Abstract
Plasmids carrying the mobilized colistin-resistance gene mcr-1 are prevalent among multidrug-resistant Gram-negative pathogens, yet their broad impact on bacterial physiology and virulence remains unclear. Here, we demonstrate that acquisition of an mcr-1 plasmid concurrently increases antimicrobial resistance and pathogenicity in Escherichia coli. On the same plasmid, the XRE-family transcriptional regulator EcaR cooperates with MCR-1 to activate the wec operon, driving biosynthesis of two surface polysaccharides: enterobacterial common antigen (ECA) and a high-molecular-weight O-chain. Expression of these surface polysaccharides increases bile resistance and virulence in a murine model and further elevates colistin resistance. MCR-1 enhances transcription of upstream genes in the wec operon, whereas EcaR directly activates an internal promoter (PwecE) to induce downstream gene expression. Thus, both components are required for surface polysaccharide expression, and deletion of either abolishes the phenotype. Genomic analysis of publicly available mcr plasmids reveals widespread co-occurrence of mcr-1 and ecaR on IncI2 and IncX4 plasmids, indicating their functional complementarity. These findings uncover a mechanism by which resistance plasmids remodel the bacterial surface, linking horizontal gene transfer to coordinated regulation of antimicrobial resistance and virulence.
Similar content being viewed by others
Introduction
The rapid emergence and global spread of multidrug-resistant (MDR) Gram-negative bacteria have compromised the effectiveness of current antimicrobial therapies1,2. Among the various resistance mechanisms, horizontal acquisition of mobile genetic elements that encode antimicrobial resistance genes is particularly alarming because these elements disseminate rapidly across bacterial populations3,4. Of particular concern is the discovery of plasmid-mediated colistin resistance, which was first reported with the description of the mobilized colistin resistance-1 (mcr-1) genes5. This finding has prompted significant concern within the medical and microbiological research communities, because colistin is frequently reserved as a last-line agent against MDR Gram-negative bacteria, particularly those resistant to β-lactams, carbapenems, quinolones, and aminoglycosides6.
Colistin is a cationic cyclic lipopeptide that binds the phosphate groups of lipid A, displaces divalent cations (Mg2+ and Ca2+), destabilizes the outer membrane, and ultimately causes cell lysis7,8. The mcr-1 gene encodes a phosphoethanolamine transferase that catalyzes the addition of phosphoethanolamine to the lipid A moiety of lipopolysaccharide (LPS)5,9, thereby diminishing the electrostatic affinity of colistin10. Since its discovery, mcr-1 has been reported worldwide in pathogenic members of Enterobacterales, including Escherichia coli11,12,13,14, Klebsiella pneumoniae11,14, and Salmonella enterica15,16. Furthermore, mcr-1 has also been identified in non-enterobacterial genera, such as Moraxella17 and Alcaligenes18, underscoring its capacity to cross taxonomic boundaries. Notably, the plasmid-borne nature of mcr-1 facilitates its rapid spread through horizontal gene transfer, posing a substantial threat to the efficacy of colistin as a last-resort antibiotic19.
Plasmids carrying antibiotic resistance genes frequently encode accessory determinants that modulate bacterial physiology20,21 and virulence22 in the host environment. For mcr-1 plasmids, such loci include genes for type IV secretion systems23, pili23, conjugation machinery24 and plasmid-copy-number regulation25. These elements are classified into incompatibility (Inc) groups IncI2, IncX4, and IncHI2, which differ in size, transfer efficiency, and genetic content. IncI2 and IncX4 plasmids are typically small and often carry only mcr-1, whereas IncHI2 plasmids frequently co-harbor multiple resistance genes and heavy metal resistance operons19,26. These differences may affect the persistence and dissemination of each plasmid type across bacterial hosts and environments. Although the mechanism of MCR-1-mediated colistin resistance is well understood9, it is important to note that the acquisition of mcr-1 plasmids can also result in physiological changes in bacteria beyond drug resistance. Appreciating these broader impacts is crucial for evaluating the public health risk posed by the dissemination of mcr-1 plasmids and for developing effective strategies to control resistant infections caused by mcr-1-positive bacteria.
The Gram-negative cell envelope, comprising the inner membrane, periplasm, and outer membrane, plays a critical role in antimicrobial resistance, virulence, and bacterial physiology27,28,29. Surface polysaccharides constitute major outer membrane components that contribute to bacterial survival in hostile environments30 and interactions with host immune systems31. Among these, the lipopolysaccharide (LPS) O-chain (O-polysaccharide), covalently linked to the LPS core, forms the outermost surface layer32. Its compositional and chain length variation influences complement evasion, host interactions, and alters antibiotic susceptibility by modulating outer membrane permeability and surface charge33,34. In addition, members of Enterobacterales produce a distinctive surface polysaccharide known as enterobacterial common antigen (ECA), which plays multifaceted roles in bacterial physiology, virulence, and host-pathogen interactions35. ECA exists in three distinct structural forms: LPS-linked ECA (ECALPS), phosphoglyceride-linked ECA (ECAPG), and cyclic ECA (ECACYC)36,37,38. Despite their structural differences, all ECA variants share a highly conserved trisaccharide repeating unit composed of N-acetyl-D-glucosamine (GlcNAc), N-acetyl-D-mannosaminuronic acid (ManNAcA), and 4-acetamido-4,6-dideoxy-D-galactose (Fuc4NAc), which is ubiquitous throughout Enterobacterales35. Among these, ECALPS is immunogenic36,39 and has been implicated in bacterial pathogenesis. Nevertheless, the molecular mechanisms underlying ECA biosynthesis and its potential role in antimicrobial resistance remain largely unexplored.
Given that MCR-1 modifies lipid A, we hypothesized that mcr-1 may influence additional aspects of envelope biology, either through direct effects or by influencing regulatory pathways. Here, we delineate how the acquisition of mcr-1-carrying plasmids affects surface polysaccharide production, thereby altering bacterial physiology, antimicrobial resistance, and virulence. Using E. coli as a model, we revealed that horizontal acquisition of mcr-1-carrying plasmids remodels surface polysaccharide production, prompting us to investigate its broader consequences for bacterial physiology and pathogenic traits.
Results
Acquisition of an mcr-1 plasmid triggers surface polysaccharide biosynthesis
Plasmids harboring mcr-1 transfer readily by conjugation among members of the order Enterobacterales19 and persist for many generations once acquired40. For this study, we used E. coli strain 15734 (GenBank accession number: CP097001), a clinical isolate obtained from human stool in South Korea and classified as enterotoxigenic E. coli (ETEC) carrying the heat-stable enterotoxin gene, as the wild-type (WT) strain. We likewise confirmed the stability of the mcr-1 plasmid (pMCR-1; pFORC82_3) in E. coli (Supplementary Fig. 1). Given that plasmid acquisition often triggers complex physiological adaptations in bacterial hosts, we examined the broader cellular consequences of mcr-1 plasmid acquisition. The acquisition of the mcr-1 plasmid altered bacterial physiology, reducing biofilm formation and enhancing swimming motility, while also modifying surface properties, such as hydrophobicity and zeta potential (Supplementary Fig. 2). These phenotypic changes suggested that the acquisition of the mcr-1 plasmid extends beyond simple antibiotic resistance to encompass fundamental alterations in bacterial surface architecture. The plasmid also increased bacterial adhesion to Caco-2 cells and virulence in the Galleria mellonella infection model (Supplementary Fig. 3), indicating that resistance plasmids may simultaneously enhance bacterial pathogenic potential.
To understand the molecular basis underlying these phenotypic changes, we performed RNA sequencing (RNA-seq) on WT, the transconjugant (TC; WT+ pmcr-1), and a Δmcr-1 mutant (TCΔmcr-1). The results revealed extensive transcriptional reprogramming upon acquisition of the mcr-1 plasmid, with 493 genes (234 up-regulated and 259 down-regulated) showing altered expression (Fig. 1a–c and Supplementary Data 1). The differentially expressed genes were functionally associated with energy production and conversion, amino acid transport and metabolism, transcription, and inorganic acid transport and metabolism (Fig. 1c). Consistent with the role of MCR-1 in lipid A modification9, genes involved in cell envelope biogenesis were also affected (Fig. 1c).
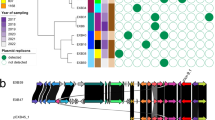
a Venn diagram showing differentially expressed genes (DEGs; fold change ≥2 or ≤−2, P < 0.05) comparing wild type (WT) vs. transconjugant (TC) and TCΔmcr-1 vs. TC based on RNA-seq analysis. The overlapping region represents mcr-1-regulated genes. b Number of mcr-1-regulated DEGs in WT (fold change ≥2 or ≤−2, P < 0.05). Red and blue indicate upregulated and downregulated genes, respectively. Gray indicates unchanged genes. c Clusters of Orthologous Groups (COG) classification of mcr-1-regulated DEGs in WT. Red and blue bars indicate upregulated and downregulated genes, respectively. d Heatmap of gene expression related to enterobacterial common antigen (ECA) synthesis in WT, TC, and TCΔmcr-1 mutant. Each column represents one biological replicate. Red indicates increased expression, and blue indicates decreased expression. e Surface polysaccharide profiles of WT (lane 1), TC (lane 2), TCΔmcr-1 mutant (lane 3), mcr-1-complemented strain (lane 4), and WT carrying the mcr-1 gene alone (lane 5). “+” indicates reintroduction of the mcr-1 gene via pACYC184. f Quantification of glycan levels by wheat germ agglutinin (WGA) staining, expressed relative to WT. Data are presented as means ± SD from n = 4 biologically independent replicates. P values were determined using one-way ANOVA with Tukey’s multiple comparison test. WT vs. TC, P < 0.0001; TC vs. TCΔmcr-1 mutant, P < 0.0001; TCΔmcr-1 mutant vs. TCΔmcr-1 mutant+pmcr-1, P < 0.0001; WT vs. WT+pmcr-1, P = 0.9415. ns not significant, ****P < 0.0001. g Transmission electron microscopy (TEM) analysis showing surface polysaccharide layer at the cell surface. Cells were stained with 1.5% phosphotungstic acid for 2 min. The experiments were repeated three times with similar results, and representative data from one of the experiments are shown (e, g). All transcriptomic data presented in a–d were obtained from the same RNA-seq dataset. The strains used were E. coli 15734 (WT), WT carrying pFORC82_3 (TC), TC harboring mcr-1-deleted pFORC82_3 (TCΔmcr-1), TC harboring mcr-1-deleted pFORC82_3 and pACYC184-mcr-1 (TCΔmcr-1 + pmcr-1), and WT harboring pACYC184-mcr-1 (WT + pmcr-1). Source data are provided as a Source Data file.
