Abstract
Immunotherapy has revolutionized cancer treatment, yet its efficacy is limited to a specific patient subset. This underscores the critical clinical demand for accessible assays capable of assessing immunotherapy outcomes and enabling timely adjustments to personalized medical care. Here, we present a multiplexed assay based on dual-target responsive DNA hydrogels for the simultaneous detection of soluble programmed death-ligand 1 and lactate dehydrogenase in lung cancer patients. The DNA hydrogel, constructed through a self-assembly strategy, achieves superior performance by reducing background noise by 33.8% and improving the signal-to-noise ratio by 61.5%. By integrating rolling circle amplification, the assay enables ultrasensitive detection at femtomolar levels. When further combining additional clinical biomarkers, the assay demonstrates strong predictive ability for immunotherapy response, achieving an area under the curve value of 0.935 in clinical cohort. Collectively, this blood-based assay offers a versatile and effective approach for advancing biomarker-based evaluations of immunotherapy outcomes.
Similar content being viewed by others
Introduction
Lung cancer (LC) is among the most prevalent cancers globally, with an estimated 2.2 million new cases and 1.8 million deaths annually1,2. Immunotherapy has opened a new era in the treatment of LC, particularly non-small cell LC, significantly increasing the 5-year survival rate of advanced cancer from approximately 5% in past decades to around 16% today3,4,5. However, only 20% to 30% of patients experience long-term benefits from immunotherapy, and some may even face accelerated disease progression6,7,8. Precise evaluation of immunotherapy efficacy would facilitate prompt adjustments to treatment strategies, including switching to alternative immunotherapy agents or combining with chemotherapy and radiotherapy, ultimately improving therapeutic outcomes9,10.
Current clinical evaluation of immunotherapy response is predominantly based on radiographic imaging (e.g., CT scans). Those methods suffer from delayed detection of therapeutic response, as tumor shrinkage may lag behind molecular immune activity by over three months, delaying clinical decisions11. In addition, frequent imaging exposes patients to cumulative radiation doses (10–20 mSv per scan), while pseudo-progression—a phenomenon where immune cell infiltration mimics tumor growth—can lead to misinterpretation and premature discontinuation of effective therapy12. These challenges highlight the urgent need for non-invasive, real-time biomarker monitoring to complement conventional imaging.
Liquid biopsy offers several advantages in disease diagnosis and prognosis, such as convenient specimen acquisition and ease of dynamic monitoring13,14,15. In particular, soluble programmed death-ligand 1 (sPD-L1) has been explored as a potential biomarker for assessing immunotherapy outcomes16,17. In addition, lactate dehydrogenase (LDH), derived from tumor cells or tumor microenvironment, has also demonstrated promise in assessing immunotherapy efficacy18,19,20. Currently, their detection primarily relies on enzyme-linked immunosorbent assay (ELISA), which face challenges such as dependence on antibodies21—often necessitating increased quantities (e.g., 5 μg/mL) for enhanced sensitivity, and difficulties in simultaneously detecting multiple biomarkers due to potential cross-reactivity22,23. Therefore, it is essential to develop a sensitive assay for the simultaneous quantitation of multiple biomarkers to improve the assessment of immunotherapy responses.
DNA hydrogels possess sequence programmability, precise framework structure, and facile chemical modification24,25,26. Especially, the structuring of DNA hydrogels enables the development of engineered nanocarriers with enhanced binding sites for biomarker detection. By precisely integrating aptamer sequences, DNA hydrogels can selectively hybridize to the targets, thereby facilitating their detection. Furthermore, cyclic reactions triggered by base pairing, such as rolling circle amplification (RCA), effectively enhance the detection signals27,28. While these characteristics establish DNA hydrogels as a robust platform for biomarker detection in the context of liquid biopsy, DNA hydrogel-based assays for multiple targets remain relatively rare due to the challenges in integrating these capabilities into a single test.
Here, we present a multiplexed assay utilizing dual-target responsive DNA hydrogels to evaluate immunotherapy efficacy. These hydrogels self-assembled from three partially complementary single-stranded DNA (ssDNA) sequences linked by a flexible DNA linker. The hydrogel reduces non-specific DNA interactions through steric hindrance and restricted diffusion, achieving an ultrasensitive detection limit down to ~ femtomolar (fM) level with lowered background noise and improved signal-to-noise ratio (SNR). This blood-based assay, demonstrated in a proof-of-concept study, assesses immunotherapy response by multiplexing variations in sPD-L1 and LDH levels.
Results
Assay design principle
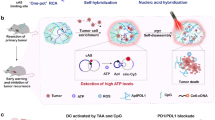
The working principle of the DNA hydrogel-based assay is illustrated in Fig. 1. We designed three ssDNA strands, P1, P2, and P3, which exhibit partial complementarity through base-pairing interactions at Binding region I/II (Fig. 1a). Each ssDNA strand (from 5’ to 3’) was functionally divided into four sequential regions: Binding region I, Binding region II, Reporter recognition region (III), and Linker binding region (inset, IV, Fig. 1a). These three ssDNA strands undergo cyclic reactions and self-assemble to form the building blocks (P1/P2/P3)n. Another building block (L1/L2/L3)n was created following a similar protocol. Both types of building blocks interacted with a dual-splint linker (C1/C2) through the Linker binding region (IV), facilitating the formation of the hydrogel structure. Consequently, DNA hydrogels were successfully synthesized with a high degree of programmability.
a Design of the self-assembled DNA hydrogels-based assay (inset: I: binding region I (1–12 base); II: binding region II (14–24 base); III: Reporter recognition region (25–50 base)); IV: Linker binding region (40–53 base). b Workflow for simultaneous detection of sPD-L1 and LDH from lung cancer (LC) samples for immunotherapy assessment. c Principle of the target-triggered rolling circle amplification (RCA) to enhance fluorescence signals. Abbreviations: MB magnetic bead; sPD-L1 soluble programmed death-ligand 1; LDH lactate dehydrogenase; Prog progressive after immunotherapy; No prog disease control after immunotherapy; Apt-P aptamer specific to sPD-L1; Apt-L aptamer specific to LDH.
In the assay, target-specific aptamers for sPD-L1 (Apt-P) and LDH (Apt-L) were immobilized on streptavidin-coated magnetic beads (MBs) via biotin-streptavidin interaction. Each aptamer partially hybridized with its corresponding DNA reporter (R1/R2) (Fig. 1b). Upon target binding in clinical samples, the respective reporters were released into the supernatant, isolated through magnetic separation, and subsequently added to the DNA hydrogels for detection. When the DNA reporters (R1/R2) paired with the circular DNA, RCA was initiated, producing long tandem-repeat DNA strands and enabling exponential amplification of R1/R2 (Fig. 1c). These amplified strands specifically recognized and hybridized with the Reporter recognition regions of (P1/P2/P3)n and (L1/L2/L3)n, forming double-stranded DNA (dsDNA) that contained restriction endonuclease (Esp3I) cleavage sites. The Reporter recognition regions and Linker binding regions of (P1/P2/P3)n were labeled with the fluorophore FAM (6-carboxy fluorescein) and the quencher BHQ-1, while those of (L1/L2/L3)n were labeled with Texas Red and BHQ-2. Cleavage by Esp3I restored the fluorescence signals of FAM and Texas Red, enabling the simultaneous sensitive detection of sPD-L1 and LDH.
Characterization of DNA hydrogel
We subsequently characterized the assembly process of the DNA hydrogel to validate its successful preparation. Statistical simulations illustrated how the ssDNAs interacted and the number of bases paired for each strand among the three designed ssDNAs. As shown in Fig. 2a, Binding region I of P1 preferentially hybridized with Binding region II of P3, while Binding region II of P1 paired with Binding region I of P2, and Binding region II of P2 bound to Binding region I of P3, enabling the DNA self-assembly process. The formation of (L1/L2/L3)n followed the same principle (Supplementary Fig. 1a).
a Statistical simulations of base-pairing probabilities and the number of paired bases for three ssDNAs (P1, P2, P3). b Predicted absolute value of minimum free energy of hybridization between pairs of P1, P2, P3 or L1, L2, L3 at equilibrium. c Free energy of the self-assembled DNA structure formed by different numbers of ssDNA. d Free energy of different DNA structures in a DNA hydrogel. a: the linker (C1/C2); b: the complex formed by P1, P2, and P3; c: the complex formed by a and b; d: the complex formed by L1, L2, and L3; e: the complex formed by a and d; f: the complex formed by a, b, and d. e Schematic illustration and predicted melting curves of the linker and the hybridization between buildings blocks (P1/P2/P3)n, (L1/L2/L3)n and the linker (C1/C2). T1: the melting temperature of the linker (C1 paired with C2). T2: the melting temperature of (P1/P2/P3)n paired with C2. T3: the melting temperature of (L1/L2/L3)n paired with C1. f Polyacrylamide gel electrophoresis (PAGE) analysis of the formation of DNA hydrogel. M: marker; Lane a: P1; Lane b: P2; Lane c: P3; Lane d: P1/P2; Lane e: (P1/P2/P3)n; Lane f: L1; Lane g: L2; Lane h: L3; Lane i: L1/ L2; Lane j: (L1/L2/L3)n; Lane k: (P1/P2/P3)n + (L1/L2/L3)n + C1/C2. g Scanning electron microscopy image of a DNA hydrogel. Scale bar: 50 µm. h Fluorescence images of the DNA hydrogel under white light (i), blue light excitation (ii, excitation: 470 nm, emission: 525 nm), and green light excitation (iii, excitation: 545 nm, emission: 605 nm), and the merged channels (iv). Scale bar: 200 µm. Exposure time: 1/1.5 s, gain: + 6 dB. The experiments in (h) were repeated independently three times with similar results.
