Abstract
Interleukin-1β (IL-1β) is a key inflammatory mediator in cancer. Its precursor, Pro-IL-1β, is conventionally considered inactive. Here we demonstrate that head and neck squamous cell carcinoma (HNSCC) cells exhibit significantly elevated Pro-IL-1β expression, driven by super-enhancer-mediated transcription of the IL1B gene. We show that intracellular Pro-IL-1β promotes tumor invasion and metastasis independent of IL-1β processing. Mechanistically, Pro-IL-1β binds RACK1 and inhibits its UBE2T-mediated ubiquitination, thereby stabilizing RACK1 and activating RhoA signaling to induce actin cytoskeleton remodeling and pseudopodia formation. Genetic inhibition of RACK1 abolishes Pro-IL-1β-induced metastasis. Clinically, RACK1 protein levels correlate with Pro-IL-1β expression in HNSCC specimens. Furthermore, we identify the natural compound Q3MG as a direct binder of Pro-IL-1β; it promotes lysosomal degradation of Pro-IL-1β and suppresses metastatic progression both in vitro and in vivo. Our study reveals a non-canonical, moonlighting function of Pro-IL-1β in tumor progression and highlights Q3MG as a promising therapeutic agent against metastatic cancer.
Similar content being viewed by others
Introduction
Interleukin-1β (IL-1β) is an important inflammatory factor in the tumor microenvironment (TME) that promotes cancer development and progression1. IL-1β is produced as a precursor, named Pro-IL-1β. The Pro-IL-1β is encoded by IL1B gene. In response to cellular stress, the transcription of IL1B is rapidly and vigorously induced. Followed by assemble of inflammasomes, the proenzyme procaspase-1 is converted to its active form caspase-1, which cleaves the Pro-IL-1β precursor, releases active IL-1β, and induces pyroptosis, an inflammatory cell death2,3. The source of IL-1β in TMEs is mainly from immune cells in response to infectious or other stressful stimuli, particularly in myeloid cells, such as macrophages4. In addition, tumor cells can produce IL-1β5,6. Both environment-derived and tumor cells-derived IL-1β have been shown to exerts pro-tumor function through various mechanisms7. Observations from the Canakinumab Anti-Inflammatory Thrombosis Outcome Study clinical trial unexpectedly found administration of canakinumab, the anti-IL-1β drug developed by Novartis, reduced the occurrence of fatal and non-fatal lung cancers in patients with a persistent pro-inflammatory response8, which fueled the enthusiasm of blocking IL-1β as a strategy for developing new cancer medicines. However, canakinumab administration failed to improve the primary endpoints of overall survival (OS) and progression-free survival in non-small cell lung cancer patients in the CANOPY-1 (NCT03631199) and CANOPY-2 (NCT03626545) phase III clinical trial9,10, raising important concerns about the validity of secreted IL-1β as a therapeutic target for cancer treatment. To date, the conventional view believes that IL-1β is solely active as an extracellular secreted cytokine, while its cytoplasmic precursor is biologically inactive11,12. Though there are many studies focused on the role of mature IL-1β, there has been no research on whether its precursor has a function in tumors.
Head and neck squamous cell carcinoma (HNSCC) are aggressive malignant diseases within high incidence of metastasis13,14. Due to lack of effective screening strategy, early detection of HNSCC remains great challenge, resulting in the majority of HNSCC patients being diagnosed in the advanced stages. Treatment of HNSCC patients consist of surgery followed by adjuvant radiation or chemotherapy plus radiation15. Mutational profiling revealed that most of frequently mutated genes in HNSCC are tumor suppressors, such as TP53, CDKN2A, FAT1, NOTCH1, KMT2D, NSD1, and TGFBR216,17. Mutations in oncogenes are rare in HNSCC, except for the case of PIK3CA18. In addition to genetic alterations, epigenetic changes also have a role in driving HNSCC oncogenesis. DNA hypermethylation-mediated inactivation of key tumor suppressor genes, including CDKN2A, RARB19,20, NOR1, DCC, and MGMT, occurs frequently21,22,23. Super enhancers (SEs) are tandem enhancer clusters that drive the expression of cell identity genes24. Cancer-specific SEs can mediate overexpression of oncogenes that sustain tumor cells proliferation and initiate metastasis25. Aberrant SEs play a critical role in maintaining the malignant behaviors of HNSCC26,27,28. Distant metastasis remains a major determinant of prognosis in HNSCC. To improve the treatment of metastatic cancers, its urgent to deepen our understanding of tumor metastasis mechanisms.
The process of cell invasion and migration requires the restructuring the cytoskeletal components, including actin, tubulin, and intermediate filaments. Remodeling of actin cytoskeleton affects the cellular architecture, enabling the formation of different protrusions such as filopodia, lamellipodia, stress fibers, and invadopodia that facilitate cell migration and invasion, which eventually leading to metastasis29,30. Rho family GTPases are key regulators of the actin cytoskeleton that are involved in cell migration, cell polarity, and membrane trafficking31. Activation of RhoA signaling pathway is well known to promote cancer cell metastasis32,33,34,35. In addition of regulating contractility in the cell body36,37, RhoA also regulates membrane protrusion38,39,40. Rho GTPase activation is conversely regulated by GTP exchange factors (GEFs) and GTPase-activating proteins (GAPs)41. In addition, proteins that do not have GEFs or GAPs activity can also regulate the activation of RhoA. For example, the receptor of activated protein kinase C1 (RACK1) interacts with RhoA and activates the RhoA/Rho kinase pathway to stimulate breast cancer cell invasion and metastasis42.
In the current study, we found upregulation of Pro-IL-1β in HNSCC tissues and cell lines, but relatively low mature IL-1β generated under resting status. High level of Pro-IL-1β predicts increased risk of distant metastasis and unfavorable prognosis in HNSCC patients. Acquisition SEs at IL1B gene is the cause leading to increased IL1B mRNA and overexpression of Pro-IL-1β in HNSCC. Tumoral Pro-IL-1β exerts an unusual role to promote tumor cell invasiveness in vitro and metastasis in vivo via activation of RhoA GTPase signaling, without needing to process into mature IL-1β. Mechanistically, Pro-IL-1β binds with RACK1 and prevents UBE2T-mediated degradation of RACK1, leading to RhoA activation and eventual cancer metastasis. We further demonstrated that Quercetin 3-O-(6”-O-malonyl)-β-D-glucoside (Q3MG) could target to intracellular Pro-IL-1β and promote its lysosomal degradation, exerting an inhibitory effect on tumor cell invasion and metastasis.
Results
High expression of Pro-IL-1β associates with metastasis and unfavorable prognosis in HNSCC cells
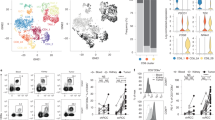
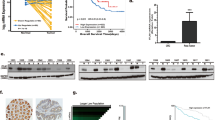
We analyzed the transcriptomic data of HNSCC and found that IL1B mRNA is increased in HNSCC (Fig. 1A). qRT-PCR assays confirmed the upregulation of IL1B mRNA in oral squamous cell carcinoma (OSCC) samples (Fig. 1B). To our surprise, Western blot assay showed that Pro-IL-1β, rather than mature IL-1β, was highly expressed in OSCC samples (Fig. 1C), albeit the antibody could recognize the mature IL-1β when cells undergo pyroptotic cell death triggered by triptolide, a pyroptosis inducer demonstrated by our previous study (Supplementary Fig. S1A)43. Our result is consistent with the previous demonstration that Pro-IL-1β was upregulated in human OSCC tumors44. We further detected Pro-IL-1β in HNSCC samples by immunohistochemistry assays and the results showed that the level of Pro-IL-1β in tumor is much higher than that of in normal counterpart (Fig. 1D and Supplementary Fig. S1B). High level of Pro-IL-1β is positively associated with advanced T stage (Fig. 1E), distant metastasis (Fig. 1F), and unfavorable prognosis in HNSCC patients (Fig. 1G). We then measured the expression of Pro-IL-1β in cancer cell lines. As shown in Supplementary Fig. S1C, IL1B mRNA was highly presented in HK1, FaDu, and BxPC3 cells, but weak in C666-1 cells. Western blot also showed Pro-IL-1β is highly expressed in HK1, FaDu, and BxPC3 cells, but absent in C666-1 cells. The level of mature IL-1β in cell lysates is undetectable by Western blot (Supplementary Fig. S1D). ELISA assays suggested that the level of secreted IL-1β in extracellular space under resting status is lower than 10 pg/mL, but it could reach to ~200 pg/mL when the NLRP3 inflammasome was activated by lipopolysaccharide (LPS) plus ATP treatment (Supplementary Fig. S1E), suggesting the NLRP3 inflammasome is intact in HNSCC cells, which is in consistent with the results reported previously45. Immunofluorescence assays showed the Pro-IL-1β protein abundantly distribute in cytoplasm of tumor cells (Supplementary Fig. S1F). Thus, our data suggested that the product of IL1B is mainly presented as Pro-IL-1β in HNSCC tumor cells under resting status.
A the mRNA levels of IL1B in HNSCC were analyzed from TCGA transcriptomic dataset. Tumor: n = 502 patient samples; Normal: n = 44 patient samples. B the mRNA levels of IL1B in fresh HNSCC samples and normal counterparts were determined by qRT-PCR. n = 3 biologically independent samples. C the protein levels of Pro-IL-1β in fresh HNSCC samples and normal counterparts were determined by Western blot assays. N normal, T tumor. The samples derive from the same experiment, but different gels for Pro-IL-1β and its loading control GAPDH were processed in parallel. D the protein levels of Pro-IL-1β in HNSCC samples were measured by IHC assays. Scale bar = 200 μm, 50 μm. E, F the protein level of Pro-IL-1β associates with advanced T stage and distant metastasis in HNSCC. T1–T3: n = 81 patient samples, T4: n = 9 patient samples; M0: n = 68 patient samples, M1: n = 22 patient samples. The box plots represent the median (center line), the 25th and 75th percentiles (bounds of the box), and the whiskers extend to the smallest and largest values within 1.5 × interquartile range (IQR) from the box. G the protein level of Pro-IL-1β predicts unfavorable overall survival in HNSCC patients. IHC scores of Pro-IL-1β greater than 6 were determined as high expression; others were considered as low expression. Student’s t-test (two-tailed) to compare the difference between two groups of data. One way ANOVA to compare the difference between three or more groups of data. These experiments were repeated at least biological triplicate. Data were presented as the means ± SD. **, P < 0.01; ***, P < 0.001. Exact P values and raw data were provided in the Source Data file.