Most remarkably, we observed dramatic upregulation of the wec operon, which is responsible for ECA synthesis (Fig. 1d and Supplementary Fig. 4). This finding was particularly intriguing because ECA is typically constitutively expressed in Enterobacterales, and its regulation by horizontally acquired elements has not been previously reported. Deletion of mcr-1 in the TC strain restored wec operon transcription to WT levels, confirming that mcr-1 is necessary for this transcriptional activation (Fig. 1d). The clinical WT strain in our study exhibits a rough phenotype due to the absence of most of the genes located between galF and gnd in the O-antigen locus (Supplementary Fig. 5). Notably, despite lacking conventional O-antigen biosynthetic machinery, acquisition of the mcr-1 plasmid enabled this rough strain to produce surface polysaccharides (Fig. 1e, lane 2). This transformation from rough to smooth phenotype represents a fundamental change in bacterial surface architecture.
To confirm that the observed polysaccharide production was indeed ECA-related, we deleted wecG, which encodes the UDP-N-acetyl-D-mannosaminuronic acid transferase essential for ECA assembly but dispensable for LPS core or O-antigen biosynthesis41. Notably, wecG deletion eliminated the glycan band, and reduced the signal of the lectin protein that detects glycans, such as β-linked GlcNAc, by wheat‑germ agglutinin (WGA) staining (Supplementary Fig. 6a, b), demonstrating that mcr-1 specifically activates ECA-related polysaccharide synthesis.
The requirement for mcr-1 in transcriptional activation was further validated through complementation studies (Fig. 1e and Supplementary Fig. 7). A TC harboring a mcr-1-deficient plasmid remained rough (Fig. 1e, lane 3), whereas re-introduction of mcr-1 restored surface polysaccharide synthesis (Fig. 1e, lane 4). WGA staining corroborated elevated glycan levels in the presence of mcr-1 (Fig. 1f), indicating that mcr-1 plasmid acquisition induced ECA production. Transmission-electron microscopy (TEM) after phosphotungstic-acid negative staining provided visual confirmation of these biochemical changes, revealing a markedly denser surface layer on the TC than on WT, whereas deletion of mcr-1 reverted to a WT-like appearance (Fig. 1g). Similarly, wecG deletion in the TC abolished the dense layer, further confirming that the observed structure corresponds to polysaccharides including ECA (Supplementary Fig. 6c).
To test whether MCR-1 alone is sufficient to drive surface polysaccharide synthesis, we introduced mcr-1 into the WT. However, whereas conjugational transfer of an mcr-1 plasmid activated surface polysaccharide production (Fig. 1e, lane 2), expression of mcr-1 alone in the WT was insufficient to induce this phenotype (Fig. 1e, lane 5), indicating that additional plasmid-encoded factor(s) are required for full activation. This observation suggested the existence of a more complex regulatory network involving multiple plasmid-encoded elements. Collectively, our findings reveal that the horizontal acquisition of an mcr-1 plasmid not only confers colistin resistance but also triggers extensive transcriptional reprogramming that activates surface polysaccharide synthesis and remodels bacterial surface properties.
Plasmid-encoded EcaR cooperates with MCR-1 to activate ECA operon transcription
Since mcr-1 alone was insufficient to activate surface polysaccharide synthesis, we next sought to identify additional regulatory factors on the plasmid. We screened mcr-1 plasmid-encoded genes predicted to contain DNA-binding domains and transcriptional regulatory functions (Supplementary Data 2). This systematic approach was based on the hypothesis that transcriptional activation of the wec operon would require a dedicated transcriptional regulator, as MCR-1 itself lacks DNA-binding capability.
Our screening strategy proved successful, revealing that the transformation of a 252 bp DNA fragment of the mcr-1 plasmid restored the synthesis of surface polysaccharides in E. coli carrying mcr-1 (Fig. 2a and Supplementary Fig. 8). Sequence analysis of this critical fragment identified an open reading frame, pFORC82_562, which encodes a putative XRE family transcriptional regulator. Given its essential role in ECA synthesis, we designated this gene ecaR (enterobacterial common antigen synthesis regulator; Supplementary Fig. 9). The discovery of ecaR represents the identification of a plasmid-encoded regulator specifically dedicated to ECA biosynthesis, expanding our understanding of ECA regulation beyond chromosomal control mechanisms.
a Surface polysaccharide profiles of wild type (WT; lane 1), WT harboring pmcr-1 or pecaR (lane 2 or 3), WT co-expressing mcr-1 and ecaR (lane 4), and transconjugant (TC; lane 5). “+” indicates reintroduction of each gene via plasmid. b Partial 1H and 1H,13C-HSQC spectrum of the high-molecular-weight polysaccharide released from WT + pmcr-1 + pecaR. The proposed disaccharide repeat is shown in the inset. c Relative mRNA expression of ECA operon genes in WT + pmcr-1 and WT + pecaR compared to WT. d Relative mRNA expression of ECA operon genes in WT + pmcr-1 + pecaR and TC compared to WT. e DNase I footprinting analysis of EcaR binding to PwecA and PwecE in the ECA operon. Red/Blue boxes mark promoter region. Blue peaks represent EcaR-protected regions; red peaks show DNase I digestion in the absence of EcaR. f β-galactosidase activity from transcriptional fusions of PwecA and PwecE with lacZ in the presence or absence of EcaR. g Schematic representation of ECA operon regulation showing genes upregulated by mcr-1 (red) and ecaR (blue). Created in BioRender. Ahn, E. (2025) https://BioRender.com/h97y857. h Surface polysaccharide profiles of WT (lane 1), WT harboring pwecE (lane 2), WT harboring both pwecE and pmcr-1 (lane 3), and WT + pmcr-1 + pecaR (lane 4). i Quantification of glycan levels in WT + pwecE + pmcr-1 by arabinose-induced wecE expression. The experiments were repeated three times with similar results, and representative data from one of the experiments are shown (a, h). Data are presented as means ± SD from n = 3 biologically independent replicates (c, d, f, i). P values were determined using two-way ANOVA with Tukey’s multiple comparison test (c, d), two-tailed Student’s t-test (f), and one-way ANOVA with Tukey’s multiple comparison test (i). Exact P values are provided in the Source Data file. ns, not significant; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. The strains used were E. coli 15734 (WT), WT carrying pFORC82_3 (Transconjugant, TC), WT harboring pACYC184-mcr-1 (WT + pmcr-1), WT harboring pBAD33-ecaR (WT + pecaR), and WT harboring both pACYC184-mcr−1 and pBAD33-ecaR (WT+pmcr-1+pecaR). Source data are provided as a Source Data file.
Functional analysis revealed a strict requirement for both regulatory elements. Neither mcr-1 nor ecaR alone was sufficient to induce surface polysaccharide synthesis (Fig. 2a, lanes 2 and 3), whereas co-expression of both genes (Fig. 2a, lane 4) or their co-localization on the same plasmid (Fig. 2a, lane 5) resulted in robust production of surface polysaccharides. This cooperative requirement indicates that MCR-1 and EcaR function as a coordinated regulatory system, with each component contributing distinct but essential roles to surface polysaccharide activation.
To definitively confirm ECA production and distinguish it from other polysaccharides, we performed Western blot analysis using an ECA-specific monoclonal antibody (mAb 898)42,43, with E. coli MG1655 as a positive control44. The results provided unambiguous evidence for ECA synthesis. The WT co-expressing mcr-1 and ecaR, and TC strains exhibited a strong immunoreactive signal comparable to MG1655, whereas the WT showed no detectable signal (Supplementary Fig. 10).
In addition to ECA, our analysis revealed the production of additional polysaccharide species. Glycosyl compositional analysis of surface polysaccharides from WT co-expressing mcr-1 and ecaR, and TC revealed a marked increase in ribose and galactose in a 1:1 molar ratio relative to WT (Supplementary Table 1). To confirm that the detected ribose originated from carbohydrate polymers rather than nucleic acid contamination, repeated nuclease treatments were performed, which failed to decrease the ribose signal in these preparations, confirming that the detected ribose represents part of a carbohydrate polymer structure. Structural characterization using 1D/2D NMR spectroscopy identified a repeating disaccharide unit [→2)-β-D-Ribf-(1 → 4)-α-D-Galp-(1→]n in the high molecular weight (HMW) region of polysaccharides released from lipid A by mild acid hydrolysis (Fig. 2b and Supplementary Fig. 11; see Supplementary Note 1 for more details). For clarity, throughout this manuscript, we use “surface polysaccharides” to refer specifically to ECA and the HMW O-chain that emerges upon mcr-1/ecaR-mediated activation. Collectively, these results demonstrate that acquisition of an mcr-1/ecaR-harboring plasmid promotes surface polysaccharide synthesis, including ECA and HMW O-chain polysaccharides, effectively converting rough strains into functionally smooth variants.
To understand the molecular mechanism underlying this dual regulation, we examined the transcriptional effects of each regulator. MCR-1 upregulated the transcription of genes upstream of wecE in the ECA operon, while EcaR upregulated genes downstream of wecD (Fig. 2c). Co-expression of mcr-1 and ecaR was necessary to induce complete transcriptional activation of the operon (Fig. 2d), confirming that both regulatory inputs are required for full wec operon activation. To further validate these findings, we also generated ecaR deletion and complementation strains in the TC (TCΔecaR and complemented TCΔecaR). Deletion of ecaR reduced ECA-operon transcript levels, surface polysaccharide production, and colistin MICs to WT levels, all of which were restored by complementation (Supplementary Fig. 12). These findings demonstrate that ecaR is essential for the mcr-1-mediated activation of surface polysaccharide synthesis.
To elucidate the direct molecular targets of EcaR, we performed DNase I footprinting analysis. This revealed that EcaR binds to a specific 500-bp region upstream of wecE, designated PwecE, located within the coding sequence of wecD (Fig. 2e and Supplementary Fig. 13a). This binding site represents a previously unrecognized internal promoter within the wec operon. Although PwecA, located upstream of wecA, is regarded as the primary promoter for the ECA operon45, our findings suggest that genes downstream of wecE in the operon are regulated by PwecE through the direct interaction with EcaR (Fig. 2c–e). In silico analysis of the PwecE region further revealed a putative EcaR binding motif (Supplementary Fig. 14), supporting a role for EcaR in activating downstream transcription from this site (Fig. 2c–e).
Functional validation of this regulatory model was provided by β-galactosidase reporter assays. The presence of EcaR significantly increased PwecE activity, consistent with its role as a transcriptional activator of PwecE (Fig. 2f). In contrast, EcaR did not affect PwecA activity (Fig. 2f), corroborating our qRT-PCR and DNase I footprinting results (Fig. 2c, e). These results establish EcaR as an activator of the internal PwecE promoter, enabling precise control of downstream ECA biosynthetic genes. Additionally, while MCR-1 upregulated genes involved in amino sugar synthesis within the ECA operon, it did not affect wecE or downstream genes (Fig. 2c, g) and does not directly bind to their promoter regions (Supplementary Fig. 13b), confirming that MCR-1 and EcaR operate through distinct molecular mechanisms.
