Abstract
Structure-based sequence redesign or inverse folding can significantly enhance structural stability but often compromises functional activity when performed using existing models. Here, we introduce ABACUS-T, a multimodal inverse folding model that improves precision and minimizes functional loss. ABACUS-T unifies several important features into one framework: detailed atomic sidechains and ligand interactions, a pre-trained protein language model, multiple backbone conformational states, and evolutionary information from multiple sequence alignment (MSA). Redesigned proteins show notable improvements: an allose binding protein achieves 17-fold higher affinity while retaining conformational change; redesigned endo-1,4-β-xylanase and TEM β-lactamase maintain or surpass wild-type activity; and OXA β-lactamase gains altered substrate selectivity. All achieve substantially increase thermostability (∆Tm ≥ 10 °C). In each test case, these enhancements are achieved by testing only a few sequences, each containing dozens of simultaneously mutated residues. ABACUS-T thus offers a promising tool for reengineering functional proteins in biotechnological applications.
Similar content being viewed by others
Introduction
Protein engineering is widely applied to enhance natural proteins in one or more attributes to overcome their limitations in various biotechnological applications1. In many circumstances, to achieve the desired enhancements (for example, to substantially increase the melting temperature of a naturally mesophilic enzyme) needs the simultaneous mutations of a large number (e.g., several dozens) of residues2. To obtain such variants is unfeasible from directed evolution campaigns, which usually use sequence libraries that contain mutations at a few random or focused sites, producing outcomes that are only a few mutated residues away from the starting sequence3,4. Moreover, these campaigns typically require multiple rounds of screening over libraries containing thousands to millions of sequence variants4. Such screenings are usually costly, and not always viable.
There have been continuous efforts in developing computational methods to ease the heavy burdens on experiment in protein engineering2. In recent years, approaches based on machine learning have attracted wide attentions5,6. One class of approaches focus on building supervised machine learning models to predict the effects of mutations on various attributes of functional proteins. To train a useful model of such type usually requires extensive experimental data on a specific system7,8. Moreover, the experimental data available in most cases can only cover single-site mutations extensively without or with only sparse data of multi-site mutations, making it difficult to include epistasis effects. This causes the performances of such models to decline rapidly with increasing number of mutated residues. Although experiments and design could be iteratively executed to explore sequences further away from the starting ones9,10,11, few cases of variants containing dozens of simultaneously changed residues have been reported.
With the recent advances in deep learning models for natural language processing, large language models (LLMs) of proteins have been trained on the large corpus of known amino acid sequences of natural proteins12,13,14,15,16,17. Protein language models fine-tuned with homologous sequences of specific protein families have been found to be able to generate functionally active family members of non-natural sequences that deviate far from the natural sequences (with the highest identities of some generated sequences to natural sequences below 40%14. Applying such models can significantly enlarge the set of amino acid sequences that possess a certain (naturally existing) function beyond the relatively limited number of natural protein sequences sharing the same function. This may bring the chances of identifying proteins that are enhanced over natural proteins in one or more attributes. So far, successful adventures along this direction have relied on the availability of a very large number of diverse natural protein sequences sharing a common target function (the numbers of homologous sequences used ranged from 4,765 for malate dehydrogenases12 to 55,948 for lysozymes14. Moreover, sequences with desired enhancements were only passively identified instead of being actively designed18 (in exception to this, a recent work incorporated thermostability prediction in the sequence generation process to enrich the generation of sequences with enhanced thermostability17.
Recently, considerable progresses have been made in structure-based de novo protein design with machine learning models19, including models for the de novo generation of designable protein backbone structures20,21,22 and for selecting (from scratch) amino sequences that can fold into a given backbone structure23,24. The latter models, which are also referred to as inverse folding models, are especially relevant to the tasks of enhancing natural proteins through extensive sequence changes. As the inverse folding models aim for sequences that fit a structure optimally, they have been repeatedly shown to produce redesigned sequences that fold into the target structure with hyper-thermostability25. However, applying an existing inverse folding model to redesign natural proteins such as enzymes more often than not produces functionally inactive proteins5,6. One cause for this is that inverse folding models by themselves do not impose sequence restraints necessary for functions such as substrate recognition or chemical catalysis, leading some functionally critical residues to be incorrectly designed. A simple approach to resolve this issue is to predetermine (either through manual expert inspections or based on a multiple sequence alignment) a set of “functionally important” residues that are to be fixed during inverse folding. While a number of successful cases using this approach have been reported9,10,26, the sets of “functionally important” residues (usually chosen according to heuristic, system-specific rules) were often quite extensive, which could overly restrict the design space (and thus lead to suboptimum solutions).
We reasoned that achieving functional preservation in protein redesign requires improving the underlying inverse folding methodology. One important aspect of improvement is the accurate description of specific molecular interactions involving sidechains as well as other molecular entities, such as substrates of enzymes. When applied to enzymes, such a model may automatically select residues required for specific interactions with substrates, alleviating the need to fix an extensive, predetermined set of “functionally important” residues. Another aspect is the consideration of multiple conformational states. Optimizing the sequence on a single structure may impair the ability of the resulting protein to adopt alternative conformations and thus disable functionally essential conformational dynamics27. This could be an important cause for the loss of catalytic activity in sequences designed through inverse folding targeting a single fixed backbone.
Such challenges are further exacerbated in enzyme systems, where the complex interplay between sequence, structure, and activity makes it particularly challenging to identify which conformational states are essential for function. In such cases, evolutionary information derived from multiple sequence alignments (MSA) provides valuable functional constraints that are missing from structural data. Conversely, structural information offers detailed spatial context and precise residue-residue interactions that cannot be reliably inferred using conventional MSA-based approaches. By integrating structural components with MSA within a multimodal inverse folding framework, the complementary strengths of both data types could be effectively harnessed to enable more robust protein redesign that enhances structural stability while maintaining functional activity.
Driven by the above rationale, here we develop ABACUS-T (A Backbone based Amino aCid Usage Survey-ligand Targeted), a model that preforms inverse folding using denoising diffusion in the amino acid sequence space28,29 and unifies several important features into a single framework: detailed considerations of atomic sidechains and ligand interactions, a pre-trained protein language model30, multiple backbone conformational states, and evolutionary information from multiple sequence alignment. These features enable ABACUS-T to achieve more precise inverse folding than previous models. More importantly, they enable the generation of functionally active sequences without having to explicitly fix certain predetermined residues. To demonstrate the capability of ABACUS-T in preserving functional dynamics while enhancing protein stability, we apply it to redesign an allose binding protein31, aiming to maintain its ligand-induced conformational transition while significantly improving thermostability. To validate the effectiveness of integrating structural and MSA data, we redesign two enzymes, the endo-1,4-β-xylanase32 and the TEM β-lactamase33, to enhance their resistance to harsh conditions while preserving or even improving catalytic activity. Finally, we apply ABACUS-T to the OXA β-lactamase34 to rationally alter its substrate specificity while simultaneously enhancing protein stability. Remarkably, experimental validation requires testing only a few numbers of designs for each of these systems, the majority of which contain several dozens of simultaneously changed residues relative to wild type enzyme. This demonstrates the use of inverse folding with evolutionary input in protein engineering to bypass exhaustive experimental screening, achieving significant structural stability enhancements while maintaining or improving functional activity.
Results
An overview of ABACUS-T
As shown in Fig. 1A, ABACUS-T adopts a sequence-space denoising diffusion probabilistic model (DDPM), which uses successive reverse diffusion steps (indexed by an integer t that decreases from T to 0) to generate amino acid sequences from a “fully noised” starting sequence (noted as xT), in which the residue types at all positions are undetermined (i.e., masked). At a given step t, ABACUS-T first produces (based on the intermediately denoised sequence x(t) a temporary sequence \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\), in which the residue types at all positions are specified. Then the residue types at a fraction of positions in \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\) are re-masked to produce x(t − 1). The re-masked positions are chosen based on model-estimated likelihoods of residue types chosen in \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\). The fraction of masked positions decreases with decreasing t and goes from 100% at t = T to 0 at t = 1. To perform inverse folding, the above multi-step sequence denoising process is conditioned on an input that is comprised of a given protein backbone structure.
A Minimum input (a target protein backbone) and additional optional inputs (structure of bound ligand, multiple backbone structures, multiple sequence alignment) of ABACUS-T. B An amino acid sequence (node X0) is generated using successive denoising steps, starting from sequence XT in which the residues of all positions are unknown or masked. At step t, the partially masked intermediate sequence Xt is denoised into intermediate sequence Xt-1 with less masked positions. This is done by first decoding the residue types for all masked positions and then re-mask those positions associated with predictions of lower confidence. C Applying ABACUS-T to redesign the sequence of a natural protein can simultaneously change a large number of residues, achieving substantial enhancements in attributes such as structural stability or desired substrate selectivity while preserving attributes such as ligand-induced conformational transitions or high catalytic activity.
One feature of ABACUS-T that differs from most deep learning inverse folding models is that at each denoising step, both the residue types and the sidechain conformations are decoded. Moreover, each denoising step is self-conditioned35 with the output amino acid sequence (embedded with the pre-retained Evolutionary Scale Modelling or ESM sequence language model30 and sidechain atomic structures of the previous step. This scheme was found to be essential for the improved accuracy of ABACUS-T over previous inverse folding models (see results below).
To facilitate the generation of proteins that are not only foldable but also functionally active, the sequence generation by ABACUS-T can be further controlled by three types of optional input information in addition to a single given backbone (Fig. 1B). These optional inputs include the atomic structures of ligand molecules bound to the given backbone, multiple conformational states of the backbone, as well as a multiple sequence alignment (MSA) of the protein of interest. More details of the ABAUC-T model including implementation and training are given in Methods. The use of ABACUS-T with different inputs to carry out various sequence redesign tasks are illustrated by examples described later in Results (Fig. 1C).
Comparisons between ABACUS-T and ablated models
To examine the contributions of the various model features in ABACUS-T to performance, we compared the full model against 5 ablated models, which were obtained by applying the following changes to the full model increasingly and in a sequential order: replacing the larger 3 billion-parameter ESM model for sequence embedding by a smaller 60 million-parameter ESM model, leaving out the ESM sequence embedding from the self-conditioning scheme, leaving out self-conditioning in the sequence denoising steps, leaving out the modeling of small molecule ligands, and replacing the multi-step DDPM decoder with a network that projects the residue types at all positions in one single step. The different models have been trained on the same dataset with the same number of steps and compared on the same validation dataset (comprised of natural proteins with no more than 30% sequence identity to proteins in the training set).
