Abstract
Precise regulation of type I interferon signaling is crucial for effective immune defense against infectious diseases. However, the molecular mechanisms governing this pathway are not fully understood. Here, we show a function for palmitoylation in enhancing anti-malarial immune responses. Our findings reveal that ZDHHC9 enhances the type I interferon signaling by palmitoylating TBK1 at cysteine 292. Following infection with Plasmodium yoelii N67, the delicate balance between palmitoylation and depalmitoylation of TBK1 is disrupted. Specifically, upregulation of APT2 promotes persistent depalmitoylation of TBK1 and triggers its selective autophagic degradation via K48-linked polyubiquitination at lysine 251/372 by E3 ligase TRIM27. This process acts as a recognition signal for the cargo receptor NDP52, resulting in inhibition of the type I interferon pathway. Notably, inhibition of APT2 using ML349 elevates type I interferon levels and improves survival rates against N67 infection. Here, we show that targeting APT2-mediated TBK1 depalmitoylation is a potential therapeutic strategy for malaria and may also be applicable to other diseases driven by dysregulated type I interferon signaling.
Similar content being viewed by others
Introduction
Malaria is a parasitic disease transmitted by mosquitoes in tropical and subtropical regions, affecting 85 countries and resulting in approximately 233 million clinical infections and 608,000 deaths each year1,2. In response to malaria infection, the immune system mounts complex and nuanced responses, with innate immunity serving as the first line of defense against Plasmodium parasite invasion3. Host innate immune cells utilize pattern recognition receptors such as Toll-like receptors (TLRs), retinoic acid-inducible gene I (RIG-I)-like receptors (RLR), NOD-like receptors (NLRs), and nucleic acid sensors to detect various components of Plasmodium, including genomic DNA (gDNA), RNA, protein-DNA complexes, phospholipids, and hemozoin-complexes4. This recognition triggers innate immune signaling pathways that ultimately activate transcription factors such as nuclear factor κB (NF-κB) and interferon regulatory factors 3/7 (IRF-3/7), leading to the production of pro-inflammatory cytokines and type I interferon (IFN-I)5. Despite these discoveries, the precise regulation of these signaling pathways during malaria is not yet fully understood.
Among the various signaling pathways activated in response to malaria, the IFN-I pathway is particularly crucial in establishing anti-malarial immunity6,7. Extensive research underscores the critical role of IFN-I signaling throughout the malaria infection cycle, including combating liver-stage infection, suppressing blood-stage parasite growth, enhancing anti-malarial immunity, and mitigating severe manifestations like cerebral malaria7,8,9,10,11. Furthermore, boost IFN-I production7 can expand malaria-specific T cell response and enhance host immunity8. TBK1 is a key kinase that phosphorylates IRF3 after being recruited by the interferon gene stimulator (STING), mitochondrial antiviral signaling protein (MAVS), and TRIF signaling pathways, ultimately leading to IFN-I production12,13. Previous studies have indicated that SHIP1 regulates the anti-malarial immune response by promoting the autophagic degradation of IRF314. However, the roles of TBK1 and its regulatory mechanisms in the context of malaria infection remain largely unexplored. Notably, TBK1 is known to undergo various post-translational modifications (PTMs), such as phosphorylation, ubiquitination, acetylation, and SUMOylation15,16,17,18, which are critical for its function. Manipulating these PTMs of TBK1 offers a promising strategy to either inhibit or enhance TBK1 signaling.
One critical PTM involved in IFN-I signaling pathway is S-palmitoylation, which entails the reversible attachment of palmitic acid to cysteine residues via thioester bonds19. This modification is primarily mediated by zinc finger DHHC (Asp-His-Cys) type (zDHHC) family palmitoyl acyltransferases (PATs), and reversed by palmitoyl-protein thioesterase (APTs)19,20,21. Palmitoylation significantly impacts Plasmodium protein localization and interactions, thus affecting development and pathogenesis of malaria22,23,24. Meanwhile, this modification is essential for the functionality and expression of a diverse array of proteins, such as receptors, adapters, enzymes, transcription factors, and effector molecules that play a critical role in host IFN-I signaling pathway. For instance, palmitoylation of signal transducer and activator of transcription 3 (STAT3) by ZDHHC7 occurs in the endoplasmic reticulum and is necessary for STAT3 to effectively recruit and activate downstream signaling components25. ZDHHC9-mediated palmitoylation of cGAS further enhances its activation, thus promoting antiviral and antitumor immune responses26. In the case of STING, its proper function and aggregation within the Golgi apparatus are contingent upon palmitoylation, which is crucial for optimal signaling responses27. Moreover, CPT1A catalyzes the C79 palmitoylation of MAVS by recruiting ZDHHC4 and ZDHHC12, affecting its ability to initiate antitumor and antiviral immunity28,29. Despite extensive research on palmitoylation’s role in antiviral and tumor immunity, little is known about its characteristics in host anti-malarial immunity.
In this study, we elucidate the role of palmitoylation in enhancing anti-malarial immune responses, specifically by targeting the reversible palmitoylation and depalmitoylation of TBK1, orchestrated by ZDHHC9 and APT2. Our findings demonstrate that following Plasmodium yoelii N67 infection, the upregulation of APT2 disrupts the equilibrium of TBK1 palmitoylation, resulting in depalmitoylation of TBK1. This process subsequently leads to selective autophagic degradation of TBK1 through K48-linked polyubiquitination at Lys 251/372, catalyzed by the E3 ligase TRIM27. This modification serves as a recognition signal for the cargo receptor NDP52, ultimately impairing the IFN-I signaling. These results provide insights into the regulatory mechanisms governing TBK1 stability through palmitoylation, highlighting its role in immune response modulation during malaria. Our study suggests therapeutic strategies aimed at modulating host immune defenses against infections.
Results
Palmitoylation inhibits immune evasion by enhancing host IFN-I signaling against malaria
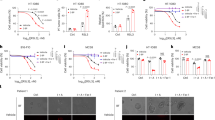
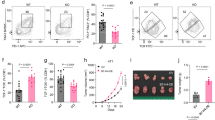
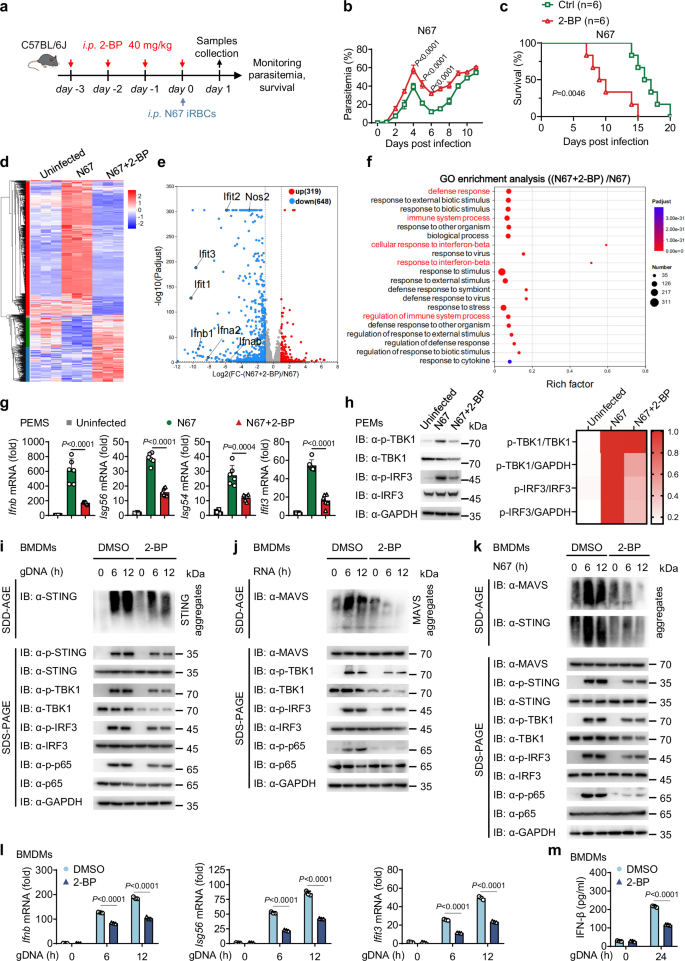
To assess the role of palmitoylation in malaria infection, we employed 2-bromopalmitate (2-BP), an inhibitor of palmitoylation that specifically targets DHHC (Asp-His-His-Cys) protein palmitoyltransferases. C57BL/6 J female mice received intraperitoneal (i.p.) injection of dimethyl sulfoxide (DMSO) or 2-BP (40 mg/kg) for four days30, followed by an injection of Plasmodium. yoelii N67 (N67) infected red blood cells (iRBCs) (1 × 106, i.p.) (Fig. 1a). Mice treated with 2-BP exhibited significantly higher parasitemia and lower survival rate compared to the control group (Fig. 1b, c). RNA-sequencing analysis of peritoneal macrophages (PEMs) revealed that 2-BP treatment down-regulated hundreds of genes, including key interferon-stimulated genes (ISGs) involved in innate immune responses against malarial infection (Fig. 1d, e). Gene Ontology (GO) analysis further indicated an enrichment of genes participating in various immune signaling pathways, notably those related to “defense response” and “response to interferon-beta” after 2-BP treatment (Fig. 1f). In addition, Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR) and immunoblotting demonstrated a marked inhibition of the IFN-I signaling pathway in PEMs from 2-BP-treated N67-infected mice when compared to the N67 infection group (Fig. 1g, h).
a–c C57BL/6 J female mice (n = 6) were injected intraperitoneally (i.p.) with dimethyl sulfoxide (DMSO) or 2-bromopalmitate (2-BP) (40 mg/kg) for 4 days, followed by an injection of Plasmodium. yoelii N67 (N67) infected red blood cells (iRBCs) (1 × 106, i.p.). Parasitemia (b) and survival rate (c) were then evaluated. This was repeated n = 3 independent times with similar results. Figure 1a was adapted from the image “Adult Mouse” by Yunan Chen (SciDraw; CC BY 4.0). d, e Heatmap (d) and volcano plot (e) display the differentially expressed genes in peritoneal macrophages (PEMs) pre-treated with DMSO or 2-BP (100 mM for 24 h), with or without stimulation following freeze-thawed iRBCs. Each group has three biological replicates. (f) Gene ontology (GO) enrichment analysis of the differentially expressed genes shown in panel (e). g, h C57BL/6 J female mice were injected i.p. with DMSO or 2-BP (40 mg/kg) for 4 days, followed by an injection of N67 iRBCs (1 × 106, i.p.) for 24 h. RT-qPCR analysis was performed to measure the mRNA expression levels of Ifnb, Isg56, Isg54 and Ifit3, data shown are from one representative experiment (n = 6 mice per group), consistent results were observed across three biological replicates (g). Immunoblotting was conducted to assess the levels of total IRF3 and TBK1, as well as phosphorylated IRF3 and TBK1, on day 1 after N67 infection in PEMs (h). Representative heatmaps display the results of three independent replicate experiments. The grayscale values from each replicate were quantified, and the average value across the three replicates was calculated. This average was then normalized using the group with the highest grayscale value as the baseline (set to 1), to generate the final heatmap. i–m Bone-marrow derived macrophages (BMDMs) were pre-treated with DMSO or 2-BP (100 mM for 24 h) and then stimulated with gDNA, RNA or freeze-thawed iRBCs at indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (i–k) and RT-qPCR analysis (n = 3 biological replicates) (l). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (m). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (b, g, l, m), and two-sided log-rank test (c). In (e) differential expression analysis between N67 + 2-BP and N67 was performed using DESeq2 (Version 1.42.0). The Wald test (two-tailed) was used, and the resulting p-values were adjusted for multiple testing using the Benjamini-Hochberg method. Genes with an adjusted |log2FC | ≥ 1 and padjust < 0.05 were considered significantly differentially expressed. In (f) Gene Ontology (GO) biological process enrichment analysis was performed on the list of genes that were significantly down-regulated in the N67 + 2-BP group compared to the N67 group (|log2FC | ≥ 1 and padjust < 0.05). The analysis was carried out using the goatools (Version 1.4.4) with Fisher’s exact test (two-tailed). All genes detected in the RNA-seq experiment were used as the background universe. The resulting p-values were adjusted for multiple testing using the Benjamini-Hochberg method. Source data are provided as a Source Data file.
To further validate the role of 2-BP, we stimulated RAW 264.7 cells and bone-marrow-derived macrophages (BMDMs) with N67 genomic DNA (gDNA), RNA or crude antigens from N67 iRBCs subjected to repeated freezing-thaw cycles. 2-BP treatment resulted in a significant inhibition of the IFN-I signaling pathway in both cell types, which is characterized by reduced oligomerization of MAVS and STING, along with decreased phosphorylation of STING, TBK1 and IRF3 (Fig. 1i–k and Supplementary Fig. 1a–c). In addition, reduced TBK1-mediated phosphorylation of p65 in the downstream NF-κB pathway was observed (Fig. 1i–k), alongside lowered expression of Ifnb and ISGs (Fig. 1l and Supplementary Fig. 1d–h) and diminished production of IFN-β (Fig. 1m and Supplementary Fig. 1i–m), indicating that 2-BP inhibits the IFN-I response against N67 infection. Therefore, these findings suggest that palmitoylation enhances the IFN-I signaling and anti-malarial immune response.