SEs drives constitutive expression of Pro-IL-1β in HNSCC cells
Acquisition of SEs contributes to overexpression of oncogene products in human cancers25. We analyzed SEs landscape of SCC cells used in this study and found that there were SEs marked by intensive H3K27ac modification at IL1B gene in HK1, BxPC3 cells within high level of Pro-IL-1β, but not in C666-1 cells without expression of Pro-IL-1β (Supplementary Fig. S2A), suggesting SEs status is well correlated with the expression level of IL1B mRNA and Pro-IL-1β in SCC cells. Treatment with JQ1, a BRD4 inhibitor known to disrupt SEs-mediated transcription, efficiently reduced the mRNA levels of IL1B and the protein levels of Pro-IL-1β in HK1, FaDu, and BxPC3 cells in a dose-dependent manner (Supplementary Fig. S2B, C). ChIP-seq data showed ΔNp63α, the major isoform of p63α expressed in SCC cells, bound with SEs-IL1B in HK1, BxPC3 cells. Deletion of TP63 by CRISPR/Cas9 dramatically reduced the H3K27ac level at SEs-IL1B in BxPC3 cells (Supplementary Fig. S3A). ChIP-qPCR assays also confirmed that silencing ΔNp63α by shRNA led to decrease of H3K27ac, H3K4me1, and MED1/RNA pol II level at SEs-IL1B in HK1 cells (Supplementary Fig. S3B–E), indicating the presence of ΔNp63α is required for maintenance of SEs-IL1B in SCC cells. We further designed sgRNAs targeting to the ΔNp63α binding sites at SEs-IL1B. When these sgRNAs were co-introduced with dCas9-KRAB into HK1 cells, the mRNA levels of IL1B and the protein levels of Pro-IL-1β were significantly reduced (Supplementary Fig. S3F, G), highlighting the essential role of ΔNp63α binding in driving the transcription of IL1B in SCC cells. Consequently, silencing ΔNp63α by shRNA resulted in reduction of the mRNA levels of IL1B and the protein levels of Pro-IL-1β in HK1, FaDu, and BxPC3 cells (Supplementary Fig. S3H, I), which is in consistent with a previous study that silencing ΔNp63α led to downregulation of IL1B mRNA in HNSCC cells46. Thus, these data indicated that the transcription of IL1B and consequent overexpression of Pro-IL-1β in SCC cells under resting status is driven by ΔNp63α-dependent SEs.
The Pro-IL-1β enhances HNSCC cells migration and invasion in vitro in a cytokine-independent manner
The phenomenon that cytosolic Pro-IL-1β exists alongside other inactive inflammasome components within tumor cells raises the intriguing possibility that it may have a moonlighting function in cancer cells, albeit Pro-IL-1β in immune cells is routinely regarded as biological inactive. To this end, we silenced the expression of Pro-IL-1β in HK1, FaDu, and BxPC3 cells by shRNAs, we also overexpressed the wild type (WT) Pro-IL-1β (wtIL1B) or a non-cleavable Pro-IL-1β mutant (mtIL1B) within point mutation at the caspase-1 and -8 cleavage site (D117) of Pro-IL-1β protein in C666-1, HK1, and BxPC3 cells. A series of experimental results confirmed that either loss of function or gain of function models were successfully established (Supplementary Fig. S4). CCK-8 assays, and colony-formation assays indicated that neither loss of Pro-IL-1β nor gain of WT Pro-IL-1β or non-cleavable Pro-IL-1β exerted little effect on SCC cells proliferation and survival (Supplementary Fig. S5). However, wound-healing assays demonstrated that loss of endogenous Pro-IL-1β significantly inhibited the migrative ability of SCC cells, whereas gain of WT Pro-IL-1β or non-cleavable Pro-IL-1β exerted opposite effect (Supplementary Fig. S6A). Transwell assays also indicated that silencing Pro-IL-1β impaired the SCC cells migration and invasion in vitro, whereas overexpression both WT Pro-IL-1β and non-cleavable Pro-IL-1β promotes SCC cells migration and invasion in a similar extent (Fig. 2A and Supplementary Fig. S6B).
A the migration and invasion ability of Pro-IL-1β was assessed in cancer cells after overexpression or knockdown of Pro-IL-1β by Transwell chamber without or with Matrigel. n = 3 biologically independent samples. B the migration and invasion of Pro-IL-1β-knockout HK1 and FaDu were detected by Transwell chamber without or with Matrigel after re-overexpression of mtIL1B or IL-1β treatment. NC Negative control. n = 3 biologically independent samples. C body weight change curve of nude mice injected with cancer cells. D lung tissue sections were stained by H&E and anti-Pan-CK. Metastatic nodules in lung tissue sections were counted under microscope (bottom), each point in the statistical graph represents the total number of lung metastatic nodules count from one mouse (n = 5 mice per group). Pan-CK was used as a tumor marker. Scale bar = 100 μm. E Representative bioluminescence images of nude mice captured using the IVIS system at 10 and 30 days after tumor cell injection (Top). The bioluminescence signals of primary tumors (tongue) and metastatic tumors (cervical lymph nodes) of nude mice in each group were quantified using the in vivo imaging system (Bottom). n = 3 mice per group. Student’s t-test (two-tailed) to compare the difference between two groups of data. One way ANOVA to compare the difference between three or more groups of data. These experiments were biologically repeated at least three times with similar results. Data were presented as the means ± SD. *, P < 0.05; **, P < 0.01; ***, P < 0.001; ns non-significant. Exact P values and raw data were provided in the Source Data file.
To rule out the possibility that phenotype changes induced by loss or gain of Pro-IL-1β is due to alteration of mature IL-1β, we measure the extracellular secreted IL-1β levels by ELISA. As shown in Supplementary Fig. S7A, B, silencing Pro-IL-1β resulted in reduction of secreted IL-1β levels from approximate 5 pg/mL to 3 pg/mL in HK1, FaDu, and BxPC3 cells, overexpression of WT Pro-IL-1β led to about onefold increase of secreted IL-1β levels, overexpression of non-cleavable Pro-IL-1β did not affect the level of secreted IL-1β. Supplement of recombinant mature IL-1β (rIL-1β) (10 pg/mL) failed to rescue the migrative or invasive phenotype of Pro-IL-1β-silenced SCC cells (Supplementary Fig. S7C). Similarly, supplement excessive IL1RA (100 ng/mL), an antagonize of mature IL-1β, did not affect the migrative or invasive phenotype of SCC cells within Pro-IL-1β or non-cleavable Pro-IL-1β overexpression (Supplementary Fig. S7D, E). Since mature IL-1β at concentration of 1–10 ng/mL has been shown to stimulate tumor cells migration and invasion in previous studies47,48, we considered that the effect of mature IL-1β on tumor cell migration and invasion is dose-dependent. So, we treated SCC cells with various concentration of rIL-1β. The results showed that when the concentration increase to 500 pg/mL, rIL-1β indeed stimulated the migration and invasion of SCC cells in vitro, but did not the case when concentration is less than 500 pg/mL (Supplementary Fig. S8), suggesting that the promotive function of IL-1β in cancer migration and invasion require a relative high concentration. This could explain why adding 10 pg/mL rIL-1β failed to rescue the invasive phenotype of Pro-IL-1β-silenced SCC cells, indicating that inhibition of migration and invasion of SCC cells by silencing Pro-IL-1β is caused by loss of Pro-IL-1β, rather than decrease of secreted IL-1β.
We further established Pro-IL-1β knockout cell models by CRISPR/Cas9 approach (Supplementary Fig. S9A), then we introduced non-cleavable Pro-IL-1β (mtIL1B) or mature IL-1β into these knockout cells by lentivirus expression system (Supplementary Fig. S9B, C). CCK−8 assays and colony formation assays demonstrated that knockout of Pro-IL-1β did not affect the proliferation and survival (Supplementary Fig. S9D, E). Wound healing assays and Transwell assays indicated that knockout of Pro-IL-1β severely impaired the migrative and invasive potential of HK1 and FaDu cells. Re-expression of non-cleavable Pro-IL-1β (mtIL1B) efficiently rescued the migrative and invasive potential of Pro-IL-1β knockout cells, but re-expression of mature IL-1β within an extracellular concentration of approximate 200 pg/mL failed to rescue the invasive phenotype of Pro-IL-1β knockout cells (Fig. 2B and Supplementary Fig. S10A, B), confirming that tumoral Pro-IL-1β exerts a cell-autonomous function to promote SCC cells migration and invasion.
Pro-IL-1β promotes HNSCC cells metastasis in vivo
We then measured the impact of Pro-IL-1β on tumorigenicity and metastasis in vivo. Subcutaneous xenograft tumor formation assays showed that stable silencing Pro-IL-1β did not affect xenograft tumor growth in nude mice (Supplementary Fig. S11A–C). Subcutaneous implantation of HK1 and FaDu cell lines in nude mice did not generate spontaneous metastases during the 8-week experimental observation period, which is consistent with the characteristics of this model. The tumor cell lines used in this experiment are difficult to metastasize under subcutaneous transplantation conditions. We have also constructed subcutaneous transplantation tumor models of SCC cells, such as HK1 in nude mice, and no distant spontaneous metastasis of tumor cells was observed during the experimental period of up to 20 weeks (data not shown). Additionally, in the subcutaneous xenograft models (HK1, FaDu) of this study, we observed no local invasion into adjacent muscle or connective tissues, consistent with extensive literature demonstrating that standard subcutaneous xenograft models of HNSCC typically fail to recapitulate local invasion and distant metastasis phenotypes49,50. As these conventional xenograft models are primarily utilized for evaluating primary tumor growth, their microenvironment appears insufficient to support the complex biological processes driving HNSCC invasion and spontaneous metastasis. All metastatic conclusions in this study are therefore derived from controlled experimental metastasis assays (tail vein—lung metastasis model and tongue—lymph node metastasis model).
By using the tail vein injection metastasis model, we found that the body weight of nude mice inoculated with Pro-IL-1β-deficient HK1 and FaDu cells during the experimental cycle was higher than that of nude mice inoculated with control cells, and the body weight of nude mice inoculated with HK1 cells within overexpression of WT Pro-IL-1β or non-cleavable Pro-IL-1β significantly decreased during the experimental cycle as compared to the control group (Fig. 2C). The gross anatomy structure of the nude mice’s lung showed there were less metastatic nodules on the surface of lung from nude mice inoculated with Pro-IL-1β knockdown cells. In contrast, there were more metastatic nodules at the surface of lung from nude mice inoculated with cells within WT Pro-IL-1β or non-cleavable Pro-IL-1β overexpression (Supplementary Fig. S11D, E). Microscopic examination further demonstrated there were less metastatic tumor nodules in the lung tissues from nude mice inoculated with Pro-IL-1β knockdown cells, whereas there were more metastatic tumor nodules in the lung tissues from nude mice inoculated with cells within WT Pro-IL-1β or non-cleavable Pro-IL-1β overexpression (Fig. 2D). This indicated that Pro-IL-1β enhanced tumor cell survival and outgrowth at distant sites in vivo.
By using an orthotopic transplantation model51, SCC cells were inoculated into the tongue, and tumor metastasis was evaluated through in vivo imaging system monitoring combined with cervical lymph node dissection analysis. Our findings showed that altered Pro-IL-1β expression exerted little effect on the growth of primary tongue tumors (Supplementary Fig. S12A). In vivo imaging revealed that overexpression of WT Pro-IL-1β or uncleavable Pro-IL-1β significantly promoted cervical lymph node metastasis, whereas silencing Pro-IL-1β inhibited lymph node metastasis (Fig. 2E). The nude mice inoculated with WT Pro-IL-1β or uncleavable Pro-IL-1β-overexpressing cells exhibited a significant increase in size of cervical lymph nodes, and anti-Pan-CK staining confirmed an increase in metastatic tumor cells within the cervical lymph nodes of nude mice inoculated with WT Pro-IL-1β or uncleavable Pro-IL-1β-overexpressing cells. Conversely, inoculation with Pro-IL-1β-deficient cells produced the opposite effect (Supplementary Fig. S12B). Thus, our data suggested that high expression of Pro-IL-1β promotes tumor metastasis in vivo.