To specifically assess whether the EcaR-mediated regulation of wecE represents the key step in surface polysaccharide synthesis, we investigated whether direct expression of wecE could compensate for the absence of ecaR. Since WecE catalyzes the synthesis of dTDP-Fuc4N, a critical precursor of Fuc4NAc46, we co-expressed wecE with mcr-1 and evaluated polysaccharide production (Fig. 2h). Remarkably, wecE expression in the presence of mcr-1 significantly enhanced surface polysaccharide synthesis (Fig. 2h), indicating that WecE represents a key bottleneck in this pathway. Furthermore, glycan levels in WT + pmcr-1 + pwecE were dependent on the level of wecE induction (Fig. 2i), demonstrating a direct dose-response relationship between WecE activity and polysaccharide production. These results highlight the essential role of both mcr-1 and ecaR in ECA operon activation and confirm wecE as a key bottleneck enzyme in surface polysaccharide biosynthesis.
The enzymatic activity of MCR-1 is critical for surface polysaccharide synthesis
Having established that MCR-1 is required for surface polysaccharide synthesis, we next investigated whether this function depends on its well-characterized enzymatic activity or represents a non-enzymatic role. MCR-1 is a membrane-associated phosphoethanolamine (pEtN) transferase comprising five hydrophobic transmembrane helices and a catalytic domain5,10. Understanding the mechanistic basis of MCR-1 function in this context is crucial for determining whether surface polysaccharide activation is an intrinsic consequence of lipid A modification or requires additional protein functions.
To address this question, we constructed a Δmcr-1 mutant of the TC, which was then complemented with either WT mcr-1 (pmcr-1) or a catalytically inactive mutant (pmcr-1E246A)10,47. The E246A mutation specifically disrupts the catalytic activity of MCR-1, while preserving its overall protein structure, thus providing an ideal tool to dissect enzymatic versus non-enzymatic functions. As expected, complementation with WT mcr-1 restored colistin resistance, increasing the colistin MIC to 2 µg ml−1 compared to the Δmcr-1 mutant (MIC = 0.125 µg ml−1) (Fig. 3a). In contrast, the catalytically inactive mcr-1E246A mutant did not restore colistin resistance, resulting in MIC values similar to those of the Δmcr-1 mutant and WT, thereby confirming that the catalytic activity of MCR-1 is essential for colistin resistance.
a Colistin susceptibility of wild type (WT), transconjugant (TC), TC harboring mcr-1-deleted pFORC82_3 (TCΔmcr-1), and TCΔmcr-1 complemented with either mcr-1 (pmcr-1) or a catalytically inactive mutant (pmcr-1E246A), was determined. b Quantification of glycan levels in wild type (WT), transconjugant (TC), TCΔmcr-1, and TCΔmcr-1 harboring either pmcr-1 or pmcr-1E246A. TCΔmcr-1 mutant vs. TCΔmcr-1 mutant + pmcr-1, P = 0.0004; TCΔmcr-1 mutant+pmcr-1 vs. TCΔmcr-1 mutant + pmcr-1E246A, P = 0.0002; TCΔmcr-1 mutant vs. TCΔmcr-1 mutant + pmcr-1E246A, P = 0.9896. c Transmission electron microscopy (TEM) analysis of the surface polysaccharide layer in each strain. Cells were stained with 1.5% phosphotungstic acid for 2 min. d Surface polysaccharide profiles of WT (lane 1), TCΔmcr-1 (lane 2), and TCΔmcr-1 harboring either pmcr-1 (TCΔmcr-1 + pmcr-1; lane 3) and peptA (TCΔmcr-1+peptA; lane 4). “+” indicates gene re-introduction via plasmid. The experiments were repeated three times with similar results, and representative data from one of the experiments are shown (c, d). Data are presented as means ± SD from n = 3 biologically independent replicates (a, b). P values were determined using one-way ANOVA with Tukey’s multiple comparison test (b). ns not significant, ****P < 0.0001. The strains used were E. coli 15734 wild type (WT), WT carrying pFORC82_3 (Transconjugant, TC), TC harboring mcr-1-deleted pFORC82_3 (TCΔmcr-1), and TC harboring mcr-1-deleted pFORC82_3 and pACYC184-mcr-1 (TCΔmcr-1 + pmcr-1) or pACYC184-mcr-1E246A (TCΔmcr-1 + pmcr-1E246A), or pBAD33-eptA (TCΔmcr-1 + peptA). Source data are provided as a Source Data file.
Consistent with this finding, the same pattern was observed for surface polysaccharide synthesis. Complementation with WT mcr-1 significantly increased glycan levels compared to the Δmcr-1 mutant, whereas the catalytically inactive mutant did not induce surface polysaccharide synthesis (Fig. 3b). These results demonstrate that surface polysaccharide activation specifically requires the enzymatic function of MCR-1. TEM analysis provided visual confirmation of these biochemical results, revealing a dense cell surface layer in the mcr-1-complemented strain, which was absent in the Δmcr-1 mutant complemented with the catalytically inactive mcr-1E246A mutant (Fig. 3c). Collectively, these findings establish that the catalytic activity of MCR-1 is a prerequisite for surface polysaccharide synthesis.
To further validate this conclusion and test the generalizability of lipid A modification in triggering surface polysaccharide synthesis, we examined whether the enzymatic function of MCR-1 could be substituted by EptA, a chromosomally encoded pEtN transferase that performs the same lipid A modification as MCR-148. This experiment was designed to determine whether surface polysaccharide activation is specific to MCR-1 or represents a general response to lipid A phosphoethanolamine modification. EptA overexpression has been shown to confer colistin resistance in E. coli49. When eptA was overexpressed in the Δmcr-1 mutant, the colistin MIC increased to levels comparable to those of the TC (Supplementary Table 2), and surface polysaccharide synthesis was also restored (Fig. 3d). These results provide compelling evidence that the activation of surface polysaccharide synthesis is not specific to MCR-1 but represents a general cellular response to lipid A phosphoethanolamine modification. This indicates that other enzymes catalyzing the same lipid A modification, such as EptA, can induce surface polysaccharide synthesis when expressed at sufficient levels. This response is consistent with the hypothesis that lipid A modification alters outer membrane properties and may require additional surface structures to maintain membrane integrity and function.
These findings confirm that the enzymatic activity of MCR-1, specifically its ability to modify lipid A, is a critical determinant for surface polysaccharide synthesis in E. coli. The inability of the catalytically inactive mcr-1E246A mutant to trigger surface polysaccharide production and the compensatory effect of EptA overexpression support the central role of lipid A modification in this process.
The global prevalence of ecaR in mcr plasmids and Enterobacterales
Given that ecaR serves as a critical regulatory partner of mcr-1, we next investigated whether this functional relationship extends beyond our laboratory strains to clinical isolates worldwide. To determine the prevalence of ecaR in mcr plasmids, we analyzed 389 mcr plasmids reported from 2010 to 2020 (Supplementary Data 3). The results revealed significant rates of mcr-1/ecaR co-occurrence: among them, 117 plasmids (30.1%) were found to carry ecaR (Fig. 4a and Supplementary Data 3). This prevalence suggests that the functional partnership between mcr-1 and ecaR has been positively selected across diverse clinical contexts.
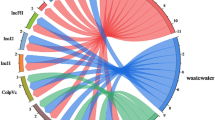
a Pangenome analysis of mcr-harboring plasmids reported from 2010 to 2020 (n = 389), performed using FriPan. Genes were clustered based on ≥95% BLASTp identity. Blue and white indicate gene presence and absence, respectively. Arrows at the top indicate the columns corresponding to mcr-1 and ecaR. The analysis reveals that 117 plasmids (30.1%) carry ecaR, with 99.1% belonging to IncI2 replicon type. b Geographic (left) and taxonomic (right) distribution of ecaR-positive mcr-harboring plasmids (n = 117), based on the dataset shown in (a). c Prevalence of the ecaR gene across Enterobacterales (n = 340), based on BLASTn analysis with 100% nucleotide identity. The left pie chart shows the genomic location of ecaR, with 91% on plasmids and 9% on chromosomes, indicating its primarily mobile nature. The right pie chart shows that among ecaR-positive plasmids, 47% also carry mcr-1 while 53% do not, revealing that ecaR has a broader distribution beyond mcr-positive contexts and may have evolved as a general ECA regulator before being recruited into mcr plasmids. Source data are provided as a Source Data file.
Geographic and taxonomic analysis revealed widespread distribution of ecaR-positive plasmids globally, with the highest prevalence detected in Asia (74.4%), followed by the Americas (11.1%), the Middle East and Europe (5.1%), Oceania and North/West Africa (1.7%), and Egypt (0.9%) (Fig. 4a, b and Supplementary Table 3). Taxonomically, ecaR was predominantly identified in E. coli (78.6%), followed by Salmonella and Shigella (each at 6.8%), Cronobacter sakazakii, Citrobacter braakii, and Klebsiella pneumoniae (each at 1.7%), and Escherichia fergusonii, Enterobacter, and Kluyvera ascorbata (each at 0.9%) (Fig. 4a, b and Supplementary Table 3). This taxonomic breadth indicates that the mcr-1/ecaR regulatory system is not species-specific but represents a transferable module that can function across diverse Enterobacterales hosts. Notably, 99.1% of these plasmids belonged to the IncI2 type, with the remaining 0.9% representing the IncI2-IncX4 hybrid type (Supplementary Table 3). Importantly, ecaR was found to coexist with various mcr variants beyond mcr-1, including mcr-1.1, mcr-3.2, and mcr-7.1 (Supplementary Data 3). This finding suggests that the regulatory partnership with ecaR may be a general feature of mcr-mediated colistin resistance rather than being specific to mcr-1.
To further characterize ecaR distribution, we expanded our analysis beyond mcr-positive plasmids. Comprehensive BLASTn analysis identified 340 ecaR-positive strains, with 91% carrying the gene on plasmids and 9% on chromosomes (Fig. 4c and Supplementary Data 4). Among the ecaR-positive plasmids, 47% also carried mcr-1, whereas 53% did not (Fig. 4c and Supplementary Data 4). While IncI2 was the most prevalent replicon type among mcr-negative ecaR plasmids, ecaR was also associated with a diverse range of replicon types, including IncY, IncX4, IncFIB, IncHI2, and IncFII (Supplementary Data 4). These results indicate that ecaR is not limited to mcr-1-harboring strains but is more broadly distributed across diverse Enterobacterales. The widespread co-localization of ecaR and mcr-1 suggests a potential functional interaction, whereby their coordinated activation of surface polysaccharide synthesis may confer a selective advantage to host bacteria.
