Abstract
Melanin synthesis within melanosomes critically depends on the fibrillar architecture formed by the pigment cell-specific protein PMEL. Although PMEL fibrils have historically been classified as functional amyloids, their native supramolecular organization in situ and detailed molecular architecture have remained unresolved. In this study, we combine in situ cryo-electron tomography (cryo-ET) and cryo-electron microscopy (cryo-EM) to elucidate the native structural organization of PMEL fibrils within human melanosomes from both patient melanoma tissues and melanocyte cell lines. We demonstrate that PMEL does not form conventional isolated amyloid fibrils, but rather assembles into highly organized lamellar sheets consisting of laterally aligned fibrils interconnected by flexible linker regions. Cryo-EM structures reveal a distinctive butterfly-shaped fibril unit composed of multiple structured domains, including both the proteolytically cleaved Mα and Mβ fragments of PMEL, which assemble into a amyloid-like β-sheet arrangement. Notably, we identify intrinsically disordered regions critical for lamellar assembly and curvature and verify key glycosylation modifications in the structure. This architecture distinguishes PMEL fibrils from canonical amyloids and elucidates the molecular basis underlying melanosome integrity and pigmentation. Moreover, our work provides molecular insights relevant for pigmentation disorders and PMEL-associated diseases, including melanoma.
Similar content being viewed by others
Introduction
Skin pigmentation is vital across animals, including humans, serving protective and communicative roles1,2,3,4. Melanin synthesis and storage occur within melanosomes, which are specialized organelles in melanocytes and retinal pigment epithelial cells essential for vertebrate survival2,3,5. The morphology of such organelles is closely tied to their function6,7,8,9,10,11. Melanosomes contain prominent fibrillar sheets (lamellae) that were first observed in the 1960s and hypothesized to serve as a structural scaffold for melanosome biogenesis and melanin deposition12. In the 1990s, PMEL (also known as gp100 or Pmel17) was identified as the protein forming these fibrillar structures, with a high expression in pigment cells and a highly conserved sequence across vertebrates13,14,15. In 2008, the sheet-like morphology of PMEL within melanosomes was first observed by Hurbain et al.16. This pioneering work was later corroborated in 2019, which further established the structural organization of native PMEL sheets17. PMEL fibrils define melanosome ultrastructure and provide a biochemical platform for organized melanin polymerization and storage16,18.
Maintaining proper PMEL expression is critical for both normal pigmentation and disease19. Disruption or absence of PMEL lamellae impairs melanin production and distribution, contributing to hypopigmentation. Numerous genetic studies have linked PMEL mutations or variants to pigmentation phenotypes in animals20,21,22,23,24,25,26,27,28. In humans, loss-of-function PMEL mutations can cause pigmentary disorders, such as ocular pigment dispersion, pigmentary glaucoma, and oculocutaneous albinism29,30. Conversely, PMEL overexpression is a hallmark of melanoma cells and is associated with tumor initiation and progression31,32,33. These observations underscore that both insufficient and excessive PMEL levels are detrimental, necessitating tight homeostatic regulation of PMEL fibril formation. Despite PMEL’s importance, the native structure of its fibrillar assemblies remained poorly characterized. PMEL has long been considered a “functional amyloid” that polymerizes into insoluble fibrils within melanosomes34,35,36,37. Prior models presumed that PMEL fibrils were typical cross-β amyloid filaments composed of the proteolytically cleaved Mα fragment of PMEL17,38,39,40,41,42,43. Until now, it was unknown how individual PMEL molecules assemble to form the observed lamellar sheets in premelanosomes.
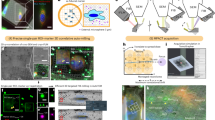
Here, we set out to determine the three-dimensional structure of native PMEL fibrils in situ and at high resolution, to resolve how these fibrils assemble into lamellae. We employ cryo-focused ion beam milling and cryo-electron tomography (cryo-ET) to directly visualize PMEL fibrils within intact melanosomes of melanoma cells, capturing their native organization. We further isolate PMEL fibrils from human melanoma cells and patient melanoma tissues and determine their structure by single-particle cryo-electron microscopy (cryo-EM). By combining these approaches, we reveal that PMEL fibrils organize into lamellar assemblies rather than existing as individual amyloid fibrils. We describe the atomic structure of the PMEL fibril lamella, identify its fundamental building blocks and symmetric assembly pattern, and elucidate how this architecture supports melanin deposition while maintaining melanosome integrity.
Results
Cryo-ET visualization of PMEL fibril structures in melanoma cells and patient tissues
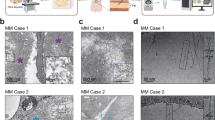
We first examined the morphology of PMEL fibrils within melanosomes from both human melanoma patient tissue and a melanoma cell line, in their native context. Conventional transmission electron microscopy of human melanoma tissue (skin melanoma) revealed abundant fibrillar material inside melanosomes, specifically in premelanosomes at stages II-III (the developmental stages where fibrils form before being fully loaded with melanin). Early stage I melanosomes contain immature fibrillar precursors, and by stage IV the fibrils are densely coated or obscured by melanin, so we focused on stage II-III organelles. Within these melanosomes, we observed two distinct fibrillar morphologies (Supplementary Fig. 1a): (i) a relatively thick structure with an uneven width, irregular fibril packing, and variable spacing, and (ii) a thinner structure of uniform width, with tightly packed parallel fibrils and a consistent inter-fibril spacing. These two forms appeared repeatedly in patient melanosomes.
To verify that these structures arise from PMEL and to observe them without the complication of melanin, we analyzed the human melanoma cell line SK-MEL-28. This cell line expresses PMEL fibrils but is amelanotic (unable to synthesize melanin), allowing clearer visualization of the fibril network44,45. Immunofluorescence confirmed that SK-MEL-28 cells indeed produce PMEL that localizes to melanosomal compartments (Supplementary Fig. 1b). Electron microscopy of SK-MEL-28 revealed fibrillar assemblies highly reminiscent of those in patient melanomas (Supplementary Fig. 1a, c). We again identified two predominant fibril organizations (Supplementary Fig. 1c): one form was thicker and irregularly arranged, while the other was thinner and more orderly, with parallel fibrils woven into a sheet-like arrangement. The consistency between patient tissue and cell line indicates that these two morphologies are inherent features of PMEL fibril assemblies, rather than artifacts of sample type.
PMEL fibrils have been widely noted to form sheet-like lamellae in premelanosomes, but until now they were often treated conceptually as individual amyloid fibrils16,46. The coexistence of two distinct morphologies led us to hypothesize that they represent different views of the same lamellar structure: specifically, a side view (edge-on perspective) corresponding to the thinner, parallel-fibril form, and a top view (face-on perspective) corresponding to the thicker, uneven form where multiple fibrils are stacked. In this model, each PMEL lamella consists of many fibrils aligned in parallel to form a sheet. When such a sheet is viewed edge-on, one would see a slender, uniform fibril profile (the width of a single fibril), whereas looking at the broad face of the sheet would show a thicker ribbon comprising many stacked fibrils. The apparent variation in thickness of the “top-view” ribbons could be explained by bending or undulation of the lamellae - flexible connections between fibrils might allow the entire sheet to curve, giving an uneven projection in 2D images while still maintaining a regular internal spacing.
To directly examine our lamella hypothesis and visualize PMEL fibrils in situ, we employed cryo-focused ion beam milling and cryo-ET on vitrified SK-MEL-28 cells. This approach enabled us to examine melanosomes within intact cells at nanometer resolution in a near-native frozen state. Tomographic 3D reconstructions of melanosomes revealed highly organized lamellar structures consistent with our proposed model. In tomographic sections, the face-on (top) view of a PMEL lamella appeared as a broad, wavy, ribbon-like density running through the melanosome (Fig. 1a, b). This sheet-like structure showed a characteristic wave-like undulation and an “up-and-down” periodic density modulation along its length (Fig. 1c–e). When viewing the tomogram cross-sections along the z-axis, we observed an array of roughly parallel fibrillar densities with consistent periodic pattern (Supplementary Fig. 2a). By segmenting the tomogram, we could isolate seven distinct lamellar sheets and two vesicles inside a single melanosome along with the bounding melanosomal membranes (Fig. 1b). The lamellae frequently appeared curved, possibly reflecting adaptation to the melanosome interior or intrinsic flexibility. In a tomographic volume rendered in a tilted orientation (a transition from top view to side view), the lamella displayed a periodic corrugation: alternate segments of the sheet rose and fell in a repeating pattern (Fig. 1c–e). These observations confirmed that native PMEL fibrils form an ordered supramolecular sheet structure within melanosomes, rather than a random tangle of fibrils. The lamellae exhibit both rigidity (maintaining regular periodic features) and flexibility (able to bend to fit the organelle’s shape).
a Original cryo-ET image of SK-MEL-28 human melanoma cells showing native PMEL fibril lamellae morphology within melanosome in situ. b–e Three-dimensional reconstructions of native PMEL fibril lamellae and melanosome from (a). b Top view of fibril lamellae (blue) and membrane (yellow). c Top-to-side view illustrating periodicity within the fibril lamella. d, e Zoomed-in images of representative PMEL fibril lamella from top and top-to-side views, highlighting periodic density patterns. f, g Cryo-EM images of SK-MEL-28 cell line extracts showing morphological characteristics of native PMEL fibril lamellae. h–j Zoomed-in images of highlighted regions in white boxes in (f, g), displaying top view, side view, and top-to-side view of native PMEL fibril lamellae. k Schematic diagram illustrating PMEL fibril lamellae as higher-order structures formed by parallel arrangement of individual PMEL fibrils through organized interactions. Data shown are representative of three independent experiments.