Pro-IL-1β induces actin cytoskeleton remodeling via activation of RhoA-ROCK signaling
Shape characteristics of cancer cells is linked with invasiveness and can be an indicator of metastatic potential52. We employed scanning electron microscope (SEM) to observe the surface morphology of tumor cells upon loss or gain of Pro-IL-1β. The data showed that stable silencing Pro-IL-1β led to alteration of SCC cell surface, which characterized by fewer burr-like protrusions on the surface and pseudopodia formation. In contrast, overexpression of either WT Pro-IL-1β or non-cleavable Pro-IL-1β facilitated pseudopodia formation and burr-like protrusions on the surface (Fig. 3A and Supplementary Fig. S13A). Since cellular protrusions and pseudopodia formation is linked with actin cytoskeleton remodeling29,30, we visualized F-actin cytoskeleton structure by immunofluorescence with phalloidin-labeled F-actin. As shown in Fig. 3B and Supplementary Fig. S13B, there were abundant F-actin rich filopodia at the surface of control tumor cells, whereas loss of Pro-IL-1β resulted in fewer F-actin rich filopodia. In contrast, overexpression of either WT or non-cleavable Pro-IL-1β promoted polymerization of F-actin and formation of F-actin rich filopodia.
A effect of Pro-IL-1β on the surface morphology of HK1 cells was visualized by scanning electron microscopy. B effect of Pro-IL-1β on F-actin skeleton was visualized by immunofluorescence with phalloidin-labeled F-actin in HK1 cells. DAPI: blue; F-actin: green; Scale bar = 20 μm. C the effects of Pro-IL-1β on Rho GTPases were determined by Rho GTPase activity assays. GTP-RhoA/Rac1/CDC42 and total RhoA/Rac1/CDC42 were detected on separate gels due to identical molecular size, using the same experiment processed in parallel. Total protein forms serve as the loading control for normalization. D subcellular distribution of RhoA protein was visualized in HK1 cells by immunofluorescence assay. DAPI: blue; RhoA: red; Scale bar = 10 μm. E F-actin skeleton structures were visualized by phalloidin after Y27632 (10 μM for 2 h) and CytB (0.1 mM for 2 h) treatment of HK1 cells overexpressing Pro-IL-1β. CytB: Cytochalasin B. DAPI: blue; F-actin: green; Scale bar = 20 μm. F migrative ability was evaluated by wound healing assay after Y27632 and CytB treatment of HK1 and BxPC3 cells overexpressing Pro-IL-1β. n = 3 biologically independent samples. G invasion was evaluated by Transwell chamber with Matrigel after Y27632 and CytB treatment of HK1 and BxPC3 cells overexpressing Pro-IL-1β. n = 3 biologically independent samples. One way ANOVA to compare the difference between three or more groups of data. All experiments were repeated three times independently with consistent results. Data were presented as the means ± SD. ***, P < 0.001. Exact P values and raw data were provided in the Source Data file.
We than asked which Rho GTPase is involved in the process of Pro-IL-1β mediated cytoskeleton remodeling. As shown in Fig. 3C, silencing Pro-IL-1β resulted in reduction of active RhoA in cancer cells, without affecting the activation of RAC1 or CDC42. In contrast, overexpression of either WT Pro-IL-1β or non-cleavable Pro-IL-1β promoted the activation of RhoA but did not affect the activation of RAC1 or CDC42. Immunofluorescence assays demonstrated that overexpression of either WT Pro-IL-1β or non-cleavable Pro-IL-1β promoted membrane localization of RhoA in HK1, C666-1, and BxPC3 cells (Fig. 3D and Supplementary Fig. S13C, D), further indicating that Pro-IL-1β promotes the activation of RhoA, since active RhoA mainly localizes at the plasma membrane53. We then treated Pro-IL-1β overexpressing cancer cells with Y27632, an inhibitor of ROCK1/2, or Cytochalasin B (CytB), a cell-permeable fungal toxin/mycotoxin that disrupt actin filaments. As shown in Fig. 3E and Supplementary Fig. S14, treatment with Y27632 or Cytochalasin B reduced F-actin rich filopodia structures and polymerization of F-actin in Pro-IL-1β overexpressing cancer cells. Furthermore, the results of silencing ROCK1 or ROCK2 by siRNA showed that both could significantly reverse the F-actin polymerization and pseudopod filamentous structure of cancer cells induced by Pro-IL-1β overexpression (Supplementary Fig. S15). Functionally, treatment with Y27632 or Cytochalasin B abolished the promotive effects on migration and invasion induced by overexpression of WT or non-cleavable Pro-IL-1β (Fig. 3F, G and Supplementary Fig. S16). Thus, our data suggested that constitutive expression of Pro-IL-1β enhanced tumor cells migration and invasion in vitro via activation of RhoA-ROCK signaling.
Pro-IL-1β binds with and stabilizes RACK1 protein in HNSCC cells
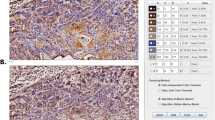
Co-immunoprecipitation assays indicated that Pro-IL-1β did not interact with RhoA protein in HK1 and BxPC3 cells (Supplementary Fig. S17A). We then employed a proximity biotinylation-based BioID strategy54 to identify Pro-IL-1β interaction partners in cells (Supplementary Fig. S17B). As shown in Supplementary Fig. S17C, D, BirA*-fused Pro-IL-1β was successfully expressed in human embryonic kidney (HEK) 293 T cells and induced cellular proteins biotinylation in the presence of exogenous biotin. We identified several differentially biotinylated proteins from two round BioID screening (Supplementary Fig. S17E and Supplementary Data 1). Among these proteins, RACK1 is known to be a regulator of RhoA in breast cancer42. Molecular docking suggested that Pro-IL-1β could directly bind to RACK1 protein (Fig. 4A). Co-immunoprecipitation assays demonstrated that the endogenous Pro-IL-1β and RACK1 proteins bind with each other in HK1 and BxPC3 cells (Fig. 4B). Immunofluorescence assays showed that Pro-IL-1β protein co-localized with RACK1 in tumor cells (Fig. 4C and Supplementary Fig. S17F). Although neither the C-terminal mature IL-1β nor the N-terminal could bind with RACK1 alone, the intermediate fragment containing the cleavage site and the full-length Pro-IL-1β protein bound with RACK1 protein. We failed to detect the N-terminal of Pro-IL-1β protein in plasmid-transfected cells, which may be due to rapid degradation since N-terminal of Pro-IL-1β protein is highly unstable55 (Fig. 4D and Supplementary Fig. S17G). Neither loss of Pro-IL-1β nor gain of Pro-IL-1β affect the mRNA levels of RACK1 in SCC cells (Supplementary Fig. S18A). However, loss of Pro-IL-1β resulted in decrease of the protein levels of RACK1 in SCC cells, whereas overexpression of either WT or non-cleavable Pro-IL-1β exerted opposite effect on RACK1 protein levels (Fig. 4E and Supplementary Fig. S18B). Pulse-chase assays indicated that silencing Pro-IL-1β shortened the half-life of RACK1 proteins. In contrast, overexpression of Pro-IL-1β prolonged the half-life of RACK1 proteins in HK1 and BxPC3 cells (Supplementary Fig. S18C). The proteasome inhibitor MG132 rescued the levels of RACK1 protein in Pro-IL-1β deficient cancer cells. Treatment with autophagy inhibitor 3-Methyladenine (3-MA) failed to rescue the protein levels of RACK1 protein in Pro-IL-1β deficient cancer cells (Fig. 4F and Supplementary Fig. S18D, E). As expected, silencing Pro-IL-1β led to increase of ubiquitinated RACK1 proteins in cancer cells (Fig. 4G and Supplementary Fig. S18F). Thus, our data suggested that loss of Pro-IL-1β accelerated the proteasomal degradation of RACK1 protein. It has been shown that UBE2T mediates ubiquitination-proteasomal degradation of RACK1 protein in cancer cells56,57. Pro-IL-1β did not affect the protein levels of UBE2T in cancer cells (Supplementary Fig. S19A). Co-immunoprecipitation assays showed that loss of Pro-IL-1β facilitated the interaction between RACK1 and UBE2T. However, overexpression of Pro-IL-1β prevented the interaction between RACK1 and UBE2T (Fig. 4H and Supplementary Fig. S19B). We silenced UBE2T by siRNAs and found that depletion of UBE2T resulted in recovery of RACK1 protein levels in Pro-IL-1β-silenced cells (Fig. 4I and Supplementary Fig. S19C). In vitro ubiquitination assay confirmed that Pro-IL-1β prevented UBE2T-mediated ubiquitination of RACK1 (Fig. 4J). Western blot assays showed that the protein levels of RACK1 protein were higher in SCC samples within high expression level of Pro-IL-1β protein (Fig. 4K). Immunohistochemical staining also indicated that the protein levels of RACK1 were positively correlated with the protein levels of Pro-IL-1β in HNSCC tissue samples (Fig. 4L and Supplementary Fig. S19D). Thus, our data suggested that Pro-IL-1β binds to RACK1 protein and prevents UBE2T-mediated ubiquitination-proteasomal degradation of RACK1 protein in HNSCC.
A the 3D spatial arrangement between Pro-IL-1β (pink) protein and RACK1 (purple) protein is shown. Key interacting residues are annotated, with red dashed lines indicating hydrogen bonds. Numeric labels adjacent to the dashed lines specify hydrogen bond lengths in angstroms (Å). GLU101, ASP116, and GLU180 on Pro-IL-1β protein form hydrogen bonds with ARG100 and LYS127 on RACK1 protein. B Co-IP assays showing the interaction between Pro-IL-1β and RACK1 proteins in HK1 and BxPC3 cells. C representative fluorescence confocal images showing co-localization of Pro-IL-1β with RACK1 proteins in HK1, FaDu, and BxPC3 cells. DAPI: blue; Pro-IL-1β: green; RACK1: red; Scale bar = 20 μm. D the binding of Pro-IL-1β with RACK1 protein was analyzed by Co-IP assays in HEK293T cells after co-transfection of myc-RACK1 overexpression vector and 3 × Flag-Pro-IL-1β full-length or truncated fragments. The Input and IP samples derive from the same experiment but were run on different gels due to the large number of samples. Because of their close molecular size and to ensure optimal antibody performance, the immunoblots for Flag-Pro-IL-1β and RACK1 were also processed on separate membranes. All gels and membranes were processed in parallel under identical conditions. E effect of Pro-IL-1β on RACK1 protein levels in HK1 cells were measured by Western blot. F the protein levels of RACK1 in loss of Pro-IL-1β HK1 cells upon MG132 (10 μM for 8 h) treatment were determined by Western blot. G the ubiquitination of RACK1 proteins in HK1 cells were assessed by immunoprecipitation and Western blot assay. H effect of Pro-IL-1β on the interactions between RACK1 with UBE2T protein in HK1 cells were assessed by Co-IP assays. I effects of UBE2T on the protein levels of RACK1 in Pro-IL-1β silenced HK1 cells were tested by Western blot. J purified RACK1 protein was subjected to in vitro ubiquitination assay with UBE2T in the ubiquitination reaction mixture in the absence or presence of Pro-IL-1β, and Western blot using the indicated antibodies. K the expression of Pro-IL-1β and RACK1 proteins in HNSCC samples were measured by Western blot. The samples derive from the same experiment, but different gels for Pro-IL-1β, RACK1, and GAPDH were processed in parallel. L representative images showing the expression of Pro-IL-1β and RACK1 proteins in HNSCC samples were measured by IHC (left). Scale bar = 200 μm, 50 μm. Pearson correlation analysis was used to verify the correlation between Pro-IL-1β and RACK1 protein expression levels in HNSCC tissues (right). R2 = 0.806, P < 0.01. These experiments were derived from three biologically independent repetitions. Each Western blots were reproduced three times with similar results. Owing to analogous molecular weights, samples for RACK1 protein (36 KDa) and GAPDH (35 KDa) were run on separate gels but processed in parallel under identical conditions. The blot for RACK1 protein is derived from the same experiment as that for GAPDH, but was processed on a separate gel. Exact P values and raw data were provided in the Source Data file.