Dual activation of colistin resistance and virulence by mcr-1 and ecaR via ECA operon-driven surface polysaccharide synthesis
With the molecular mechanisms underlying mcr-1/ecaR-mediated surface polysaccharide synthesis established, we next investigated the biological consequences of this regulatory circuit for bacterial pathogenesis and antibiotic resistance. We conducted RNA-seq analysis to examine the transcriptional effects of mcr-1 and ecaR. The results demonstrated that the presence of both mcr-1 and ecaR led to significant changes in the transcription of numerous genes (Supplementary Fig. 15a and Supplementary Data 5), especially those related to cell wall/membrane/envelope biogenesis (Supplementary Fig. 15b). Notably, upregulation of the ECA operon genes was observed only when both mcr-1 and ecaR were present, confirming that ECA operon activation requires their combined action (Supplementary Fig. 15c). In addition, co-expression of mcr-1 and ecaR increased both the net charge and hydrophilicity of the bacterial surface, while ecaR alone had no measurable impact on surface properties (Supplementary Fig. 15d, e).
To assess how surface polysaccharide synthesis induced by mcr-1 and ecaR affects pathogenicity, we employed a mouse sepsis model. Mice inoculated with strains producing surface polysaccharides (TC or WT co-expressing mcr-1 and ecaR) all died within 48 h (Fig. 5a). In contrast, mice infected with WT or with WT strains harboring either mcr-1 or ecaR alone survived (Fig. 5a). Bacterial clearance analysis revealed distinct patterns between treatment groups. WT and WT harboring either mcr-1 or ecaR alone were rapidly cleared from liver tissue (Fig. 5b). By contrast, TC and WT co-expressing mcr-1 and ecaR strains exhibited delayed clearance, with substantial bacterial levels (6.5 × 104–1.2 × 105 and 1.7 × 106–2.0 × 106 CFU g−1), still detectable at 72 h post-infection (Fig. 5b).
a Survival curves of mice (n = 6) over 72 h post-infection with wild type (WT), WT harboring either pmcr-1 or pecaR (WT + pmcr-1 or WT + pecaR), WT co-expressing mcr-1 and ecaR (WT + pmcr-1 + pecaR), and the transconjugant (TC), following inoculation with 2 × 1010 colony-forming units (CFU). WT vs. TC, P = 0.0089; WT vs. WT + pmcr-1 + pecaR, P = 0.0051; WT vs. WT + pmcr-1, P = 0.3854. Bacterial loads in liver (b), spleen (c), and kidney (d) of mice (n = 6 per group) at the indicated time points post-infection. Exact P values are provided in the Source Data file. e Minimal inhibitory concentrations (MICs) of bile salts for WT and derivative strains. f Growth kinetics of WT, TC, and WT + pmcr-1 + pecaR in LB broth with 0% (left) or 0.05% (right) bile salts. WT vs. TC, P = 0.8636; WT vs. WT + pmcr-1 + pecaR, P = 0.1563 (0% bile salts); WT vs. TC, P = 0.0012; WT vs. WT + pmcr-1 + pecaR, P = 0.0044 (0.05% bile salts). g Colistin susceptibility of WT and derivative strains. h Colistin susceptibility of rough E. coli isolates E17, E18, E20 harboring pmcr-1 and/or pecaR. Data are presented as mean ± SD from n = 10 (a), n = 6 (b–d), and n = 3 (e–h) biologically independent replicates. P values were determined using log-rank (Mantel–Cox) test (a), two-way ANOVA with Tukey’s multiple comparison test (b–d), and one-way ANOVA with Tukey’s multiple comparison test (f). ns not significant, *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. Except for h, the strains used were E. coli 15734 wild type (WT), WT carrying pFORC82_3 (Transconjugant, TC), WT harboring pACYC184-mcr-1 (WT + pmcr-1), WT harboring pBAD33-ecaR (WT + pecaR), and WT harboring both pACYC184-mcr-1 and pBAD33-ecaR (WT + pmcr-1 + pecaR). Source data are provided as a Source Data file.
Similar patterns were observed in spleen and kidney tissues. WT and WT harboring either mcr-1 or ecaR alone were rapidly cleared, while TC and WT co-expressing mcr-1 and ecaR showed delayed clearance, with 9.8 × 102–4.4 × 103 and 8.7 × 103–3.4 × 104 CFU g−1, respectively, remaining at 72 h (Fig. 5c). In the kidneys, bacterial counts in mice infected with WT or WT harboring either mcr-1 or ecaR alone fell below detectable levels at 72 h (Fig. 5d). By contrast, TC and WT co-expressing mcr-1 and ecaR strains showed delayed clearance, with 4.3 × 102–1.2 × 103 and 4.2 × 103–7.4 × 103 CFU g−1, respectively, still detectable at 72 h post-infection (Fig. 5d).
Surface polysaccharides are known to enhance the resistance of Enterobacterales to bile salts, which is crucial for their survival in the gastrointestinal tract, as exposure to bile salts can disrupt the bacterial membrane50. To assess the impact of increased ECA and O-chain synthesis on bacterial bile resistance, we measured bile salt MICs. Our findings revealed that TC or WT co-expressing mcr-1 and ecaR exhibited a four-fold increase in resistance to bile salts compared to WT (Fig. 5e). Moreover, while bile salts inhibited the growth of WT, they had minimal effect on strains expressing both genes (Fig. 5f), indicating that activation of surface polysaccharide synthesis enhances resistance to bile stress.
Bacterial surface polysaccharides form a significant barrier against antimicrobial agents51, a property that becomes particularly important when MCR‑1 diminishes colistin binding through lipid‑A modification9. We therefore hypothesized that the activation of ECA and O-chain synthesis by MCR-1 and EcaR further augments colistin resistance. In the WT background, expression of mcr-1 alone increased the MIC of colistin from 0.125 μg ml−1 to 1 μg ml−1 (Fig. 5g), whereas ecaR alone had no effect (MIC = 0.125 μg ml−1) (Fig. 5g). By contrast, the co-existence of mcr-1 and ecaR significantly elevated colistin resistance (MIC = 2 μg ml−1) to levels similar to those observed in the TC (Fig. 5g). These results suggest that this additional protection result from surface polysaccharides, which form an outer‑layer polysaccharide shield that limits colistin access to modified lipid A. Supporting this, deletion of ecaR in the TC strain or disruption of ECA and O-chain synthesis via wecG deletion reduced the MIC to 0.5 μg ml−1 despite the presence of mcr-1 (Supplementary Fig. 16). These strains, which lack ECA and O-chain but retain mcr-1, exhibited reduced susceptibility, indicating that surface polysaccharide synergizes with MCR-1-mediated lipid A modification to confer high-level colistin resistance. Significant increases in colistin resistance caused by the coexistence of mcr-1 and ecaR were observed in multiple rough E. coli isolates (Fig. 5h), which exhibited surface polysaccharide synthesis via the acquisition of an mcr-1 plasmid (Supplementary Fig. 17).
To further validate the role of ecaR, we evaluated its contribution to surface polysaccharide biogenesis and colistin resistance. Acquisition of mcr-1 plasmids lacking ecaR did not induce ECA or O-chain production in WT, whereas introducing an ecaR-positive mcr-1 plasmid (pJSMCR1_4) into WT restored their production (Supplementary Fig. 18a, b). Furthermore, the transfer of an ecaR-negative mcr-1 plasmid into WT resulted in lower colistin MICs compared to those observed with the transfer of ecaR-positive mcr-1 plasmids, indicating that mcr-1 alone cannot significantly increase colistin resistance in the absence of ecaR (Supplementary Fig. 18c and Supplementary Table 4). These findings demonstrate that both ecaR and mcr-1 are critical for ECA and O-chain production and that the resulting synthesis of surface polysaccharides contributes to the development of high-level colistin resistance.
We further examined whether surface polysaccharides, formed through the combined expression of mcr-1 and ecaR, confer resistance to other classes of antibiotics. We assessed susceptibility to six representative antibiotics, including penicillin (β-lactam), meropenem (carbapenem), erythromycin (macrolide), rifampicin (rifamycin), and gentamicin and kanamycin (aminoglycosides). Strains expressing both mcr-1 and ecaR, as well as the TC, exhibited increased resistance compared to the WT strain across all antibiotics tested (Supplementary Fig. 19). These results suggest that synthesis of surface polysaccharide alters membrane permeability in a way that broadly affects antibiotic susceptibility beyond colistin. These findings reveal that the cooperative action of MCR-1 and EcaR activates surface polysaccharide synthesis, thereby enhancing pathogenicity, bile salt tolerance, and antimicrobial resistance in E. coli.
Discussion
Our findings reveal a mechanism by which horizontally acquired resistance genes substantially alter core cellular processes, challenging the assumption that essential pathways, such as surface polysaccharide synthesis, are governed exclusively by chromosomal determinants51,52. This study demonstrates that the acquisition of mcr-1 plasmids triggers a sophisticated regulatory cascade that simultaneously enhances antimicrobial resistance and bacterial virulence through the coordinated activation of surface polysaccharide biosynthesis. This discovery advances our understanding of mobile genetic elements from simple vehicles of resistance gene dissemination to complex regulatory platforms capable of orchestrating extensive physiological changes.
The discovery of cooperative regulation between MCR-1 and EcaR represents an important advance in our understanding of surface polysaccharide biosynthesis, revising the classical model of ECA synthesis as a constitutively expressed pathway controlled solely by the PwecA promoter upstream of wecA45. Our results reveal a sophisticated dual-promoter system where MCR-1 enhances transcription of upstream genes in the wec operon, while EcaR functions as a transcriptional activator that specifically targets the PwecE promoter (Fig. 2). This dual regulatory mechanism enables precise control over WecE-mediated synthesis of dTDP-Fuc4N46, a critical precursor for the Fuc4NAc component of the ECA trisaccharide repeat unit. The conservation of the PwecE sequence across all four rough E. coli strains examined in this study (E. coli 15734, E17, E18, and E20) (Supplementary Fig. 20) suggests that this regulatory architecture may be broadly exploitable. This conservation implies that the capacity for plasmid-mediated ECA regulation may be more widespread than previously recognized, indicating a potentially universal mechanism for envelope remodeling upon acquisition of mcr-1/ecaR plasmids. Further studies are needed to assess the genome-wide distribution of EcaR binding sites in diverse E. coli strains and to explore its regulatory functions beyond the ECA operon. Additionally, the precise molecular mechanisms linking MCR-1’s pEtN transferase activity to transcriptional regulation of upstream wec genes remain to be fully elucidated and warrant further investigation through biochemical and structural approaches.
