Abstract
Medulloblastoma (MB) is the most prevalent malignant brain tumor in children, exhibiting clinical and genomic heterogeneity. Of the four major subgroups, Group 3 tumors (MYC-MB), display high levels of MYC and metastasis rates. Despite treatment with surgery, radiation, and chemotherapy, patients with Group 3 MB are more likely to develop aggressive recurrent tumors with poor survival. To examine resistance mechanisms in this study, we perform single nuclei multiome analysis of matched primary and recurrent tumors. Therapy resistant Medulloblastoma demonstrates an expanded persistent progenitor population. Additionally, distinct chromatin landscapes link to altered transcription and correspond with metabolic reprogramming. In vivo modeling of radiation resistance exhibits similar chromatin-based metabolic reprogramming focused on wild-type isocitrate dehydrogenase (IDH1) activity. IDH1 inhibition reverses resistance-mediated chromatin changes and enables radiation re-sensitization. Ultimately, these findings demonstrate the efficacy of single-cell multiome analysis in elucidating resistance mechanisms and identifying targetable pathways for MYC-driven medulloblastoma.
Similar content being viewed by others
Introduction
Medulloblastoma (MB) is the prevailing malignant brain tumor in children, accounting for 20% of all pediatric brain tumors1,2. MB is clinically heterogeneous, with varying prognoses depending on the molecular subgroup. Group 3 tumors with Myc amplification exhibit higher rates of metastasis and recurrence3,4,5. Treatments for these high-risk patients include craniospinal irradiation (36 Gy) after surgical resection; however, most patients relapse within 1 year1,6,7. Despite this, radiation remains the primary therapeutic option for high-risk MB patients, culminating in <5% long-term survival1,8,9,10.
Tumor heterogeneity driven by clonal evolution and tumor-initiating cell diversity is thought to contribute to therapy failure11,12,13. Recent single-cell RNA-seq (scRNA-seq) studies have validated the intra- and inter-tumoral heterogeneity of medulloblastoma14,15,16. We previously identified a stem-like population in Group 3 tumors with a gene signature that predicts poor outcomes14. Although, MB molecular subtypes present differing relapse patterns and evolution of chromosomal changes, new genetic events are rare in Group 3 medulloblastoma17.
Cellular plasticity is another major driver of therapy resistance. Cancer cells can alter their lineage by reverting to a stem-like state, evading therapies targeting the original lineage-directed survival programs. A MB disease progression model of relapse highlighted this phenomenon, noting a population of MB stem-like cancer cells averted standard therapies by increasing self-renewal18. Although many genomic aberrations have been associated with lineage plasticity, few tumors exhibit these alterations, suggesting that resistance due to lineage plasticity occurs through alternative mechanisms. Moreover, epigenetic reprogramming impacts cellular lineage decisions and cell state changes that reduce responsiveness to therapy19,20,21,22,23. Given that no current therapies effectively target lineage plasticity-driven resistance in tumors, an urgent need exists to identify key lineage plasticity modifiers as potential therapeutic targets.
In this study, we perform single-cell multiome sequencing of matched primary and recurrent MB to explore the heterogeneous cell populations' response to therapy. Our data identifies cell populations that demonstrate lineage plasticity and chromatin remodeling during resistance to therapy. Further, we determine that chromatin remodeling reprograms the metabolic state, demonstrating the potential for phenotypic targeting of a resistant cell population by disrupting lineage-directed plasticity.
Results
scMultiome-seq profiling of matched relapsed and primary MB tumors
To understand the differences between primary and relapsed tumors we performed scMulti-omic sequencing of patient-matched primary and relapsed MB tumors. Single nuclei were isolated from three pairs of primary and recurrent MB tumor samples, classified as Group 3 and Group 3/4 (Supplementary Data 1). Unsupervised clustering identified 20 clusters with initial cell-type annotations based on differentially expressed genes (DEG; Fig. 1A, Supplementary Data 1). SnATAC and snRNA-seq data were integrated to identify cell states based on transcriptomic and chromatin accessibility changes; clustering retained 20 cell-state identities, which further consolidated into 12 major cell groups (Fig. 1B, C). Cell-state annotation was based on gene expression patterns from early cerebellar development and prior single-cell MB patient analysis24,25,26,27,28,29,30,31. Normal diploid cells were identified by inferCNV32,33 (Supplementary Fig. 1A–C) and cell identification markers for microglia (C1qb), radial glial cells (RGC) (AQP4, SOX2, NES, and AMOT1L), retinal pigment epithelial cells (RPE), neuronal epithelial cells (NEC) (NES, YAP1, FAT4, and OCLN), and Purkinje cells (Fig. 1B, C, Supplementary Data 1). Malignant cells expressing TBR1, NEUROD1, and STMN2 resembled GABAergic-like (GAD1, GAD2, SLC6A1), glutamatergic-like interneurons (CALB1, CALB2) and neurons (SLC17A7, GRIN1) (Fig. 1D, Supplementary Fig. 1D, 2A). We identified malignant cells comprising cycling neural progenitors expressing MKI67, MYC, and TOP2A. Non-cycling neural progenitors expressed genes from the subventricular zone of the Rhombic Lip and early photoreceptor (NRL, CRX, OTX2, and TULP1) and unipolar brush cell (UBC) lineages (PAX6 and EOMES; Fig. 1D, Supplementary Fig. 1E). The proportion of the progenitor_photoreceptor cell state increased in Group 3 recurrences, whereas the proportion of more differentiated neuron-like cells are relatively unchanged (Fig. 1E). Spatial transcriptomic in situ analysis of the same tumor samples corroborated the amplification of the progenitor_photoreceptor cell state (Fig. 1F, Supplementary Fig. 3A).