Thermodynamic calculations demonstrated that the predicted absolute minimum free energies of P1/P2 (33.23 kcal/mol), P1/P3 (30.57 kcal/mol), and P2/P3 (30.81 kcal/mol) hybrids were significantly higher than those of the corresponding self-dimers (P1/P1: 8.84 kcal/mol; P2/P2: 11.49 kcal/mol; P3/P3: 7.46 kcal/mol) (Fig. 2b). This finding indicates a preference for interstrand pairing over self-hybridization. A similar trend was observed for L1, L2, and L3, with interstrand hybrids demonstrating higher absolute free energy compared to self-dimers. Modeling of self-assembled structures with one to six ssDNA strands showed that as the number of strands increased, the absolute free energy also rose, indicating enhanced structural stability (Fig. 2c). In addition, connecting two building blocks with complementary linkers resulted in a more stable structure, as evidenced by a further increase in absolute free energy (Fig. 2d). The cleavage within a similar temperature range suggested conditions favorable for hydrogel formation (Fig. 2e).
The formation of the DNA assembly was confirmed by polyacrylamide gel electrophoresis (PAGE). As depicted in Fig. 2f, the molecular weights of the assembled structures, (P1/P2/P3)n and (L1/L2/L3)n (lane e and lane j), were significantly higher than those of the individual ssDNA strands. The successful synthesis of the linker was validated via PAGE analysis (Supplementary Fig. 1b). When the two building blocks (lane 2 and 3) were connected with the linker, the resulting hydrogel (lane k) exhibited an even larger molecular weight (Fig. 2f).
Visual observations confirmed the successful DNA hydrogels, as the DNA hybridization solution lost its fluidity and adhered to the tube wall upon inversion (Supplementary Fig. 1c). Scanning electron microscopy (SEM) at varying magnifications consistently illustrated the porous structure of the DNA hydrogel (Fig. 2g and Supplementary Fig. 1d). In addition, the fluorescence properties of FAM and Texas Red in the DNA strands were effectively retained following hybridization and hydrogel formation (Fig. 2h, exposure time: 1/1.5 s, gain: + 6 dB).
These results demonstrate the successful formation of the DNA hydrogel with a cross-linked network structure, which provides abundant binding sites for DNA reporters and facilitates dual-target detection in complex biosamples.
Assay feasibility
We initially investigated the background fluorescence signals of the DNA hydrogel, as elevated background noise can compromise measurement accuracy. Exonuclease I (Exo I) was used to digest the DNA complexes in both hydrogel and non-gel solutions. We conducted a dose-response analysis by varying FAM probe concentrations (0.3, 0.5, and 0.7 mM) while maintaining a fixed reaction volume of 5 μL. Using FAM as a representative fluorophore, we found that the hydrogel assay exhibited a maximum 33.8% lower background fluorescence and a 61.5% higher SNR compared to the solution assay (Fig. 3a). These results underscore the enhanced reaction efficiency of the hydrogel across tested probe concentrations.
a Comparison of signal-to-noise ratio (SNR) between DNA hydrogels and DNA solutions at different concentrations (0.3 mM, 0.5 mM, and 0.7 mM) after digestion with 5 µL of Exonuclease I at 37 °C for 1 h. The SNR was calculated as the ratio of post-digestion to pre-digestion fluorescence intensity. Data represent mean ± s.d. (n = 3 independent experiments). b, c Decay profiles (b) and fluorescence reduction ratios (c) of two fluorophores under sustained light excitation in DNA hydrogel and solution. Data represent mean ± s.d. (n = 3 independent experiments). d Free energy of different ssDNAs (P1, P2, P3, L1, L2, and L3) paired with R1 and R2, respectively. The rate of fluorescence reduction is defined as the ratio of the reduced fluorescence intensity to the initial intensity. e Schematic of fluorescence quenching due to hybridization of two ssDNAs in DNA hydrogel and solution. f Kinetics of fluorescence produced by the hybridization of P1–2 and R2–2 in DNA hydrogel and solution. Fluorescence was monitored at excitation wavelengths of 492 nm and emission wavelengths of 518 nm. g, h Fluorescence signals of the assay for detecting 1000 pg/mL sPD-L1 (g) and 2000 U/L LDH (h) with and without Esp3I. Blank: fluorescence of hydrogel alone. i Fluorescence signal changes of the assay with and without fluorophore/quencher modification for detecting 80 pg/mL sPD-L1 and 20 U/L LDH. Data represent mean ± s.d. (n = 3 independent experiments). j, k Fluorescence signals (j) and corresponding linear correlation (k) for serial concentrations of sPD-L1 detected using DNA hydrogel with RCA reaction as a reference for evaluating RCA amplification efficiency. The sPD-L1 concentrations: A, 1000 pg/mL; B, 200 pg/mL; C, 60 pg/mL; D, 10 pg/mL; a, 800 pg/mL; b, 200 pg/mL; c, 40 pg/mL; d, 12 ng/mL. Data represent mean ± s.d. (n = 3 independent experiments). l Optimization of RCA reaction time. Data represent mean ± s.d. (n = 3 independent experiments).
To further evaluate background interference for sample analysis, we collected fluorescence microscopy images under the strictly controlled parameters (exposure time: 1/1.5 s, gain: +6 dB) for both blank controls and target-containing samples (Supplementary Fig. 2). The blank control exhibits negligible background fluorescence, while the target-dependent fluorescence observed in other wells increases predictably with sPD-L1/LDH concentrations (8–6400 pg/mL, 20–2000 U/L). These results confirm the assay’s low background interference.
We subsequently examined the stability of fluorescence under continuous laser irradiation, which typically leads to photobleaching and compromises detection reliability29,30,31. We measured the signal attenuations of Texas Red and FAM in both hydrogel and solution states (Fig. 3b). As demonstrated in Fig. 3c, Texas Red exhibited lower fluorescence decay in the hydrogel (24% ± 1.9%) compared to the solution (31% ± 1.6%), while FAM displayed similar levels of attenuation in both states (see detailed experimental protocol in Supplementary Information). The hydrogel appears to protect Texas Red but not FAM. This selective photoprotection arises from the intrinsic differences in photophysical properties of fluorophores rather than measurement bias. We thus conducted controlled photostability experiments using single-stranded DNA (P1) labeled individually with FAM or Texas Red under identical irradiation conditions (5000 s of continuous laser exposure). FAM exhibited a fluorescence intensity reduction of 24.9% over the irradiation period, whereas Texas Red showed only 7.8% reduction (Supplementary Fig. 3). This discrepancy aligns with established photophysical principles indicating that FAM, a member of the fluorescein family, is inherently more susceptible to photobleaching due to its chemical structure and excitation dynamics, which render it prone to reactive oxygen species-mediated degradation32. In contrast, Texas Red, a rhodamine-based fluorophore, demonstrates superior intrinsic photostability, allowing the hydrogel to effectively maintain its fluorescence. Together, these findings suggest that the structure of the DNA hydrogel contributes to a reduction in fluorescence decay.
Furthermore, we explored the cross-reactivity between hydrogel building blocks and non-target reporters. Thermodynamic predictions revealed that P1, P2, and P3 exhibited a stronger binding affinity for R1 than for R2, with a 1.2-fold increase in absolute values of free energy (Fig. 3d). In contrast, L1, L2, and L3 preferred binding to R2, showing 1.6-fold higher absolute free energy values than when bound to R1. Simulations confirmed that in the presence of both reporters, P1, P2, and P3 exclusively bound to R1, while L1, L2, and L3 paired solely with R2 (Supplementary Fig. 4a, b), demonstrating minimal cross-reactivity.
For further validation, we removed the fluorescence modification from the ssDNA and selectively modified the 3’ end of P1 with the fluorophore FAM (designated P1-2) while labeling the 5’ end of the interfering reporter R2 with the quencher BHQ-1 (designated R2-2). When P1-2 hybridized with R2-2, FAM and BHQ-1 were brought into close proximity, resulting in quenching via energy transfer (Fig. 3e). The fluorescence intensity decreased by approximately 8% in the hydrogel compared to a 21% decrease in the solution (Fig. 3f). We attributed the reduction in non-specific DNA interactions to steric hindrance and restricted diffusion33,34,35.