The acquisition of mcr-1 plasmids facilitates extensive remodeling of the bacterial cell envelope through two complementary mechanisms. First, MCR-1-mediated addition of pEtN to lipid A reduces the net negative surface charge of the outer membrane9,10, as evidenced by decreased zeta potential measurements (Supplementary Fig. 2f). This charge neutralization alters membrane biophysical properties, including fluidity and rigidity, with cascading effects on bacterial motility and biofilm architecture53. Second, the coordinated upregulation of surface polysaccharide biosynthesis substantially increases the density of surface glycans, significantly altering cell surface hydrophobicity and adhesion properties (Supplementary Figs. 2e and 3a). Together, lipid A modification by MCR-1 and increased surface polysaccharide expression synergize to drive the observed changes in biofilm formation, swimming motility, and zeta potential.
These envelope modifications manifest as substantial changes in bacterial behavior and physiology. The observed reductions in swimming motility and biofilm formation reflect the complex interplay between altered membrane charge distribution and increased polysaccharide burden (Supplementary Fig. 2a–c). Simultaneously, mcr-1 plasmid acquisition induces overproduction of surface polysaccharides, which alters cell surface hydrophobicity and bacterial adhesion (Fig. 1g; Supplementary Figs. 2e and 3a). The combined effects of lipid A modification and surface polysaccharide production account for the observed phenotypes.
Our data support a model in which ECA and O-chain biosynthesis are metabolically coupled via shared pools of the lipid carrier undecaprenyl phosphate (Und-P) and nucleotide-sugar precursors. In the TC, deletion of wecG abolished not only ECA but also O-chain, consistent with depletion of shared precursors and/or carrier availability (Supplementary Fig. 6a). RNA-seq results of the TC indicated an expanded precursor-supply network: galactose uptake (mglABC), PRPP generation (prs), UDP-hexose biosynthesis (galF), and dTDP-sugar pathways (e.g., rffGH) were transcriptionally upregulated, consistent with the increased demand imposed by ECA overproduction (Supplementary Table 5). By contrast, Leloir-pathway genes (galT/galK/galE) were unchanged or decreased, implying diversion of galactose flux into nucleotide-sugar pools rather than catabolism (Supplementary Table 5), with intracellular galactose/UDP-Gal accumulation as a likely consequence. Collectively, these observations suggest that, when Und-P and nucleotide-sugar precursors are sufficiently supplied, ECA and the O-chain can be synthesized concomitantly. This metabolic-coupling model explains the coordinated changes we observe and underscores the need for further study.
The correlation between colistin resistance and increased surface polysaccharide production reveals a notable dual threat that extends beyond the well-established resistance mechanisms. A previous study indicated a potential link between surface polysaccharides and antibiotic resistance54, though this relationship requires further exploration. While mcr-1 acquisition alone elevates colistin MICs above clinical breakpoints, the co-mobilization of ecaR further amplifies both surface polysaccharide synthesis and colistin resistance levels (Fig. 5g, h). This synergistic effect suggests that many colistin-resistance phenotypes previously attributed solely to mcr-1 may result from the combined activity of mcr-1 and ecaR, given their widespread co-occurrence on IncI2 and IncX4 plasmids.
The production of ECA adds additional biological complexity36,39. Unlike the highly variable O-antigen, ECA is a conserved surface polysaccharide across Enterobacterales, potentially serving as a universal virulence factor35,55. In parallel, co-production of HMW O-chain is poised to further remodel the cell surface, with consequences for complement evasion, bile-salt tolerance, and biofilm architecture. The increased ECA expression observed in mcr-1/ecaR-positive strains, together with O-chain production (Figs. 1e and 2b), may enhance immune evasion capabilities while simultaneously providing structural integrity to the outer membrane, particularly in rough strains that lack complete LPS structures. The enhanced virulence observed in murine infection models (Fig. 5a–d), coupled with improved bile salt resistance (Fig. 5e, f), suggests that mcr-1/ecaR-positive strains may exhibit a survival advantage in gastrointestinal environments. This physiological advantage facilitates both colonization and transmission, potentially accelerating the dissemination of these dual-threat plasmids in clinical and community settings. The ability to survive bile stress while maintaining enhanced pathogenic properties positions these strains as challenging pathogens capable of establishing persistent infections that are difficult to eradicate with current therapeutic approaches.
The clinical significance of our findings extends across multiple dimensions of infectious disease management, requiring reassessment of how we approach mcr-1-positive infections. The widespread presence of mcr-1 and ecaR across diverse pathogenic members of Enterobacterales (Fig. 4) indicates that this resistance-virulence coupling mechanism may extend beyond E. coli to include other major pathogens, such as Klebsiella pneumoniae and Salmonella enterica. The universal nature of ECA among Enterobacterales suggests that this coupling mechanism may be broadly exploitable across diverse pathogenic species, significantly amplifying the public health threat posed by mcr plasmids.
Although rough E. coli strains are not the predominant cause of invasive infections, our findings reveal how mcr-1 plasmid acquisition can restore virulence and resistance in attenuated isolates. We demonstrate that this horizontal gene transfer compensates for physiological deficits in rough strains, shifting them toward enhanced virulence and colistin resistance. This mechanism highlights how mobile genetic elements can expand the pathogenic and ecological range of bacterial populations. Intestinal E. coli lineages, including ETEC, act as critical reservoirs for resistance plasmids despite not being primary colistin therapy targets. From a One Health perspective, characterizing these reservoirs is essential for understanding how resistance determinants persist and spread across diverse environmental and host contexts.
Our genomic survey of nearly 400 plasmids documents frequent mcr-1 and ecaR co-occurrence, substantiating the broader significance of this regulatory architecture (Fig. 4). These data suggest widespread potential for coordinated control of surface polysaccharide biosynthesis, though systematic epidemiological studies are required to establish clinical prevalence. The frequency with which rough or other clinical isolates employ this regulatory circuit, and its population-level impact on resistance and virulence, remains to be determined. Establishing its distribution and clinical consequences across diverse E. coli lineages represents an important avenue for future investigation.
This dual augmentation of resistance and virulence fosters the emergence of bacterial strains that are simultaneously more difficult to eradicate and capable of causing more severe disease. Such strains present significant challenges to current treatment strategies and infection control measures, potentially necessitating more aggressive therapeutic approaches, including higher antibiotic dosages or combination therapies. However, these intensified treatment regimens may paradoxically exacerbate antimicrobial resistance, creating a vicious cycle of escalating resistance and treatment complexity.
Moreover, the co-localization of mcr-1 and ecaR in various members of Enterobacterales (Fig. 4) indicates their functional complementarity. These results suggest that the activation of surface polysaccharide synthesis by these two genes is critical for the fitness of bacteria in host environments. Rough mutants exhibit increased susceptibility to environmental stressors and antimicrobial agents, particularly lipophilic agents such as bile salts and hydrophobic antibiotics34. Thus, the acquisition of surface polysaccharide-producing capability represents a significant survival advantage. Since bile resistance is essential for Enterobacterales to survive and colonize the gastrointestinal tract56, the mcr-1/ecaR-mediated enhancement of ECA and O-chain production provides a crucial fitness benefit that may drive the maintenance and dissemination of these plasmids in clinical populations.
Furthermore, this study underscores the importance of considering the broader physiological implications of resistance genes when assessing their potential impact on public health. Traditional approaches to studying antibiotic resistance typically focus on the direct mechanism of resistance conferred by a particular gene57,58. However, this study demonstrates that the effects of resistance genes can be extensive, influencing fundamental aspects of bacterial biology. This suggests that a more comprehensive approach to studying antibiotic resistance is required, one that considers not just the primary resistance mechanism but also secondary effects on bacterial physiology and virulence. Our discovery of the association between mcr-1 plasmids and surface polysaccharide production unveils a previously unknown complexity in understanding and controlling antibiotic resistance.
In conclusion, our findings elucidate a remarkable association between antibiotic resistance, virulence, and fundamental cellular processes in the context of surface polysaccharide synthesis. The discovery that mcr-1 plasmids can simultaneously enhance both antibiotic resistance and virulence in Enterobacterales represents an important advancement in our understanding of bacterial pathogenesis and the multifaceted consequences of antimicrobial resistance. This work underscores the necessity for a comprehensive approach to investigating antibiotic-resistant bacteria, considering not only their ability to withstand antimicrobials but also their potential to cause more severe infections. Our results highlight the complex interplay between horizontally acquired resistance determinants and core bacterial physiology, emphasizing the need for integrated strategies in combating the global threat of antimicrobial resistance.
Methods
Bacterial strains and culture conditions
The strains used in this study are listed in Supplementary Table 6. E. coli 15734 (accession number CP097001), a clinical E. coli isolate obtained from human stool in South Korea, was used as the wild type in this study. This strain was obtained from the National Culture Collection for Pathogens (NCCP) in South Korea. This strain is classified as an enterotoxigenic E. coli (ETEC) carrying the heat-stable enterotoxin (ST) gene. In addition, ResFinder (v. 4.6.0) analysis revealed the presence of multiple antibiotic resistance genes, including aadA2, ant(2”)-Ia, and mph(A). The mcr-1 plasmid (pFORC82_3) used in this study was found in E. coli FORC82, isolated from raw chicken purchased in South Korea12. All E. coli strains were grown in Luria–Bertani (LB) medium (Difco, USA) at 37 °C. When necessary, the following concentrations of antibiotics or supplements were used: colistin (2–4 µg ml−1), chloramphenicol (25 µg ml−1), kanamycin (50 µg ml−1), tetracycline (10 µg ml−1), ampicillin (50 µg ml−1), and L-arabinose (0.2% w/v).
Bacterial strain construction
Plasmids and primers used in this study are listed in Supplementary Tables 6 and 7, respectively. Standard cloning plasmids (pACYC184 and pBAD33) were obtained from laboratory stocks. To construct the transconjugant, the mcr-1 plasmid (pFORC82_3; pMCR-1) from FORC82 was introduced into the wild type by conjugation. The Δmcr-1 mutant and ΔecaR mutant strains were generated by Lambda Red homologous recombination59 and subsequently introduced into the wild type by conjugation.
For complementation studies, the coding region of mcr-1, including its native promoter, was PCR-amplified from FORC82 and cloned into the pACYC184. The resulting construct (pACYC184-mcr-1, designated pmcr-1) was transformed into the Δmcr-1 mutant. Similarly, the coding region of ecaR was PCR-amplified from FORC82 and cloned into pBAD33. The resulting plasmid (pBAD33-ecaR, designated pecaR) was transformed into the ΔecaR mutant.
To generate the WT harboring mcr-1 and ecaR (WT + pmcr-1 + pecaR), both plasmids were co-transformed into the wild-type strain. Empty pACYC184 and pBAD33 vectors were also transformed into each strain as negative controls.
Site-directed mutagenesis of mcr-1 was performed using the overlap extension PCR method to obtain the MCR-1(E246A) variant (pmcr-1E246A), which was subsequently transformed into the Δmcr-1 mutant.