ZDHHC9 mediates TBK1 palmitoylation during malarial infection
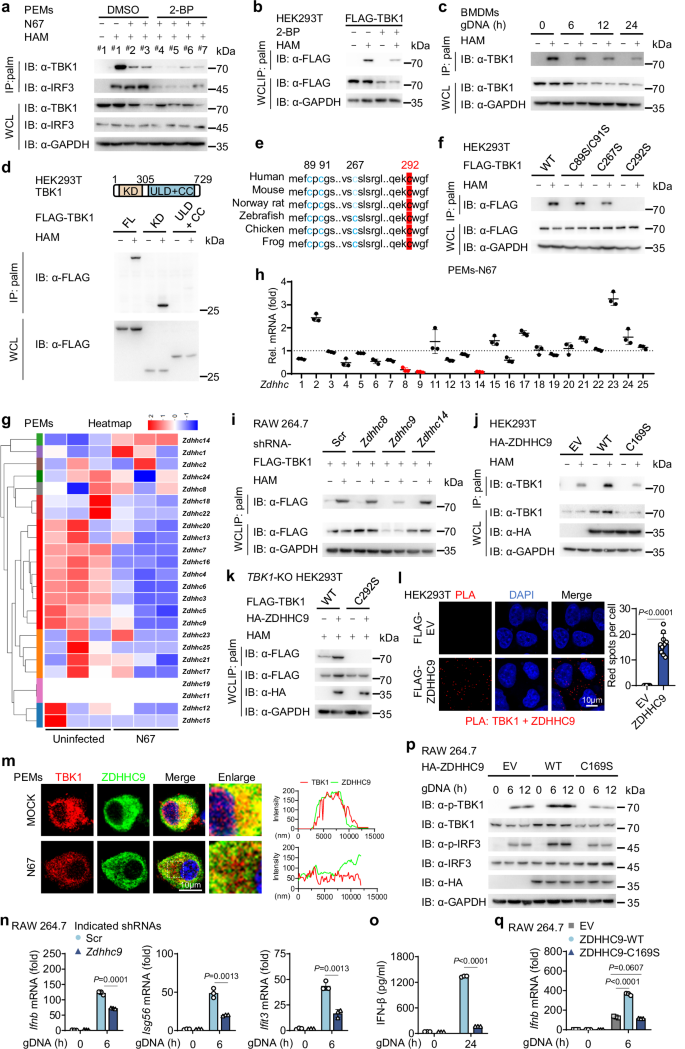
In order to identify the molecular mechanism through which 2-BP inhibits the IFN-I pathway, we conducted an acyl-biotin exchange (ABE) assay to investigate the palmitoylation status of key molecules involved in the IFN-I signaling pathway undergo palmitoylation. The persistence of the signal after hydroxylamine (HAM) treatment indicated that these proteins are palmitoylated via thioester bond formation31. Our results revealed that both TBK1 and IRF3 are susceptible to palmitoylation, while only TBK1 exhibited a decrease in palmitoylation levels following N67 infection (Fig. 2a). Treatment with 2-BP during N67 infection resulted in nearly undetectable palmitoylation levels of TBK1 (Fig. 2a). Exogenous experiments reaffirmed TBK1’s susceptibility to palmitoylation, with treatment with 2-BP substantially reducing its palmitoylation levels (Fig. 2b). Furthermore, stimulation of BMDMs with gDNA, RNA or freeze-thawed iRBCs revealed a time-dependent reduction in TBK1 palmitoylation (Fig. 2c and Supplementary Fig. 2a, b). These results indicate that TBK1 undergoes palmitoylation, which declines during N67 infection.
a Cell lysates from experiments in Fig. 1 (h) were subjected to acyl-biotin exchange (ABE) assays and immunoblot analysis to assess TBK1 palmitoylation. This was repeated n = 3 independent times with similar results. b Human embryonic kidney (HEK) 293 T cells were transfected with Flag-TBK1 for 24 h, followed by the treatment of DMSO (vehicle) or 2-BP (100 mM for 12 h). TBK1 palmitoylation and protein levels were assessed by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. c BMDMs were stimulated with gDNA at the indicated time points, and TBK1 palmitoylation levels were measured using ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. d HEK293T cells were transfected with wild-type (WT) or deletion mutants of Flag-TBK1 for 24 h, and TBK1 palmitoylation levels were measured using ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. e Comparative analysis of potential palmitoylation sites within TBK1 sequences across different species. f HEK293T cells were transfected with WT FLAG-TBK1 or TBK1 palmitoylation-deficient mutants for 24 h. TBK1 palmitoylation levels were detected by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. g Heatmap analysis of expression changes in the ZDHHC gene family in PEMs, with or without stimulation following freeze-thawed iRBCs. Each group has three biological replicates. h The relative mRNA levels of Zdhhc genes in PEMs were determined by RT-qPCR following N67 infection, with mRNA levels was normalized to Gapdh (n = 3 biological replicates). i RAW 264.7 cells were transfected with Scr shRNA or Zdhhc shRNAs for 24 h, followed by transfection with FLAG-TBK1. TBK1 palmitoylation and protein levels were measured using the ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. j, k HEK293T cells were transfected with either WT HA-ZDHHC9 or the enzymatically inactive mutant ZDHHC9 (C169S) for 24 h (j). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (k). Cell lysates were harvested for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. l Proximity ligation assay (PLA) was performed on HEK293T cells overexpressing FLAG-ZDHHC9 to examine the colocalization of TBK1 and FLAG. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are presented next to the image. This was repeated n = 3 independent times with similar results. m PEMs were stimulated with freeze-thawed iRBCs for 12 h. The colocalization between TBK1 (red) and ZDHHC9 (green) was examined by confocal microscopy. The nucleus was stained with DAPI (blue). Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. n, o RAW 264.7 cells were transfected with Scr shRNA or Zdhhc9 shRNA for 24 h, followed by stimulation with gDNA at the indicated time points. Cell lysates were used for RT-qPCR analysis (n = 3 biological replicates) (n). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (o). p, q RAW 264.7 cells were transfected with HA-EV, WT HA-ZDHHC9, or the enzymatically inactive mutant of ZDHHC9 (C169S) for 24 h, and then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblotting. This was repeated n = 3 independent times with similar results (p) and RT-qPCR analysis (n = 3 biological replicates) (q). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (l, n, o, q). Source data are provided as a Source Data file.
TBK1 consists of three main domains: a kinase domain (KD), an ubiquitin-like domain (ULD), and a coiled-coil (CC) domain16. To pinpoint the region of TBK1 that undergoes palmitoylation, HEK293T cells were transiently transfected with TBK1 full-length (FL), TBK1-KD, and TBK1-ULD-CC constructs. Immunoblotting confirmed that both TBK1-FL and TBK1-KD were palmitoylated, while TBK1-ULD-CC was not (Fig. 2d). Notably, four conserved cysteine residues in the KD domain-C89, C91, C267, and C292-were identified across species (Fig. 2e). Only mutating C292 led to a significant reduction in TBK1 palmitoylation levels (Fig. 2f), suggesting that C292 is critical for TBK1’s palmitoylation.
Palmitoylation is catalyzed by DHHC-palmitoyltransferases32. The decrease in TBK1 palmitoylation following N67 infection prompted us to investigate the expression levels of palmitoyltransferases. Heat map analysis in RNA-seq indicated that multiple enzymes of the ZDHHC family in PEMs were downregulated after N67 infection (Fig. 2g). Further verification by RT-qPCR revealed significant down-regulation of Zdhhc8, Zdhhc9, and Zdhhc14 in PEMs after N67 infection (Fig. 2h). Using short hairpin-mediated RNA (shRNA) (Supplementary Fig. 2c) to knockdown Zdhhc9 but not Zdhhc8 or Zdhhc14 markedly decreased TBK1 palmitoylation (Fig. 2i). To further validate the relationship between members of the ZDHHC family and TBK1, exogenous co-immunoprecipitation (co-IP) experiments revealed that, in addition to ZDHHC9, several other members, pacifically ZDHHC13, ZDHHC16, ZDHHC21, and ZDHHC23, demonstrated significant interactions with TBK1. Building on these findings, we simultaneously assessed the impact of these interactions on the palmitoylation level of TBK1. Results indicated that only ZDHHC9 was capable of enhancing the palmitoylation level of TBK1 (Supplementary Fig. 2d, e), indicating that ZDHHC9 is a major palmitoyltransferase for TBK1 palmitoylation. To further explore the role of ZDHHC9 in TBK1 palmitoylation, we generated an enzymatically inactive mutant of ZDHHC9 (C169S)33 and confirmed that enzymatic activity of ZDHHC9 was crucial for TBK1 palmitoylation (Fig. 2j). Moreover, ZDHHC9 was unable to catalyze the palmitoylation of the TBK1 C292S mutant (Fig. 2k). Notably, both proximity ligation assays (PLA) and confocal microscopy consistently demonstrated a strong interaction between flag-ZDHHC9 and endogenous TBK1 (using flag-ZDHHC3 and endogenous TBK1 as controls) (Supplementary Fig. 2f). Furthermore, confocal microscopy demonstrated that stimulation with freeze-thawed iRBCs significantly decreased the co-localization of endogenous ZDHHC9 and endogenous TBK1 (Fig. 2l, m). Therefore, these results suggest that ZDHHC9 interacts with TBK1 and serves as the primary palmitoyltransferase for TBK1, maintaining its palmitoylation levels in the resting state. During N67 infection, the down-regulation of ZDHHC9 leads to a decrease in the interaction between ZDHHC9 and TBK1, which consequently results in a simultaneous reduction in the palmitoylation levels of TBK1.
Next, we investigated whether ZDHHC9 mediated TBK1 palmitoylation affects its function. After silencing ZDHHC9, we observed a reduction in the activity of TBK1-induced IFN-β and ISRE promoter (Supplementary Fig. 2g). Similar results were observed in RAW 264.7 cells when silencing ZDHHC9 expression with shRNA led to decrease activation of IFN-I signaling after stimulation with gDNA, RNA, or freeze-thaw iRBCs (Fig. 2n, o and Supplementary Fig. 2h–m). In contrast, the expression of wild-type (WT) ZDHHC9 markedly augmented the IFN-I signaling pathway, whereas expressing the enzymatically inactive mutant (C169S) ZDHHC9 did not produce this effect (Fig. 2p, q). Thus, these results suggest that ZDHHC9 plays a critical role in the palmitoylation of TBK1, thus activating the IFN-I signaling pathway.
APT2 mediates the depalmitoylation of TBK1 during Plasmodium infection
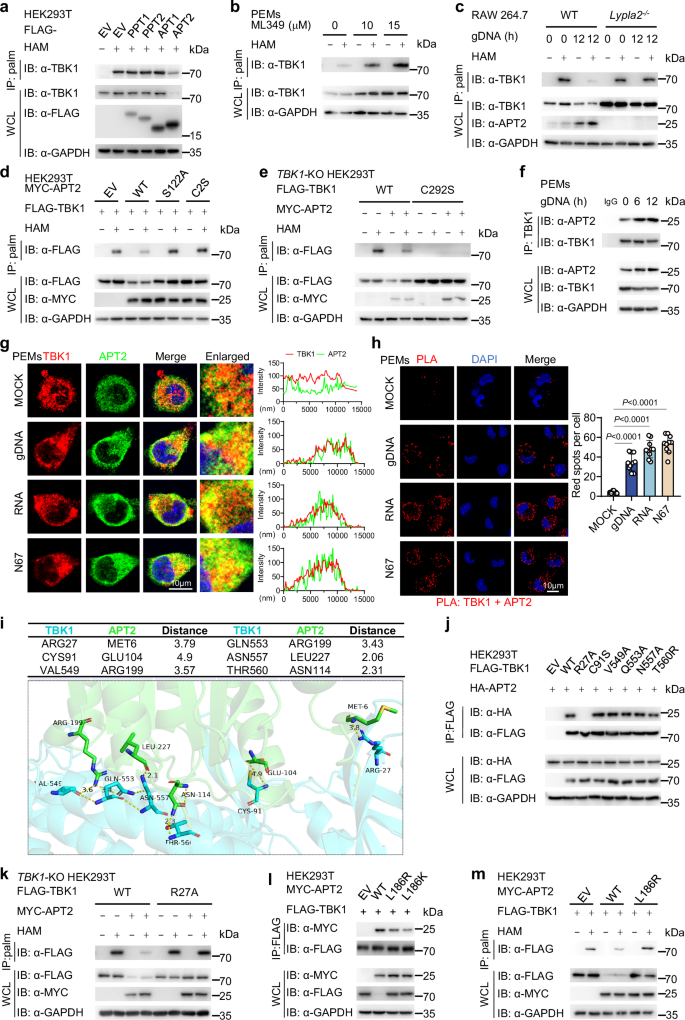
S-palmitoylation is a dynamic process regulated by two opposing classes of enzymes: palmitoyl acyltransferases (PATs) which add palmitate to proteins and acyl protein thioesterases, which remove it. Previous studies have identified several mammalian acyl-protein thioesterases, including APT1 and APT2, as well as palmitoylprotein thioesterases 1 and 2 (PPT1 and PPT2)34. To determine which enzyme is responsible for the depalmitoylation of TBK1 during Plasmodium infection, we expressed four known acyl protein thioesterases and measured the palmitoylation level of endogenous TBK1. Our results demonstrated that only the expression of APT2 led to a significant reduction in TBK1 palmitoylation, while the other APTs showed no such effect (Fig. 3a). Consistent with this finding, pharmacological inhibition of APT2 using ML349 and silencing of Lypla2 with small interfering RNA (siRNA) resulted in increased palmitoylation of TBK1 (Fig. 3b and Supplementary Fig. 3a, b).
