.M.L., K.A.M., F.H., K.E.N., R.D.B., and R.C.K. reviewed and edited the manuscript. R.C.K. supervised the project and assisted with data access. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Fernando Holguin is a member of the adjudication committee for Insmed Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jonathan Thorsen who co-reviewed with Anders Eliasen and Anton Kjellberg; Bor-Luen Chiang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Stanislawski, M.A., Litkowski, E., Arehart, C.H. et al. Relationships among host genetics, gut microbiota, and asthma in US Hispanic/Latino adults. Nat Commun 16, 10223 (2025). https://doi.org/10.1038/s41467-025-65028-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65028-z