Abstract
Plant cell differentiation incorporates environmental cues over time to optimize overall growth. Iron deficiency influences development, such as root hair, cortical, and endodermal cell differentiation. However, the mechanisms by which iron deficiency regulates cell differentiation are not well characterized. Root sieve elements serve as an excellent model for cell differentiation since all cells, from undifferentiated to differentiated, are present in a distinct cell file. Here, we use semi-automated image analysis to show that iron deficiency delays sieve element differentiation, particularly enucleation and cell wall thickening, and consequently delays phloem sap unloading to roots. Using Dynamic Bayesian modeling we also characterize how iron deficiency changes the fundamental structure of the gene regulatory network associated with sieve element differentiation. We identify DOF1.5 as a positive regulator of sieve element enucleation and, consequently, of root sap translocation. These results clarify how abiotic stress can influence overall plant growth as a consequence of negatively influencing vascular differentiation.
Similar content being viewed by others
Introduction
Plants respond to both internal and external cues to optimize growth, development, and reproduction within the constraints of the local environment1. Abiotic stress alters the regulation of plant cell differentiation and patterning to mediate plant adaptation to environmental stressors1,2,3. Much work has been done to improve our understanding of how cell differentiation responds to developmental cues and abiotic stressors. For example, iron deficiency controls root hair or transfer cell differentiation in tomato4, and cortex, endodermal, and palisade cell differentiation in Arabidopsis5. Cell differentiation is also required for several iron deficiency responses6. Yet, many questions remain about how specific stress conditions influence the temporal progression of cellular differentiation, especially in vascular tissues, which is a challenge that can be tackled using tissue-specific tools and model systems3,7,8,9.
The phloem vascular tissue in Arabidopsis roots is a useful model in assessing the progression of cell differentiation because the protophloem consists primarily of a single file of only 20–25 cells that originate from the root stem cells in the Quiescent Center (QC)10,11. Consequently, each progressive differentiation stage is present and can be analyzed spatially at various time points and under different conditions, especially with respect to each root development zone10,11,12. Phloem initials, derived from the root stem cells, divide to form cells that contribute to the transit-amplifying region of the root meristematic zone10,13. Transit-amplifying cells that have entered the elongation zone then begin to differentiate into sieve elements (SEs), the functional units of the phloem10,13,14. SE differentiation is defined by three major processes: enucleation, cell wall thickening, and sieve pore formation10,14,15,16,17,18.
The phloem transcriptional regulator, ALTERED PHLOEM DEVELOPMENT (APL), controls SE enucleation via NAC DOMAIN CONTAINING PROTEIN 86 (NAC86). NAC86 then positively regulates NAC45/86-DEPENDENT EXONUCLEASE-DOMAIN PROTEIN 1–4 (NEN1-4), a family of nucleases that mediate degradation of nuclei in maturing SE cells10,16,19. Phloem sap, containing photosynthetic products contributing to plant growth, can only move between enucleated sieve tubes. Consequently, enucleation is directly required for phloem sap translocation10,20. Interestingly, enucleation is also required for terminal erythroid differentiation of mammalian red blood cells and is dependent on intracellular iron concentration21. Without enucleation, mammalian red blood cells would not be able to transport oxygen to sink tissues21. Therefore, it is possible that iron is a fundamental component of enucleation-mediated cell differentiation regardless of taxonomy.
Here, we examine how iron deficiency regulates enucleation and other processes involved in SE differentiation in Arabidopsis roots. To do this, we extracted shape metrics, a key proxy for measuring enucleation progression, from microscopy images using a modified version of the BioVisionTracker (BVT)22, which showed that iron deficiency delays nuclear breakdown in differentiating SEs. We also isolated genes expressed in the major stages of SE differentiation using GFP marker lines representing initial, transit-amplifying, and terminally differentiating SE cells. We then selected the set of genes whose enrichment in each stage of differentiation was different between iron sufficiency and iron deficiency and inferred Gene Regulatory Networks (GRN) associated with SE differentiation under iron sufficiency and iron deficiency conditions. We found that while the iron sufficiency and iron deficiency networks both had similar and unique master regulators, DOF1.5 is a significant and likely positive regulator of SE differentiation, especially of enucleation, under both iron sufficiency and iron deficiency conditions. Thus, abiotic stress induces intrinsic mechanisms that control SE differentiation, and consequently, the perception of phloem-derived signals for root elongation.
Results
Iron deficiency delays the progression of enucleation and overall SE differentiation
To observe the impact that iron deficiency has on the progression of enucleation in differentiating SEs, we measured the shape of enucleating nuclei in different cross-sectional locations of roots under iron sufficiency and iron deficiency treatments using the BVT22. Because the shape of enucleating nuclei gives an approximation of the extent of nuclear breakdown in each nucleus16, we imaged the reporter line, pNAC86:HB2-YFP, which marks NAC86 expression in the nuclei of SE cells cued for differentiation and of companion cells. To capture changes to nuclear breakdown during seedling development, pNAC86:HB2-YFP seedlings were grown for 4 days under iron sufficiency conditions then imaged at 24 h, 48 h, or 72 h after being transferred to either iron sufficiency or iron deficiency conditions (Supplementary Fig. 1). To isolate signals that were specific to SEs (i.e., filtering out signal associated with companion cells), we used the BVT software to locate and measure NAC86 expression and then applied a neural network classifier to distinguish between the two cell type classes (Supplementary Fig. 2). We mapped the progression of enucleation in SEs using two metrics for each SE nucleus: (1) an elongation index, which measures the shape of a nucleus as it is distorted from spherical to ovoid by calculating the spatial covariance matrix of pixel values for a segmented nucleus (Fig. 1a, b, see Methods), and (2) the longitudinal distance measured from the QC to each SE nucleus (Fig. 1a–c), which generally gives a proxy for when, through the course of meristematic cell division, SEs initiate or terminate enucleation16,23. Comparing the elongation index to the distance from the QC quantifies the progression of enucleation with respect to cell elongation zones23. SE differentiation typically occurs within 20–25 cells from the QC, so any deviation from this cell count or its corresponding distance from the QC indicates whether SE differentiation is delayed (>20–25 cells) or promoted (<20–25 cells)10,12.
a Representative image of the detection of SE nuclei and Epidermal Cell Lengths using BioVision Tracker (BVT). Scale bar of root panel = 30 µm. Scale bar of “Elongation index” panel and of “Epidermal Cell Length” panel = 10 µm. b Representative image of the mapping of each SE nuclear Elongation Index to c the average Epidermal Cell Length at the same Distance from the Quiescent Center (QC). d Ratio of Elongation Index to distance from the QC at 24 h, 48 h, and 72 h under iron sufficiency and iron deficiency conditions. Sample sizes are n = cells, n = 241 (24 h iron sufficiency, +Fe), n = 407 (24 h iron deficiency, -Fe), n = 154 (48 h iron sufficiency), n = 187 (48 h iron deficiency), n = 198 (72 h iron sufficiency), and n = 166 (72 h iron deficiency). The statistical analysis of the ratio between elongation index and distance to the QC in WT was determined using a two-way Student’s T Test. For 24 h iron sufficiency, the minimum = 0.01337, maximum = 2.214, median = 0.9761, 25% percentile = 0.6753, 75% percentile = 1.329. For 24 h iron deficiency, the minimum = 0.008458, maximum = 2.628, median = 1.036, 25% percentile = 0.6038, 75% percentile = 1.421. For 48 h iron sufficiency, the minimum = 0.9997, maximum = 1.969, median = 1.02, 25% percentile = 0.7138, 75% percentile = 1.277. For 48 h iron deficiency, the minimum = 0.2857, maximum = 2.517, median = 1.243, 25% percentile = 0.8914, 75% percentile = 1.557. For 72 h iron sufficiency, the minimum = 0.03601, maximum = 2.276, median = 0.9349, 25% percentile = 0.6625, 75% percentile = 1.322. For 72 h iron deficiency, the minimum = 0.07271, maximum = 1.171, median = 0.5657, 25% percentile = 0.451, 75% percentile = 0.7419. e Ratio of elongation index to epidermal cell length at 24 h, 48 h, and 72 h under iron sufficiency or iron deficiency conditions. Sample sizes are n = cells, n = 241 (24 h iron sufficiency), n = 407 (24 h iron deficiency), n = 155 (48 h iron sufficiency), n = 187 (48 h iron deficiency), n = 198 (72 h iron sufficiency), n = 167 (72 h iron deficiency). The statistical analysis of the ratio between elongation index and epidermal cell length in pNAC86:HB2-YFP was determined using a two-way Student’s T Test. For 24 h iron sufficiency, the minimum = 0.01337, maximum = 2.214, median = 0.9761, 25% percentile = 0.6753, 75% percentile = 1.329. For 24 h iron deficiency, the minimum = 0.008458, maximum = 2.628, median = 1.036, 25% percentile = 0.6038, 75% percentile = 1.421. For 48 h iron sufficiency, the minimum = 0.09922, maximum = 2.152, median = 1.015, 25% percentile = 0.7115, 75% percentile = 1.268. For 48 h iron deficiency, the minimum = 0.2835, maximum = 2.499, median = 1.233, 25% percentile = 0.8848, 75% percentile = 1.545. For 72 h iron sufficiency, the minimum = 0.07863, maximum = 4.97, median = 2.042, 25% percentile = 1.447, 75% percentile = 2.886. For 72 h iron deficiency, the minimum = 0.1588, maximum = 2.612, median = 1.236, 25% percentile = 0.9898, 75% percentile = 1.625. f Representative image of WT roots under 72 h of iron sufficiency conditions. “S” represents distance to the first sieve element with cell wall thickening, while “E” represents distance to the first elongated epidermal cell. Scale bars = 20 µm. Inset highlights cell wall thickening in the first differentiating sieve element. g Model depicting the derivation of Ratio-S/Ep. “S” denotes the distance from QC to the first sieve element with cell wall thickening while “Ep” denotes the distance from QC to the first elongated epidermal cell. QC = yellow cells, Undifferentiated phloem cells = green, Differentiating sieve elements = red, Epidermal cells = red outer cell file, Meristematic zone = boxed light orange region, Elongation zone = boxed orange region. h Ratio-S/Ep in WT roots at 24 h, 48 h, and 72 h of iron sufficiency and iron deficiency conditions. + represents iron sufficiency, while—represents iron deficiency. +Fe represents iron sufficiency. Sample sizes are n = roots, n = 13 (24 h iron sufficiency), n = 14 (24 h iron deficiency), n = 24 (48 h iron sufficiency), n = 20 (48 h iron deficiency), n = 19 (72 h iron sufficiency), and n = 18 (iron deficiency). For 24 h iron sufficiency (+Fe), the minimum = 0.5073, maximum = 1.453, median = 1.036, 25% percentile = 0.8066, 75% percentile = 1.163. For 24 h iron deficiency (−Fe), the minimum = 0.5142, maximum = 1.453, median = 1.036, 25% percentile = 0.8066, 75% percentile = 1.163. For 48 h iron sufficiency, the minimum = 0.4312, maximum = 1.501, median = 0.9786, 25% percentile = 0.7813, 75% percentile = 1.209. For 48 h iron deficiency, the minimum = 0.4157, maximum = 1.343, median = 0.8373, 25% percentile = 0.626, 75% percentile = 1.171. For 72 h iron sufficiency, the minimum = 0.4177, maximum = 1.735, median = 0.9315, 25% percentile = 0.626, 75% percentile = 1.192. For 72 h iron deficiency, the minimum = 0.2843, maximum = 1.324, median = 0.7969, 25% percentile = 0.5583, 75% percentile = 0.8986. Statistical analysis was performed using a two-way Student’s T Test (p < 0.05).
