Abstract
Strigolactones are plant hormones that regulate development and mediate interactions with soil organisms, including the germination of parasitic plants such as Striga hermonthica. Strigolactone perception by receptors initiates the degradation of transcriptional repressors via E3 ubiquitin ligases, but the mechanistic link between hormone binding and substrate ubiquitination has remained unclear. We determine cryogenic electron microscopy structures of the receptor–ligase–substrate complex, composed of Arabidopsis ASK1 and substrate, and Striga F-box and receptor proteins. Strigolactone hydrolysis by the receptor, which covalently retains the D-ring, is a prerequisite for complex formation. The substrate engages the complex through two domains, forming a dynamic interface that stabilises the receptor–ligase assembly and repositions the ASK1, suggesting a mechanism for efficient ubiquitination. Here, we show how dynamic, multivalent interactions within the receptor–ligase–substrate complex translate hormone perception into targeted protein degradation, providing insight into how plants integrate hormonal signals into developmental decisions.
Similar content being viewed by others
Introduction
Eukaryotes utilise a cascade of enzymes, known as E1, E2, and E3, to post-translationally modify substrate proteins with ubiquitin1,2,3. Ubiquitination often leads to proteasome-mediated degradation of the substrate, but can also result in altered location or activity4,5,6. This regulatory versatility makes ubiquitination essential for proteome homoeostasis and enables rapid cellular adaptation to environmental and developmental cues. In plants, the E3 ubiquitin ligase family has undergone significant expansion, with over 1000 predicted members, underscoring their critical role in signalling and adaptation in these sessile organisms7,8,9.
E3 ligases facilitate substrate ubiquitination by bridging ubiquitin-charged E2 enzymes with specific target proteins. A major family of E3 ligases in both plants and animals is the multisubunit SCF complex, named after its canonical core components Skp1, cullin, and F-box proteins10,11,12,13. Within this modular assembly, cullin binds the E2–ubiquitin conjugate (via an associated RING-box protein), F-box proteins bind to the substrate, and Skp1 serves as a linker between the two modules.
Reflecting their role in recognising diverse substrates, nearly 900 different F-box proteins have been identified in plants14,15,16. Whereas animal F-box proteins primarily recognise post-translationally modified substrates17, recent findings suggest that plant F-box proteins have evolved sophisticated recognition mechanisms to also function as environmental sensors7,18. These sensors link phytohormone perception with proteomic changes. Initial structural analyses have revealed how SCF E3 ligases mediate the perception of plant hormones such as auxin and jasmonate18,19,20. In both cases, the hormone associates with the leucine-rich repeat (LRR) domain of the F-box protein, creating a shared binding site for the substrates. The subsequent ubiquitination and degradation of these substrates, which are transcriptional repressors, lead to changes in gene expression.
The perception of strigolactones (SLs) also involves SCF E3 ligases, but with additional layers of complexity. Strigolactones are plant hormones that play diverse roles in regulating plant architecture, adaptation, defence, and plant–rhizospheric communications21,22. SLs are carotenoid derivatives characterised by a butenolide ring (D-ring) linked by an enol ether bridge to a tricyclic lactone (ABC-ring) in canonical SLs, or to a different moiety in non-canonical SLs23,24,25. Unlike auxin and jasmonate, SLs do not bind directly to their F-box protein—MORE AXILLARY GROWTH 2 (MAX2) in Arabidopsis or DWARF3 (D3) in rice—but instead to the α/β-hydrolase DWARF14 (D14)26,27. D14, derived from an α/β-hydrolase, retains low hydrolase activity, allowing cleavage of the SL enol ether bridge while retaining the D-ring28,29. SL recognition destabilises the lid domain of D14, enabling it to associate with MAX228,30. The resulting F-box–D14 complex recruits substrates, which are the transcriptional repressors DWARF53 in rice or SUPPRESSOR OF MAX2 1 (SMAX1)–like (SMXL) 6, 7, and 8 in Arabidopsis31,32,33,34,35. These substrates all share a common domain structure composed of an N-terminal double Clp-N motif (N), a first ATPase domain (D1), a middle domain (M), and a C-terminal second ATPase domain (D2). Assembly of the receptor–ligase–substrate complex enables SCF-mediated ubiquitination and proteasomal degradation of the repressor, thereby activating downstream signalling. The specific signalling outcome is determined by the repressor’s function.
SLs exuded from the roots of host plants also serve as germination cues for the parasitic plant Striga hermonthica36. Upon germination, S. hermonthica attaches to host roots and syphons off water and nutrients, leading to severe crop losses that amount to billions of dollars annually and contribute significantly to food insecurity in affected regions37,38,39. To exploit this host-derived signal, S. hermonthica has adapted the SCF–D14 signalling pathway by evolving a family of D14 homologues known as HYPOSENSITIVE TO LIGHT (HTL) proteins40. These HTLs connect the F-box protein ShMAX2 to transcriptional repressors that regulate seed germination, likely paralogues of SMAX1. Among these, ShHTL7 exhibits the highest sensitivity to strigolactones and is a key player in germination signalling41,42. Efforts to develop chemical inhibitors that block S. hermonthica germination have been hampered by the high degree of conservation between the SL perception machinery of the parasite and that of its host plants, complicating selective targeting39,43,44,45.
Although there is general agreement on the SLs’ response mechanism in plants, several key aspects remain unresolved and controversial. These include the precise role of SL hydrolysis in signal transduction, the identity of the SL degradation product retained by D14/HTL during complex formation with MAX2/D346,47, and the mechanism by which receptor–F-box association leads to substrate recognition and ubiquitination48,49. These uncertainties largely stem from the absence of structural information on the complete receptor–ligase–substrate complex. While accumulating evidence suggests that the substrate contributes to both the stability and specificity of the SCF–D14/HTL–substrate complex, structural studies to date have only captured partial assemblies, such as Skp1–F-box alone or Skp1–F-box–D1428,50,51. Notably, a recent cryo-electron microscopy (cryo-EM) study aimed at resolving the full SCF–D14/HTL–substrate complex failed to detect density for the D53 substrate, even though D53 was necessary for stabilising the MAX2–D14 interaction52. This highlights a central paradox: the substrate is essential for complex formation and is ultimately ubiquitinated, yet its interactions are so transient and dynamic that they cannot be captured by structural methods.
In this study, we leveraged species-specific idiosyncrasies to identify an active signalling complex that is sufficiently stable to resolve the substrate when cryogenic grid preparation for cryo-EM analysis is carefully timed.
This complex, composed of Arabidopsis SKP1 (ASK1), ShMAX2, ShHTL7, and AtSMAX1, demonstrates how a bidentate and highly dynamic interaction with AtSMAX1 confers both stability and specificity to the SCFMAX2–HTL complex. SMAX1 also allosterically influences its positioning to the E2 enzyme, suggesting a mechanism for efficient ubiquitination. Our findings uncover a previously uncharacterised mechanism of SCF-mediated phytohormone signalling and provide a structural framework that reconciles earlier biochemical and functional observations.
Results
Identification of stable complexes
Previous work suggested that the interaction between D14/HTLs and the F-box proteins requires the slow hydrolysis of SLs by the receptor and is only stable for a few hours in vitro26,29,53. To capture this slow-onset transient complex, we used single-particle analysis (SPA) with cryogenic electron microscopy (cryo-EM) to image samples vitrified within a 2-3 h time window after addition of GR24, a synthetic SL analogue. Complexes were reconstituted by combining D14/HTL receptors produced in Escherichia coli with SKP1 and F-box proteins co-expressed in insect cells (see “Methods”). Our initial cryo-EM structural analysis focused on Striga HTL7, rice D3, and Arabidopsis ASK1, as this complex showed GR24-dependent stable binding on size-exclusion chromatography (SEC) and pull-down studies (Supplementary Fig. 1A–C). Cryo-EM grids showed severe preferred particle orientation, which we addressed by tilting the stage. Nonetheless, these grids only yielded the structure of the AtASK1–OsD3 complex to a maximum resolution of 4.3 Å (Supplementary Fig. 1D–J), without the SL-bound ShHTL7 receptor, suggesting that the tripartite complex was too unstable under the cryogenic preparation conditions used. Attempts to stabilise the complex through chemical cross-linking were unsuccessful.
Guided by previous reports suggesting that substrate binding can stabilise the receptor–F-box interaction52,54, we next included the ubiquitination substrate to stabilise the complex for cryo-EM. We selected AtSMAX1 as the substrate based on prior evidence of its interaction with ShHTL7 and both Striga and Arabidopsis MAX254,55, and because reliable Striga SMAX1 orthologues were not yet available at the time (our work began prior to the release of the annotated Striga proteome56). SEC and pull-down experiments indicated that the most stable complex in the presence of GR24 was formed by HTL7 and MAX2 from S. hermonthica, together with A. thaliana ASK1 and SMAX1 (Fig. 1A, B). Conversely, AtSMAX1 interacted only weakly with OsD3–AtASK1–ShHTL7 in the presence of GR24 (Supplementary Fig. 1K). We therefore focused on AtASK1, ShMAX2, ShHTL7, and AtSMAX1, which we refer to hereafter as ASK1, MAX2, HTL7, and SMAX1, respectively, unless disambiguation is required.
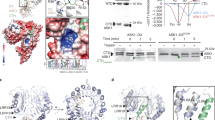
A SEC profile (top) and corresponding Coomassie-stained SDS-PAGE (bottom) of ASK1–MAX2–HTL7–SMAX1 with (orange; right) and without (blue; left) the addition of GR24. Solid lines show the absorbance at 280 nm. Roman numerals indicate the peak fractions analysed. The approximate molecular weight of each protein is labelled next to the reference ladder. The dotted black box highlights the fractions containing the entire complex. Representative of three independent experiments (n = 3). B Coomassie-stained SDS-PAGE of GST pull-down assays using GST-HTL7 as the bait and combinations of ASK1–MAX2, SMAX1, GR24, and MP3 as targets (n = 1, findings supported by complementary experiments). Plus signs denote the presence and minus signs the absence of species in each pull-down assay. Approximate molecular weights are labelled. C–E Compositional assemblies of ASK1–MAX2–HTL7–SMAX1 in the presence of GR24. ASK1, MAX2, and HTL7 are coloured yellow, green and purple, respectively. SMAX1N is coloured light pink, and SMAX1D2 is coloured dark pink. Unassigned cryo-EM density is coloured white. C Cryo-EM density of apo-ASK1–MAX2 (EMD-62408). D Cryo-EM density of the 2:2 ASK1–MAX2 dimer (EMD-62414). E Cryo-EM density of ASK1–MAX2–HTL7 classes. Class 3 is a subclass of Class 2, following masked classification around SMAX1N. (Class 1, EMD-62397; Class 2, EMD-62401; Class 3, EMD-62417; Class 4, EMD-62407). F Complete atomic model of ASK1–MAX2–HTL7–SMAX1 from Class 3 (PDB 9KLV), coloured as in (C–E). Cryo-EM density of Class 3 is shown as a transparent grey surface. The GR24 D-ring atoms are shown as red spheres and highlighted by black arrows.
Structural Landscape Of The SCFMAX2–Receptor–Substrate Complexes
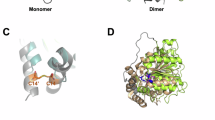
For cryo-EM analysis, purified ASK1, MAX2, HTL7, and SMAX1 were incubated with GR24 and subjected to SEC. The peak fraction was concentrated and used for cryo-EM grid preparation. SPA of these grids yielded structures of three compositional assemblies: (1) the unliganded ASK1–MAX2 complex determined to 2.7 Å resolution from 524k particles; (2) a 2:2 ASK1–MAX2 dimer, with a strongly anisotropic resolution of maximal 3.3 Å obtained from 309k particles; and (3) the ASK1–MAX2–HTL7 complex resolved to 2.7 Å from 710k particles (Fig. 1C, D and Supplementary Fig. 2–5).
The ASK1–MAX2–HTL7 complex adopted the same seahorse-shaped topology previously observed in the crystal structure of the ASK1–OsD3–AtD14 complex (PDB 5HZG)28. In this configuration, HTL7 forms the head, the solenoid-shaped leucine-rich repeat (LRR) domain of MAX2 constitutes the body, and the N-terminal F-box motif of MAX2 together with ASK1 forms the tail (Fig. 1E, F).
Further 3D classification of the ASK1–MAX2–HTL7 assembly revealed additional densities corresponding to domains of SMAX1 (Supplementary Fig. 2). Class 1 (with 60k particles, refined to 2.8 Å resolution) contained additional density docked on the leucine-rich repeat (LRR) domain of MAX2 in proximity to ASK1 (Fig. 1E). Based on secondary structure features and AlphaFold structure predictions, we identified this density as the SMAX1 N domain (residues 10-167) (Fig. 1F). Class 2 (with 80k particles refined to 2.9 Å resolution) showed weaker density for the N domain, but featured a clear additional density between HTL7 and MAX2, which we identified as the SMAX1 D2 domain (Fig. 1E, F). The observation that Class 1 and Class 2 each resolved only one of the SMAX1 domains raised the question of whether both domains can bind simultaneously. Further subclassification of Class 2 identified a subset of 17k particles (termed Class 3) that produced a structure with density for both N and D2 domains, but at a lower resolution of 3.3 Å (Fig. 1E). A final Class 4 contained the complex that only showed density for ASK1, MAX2 and HTL7 (44k particles, 2.9 Å) (Fig. 1E).