In Fig. S1A, the native residue type recovery rates of the different models are compared. Besides the recovery rates for all residues, the recovery rates for pocket residues (determined as residues containing at least one atom within 5 Å from any ligand atom) and for catalytic residues (taken from the M-CSA database36, which contained such residues annotated by human experts) were compared as well. The results show that starting from the most ablated model that used a projection network for one-step residue type projection, the increasingly added model features monotonically improved performance. Notably, including the structures of ligands to condition the DDPM improved the median recovery rate of pocket residues from 0.56 to 0.62, verifying that the model can sensitively capture the contexts of bound ligand. The median recovery rate of pocket residues was further improved to 0.66 by introducing self-conditioning of the DDPM with decoded atomic structures of sidechains (FS1. S1A). Fig. S1B shows that this treatment also produced sequences with lowered ABACUS2 energies (indicating improved sidechain packing) relative to the sequences produced by DDPM without self-conditioning. These improvements represent the benefits of considering atomic structures of sidechains in inverse folding. The results in Figs. S1A and S1C show that incorporating the ESM language model further substantially improved the recovery rates of the pocket residues (median value from 0.66 to 0.76) and of the catalytic residues (median value from 0.8 and 1.0), which suggest that functionally important residues could be largely recovered by the final model. This ability of the final model is expected to be crucial for the generation of activity-preserving protein sequences without manually fixing a pre-determined set of residues.
Comparing ABACUS-T against other inverse folding models
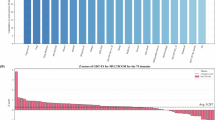
The results are summarized in Fig. 2A. To ensure fair comparisons, the benchmark dataset used here comprised PDB entries released after January 1, 2023 and with less than 30% sequence identity to any proteins released earlier (see PDB ID list in Supplementary Table 1). With this treatment, no benchmark protein or similar ones should have been included in the training sets of the methods being compared. The metrics compared include the native residue type recovery rates for all residues and for pocket residues, the Kullback–Leibler (KL) divergence between the residue type composition of the sequences designed by a method and the residue type composition derived from the native sequences, the root mean square deviations between the backbone structures predicted (with the ESMfold program30 for the designed sequences and the targeted backbone structures (the scRMSDs), and the predicted local distance difference test (pLDDT) scores of the ESMfold predictions, which reflect the physical plausibility of the structures predicted from the designed sequences. The results (Fig. 2A) indicate that while ABACUS-T performs comparably as LigandMPNN and CarbonDesign in the pLDDT and scRMSD metrics and as LM-design in the residue type composition metric, ABACUS-T outperforms the other methods by notable margins in the remaining comparisons. Especially, the high recovery rates of pocket residues combined with the small scRMSDs of ABACUS-T may critically advance its competence in generating functionally active sequences relative to previous methods (Fig. 2B). Besides natural protein backbones, we also compared the scRMSD distributions of sequences designed on a set of de novo backbones20 by different methods (Fig. S1D). The distributions for sequences generated by ABACUS-T spanned lower scRMSD ranges than those for sequences generated by the other methods.
A Performance of sequence design methods on 124 proteins released after Jan. 1, 2023. For each backbone and method, 10 sequences were generated; the one with the highest overall sequence recovery rate (relative to the native sequence) was analyzed. Kullback-Leibler (KL) divergence measures differences in residue composition between native and designed sequences. pLDDT scores reflect the predicted local structure quality using ESMFold. scRMSD is the root mean square deviation between predicted structure and target backbone. B Native residue recovery rates for ligand-binding pocket residues, grouped by ligand type. Boxplots show median, interquartile range, and min/max values excluding outliers ( >1.5 times the interquartile range beyond the box), n = 124. C Accuracy in predicting sidechain torsional angles \({\chi }_{1}\) and \({\chi }_{2}\) (within 20° error), reported separately for all residues and pocket residues. Boxplots show median, interquartile range, and min/max values excluding outliers ( >1.5 times the interquartile range beyond the box), n = 124. D Multi-state vs. single-state design on 140 proteins with multiple experimental conformations. Dynamic residues are those with >1.0 Å RMSD between at least two states. For multi-state design, 10 sequences per protein were generated; for single-state, 5 sequences per conformation. The sequence with the highest recovery rate was selected. TM-scoreensemble measures similarity between experimental and PVQD-predicted conformational ensembles of 20 structures. RMSF correlations are Pearson coefficients between experimental and predicted residue fluctuations. E Spearman correlations between zero-shot model predictions and Deep Mutational Scanning (DMS) results for 21 ProteinGym proteins. Source data are provided as a Source Data file.
We observe that the distinct framework used for inverse folding in ABACUS-T, as compared to previous methods, may lead to designed sequences that differ substantially from those generated by earlier deep learning-based inverse folding approaches. For instance, the average sequence identity between sequences designed by ABACUS-T and those designed by ABACUS-R is only 0.53 (Supplementary Table 2), which is lower than the average identity of 0.63 between ABACUS-T-designed sequences and native sequences.
We examined the frequencies of residue type substitutions between native sequences and ABACUS-T designed sequences for the 1049 natural proteins in our validation set. Residue type preferences in redesigned sequences were compared to those in wild-type sequences (Fig. S2A), with differences further categorized by solvent accessibility (Fig. S2B) and secondary structures (Fig. S2C). Substitution probabilities are summarized in Supplementary Table 3 according to structural context, and bidirectional mutation frequencies for each residue type are listed in Supplementary Table 4. Detailed substitution matrices related to solvent accessibility and secondary structure are shown in Figs. S2D and S2E, respectively.
Key trends emerged from this analysis include: substitutions occur more frequently at solvent-exposed sites than buried ones (Supplementary Table 3), and substitution rates are relatively uniform across secondary structure types (Supplementary Table 3). Exposed non-polar residues tend to be replaced by polar ones, while exposed polar residues often become charged (e.g., Lys or Glu). Ala and Leu are favored in helices, and Val in strands, likely enhancing sequence compatibility with backbone conformations. Mutation rates for Pro and Gly are low, underscoring their role in maintaining backbone geometry. Common substitutions such as Arg to Lys, Met to Leu, and Ile to Val may reflect a general tendency to replace fewer common residues with more prevalent ones sharing similar physicochemical properties.
While these trends are interpretable, they account for only a small fraction of all observed substitutions (e.g., Gln to Glu at exposed sites accounts for <2%). Ultimately, ABACUS-T substitutions are determined by a deep network that integrates complex structural contexts—factors that cannot be fully captured by simplified rules or patterns.
Interestingly, the ABACUS-T sequences yielded lower scRMSDs (according to ESMFold predictions) than the corresponding native sequences for more than 80% of target backbones (Fig. S1E). Moreover, when repacked on the target backbones, the ABACUS-T sequences yielded a lower average ABACUS2 energy per residue (−0.04 in arbitrary unit) than the native sequences (0.12 in arbitrary unit). On a benchmark of protein mutants with experimentally measured changes in the free energy of folding (ΔΔG)37, the changes in the pseudo-log-likelihoods predicted by ABACUS-T correlate with ΔΔG with a Spearman correlation coefficient of 0.54 (Fig. S1F), which is higher than the correlation coefficients obtained with several other methods including ABACUS2 (0.47), ABACUS-R (0.46), ESM-IF (0.46) and MIF38 (0.50) (see Supplementary Table 5).
The sequences redesigned with ABACUS-T also show large improvements over native sequences in total Rosetta energy. Supplementary Fig. S3A shows that 90% of the designed sequences exhibiting lower energies compared to their native counterparts, indicating improved structural stability. When individual energy components were compared, 98% of redesigned sequences exhibited more favorable van der Waals interactions (Fig. S3B). This improvement can be attributed to ABACUS-T’s all-atom modeling framework, which enhances sidechain packing. Additionally, 69% of redesigned sequences showed better electrostatic energies than the native sequences (Fig. S3C), consistent with the observed trends in environment-dependent residue substitutions.
No significant differences were observed in the frequency of Gly and Pro residues in loop regions (Gly: 0.086 vs. 0.082; Pro: 0.127 vs. 0.124 between redesigned and native sequences). Therefore, any potential changes in loop flexibility are not simply due to altered usage of these residues.
These results together support that ABACUS-T can be applied to redesign natural proteins to improve their compatibility with given backbone structures.
The improved accuracy of ABACUS-T should have come partially from the explicit consideration of sidechain conformations. To evaluate the accuracy of sidechain conformation modeling by ABACUS-T, we performed sidechain repacking of native sequences on natural backbones with ABACUS-T and compared the results against several previous methods including two energy function-based methods Rosetta39 and ABACUS240, and three recent deep-learning-based methods DiffPack41, DLPacker42 and AttnPacker43. Fig. 2C shows for different methods the distributions of the fraction of sidechains with correctly predicted torsional angles χ1 or χ2. To show the dependence of the prediction accuracy on the structural contexts of individual residues, the residues were roughly classified according to their relative solvent-accessible surface areas (RSA): core residues (RSA below 0.25), intermediately buried residues (RSA between 0.25 and 0.75), and exposed residues (RSA above 0.75). For residues in all three classes, ABACUS-T achieved substantially improved predictions of χ1 and χ2 relative to the other methods (see Fig. S1G).
Sequence design with inputs of multiple conformational states or a multiple sequence alignment
In order to design sequences that can adopt multiple conformations, we extended the basic ABACUS-T model to accept more than one backbone structures as input to control the sequence generation process. As described in Methods, we adjusted the model structure to handle the input of multiple backbone structures in different conformational states (multi-state design or MSD). The parameters of the resulting model, named ABACUS-TMSD, were fine-tuned from the basic model using a dataset of natural proteins with experimentally determined multiple conformational states.
To benchmark ABACUS-TMSD, we used a home-made set of 140 test proteins (see PDB ID list in Supplementary Table 6), each having multiple experimentally determined crystal structures with the largest RMSDs between different structures exceeding 1.0 Å. ABACUS-TMSD were compared against single-state designs with ABACUS-T as well as with ABACUS-TMSD on separate individual states. The results were also compared against single-state designs with ProteinMPNN23 and multi-state designs with ProteinMPNN-MSD44. The different methods were compared based on the native sidechain type recovery rates for all residues and separately for residues displaying large structural variations between the different conformational states (root mean square atomic displacements above 1.0 Å between at least two states. These residues are referred to as dynamic residues) (Fig. 2D). While the recovery rates for dynamic residues are lower than for all residues by all methods, the multi-state designs with ABACUS-TMSD or ProteinMPNN-MSD outperformed single-state designs with the corresponding base models. Moreover, ABACUS-TMSD showed substantial improvements over the other models (the improvements are more than 0.05 in absolute recovery rate for both all residues and dynamic residues).