We developed a multiplexed assay for the detection of sPD-L1 and LDH, utilizing restriction endonuclease Esp3I for cleavage. The specific cleavage sites (blue region, Supplementary Fig. 5a) were activated upon dsDNA formation between P1 and R1 or L1 and R2 (Supplementary Fig. 5b, c). The fluorescence curves indicated an increase in signal at the maximum emission wavelength of 518 nm for FAM and 615 nm for Texas Red, which occurred exclusively after Esp3I cleavage, thus confirming that the fluorescence originated from enzymatic activity (Fig. 3g, h). In addition, significant signal changes (P < 0.001) were observed only when the building blocks were modified with both fluorophore and quencher, demonstrating the effectiveness of this configuration (Fig. 3i). In the absence of the target biomarkers, no significant changes were detected, confirming the feasibility of the assay for target detection (Supplementary Fig. 6a, b).
RCA was employed to enhance sensitivity for low-concentration targets36,37,38. Supplementary Fig. 6c, d confirmed the successful synthesis of circular DNA. Exo I effectively degraded ssDNA but had no effect on circular DNA39,40,41. The electrophoresis bands of ssDNA remained intact after cyclization and Exo I digestion, whereas untreated ssDNA was completely degraded by Exo I. To evaluate the amplification efficiency of RCA process, we analyzed RCA reactions within DNA hydrogel using a serial dilution of target sPD-L1 (Fig. 3j, k). Under the RCA reaction conditions, four sPD-L1 concentrations (1000, 200, 60, and 10 pg/mL, denoted as points A–D) were tested (Fig. 3j), resulting in fluorescence signals that linearly correlated with concentration (calibration curve: Y = 1591.9·X + 417.26; Fig. 3k). To isolate RCA’s amplification effect, we further measured non-RCA fluorescence at four higher sPD-L1 concentrations (800, 200, 40, and 12 ng/mL, labeled a–d in Fig. 3j) and calculated their equivalent RCA-amplified sPD-L1 concentrations using RCA calibration curve. The ratio of non-RCA to RCA concentrations revealed that the hydrogel-supported RCA achieved 1262.0 ± 125.06-fold amplification (Supplementary Table 2). This substantial amplification validated the critical role of the platform in achieving high sensitivity. Optimization experiments indicated that a 100-minute RCA reaction time and a phi29 DNA polymerase concentration of 0.5 U/μL produced the highest signal intensity, establishing these parameters as optimal conditions (Fig. 3l and Supplementary Fig. 6e).
To evaluate enzymatic accessibility, we compared Esp3I endonuclease digestion efficiency between hydrogel and solution assays (Supplementary Fig. 7a). Fluorescence measurements after 30 min of cleavage revealed no statistically significant difference in signal intensity (P > 0.05; Supplementary Fig. 7b, c), demonstrating that the porous structure of hydrogel allows unrestricted enzyme diffusion and substrate access, preserving cleavage efficiency. These findings align with previous studies showing that the interconnected 3D networks of hydrogels support enzymatic activity by facilitating unimpeded molecular transport42,43.
Evaluation of analytical performance
The analytical performance of the DNA hydrogel-based assay was systematically evaluated through serial dilutions of the target proteins (sPD-L1 and LDH) under optimized conditions. As illustrated in Fig. 4a, the fluorescence intensity increased progressively with the concentration of sPD-L1, exhibiting a linear dynamic response ranging from 8 pg/mL to 16 ng/mL (Fig. 4b). The limit of detection (LOD) for sPD-L1 was determined to be 1.33 pg/mL, corresponding to fM levels. Similarly, LDH detection exhibited a linear range of 5 U/L to 2000 U/L with a LOD of 0.72 U/L (Fig. 4c, d). Stable fluorescence signals were achieved, with a maximum relative standard deviation (RSD) of 1.4% for sPD-L1 (Fig. 4e) and 2.0% for LDH (Supplementary Fig. 8a).
a, b Fluorescence signals of DNA hydrogel (a) and the corresponding linear calibration curve of sPD-L1 (b) at various concentrations. Data represent mean ± s.d. (n = 3 independent experiments). c, d Fluorescence signals of DNA hydrogel (c) and the corresponding linear calibration curve of LDH (d) at various concentrations. Data represent mean ± s.d. (n = 3 independent experiments). e Repeatability of the assay for detecting sPD-L1. f Specificity of the assay for detecting sPD-L1 (6400 pg/mL) and LDH (2000 U/L) against different interferents. 5’-Nucleotidase (5’-NT) and deoxyribonuclease I (DNase I) were tested at 10 U/L, and other interferents were evaluated at 100 ng/mL. g Stability of the DNA hydrogel for detecting 800 pg/mL sPD-L1 after storage at 4 °C for different days. Data represent mean ± s.d. (n = 3 independent experiments). h Correlation between the parallel measurements by DNA hydrogel and enzyme-linked immunosorbent assay (ELISA) for sPD-L1. i Correlation between the parallel measurements by DNA hydrogel and colorimetry for LDH. Linear fitting was performed at the 95% confidence level.
Comparative performance analyses between hydrogel-vs-solution systems for detecting different concentrations of sPD-L1 and LDH were performed with integrated RCA steps under identical reaction conditions (Supplementary Fig. 9a). As a result, the hydrogel system outperformed the solution-phase system (P < 0.001, Supplementary Fig. 8b, c), exhibiting a significant overall advantage in detection performance. These findings align with our previous comparisons (Fig. 3a–f), further validating the robustness of the hydrogel approach.
Common interferents and endogenous nucleases typically present in human serum, including phosphodiesterase (PDE), 5’-nucleotidase (5’-NT), and deoxyribonuclease I (DNase I), were utilized to assess the specificity of the assay. As shown in Fig. 4f, robust fluorescence signals were detected for both sPD-L1 and LDH, while minimal signals (< 3.6% for sPD-L1 and < 4.3% for LDH) were observed from interferents (e.g., nucleases or other serum components). Furthermore, the hydrogel encapsulation strategy effectively shields the probes from nonspecific enzymatic degradation, as evidenced by the consistent signal intensity in both blank and nuclease-spiked samples (P > 0.05, Supplementary Fig. 8b, c). These results validate the assay’s specificity in complex biological matrices and reinforce its reliability for clinical use.
The stability of the DNA hydrogel was further evaluated by storing it at 4 °C. The hydrogel used for detecting sPD-L1 (800 pg/mL) retained 94.2% of its original fluorescence intensity after 30 days of storage (Fig. 4g). In addition, the fluorescence intensity was preserved following four cycles of lyophilization and rehydration, exhibiting a retention rate of 95.6% (Supplementary Fig. 8d).
As a result, the DNA hydrogel assay demonstrated strong correlations with ELISA kits for sPD-L1 (Pearson’s r = 0.982) and with biochemical kits for LDH (Pearson’s r = 0.977) (Fig. 4h, i). Notably, the linear detection range of our assay for sPD-L1 (8–16,000 pg/mL, Supplementary Fig. 8e) was significantly broader, offering a maximum 10-fold enhancement compared to commercial ELISA kits (e.g., R&D, 25–1600 pg/mL; see additional comparisons in Supplementary Table 3) and state-of-the-art assays44,45. The assay also exhibited a maximum 5-fold increase in the linear detection range for LDH (5–2000 U/L) compared to reported platforms (Supplementary Table 4, Supplementary Fig. 8f)46. Regarding clinical relevance, the linear detection range of our assay (8–16,000 pg/mL for sPD-L1 and 5–2000 U/L for LDH) fully encompasses clinically reported biomarker levels16,47. Furthermore, our assay achieves an ultra-low LOD of 1.33 pg/mL for sPD-L1 and 0.72 U/L for LDH, representing an up to 35-fold improvement in sensitivity compared to conventional clinical methods (Supplementary Table 3, 4). This significant enhancement in LOD—particularly for low-abundance biomarkers like sPD-L1—validates the system’s ability to detect clinically relevant concentrations that existing techniques may overlook.
To evaluate multi-target capacity, we demonstrated modular incorporation of additional biomarker such as carcinoembryonic antigen (CEA). For example, by replacing partial sequences in the sPD-L1-targeting module with CEA-specific sequences, a CEA detection module can be incorporated into the hydrogel to enable simultaneously detection of these biomarkers (Supplementary Fig. 10a). The CEA-specific fluorescence showed a linear range of 5 pg/mL–20 ng/mL (R² = 0.991) and a low LOD of 0.84 pg/mL (Supplementary Fig. 10b, c). The hydrogel enabled simultaneous detection of sPD-L1, LDH, and CEA within a single assay (Supplementary Fig. 10d), validating its multiplexing capability. Our DNA hydrogels address a critical gap in traditional assays: their modular architecture allows easy substitution of functional units (e.g., aptamers, primers) to target diverse biomarkers, a feature absent in conventional hydrogel or solution-based systems. This modular architecture is critical for clinical applications requiring scalable, multiplexed biomarker panels.