These structures provided atomistic information on an SMXL transcription factor. They suggested that the N and D2 domains of SMAX1 bind to the ASK1–MAX2–HTL7 complex with a significant flexibility (conformational heterogeneity), where focused refinement results in different degrees of visibility of the N and D2 domain density. However, our data also supports the existence of compositional heterogeneous complexes, where either N or D2 are bound. We did not identify density for the SMAX1 D1 and M domains, suggesting that all SMAX1 domains are flexibly linked, in agreement with AlphaFold predictions (model AF-Q9FHH2-F1-v4).
The closed D-ring is bound to the catalytic histidine in the signalling state
We first focused our analysis on the receptor component of the complex. In our cryo-EM structures, HTL7 is bound to the C-terminal LRRs 17-20 of MAX2 (Fig. 1F). The lid domain of HTL7 adopts a closed conformation, in contrast to its previously reported open state (PDB 5Z8P) (Fig. 2A). This positioning and conformation of HTL7 closely resemble those observed for AtD14 in its crystallographic complex with ASK1–OsD3 (PDB 5HZG)28 (Supplementary Fig. 6A). Compared to apo-HTL7, MAX2-bound HTL7 clearly displays an additional density attached to the catalytic histidine H246 (Fig. 2B). This density is consistent with a covalently bound closed D-ring (termed the C5H5O2 modification)42,55,57, but not with the open CLIM linked to both S95 and H246, as proposed for the ASK1–OsD3–AtD14 crystal structure (PDB 5HZG) (Supplementary Fig. 6B)28. Our cryo-EM structure was captured within the hour-long SL signalling window, suggesting that the covalently attached closed D-ring represents the active signalling state. Mass spectrometry analysis of HTL7 preincubated with GR24 confirmed the presence of a C5H5O2 modification covalently linked to H246. This modification was absent in control samples lacking GR24 (Supplementary Fig. 7), and aligns with previously published mass spectrometry data for HTL7 incubated with GR2442,55.
A Superimposition of HTL7 from our cryo-EM structure (PDB 9KKX; purple) and the open-conformation crystal structure of HTL7 (5Z8P; pink). The collapsed lid domain of our model is coloured grey, and the D-loop is coloured magenta. The catalytic triad residues are shown as sticks. The black box outlines the active site and magnified views in the following panels. B Fit of the D-ring covalently attached to H246 (H246–D-ring) in the cryo-EM density. Catalytic triad residues are shown as sticks and labelled. C Hydrophobic enclosure (mesh outline) in the open (top) and our closed (bottom) HTL7 active site. The GR24 D-ring is shown as orange spheres. Red arrows highlight the active site entrance. The collapsed lid domain is coloured grey. Catalytic triad residues are shown as sticks and labelled (top). Residues that form the hydrophobic enclosure in the closed conformation are shown as sticks and labelled (bottom). D Superimposition of catalytic triad residues in the closed and open conformations with the H246–D-ring modification added to the open conformation. Yellow dashed lines depict clashes between atoms closer than 0.6 Å. E Superimposition of catalytic triad residues in closed and open conformations. Distances between atoms are shown as dotted lines and labelled.
The D-ring is tightly encapsulated by hydrophobic interactions involving residues Y26, M139, Y174, L178, and I193. These residues are distant from the catalytic residues and form a 238 Å3 cavity in the open conformation of HTL7, but collapse to 18 Å3 in the MAX2-bound closed conformation, leaving no space for the ABC rings of an uncleaved SL (Fig. 2C and Supplementary Fig. 6C). Accordingly, our cryo-EM data show no density corresponding to the ABC rings of GR24, indicating that this moiety was released from the active site cleft.
In the MAX2-bound HTL7, the loop containing H246 is shifted away from S95 to a distance of 3.9 Å, compared to 3.0 Å in the open state. This shift is likely the result of the covalently attached D-ring, as a D-ring–modified H246 would clash with S95 in the open receptor conformation (Fig. 2D). The catalytic D217, located on the eponymous D-loop58, is also displaced by 2.9 Å relative to open HTL7. The combined movement of H246 and D217 results in a distance of more than 7 Å between their side chains, compared to 2.7 Å in the open form (Fig. 2E) (PDB 5Z8P)59, disrupting the catalytic triad. Along with the tight hydrophobic enclosure that prevents nucleophilic attack by a water molecule, this architecture stabilises the covalently bound D-ring within a catalytically inactive conformation of HTL7.
In the crystal structure of OsD3–AtD14, the AtD14 D-loop was not resolved and was proposed to move out to associate with D53/SMXLs29. However, in our structure, the D-loop was well resolved and positioned more than 10 Å away from the SMAX1 D2 domain (Supplementary Fig. 6D). No unattributed density was visible near the D-loop, but we cannot exclude transient interactions with SMAX1 regions other than the N or D2 domains. Collectively, these observations inferred that SL cleavage and covalent capture of the D-ring must occur before the docking of HTL7 onto the MAX2 LRR surface can be completed.
To further substantiate the requirement of hydrolysis for signalling complex formation, we generated two catalytically inactive HTL7 mutants: HTL7S95C (a conservative substitution of the catalytic serine) and HTL7D217A (a non-homologous substitution of the catalytic aspartate). Using differential scanning fluorimetry (DSF) and nanoDSF, we observed that while HTL7S95C retained thermal stability similar to wild-type HTL7, HTL7D217A was inherently unstable (Supplementary Fig. 8A–D). Importantly, SL addition reduced the melting temperature, Tm, of wild-type HTL7 but had no effect on either mutant, consistent with impaired hydrolysis. Functionally, the HTL7S95C and HTL7D217A mutants were defective in YLG hydrolysis, and HTL7S95C failed to form a complex with MAX2 and SMAX1 in the presence of GR24 (Supplementary Fig. 8E, F). The HTL7D217A mutant, due to its instability, could only be tested in pull-down assays, where it also failed to form the signalling complex (Supplementary Fig. 8G). These findings underline the requirement of HTL7 catalytic activity for formation of the SCFMAX2–HTL7–SMAX1 complex described in our cryo-EM data.
Together, these observations substantiate that SL-hydrolysis is required for the association of the ASK1–MAX2–HTL7–SMAX1 signalling complex displayed in Fig. 1F.
Stabilisation of the MAX2–HTL7 interaction by the substrate
The D2 domain of SMAX1 is essential for its ubiquitination and degradation60. However, the structural basis for this requirement has remained unclear, as the closest experimentally resolved homologues—AAA + disaggregase ClpB proteins—share less than 24% sequence identity. Our cryo-EM maps now provide a clear resolution of the D2 α/β core domain (residues 607–767), corresponding to the structured region of the annotated D2a subdomain60, as well as the C-terminal portion of D2b (residues 860–885), which contributes an additional helix–strand motif to the D2a α/β core (Fig. 3A, B). Density for the remaining D2b domain residues 774–798 and 886–990 (forming a four-helix structure capped by a two-stranded β-sheet) only appears at lower thresholds in the cryo-EM map, indicating that this domain is more flexible (Supplementary Fig. 9A).
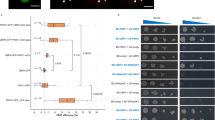
A Reference view of ASK1–MAX2–HTL7–SMAX1 (EMD-62417); grey box outlines the region magnified in panels (B–E). Close-up box shows SMAX1D2 domain fitted into the cryo-EM density of Class 2 (grey, EMD-62401). Cryo-EM densities shown as transparent grey surfaces. Proteins and domains coloured as in Fig. 1. The Class 2 model (PDB 9KLD) is used for panels (B–E). B SMAX1D2a (residues 607–767; dark pink) and SMAX1D2b (residues 860–885; light pink) superimposed on the nucleotide binding domain (NBD) of ClpB (4LJ8; white). The GYGV loop of ClpB (orange) and the R700GKT motif of SMAX1D2 (yellow) are highlighted. Bound ADP shown as sticks and the Walker A fold of ClpB are coloured blue. The Walker A fold is absent in SMAX1D2. C Polar interface of SMAX1D2–MAX2. MAX2 depicted as a surface coloured by electrostatic potential (red: negatively charged; blue: positively charged). SMAX1D2 interacting residues (light pink sticks) and the RGKT motif residues (in grey) are labelled. D Hydrophobic interface of HTL7–SMAX1D2. The R700GKT motif residues of SMAX1D2 are coloured grey and labelled. Hydrophobic residues are shown as sticks and labelled. E Interaction of MAX2–HTL7–SMAX1D2 through R700 of the SMAX1D2 R700GKT motif. The R700GKT motif residues are coloured grey and labelled. Black arrows indicate LRR loops 17 and 18 of MAX2, and dotted lines show hydrogen bonds and salt bridges with R700. Interacting residues are shown as sticks and labelled. F Coomassie-stained SDS-PAGE of GST pull-downs using GST-HTL7 as bait and combinations of ASK1–MAX2, full-length SMAX1, full-length SMAX1∆RGKT, SMAX1D2, SMAX1D2∆RGKT, and GR24 (n = 1). Plus/minus signs denote the presence and absence of species in each lane. Approximate molecular weights are labelled. G SEC profiles (left) and Coomassie-stained SDS-PAGE (right) of ASK1–MAX2–HTL7–SMAX1 and GR24 with (orange; n = 3) and without (green; n = 1) the SMAX1 RGKT motif versus MAX2–HTL7 and GR24 alone (blue; n = 1). Solid lines show the absorbance at 280 nm. Asterisk and black dashed line, and boxes highlight fractions analysed by SDS-PAGE. The gels outline matches the SEC profile colours. See Supplementary Fig. 9E for the SDS-PAGE gel for MAX2–HTL7 and GR24 alone. The observations from experiments with n = 1 were supported by complementary assays.
Structure superimposition of SMAX1D2 with the second nucleotide-binding domain (NBD2) of ClpB from T. thermophilus (PDB 4LJ8)61 produces an RMSD of 1.3 Å for the conserved α/β core (85 residues), but exceeds 6 Å when all 166 residues are considered (Fig. 3B). This indicates substantial structural divergence beyond the conserved core. Notably, SMAX1D2 differs from canonical NBDs in several key functional elements. Firstly, we observed no evidence of a bound nucleotide in our structure. This absence is attributable to a degenerate Walker A motif, which adopts a helical conformation that occludes the canonical nucleotide-binding site (Supplementary Fig. 9B, C). Secondly, the GYGV motif, which forms a pore loop in the ClpB hexamer, is replaced by the R700GKT motif in SMAX1D2 (Fig. 3B and Supplementary Fig. 9C). This RGKT motif is conserved among SMAX1 homologues (Supplementary Fig. 9D)60. Its deletion in SMAX1, D53, SMXL6, and SMXL7 abrogates GR24-induced degradation of these proteins in planta, suggesting that this motif is either ubiquitinated or stabilises the SCF–D14/HTL complex32,34,35,60. These findings show that both the structure and function of SMAX1D2 have significantly diverged from NBDs of hexameric AAA + ATPases.
In our cryo-EM model of the E3–substrate complex, D2 associates with an LRR surface formed through mostly charged and polar contacts (D2 residues R700, K702, R744, R746, H749, R751, and E752) with only two hydrophobic residues making significant contacts (P716 and F717) (Fig. 3C). This interaction buries 928 Å2 of solvent exposed surface. Conversely, the D2 contact surface with HTL7 is smaller (430 Å2 buried in total) and more hydrophobic, formed mostly by D2 residues G701, T703, and L731, engaging HTL7 residues L35, Y39, L250, M252, P253, V255, and P258) (Fig. 3D). In addition, HTL7 E254 is reaching into the SMAX1D2 loop formed by residues 686 to 702, where HTL7 E254 forms a hydrogen bond with the backbone carbonyl of D694, and an ionic interaction with K707 (Fig. 3E). The SMAX1D2 residue K702 is mostly buried in the complex, making it an unlikely substrate for ubiquitination (Fig. 3C, E). However, SMAX1D2 R700 stabilises the association between HTL7 and MAX2 by wedging between the MAX2 loops of LRRs 17 and 18 (residues 604–616 and 642–650, respectively)(Fig. 3E). In this position, R700 forms ionic bonds with MAX2 D643 and D645, and hydrogen bonds with T609. The LRR17 and LRR18 loops are in a similar position in the crystallographic ASK1–OsD3–AtD14 complex (PDB 5HZG, where SMAX1 was absent), but the LRR17 loop was arranged differently in our ASK1–MAX2 apo structure (Supplementary Fig. 6F), indicating that the LRR17 loop adopts a D14/HTL-bound conformation that is further stabilised by the D2 domain.