We further evaluated the multi-state designs using structure predictions with a recently developed approach PVQD (Protein Vector Quantization and Diffusion)45. For each designed sequence, PVQD was applied to predict an ensemble of 20 conformations, from which we could compute a TM-scoreensemble (defined as the mean value of the highest TM-score of aligning each of the individual targeted backbone states against the ensemble of predicted conformations). We also determined for each designed sequence the Pearson correlation coefficient between the residue-wise RMSFs computed on the PVQD-predicted ensemble and those computed on the multiple experimental structures used as design targets. The multi-state designs exhibited clear improvements in these two metrics over the single-state designs, with multi-state design by ABACUS-TMSD outperformed the other models (Fig. 2D).
To further boost the ability of ABACUS-T in generating sequences that preserve function, we extended the model to incorporate protein family-specific evolution information contained in multiple sequence alignments (MSA) in two alternative models. One is named ABACUS-Tprior, in which the residue type distributions at individual positions are derived from the MSA and combined with the distributions derived from the structure-based residue type decoder with a user-specified weight (the prior weight, with a default value of 0.5). The other model, named as ABACUS-TMSAT, uses the MSA-transformer46 to produce a representation of the MSA for use as an additional input to condition the sequence generation. Results in Supplementary Table 7, show that incorporating the MSA information led to sequences closer to the native ones as expected: compared with base ABACUS-T model, the two models considering MSA substantially raised the native residue recovery rates for both all residues (from 63% to 70%) and pocket residues (from 75% to 85%), in doing so without distorting the overall residue type compositions (the KL-divergence from native compositions remained around 0.008). To examine if the combination of evolutionary and structural information in ABACUS-Tprior and ABACUS-TMSAT could in general promote the design of proteins with enhanced attributes, we benchmarked ABACUS-T, ABACUS-Tprior and ABACUS-TMSAT with the ProteinGym47 deep mutational scanning (DMS) datasets, which cover experimentally estimated mutational effects on diverse protein families and functions. We compared the results of our models against the results of four other methods (the MSA-transformer46, ESM2 (650 M)30, ProGen2 base15, and ProteinMPNN23 for 21 DMS datasets reporting biochemical functions47. All methods were under an unsupervised-learning (or zero-shot prediction) set up, which means no protein-specific experimental data was used to train the computational model. According to the Spearman correlation coefficients between the model predictions and the experimental results (shown in Fig. 2E), ABACUS-T showed large improvements over the previous inverse folding method ProteinMPNN, probably because of the various accuracy-improving features of ABACUS-T. ABACUS-T also outperformed the sequence-only models ESM2, ProGen2 and MSA-transformer. The additionally incorporation of MSA information in ABACUS-Tprior and ABACUS-TMSAT further improved the performance over ABACUS-T without using MSA.
Multi-state design on an allose binding protein
In this section, we present a case study of multi-state state design on the allose binding protein. The natural allose binding protein can exist in a ligand-free apo state and a ligand-bound holo state, both with available experimental structures (PDB IDs 1gud and 1rpj31, respectively). The structures exhibit a large inter-domain movement induced by allose binding, as exemplified by the change of the inter-Cα distance between E42 and D176 from 17.7 Å to 7.4 Å (see Fig. 3A) (Unless specified otherwise, residue numbers from the original PDB are used). We set the design goal as to obtain sequences with substantially enhanced structural stability while retaining the function of not only binding allose but also changing conformation from the open to the closed state upon allose binding.
A Left: a diagram of apo-to-holo conformational change. Right: PDB structure of the allose binding protein in apo and holo states. Distances between the Cα atoms of E42 and D176 are indicated. B Left: overall structure of holo state showing the distributions of residues of distinctive types (colored in salmon) in sequences designed on multiple conformational states relative to that on individual conformational states. The enlarged views show one of the spatial regions enriched with residues of distinctive types (shown as sticks). Right: Sequence logos for positions in this region computed from the sequence groups designed in different ways, colored by chemical properties (black: non-polar; magenta/green: polar neutral; red: negative; blue: positive. The native amino acid types are indicated above the logos. C Distributions of the values of the Pearson correlation coefficients of the RMSFs and of RMSDensemble computed from the native sequences and from sequences designed by different methods and/or in different ways. The boxplots show median, interquartile range, and minimum and maximum values excluding outliers ( >1.5 times the interquartile range beyond the box), n = 20. D Overall sequence recovery rates and KD measured by isothermal calorimetry titration (ITC) for binding to allose for native proteins and for seven proteins designed with ABACUS-TMSD. For each protein, the ITC experiment was conducted once, with the binding affinity constant and its error calculated by fitting the titration curve. E Left and middle: Superimpositions of the crystal structures (colored in blue) determined for designed proteins MSD1 and MSD3 with the corresponding structures of the natural allose binding protein (colored in gray). Right: enlarged views of the holo state structures showing the allose binding pockets with ligand and sidechains. F Thermal melting curves of the natural allose binding protein and the designed proteins MSD1 and MSD7 based on the circular dichroism (CD) signals at 222 nm. MRE: mean residue ellipticity. The Tm values are indicated. The ligand is shown as salmon sticks in all holo state structures. Source data are provided as a Source Data file.
We performed multi-state design with ABACUS-TMSD targeting simultaneously the apo and the holo states without fixing amino acid types for any residues. For comparisons, we also performed multi-state design with ProteinMPNN-MSD, as well as single-state design with either ABACUS-T (without fixing amino acid types for any residues) or ProteinMPNN based on either the apo or the holo state alone. The ligand allose was included in the holo state during designing with ABACUS-T or ABACUS-TMSD, while the residues forming the allose binding pocket were fixed during designing with ProteinMPNN or ProteinMPNN-MSD since these two models are unable to design sequences considering the ligand context.
As shown in Supplementary Table 8, the average native residue type recovery rate of the multi-state designs with ABACUS-TMSD was 74%. This mean rate decreased to 56% and 69% for ABACUS-T designs on the separated apo and holo states, respectively. For comparisons, this average rate was 53% for ProteinMPNN-MSD designs, which decreased further to 48% and 49% for ProteinMPNN designs on the separated apo and holo states, respectively. Comparing the sets of sequences generated by ABACUS-TMSD and by ABACUS-T revealed that the positions of clearly varied residue type choices were dispersed throughout the protein structures. Fig. S4 shows sequence positions where designs based on both the apo and holo states deviate significantly from those based on either state alone. These positions cluster spatially into four regions as indicated in the overall structure in Fig. S4, and the local structures of these regions in both the apo and the holo states are shown in details in the zoom-in views of Fig. 3B (for region 1) and Fig. S4 (for all regions) to illustrate the differences in the structural environments of these regions between the apo and the holo states. Some of the residue type choices preferred only in the single-state designs could be explained by the selective stabilization of a specific conformational state. For examples, the sequences designed on the holo state alone contained hydrophobic residues that stabilized the inter-domain packing (region 1) in the closed state but would disfavor the open state, while the sequences designed on the apo state alone contained a proline in a hinge loop (in region 2 shown in Fig. S4) which could increase the conformational rigidity of the loop. These features could prevent conformational change between the two states, and were absent from the multi-state designs.
We further predicted possible multiple conformations of the designed sequences with PVQD and computed the RMSDensemble (defined as the mean value of the lowest RMSD of aligning each of the individual targeted backbone states against the ensemble of predicted conformations) and the RMSF correlation coefficient metrics as described earlier. The results are shown and compared with those computed on the native sequence in Fig. 3C. Among the sequences designed with the various methods, only some designed with ABACUS-TMSD showed comparable metric values as the native sequence. The sequences designed with the other protocols produced inferior values, with the sequences designed on the apo state generally surpassing those designed on the holo state. This could be explained by that the holo state is structurally more compact than the apo state, leading the sequences designed on the holo state to have a deeper free energy minimum at the holo state, hindering conformational transitions to the apo state.
Based on the computational evaluations, we chose seven sequences designed on multiple states by ABACUS-TMSD (MSD1 to MSD7) for experimental characterization. For comparisons, we also experimentally characterized three sequences designed on the apo state (apoSD1 to apoSD3) and three sequences designed on the holo state (holoSD1 to holoSD3) with ABACUS-T, and four sequences designed with ProteinMPNN-MSD (with residues forming the allose binding pocket fixed). All designed proteins were successfully expressed, purified and confirmed to be in monomeric state by size exclusion chromatography (SEC) (see example results in Fig. S5). The binding constants to allose were measured using isothermal calorimetry titration (ITC) (Fig. 3D and S6). All the proteins designed with ABACUS-TMSD and those designed with ABACUS-T on the holo state exhibited binding affinities detectable by ITC. All the other sequences, including those designed with ProteinMPNN-MSD and pocket residues fixed to native types, showed no detectable binding affinity (Fig. S6). Interestingly, the KD values of the seven sequences produced by ABACUS-TMSD spanned a wide range corresponding to 0.01 to 17-fold changes in binding affinity relative to the natural allose binding protein (Fig. 3D). As the pocket residues are the same in the different ABACUS-TMSD designs, the variations in their affinity to allose were probably caused by shifts of the conformational equilibrium between the apo and the holo states rather than by changed molecular interactions with the ligand. Compared with the multi-state designs, the single-state designs based on the holo state structure would substantially shift the conformational equilibrium in favor of the holo state. Indeed, the three sequences designed on the holo state alone with ABACUS-T exhibit binding affinities 9 to 61-folds stronger than the natural protein (Fig. S6).
To verify that the ABACUS-TMSD designs can indeed change conformation from the apo to the holo state upon allose binding, we attempted to grow protein crystals for all designs at both the presence and the absence of allose. We obtained both apo and holo state crystals for MSD1 and MSD3 suitable for X-ray crystallography analysis and solved their structures at resolutions of 1.6 Å to 2.4 Å, respectively. The crystal structures of the apo and holo states matched with their respective design targets with backbone RMSDs from 0.59 to 1.67 Å (Fig. 3E). The all-atom RMSDs of the binding pocket residues are 0.38 Å and 0.43 Å for the holo state structures of MSD1 and MSD3, respectively, confirming the accuracy of sidechain-ligand interactions.