a HEK293T cells were transfected with FLAG-EV, -PPT1, -PPT2, -APT1, or -APT2 for 24 h. TBK1 palmitoylation levels were detected by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. b PEMs were treated with ML349 at the indicated concentrations for 12 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. c WT or Lypla2-knockout (KO) RAW 264.7 cells were stimulated with gDNA for 12 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. d HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, MYC-WT APT2, or its catalytically inactivated variants (S122A and C2S) for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. e TBK1-KO HEK293T cells were reconstituted with plasmids encoding FLAG-WT TBK1 or FLAG-TBK1 C292S mutant and MYC-APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. f PEMs were stimulated with gDNA at the indicated time points. Cell lysates were collected for IP and immunoblot analysis. This was repeated n = 3 independent times with similar results. g PEMs were stimulated with gDNA, RNA, or repeated freeze-thaw cycles of N67 iRBCs for 12 h. The colocalization between TBK1 (red) and APT2 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. h PEMs were stimulated with gDNA, RNA, or repeated freeze-thaw cycles of N67 iRBCs for 12 h. The colocalization between TBK1 and APT2 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are displayed next to the image. This was repeated n = 3 independent times with similar results. i Protein structure and molecular docking maps of TBK1 and APT2. j HEK293T cells were transfected with various combinations of plasmid encoding FLAG-WT TBK1, -R27A mutant, -C91S mutant, -V549A mutant, -Q553A mutant, -N557A mutant, or -T560R mutant of TBK1 and HA-APT2 for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. k TBK1-KO HEK293T cells were reconstituted with plasmids encoding FLAG-WT TBK1 or FLAG-TBK1 R27A mutant and MYC-APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. l HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, -WT APT2, -L186R APT2, or -L186K APT2 for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. m HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, -WT APT2, or -L186R APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. Data are presented as the mean ± SD. P-values were determined by unpaired two-tailed Student’s t tests (h). Source data are provided as a Source Data file.
To further elucidate the in vivo function of APT2, we employed CRISPR/Cas9 technology to create Lypla2-knockout (KO) RAW 264.7 cells. Upon stimulation with gDNA, RNA, or freeze-thawed iRBCs, we observed a significant decrease in TBK1 palmitoylation levels in WT cells but not in Lypla2-KO cells (Fig. 3c and Supplementary Fig. 3c,d). In addition, catalytically inactive mutants of APT2 (S122A and C2S)35 were engineered and it was observed that the expression of WT APT2 significantly decreased TBK1 palmitoylation, while the inactive mutants did not (Fig. 3d). Furthermore, APT2 was ineffective in reducing the palmitoylation of the TBK1 C292S mutant (Fig. 3e). Together, these findings indicate that APT2 is responsible for the depalmitoylation of TBK1.
Furthermore, exogenous Co-IP experiments demonstrated that APT2 can interact with TBK1 when overexpressed (Supplementary Fig. 3e), and continued to bind to a TBK1 mutant with a phosphorylation site alteration (S172A), indicating that APT2 acts on the non-phosphorylated form of TBK1 (Supplementary Fig. 3e). In addition, endogenous Co-IP assay indicated that this physiological interaction is enhanced in PEMs under gDNA or RNA stimulation conditions (Fig. 3f and Supplementary Fig. 3f). Confocal microscopy and PLA analyses confirmed increased colocalization of APT2 with TBK1 in PEMs and BMDMs following stimulation with gDNA, RNA, or freeze-thawed iRBCs (Fig. 3g, h and Supplementary Fig. 3g). Additionally, molecular docking studies identified several potential binding sites on TBK1, including R27, C91, V549, Q553, N557, and T560, as likely candidates for its interaction with APT2 (Fig. 3i). To explore these interactions, we generate mutants of TBK1 and performed Co-IP assays, which revealed that the TBK1 R27 mutant did not bind to APT2 (Fig. 3j). Moreover, APT2 was unable to downregulate palmitoylation in the TBK1 R27A mutant (Fig. 3k). Previous studies indicated that the L186 residue on APT2 as crucial for both substrate binding and catalytic activity35. Our experiments further demonstrated that mutation at this site led to a reduced ability to bind to TBK1 and resulted in the loss of its depalmitoylation function (Fig. 3l, m). Therefore, APT2 mediates the depalmitoylation of TBK1 by binding to the R27 residue on TBK1.
APT2 inhibits TBK1-mediated IFN-I responses
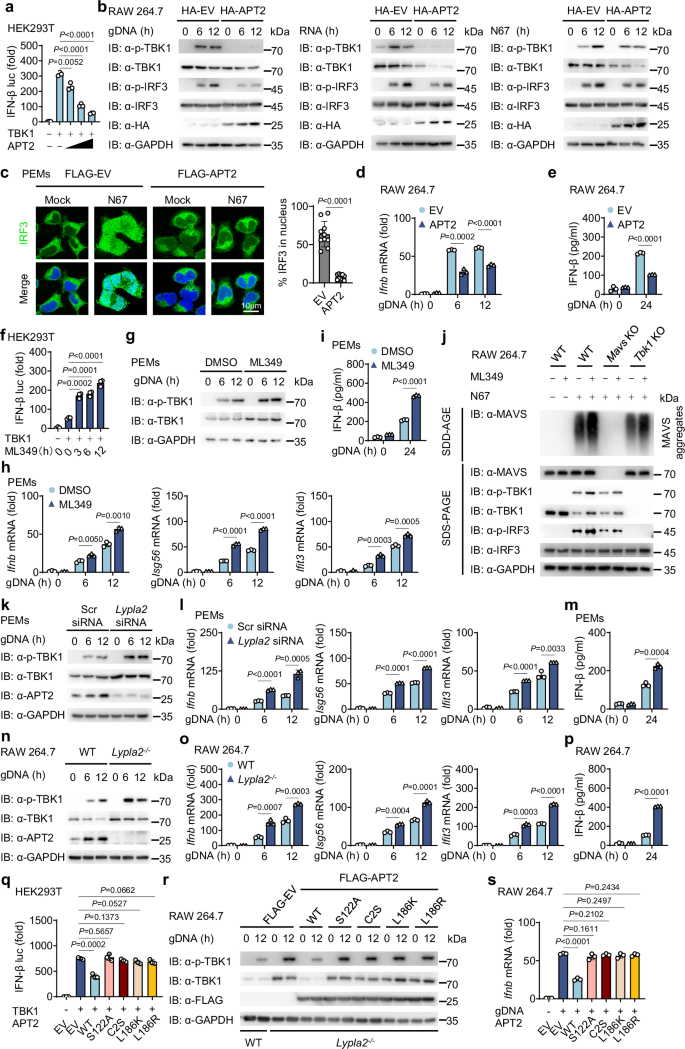
We next investigated whether depalmitoylation of TBK1 by APT2 affects its role in activating IFN-I responses. Overexpressing APT2 in HEK293T cells resulted in a marked inhibition of TBK1-mediated activation of IFN-β reporter gene (Fig. 4a). In parallel, APT2 overexpression significantly suppressed the phosphorylation of TBK1 and IRF3 and inhibited the nuclear translocation of IRF3, as well as the transcription of IFN-responsive genes, including Ifnb, Isg56, and Ifit3, along with the release of IFN-β in RAW 264.7 cells stimulated with gDNA, RNA or freeze-thawed iRBCs (Fig. 4b–e and Supplementary Fig. 4a–d).
a HEK293T cells were transfected with a luciferase reporter for IFN-β-luc, FLAG-TBK1 and increasing doses of MYC-APT2 for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). b RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with gDNA, RNA or freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. c PEMs were transfected with FLAG-EV or FLAG-APT2 for 24 h, then stimulated with freeze-thawed iRBCs for 12 h. IRF3 (green) and nucleus (DAPI, blue) was examined by confocal microscopy. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are presented next to the image. This was repeated n = 3 independent times with similar results. d, e RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates) (d). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (e). f HEK293T cells were transfected with a luciferase reporter for IFN-β-luc and FLAG-TBK1 for 24 h, followed by treatment with ML349 at different time points. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). g–i PEMs were treated with ML349 (10 μM) for 12 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (g) and RT-qPCR analysis (n = 3 biological replicates) (h). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (i). j WT, Mavs-KO, or Tbk1-KO RAW 264.7 cells were treated with or without ML349 (10 μM) for 12 h and subsequently stimulated with freeze-thawed iRBCs for 6 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. k–m PEMs were transfected with Scr-siRNA or Lypla2-siRNA for 36 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (k) and RT-qPCR analysis (n = 3 biological replicates) (l). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (m). n–p WT or Lypla2-KO RAW 264.7 cells were stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (n) and RT-qPCR analysis (n = 3 biological replicates) (o). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (p). q HEK293T cells were transfected with a luciferase reporter for IFN-β luc, FLAG-TBK1 and FLAG- WT APT2 or APT2 mutants for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). r, s Lypla2-KO RAW 264.7 cells were reconstituted with FLAG- WT APT2 or APT2 mutants for 36 h, then stimulated with gDNA for 12 h. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (r) and RT-qPCR analysis (n = 3 biological replicates) (s). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, c–f, h, i, l, m, o–q, s). Source data are provided as a Source Data file.
To further elucidate APT2’s role in the IFN-I signaling pathway, we used the inhibitor ML349 to suppress APT2 in HEK293T cells. This manipulation significantly boosted the activity of the IFN-β promoter following the introduction of exogenous TBK1 (Fig. 4f and Supplementary Fig. 4e). In addition, ML349 treatment resulted in increased TBK1 phosphorylation and a strengthened IFN-I signaling response to N67 stimulation (Fig. 4g–i and Supplementary Fig. 4f–i). While ML349 is known to enhance IFN-I signaling pathway via increasing MAVS oligomerization36, the activation of IFN-I by N67 stimulation was markedly diminished in Mavs-KO RAW 264.7 cells compared to WT cells. Nevertheless, ML349 still enhanced IFN-I activation in Mavs-KO RAW 264.7 cells. In contrast, in Tbk1-KO RAW 264.7 cells, ML349 failed to enhance IFN-I activation, resulting in undetectable levels of both basal and stimulated IFN-I activation (Fig. 4j and Supplementary Fig. 4j). In addition, inhibition of APT2, either through knockdown or knockout approaches, led to an enhanced response of IFN-I signals to N67 stimulation (Fig. 4k–p and Supplementary Fig. 4k–r). Notably, WT APT2 overexpression inhibited TBK1-mediated activation of IFN-I, whereas APT2 mutants lacked this inhibitory effect (Fig. 4q), which is consistent with its palmitoyl acyltransferases activity. Next, in Lypla2-KO RAW 264.7 cells, reconstitution of WT APT2 significantly reduced TBK1 phosphorylation and the transcription levels of Ifnb after gDNA stimulation, but this effect was absent with APT2 mutants reconstitution (Fig. 4r, s). Moreover, while APT2 effectively suppressed the activation of WT TBK1-mediated IFN-I, it did not impact the activation of IFN-I mediated by the TBK1 C292S mutant (Supplementary Fig. 4s). Collectively, these results suggest that APT2 promotes the depalmitoylation of TBK1, thereby attenuating TBK1-mediated IFN-I responses.