Our structural analysis suggested that the conserved R700GKT motif is used by SMXL proteins as a part of the interface with SCFMAX2 and the SL-bound receptor. To test this model, we first generated RGKT deletion (∆RGKT) mutants in both full-length SMAX1 and the isolated SMAX1D2 to mimic the original phenotypic d53 mutant reported by Zhou et al.34. Based on our structural data and prior observations of d53 stability in Col-0 cells50, deletion of these four residues from the 18-residue loop was not expected to compromise the D2 domain structure. We tested the ability of wild-type and ΔRGKT variants of full-length SMAX1 and SMAX1D2 to stabilise the ASK1–MAX2–HTL7–SMAX1 complex in SEC and pull-down experiments. MAX2 and GST-HTL7 bound only weakly in GST pull-down assays in the presence of GR24, and did not co-elute in SEC without SMAX1 (Fig. 1B and Supplementary Fig. 9E). The presence of SMAX1 increased GR24-induced co-precipitation and co-elution of MAX2 and HTL7, further confirming the stabilising role of SMAX1 (Fig. 1A, B). We further observed that D2 alone increased GR24-induced complex coprecipitation less than full-length SMAX1 in our pull-down assay (Fig. 3F), and that only full-length SMAX1, but not D2 alone, stabilised the complex on SEC (Supplementary Fig. 10A). These observations confirmed that the presence of SMAX1N stabilises the complex, presumably through its additional ASK1–MAX2 interactions, and that SEC provides a more stringent assay than pull-down.
Both SEC and pull-down assays showed reduced complex stabilisation by the SMAX1ΔRGKT mutant compared to wild-type. (Fig. 3F, G). In agreement, SMAX1D2ΔRGKT was less effective than wild-type SMAX1D2 in co-precipitating the complex. In contrast, an RGKT to AGAA substitution had no detectable effect on complex formation in SEC and in pull-down (Supplementary Fig. 10B, C), suggesting that complete deletion, rather than conservative substitution, is required to disrupt the interface. To rule out whether the effect of the ΔRGKT mutants was based on their instability, we designed a triple SMAX1 mutant (termed SMAX1mD2) carrying the R700E, L731D, and R744E substitutions that target the D2–MAX2 and D2–HTL7 interfaces (Fig. 3C–E). This mutant failed to stabilise the complex on SEC, in agreement with our structural model (Supplementary Fig. 10D).
Together, these findings demonstrated that SMAX1D2 is an ATPase domain repurposed to function as a non-catalytic binding module, ensuring proper assembly and positioning of the MAX2–HTL–substrate complex for efficient ubiquitination and signal transduction. Within this complex, the SMAX1D2 RGKT motif contributes to a composite interface involving multiple domains and proteins, rather than serving as a direct ubiquitination site.
Idiosyncrasies in the interface between MAX2 and the N domain
Our cryo-EM data for Class 1 and 3 revealed an additional density different from SMAX1D2 (Fig. 4A). The number and topology of helical elements observed in this density were inconsistent with the remaining D1 and M domains of SMAX1, but produced a good fit for the AlphaFold prediction of the SMAX1 N domain. SMAX1N belongs to the Clp repeat (ClpR) family62,63. Its closest structurally characterised homologue is the N domain of the AAA + protein ClpC1 from Mycobacterium tuberculosis (PDB 6PBA), which shares only 17.3% sequence identity64. Despite the low identity, SMAX1N retains the hallmark architecture of ClpR domains, namely two four-helix bundle repeats (residues 12–83 and 97–167), arranged with pseudo-twofold symmetry, as previously described for other ClpR family members65. Structural superposition of SMAX1N with M. tuberculosis ClpC1 N domain reveals a close alignment of the conserved helical elements within each repeat (RMSD ≈ 1 Å), but a larger deviation across the full domain (RMSD ≈ 5 Å) (Fig. 4B and Supplementary Fig. 11A). These differences in both sequence and overall domain architecture suggest that SMAX1N has functionally significantly diverged from canonical ClpR proteins.
A Overview of the ASK1–MAX2–HTL7–SMAX1 complex, with a grey box highlighting the region examined in panels (B–G). A magnified view shows the fit of the SMAX1N in the cryo-EM density of Class 1 (grey, EMD-62397). Arrows indicate the orientation and perspective corresponding to each subsequent panel. Proteins and domains are coloured as in Fig. 1. The model corresponding to Class 1 (PDB 9KKX) is used for panels (B–G). B Repeat 1 (grey) and Repeat 2 (pink) of SMAX1N superimposed on the N-terminal domain (NTD) of ClpC1 (6PBA; light blue). SMAX1N helices are labelled, with prime marking the pseudosymmetric helix in Repeat 2. C Interface of MAX2–SMAX1N through Repeat 2. The pseudosymmetric axis of SMAX1N is shown as a dotted line. The interacting LRR1-3 of MAX2 and H1’ of SMAX1N are labelled. D Hydrophobic interface of MAX2–SMAX1N. Interface residues are coloured grey, shown as sticks, and labelled. Additional hydrogen bonds between SMAX1N residues E162 and R113 with MAX2 H130 and the carbonyl oxygen of P133 are shown as dotted lines. E Electrostatic interface of ASK1–SMAX1N. Red represents negatively and blue represents positively charged surfaces. Interface residues are coloured grey, shown as sticks, and labelled. F Unassigned cryo-EM density (black box) adjacent to SMAX1N and LRR3-5 of MAX2. Cryo-EM density is shown as a transparent grey surface. G. Position of MAX2 C186 in ASK1–MAX2–HTL7–SMAX1 (PDB 9KKX; light green), apo-ASK1–MAX2 (PDB 9KLL; dark green) and OsD3 (PDB 5HZG; grey). Cryo-EM density of Class 1 (EMD-62397) is shown as a transparent grey surface. C198 of OsD3 and C186 of MAX2 are shown as sticks and labelled. The distance between C186 in apo-ASK1–MAX2 and Class 1 is shown as a dotted line. H SEC profiles (left) and corresponding Coomassie-stained SDS-PAGE (right) of ASK1–MAX2–HTL7 complex with SMAX1ND2 with (blue; bottom) and without GR24 (orange; top). Solid lines show the absorbance at 280 nm. The approximate molecular weight of each protein is labelled next to the reference ladder. The dotted black box highlights the fractions containing the entire complex. n = 1, findings supported by complementary experiments.
Although SMAX1N repeats 1 and 2 are structurally similar, our cryo-EM data clearly identify repeat 2 as the region that binds MAX2. First, several side-chain features in the density map match repeat 2 residues, whereas repeat 1 yields a poor fit (Supplementary Fig. 12A–C). Second, repeat 2 produces a two-fold higher model-to-map correlation coefficient than repeat 1, and a better secondary structure fit to cryo-EM density, when placed at the MAX2 interface (Supplementary Fig. 12D–G).
SMAX1N repeat 2 binds to MAX2 LRRs 1 and 2, close to the ASK1 interface, burying 558 Å2 of surface area (Fig. 4C and Supplementary Fig. 12). The hydrophobic side of the C-terminal SMAX1N helix (H4’, residues S153 to N166) interacts with a hydrophobic patch on the first and second LRR of MAX2 (residues F89, M90, L123, and L127) (Fig. 4D). This interaction is further stabilised by hydrogen bonds and charge–charge interactions between R145 and R148 of the SMAX1N H3’ helix and E146, E147, and E150 of ASK1 (Fig. 4E). An additional density, which we could not attribute to any protein region, is observed between the SMAX1N H1’–H2’ loop and the loops connecting the helices of LRRs 3 to 5 with their following β strands (Fig. 4F). Notably, the helix–β loop of LRR4 (residues 183–189) is displaced outward by up to 6.9 Å (for the Cα of C186) compared to the apo ASK1–MAX2 structure and the ASK1–OsD3–AtD14 crystal structure (Fig. 4G), suggesting that this conformational shift is induced by SMAX1 binding.
To investigate the role of the N domain in complex assembly, we generated both the isolated N domain and a truncated construct comprising the N and D2 domains (termed SMAX1ND2). The domain boundaries, based on Khosla et al.60, include a portion of the intrinsically disordered region following the helical N domain. In the SMAX1ND2 construct, this region serves as a flexible linker, allowing the N and D2 domains to bind independently within the HTL7–MAX2 complex (Supplementary Fig. 13A). Unlike SMAX1D2, the SMAX1ND2 construct remained stably associated with MAX2 and HTL7 in the presence of GR24, as shown by SEC (Fig. 4H). This interaction was further validated by GST pull-down assays in which we included SMAX1N and SMAX1D2 (Supplementary Fig. 13B). After extensive washes, both full-length SMAX1 and SMAX1ND2 showed strong association with MAX2 and HTL7 in the presence of GR24. Interestingly, SMAX1D2 co-precipitated more MAX2 when SMAX1N was added in trans, indicating that binding of the two domains is cooperative, possibly through stabilising MAX2. However, significantly more MAX2 was co-precipitated when the N and D2 domains were covalently linked in the SMAX1ND2 construct than when supplied separately. This indicates that while the N domain alone binds MAX2 weakly, its tethering to the D2 domain is necessary for robust complex stabilisation (Supplementary Fig. 13B). Mutational analysis targeting either repeat 1 or repeat 2 of the N domain did not significantly impair complex assembly (Supplementary discussion and Supplementary Fig. 13C–E). This minimal effect in our assays is likely due to two factors: the dominant role of the D2 domain in stabilising the complex, and the incomplete disruption of the N domain interface by the point mutations introduced. In addition, the unexplained density adjacent to the N domain (white density in Fig. 1E, Class 1) may reflect additional interactions, possibly from the flexible linker regions flanking the N domain, not fully captured by the current structural model.
Biochemical evidence for the dynamic features of the complex
Our cryo-EM analysis supported several conclusions: (i) SL hydrolysis by HTL7 destabilises its lid domain, which is a prerequisite for binding to MAX2, (ii) SMAX1 binds to the complex with at least two domains (N and D2) in a flexible and dynamic manner, and (iii) D2 binding stabilises the association of HTL7 with MAX2. To further investigate these findings in solution, we employed hydrogen-deuterium exchange (HDX) coupled with mass spectrometry.
We compared the changes in HTL7 when complexed with ASK1–MAX2 alone, and in combination with full-length SMAX1 and the D2 domain only, all in the presence of GR24. We obtained a high peptide coverage for all the components (95% for ASK1, 97% for MAX2, 78% for HTL7, and 90% for SMAX1), enabling us to assess the dynamics of all components (Fig. 5A, Supplementary Figs. 14, 15 and Supplementary Data 1).
A Each black bar represents a peptide of HTL7 identified in hydrogen-deuterium exchange (HDX) coupled with mass spectrometry (HDX-MS) experiments. The difference in HDX of pairwise comparisons between HTL7-containing samples is displayed on its amino acid sequences. Different tones of red and blue indicate HTL7 residues that incorporate more or less deuterium, respectively, in the first state of the pairwise comparison. A numbered list of tested conditions is provided, with each component name coloured based on the reference view in panel (D). B Deuterium-enriched peptides mapped on the open (PDB 5Z8P) and closed structure of HTL7 from this study (PDB 9KKX). C Protected regions of HTL7 + GR24 and MAX2 in complex with SMAX1 mapped on the structure (PDB 9KLV). The black coloured regions had no peptide coverage. D Reference view (left) of ASK1–MAX2–HTL7–SMAX1 (EMD-62417) with a white box outlining the focal area. Interface of MAX2–HTL7–SMAX1D2 (middle, 9KLD) and OsD3–AtD14–SMXL7D2 (right, PDB 5HZG). The SMAX1D2/SMXL7D2 (dark pink/light pink) RGKT motif and interface residues with HTL7/AtD14 are shown as sticks and coloured grey. HTL7/AtD14 are shown as a surface and coloured by electrostatic potential from red (negatively charged) to blue (positively charged). MAX2/OsD3 are coloured green.
GR24 treatment induced pronounced HDX deprotection in HTL7 compared to the DMSO control (Fig. 5A, B and Supplementary Fig. 14A, B). The highest deprotection occurred in the lid domain and in the structural elements that support the loops carrying the catalytic residues D217 and H246 (Fig. 5B). Thus, exposure to GR24 alone increased the malleability of HTL7 already in the absence of MAX2, in particular in these structural elements that are conformationally changed when bound to MAX2. Samples were preincubated with SL, and the HDX data collection proceeded over 24 h, inferring that the measurement captured HTL7 post SL hydrolysis (“Methods”). The GR24-induced flexibility is consistent with our model and with prior reports of SL-induced thermal destabilisation of D14 and HTL729,30.
HDX of HTL7, and thus the increased flexibility, was partially suppressed by MAX2, and further reduced in the presence of SMAX1D2, with the strongest protection observed when full-length SMAX1 was present. The presence of MAX2 and SMAX1 stabilised the HTL7 lid domain, particularly the helices that change between the open and closed lid conformations, in line with our structural model (Fig. 5B, C). These results confirm that the D2 domain stabilises GR24-bound HTL7 on MAX2 (Fig. 5A–C and Supplementary Fig. 14A, B). MAX2 exhibited reduced HDX in the presence of GR24–HTL7, with further protection upon addition of SMAX1, and to a lesser extent, SMAX1D2 (Supplementary Fig. 14C, D). On MAX2, the region stabilised involved the LRRs 17-20, including the loops 17 and 18, which were associated with SMAX1D2 and HTL7 in our structural model.