Temperature-dependent circular dichroism (CD) spectra were used to examine two of the designed proteins, MSD1 (one with solved crystal structures) and MSD7 (the one showing the highest affinity to allose) to confirm that their structural stability were indeed enhanced relative to the wild-type protein (as the generation of proteins with enhanced stability is a well-established property of backbone-based sequence design tools, the melting temperature analysis was performed only for these two representative designed proteins to balance cost and conclusion robustness). The results in Fig. 3F shows that the melting temperatures of both MSD1 and MSD7 (66 °C and 68 °C, respectively) are substantially higher than that of the wild-type protein (58 °C).
Designing active enzymes that resist adversarial conditions
Encouraged by the relatively high correlations between the ABACUS-Tprior and ABACUS-TMSAT scores with the DMS results including mutational effects on enzyme activity, we attempted to apply these approaches to redesign complete enzyme sequences without pre-fixing the residue type at any position. We considered as test cases two enzymes, endo-1,4-β-xylanase (xylanase in short, PDB: 4pn2) and TEM β-lactamase (PDB: 1jvj, Fig. 4A, B). The MSAs of these enzymes were constructed as described in Methods. The experimentally determined structures of these proteins in complexes with their small molecule substrates32,33 were used as the backbone targets. For comparisons, we also performed ABACUS-T designs without using MSA information.
A Left: A structure of the natural xylanase (PDB ID: 4pn2) shown with the chemical formulas of the substrate and product (salmon sticks in the shown structures) of the enzymatic reaction; Right: The summary of experimental results of designed xylanase. The three sets of designed sequences were compared for the number of mutated residues, the number of designs with detectable activity at 35 °C, the number of designs with enhanced activity at 35 °C and the number of designs exhibiting above 50% residual activity after 60 °C heat treatment for 10 min (the denominators indicate the total numbers of experimentally characterized designs). B The same as A, but for designs of TEM β-lactamase (PDB ID: 1jvj) and changed experimental parameters as indicated. C Left: The relative activity at 35 °C of wild-type (normalized to 1.0) and redesigned xylanases with different methods. The bars and dots represent mean values and three independent biological replicates, and the error bars represent the standard deviations (SD).; Middle: The residual activity of wild-type xylanase and pxyl3-0.5 after heat treatment at 80 °C for a range of time from 0 min (normalized to 1.0) to 60 min. The dots represent mean values of three independent biological replicates, and the error bars represent SD; Right: The thermal melting curves of pxyl3-0.5 (colored in blue), txyl4 (colored in red), and native xylanase (colored in gray) according to CD signals at 222 nm. MRE: mean residue ellipticity. The values of Tm are indicated. D The same as (C) but Left: the relative activity at 37 °C of wild-type (normalized to 1.0) and redesigned lactamases with different methods; Middle: The residual activity of wild-type TEM β-lactamase, tlact2, plact2-0.2, plact2-0.5 after heat treatment at 50 °C for a range of time from 0 min (normalized to 1.0) to 60 min; Right: The thermal melting curves of wild-type TEM β-lactamase, tlact2, plact2-0.2, plact2-0.5 according to CD signals at 222 nm. Source data are provided as a Source Data file.
For xylanase, we selected in total 12 designs that are most different from each other within each group of sequences designed with a given method for experimental characterization. The sequences include pxyl1-0.5 to pxyl5-0.5 designed with ABACUS-Tprior, mxyl1 to mxyl3 designed with ABACUS-TMSAT, and txyl1 to txyl4 designed with ABACUS-T (see Supplementary Table 9 for detailed mutation rates of these variants). All these sequences have recovered the native residue types for all residues at the active site, while the total numbers of changed residues compared with the corresponding natural enzymes ranged from 56 to 80 (Fig. 4A). All designed proteins could be expressed and purified in soluble monomeric forms (see example SEC results in Fig. S5).
The hydrolytic activity on the xylo-oligosaccharide substrate of the purified proteins were measured as described in Methods. While all tested designs exhibited detectable activity, the designs generated with ABACUS-Tprior and ABACUS-TMSAT showed higher activity than those generated with ABACUS-T (Fig. 4A, C and Supplementary Table 9). Two designs, pxyl3-0.5 (generated by ABACUS-Tprior) and mxyl1 (generated by ABACUS-TMSAT) exhibited catalytic activities comparable to that of the natural enzyme. For comparisons, we also experimentally tested the catalytic activity of two sequences designed with ProteinMPNN based on the xylanase backbone structure (during the design, we fixed the sidechain types for 12 residues that were heuristically determined to be required for substrate recognition and catalysis, see Fig. S7A). Purified proteins of these two designs did not show detectable catalytic activity (Fig. S7A).
We compared the designed proteins against the natural protein in their resistance to treatments at raised temperatures. Fig. S8A shows that while the natural protein lost 86%/99.9% activity after 10 min treatment at 40 °C/60 °C, the redesigned proteins retained 55%/2% to 100%/100% activity after the same treatments (the only exception is mxyl3 generated with ABACUS-TMSAT, which lost activity after the treatments at 40 °C or 60 °C). The designed pxyl3-0.5, which is of comparable activity to the natural protein, showed the strongest resistance to high temperature treatment, maintaining over 50% residual activities after treatment at 80 °C for 50 min (Fig. 4C). The thermal melting curves in Fig. 4C indicates that the Tm has increased from 41 °C of the natural proteins to 60 °C of pxyl3-0.5. Moreover, the CD spectra in Fig. S9A showed that while the thermal denaturation of the natural protein is irreversible, the thermal denaturation of pxyl3-0.5 is fully reversible, being able to recover 100% activity after thermal unfolding-refolding. We note that the effects on structural stability of some individual residue changes from the sequence of the natural xylanase to the sequence of pxyl3-0.5 could be rationalized (Fig. S9B), including R108P, which may increase the backbone rigidity of a loop, K184R, which may introduce polar interactions, and E222A, which may improve hydrophobic packing. However, as the total number of replaced residues from wild-type sequence to pxyl3-0.5 is 63, the considerable overall stability enhancement may not be easily attained by step-wise accumulation of individual mutations.
We further examined the resistance of pxyl3-0.5 to other adversarial conditions including extreme pH and high concentrations of denaturant (guanidine hydrochloride or GdnHCl), organic solvent (ethanol), or salt (NaCl). The results (Fig. S10) indicate substantial enhancements of the designed protein in the resistance to these conditions relative to the natural proteins.
For the TEM β-lactamase, we designed sequences with ABACUS-Tprior, ABACUS-TMSAT or ABACUS-T based on an experimentally determined structure of the enzyme-substrate complex. We found that with the prior weight parameter of the MSA distribution setting to 0.5, ABACUS-Tprior generated sequences containing only 12 residues relative to the natural protein. Thus, we also considered sequences designed with the prior weight reduced to 0.2 in ABACUS-Tprior. The set of 11 experimentally tested sequences included plact1-0.5 and plact2-0.5 (designed with ABACUS-Tprior with prior weight 0.5), plact1-0.2 and plact2-0.2 (designed with ABACUS-Tprior with prior weight 0.2), mlact1 to mlact4 (designed with ABACUS-TMSAT), and tlact1 to tlact3 (designed with ABACUS-T) (See Supplementary Table 9 for detailed mutation rates of these variants). All proteins could be expressed and purified in their soluble, monomeric forms (see example SEC results in Fig. S5). We measured the hydrolytic activity on ampicillin of each purified protein as described in Methods and compared the results with those of the natural protein. Figur. 4B shows that 10 out of 11 designs exhibited detectable catalytic activity. Interestingly, the two designs generated by ABACUS-Tprior with prior weight 0.5 showed more than two times higher activity than the natural protein, while the two designs from ABACUS-Tprior with prior weight 0.2 showed activity comparable to the natural protein. The designs generated with ABACUS-T or ABACUS-TMSAT were of lower activity than the natural protein (Fig. 4D). For comparisons, we also experimentally tested the catalytic activity of two sequences designed with ProteinMPNN based on the TEM β-lactamase backbone structure (during the design, we fixed the sidechain types for 7 residues that were heuristically determined to be required for substrate recognition and catalysis, see Fig. S7B). Purified proteins of these two designs did not show detectable catalytic activity (Fig. S7B).
The resistance to treatments at raised temperatures of the designed proteins was compared against the natural protein in Fig. 4D and Fig. S8B. All the sequences designed with ABACUS-Tprior and ABACUS-T showed enhanced resistance relative to the natural protein, while the sequences designed with ABACUS-TMSAT showed weakened resistance (Fig. S8B). Interestingly, the two sequences designed with ABACUS-Tprior at prior weight 0.2 appeared to resist treatments at raised temperature better than the two sequences designed with ABACUS-Tprior at prior weight 0.5 (Fig. S8B). However, the latter two sequences exhibited higher absolute activities than the former sequences at the room temperature (Fig. 4D). These results suggest that the prior weight parameter, which grossly determines the weight of evolution relative to structural information in the ABACUS-Tprior model, implicitly controlled the tradeoff between retaining catalytic activity and enhancing structural stability. To examine this further, we measured the residual activities of the natural protein and the designed ones plact2-0.5, plact2-0.2 and tlact2 after treatment at 50 °C for varied time periods. The results in Fig. 4D show that plact2-0.2 (designed with prior weight 0.2) and tlact2 (which could be viewed as having been designed with prior weight 0.0) both retained around 75% activity after 60 min treatment, while plact2-0.5 (designed with prior weight 0.2) lost 60% activity after 20 min treatment. The melting curves determined from CD spectra (Fig. 4D) showed that the Tm of the proteins increased from 49 °C of the natural protein to 53 °C of plact2-0.5, 61 °C of plact2-0.2, and 72 °C of tlact2. The ΔTm results for the designed constructs show varying degrees of improvement compared to the wild-type. Specifically, the Tm of plact2-0.5 (53 °C) is not significantly higher than that of the native protein (49 °C). In contrast, the Tm values of plact2-0.2 (61 °C) and tlact2 (72 °C) are markedly higher than the native Tm value, indicating substantial improvements in thermal stability. The relatively modest improvement in plact2-0.5, which was designed with a greater emphasis on evolutionary constraints (higher MSA weight), highlights the trade-off between adhering to natural sequence preferences and optimizing structural stability in the design process.
Notably, through comparing the sequence profiles generated by raw ABACUS-T without evolutionary guidance and those derived using MSA-transformer for the xylanase and the TEM β-lactamase (Fig. S11), we found that most of residues showing changed design choices are surface-exposed and located far from the substrate-binding pockets. This suggests that evolutionary constraints related to direct interactions with substrates or catalytic residues are already captured by the raw ABACUS-T model, which explicitly models protein-substrate interactions. The incorporation of MSA data primarily adds evolutionary constraints that cannot be attributed to these direct interactions, likely influencing properties such as protein dynamics or other indirect functional aspects.