APT2-mediated depalmitoylation promotes TBK1 autophagic degradation
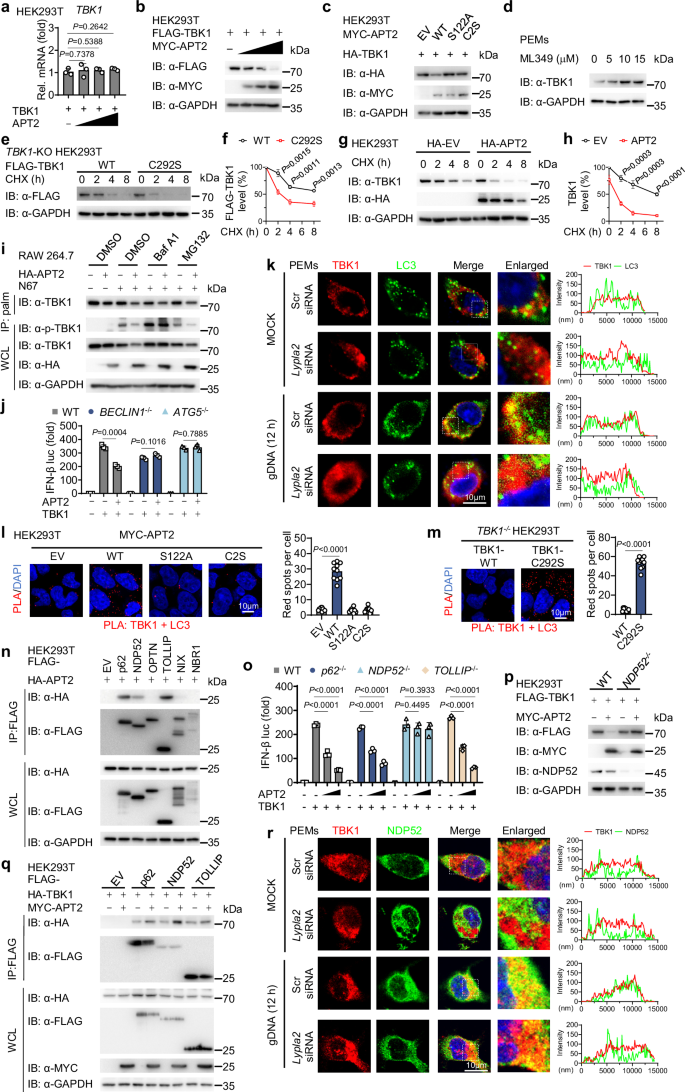
To elucidate the inhibitory effects of APT2 on IFN-I signaling through TBK1, we considered the role of palmitoylation in protein stability, as previous studies have indicated that this modification is crucial for the stability of proteins such as NLRP337 and MAVS28. We found that increased levels of APT2 led to a significant reduction in TBK1 protein levels without affecting TBK1 mRNA expression (Fig. 5a, b). Notably, only WT APT2, and not the catalytically inactive variant (S122A and C2S), induced degradation of TBK1 (Fig. 5c). Consistently, treatment with APT2 inhibitor ML349 or silencing Lypla2 enhanced TBK1 stability (Fig. 5d and Supplementary Fig. 5a, b). The cycloheximide (CHX) chase assay also revealed that the TBK1 depalmitoylated C292S mutation was more prone to degradation compared to WT TBK1 (Fig. 5e, f). Moreover, APT2 overexpression accelerated endogenous TBK1 degradation, and ML349 treatment stabilized TBK1 protein (Fig. 5g, h and Supplementary Fig. 5c). These findings suggest that APT2-mediated depalmitoylation promotes TBK1 degradation.
a, b HEK293T cells were transfected with FLAG-TBK1 and increasing doses of MYC-APT2 for 24 h. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates) (a) and immunoblot analysis, this was repeated n = 3 independent times with similar results (b). (c) HEK293T cells were transfected with HA-TBK1 and WT MYC-APT2 or its catalytically inactivated variants (S122A and C2S) for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. d PEMs were treated with ML349 at indicated concentrations for 12 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. e, f TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h, then treated with cycloheximide (CHX, 100 mg/mL) at the indicated time points. Cell lysates were collected for immunoblot analysis (e). Quantitative analyses (n = 3 biological replicates) (f) of FLAG-TBK1 signals. g, h HEK293T cells were transfected with HA-EV or HA-APT2 for 24 h, then treated with CHX (100 mg/mL) at the indicated time points. Cell lysates were collected for immunoblot analysis (g). Quantitative analyses (n = 3 biological replicates) (h) of TBK1 signals. i RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with freeze-thawed iRBCs for 12 h and then DMSO (control), bafilomycin A1 (Baf A1) (0.2 μM), or MG132 (10 μM) treatment for 6 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. j WT, BECLIN1-KO, or ATG5-KO HEK293T cells were transfected with the indicated plasmids. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). k PEMs were transfected with Scr-siRNA or Lypla2 siRNA for 36 h, then stimulated with gDNA for 12 h. The colocalization between TBK1 (red) and LC3 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. l, m HEK293T cells were transfected with MYC-EV, WT MYC-APT2 or its catalytically inactivated variants (S122A and C2S) for 24 h (l). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (m). The colocalization between TBK1 and LC3 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) is next to the image. This was repeated n = 3 independent times with similar results. n HEK293T cells were transfected with the indicated FLAG-tagged cargo receptors along with HA-APT2 for 24 h, then treated with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. o WT, p62-KO, NDP52-KO or TOLLIP-KO HEK293T cells were transfected with the indicated plasmids. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). p WT or NDP52-KO HEK293T cells were transfected with FLAG-TBK1 in the presence of MYC-APT2 or not for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. q HEK293T cells were transfected with HA-TBK1 and indicated FLAG-tagged cargo receptors in the presence of MYC-APT2 or not for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. r PEMs were transfected with Scr-siRNA or Lypla2 siRNA for 36 h, then stimulated with gDNA for 12 h. The colocalization between TBK1 (red) and NDP52 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, f, h, j, l, m, o). Source data are provided as a Source Data file.
TBK1 degradation primarily occurs through the autophagy-lysosomal and proteasome pathways16,38,39. Our immunoblotting results indicated that the degradation of TBK1 mediated by 2-BP or APT2 was inhibited by autophagy-lysosome inhibitors such as 3-methyladenine (3-MA) and bafilomycin A1 (Baf A1), but not by the proteasome inhibitor MG132 (Fig. 5i and Supplementary Fig. 5d), which suggests that 2-BP and APT2-mediated TBK1 degradation occurs predominantly through the autophagy-lysosomal pathway. Meanwhile, our findings revealed that following treatment with Baf A1, APT2 or 2-BP no longer lead to a decrease in p-TBK1 levels (Fig. 5i and Supplementary Fig. 5d). Moreover, upon application of CHX to inhibit new protein synthesis, we observed that the palmitoylation-deficient mutant TBK1-C292S was more prone to degradation compared to WT TBK1, and its p-TBK1 levels also exhibited a concurrent decline. Importantly, when using Baf A1 to stabilize TBK1 protein levels, the reduction in p-TBK1 was effectively blocked (Supplementary Fig. 5e). These results collectively indicate that the degradation of TBK1 protein is a direct cause for the decrease in its phosphorylated form (p-TBK1). To further verify the role specificity of the proteasome and autophagosome in APT2-mediated TBK1 degradation, we used Lactacystin, a more selective proteasome inhibitor than MG132. Western blot analysis showed that treatment with Lactacystin also failed to effectively block APT2-induced TBK1 degradation. However, when we added Baf A1 on the basis of the presence of Lactacystin, the degradation of TBK1 by APT2 was significantly blocked (Supplementary Fig. 5f). These results indicate that simultaneously inhibiting the autophagic lysosomal pathway can effectively prevent APT2-mediated TBK1 degradation, suggesting that the autophagic pathway is crucial for TBK1 degradation mediated APT2. To further assess the degradation of TBK1 by APT2 via autophagy, we examined cells deficient in autophagy-related genes. In BECLIN1- or ATG5-KO HEK293T cells, APT2 lost its ability to impair IFN-I signaling activation (Fig. 5j). Confocal microscopy revealed an interaction between TBK1 and the autophagy marker LC340 following gDNA stimulation, and APT2 knockdown reduced the colocalization between TBK1 and LC3 (Fig. 5k). Subsequent immunoblotting and PLA demonstrated that overexpression of WT APT2 significantly enhanced the co-localization of TBK1 and LC3, whereas the catalytically inactive variants (S122A and C2S) did not (Fig. 5l and Supplementary Fig. 5g). Furthermore, the TBK1 C292S mutant exhibited increased co-localization with LC3 compared to WT TBK1 (Fig. 5m and Supplementary Fig. 5h). Taken together, these findings indicate that APT2-mediated depalmitoylation leads to TBK1 degradation via the autophagy pathway.
Emerging evidence underscores the essential role of cargo receptors in the selective autophagy degradation of substrates16,41,42,43. To investigate this further, we aimed to identify cargo receptors that mediate APT2-induced autophagic degradation of TBK1. Co-IP experiments revealed that APT2 interacts with several cargo receptors, p62, NDP52, and TOLLIP (Fig. 5n). Using the IFN-β luciferase reporter assay and immunoblotting, we found that APT2 failed to inhibit IFN-I signaling or promote TBK1 degradation in NDP52-KO cells, but not in p62- or TOLLIP-KO cells (Fig. 5o, p and Supplementary Fig. 5i, j). In addition, APT2 enhanced the interaction between TBK1 and NDP52, with no significant effects on TBK1 interactions with p62 or TOLLIP (Fig. 5q). Confocal microscopy confirmed the interaction between TBK1 and NDP52 in response to gDNA, and silencing APT2 expression reduced their co-localization (Fig. 5r). Moreover, overexpression of WT APT2 significantly increased the TBK1-NDP52 interaction, while catalytically inactivated variants (S122A and C2S) did not (Supplementary Fig. 5g). Interestingly, the interaction between TBK1 C292S mutant and NDP52 was stronger than that of WT TBK1 (Supplementary Fig. 5h), indicates that depalmitoylated TBK1 is more susceptible to cargo-mediated autophagic degradation. These findings suggest that APT2 facilitates the interaction between TBK1 and NDP52, thereby promoting the selective autophagy degradation of TBK1.
APT2 modulates K48-linked polyubiquitination of TBK1 through TRIM27
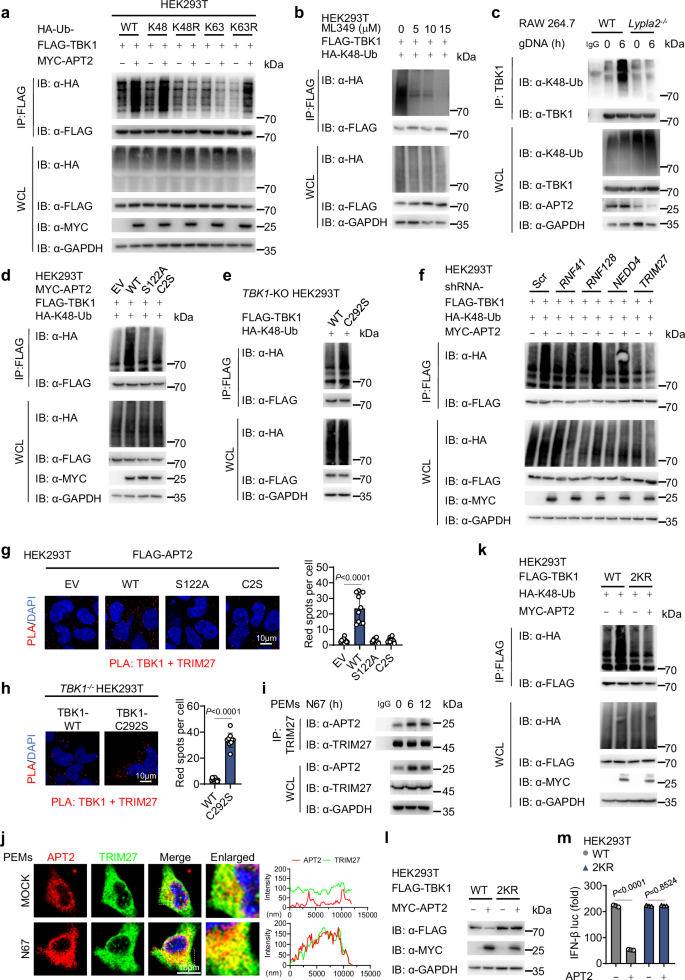
A key step in the selective autophagy degradation process is cargo proteins recognize ubiquitinated substrates, which signals for autophagy-lysosomal degradation. To elucidate APT2’s role in this process, we investigated its effect on TBK1 polyubiquitination and the results revealed that APT2 specifically enhances K48-linked polyubiquitination of TBK1 without affecting other ubiquitin linkages (Fig. 6a and Supplementary Fig. 6a). We also investigated the impact of ZDHHC9 on the ubiquitination of TBK1 and discovered that ZDHHC9 specifically reduced K48-linked polyubiquitination of TBK1 without affecting K63 ubiquitin linkage (Supplementary Fig. 6b). Furthermore, in an in vivo infection model using N67, we observed a notable increase in K48-linked polyubiquitination of TBK1 in the spleen tissues of mice 12 to 24 h post-infection, while levels of K63 ubiquitination remained relatively unchanged (Supplementary Fig. 6c). These results indicate that Plasmodium infection disrupts the dynamic balance between palmitoylation and depalmitoylation of TBK1, and APT2 further depalmitoylates TBK1, rendering it more susceptible to recognition and modification by K48-linked ubiquitin chains, ultimately leading to autophagic degradation pathways. Furthermore, treatment with ML349 decreased K48-linked polyubiquitination of TBK1 in both concentration- and time-dependent manners (Fig. 6b and Supplementary Fig. 6d), further confirming the role of APT2, as silencing APT2 with siRNA also led to reduced K48-linked polyubiquitination of endogenous TBK1 (Supplementary Fig. 6e). In addition, Lypla2 depletion significantly diminished K48-linked polyubiquitination of TBK1 induced by gDNA in RAW 264.7 cells (Fig. 6c). Consistently, APT2 enhances the K48-linked polyubiquitination of TBK1, whereas catalytically inactivated variants (S122A and C2S) failed to produce this effect (Fig. 6d). Notably, the TBK1 C292S mutant exhibited a higher level of K48-linked polyubiquitination compared to WT TBK1 (Fig. 6e), underscoring APT2’s role in promoting the K48-linked polyubiquitination of TBK1.