To compare enucleation with respect to the distance from the QC between iron sufficiency and iron deficiency conditions, we calculated the ratio of the elongation index over the distance from the QC of each enucleating nucleus for the time points 24 h, 48 h, and 72 h, respectively, and normalized each ratio relative to the values at 24 h of iron sufficiency (Fig. 1d). We observed differences in this ratio between the iron sufficiency and iron deficiency conditions at 48 h and 72 h (Fig. 1d). At 48 h, this ratio was higher but at 72 h, this ratio was lower. This suggests that the progression of nuclear breakdown in SEs with respect to their spatial distribution within the root was affected by iron deficiency. A low ratio here could suggest that the SEs and elongating epidermal cells are differentiating at different positions in the root with respect to each other. Either the SEs could be comparatively closer to the QC, or the elongating epidermal cells could be further from the QC compared to the SEs.
Since both SE differentiation and root differentiation zones may be affected by iron deficiency, we compared changes to SE differentiation with changes to root elongation. To capture iron deficiency-mediated changes to the root differentiation zones, we measured the lengths of cortical and epidermal cells in wildtype (WT) roots imaged at 24 h, 48 h, and 72 h of iron sufficiency and iron deficiency treatment (Supplementary Figs. 3–4). Each cell file and cell length was autonomously detected by the BVT software using the local phase symmetry of each image24,25. The measured cell lengths were used as input data for the BVT software to plot the trend of cell lengths compared to the longitudinal distance from the QC (Fig. 1a, c). We observed that in WT roots at 48 h and 72 h, cortical and epidermal cells were significantly longer under iron deficiency than under iron sufficiency within 300 μm from the QC (Supplementary Fig. 4). We found similar results in the pNAC86:HB2-YFP reporter line (Supplementary Fig. 5). Since cells elongate as they enter the root transition and elongation developmental zones, these data suggest that 48–72 h of iron deficiency results in the establishment of root developmental zones closer to the QC. This suggests that epidermal cell elongation under iron deficiency is initiated spatially earlier in the root under iron-sufficient conditions. Thus, although enucleation occurs at a similar position within each root with respect to the QC between iron sufficiency and iron deficiency conditions, the position at which enucleation occurs is developmentally later. Since enucleation is typically initiated around the transition zone (e.g., between the meristematic zone and elongation zone), our findings indicate that iron deficiency causes a delay in SE enucleation.
To compare the progression of enucleation to the location of developmental zones at a similar distance from the QC, we collected the elongation index of every enucleating cell and the average epidermal cell length (a proxy for where developmental zones occur), measured using BVT (See Methods), at the same cross-section of the root (Fig. 1b-c). We then took the ratio of the elongation index over the corresponding epidermal cell length, normalized either set of ratios to iron sufficiency at 24 h, and compared each ratio across each of the three time points. We observed that the resulting ratio was significantly higher in iron deficiency after 48 h and significantly smaller in iron deficiency after 72 h of treatment (Fig. 1e). These data suggest that the progression of enucleation compared to the progression of epidermal cell elongation was promoted by 48 h of iron deficiency exposure but delayed by 72 h of iron deficiency exposure because enucleation is associated with significantly more elongated epidermal cells by 72 h of iron deficiency conditions than under 72 h of iron sufficiency. These data also suggest that the regulatory mechanisms underlying SE differentiation with respect to development dynamically change over time under iron deficiency treatment.
Next, we assessed whether iron deficiency delays initiation of SE cell wall thickening, another major characteristic of SE differentiation14. In WT plants, we manually measured the Ratio-S/Ep. In Ratio-S/Ep, the variable “S” is the distance of the first SE from the QC and “Ep” is the vertical length of the Epidermal cell within the same cross-sectional position as the first SE (Fig. 1f, g). The first differentiating SE is identified by the earliest phloem cell with thick cell walls and a narrowed, elongated shape13,26. Each “S” and “Ep” distance was measured to calculate Ratio-S/Ep in each root. At 72 h of iron deficiency conditions, the Ratio-S/Ep was statistically smaller than that under iron sufficiency conditions, while not significantly different under 24 h or 48 h of iron deficiency conditions (Fig. 1h). This suggests that SE cell wall thickening was delayed relative to overall root differentiation by 72 h of iron deficiency conditions. Overall, these data suggest that both enucleation and cell wall thickening are delayed by at least 72 h of iron deficiency.
Iron deficiency changes the regulatory transcriptional relationships associated with SE differentiation
Since our data suggest that iron deficiency delays the progression of enucleation and the initiation of cell wall thickening in differentiating SEs, we investigated the regulatory mechanisms underpinning this delay. We first selected three marker lines as proxies that represent three broad stages of SE differentiation: “early” phloem initials (pPEAR1::dBOX-YFP), “middle” transit-amplifying protophloem (pPEAR1-erGFP), and “late” differentiating SEs (pC2-YFP)8,12. We then grew each marker line (Supplementary Fig. 6a) on iron sufficiency media for 4 days and then transferred them to iron sufficiency or iron deficiency media for 24 h. At 24 h, we performed Fluorescence-Activated Cell Sorting (FACS) in each marker line to isolate cells at each major stage of SE differentiation and iron condition. To decipher the initial regulatory factors underlying the temporal progression of phloem development, we performed RNA-Seq analysis and characterized changes in the phloem transcriptome in the isolated cells. Between iron sufficiency and iron deficiency conditions, 8058 genes, of which 595 were transcription factors, were differentially expressed (q < 0.01, FC > 2) (Supplementary Table 1). We also confirmed the induction of seven genes that regulate phloem differentiation (Supplementary Table 2). These genes include BRX, C2, sAPL, NEN1, NEN4, CaLS7, and APL, all of which are involved in key aspects of phloem differentiation, including the regulation and timing of differentiation, sieve pore formation and gating, and enucleation13,14,16,27. As expected, expression of these genes was enriched in the “late” stage of differentiation under iron sufficiency conditions corresponding to actively differentiating SEs. However, these genes were not enriched in the “late” stage of differentiation under iron deficiency conditions (Fig. 2a, boxed in Fig. 2b). Our observation suggests that iron deficiency may cause a delay in their expression or enrichment in the “late” differentiation stage such that their expression may be shifted beyond the “late” differentiation stage marker used for transcriptomics analysis. Such a transcriptional delay could be the causative mechanism for delayed SE differentiation. Additionally, we confirmed the induction of 83 differentially expressed genes (DEGs, 14 of which are TFs) involved in iron homeostasis in the iron deficiency dataset (Supplementary Table 3). These genes include FIT, BTS and many of their targets, all of which are significant regulators of iron deficiency responses and homeostasis28. 24 iron deficiency-responsive genes, including FIT, IRT1, and FRO2, are enriched (Supplementary Fig. 6b) in one or more SE differentiation stages. Moreover, five differentially expressed phloem-regulating genes, NEN4, APL, C2, CaLS7, and NEN1, (Supplementary Table 2, Fig. 2a) are enriched in one or more SE differentiation stages. Overall, we have accumulated data showing that iron deficiency responses can be upregulated in the phloem.
a Heat map of expression of phloem differentiation-related genes from RNA-Seq data. All genes have reduced expression in all markers under iron deficiency. b Heat map of genes selected by the modified Shannon entropy approach in the isolated cells of each developmental stage and condition. The highlighted section includes phloem differentiation-related genes. c, d Relationships between the nodes of the c iron sufficiency and d iron deficiency Gene Regulatory Networks (GRNs) inferred from developmentally-shifted genes using a Dynamic Bayesian Model. Red arrows show an inferred positive regulatory relationship, blue flat arrows indicate an inferred negative regulatory relationship. Arrows with a slash-dot pattern are edges that are the same in directionality between each network. Arrows with a heavier weight are edges with a different directionality in the iron deficiency network. Yellow nodes are similar between both networks, blue nodes are unique to the iron sufficiency network, and orange nodes are unique to the iron deficiency network. DOF1.5 is emphasized by a thick line weight. e Tabular output from the Node Analyzer application showing each node and its corresponding weight and impact (see Methods) in the e iron sufficiency and f iron deficiency network. +Fe represents iron sufficiency, while −Fe represents iron deficiency.