Stabilization of RACK1 protein contributes to RhoA activation and metastasis induced by Pro-IL-1β
We silenced RACK1 expression by shRNAs in HK1, FaDu, and BxPC3 cells (Supplementary Fig. S20A–C). Stable silence of RACK1 did not affect the expression level of Pro-IL-1β protein and release of IL-1β in cancer cells (Supplementary Fig. S20C, D). Stable silence of RACK1 reduced the levels of active RhoA in HK1, FaDu, and BxPC3 cells (Fig. 5A). As expected, inhibition of RACK1 inhibited polymerization of F-actin and reduced F-actin rich filopodia at the surface of tumor cells (Supplementary Fig. S20E), confirming the regulatory role of RACK1 in RhoA signaling and cytoskeleton remodeling. Co-immunoprecipitation assays revealed that silence of Pro-IL-1β attenuated RACK1 protein binding to RhoA, whereas overexpression of WT or non-cleavable Pro-IL-1β exerted opposite effect (Fig. 5B, C). Depletion of RACK1 by siRNAs resulted in reduction of active RhoA levels in WT or non-cleavable Pro-IL-1β overexpressing tumor cells, whereas overexpression of RACK1 in Pro-IL-1β silenced cancer cells rescued the active RhoA levels (Fig. 5D, E and Supplementary Fig. S20F, G). Consequently, depletion of RACK1 abolished the promotive effect of Pro-IL-1β on F-actin rich filopodia structures and polymerization of F-actin in HK1, C666-1, and BxPC3 cells (Fig. 5F and Supplementary Fig. S21A). Functionally, depletion of RACK1 attenuated the migration and invasion of WT or non-cleavable Pro-IL-1β overexpressing tumor cells in vitro (Fig. 5G and Supplementary Fig. S21B), but did not affect proliferation and survival of these cells (Supplementary Fig. S22).
A effect of RACK1 silencing on the activation of RhoA was tested by Rho GTPase activity assays. GTP-RhoA and total RhoA were detected on separate gels due to identical molecular size, using the same lysates processed in parallel. Total RhoA serves as the loading control for normalization. B, C effect of Pro-IL-1β on the interactions between RACK1 with RhoA protein in cancer cells were assessed by Co-IP assays. D, E the activation of RhoA was examined by Rho GTPase activity assays after overexpression or knockdown of Pro-IL-1β or RACK1. GTP-RhoA and total RhoA were detected on separate gels due to identical molecular size, using the same experiment processed in parallel. Total RhoA serves as the loading control for normalization. F effect of RACK1 knockdown on F-actin skeleton was visualized by phalloidin in HK1 cells overexpressing Pro-IL-1β. Scale bar = 20 μm. G the migration and invasive ability of cancer cells was examined by Transwell chamber without or with Matrigel after overexpression of Pro-IL-1β or knockdown of RACK1. n = 3 biologically independent samples. Student’s t-test (two-tailed) to compare the difference between two groups of data. ANOVA to compare the difference between three or more groups of data. These experiments were repeated three times biologically with consistent results. Data were presented as the means ± SD. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P values and raw data were provided in the Source Data file.
To further validate the effect of RACK1 on Pro-IL-1β-induced tumor cell metastasis in vivo, we established animal models. By using a tail vein injection lung metastasis model, we provide evidence that nude mice injected with either WT or non-cleavable Pro-IL-1β overexpressing HK1 cells lose body weight during the experiment period. However, when RACK1 was silenced in these cells, the body weight of nude mice injected with tumor cells remained stable (Fig. 6A). Overexpression of WT or non-cleavable Pro-IL-1β in HK1 cells promoted lung metastasis in nude mice, whereas depletion of RACK1 abolished the promotive effect of Pro-IL-1β on lung metastasis in vivo (Fig. 6B–D and Supplementary Fig. S23A). The expression of Pro-IL-1β or RACK1 in lung metastatic tumors were measured by immunohistochemical staining (Fig. 6D and Supplementary Fig. S23B). In the orthotopic transplantation model, we found that overexpression of WT or non-cleavable Pro-IL-1β promoted cervical lymph node metastasis through in vivo imaging, while depletion of RACK1 significantly inhibited lymph node metastasis and reversed the effect of Pro-IL-1β overexpression (Fig. 6E). Silencing RACK1 significantly attenuated Pro-IL-1β-induced lymph node enlargement and reduced tumor cell infiltration in lymph nodes (Fig. 6F and Supplementary Fig. S23C), without affecting orthotopic tongue tumor growth in mice (Supplementary Fig. S23D). Thus, these data indicated that Pro-IL-1β promotes invasion and metastasis of cancer cells through upregulation of RACK1 protein in HNSCC cells.
A body weight change curve of nude mice injected with HK1 cells treated with Pro-IL-1β overexpression or RACK1 knockdown (n = 5 mice per group). B the gross view of lungs from mice injected with HK1 cells in each group at the end point of experiment (n = 5 mice per group). C the number of metastatic nodules at lung surface was counted for each group (n = 5 mice per group). D representative images of H&E staining of metastatic lung nodules and the expression of Pro-IL-1β, RACK1, and Pan-CK by IHC assays (left), Pan-CK is used as a tumor marker. Scale bar = 100 μm. Metastatic nodules in H&E-stained lung tissue sections were counted under microscope (right), each point in the statistical graph represents the total number of lung metastatic nodules count from one mouse (n = 5 mice per group). E BLI showed that silencing RACK1 significantly attenuated the promoting effect of Pro-IL-1β overexpression on lymph node metastasis in mice. Top, representative bioluminescence images of nude mice captured using the IVIS system at 10 and 30 days after tumor cell injection. Bottom, the bioluminescence signals of primary tumors (tongue) and metastatic tumors (cervical lymph nodes) of nude mice in each group were quantified using the in vivo imaging system (n = 3 mice per group). F images of cervical lymph nodes of nude mice injected with HK1 cells treated with Pro-IL-1β overexpression or RACK1 knockdown from the orthotopic metastasis model. Bottom panel, quantification of lymph node volume (n = 5 mice per group). ANOVA to compare the difference between three or more groups of data. These experiments were repeated at least biological triplicate. Data were presented as the means ± SD. *, P < 0.05; **, P < 0.01; ***, P < 0.001. Exact P values and raw data were provided in the Source Data file.
Q3MG targets Pro-IL-1β protein and suppresses metastatic potential of HNSCC cells
We employed high-throughput virtual screening (HTVS) strategy to search small molecules that have the potential to bind with Pro-IL-1β protein. After consecutive SP and XP docking analysis, we obtained the top 50 compounds with the highest docking scores (Supplementary Data 2). In silico docking showed that Q3MG bound to the allosteric pocket on the surface of IL1B protein (PDB ID: 5R8Q) through hydrogen bonding, and the key residues of the pocket were L80/L134/K74/V132 (Supplementary Fig. S24). We then tested the effect of Q3MG on Pro-IL-1β expression. Q3MG treatment decreased the levels of endogenous Pro-IL-1β proteins in HK1, FaDu, and BxPC3 tumor cells in a dose- and time- dependent manner (Fig. 7A), without affecting the mRNA levels and inflammasome activation to secrete IL-1β (Supplementary Fig. S25A, B). After treating SCC cells with lysosomal inhibitor CQ or proteasome inhibitor MG132, Western blotting results showed that Q3MG mediated Pro-IL-1β protein degradation through the lysosomal pathway, rather than through the proteasomal pathway (Fig. 7B, Supplementary Fig. S25C). Cellular thermal shift assay (CETSA) further confirmed the direct interaction between Pro-IL-1β protein and Q3MG (Fig. 7C and Supplementary Fig. S25D).
A Western blot showed that Q3MG significantly inhibited the expression of Pro-IL-1β protein in HK1, FaDu, and BxPC3 cells in a dose- and time-dependent manner. The concentration and time of the drug in the in vitro cell experiment were finally determined to be 50 μM and 24 h. Q3MG: Quercetin 3-O-(6”-O-malonyl)-β-D-glucoside. B the effect of Q3MG on Pro-IL-1β protein degradation was detected by Western blot after treatment of HK1, FaDu, and BxPC3 using CQ. CQ Chloroquine. C CETSA curves for Pro-IL-1β protein with or without Q3MG in HK1, FaDu, and BxPC3 cell lysates. D the effect of Q3MG on cell migration and invasion in Pro-IL-1β knockout HK1 and FaDu cells was detected by Transwell chamber without or with Matrigel. n = 5 biologically independent samples. E body weight change curve of nude mice with or without Q3MG treatment in tail vein-lung metastasis model (n = 5 mice per group). F gross view of lung tissue removed from nude mice at the end of experiment (n = 5 mice per group), arrows point to metastatic nodules on the lung surface. G representative images of H&E staining of metastatic lung nodules and the expression of Pro-IL-1β and Pan-CK by IHC assays, Pan-CK is used as a tumor marker. Scale bar = 200 μm. On the right, metastatic nodules in H&E-stained lung tissue sections were counted under microscope (n = 5 mice per group). Student’s t-test (two-tailed) to compare the difference between two groups of data. One way ANOVA to compare the difference between three or more groups of data. These experiments were repeated at least biological triplicate. Data were presented as the means ± SD. *, P < 0.05; **, P < 0.01; ***, P < 0.001; ns non-significant. Exact P values and raw data were provided in the Source Data file.
As shown in Supplementary Figs. S25E–G and S26A, Q3MG treatment dramatically impaired the migration and invasion of HK1, FaDu, and BxPC3 cells within Pro-IL-1β expression, without affecting cell proliferation and survival. However, in the case of Pro-IL-1β knockout HK1 and FaDu cells, Q3MG did not suppress cell migration and invasion (Fig. 7D and Supplementary Fig. S26B), suggesting a Pro-IL-1β-dependent effect. Treatment with Q3MG effectively prevented weight loss in nude mice injected with HK1 and FaDu tumor cells (Fig. 7E) without obvious toxic side effects (Supplementary Fig. S26C). Importantly, Q3MG treatment effectively reduced the number of lung metastatic tumor nodules (Fig. 7F and Supplementary Fig. S26D, E). H&E and IHC staining of paraffin sections of lung tissue showed that Pro-IL-1β expression signal in Q3MG treatment group was significantly weakened, and the number and area of lung metastatic lesions were significantly lower than those in the control group (Fig. 7G).