A Schematic representation of matched primary and recurrent tumor processing for scMultiome. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/5s3qjvg. Joint Uniform Manifold and Projection (UMAPs) were generated using principal component analysis (PCA) dimensional reduction for gene expression and Latent Semantic Indexing (LSI) indexing for DNA accessibility. (Below) UMAP visualization of snRNAseq data from patient primary (n = 3) and recurrent tumor samples (n = 3). Coloring based on clustering. (Above) UMAP visualization of integrated snATAC and snRNA-seq datasets. Color designates primary cells (pink) and recurrent cells (turquoise). Total nuclei count=50,184. B Cell state designation of 20 clusters based on FindClusters. Non-malignant, diploid cells are circled. C Merged cell states of 12 major clusters. D snGene expression marker dot plot from patient primary (n = 3) and recurrent tumor samples (n = 3). Color indicates average expression and dot size refers to the percentage of cells within the cluster expressing the gene. E Cell proportions in 12 major clusters and 20 minor clusters. Colored by cell type. Violin plot of cell proportions of CNP and Progenitor-Photoreceptor in primary and recurrent tumors. F Xenium in situ spatial transcriptomic analysis of patient primary (n = 2) and recurrent (n = 2) Group 3 tumor. Subpopulations segmented based on gene expression signatures from14. A-Mitotic (MKI67, TOP2A, CENPF, red), B-Progenitor (UQCRB, FOXF2, blue), C1-Differeniated (LUC7L3, PNISR, green), C2-Photoreceptor (NRL, IMPG2, white). H&E staining post Xenium workflow. Scale bar=2 mm. Quantification of subpopulations = % of neoplastic. G fGSEA dot plot of top 10 enriched paths in primary (n = 3) and recurrent tumor (n = 3) populations. The statistical significance of the enrichment score is reported as -Log10(Pval) and the normalized enrichment score (dot size), is also shown. Enrichment score calculations are based on permutation tests. H Representative UMAP marker gene expression from progenitor-photoreceptor cell state, cycling neural progenitor cell state, and Interneuron/Neuron cell state. Total nuclei count=50,184. snMultiome data from patient primary (n = 3) and recurrent tumor samples (n = 3). See also Supplementary Figs. 1–3.
Comparison of the gene expression differences between recurrent tumor populations and primary tumors by GSEA revealed an enhanced association with metabolism, stemness, the epithelial- mesenchymal transition (EMT), and DNA repair (Fig. 1G, Supplementary Fig. 3B). Additional profiling of the progenitor_photoreceptor populations in primary versus recurrent tumors identified enrichment of genes normally associated with high CpG density and H3K4me2/3 and H3K27me3 alterations in recurrences (Fig. 1G and Supplementary Fig. 2B). Indeed, expression of key determinants of neural progenitor fate and maintenance (SOX6, TCF4, and TEAD1) was observed primarily in progenitor_photoreceptor clusters and elevated in recurrent samples (Fig. 1H). Further, expression of genes that modulate metabolic responses and energy production (IDH1, HK2, HSPH1, GLS, and NFE2L1) are increased in recurrent tumors and acutely expressed in the recurrent progenitor_photoreceptor cluster (Supplementary Fig. 1E).Specifically, IDH1 expression is concentrated within the progenitor_photoreceptor cluster whereas additional enzymes that contribute to α-KG production (GOT1/2, BCAT1, KYAT1, GPT2, and IDH2) are more prevalent in differentiated clusters (Supplementary Fig. 3C). These genes were used to define the progenitor_photoreceptor populations based on a gene signature amplified in recurrent tumor samples (Fig. 1H, Supplementary Fig. 1E, Supplementary Data 1).
Chromatin restructuring in recurrent samples promote a switch in network programming
We used inferCNV33 to assess the clonal succession of large-scale copy number alterations in primary vs recurrent tumors. Primary tumors show a gain of Ch7 and Ch8, loss of Ch10, and gain of Ch17, consistent with known Group 3 and Group 3/4 amplifications3,34. Recurrent tumors maintain large-scale CNVs and display subsidiary alterations to expression patterns based on changing cell state. (Supplementary Fig. 1A–C). Comparison between snATAC and snGEX patterns revealed that shifting chromatin accessibility was positively correlated with altered gene expression (Fig. 2A, Supplementary Data 2). Furthermore, the chromatin accessibility matrix showed open chromatin regions associated with the cell-state marker genes we identified, further demonstrating the correspondence between transcriptomic and chromatin structure (Fig. 2B, Supplementary Data 2). Gains were observed in 598 peaks and a decrease in 1087 peaks in recurrent tumors compared to primary tumor samples (Fig. 2C). The proximal cis-regulatory regions associated with gained peaks were predominately associated with the regulation of chromatin silencing and metabolic program regulation (cellular amide and methylglyoxal conversion to D-lactate via glutathione) (Fig. 2D). The decreased peaks were associated with oxygen transport and differentiation (Fig. 2E). These data suggest, recurrent tumors show signs of chromatin restructuring and utilize alternative metabolic pathways to enable survival with the downregulation of oxygen availability (i.e., hypoxia) and differentiation programs.
A Two-tailed Pearson’s correlation analysis between peak accessibility (ATAC-seq, n = 187,779 peaks) and DEGs (RNA-seq, n = 36601). Log2FC in primary vs recurrent tumors is plotted. P-value represents a one-sample proportion test. B Chromatin accessibility matrix with gene expression markers (n = 32) by cell state (n = 20). Color indicates gene accessibility, and dot size refers to the percent of cells in the cluster. C Volcano plot depicts differential peak accessibility (Log2FC) in primary vs recurrent tumor cells. The statistical significance of altered peak accessibility is reported as -Log10(Padj). Padj corrected for multiple comparisons. Significant peaks (n = 20832). D Functional enrichment of gained peaks as determined by GREAT v4.0.4. Metabolic and epigenetic associations are highlighted in pink. -Log10(Binomial FDR Q) is reported. Gained peaks (n = 598). E Functional enrichment of decreased peaks. -Log10(Binomial FDR Q) is reported. Decreased peaks (n = 1087). F Peak accessibility profiles by cell state with proximal gene linkages for genes (NFE2L1, NRL, MYC, TCF4, and HLX) representative of the changing cell states in primary (n = 3) and recurrent tumor (n = 3). The height and color intensity of linkages indicate the accessibility score. GABA (GABAergic), GABA-I (GABAergic-Interneuron), Glut-I (Glutamatergic-Interneuron), Glut (Glutamatergic), PN (Purkinje), CNP (Cycling Neural Progenitor), P-Photo (Progenitor-photoreceptor), P-UBC (Progenitor-UBC), RPE (Retinal Pigment Epithelium). G Top ten gene regulatory networks in primary (n = 3) compared to recurrence (n = 3). Significance of the gene network association is reported as -Log10(Adj pVal). Padj corrected for multiple comparisons. H Dot plot of the top five active gene regulatory networks by cell state (n = 12). p < 0.01. See also Supplementary Fig. 3.