a HEK293T cells were transfected with FLAG-TBK1 and HA-tagged WT ubiquitin (HA-Ub) or its mutants, together with MYC-EV or MYC-APT2 for 24 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. b HEK293T cells were transfected with FLAG-TBK1 and HA-K48-linked ubiquitin for 24 h, followed by treatment with ML349 at indicated concentrations for 12 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. c WT or Lypla2-KO RAW 264.7 cells were stimulated with gDNA for 6 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-TBK1) and immunoblot analysis. This was repeated n = 3 independent times with similar results. d, e WT (d) and TBK1-KO (e) HEK293T cells were transfected with the indicated plasmids, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. f HEK293T cells were transfected with Scr shRNA or other E3 ligase-specific shRNAs for 24 h, followed by transfected with FLAG-TBK1, together with HA-K48-linked ubiquitin and MYC-EV or MYC-APT2 for 24 h, and then followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. g, h HEK293T cells were transfected with FLAG-EV, WT FLAG-APT2 or its catalytically inactivated variants for 24 h (g). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (h). The colocalization between TBK1 and TRIM27 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) is next to the image. This was repeated n = 3 independent times with similar results. i PEMs were stimulated with freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for IP and immunoblot analysis. This was repeated n = 3 independent times with similar results. j PEMs were stimulated with freeze-thawed iRBCs for 12 h. The colocalization between APT2 (red) and TRIM27 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. k HEK293T cells were transfected with WT FLAG-TBK1 or its K251R/K372R (2KR) mutant, together with HA-K48-linked ubiquitin and MYC-EV or MYC-APT2 for 24 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. l HEK293T cells were transfected with WT FLAG-TBK1 or its 2KR mutant, together with MYC-EV or MYC-APT2 for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. m HEK293T cells were transfected with WT FLAG-TBK1 or its 2KR mutant, together with a luciferase reporter for IFN-β-luc and MYC-EV or MYC-APT2 for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (g, h, m). Source data are provided as a Source Data file.
Ubiquitin ligase (E3) plays a pivotal role in selecting substrates, and this process is essential for promoting the ubiquitination and subsequent degradation of the substrate protein44. However, APT2 lacks E3 ligase activity, leading us to hypothesize that APT2 enhances TBK1’s K48-linked polyubiquitination by recruiting an E3 ligase. Previous studies have identified several E3 ligases, including RNF41, RNF128, NEDD4, TRIM27, DTX4, TRIP, and TRIM23, as mediators of TBK1 polyubiquitination and degradation15,16,41,45,46. We tested this hypothesis by using shRNA to knockdown the expression of these E3 ligases. Notably, silencing TRIM27 reversed the effects of APT2 on TBK1-induced IFN-β promoter activity, while knockdowns of RNF41, RNF128, and NEDD4 offered only partial rescue (Supplementary Fig. 6f, g). Furthermore, our investigation revealed that APT2 did not enhance K48-polyubiquitination of TBK1 after TRIM27 silencing (Fig. 6f). Immunoblotting analysis and PLA indicated that APT2 facilitates the binding between TBK1 and TRIM27, whereas APT2 S122A and C2S mutants had no effect (Fig. 6g and Supplementary Fig. 6h). In addition, the TBK1 C292S mutant showed increased interaction with TRIM27 (Fig. 6h). Exogenous and endogenous Co-IP experiments directly confirmed the specific binding of APT2 to TRIM27 (Fig. 6i and Supplementary Fig. 6i). In addition, confocal microscopy analysis clearly demonstrated that the interaction between APT2 and TRIM27 was significantly enhanced following stimulation with freeze-thawed iRBCs (Fig. 6j), suggesting that APT2 inhibits the IFN-I signaling pathway by promoting K48-linked polyubiquitination of TBK1 through recruiting TRIM27.
Prior research has established that TRIM27 catalyzes ubiquitination of TBK1 at the K251 and K372 residues47. Our immunoblotting results indicated that mutating these two sites abolished APT2’s ability to enhance K48 ubiquitination of TBK1, culminating in impaired degradation of TBK1 and activation of IFN-I signaling (Fig. 6k–m). These findings highlight the critical role of K48-linked polyubiquitination at K251 and K372 of TBK1 for APT2-mediated autophagic degradation.
ML349 enhances mice resistance to N67 infection
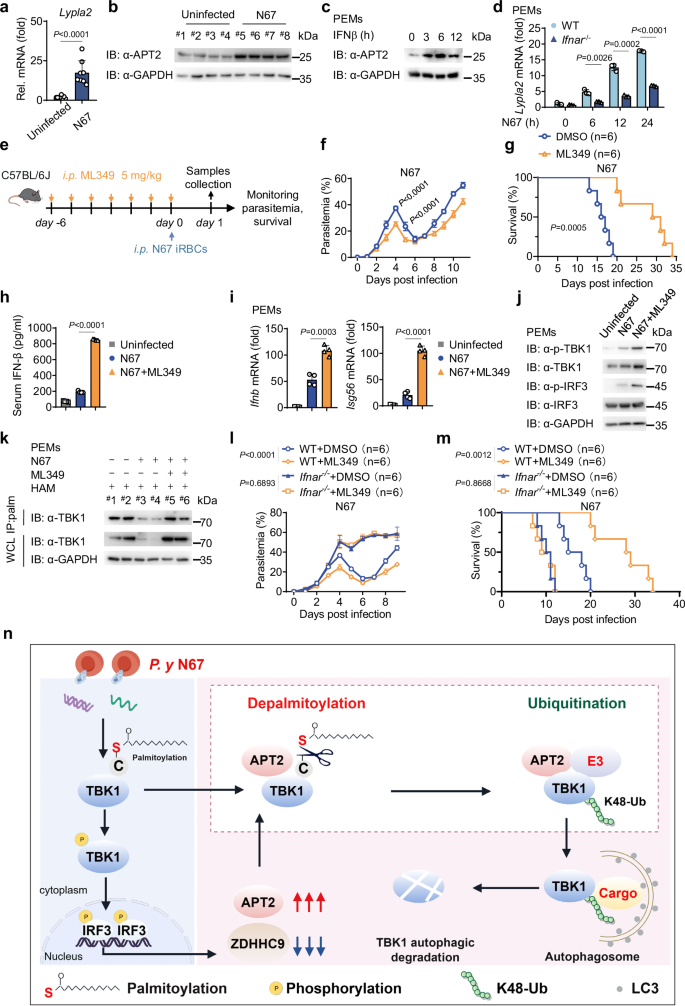
Finally, we examined the impact of N67 infection on the expression of APT2. Both RT-qPCR and immunoblotting analyses revealed a significant increase in APT2 mRNA and protein levels following N67 infection (Fig. 7a, b). Furthermore, stimulation of RAW 264.7 cells or PEMs with freeze-thawed iRBCs also resulted in elevated APT2 expression (Supplementary Fig. 7a–c), confirming that Plasmodium infection promotes the expression of APT2. Notably, recombined mouse IFN-β was able to induce APT2 expression in PEMs (Fig. 7c), and freeze-thawed iRBCs stimulation-induced expression of APT2 was markedly reduced in PEMs derived from Ifnar-deficient (Ifnar-/-) mice (Fig. 7d and Supplementary Fig. 7d). These results indicate that IFN-I can up-regulate the expression of APT2.
a, b C57BL/6 J female mice treated with phosphate-buffered saline (PBS) or infected with N67 via intraperitoneal injection for 24 h. The RT-qPCR analysis of APT2 mRNA presented here is derived from one representative experiment (n = 8 mice per group), and consistent results were observed across three biological replicates (a). In addition, immunoblotting (b) analysis was conducted to assess APT2 protein expression levels, this was repeated n = 3 independent times with similar results. c PEMs were treated with recombined mouse IFN-β at the indicated time points. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. d WT and Ifnar-/- PEMs were stimulated with freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates). e–g C57BL/6 J female mice (n = 6 per group) were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.). Parasitemia (f) and survival rate (g) were then evaluated. This was repeated n = 3 independent times with similar results. Figure 7e was adapted from the image “Adult Mouse” by Yunan Chen (SciDraw; CC BY 4.0). (h) C57BL/6 J female mice were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.) for 24 h. IFN-β in serum was determined by ELISA. Data shown are from one representative experiment (n = 4 mice per group), consistent results were observed across three biological replicates. i–k PEMs from mice as in (h). Cell lysates were then collected and analyzed by RT-qPCR; the data shown are from one representative experiment (n = 4 mice per group). Consistent results were observed across three biological replicates (i). Lysates were also subjected to immunoblot analysis (j, k), this was repeated n = 3 independent times with similar results. l, m WT and Ifnar-/- C57BL/6 J female mice (n = 6 per group) were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.). Parasitemia (l) and survival rate (m) were then evaluated. This was repeated n = 3 independent times with similar results. n Graphical abstract created with Figdraw (www.figdraw.com) depicting the mechanism by which Plasmodium evades the host immune response. Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, d, f, h, i), Two-way ANOVA (two tailed) with Tukey’s post hoc test (l) and two-sided log-rank test (g, m). Source data are provided as a Source Data file.
To further explore the role of APT2 in the host immune response to N67 infection, we employed ML349 as a model drug for pharmacological inhibition of APT225. Pretreatment with ML349 significantly reduced parasitemia, improved survival rates, and enhanced the IFN-I response after N67 infection, suggesting that ML349 treatment effectively promoted anti-malarial immune responses (Fig. 7e–j). Subsequent immunoblotting revealed that pretreatment with ML349 markedly increased the palmitoylation level and protein stability of TBK1 (Fig. 7k), suggesting that ML349 exerts its protective effects through modulating TBK1 stability. Next, we investigated whether the resistance conferred by ML349 to N67 infection is influenced by IFN-I by comparing the effect of ML349 in WT and Ifnar-/- mice. Our findings indicated that in the WT mice group, ML349 pretreatment resulted in a lower infection rate and higher survival rate after N67 infection, whereas these benefits were not observed in the Ifnar-/- group (Fig. 7l, m). These findings highlight that the protective effects of ML349 on host anti-malarial immunity depend on the IFN-I signaling pathway. Collectively, these data demonstrate that Plasmodium evades the host immune response by up-regulating APT2 and inhibiting the palmitoylation of TBK1 (Fig. 7n).
Discussion
Palmitoylation is crucial for protein localization and interactions in Plasmodium parasites, influencing the targeting of key proteins such as ISP1 and ISP3 to the inner membrane complexes (IMCs) during parasite division48. Despite its significance in these processes, the role of palmitoylation in shaping host immune responses to malaria remains underexplored, with most studies mainly focusing on antiviral and anti-tumor immunity26,28,30. In our research, we discovered that administration of the palmitoyltransferase inhibitor 2-BP resulted in decreased survival rates and increased susceptibility to infection in mice, underscoring the importance of palmitoylation in the host immune response to malaria and highlighting the need for further research into therapeutic strategies that target palmitoylation pathways in malaria treatment. Furthermore, malaria infection disrupted the balance between ZDHHC9-mediated palmitoylation and APT2-mediated depalmitoylation of the immune kinase TBK1, leading to its degradation via selective autophagy and subsequent inhibition of IFN-I production. Notably, treatment with APT2 inhibitor ML349 enhanced the IFN-I response, suggesting a potential therapeutic avenue for enhancing anti-malarial immunity. Our findings contribute to the broader understanding of palmitoylation’s role in various biological processes, including immune regulation, metabolic diseases, and neurodegenerative diseases21,32,49,50, by demonstrating its capacity to support anti-parasitic immune responses through IFN-I signaling activation.
TBK1 is known to be tightly regulated by various post-translational modifications in the context of infectious diseases and cancers38,51,52; however, its regulatory mechanisms in malaria have remained unclear. We found that the level of TBK1 palmitoylation is significantly down-regulated following Plasmodium infection. Through a detailed analysis of palmitoyltransferases and acyl protein thioesterases, we identified that the palmitoyltransferase ZDHHC9 is significantly down-regulated, while the acyl protein thioesterase APT2 is notably up-regulated after infection. Our findings indicate that ZDHHC9 acts as the primary palmitoyltransferase for TBK1 in resting conditions, maintaining a high palmitoylation state necessary for its stable and effective function. However, during Plasmodium infection, the down-regulation of ZDHHC9, coupled with the up-regulation of APT2, suggests a shift in the regulatory landscape. APT2 mediates the depalmitoylation of TBK1, thereby inhibiting the IFN-I pathway and playing a negative feedback role that prevents sustained activation of anti-malarial immunity. Here, we represent the report on the palmitoylation of TBK1, highlighting the significant changes in this PTM during malaria infection. By elucidating the intricate balance of palmitoylation and depalmitoylation in TBK1 regulation during malaria, our study provides insights into how these mechanisms may be manipulated to enhance host immune responses against the parasite.