Extraction and cryo-EM structural determination of PMEL fibrils from cells and tissues
The in situ cryo-ET data established the lamellar organization of PMEL fibrils. To analyze their structure at higher resolution, we next isolated PMEL fibrils from melanoma cells and tissues for cryo-EM imaging. We extracted fibrils from SK-MEL-28 cells and from melanoma patient tissue by biochemical fractionation of melanosome-rich fractions (Supplementary Fig. 3a, b). The extracted material was applied to EM grids for plunge-freezing and examination by cryo-EM (Supplementary Fig. 3a).
Cryo-EM micrographs of the enriched PMEL fraction revealed three predominant fibrillar morphologies—Type 1, Type 2, and Type 3 (Fig. 1f–j and Supplementary Fig. 4). Type 1 appeared as single, long fibrils with a relatively dark, dense appearance and visible internal periodic features. Interestingly, along a Type 1 fibril, we often saw a transition: one end of the fibril displayed a dark, thick ribbon-like aspect with pronounced periodic striations, whereas moving along its length it gradually thinned and became lighter and more diffuse (Fig. 1f, h, j and Supplementary Fig. 4a, c, e). This gave the impression that a lamella (multiple fibrils together) was peeling apart into a single fibril—reminiscent of the top view of a lamella transitioning into an isolated fibril. Type 2 appeared as multiple fibrils aligned in parallel, with a lighter overall contrast and a fixed, relatively large spacing between neighboring fibrils (Fig. 1g, i and Supplementary Fig. 4b, d). This morphology corresponds well to an intact side view of a lamella, where the parallel fibrils (seen edge-on) are separated by uniform gaps (likely containing solvent or disordered regions). Type 3 structures were intermediate: they consisted of a small bundle of parallel fibrils with spacing that was variable and generally smaller than in Type 2, often appearing at the ends of Type 1 fibrils (Fig. 1f, j and Supplementary Fig. 4a, e). Type 3 can be interpreted as a partial lamella tilted relative to the grid - essentially a top-to-side intermediate view of the lamella, where we see a few fibrils grouped (more than one, but not a fully flat sheet). The coexistence of these three forms in our cryo-EM sample strongly supports our model that Types 1 and 2 are simply different orientations of the same lamellar assembly (face-on vs edge-on), and Type 3 represents a lamella caught at an oblique angle or a lamella that is flexing. The ability to observe continuous transitions between these forms (e.g., a Type 1 fibril end merging into a Type 3 small bundle, then into a wider Type 2 bundle) suggests that PMEL lamellae are flexible sheets that can partially dissociate or bend without losing their periodic internal structure. Taken together, these cryo-EM observations are fully consistent with the in situ cryo-ET results (Fig. 1a–e), reinforcing the idea that PMEL “fibrils” in melanosomes are arranged as lamellae that can appear differently depending on orientation.
Having visually confirmed the lamellar nature of PMEL assemblies, we proceeded to determine their 3D structure by single-particle cryo-EM analysis. The lamella is a complex supramolecular assembly that does not fall neatly into standard single-particle or helical reconstruction categories. We developed a hybrid strategy to tackle this: we treated segments of individual fibrils as particles for single-particle analysis, while recognizing that these fibrils are part of a larger sheet. From the SK-MEL-28 fibril preparations, we manually picked all discernible single fibrils in lamellar clusters and extracted particle images of these fibril segments. Two-dimensional (2D) classification of these segments in cryoSPARC cleanly separated them into three groups corresponding to the fully edge-on (side) view, the fully face-on (top) view, and a range of intermediate tilt views of the lamella (Fig. 2a and Supplementary Fig. 5a). This classification recapitulated the three morphological types seen in raw images. Notably, even in side-view classes, a faint but regular density between adjacent fibrils was apparent, hinting at specific interactions linking fibrils in the lamella (Fig. 2a, side-view classes). The top-view classes showed pronounced periodic striations along the fibril length, reflecting the internal repeating unit of the fibril structure. We interpreted these classes as projections of a lamellar sheet from different tilt angles. By arranging representative class averages from the flattest (side view) to the most upright (top view), one can imagine the lamella gradually tilting relative to the viewing direction (Fig. 2a, schematic). As the lamella tilts, the apparent spacing between fibrils in the projection decreases and the fibril density (contrast) increases, transitioning smoothly from the separated-fibril view to the overlapping-fibril view. This analysis further cemented our understanding of the lamella’s geometry and confirmed that the dataset captured views spanning the full 0°–90° range of lamella tilt.
a Representative morphologies of PMEL fibril lamella extracted from SK-MEL-28 cells in the cryo-EM 2D classification. Schematic diagrams above each 2D images illustrate relationship between 2D and 3D morphologies of the PMEL fibril lamella. b–j Typical views of the overall architecture of native PMEL fibril lamellae. Side view (b), top view (e) and top-to-side view (g) displayed. Unit −1, 0, and +1 in fibril 0 colored in yellow, orange, and red, respectively. Linker regions marked in (c). h–j Zoomed-in images of the corresponding highlighted regions in (b, e), with cartoons depicting two-fold rotational symmetries in (h–j). The central red circular arrows in the figure denote the symmetrical arrangement.
Using the classified particle images, we attempted an ab initio 3D reconstruction treating the lamella as a single particle. The initial 3D reconstruction, using a subset of well-aligned particles, produced a volume consistent with a lamella comprising several parallel fibrils (Supplementary Fig. 5a). We identified a two-fold symmetry axis (C2 symmetry) down the center of a fibril when viewed from the top, suggesting an internal symmetry in the fibril’s subunit organization. Imposing this C2 symmetry during refinement improved the map quality. Through iterative rounds of heterogeneous refinement and addition of more particle images (including a second round of particle picking to bolster under-represented views), we converged on a 3D reconstruction of the PMEL fibril lamella at an overall resolution of ~4.9 Å (Supplementary Fig. 5a). This reconstruction resolved a segment of the lamella containing three parallel fibrils, each fibril consisting of three repeating units along its length (Fig. 2b). The map was of sufficient quality to discern β-sheet architecture and helical secondary structures in many regions, enabling us to begin atomic modeling. To validate the C2 symmetry of the native PMEL fibril, we performed an identical data processing workflow imposing C1 symmetry, which yielded a PMEL fibril lamella map at an overall resolution of ~5.7 Å (Supplementary Fig. 6a). The resulting map was virtually indistinguishable in morphology and detail from that obtained under C2 symmetry, thereby confirming the C2 symmetry of the native PMEL fibril lamella.
To ensure that the structure we obtained from SK-MEL-28 material is representative of native PMEL fibrils in general, we performed the same cryo-EM analysis on fibrils extracted from an independent melanoma source. We purified fibrils from melanoma patient tissue (the same source as the initial TEM) and from MNT-1 cells, another well-established pigmented human melanoma cell line16. Cryo-EM imaging and 3D classification of these samples revealed lamellar fibril structures indistinguishable from those in SK-MEL-28 (Supplementary Figs. 5a and 7a, b). Because PMEL fibrils derived from patient tissues and MNT-1 cells are heavily loaded with melanin, making it difficult to clearly and fully discern the underlying PMEL fibrils, preventing reconstruction of the fine structural details of the fibrils at atomic resolution (Supplementary Fig. 7a, b). However, the lamellae from patient tissue and MNT-1 cells exhibited the same three morphological views and the same fibril spacing and periodicity. They all consist of parallel fibrils of identical morphology, directly interconnected with each other, and exhibiting the same C2 symmetry (Supplementary Figs. 5a and 7a, b). Thus, the PMEL fibril lamella structure is conserved across different human sources, and the high-resolution insights gained from SK-MEL-28 fibrils are biologically relevant and not an artifact of a single cell line.
With a 3D map of the lamella in hand, we examined its overall architecture. The reconstruction (Fig. 2b–g) confirmed that PMEL fibrils are organized in parallel within the lamella and uncovered multiple symmetry relationships. In the clearest side-view representation (Fig. 2b), we see three parallel fibrils (labeled fibril −1, 0, +1 from top to bottom). Each fibril contains three copies of a characteristic butterfly-shaped density (labeled unit −1, 0, +1 from left to right within fibril 0). These units repeat along the fibril’s length. Fibril 0’s units are colored yellow (unit −1), orange (unit 0), and red (unit +1) in Fig. 2b for clarity. A 90° rotation to look at the lamella end-on (Fig. 2c) highlights the regions that connect adjacent fibrils. These inter-fibril linkers appear as small bridging densities between fibrils, and their positions correspond well with the faint densities between fibrils observed in cryo-ET (compare Fig. 2c with Fig. 1e).
Examining the lamella from the top (Fig. 2e), we observe the packed arrangement of multiple fibrils side by side, consistent with the 2D class averages of top-view lamellae. Each fibril unit (the butterfly shape) exhibits its own internal two-fold symmetry. Indeed, when focusing on a single unit (Fig. 2h), we found that it consists of two equivalent halves (we term these chain A and chain B for the two symmetric halves of a unit). Thus, how do these units relate to one another to build the fibrils and lamella? We found that adjacent units along a single fibril are related by a 180° rotation (a two-fold symmetry) along the fibril axis (compare Fig. 2f with Fig. 2b). This implies that each fibril is constructed by a repeating two-fold symmetry operation, which flips each unit upside-down relative to its neighbor. Notably, two adjacent units in a fibril also exhibit a two-fold rotational symmetry within the lamella plane, see unit −1(yellow) and unit 0 (orange) in Fig. 2i. Finally, when comparing neighboring fibrils, we observed a symmetry relationship between units in fibril 0 and fibril +1 as well. Specifically, unit 0 of fibril 0 and unit −1 of fibril +1 are arranged as approximate two-fold symmetric images across the inter-fibril linker (Fig. 2j). The same holds for unit −1 of fibril 0 and unit 0 of fibril +1 (Fig. 2j). This indicates that the linker region between fibrils lies on a symmetry axis that relates the units in adjacent fibrils. In summary, the lamella displays three levels of C2 symmetry: within each unit (chain A vs chain B), along each fibril (unit-to-unit repetition), and between adjacent fibrils (across the linker). Such a highly symmetric assembly is remarkable for a functional amyloid scaffold and underscores the regular, reproducible nature of PMEL fibril packing. Having defined the overall morphology and symmetry of the lamella, we next sought to build an atomic model to explain the density map and these symmetry relationships in molecular terms.