To identify genes enriched at specific stages of phloem development in iron sufficiency or iron deficiency conditions, we performed a modified Shannon Entropy (MSE) analysis to identify the subset of genes that were enriched in a different set of differentiation stages under iron deficiency conditions than under iron sufficiency conditions29,30,31. The MSE allows for the unbiased selection of genes with enriched expression in a particular developmental stage. 1311 genes with a high entropy were selected and then assigned scores of: enriched (assigned a value of 1), non-enriched (0), or under-enriched (−1), for each developmental stage and iron condition29,30,31 (Supplementary Table 4). To identify genes that showed a different developmental expression under the two iron conditions, we specifically selected high-entropy genes that showed a delayed or accelerated enrichment during phloem development (Fig. 2b, Supplementary Table 5). Interestingly, 24 genes significant to iron deficiency responses or iron homeostasis, including FIT, IRT1, and FRO2, are enriched (Supplementary Table 6, Supplementary Fig. 6b) in one or more SE differentiation stages while five genes, NEN4, APL, C2, CaLS7, and NEN1, (Supplementary Table 7, Supplementary Fig. 6c) are enriched in one or more SE differentiation stages. Thus, we have accumulated data suggesting that the regulation of either iron deficiency responses or SE differentiation is enriched in cells associated with SE cell differentiation.
To unravel the regulatory mechanisms regulating the course of phloem differentiation under iron sufficiency or iron deficiency conditions, we used a Dynamic Bayesian Model to infer causal relationships among the set of genes enriched in a different developmental stage under each of two iron conditions and visualized the two inferred networks using Cytoscape 3.8.0 (Fig. 2c, d, Supplementary Figs. 7–8)32,33. Interestingly, the relationships between the master regulators, or genes with the most network connections, of either network have major differences, which further suggests that iron deficiency changes the regulation of SE differentiation. Specifically, the iron sufficiency network (Fig. 2c) has its distinct set of master regulators: DAG1, UNE12, UK2, and FIT, while the iron deficiency (Fig. 2d) network has its distinct set of master regulators: HHO1, CCA1, and WRKY21. The master regulators in common are UK1, DOF1.5, and MYB63. To determine which master regulators of either network have the largest impact on network stability when perturbed, we quantified node importance using Node Analyzer34, an R-based Shiny application (Fig. 2e, f). UK1 and DOF1.5 showed a high importance in both networks. DAG1 and MYB63 had a high impact specifically in the iron sufficiency GRN, while HHO1 and CCA1 showed a high impact specifically in the iron deficiency GRN (Fig. 2e, f). This suggests that iron deficiency significantly changes the key regulators of phloem differentiation. Interestingly, upon visualizing the expression dynamics, we noticed that the enrichment of most nodes included a later differentiation stage under iron deficiency than under iron sufficiency conditions (Supplementary Fig. 6c). This further suggests that iron deficiency influences the spatial enrichment of genes across the major stages of SE differentiation.
DOF1.5 is involved in the positive regulation of SE differentiation
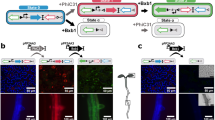
DOF1.5/COG1/PAPL1 (AT1G29160) is the most significant node in both the iron sufficiency and iron deficiency networks, thus we reasoned that it may be a significant factor underlying the change to SE differentiation under iron deficiency (Fig. 2e, f). Transcriptional analysis indicates that DOF1.5 is enriched in the root phloem, phloem pericycle, and the protophloem and metaphloem tissues (Supplementary Fig. 9a)35,. Further analyses have confirmed DOF1.5 expression in the phloem, pericycle, and companion cell tissues and even the epidermis as early as 40 μm from the QC36. pDOF1.5-GFP is also localized in the phloem (Fig. 3a). To evaluate the localization of DOF1.5 expression, we quantified the point in pDOF1.5-GFP roots at which DOF1.5 is expressed by measuring the distance from the QC to the beginning of pDOF1.5-GFP signal at 72 h under iron sufficiency and iron deficiency conditions (Fig. 3a, b). Under both conditions, pDOF1.5-GFP is expressed from 0-400 µm from the QC. To confirm that DOF1.5 is expressed within both the meristematic and elongation zones, we also measured the distance between the QC and the beginning of the elongation zone (Fig. 3b). The meristematic zone ranged between 200 and 500 µm under iron sufficiency conditions and between 200-400 µm under iron deficiency conditions, thus DOF1.5 is expressed across both developmental zones. Moreover, DOF1.5 has recently been shown to be a member of a PXY-mediated root vascular development regulatory network that was inferred to capture genes that regulate the proliferation of vascular tissue, thereby controlling overall plant biomass by enabling the proper allocation of photosynthate from the shoot to the root36,37. Since DOF1.5 is associated with the regulation of vascular and phloem differentiation37 and carbon biomass allocation38, we evaluated how either loss- or gain-of-function of DOF1.5 influences SE differentiation by controlling enucleation, sap translocation, and cell wall thickening and, consequently, root growth in the presence or absence of iron.
a Expression domain of pDOF1.5-GFP at 72 h of either iron deficiency or sufficiency. Representative images include an iron-sufficient root (left) and an iron-deficient root (right). b The distance from the QC to the start of pDOF1.5-GFP signal is quantified, as is the distance from the QC to the elongation zone (EZ). Sample sizes are: n = roots, n = 19 for +Fe and n = 15 for −Fe. For 72 h iron sufficiency (+Fe), the minimum = 226.1, maximum = 361.8, median = 294, 25% percentile = 226.1, 75% percentile = 361.8. For 72 h iron deficiency, the minimum = 220.3, maximum = 276, median = 248.2, 25% percentile = 220.3, 75% percentile = 276. Statistical analyses were performed using a two-way Student’s t test (p < 0.05). c dof1.5-1 pNAC86:HB2-YFP SE Enucleation is quantified by the ratio of elongation index to epidermal cell length at 24 h, 48 h, and 72 h under iron sufficiency or iron deficiency conditions using BVT. Sample sizes are n = cells, n = 303 (24 h iron sufficiency), n = 153 (24 h iron deficiency), n = 269 (48 h iron sufficiency), n = 124 (iron deficiency), n = 309 (72 h iron sufficiency), n = 64 (72 h iron deficiency). The statistical analysis of the ratio between elongation index and epidermal cell length in dof1.5-1 pNAC86:HB2-YFP was determined using a two-way Student’s T Test. *p < 0.05, **p < 0.01, and ***p < 0.001. For 24 h iron sufficiency, the minimum = 0.08912, the maximum = 3.069, median = 1.26, 25% percentile = 0.8163, 75% percentile = 1.875. For 24 h iron deficiency, the minimum = 0.03638, maximum = 2.804, median = 1.005, 25% percentile = 0.6303, 75% percentile = 1.521. For 48 h iron sufficiency, the minimum = 0.00541, maximum = 2.106, median = 0.7742, 25% percentile = 0.6303, 75% percentile = 1.216. For 48 h iron deficiency, the minimum = 0.003790, the maximum = 1.929, median = 0.665, 25% percentile = 0.3519, 75% percentile = 1.007. For 72 h iron sufficiency, the minimum = 0.007856, maximum = 2.592, median = 0.9125, 25% percentile = 0.5601, 75% percentile = 1.381. For 72 h iron deficiency, the minimum = 0.1126, maximum = 1.877, median = 0.7584, 25% = 0.3983, 75% percentile = 1.082. d Fold change in APL, NAC86, and NEN4 expression in WT, dof1.5-1, and dof1.5-2D whole roots at 24 h under iron sufficiency and deficiency conditions. Fold change is relative to the WT +Fe. Standard error bars are included. +Fe represents iron sufficiency, while −Fe represents iron deficiency. n = 3 biological reps of pooled seedlings. Statistical analyses were performed using a two-way Student’s T Test (p < 0.05). e Initiation of cell wall thickening quantified using the Ratio-S/Ep, in WT, dof1.5-1, and dof1.5-2D roots at 72 h under iron sufficiency and iron deficiency conditions. Sample sizes are n = roots, n = 29 (WT iron sufficiency), n = 34 (dof1.5-1 iron sufficiency), n = 18 (dof1.5-2D iron sufficiency), n = 20 (WT -Fe), n = 21 (dof1.5-1 -Fe), n = 16 (dof1.5-2D −Fe). Statistical analysis of plants dyed with esculin was performed using an ANOVA/Tukey HSD. For WT iron sufficiency (+Fe), the minimum = 0.5695, maximum = 1.566, median = 1.02, 25% percentile = 0.7658, 75% percentile = 1.194. For WT iron deficiency (−Fe), the minimum = 0.308, maximum = 1.367, median = 0.6344, 25% percentile = 0.4865, 75% percentile = 0.8643. For dof1.5-1 iron sufficiency, the minimum = 0.5206, maximum = 1.438, median = 0.9425, 25% percentile = 0.7402, 75% percentile = 1.134. For dof1.5-1 iron deficiency, the minimum = 0.6065, maximum = 1.511, median = 0.957, 25% percentile = 0.7591, 75% percentile = 1.173. For dof1.5-2D iron sufficiency, the minimum = 0.6746, maximum = 1.512, median = 1.169, 25% percentile = 0.9609, 75% percentile = 1.225. For dof1.5-2D iron deficiency, the minimum = 0.4081, maximum = 1.267, median = 0.7256, 25% percentile = 0.5547, 75% percentile = 1.095. Statistical test was performed using an ANOVA Tukey HSD test (p < 0.05). f Representative image (left) of a pNAC86:HB2-YFP root at 72 h under iron sufficiency conditions that was stained with esculin (circled in white) 15 min prior to imaging, and Model (right) depicting Distance EN-EF metric. Orange regions = SE nuclei marked by pNAC86:HB2-YFP, “EN” = last SE nucleus expressing pNAC86:HB2-YFP, and dark blue region, marked “EF” = esculin dye front. See Fig. 1g for the extended model legend. g Sap translocation through mature SEs quantified by the Distance EN-EF in pNAC86:HB2-YFP and in dof1.5-1 pNAC86:HB2-YFP at 72 h under iron sufficiency and iron deficiency conditions. Sample sizes are n = roots, n = 20 (pNAC86:HB2-YFP iron sufficiency), n = 20 (pNAC86:HB2-YFP iron deficiency), n = 19 (dof1.5-1 pNAC86:HB2-YFP iron sufficiency), n = 20 (dof1.5-1 pNAC86:HB2-YFP iron deficiency), n = 20 (dof1.5-2D pNAC86::HB2-YFP iron sufficiency), n = 19 (dof1.5-2D pNAC86:HB2-YF iron deficiency). For pNAC86:HB2-YFP iron sufficiency, the minimum = 11.03, maximum = 151.1, median = 80.06, 25% percentile = 53.97, 75% percentile = 130.6. For dof1.5-1 pNAC86:HB2-YFP iron sufficiency, the minimum = 53.74, maximum = 259.7, median = 159.5, 25% percentile = 110.8, 75% percentile = 211. For dof1.5-2D pNAC86::HB2-YFP iron sufficiency, the minimum = 20.69, maximum = 342.7, median = 157.5, 25% percentile = 108.5, 75% percentile = 208. For pNAC86:HB2-YFP iron deficiency, the minimum = 42.14, maximum = 280.6, median = 136, 25% percentile = 80.75, 75% percentile = 189.1. For dof1.5-1 pNAC86:HB2-YFP iron deficiency, the minimum = 62.71, maximum = 295.6, median = 160, 25% percentile = 135.6, 75% percentile = 201.7. For dof1.5-2D pNAC86:HB2-YFP iron deficiency, the minimum = 45.78, maximum = 288.1, median = 120.1, 25% percentile = 74.76, 75% percentile = 210.8. Statistical test was performed using an ANOVA Tukey HSD test (p < 0.05).