Gene-specific peak profiles were evaluated to determine the differences in gene accessibility profiles in primary and recurrent tumors based on cell state. Open chromatin regions in primary tumors indicated cell-state specificity of peaks for NFE2L1, TCF4, and HLX. Metabolic genes, IDH1, HK2, HSPH1, GLS, and NFE2L1 show decreased accessibility within CNP populations (Supplementary Fig. 3C). Chromatin accessibility increased to all cell states and condensed in specific cell states in recurrent tumors, modulating gene expression (Fig. 2F, Supplementary Fig. 3D). Additionally, proximal gene linkages between regulatory elements and gene promoters were established for TCF4 and MYC, manipulating gene expression by altering the chromatin landscape.
To then understand how the gene regulatory networks (GRNs) in primary tumors change with respect to recurrence, the integrated snATAC and snGEX data was used to determine the active GRNs. The top GRNs in primary tumors were TCF4, TAGLN2, FOXK1, and BHLHE41, whereas the top upregulated GRNs in recurrent tumor cells were FOXP2, FOXD1, SIX4, and HMGB1 (Fig. 2G). The top ten network regulon activity was mapped to the 12 major cell-states (Fig. 2H). Non-cycling progenitor populations showed specific GRN activity with TBX4, DRGX, and EN2 in UBC-linked progenitors, whereas HOXC10, NRL, and RAX regulons were most active in photoreceptor-linked progenitors. The most active GRNs in the cycling neural progenitors were HMGB1, ZNF695, and DNMT1 (Fig. 2H). In total, analysis of integrated snGEX and snATAC in primary and recurrent tumors suggests that recurrences undergo chromatin restructuring, facilitating a switch in network programming that favors pluripotency and metabolism despite oxygen depletion.
Trajectory analysis suggests the amplification of the progenitor_photoreceptor lineage in relapsed Grp3-MB
To evaluate the differences in gene expression dynamics as primary tumor cells transition to a resistant state, cell-state progression was mapped along a pseudotemporal trajectory for Group 3 primary and recurrent tumors (Fig. 3A, B, Supplementary Fig. 4A). In primary tumors, the pre-branch was occupied by progenitor photoreceptor-like cells that bifurcate into two cell-states: more differentiated cells (Cell fate 1) and cycling progenitor cells (Cell fate 2; Fig. 3A). Consequently, in recurrences, the early progenitor photoreceptor-like state occupied the pre-branch and contributed to the amplification of cycling progenitor and more differentiated states in Cell fate 2, similar to the primary tumor. However, in Cell fate 1, the progenitor photoreceptor-like cells lead to a less differentiated state (Fig. 3B, Supplementary Fig. 4B).
A Pseudotime ordering of primary tumor cell states (CNP, GABA-I, Pro-photo, Pro-UBC) B. Recurrent tumor cell states (CNP, GABA-I, Pro-photo, GABA) on a bifurcated cell trajectory, (above) color scale based on pseudotime, (below) separated and colored by cell state. C Heatmap of hierarchically clustered expression of top 100 branch-dependent genes based on branched expression analysis modeling (BEAM) for primary (genes, n = 100) D. Recurrence (genes, n = 100). The direction of pseudotime starts at the pre-branch and follows either cell fate 1 or cell fate 2. E Primary ATAC accessibility score (above) and Gene expression (below) along pseudotime trajectory for marker genes (n = 44). F Recurrence ATAC accessibility score (above) and Gene expression (below) along pseudotime trajectory for marker genes (n = 44). Color scale indicates accessibility or relative expression. See also Supplementary Fig. 4.
Hierarchical clustering of differential expression between individual cell-state branches revealed upregulated expression of RGS7, TOX, SEMA3A, SOX6, and ESRRG within the recurrent pre-branch (Fig. 3C and D). Enrichment analysis suggested that these cells aligned with retinal progenitor cells (RPCs) (cluster 3) and promoted a photoreceptor-like state in Cell fate 2 (cluster 1) (Supplementary Fig. 4B). Additionally, genes mediating the pentose-phosphate shunt and metabolic processes were amplified, as is often exhibited by neural stem cells (NSCs) and retinal progenitors. In recurrent tumors, this population was heavily maintained at Cell fate 2, whereas in primary tumors, the cells dissipate beyond the branch point.
Next, ATAC and GEX profiles were mapped onto the pseudotime trajectory for the cell-state markers. In primary tumors, marker gene accessibility was limited in progenitor_photoreceptor and interneuron cells states, whereas in recurrent tumors, chromatin accessibility was open and extended across the length of the trajectory (Fig. 3E, F). Additionally, the expression of key markers such as NRL, CRX, LHX1, and OTX2 aligned with the progenitor_photoreceptor and interneuron cell states. Recurrences showed amplification and extended NRL, OTX2, and TTR expression throughout the trajectory. Hence, the progenitor_photoreceptor-like cells are present in both primary and recurrent tumors and represent two cell-states: an early state that resembles multipotent RPCs and a later, more determined photoreceptor-like state.
Progenitor_Photoreceptor signature maps to relapse MB
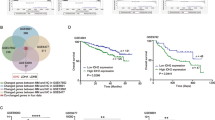
To assess if the progenitor_photoreceptor signature was present in a larger cohort of patient transcriptomic data, the Cavalli dataset3 (RNA-seq of 763 MB tumors) and Okonechnikov dataset35 (84 primary and recurrent MB tumors) were assessed. Based on gained peak association with metabolic programs, glutathione metabolism, TCA, and hypoxia gene signatures were aligned along with the progenitor_photoreceptor gene signature. The progenitor_photoreceptor, metabolic, and hypoxia signatures correlated primarily with Group 3 (Fig. 4A). A higher probability of survival associated with lower expression of the progenitor_photoreceptor signature in Group 3 and 4 tumors (Fig. 4B). Cox regression analysis identified a significant hazard ratio for genes within the progenitor_photoreceptor signature in all MB subgroups and in Group 3 and 4 tumors (Fig. 4C). Examination of the gene expression differences between the primary and recurrent groups showed the progenitor_photoreceptor signature was overrepresented in recurrences (Fig. 4D, E). Additionally, the progenitor_photoreceptor gene signature similarly aligned with metabolic and hypoxia-inducible factor (HIF) regulation gene signatures (Fig. 4E). Crucially, survival probability improved in patients with lower expression of the progenitor_photoreceptor signature in Group 3 and 4 recurrent tumors (Fig. 4F). This data suggests a cell-state change in early progenitor cells may be specified by gene signatures modulating metabolic and pluripotency programs, often observed in normal retinal progenitor cells and quiescent adult NSCs (Fig. 4G).