In contrast, SMAX1 (full-length or D2) showed no significant HDX changes upon complex formation, consistent with a transient and dynamic mode of interaction (Supplementary Fig. 15A, B). Finally, ASK1 displayed no detectable HDX changes in any of the tested complexes, in agreement with the structural data (Supplementary Fig. 15C, D).
Determinants of specificity
Different hormone-bound receptors recruited to MAX2 elicit distinct biological outcomes32. In our structural analysis, only the D2 domain of SMAX1 makes direct contact with the HTL7 receptor. Although the interface is relatively small (~ 430 Ų), homology modelling suggests that these contacts are more complementary within cognate pairs—SMAX1–HTL7 and SMXL7–D14—than across non-cognate combinations (Fig. 5D), indicating that D2–receptor compatibility may contribute to substrate specificity.
However, specificity is not absolute in vitro SMXL6 and SMXL7, both considered D14-specific substrates, were also able to co-precipitate MAX2 with GST-HTL7 in the presence of GR24 in our pull-down assay (Supplementary Fig. 16A). Conversely, the homologous complexes OsD3–HTL7 and AtMAX2–HTL7 showed stable association in SEC even in the absence of SMAX1 homologues (Supplementary Figs. 1A–C and 16B). In addition, the non-canonical strigolactone analogue MP366 was markedly less effective at promoting complex formation between MAX2–HTL7 and OsD3–HTL7 (Fig. 1B and Supplementary Fig. 1A–C), suggesting that both receptor–ligand and receptor–substrate interactions contribute to specificity, and that these vary across paralogues and hormone isoforms.
Our structural data also show that auxiliary interactions involving the SMAX1 N domain help stabilise the D2-mediated association with MAX2 and HTL7, suggesting that the N domain contributes to overall complex stability. A BLAST search and structural modelling for A. thaliana SMAX1 homologues in the S. hermonthica proteome identified three orthologs—ShSMXL1 (47% identity, UniProtKB A0A9N7R3H5), ShSMXL2 (39%, A0A9N7NT54), and ShSMXL3 (56%, A0A9N7RQY7)(Supplementary Figs. 17 and 19). Notably, the N domain is more conserved between SMAX1 and these Striga orthologs (91%, 71%, and 87% identity, respectively) than between SMAX1 and A. thaliana SMXL6 (47%), SMXL7 (47%), or Oryza sativa D53 (45%) (Supplementary Fig. 17). Considering that the SMXL N domains are sufficient to specify developmental functions63, at least in part through interactions with cognate transcription factors, these similarities may indicate similar biological roles. However, the regions conserved across the N domains of SMAX1, ShMXL1, 2, and 3 also include the helices H3’ and H4’ used by SMAX1N to associate with MAX2 (Supplementary Fig. 17A), suggesting conserved MAX2 association. Structural modelling of MAX2 complexes further indicates that while the N domain sequences of SMXL6 and SMXL7 are compatible with MAX2 binding, they may differ in affinity or dynamics (Supplementary Fig. 18A, B). This is consistent with previous HDX data showing that the N domain of OsD53 associates with OsD352, supporting the idea that N domain–MAX2 interactions are conserved across orthologous signalling pathways in related species. Together, these findings suggest a model where the DNA-associated SMXL binding partners govern the gene transcription pathways to be controlled, whereas the SMXL interaction with the SCF–receptor complex dictates its ubiquitination efficiency. Therein, the specificity of the SCFMAX2–receptor–substrate complex is not dictated by a single interaction, but rather emerges from the cumulative stability provided by multiple, coordinated contacts.
SMAX1 dynamically stabilises SCFMAX2 for ubiquitination
Our data revealed a hinge movement in the positioning of ASK1 relative to MAX2 across different cryo-EM classes (Fig. 6A). In the class only showing density for ASK1 and MAX2, the tip of an extended helix-turn-helix element of ASK1 (residues 74–81) contacts the MAX2 LRR11 and LRR12 opposite of its root interaction with the F-box domain (residues 35–77) and first LRR (residues 78–95) of MAX2 (Supplementary Fig. 6E). In this conformation (which we call the “resting state” of ASK1), the C-terminal helix element (CTH) of MAX2 (residues 704–728) was not visible, suggesting it was fully disengaged from the LRR domain (Supplementary Fig. 6F, G). Conversely, in the complexes with SMAX1 and HTL7, ASK1 is pulled backwards, losing the contact to the LRRs (the “retracted state” of ASK1). In addition, the cryo-EM density of the ASK1 helix-turn-helix tip loses definition, indicative of increased dynamics (Fig. 6A, right), and the CTH becomes visible in its “engaged” state50 within the LRR domain (Fig. 6B). In the crystal structure of ASK1–OsD3–AtD14 (PDB 5HZG), ASK1 is also retracted, and the CTH is engaged, suggesting that this conformation is caused by the presence of the SL receptor (D14/HTL7) (Supplementary Fig. 6A).
A Cryo-EM density reconstructed from the tail clusters of one variability eigenvector from 3D variability analysis (3DVA). Proteins and domains are coloured as in Fig. 1. Arrows depict ASK1 motion and correspond to Supplementary Movie 1. B Same as in panel A, but rotated 90°. The MAX2 CTH is coloured blue and further indicated with a white arrow. C, D Model of a complete A. thaliana SCF–HTL7 ubiquitination complex in the resting (panel C) and retracted (panel D) ASK1 state. The ASK1–MAX2–HTL7–SMAX1 complex is coloured as in Fig. 1. CUL1 is coloured white, its Nedd/RUB modification is coloured pale pink. RBX1A is coloured light blue. UBC8 is coloured bright yellow. Ubiquitin is coloured orange. Arrows show the relative motion between resting and retracted states. E The transposition of SMAX1D2 in the resting (dark pink) and retracted (light pink) ASK1 conformations and the proximity to the E2 active site and ubiquitin. The conserved lysine residues of SMAX1D2 are coloured grey, shown as sticks, and labelled. The distance between resting and retracted states are shown as dotted lines and labelled. The gradient arrow indicates the relative motion between the two states. The E2 active site cysteine is coloured grey and shown as sticks. F Model of the 2:2 ASK1–MAX2 dimer fit in cryo-EM density (EMD-62414). Coloured as in Fig. 1.
A 3D variability analysis of the ASK1–MAX2–HTL7–SMAX1 particle stack revealed that both ASK1 retraction and CTH engagement correlate with the presence of SMAX1N density, suggesting that SMAX1N helps stabilise this conformation (Fig. 6A, B and Supplementary Movie 1, 2). In the cryo-EM models based on Class 1 (with maximal SMAX1N density), arginine residues of SMAX1N are positioned to form ionic interactions with ASK1 glutamic acids, as described above (Fig. 4E). Comparison of the apo ASK1–MAX2 structure with the Class 1 complex shows a 1.5–2 Å shift of the LRR1 and LRR2 helices toward HTL7 (Supplementary Fig. 6G), further supporting the SMAX1N-promoted formation of contacts between the MAX2 CTH and the C-terminal helix of ASK1 (residues 144–160) in the engaged/retracted state.
We next wanted to assess the impact of ASK1 movements on the positioning of the SMAX1 substrate within the full E2–SCFMAX2–receptor–substrate complex. Using template-based modelling on PDB 8RX067 (“Methods”), we produced the Arabidopsis complex formed by UBC8 (E2), RBX1A, CUL1, and ASK1. We then superimposed the experimental ASK1–MAX2–HTL7–SMAX1 complex onto the SCF scaffold in both the resting and retracted ASK1 states (Fig. 6C, D). In this context, the SMAX1 D2a subdomain is closest to Ub-conjugated E2, suggesting that ubiquitination occurs preferentially on this subdomain. Several lysines cluster in the D2a region facing the E2 ligase (K597, K610, K643 and K650), providing likely candidates for ubiquitination (Fig. 6E), based on the observation that SCF ligases ubiquitinate E2-proximal lysines67. Ubiquitinable lysines are placed in the corresponding region in the D2 domains of other SMXLs, suggesting conservation of the mechanism (Supplementary Fig. 19A, B). The hinging motion of ASK1 between resting and retracted states is amplified through the CUL1 lever arm, displacing the SMAX1 D2 domain by up to 7 Å relative to the E2. The retracted state places the D2 domain further from the Ub-loaded E2, which is counterintuitive as ubiquitination is driven by proximity. However, the transient binding of the SMAX1 N domain and associated conformational changes in ASK1 would result in tapping motions and positional variations that may be required for polyubiquitination to occur.
ASK1–MAX2 dimers
Although we did not observe dimerisation on SEC, our cryo-EM dataset revealed a dimeric ASK1–MAX2 class, representing a third of all particles (Fig. 1D and Supplementary Fig. 2). Native PAGE gels showed a weak band corresponding to a dimer, alongside a strong monomer band (Supplementary Fig. 4A). Mass photometry revealed a mixed population of monomers and dimers in vitro (Supplementary Fig. 4B). Despite achieving a nominal maximum resolution of 3.3 Å in 3D reconstructions, detailed structural modelling was hindered by strong anisotropy in the maps due to preferred particle orientation (Supplementary Fig. 4C–E). AI-based correction for anisotropy without misalignment correction (spIsoNet) led to minor improvements in map interpretability. In the resulting reconstruction, obtained without imposing a C2 symmetry, the map coverage of the two ASK1 was unequal (Fig. 6F). However, a C2-symmetric ASK1–MAX2 dimer model generated by AlphaFold fitted the map well, and was adjusted through rigid-body fitting (Fig. 6F). The resulting model suggested that dimerisation arises from a C2 symmetric association between the hydrophobic N domain-binding patches on LRR1 and LRR2 of MAX2, with additional charge-charge compensation between R86 from LRR1 and E146, E147, and E150 from the ASK1 C-terminal helix (Fig. 6F and Supplementary Fig. 4F, G). We did not observe density for the MAX2 CTH in the dimer, suggesting it is dislodged as in the monomeric apo-ASK1–MAX2 complexes. The overlap between the dimer interface and the N domain binding site suggests competition between dimerisation and SMAX1 binding, potentially reducing the number of SMAX1-containing ASK1–MAX2(–HTL7) particles in vitro.
Additional analyses presented both supporting and opposing evidence regarding the biological relevance of ASK1–MAX2 dimers. While analysis of additional cryo-EM data and PDBePISA interface analysis support a potential ASK1–ShMAX2 dimer, several lines of evidence—including lack of conservation across orthologues, condition-dependent dimer formation, and absence of dimers in rice D3 crystal structures—argue against a conserved or functionally relevant dimer (Supplementary Discussion, Supplementary Fig. 5).
Discussion
The mechanism by which SL binding and hydrolysis by their D14/HTL receptors leads to the ubiquitination of transcriptional repressors by the SCFMAX2 E3 ligase has remained elusive due to the transient and dynamic nature of the interactions involved. By identifying a sufficiently stable inter-species complex and preparing cryo-grids within the SCFMAX2 signalling time window, we were able to obtain cryo-EM structures that reveal the conformational and compositional landscape of the SL signalling complex. This complex contains A. thaliana SMAX1 as the substrate, rather than its S. hermonthica orthologue. Several lines of evidence support the relevance of this interspecies complex in elucidating gemination-related SL signalling. Functionally, AtSMAX1 phenocopies its Striga counterpart: it is responsive to SLs, undergoes degradation in a D14-dependent manner31,68, and acts as a repressor of seed germination and seedling development63,69. Furthermore, SMAX1 can form an SL-induced complex with both Arabidopsis and Striga MAX2 proteins via HTL754,55, and HTL7 expression in a kai2 mutant Arabidopsis restores SL-dependent germination through SMAX1 inactivation41,68. Finally, we identified close Striga homologues to AtSMAX1, based on sequence conservation and structural modelling (termed ShSMXL1, 2 and 3, see Supplementary Figs. 17 and 19), suggesting functional conservation. Nonetheless, future structural analyses of native ASK1–MAX2/D3–D14/HTL7–SMXL6,7,8/D53 complexes are essential to fully elucidate the SL signalling mechanism.
Our structural and biochemical findings further elucidate several aspects of SL signalling, which have remained controversial. Among these are the nature of the SL cleavage product associated with the SCFMAX2–D14/HTL–substrate complex, and the role and timing of SL cleavage during SCFMAX2–D14/HTL signalling. Our HDX data show that exposure to GR24 is sufficient to destabilise HTL7, in agreement with computational and biochemical evidence29,30,58,70, whereas the added presence of MAX2 and SMAX1 increasingly stabilises HTL7. Our structures reveal that the HTL7 lid domain closes to dock onto the LRR of MAX2, showing that the structural transition previously observed for D14 in a complex with ASK1–OsD3 (PDB 5HZG)28 is conserved among receptor homologues. However, our structures resolve the receptor D-loop, which could not be modelled in the 3.3 Å crystal structure of the ASK1–OsD3–AtD14 complex (PDB 5HZG). The remaining space in the HTL7 ligand binding pocket is too constricted to accommodate a non-hydrolysed SL, challenging the hypothesis that an intact SL could occupy the space of a disengaged D-loop29.