We compared the progression of enucleation with the location of developmental zones, as in Fig. 1h, by expressing pNAC86:HB2-YFP in a DOF1.5 loss-of-function background (dof1.5-1 pNAC86:HB2-YFP). We then divided the nuclear elongation index by the epidermal cell length at a similar distance from the QC using the BVT software, normalizing it with respect to the 24 h iron sufficiency data (Fig. 3c). We observed that this metric was significantly lower at 24 h, 48 h, and 72 h in dof1.5-1 pNAC86:HB2-YFP under iron deficiency conditions (Fig. 3c, Supplementary Figs. 10–11). This suggests that the ratio of the elongation index over the epidermal cell length, or progression of enucleation, in a DOF1.5 loss-of-function background is delayed after 24 h, while, in a WT background (Fig. 1e), the progression of enucleation is only delayed at 72 h of iron deficiency conditions. We also compared the progression of enucleation in a dof1.5-1 background (Fig. 3c) with that of the WT background (Fig. 1e). At all time points, the progression of enucleation was lower in a dof1.5-1 background in iron-sufficient conditions. From 48–72 h, the progression of enucleation was also lower under iron-deficient conditions. These data suggest that, in a dof1.5-1 background, enucleation is delayed and that iron deficiency compounds this delay. Thus, DOF1.5 likely positively regulates enucleation under both conditions to a different extent.
We then evaluated whether transcriptional regulation of SE differentiation was influenced by the loss of function of DOF1.5. We performed qRT-PCR on whole roots to identify changes to the expression of APL, NAC86, and NEN4 on WT, dof1.5-1, and dof1.5-2D roots that had undergone 72 h of either iron sufficiency or iron deficiency (Fig. 3d). In response to iron deficiency in WT roots, APL, NAC86, and NEN4 were all downregulated. In dof1.5-1 under iron sufficiency, NAC86 was downregulated, but APL and NEN4 had basal levels of expression. In dof1.5-1 under iron deficiency, APL, NAC86, NEN4 were all downregulated. In dof1.5-2D under iron sufficient conditions, all genes were expressed at basal levels. However, in dof1.5-2D under iron-deficient conditions, APL and NEN4 were upregulated. These data provide further evidence that iron deficiency inhibits or delays SE differentiation and enucleation since APL, NAC86, and NEN4 were all downregulated in WT. These data also support previous studies, which suggest that DOF1.5 is a positive regulator of SE differentiation and enucleation since APL and NEN4 are upregulated in the dof1.5-2D background16. This is also suggested by the downregulation of APL, NAC86, and NEN4 in the dof1.5-1 iron-deficient background. However, in the dof1.5-1 iron-sufficient background, only NAC86 was downregulated. This suggests that there are additional positive regulators of SE differentiation or enucleation14 that may regulate APL and NEN4 under iron sufficiency.
We then assessed whether DOF1.5 has any role in regulating the initiation of SE cell wall thickening by manually measuring the normalized Ratio-S/Ep, as in Fig. 1h, in dof1.5-1 and in dof1.5-2D, roots at 72 h of either iron sufficiency or iron deficiency conditions (Fig. 3e, Supplementary Fig. 13). Under iron sufficiency conditions, the Ratio-S/Ep was similar across all mutants, suggesting no change to the timing of SE cell wall thickening. As expected, iron deficiency resulted in a lower Ratio-S/Ep in WT roots, suggesting that iron deficiency delays SE cell wall thickening. In dof1.5-1 roots, the ratio S/Ep remained similar between both iron deficiency and sufficiency conditions. However, the Ratio S/Ep of dof1.5-1 roots was higher than in WT roots under iron-deficient conditions. This suggests that cell wall thickening was delayed in dof1.5-1 roots. Moreover, in dof1.5-2D roots, the Ratio-S/Ep is lower in iron-deficiency conditions than in iron-sufficiency conditions. Also, the Ratio-S/Ep in iron-deficient dof1.5-2D roots is similar to that of WT iron-deficient roots. These data suggest that DOF1.5 plays a critical role in the iron deficiency-induced delay in cell wall thickening since the delay occurs in dof1.5-2D roots but not in dof1.5-1 roots under iron deficiency.
Since phloem sap unloading requires enucleation20,36, we evaluated how iron deficiency and DOF1.5 both influence sap unloading. DOF1.5 is involved in regulating starch accumulation and allocation to sink tissues36,38. To evaluate whether DOF1.5 influences source to sink translocation of sucrose, we applied esculin, a phloem-mobile fluorescent probe to a cut cotyledon of each marker line, pNAC86:HB2-YFP and dof1.5-1 pNAC86:HB2-YFP, and imaged each root at 72 h of iron sufficiency or iron deficiency conditions (Supplementary Fig. 12). We then measured the distance, or Distance (Distance EN-EF), between the last enucleating cell, “EN”, and the esculin dye front, “EF” (Fig. 3f, g). We found that the Distance EN-EF was statistically longer in the WT background under iron deficiency conditions than in the WT background under iron sufficiency conditions (Fig. 3g). This suggests that iron deficiency delays the transport of phloem sap through the differentiating cells of the SE cell file, likely by delaying enucleation and cell wall thickening (Fig. 3c, e). Moreover, the Distance EN-EF in the dof1.5-1 and dof1.5-2D backgrounds in both iron sufficiency or iron deficiency conditions was also longer than that in the WT background under iron deficiency or sufficiency conditions (Fig. 3f). Since DOF1.5 positively regulates regulators of enucleation under iron deficiency (Fig. 3c), phloem transport to roots is consequently delayed in a dof1.5-1 background (Fig. 3f). However, sap movement is also delayed in a dof1.5-2D background, likely because SE cell wall thickening is not also promoted (Fig. 3d), which makes SE cells less capable of transporting sap that has a high turgor pressure. These data may also suggest a dose-dependence of DOF1.5 on regulating enucleation and sap transport to roots, in which an under- or overexpression of DOF1.5 misregulates the process entirely.
To assess the developmental consequences of knocked-down or enhanced expression of DOF1.5, we measured the root lengths of WT, dof1.5-1, and dof1.5-2D 7d-old seedlings at 72 h after induction of iron deficiency (Fig. 4b, c). We first evaluated two complemented lines in which dof1.5-2D was crossed into a dof1.5-1 background in one line (dof1.5-1 dof1.5-2D) and in which dof1.5-1 was crossed into the dof1.5-2D background in the other line (dof1.5-2D dof1.5-1). We performed qRT-PCR on all lines to quantify their corresponding DOF1.5 expression (Fig. 4a). In dof1.5-1 of both iron-deficient and iron-sufficient conditions, DOF1.5 expression was below WT. In WT roots, DOF1.5 was also downregulated under iron deficiency. In dof1.5-2D roots, DOF1.5 expression is higher than that of WT regardless of iron sufficiency or iron deficiency. In the complemented lines, DOF1.5 expression is similar to WT under iron-sufficient conditions. Moreover, under iron-deficient conditions, dof1.5-1 in a dof1.5-2D results in increased DOF1.5 expression, but activating dof1.5-2D in a dof1.5-1 background doesn’t result in increased DOF1.5 expression. After 72 h of iron deficiency, dof1.5-2D root growth was reduced while dof1.5-1 root growth was increased compared to WT root growth (Fig. 4b–d). This suggests that when DOF1.5 expression is decreased, roots grow longer, but when DOF1.5 expression is enhanced, roots grow shorter under iron deficiency conditions. Moreover, in either complemented line, root growth was longer than WT and dof1.5-2D but shorter than dof1.5-1. This suggests that adding DOF1.5 to a dof1.5-1 background attenuates root growth, but that knocking down DOF1.5 expression in a dof1.5-2D background increases root growth under iron sufficiency and deficiency. Moreover, in dof1.5-1, enucleation and sap transport are delayed (Fig. 3c, g), but cell wall thickening is not delayed (Fig. 3e) by iron deficiency. In dof1.5-2D under iron deficiency conditions, enucleation is promoted (Fig. 3c), but cell wall thickening is delayed as in WT (Fig. 3e), also resulting in delayed sap transport (Fig. 3g). However, in iron sufficient roots, all lines grew similarly to WT except for dof1.5-2D, which grew significantly shorter (Fig. 4b, c). We conclude that, dof1.5-2D may have a greater proportion of enucleated SEs that aren’t able to carry as much sap to roots since cell wall thickening wasn’t promoted at the same rate as enucleation, resulting in shorter roots regardless of iron condition. In iron-deficient dof1.5-1 roots, enucleation was delayed, but cell wall thickening occurred at the rate of WT iron-sufficient roots, allowing sufficient phloem sap unloading in roots to promote root growth.