A GSVA of Cavalli data set (n = 763) compared to the Progenitor-photoreceptor gene signature, REACTOME_CITRIC_ACID_CYCLE_TCA CYCLE, REACTOME_REGULATION_OF_GENE_EXPRESSION_BY_HYPOXIA_WP_GLUTATHIONE_METABOLISM, WP_HIF1A_AND_PPARG_REGULATION_OF_GLYCOLYSIS, WP_TCA_CYCLE_AKA_KREBS_OR_CITRIC_ACID_CYCLE. Age: 0-3,4-10,10-17, 18 + , NA; Gender: M, F; Histology: Classic, Desmoplastic, LCA, MBEN, NA; Metastasis: 0=pink, 1=maroon, NA=gray. B Kaplan-Meier survival curves in All Groups (n = 611), Group 3 (n = 113), and Group 3/ Group 4 MB (n = 374) for high and low progenitor-photoreceptor gene signature expression groups. High >median cutoff, Low<median; Log rank Mantel-Cox test; Group 3_4, p = 0.049. C Cox regression analysis for the Progenitor-photoreceptor gene signature in all MB subgroups. Cavalli data set (n = 763). (Right) In Groups 3 and 4, the Cox proportional Hazard ratio is graphed with p-value on the right. D Differential gene expression volcano plot of primary vs recurrence from the Okonechnikov data set (n = 84). Significant changes in gene expression (p < 0.05) are reported as -Log(pVal). Progenitor-photoreceptor gene signature is overlayed in red. E Gene signature heatmap comparison to Group 3 and 4 primary (n = 19) and recurrent (n = 19) tumors in Okonechnikov data set, and REACTOME_CITRIC_ACID_CYCLE_TCA CYCLE, REACTOME_REGULATION_OF_GENE_EXPRESSION_BY_HYPOXIA_WP_GLUTATHIONE_METABOLISM, WP_HIF1A_AND_PPARG_REGULATION_OF_GLYCOLYSIS, WP_TCA_CYCLE_AKA_KREBS_OR_CITRIC_ACID_CYCLE. F Kaplan Meier survival curves in All Groups (n = 86) and in Group 3 and Group 4 (n = 37). Primary and recurrent tumors are separated based on high and low expression of the progenitor-photoreceptor gene signature. High>median cutoff, Low<median; Log rank Mantel-Cox test. Recurrence high vs. low Group 3_4, p = 0.05. G Model of the development of recurrence from a resilient cell resembling the progenitor-photoreceptor population. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/awlzjnn.
Photoreceptor_progenitor populations persist in radiation resistance murine models
To explore the development of recurrence in Group 3 MB, recurrence after radiation exposure was modeled in vivo with two separate cell lines. Group 3 mouse orthotopic xenografts were created using the D458 cell line and the MED411 PDX. Tumors were exposed to 0.5 Gy/day for 5 days to generate a model that developed recurrent tumors (Fig. 5A). Tumors responded to radiation but subsequently recurred after 3 weeks. Tumors were recovered at the endpoint and subjected to scRNA-seq (D458) and scMultiome-seq (MED411).
A Schematic of the radiation resistance model development. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/miahqmf. B UMAP visualization of control tumors(n = 2) and radiation-resistant tumors (n = 4). Total cells (n = 15982). C Cell grouping of clusters is based on ORA GSEA. Color designation is based on clustering. Control tumors(n = 2) and radiation-resistant tumors (n = 4). Total cells (n = 15982). D Heatmap of top 10 DEGs by cluster. Control tumors(n = 2) and radiation-resistant tumors (n = 4). E Over-representation analysis (ORA) heatmap based on individual clusters. Control tumors (n = 2) and radiation-resistant tumors (n = 4). F GSEA comparing radiation-resistant tumors (n = 4) to control (n = 2). The top 10 gene sets with >1 Normalized Enrichment Scores and <−1 are shown. p < 0.001. P-value is calculated by a permutation test. G UMAP visualization of PDX411 radiation-resistant tumors (914, 915, 916); (n = 3) and 411FH control (Hovestadt) and 918 (n = 2). Total nuclei =12614. H Progenitor-photoreceptor gene signature expression on UMAP of PDX411 radiation resistant (n = 3) and 411FH control (n = 2). Color scale indicates Log2 gene expression. Total nuclei n = 12614. I Violin plot of Progenitor-photoreceptor gene signature expression in radiation-resistant tumors (n = 3) compared to control (n = 2). The widest part of the violin plot corresponds with the average single-cell expression levels. J Heatmap of top three DEGs by cluster in PDX411 tumors (n = 5). Gene expression color indicates Log2 scale. K Gene ontology analysis of the top 100 genes in cluster 6. Significant GO enrichment terms are determined by the hypergeometric distribution and reported as -Log10(Pval). L Functional enrichment on genomic regions associated with gained peaks (n = 195098 genomic regions). Metabolic and epigenetic associations are highlighted in color. Significance of enrichment is determined by a binomial test after multiple testing correction (false discovery rate) applied, shown as -Log10(Binomial FDR Q). M UMAP visualization of ATAC-seq clusters from radiation-resistant tumors (n = 3) and control (n = 1). Total nuclei = 12528. The progenitor-photoreceptor and cycling neural progenitor gene expression signatures are mapped onto clusters. Color scale indicates Log2 gene expression. (Left) Representative ATAC peak profile for NFE2L2 accessibility in radiation-resistant (n = 3) and control (n = 1) tumors by cluster. Link arc plot is shown between peaks, Significance reported as -Log10(Pval)=9.43, correlation = 0.28. Linkage correlation, ranging from −1 to 1. Linkage significance: -log10 p-value after multiple testing correction (false discovery rate). Peaks are highlighted in black boxes. See also Supplementary Fig. 5.
Within the D458RT (radiation-treated) model, Seurat clustering analysis identified ten distinct clusters in the D458 dataset (Fig. 5B, C). Enrichment analysis classified clusters similar to the recurrent patient samples demonstrating the fidelity of the model. Clusters 1/2/4/5 were associated with cell cycle and DNA replication, Clusters 0/7 were associated with translation, Cluster 3 with MYC expression, and Clusters 6/7/8 with glycolysis and hypoxia (Fig. 5C–E). The progenitor_photoreceptor gene activity was mapped to D458RT model and associated with metabolically distinct clusters (Supplementary Fig. 5A–E). Enrichment analysis in D458RT tumors, established that metabolic programs mediating cellular respiration, modulation of stem cell differentiation, and DNA repair were enhanced compared to radiation-naive tumors. Negative enrichment was observed for differentiation programs and cell fate commitment (Fig. 5F and Supplementary Fig. 5F).