During malaria infection, serum levels of IFN-α/β peak between 18 and 24 h post-infection but then rapidly returning to baseline levels53, suggesting the presence of a negative regulatory mechanism. Recent research has identified several regulators of the IFN-I response during malaria, including suppressor of cytokine signaling 1 (SOCS1), Fos-like antigen 1 (FOSL1), CD40, membrane-associated ring finger protein 1 (MARCH1), and receptor transporter 4 (RTP4)7,54,55,56,57. Our study indicates that Plasmodium infection promotes the expression of APT2, and experiments using Ifnar-/- mice confirm that this up-regulation is dependent on IFN-I, highlighting APT2’s role as a negative feedback regulator of the IFN-I signaling pathway. APT2 is known to interact with various molecules, including MAVS, p-STAT3, and GSDMD25,28,36,58. Notably, APT2 inhibits the antiviral immune response by depalmitoylating MAVS, while its action on p-STAT3 and GSDMD enhances their nuclear translocation and splicing, respectively. In contrast, our findings reveal that APT2-mediated depalmitoylation of TBK1 increases TBK1’s susceptibility to degradation via selective autophagy. This suggests that APT2 does not have a uniform function; instead, its effects vary depending on the substrate involved. Importantly, APT2 targets both upstream (MAVS) and downstream (TBK1) components of the IFN-I signaling pathway, both of which help inhibit IFN-I activation. However, while previous literature36 showed no significant change in TBK1 stability following treatment with ML349, our finding highlights a discrepancy that may stem from differences in treatment duration or concentration.
Emerging evidence indicates that autophagy has both positive and negative regulatory roles in the IFN-I signaling pathway, helping to prevent excessive inflammation during acute infection43,59,60,61,62. This positions autophagy as a crucial regulator of IFN-I signaling. Notably, APT2 lacks the classical cargo receptor motif and E3 ligase activity, necessitating the involvement of additional molecules for its function. APT2 interacts with the cargo receptor NDP52 and the E3 ligase TRIM27, which facilitates the association between TBK1 and these proteins. TRIM27 mediates K48-linked ubiquitination of TBK1 at lysine residues 251 and 372, serving as a recognition signal that binds to NDP52 and promotes the selective autophagic degradation of TBK1. Interestingly, ZDHHC4 enhances MAVS stability and activity by palmitoylating MAVS at Cys79, which inhibits K48-linked ubiquitination while promoting K63-linked ubiquitination28. In addition, the palmitoylation of NOD2 prevents its degradation through the p62/SQSTM1-mediated autophagy pathway63, whereas the palmitoylation of NLRP3 promotes its degradation via the chaperone-mediated autophagy (CMA) pathway37. These varying effects of palmitoylation on autophagy may arise from differences in substrate-enzyme interactions, membrane domain localization, and protein conformation, highlighting the complexity and diversity of palmitoylation mechanisms.
IFN-α/β produced in response to various malaria species plays a dual role in the immune response against malaria7,64,65,66. Research indicates that the TLR7-IRF7 pathway facilitates the early production of IFN-α/β following infection with Plasmodium yoelii 17XL, thereby enhancing the anti-malarial immune response. In contrast, the late production of IFN-α/β induced by TLR9-IRF7 and STING-IRF3 signaling has been shown to inhibit the generation of malaria antigen-specific CD4+ T cells while promoting the development of PD-1+CD8+ T cells, ultimately suppressing the host immune response during Plasmodium berghei ANKA infection8. Beyond the STING/MAVS pathway, TBK1 also serves as a pivotal kinase in the TLR-dependent signaling cascade13. This pathway has the capacity to induce IFN-α/β production in response to Plasmodium stimulation67,68. Plasmodium infection may coordinately regulate the expression of the depalmitoylating enzyme APT2 and the palmitoyltransferase ZDHHC9 by activating multiple innate immune signaling pathways, including TLR-TRIF, MAVS, and STING, thereby mediating the reduction in TBK1 palmitoylation levels. In our study, we found that IFN-β production upregulated APT2 in the early stages of N67 infection, where APT2 suppressed the anti-malarial immune response by promoting the depalmitoylation and selective autophagic degradation of TBK1. Given that different parasites exhibit distinct infection patterns, further investigation is necessary to understand how palmitoylation modification affects other strains of Plasmodium and various pathogens. In addition, the role of APT2 in inhibiting the IFN-I signaling pathway and its potential as a determinant across different Plasmodium strains requires further validation, which could support the development of targeted therapies for malaria. Importantly, several autoimmune diseases, including systemic lupus erythematosus (SLE), Sjögren’s syndrome, and systemic scleroderma69,70,71, require precise regulation of the IFN-I response, suggesting that APT2 may also represent a potential therapeutic target for these conditions.
The resurgence of malaria in many African and South American countries poses significant public health challenges. In the Greater Mekong region, some Plasmodium falciparum strains have developed resistance to artemisinin, a trend now emerging in Africa as well. In addition, Anopheles mosquitoes are evolving, exhibiting both physiological and behavioral resistance to insecticides1. This situation underscores the urgent need for more effective therapeutic options. Our study reveals that treatment with the APT2 inhibitor ML349 significantly reduced infection rates in WT mice, a result dependent on IFN-I signaling, as evidenced by the lack of effect in Ifnar-/- mice. We also observed that TBK1 palmitoylation levels decreased significantly in WT mice post-infection but remained unchanged with ML349 prophylaxis. This indicates that inhibiting APT2 enhances TBK1 palmitoylation, thereby activating IFN-I and reducing parasitemias. Our findings indicate that targeting APT2 may offer a strategy for malaria prevention and treatment, contributing to the understanding of the underlying mechanisms involved.
Collectively, palmitoylation contributes to a protective role in anti-malarial immunity. During malaria parasite infection, early IFN-I production disrupts the palmitoylation balance between ZDHHC9-TBK1-APT2. The upregulation of APT2 promotes the selective autophagic degradation of TBK1 through its depalmitoylation, thereby establishing a negative feedback mechanism. Treatment with APT2 inhibitor ML349 enhances the anti-malarial immune response by elevating the palmitoylation level of TBK1 and increasing IFN-I production. Consequently, the discovery of the immunosuppressive molecule APT2 during the Plasmodium infection process not only serve as a molecular marker for prognostic assessment in malaria patients, but also offer a target for immune intervention against malaria and supports the theoretical foundation for designing effective malaria vaccines.
Methods
Mice
All animal experiments were approved by the Institutional Animal Care and Use Committee of Southern Medical University (Approval number: SMUL2019243). All animal experiments were performed in specific pathogen-free levels using 6- to 8-week-old female C57BL/6 J mice. Females were selected because male mice are known to be more resistant to infection with Plasmodium species, exhibiting lower infection rates and highly variable parasitemia compared to females72. The use of female mice ensured the establishment of a robust, consistent, and reproducible infection model, thereby minimizing sex-based variability. Mice were kept with 50% to 60% humidity and daily cycles of 12 h of light at an ambient temperature of 21 °C to 23 °C. At the times indicated in the figure legends, the mice were anesthetized with isoflurane and euthanized by cervical dislocation.
Cell lines and culture methods
Human embryonic kidney (HEK) 293 T (Cat#CRL-321) and murine macrophage RAW 264.7 cells (Cat#TIB-7) were obtained from the American Type Culture Collection (ATCC). HEK293T and RAW 264.7 cells were cultured in DMEM (Corning, Cat#10-013-CV) containing 10% fetal bovine serum (FBS) (HyClone, Cat#BS-1101-3) incubated in a 5% CO2 chamber (Thermo Fisher Scientific). BMDMs were obtained from the rear-leg bone marrow of two-month-old littermate and female C57BL/6 mice (Guangdong Medical Laboratory Animal Center, Guangzhou, Guangdong, China), and differentiated into macrophages for six days with 20% filtered L929 cell conditional medium (a source of macrophage colony-stimulating factor). Mouse PEMs were acquired from ascites of female C57BL/6 mice, which were injected intraperitoneally with 4% Brewer thioglycolate medium (BD Biosciences, Cat#211716) for three consecutive days before sacrifice, then cold PBS (Bio-Channel, Cat#BC-BPBS-08) was injected into the peritoneum of sacrificed mice, and fluid containing PEMs was aspirated from the peritoneum after the shake and collected after centrifugation at 300 × g for 5 min.
Reagents
Anti-APT-2 (Cat#ab151578, 1:1000, polyclonal), anti-rabbit immunoglobulin G (IgG) H&L horseradish peroxidase (Cat#ab6721, 1:10000, polyclonal), goat anti-mouse IgG H&L GRP (Cat#ab6789, 1:10000, polyclonal), goat anti-mouse IgG H&L AF594 (Cat#ab150116, 1:200, polyclonal), goat anti-rat IgG H&L AF488 (Cat#ab150165, 1:200, polyclonal) were purchased from Abcam. Anti-phosphor-TBK1/NAK (Ser172) (Cat#5483, 1:1000, clone name D52C2), anti-phosphor-IRF3 (Ser396) (Cat#4947, 1:1000, clone name 4D4G), anti-IRF3 (Cat#11904, 1:1000, clone name D6I4C), anti-hemagglutinin (HA) (Cat#5017, 1:1000, clone name C29F4), anti-MYC (Cat#2276, 1:1000, clone name 9B11), anti-p62 (Cat#88588, 1:1000, clone name D5L7G), anti-K48-Ub (Cat#8081, 1:1000, clone name D9D5), anti-phosphor-STING (Ser365) (Cat#72971, 1:1000, clone name D8F4W) were purchased from Cell Signaling Technology. Anti-NDP52 (Cat#12229-1-AP, 1:1000, polyclonal), anti-TOLLIP (Cat#11315-1-AP, 1:1000, polyclonal), anti-TBK1 (Cat#83686-3-RR, 1:10000, clone name 240769G5), anti-STING (Cat# 19851-1-AP, 1:20000, polyclonal), anti-p65 (Cat#80979-1-RR, 1:1000, clone name 4C7), anti-ZDHHC9 (Cat#82739-1-RR, 1:1000, clone name 9G5) were purchased from Proteintech. Anti-FLAG (Cat#A8592, 1:1000, clone name M2) was purchased from Sigma-Aldrich. Anti-TRIM27 (Cat#823342, 1:1000, polyclonal), anti-GAPDH (Cat#200306-7E4, 1:10000, clone name 7E4-H6-H6), anti-MAVS (Cat#161894, 1:1000, polyclonal), anti-phosphor-p65 (Ser536) (Cat#340830, 1:1000, polyclonal), anti-Histone H3 (Cat#R24572, 1:1000, clone name R06-4I8), anti-GFP (Cat#250065, 1:1000, clone name 6A2-1H8-5G7), anti-K63-Ub (Cat#R381564, 1:1000, clone name R06-2A6), anti-Ub (Cat#R26025, 1:1000, clone name R02-4I6) were purchased from Zen-Bio. 2-bromopalmitate (2-BP) (Cat#21604), Hydroxylamine (HAM) (Cat#379921), NeutrAvidin agarose beads (Cat#29200), Flag-beads (Cat#A2220) were purchased from Sigma-Aldrich. 3-MA (Cat#S2767) and Baf A1 (Cat#S1413) were purchased from Selleck. MG132 (Cat#S2619) was purchased from Ambeed (A181909). Cycloheximide (CHX) (Cat#2112S) was purchased from Cell Signaling Technology. ML349 (Cat#HY-100737), protein A and G beads (Cat#HY-K0202) were purchased from MedChem Express.
Construction of plasmids and transfection
Plasmids and their domain constructs were cloned into the pcDNA3.1 vector, incorporating FLAG-, HA-, and MYC-tags for transient expression, utilizing In-Fusion HD Cloning kits (Takara, Cat#639650). Site-directed mutagenesis (SDM) was performed using the MutanBEST Kit (Takara, Cat#R401) to generate relevant mutants. Transfection of HEK293T cells was conducted with either StarFect Transfection Reagent (Genstar, Cat#C101), Lipomaster 3000 Transfection Reagent (Vazyme, Cat#TL301) or Lipofectamine 2000 (Invitrogen, Cat#11668019), following the protocols recommended by the manufacturers, with primers detailed in Supplementary Data 1.