a FC in DOF1.5 expression in WT, dof1.5-1, dof1.5-2D, dof1.5-2D dof1.5-1, and dof1.5-1 dof1.5-2D iron sufficient and iron-deficient roots at 24 h after induction of iron deficiency. Standard deviation bars are shown. Statistical analyses were performed using a two-way Student’s T Tests (p < 0.05). Unique all caps letters signify statistical differences between +Fe (iron sufficient) samples. Unique lowercase letters signify statistical differences between −Fe (iron-deficient) samples. Asterisks (*) denote significant differences (p < 0.05) of -Fe samples from WT +Fe samples. n = 3 independent biological reps of pooled seedlings for mutants and n = 6 biological reps of pooled seedlings for WT samples. Each biological rep is a pool of seedlings. b Root length of WT, dof1.5-1, dof1.5-2D, dof1.5-2D dof1.5-1, and dof1.5-1 dof1.5-2D after 72 h of iron sufficiency. Sample sizes are n = roots, n = 28 (WT), n = 27 (dof1.5-1), n = 28 (dof1.5-2D), n = 28 (dof1.5-2D dof1.5-1), and n = 28 (dof1.5-1 dof1.5-2D). Root length of WT, dof1.5-1, dof1.5-2D, dof1.5-2D dof1.5-1, and dof1.5-1 dof1.5-2D after 72 h of iron deficiency. Sample sizes are n = 30 (WT), n = 30 (dof1.5-1), n = 30 (dof1.5-2D), n = 29 (dof1.5-2D dof1.5-1), and n = 30 (dof1.5-1 dof1.5-2D). Statistical analysis was performed using an ANOVA/Tukey HSD test (p < 0.05). For WT iron sufficiency, the minimum = 1.423, maximum = 2.556, median = 1.671, 25% percentile = 1.536, 75% percentile = 2.015. For dof1.5-1 iron sufficiency, the minimum = 1.126, maximum = 2.151, median = 1.721, 25% percentile = 1.637, 75% percentile = 1.962. For dof1.5-2D iron sufficiency, the minimum = 0.831, maximum = 1.879, median = 1.408, 25% percentile = 1.236, 75% percentile = 1.574. For dof1.5-2D dof1.5-1 iron sufficiency, the minimum = 1.415, maximum = 2.25, median = 1.822, 25% percentile = 1.66, 75% percentile = 1.975. For dof1.5-1 dof1.5-2D iron sufficiency, the minimum = 1.375, maximum = 2.648, median = 1.725, 25% percentile = 1.612, 75% percentile = 2.019. For WT iron deficiency, the minimum = 0.459, maximum = 1.224, median = 0.899, 25% percentile = 0.6865, 75% percentile = 1.081. For dof1.5-1 iron deficiency, the minimum = 1.094, maximum = 1.847, median = 1.487, 25% percentile = 1.342, 75% percentile = 1.603. For dof1.5-2D iron deficiency, the minimum = 0.197, maximum = 1.222, median = 0.6185, 25% percentile = 0.5065, 75% percentile = 0.7283. For dof1.5-2D dof1.5-1 iron deficiency, the minimum = 0.325, maximum = 1.561, median = 1.171, 25% percentile = 1.019, 75% percentile = 1.286. For dof1.5-1 dof1.5-2D iron deficiency, the minimum = 0.523, maximum = 1.624, median = 1.156, 25% percentile = 0.908, 75% percentile = 1.325. c Representative images of root and shoot phenotypes in WT, dof1.5-1, and dof1.5-2D iron sufficiency at 72 h after induction of iron deficiency. Scale bar = 1 mm. +Fe represents iron sufficiency. Phenotypes are consistent across all replicates. d Representative images of root and shoot phenotypes in WT, dof1.5-1, and dof1.5-2D iron sufficiency at 72 h after induction of iron deficiency. Scale bar = 1 mm. -Fe represents iron deficiency. Phenotypes are consistent across all replicates.
Discussion
Cellular differentiation is a complex process that must be responsive to developmental and environmental cues to optimize overall plant viability. Much work has been done to characterize the regulatory mechanisms of tissue differentiation, such as hormonal, peptide signaling, molecular, and genetic mechanisms1,3,9,10,12,37,38,39,40,41,42. However, less is known about how abiotic stress impacts the regulatory mechanisms involved in tissue differentiation. Here, we used BioVision Tracker (BVT) to capture subtle spatiotemporal changes to the shape of nuclei in SE cells undergoing enucleation. Although other tracking software can segment and quantify characteristics of nuclear architecture, BVT is particularly useful in quantifying and mapping nuclear characteristics with respect to their position in roots and in filtering nuclei outside of the region of interest, such as companion cells, with a high throughput. Thus, BVT enabled us to capture how iron deficiency resulted in delayed SE enucleation. This tool may also have broader applicability in capturing spatial dynamics associated with subtle changes in nuclear morphology using nuclear-localized fluorescent markers in future studies.
DOF1.5 is found in both iron sufficiency and iron deficiency networks inferred in this study. DOF1.5 is involved in SE differentiation and sucrose mobilization (Fig. 3). DOF1.5/PAPL1 is expressed in early phloem cells as close as 40 μm from the QC as well as in the epidermis36. In studies of triple mutants of PAPL genes, including DOF1.5, seedling roots grew poorly without exogenous sucrose supplementation, suggesting a misregulation of sucrose translocation to roots36. Previous studies have also shown that DOF1.5 is heavily associated with phytochrome and gibberellin signaling, which is also associated with establishing the cell maturation zone43,44. DOF1.5 also has additional roles in regulating seed, seedling, and plant development as well, including: upregulating peroxisome- and suberin-mediated seed coat permeability44,45, downregulating phytochrome-mediated light perception43, enhancing photosynthetic capacity38, and promoting leaf starch accumulation38. Moreover, in a network inferring the regulators of vascular proliferation from cambial tissue, DOF1.5 was identified as an interactor with three nodes, PHB, PHV, and REV, of the network that had the highest degree of in-degree connectivity46. This may also suggest that DOF1.5 action is also related to overall vascular differentiation. With respect to SE differentiation, DAP-seq data47 suggest that DOF1.5 can bind the promoters of several key genes associated with phloem differentiation, CVP2, BRX, EPM, SAPL, MYR1, and C2 while HCA2 can bind the promoter of DOF1.510,14. A DOF-CLE peptide circuit also maintains phloem cell patterning by modulating expression of PEAR transcription factors that then regulate SE differentiation37. For example, DOF1.1/OBP2 promotes APL-mediated SE differentiation while CLE25 inhibits DOF1.1 expression within the circuit. DOF1.1 also promotes DOF1.5 expression, suggesting that DOF1.5 may also be a positive regulator of SE differentiation within the circuit. Moreover, a gradient of PLETHORA (PLT) transcription factors is also expressed along the protophloem SE file to regulate which cells differentiate12,36,37. In differentiating SE cells, PLT2 is degraded, releasing repression of SE differentiation genes and releasing repression of DOF1.5. This further suggests that DOF1.5 is a positive regulator of SE differentiation. However, DOF1.5 may not regulate each process underlying SE differentiation, such as enucleation and cell wall thickening, in the same way. We found that enucleation was delayed in dof1.5-1 mutants regardless of iron availability and that NAC86 expression was reduced in iron-sufficient conditions. However, iron-deficient dof1.5-1 roots did not have delayed cell wall thickening as WT roots normally would under iron deficiency. Similarly, NAC45/86 knockout mutants do not exhibit changes to cell wall thickening, but their SE cells have inhibited enucleation, suggesting that the two processes are regulated by different regulatory mechanisms16,19,37. Whereas it is logical that a nutrient-limiting condition like iron deficiency would slow metabolism and inhibit developmental processes such as SE differentiation, our data indicate that DOF1.5 positively regulates SE enucleation while repressing cell wall thickening that occurs in response to iron deficiency.