Proof-of-concept application for clinical use
To demonstrate the clinical translational potential of our DNA hydrogel analysis, we conducted analyses using patient-derived clinical serum samples. A CONSORT flowchart for patient enrollment and analysis subsets is shown in Supplementary Fig. 11. Detailed inclusion and exclusion criteria for patient recruitment, baseline clinical characteristics, such as patient treatment regimens and sampling time points, are summarized in Supplementary Table 5 and detailed in the Supplementary Information.
Before evaluating the assay’s capability to assess immunotherapy responses, we first benchmarked its performance against established diagnostic assays for clinical sample analysis. We conducted a direct, sample-by-sample comparison for every patient in the study cohort between our hydrogel-based assay and established diagnostic assays. For sPD-L1 quantification, we implemented the standard ELISA as a reference method and compared its results to those obtained with our hydrogel platform. Likewise, for LDH, we employed a standard colorimetric assay based on the lactate–pyruvate reaction and performed a parallel comparison. As illustrated in Supplementary Table 6 and Supplementary Fig. 12a, b, our method demonstrated strong concordance with standard assays (sPD-L1 via ELISA; LDH via colorimetry) across all clinical samples, achieving agreement rates of 96.9% for sPD-L1 and 97.6% for LDH. This confirms that our platform delivers results consistent with established assays at the individual biomarker level.
We then determined its feasibility in detecting sPD-L1 and LDH for differentiating between benign and malignant tumors. We collected clinical serum samples from healthy donors (HD group, n = 22) and patients with LC (n = 22). All HD participants had no history of malignancy. All LC patients were pathologically and radiologically diagnosed, with the exclusion of those with secondary primary cancers.
Our analysis revealed a significant elevation of sPD-L1 concentrations in LC patients compared to healthy controls (P < 0.0001, Fig. 5a, b)48,49. Moreover, sPD-L1 levels in patients with metastatic LC were notably higher than those with localized LC (n = 11, P < 0.05, Supplementary Fig. 13a), in line with the previous studies50. A similar trend was observed for LDH, with significantly increased levels in LC patients versus the HD group (P < 0.05, Fig. 5c, d), and notably higher levels in metastatic patients compared to those with localized tumors (n = 11, P < 0.05, Supplementary Fig. 13b)51,52. These findings suggest the real-time regulation of sPD-L1 and LDH with tumor development and progression, highlight their potential as biomarkers for indicating and evaluating therapeutic outcomes.
a, b Heatmap (a) and corresponding scatter plots (b) of sPD-L1 levels in serum samples from healthy donors (HD) and patients with LC. Data represent mean ± s.d. (n = 22 for each group, two-tailed Mann–Whitney tests, P = 0.000031). c, d Heatmap (c) and corresponding scatter plots (d) of LDH levels in serum samples from HD and LC groups. Data represent mean ± s.d. (n = 22 for each group, two-tailed Mann–Whitney tests, P = 0.041). e, f sPD-L1 (e) and LDH levels (f) in patients who achieved disease control before and after immunotherapy (n = 9 for each group, two-tailed paired t test, P = 0.000214 for (e), P = 0.001 for (f)). g, h sPD-L1 (g) and LDH levels (h) of LC patients with progression (prog and non-progression (No prog). Data represent mean ± s.d. (n = 15 for each group, two-tailed Student’s t test, P = 0.003 for (g); two-tailed Welch’s t test, P = 0.005 for (h)). i Comparison of 1-year progression rates with different changes in sPD-L1 and LDH levels after immunotherapy (n = 38 patients). “-” indicates no increase of sPD-L1 or LDH. “+” indicates the increase of sPD-L1 or LDH. j Clinical outcomes of patients by changes in sPD-L1 and LDH levels after immunotherapy. k Receiver operating characteristic curves of different biomarkers and the combined multiple biomarkers (SUM) for predicting progression after immunotherapy. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
After validating the feasibility of detecting sPD-L1 and LDH, we adapted a DNA hydrogel-based assay to evaluate immunotherapy responses, serving as a proof-of-concept demonstration for clinical application. Tumor responses in LC patients after immunotherapy were assessed based on the revised Response Evaluation Criteria in Solid Tumors, with responses categorized as complete response (CR), partial response (PR), stable disease (SD), and progressive disease (PD). Patients with disease control (CR + PR + SD) showed a decrease in sPD-L1 levels post-immunotherapy compared to pre-treatment levels, consistent with previous findings (n = 9, P < 0.001, Fig. 5e)16. A similar trend was noted for LDH levels (Fig. 5f)53. In contrast, patients with disease progression after immunotherapy had increased sPD-L1 levels compared with pretreatment, with a similar trend in LDH levels (n = 9, P < 0.01 and P < 0.0001, Supplementary Fig. 13c, d). In addition, serum samples from progressive patients (n = 15) were analyzed and showed that they had higher levels of sPD-L1 and LDH compared to non-progressive individuals (n = 15, P < 0.01, Fig. 5g, h).
Furthermore, we evaluated the association between biomarker dynamics and clinical progression in a clinical cohort of LC patients (n = 38, Fig. 5i, j). Patients exhibiting increasing sPD-L1 levels following immunotherapy had a significantly higher progression rate (58.3%) compared to those with stable levels (16.7%). Similarly, elevated LDH levels were associated with a higher risk of progression (P < 0.05). Patients demonstrating simultaneous increases in both sPD-L1 and LDH showed a notable progression rate of 75%, compared to those experiencing an increase in only one biomarker (P < 0.05).
The assay’s performance in evaluating immunotherapy outcomes was assessed using the receiver operating characteristic (ROC) curve in a clinical cohort (n = 36, Fig. 5k and Supplementary Table 7). The area under the curve (AUC) for sPD-L1 and LDH in distinguishing progressive patients from non-progressive controls was 0.794 and 0.755, respectively (Supplementary Table 8). In contrast, traditional single-biomarker detection, including CEA, cytokeratin fragment 19 (CYF21-1), interleukin-6 (IL-6), squamous cell carcinoma antigen (SCCA), and neuron-specific enolase (NSE), yielded modest AUC values, ranging from 0.581 (95% confidence interval (CI), 0.387–0.775) for IL-6 to 0.761 (95% CI, 0.597–0.926) for SCCA. Notably, the combination of sPD-L1 and LDH improved the AUC to 0.880 (95% CI, 0.764–0.996), with a sensitivity of 81.8% and a specificity of 92.9%. Importantly, the comprehensive combination of all clinical accessible biomarkers optimally differentiated progressive patients from non-progressive patients, achieving an AUC of 0.935 (95% CI, 0.858–1.000), with a sensitivity of 90.9% and a specificity of 85.7%. These findings underscore the high sensitivity, specificity, and robustness of our method for biomarker analysis in liquid biopsy-based immunotherapy prognosis.
We further performed a confusion matrix analysis comparing our platform’s multiplexed predictions (sPD-L1 + LDH) against “gold standard” clinical imaging results (e.g., CT for tumor burden). Our method achieved a prediction accuracy of 86.1%, surpassing the 77.8% accuracy of the combined conventional ELISA (for sPD-L1)/colorimetry assays (for LDH) (Supplementary Fig. 14a–c).
We also compared the performance of our hydrogel system with traditional DNA solution-based assays in the same clinical cohort (n = 36) for both sPD-L1 and LDH detection. For sPD-L1, the hydrogel-based assay demonstrated higher sensitivity (63.6% vs. 59.1%) and higher specificity (100% vs. 85.7%) compared to the solution-based method (Supplementary Fig. 14d, e and Supplementary Table 8). Similarly, for LDH, the hydrogel assay showed improved performance with enhanced sensitivity (72.7% vs. 68.2%) and specificity (78.6% vs. 71.4%) relative to the solution method (Supplementary Fig. 14d, e and Supplementary Table 8). When integrating sPD-L1 and LDH results, the hydrogel-based assay further outperformed the solution-based approach, yielding a higher AUC (0.880 vs. 0.825), increased sensitivity (81.8% vs. 77.3%), and improved specificity (92.9% vs. 85.7%) (Supplementary Fig. 14f). Collectively, these findings indicate that the DNA hydrogel-based assay offers improved predicting sensitivity and specificity compared to traditional DNA solution assays.
Discussion
Immunotherapy has become the standard treatment for advanced or metastatic LC. However, only approximately 20% of patients experience sustained positive responses, while the remainder may encounter severe side effects or no response at all54,55. We developed a DNA hydrogel-based assay employing self-assembled building blocks ((P1/P2/P3)n and (L1/L2/L3)n) for the sensitive and multiplexed detection of dual targets, sPD-L1 and LDH, to assess the response to immunotherapy.
There are two unique advantages of employing DNA hydrogels instead of a solution-based system, from the perspective of amplification efficiency and detection specificity. For amplification, the hydrogel-based systems demonstrated a maximum 47% increase in fluorescence intensity compared to the solution-phase systems (Supplementary Fig. 8). For specificity, while solution-phase systems can support basic signal amplification, they are limited by signal crosstalk and lack of compartmentalization. In contrast, the porous structure of the hydrogel spatially restricts the diffusion of RCA products, leading to a 61.9% reduction in mismatch rate (Fig. 3e, f) and a 61.5% improvement in SNR compared to solution-phase systems (Fig. 3a). Consequently, the DNA hydrogel achieved a broad detection range of 8 pg/mL to 16 ng/mL for sPD-L1 (LOD of 1.33 pg/mL) and 5 U/L to 2000 U/L for LDH (LOD of 0.72 U/L). These metrics outperformed those of commercial assays and state-of-the-art sensors (Supplementary Table 3 and Supplementary Table 4).