Designing enhanced enzymes with changed substrate selectivity
A common objective of enzyme engineering is to change the substrate spectrum. ABACUS-T can be employed to carry out such a task by conditioning its sequence generation process with different ligands in complex with the protein backbone. We demonstrated this by redesigning the substrate selectivity of a broad-spectrum OXA β-lactamase34, which is known to hydrolyze three β-lactam substrates of distinct molecular structures: cefotaxime, ceftazidime and imipenem (Fig. 5A). Starting from a structure of the natural OXA β-lactamase in complex with cefotaxime (PDB ID: 5wi3), we docked the other two substrates ceftazidime and imipenem into the 5wi3 backbone by superimposing the shared lactam motif (highlighted in Fig. 5A) (the used atomic structures of ceftazidime and imipenem were taken from two other PDB structures 4 × 56 and 7t7g, respectively. The thiane/aniline moiety in ceftazidime and the ester moiety in cefotaxime is not solved in the corresponding crystal structures. Protein-ligand modeling indicate that the aniline moiety of ceftazidime and the ester moiety of cefotaxime are located in solvent-exposed regions, with no direct contacts to binding pocket residues (minimum distance > 4.5 Å). These moieties were thus omitted for the ligand structures during sequence design). We then used ABACUS-Tprior (with prior weight 0.2) to redesign sequences on each of the three backbone-substrate complexes. In the first round of design, we only aim to change the substrate selectivity by allowing only the first-shell residues surrounding the substrate to change while fixing the rest of the protein to native residue types.
A Left: Structures of three β-lactam substrates (cefotaxime, ceftazidime and imipenem) with the β-lactam ring in red; actually modeled structures are boxed in blue. Middle: Sequence logos of redesigned active site residues for each substrate, colored by chemical properties (black: non-polar; magenta/green: polar neutral; red: negative; blue: positive). Native (blue) and top-performing designed (grey) residue types are labeled above. Right: Enlarged views of redesigned active sites, with pocket residues and substrates shown as sticks; substrate carbon skeletons are salmon-colored. B Hydrolytic activities (normalized to wild-type enzyme) of seven designs per substrate. Dots show means of three independent biological replicates; bar heights indicate averages; error bars represent standard deviation (SD). Statistical significance was assessed by two-sample two-sided t-tests (assuming equal variance): cefotaxime, t12 = 1.99, p = 0.0693, Cohen’s d = 1.07, 95% CI [ − 0.022, 0.498] and t12 = 1.23, p = 0.242, d = 0.658, CI[−0.120,0.429]; ceftazidime, t12 = 5.45, p = 0.000147, d = 2.91, CI [4.65, 10.8] and t12 = 5.12, p = 0.000253, d = 2.74, CI [4.22, 10.5]; imipenem, t12 = 2.44, p = 0.031, d = 1.31, CI [0.032, 0.557] and t12 = 2.76, p = 0.0172, d = 1.44, CI [0.070, 0.592]. Significance: N.S. p ≥ 0.1; * p < 0.1; ** p < 0.05; *** p < 0.01. C Relative activities (normalized to wild-type enzyme) of overall sequence-redesigned enzymes (three sets) against each substrate, shown as average ± SD of three independent biological replicates. D Residual activity of native and overall sequence-redesigned enzymes after incubation at indicated temperatures, normalized to activity at 37 °C. Dot and error bar indicate respectively average and SD of three independent biological replicates. E Thermal melting curves monitored by CD at 222 nm for native enzyme, cefo3-0.2, cefta2-0.2, imip2-0.2. MRE: mean residue ellipticity. The values of Tm are indicated. Source data are provided as a Source Data file.
Fig. 5 A shows that the designed sequences exhibited substrate-specific sidechain type profiles for the pocket residues. Some specifically preferred sidechain types may enhance hydrophobic packing with the corresponding substrate (e.g., Y167 with cefotaxime). Some other preferences may reduce steric strains with the corresponding substrate (e.g., G221 with ceftazidime and L110 with imipenem). Other preferences may increase the polarity of solvent-accessible sidechains in the corresponding enzyme-substrate complexes (e.g., E109, R109 and R221 in the cefotaxime, ceftazidime, and imipenem complexes, respectively).
Based on the computational analysis, we chose seven sequences designed for each of three substrates for experimental characterization, including cefo1 to cefo7 designed for cefotaxime, cefta1 to cefta7 designed for ceftazidime, and imip1 to imip7 designed for imipenem. All 21 proteins were successfully expressed and purified in soluble, monomeric forms (see SEC results in Fig. S5). The hydrolytic activity on the three substrates of each purified protein were measured as described in Methods. Figure. 5B and S12A showed the three sets of sequences on average exhibited the intended substrate selectivity.
We then performed a second round of design with ABACUS-T to examine whether the substrate selective enzymes could be enhanced in their thermostability through sequence redesign on the rest of the protein. In this design round, we fixed the pocket residues to be of the same type as those in the particularly high activity variants from the first round (cefo2 for cefotaxime, cefta7 for ceftazidime, and imip2 for imipenem), and allowed all the other residues to be redesigned. Three sequences were designed for each substrate with ABACUS-Tprior at prior weight 0.2 (named as cefo1-0.2 to cefo3-0.2, cefta1-0.2 to cefta3-0.2 and imip1-0.2 to imip3-0.2). These proteins exhibited similar substrate preferences as their corresponding precursor proteins (Fig. 5C).
We examined the most active redesigned proteins for their resistance to treatments (for 5 min) at temperatures from 37 °C to 70 °C by measuring their residual activities on the corresponding target substrates (for the natural protein, the substrate used was cefotaxime). The results in Fig. 5D and S12B shows that while the natural protein retained only 15% of activity after treatment for 5 min at 50 °C, all the tested designed proteins retained over 60% of residual activity after 5 min treatment at 70 °C. The thermal melting curves from CD measurements show that while the native protein unfolded with a Tm of around 52 °C, the redesigned proteins showed hyper-thermostability and did not unfold until a very high temperature (Tm above 90 °C, see Fig. 5E).
Rosetta energy analysis of experimentally validated redesigned proteins
To rationalize the mechanisms underlying the improved thermostability of the proteins redesigned using ABACUS-T, we analyzed the Rosetta energies of 10 redesigned proteins that were experimentally characterized to show varying increases in their unfolding temperatures compared to their native counterparts. The results are summarized in Fig. S13A. For most of the re-designed proteins (9 out of 10), their total Rosetta energies were lower than those of their native counterparts (ΔEtotal < 0), consistent with the observed enhancement in thermostability. The only exception was plact2-0.5, which showed a small positive ΔEtotal. This protein also exhibited a relatively modest increase in its unfolding temperature (4 °C, compared to 12 °C for plact2-0.2 and 23 °C for tlact2).
To further investigate whether the changes in total energy (ΔEtotal) were primarily driven by a few key inter-residue interactions or by more broadly distributed interactions, we calculated residue-wise energy changes (ΔEresidue) and compared the contributions of residues within the 5%−95% range of the ΔEresidue distribution to those outside this range. The results in Fig. S13A show that residues within the 5%-95% range generally contributed significantly, particularly in redesigned proteins with substantial increases in unfolding temperatures. However, key mutations can also play a critical role, depending on the specific structural and sequence context of the protein. In some cases, these mutations may contribute equally or even more than the broader residue network. Fig. S13B shows the local structural details of selected residues with large negative ΔEresidue values, illustrating possible improvements in local interactions.
The energy calculations thus suggest that the enhanced thermostability observed in ABACUS-T redesigned sequences arises from a synergistic combination of key mutations and extensive sequence changes. This underscores the effectiveness of the deep learning-based redesign approach in balancing local and global energetic contributions to achieve improved protein stability.
The results presented in Supplementary Table 5 further highlight the practical distinctions between deep learning-based inverse folding models and traditional energy-based models. This table compares the correlations between scores computed by various methods and the experimentally determined effects on changes in the free energy of folding (ΔΔG) for a dataset comprised mainly of single mutations. As shown by data in this table, deep learning models, including ABACUS-T, do not surpass traditional models in predicting the effects of single mutations. However, the experimental results of this and other studies39 suggest that the probability of successful predictions by deep learning models does not exhibit exponential decay as deviations from native sequences increase, which is distinct from traditional methods. Thus the strength of deep learning approaches lies in their ability to design sequences with multiple mutations or to redesign entire sequences de novo.
Discussion
While holding considerable potential for protein engineering, previous inverse folding models often fall short of producing functionally active proteins. There are several possible causes. Besides the obvious cause that inverse folding does not impose the preserving of residues obligatory for functions, other possible causes include insufficient accuracy of inverse folding and impaired conformational dynamics by choosing to stabilize a single target structure. ABACUS-T performs inverse folding using a self-conditioned sequence denoising diffusion network, with the language model-embedded sequence representation and atomic sidechain structures used for self-conditioning. ABACUS-T have also been devised to consider multimodal inputs, including the structures of bound ligands, multiple conformational states, and MSA of natural proteins, which are expected to further promote the generation of functionally active sequences. With the sequence language models capturing sequence statistics and global dependencies, the atomic sidechain representations refining local interactions, and the multiple backbone conformations enhancing robustness by accounting for protein dynamics, we expect this integration to provide a more comprehensive representation of protein structural features and can improve generalization of inverse folding models beyond training examples. Potential trade-offs of this integration include increased model complexity, compromises in stability when balancing different modalities, and data limitations, as structures of inter-molecular complexes and diverse backbone states are not always available.
The above reasoning suggests two main factors to guide the choice of ABACUS-T variant: the availability of input data and the specific design goals for the target protein. While standard ABACUS-T works for backbones lacking evolutionary constraints or conformational ensembles, ABACUS-Tprior (Evolutionary Constraint-Integrated) should be preferred for functional proteins requiring evolutionary conservation, especially enzymes. The variant ABACUS-TMSD (Multi-State Design) is essential for conformationally dynamic targets with multiple resolved or predicted structures, such as allosteric regulators. The flexibility of ABACUS-T variants allows users to tailor their pipelines based on specific goals, whether prioritizing stability, functionality, or multi-state behavior.