The fibril unit structure of PMEL lamella
To elucidate the fundamental repeating unit structure within PMEL lamellae, we focused on resolving the butterfly-shaped density segment corresponding to one fibril unit. Utilizing advanced local refinement techniques, including per-particle defocus (CTF) refinement and masked sub-volume refinement, we successfully improved the resolution of the map for fibril 0, comprising three units, to ~3.5 Å and local resolution to ~2.7 Å (Supplementary Fig. 5b, c and 8a). This enhanced resolution allowed us to perform detailed de novo atomic model building (Supplementary Fig. 8b), revealing how individual PMEL polypeptides fold and assemble into fibrils. We also performed identical local refinement under C1 symmetry, which yielded a map at an overall resolution of ~4.1 Å, with local resolution extending to 3.1 Å (Supplementary Fig. 6b, c). The map density obtained under C1 symmetry was indistinguishable from that derived under C2 symmetry, except for the moderate resolution improvement achieved with C2, further corroborating the C2 symmetry of the native PMEL fibril (Supplementary Figs. 5b, c and 6b, c).
Each fibril unit consists of two identical PMEL polypeptide chains (chain A and chain B) related by two-fold symmetry. For simplicity, we describe the unit in terms of chain A, which is structurally identical to chain B. Chain A comprises two distinct PMEL molecules, referred to as horizontal and vertical subunits, based on their orientation relative to the fibril axis (Fig. 3a–f). Each PMEL molecule adopts a compact conformation, and two molecules (one horizontal, one vertical) pair through tight β-sheet interactions near their C-terminal regions, forming a contiguous and stable density. Therefore, a single PMEL fibril unit contains a total of four PMEL molecules-two each in chains A and B-organized symmetrically into horizontal and vertical orientations. Given the remarkable structural similarity between horizontal and vertical subunits, we primarily utilized the horizontal subunit for model building and detailed analysis (Fig. 3e–h).
a, b Cryo-EM map density of one unit of PMEL fibrils extracted from SK-MEL-28 cells. Two chains (chain A and chain B) within a unit colored in green and purple, respectively. c, d Atomic model of a unit and its corresponding map density. The two chains colored in green and purple are chain A and chain B, respectively. e, f Atomic model of chain A within a unit. Chain A is composed of a horizontal subunit and a vertical subunit. Each subunit is a PMEL molecule. g, h The atomic model of the horizontal subunit of chain A within a unit. i The atomic model of the four polypeptides within a subunit. j A planar diagram of a subunit illustrating the spatial relationship among β-sheets in a subunit. The colors of β-sheets are corresponding to those in the atomic model (e–i). k A mapping diagram showing four chains and three IDRs identified in this study, signal peptide (SP), enzyme cleavage sites, and domains reported previously in the sequence of the full-length PMEL protein.
Notably, each PMEL molecule is not represented as continuous density from N- to C-terminus. Instead, the density corresponding to a single molecule is segmented into four ordered regions interconnected by flexible linkers. We designated these structured segments as chains α, β, γ, and δ (Fig. 3i–k and Supplementary Fig. 9a–c), separated by intrinsically disordered regions (IDR1, IDR2, and IDR3). Based on sequence alignment and known PMEL domain architecture, we assigned chain α to residues ~V66-N89, chain β to ~S148-L222, chain γ to ~H231-L298, and chain δ to ~C475-Q491 (Fig. 3k). These assignments align with PMEL’s known domain organization: chain α belongs to N-terminal fragment (NTF), chain β precisely matches the core amyloid fragment (CAF), chain γ corresponds to the entire polycystic kidney disease (PKD) domain, chain δ in located within the Mβ fragment preceding the transmembrane domain, and IDR3 encompasses the repeat (RPT) region38,39,41,42,47,48 (Fig. 3k).
All four structured segments predominantly adopt β-sheet-rich conformations, forming a stable core supplemented by loops and turns. Chain β bridges the N-terminal (α + β) and C-terminal (γ + δ) domains, creating a β-sheet sandwich structure involving substantial inter-chain interactions (Fig. 3g, h). Structural examination revealed additional density consistent with an N-linked glycan at residue N81 within chain α, validating earlier studies that residue N81 carries an N-linked glycosylation modification43,45,49,50 (Supplementary Fig. 9d).
An interesting observation from our structural data is the incorporation of the Mβ fragment (chain δ) within the fibril core. Contrary to previous assumptions that only the Mα fragment polymerizes, we observed chain δ forming integral β-sheet interactions with chain γ, indicating that nearly the entire luminal region of PMEL participates in fibril assembly5,41.
Unlike previous conceptions that PMEL was classical amyloid fibril with β-sheet stacks, our study reveals that the native PMEL fibril is fundamentally built from well-folded PMEL protein molecules. To further explore the potential correspondence between the PMEL monomer and its fibrillar assembly, we employed AlphaFold 3 to predict the structure of the full-length protein (Supplementary Fig. 8c). The cryo-EM-resolved segments (chains α, β, γ, and δ) exhibited striking consistency with the AlphaFold 3-predicted structure (Supplementary Fig. 8d, f). Moreover, AlphaFold 3 predictions captured the intrinsic disorder of inter-chain regions, with IDR1 (between chains α and β) showing partial disorder and IDR3 (between chains γ and δ) appearing fully disordered (Supplementary Fig. 8e). These findings indicate a remarkable structural coherence between PMEL in its monomeric state and within the fibril architecture.
Our comparative sequence analysis demonstrated high conservation of structured segments (α-δ) across diverse vertebrate species, indicating strong evolutionary constraints and underscoring the critical functional roles of these fibrillar regions (Supplementary Fig. 10a). Intriguingly, the glycosylation site at N81 shows variable conservation, suggesting adaptive roles in higher vertebrates (Supplementary Fig. 9e). Although N81 is indeed N-glycosylated in human PMEL fibrils, both previous studies and our mutagenesis study demonstrate that this modification is not essential for PMEL fibril formation or architecture43,47 (Supplementary Fig. 11a, b).
Structural architecture of individual PMEL fibrils
With the fibril unit structure defined, we next investigated how these units stack and organize into a single PMEL fibril. Each fibril consists of repeating units spaced evenly (~108 Å apart), forming an elongated fibril with distinctive symmetry (Fig. 4a–h). Specifically, each fibril is composed of two intertwined protofilaments (designated as chain A and chain B) that twist around each other to form successive structural units (Fig. 4f–h). Within this framework, each structural unit comprises two symmetrically related chains (chain A and chain B), each composed of paired horizontal and vertical subunits (Fig. 4a–e). Consequently, the fibril presents as an alternating stack of these paired subunits, creating an extended and symmetrical structural motif throughout the fibril length (Fig. 4d, f). Although adjacent units within the fibril are inverted relative to each other, each maintains identical internal organization, explaining the repetitive appearance of units within the three-dimensional cryo-EM maps (Fig. 2b, f).
a–h A schematic diagram illustrating step-by-step assembly of the native PMEL fibril extracted from SK-MEL-28 cells by unit. a The atomic model of horizontal and vertical subunits of chain A in unit 0. b The atomic model of chain A in unit 0. c The atomic model of chain B in unit 0, which exhibits C2 symmetry with chain A in unit 0 along the Z axis. d The atomic model of unit 0, composed of chain A and chain B. e The atomic model of chain B in unit −1, which exhibits C2 symmetry with chain A in unit 0 along the Y axis. f The atomic model of a native PMEL fibril, showing three adjacent units. g, h Chain A colored in green and chain B colored in purple in the native PMEL fibril. i–o Binding sites both within a unit and between units in the native PMEL fibril. i The atomic model of chain A comprising three units, illustrating the binding details between units (j, k) and between subunits (l). m The atomic model of the unit 0, highlighting the binding details between chain A and chain B within one unit (n, o).
We next examined the interactions responsible for stabilizing the stacking of fibril units within a single PMEL fibril. First, at the N-terminal boundary, fibril stacking is primarily maintained by relatively modest yet specific hydrogen bonding interactions involving Tyr189 residues from chain β of adjacent fibril units (Fig. 4i–k). To verify the essential role of Tyr189 in PMEL fibril assembly, we expressed the Tyr189Ala mutant in PMEL-knockout SK-MEL-28 cells. Both immunofluorescence and TEM analyses showed that the Tyr189Ala mutant failed to form structurally normal PMEL fibrils (Supplementary Fig. 11a, b), consistent with previous reports42. These symmetric Tyr189-Tyr189 interactions at the N-terminal β-sheet edge provide flexibility and act as molecular hinges, accommodating minor positional adjustments as the lamellar assembly curves or bends.
Conversely, at the C-terminal interface, adjacent fibril units engage in extensive β-sheet interactions, forming a robust structural link. Specifically, residues from the horizontal and vertical subunit pairs of successive units create an extensive, heterogeneous eight-stranded β-sheet spanning the boundary between subunit pairs. Each subunit contributes four β-strands to this extended β-sheet, forming a continuous eight-stranded arrangement across the interface (Fig. 4l; Supplementary Fig. 12a–g). Such interfaces are structurally analogous to classical amyloid cross-β motifs but involve different β-sheets rather than homotypic stacking of identical segments. We refer to these as heterotypic amyloid-like β-sheets, which confer remarkable mechanical stability by strongly anchoring adjacent paired subunits together34,51. Classical cross-β sheet amyloid fibrils are formed by layers of identical polypeptides stacked through β-sheet interactions. In contrast, the eight peptide chains at the core of a PMEL fibril unit form β-sheets in which each layer is distinct, giving rise to the term heterotypic amyloid-like β-sheets (Supplementary Fig. 12h). Therefore, native PMEL fibrils represent a structural class of amyloid-related assemblies, blending globular, folded domains with localized amyloid-like interactions.