Furthermore, the DNA hydrogel enables simultaneous detection of multiple biomarkers. The lower cross-reactivity and noise suppression within the hydrogel matrix enable reliable signal detection and thus facilitate rapid adaptation to multi-biomarker detection (Supplementary Fig. 10a). For instance, replacing the sPD-L1 recognition sequence with a CEA-specific sequence enabled simultaneous detection of three biomarkers within the same matrix without cross-interference (Supplementary Fig. 10b–d). The modular architecture of DNA hydrogels provides enhanced specificity, which is critical for clinical applications that rely on scalable, multiplexed biomarker panels. The combination of sPD-L1 and LDH exhibited superior performance in differentiating progression patients from non-progression controls, achieving an AUC of 0.880 (see Supplementary Table 8). In contrast, lower AUCs ranging from 0.581 to 0.761 were observed for clinically approved biomarkers such as CEA, CYF21-1, IL-6, SCCA, and NSE. Notably, the integration of these two biomarkers with additional clinical markers further enhanced assessment, yielding an AUC of 0.935 and improving accuracy in evaluating immunotherapy progression.
Currently, tumor shrinkage assessed through computed tomography (CT) or magnetic resonance imaging (MRI) remains the standard method for evaluating treatment response. However, these approaches often delay progression detection by three months or more and struggle to identify early signs of relapse compared to blood biomarkers56. Approximately 14% of patients may experience “pseudo-progression”, where tumors appear larger despite effective treatment, complicating clinical decision-making and therapeutic evaluations57,58. In contrast, this assay provides a blood-based alternative for early and continuous monitoring. Utilizing readily accessible blood samples enables real-time assessment of immunotherapy responses, which promises to reduce the risk of progression. The modular structure of the hydrogel allows for adaptation to other biomarkers, indicating its potential in personalized medicine.
While this study shows promising, it is not without limitations. Hydrogel preparation involves multi-step protocols. However, this can be addressed through batch production using standardized parallel reactors (such as hybrid reactions of P1, P2, and P3) or integrating with microfluidic automation for reproducibility and scalability (such as batch addition of phi29 DNA polymerase). Overlapping signals from multiple fluorophores may occur. This can be addressed by using microplate-based detection, where spatial separation minimizes interference. In addition, this study exclusively enrolled patients receiving PD-1/PD-L1 therapy, and thus the assay’s performance remains unvalidated in populations treated with newer bispecific agents (e.g., PD-1/VEGF therapies). Given the increasing clinical adoption of bispecific antibodies, future validation in such cohorts is essential to confirm generalizability. Ongoing efforts will concentrate on expanding the assay’s applicability to diverse treatment regimens and integrating it into portable devices for point-of-care applications, thereby facilitating its translation into clinical practice.
Methods
Materials and reagents
DNA oligonucleotides (listed in Supplementary Table 1), TE (Tris-EDTA) buffer, and Tris-borate-EDTA (TBE) buffer were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). All oligonucleotides were dissolved in 10 mM TE buffer (pH 8.0) and stored at − 20 °C before use. Streptavidin-coated magnetic beads and DNA ladders were purchased from Thermo Fisher (USA). T4 DNA Ligase, T4 Polynucleotide Kinase, phi29 DNA polymerase, and the restriction endonuclease Esp3I were obtained from New England Biolabs (USA). Exonuclease I and dNTP mixture were purchased from Takara Co., Ltd. (China). DNA circligase was purchased from Lucigen (UK). Programmed death-ligand 1 (PD-L1) was purchased from Sino Biological (China). Lactate dehydrogenase (LDH) was purchased from Sinopharm Chemical Reagents (China). The enzyme-linked immunosorbent assay (ELISA) kit for sPD-L1 was purchased from Multi Sciences (China). Bovine serum albumin (BSA) was purchased from Yeasen Biotech (China). Tumor biomarkers were purchased from Tellgen Co. Ltd. (China), including carcinoembryonic antigen (CEA), neuron-specific enolase (NSE), squamous cell carcinoma antigen (SCCA), pro-gastrin-releasing peptide (ProGRP), and cytokeratin fragment 19 (CYF21-1). IL-6 was purchased from Cellgene (China). Ammonium persulphate and N, N, N’, N’-Tetramethylethylenediamine (TEMED) were purchased from Beyotime (China). Acrylamide-bisacrylamide solution was obtained from Epizyme (China). Immunoglobulin G (IgG) and human chorionic gonadotropin (hCG) were purchased from Sigma-Aldrich (USA). All reagents were of analytical reagent grade and used without further purification.
Apparatus
Scanning electron microscopy (SEM) was done on a Sigma 500 SEM (Zeiss, Germany). Atomic Force Microscope (AFM) was undertaken using a Dimension ICON system (Bruker, Germany). The DNA hydrogel was synthesized in a thermal cycler (Thermo Fisher, USA). The fluorescence images of the DNA hydrogel were observed with a BZ-X810 fluorescence microscope (Keyence, China). Fluorescence measurements were performed on a R-7000 spectrofluorophotometer (Hitachi, Japan). Colorimetric detection of LDH for parallel analysis were obtained by a Beckman Coulter AU biochemistry analyzer at Shanghai Chest Hospital. The detection of sPD-L1 by ELISA was performed on an EPOCH2 microplate reader (BioTek, USA).
Synthesis of DNA hydrogel
For the synthesis of the building block (P1/P2/P3)n, 2 mM of P1, P2, and P3 were mixed in a buffer containing 10 mM Tris-HCl and 50 mM NaCl. The mixtures were then heated to 95 °C for 5 min and cooled slowly to room temperature. Similarly, the building block (L1/L2/L3)n was formed by mixing L1, L2, and L3 following the same annealing procedure. For the synthesis of the linker (C1/C2), oligonucleotide strands C1 and C2 were each individually mixed with 10 mM ATP solution, 10 U/μL of T4 polynucleotide kinase, and 4 μL of T4 polynucleotide kinase reaction buffer, and then incubated at 37 °C for 2 h. Subsequently, the two mixtures were heated at 70 °C for 10 min. After that, the two mixtures were added simultaneously to a buffer containing 10 mM Tris-HCl and 50 mM NaCl, heated to 95 °C for 5 min, and then cooled to room temperature to form the linker. The two aforementioned building blocks were mixed with the linker, and the mixture incubated at 50 °C for 2 h, then cooled to room temperature. Subsequently, 5 μL of T4 DNA ligase was added, and the mixture was incubated overnight at 16 °C to obtain the DNA hydrogel. For the comparative experiment, all the previously mentioned components were mixed together at room temperature for analysis.
Preparation of circular DNA
A 10 μM solution of circular template oligonucleotide (cirDNA-P/cirDNA-L) was mixed with 2 µL of circligase reaction buffer, 1 µL of 1 mM ATP, 1 µL of 50 mM MnCl2, and 1 µL of DNA circligase. The mixture was incubated at 60 °C for 3 h. Subsequently, the circligase was inactivated by heating at 80 °C for 10 min. Next, 1 µL of Exonuclease I was added to the mixture and incubated at 37 °C for 45 min to digest the linear single-stranded DNA (ssDNA), followed by inactivation by heating at 80 °C for 15 min. The resulting reaction product was finally stored at 4 °C.
Fabrication of DNA hydrogel-based assay
A 50 μM solution of aptamer (Apt-P/Apt-L) was incubated with 50 μM reporter (R1/R2) at 37 °C for 1 h to form the Apt-P@R1 and Apt-L@R2 duplexes. The DNA duplex mixture (1 μM) was then incubated with 100 µL of streptavidin-coated magnetic beads (10 mg/mL) at room temperature for 30 min. Then, the mixture was washed several times with 2 M NaCl and resuspended in 100 µL TE buffer to form dsDNA-modified magnetic beads. Different concentrations of targets (sPD-L1/LDH) were added to the modified beads for 1 h, where they were captured by the aptamer (Apt-P/Apt-L), releasing the reporter (R1/R2) into the solution. The mixture was exposed to a magnet for 5 min, and the sediment was isolated using a magnet. The supernatant was added to the synthesized DNA hydrogel, along with 2 mM dNTPs, 0.5 U/μL phi29 DNA polymerase, 1 × phi29 buffer, 2 μL of circular DNA, and reacted at 37 °C for 100 min. Subsequently, Esp3I (10 U/µL) was added to the mixture and reacted for 1 h. The mixture was then diluted to 800 μL, and the fluorescence of the solution was detected with a Hitachi R-7000 fluorescence spectrophotometer. The maximum fluorescence signal of FAM was recorded at 518 nm, and the maximum fluorescence signal of Texas Red was recorded at 615 nm.