The ΔwecG mutant strain was generated using pKO360. The in-frame wecG deletion cassette was constructed using overlap PCR. An upstream fragment (~700 bp) containing the start codon (ATG) of wecG and a downstream fragment (~700 bp) starting from the stop codon (TAA) were designed, with primers incorporating a 24 bp overlapping region between the two fragments. The amplified wecG deletion cassette and the pKO3 vector were digested with BamHI and ligated using a Rapid DNA Ligation Kit (Roche, Switzerland). The constructed plasmid was introduced into the WT by electroporation, and the cells were recovered at 30 °C for 1 h. Recovered cells were plated on LB agar containing chloramphenicol, and on the following day, serially diluted onto LB agar plates containing chloramphenicol at 43 °C. Colonies were picked, serially diluted, and plated onto LB agar supplemented with 5% (w/v) sucrose. The colonies were suspended in PBS and simultaneously spotted onto LB agar plates with and without chloramphenicol, followed by incubation at 30 °C overnight. The chloramphenicol-susceptible colonies were then selected and the deletion of the wecG gene was confirmed by PCR. To generate the ΔwecG mutant of TC, the mcr-1 plasmid in FORC82 was introduced into the ΔwecG mutant by conjugation. All constructed strains were verified by Sanger sequencing of the targeted gene (Bionics, Seoul, South Korea). Materials generated in this study are available from the corresponding author upon request.
Plasmid stability test
Transconjugant (TC; WT harboring mcr-1 plasmid (pFORC82_3)) was grown overnight in LB broth supplemented with colistin. Ten microliters of overnight seed cultures were inoculated into 10 ml of fresh LB broth without colistin. Subculturing was performed after 24 h of incubation and repeated for 10 consecutive days. For each subculture, the culture was serially diluted, spread on LB agar plates, and incubated at 37 °C overnight. The number of mcr-1-deficient cells within each culture was determined by transferring 96 colonies from LB agar plates to 4 µg ml−1 colistin-containing LB broth in each well of a 96-well plate. The proportion of plasmid-containing cells was calculated by counting the number of wells with visible bacterial growth. FORC82, which originally harbored the mcr-1 plasmid (pFORC82_3), was used as the control strain.
Zeta potential measurement
Overnight cultures were subcultured 1:100 into 10 ml of LB broth and incubated at 37 °C until the OD600 reached 1.0. The bacterial cells (5 × 104 CFU ml−1) were harvested by centrifugation at 5000 × g for 5 min at 4 °C. The pellets were washed twice with ddH2O and resuspended in 1 ml of ddH2O. The zeta potential was measured using a Zetasizer Nano ZS90 analyzer (Malvern Panalytical, Malvern, UK).
Cell surface hydrophobicity test
Overnight cultures were harvested by centrifugation at 16,000 × g for 1 min at 4 °C and diluted to an OD600 of 1.0 using PBS. Subsequently, 2 ml of bacterial suspension was transferred to a tube, and the OD600 was measured. Then, 200 µl of hexane was added to the suspension, and the mixture was vortexed for 2 min. The mixture was allowed to rest for 10 min at room temperature to achieve phase separation, and the OD600 of the lower aqueous phase was measured. Cell surface hydrophobicity (%) was calculated as the relative decrease in OD600 compared to the initial OD600 value.
Autoagglutination assay
Overnight cultures were subcultured 1:100 into 10 ml of LB broth and incubated at 37 °C until the OD600 reached 1.0. Subsequently, 3 ml of the suspension was transferred to a tube and mixed vigorously for 15 s. After every 1 h for 4 h, the OD600 of the suspension from the liquid surface was measured.
Swimming motility assay
Overnight cultures of E. coli strains were diluted with fresh LB broth to an OD600 of 0.6. Two microliters of each diluted culture were inoculated into swimming motility agar (a 0.3% agar plate containing 1% tryptone and 0.25% NaCl). After incubation for 30 h at 37 °C, the swimming halos were measured.
Biofilm formation assay
To quantify biofilms of the E. coli strains, a biofilm formation assay was performed. Briefly, overnight cultures of E. coli strains were diluted with fresh LB broth to an OD600 of 0.05 and cultured in 96-well polystyrene plates for 72 h at 37 °C. The cell cultures were washed three times with PBS to remove nonadherent cells. Biofilms were stained with 0.1% crystal violet for 20 min and washed three times with PBS. The remaining crystal violet was solubilized with 33% acetic acid. The OD570 was measured using a SpectraMax i3 multimode microplate reader (Molecular Devices, CA, USA). For SEM analysis, biofilms on polystyrene coverslips incubated for 72 h at 37 °C were fixed and dehydrated, then coated with platinum and visualized using SEM (Zeiss-SIGMA, UK).
Cell adhesion assay
The adhesion assay was performed using the human colorectal adenocarcinoma cell line Caco-2 (ATCC HTB-37, American Type Culture Collection, USA). The cell line was obtained directly from ATCC, which provides authenticated cell lines, and was not further authenticated by the authors. The cell line tested negative for mycoplasma contamination, and no commonly misidentified cell lines listed in the ICLAC Register were used. Briefly, a monolayer of 1 × 105 Caco-2 cells was infected with E. coli strains at a multiplicity of infection (MOI) of 10. After 3 h of incubation at 37 °C, the cells were washed three times with prewarmed PBS, lysed in 1% Triton X-100 for 15 min, and then serially diluted in PBS. The suspensions were plated on LB agar plates to enumerate the CFUs. Data were obtained from three biological replicates.
Galleria mellonella infection model
Overnight cultures were subcultured 1:100 into 10 ml of LB broth and incubated at 37 °C until the OD600 reached 1.0. The bacteria were harvested by centrifugation at 16,000 × g for 1 min at 4 °C. The pellets were washed and then diluted to 108 CFU ml−1 with PBS. Using a 50 μl Hamilton syringe, 10 μl of bacterial suspension was injected into the pro-legs of 10 larvae, which were randomly selected. After injection, the larvae were incubated at 37 °C, and their survival was monitored for 3 days. A control group injected with PBS was included, and no larval mortality was observed in the control group.
Total RNA extraction, RNA sequencing, and analysis
Overnight cultures of E. coli strains were subcultured at a ratio of 1:100 into 10 ml of LB broth without antibiotics and grown at 37 °C to an OD600 of 0.5. Total RNA was extracted using the RNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions and quantified and assessed for quality using a NanoPhotometer N60 (Implen, CA, USA). Two or three biological replicates of RNA samples were submitted to Macrogen (Seoul, South Korea), and RNA sequencing was performed on the Illumina NovaSeq 6000 platform. The expression level of each gene was normalized by calculating the reads per kilobase (kb) per million mapped reads (RPKM) using the CLC Genomics Workbench (v9.0). Differential expression was assessed using the Empirical Analysis of DGE tool (edgeR test, two-sided) implemented in CLC Genomics workbench, with P-values adjusted for multiple testing using the Benjamini–Hochberg false discovery rate (FDR). The fold change was determined by comparison to the wild type, and the differentially expressed genes (DEGs, log 2 (fold change) ≥1 or ≤−1, P < 0.05) were filtered. The raw RNA sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive under accession codes PRJNA1094456 and PRJNA1092527.
Surface polysaccharide extraction and analysis
Surface polysaccharides (ECA and LPS) were extracted from cultures using the hot phenol-water microextraction method61,62. Bacterial cells (2 × 109 CFU ml−1) were harvested, washed with 1 ml of PBS containing 0.15 mM CaCl2 and 0.5 mM MgCl2, and centrifuged at 16,000 × g for 1 min at 4 °C. The pellets were resuspended in 100 µl of PBS and sonicated. Ten microliters of proteinase K (1 mg ml−1) were added, and the samples were incubated at 37 °C for 1 h. Then, 200 µl of ddH2O and 300 µl of preheated (68 °C) phenol solution were added, and the samples were incubated at 68 °C for 15 min. The mixture was incubated on ice for 5 min and centrifuged at 10,000 × g for 5 min at 4 °C. The surface polysaccharides were extracted by adding 300 µl of ddH2O. Then, 500 µl of 1 M sodium acetate and 10 ml of 95% ethanol were added, and the mixture was incubated at −20 °C overnight. The surface polysaccharides were harvested by centrifugation at 10,000 × g for 5 min at 4 °C, resuspended in 100 µl of ddH2O, and precipitated with 95% ethanol. The surface polysaccharide was dissolved in 50 µl of ddH2O. The surface polysaccharide extracts were analyzed by deoxycholate-polyacrylamide gel electrophoresis (DOC-PAGE), and the gels were fluorescently stained using the Pro-Q® Emerald 300 Lipopolysaccharide Gel Stain Kit (Cat. No. P20495, Molecular Probes, OR, USA) according to the manufacturer’s instructions.
Wheat germ agglutinin staining
Glycan levels were quantified by WGA staining. Bacteria (2 × 109 CFU ml−1) were harvested by centrifugation at 4000 × g for 3 min at 4 °C and washed with PBS. After washing, the cells were combined with 10 μg ml−1 WGA in PBS and incubated in the dark at room temperature for 10 min. Subsequently, the cells were washed twice and resuspended in PBS. The fluorescence (excitation at 485 nm, emission at 519 nm) and OD600 were measured. The fluorescence measurements were normalized to the OD600.
Transmission electron microscopy (TEM) analysis
Overnight cultures were harvested and washed with PBS. Bacterial cells were stained with 1.5% phosphotungstic acid for 2 min and examined via transmission electron microscopy (TEM) (JEM1010, JEOL, Japan) at an accelerating voltage of 80 kV at the National Instrumentation Center for Environmental Management (Seoul, South Korea).
Western blotting with anti-ECA monoclonal antibody
Western blot was performed using whole cell lysates63. Overnight cultures were subcultured 1:100 into 10 ml of fresh LB broth and incubated at 37 °C for 24 h. Cells were harvested by centrifugation at 16,000 × g for 10 min at 4 °C, and the pellets were resuspended in sample buffer to adjust OD600 of 5.0. The suspensions were then boiled at 100 °C for 10 min. E. coli K-12 MG1655 was included as a positive control. The whole cell lysates were resolved by PAGE, transferred to a PVDF membrane, and blocked with Dot blot buffer containing 10% skim milk. The membrane was incubated with monoclonal anti-ECA antibody (mAb ECA 898; 1:200 dilution) for 1 h, washed twice with dot blot buffer, and subsequently incubated with alkaline phosphatase-conjugated goat anti-mouse secondary antibody (D0486, DAKO, 1:2000 dilution) for 1 h at room temperature. The membrane was washed (five times for 5 min each in dot blot buffer), and the substrate (5-bromo-4-chloro-3-indolyl-phosphate, BCIP, and p-nitroblue tetrazolium chloride, NBT in AP buffer) was added (15 min incubation performed in darkness without shaking) to develop a visible reaction. The reaction was stopped by removing the substrate and adding Millipore water (at 20 °C for 5 min).