Discussion
In this study, we unveiled an unrecognized moonlighting function of Pro-IL-1β to promote cancer invasion and metastasis, a process which RACK1-RhoA signaling mediated cytoskeleton remodeling was involved. Degradation of Pro-IL-1β using Q3MG exerted therapeutic effectiveness against Pro-IL-1β expressing SCC cancer cells in vitro and in vivo.
Roles of the pro-inflammatory cytokine IL-1β contributing to tumorigenesis, metastasis, angiogenesis, and immune evasion are well recognized7. Production of IL-1β requires two signals, namely “priming” that corresponds to transcription of IL1B gene and cleavage by proteases. Activation of inflammasome is the most important step involved in IL-1β maturation1. Several studies reported that Pro-IL-1β is highly expressed in HNSCC cells, but the mature IL-1β remains a relative low level less than 20 pg/mL44,45. This phenomenon is not due to lack of inflammasome machinery in these cancer cells, because our data along with other’s observation, showed that activation of NLRP3 inflammasome by LPS plus ATP effectively induced release of mature IL-1β in HNSCC cells, that means inflammasome machinery is intact in these cancer cells but remains inactive without stimuli. Therefore, it is reasonable to consider whether Pro-IL-1β have a cell autonomous role in tumor cells. We provide evidence that depletion of tumoral Pro-IL-1β impaired tumor cells invasion and metastatic potential, which could not be recovered by adding physiological level of mature IL-1β (< 20 pg/mL). Overexpression of non-cleavable Pro-IL-1β, which have a point mutation at the caspase-1, caspase-8 cleavage site promoted tumor cells invasion in vitro and metastasis in vivo. Thus, the current study highlights a previous unrecognized function of tumoral Pro-IL-1β itself, rather than secreted IL-1β, to promote cancer progression. The current study guarantees more research using genetic animal models to further verify the biological activities of tumoral Pro-IL-1β in cancer initiation and progression. However, this study does not negate the roles of mature IL-1β in cancer development and progression. Also, this study does not speculate on the function of Pro-IL-1β in immune cells, because Pro-IL-1β is rapidly turned over by ubiquitin-proteasomal degradation in immune cells to avoid collateral damage58. We found that the promotive effect of mature IL-1β on tumor cell migration and invasion is dose-dependent. IL-1β was used up to 1 ng/mL-10 ng/mL in many in vitro studies during the past decades47,48. In this study, when the concentration is higher than 500 pg/mL, IL-1β do stimulate the movement and invasion of tumor cells. Though it is difficult to measure the accurate level of IL-1β in the local extracellular space of tumor cells in TME, we speculate that when inflammasomes are activated or cells undergo pyroptosis, local IL-1β may reach high concentrations.
In this study, we found that the transcription of IL1B in HNSCC cells under resting status attributes to acquisition of ΔNp63α dependent SEs. Disruption of SEs by BRD4 inhibitor or CRISPR interference targeting to ΔNp63α binding sites repressed the mRNA levels of IL1B and eventually reduced the Pro-IL-1β protein levels in SCC cells. The ΔNp63α is the predominant isoform of TP63 expressed in epidermal cells and SCC. As a lineage survival oncogene for SCC, ΔNp63α is highly expressed nearly in all SCC samples. This explains to some extent why there are high levels of IL1B mRNA and Pro-IL-1β protein expression in HNSCC cells even in the unstimulated status. At present, we cannot infer whether ΔNp63α is involved in the “priming” stage of transcription of IL1B gene upon inflammatory stimuli. Previous study reported that ΔNp63α interacts with NF-κB family members to regulate pro-inflammatory gene expression in HNSCC46. We propose that active SEs shaped by ΔNp63α may serve as a “pre-priming” mechanism to orchestrate IL1B transcription program.
By using a proximity biotinylation approach, we identified that Pro-IL-1β interacts with and stabilizes RACK1 scaffolding protein in SCC cells. Interestedly, neither the N-terminus of Pro-IL-1β nor the C-terminal mature IL-1β reserve the ability to bind with RACK1 protein, suggesting a precursor-specific role. A previous study demonstrated RACK1 interacts with NLRP3 and NEK7 in macrophage treated with LPS alone or LPS plus ATP stimulation. Silencing RACK1 attenuates activation of caspase-1 and release of mature IL-1β in macrophage in response to NLRP3-activating stimuli59. However, silencing RACK1 did not affect the protein level of Pro-IL-1β and release of mature IL-1β in resting SCC cells, suggesting the role of RACK1 in cancer cells is different to that of in macrophage. The interaction between Pro-IL-1β and RACK1 seem to be not linked to NLRP3, because HEK293T cells used for proximity biotinylation screening do not express endogenous NLRP359. RACK1 protein is overexpressed in OSCC and predicts unfavorable prognosis in OSCC60,61,62. Binding with Pro-IL-1β protect RACK1 from UBE2T-mediated ubiquitination and proteasomal degradation. In OSCC samples, the level of Pro-IL-1β is positively correlated with that of RACK1 protein. Thus, high level of tumoral Pro-IL-1β is a cause of aberrant expression of RACK1 in cancer cells. Upregulation of RACK1 protein promotes migration and invasion in a variety of human cancers63. As a scaffolding protein, RACK1 promotes the activation of RhoA/Rho kinase signaling in cancer cells42. We provide evidence showing that the promotive effects of Pro-IL-1β on RhoA signaling, F-actin cytoskeleton rearrangement and invasive behaviors were mediated by RACK1 protein. Therefore, our study uncovered a Pro-IL-1β/RACK1/RhoA regulatory axis which contributes to acquisition of invasiveness in cancer.
As a monoclonal neutralizing antibody of mature IL-1β, canakinumab achieved glamorous success in immune-related disorders64. The failure of the CANOPY-1 and 2 trials cast a shadow on the application of canakinumab for cancer treatment10, which raised concerns about the effectiveness of secreted IL-1β as a target for cancer treatment. Previous studies have not recognized the moonlighting function of tumoral Pro-IL-1β to assist tumor progression, so prior to this, no research has attempted to determine whether targeting the intracellular Pro-IL-1β protein have potential as therapeutic strategy for cancer. Compounds for degrading Pro-IL-1β are also largely unavailable. In fact, drug screening targeting Pro-IL-1β may provide a direction for anti-tumor therapy. Structure-based HTVS found that the quercetin derivative Q3MG could bind to Pro-IL-1β protein. Our data showed that Q3MG accelerated the degradation of Pro-IL-1β protein via the lysosomal pathway without affecting IL-1β secretion, resulting in a significant inhibitory effect on tumour cell invasion and metastasis in vitro and in vivo. Reportedly, baicalein can enhance the interaction between CD274 and LC3 in the form of a molecular “glue,” thereby promoting the autophagic-lysosomal degradation of CD27465. The promotion of Pro-IL-1β lysosomal degradation by Q3MG may also function as a molecular “glue” similar to baicalein, but further verification is needed. Though our preliminary data could not guarantee the utilization of Q3MG in human cancer treatments, it shed light on the prospect of intracellular tumoral Pro-IL-1β as a therapeutic target. Developing small molecule drugs targeted to tumoral Pro-IL-1β may be an alternative strategy.
In conclusion, our study uncovers the moonlighting function of tumoral Pro-IL-1β in promoting HNSCC cell invasion and metastasis. Pro-IL-1β binds with RACK1 and inhibits UBE2T-mediated RACK1 degradation, thereby activating RhoA and promoting cytoskeletal remodeling. Targeting Pro-IL-1β by Q3MG, our identified Pro-IL-1β inhibitor, produced significant anti-metastatic efficacy (Supplementary Fig. S27). Taken together, Pro-IL-1β is a promising prognostic factor and therapeutic target for HNSCC patients.
Methods
Ethics approval and consent to participate
This research complies with all relevant ethical regulations. Approval for the use of clinical samples and information was obtained from each patient and the Research Ethics Committee of Central South University. Written informed consent was received prior to patient participation. The animal experiments were conducted according to the protocol approved by the Animal Welfare Committee of Central South University.
Patient samples
Ten fresh OSCC samples and matched adjacent normal tissue samples used for RNA and protein extraction were collected by surgery from the Department of Oral and Maxillofacial Surgery, The Second Xiangya Hospital, Central South University, within informed consents obtained from all participants. A cohort comprised 90 formalin-fixed, paraffin-embedded HNSCC specimens and 90 adjacent normal tissue samples was collected from the Department of Pathology, The Second Xiangya Hospital, Central South University (Supplementary Data 3). There was no restriction on sex and age in our study. The usage of tumor samples was approved by the Joint Ethics Committee of Central South University, and informed consents were obtained for both the research and the reporting/sharing of individual-level data.
Cell culture and reagents
HK1 (RRID: CVCL_7084), a well differentiated squamous cell carcinoma cell line from nasopharynx, was maintained in our lab66. FaDu (RRID: CVCL_1218), a hypopharyngeal carcinoma cell line, was purchased from the National Infrastructure of Cell Line Resource (Shanghai, China)67. C666-1 (RRID: CVCL_7949) is an undifferentiated nasopharyngeal carcinoma cell line maintained in our lab68. BxPC3 (RRID: CVCL_0186) is a pancreatic squamous cell carcinoma cell line purchased from the National Infrastructure of Cell Line Resource (Shanghai, China). All these cells were maintained in RPMI 1640 medium (Gibco, Grand Island, NY, USA) containing 10% fetal bovine serum (FBS, Gibco) and 1% penicillin-streptomycin. All cell lines were maintained in a humidified atmosphere consisting of 5% CO2 and 95% air at 37 °C. For stable overexpression, Pro-IL-1β, non-cleavable Pro-IL-1β, or mature IL-1β-expressing lentivirus were introduced into cancer cells and then the cells were selected by puromycin. Human IL-1RA/IL1RN Protein (Cat No.10123-HNAE) and Human IL-1β (Cat No.10139-HNAE) were purchased from Sino Biological, Inc.
Hematoxylin-Eosin staining (H&E) and immunohistochemistry (IHC)
Tissue sections were firstly heated at 65 °C for 2 h and then dewaxed by xylene. After deparaffinization, tissue sections were serially hydrated. Nucleus were visualized by hematoxylin solution (Biosharp, Anhui, China), and then cytoplasm was stained by eosin staining solution (Biosharp, Anhui, China). For IHC, tissue sections were deparaffinized and rehydrated. Antigen retrieval was achieved by boiling tissue section in citrate buffer (10 mM Sodium Citrate, 0.05% Tween 20, pH 6.0). After antigen retrieval, tissue sections were blocked with normal goat serum and then incubated with the primary antibodies overnight at 4 °C. Immuno-staining signal was developed by Horseradish Peroxidase (HRP) conjugated secondary antibodies and chromogen 3,3′-diaminobenzidine. Tissue sections were counterstained with hematoxylin. Each tissue section was comprehensively evaluated by whole-slide scanning using the 3DHISTECH Pannoramic SCAN II digital pathology system. As for IHC scoring criteria: All sections were independently scored by two pathologists who were blind to the clinicopathological features of the samples. A semi-quantitative scoring criterion was used for Pro-IL-1β and RACK1 based on the staining intensity and the proportion of positive cells. When the tissue was not stained, scoring 0. When the tissue was pale yellow, scoring 1; yellow, scoring 2; light brown, scoring 3; dark brown, scoring 4. On the other hand, when less than 10% of cells were positive, scoring 1; 10%–50% positive, scoring 2; and more than 50% positive, scoring 3. Finally, a comprehensive score was obtained by multiplying the staining intensity score and the positive ratio score (Supplementary Data 3).