Iron deficiency is required for erythropoiesis of enucleating red blood cells. Both iron enrichment and tight reactive oxygen species (ROS) homeostasis are necessary to induce enucleation and prevent cell death48. Enucleating SEs likely have similar pressures guiding enucleation since iron deficiency fundamentally alters the redox state of each cell and could ultimately slow the progression of enucleation; however, future studies will confirm this21,48,49. In addition to delayed SE enucleation, iron deficiency also delays the initiation of cell wall thickening and reduces sap transport to roots, likely by changing the network of regulators underlying SE differentiation. When we inferred GRN associated with SE differentiation under iron sufficiency and iron deficiency conditions and evaluated the importance to network stability of each node using Node Analyzer, we found that iron deficiency fundamentally changed the transcriptional regulation associated with SE differentiation. This has also provided an extensive network of genes to probe in future studies that will help explain how extrinsic conditions are linked to controlling vascular development. For example, we found that the master transcriptional regulator of iron deficiency responses, FIT, is a prominent hub in the iron sufficiency network. While other important iron homeostasis genes such as PYE and BTS have previously been shown to be transcribed in internal root tissues under iron deficiency50,51,52, we confirm here that FIT is transcribed in SE cells under iron sufficiency even though its main function is served in epidermal tissues under iron deficiency53,54 (Supplementary Fig. 6b). The potential role of FIT within the phloem requires further study51,55. HHO1 is a key network hub found only in the iron deficiency GRN. HHO1, in concert with HRS1, coordinates sensing of nitrate and phosphorus levels at the root tip and, thus, may be a significant regulator underpinning the iron deficiency-induced change of the regulation of SE differentiation and, consequently, root elongation56,57. Moreover, increased NH4+ levels result in an accumulation of iron deposits in the phloem that increase ROS levels and, consequently, reduce root elongation55. Since root responses to iron, nitrogen, and phosphorus cycles are linked, regulators such as HHO1 may influence phloem differentiation in response to deficiencies in one or more of these nutrient levels. DOF3.7/DAG1 is a DOF1.5/COG1/PAPL1 homolog and is a master regulator of embryonic vascular divisions under normal growing conditions46,58,59. DAG1, DOF1.5, and other DOF transcription factors also regulate embryonic differentiation, seedling development, hypocotyl elongation, and vascular differentiation43,46,58,59,60,61. Thus, it is tempting to speculate that DOF1.5 may act in concert with DAG1 and other DOF regulators in the seedling stage to regulate SE differentiation, particularly under iron sufficiency. Further studies would confirm if the interaction between DOF1.5 and either DAG1 or HHO1 regulates the differences in SE differentiation under iron sufficiency and iron deficiency, respectively.
Both DOF1.5 overexpression and knockout resulted in delayed sap transport but modulated root growth differentially36. DOF1.5 has been implicated in maintaining root sucrose nutrition, which is mediated by sap unloading into roots, and promoting starch accumulation in rosette leaves in triple mutants36. While previous studies have implicated DOF1.5 in various processes, our work presents the first characterization of single mutant phenotypes. Notably, the emergence of this phenotype under iron-deficient conditions suggests a distinct and previously unrecognized role for DOF1.5 in the seedling establishment response to abiotic stress. The Münch hypothesis suggests turgor pressure differentials drive sap movement from source to sink tissues20,62,63. SE cell wall thickening may help accommodate the high turgor pressures caused by bulk sap movement64. In root SEs, cell wall thickening and enucleation typically occur in the root transition zone in overlapping regions directly under the phloem unloading zone, possibly so that as both enucleation and cell wall thickening progress, the unloading zone further facilitates sucrose unloading closer to the root apical meristem20,62. In dof1.5-2D iron-deficient roots, cell wall thickening timing was neither delayed nor promoted, while NEN4, a regulator of enucleation, was promoted. However, sap transport was still delayed, and root growth was inhibited despite iron deficiency or sufficiency. This suggests that there may be some relationship to cell wall thickening and efficient sap mobilization. Similarly, NAC45/86 knockout mutants impair phloem transport and unloading into the root, likely due to incomplete enucleation despite normal cell wall thickening16. In dof1.5-2D leaves, starch accumulates, further suggesting that DOF1.5 overexpression disrupts sucrose mobilization to roots38. Also, ectopic NEN4 expression results in cell death12, which could suggest that dof1.5-2D iron-deficient roots, which had high levels of NEN4 expression, grew poorly under iron deficiency and iron sufficiency due to excessive initiation of enucleation that may have resulted in cell death rather than SE differentiation. Conversely, in dof1.5-1 iron-deficient roots, cell wall thickening was promoted while enucleation was delayed. As a result, sap transport was delayed despite increased root length compared to WT. This suggests that starch degradation and mobilization to roots may have been promoted by the loss of function of DOF1.5 and promoted cell wall thickening, but that sap transport was still disrupted, likely due to delayed enucleation. These findings suggest that loss of DOF1.5 function may enhance starch mobilization to roots, thereby promoting cell wall thickening, which may partially compensate for impaired sap transport resulting from delayed enucleation and consequently result in increased root elongation compared to WT. In iron sufficient dof1.5-1 roots, enucleation and NAC86 expression were delayed, but cell wall thickening was not different from WT. While sap transport was delayed, dof1.5-1 iron-sufficient roots grew similarly to WT. This also suggests that although enucleation and sap transport were delayed, roots were able to acquire enough sugars to grow at a normal rate, possibly due to normal cell wall thickening. As plants transition from iron sufficiency to deficiency, the presence of a specific range of DOF1.5 may be essential for the precise timing of both enucleation and cell wall thickening that enables SE differentiation and, consequently, sap release into SEs for proper root elongation.
Notably, dof1.5-1 typically does not result in root growth phenotypes36. However, here we show that iron-deficient dof1.5-1 mutants exhibit increased root growth, further suggesting that iron deficiency triggers a change in both SE differentiation and root development that may be mediated by DOF1.5. Further observations, such as agravitropism (Supplementary Fig. 9b) in dof1.5-2D iron sufficient roots and delayed flowering (Supplementary Fig. 9c), suggest that DOF1.5 is a critical DOF in proper sensing and response to environmental conditions. Thus, the observed root length phenotypes suggest that DOF1.5 also acts at a nexus between root phloem-mediated transport38 of either sugars and systemic iron deficiency stress signals.
Methods
Media and seedling preparation
All seeds were wet-sterilized by rotation in 70% ethanol for 5 min followed by a rotation in 30% bleach and 0.02% Triton X-100 for 15 min. Seeds were then rinsed 3 times in sterile distilled water and then stratified for at least 2 days in the dark at 4 °C. The seeds were then sown on iron sufficiency media and then transferred to either iron sufficiency or iron deficiency media. Iron sufficiency plates consisted of standard 1% Sucrose, Murashige and Skoog (MS) medium without Iron, containing 0.1 mM FeEDTA substituted for iron sulfate. Iron deficiency plates consisted of MS agar media containing 1% Sucrose, standard MS medium containing 300 μM Ferrozine as an iron chelator. All plants were grown at 22 °C under a 16h-light/8h-dark period in environmentally-controlled plant growth chambers (Percival Scientific). dof1.5-1 was generated by T-DNA insertion (SALK_045465C). dof1.5-1 pNAC86:HB2-YFP was generated by crossing, and analysis was performed in the F3 generation. dof1-5-1 and dof1.5-2D seeds were provided by Eduardo Bueso. pPEAR1::dBOX-YFP, pPEAR1-erGFP, and pC2-YFP were provided by Antia Villalon.
Confocal microscopy imaging
Confocal microscopy was performed using a Zeiss LSM 710. Cell walls were stained using 10 μM Propidium Iodide (PI) in water. Each WT, marker, or mutant line was excited using a 561 nm laser to visualize the PI stain and the collection was detected at 415–735 nm. Each marker line was excited using a 561 nm and 488 nm laser to visualize the PI stain and YFP fluorophore, respectively. Collection was detected at 620–719 and 490–556 nm, respectively. pDOF1.5-GFP was visualized using a 488 nm and 561 nm laser, and detection occurred at 493–540 and 621–719 nm, respectively. For the esculin assay, each marker was labeled with esculin (Millipore Sigma, E8250), then excited using a 561, 488, and 405 nm laser to visualize PI, YFP, and esculin, respectively. Collection was detected at 621–689, 523–572, and 420–480 nm, respectively. In each line, 3–4 biological replicates were imaged. The sap transport assay was performed using 10 μM esculin in water. To bypass the cuticle, a cotyledon of each seedling was cut at least to the midrib. Then, esculin was applied to the wound. Confocal imaging was performed no earlier than 15 minutes after application.
Root growth assay
Seedlings were sown on sterilized nylon mesh sheets placed on 1% sucrose, standard 1x MS iron sufficiency or iron deficiency plates. At day 4 after sowing, plants were transferred to either iron sufficiency or iron deficiency plates. Root lengths between day 4 and 7 were measured using ImageJ (National Institutes of Health; http://rsb.info.nih.gov/ij/)65. 3 biological replicates were measured. Statistical analysis was performed using an ANOVA/Tukey HSD Test using JMP Pro 16 between all iron sufficiency and iron deficiency samples (p < 0.05).
Polymerase chain reaction (PCR)
The homozygous genotype of the dof1.5-1 lines was confirmed using PCR. DNA was amplified using Apex 2.0X Taq RED Master Mix (Apex BioResearch Products) and 5 µM of the right and left primers associated with DOF1.5 and the left border primer associated with the T-DNA insert. The associated primers are listed in Supplementary Table 8.
Quantitative reverse transcriptase PCR (qRT-PCR)
RNA was extracted from pools of whole roots using an RNeasy Plant Mini Kit (Qiagen), and RNA quality was validated using a Nanodrop (ThermoScientific). Three biological replicates of pooled seedlings were collected. From each sample, 1 µg of RNA was then suspended in DEPC-treated water (Invitrogen) for use in cDNA conversion. Then, cDNA conversion was performed using the SuperScript III First-Strand Synthesis System (ThermoFisher Scientific). Next, qPCR was performed using iTaq Universal SYBR Green Supermix (Bio-Rad), the converted cDNA, and the primers associated with amplifying each target sequence. β-TUB was amplified as the housekeeping gene. The associated primers are listed in Supplementary Table 8. All qRT-PCR reactions were run using a StepOnePlus Real-Time PCR System cycler (Applied Biosystems). Finally, fold change (FC) was calculated by dividing the mean 2(−ΔCT) of each sample by the mean 2(−ΔCT) of the WT iron sufficiency sample. ΔCT was calculated by subtracting the CT value of the housekeeping gene from that of the gene of interest. Statistical significance was determined by a Student’s T Test (p < 0.05).