Our structures also resolve the nature of the SL cleavage product bound to HTL7 in the signalling state. In the initial crystal structure of the ASK1–OsD3–AtD14 complex (PDB 5HZG), features of the electron density led the authors to conclude that the SL cleavage product is the open D-ring covalently bound to both the catalytic Ser and His of D1428. However, subsequent reanalysis of these data by other researchers refuted this claim, suggesting that the observed electron density rather corresponds to either an iodide or a closed D-ring bound to the catalytic His46,47. Our findings strongly support that the closed D-ring, covalently attached to the catalytic His246, is the receptor-associated SL cleavage product during the signalling stage. This visual conclusion is in line with previous biochemical evidence28,55,57. Importantly, modelling shows that attachment of the D-ring in the apo (lid-open) HTL7 confirmation is not possible, as it would lead to clashes with the Ser95 residue of the catalytic triad. In the MAX2-bound lid-closed structure, the catalytic triad of HTL7 is distorted and can accommodate the D-ring modification of His246. This distortion, however, separates the three catalytic residues sufficiently to block catalysis. Thus, MAX2, SMAX1 and the covalently His246-linked closed D-ring jointly retain the receptor in a catalytically inactive conformation. Collectively these findings substantiate that SL hydrolysis precedes, and is required for, SCFMAX2–receptor–substrate complex assembly. This “hydrolysis-first” model is consistent with literature showing that catalytic triad mutants in AtD14 (e.g., S97C, S97A, H247A) are inactive in planta and fail to form signalling complexes or undergo D-ring modification28,29,57. Similar results have been reported for HTL homologues, including HTL7S95A and HTL5S95A/H246A 54,71. Conversely, catalytic D217 mutations produced mixed results in the literature. Whereas AtD14D218A has been reported to retain signalling activity in some contexts29, HTL5D217M did not induce germination in another study71. In our hands, HTL7D217A was unstable and unresponsive to SL, consistent with prior reports of instability in the corresponding AtD14D218A variant29. A plausible explanation is that D218A partially mimics the receptor lid destabilisation normally induced by SL hydrolysis, potentially enabling hydrolysis-independent signalling under certain conditions.
This hydrolysis-first mechanism does not rule out a biological function of lid-open D14/HTL bound to an intact SL29, but that function would need to occur prior to fully engaging the MAX2 LRR surface. For instance, previous studies revealed that the MAX2 CTH can transition from a helical conformation associated with MAX2 LRRs, through a more flexible intermediate, to a fully dislodged state50,51. These findings are compatible with our ASK1–MAX2 structures, where the CTH is unresolved/dislodged in the absence of the receptor and substrate, and present as a helix in the assembled SCFMAX2–HTL7–SMAX1 complex. Therefore, our results are compatible with an initial encounter complex between the dislodged MAX2 CTH and a non-collapsed SL-bound receptor50,51, which then transitions to the fully assembled SCFMAX2–receptor–substrate complex where CTH is engaged with MAX2. An interesting aspect for future research is to determine how small non-hydrolysable compounds such as KAR1, DCL and Cyclo(Leu-Pro) act through KAI2 and related receptors72,73,74.
Unexpectedly, our cryo-EM data contained a significant amount of ASK1–ShMAX2 dimers. However, the evidence for dimerisation was ambiguous and dependent on sample preparation and the analytical approach used. Moreover, our analysis does not support dimerisation of Arabidopsis or rice MAX2 orthologues, collectively arguing against a biological role of MAX2 dimerisation.
Our analysis provides insights into SMAX1, shedding light on how the SMXL family has diverged from hexameric AAA + protein to fulfil their roles in transcriptional regulation and protein–protein interactions. We show that SMAX1 uses its N and D2 domains to engage in a dynamic cooperative association with SCFMAX2–HTL7. The involvement of the D2 domain in the SCFMAX2–SL signalling complex is strongly supported by various lines of evidence50,52,60, whereas support for the involvement of the N domain was only weak52,63. The SMAX1N domain (and possibly an additional unidentified SMAX1 region) binds to a MAX2 surface close to ASK1, while SMAX1D2 engages with HTL7 and another site of the MAX2 LRR. For this role, SMAX1D2 has been repurposed from an ATPase domain into a binding module through the loss of its nucleotide binding site and the use of its pore loop for allosteric stabilisation of the association between the F-box and the SL-bound receptor. This pore loop has acquired an RGKT motif that is conserved in other SMXL family members. Our data reveal that the conserved RGKT motif plays a dual role in the complex, by mediating direct contacts between SMAX1D2 and MAX2, and by allosterically stabilising MAX2 LRR17 in its HTL7-binding position. In D53, the rice orthologue of SMAX1, deletion of this motif abrogates D53 ubiquitination and degradation34,35. The occluded positioning of the RGKT motif in the ASK1–MAX2–HTL7–SMAX1 complex makes it an unlikely ubiquitination site, suggesting that its loss impairs D53 ubiquitination by destabilising the complex. Results from our semi-quantitative pull-down assays show only a slightly reduced capability of SMAX1D2∆RGKT to stabilise the complex (Fig. 3F), and full-length D53∆RGKT was still able to interact with D14 in pull-down assays by Jiang et al.34. Thus, we speculate that the effects of the RGKT deletion on the stability of the D2 domain, and/or the SCFMAX2–receptor–substrate complex are too subtle to produce strong effects in vitro, but reduce the longevity and selectivity of the complex sufficiently to decrease ubiquitination efficacy in planta.
Indeed, a striking aspect of the SCFMAX2–receptor–substrate complex is that it is composed of individually weak associations that jointly produce a cooperative complex stable enough to persist over SEC. The mutually stabilising association between SMAX1D2, MAX2, and HTL7 is not the only instance of this mechanism. Also, SMAX1N and SMAX1D2 appear mechanistically connected through the MAX2 CTH in its “engaged” helical form, although both domains are separated by more than 10 Å when bound to MAX2. This allosteric connection may explain the cooperative in-trans binding of SMAX1N and SMAX1D2 to the SCFMAX2–HTL7 complex (see Supplementary Fig. 13B).
Another, related, aspect is the highly dynamic nature of the complex. Our structural and biochemical analyses show that the weak individual association of SMAX1N and SMAX1D2 with MAX2–HTL7 fluctuate around the bound position and may temporarily detach from the complex. On one hand, the lack of rigid structural coupling between the N and D2 domains allows the bidentate interaction to avoid excessive affinity gains or structural rigidity through avidity, while achieving overall higher affinity and specificity due to increased local concentrations of one domain when the other is bound. On the other hand, these positional fluctuations may also directly enhance the ubiquitination efficacy: we show that the presence of density for SMAX1N correlates with the retraction of ASK1 from its “resting” position on MAX2, inferring that the presence of the N domain modifies the position of ASK1. Within the fully assembled E2–E3–substrate complex, where ASK1 movements are amplified by the long cullin lever arm, these N-domain binding dynamics would result in a tapping encounter between the ubiquitin-conjugated E2 and the D2 domain (Fig. 7). We hypothesise that these structural dynamics are essential for efficient substrate polyubiquitination. Thus, our analysis provides a mechanistic explanation for previous indirect observations that dynamics in the SCFMAX2–receptor–substrate complex are crucial for substrate polyubiquitination and release48,50,51,75.
In the pre-signalling state (left), the MAX2 CTH is dislodged, as seen in apo-ASK1–MAX2 (PDB 9KLL). Strigolactone perception and hydrolysis enable HTL7 to close and bind to MAX2, trapping the D-ring in the active site. This HTL7 conformation creates the interface for SMAX1D2 to mutually stabilise the tripartite MAX2–HTL7–SMAX1D2 complex. Simultaneously, SMAX1N engages another site on the MAX2 LRR, close to the MAX2 CTH and ASK1, promoting the “retracted” position of ASK1. The transient interactions of SMAX1N lead to a tapping motion of the E2 towards SMAX1D2, while the interaction dynamics of SMAX1D2 with MAX2 lead to positional variations towards the E2. Together, these oscillating interactions flexibly position SMAX1D2 for polyubiquitination, leading to proteasomal degradation of SMAX1, thereby enabling transcription to occur.
An important question is how the affinity and dynamics of the complex cause specific outcomes. Although D14 and HTLs (and the homologous KAI2) engage SCFMAX2, they trigger different biological outcomes, with only HTL/KAI2 being associated with germination43,74,76. Previous work evidenced a level of selectivity of the SCFMAX2–D14 for the substrates SMXL6, SMXL7, and SMXL8 in response to SL, whereas SCFMAX2–HTL/KAI2 preferentially targets SMAX1 and SMXL2 in response to karrikins32,60. However, cross-talk between D14 and SMAX1 was reported, demonstrating that both SMAX1 and SMXL2 can be targeted for degradation via D14 in response to SL31,68. Despite significant efforts from several groups, the determinants for specificity remained ill-defined and controversial, with N, D1, M, D2 domains and combinations thereof, proposed as key elements35,50,60. Moreover, in many experiments, the selection of one substrate over another appears preferential rather than stringent. Indeed, we observed that GR24-supplemented GST-HTL7 co-precipitated ASK1–MAX2 not only in the presence of SMAX1 but also in the presence of the D14-specific SMXL6 and SMXL7. Our structural analysis showed that only the D2 domain of SMAX1 contacts HTL7, with better complementarity in cognate pairs like SMAX1–HTL7 and SMXL7–D14, but only with a modest interface area (430 Ų). Although our structures were prepared with full-length SMAX1, we did not see another density in contact with HTL7 to indicate additional determinants of receptor specificity. The only non-attributed density is adjacent to the N domain, away from the D14/HTL7 binding site on MAX2. The N domains of SMXLs have been shown to be involved in developmental control in A. thaliana63. Our analyses support that the N domains of AtSMXL6 and 7 can associate with MAX2 in the same way, but with altered affinities and dynamics.
Moreover, the association between SMAX1 and SCFMAX2–receptor complexes may directly compete with the association between SMAX1 and DNA/transcriptional regulators. Chang et al. reported that N-terminal domains of SMXL-family proteins interact with TCP-family transcription factors to repress gene expression63. Interestingly, AlphaFold predictions suggest that the surface of the SMAX1 N domain used for MAX2 binding is also involved in TCP interaction (Supplementary Fig. 11B, C). Hence, the bidentate association with MAX2 may need to be strong enough to disengage SMAX1 from DNA and TCP family proteins, to release gene repression.
Together, these findings support a model in which DNA-associated SMXL partners define transcriptional outcomes, while SMXL interactions with the SCF–receptor complex govern ubiquitination efficiency. The specificity and function of the SCFMAX2–receptor–substrate complex are not only determined by the receptor–substrate match, but rather emerge from the combined affinities and cooperative potential of multiple, coordinated contacts. It is this cumulative stability and dynamic interplay that ultimately dictate substrate recruitment, ubiquitination, and biological specificity.
Through this mechanistic framework, the SL perception machinery may also be sensitive to additional regulatory layers, such as the controlled co-expression of the dozens of proteins involved in substrate ubiquitination and degradation77, small molecules (e.g., citrate or inositol phosphates)19,48,51 or post-translational modifications78. The complexity of the SCFMAX2-based SL perception, which, remarkably, occurs in Striga only after a decade-long seed dormancy and conditioning, may reflect the level of signal integration required for Striga to commit to germination only in the presence of a host and under favourable conditions.
In summary, our work fills an important knowledge gap by capturing the long-elusive SCFMAX2–receptor–substrate complex that mediates SL perception. Our results help rationalise and resolve previous observations pertaining to the mechanism of SL perception, and may inspire chemical or genetic interventions to control Striga germination. In addition, these findings highlight the importance of finely tuned dynamics in E3-based hormone perception (Fig. 7). These insights significantly advance our understanding of how E3 ligases are used in plants to translate hormone perception into genetic adaptations.
Methods
Plasmid construction
The Multibac™ expression system from Geneva Biotech was used to co-express in insect cells each of the full-length proteins OsD3, AtMAX2, and ShMAX2 with AtASK1. The genes encoding OsD3 (UniprotKB Q5VMP0), AtMAX2 (UniprotKB Q9SIM9), ShMAX2 (UniprotKB T1RVG4) and AtASK1 (UniprotKB Q39255) were codon optimised for insect cells expression, and ordered from Twist Bioscience, with an N-terminal cleavable tandem 9xHis tag and Strep-tag for the AtMAX2 and ShMAX2 inserts. OsD3 insert was amplified with an N-terminal cleavable Strep-tag and a C-terminal 6xHis tag and was cloned in the pIDK donor vector. The AtMAX2 and ShMAX2 inserts were cloned in the pACEBac1 acceptor vector, and the untagged AtASK1 insert was cloned in both the pIDS and pACEBac1 vectors. The single transfer vectors with different MAX2 and AtASK1 inserts were generated using Cre recombinase according to the MultiBac™ expression system user manual, and the recombinant transfer vectors were transformed into DH10EMBacY™ cells, with a constitutively expressing YFP expression cassette. The resulting bacmid DNA was isolated by isopropanol precipitation.