Seurat analysis of MED411 PDX control and RT-exposed tumors revealed 11 clusters (Fig. 5G). The progenitor_photoreceptor gene signature was mapped to Cluster 6, comprising MED411RT (radiation-resistant) cells (Fig. 5H), and was significantly upregulated in MED411RT tumors compared to control (Fig. 5I). The top three expressed genes in Cluster 6 were DNAJB1, HSPH1, and HSPA1B, reiterating the stress response genes found in patient tumors. These heat shock proteins have pivotal roles in the unfolded protein response and are linked with enhanced resistance of CSCs to genotoxic stress36,37. Gene set enrichment analysis of the top 50 genes in Cluster 6 revealed enhanced metabolic and hypoxic pathways as well as increased expression of genes responsive to 2-oxoglutarate dioxygenase activity (Fig. 5J, K, Supplementary Fig. 5G). Genes proximal to ATAC peaks were associated with metabolic processes, pattern specification, and chromatin modifications, phenocopying the peak locations in recurrent patient samples (Fig. 5L). Additionally, ATAC peak profiles showed proximal gene linkages in clusters mediating the progenitor_photoreceptor signature and cycling progenitors for NFE2L2, HLX, and HSPH1, associating chromatin restructuring in progenitor_photoreceptor populations with enhanced radiation resistance (Fig. 5M, Supplementary Fig. 5H). In both (D458, MED411) radiation-resistant models, we observed gene expression and peak association with genes that mediate metabolic processes, specifically α-ketoglutarate dioxygenase activity, cellular respiration, and metabolite and energy generation (Fig. 5E, K–M, Supplementary Fig. 5G-H). A decrease in neuronal differentiation programs and an increase in the regulation of stemness differentiation programs and epigenetic modulators that affect histone methylation were also observed (Fig. 5F, Supplementary Fig. 5G). These results recapitulate the progenitor photoreceptor signature in in vivo models and suggest that therapeutically targeting the metabolic and epigenetic dysregulation in this proportion of IR-resistant cells could block a source of resistance development.
Disruption of IDH1 activity suppresses radiation resistance and MB proliferation by inhibiting reductive glutamate carboxylation
Metabolic dysregulation in the progenitor_photoreceptor population from patient tumors implicates multiple metabolic mechanisms involving IDH1 activity. As an essential participant in metabolic and epigenetic programs, we reasoned that the disruption of IDH1 may attenuate resistance by disturbing α-KG-responsive enzymes and stress responses through regulation of metabolic networks. To evaluate this, radiation-resistant MYC-amplified MB cell lines, D458IR and D425IR, were generated. A global metabolomic analysis of D458/D458IR and D425/D425IR cells validated the depletion of D-glucose, nucleic acids, and lactate, indicating that the model radiation-resistant cells had a higher rate of metabolism, causing carbon sources to become rapidly exhausted and cells to behave as nutrient-deprived (Fig. 6A, Supplementary Fig. 6A, B). The elevated levels of metabolites (citrate, acyl-C18, ADP) are affiliated with IDH1 activity and alter the TCA cycle, glutamate, and lipid metabolism. These metabolomic results aligned with the in vivo resistant models and recurrent patient samples (Fig. 6A, B).
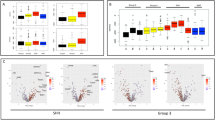
A Hierarchical clustering heatmap between radiation resistant (D458IR) n = 3 and sensitive (D458) n = 3 cells. Measured using Euclidean distance. B Metabolite enrichment summary Over Representation Analysis (ORA) in radiation resistant (D458IR) n = 3 compared to sensitive (D458) n = 3 cells. A one-sided hypergeometric distribution test was used. Color scale indicates significance of enrichment C (Above). Neurosphere assay growth in IDH1-depleted MB cell lines. Total area (μM2) of neurospheres is shown, n = 3, Mean ± SD. Significance determined by unpaired t-test, D458IR p = 0.008, D425IR p = 0.0069. (Below) Box plot of neurosphere size. Whiskers represent min and max. All data points shown. The box bounds represent the 25th and 75th percentile and center is median. Each dot represents a neurosphere. Significance determined by unpaired Welch’s t-test; D458IR p < 0.0001, D425IR p = 0.00347p. D Log IC50 determination of IDH305, GSK321, and IDH1−13 on D458, D425 sensitive and D458IR, D425IR cell lines. n = 3,biological replicates. Mean ± SEM. See also Supplementary data Fig. 6G. E Box and whisker plot of α-Ketoglutarate assay on D458, D458IR, D425, D425IR cells treated with IDH305 (7μM) Whiskers represent min and max. The box bounds represent the 25th and 75th percentile and center is median. Paired t-test. D458IR p = 0.0367, D425IR p = 0.0027 or GSK321 (1μM) D425IR p = 0.0121. n = 3. F Scatter dot plot of ALDH+ cells in D458, D458IR, D425, D425IR treated with DMSO or IDH305 (7μM). Mean ± SD. Two-way Anova, multiple comparisons; D458 p = 0.0043, D458IR p = 0.0054, D425IR p = 0.003. n = 3, biological replicates. G Extreme limiting dilution assay in D458IR and D425IR treated with IDH305 (7μM) or GSK321 (1μM). Log-fraction plot of the limiting dilution model, where the slope of the line is the log-active cell fraction. Pairwise Chi-square Goodness of fit test, CI = 0.95, D458IR IDH305 p = 0.0028, D458IR GSK321 p = 0.00132; D425IR IDH305 p = 0.06, D425IR GSK321 p = 0.00274. n = 8, independent replicates. H Survival fraction plot. (Right) Survival enhancement ratio. n = 3, biological replicates. Two-way Anova, D458IR GSK321 p < 0.0001, D458IR IDH305 p = 0.0001. I Neurosphere growth assay of D458IR cells treated with DMSO, 1.5μM IDH1−13, 2 mM α-KG, or combined. Box plot of the average area (μM2) on day 6.5. Whiskers represent min and max. The box bounds represent the 25th and 75th percentile and center is median. D458IR p = 0.0367, D425IR p = 0.0027 or GSK321 (1μM) D425IR p = 0.0121. One-way Anova, multiple comparisons, p < 0.0001 all comparisons. n = 9, independent replicates. J Model of citrate production from glutamine. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/ywuiesl. 13C5, 15N2 L-glutamine is catabolized into α-ketoglutarate (C5) which contributes to citrate production through reductive carboxylation (orange arrows, C5) and oxidative metabolism (blue arrows, C4). K Scatter bar plot of labeled 13C5, 15N2 L-Glut. 13C4, 13C5, and 13C6 are derived from labeled 13C5, 15N2 L-Glut. Total carbon is 12 C. n = 3, biological replicates. Mean ± SD. Two-way Anova, multiple comparisons. Citrate p = 0.0265, p = 0.0346; Aspartate p = 0.0369, p = 0.0422; α-KG p < 0.0001, p = 0.0209, p = 0.0234, p < 0.0001; Glutamate p = 0.0486, p = 0.0012, p = 0.0001, p < 0.0001; Malate p = 0.0341, p = 0.0005; Fumarate p < 0.0001, p = 0.0059; Succinate p < 0.0001, p = 0.0003, p = 0.0054. See also Supplementary Fig. 6.