Our computational benchmarks indicated that the accuracy for inverse folding has been substantially improved by ABACUS-T over a number of previous models. Computationally benchmarking on the ProteinGym DMS datasets have shown that ABACUS-T augmented with MSA outperformed a range of structure-based or none-structure-based models for the zero-shot prediction of mutational effects of protein attributes. These results inspired us to apply ABACUS-T to the challenging task of engineering functionally active proteins with sizeable enhancements in structural stability-related attributes by designing many simultaneously mutated residues in one step. In each our test cases, this design goal has been achieved by experimentally testing only a few designs, the majority of which contain several dozens of changed residues compared with wild-type protein. The OXA β-lactamase presented an example of designing for altering substrate specificity with ABACUS-T.
We note that some aspects of ABACUS-T have been considered in different recently reported inverse folding models. These include the use of pretrained protein language models48 and the explicit modeling of sidechain conformation49, which were found to improve the accuracy of inverse folding. Interactions with ligands50 and designing for multiple conformational states44 have also been considered but in separate models. Not considering inverse folding, a recent study investigated the sampling of sequence variants guided solely by evolutionary coupling analyses on MSA18. In ABACUS-T, a unified framework was devised to integrate the different modalities. Our computational benchmarks confirmed the improved accuracy of ABACUS-T over previous models. More importantly, our experimental results demonstrate that these improvements and the multimodal capability of ABUCAS-T lead to the efficient generation of functionally active proteins including enzymes and allosteric ligand-binding proteins, which has been a long-standing challenge for inverse folding models.
We acknowledge that ABACUS-T still relies on high quality structures of the holo state for designing active proteins. We emphasize that the complexity of enzymes with multiple conformational states poses a significant challenge for protein design. While ABACUS-T enables sequence design based on multiple conformations, this represents only the initial step in tackling this issue. It remains challenging to determine experimentally or to model computationally the catalytically critical conformational states for enzymes of multiple conformational states. Multi-conformation-based sequence design combined with experimental validation of enzyme activity can play important roles in driving progresses of this direction. In this study, evolutionary information extracted from MSA is used as a practical alternative to tackle the complexity associated with factors such as conformational dynamics. This information serves as input for ABACUS-T, enabling the generation of sequences with high activity levels. Although some of the designed sequences reported here demonstrated higher catalytic efficiency than their wild-type counterparts, along with significantly improved structural stability, it is important to note that the design process itself does not yet actively target enhancements in catalytic efficiency. Moreover, for the enzymes tested here, improvements in inverse folding achieved by ABACUS-T in the absence of evolutionary information are insufficient for designing extensive stabilizing mutations without compromising catalytic activity.
While the catalytic efficiency of some designs has been improved, the gains remain modest compared to increases in thermodynamic stability. This likely arises from two key factors: first, the single state structure input misses dynamic conformational changes essential for catalysis; second, the evolutionary conservations from MSA could also emphasize stability over activity. Possible improvements on this aspect include incorporating multi-state designs for both substrate-free and substrate-bound conformations, and using activity-aware scoring functions trained on experimental data to prioritize designs with enhanced activity alongside stability.
In summary, ABACUS-T has the following limitations. As an inverse folding method, it requires a predefined backbone structure, and the sequence novelty is limited by the structural novelty of the input. Although the fact that ABACUS-T is trained on natural protein structure is not expected to prevent it from generating sequences compatible with backbones, the resulting sequences may not support the desired function, especially when functional requirements are not fully captured by structural constraints. Although the method can incorporate evolutionary information and account for ligand atoms and multiple conformations to enhance functional compatibility, it does not explicitly model functional activity during design. This limits its ability to consistently generate fully functional sequences.
One advantage of repurposing structure-based de novo protein design models for protein engineering is that these models do not need supervised training with extensive, system-specific experimental data. Moreover, the ability of ABACUS-T to design simultaneous mutations of many residues in one step to achieve sizeable enhancements of structure-related attributes represents a notable progress over previous methods. Therefore, methods like ABACUS-T are promising in various biotechnological applications.
Methods
Sequence generation based on denoising diffusion in ABACUS-T
As shown in Fig. 1A, sequence generation in ABACUS-T is based on modeling a discrete space diffusion process in reverse28,29,51. Here, the amino acid sequence is abstracted as a sequence of N tokens, \({\left\{{{{\boldsymbol{x}}}_{0}}^{n}\right\}}_{n=1}^{N}\), which follows the distribution pdata. Each token in the sequence takes one of K possible states including an absorbing state (i.e., [MASK]), which is introduced as a way to consider noise in the sequence of discrete data. In the forward diffusion process, each token of the sequence transits independently from its initial state x0 into its final state xT in a total number of T steps with a Markov transition matrix \({\left[{{\boldsymbol{Q}}}_{t}\right]}_{{ij}}=q\left({x}_{t}=j|{x}_{t-1}=i\right)\) at step t. This leads to the categorical conditional distribution q(xt|xt−1) = Cat(xt; p = xt−1Qt), with the series of states represented by one-hot row vectors \({{\boldsymbol{x}}}_{1},\ldots,{{\boldsymbol{x}}}_{T}{\boldsymbol{\in }}{\left\{\mathrm{0,1}\right\}}^{K}\). Following Austin et al.28, we considered the absorbing state transition matrix \({\left[{{\boldsymbol{Q}}}_{t}\right]}_{{ij}}\), which is defined as that the token retains its current state with a probability of \(1-{{\beta }}_{t}\) while transits into the38 state with a probability of \({{\beta }}_{t}\). The noise schedule \({{\beta }}_{t}\) was chosen so that the probability of the [MASK] state increases linearly from 0 to 1 with t from 0 to T. To determine the likelihood of the observed data \({{\boldsymbol{x}}}_{0}\), a reverse model pθ(xt−1|xt) with the learnable parameter set θ was used to predict the probability of state \({{\boldsymbol{x}}}_{t-1}\) of each token given the states of all tokens \({\left\{{{{\boldsymbol{x}}}_{t}}^{n}\right\}}_{n=1}^{N}\) at step t. In practice, this was achieved by learning a model \({p}_{{\rm{\theta }}}\left({\widetilde{{\boldsymbol{x}}}}_{0}|{{\boldsymbol{x}}}_{t}\right)\) and then applying the relation \({{p}_{{\rm{\theta }}}\left({{\boldsymbol{x}}}_{t-1}|{{\boldsymbol{x}}}_{t}\right)=\sum _{{\widetilde{{\boldsymbol{x}}}}_{0}}q\left({{\boldsymbol{x}}}_{t-1}|{\widetilde{{\boldsymbol{x}}}}_{0}\right)p}_{{\rm{\theta }}}\left({\widetilde{{\boldsymbol{x}}}}_{0}|{{\boldsymbol{x}}}_{t}\right)\). The objective for learning \({p}_{{\rm{\theta }}}\left({{\boldsymbol{x}}}_{t-1}|{{\boldsymbol{x}}}_{t}\right)\) can be set as to maximize the evidence lower bound (ELBO) of the log likelihood of the training data29. It has been shown that this objective can be substituted by minimizing the following loss function,
in which \({{\lambda }}_{t-1}=\frac{{{\rm{\alpha }}}_{t-1}-{{\rm{\alpha }}}_{t}}{1-{{\rm{\alpha }}}_{t}},{{\alpha }}_{t}=\mathop{\prod }\limits_{i=1}^{t}{{\rm{\beta }}}_{i}\) represents the weight of the loss of step t, \({\text{f}}_{{\rm{\theta }}}^{n}\left({{\boldsymbol{x}}}_{t}\right)\) represents the model that predicts the categorical distribution of the nth token conditioned on the partially masked sequence \({{\boldsymbol{x}}}_{t}\) at time t.
With the trained denoising model, sequences can be generated by iteratively applying the following two steps, starting from sampling a sequence of all tokens in the [MASK] state,
-
1.
generating a sequence of N tokens at time step t, with a temperature \(\tau\), namely, sample each \({{\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)}^{n}\) from \({\text{Cat}}({{\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)}^{n}{;}p\propto l{og}{\text{f}}_{{\rm{\theta }}}^{n}\left({{\boldsymbol{x}}}_{t}\right)/\tau )\);
-
2.
predicting the partially masked sequence \({\left\{{{{\boldsymbol{x}}}_{t-1}}^{n}\right\}}_{n=1}^{N}\) by predicting \({{\boldsymbol{x}}}_{t-1}\) based on \({{\boldsymbol{x}}}_{t}\) and \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\) for each token.
In ABACUS-T, sequence diffusion is performed under a given protein backbone. For simplicity, the training weight \({{\lambda }}_{t-1}\) was set to a constant independent of t, meaning denoising losses on samples containing different factions of masked positions were weighed equally. During inference, a greedy scheme was employed to sample \({p}_{{\rm{\theta }}}\left({{\boldsymbol{x}}}_{t-1}|{{\boldsymbol{x}}}_{t}\right)\), meaning that residues with the lowest confidence in \({\left\{{{\widetilde{{\boldsymbol{x}}}}_{0}}^{n}\right\}}_{n=1}^{N}\) (namely, of the smallest likelihoods \({\text{f}}_{{\rm{\theta }}}^{n}\left({{\boldsymbol{x}}}_{t}\right)\)) were re-masked and refined in the next step, with the fraction of re-masked residues decreases linearly with t.
The architecture of the ABACUS-T network
As illustrated in Fig. S14, the overall network comprises an encoder module, a decoder module, and an iterator module. The encoder module represents the structural contexts as the conditions for sequence generation, the iterator module implements the denoising diffusion iterations to generate iteratively refined sequences, while the decoder module predicts the residue types (and sidechain conformations) at masked positions.
The encoder module consists of a backbone encoder block, a ligand encoder block, and a multi-modal block.