Finally, we analyzed the interactions between the two intertwined protofilaments (chains A and B) within each fibril. Within each fibril unit, these protofilaments are positioned side-by-side, stabilized by reciprocal hydrogen bonding networks between their C-terminal domains. We found that the chain B vertical subunit interacts with both the horizontal and vertical subunits of chain A, and vice versa. Owing to this symmetry, we analyzed the chain B vertical subunit as a representative (Fig. 4m). Specifically, Ser482 and Ser484 of the chain B vertical subunit form stable hydrogen bonds with Arg272 of the chain A horizontal subunit (Fig. 4n and Supplementary Fig. 12i, j). In addition, Asp255 of the chain A horizontal subunit interacts with Ser484 of the chain B vertical subunit and with Arg272 of the chain A horizontal subunit, likely reinforcing the stability of this interface (Fig. 4n). Meanwhile, Tyr480 of the chain B vertical subunit forms a hydrogen bond with Arg272 of the chain A vertical subunit. Within the chain A vertical subunit, Asp255 and Ser257 also interact with Arg272, further stabilizing the Arg272-Tyr480 interaction (Fig. 4o and Supplementary Fig. 12k, l).
Strikingly, both the horizontal and vertical subunits of chain A interact with the chain B vertical subunit through the same triad of residues-Arg272, Asp255, and Ser257-at two distinct sites. These observations suggest that single mutations of Arg272, Asp255, or Ser257 may partially impair PMEL fibril formation, whereas combined mutations could disrupt fibril assembly. To test this, we expressed single or combined PMEL point mutants in PMEL-knockout SK-MEL-28 cells (Supplementary Fig. 11a). TEM showed that Arg272Ala and Asp255Ala; Ser257Ala mutants failed to assemble intact fibril lamellae, whereas the Arg272Ala; Asp255Ala; Ser257Ala triple mutant completely abolished fibril formation (Supplementary Fig. 11a, b). Immunofluorescence staining with HMB45 further confirmed these findings: weak signals were still detected in Arg272Ala and Asp255Ala; Ser257Ala mutants, but were completely lost in the Arg272Ala; Asp255Ala; Ser257Ala triple mutant, similar to the Tyr189Ala mutant (Supplementary Fig. 11a, b). Together, these results demonstrate that Arg272, Asp255, and Ser257 are essential for stabilizing the interactions between chain A and chain B. The involvement of multiple interaction sites likely enhances fibril robustness, providing a mechanism to buffer against single-point mutations.
Collectively, our structural analyses reveal that PMEL fibrils assemble through a coordinated interplay of flexible N-terminal contacts and robust C-terminal β-sheet interactions, generating a fibril structure that balances stability with flexibility. This precise molecular architecture explains how PMEL fibrils form the lamellar scaffolds essential for melanin deposition within melanosomes.
Assembly of PMEL fibrils into lamellae
Analysis of the junctions between adjacent fibrils revealed that the map density at the linker region, although insufficient for unambiguous model building, aligns well with the expected position of IDR1 (residue ~90–147, yellow, Fig. 5a, b). Guided by the validated consistency between the AlphaFold 3-predicted and experimentally resolved PMEL structures, we used the AlphaFold 3 model of chains α-δ plus IDR1 to probe the linker architecture (Fig. 5c, with IDR1 shown in red). Fitting into the fibril lamella map showed that the linker region is formed by IDR1 from four neighboring subunits, arranged as two C2-symmetric pairs (Fig. 5d–f). Specifically, the fibril −1 chain B unit −1 horizontal subunit (light green) and the fibril 0 chain A unit 0 horizontal subunit (purple) exhibit C2 symmetry, while the fibril −1 chain B unit 0 vertical subunit (dark green) and the fibril 0 chain A unit −1 vertical subunit (gray green) also form a C2-symmetric pair (Fig. 5f). We therefore modeled IDR1 schematically, revealing that the linker serves as the symmetric hub connecting adjacent fibrils (Fig. 5g). This linker is not fully disordered, as density is still visible, but it retains flexibility that enables the fibril lamella to bend and accommodate the spherical interior of the melanosome.
a, b The atomic model of native PMEL lamella and its cryo-EM map density, showing three paralleled fibrils. The linker region located between fibril −1 and fibril 0, and unit −1 and unit 0 is highlighted in yellow. c Comparison of the AlphaFold 3-predicted PMEL core structure with the experimentally resolved structure, highlighting the predicted IDR1 (red). d–f Fitting of the AlphaFold 3-predicted PMEL structure into the linker region between adjacent fibrils, shown in side and top views. Enlarged view of the linker, illustrating contributions from four neighboring subunits (f). g A magnified view of the linker region is provided, with the schematic structure of IDR 1 represented by red dashed lines. h A mapping diagram highlighting IDR 1 (red) within the full-length PMEL protein sequence.
Another large disordered segment of PMEL is IDR3 (residue ~299–474), encompassing the RPT region known to be rich in tandem repeats and heavily O-glycosylated. IDR3 is entirely invisible in our density, implying it protrudes away from the ordered core without a defined structure. However, functional studies have shown that modifying or deleting portions of the RPT region dramatically affects fibril sheet architecture and melanosome morphology17,30. This suggests that, although not part of the rigid fibril core, the RPT region likely plays a regulatory or spacing role. It could act as a molecular spacer or brush that keeps lamellae apart at an optimal distance or that controls how melanin intermediates access the fibril surface. Its heavy glycosylation could also influence the hydration and charge environment between fibrils. We speculate that IDR3 helps prevent fibrils from aggregating uncontrollably and ensures the lamellar sheets are appropriately organized for melanin to be deposited on their surfaces. In summary, adjacent PMEL fibrils are joined into lamellae primarily through N-terminal disordered region interactions (IDR1 as a flexible linker), while the large RPT domain (IDR3) likely modulates higher-order packing and melanin interactions. This assembly strategy—using flexible linkers to build a pliable sheet—is well-suited to the dynamic environment of the maturing melanosome, where the matrix must accommodate increasing melanin content and possibly undergo remodeling.
To assess the requirement of IDR1 and IDR3 in PMEL fibril lamella formation, we generated PMEL truncation mutants lacking either IDR1 or IDR3 and expressed them in PMEL-knockout SK-MEL-28 cells (Supplementary Fig. 11a). Immunofluorescence and TEM analyses revealed that deletion of IDR1 or IDR3 abolished HMB45 staining of PMEL fibrils and prevented the formation of intact fibril lamellae within melanosomes (Supplementary Fig. 11a, b). These findings demonstrate that both IDR1 and IDR3 are essential for PMEL fibril lamella assembly.
Discussion
In this study, we provide a comprehensive structural analysis of native PMEL fibrils within human melanosomes, integrating in situ cryo-ET and high-resolution cryo-EM. We show that PMEL fibrils do not exist as isolated amyloid filaments in cells, but rather form a network of regularly arranged lamellar sheets. We resolved the architecture of these lamellae from both melanoma patient tissue and cell culture, finding a highly conserved architecture in all cases. The phrase “points form lines, and lines form planes” aptly summarizes the hierarchical assembly we uncovered: individual PMEL molecules (points) assemble into fibrils (lines), and fibrils align into lamellae (planes). This work delivers the atomic-level insight into the PMEL fibril lamella and elucidates how this supramolecular scaffold is built and organized within premelanosomes (Fig. 6). We note that a recent study reported the cryo-EM structure of cell-derived PMEL fibrils, revealing a classical cross-β sheet amyloid architecture52. In that study, the extraction process involved de-glycosylation, protease digestion, and sonication52, whereas our extraction protocol was much milder and did not include these steps (Supplementary Fig. 3a). It’s possible that these harsher treatments may result in disruption of the native lamellar structure of PMEL fibrils and induction of the new amyloid fibril formation. Nevertheless, we do not exclude the possibility that, under specific cellular or pathological conditions, PMEL fibrils within melanosomes may adopt cross-β sheet amyloid morphologies.
PMEL fibril lamellae are broadly conserved across animal species, exhibiting both physiological and pathological significance. This study presents the atomic resolution structure of the human native PMEL fibril lamella. PMEL fibrils are formed from well-folded protein domains that pack together into a larger assembly. The fundamental unit of the lamella is a single PMEL molecule, which forms either a horizontal or a vertical subunit. A horizontal subunit and a vertical subunit interact via their C-terminal domains to assemble chain A of a unit. Two symmetric chains, Chain A and Chain B, engage in interactions to form a single unit, exhibiting a butterfly-like shape. A unit extends by interactions at its N-terminal domains, contributing to the formation of a single PMEL fibril. Individual PMEL fibrils interact through their intrinsically disordered regions (IDR 1, red dashes) near the N-terminal domains, forming a network structure that constitutes the complete PMEL fibril lamella. The flexible interactions at the binding sites enable the PMEL lamella to bend and conform to the internal morphology of the melanosome. Created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Our findings revise the previous models of PMEL fibril structure. Unlike canonical amyloid fibrils, which typically consist of a single repeating peptide stack forming a cross-β spine, PMEL fibrils are formed from well-folded protein domains that pack together into a larger assembly5,16,34,36,37,40,41,48,53,54,55,56,57. Each PMEL molecule contributes multiple β-sheet-rich segments (chains α-δ) that fold into two domains, connected by a linker. Two PMEL molecules interact via C-terminal β-sheets (horizontal + vertical subunits), creating a subunit pair that is far more globular and complex than a typical amyloid protofilament. These paired subunits then polymerize with C2 symmetries to form a fibril, and pairs of fibrils associate via symmetric linkers to form the lamella. Thus, the PMEL fibril lamella is a hybrid structure: it contains amyloid-like β-sheet elements (specifically, the inter-subunit β-sheet that yields an eight-stranded cross-β structure) embedded in a framework of globular protein assembly. This reconciles a longstanding paradox: PMEL was known to form amyloid fibrils, yet parts of the protein (e.g., the PKD domain) are predicted to be structured and not typical amyloid sequences. Our structure shows that indeed PMEL fibrils are not purely β-steric zippers of a short motif; instead, they rely on large structured domains assembling much like pieces of a puzzle, supplemented by local amyloid-like bonding to lock them together. The result is a heterogeneous protein assembly architecture, where a mix of classical cross-β interactions and domain-domain interactions coexists in one complex. This insight expands our understanding of what functional protein fibrillar polymer structures can look like, blurring the line between amyloid fibrils and ordered protein polymers.