Optimization of experimental conditions
Key experimental parameters, including the duration of the rolling circle amplification (RCA) process and the concentration of phi29 DNA polymerase, were optimized. The optimal experimental conditions were evaluated by monitoring the fluorescence signal at a constant target concentration. The duration of the RCA process was explored from 40 min to 120 min. The fluorescence signal responses at the maximum emission wavelength of 518 nm were recorded for detecting 80 pg/mL of sPD-L1. To optimize the appropriate DNA polymerase concentration, different concentrations of phi29 DNA polymerase (0.1 U/μL, 0.25 U/μL, 0.5 U/μL, 0.75 U/μL, and 1 U/μL) were analyzed, and the fluorescence responses for 40 pg/mL sPD-L1 at 518 nm were documented.
Theoretical simulations of thermodynamic properties
A thermodynamic analysis tool, Nucleic Acids Package (NUPACK), was used to analyze the thermodynamic properties of the secondary structure of multiple interacting DNA strands. The equilibrium base-pairing probabilities between DNA strands were estimated by empirically sampling sequence-level secondary structures from the Boltzmann distribution. In the plot of pair probabilities, the numerical values on the lower horizontal axis and the left vertical axis denote the sequential order of DNA bases from the 5’ to the 3’ end. The labels on the upper horizontal axis and the right vertical axis correspond to the designations of various DNA strands. The color and size of each dot scale with the equilibrium probability of the corresponding base pair. For the calculation of free energy of complexes formed by the hybridization of two ssDNA strands, the model parameters were set to T = 37 ° C, 1 M NaCl, 1 µM for each ssDNA strand, and a complex size of 2. To calculate the melting profile, the reaction temperature was set from 26 ° C to 84 ° C at a 2 ° C interval. The concentration of each DNA strand was set to 1 µM, and the maximum complex size was set to 2.
Polyacrylamide gel electrophoresis analysis (PAGE)
Native PAGE was used to verify the hybridization of the DNA. The native PAGE gel was prepared using 30% acrylamide/bis-acrylamide (29:1) stock solution. An 8% polyacrylamide gel, which effectively resolves both fragment sizes while ensuring molecular weight markers align accurately across panels, was prepared by mixing 8 mL of 30% acrylamide/bis-acrylamide (29:1) stock solution, 6 mL of 5 × TBE buffer, 210 μL of 10% ammonium persulphate, 20 μL of TEMED, and 15.8 mL of distilled water. The loading samples were prepared by mixing 8 μL of the DNA sample with 2 μL of 5 × GelRed pre-stained loading buffer. Gel electrophoresis was performed on a PowerPac power system from Bio-Rad at 100 V for 80 min in 0.5 × TBE buffer. The DNA bands were then visualized using a Bio-Rad GelDoc XR gel imaging analyzer (Fig. 2f and Supplementary Figs. 1b, 5b, c, 6c, d).
Digestion of DNA solutions and DNA hydrogels by exonuclease I
To assess the enzymatic digestion of DNA hydrogels and DNA solutions, DNA hydrogel and DNA solution samples at varying concentrations (0.3 mM, 0.5 mM, and 0.7 mM) were incubated with 10 µL of 10 × Exonuclease I reaction buffer, 85 µL of DEPC-treated water, and 5 µL of Exonuclease I. Reactions were carried out at 37 °C for 1 h. Fluorescence intensities were recorded before and after enzymatic digestion for both hydrogel and solution-based systems. The signal-to-noise ratio (SNR) was calculated by dividing the post-digestion fluorescence intensity by the pre-digestion fluorescence intensity to evaluate the signal output capacity of the hydrogel and solution. The pre-digestion fluorescence signal represents the baseline background, while the post-digestion signal corresponds to the fluorophore release following enzymatic degradation of DNA. A higher SNR indicates greater signal strength relative to background noise, reflecting enhanced detection sensitivity and stability of the system.
Photobleaching analysis of DNA hydrogels and DNA solutions
For the comparison of the photostability between DNA hydrogels and DNA solutions, 5 μL of each sample was added to a 96-well microplate and subjected to continuous laser irradiation. To ensure these findings are not confounded by measurement bias, we standardized experimental procedures. Sample preparation, handling, and irradiation protocols were strictly controlled across replicates. Fluorescence decay kinetics were monitored using a microplate reader with fixed parameters: 5 μL of each sample (P1 modified FAM and P1 modified Texas Red) was loaded into a 96-well microplate and continuously irradiated. The fluorescence intensity was recorded every 5 s over 4000 cycles (20,000 s total) using specific excitation wavelengths of 492 nm for FAM and 596 nm for Texas Red. The fluorescence reduction rate was quantified as the ratio of reduced intensity to initial intensity. All experiments involving FAM and Texas Red measurements were performed in triplicate.
Comparison of photostability between FAM and Texas Red
To evaluate the intrinsic photostability of FAM and Texas Red, single-stranded DNA (P1) was independently labeled at the 3’ end with either FAM or Texas Red. For each condition, 100 µL of 1 µM fluorophore-labeled P1 was subjected to continuous laser irradiation (5000 s of continuous laser exposure) using a CLARIOstar microplate reader (BMG LABTECH Inc., USA). FAM-labeled samples were excited at 492 nm, and Texas Red-labeled samples at 596 nm. Fluorescence intensity was recorded every 5 s for a total of 1000 cycles.
Evaluation of cross-reactivity between building blocks and non-target reporters
To assess the potential cross-reactivity between building blocks and non-target reporters, fluorescent modifications on all ssDNA in building blocks were removed except for P1. The 3’ end of P1 was labeled with 6-carboxyfluorescein (FAM), generating a modified oligonucleotide designated as P1-2. In parallel, an ssDNA sequence labeled at the 5’ end with the quencher BHQ-1 was synthesized and designated as R2–2, serving as the interfering reporter. Upon hybridization between P1–2 and R2–2, the spatial proximity of FAM and BHQ-1 facilitated fluorescence resonance energy transfer, resulting in quenching of the FAM signal. For the reaction, 10 μL of R2–2 was added to 1 μL of DNA hydrogel and incubated at 37 °C. Fluorescence measurements were recorded every 5 s using an excitation wavelength of 492 nm and an emission wavelength of 518 nm. To compare the cross-reactivity of DNA hydrogels and DNA solutions, identical experimental conditions were employed using aqueous DNA solutions in place of hydrogels.
LOD and RCA amplification calculation
The LOD was determined using the formula:
where S represents standard deviation of the fluorescence intensity of the blank sample, and K refers to slope of the calibration curve1.
The RCA amplification efficiency was defined as the ratio of equivalent non-RCA-amplified sPD-L1 concentrations to RCA-amplified sPD-L1 concentrations:
This ratio quantifies the fold-reduction in sPD-L1 required to achieve comparable signal intensity when RCA is employed.
Comparison of enzymatic reactions of Esp3I in DNA hydrogels and solutions
To compare the enzymatic digestion efficiency of Esp3I in different environments, 8 μg/mL of sPD-L1 was detected using hydrogels and aqueous solutions and then cleaved using Esp3I. Briefly, 50 µL of 8 μg/mL sPD-L1 was added to 100 µL of dsDNA-modified magnetic beads and incubated for 1 h, and separated using a magnet to isolate the supernatant. The supernatant was mixed with 2 μL of either DNA hydrogel or DNA solution, 10 μL of 10 × Esp3I reaction buffer, 85 μL of DEPC-treated water, and 5 μL of Esp3I enzyme. The mixtures were incubated at 37 °C in a 96-well microplate, and fluorescence intensities were recorded every 5 s both before and after enzymatic treatment under an excitation wavelength of 492 nm and an emission wavelength of 518 nm using a CLARIOstar microplate reader (BMG LABTECH Inc., USA).
ELISA detection of sPD-L1
ELISA detection of sPD-L1 was performed according to the manufacturer’s instructions. First, 100 μL of standards with various concentrations and serum samples were added to the microplate wells and incubated at room temperature for 1.5 h. After washing, 100 μL of antibody solution was added and incubated for 30 min. Then, 100 μL of streptavidin solution was added and incubated for another 30 min. After washing, 100 μL of signal enhancer solution was added and incubated for 15 min, followed by incubation with 100 μL of streptavidin solution for 15 min. Finally, 100 μL of 3,3’,5,5’-Tetramethylbenzidine (TMB) substrate was added, incubated in the dark for 10 min, and the reaction was quenched with 100 μL of stop solution. Optical Density (OD) values were measured with a microplate reader at 450 nm, using 630 nm as the reference wavelength (BioTek, USA). Sample concentrations were determined from a standard curve plotted based on OD values and standard concentrations.
Colorimetric detection of LDH
For methodological comparison, serum levels of LDH were quantified using a standard colorimetric assay based on the lactate-pyruvic acid method. In this reaction, LDH catalyzes lactate oxidation coupled with NAD⁺ reduction to NADH, whose absorbance is correlate directly to the activity of LDH. The assay was detected with a commercially available biochemical kit (cat.OSR6128, Beckman Coulter, USA), which quantified based on the absorbance intensity of NADH at 340 nm. All experiments were performed on an automated biochemical analyzer (AU5831, Beckman Coulter, USA) following the manufacturer’s protocols. All reagents (including methylglucosamine, lithium lactate, and β-nicotinamide adenine dinucleotide) and instrumentation were provided and validated by the Department of Clinical Laboratory Shanghai Chest Hospital, ensuring compliance with clinical diagnostic standards.