Enzyme-linked immunosorbent assay (ELISA)
The concentration of extracellular IL-1β was measured by using human IL-1β instant ELISA Kit (Invitrogen) according to the manufacture’s instruction.
Immunofluorescence (IF) assays
Cells were seeded on glass cover slides to reach 30% confluency and washed with pre-cold PBS for three times, after which they were fixed at RT with 4% paraformaldehyde for 30 min, and permeabilized with 0.1% Triton ×-100 for 10 min. The cells were then blocked with 3% BSA for 1 h. Primary antibodies were visualized using a secondary antibody conjugated to Alexa Fluor-492, or -550 (Beyotime, Shanghai, China). For assays involving actin cytoskeleton staining, cells were treated with 100 nM F-actin Phalloidin (Yesen, Shanghai, China) for 30 min and protected from light. Finally, the nucleus was stained with DAPI (Solarbio, Beijing, China), and the tablet was sealed with an anti-fluorescent quench agent (Solarbio, Beijing, China) to fluorescence microscopy observation.
Cell counting kit 8 (CCK8) assay
Cell viability was determined by CCK8 assay kit (TargetMol, Beijing, China) as described previously69. Briefly, single cell suspension (1 × 103 cells/100 μL) was seeded into 96-well plates and allowed to grow. CCK-8 assays were performed at the indicated time points.
Colony formation assay
Cell survival was assessed by colony formation assay as formerly described70. Briefly, 1 × 103 cancer cells in 2 mL culture medium were seeded into 6-well plates and allowed to grow for about two weeks. When the colonies are visible, cells were fixed with 4% paraformaldehyde and stained with 1% crystal violet. Visible colonies were then counted and photographed. These assays were performed in triplicate.
Cell migration and invasion assays
Cell mobility was determined by wound-healing assay. A scratch was created in confluent monolayer cells by using microtips, then cells were cultured in medium containing 1% FBS. Images at the beginning and at indicated intervals during cell migration to close the wound were captured. Transwell assays were used to evaluate cell migration and invasion as described previously71,72. Transwell inserts (Corning, Beijing, China), precoated with or without Matrigel™ Basement Membrane Matrix (BD Biosciences, Beijing, China), were put into 24-well plates within 700 μL culture medium containing 15% FBS. Tumor cells in serum-free medium (1 × 104 cells/200 μL) were seeded into transwell inserts and incubated at 37 °C for 24–48 h, allowing cells to migrate or invade across the transwell membrane. Then the cells were fixed with 4% paraformaldehyde. Cells retained in the upper layer of inserts were carefully removed by cotton swab, whereas cells crossed the transwell membrane were visualized with crystal violet and counted under a microscope.
RNA extraction and quantitative real-time polymerase chain reaction (qRT-PCR)
Extraction of RNA from cultured cells was performed by using TRIzol Reagent (Life Technologies) as previously described73,74. DNase I (Takara, Beijing, China) was used to digest the trace amount of genomic DNA in cellular RNA samples. Reverse transcription was performed by using the RevertAid First Strand cDNA Synthesis Kits (Thermo Fisher Scientific, Beijing, China). qRT-PCR assays were performed by using SYBR Green reagent (Bimake, Shanghai, China) and CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad Laboratories, Richmond, CA, USA). We employed the 2−ΔΔCT method to assess the relative expression levels of genes. The sequences of primers used in qRT-PCR assays are provided in Supplementary Data 4.
Chromatin Immunoprecipitation quantitative real-time PCR (ChIP-qPCR)
The histone-DNA complexes from HK1 cells were cross-linked with formaldehyde and fragmented into 200–1000 bp DNA by ultrasound. Anti-H3K27ac (active motif; Cat No. 39685), Anti-H3K4me1 (active motif; Cat No. 39635), Anti-MED1 (active motif; Cat No. 61065), Anti-pol II (active motif; Cat No. 61667), were used to precipitate histone-DNA complexes, within normal human immunoglobulin G (IgG) as negative control. The DNA fragments were isolated from the precipitate and were detected by qRT-PCR with specific Primers.
siRNA, shRNA, and gene transfection
The siRNAs targeted to TP63, RACK1, and UBE2T were purchased from GenePharma (Shanghai, China). Transfection of siRNAs into cancer cells were performed by using Lipofectamine RNAiMAX (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. Transfection of plasmids into cancer cells was performed by using Lipofectamine 3000 Reagent (Invitrogen) according to the manufacture’s protocols. Stable inhibition was achieved by infecting cancer cells with shRNAs expressing lentivirus and then selected by puromycin. Sequences of siRNAs or shRNAs used in this study were listed in Supplementary Data 5.
Western blot analysis
Western blot analysis was utilized to measure the expression levels of cellular proteins75. Cell lysates were prepared by using RIPA buffer (Beyotime, Jiangsu, China) containing protease inhibitor cocktail (Roche Applied Science, USA). Proteins separation was accomplished by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Proteins were transferred from gel to PVDF membranes (Millipore, Billerica, MA), and then the PVDF membranes were blocked by incubation with 5% non-fat milk for 1.5 h. Incubation of primary antibodies was performed at 4 °C overnight. After washing with four times with PBS, the PVDF membranes were incubated with a secondary antibody coupled to HRP for 1 h. Chemiluminescent signal were developed by using enhanced chemiluminescent substrates (Thermo Fischer Scientific, Waltham, MA, USA) and recorded by ChampChemi500 system (Sagecreation, Beijing, China). The information of antibodies used in this article were provided in Supplementary Data 6.
Affinity capture of biotinylated proteins and protein identification by mass spectrometry
For BirA* proximity-labeling experiments, the full-length cDNA encoding for human Pro-IL-1β was cloned in frame 5′ of myc-BirA* in the plasmid PB510-mycBioID, creating plasmid PB510-Pro-IL-1β-mycBioID. HEK293T (RRID: CVCL_0063) cells were transfected with PB510-Pro-IL-1β-mycBioID using the polyethylenimine transfection reagent according to the manufacturer’s instructions (Polysciences, Inc., Beijing, China). Stable transfectants were established by selecting with puromycin. HEK293T cells stably transfected with PB510-mycBioID was used as control. Cells were incubated with 50 μM biotin for 24 h, then the cells were harvested, lysed. Biotinylated proteins were captured by streptavidin-conjugated beads as described previously54. The beads were washed with PBS four times, the bound proteins were eluted by boiling in the SDS-containing sample buffer and separated by SDS-PAGE. 10% of the sample was used for immunoblotting. Separated proteins were visualized by Coomassie Blue staining. The whole gel lanes were submitted to in-gel trypsin digestion and LC-MS/MS analysis at the Shanghai Zhongke New Life Biological Technology. The raw proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository76,77 with the dataset identifier PXD061201.
Co-immunoprecipitation (Co-IP)
Co-immunoprecipitation was performed as described previously14. Cancer cells were lysed with immunoprecipitation buffer (20 mM Tris, pH 7.5, 150 mM NaCl, 1% Triton ×-100) containing protease inhibitor cocktail (Roche Applied Science, USA). After centrifugation at 15,000 × g, the supernatant was incubated with primary antibodies and protein A/G agarose beads (Sigma) at 4 °C overnight. The beads were washed with PBS solution four times, boiled in SDS-PAGE sample buffer, and then subjected to Western blot analysis.
In vitro ubiquitylation assay
In vitro ubiquitination assays were performed as previously described56,78. Briefly, purified RACK1 (12498-H10E, sinobiological) and UBE2T (12440-H08E, sinobiological) were incubated together at 37 °C for 2 h in the presence or absence of Pro-IL-1β (10139-H07E, sinobiological) protein, along with E1 (UBA1, 11990-H20B, sinobiological), Ubiquitin (U06-54N, sinobiological), and ubiquitination buffer (20 mM Tris-HCl, pH 7.5, 5 mM MgCl2, 0.5 mM DTT, 4 mM ATP). The reaction was terminated by adding loading buffer and boiling for 5–10 min, followed by SDS-PAGE and Western blot analysis.
Scanning electron microscope (SEM)
The surface morphologies of the tumor cells were observed by the Hitachi SEM S-3400N (Hitachi Ltd., Tokyo, Japan). For SEM, cells were seeded on glass cover slides to reach 70%–80% confluency and fixed in 2.5% glutaraldehyde solution overnight. After that, the samples were postfixed in 1% osmic acid solution, dehydrated in different concentrations of acetone, and then infiltrated using isoamyl acetate. Probes were prepared with the Critical Point Dryer (Hitachi Ltd., Tokyo, Japan) and EIKO IB-3 gold coating (Hitachi Ltd., Tokyo, Japan).
Rho GTPase activity assay
The GTPase pull-down assay was performed using the Rho Activation Assay Kit (EMD Millipore Corporation, Temecula, CA, USA) or Rac/cdc42 Assay Reagent (EMD Millipore Corporation, Temecula, CA, USA) according to the manufacturer’s protocol. Briefly, tumor cells were cultured to approximately 80%–90% confluence in 100 mm plates. Mg2+ containing lysate buffer (125 mM HEPES, pH 7.5, 750 mM NaCl, 5% lgepal CA-630, 50 mM MgCl2, 5 mM EDTA, and 10% glycerol) was incubated together with agarose coupled to the Rhotekin Rho binding domain or incubated with agarose coupled to the Rac/Cdc42 (p21) binding domain (PBD) of PAK1 protein. After washing, the bound GTPase is eluted from the glutathione resin by the GST-fusion protein. Precipitated GTP-bound Rho or GTP-bound Rac1 and Cdc42 was subjected to the Western blot procedure described above.
CRISPR/Cas9-mediated enhancer repression
Enhancer repression was achieved by dCas9-KRAB based CRISPR interference as described previously79,80. Firstly, dCas9-KRAB was stably introduced into HK1 cells by lentivirus. HK1 cells expressing dCas9-KRAB cells were then infected with lentivirus containing sgRNAs targeted to TP63 binding sites on IL1B SEs. The sequences of sgRNAs used in this study were listed in Supplementary Data 5.
Protein-protein docking
The sequences of Pro-IL-1β protein and RACK1 protein were obtained from the Uniprot database. HDOCK server (Template-Free) (RRID: SCR_024799, http://hdock.phys.hust.edu.cn/) was used for rigid docking and adjusting the initial conformations of the two proteins. Subsequently, based on the rigid docking results, RosettaDock (RRID:SCR_013393) was employed for flexible docking, with the final models selected using Rosetta’s built-in score module for evaluation. The overall binding energy of the protein-protein interaction was analyzed using Rosetta’s Interface_analyzer module. The bonding interaction between the two proteins was analyzed using LigPlot+ (RRID: SCR_018249). The docked conformations were visualized using PyMOL (https://pymol.org/2/) (RRID: SCR_000305).
High-throughput virtual screening (HTVS)
Structure-based HTVS was performed using Schrödinger Maestro version 12.8, with the expectation of obtaining small molecule compounds with strong binding to Pro-IL-1β protein. It should be noted that the C-terminal mature domain (117–269 aa, IL-1β) of Pro-IL-1β has a resolved co-crystal structure (PDB ID: 5R8Q) with a small-molecule ligand, and can be virtually screened based on the ligand-binding pocket occupied by Z2643472210 in the mature domain. Therefore, subsequent screening and prediction were performed using the key structural domain (117–269 aa) of the full-length protein, with the ultimate goal of identifying high-affinity small-molecule binders of Pro-IL-1β.