The key structural domains identified in this study show a remarkable concordance with those previously characterized by mass spectrometry in native PMEL fibrils. Notably, the β chain identified here (aa 148-222) exhibits striking similarity to the CAF region (aa 148-223) reported by Hee et al.42, providing cross-validation from structural biology and mass spectrometry for the key contribution of this domain to PMEL fibril architecture. Likewise, the α chain identified in this study (aa 66-89) aligns closely with the aa 70-90 region described by Leonhardt et al.47, again providing dual confirmation of the PMEL fibril structure.
Furthermore, previous studies have identified several critical residues within the NTF and CAF regions, including Asp73, Pro75, Leu77, and Phe84 in the NTF, and Phe149, Tyr151, Trp153, Trp160, Leu170, Ile172, Tyr189, Phe207, and Ile209 in the CAF42,43. In the native PMEL fibril structure resolved in this study, Phe149, Tyr151, Trp153, Leu170, Trp160, Phe84, Ile172, Phe207, and Ile209 form extensive hydrophobic interactions, collectively establishing a hydrophobic core within the N-terminal β-sandwich, which is critical for the stability of this region (Supplementary Fig. 13a). Notably, Tyr151 is positioned between Phe149 and Trp153, with their aromatic side chains engaging in π-π stacking (Supplementary Fig. 13b). This explains why substitution of Tyr151 with another aromatic residue (phenylalanine) preserves fibril formation, whereas replacement with an aliphatic hydrophobic residue (leucine) disrupts assembly42. Similarly, Tyr189 serves as a key residue mediating inter-unit interactions, likely contributing both hydrogen bonding and L-type π-π stacking between adjacent units (Fig. 4j, k). This structural role explains why mutations of Tyr189 to other aromatic residues maintain fibril stability42. In addition, Pro75 is located at a loop turn, where substitution with other residues could disrupt the local loop conformation (Supplementary Fig. 13c). Leu77 engages in hydrophobic interactions with Val213, Leu251, and Tyr250, thereby stabilizing the interface between chains α and β with chain γ (Supplementary Fig. 13d). Asp73 lies at the interface between chains α and γ, with its side chain closely juxtaposed to those of Pro247 and His245. Substitution with lysine or glutamic acid could introduce steric clashes due to longer side chains, thereby compromising the stability of the α-γ interface47 (Supplementary Fig. 13e). Collectively, the native PMEL fibril structure determined in this study is highly consistent with prior mutagenesis-based cellular studies, further validating the accuracy of our structural model.
Establishing that the PKD domain directly contributes to the native PMEL fibril core represents a key conceptual advance of this study. This possibility had long been speculated for several reasons. First, PMEL fibrils were shown to react with the antibodies HMB50 and NKI-beteb58, although the poorly defined epitopes of these reagents left open the possibility that they instead recognize the extreme C-terminus of the CAF. Second, the PKD domain must have been at least transiently associated with the nascent amyloid, given its inclusion in the MαC fragment59. Yet, because no antibody was available to detect the PKD domain by Western blotting, it remained unclear whether the domain was subsequently released from the amyloid through proteolysis. Interestingly, Watt and colleagues showed in 2009 that the PKD domain is highly amyloidogenic in vitro39, leading them to propose that it might either form or contribute to the PMEL amyloid core. This study validated this prediction by cryo-EM structure.
This study provides direct structural evidence to resolve a long-standing controversy in the field, namely the nature of the PMEL amyloid core. For years, numerous reports proposed that the RPT domain constitutes the fibrillar core, primarily based on in vitro assays with non-glycosylated, bacterially expressed protein40,56,60,61,62,63,64. This view persisted despite subsequent evidence demonstrating that the RPT domain is dispensable for amyloid formation in vitro39 and in vivo17,47, where its deletion does not impair fibril formation but disrupts the characteristic sheet morphology. By resolving the native PMEL fibril structure, our study provides a structural resolution of this fundamental question.
A notable discovery is the structural integration of the Mβ fragment into the fibril architecture. Previously, Mβ was thought to primarily function as a transient anchor for Mα before being cleaved and discarded38,41,65. However, our structural data show that a segment of Mβ (chain δ, residues 475–491) is stably incorporated into the fibril, forming key β-strand interactions with the C-terminal domain. This suggests that after initial proteolysis, the liberated PMEL unit is not solely Mα but rather an Mα-Mβ complex, which then undergoes polymerization. The presence of Mβ contributes to fibril stability and may regulate fibril assembly, mirroring proteolytic control mechanisms seen in other amyloidogenic systems. This finding redefines our understanding of PMEL fibrillogenesis and suggests broader biological parallels in how cells control functional amyloid assembly through stepwise proteolysis.
The PMEL fibril lamella acts as a scaffold that optimizes melanin polymerization while maintaining melanosome integrity5. The flexible IDR1 linkers allow fibrils to form curved, adaptable lamellae that can accommodate growing melanin deposits. This ability to bend ensures that melanosomes can efficiently pack fibrils while preventing the formation of large, amorphous aggregates. Additionally, the repeat domain (IDR3), although unresolved in our maps, likely functions as a spacer that regulates fibril packing. Its glycosylation may prevent premature aggregation, ensuring that fibril networks remain organized until melanin deposition is complete17.
Notably, a single PMEL fibril unit may comprise the four chains (α, β, γ, and δ) derived from different PMEL protein molecules. During maturation, PMEL undergoes multiple proteolytic cleavages: Mα separates from Mβ while remaining linked via a disulfide bond38, followed by further cleavages that generate MαN (NTF + CAF) and MαC (PKD + RPT) fragments59 and separate NTF from CAF and PKD from RPT39,42,59. Previous work also showed that distinct PMEL variants can co-assemble into the same fibril47, supporting the notion that chains within a single fibril unit may originate from different PMEL protein molecules. Thus, the mature PMEL fibril is not a single intact protein but a structural assembly of four proteolytically separated chains, potentially derived from multiple PMEL molecules.
Finally, known PMEL mutations associated with pigmentary disorders, such as pigmentary glaucoma, can now be mapped onto our atomic model, offering mechanistic insights into how structural disruptions affect fibril assembly30. Given that PMEL is overexpressed in melanoma, our findings also provide a framework for investigating whether its fibril structure contributes to tumor progression by altering intracellular signaling or mechanical properties of the tumor microenvironment.
Methods
Tissues and cells
Melanoma patient tissues were obtained from the Fudan University Shanghai Cancer Center with informed consent and approved by institute review board (#050432-4-2108). The patient was a 70-year-old male with a primary melanoma on the left plantar region, and tumor tissues were collected during surgical treatment. Tumor tissues were flash-frozen in liquid nitrogen and stored at −80 °C. The human melanoma cell lines SK-MEL-28 and MNT-1 were purchased from ATCC. SK-MEL-28 cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin (PS). MNT-1 cells were cultured in DMEM medium supplemented with 10% FBS and 1% PS. All cells were maintained at 37 °C in a 5% CO2 atmosphere.
PMEL in SK-MEL-28 was knocked out using the CRISPR/Cas9 system by Haixing Biosciences (China). The sequences of gRNAs were gRNA1: TCACCCGTACCCATCCCAAGAGG, and gRNA2: AGTCTTGCTTGTAGGACCTGGGG according to http://crispor.tefor.net. Plasmid pLV-C-GFPSpark-PMEL (HG13693-ACGLN) was bought from SinoBiological, Inc. (China). All PMEL mutants were constructed using this vector. For transfection, plasmids were mixed with Polyplus jetPRIME reagent (101000046) at a ratio of 1 μg DNA to 2 μL reagent in jetPRIME buffer and incubated at room temperature for 10 min. The resulting mixture was then added to SK-MEL-28 PMEL-KO cells and cultured under standard conditions. After 48 h, the cells were processed for immunofluorescence and TEM analyses.
Immunofluorescence and confocal microscopy
The IF protocol follows this ref. 66. SK-MEL-28 cells were cultured on glass-bottom 3 cm dishes and fixed with 4% paraformaldehyde for 5 min. The cells were then treated with 2 mg/mL glycine in PBS for 5 min, permeabilized with 0.2% Triton X-100 for 5 min, and blocked with 10% goat serum for 15 min. Specific primary antibodies (PMEL-N: ab137078; HMB45: ab787, both from Abcam) were applied, and samples were incubated overnight at 4 °C. After five 5-min washes with PBST (0.05% Tween-20), secondary antibodies were added for 1 h at room temperature, followed by another set of five 5-min PBST washes. Samples were stained with DAPI for 2 min at room temperature, washed with PBST for 5 min, and then imaged using confocal microscopy. The images were acquired by Olympus FV3000.