Chemical characterization of the high-molecular-weight O-chain polysaccharide
For chemical characterization, LPS was extracted using the hot phenol water method64. Purified LPS was methanolyzed with 1 M HCl in methanol at 80 °C for 18 h, re-N-acetylated at 100 °C for 1 h, and derivatized to O-trimethylsilyl (TMS) methyl glycosides65 using Tri-Sil reagent (Thermo Fisher, USA) at 80 °C for 30 min. The TMS derivatives were analyzed using GC–MS (Agilent 7890 A GC interfaced to 5975B MSD) equipped with an Equity-1 column (30 m × 0.25 mm × 0.25 μm). Gas chromatography (GC) conditions were as follows: 80 °C for 2 min, ramped to 140 °C at 20 °C min−1 (2 min hold), to 200 °C at 2 °C/min, then to 250 °C at 30 °C min−1 (5 min hold). Data were processed with Enhanced MSD ChemStation software (vF.01.03.2357; Agilent Technologies). The O-polysaccharide portion was released by mild acid hydrolysis with 1% acetic acid at 100 °C for 1 h. Lipid A was removed by three rounds of chloroform extraction. The aqueous phase was lyophilized and fractionated via size-exclusion chromatography using a Superdex 75 Increase 10/300 column (GE Healthcare), assembled with an Agilent Technologies 1200 Series System, with 50 mM ammonium acetate buffer at 0.5 ml min−1 used as eluent. Fractions were monitored by refractive index (RID-10A, Shimadzu), and high molecular weight (HMW) fractions were collected and lyophilized. Purified polysaccharides were exchanged twice in D2O and analyzed by NMR spectroscopy. Spectra were recorded at 298 K on a Bruker Avance NEO 800 MHz NMR spectrometer equipped with a 1.7 mm cryoprobe. Standard 1D and 2D experiments, including COSY, TOCSY, HSQC, HMBC, and NOESY were performed to assign structural features and glycosidic linkages. Chemical shifts were referenced to the DSS signal (δH = 0.00 ppm; δC = 0.00 ppm). NMR data were processed using MestReNova v14.2.1-27684 (Mestrelab Research S.L., Spain) and Bruker Topspin 4.1.3.
Quantitative real-time (qRT)-PCR
RNA was extracted as described above, and cDNA was synthesized from the extracted RNA using cDNA EcoDry Premix (Clontech, CA, USA). Real-time PCR was conducted on cDNA using 2× iQ SYBR Green Supermix (Bio-Rad, CA, USA) and the CFX ConnectTM Real-time PCR system (Bio-Rad, CA, USA), and the data for the samples were normalized to 16S rRNA expression. The data are shown as the fold change compared with that of the wild type.
DNase I footprinting assay
The promoter regions were amplified by PCR with 6-FAM-labeled primers (Supplementary Table 7) and purified. Labeled DNA fragments (200 ng) and EcaR proteins (800 ng) were incubated in binding buffer (10 mM Tris-HCl (pH 8.0), 50 mM KCl, 8 mM MgCl2, 50 ng μl−1 BSA, 5% glycerol, and 1 mM dithiothreitol) for 15 min at room temperature. Subsequently, 0.08 U of DNase I (NEB), 10 mM CaCl2, and 10 mM MgCl2 were added to the reaction mixture and incubated for 30 s at room temperature. The reaction was stopped by heating at 65 °C for 10 min in the presence of 30 mM ethylenediaminetetraacetic acid (EDTA). The DNA fragments were purified and analyzed using an ABI3730xI DNA Analyzer (Applied Biosystems) and visualized with Peak Scanner software v1.0 (Applied Biosystems).
β-galactosidase assay
β-galactosidase activity was measured in E. coli K-12 MG1655 harboring the pMW10 recombinant plasmid carrying the 500-bp region upstream of the ATG start codon of wecA and wecE. E. coli MG1655 with pMW10 alone was used as a negative control. Overnight cultures of strains carrying promoter and lacZ fusion plasmids were subcultured at a ratio of 1:100 into 10 ml of LB broth and grown at 37 °C for 4 h. Bacteria were harvested by centrifugation at 13,000 × g for 1 min at 4 °C and resuspended in 1 ml of Z-buffer. Subsequently, the OD600 was measured. Then, 100 μl of each sample was resuspended in 900 μl of Z-buffer, 40 μl of chloroform and 20 μl of 0.1% SDS were added, and the mixture was vortexed. After incubating for 10 min, 200 μl of o-nitrophenyl-β-D-galactopyranoside (ONPG) was added, and the mixture was vortexed. When a color change was observed (time t), the reactions were terminated with 500 μl of 1 M Na2CO3, and the OD420 and OD550 were measured to calculate the enzyme activity.
Pangenome analysis of the mcr plasmid
The sequences of the mcr plasmids reported from 2010 to 2020 were collected, and pangenome analysis was performed using Roary Pangenome Pipeline (v3.12.0; 95% BLASTp percentage identity)66 and visualized using Fri-pan (v.1.2.1).
Antimicrobial susceptibility testing
Minimum inhibitory concentrations (MICs) of colistin were determined by the gradient diffusion method using E-test strips (bioMérieux, France). Colistin (0.016–256 mg l−1) was applied onto the Muller-Hinton (MH) agar plates inoculated with bacterial suspensions adjusted to a McFarland standard of 0.5 and incubated at 37 °C for 16–20 h. MIC values were recorded at the point where the ellipse of inhibition intersected the strip, following the manufacturer’s instructions.
In addition, MICs of other antibiotics were measured using the broth microdilution method in accordance with Clinical and Laboratory Standards Institute (CLSI) guidelines. Six representative antibiotics from different drug classes, including penicillin (β-lactam), meropenem (carbapenem), erythromycin (macrolide), rifampicin (rifamycin), gentamicin and kanamycin (aminoglycosides), were tested. Bacterial suspensions were prepared in cation-adjusted MH Broth Ⅱ and applied to two-fold serial dilutions of each antibiotic in 96-well microtiter plates. Plates were incubated at 37 °C for 16–20 h, and MICs were determined as the lowest concentration that inhibited visible bacterial growth.
Animal experiments
Six-week-old female BALB/c mice (18–22 g) were purchased from SPF (Beijing, China) Biotechnology Co., Ltd. and used for all experiments (6 mice per group). The mice were acclimated for 7 days after arrival before the experiments began. Mice were acclimated in a specific pathogen-free room for 1 week to minimize stress, with access to antibiotic-free feed and water. The light cycle was set to 12 h of light and 12 h of darkness, with a temperature range of 18–23 °C and humidity maintained between 40% and 60%. The environmental conditions were monitored and maintained daily by vivarium staff. Bacterial strains were grown in LB broth for 6–8 h at 37 °C, harvested by centrifugation (3000 × g, 10 min), washed three times in sterile PBS and resuspended in physiological saline (PBS) and adjusted to a concentration of 2 × 1010 CFU in 0.2 ml PBS. Each group of mice (n = 6) was then injected intraperitoneally (i.p.) with bacterial strains to determine the 50% lethal dose (LD50). Mortality was recorded daily for 3 days. LD50 values were calculated by the Reed-Muench method. For subsequent virulence experiments, the challenged mice will be monitored, and deaths will be recorded daily for 3 days. For subsequent experiments to assess virulence, an inoculum of 2 × 109 CFU in 0.2 ml PBS was administered i.p. to each mouse. Spleen, liver, and kidney tissues were collected at 12, 24, 48, and 72 h post-infection, homogenized, and serially diluted for CFU enumeration (detection limit = 100 CFU g−1). Survival curves were compared by the log-rank (Mantel–Cox) test. Bacterial loads were assessed by two-way ANOVA with Tukey’s posthoc test. A P < 0.05 was considered significant. All experiments were performed in female mice; sex was not considered as a variable in the study design, and data disaggregation by sex is not applicable.
Growth rates with bile salts
Overnight cultures were diluted 1:100 with fresh LB broth with or without 0.05% bile salts. Bacterial growth was monitored every 30 min for 12 h at 37 °C. Growth curves were determined by measuring the OD600 using a SpectraMax i3 multimode microplate reader (Molecular Devices, CA, USA).
Statistical analysis
All statistical analyses were performed on three or more independent experiments, unless otherwise stated, using GraphPad Prism (version 8.0.1, USA). Sample size and information about statistical tests are reported in the figure legends and “Methods”. Data are presented as mean ± SD.
Ethical statement
All animal studies were approved by the Committee on Animal Welfare and Ethics at China Agricultural University (AW52114202-2-01).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive under accession codes PRJNA1094456 and PRJNA1092527. Source data are provided with this paper.
References
van Duin, D. & Paterson, D. L. Multidrug resistant bacteria in the community: an update. Infect. Dis. Clin. N. Am. 34, 709 (2020).
Murray, C. J. et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655 (2022).
Acman, M. et al. Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene bla NDM. Nat. Commun. 13, 1131 (2022).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 31, https://doi.org/10.1128/cmr.00088-00017 (2018).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Thomas, R. et al. The use of polymyxins to treat carbapenem resistant infections in neonates and children. Expert Opin. Pharmacother. 20, 415–422 (2019).
Sabnis, A. et al. Colistin kills bacteria by targeting lipopolysaccharide in the cytoplasmic membrane. elife 10, e65836 (2021).
El-Sayed Ahmed, M. A. E.-G. et al. Colistin and its role in the era of antibiotic resistance: an extended review (2000–2019). Emerg. Microbes Infect. 9, 868–885 (2020).
Gao, R. et al. Dissemination and mechanism for the MCR-1 colistin resistance. PLoS Pathog. 12, e1005957 (2016).
Hinchliffe, P. et al. Insights into the mechanistic basis of plasmid-mediated colistin resistance from crystal structures of the catalytic domain of MCR-1. Sci. Rep. 7, 39392 (2017).
Quan, J. et al. Prevalence of mcr-1 in Escherichia coli and Klebsiella pneumoniae recovered from bloodstream infections in China: a multicentre longitudinal study. Lancet Infect. Dis. 17, 400–410 (2017).
Kim, J. et al. Characterization of mcr-1-harboring plasmids from pan drug-resistant Escherichia coli strains isolated from retail raw chicken in South Korea. Microorganisms 7, 344 (2019).
McGann, P. et al. Escherichia coli harboring mcr-1 and bla CTX-M on a novel IncF plasmid: first report of mcr-1 in the United States. Antimicrob. Agents Chemother. 60, 4420–4421 (2016).
Kieffer, N., Aires-de-Sousa, M., Nordmann, P. & Poirel, L. High rate of MCR-1–producing Escherichia coli and Klebsiella pneumoniae among pigs, Portugal. Emerg. Infect. Dis. 23, 2023 (2017).