The backbone encoder block consists of repeated sub-blocks of graph neural networks (GNN) processing residue-wise node features and residue pair-wise edge features. The initial residue-wise features include the scalar values \(\left\{\phi,\varphi,\omega \right\}\) referring to dihedral angles computed from the atoms \({C}_{i-1},{N}_{i},{C}_{{{\rm{\alpha }}}_{i}}\) and \({N}_{i+1}\) for residue i as well as a unit vector representing the direction of \({C}_{{{\beta }}_{i}}{-C}_{{{\rm{\alpha }}}_{i}}\) (computed as \(\sqrt{\frac{1}{3}}\left({\boldsymbol{n}}\times {\boldsymbol{c}}\right)/{{||}{\boldsymbol{n}}\times {\boldsymbol{c}}{||}}_{2}-\sqrt{\frac{1}{3}}\left({\boldsymbol{n}}+{\boldsymbol{c}}\right)/{{||}{\boldsymbol{n}}+{\boldsymbol{c}}{||}}_{2}\), with \({\boldsymbol{n}}={N}_{i}{-C}_{{{\alpha }}_{i}}\) and \({\boldsymbol{c}}={C}_{i}{-C}_{{{\alpha }}_{i}}\)) (in our formulations, atom names denote their vector coordinates in the three-dimensional space). The initial residue pair-wise features include the Gaussian radial basis function encodings of the inter-residue atomic distances involving the five backbone atoms of each residue (\({C}_{i-1},{N}_{i},{C}_{{{\alpha }}_{i}}\) and \({N}_{i+1}\)), the unit vectors representing the cross-residue directions \({C}_{{{\alpha }}_{i}}-{C}_{{{\alpha }}_{j}}\) and \({C}_{{{\alpha }}_{i}}-{C}_{{{\alpha }}_{j}}\), as well as a quaternion representing the rotation between the three-atom rigid bodies \({N}_{i}-{C}_{{{\alpha }}_{i}}-{C}_{i}\) and \({N}_{j}-{C}_{{{\alpha }}_{j}}-{C}_{j}\). The node features from the last layer of the backbone encoder block were used as residue-wise representations of the protein backbone.
The ligand encoder block was built upon the SchNet network52 and the pre-trained Uni-Mol network53, both processing the chemical structures of a ligand molecule through atomic and atom pair-wise features. The atomic features include atom type, implicit valence number, charge state, hybridization state, aromaticity, and whether an atom belongs to a ring structure. The atom pair-wise features include covalent atomic bond types and inter-atomic distances (encoded with Gaussian radial basis function). The outputs of the SchNet with trainable parameters and the Uni-Mol network with fixed parameters were concatenated to form atom-wise representations of the ligand molecule.
The residue-wise backbone representations and the atom-wise ligand representations are integrated by the multi-modal encoder block, which is a GNN with nodes corresponding to protein residues plus ligand atoms. The initial node features were provided by the backbone encoder and the ligand encoder, while the edge features include encodings of the type of the edge (protein-protein, protein ligand, or ligand-ligand) and the Gaussian basis function-encoded inter-atomic distances from a ligand atom to the backbone atoms of a protein residue.
The iterator module processes the partially masked sequence xt and the temporarily decoded sequence \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\) (with predicted sidechain conformations) by using a residue type embedder block, an overall sequence encoder block and a sidechain structure encoder block. The residue type embedder block processes the sidechain type states in xt, the overall sequence encoder block uses a pre-trained protein language model to encode \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t\right)\), while the sidechain structure encoder block processes the distances from the sidechain atoms to the backbone and the ligand atoms. The outputs of these blocks are added with the backbone and ligand structure information processed by the encoder module as input to the decoder module, with the contribution of the temporarily decoded sequence and sidechain structures serve the purpose of self-conditioning at each denoising diffusion step.
The decoder module contains three blocks: a protein representation updating block, a residue type projection block and a sidechain conformation decoder block. The protein representation updating block uses the merged representations from the overall encoder and the iterator modules as input. It updates the residue-wise node data using a massage passing neural network without updating the edge data. The updated node data from the last layer of the message passing network are passed to the other two blocks of this module: the residue type projection block that predicts the residue types for the masked residues, and the sidechain conformation decoder block that decode the sidechain torsional angles of the re-predicted residue types. The torsional angles χ1, χ2, χ3 and χ4 for each residue were auto-regressively predicted as discretized classes in 10-degree intervals.
To train the ABACUS-T network, we built a database of protein structures including bound ligands from PDB by first selecting all crystal structures containing proteins with resolution below 4.0 Å released before Apr. 30, 2022, and then clustering the amino acid sequences of the selected protein chains using MMSeq254 with a cutoff of 30% sequence identity. This yielded 40291 clusters for training and 1145 clusters for testing. Each training sample was a continuous peptide segment of at most 512 residues cropped from a randomly selected protein of a randomly selected training cluster plus any ligand units that were spatially close to the remaining protein part after cropping. The retained ligand units included small molecules (excluding a list of common solvents, see Supplementary Table 10), covalently modified sidechains, ions and nucleic acid segments, with at least one atom of the ligand unit within 5 Å from at least one atom of the protein part. Following Dauparas et al.23, Gaussian noise of standard deviation 0.02 Å were added to the atomic positions of the training backbones on the fly to improve the robustness of the resulting model to inaccuracies of input conditions. We also trained another set of parameters with increased Gaussian noise (standard deviation standard deviation 0.1 Å) added during training, and the resulting model was used for designing sequences on computationally generated de novo backbones.
The training process was divided into an initial phase and a fine-tuning phase. The initial phase lasted for 35k training steps (one step corresponded to the processing of a batch of 448 training samples) without considering any self-conditioning. The fine-tuning phase was started with the parameter values learned in the initial phase, lasted 10k steps with self-conditioning turned off for processing 50% of the training samples and turned on for processing the remaining 50% of the samples. To enable self-conditioning at step t, a forward step was firstly performed on the current xt to obtain a temporary \({{\boldsymbol{x}}}_{t+1}\), then a denoising step was performed (without backpropagating the gradients) to decode a temporary \({\widetilde{{\boldsymbol{x}}}}_{0}\left(t+1\right)\) including predicted sidechain conformations. The results were embedded by the pre-trained language model and sidechain encoder as \({embedding}(\widetilde{{\boldsymbol{x}}}_{0}(t+1))\), which was used as additional input to the network denoising the current \({{\boldsymbol{x}}}_{t}\), which may be denoted as \({\text{f}}_{{\rm{\theta }}}\left({{\boldsymbol{x}}}_{t},{embedding}(\widetilde{{\boldsymbol{x}}}_{0}\left(t+1\right))\right)\). The model was trained using an Adam optimizer with 1k warm-up steps and the learning rate decayed exponentially from 8 × 10−4. The entire training process took around 3 days on 8 A40 GPUs (each with 48 G memory).
The network architecture and training of ABACUS-TMSD
The architecture of ABACUS-TMSD (see Fig. S14E) was devised to design one amino acid sequence for multiple conformational states of the backbone and to generate a set of sidechain conformation for each backbone conformational state. Each of the backbone conformational states (some of them could include structures of bound ligands) was encoded using the same encoder module as in ABACUS-T. Compared with ABACUS-T, ABACUS-TMSD uses an additional inter-state updating layer after the repeated intra-state updating layers in the protein representation updating block. Moreover, a state-pooling layer was introduced in the residue type projection bock of ABACUS-TMSD to produce one amino acid sequence from the projections of the protein representations of different conformational states. During the fine-tuning from ABACUS-T to ABACUS-TMSD, the parameters of the encoder module are fixed. The network was fine-tuned for 7k steps (including 1k warm-up steps) with a peak learning rate of 2 × 10−4.
The dataset for fine-tuning comprised of natural proteins with experimentally determined structures in multiple conformational states. To build this dataset, we first clustered the protein chains in the PDB database using MMSeq2 with a cutoff of 100% sequence identity. All clusters containing at least two structures with a mutual RMSD above 1.0 Å for one sequence were considered as valid candidates (we note that the alternative structural models in one NMR structure entry are counted as different conformational states in this process). After removing those clusters containing no X-ray structures but only NMR structures, the candidate proteins were further clustered with a sequence identity cutoff of 30%. From the resulting clusters, we selected a test set of 100 clusters containing in total 140 proteins, each having at least two conformational states with a mutual RMSD above 2.0 Å. All proteins in the remaining 11195 clusters were used for training.
To assess whether the140-protein test dataset is large enough, we performed bootstrap sampling experiments, repeating the analysis 100 times (nbootstrap = 100) with half of the samples (nsample = 70). The results, summarized in Supplementary Table 11, demonstrate that the averaged recovery rates for the sampled datasets are consistent with the original values (e.g., 0.62 for ABACUS-TMSD on multiple states), with small standard deviations (0.01–0.02). These findings suggest that the dataset size is sufficient for robust evaluation, as the performance metrics remain stable even when subsampling the data.
The network architecture and training of ABACUS-Tprior and ABACUS-TMSAT
ABACUS-Tprior was obtained by averaging for each sequence position the residue type distribution produced by ABACUS-T (without fine tuning of the parameters) with weight 1-α with the same distribution produced by the MSA-transformer model46 with weight α (see Fig. S14F). The value of α (i.e., the prior weight) was set to 0.5 unless specified otherwise.
ABACUS-TMSAT was obtained by extending the ABACUS-T decoder block to accept the residue representation from the pre-trained MSA-transformer model as additional input (see Fig. S14F). The parameters of ABACUS-TMSAT were fine-tuned from the parameters of ABACUS-T. To avoid leaking the amino acid context for the masked residues from the MSA-transformer during training, the raw MSA curated from the sequence databases were subjected to a mask operation on the fly before it was feeding into the MSA-transformer model during training. Specifically, the residue types in the wild-type sequence and in randomly selected 97% homolog sequences in the MSA were changed to [MASK] at the currently masked positions. The output of the pre-trained MSA-transformer applied to the masked MSA were added to the one-dimensional output from the iterator module for use as a part of the conditions for the next sequence denoising step. The parameters of ABACUS-TMSAT were fine-tuned from ABACUS-T for an additional 10k steps (including 1k warm-up steps) with a peak learning rate of 1◊10−4 on the same dataset as used in the training of ABACUS-T. The MSAs were obtained by searching against the Big Fantastic Database (https://bfd.mmseqs.com) and UniRef30 database (https://uniclust.mmseqs.com) using MMSeq2 with default settings. The final MSAs consist of the curated sequences filtered using hhfilter55 with the configurations of a minimum sequence coverage of 60% and a maximum pair-wise sequence identity of 90%.
Computational details and experimental characterizing specific proteins
Detailed descriptions of methods are provided in Supplementary Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Protein structures for training the models have been downloaded from the Protein Data Bank. The experimentally solved protein structures have been deposited in the PDB under accession codes: 9JWL (MSD1 holo state); 9JWO (MSD1 apo state); 9JWT (MSD3 holo state); 9JWQ (MSD3 apo state). We referenced the structures 1GUD and 1RPJ from PDB for the redesign of allose binding proteins; 4PN2 and 1JVJ for xylanase and TEM β-lactamase, respectively; and 5WI3, 4X56 and 7T7G for OXA β-lactamase. The amino acid sequences of the experimentally examined proteins are available in Supplementary Tables 14 to 17. The source data of experimental results (enzyme activity, SEC, ITC, CD, validation reports for experimentally solved protein structures) and all in silico experimental results are provided as a Source Data file. Source data are provided with this paper.