Transmission electron microscopy
Melanoma patient tissues and SK-MEL-28 cells were fixed with 2.5% glutaraldehyde. Then, 500 μL of 0.1 M phosphate buffer (PB) was added, incubated for 10 min, centrifuged at 1000 × g for 1 min, and the supernatant was discarded. This step was repeated four times. The samples were subsequently fixed with 1% osmium tetroxide for 1.5 h and washed three times with 0.1 M PB using the same procedure. Next, ethanol solutions of 30%, 50%, 70%, and 90% were added sequentially, with each step incubated for 15 min. Ethanol:acetone mixtures (90% ethanol:90% acetone at a 1:1 ratio) and 90% acetone were added sequentially, each incubated for 20 min. Subsequently, 100% acetone was added, incubated for 10 min, centrifuged, and the supernatant discarded. This step was repeated three times. A mixture of 0.3 mL acetone:resin (1:1) was added, incubated for 1 h, centrifuged, and the supernatant discarded. A mixture of 0.3 mL acetone:resin (1:2) was added, incubated for 1 hour, centrifuged, and the supernatant discarded. A mixture of 0.3 mL acetone:resin (1:3) was added and left overnight, followed by centrifugation and removal of the supernatant. The next day, 0.2 mL of pure resin was added and incubated for 3 h. The supernatant was discarded, and 0.2 mL of pure resin was added and incubated for 4 h. The samples were embedded in embedding molds and placed in a 60 °C oven for 48 h. The embedded samples were sectioned into 80 nm-thick slices and transferred onto Formvar-coated copper grids. Finally, the samples were observed using a transmission electron microscope (Hitachi H7800) at 80 kV.
Grid preparation for cryo-ET of SK-MEL-28 cell
Quantifoil Au grids (R2/2, 200 mesh) were glow-discharged for 45 s. Prior to vitrification, SK-MEL-28 cells were incubated with Hoechst ((Invitrogen)) for 10 min for later targeting, followed by PBS wash. The vitrification process was performed using a Vitrobot Mark IV (FEI) under the following conditions: temperature, 15 °C; humidity, 95%; blot force, 8; and blot time, 10 s. Grids were blotted from the back using filter paper (Sigma–Aldrich) and rapidly plunged into a ethane:propane mixture pre-cooled to liquid nitrogen temperature. The vitrified samples were then transferred to cryo-EM storage boxes and stored in liquid nitrogen before FIB milling.
Cryo-focused ion beam milling
Plunge-frozen grids, fixed in custom-made AutoGrid frames, were mounted onto a 35° pre-tilt transfer device and transferred to an Aquilos2 cryo-focused ion beam/scanning electron microscope/integrated Fluorescence Light Microscopy (FIB/SEM/iFLM microscope, Thermo Scientific™). During FIB operation, the samples were maintained at near-liquid nitrogen temperature. To enhance sample conductivity and reduce charging artifacts during milling, the samples were first sputter-coated with platinum, followed by the deposition of an organometallic platinum layer using a gas injection system. The combination of iFLM software and MAPS software enabled precise localization of the target cellular regions for thinning. The milling process was carried out using a 30 kV Ga+ ion beam, with the details as follows: Rough milling was performed using a 1 nA ion beam current, which was gradually reduced to lower currents, with a final fine polishing step at 50 pA. The milling progress was monitored in real-time using SEM (10 kV, 50 pA). To further improve lamella conductivity, an additional platinum sputter coating was applied after milling. Finally, approximately 150–250 nm thick cellular lamellae were obtained.
Cryo-ET data collection
Cryo-FIB lamellae were imaged using a Titan Krios cryo-transmission electron microscope, equipped with a Selectris energy filter (slit width: 20 eV) and a BioContinuum K3 direct detector (Gatan). Low-magnification search maps of lamellas were recorded at ×11,500 magnification, with clearly visible melanosomes structural features serving as reference regions for data acquisition. Multiple tilt series were acquired at ×33,000 magnification (pixel size: 2.66 Å) or ×64,000 magnification (pixel size: 1.37 Å), spanning a tilt range of −50° to 50° in 2° increments, using a dose-symmetric acquisition scheme in Tomography (v5.12). The total electron dose was maintained at approximately 110–140 e−/Ų, with the defocus set to 7 µm.
Tomogram reconstruction and segmentation
For tomographic image processing, raw tilt series images were motion-corrected and CTF-estimated using Warp67. After manually filtering out poor-quality tilt images, the tilt series were automatically aligned using the patch-tracking method, and reconstructed into tomograms via back-projection in the AreTomo package (v2.0)68. To enhance contrast in the regions of interest, CTF deconvolution and missing-wedge correction were performed using IsoNet (v0.2)69. Membrane segmentation was first detected with the Membrain-Seg package70, and analyzed using Amira software (Thermo Scientific™). For the visualization of the lamellar structures formed by PMEL within melanosomes, Amira software (Thermo Scientific™) was used to render different Z-slices. Further 3D representation and segmentation was performed using ChimeraX71.
Extraction of native PMEL fibril lamellae
To obtain intact native PMEL fibril lamellae, extraction procedures were based on published protocols with appropriate modifications to optimize for cryo-EM sample preparation18,38,44. Specifically, 50 mg of melanoma patient tissues were used, while for cultured cells, twelve 10-cm dishes of cells were required. The cells were gently scraped off the culture dishes, centrifuged to collect cell pellets, placed on ice, and immediately processed for extraction. The cell pellets were resuspended in HEE buffer (25 mM HEPES, 1 mM EDTA, 0.1 mM EGTA, proteases inhibitors (PI), 0.25 M sucrose, pH 7.4. PI was bought from Roche) and gently homogenized on ice using a glass pellet pestle. The homogenized cell suspension was centrifuged at 700 g for 10 minutes, and the supernatant (S1) was carefully collected while discarding the pellet. S1 was gently layered onto HEE buffer containing 2 M sucrose and centrifuged again at 11,000 × g for 10 min. The upper phase (light membrane fraction) and intermediate phase (dense membrane fraction) were separately collected and centrifuged at 100,000 × g for 60 min to obtain the pellets (P2 and P3). P2 and P3 were resuspended in 600 μL of buffer TN (100 mM Tris, 150 mM NaCl, PI, pH 7.4) supplemented with 1% Triton X-100 and gently rotated at 4 °C for 2 h. The resulting solutions were centrifuged at 100,000 g for 30 min to obtain pellets P4 and P5. P4 and P5 were resuspended in 600 μL of buffer TN (100 mM Tris, 150 mM NaCl, PI, pH 7.4) supplemented with 1% sarcosyl and gently rotated at 4 °C for 2 h. The resulting solutions were centrifuged at 100,000 × g for 30 min to obtain pellets P6 and P7. P6 and P7 were resuspended in 50 μL of TN buffer and subsequently prepared for cryo-EM analysis (Supplementary Fig. 3a).
Cryo-EM sample preparation and data collection
Sample preparation and data collection were based on this referenced study with further optimizations72. Cryo-EM grids were prepared by depositing 4 μL of extracted native PMEL fibril lamellae onto glow-discharged holey carbon grids (Quantifoil R1.2/1.3, 300 mesh). Excess liquid was removed by blotting the grids for 2 seconds using filter paper in a Vitrobot Mark IV (FEI) operated at 16 °C and 100% humidity. The grids were then rapidly vitrified by plunging into liquid ethane.
Data acquisition was conducted using a BioContinuum K3 direct electron detector (Gatan, Inc.) operating in counting mode, integrated with a Thermo Fisher Titan Krios G4 cryo-transmission electron microscope (Thermo Scientific™) at 300 kV. The microscope was equipped with a GIF Quantum energy filter (Gatan, Inc.) employing a slit width of 20 eV to exclude inelastically scattered electrons. Super-resolution movies were acquired at a magnification of ×105,000, corresponding to a pixel size of 0.83 Å. The total electron dose was approximately 55 e⁻/Å2, with an exposure time of 2.0 s. Automated data collection was managed using EPU software (Thermo Scientific™), with defocus values ranging from −1.0 to −2.4 μm.
Cryo-EM data processing
For image pre-processing, each micrograph consisting of 40 movie frames was subjected to beam-induced motion correction, alignment, dose-weighting, and subsequent binning to a physical pixel size of 0.83 Å using MotionCorr273. The contrast transfer function (CTF) parameters were estimated using CTFFIND-4.1.8 based on motion-corrected images74.
In cryo-EM micrographs, samples derived from the light membrane fraction (P6) and dense membrane fraction (P7) both displayed native PMEL fibril lamellae. Notably, due to the lack of melanin synthesis in SK-MEL-28 cells, the PMEL fibril lamellae appeared more distinct and abundant. In contrast, PMEL fibril lamellae from melanoma patient tissues and MNT-1 cells were less distinct, fewer in number, and predominantly consisted of fibrils heavily coated with melanin, rendering particle picking difficult.
Based on our morphological analysis of native PMEL fibril lamellae, we employed a manual picking method targeting the termini of individual fibrils during the particle picking stage, manually picking all resolvable fibrils within a PMEL fibril lamella in RELION75. Particles were extracted using a 512-pixel box with an inter-box distance of 56 Å, resulting in 1,150,453 particles from the dataset derived from SK-MEL-28 cells (Supplementary Fig. 5a). These particles were imported into cryoSPARC for two rounds of 2D classification, revealing top views, side views, and various top-to-side views of PMEL fibril lamellae76. A subset of 659,107 particles was selected for ab initio construction of an initial 3D model, where one class resembled a 2D side view and aligned with our hypothesis regarding the morphology of native PMEL fibril lamellae (Fig. 1k). Subsequently, heterogeneous refinement of the three initial 3D models further confirmed that the third class matched the hypothesized morphology. This class, comprising 432,105 particles, was selected for homogeneous refinement, yielding a map with a resolution of 6.5 Å. Examination of the map revealed C2 symmetry, with the symmetry center corresponding to the central position of each unit within a fibril. Using the volume alignment tool, the volume was centered within the box to enable proper recognition of C2 symmetry during homogeneous refinement. After that, homogeneous refinement with C2 symmetry on the aligned particles and volume produced a map with a resolution of 6.1 Å (Supplementary Fig. 5a).