To directly address the contribution of IDH1 in radiation resistance, wtIDH1 expression was inhibited by shRNA knockdown under NSC conditions (Supplementary Fig. 6C-E). Examination of proliferation and self-renewal of shIDH1 neurospheres showed a significant reduction in growth and self-renewal indicating that blocking IDH1 expression affects stem cell-like characteristics (Fig. 6C). Additional enzymes support α-KG production, to establish IDH1 activity as the major source of enhanced α-KG in IR-resistant cells, α-KG production was measured in parent, radiation resistant, and shIDH1 KDs. The levels of α-KG increased on average by 1.5-2-fold in radiation-resistant cells, and shIDH1 KDs show a similar decrease in α-KG compared to shNull IR-resistant cells (Supplementary Fig. 6F). To assess whether therapeutic inhibition can similarly limit IR-resistant cell viability, three IDH1 inhibitors (IDH1i) were examined. IDH305 and GSK321 are selective brain-penetrant allosteric inhibitors designed to target the R132H mutation. However, both exhibit considerable activity against wild-type IDH138,39, and inhibited growth of D458IR and D425IR lines at 7-10μM and 1μM, respectively (Fig. 6D). The third inhibitor, IDH1-13, specifically inhibits wtIDH1, and reduced the viability of radiation-resistant cells at 1.5 μM (Fig. 6D, Supplementary Fig. 6G)39. Consideration was given to the mutant IDH1 inhibitor, Ivosidenib/AG-120, currently in phase II clinical trials for AML and solid tumors. However, AG-120 was ineffective in parent and radiation-resistant MB cells (Supplementary Fig. 6G). Treatment with IDH305 or GSK321 at IC50 depleted α-KG production in D458IR by ~50%, and 40%-50% in D425IR cells (Fig. 6E). Similar to IDH1 KD, treatment with IDH1i’s reduced ALDH+ (stem-like marker) cells by 38% in D458 and D458IR cells, and by 60% in D425IR cells (Fig. 6F). An extreme limiting dilution assay (ELDA) showed a 50% reduction in stem cell frequency in radiation-resistant and radiation-sensitive D458 and D425 cells upon IDH1i (Fig. 6G, Supplementary Fig. 6H, I). Radiation-resistant cells exhibited an enhanced Survival Fraction (SF), requiring up to 6 Gy to reduce the same number of colonies as parental cells. Following treatment with IDH1i, the amount of radiation required to reduce 70% of colonies decreased to 4 Gy. Additionally, the sensitivity enhancement ratio significantly increased in the presence of IDH1 inhibitors (Fig. 6H, Supplementary Fig. 6J). We tested whether exogenous α-KG could reverse the effects of IDH1 inhibition. Exposure to exogenous α-KG significantly improved cell growth and maintained cell viability after exposure to 4 Gy IR (Fig. 6I, Supplementary Fig. 6K-L). These data further support our hypothesis that IDH1 production of α-KG promotes resilience against radiation exposure.
SnRNA-seq and snMultiome-Seq revealed a heterogeneous tumor with pockets of hypoxia and metabolically altered programming. To determine the effect of IDH1 inhibition on metabolism in radiation-resistant cells, 13C5,15N2-labeled glutamine was used to track glutamine under NSC conditions (Fig. 6J). Compared to the parent cells, radiation-resistant D458 cells showed an increase in total carbon (12 C) incorporation for succinate, citrate, α-KG, and aspartate; these were depleted upon IDH1 suppression. Labeled α-KG, citrate, and aspartate increased in radiation-resistant cells compared to sensitive (parent) cells (Fig. 6K). Whereas treatment with IDH305 in IR resistant cells blocked the conversion of α-KG back to citrate as noted by the significant decrease in incorporated 13C5. This data suggested that inhibiting IDH1 blocks α-KG production and the use of glutamate as an alternate carbon source specifically altering the availability of citrate and acetyl-coA.
Epigenetic restructuring in radiation-resistant cells triggers bivalency at pluripotent gene promoters, which are partially restored to the parental state by IDH1 inhibition
To understand the effects of altered metabolism on epigenetic reprogramming, we performed CUT&RUN analysis to examine H3K4me3, H3K27ac, and H3K27me3 occupancy (Fig. 7A). Radiation-resistant cells exhibited depleted H3K4me3 deposits and enhanced H3K27ac and H3K27me3 peaks at promoter regions (Fig. 7B) while depletion of H3K27ac and H3K27me3 was observed in distal intergenic and intron regions (Fig. 7C, D). IDH1 suppression restored global H3K4me3 promoter occupancy, whereas H3K27ac and H3K27me3 levels were reduced below the parent cell line at the transcriptional start sites (TSS; Fig. 7B–D). Radiation-resistant cells treated with the IDH1i showed a reversal in the diminished peaks compared to the isogenic parent cell line. H3K4me3 peaks were enhanced in the promoter regions, and H3K27ac and H3K27me3 peaks were enhanced in the distal intergenic and intron regions (Fig. 7B–D, Supplementary Fig. 7A). Given the decrease in H3K27ac at intergenic regions, H3K27ac levels were evaluated at enhancer regions, revealing a global increase in H3K27ac levels at enhancers, which remained unchanged under IDH1 suppression (Supplementary Fig. 7B, C).