Protein preparation: The three-dimensional (3D) structure of Human IL1B (PDB ID: 5R8Q) was downloaded from the RCSB PDB database. Protein Preparation Wizard module was used to protonate the protein, delete water molecules and SO42−, retain the A chain and the small molecule JGY (i.e., Z2643472210) binding pocket, and perform energy optimization (OPLS2005 force field, RMSD of 0.30 Å). Receptor Grid Generation module was then used to create a grid file, with the pocket occupied by Z2643472210 set as the grid center (box size: 20 Å × 20 Å × 20 Å). Compound preparation: Compound library HY-L001 (MCE Bioactive Compound Library Plus, containing 19.1 K compounds) was processed for protonation and energy optimisation by LigPrep Module of Schrödinger to output 3D structures. Molecular docking: These optimized compounds were then subjected to Virtual Screening Workflow module, and the Glide module is used for molecular docking, that is, the receptor and ligand molecules are docked with each other through geometric matching and energy matching. Firstly, compounds were screened in Glide HTVS mode, where the top-scored 15% compounds were subjected to Glide Standard Precision (SP) docking. Subsequently, the top 15% from Glide SP were selected for further screening using Glide Extra Precision (XP), and the top 50 compounds with the highest absolute docking scores were output as candidate molecules (Supplementary Data 2). An SDF file containing the top 50 compounds has been uploaded as supplementary data (Supplementary Data 7). The higher the absolute value of the predicted docking score, the stronger the binding affinity. Maestro (RRID:SCR_016748) and PyMOL were used for visualization analysis.
Cellular thermal shift assay (CETSA)
The binding of Q3MG to Pro-IL-1β was verified using the CESTA81,82. CETSA was performed as previously described83,84. Briefly, soluble protein lysates from HK1, FaDu, and BxPC3 cells were aliquoted into PCR tubes and treated with Q3MG (250 μmol/L) or DMSO for 1 h at room temperature before the CETSA heat pulse. The solution was heated at the indicated temperature (37–73 °C) for 3 min and then cooled at 4 °C for 3 min. After centrifugation for 20 min (20,000 × g, 4 °C), the soluble supernatant was subjected to Western blot analysis.
Animal experiments
Mice (BALB/c nude, Female, 4–6 weeks old) were obtained by the Department of Experimental Animals of Central South University. Xenograft tumor formation assays were performed as described previously85. Single cell suspensions (1.0 × 106 cells/200 μL) were subcutaneously inoculated into nude mice. Xenograft tumor volumes were measured and recorded weekly. Mice were euthanized once the primary tumor measured a maximal size of 2 cm in at least one dimension. Maximal tumor size was not exceeded in this study. For the tail vein metastasis model, tumor cells suspensions (1.0 × 106 cells/200 μL) were injected into nude mice via the tail vein. The body weight of nude mice injected with tumor cells were recorded weekly. Mice were euthanized if significant body weight loss (>20% of initial body weight) occurred in metastasis model. No animals reached this weight loss threshold in the study. Lungs and other main organs were isolated from the mice. Metastatic nodules at the surface of lung were observed and photographed. Tissues were then fixed in 4% formalin and embedded in paraffin. Lung sections are then sliced from paraffin-embedded lungs for H&E and other histologic staining. Metastatic nodules in each lung section were carefully examined and counted. 20 non-consecutive sections (spaced at 30–50 μm intervals) were analyzed for each mice to represent the overall heterogeneity (covering the entire lung), and each section systematically scanned the entire lung area and finally performed counting analysis. For Q3MG treatment, PBS or 15 mg/kg Q3MG was administered intravenously every 2 days starting on week 2–3 after tumor cell injection for a total of 7 times. For the orthotopic metastasis model51, tumor cell suspensions (1.0 × 106 cells/50 μL) were injected into the middle-posterior dorsal tongue, and nude mice were sacrificed 30 days later to obtain cervical lymph nodes for tumor metastasis assessment, statistics, and staining. To observe the size of primary tumors in the tongue and metastatic tumors in the cervical lymph nodes, bioluminescence imaging (BLI) was performed after intraperitoneal injection of D-luciferin and measured using the Xenogen IVIS Spectrum Living Imaging System (PerkinElmer). BLI data were analyzed using Living Image software (PerkinElmer). Animals were euthanized immediately if any of the following humane endpoint criteria were met: sustained weight loss >20%, lethargy, inability to access food or water, or any signs of severe distress. Animal experiments were approved by the Animal Welfare Committee of Central South University and conducted according to the guidance.
Statistical analysis
Statistical analysis was performed using the GraphPad Prism 8.0 (RRID:SCR_002798) and SPSS statistical software (RRID: SCR_002865). Statistical comparisons were analyzed using two-tailed Student’s t-test for two groups and analysis of variance (ANOVA) for three or more groups. All experiments were repeated at least biological triplicate. Data are expressed as mean ± standard deviation from at least three independent experimental replicates. P-value < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are available within the article, its Supplementary Information, and the associated Source Data files. Source Data are provided with this paper. The raw protein mass spectrometry data from this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository. The dataset identifier is PXD061201. Source data are provided with this paper.
References
Karki, R. & Kanneganti, T. D. Diverging inflammasome signals in tumorigenesis and potential targeting. Nat. Rev. Cancer 19, 197–214 (2019).
Thornberry, N. A. et al. A novel heterodimeric cysteine protease is required for interleukin-1 beta processing in monocytes. Nature 356, 768–774 (1992).
Tan, Y. et al. Pyroptosis: a new paradigm of cell death for fighting against cancer. J. Exp. Clin. Cancer Res. 40, 153 (2021).
Dinarello, C. A. Overview of the IL-1 family in innate inflammation and acquired immunity. Immunol. Rev. 281, 8–27 (2018).
Das, S., Shapiro, B., Vucic, E. A., Vogt, S. & Bar-Sagi, D. Tumor cell-derived IL1beta promotes desmoplasia and immune suppression in pancreatic cancer. Cancer Res. 80, 1088–1101 (2020).
Chen, L. C. et al. Tumour inflammasome-derived IL-1beta recruits neutrophils and improves local recurrence-free survival in EBV-induced nasopharyngeal carcinoma. EMBO Mol. Med. 4, 1276–1293 (2012).
Garlanda, C. & Mantovani, A. Interleukin-1 in tumor progression, therapy, and prevention. Cancer Cell 39, 1023–1027 (2021).
Ridker, P. M. et al. Effect of interleukin-1beta inhibition with canakinumab on incident lung cancer in patients with atherosclerosis: exploratory results from a randomised, double-blind, placebo-controlled trial. Lancet 390, 1833–1842 (2017).
Paz-Ares, L. et al. Canakinumab (CAN) plus docetaxel (DTX) for the second- or third-line (2/3L) treatment of advanced non-small cell lung cancer (NSCLC): CANOPY-2 phase III results. Ann. Oncol. 32, S953–S954 (2021).
Lythgoe, M. P. & Prasad, V. Repositioning canakinumab for non-small cell lung cancer-important lessons for drug repurposing in oncology. Br. J. Cancer 127, 785–787 (2022).
Apte, R. N. et al. The involvement of IL-1 in tumorigenesis, tumor invasiveness, metastasis and tumor-host interactions. Cancer Metastasis Rev. 25, 387–408 (2006).
Voronov, E. et al. Unique versus redundant functions of IL-1alpha and IL-1beta in the tumor microenvironment. Front. Immunol. 4, 177 (2013).
Yi, M. et al. TP63 links chromatin remodeling and enhancer reprogramming to epidermal differentiation and squamous cell carcinoma development. Cell Mol. Life Sci. 77, 4325–4346 (2020).
Ban, Y. Y. et al. LNCAROD is stabilized by m6A methylation and promotes cancer progression via forming a ternary complex with HSPA1A and YBX1 in head and neck squamous cell carcinoma. Mol. Oncol. 14, 1282–1296 (2020).
Johnson, D. E. et al. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Prim. 6, 92 (2020).
Agrawal, N. et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science 333, 1154–1157 (2011).
Cancer Genome Atlas, N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
Nichols, A. C. et al. High frequency of activating PIK3CA mutations in human papillomavirus-positive oropharyngeal cancer. JAMA Otolaryngol. Head. Neck Surg. 139, 617–622 (2013).
Foy, J. P. et al. New DNA methylation markers and global DNA hypomethylation are associated with oral cancer development. Cancer Prev. Res. (Philos.) 8, 1027–1035 (2015).
Li, W. et al. Tumor suppressor gene oxidored-nitro domain-containing protein 1 regulates nasopharyngeal cancer cell autophagy, metabolism, and apoptosis in vitro. Int. J. Biochem. Cell Biol. 45, 2016–2026 (2013).
Viswanathan, M., Tsuchida, N. & Shanmugam, G. Promoter hypermethylation profile of tumor-associated genes p16, p15, hMLH1, MGMT and E-cadherin in oral squamous cell carcinoma. Int. J. Cancer 105, 41–46 (2003).
Ha, P. K. & Califano, J. A. Promoter methylation and inactivation of tumour-suppressor genes in oral squamous-cell carcinoma. Lancet Oncol. 7, 77–82 (2006).
Li, W. et al. NOR1 is an HSF1- and NRF1-regulated putative tumor suppressor inactivated by promoter hypermethylation in nasopharyngeal carcinoma. Carcinogenesis 32, 1305–1314 (2011).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Bacabac, M. & Xu, W. Oncogenic super-enhancers in cancer: mechanisms and therapeutic targets. Cancer Metastasis Rev. 42, 471–480 (2023).
Cai, J. et al. DeltaNp63alpha is a super enhancer-enriched master factor controlling the basal-to-luminal differentiation transcriptional program and gene regulatory networks in nasopharyngeal carcinoma. Carcinogenesis 41, 1282–1293 (2020).
Dong, J. et al. Transcriptional super-enhancers control cancer stemness and metastasis genes in squamous cell carcinoma. Nat. Commun. 12, 3974 (2021).
Zhang, M. et al. FOSL1 promotes metastasis of head and neck squamous cell carcinoma through super-enhancer-driven transcription program. Mol. Ther. 29, 2583–2600 (2021).
Aseervatham, J. Cytoskeletal remodeling in cancer. Biology (Basel) 9, 385 (2020).
Balta, E. et al. Spatial oxidation of L-plastin downmodulates actin-based functions of tumor cells. Nat. Commun. 10, 4073 (2019).
Hodge, R. G. & Ridley, A. J. Regulating Rho GTPases and their regulators. Nat. Rev. Mol. Cell Biol. 17, 496–510 (2016).
Kosla, J. et al. Metastasis of aggressive amoeboid sarcoma cells is dependent on Rho/ROCK/MLC signaling. Cell Commun. Signal. 11, 51 (2013).
Takami, Y. et al. The activity of RhoA is correlated with lymph node metastasis in human colorectal cancer. Digest. Dis. Sci. 53, 467–473 (2008).
Kamai, T. et al. Overexpression of RhoA, Rac1, and Cdc42 GTPases is associated with progression in testicular cancer. Clin. Cancer Res. 10, 4799–4805 (2004).