Fluorescence-activated cell sorting and RNA-sequencing
Seeds from the marker lines indicated for FACS were sown on iron-sufficient media, then transferred to fresh iron-sufficient or deficient media after 4 days. Roots were then harvested on day 5, digested and subjected to FACS as described previously66. Cell sorting was then performed by the Flow Cytometry and Cell Sorting Facility (NCSU), and 3–5 biological replicates were collected. RNA was extracted from FACS-sorted protoplasts corresponding to each marker line and iron condition using the RNeasy Micro Kit (Qiagen). RNA quality was then validated using a BioAnalyzer Tapestation (Genomic Sciences Laboratory, NCSU). The cDNA library was then prepared using the Clontech SMARTer Ultra Low Input RNA kit v3. Finally, genes were sequenced using Illumina Sequencing. Sequencing data was cleaned using fastq-mcf and mapped to the Arabidopsis reference genome (TAIR 10) using Tophat. Then, featureCounts was used to determine each raw read count. Finally, edgeR was used to determine the 456 DEGs (q < 0.01).
Shannon entropy
Previous methods were adapted for use in this study to identify which genes were enriched in a different set of one/more differentiation stages under iron deficiency than under iron sufficiency. In this study, enriched genes had an entropy30,48 (H) < 2. Output tables used in this study are provided in Supplementary Tables 4 and 5.
Network inference
All networks were inferred with GENIST using a Dynamic Bayesian Model32, and clustering prior to network inference was not performed. Output tables from GENIST are provided in Supplementary Tables 9 and 10. Output networks are provided in Supplementary Figs. 7 and 8. Nodes and edges (Fig. 2c, d) were visualized using Cytoscape 3.8.0.
BVT image analysis
The BVT software was used to analyze all images according to the protocol in Buckner et al.22. This software was used to segment and map individual pNAC86:HB2-YFP and dof1.5-1 pNAC86::HB2-YFP regions in roots. This information was then expanded upon with the creation of the elongation index and epidermal cell length measurements:
Elongation index
For every segmented NAC86::YFP instance detected by BVT, a 3D subimage was extracted that completely encapsulated all portions of that segmented volume. A maximum projection of this image was calculated and defined as Ii (x,y) where i is an identifier for a pNAC86:HB2-YFP region.
Then \({I}_{i}\left(x,y\right)\) was normalized using the following equation.
Finally, the elongation index was calculated by rotating the image from −45° to 45° while taking the covariance \({C}_{{xy}}\) of \({I}_{i}^{{\prime} }(x,y)\). The covariance was then plotted with respect to the angle, squared, and integrated. Regions that were more elongated created larger fluctuations in the covariance plot, meaning the integrated value was larger for more elongated regions.
Enucleation classifier
We manually classified 8,023 nuclei expressing pNAC86:HB2-YFP as enucleating or non-enucleating. Eight features were gathered for each nucleus (see list below) to train a neural network with a single 20-node hidden layer. Training data and testing data were split into 5617 and 2406, respectively.
Feature description
-
1.
The elongation index, as described previously
-
2.
Radial distance from the roots longitudinal axis in microns
-
3.
Distance to the closest expressing nucleus (nearest neighbor) in microns
-
4.
Number of expressing nuclei within the same cell file
-
5.
Number of expressing nuclei closer to the QC than the current nucleus
-
6.
Volume of the expressing nucleus in microns^3
-
7.
Distance to the surface of the root in microns
-
8.
Longitudinal distance from the QC in microns
Measurement of epidermal cell length
We used a local image symmetry methodology to detect cell files and cell lengths within the root24,25. We expanded on this method by introducing a machine learning approach that uses features of cell files to classify the cell files according to cell type (epidermal cells vs. non-epidermal cells). The features collected from these cell files were the standard deviation of the location of the file in the radial direction, the average distance of the cell file from the outer edge of the root, and the length of the cell file in the axial direction. These features were collected from 714 cell files and trained on a K-Nearest Neighbors classification model and used to detect epidermal cell files. The files were classified with this model, and false detections were corrected by the user.
Once the epidermal cell lengths and locations were detected, we integrated the capabilities of the BVT software to map the longitudinal distance of each epidermal cell from the QC. For each cell file, an exponential regression model was calculated to model the average increase in epidermal cell length with respect to distance from the QC. An exponential regression model was chosen since the epidermal cell lengths followed exponential growth with respect to distance from the QC in the area of the root we were analyzing. Supplementary Fig. 14 shows an array of examples in which an exponential regression model fits these data.
Elongation index vs. epidermal cell length
The exponential regression model of each cell file was used as a mapping tool to compare the elongation index of enucleating nuclei to the corresponding epidermal cell size. For each enucleating nucleus, the elongation index and distance from the QC were collected. That distance from the QC was plugged into the exponential regression model to estimate a corresponding epidermal cell length. The elongation index and epidermal cell length were then collected for each enucleating nucleus and plotted in 2-dimensional space across all conditions (24 h iron sufficiency, 24 h iron deficiency, 48 h iron sufficiency, 48 h iron deficiency, 72 h iron sufficiency, and 72 h iron deficiency).
Modified Shannon entropy
MSE was used to select differentially expressed genes (DEGs). Importantly, MSE offers an unbiased, combinatorial approach to DEG selection by replacing user-selected FC and false discovery rate threshold with an entropy cutoff and subsequent outlier calculation. First, an entropy score is calculated for each gene based on its expression values, which can be interpreted as a rating of how volatile or unpredictable the values are29. The entropy score for a gene is given by the following equation:
where t is a given sample, N is the total number of samples, and pt is the relative expression of a given gene in t.
Next, for each gene meeting entropy selection criteria, outlier and non-outlier scores are assigned to label samples58. An outlier score of 1 for a particular sample indicates that the gene is enriched when compared to its entire expression profile. A non-outlier score of 0 indicates “normal” expression and corresponds to where the gene is not differentially expressed. Each possible combination of outlier and non-outlier scores for the samples of a given gene is generated and evaluated. The statistic for evaluating outlier assignments is given by the following equation:
where s is the number of outlier candidates, n is the number of non-outlier candidates, and is the standard deviation of the expression values of the non-outlier candidates. The combination of scores that results in the lowest U value is accepted. Outlier scoring enables high-resolution combinatorial DEG selection via the selection of genes that are labeled as outliers or non-outliers in specific user-chosen samples.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sufficient Raw RNA-sequencing data for pooled (bulk-sorted) protophloem SE cells are deposited at the Gene Expression Omnibus of the National Center for Biotechnology Information (Submission GSE142259). All iron-deficient RAW RNA-sequencing data for pooled (bulk-sorted) protophloem SE cells are deposited at the EMBL-EBI Annotare research center (E-MTAB-15703). The source data for Figs. 1d, e, h, 2a, 3b–e, g and 4a, b and Supplementary Fig. 6b–c are provided with this paper. Source data are provided with this paper.
References
Iyer-Pascuzzi, A. S. et al. Cell identity regulators link development and stress responses in the arabidopsis root. Dev, Cell 21, 770–782 (2011).
Bloch, D., Puli, M. R., Mosquna, A. & Yalovsky, S. Abiotic stress modulates root patterning via ABA-regulated microRNA expression in the endodermis initials. Development 146, dev177097 (2019).
Dinneny, J. R. et al. Cell identity mediates the response of arabidopsis roots to abiotic stress. Science 320, 942–945 (2008).
Offler, C. E., McCurdy, D. W., Patrick, J. W. & Talbot, M. J. Transfer cells: cells specialized for a special purpose. Annu. Rev. Plant Biol. 54, 431–454 (2003).
Henriques, R. et al. Knock-out of Arabidopsis metal transporter gene IRT1 results in iron deficiency accompanied by cell differentiation defects. Plant Mol. Biol. 50, 597–597 (2002).
Feng, H. et al. Light-regulated, tissue-specific, and cell differentiation-specific expression of the arabidopsis Fe(III)-chelate reductase Gene AtFRO6. Plant Physiol. 140, 1345–1354 (2006).
Bruex, A. et al. A gene regulatory network for root epidermis cell differentiation in arabidopsis. PLoS Genet. 8, e1002446 (2012).
Drapek, C., Sparks, E. E. & Benfey, P. N. Uncovering gene regulatory networks controlling plant cell differentiation. Trends Genet. 33, 529–539 (2017).
López-Salmerón, V., Cho, H., Tonn, N. & Greb, T. The phloem as a mediator of plant growth plasticity. Curr. Biol. 29, R173–R181 (2019).
Blob, B., Heo, J. & Helariutta, Y. Phloem differentiation: an integrative model for cell specification. J. Plant Res. 131, 31–36 (2018).
Dolan, L. et al. Cellular organisation of the Arabidopsis thaliana root. Development 119, 71–84 (1993).
Roszak, P. et al. Cell-by-cell dissection of phloem development links a maturation gradient to cell specialization. Science 374, eaba5531 (2021).
Gujas, B. et al. A reservoir of pluripotent phloem cells safeguards the linear developmental trajectory of protophloem sieve elements. Curr. Biol. 30, 755–766.e4 (2020).
Rodriguez-Villalon, A. Wiring a plant: genetic networks for phloem formation in Arabidopsis thaliana roots. N. Phytol. 210, 45–50 (2016).
Esau, K. Development and structure of the phloem tissue. Bot. Rev. 5, 373–432 (1939).
Furuta, K. M. et al. Arabidopsis NAC45/86 direct sieve element morphogenesis culminating in enucleation. Science 345, 933–937 (2014).
Torode, T. A. et al. Branched pectic galactan in phloem-sieve-element cell walls: implications for cell mechanics. Plant Physiol. 176, 1547–1558 (2018).
Esau, K. & Cheadle, V. I. Wall thickening in sieve elements. Proc. Natl. Acad. Sci. 44, 546–553 (1958).
Heo, J., Blob, B. & Helariutta, Y. Differentiation of conductive cells: a matter of life and death. Curr. Opin. Plant Biol. 35, 23–29 (2017).