A Experimental schematic. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/5nsqycy. B Profile heatmap of H3K4me3, H3K27ac, and H3K27me3 -3Kb upstream and 3Kb downstream of the TSS. D458 (parent), D458IR were treated with IDH305 (7μM) or GSK321 (1μM) for 48 hours. C Annotation of percent H3K27ac, H3K4me3, and H3K27me3 increased and decreased peak occupancy in D458IR(resistant) compared to D458(parent). (Below) Increased and decreased peak occupancy in D458IR(resistant) treated with IDH305 compared to DMSO treated D458IR. D Summary of H3K4me3, H3K27ac, and H3K27me3 peak occupancy changes at the promoter and distal intergenic and intron regions. E Schematic representation of (F and G). Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/5nsqycy. F Scatter plots of D458IR/D458 Log2FC ratio compared to D458IR + IDH1i/D458IR for H3K27ac (correlation coefficient = -.58, p = 2.2e−16) and G H3K27me3 (correlation coefficient = -.35, p = 5.758e-08). Two-tailed Pearson’s correlation analysis between assigned peaks in (IR/Ctrl) and (IDH1i+IR/IR). P-value represents one sample proportions test. H Scatter plots of D458IR/D458 Log2FC ratio of H3K4me3 compared to H3K27me3. DEGs from scRNA-seq is overlayed on gene identified peaks, red genes up and blue genes downregulated. Green box indicates increased peak occupancy for H3K4me3 and H3K27me3. I Gene set enrichment analysis of dual occupied H3K4me3/H3K27me3 peaks. The significance of enrichment is reported as(-Log10(Pval)). J Scatter plots of D458IR + IDH1i/D458IR Log2FC H3K4me3 compared to H3K27me3. K Genome browser view of representative gene loci from the progenitor_photoreceptor signature. Chromatin accessibility (ATAC-seq), H3K4me3, H3K27ac, H3K27me3 occupancy in D458 (sensitive), D458IR (IR-resistant), D458IR (IR-resistant) + IDH305 (7μM) or + GSK321 (1μM) are displayed. Scale for each histone mark is shown. All CUT and RUN experiments represent a n = 2. See also Supplementary Fig. 7.
To examine whether the decreased gene-associated peaks in radiation-resistant cells were associated with the peaks enhanced by IDH1 inhibition, H3K27ac and H3K27me3 peaks in radiation-resistant cells were compared to those in radiation-resistant cells treated with an IDH1i (Fig. 7E). An inverse correlation between radiation-resistant and enhanced peaks after IDH1 inhibition suggested that the redistribution of H3K27ac and H3K27me3 peaks in radiation-resistant cells was partially recovered by IDH1 suppression (Fig. 7F, G).
Bivalent promoters are commonly associated with the regulation of pluripotent stem cells, but also contribute to cancer development40,41,42,43. To evaluate the overlap between H3K4me3 and H3K27me3, we mapped peaks and DEGs from scRNA-seq of resistant MB tumors generated in vivo (Fig. 7H). Genes with enhanced H3K4me3 and depleted H3K27me3 levels were positively expressed, whereas those with enhanced H3K27me3 and depleted H3K4me3 levels were repressed. Considerable overlap between amplified H3K4me3 and H3K27me3 deposits was detected for up-and downregulated genes (Fig. 7H). Similarly, H3K27ac and H3K27me3 overlap at promoter regions was also enhanced in radiation-resistant cells (Supplementary Fig. 7D). Gene ontology of the dual-occupied genes was examined to understand which gene-associated regions showed increased bivalency. H3K4me3 and H3K27me3 dual-occupied genes mediated glucose consumption (SRF, SLC2A5), TGFβ signaling (ID2, LEFTY), Hippo signaling (TEAD2), and the polycomb repressive complex (BCORL1; Fig. 7I). Similarly, H3K27ac and H3K27me3 bivalency increased at the promoters of genes regulating stem cell pluripotency (WNT9a and TBX3), Hippo signaling (TEAD2 and BBC3), and pyruvate metabolism (FH and PC) (Supplementary Fig. 7E). We then examined the gene-associated peak location after IDH1 inhibition to understand the response to IDH1 suppression. H3K27me3 peaks were reduced, whereas H3K4me3 peaks remained largely enhanced, excluding several key genes (FZR1, and SPRY4) which reverse the epigenetic profile (Fig. 7J). Comparably, the fold-change after treatment showed a minimal decline and enhancement in both H3K27ac and H3K27me3 peaks (Supplementary Fig. 7F).
To determine whether bivalency was present on genes within the progenitor_photoreceptor gene signature, individual peak profiles were examined. ATAC-seq was conducted on parent, and radiation-resistant cells after exposure to an IDH1i. ATAC sequencing of resistant cells revealed an opening of chromatin regions, whereas IDH1 treatment diminished chromatin accessibility (Supplementary Fig. 7G). Comparison of ATAC and histone peak profiles of several genes in the progenitor_photoreceptor signature showed radiation-resistant cell amplification of H3K4me3 and H3K27ac for NFE2L1, SOX11, ID4, RAX2, and POU3F1. Similarly, H3K27me3 levels increased compared to sensitive (parent) cells; treatment with IDH1i reversed and/or diminished these effects (Fig. 7K). These results suggested that radiation-resistance may be facilitated by a bivalency switch on pluripotent gene promoters mediating Hippo signaling, TGFβ signaling, and metabolism. However, this can be disrupted by IDH1 suppression reinstating the epigenetic profile of the parent cells. Suppression of IDH1 as a mechanistic key to supplying the substrates and energy source for this transition can impede epigenetic and metabolic reprogramming.
Secondary radiation therapy combined with IDH1 inhibition, extends survival benefit of radiation alone
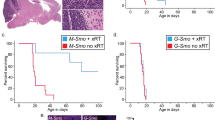
To establish the proof of concept for targeting wtIDH1 in vivo, D458IR shIDH1 or D458IR shNull cells were injected into the cerebellum of nude mice to establish orthotopic xenografts. Secondary radiation was administered at 0.5 Gy/day for 5 consecutive days, 10 days post injection (Fig. 8A). IDH1 KD cells increased survival in mice with and without secondary radiation compared to shNull and shNull+IR (Fig. 8B). The therapeutic efficacy of targeting IDH1 was further examined in D458IR flank tumors. A flank model was selected because the chemical inhibitor IDH1-13 does not effectively penetrate the CNS. Treatment with IDH1-13 (10 mg/kg) once daily was initiated and concurrent radiation at 0.5 Gy/day was administered for one week (Fig. 8A). Tumor growth is delayed in treated mice, and co-treatment with radiation further suppresses tumor volume (Fig. 8C–E). These results support IDH1 co-inhibition with radiation as a method of mitigating radiation resistant tumor growth.