Quantification of CEA, CYF21-1, IL-6, SCCA, and NSE
The biomarker contents, including CEA, cytokeratin fragment 19 (CYFRA 21-1), interleukin-6 (IL-6), squamous cell carcinoma antigen (SCCA), and neuron-specific enolase (NSE), were sourced from routine clinical assays performed in compliance with standard diagnostic protocols. Serum concentrations of CEA, CYF21-1, and NSE were measured with an immunoassay kit (cat.LP120460, Tellgen, China) on an automated immunoanalyzer (TESMIF4000, Tellgen, China). SCCA was quantified with an immunoassay kit (cat.LP121160, Tellgen, China) on the same platform. IL-6 levels were measured using a cytokine co-detection kit (cat.P110100403, Cellgene, China) on a flow cytometer (BriCyte E6, Mindray, China). These markers were selected based on their well-documented roles as indicators of tumor immune response and therapy efficacy, as previously reported2,3.
Study population and sample collection
In this study, a retrospective analysis was encompassed involving a total of 166 participants at Shanghai Chest Hospital from September 2023 to December 2024, including 22 healthy individuals and 144 lung cancer (LC) patients. All participants were enrolled randomly without prior sex-based selection or stratification. Gender was determined based on biological attribution (also consistent with the self-report). When collecting clinical information, all cases submitted informed consent. All cancer diagnoses were confirmed by pathological results. The tumor, node, and metastasis (TNM) staging criteria were applied to assess LC metastasis. Inclusion criteria for the LC patients were as follows: (1) confirmed diagnosis of NSCLC through pathological assessment; (2) TNM stage III to IV; (3) Age ≥ 18 years; (4) presence of at least one measurable lesion; (5) receipt of immunotherapy alone or in combination with other agents such as chemotherapies.
Exclusion criteria were as follows: (1) presence of known driver gene mutations (e.g., EGFR/ALK/ROS1); (2) undergoing surgical resection after medical therapy initiation; (3) occurrence of infection within 10 days before blood collection; (4) failure to complete essential systemic examinations such as chest CT imaging; (5) severe chronic comorbidities (e.g., cardiovascular, cerebrovascular, or autoimmune diseases) that could influence the safety or efficacy of immunotherapy.
Patients received immunotherapy agents—pembrolizumab, tislelizumab, or sintilimab—administered intravenously at a dose of 200 mg every three weeks according to the guideline of the Chinese Society of Clinical Oncology (CSCO) non-small cell lung cancer. Chemotherapy regimens, determined based on clinical judgment and patient-specific factors, included pemetrexed, paclitaxel or nab-paclitaxel, vinorelbine, docetaxel, gemcitabine, and platinum-based agents. Tumor responses in LC patients were assessed according to the revised Response Evaluation Criteria in Solid Tumors (RECIST). Assessments were performed by a radiologist blinded to patient identifiers and biomarker data to ensure objective evaluation. Treatment responses were classified into four categories: complete response (CR), partial response (PR), stable disease (SD), and progressive disease (PD).
Blood samples from healthy donors and all patients before treatment were collected prior to the initiation of therapy, with serum being used for biomarker detection. After immunotherapy, the blood samples were collected every 8 weeks after immunotherapy, and disease response was assessed according to RECIST guidelines. Blood samples utilized in this study were residual specimens obtained from routine medical examinations conducted during regular clinical visits, and didn’t pose any additional risk or burden to the participants.
All blood samples, collected in coagulation tubes (REF.367955, BD, USA) through venipuncture, were centrifuged at 1600 g for 5 min to isolate the serum, which was then reserved for subsequent analysis. All human samples were collected from the Shanghai Chest Hospital and approved by the institutional ethics committee of the Shanghai Chest Hospital (No. KS22025). Patient treatment protocols were conducted independently of the research study. The study did not interfere with or influence any aspect of the patient treatment or management.
Statistical analysis
Statistical analyses and illustrations were conducted using GraphPad Prism (Version 8) and Origin (Version 2024). A two-tailed Student’s t test with Welch correction was used to assess statistical differences between two groups. For all statistical tests, P < 0.05 were considered significant. Power analysis was conducted to predetermine sample size on Gpower (Version 3.1). Randomization and blinding were used in this study. Sample sizes are specified in the figure captions, where n refers to the number of distinct samples. All values are expressed as the mean ± standard deviation. Receiver operating characteristic (ROC) curve analysis was performed to determine the area under the curve (AUC) using IBM SPSS Statistics software (Version 24.0). The biomarker (CEA, CYF21-1, SCCA, and NSE) datasets used for ROC analysis were obtained from Shanghai Chest Hospital.
Ethical statement
This retrospective cohort study was approved by the institutional ethics committee of the Shanghai Chest Hospital (No. KS22025). All participants in this study provided written consent and approved the use of clinical data and serum samples for analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data generated in the main manuscript and Supplementary Information are provided in the Source Data file. Source data are provided in this paper.
References
Thai, A. A., Solomon, B. J., Sequist, L. V., Gainor, J. F. & Heist, R. S. Lung cancer. Lancet 398, 535–554 (2021).
Qi, J. et al. National and subnational trends in cancer burden in china, 2005-20: An analysis of national mortality surveillance data. Lancet Public Health 8, e943–e955 (2023).
Garon, E. B. et al. Five-year overall survival for patients with advanced non‒small-cell lung cancer treated with pembrolizumab: Results from the phase i keynote-001 study. J. Clin. Oncol. 37, 2518–2527 (2019).
Gettinger, S. et al. Five-year follow-up of nivolumab in previously treated advanced non-small-cell lung cancer: Results from the ca209-003 study. J. Clin. Oncol. 36, 1675–1684 (2018).
Wang, Y. et al. Deep learning analysis of histopathological images predicts immunotherapy prognosis and reveals tumour microenvironment features in non-small cell lung cancer. Br. J. Cancer 131, 1833–1845 (2024).
Ferrara, R. et al. Hyperprogressive disease in patients with advanced non–small cell lung cancer treated with pd-1/pd-l1 inhibitors or with single-agent chemotherapy. JAMA Oncol. 4, 1543–1552 (2018).
Chubachi, S. et al. A case of non-small cell lung cancer with possible “disease flare” on nivolumab treatment. Case Rep. Oncol. Med. 2016, 1075641 (2016).
Champiat, S. et al. Hyperprogressive disease is a new pattern of progression in cancer patients treated by anti-pd-1/pd-l1. Clin. Cancer Res. 23, 1920–1928 (2017).
Zhou, C. et al. A global phase 3 study of serplulimab plus chemotherapy as first-line treatment for advanced squamous non-small-cell lung cancer (astrum-004). Cancer Cell 42, 198–208 (2024).
Ohri, N. et al. Selective personalized radioimmunotherapy for locally advanced non-small-cell lung cancer trial (sprint). J. Clin. Oncol. 42, 562–570 (2024).
Frank, M. S. et al. Circulating tumor DNA monitoring reveals molecular progression before radiologic progression in a real-life cohort of patients with advanced non-small cell lung cancer. Cancer Res. Commun. 2, 1174–1187 (2022).
Fujimoto, D. et al. Pseudoprogression in previously treated patients with non-small cell lung cancer who received nivolumab monotherapy. J. Thorac. Oncol. 14, 468–474 (2019).
Li, S. et al. Sensitive detection of tumor mutations from blood and its application to immunotherapy prognosis. Nat. Commun. 12, 4172 (2021).
Tan, A. C., Cook, S. L. & Khasraw, M. Soluble immune-checkpoint factors: A potential immunotherapy biomarker. J. Clin. Invest. 134, e179352 (2024).
Wang, R. et al. A sustainable approach to universal metabolic cancer diagnosis. Nat. Sustain. 7, 602–615 (2024).
Hayashi, H. et al. Soluble immune checkpoint factors reflect exhaustion of antitumor immunity and response to pd-1 blockade. J. Clin. Invest. 134, e168318 (2024).
Mahoney, K. M. et al. Soluble pd-l1 as an early marker of progressive disease on nivolumab. J. Immunother. Cancer 10, e003527 (2022).
Kazandjian, D., Gong, Y., Keegan, P., Pazdur, R. & Blumenthal, G. M. Prognostic value of the lung immune prognostic index for patients treated for metastatic non-small cell lung cancer. JAMA Oncol. 5, 1481–1485 (2019).
Pan, C. X. et al. Association between baseline lactate dehydrogenase and immune-related adverse events among patients with melanoma varies by tumor stage at immune checkpoint inhibitor initiation. J. Am. Acad. Dermatol. 89, 1264–1268 (2023).