Struckhoff, A. P., Rana, M. K. & Worthylake, R. A. RhoA can lead the way in tumor cell invasion and metastasis. Front. Biosci.-Landmark 16, 1915–1926 (2011).
Raftopoulou, M. & Hall, A. Cell migration: rho GTPases lead the way. Dev. Biol. 265, 23–32 (2004).
Burridge, K. & Wennerberg, K. Rho and Rac take center stage. Cell 116, 167–179 (2004).
Fukata, Y. et al. Phosphorylation of adducin by Rho-kinase plays a crucial role in cell motility. J. Cell Biol. 145, 347–361 (1999).
Palazzo, A. F., Cook, T. A., Alberts, A. S. & Gundersen, G. G. mDia mediates Rho-regulated formation and orientation of stable microtubules. Nat. Cell Biol. 3, 723–729 (2001).
Pertz, O., Hodgson, L., Klemke, R. L. & Hahn, K. M. Spatiotemporal dynamics of RhoA activity in migrating cells. Nature 440, 1069–1072 (2006).
Ridley, A. J. Rho GTPase signalling in cell migration. Curr. Opin. Cell Biol. 36, 103–112 (2015).
Cao, X. X. et al. RACK1 promotes breast carcinoma migration/metastasis via activation of the RhoA/Rho kinase pathway. Breast Cancer Res. Treat. 126, 555–563 (2011).
Cai, J. et al. Natural product triptolide induces GSDME-mediated pyroptosis in head and neck cancer through suppressing mitochondrial hexokinase-IotaIota. J. Exp. Clin. Cancer Res. 40, 190 (2021).
Lee, C. H. et al. IL-1beta promotes malignant transformation and tumor aggressiveness in oral cancer. J. Cell Physiol. 230, 875–884 (2015).
Huang, C. F. et al. NLRP3 inflammasome activation promotes inflammation-induced carcinogenesis in head and neck squamous cell carcinoma. J. Exp. Clin. Cancer Res. 36, 116 (2017).
Yang, X. et al. DeltaNp63 versatilely regulates a broad NF-kappaB gene program and promotes squamous epithelial proliferation, migration, and inflammation. Cancer Res. 71, 3688–3700 (2011).
Eyre, R. et al. Microenvironmental IL1beta promotes breast cancer metastatic colonisation in the bone via activation of Wnt signalling. Nat. Commun. 10, 5016 (2019).
Tulotta, C. et al. Endogenous production of IL1B by breast cancer cells drives metastasis and colonization of the bone microenvironment. Clin. Cancer Res. 25, 2769–2782 (2019).
Myers, J. N., Holsinger, F. C., Jasser, S. A., Bekele, B. N. & Fidler, I. J. An orthotopic nude mouse model of oral tongue squamous cell carcinoma. Clin. Cancer Res. 8, 293–298 (2002).
Sobral, L. M. et al. Stable SET knockdown in head and neck squamous cell carcinoma promotes cell invasion and the mesenchymal-like phenotype in vitro, as well as necrosis, cisplatin sensitivity and lymph node metastasis in xenograft tumor models. Mol. Cancer 13, 32 (2014).
Chen, H. H., Yu, H. I., Yang, M. H. & Tarn, W. Y. DDX3 activates CBC-eIF3-mediated translation of uORF-containing oncogenic mRNAs to promote metastasis in HNSCC. Cancer Res. 78, 4512–4523 (2018).
Lyons, S. M. et al. Changes in cell shape are correlated with metastatic potential in murine and human osteosarcomas. Biol. Open 5, 289–299 (2016).
Garcia-Mata, R., Boulter, E. & Burridge, K. The ‘invisible hand’: regulation of RHO GTPases by RHOGDIs. Nat. Rev. Mol. Cell Biol. 12, 493–504 (2011).
Reis, A. H., Xiang, B., Ossipova, O., Itoh, K. & Sokol, S. Y. Identification of the centrosomal maturation factor SSX2IP as a Wtip-binding partner by targeted proximity biotinylation. PLoS ONE 16, e0259068 (2021).
Hailey, K. L. et al. Pro-interleukin (IL)-1 beta shares a core region of stability as compared with mature IL-1 beta while maintaining a distinctly different configurational landscape A COMPARATIVE HYDROGEN/DEUTERIUM EXCHANGE MASS SPECTROMETRY STUDY. J. Biol. Chem. 284, 26137–26148 (2009).
Yu, Z. et al. A novel UBE2T inhibitor suppresses Wnt/beta-catenin signaling hyperactivation and gastric cancer progression by blocking RACK1 ubiquitination. Oncogene 40, 1027–1042 (2021).
Liu, G. Y. et al. Hepatic RACK1 deficiency protects against fulminant hepatitis through myeloid-derived suppressor cells. Theranostics 12, 2248–2265 (2022).
Vijayaraj, S. L. et al. The ubiquitylation of IL-1beta limits its cleavage by caspase-1 and targets it for proteasomal degradation. Nat. Commun. 12, 2713 (2021).
Duan, Y. et al. RACK1 mediates NLRP3 inflammasome activation by promoting NLRP3 active conformation and inflammasome assembly. Cell Rep. 33, 108405 (2020).
Liu, S. et al. RACK1 is an organ-specific prognostic predictor in OSCC. Oral. Oncol. 76, 22–26 (2018).
Dan, H. et al. RACK1 promotes cancer progression by increasing the M2/M1 macrophage ratio via the NF-kappaB pathway in oral squamous cell carcinoma. Mol. Oncol. 14, 795–807 (2020).
Wang, Z. et al. RACK1, an excellent predictor for poor clinical outcome in oral squamous carcinoma, similar to Ki67. Eur. J. Cancer 45, 490–496 (2009).
Duff, D. & Long, A. Roles for RACK1 in cancer cell migration and invasion. Cell Signal 35, 250–255 (2017).
Arnold, D. D., Yalamanoglu, A. & Boyman, O. Systematic review of safety and efficacy of IL-1-targeted biologics in treating immune-mediated disorders. Front. Immunol. 13, 888392 (2022).
Hao, B. et al. Baicalein tethers CD274/PD-L1 for autophagic degradation to boost antitumor immunity. Autophagy 21, 917–933 (2025).
Chan, S. Y. et al. Authentication of nasopharyngeal carcinoma tumor lines. Int. J. Cancer 122, 2169–2171 (2008).
Chen, S. et al. EGFR-PKM2 signaling promotes the metastatic potential of nasopharyngeal carcinoma through induction of FOSL1 and ANTXR2. Carcinogenesis 41, 723–733 (2020).
Cheung, S. T. et al. Nasopharyngeal carcinoma cell line (C666-1) consistently harbouring Epstein-Barr virus. Int. J. Cancer 83, 121–126 (1999).
Li, J. et al. FOXA1 prevents nutrients deprivation induced autophagic cell death through inducing loss of imprinting of IGF2 in lung adenocarcinoma. Cell Death Dis. 13, 711 (2022).
Ban, Y. et al. LNCAROD is stabilized by m6A methylation and promotes cancer progression via forming a ternary complex with HSPA1A and YBX1 in head and neck squamous cell carcinoma. Mol. Oncol. 14, 1282–1296 (2020).
Mo, Y. et al. Circular RNA circPVT1 promotes nasopharyngeal carcinoma metastasis via the beta-TrCP/c-Myc/SRSF1 positive feedback loop. Mol. Cancer 21, 192 (2022).
Li, J. et al. FOXA1 reprograms the TGF-beta-stimulated transcriptional program from a metastasis promoter to a tumor suppressor in nasopharyngeal carcinoma. Cancer Lett. 442, 1–14 (2019).
Cai, J. et al. Natural product triptolide induces GSDME-mediated pyroptosis in head and neck cancer through suppressing mitochondrial hexokinase-ΙΙ. J. Exp. Clin. Cancer Res. 40, 190 (2021).
Wu, P. et al. Circular RNA circRILPL1 promotes nasopharyngeal carcinoma malignant progression by activating the Hippo-YAP signaling pathway. Cell Death Differ. 30, 1679–1694 (2023).
Mo, Y. et al. Circular RNA circPVT1 promotes nasopharyngeal carcinoma metastasis via the β-TrCP/c-Myc/SRSF1 positive feedback loop. Mol. Cancer 21, 192 (2022).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Zhang, N. et al. FBXO31-mediated ubiquitination of OGT maintains O-GlcNAcylation homeostasis to restrain endometrial malignancy. Nat. Commun. 16, 1274 (2025).
Yeo, N. C. et al. An enhanced CRISPR repressor for targeted mammalian gene regulation. Nat. Methods 15, 611–616 (2018).
Chen, C. H. et al. Inhibition of super enhancer downregulates the expression of KLF5 in basal-like breast cancers. Int. J. Biol. Sci. 15, 1733–1742 (2019).
Wang, Y. et al. MCM6 is a critical transcriptional target of YAP to promote gastric tumorigenesis and serves as a therapeutic target. Theranostics 12, 6509–6526 (2022).
Xiao, H. et al. Gentiopicroside targets PAQR3 to activate the PI3K/AKT signaling pathway and ameliorate disordered glucose and lipid metabolism. Acta Pharm. Sin. B 12, 2887–2904 (2022).
Dai, L. et al. Modulation of protein-interaction states through the cell cycle. Cell 173, 1481–1494 e13 (2018).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Li, J. et al. FOXA1 reprograms the TGF-β-stimulated transcriptional program from a metastasis promoter to a tumor suppressor in nasopharyngeal carcinoma. Cancer Lett. 442, 1–14 (2019).
Acknowledgements
This study was supported by grants from The National Natural Science Foundation of China (No. U23A20456, 82272631, 82072596, 82173339, 82403859), the National “111” Project (No. Project #111-2-12), the Key Fundamental Research Program of Hunan Province (No. 2025JC0003), the scientific research program of FuRong laboratory (No. 2023SK2094), the Natural Science Foundation of Hunan Province, China (No. 2024JJ3036, 2022JJ40726, 2024JJ6544), the Key Research Project of Hunan Health Commission, China (No. 20256325), the Science and Technology Innovation Program of Hunan Province, China (No. 2024RC3029), the Beijing Xisike Clinical Oncology Research Foundation (No. YHR2020ZD-0052), the Natural Sciences Foundation of Changsha (No. kq2402235), the China Postdoctoral Science Foundation (No. 2025T180398, 2024M753678), the Postdoctoral Fellowship Program of CPSF (No. GZC20233193), the Independent Exploration and Innovation Program of Central South University (No. 1053320221155). Certain diagrams were generated using BioRender (www.biorender.com) under Academic License terms (GE28P9T7XH).
Author information
Authors and Affiliations
Contributions
Conception and design: B.X., M.Y. Development of methodology and writing of manuscript: Q.C., P.W. Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc): J.C., X.L. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): Q.C., P.W. Review and/or revision of the manuscript: Z.G., P.C., Z.Z., G.L., W.X., M.Y., B.X. Study supervision: M.Y., B.X. M.Y. and B.X. are the guarantors of this study. They accept full responsibility for the conduct of the research and had access to the data. They controlled the decision to publish.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jose Luis Orgaz Bueno, and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.”
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Q., Wu, P., Cai, J. et al. A moonlighting function of tumoral interleukin-1β precursor promotes metastasis via RACK1-mediated actin remodeling. Nat Commun 16, 10148 (2025). https://doi.org/10.1038/s41467-025-65220-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65220-1