Code availability
Executable computer programs and source codes of ABACUS-T (version 1.0) are publicly available from Zenodo at [https://doi.org/10.5281/zenodo.17089342] (ref. 56) and can be freely used for non-commercial purposes.
References
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. (Int. Ed. Engl.) 57, 4143 (2018).
Arnold, F. H. Combinatorial and computational challenges for biocatalyst design. Nature 409, 253–257 (2001).
Packer, M. S. & Liu, D. R. Methods for the directed evolution of proteins. Nat. Rev. Genet. 16, 379–394 (2015).
Cobb, R. E., Chao, R. & Zhao, H. Directed evolution: past, present, and future. AIChE J. 59, 1432–1440 (2013).
Notin, P., Rollins, N., Gal, Y., Sander, C. & Marks, D. Machine learning for functional protein design. Nat. Biotechnol. 42, 216–228 (2024).
Listov, D., Goverde, C. A., Correia, B. E., & Fleishman, S. J Opportunities and challenges in design and optimization of protein function. Nat. Rev. Mole. Cell Biol. 25, 1–15 (2024).
Yang, K. K., Wu, Z. & Arnold, F. H. Machine-learning-guided directed evolution for protein engineering. Nat. methods 16, 687–694 (2019).
Gelman, S., Fahlberg, S. A., Heinzelman, P., Romero, P. A. & Gitter, A. Neural networks to learn protein sequence-function relationships from deep mutational scanning data. Proc. Natl Acad. Sci. USA. 118, e2104878118 (2021).
Cui, Y. et al. Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy. Acs Catal. 11, 1340–1350 (2021).
Lu, H. et al. Machine learning-aided engineering of hydrolases for PET depolymerization. Nature 604, 662–667 (2022).
Zhou, B. et al. Protein engineering with lightweight graph denoising neural networks. J. Chem. Inf. Modeling 64, 3650–3661 (2024).
Repecka, D. et al. Expanding functional protein sequence spaces using generative adversarial networks. Nat. Mach. Intell. 3, 324–333 (2021).
Hawkins-Hooker, A. et al. Generating functional protein variants with variational autoencoders. PLoS Comput. Biol. 17, e1008736 (2021).
Madani, A. et al. Large language models generate functional protein sequences across diverse families. Nat. Biotechnol. 41, 1099–1106 (2023).
Nijkamp, E., Ruffolo, J. A., Weinstein, E. N., Naik, N. & Madani, A. Progen2: exploring the boundaries of protein language models. Cell Syst. 14, 968–978. e963 (2023).
S. R. Johnson et al., Computational scoring and experimental evaluation of enzymes generated by neural networks. Nat. Biotechnol. 43, 396–405 (2024).
Chu, H. et al. High-temperature tolerance protein engineering through deep evolution. BioDesign Res. 6, 0031 (2024).
Fram, B. et al. Simultaneous enhancement of multiple functional properties using evolution-informed protein design. Nat. Commun. 15, 5141 (2024).
Chu, A. E., Lu, T. & Huang, P.-S. Sparks of function by de novo protein design. Nat. Biotechnol. 42, 203–215 (2024).
Watson, J. L. et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023).
Ingraham, J. B. et al. Illuminating protein space with a programmable generative model. Nature 623, 1070–1078 (2023).
Liu, Y. et al. De novo protein design with a denoising diffusion network independent of pretrained structure prediction models. Nat. Methods 21, 2107–2116 (2024).
Dauparas, J. et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Liu, Y. et al. Rotamer-free protein sequence design based on deep learning and self-consistency. Nat. Comput. Sci. 2, 451–462 (2022).
Liu, R., Wang, J., Xiong, P., Chen, Q. & Liu, H. De novo sequence redesign of a functional Ras‐binding domain globally inverted the surface charge distribution and led to extreme thermostability. Biotechnol. Bioeng. 118, 2031–2042 (2021).
Sumida, K. H. et al. Improving protein expression, stability, and function with ProteinMPNN. J. Am. Chem. Soc. 146, 2054–2061 (2024).
Corbella, M., Pinto, G. P. & Kamerlin, S. C. Loop dynamics and the evolution of enzyme activity. Nat. Rev. Chem. 7, 536–547 (2023).
Austin, J., Johnson, D. D., Ho, J., Tarlow, D. & Van Den, R. Berg, Structured denoising diffusion models in discrete state-spaces. Adv. Neural Inf. Process. Syst. 34, 17981–17993 (2021).
L. Zheng, J. Yuan, L. Yu, L. Kong, A reparameterized discrete diffusion model for text generation. In Conference on Language Modeling (COML, 2024).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Chaudhuri, B. N., Ko, J., Park, C., Jones, T. A. & Mowbray, S. L. Structure of D-allose binding protein from Escherichia coli bound to D-allose at 1.8 A resolution. J. Mol. Biol. 286, 1519–1531 (1999).
Santos, C. R. et al. Molecular mechanisms associated with xylan degradation by Xanthomonas plant pathogens. J. Biol. Chem. 289, 32186–32200 (2014).
Maveyraud, L. et al. Crystal structure of 6α-(hydroxymethyl) penicillanate complexed to the TEM-1 β-lactamase from Escherichia coli: Evidence on the mechanism of action of a novel inhibitor designed by a computer-aided process. J. Am. Chem. Soc. 118, 7435–7440 (1996).
Harper, T. M. et al. Multiple substitutions lead to increased loop flexibility and expanded specificity in Acinetobacter baumannii carbapenemase OXA-239. Biochemical J. 475, 273–288 (2018).
Chen, T., Zhang, R. & Hinton, G. Analog bits: Generating discrete data using diffusion models with self-conditioning. arXiv preprint, arXiv:2208.04202 (2022).
Ribeiro, A. J. M. et al. Mechanism and Catalytic Site Atlas (M-CSA): a database of enzyme reaction mechanisms and active sites. Nucleic acids Res. 46, D618–D623 (2018).
Dehouck, Y., Kwasigroch, J. M., Gilis, D. & Rooman, M. PoPMuSiC 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinforma. 12, 1–12 (2011).
Yang, K. K., Zanichelli, N. & Yeh, H. Masked inverse folding with sequence transfer for protein representation learning. Protein Eng., Des. Selection 36, gzad015 (2023).
Park, H. et al. Simultaneous optimization of biomolecular energy functions on features from small molecules and macromolecules. J. Chem. theory Comput. 12, 6201–6212 (2016).
Xiong, P. et al. Protein design with a comprehensive statistical energy function and boosted by experimental selection for foldability. Nat. Commun. 5, 5330 (2014).
Zhang, Y., Zhang, Z., Zhong, B., Misra, S., &Tang, J. Diffpack: A torsional diffusion model for autoregressive protein side-chain packing. Adv. Neural Inf. Process. Syst. 36, 48150– 48172 (2024).
Misiura, M., Shroff, R., Thyer, R. & Kolomeisky, A. B. DLPacker: Deep learning for prediction of amino acid side chain conformations in proteins. Proteins: Struct., Funct., Bioinforma. 90, 1278–1290 (2022).
McPartlon, M. & Xu, J. An end-to-end deep learning method for protein side-chain packing and inverse folding. Proc. Natl Acad. Sci. 120, e2216438120 (2023).
Praetorius, F. et al. Design of stimulus-responsive two-state hinge proteins. Science 381, 754–760 (2023).
Liu, Y. et al. Designing flexible protein structures and sampling protein conformations with a unified model using vector quantization and diffusion. Natl. Sci. Rev., nwaf290 https://doi.org/10.1093/nsr/nwaf290 (2025).
Rao R. M. et al. MSA Transformer. In International Conference on Machine Learning. 8844–8856 (PMLR, 2021).
Notin, P. et al. Proteingym: Large-scale benchmarks for protein fitness prediction and design. Adv. Neural Inf. Process. Systems 36, 64331–64379 (2023).
Ren, M., Yu, C., Bu, D. & Zhang, H. Accurate and robust protein sequence design with CarbonDesign. Nat. Mach. Intell. 6, 536–547 (2024).
Liu, J., Guo, Z., You, H., Zhang, C.,& Lai, L. All-atom protein sequence design based on geometric deep learning. Angewandte Chemie, 63, e202411461 (2024).
Dauparas, J. et al. Atomic context-conditioned protein sequence design using LigandMPNN. Nat Methods 22, 717–723 (2025).
Li, X., Thickstun, J., Gulrajani, I., Liang, P. S. & Hashimoto, T. B. Diffusion-lm improves controllable text generation. Adv. Neural Inf. Process. Syst. 35, 4328–4343 (2022).
Schütt K. et al. Schnet: a continuous-filter convolutional neural network for modeling quantum interactions. Adv. Neural Inf. Process. Systems 30, 992–1002 (2017).
Zhou G. et al. Uni-mol: a universal 3d molecular representation learning framework. In International Conference on Learning Representations (ICLR, 2022).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinforma. 20, 1–15 (2019).
Wang, X. Enhancing functional proteins through multimodal inverse folding with ABACUS-T. Zenodo https://doi.org/10.5281/zenodo.17089342 (2025).
Acknowledgements
We thank Dr. Chao Xu and the staff from the BL19U1 beamlines of the National Facility for Protein Science in Shanghai (NFPS) for their assistance during crystallographic data collection. We also thank Yuzhu Wang and Lei Wang for their help with experimental techniques, and Dr. Qingguo Gong and Dr. Lin Xue for their insightful discussions and suggestions. This work was supported by the National Key R&D Program of China (2022YFF1203100 to Q.C. and 2022YFA1303700 to H.L.), National Natural Science Foundation of China (T2221005, 92253302, 22177107 to H.L. and 32371487, 32171411 to Q.C.), CAS Strategic Priority Research Program (XDB0500201 to H.L.), CAS Project for Young Scientists in Basic Research (YSBR-072 to Q.C.), Anhui Provincial Natural Science Foundation (2308085J01 to Q.C.), and Research Funds of Center for Advanced Interdisciplinary Science and Biomedicine of IHM (QYPY20230035 to Q.C.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.L. developed computational models and codes with the assistance of L.C. R.W. and X. W. carried out the experimental work with the help of S.W. F.L. helped analyzing the crystal structural data. H.L. and Q.C. supervised the project. H.L., Y.L. and Q.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks Markos Koutmos, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Wu, R., Wang, X. et al. Enhancing functional proteins through multimodal inverse folding with ABACUS-T. Nat Commun 16, 10177 (2025). https://doi.org/10.1038/s41467-025-65175-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65175-3