Ross-Elliott, T. J. et al. Phloem unloading in Arabidopsis roots is convective and regulated by the phloem-pole pericycle. eLife 6, e24125 (2017).
Byrnes, C. et al. Iron dose-dependent differentiation and enucleation of human erythroblasts in serum-free medium: iron-dependent erythroblast differentiation. J. Tissue Eng. Regen. Med. 10, E84–E89 (2016).
Buckner, E. et al. BioVision tracker: a semi-automated image analysis software for spatiotemporal gene expression tracking in Arabidopsis thaliana. Methods Cell Biol. 160, 419–436 (2020).
Ruiz Sola, M. A. et al. OCTOPUS-LIKE 2, a novel player in Arabidopsis root and vascular development, reveals a key role for OCTOPUS family genes in root metaphloem sieve tube differentiation. N. Phytol. 216, 1191–1204 (2017).
Kovesi, P. Symmetry and asymmetry from local phase. https://www.peterkovesi.com/papers/ai97.pdf (1997).
Quelhas, P. et al. Arabidopsis thaliana automatic cell file detection and cell length estimation. In Image Analysis and Recognition (Springer, Burnaby, BC, Canada, 2011).
Truernit, E. et al. High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of phloem development and structure in Arabidopsis. Plant Cell 20, 1494–1503 (2008).
Bonke, M., Thitamadee, S., Mähönen, A. P., Hauser, M.-T. & Helariutta, Y. APL regulates vascular tissue identity in Arabidopsis. Nature 426, 181–186 (2003).
Colangelo, E. P. & Guerinot, M. L. The essential basic helix-loop-helix protein FIT1 is required for the iron deficiency response. Plant Cell 16, 3400–3412 (2004).
Xiong, M., Heruth, D. P., Zhang, L. Q. & Ye, S. Q. Identification of lung-specific genes by meta-analysis of multiple tissue RNA-seq data. FEBS Open Bio 6, 774–781 (2016).
Kadota, K., Ye, J., Nakai, Y., Terada, T. & Shimizu, K. ROKU: a novel method for identification of tissue-specific genes. BMC Bioinformatics 7, 294 (2006).
Schug, J. et al. Promoter features related to tissue specificity as measured by Shannon entropy. Genome Biol. 6, R33 (2005).
de Luis Balaguer, M. A. et al. Predicting gene regulatory networks by combining spatial and temporal gene expression data in Arabidopsis root stem cells. Proc. Natl. Acad. Sci. USA 114, E7632–E7640 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Van den Broeck, L. et al. A hybrid model connecting regulatory interactions with stem cell divisions in the root. Quant. Plant Biol. 2, e2 (2021).
Li, S., Yamada, M., Han, X., Ohler, U. & Benfey, P. N. High-resolution expression map of the arabidopsis root reveals alternative splicing and lincRNA regulation. Dev.Cell 39, 508–522 (2016).
Otero, S. et al. A root phloem pole cell atlas reveals common transcriptional states in protophloem-adjacent cells. Nat. Plants 8, 954–970 (2022).
Qian, P. et al. A Dof-CLE circuit controls phloem organization. Nat. Plants 8, 817–827 (2022).
Wei, Z. et al. The Dof transcription factor COG1 acts as a key regulator of plant biomass by promoting photosynthesis and starch accumulation. Mol. Plant 16, 1759–1772 (2023).
Marhava, P. et al. A molecular rheostat adjusts auxin flux to promote root protophloem differentiation. Nature 558, 297–300 (2018).
Depuydt, S. et al. Suppression of Arabidopsis protophloem differentiation and root meristem growth by CLE45 requires the receptor-like kinase BAM3. Proc. Natl. Acad. Sci. 110, 7074–7079 (2013).
Durgaprasad, K. et al. Gradient expression of transcription factor imposes a boundary on organ regeneration potential in plants. Cell Rep. 29, 453–463.e3 (2019).
Miyashima, S. et al. Mobile PEAR transcription factors integrate positional cues to prime cambial growth. Nature 565, 490–494 (2019).
Park, D. H. et al. The Arabidopsis COG1 gene encodes a Dof domain transcription factor and negatively regulates phytochrome signaling. Plant J. 34, 161–171 (2003).
Bueso, E. et al. Arabidopsis COGWHEEL1 links light perception and gibberellins with seed tolerance to deterioration. Plant J. 87, 583–596 (2016).
Renard, J. et al. PRX2 and PRX25, peroxidases regulated by COG1, are involved in seed longevity in Arabidopsis. Plant Cell Environ. 43, 315–326 (2020).
Smit, M. E. et al. A PXY-mediated transcriptional network integrates signaling mechanisms to control vascular development in Arabidopsis. Plant Cell 32, 319–335 (2020).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Taoka, K. et al. The effect of iron overload and chelation on erythroid differentiation. Int. J. Hematol. 95, 149–159 (2012).
Kempe, D. S. et al. Enhanced programmed cell death of iron-deficient erythrocytes. FASEB J. 20, 368–370 (2006).
Long, T. A. et al. The bHLH transcription factor POPEYE regulates response to iron deficiency in Arabidopsis roots. Plant Cell 22, 2219–2236 (2010).
Gayomba, S. R., Zhai, Z., Jung, H. & Vatamaniuk, O. K. Local and systemic signaling of iron status and its interactions with homeostasis of other essential elements. Front. Plant Sci. 6, 716 (2015).
Gao, F., Li, M. & Dubos, C. bHLH121 and clade IVc bHLH transcription factors synergistically function to regulate iron homeostasis in Arabidopsis thaliana. J. Exp. Bot. 75, 2933–2950 (2024).
Jakoby, M., Wang, H.-Y., Reidt, W., Weisshaar, B. & Bauer, P. FRU (BHLH029) is required for induction of iron mobilization genes in Arabidopsis thaliana. FEBS Lett. 577, 528–534 (2004).
Wild, M. et al. Tissue-specific regulation of gibberellin signaling fine-tunes arabidopsis iron-deficiency responses. Dev. Cell 37, 190–200 (2016).
Liu, X. X. et al. Phloem iron remodels root development in response to ammonium as the major nitrogen source. Nat. Commun. 13, 561 (2022).
Medici, A. et al. AtNIGT1/HRS1 integrates nitrate and phosphate signals at the Arabidopsis root tip. Nat. Commun. 6, 6274 (2015).
Liu, H. et al. Overexpressing HRS1 confers hypersensitivity to low phosphate-elicited inhibition of primary root growth in Arabidopsis thaliana. J. Integr. Plant Biol. 51, 382–392 (2009).
Le Hir, R. & Bellini, C. The plant-specific dof transcription factors family: new players involved in vascular system development and functioning in arabidopsis. Front. Plant Sci. 4, 51818 (2013).
Boccaccini, A. et al. Independent and interactive effects of DOF affecting germination 1 (DAG1) and the Della proteins GA insensitive (GAI) and Repressor of ga1-3(RGA) in embryo development and seed germination. BMC Plant Biol. 14, 200 (2014).
Ruta, V. et al. The DOF transcription factors in seed and seedling development. Plants 9, 218 (2020).
Papi, M. et al. Inactivation of the phloem-specific dof zinc finger gene DAG1 affects response to light and integrity of the testa of arabidopsis seeds. Plant Physiol. 128, 411–417 (2002).
Anne, P. & Hardtke, C. S. Phloem function and development — biophysics meets genetics. Curr. Opin. Plant Biol. 43, 22–28 (2018).
Knoblauch, M. et al. Testing the Münch hypothesis of long distance phloem transport in plants. eLife 5, e15341 (2016).
Mokshina, N., Ibragimova, N., Ageeva, M. & Mikshina, P. Changes in the cell wall of flax phloem fibers during gravitropic reaction. Acta Physiol. Plant 46, 93 (2024).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Drapek, C. et al. Minimum requirements for changing and maintaining endodermis cell identity in the Arabidopsis root. Nat. Plants 4, 586–595 (2018).
Acknowledgements
We thank the Flow Cytometry and Cell Sorting Laboratory at North Carolina State University (NCSU) for their assistance with cell sorting. Images in this manuscript were generated using the instruments and services at the Cellular and Molecular Imaging Facility (CMIF) at NCSU. This work was supported by a Southern Regional Education Board (SREB) Doctoral Scholars Program fellowship, an NIH Molecular Biotechnology Training Award (5T32GM133366-02), and an NSF Postdoctoral Research Fellowship in Biology (IOS-2305774) awarded to I.M., Research in the T.A.L. lab was funded by the National Science Foundation and the Biotechnology and Biological Sciences Research Council (BBSRC) (NSF MCB-1517058), T.A.L. is also funded by and the USDA National Institute of Food and Agriculture, Hatch Project (Accession Number 101090). E.D.B. was supported by the GAANN Fellowship in Molecular Biotechnology (grant #P200A160061).
Author information
Authors and Affiliations
Contributions
I.M., C.W., R.S., and T.A.L. conceived and designed the study. E.D.B. conducted the automated BioVision Tracker image analyses. I.M. and D.Sr. conducted and analyzed manual experimental data, D.Se. conducted the Fluorescence-Activated Cell Sorting and RNA-Sequencing. M.A., d.L.B., and J.S. contributed to the GRN inference and to developing the Shannon Entropy approach. A.H. created the scientific diagrams. E.B. provided mutant seed materials and related guidance. I.M. and E.D.B. wrote the manuscript, and all authors contributed to editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
Rosangela Sozzani declares a competing interest with Raleigh BioSciences. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Lenwood Heath, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Madison, I., Buckner, E.D., de Luis Balaguer, M.A. et al. Iron deficiency changes regulatory mechanisms governing sieve element cell differentiation. Nat Commun 16, 10196 (2025). https://doi.org/10.1038/s41467-025-65428-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65428-1