Dercle, L. et al. High serum ldh and liver metastases are the dominant predictors of primary cancer resistance to anti-pd(l)1 immunotherapy. Eur. J. Cancer 177, 80–93 (2022).
Zhand, S. et al. Metal-organic framework-enhanced elisa platform for ultrasensitive detection of pd-l1. ACS Appl. Bio. Mater. 3, 4148–4158 (2020).
Zhang, S. J., Wu, C. & Walt, D. R. A multiplexed digital platform enables detection of attomolar protein levels with minimal cross-reactivity. ACS Nano 18, 29891–29901 (2024).
Wu, C., Dougan, T. J. & Walt, D. R. High-throughput, high-multiplex digital protein detection with attomolar sensitivity. ACS Nano 16, 1025–1035 (2022).
Peng, Y. H. et al. Dynamic matrices with DNA-encoded viscoelasticity for cell and organoid culture. Nat. Nanotechnol. 18, 1463–1473 (2023).
Yao, C., Zhang, R., Tang, J. & Yang, D. Rolling circle amplification (rca)-based DNA hydrogel. Nat. Protoc. 16, 5460–5483 (2021).
Wang, D. et al. Enrichment and sensing tumor cells by embedded immunomodulatory DNA hydrogel to inhibit postoperative tumor recurrence. Nat. Commun. 14, 4511 (2023).
Al Sulaiman, D., Juthani, N. & Doyle, P. S. Quantitative and multiplex detection of extracellular vesicle-derived microrna via rolling circle amplification within encoded hydrogel microparticles. Adv. Healthc. Mater. 11, e2102332 (2022).
Zhang, D. et al. Label-free multiplexed detection based on core-shell photonic barcodes integrated rca. Biosens. Bioelectron. 271, 117037 (2025).
Ludvikova, L. et al. Near-infrared co-illumination of fluorescent proteins reduces photobleaching and phototoxicity. Nat. Biotechnol. 42, 872–876 (2024).
Zhang, Y. et al. General strategy to improve the photon budget of thiol-conjugated cyanine dyes. J. Am. Chem. Soc. 145, 4187–4198 (2023).
Lee, J. et al. Bright and photostable yellow fluorescent proteins for extended imaging. Nat. Commun. 16, 3241 (2025).
Wang, Y., Lang, Y., Yang, Q. & Wu, P. Breaking the photostability and ph limitation of halo-fluoresceins through chitosan conjugation. Adv. Mater. 35, e2210956 (2023).
Al Sulaiman, D., Chang, J. Y. H. & Ladame, S. Subnanomolar detection of oligonucleotides through templated fluorogenic reaction in hydrogels: Controlling diffusion to improve sensitivity. Angew. Chem. Int. Ed. Engl. 56, 5247–5251 (2017).
Huang, Y. et al. Colloidal hydrogel with staged sequestration and release of molecules undergoing competitive binding. ACS Nano 18, 25841–25851 (2024).
Li, X. et al. Remarkable sol-gel transition of pnipam-based nanogels via large steric hindrance of side-chains. Mater. Horiz. 10, 4452–4462 (2023).
Xu, H. et al. A universal strategy for enhancing the circulating mirnas’ detection performance of rolling circle amplification by using a dual-terminal stem-loop padlock. ACS Nano 18, 436–450 (2024).
Yan, H. et al. A one-pot isothermal cas12-based assay for the sensitive detection of micrornas. Nat. Biomed. Eng. 7, 1583–1601 (2023).
Chaibun, T. et al. Rapid electrochemical detection of coronavirus sars-cov-2. Nat. Commun. 12, 802 (2021).
Lu, D., Myers, A. R., George, N. P. & Keck, J. L. Mechanism of exonuclease i stimulation by the single-stranded DNA-binding protein. Nucleic Acids Res. 39, 6536–6545 (2011).
Shi, Y., Hellinga, H. W. & Beese, L. S. Interplay of catalysis, fidelity, threading, and processivity in the exo- and endonucleolytic reactions of human exonuclease i. Proc. Natl. Acad. Sci. USA 114, 6010–6015 (2017).
Orans, J. et al. Structures of human exonuclease 1 DNA complexes suggest a unified mechanism for nuclease family. Cell 145, 212–223 (2011).
Zhang, L. et al. Multifunctional quantum dot DNA hydrogels. Nat. Commun. 8, 381 (2017).
Hu, Y. et al. Reversible modulation of DNA-based hydrogel shapes by internal stress interactions. J. Am. Chem. Soc. 138, 16112–16119 (2016).
Xing, Y. et al. A dual-channel intelligent point-of-care testing system for soluble programmed death-1 and programmed death-ligand 1 detection based on folding paper-based immunosensors. ACS Sens. 7, 584–592 (2022).
Saputra, H. A., Chung, J. H., Sahin, M. A. Z., Park, D.-S. & Shim, Y.-B. Dual-signal output biosensor for the detection of program death-ligand 1 and therapy progress monitoring of cancer. Biosens. Bioelectron. 262, 116565 (2024).
Zhu, L., Liu, X., Yang, J., He, Y. & Li, Y. Application of multiplex microfluidic electrochemical sensors in monitoring hematological tumor biomarkers. Anal. Chem. 92, 11981–11986 (2020).
Stromme, J. H., Rustad, P., Steensland, H., Theodorsen, L. & Urdal, P. Reference intervals for eight enzymes in blood of adult females and males measured in accordance with the international federation of clinical chemistry reference system at 37 degrees c: Part of the nordic reference interval project. Scand. J. Clin. Lab. Invest. 64, 371–384 (2004).
Okuma, Y. et al. Soluble programmed cell death ligand 1 as a novel biomarker for nivolumab therapy for non-small-cell lung cancer. Clin. Lung Cancer 19, 410–417 (2018).
Pasello, G. et al. Sex-related differences in serum biomarker levels predict the activity and efficacy of immune checkpoint inhibitors in advanced melanoma and non-small cell lung cancer patients. J. Transl. Med. 22, 242 (2024).
Ancin, B. et al. The correlation of serum spd-1 and spd-l1 levels with clinical, pathological characteristics and lymph node metastasis in nonsmall cell lung cancer patients. Turk. J. Med. Sci. 52, 1050–1057 (2022).
Sagman, U. et al. The prognostic significance of pretreatment serum lactate dehydrogenase in patients with small-cell lung cancer. J. Clin. Oncol. 9, 954–961 (1991).
Mezquita, L. et al. Association of the lung immune prognostic index with immune checkpoint inhibitor outcomes in patients with advanced non-small cell lung cancer. JAMA Oncol. 4, 351–357 (2018).
Sung, M. et al. Prognostic value of baseline and early treatment response of neutrophil-lymphocyte ratio, c-reactive protein, and lactate dehydrogenase in non-small cell lung cancer patients undergoing immunotherapy. Transl. Lung Cancer Res. 12, 1506–1516 (2023).
Borghaei, H. et al. Five-year outcomes from the randomized, phase iii trials checkmate 017 and 057: Nivolumab versus docetaxel in previously treated non-small-cell lung cancer. J. Clin. Oncol. 39, 723–733 (2021).
Brahmer, J. R. et al. Five-year survival outcomes with nivolumab plus ipilimumab versus chemotherapy as first-line treatment for metastatic non–small-cell lung cancer in checkmate 227. J. Clin. Oncol. 41, 1200–1212 (2023).
Anagnostou, V. et al. Dynamics of tumor and immune responses during immune checkpoint blockade in non-small cell lung cancer. Cancer Res. 79, 1214–1225 (2019).
Masse, M. et al. [18f]FDG-PET/CT atypical response patterns to immunotherapy in non-small cell lung cancer patients: Long term prognosis assessment and clinical management proposal. Eur. J. Nucl. Med. Mol. Imaging 51, 3696–3708 (2024).
Humbert, O. et al. 18FDG PET/CT in the early assessment of non-small cell lung cancer response to immunotherapy: Frequency and clinical significance of atypical evolutive patterns. Eur. J. Nucl. Med. Mol. Imaging 47, 1158–1167 (2020).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (grants 82422042 (L.H.), 82372148 (L.H.), 82272679 (T. C.)) and the Open Research Project of Beijing Key Laboratory from Beijing Youan Hospital, Capital Medical University (grant BJYAHKF2023002 (L.H.))
Author information
Authors and Affiliations
Contributions
J.W., L.H., and T.C. conceived and supervised the project. Y.Z. and F.M. designed and performed the experiments. Z.G., Y.D., and T.L. analyzed the data. H.J. assisted in the clinical studies. Y.Z., F.M., and Z.G. wrote the manuscript. J.W., L.H., and T.C. revised the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.W., L.H., Y.Z., F.M., and Z.G. have filed patents for both the technology and the use of the technology to detect biosamples. These patents are owned and managed by Shanghai Chest Hospital. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Shengxiang Ren, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Y., Meng, F., Gu, Z. et al. A multiplexed assay by self-assembled dual-target responsive DNA hydrogels for efficacy evaluation of immunotherapy. Nat Commun 16, 10132 (2025). https://doi.org/10.1038/s41467-025-65075-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65075-6