Lu, X. et al. Epidemiologic and genomic insights on mcr-1-harbouring Salmonella from diarrhoeal outpatients in Shanghai, China, 2006–2016. EBioMedicine 42, 133–144 (2019).
Yi, L. et al. mcr-1− Harboring Salmonella enterica serovar Typhimurium sequence type 34 in pigs, China. Emerg. Infect. Dis. 23, 291 (2017).
Che, Y. et al. Characterization of two novel colistin resistance gene mcr-1 variants originated from Moraxella spp. Front. Microbiol. 14, 1153740 (2023).
Ngbede, E. O. et al. Identification of mobile colistin resistance genes (mcr-1.1, mcr-5 and mcr-8.1) in Enterobacteriaceae and Alcaligenes faecalis of human and animal origin, Nigeria. Int. J. Antimicrob. Agents 56, 106108 (2020).
Wang, R. et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 9, 1179 (2018).
Risely, A. et al. Host-plasmid network structure in wastewater is linked to antimicrobial resistance genes. Nat. Commun. 15, 555 (2024).
Newbury, A. et al. Fitness effects of plasmids shape the structure of bacteria–plasmid interaction networks. Proc. Natl. Acad. Sci. USA 119, e2118361119 (2022).
Carattoli, A. Plasmids and the spread of resistance. Int. J. Med. Microbiol. 303, 298–304 (2013).
Strepis, N. et al. Genetic analysis of mcr-1-carrying plasmids from gram-negative bacteria in a dutch tertiary care hospital: evidence for intrapatient and interspecies transmission events. Front. Microbiol. 12, 727435 (2021).
Yi, L. et al. PixR, a novel activator of conjugative transfer of IncX4 resistance plasmids, mitigates the fitness cost of mcr-1 carriage in Escherichia coli. mBio 13, e03209–e03221 (2022).
Yang, J. et al. A ProQ/FinO family protein involved in plasmid copy number control favours fitness of bacteria carrying mcr-1-bearing IncI2 plasmids. Nucleic Acids Res. 49, 3981–3996 (2021).
Matamoros, S. et al. Global phylogenetic analysis of Escherichia coli and plasmids carrying the mcr-1 gene indicates bacterial diversity but plasmid restriction. Sci. Rep. 7, 15364 (2017).
Costerton, J. & Cheng, K.-J. The role of the bacterial cell envelope in antibiotic resistance. J. Antimicrob. Chemother. 1, 363–377 (1975).
Zgurskaya, H. I., López, C. A. & Gnanakaran, S. Permeability barrier of Gram-negative cell envelopes and approaches to bypass it. ACS Infect. Dis. 1, 512–522 (2015).
Silhavy, T. J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414 (2010).
Ophir, T. & Gutnick, D. L. A role for exopolysaccharides in the protection of microorganisms from desiccation. Appl. Environ. Microbiol. 60, 740–745 (1994).
Lerouge, I. & Vanderleyden, J. O-antigen structural variation: mechanisms and possible roles in animal/plant–microbe interactions. FEMS Microbiol. Rev. 26, 17–47 (2002).
Bertani, B. & Ruiz, N. Function and biogenesis of lipopolysaccharides. EcoSal 8, ESP-0001-2018 (2018).
Murray, G. L., Attridge, S. R. & Morona, R. Altering the length of the lipopolysaccharide O antigen has an impact on the interaction of Salmonella enterica serovar Typhimurium with macrophages and complement. J. Bacteriol. 188, 2735–2739 (2006).
Delcour, A. H. Outer membrane permeability and antibiotic resistance. Biochim. Biophys. Acta 1794, 808–816 (2009).
Kuhn, H.-M., Meier-Dieter, U. & Mayer, H. ECA, the enterobacterial common antigen. FEMS Microbiol. Rev. 4, 195–222 (1988).
Kiss, P., Rinno, J., Schmidt, G. & Mayer, H. Structural studies on the immunogenic form of the enterobacterial common antigen. Eur. J. Biochem. 88, 211–218 (1978).
Rick, P., Mayer, H., Neumeyer, B., Wolski, S. & Bitter-Suermann, D. Biosynthesis of enterobacterial common antigen. J. Bacteriol. 162, 494–503 (1985).
Dell, A. et al. The enterobacterial common-antigen, a cyclic polysaccharide. Carbohydr. Res. 133, 95–104 (1984).
Kunin, C. M., Beard, M. V. & Halmagyi, N. E. Evidence for a common hapten associated with endotoxin fractions of E. coli and other Enterobacteriaceae. Proc. Soc. Exp. Biol. Med. 111, 160–166 (1962).
Choi, Y. et al. Comparison of fitness cost and virulence in chromosome-and plasmid-mediated colistin-resistant Escherichia coli. Front. Microbiol. 11, 798 (2020).
Maczuga, N., Tran, E. N. & Morona, R. Subcellular localization of the enterobacterial common antigen GT-E-like glycosyltransferase, WecG. Mol. Microbiol. 118, 403–416 (2022).
Peters, H. et al. Monoclonal antibodies to enterobacterial common antigen and to Escherichia coli lipopolysaccharide outer core: demonstration of an antigenic determinant shared by enterobacterial common antigen and E. coli K5 capsular polysaccharide. Infect. Immun. 50, 459–466 (1985).
Kuhn, H. M., Basu, S. & Mayer, H. Comparison of enterobacterial common antigen from different species by serological techniques. Eur. J. Biochem. 162, 69–74 (1987).
Maciejewska, A., Kaszowska, M., Jachymek, W., Lugowski, C. & Lukasiewicz, J. Lipopolysaccharide-linked enterobacterial common antigen (ECALPS) occurs in rough strains of Escherichia coli R1, R2, and R4. Int. J. Mol. Sci. 21, 6038 (2020).
Rai, A. K., Carr, J. F., Bautista, D. E., Wang, W. & Mitchell, A. M. ElyC and cyclic enterobacterial common antigen regulate synthesis of phosphoglyceride-linked enterobacterial common antigen. mBio 12, e02846–02821 (2021).
Hwang, B.-Y., Lee, H.-J., Yang, Y.-H., Joo, H.-S. & Kim, B.-G. Characterization and investigation of substrate specificity of the sugar aminotransferase WecE from E. coli K12. Chem. Biol. 11, 915–925 (2004).
Yang, Q. et al. Balancing mcr-1 expression and bacterial survival is a delicate equilibrium between essential cellular defence mechanisms. Nat. Commun. 8, 2054 (2017).
Lee, H., Hsu, F.-F., Turk, J. & Groisman, E. A. The PmrA-regulated pmrC gene mediates phosphoethanolamine modification of lipid A and polymyxin resistance in Salmonella enterica. J. Bacteriol. 186, 4124–4133 (2004).
Janssen, A. B. et al. Nonclonal emergence of colistin resistance associated with mutations in the BasRS two-component system in Escherichia coli bloodstream isolates. mSphere 5, https://doi.org/10.1128/msphere. 00143–00120 (2020).
Begley, M., Gahan, C. G. & Hill, C. The interaction between bacteria and bile. FEMS Microbiol. Rev. 29, 625–651 (2005).
Whitfield, C. & Trent, M. S. Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 83, 99–128 (2014).
Wyres, K. L., Lam, M. M. & Holt, K. E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 18, 344–359 (2020).
Needham, B. D. & Trent, M. S. Fortifying the barrier: the impact of lipid A remodelling on bacterial pathogenesis. Nat. Rev. Microbiol 11, 467–481 (2013).
Mitchell, A. M., Srikumar, T. & Silhavy, T. J. Cyclic enterobacterial common antigen maintains the outer membrane permeability barrier of Escherichia coli in a manner controlled by YhdP. mBio 9, 01321–01318 (2018).
Rai, A. K. & Mitchell, A. M. Enterobacterial common antigen: synthesis and function of an enigmatic molecule. mBio 11, 01914–01920 (2020).
Urdaneta, V. & Casadesús, J. Interactions between bacteria and bile salts in the gastrointestinal and hepatobiliary tracts. Front. Med. 4, 163 (2017).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Link, A. J., Phillips, D. & Church, G. M. Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: application to open reading frame characterization. J. Bacteriol. 179, 6228–6237 (1997).
Kim, M. & Ryu, S. Spontaneous and transient defence against bacteriophage by phase-variable glucosylation of O-antigen in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 86, 411–425 (2012).
Kuhn, H. M., Adamus, G., Romanowska, E. & Mayer, H. Effect of proteins on the immunogenicity of enterobacterial common antigen. Infect. Immun. 34, 373–377 (1981).
Rabsztyn, K. et al. Characterization of anti-ECA antibodies in rabbit antiserum against rough Yersinia enterocolitica O:3. Biochemistry 76, 832–839 (2011).
Westphal, O. Methods Carbohydrate Chemistry 5 (ed. Whistler R. L.) 83–91 (Academic Press, 1965).
York, W. S., Darvill, A. G., McNeil, M., Stevenson, T. T. & Albersheim, P. Methods in Enzymology. Vol. 118 (eds Packer, L. & Glazer, A. N.) 3–40 (Academic Press, 1986).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Acknowledgements
We thank Prof. Do Yup Lee from Seoul National University for assistance with metabolite analysis. The chemical and structural characterization of LPS was supported by the U.S. Department of Energy, Office of Science, Basic Energy Sciences, Chemical Sciences, Geosciences and Biosciences Division, under award DE-SC0015662 to DOE Center for Plant and Microbial Complex Carbohydrates at the CCRC and NIH R24GM137782 award to National Glycoscience Resource-CCRC Service and Training to P.A. B.J. is supported by MnDRIVE (Minnesota’s Discovery, Research, and InnoVation Economy). This work was supported by the Ministry of Education of the Republic of Korea and the National Research Foundation of Korea (NRF-2023R1A2C1006359).
Author information
Authors and Affiliations
Contributions
E.A. and J.K. (Jinshil Kim) conceived the research, designed experiments, performed data analysis, and wrote the manuscript. J.J., Y.C., and Y.W. conducted mouse sepsis infection and wrote the manuscript. J.K. (Joonbeom Kim) contributed to mutant construction. A.M. and K.K. conducted western blotting and wrote the manuscript. H.S. contributed to pangenome analysis and LPS/ECA extraction. Y.S. assisted with LPS/ECA extraction. Y.P. conducted metabolite analysis. A.M., C.H., L.F., and P.A. performed LPS extraction, chemical, and NMR experiments. B.J. and S.R. conceived the research, interpreted data, and edited the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Toyotaka Sato, Bruno Gonzalez-Zorn, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ahn, E., Kim, J., Jiang, J. et al. Colistin resistance plasmids dually enhance bacterial virulence and antibiotic resistance via surface polysaccharide biosynthesis. Nat Commun 16, 9966 (2025). https://doi.org/10.1038/s41467-025-65412-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65412-9