Leveraging a deeper understanding of the native PMEL fibril lamella structure, we conducted a second round of manual picking on cryo-EM micrographs, extracting particles with a 512-pixel box and obtaining a total of 2,389,363 particles. These particles were imported into cryoSPARC for 2D classification, yielding 1,611,060 particles. Using the previously obtained 6.1 Å PMEL fibril lamella map as the initial volume, homogeneous refinement with C2 symmetry produced a map with a resolution of 5.51 Å. We then performed 3D classification on this map and its corresponding particles, dividing them into five classes. From the 3D classification results, the most plausible class was selected for homogeneous refinement, yielding a map with a resolution of 4.94 Å. This map revealed three PMEL fibrils, each showing three adjacent units, providing a clear depiction of the overall morphology of native PMEL fibril lamellae (Supplementary Fig. 5a).
To obtain a high-resolution map of a single fibril, we constructed a mask and conducted local refinement. After two rounds of global/local CTF refinement and local refinement, yielding a map with a resolution of 3.48 Å. This map encompassed three adjacent units. The reported resolutions were determined based on the FSC 0.143 criterion. Local resolution was estimated using cryoSPARC, with the highest resolution reaching 2.7 Å (Supplementary Fig. 5b, c).
In addition, we performed cryo-EM data processing and reconstruction for samples derived from melanoma patient tissue and MNT-1 (Supplementary Fig. 6a, b). The main steps were consistent with those used for SK-MEL-28. For tissues and MNT-1, manual picking and extraction yielded 1,239,841 particles and 995,233 particles, respectively, which were imported into cryoSPARC for 2D classification. Side views similar to the native PMEL fibril lamellae of SK-MEL-28 were observed. Moreover, this side-view class contained the largest number of particles among all classes, confirming that the morphology of native PMEL fibril lamellae in tissues and MNT-1 is consistent with that in SK-MEL-28. A total of 45,560 particles from the tissue sample and 60,845 particles from MNT-1 were selected for ab initio initial model construction. Both tissue and MNT-1 samples successfully reconstructed the same morphology as the native PMEL fibril lamellae observed in SK-MEL-28. Subsequently, heterogeneous refinement was performed, yielding 25,416 particles for the tissue sample and 45,448 particles for MNT-1. Homogeneous refinement was then performed for each sample. The volume alignment tool was used to center the symmetry axis within the box, followed by homogeneous refinement with C2 symmetry. The reconstructed map of the native PMEL fibril lamellae reached a resolution of 7.41 Å for the tissue sample and 7.2 Å for MNT-1. The final 3D morphologies of both samples confirmed their consistency with the native PMEL fibril lamellae observed in SK-MEL-28.
Model building and refinement
The initial structural model of a single PMEL molecule (horizontal subunit and vertical subunit) was obtained using ModelAngelo77. Subsequently, the structure was manually adjusted and rebuilt using COOT78. During the model building process for the single PMEL molecule, the four chains (Chain α, β, γ, and δ) were identified, along with three regions of unresolved density: IDR1, IDR2, and IDR3. Notably, IDR1, located between Chain α and Chain β, corresponds to the region connecting adjacent PMEL fibrils. Additionally, a distinct glycosylation density was observed at position N81, and the glycan model was built accordingly. The final model was refined against the corresponding maps in real space using PHENIX, with secondary structure and geometry restraints applied79. Final model validation was performed using the “Comprehensive Validation” feature in PHENIX79. Visualization of density maps and atomic models was achieved using UCSF Chimera X71.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM map of native PMEL fibril lamella and the corresponding atomic coordinate have been deposited in the Electron Microscopy Data Bank [https://www.ebi.ac.uk/pdbe/emdb/] and the Protein Data Bank [http://www.rcsb.org], under the accession codes EMD-63119 (native PMEL fibril lamella) [https://www.ebi.ac.uk/emdb/EMD-63119] and 9LIP, respectively. All materials are available from the corresponding authors upon request.
References
Lin, J. Y. & Fisher, D. E. Melanocyte biology and skin pigmentation. Nature 445, 843–850 (2007).
Benito-Martinez, S., Salavessa, L., Raposo, G., Marks, M. S. & Delevoye, C. Melanin transfer and fate within keratinocytes in human skin pigmentation. Integr. Comp. Biol. 61, 1546–1555 (2021).
Le, L., Sires-Campos, J., Raposo, G., Delevoye, C. & Marks, M. S. Melanosome biogenesis in the pigmentation of mammalian skin. Integr. Comp. Biol. 61, 1517–1545 (2021).
Brenner, M. & Hearing, V. J. The protective role of melanin against UV damage in human skin. Photochem Photobio. 84, 539–549 (2008).
Raposo, G. & Marks, M. S. Melanosomes–dark organelles enlighten endosomal membrane transport. Nat. Rev. Mol. Cell Biol. 8, 786–797 (2007).
Delevoye, C., Marks, M. S. & Raposo, G. Lysosome-related organelles as functional adaptations of the endolysosomal system. Curr. Opin. Cell Biol. 59, 147–158 (2019).
Ma, L. et al. Discovery of the migrasome, an organelle mediating release of cytoplasmic contents during cell migration. Cell Res. 25, 24–38 (2015).
Lyon, A. S., Peeples, W. B. & Rosen, M. K. A framework for understanding the functions of biomolecular condensates across scales. Nat. Rev. Mol. Cell Biol. 22, 215–235 (2021).
Tabara, L. C., Segawa, M. & Prudent, J. Molecular mechanisms of mitochondrial dynamics. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-024-00785-1 (2024).
Zhao, Y. G., Codogno, P. & Zhang, H. Machinery, regulation and pathophysiological implications of autophagosome maturation. Nat. Rev. Mol. Cell Biol. 22, 733–750 (2021).
Rousseau, A. & Bertolotti, A. Regulation of proteasome assembly and activity in health and disease. Nat. Rev. Mol. Cell Biol. 19, 697–712 (2018).
Seiji, M., Fitzpatrick, T. B., Simpson, R. T. & Birbeck, M. S. C. Chemical composition and terminology of specialized organelles (melanosomes and melanin granules) in mammalian melanocytes. Nature 197, 1082–1084 (1963).
Zhou, B. K. et al. Identification of a melanosomal matrix protein encoded by the murine si (silver) locus using” organelle scanning. Proc. Natl Acad. Sci. USA 91, 7076–7080 (1994).
Kobayashi, T. et al. The Pmel 17/silver locus protein. Characterization and investigation of its melanogenic function. J. Biol. Chem. 269, 29198–29205 (1994).
Adema, G. J., de Boer, A. J., Vogel, A. M., Loenen, W. A. & Figdor, C. G. Molecular characterization of the melanocyte lineage-specific antigen gp100. J. Biol. Chem. 269, 20126–20133 (1994).
Hurbain, I. et al. Electron tomography of early melanosomes: Implications for melanogenesis and the generation of fibrillar amyloid sheets. Proc. Natl Acad. Sci. USA 105, 19726–19731 (2008).
Graham, M., Tzika, A. C., Mitchell, S. M., Liu, X. & Leonhardt, R. M. Repeat domain-associated O-glycans govern PMEL fibrillar sheet architecture. Sci. Rep. 9, 6101 (2019).
Kushimoto, T. et al. A model for melanosome biogenesis based on the purification and analysis of early melanosomes. Proc. Natl Acad. Sci. USA 98, 10698–10703 (2001).
D’Alba, L. & Shawkey, M. D. Melanosomes: biogenesis, properties, and evolution of an ancient organelle. Physiol. Rev. 99, 1–19 (2019).
Tzika, A. C., Ullate-Agote, A., Helleboid, P. Y. & Kummrow, M. PMEL is involved in snake colour pattern transition from blotches to stripes. Nat. Commun. 15, 7655 (2024).
Wang, J. et al. Genome-Wide Association Analysis Identifies the PMEL Gene Affecting Coat Color and Birth Weight in Simmental x Holstein. Animals 13, https://doi.org/10.3390/ani13243821 (2023).
Liu, F., Sun, F., Kuang, G. Q., Wang, L. & Yue, G. H. The insertion in the 3’ UTR of Pmel17 is the causal variant for golden skin color in Tilapia. Mar. Biotechnol. 24, 566–573 (2022).
Kimura, S. et al. PMEL p.Leu18del dilutes coat color of Kumamoto sub-breed of Japanese Brown cattle. BMC Genom. 23, 694 (2022).
Chrystal, P. W. et al. Functional domains and evolutionary history of the PMEL and GPNMB family proteins. Molecules, 26, https://doi.org/10.3390/molecules26123529 (2021).
Hellstrom, A. R. et al. Inactivation of Pmel alters melanosome shape but has only a subtle effect on visible pigmentation. PLoS Genet. 7, e1002285 (2011).
Schonthaler, H. B. et al. A mutation in the silver gene leads to defects in melanosome biogenesis and alterations in the visual system in the zebrafish mutant fading vision. Dev. Biol. 284, 421–436 (2005).
Brunberg, E. et al. A missense mutation in PMEL17 is associated with the Silver coat color in the horse. BMC Genet. 7, 46 (2006).
Kerje, S. et al. The Dominant white, Dun and Smoky color variants in chicken are associated with insertion/deletion polymorphisms in the PMEL17 gene. Genetics 168, 1507–1518 (2004).
AlAbdi, L., Alshammari, M., Helaby, R., Khan, A. O. & Alkuraya, F. S. PMEL is mutated in oculocutaneous albinism. Hum. Genet. 142, 139–144 (2023).
Lahola-Chomiak, A. A. et al. Non-synonymous variants in premelanosome protein (PMEL) cause ocular pigment dispersion and pigmentary glaucoma. Hum. Mol. Genet. 28, 1298–1311 (2019).
Matafora, V. et al. Amyloid aggregates accumulate in melanoma metastasis modulating YAP activity. EMBO Rep. 21, e50446 (2020).