Full-length AtSMAX1 gene (UniprotKB Q9FHH2) was codon optimised for E. coli expression, and ordered from Twist Bioscience and cloned into the pQlinkH vector (with an N-terminal 7xHis tag) by restriction cloning between the BamHI and NotI restriction sites. An additional C-terminal Strep-tag was added by overhang PCR amplification with phosphorylated primers, followed by ligation. AtSMAX1 mutants were ordered from Twist Bioscience with an N-terminal 7xHis tag and a C-terminal Strep-tag in the pJEx411c vector. The corresponding truncations of AtSMAX1 wild-type and mutant constructs were generated by overhang PCR amplification with phosphorylated primers, followed by ligation. AtSMXL6 (UniprotKB Q9LML2) and AtSMXL7 (UniprotKB O80875) were codon optimised for E. coli expression, and ordered from Twist Bioscience with an N-terminal tandem Strep and a C-terminal 7xHis tag in the pJex411c vector. ShHTL7 gene (UniprotKB A0A0M3PNA2) cDNA59 was cloned into pQlinkH and pGEX6P-1 (with an N-terminal GST tag) vectors by restriction cloning between the BamHI and NotI restriction sites. The ShHTL7 mutants were ordered from Twist Bioscience in the pGEX6P-1 vector. All primers used for cloning were obtained from Integrated DNA Technologies (IDT) and are listed in Supplementary Data 2.
Protein preparation
For insect cell expression, Sf9 cells (Novagen Merk Millipore, 71104-3) were cultured in ESF 921 medium (Expression Systems) at 27 °C. The bacmid DNA was transfected into Sf9 cells in adherent culture using FuGENE® HD (Promega), and the YFP fluorescence was monitored for baculovirus production. The baculovirus was amplified twice in suspension culture to obtain a higher viral titre. For the expression of the MAX2–ASK1 proteins, the amplified virus was used to infect Sf9 cell cultures at a density of 2 × 106 cells/mL. The cells were harvested at 70–80% cell death post-infection within 66–72 h by centrifugation, and the pellet was gently washed with PBS or purification buffer and was stored at −80 °C if purification was not done immediately after.
The AtSMAX1 plasmid was transformed into E. coli BL21 (DE3) cells grown on agar plates with ampicillin selection. A single colony was used to grow an overnight starter culture in 2YT medium. For expression, flasks containing 1 L culture media were inoculated with the starter culture and the cells were grown at 37 °C and 200 rpm to reach the OD600 of 0.6, when protein expression was induced by adding isopropyl β-D-1-thiogalactopyranoside (IPTG) to 0.2 mM at 16 °C for 12–16 h. A similar protocol was used for expressing all the AtSMAX1 constructs, AtSMXL6, AtSMXL7, ShHTL7 and its corresponding mutants from BL21 (DE3) cells. LB media was used for ShHTL7 expression, and kanamycin selection was used for the constructs in the pJEx411c vector.
For all MAX2 and AtSMAX1 proteins, purification was done on ÄKTA pure™ systems, first by affinity purification on HisTrap™ HP 5 mL (Cytiva) column, followed by StrepTrap™ XT 5 mL (Cytiva) column and then by size-exclusion chromatography (SEC) on HiLoad 16/600 Superdex 200 pg (Cytiva). AtASK1 co-eluted with all MAX2 proteins after all the purification steps. Cell pellets were first homogenised in binding buffer (50 mM Tris-HCl, pH 7.5, 500 mM NaCl, 1 mM TCEP, 20 mM imidazole, 4% (v/v) glycerol) with protease inhibitors (SIGMAFAST™ EDTA free) and benzonase (MilliporeBenzonase® Nuclease) by gentle sonication or using a manual cell homogeniser, and then lysed using a cell disruptor. The lysate was clarified by centrifugation at 50,000 × g and passed through the His-affinity column equilibrated with binding buffer, and the protein was eluted with 250 mM imidazole after an initial washing step with 40 mM imidazole. The protein fractions were then loaded on the Step-affinity column equilibrated with binding buffer (with adjusted imidazole composition) and eluted after washing with 50 mM D-(+)-Biotin (Thermo Scientific) after 1 h incubation on the bead. The protein fractions were injected into the SEC column equilibrated with 20 mM HEPES, pH 7.5, 200 mM NaCl, 1 mM TCEP and 4% (v/v) glycerol (for AtSMAX1 300 mM NaCl was used), and the pure fractions were concentrated and snap-frozen for storage at −80 °C. An SDS-PAGE gel was run after each purification step. All purification steps were performed at 4 °C and as quickly as possible, particularly for AtSMAX1, which showed signs of degradation.
His-tagged and GST-tagged ShHTL7 (and mutants) were purified by gravity flow chromatography in a glass Econo-Column® from Bio-Rad. Cell pellets expressing His-tagged ShHTL7 were lysed by sonication in binding buffer containing 50 mM Tris-HCl, pH 8.0, 200 mM NaCl, 1 mM DTT, 20 mM imidazole with 0.1% (v/v) Tween-20 added, and the clarified lysate was loaded onto Ni-NTA Agarose resin (QIAGEN). The resin was washed with binding buffer containing 40 mM imidazole and eluted with binding buffer containing 250 mM imidazole. The eluted protein fractions were further purified on the SEC column equilibrated with 20 mM HEPES, pH 7.5, 150 mM NaCl, and 3 mM DTT. Protein purity was evaluated using SDS-PAGE. The purified proteins were concentrated and stored at −80 °C. The GST-tagged ShHTL7 was purified similarly using 50 mM Tris-HCl, pH 8.0, 200 mM NaCl, 1 mM DTT, with 0.1% (v/v) Tween-20 added for lysis, and loaded on Glutathione Sepharose 4B (Cytiva) resin. After washing with the binding buffer, the protein was eluted with 20 mM reduced glutathione (VWR) in the binding buffer. The eluted fraction was diluted in 20 mM HEPES buffer to 100 mM NaCl and further purified by anion exchange on HiTrap Q HP column (Cytiva), followed by SEC on HiLoad 16/600 Superdex 75 pg (Cytiva). The pure fractions were concentrated and snap-frozen for storage at − 80 °C.
Size-exclusion chromatography (SEC) assay
Purified ShMAX2–AtASK1 (17 μM), ShHTL7 (25 μM) and AtSMAX1 (20 μM) in 150 μL reaction volume were incubated at 4 ˚C for 30 min with 200 μM GR24 or an equal amount of DMSO as the solvent control, in a buffer containing 20 mM HEPES pH 7.5, 200 mM NaCl, 1 mM TCEP. The reactions were injected into a Superdex 200 10/300 column (Cytiva) for analysis at a flow rate of 0.5 mL/min. The fractions were analysed by SDS-PAGE and visualised by Coomassie staining. For testing the interaction of other MAX2–HTL(–SMXL) components, the proteins were mixed in a 1:1.5:1.2 molar ratio to a final 150 μL volume with roughly 0.25 mg of the MAX2 protein, and corresponding amounts of the binding partners, following the same procedure as above.
Pull-down assay
For testing the ShHTL7 interaction with ShMAX2–AtASK1 and AtSMAX1 (full length, D2 domain and corresponding mutants) or AtSMXL6/AtSMXL7 using GST-ShHTL7 as bait, reaction mixtures of 100 µL in 20 mM HEPES pH 7.5, 200 mM NaCl, 0.5 mM TCEP buffer were prepared for a final concentration of roughly 3.5 µM GST-ShHTL7 and 5 µM for each of the tested binding partners. Purified GST-ShHTL7 was first bound to beads (Glutathione Sepharose 4B resins), and the beads were aliquoted to each reaction mixture. The reactions were incubated with 100 µM GR24, MP3 or their solvent DMSO as the control for 30 min on a rotator at 4 °C and protected from light. The beads were then rinsed with 150 µL buffer and washed three more times with 5 min incubation on the rotator, before elution with 40 µL buffer with 20 mM glutathione by incubation for 20 min. The washes were monitored by Bradford assay (Bio-Rad) and continued until no protein was visibly detected in the flow-through. The eluted fractions were analysed by SDS-PAGE. A similar procedure was used for testing ShHTL7 interaction with OsD3–AtASK1 with GR24 and MP3. The eluted fractions were tested for OsD3 presence by western blot using anti-6xHis HRP-conjugated antibody (Abcam). For testing OsD3–AtASK1 interaction, ShHTL7 using Strep-tagged OsD3 as bait, 200 µL reaction mixtures (in the same buffer) of the two proteins of roughly 10 µM OsD3–AtASK1 and 17 µM ShHTL7 were prepared and incubated for 30 min with GR24 and MP3, and then loaded to Strep-Tactin® XT beads and incubated for another 30 min. Washes were performed as above, and the proteins were eluted in buffer with 50 mM biotin, and the fractions were analysed by SDS-PAGE.
Cryo-EM specimen preparation
Cryo-EM grids (Quantifoil R1.2/1.3, 400 mesh Au or UltrAuFoil, R1.2/1.3, 300 mesh) were washed with acetone for 30 s, isopropyl alcohol for 10 s and left to dry. Washed grids were then glow-discharged in air using either a Quorum GloQube (60 s at 35 mA current) or a Pelco easiGlow (30 s at 30 mA current). 3 µL of the sample (AtASK1–OsD3–ShHTL7 at 0.5 mg/mL or AtASK1–ShMAX2–ShHTL7–AtSMAX1 at 0.35 mg/mL) was applied to freshly glow-discharged grids, blotted for 2-3 s and plunge-frozen using a TFS Vitrobot Mark IV at 4 ˚C and 100% humidity. Samples with detergent were mixed with detergent (final 0.005% (v/v) Tween-20) just before vitrification.
Cryo-EM data collection
Cryo-EM data were recorded on a TFS Krios G4 operated at 300 kV, equipped with a Selectris X energy filter and Falcon 4i direct detector. For the AtASK1–ShMAX2–ShHTL7–AtSMAX1 dataset, 20,175 EER movies were collected at 165k magnification, yielding a pixel size of 0.73 Å/pixel at the specimen level. A flux of 7.8 e-/pixel/s resulted in a total fluence of 44 e-/Å2 on the specimen, over a 3 s exposure time. Data were collected using aberration-free image shift (AFIS) of +/- 12 µm in EPU (v3.5) with a nominal defocus range − 2.7 µm to − 1.5 µm in 0.3 µm intervals and an energy filter slit width of 10 eV, with automatic re-centring of the zero-loss peak every 2 h.
For the AtASK1–OsD3–ShHTL7, untilted data set of 6,741 movies were recorded similarly, at a flux of 4.76 e-/pixel/s, resulting in total fluence on the specimen of 40 e-/Å2 over a 4.5 s exposure time. Similar collection parameters were used for the tilted dataset (7,372 movies), except for a reduction of AFIS to +/− 6 µm, a single nominal defocus value of −2 µm, and a + 30° stage tilt.
Cryo-EM image processing
For the AtASK1–ShMAX2–ShHTL7–AtSMAX1 dataset, an overview of the data processing can be found in Supplementary Fig 2. Initial processing was performed in RELION (version 5.0-b commit-90d239)79. Movies were motion-corrected using RelionCorr with an EER fractionation of 21 and electron exposure of 1.02 e-/Å2 per fraction. CTF estimation was carried out using CTFFind480 and images were curated based on resolution (< 6 Å), defocus (− 1.0 to − 3.0 µm), and relative figure of merit, resulting in 15,184 accepted images. Particles were picked using the default model of Topaz81 on a subset of 1k images to generate references for template-based picking. After picking on the entire data set by template-based picking and doing several rounds of 2D classification-based cleaning on particles extracted with 360 pix and 4x binning, we obtained a particle stack of 1.8 m particles. The cleaned particle stack was re-extracted with 2x binning and imported into CryoSPARC (version 4.4)82. Multi-class Ab-Initio Reconstruction followed by Homogeneous Refinement and 3D Classification jobs were used to generate high-quality reference volumes for each compositional assembly. In addition, 4 decoy classes of pure noise were generated by sampling a very small number of particles using Ab-Initio Reconstruction. Concomitantly, the images from RelionCorr were imported into CryoSPARC and curated similarly. The images were over-picked using blob picking (diameter 80 Å to 170 Å) to generate a particle stack of 7.4 m particles with a box of 360 pix with 2x binning.
The over-picked particle stack with high-quality references and decoy volumes was used for iterative Heterogeneous Refinement in CryoSPARC. Only particles retained in the high-quality reference classes were used as input for each sequential Heterogeneous Refinement until >90% of particles were retained in these classes. Particles corresponding to each compositional assembly were grouped and extracted without binning, and Non-Uniform Refinement (C1 with global and local CTF refinements) was performed to generate the consensus refinements.
These consensus particle stacks were imported back to RELION using pyem83 where alignment-free 3D classification using Blush84 was used to separate different conformations. For selected classes, iterative CTF Refinement, Bayesian Polishing, and Refine3D using Blush and local searches only was performed to generate the final volumes. For Class 2 of the AtASK1–ShMAX2–ShHTL7 complex, an additional round of alignment-free 3D classification was performed with a large mask around AtSMAX1N to identify a subset of particles with the best AtSMAX1N density. The Dimer particle stacked comprised primarily side views, causing anisotropic reconstructions. Selecting classes with the highest isotropy and attempting to rebalancing orientations were unsuccessful at alleviating the anisotropy.