Western blotting and Co-immunoprecipitation
Western blotting: For total protein extraction, cells were lysed in RIPA buffer (composed of 50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS) supplemented with protease inhibitor cocktail (MCE, Cat#HY-K0010). Following lysis, the samples were centrifuged at 16,800 × g for 10 minutes. The supernatant obtained was collected and subsequently combined with an appropriate volume of 5 × loading buffer. Equal quantities of protein from each experimental group were resolved using SDS-PAGE and electrotransferred onto a polyvinylidene difluoride (PVDF) membrane at 100 V for a duration of 1.5 h. Following transfer, the PVDF membrane was blocked with 5% non-fat powdered milk for 1 h at room temperature. The membrane was then incubated with primary antibodies in universal antibody diluent (NCM Biotech, Cat#WB500D) overnight at 4 °C, followed by incubation with an HRP-conjugated secondary antibody. Protein signals were detected using enhanced chemiluminescence (ECL) detection systems.
Co-immunoprecipitation (Co-IP): Cells were washed three times with ice-cold PBS and lysed on ice for 10 min using RIPA lysis buffer supplemented with protease inhibitor cocktail. A small aliquot of the lysate was reserved as input, boiled at 100 °C for 5 min in 1 × SDS loading buffer. The remaining lysate was subjected to immunoprecipitation (IP) by incubation with either anti-FLAG (Sigma-Aldrich, Cat#A8592) or anti-HA (Cell Signaling Technology, Cat#5017) antibody. For endogenous IP, lysates were incubated with the indicated antibodies or IgG control overnight at 4 °C, followed by incubation with Protein A/G magnetic beads (MedChemExpress, Cat#HY-K0202) for 4–6 h. Protein-bound magnetic beads were then collected using a magnetic stand and washed gently three times with lysis buffer. Finally, the bound proteins were eluted, resolved by SDS-PAGE, and subjected to immunoblotting analysis with the indicated antibodies. The acquisition and quantitative analysis of protein bands were carried out using Image Lab software (version 6.0.1, Bio-Rad).
Confocal analysis
HEK 293 T, BMDMs or PEMs cultured on glass coverslips were subjected to various treatments as depicted in the figure. The cells underwent three washes with PBS, followed by fixation with 4% paraformaldehyde diluted in PBS for 20 min, permeabilization with 0.2% Triton X-100 (Sangon Biotech, Cat#A600198) for 20 min, and blocking with 3% BSA (Sangon Biotech, Cat#A600903) at room temperature for 30 min. Subsequently, they were incubated at 4 °C for 12 h with the corresponding antibodies, followed by one-hour incubation at room temperature with fluorescently labeled secondary antibodies. Finally, the cell nuclei were stained with DAPI (Sigma-Aldrich, Cat#F6057) for 5 min prior to observation under confocal microscopy. The obtained original image data were subsequently processed and analyzed using ZEISS ZEN 3.9 software.
Proximity ligation assay (PLA)
The cells cultured on glass coverslips were treated as illustrated in the figure. They underwent three washes with PBS, followed by fixation using 4% paraformaldehyde diluted in PBS for 20 minutes. Subsequently, permeabilization was performed with 0.2% Triton X-100 for an additional 20 mins, and blocking was conducted at room temperature using 3% BSA for 30 min. Following these steps, the cells were incubated at 4 °C for 12 h with the appropriate antibodies. After incubation, the cells were washed twice with TBST for five minutes each. This was followed by a ligation reaction (Sigma-Aldrich, Cat#DUO92008) at 37 °C lasting for 30 min and an amplification reaction at the same temperature lasting approximately 100 min. After washing with PLA wash buffer B for ten minutes, cell nuclei were stained with DAPI for five minutes prior to observation under confocal microscopy. The obtained original image data were subsequently processed and analyzed using ZEISS ZEN 3.9 software.
Acyl-biotin exchange (ABE) assay
The indicated cells were harvested using a low-salt lysis buffer and subsequently placed on a rocker with ice for 30 min. Following this, centrifugation was performed at 16,800 × g at 4 °C for 15 min. The cell lysates were then treated with TCEP (500 mM) (Sigma-Aldrich, Cat# C4706) at 55 °C for one hour and incubated overnight with NEM (50 mM) (Sigma-Aldrich, Cat# E3876) at 4 °C. Removal of NEM from the cell lysates was accomplished through repeated methanol-chloroform precipitation. The resulting protein pellets were dried at room temperature and dissolved in SDS buffer containing 4% SDS, 5 mM EDTA, and 50 mM Tris (pH 8.0), along with a protease inhibitor at 37 °C. Each sample was divided into two portions and mixed separately with the thiol-reactive biotin molecule HPDP-biotin (0.4 mM) (Thermo Fisher Scientific, Cat#21341). The portion that excluded the hydroxylamine hydrochloride (HAM) (Sigma-Aldrich, Cat#379921) cleavage step (-HAM) was supplemented with lysis buffer, while the portion including the HAM step (+HAM) received an addition of 1 M HAM. After 5 h of incubation at 4 °C, samples were pelleted via methanol-chloroform precipitation and then resuspended in a solution of SDS buffer. Following dissolution, one-fifth of each sample was eluted using a 5 × loading buffer and boiled at 100 °C for five minutes. Meanwhile, the remaining samples underwent incubation overnight with neutravidin agarose beads at 4 °C. The beads were washed four times using lysis buffer before being eluted in 2 × loading buffer and subsequently resolved by SDS-PAGE.
Enzyme-linked immunosorbent assay (ELISA)
IFN-β levels in cell supernatants and mouse serum were quantified using the Mouse IFN-β DuoSet ELISA kit (R&D Systems, Cat#DY8234-05) according to the manufacturer’s protocol. Absorbance was measured at 450 nm using the Multiskan FC (Thermo Fisher Scientific).
RNA-seq data analyses and functional interpretations
Total RNA extracted from PEMs subjected to the indicated treatments was analyzed using RNA sequencing (RNA-seq). The quality of the RNA was assessed with a 2100 Bioanalyzer (Agilent) and quantified using the ND-2000 spectrophotometer (NanoDrop Technologies). The RNA-seq transcriptome library was prepared according to the NovaSeq X Series 25B Reagent Kit (100 cycles) (Illumina, Cat#20125967), utilizing 1 μg of total RNA. Subsequently, the paired-end RNA-seq library was sequenced on an Illumina NovaSeq X Plus Sequencing System.
Total RNA extraction and quantitative real-time PCR analysis
Total RNA was extracted from tissue or cells using TRIzol reagent (Invitrogen, Cat#15596026CN). Subsequently, complementary DNA (cDNA) was synthesized employing the Starscript II first-strand cDNA synthesis kit (GenStar, Cat#A240). Real-time PCR was conducted on a QuantStudio 6 Flex system (Thermo Fisher Scientific) utilizing a RealStar Green Power Mixture (GenStar, Cat#A308), with primers detailed in Supplementary Data 1.
Luciferase and reporter assays
HEK293T cells (1 × 105) were seeded in 24-well plates (Vazyme, Cat#CCP01024) and transfected with plasmids encoding the IFN-β/ISRE luciferase reporter (Firefly luciferase, 30 ng) and pRL-TK (Renilla luciferase, 10 ng), along with various additional plasmids or stimuli. Samples were collected, and luciferase activity was measured using the Dual Luciferase Assay (Promega, Cat#E1910) on a Luminoskan Ascent luminometer (Thermo Fisher Scientific). Reporter gene activity was assessed by normalizing firefly luciferase activity to Renilla luciferase activity. The values presented are mean ± SD from three independent experiments.
Gene knockdown by shRNA
The shRNA oligonucleotide sequences and their corresponding target genes are detailed in Supplementary Data 1. These sequences were designed using the website (https://rnaidesigner.thermofisher.com/rnaiexpress/sort.do), with selection based on the highest scoring, followed by annealing, cloning into the PLKO.1-Puro vector, and confirmation through sequencing.
Small interfering RNA (siRNA) transfection
For knockdown experiments, the small interfering RNA (siRNA) targeting specific genes were procured from Sangon (Shanghai, China) and transfected using Lipofectamine RNAiMAX (Invitrogen) in accordance with the manufacturer’s instructions. Sequences of siRNAs oligonucleotides targeting the corresponding genes are presented in Supplementary Data 1.
Generation of knockdown and reconstitution cell lines
Sequences of guide RNA (gRNA) oligonucleotides targeting the corresponding genes are presented in Supplementary Data 1. These sequences were designed using the online tool available at https://portals.broadinstitute.org/gpp/public/analysis-tools/sgrna-design. The sgRNA primers were annealed and subsequently cloned into the lenti-sgV2 vector to generate Lypla2-targeting vectors, which were confirmed by sequencing. Following this, these vectors were transfected into RAW 264.7 cells for 48 h. Subsequently, the supernatant was replaced with fresh culture medium containing puromycin (5 mg/mL) (Sigma-Aldrich, Cat#540411). Live cells were screened after an additional 48 h and subjected to Western blot analysis to assess knockout efficiency.
Extraction of malarial gDNA and RNA
Parasite-infected mice blood was collected in saline solution and filtered to deplete white blood cells. Parasites were spun down after RBC lysis buffer treatment, and the lysate was incubated with buffer A (150 mM NaCl, 25 mM EDTA, 10% SDS, and protein kinase) overnight. gDNAs were isolated using phenol/chloroform, and RNAs were isolated using TRIzol reagent (Invitrogen, CA, USA).
Molecular docking
Multiple sequence alignment and structural comparison. The amino-acid sequences of Mus musculus Lypla2 (Q9WTL7) and Mus musculus Tbk1 (A2A9T0) were retrieved and aligned online at the alphafold website (https://alphafold.ebi.ac.uk/), and the targeted amino acid residues were determined according to the interaction distance within 5 Å. Structures and maps in the figures were rendered with PyMOL (The PyMOL Molecular Graphics System, v.4.0, Schrödinger).
Malaria parasite infection procedures
The parasites N67 were initially obtained from the Malaria Research and Reference Reagent Resource Center (MR4, https://www.beiresources.org/About/MR4.aspx). For N67 infection, 1 × 106 iRBCs suspended in 200 μL PBS from the donor mice were injected i.p. into experimental mice (6- to 8-week-old female C57BL/6 J mice). All animal experiments were approved by the Southern Medical University Animal Care and Use Committee (SMUL2019243).
Giemsa staining
Blood was collected from the tail vein of mice infected with the N67 parasite strain, and thin blood smears were immediately prepared on clean glass slides. After the blood smears were completely air-dried, they were fixed with 100% methanol for 2–5 min. Then, they were stained with diluted Giemsa stain (Sigma-Aldrich, Cat#51811-82-6) for 30 min. After staining, the glass slides were gently rinsed with running water to remove excess stain and then air-dried naturally.
The stained blood smears were examined under an optical microscope using an oil immersion lens (100 × objective). The parasitemia was calculated by counting the number of infected red blood cells (iRBCs) among at least 1000 red blood cells (RBCs). The parasitemia (%) was calculated according to the following formula: Parasitemia (%) = (Number of infected RBCs / total number of RBCs counted) × 100%. Regarding reproducibility: For each sample, 10 fields of view were counted, or multiple blood smears were prepared, and the average value was taken as the final parasitemia result.
Statistical analysis
Data are representative of three independent experiments. All statistical analyses were performed using GraphPad Prism 9.4.1 (GraphPad software, inc.). All data are shown as mean ± SD. Statistical analyses were performed by unpaired two-tailed Student’s t tests, or by the two-sided log-rank test and Two-way ANOVA with Tukey’s post hoc test. P-values < 0.05 were considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq datasets generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under the BioProject accession PRJNA1288125. All other data are available in the article and its Supplementary files or from the corresponding author upon request. All newly generated materials in this study are available from the corresponding author upon request. Source data are provided in this paper.
References
Poespoprodjo, J. R., Douglas, N. M., Ansong, D., Kho, S. & Anstey, N. M. Malaria. Lancet 402, 2328–2345 (2023).
Monroe, A., Williams, N. A., Ogoma, S., Karema, C. & Okumu, F. Reflections on the 2021 World Malaria Report and the future of malaria control. Malar. J. 21, 154 (2022).
Boyle, M. J., Engwerda, C. R. & Jagannathan, P. The impact of Plasmodium-driven immunoregulatory networks on immunity to malaria. Nat. Rev. Immunol. 24, 637–653 (2024).
Parroche, P. et al. Malaria hemozoin is immunologically inert but radically enhances innate responses by presenting malaria DNA to Toll-like receptor 9. Proc. Natl. Acad. Sci. USA 104, 1919–1924 (2007).
Schofield, L. & Grau, G. E. Immunological processes in malaria pathogenesis. Nat. Rev. Immunol. 5, 722–735 (2005).
Yu, X. et al. Inflammasome activation negatively regulates MyD88-IRF7 type I IFN signaling and anti-malaria immunity. Nat. Commun. 9, 4964 (2018).
Yu, X. et al. Cross-regulation of two type I interferon signaling pathways in plasmacytoid dendritic cells controls Anti-malaria immunity and host mortality. Immunity 45, 1093–1107 (2016).