Farris, F. et al. Unveiling the mechanistic link between extracellular amyloid fibrils, mechano-signaling and YAP activation in cancer. Cell Death Dis. 15, 28 (2024).
Zhang, S. et al. PMEL as a prognostic biomarker and negatively associated with immune infiltration in skin cutaneous melanoma (SKCM). J. Immunother. 44, 214–223 (2021).
Sawaya, M. R., Hughes, M. P., Rodriguez, J. A., Riek, R. & Eisenberg, D. S. The expanding amyloid family: structure, stability, function, and pathogenesis. Cell 184, 4857–4873 (2021).
Maji, S. K. et al. Functional amyloids as natural storage of peptide hormones in pituitary secretory granules. Science 325, 328–332 (2009).
Otzen, D. & Riek, R. Functional amyloids. Cold Spring Harb. Perspect. Biol. 11, a033860(2019).
Fowler, D. M. et al. Functional amyloid formation within mammalian tissue. PLoS Biol. 4, e6 (2006).
Berson, J. F. et al. Proprotein convertase cleavage liberates a fibrillogenic fragment of a resident glycoprotein to initiate melanosome biogenesis. J. Cell Biol. 161, 521–533 (2003).
Watt, B. et al. N-terminal domains elicit formation of functional Pmel17 amyloid fibrils. J. Biol. Chem. 284, 35543–35555 (2009).
McGlinchey, R. P. et al. The repeat domain of the melanosome fibril protein Pmel17 forms the amyloid core promoting melanin synthesis. Proc. Natl Acad. Sci. USA 106, 13731–13736 (2009).
Watt, B., van Niel, G., Raposo, G. & Marks, M. S. PMEL: a pigment cell-specific model for functional amyloid formation. Pigment Cell Melanoma Res. 26, 300–315 (2013).
Hee, J. S., Mitchell, S. M., Liu, X. & Leonhardt, R. M. Melanosomal formation of PMEL core amyloid is driven by aromatic residues. Sci. Rep. 7, 44064 (2017).
Mitchell, S. M., Graham, M., Liu, X. & Leonhardt, R. M. Identification of critical amino acid residues in the regulatory N-terminal domain of PMEL. Sci. Rep. 11, 7730 (2021).
Watabe, H., Kushimoto, T., Valencia, J. C. & Hearing, V. J. Isolation of melanosomes. Curr. Protoc. Cell Biol. Chapter 3, Unit 3 14 (2005).
Yasumoto, K. et al. Epitope mapping of the melanosomal matrix protein gp100 (PMEL17): rapid processing in the endoplasmic reticulum and glycosylation in the early Golgi compartment. J. Biol. Chem. 279, 28330–28338 (2004).
van Niel, G. et al. Apolipoprotein E regulates amyloid formation within endosomes of pigment cells. Cell Rep. 13, 43–51 (2015).
Leonhardt, R. M., Vigneron, N., Hee, J. S., Graham, M. & Cresswell, P. Critical residues in the PMEL/Pmel17 N-terminus direct the hierarchical assembly of melanosomal fibrils. Mol. Biol. Cell 24, 964–981 (2013).
Bissig, C., Rochin, L. & van Niel, G. PMEL amyloid fibril formation: the bright steps of pigmentation. Int. J. Mol. Sci. 17, 1438 (2016).
Berson, J. F., Harper, D. C., Tenza, D., Raposo, G. & Marks, M. S. Pmel17 initiates premelanosome morphogenesis within multivesicular bodies. Mol. Biol. Cell 12, 3451–3464 (2001).
Maresh, G. A. et al. Differential processing and secretion of the melanoma-associated ME20 antigen. Arch. Biochem. Biophys. 311, 95–102 (1994).
Mompean, M. et al. The structure of the necrosome RIPK1-RIPK3 core, a human hetero-amyloid signaling complex. Cell 173, 1244–1253.e1210 (2018).
Yanagisawa, H. et al. Cryo-EM of wild-type and mutant PMEL amyloid cores reveals structural mechanism of pigment dispersion syndrome. Nat. Commun. 16, 5411 (2025).
Sergeeva, A. V. & Galkin, A. P. Functional amyloids of eukaryotes: criteria, classification, and biological significance. Curr. Genet. 66, 849–866 (2020).
Greenwald, J. & Riek, R. Biology of amyloid: structure, function, and regulation. Structure 18, 1244–1260 (2010).
Dean, D. N. & Lee, J. C. Defining an amyloid link between Parkinson’s disease and melanoma. Proc. Natl Acad. Sci. USA 117, 22671–22673 (2020).
Dean, D. N. & Lee, J. C. Modulating functional amyloid formation via alternative splicing of the premelanosomal protein PMEL17. J. Biol. Chem. 295, 7544–7553 (2020).
McGlinchey, R. P. & Lee, J. C. Why study functional amyloids? Lessons from the repeat domain of Pmel17. J. Mol. Biol. 430, 3696–3706 (2018).
Harper, D. C. et al. Premelanosome amyloid-like fibrils are composed of only golgi-processed forms of Pmel17 that have been proteolytically processed in endosomes. J. Biol. Chem. 283, 2307–2322 (2008).
Hoashi, T. et al. The repeat domain of the melanosomal matrix protein PMEL17/GP100 is required for the formation of organellar fibers. J. Biol. Chem. 281, 21198–21208 (2006).
McGlinchey, R. P. et al. Repeat domains of melanosome matrix protein Pmel17 orthologs form amyloid fibrils at the acidic melanosomal pH. J. Biol. Chem. 286, 8385–8393 (2011).
Jiang, Z. & Lee, J. C. Lysophospholipid-containing membranes modulate the fibril formation of the repeat domain of a human functional amyloid, pmel17. J. Mol. Biol. 426, 4074–4086 (2014).
Pfefferkorn, C. M., McGlinchey, R. P. & Lee, J. C. Effects of pH on aggregation kinetics of the repeat domain of a functional amyloid, Pmel17. Proc. Natl Acad. Sci. USA 107, 21447–21452 (2010).
Dean, D. N. & Lee, J. C. Purification and characterization of an amyloidogenic repeat domain from the functional amyloid Pmel17. Protein Expr. Purif. 187, 105944 (2021).
Dean, D. N. & Lee, J. C. pH-dependent fibril maturation of a Pmel17 repeat domain isoform revealed by tryptophan fluorescence. Biochim. Biophys. Acta Proteins Proteom. 1867, 961–969 (2019).
Rochin, L. et al. BACE2 processes PMEL to form the melanosome amyloid matrix in pigment cells. Proc. Natl Acad. Sci. USA 110, 10658–10663 (2013).
Ma, B. et al. Albumosomes formed by cytoplasmic pre-folding albumin maintain mitochondrial homeostasis and inhibit nonalcoholic fatty liver disease. Signal Transduct. Target Ther. 8, 229 (2023).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Zheng, S. et al. AreTomo: An integrated software package for automated marker-free, motion-corrected cryo-electron tomographic alignment and reconstruction. J. Struct. Biol. X 6, 100068 (2022).
Liu, Y. T. et al. Isotropic reconstruction for electron tomography with deep learning. Nat. Commun. 13, 6482 (2022).
Lamm, L. et al. MemBrain v2: an end-to-end tool for the analysis of membranes in cryo-electron tomography. https://doi.org/10.1101/2024.01.05.574336 (2024).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Liu, K. et al. Binding adaptability of chemical ligands to polymorphic alpha-synuclein amyloid fibrils. Proc. Natl Acad. Sci. USA 121, e2321633121 (2024).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature 628, 450–457 (2024).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank the Cryo-EM Center at the Interdisciplinary Research Center on Biology and Chemistry, Shanghai Institute of Organic Chemistry for help with cryo-EM data collection, and the Instrumental Analysis Center of Shanghai Jiao Tong University and the Cell Biology Facility of the Center of Biomedical Analysis of Tsinghua University for help with TEM analysis. We thank Dr. Guoqiang Huang (Tsinghua University) for help with cryo-EM data processing and analysis. This work was supported by the National Natural Science Foundation (NSF) of China (32494764, 92353302, and 32170683 to D.L.; 22425704, 82188101 and 32171236 to C.L.), Shanghai Basic Research Pioneer Project to C.L., Shanghai Municipal Science and Technology Major Project to C.L., the Shanghai Pilot Program for Basic Research - Chinese Academy of Science, Shanghai Branch (Grant No. JCYJ-SHFY-2022-005 to C.L.), the CAS Project for Young Scientists in Basic Research (Grant No.YSBR-095 to C.L.), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant No. XDB1060000 to C.L.), the Shanghai Key Laboratory of Aging Studies (Grant No. 19DZ2260400 to C.L.), the Postdoctoral Fellowship Program of CPSF (Grant No. GZB20230801 to B.M.), the China Postdoctoral Science Foundation (Grant No. 2024M753382 to B.M.), and the Shanghai Postdoctoral Excellence Program (Grant No. 2023725 to B.M.). Dr. Cong Liu is a SANS Exploration Scholar.
Author information
Authors and Affiliations
Contributions
Conceptualization: B.M., D.L., and C.L.; Methodology: B.M., Y.Y., H.D., L.Y., D.N.L., and Q.Z.; Investigation: B.M., Y.Y., and C.L.; Writing and editing and reviewing: B.M., Y.Y., H.D., L.Y., D.N.L., Q.Z., B.S., Y.C., D.L., and C.L.; Supervision: D.L. and C.L. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ralf Leonhardt, Hannan Xu, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, B., Yao, Y., Dong, H. et al. Atomic structure and in situ visualization of native PMEL lamellae in melanosomes. Nat Commun 16, 10300 (2025). https://doi.org/10.1038/s41467-025-65221-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65221-0
This article is cited by
-
Microbial melanin: biosynthesis, functional Insights, and emerging research challenges
World Journal of Microbiology and Biotechnology (2026)