To improve the interpretability of maps for the initial stages of model building, unsharpened maps were subjected to the EMReady algorithm85. To improve the isotropy of the Dimer map, the unsharpened map was subjected to Anisotropy Correction without Misalignment Correction using the SpIsoNet algorithm86. The corrected Dimer map was then further modified using EMReady to improve the interpretability.
The untitled and tilted AtASK1–OsD3–ShHTL datasets were processed in RELION (version 4.0.0-commit-138b9c)79. Both data sets were processed independently to the stage of 3D classification before joining the curated particle stacks for further processing. An overview of the data processing can be found in Supplementary Fig 1. Movies were motion-corrected using RelionCorr with an EER fractionation of 33 and an electron exposure of 0.96 e-/Å2 per fraction. CTF estimation was carried out using CTFFind480, and images were curated based on resolution (<6 Å) and relative figure of merit, resulting in 5949 accepted images for the untilted and 3175 movies for the tilted datasets. The default model of Topaz81 was used to pick particles, which were extracted with a box size of 220 pix with 2x binning. Particles were curated through iterative 2D and 3D classifications, yielding a final stack of 227k particles. Refinement of this particle stack reached 4.4 Å, which was further improved to 4.3 Å after CTF refinement. Further polishing or CTF refinement did not improve the quality of the map.
Cryo-EM variability analysis
3DVA87 on AtASK1–ShMAX2–ShHTL7–AtSMAX1 was performed from the consensus refinement using a filter resolution of 6 Å and a mask containing all domains. The heterogeneity was represented along 3 eigenvectors. 3DVA Display in Simple mode was used to generate 20 volumes along each eigenvector. Movies were then generated for each volume series in ChimeraX88. To discretise the heterogeneity, 3DVA Display in Cluster mode was used to generate 5 clusters. The tail clusters from the variability distribution along each eigenvector were refined using Local Refinement with the same mask. These end-states were used to interpret the extreme ranges of heterogeneity present in the particle stack. 3DVA of apo-AtASK1–ShMAX2 did not suggest substantial heterogeneity. Due to the high anisotropy of the Dimer, 3DVA was not performed.
Model generation, fitting and refinement
Initial models were generated with AlphaFold89 and SWISS-MODEL90 using 5HZG as a template for the collapsed form of ShHTL7. Models were rigid-body docked into the cryo-EM densities using ChimeraX (v1.5)88 and flexibly fit using Isolde (v1.8)91. Iterative manual adjustments of the models were performed in Coot (v0.9.8.92)92 alongside optimisation of the geometry and fit to density through Real Space Refinement in Phenix93, including the ligand restraint file when necessary. The restraint file for the ShHTL7 ligand (D-ring) was generated using eLBOW94 after using the Ligand Builder tool in Coot92 to generate the SMILES string.
Model of the E2–SCFMAX2 E3–receptor–substrate complex
The SCF–E2–Ubiquitin scaffold was obtained by assembling Arabidopsis CUL1 and RBX1A (modelled with SWISS-MODEL90 on the template PDB 8RX0), with the ubiquitin chain (chain U) from PDB 8RX0, the Arabidopsis E2 UBC8 attached to an Ub-like molecule (chains A and B of PDB 4×57) which superimposed well (Cα RMSD 0.751 Å) with the E2 and NEDD8 of 8RX0 (chains N and G), and AtASK1 from the apo-AtASK1–ShMAX2 model. The PDB 8OR0 (structure of a HsCullin-1 complex) superimposed onto the HsCullin-2 (chain E) of PDB 8RX0 (Cα RMSD 1.224 Å) was used as a reference to position AtASK1 on the modelled AtCUL1. The experimental AtASK1–ShMAX2–ShHTL7–AtSMAX1 complex was superimposed onto the generated SCF scaffold, and the full-length AlphaFold predicted model of AtSMAX1D2 was used.
Hydrogen/deuterium exchange mass spectrometry (HDX-MS)
The individually purified ShHTL7, AtSMAX1 and ShMAX2–AtASK1 proteins were dialysed in 20 mM HEPES pH 7.5, 200 mM NaCl, 0.5 mM TCEP buffer and then mixed in equimolar ratios to reach final concentrations of 25 µM per protein in 200 µL final volume (see Supplementary Data 1). Prior to HDX-MS, the protein stock solution was supplemented with 0.8 µL of 50 mM GR24 dissolved in DMSO or a DMSO control, yielding 200 µM GR24 final concentration. This stock solution was maintained at 1 °C for the entire HDX data acquisition time of approx. 24 h. The preparation of individual HDX reactions was aided by a two-arm robotic autosampler (LEAP Technologies) as described previously95. In brief, 7.5 μL of sample stock solution were pre-dispensed in a plate maintained at 25 °C, then 67.5 μL of HDX buffer (20 mM HEPES-Na, pH 7.5, 200 mM NaCl, 0.5 mM TCEP) prepared with 99.9% D2O was added and incubated for 10, 100, 1,000 or 10,000 s at 25 °C. 55 µL of the HDX reaction were then withdrawn and added to 55 µL of quench buffer (400 mM KH2PO4/H3PO4, pH 2.2, 2 M guanidine-HCl) pre-dispensed in another plate and kept at 1 °C. After mixing, 95 µL of the quenched reaction were injected into an ACQUITY UPLC M-Class System with HDX Technology (Waters)96. Non-deuterated HDX samples were prepared similarly by 10-fold dilution of sample stock solutions with HDX buffer prepared with H2O (incubation for approximately 10 s at 25 °C). All measurements were conducted in n = 3 technical replicates (individual HDX reactions) per time-point of HDX. The protein samples were digested online with porcine pepsin and separated by reversed-phase HPLC, followed by mass spectrometric analysis of three technical replicates (individual HDX reactions) per HDX time-point as described95. In brief, injected samples were flushed out of the 50 µL-loop with H2O + 0.1% (v/v) formic acid at 100 µL/min flow rate and guided to a column (2 mm × 2 cm, 12 °C) containing porcine pepsin immobilised to beads. The generated peptic peptides were collected on an AQUITY UPLC BEH C18 1.7 µm 2.1 × 5 mm VanGuard Pre-column (Waters) kept at 0.5 °C. After 3 min of digestion and trapping, peptides were eluted at 0.5 °C by a gradient of eluents A (H2O + 0.1% (v/v) formic acid) and B (acetonitrile + 0.1% (v/v) formic acid) at 30 μL/min flow rate and separated on an ACQUITY UPLC BEH C18 1.7 μm 1.0 × 100 mm column (Waters) as follows: 0–7 min: 95-65% A; 7-8 min: 65-15% A; 8-10 min: 15% A; 10-11 min: 5% A; 11-16 min: 95% A. Eluting peptides were delivered to a G2-Si HDMS mass spectrometer with ion mobility separation (Waters), ionised by electrospray ionisation (250 °C capillary temperature, 3.0 kV spray voltage) and mass spectra acquired in positive ion mode over a range of 50 to 2000 m/z in enhanced high definition MS (HDMSE) or high definition MS (HDMS) mode for non-deuterated and deuterated samples, respectively. [Glu1]-Fibrinopeptide B standard (Waters) was employed for lock-mass correction. During separation of the peptide mixtures, the loop and pepsin column were washed three times with 80 µL of wash solution (0.5 M guanidine hydrochloride in 4% (v/v) acetonitrile), and blank injections performed between each sample to reduce peptide carry-over. HDX-MS data collection utilised the MassLynx 4.1 software (Waters).
Peptide ions were identified from the non-deuterated samples with HDMSE by the ProteinLynx Global Server 3.0.1 software (PLGS, Waters) employing low energy, elevated energy, and intensity thresholds of 300, 100 and 1000 counts, respectively, and matched to peptides against a database containing the amino acid sequences of ShHTL7, AtSMAX1, ShMAX2, AtASK1, porcine pepsin, and their reversed sequences with the following search parameters: peptide tolerance = automatic; fragment tolerance = automatic; min fragment ion matches per peptide = 1; min fragment ion matches per protein = 7; min peptide matches per protein = 3; maximum hits to return = 20; maximum protein mass = 250,000; primary digest reagent = non-specific; missed cleavages = 0; false discovery rate = 100. For subsequent quantification of deuterium incorporation with DynamX 3.0 software (Waters), peptides had to fulfil the following criteria: minimum intensity of 10,000 counts; length of 5–40 amino acids; minimum number of products of two; maximum mass error of 25 ppm; retention time tolerance of 0.5 min. After automated data processing by DynamX 3.0, the mass spectra were manually inspected. The observable maximal deuterium uptake of a peptide was calculated by the number of amino acid residues minus one (for the N-terminal residue) minus the number of proline residues. For rendering of the residue-specific HDX and differences thereof from overlapping peptides, the shortest peptide covering a residue was employed. Where multiple peptides were of the shortest length, the peptide with the residue closest to the peptide’s C-terminus was utilised. Peptide and residue-level HDX values are provided as Supplementary Data 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM reconstructions and atomic models have been deposited in the Electron Microscopy Data Bank (EMDB) and Protein Data Bank Japan (PDBj), respectively, under the following accession numbers: EMD-62397, PDB 9KKX (ASK1–MAX2–HTL7–SMAX1 Class 1); EMD-62401, PDB 9KLD (ASK1–MAX2–HTL7–SMAX1 Class 2); EMD-62417, PDB 9KLV (ASK1–MAX2–HTL7–SMAX1 Class 3); EMD-62407, PDB 9KLK (ASK1–MAX2–HTL7–SMAX1 Class 4); EMD-62408, PDB 9KLL (apo-ASK1–MAX2); EMD-62414 (ASK1–ShMAX2 dimer); EMD-62415 (AtASK1–OsD3). The HDX–MS data were deposited to the ProteomeXchange Consortium via the PRIDE97 partner repository with the dataset identifier PXD068313. Additional processed HDX-MS data are provided in the Supplementary Data 1 file. Source data are provided in this paper.
References
Damgaard, R. B. The ubiquitin system: from cell signalling to disease biology and new therapeutic opportunities. Cell Death Differ. 28, 423–426 (2021).
Ebner, P., Versteeg, G. A. & Ikeda, F. Ubiquitin enzymes in the regulation of immune responses. Crit. Rev. Biochem. Mol. Biol. 52, 425–460 (2017).
Mulder, M. P. C. et al. A cascading activity-based probe sequentially targets E1–E2–E3 ubiquitin enzymes. Nat. Chem. Biol. 12, 523–530 (2016).
Komander, D. & Rape, M. The Ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Zhang, Z., Li, J., Liu, H., Chong, K. & Xu, Y. Roles of ubiquitination-mediated protein degradation in plant responses to abiotic stresses. Environ. Exp. Bot. 114, 92–103 (2015).
Yu, F., Wu, Y. & Xie, Q. Ubiquitin–proteasome system in ABA signaling: from perception to action. Mol. Plant 9, 21–33 (2016).
Al-Saharin, R., Hellmann, H. & Mooney, S. Plant E3 ligases and their role in abiotic stress response. Cells 11, 890 (2022).
Kelley, D. R. E3 Ubiquitin ligases: key regulators of hormone signaling in plants *. Mol. Cell. Proteom. 17, 1047–1054 (2018).
Zheng, N. & Shabek, N. Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 (2017).
Cardozo, T. & Pagano, M. The SCF ubiquitin ligase: insights into a molecular machine. Nat. Rev. Mol. Cell Biol. 5, 739–751 (2004).
Petroski, M. D. & Deshaies, R. J. Function and regulation of cullin–RING ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 6, 9–20 (2005).
Willems, A. R., Schwab, M. & Tyers, M. A hitchhiker’s guide to the cullin ubiquitin ligases: SCF and its kin. Biochim. Biophys. Acta 1695, 133–170 (2004).
Hua, Z. & Vierstra, R. D. The cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 62, 299–334 (2011).
Gagne, J. M., Downes, B. P., Shiu, S.-H., Durski, A. M. & Vierstra, R. D. The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl. Acad. Sci. USA 99, 11519–11524 (2002).
Kuroda, H. et al. Classification and expression analysis of arabidopsis F-box-containing protein genes. Plant Cell Physiol. 43, 1073–1085 (2002).
Kuroda, H., Yanagawa, Y., Takahashi, N., Horii, Y. & Matsui, M. A Comprehensive analysis of interaction and localization of arabidopsis SKP1-LIKE (ASK) and F-Box (FBX) proteins. PLOS ONE 7, e50009 (2012).
Skaar, J. R., Pagan, J. K. & Pagano, M. Mechanisms and function of substrate recruitment by F-box proteins. Nat. Rev. Mol. Cell Biol. 14, 369–381 (2013).
Tal, L., Gil, M. X. A., Guercio, A. M. & Shabek, N. Structural aspects of plant hormone signal perception and regulation by ubiquitin ligases. Plant Physiol. 182, 1537–1544 (2020).
Sheard, L. B. et al. Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor. Nature 468, 400–405 (2010).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Al-Babili, S. & Bouwmeester, H. J. Strigolactones, a novel carotenoid-derived plant hormone. Annu. Rev. Plant Biol. 66, 161–186 (2015).
Brewer, P. B., Koltai, H. & Beveridge, C. A. Diverse roles of strigolactones in plant development. Mol. Plant 6, 18–28 (2013).