Lu, J. et al. Dual nature of type I interferon responses and feedback regulations by SOCS1 dictate malaria mortality. J. Adv. Res. 73, 295–310 (2025).
Liehl, P. et al. Host-cell sensors for Plasmodium activate innate immunity against liver-stage infection. Nat. Med. 20, 47–53 (2014).
Patel, H. et al. Malaria blood stage infection suppresses liver stage infection via host-induced interferons but not hepcidin. Nat. Commun. 15, 2104 (2024).
Vigário, A. M. et al. Recombinant human IFN-alpha inhibits cerebral malaria and reduces parasite burden in mice. J. Immunol. 178, 6416–6425 (2007).
Ding, C., Song, Z., Shen, A., Chen, T. & Zhang, A. Small molecules targeting the innate immune cGAS‒STING‒TBK1 signaling pathway. Acta Pharm. Sin. B 10, 2272–2298 (2020).
Fitzgerald, K. A. et al. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 4, 491–496 (2003).
Li, H. et al. SHIP1 modulates antimalarial immunity by bridging the crosstalk between type I IFN signaling and autophagy. mBio 14, e0351222 (2023).
Zhang, M. et al. TRAF-interacting protein (TRIP) negatively regulates IFN-β production and antiviral response by promoting proteasomal degradation of TANK-binding kinase 1. J. Exp. Med. 209, 1703–1711 (2012).
Xie, W. et al. Selective autophagy controls the stability of TBK1 via NEDD4 to balance host defense. Cell Death Differ. 29, 40–53 (2022).
Li, X. et al. Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat. Immunol. 17, 806–815 (2016).
Ma, X. et al. Molecular basis of Tank-binding kinase 1 activation by transautophosphorylation. Proc. Natl. Acad. Sci. USA 109, 9378–9383 (2012).
Elliot Murphy, R. & Banerjee, A. In vitro reconstitution of substrate S-acylation by the zDHHC family of protein acyltransferases. Open Biol. 12, 210390 (2022).
Wang, Y. & Yang, W. Proteome-scale analysis of protein S-acylation comes of age. J. Proteome Res. 20, 14–26 (2021).
Das, T., Yount, J. S. & Hang, H. C. Protein S-palmitoylation in immunity. Open Biol. 11, 200411 (2021).
Wang, X., Qian, P., Cui, H., Yao, L. & Yuan, J. A protein palmitoylation cascade regulates microtubule cytoskeleton integrity in Plasmodium. Embo J. 40, e109070 (2021).
Counihan, N. A., Chernih, H. C. & de Koning-Ward, T. F. Post-translational lipid modifications in Plasmodium parasites. Curr. Opin. Microbiol. 69, 102196 (2022).
Jones, M. L., Collins, M. O., Goulding, D., Choudhary, J. S. & Rayner, J. C. Analysis of protein palmitoylation reveals a pervasive role in Plasmodium development and pathogenesis. Cell Host Microbe 12, 246–258 (2012).
Zhang, M. et al. A STAT3 palmitoylation cycle promotes T(H)17 differentiation and colitis. Nature 586, 434–439 (2020).
Fan, Y. et al. Targeting LYPLAL1-mediated cGAS depalmitoylation enhances the response to anti-tumor immunotherapy. Mol. Cell 83, 3520–3532 (2023).
Hansen, A. L., Mukai, K., Schopfer, F. J., Taguchi, T. & Holm, C. K. STING palmitoylation as a therapeutic target. Cell Mol. Immunol. 16, 236–241 (2019).
Zhang, G. et al. CPT1A induction following epigenetic perturbation promotes MAVS palmitoylation and activation to potentiate antitumor immunity. Mol. Cell 83, 4370–4385 (2023).
Wang, L. et al. Palmitoylation acts as a checkpoint for MAVS aggregation to promote antiviral innate immune responses. J. Clin. Invest. 134, e177924 (2024).
Yao, H. et al. Inhibiting PD-L1 palmitoylation enhances T-cell immune responses against tumours. Nat. Biomed. Eng. 3, 306–317 (2019).
Brigidi, G.S. & Bamji, S.X. Detection of protein palmitoylation in cultured hippocampal neurons by immunoprecipitation and acyl-biotin exchange (ABE). J. Vis. Exp.72, https://doi.org/10.3791/50031 (2013).
Ko, P.J. & Dixon, S.J. Protein palmitoylation and cancer. EMBO Rep. 19, e46666 (2018).
Swarthout, J. T. et al. DHHC9 and GCP16 constitute a human protein fatty acyltransferase with specificity for H- and N-Ras. J. Biol. Chem. 280, 31141–31148 (2005).
Hornemann, T. Palmitoylation and depalmitoylation defects. J. Inherit. Metab. Dis. 38, 179–186 (2015).
Abrami, L. et al. Palmitoylated acyl protein thioesterase APT2 deforms membranes to extract substrate acyl chains. Nat. Chem. Biol. 17, 438–447 (2021).
Bu, L. et al. Targeting APT2 improves MAVS palmitoylation and antiviral innate immunity. Mol. Cell 84, 3513–3529 (2024).
Wang, L. et al. Palmitoylation prevents sustained inflammation by limiting NLRP3 inflammasome activation through chaperone-mediated autophagy. Mol. Cell 83, 281–297 (2023).
Zhao, X. et al. USP19 (ubiquitin specific peptidase 19) promotes TBK1 (TANK-binding kinase 1) degradation via chaperone-mediated autophagy. Autophagy 18, 891–908 (2022).
Sui, C. et al. SARS-CoV-2 NSP13 Inhibits type I IFN production by degradation of TBK1 via p62-dependent selective autophagy. J. Immunol. 208, 753–761 (2022).
Mizushima, N. & Yoshimori, T. How to interpret LC3 immunoblotting. Autophagy 3, 542–545 (2007).
He, X. et al. RNF34 functions in immunity and selective mitophagy by targeting MAVS for autophagic degradation. Embo J. 38, e100978 (2019).
Wu, Y. et al. Selective autophagy controls the stability of transcription factor IRF3 to balance type I interferon production and immune suppression. Autophagy 17, 1379–1392 (2021).
Hu, Z. et al. VANGL2 inhibits antiviral IFN-I signaling by targeting TBK1 for autophagic degradation. Sci. Adv. 9, eadg2339 (2023).
Toma-Fukai, S. & Shimizu, T. Structural diversity of Ubiquitin E3 ligase. Molecules 26, https://doi.org/10.3390/molecules26216682 (2021).
Lin, M. et al. USP38 Inhibits type I interferon signaling by editing TBK1 ubiquitination through NLRP4 signalosome. Mol. Cell 64, 267–281 (2016).
Sparrer, K. M. J. et al. TRIM23 mediates virus-induced autophagy via activation of TBK1. Nat. Microbiol. 2, 1543–1557 (2017).
Zheng, Q. et al. Siglec1 suppresses antiviral innate immune response by inducing TBK1 degradation via the ubiquitin ligase TRIM27. Cell Res. 25, 1121–1136 (2015).
Wang, X., Qian, P., Cui, H., Yao, L. & Yuan, J. A protein palmitoylation cascade regulates microtubule cytoskeleton integrity in Plasmodium. Embo J. 39, e104168 (2020).
Gu, M. et al. Palmitoyltransferase DHHC9 and acyl protein thioesterase APT1 modulate renal fibrosis through regulating β-catenin palmitoylation. Nat. Commun. 14, 6682 (2023).
Balasubramanian, A. et al. The palmitoylation of gasdermin D directs its membrane translocation and pore formation during pyroptosis. Sci. Immunol. 9, eadn1452 (2024).
Lin, Y. et al. PTK2B promotes TBK1 and STING oligomerization and enhances the STING-TBK1 signaling. Nat. Commun. 14, 7567 (2023).
Hu, Z. et al. The toxoplasma effector GRA4 hijacks host TBK1 to oppositely regulate anti-T. gondii immunity and tumor immunotherapy. Adv. Sci. 11, e2400952 (2024).
He, X., Xia, L., Tumas, K. C., Wu, J. & Su, X. Z. Type I interferons and Malaria: A double-edge sword against a complex parasitic disease. Front. Cell Infect. Microbiol. 10, 594621 (2020).
Cai, B., Wu, J., Yu, X., Su, X.Z. & Wang, R.F. FOSL1 Inhibits type I interferon responses to Malaria and viral infections by blocking TBK1 and TRAF3/TRIF interactions. mBio 8, https://doi.org/10.1128/mbio.02161-16 (2017).
Yao, X. et al. Increased CD40 Expression enhances early STING-mediated type I interferon response and host survival in a rodent Malaria model. PLoS Pathog. 12, e1005930 (2016).
Wu, J. et al. The E3 ubiquitin ligase MARCH1 regulates antimalaria immunity through interferon signaling and T cell activation. Proc. Natl. Acad. Sci. USA 117, 16567–16578 (2020).
He, X. et al. RTP4 inhibits IFN-I response and enhances experimental cerebral malaria and neuropathology. Proc. Natl. Acad. Sci. USA 117, 19465–19474 (2020).
Zhang, N. et al. A palmitoylation-depalmitoylation relay spatiotemporally controls GSDMD activation in pyroptosis. Nat. Cell Biol. 26, 757–769 (2024).
Li, J. et al. Porcine reproductive and respiratory syndrome virus degrades DDX10 via SQSTM1/p62-dependent selective autophagy to antagonize its antiviral activity. Autophagy 19, 2257–2274 (2023).
Xie, B. et al. Induction of autophagy and suppression of type I IFN secretion by CSFV. Autophagy 17, 925–947 (2021).
Zhang, X., Bai, X. C. & Chen, Z. J. Structures and mechanisms in the cGAS-STING innate immunity pathway. Immunity 53, 43–53 (2020).
Hopfner, K. P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
Zhou, L. et al. Palmitoylation restricts SQSTM1/p62-mediated autophagic degradation of NOD2 to modulate inflammation. Cell Death Differ. 29, 1541–1551 (2022).
Wu, J. et al. Strain-specific innate immune signaling pathways determine malaria parasitemia dynamics and host mortality. Proc. Natl. Acad. Sci. USA 111, E511–E520 (2014).
Haque, A. et al. Type I interferons suppress CD4⁺ T-cell-dependent parasite control during blood-stage Plasmodium infection. Eur. J. Immunol. 41, 2688–2698 (2011).
Ball, E. A. et al. IFNAR1 controls progression to cerebral malaria in children and CD8+ T cell brain pathology in Plasmodium berghei-infected mice. J. Immunol. 190, 5118–5127 (2013).
Couper, K. N. et al. Parasite-derived plasma microparticles contribute significantly to malaria infection-induced inflammation through potent macrophage stimulation. PLoS Pathog. 6, e1000744 (2010).
Mantel, P. Y. et al. Malaria-infected erythrocyte-derived microvesicles mediate cellular communication within the parasite population and with the host immune system. Cell Host Microbe 13, 521–534 (2013).
Fernandez-Ruiz, R. & Niewold, T. B. Type I interferons in autoimmunity. J. Invest. Dermatol. 142, 793–803 (2022).
Crow, M. K. Pathogenesis of systemic lupus erythematosus: risks, mechanisms and therapeutic targets. Ann. Rheum. Dis. 82, 999–1014 (2023).
Lazar, S. & Kahlenberg, J. M. Systemic lupus erythematosus: new diagnostic and therapeutic approaches. Annu. Rev. Med. 74, 339–352 (2023).
Duncombe, C.J. et al. Androgens inhibit protective CD8(+) T cell responses against pre-erythrocytic malaria parasites in mice. Nat. Commun. 16, 5172 (2025).
Acknowledgements
This work was supported by grants from the National Key R&D Program of China (2024YFC2309700) to X.Y., the National Natural Science Foundation of China (82572009, 82371761 and 82171741) to X.Y., the Guangdong Basic and Applied Basic Research Foundation (2023A1515010421) to X.Y. and the Guangzhou Science and Technology Research Program (2025A04J7166) to X.Y. Figures 1a and 7e were adapted from the image “Adult Mouse” by Yunan Chen, available from the SciDraw repository (https://scidraw.io/) under a Creative Commons Attribution 4.0 International (CC BY 4.0) license. Figure 7n was created with materials from Figdraw (https://www.figdraw.com). We are grateful to Dr. Jun Cui (Sun Yat-sen University) for kindly providing the ZDHHCs plasmids.
Author information
Authors and Affiliations
Contributions
X.Y. and Z.X.H. performed the experiment design. Z.X.H., S.Y.X., K.Z. and Z.L.X. performed major experiments, performed data analysis and wrote the original draft. L.Y.Z., Y.F.Z., J.Y.G., W.Q.P., Y.C.X. and W.W.L. supported the experiments of molecular and cellular biology. X.Y. revised the manuscript. X.Y. supervised all experiments and the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chul-Su Yang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, Z., Xiong, S., Zeng, K. et al. Palmitoylation of TBK1 enhances the type I interferon signaling and strengthens anti-malarial immunity in mice. Nat Commun 16, 10109 (2025). https://doi.org/10.1038/s41467-025-65081-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65081-8