Wang, Y. & Bouwmeester, H. J. Structural diversity in the strigolactones. J. Exp. Bot. 69, 2219–2230 (2018).
Jia, K.-P., Baz, L. & Al-Babili, S. From carotenoids to strigolactones. J. Exp. Bot. 69, 2189–2204 (2018).
Al-Babili, S. A dirigent of the ring for strigolactone stereochemistry. Proc. Natl. Acad. Sci. USA 121, e2410953121 (2024).
Zhao, L.-H. et al. Crystal structures of two phytohormone signal-transducing α/β hydrolases: karrikin-signaling KAI2 and strigolactone-signaling DWARF14. Cell Res. 23, 436–439 (2013).
Arite, T. et al. d14, a Strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol. 50, 1416–1424 (2009).
Yao, R. et al. DWARF14 is a non-canonical hormone receptor for strigolactone. Nature 536, 469–473 (2016).
Seto, Y. et al. Strigolactone perception and deactivation by a hydrolase receptor DWARF14. Nat. Commun. 10, 191 (2019).
Zhao, L.-H. et al. Destabilization of strigolactone receptor DWARF14 by binding of ligand and E3-ligase signaling effector DWARF3. Cell Res. 25, 1219–1236 (2015).
Li, Q. et al. The strigolactone receptor D14 targets SMAX1 for degradation in response to GR24 treatment and osmotic stress. Plant Commun. 100303 https://doi.org/10.1016/j.xplc.2022.100303 (2022).
Soundappan, I. et al. SMAX1-LIKE/D53 Family Members Enable Distinct MAX2-Dependent Responses to Strigolactones and Karrikins in Arabidopsis. Plant Cell 27, 3143–3159 (2015).
Li, Q. et al. SMXL5 attenuates strigolactone signaling in Arabidopsis thaliana by inhibiting SMXL7 degradation. Mol. Plant 17, 631–647 (2024).
Jiang, L. et al. DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 504, 401–405 (2013).
Zhou, F. et al. D14–SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 504, 406–410 (2013).
Xie, X., Yoneyama, K. & Yoneyama, K. The Strigolactone Story. Annu. Rev. Phytopathol. 48, 93–117 (2010).
Vurro, M., Bonciani, B. & Vannacci, G. Emerging infectious diseases of crop plants in developing countries: impact on agriculture and socio-economic consequences. Food Sec. 2, 113–132 (2010).
David, O. G., Ayangbenro, A. S., Odhiambo, J. J. O. & Babalola, O. O. Striga hermonthica: A highly destructive pathogen in maize production. Environ. Chall. 8, 100590 (2022).
Chesterfield, R. J., Vickers, C. E. & Beveridge, C. A. Translation of strigolactones from plant hormone to agriculture: achievements, future perspectives, and challenges. Trends Plant Sci. 25, 1087–1106 (2020).
Conn, C. E. et al. Convergent evolution of strigolactone perception enabled host detection in parasitic plants. Science 349, 540–543 (2015).
Toh, S. et al. Structure-function analysis identifies highly sensitive strigolactone receptors in Striga. Science 350, 203–207 (2015).
Uraguchi, D. et al. A femtomolar-range suicide germination stimulant for the parasitic plant Striga hermonthica. Science 362, 1301–1305 (2018).
Xu, Y. et al. Structural analysis of HTL and D14 proteins reveals the basis for ligand selectivity in Striga. Nat. Commun. 9, 3947 (2018).
Khosla, A. & Nelson, D. C. Strigolactones, super hormones in the fight against Striga. Curr. Opin. Plant Biol. 33, 57–63 (2016).
Nelson, D. C. The mechanism of host-induced germination in root parasitic plants. Plant Physiol. 185, 1353–1373 (2021).
Carlsson, G. H., Hasse, D., Cardinale, F., Prandi, C. & Andersson, I. The elusive ligand complexes of the DWARF14 strigolactone receptor. J. Exp. Bot. 69, 2345–2354 (2018).
Bürger, M. & Chory, J. The many models of strigolactone signaling. Trends Plant Sci. 25, 395–405 (2020).
Tal, L. et al. C-terminal conformational changes in SCF-D3/MAX2 ubiquitin ligase are required for KAI2-mediated signaling. N. Phytol.239, 2067–2075 (2023).
Hu, Q. et al. DWARF14, A receptor covalently linked with the active form of strigolactones, undergoes strigolactone-dependent degradation in rice. Front. Plant Sci. 8, https://doi.org/10.3389/fpls.2017.01935 (2017).
Shabek, N. et al. Structural plasticity of D3–D14 ubiquitin ligase in strigolactone signalling. Nature 563, 652–656 (2018).
Tal, L. et al. A conformational switch in the SCF-D3/MAX2 ubiquitin ligase facilitates strigolactone signalling. Nat. Plants 8, 561–573 (2022).
Liu, S. et al. Conformational dynamics of the D53 − D3 − D14 complex in strigolactone signaling. Plant Cell Physiol. 64, 1046–1056 (2023).
Hamiaux, C. et al. DAD2 Is an α/β hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone. Curr. Biol. 22, 2032–2036 (2012).
Wang, Y. et al. Molecular basis for high ligand sensitivity and selectivity of strigolactone receptors in Striga. Plant Physiol. 185, 1411–1428 (2021).
Yao, R. et al. ShHTL7 is a non-canonical receptor for strigolactones in root parasitic weeds. Cell Res. 27, 838–841 (2017).
Qiu, S. et al. Genome-enabled discovery of candidate virulence loci in Striga hermonthica, a devastating parasite of African cereal crops. N. Phytol. 236, 622–638 (2022).
de Saint Germain, A. et al. An histidine covalent receptor and butenolide complex mediates strigolactone perception. Nat. Chem. Biol. 12, 787–794 (2016).
Chen, J., White, A., Nelson, D. C. & Shukla, D. Role of substrate recognition in modulating strigolactone receptor selectivity in witchweed. J. Biol Chem. 297, https://doi.org/10.1016/j.jbc.2021.101092 (2021).
Shahul Hameed, U. et al. Structural basis for specific inhibition of the highly sensitive ShHTL7 receptor. EMBO Rep. 19, e45619 (2018).
Khosla, A. et al. Structure–function analysis of SMAX1 reveals domains that mediate its karrikin-induced proteolysis and interaction with the receptor KAI2. Plant Cell 32, 2639–2659 (2020).
Zeymer, C., Barends, T. R. M., Werbeck, N. D., Schlichting, I. & Reinstein, J. Elements in nucleotide sensing and hydrolysis of the AAA+ disaggregation machine ClpB: a structure-based mechanistic dissection of a molecular motor. Acta Cryst. D. 70, 582–595 (2014).
Ma, B., Zhu, J. & Huang, X. Diversification of plant SUPPRESSOR OF MAX2 1 (SMAX1)-like genes and genome-wide identification and characterization of cotton SMXL gene family. BMC Plant Biol. 23, 419 (2023).
Chang, S. H., George, W. J. & Nelson, D. C. An N-terminal domain specifies developmental control by the SMAX1-LIKE family of transcriptional regulators in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 122, e2412793122 (2025).
Wolf, N. M. et al. Structure of the N-terminal domain of ClpC1 in complex with the antituberculosis natural product ecumicin reveals unique binding interactions. Acta Cryst. D. 76, 458–471 (2020).
Guo, F., Maurizi, M. R., Esser, L. & Xia, D. Crystal structure of ClpA, an Hsp100 chaperone and regulator of ClpAP protease*. J. Biol. Chem. 277, 46743–46752 (2002).
Jamil, M. et al. Methyl phenlactonoates are efficient strigolactone analogs with simple structure. J. Exp. Bot. 69, 2319–2331 (2018).
Crowe, C. et al. Mechanism of degrader-targeted protein ubiquitinability. Sci. Adv. 10, eado6492 (2024).
Bunsick, M. et al. SMAX1-dependent seed germination bypasses GA signalling in Arabidopsis and Striga. Nat. Plants 6, 646–652 (2020).
Stanga, J. P., Smith, S. M., Briggs, W. R. & Nelson, D. C. SUPPRESSOR OF MORE AXILLARY GROWTH2 1 controls seed germination and seedling development in arabidopsis. Plant Physiol. 163, 318–330 (2013).
Chen, J., Nelson, D. C. & Shukla, D. Activation mechanism of strigolactone receptors and its impact on ligand selectivity between host and parasitic plants. J. Chem. Inf. Model. 62, 1712–1722 (2022).
Arellano-Saab, A. et al. Structural analysis of a hormone-bound Striga strigolactone receptor. Nat. Plants 9, 883–888 (2023).
Han, S. et al. Naturally occurring dehydrocostus lactone covalently binds to KARRIKIN INSENSITIVE 2 by dual serine modifications in orobanche cumana and arabidopsis. J. Agric. Food Chem. 72, 19920–19930 (2024).
Zhang, J. et al. Root microbiota regulates tiller number in rice. Cell 188, 3152–3166 (2025).
Guo, Y., Zheng, Z., La Clair, J. J., Chory, J. & Noel, J. P. Smoke-derived karrikin perception by the α/β-hydrolase KAI2 from Arabidopsis. Proc. Natl. Acad. Sci. USA 110, 8284–8289 (2013).
Guercio, A. M., Gilio, A. K., Pawlak, J. & Shabek, N. Structural insights into rice KAI2 receptor provide functional implications for perception and signal transduction. J. Biol. Chem. 300, 107593 (2024).
Waters, M. T. et al. Specialisation within the DWARF14 protein family confers distinct responses to karrikins and strigolactones in Arabidopsis. Development 139, 1285–1295 (2012).
Yap, J. X. & Tsuchiya, Y. Gibberellins promote seed conditioning by up-regulating strigolactone receptors in the parasitic plant striga hermonthica. Plant Cell Physiol. 64, 1021–1033 (2023).
Struk, S. et al. Unraveling the MAX2 protein network in Arabidopsis thaliana: identification of the protein phosphatase PAPP5 as a Novel MAX2 Interactor. Mol. Cell. Proteom. 20, https://doi.org/10.1074/mcp.ra119.001766 (2021).
Scheres, S. H. W. R. E. L. I. O. N. Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Asarnow, D., Palovcak, E. & Cheng, Y. asarnow/pyem: UCSF pyem v0.5. Zenodo https://doi.org/10.5281/zenodo.3576630 (2019).
Kimanius, D. et al. Data-driven regularization lowers the size barrier of cryo-EM structure determination. Nat. Methods 21, 1216–1221 (2024).
He, J., Li, T. & Huang, S.-Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Liu, Y.-T., Fan, H., Hu, J.J. & Zhou, Z.H. Overcoming the preferred-orientation problem in cryo-EM with self-supervised deep learning. Nat. Methods 22, 113–123 (2025).
Punjani, A. & Fleet, D. J. 3D variability analysis: Resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Cryst. D. 74, 519–530 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Cryst. D. 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Cryst. D. 75, 861–877 (2019).
Moriarty, N. W., Grosse-Kunstleve, R. W. & Adams, P. D. Electronic Ligand Builder and Optimization Workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Cryst. D 65, 1074–1080 (2009).
Düring, J. et al. Structural mechanisms of autoinhibition and substrate recognition by the ubiquitin ligase HACE1. Nat. Struct. Mol. Biol. 31, 364–377 (2024).
Wales, T. E., Fadgen, K. E., Gerhardt, G. C. & Engen, J. R. High-speed and high-resolution UPLC separation at zero degrees Celsius. Anal. Chem. 80, 6815–6820 (2008).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
We thank L. Zhao for help with cryo-EM data recording. We thank S. Al-Babili and A. De Biasio for valuable discussions and S. Al-Babili for reagents. For computer time, this research used the resources of the KAUST Supercomputing Laboratory, and experimental research was supported by the Bioscience Core Lab, ACL Proteomics Lab and the Imaging and Characterisation Core Lab at King Abdullah University of Science & Technology (KAUST) in Thuwal, Saudi Arabia. This research was supported by the King Abdullah University of Science and Technology (KAUST) through the baseline fund to STA and the Award No. URF/1/4039-01-01 and URF/1/4080-01-01. We acknowledge the support by the German Research Council (DFG) through the core facility for HDX-MS (projects 260989694 and 324652314 to Gert Bange, Philipps-University Marburg).
Author information
Authors and Affiliations
Contributions
Study design: A.I.V, U.F.S.H, and S.T.A. Protein production and biochemical analysis: A.I.V. HDX data collection: W.S. HDX analysis: A.I.V., W.S., and S.T.A. Cryo-EM sample preparation and data collection: A.I.V. and C.G.S. Biochemical and structural data analysis: A.I.V., B.H., C.G.S., and S.T.A. Manuscript writing: A.I.V., B.H., and S.T.A. All authors read the manuscript and provided comments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Peter McCourt, Bing Wang and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vancea, A.I., Huntington, B., Steinchen, W. et al. Mechanism of cooperative strigolactone perception by the MAX2 ubiquitin ligase–receptor–substrate complex. Nat Commun 16, 10291 (2025). https://doi.org/10.1038/s41467-025-65205-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65205-0