A Radiation therapy and IDH1−13 treatment schedule xenograft and flank in vivo studies. Created in BioRender. Vibhakar, R. (2025) https://BioRender.com/miahqmf. Treated with 0.5 Gy for 5 days starting on day 10. B Kaplan-Meier survival curve of D458IR shIDH1 (n = 11) and shNull (n = 9) xenograft mice and D458IR shIDH1+IR (n = 13) and shNull+ IR (n = 9). Statistical analysis, log-rank (Mantel-Cox) test, p = 0.007, p = 0.037, p = 0.0009, p < 0.0001. C Box and whiskers plot of Tumor volume (mm2) from vehicle (n = 4), IDH1−13 (n = 6), Vehicle+IR (n = 4), and IDH1−13 + IR (n = 6). Whiskers represent min and max. The box bounds represent the 25th and 75th percentile and center is median. Statistical analysis, Mixed-effects analysis, Tukey’s multiple comparisons test; p = 0.002, p = 0.0076, p = 0.0098. D Tumor images, centimeters and millimeters. vehicle (n = 4), IDH1−13 (n = 6), Vehicle+IR (n = 4), and IDH1−13 + IR (n = 5). E Box and whiskers plot of Tumor weight (grams) from vehicle (n = 4), IDH1−13 (n = 5), Vehicle+IR (n = 4), and IDH1−13 + IR (n = 5). Whiskers represent min and max. The box bounds represent the 25th and 75th percentile and center is median. Statistical analysis, One-way Anova; p = 0.0280.
Discussion
Therapy resistance remains a key challenge in pediatric brain tumors and is the major contributor to tumor driven morbidity and mortality in this vulnerable population. In the present study, we evaluated the key differences in recurrent medulloblastoma compared to primary tumor sites utilizing scMulti-omic sequencing. This enabled the characterization of the genetic and epigenetic intratumor heterogeneity with further attention placed on the development of radiation tolerance. Therapy tolerant cells (TTC) also called drug tolerant persister cells (DTP) are phenotypically and genomically heterogeneous44. DTP cells are generally characterized by quiescence and slow proliferation, altered metabolism, altered epigenetic programming, resistance to apoptosis, and immune evasion45. The current study presents evidence Group 3 MB heterogeneity arises from differential lineage commitment controlled by distinct transcription factor_driven gene regulatory networks (TF-GRNs). We show that these GRNs are altered in therapy-resistant tumors, resulting in an enrichment of DTP cells mediated by a neural progenitor_photoreceptor GRN. Okonechnikov and colleagues recently identified similar specific TF-GRNs in MB subpopulations46. Our snATAC seq data further showed that the chromatin of therapy tolerant medulloblastoma tumors was restructured to exhibit increased accessibility, particularly at cis-regulatory elements associated with regulating chromatin silencing and metabolic programs. The data suggest that as the primary tumor progresses to recurrence, the early progenitors undergo a cell-state change, further amplifying a multipotent state of early photoreceptor-like progenitor tumor cells.
Harnessing two in vivo radiation resistance models substantiated patient tumor observations adding weight to the study and focused more distinctly on the contribution of radiation resistance to recurrence. The results showed a rewiring from differentiation programs and cell fate commitment to regulatory networks controlling metabolic programs mediating stem cell differentiation in radiation-resistant tumors. GRNs associated with cellular respiration and α-KG metabolism were enriched in resistant tumors, particularly in the progenitor_photoreceptor population. Metabolomics confirmed that radiation-resistant cells had a higher rate of metabolism with enriched metabolites associated with the TCA cycle, glutamate metabolism and IDH1 activity. Interestingly, wtIDH1 enzymatically regulates energy production and glutamine metabolism while producing α-KG, a crucial substrate for iron-dependent enzymes involved in epigenetic programming and maintenance of pluripotent embryonic stem cells47,48,49. A recent study reported that IDH1 regulated DLAT to prime Group 3 medulloblastoma for cuproptosis50. Here we demonstrate that the inhibition of wtIDH1 reversed radiation resistance and depletion of a stem cell state in IR-resistant MB cells. Furthermore, chromatin was remodeled during radiation resistance, a phenomenon that can be reversed upon wtIDH1 inhibition. This provides additional evidence supporting the assertion that epigenetic reprogramming plays a crucial role in driving therapy resistance. Our data emphasizing the role of wild type IDH1 in therapy resistance in medulloblastoma is in line with previous reports showing the importance of wt IDH1 in Glioblastoma, Lung Cancer, and Pancreatic Cancer49,51,52,53. In Glioblastoma wtIDH1 regulates histone methylation and expression of glial differentiation markers. Similarly in NSCLC wtIDH1 interacts with FXR1 to promote a stem cell phenotype by altering serine metabolism. In pancreatic cancer low magnesium levels facilitate allosteric inhibition of wtIDH1 which results in cancer cell death under nutrient limited conditions54. These and other studies corroborate the central role of IDH1 in cancer cell metabolism and establish the complex nature of the underlying mechanisms.
Tumor initiating cells exhibit metabolic and epigenetic plasticity and can reprogram their cell state to adapt to external stimuli55. Indeed, metabolic adaptation is a way for tumor cells to acquire stem-like features56. Thus, our insights into the metabolic dependencies of progenitor cells in medulloblastoma and the central role of IDH1 in mediating the metabolic and epigenetic switch during adaptation to therapy may provide potential therapeutic strategies for targeting these progenitor populations.
While obtaining quality-matched medulloblastoma primary and recurrent pairs limited the current size of this multi-omic study, the continual addition of matched pairs to increase the sample size and strength of the study is ongoing. Where multiple differentially active GRNs were identified in specific cell populations, the underlying theme of metabolic and epigenetic program dysregulation presented a comprehensive vulnerability for targeting. Additional validation of cell state GRNs will be necessary to fully appreciate the differing cellular states’ epigenetic and transcriptional alterations promoting resistance development in Group 3 MB. Collectively, this study links the persistence of the progenitor_photoreceptor GRN within the radiation-resistance model and recurrent patient tumors to the rewiring of epigenetically defined pluripotency programs via IDH1-mediated metabolic reprogramming. These data provide evidence that targeting wtIDH1-regulated mechanisms in relapsed MB may be an effective therapeutic approach for this complex tumor.
Methods
Our research complies with all ethical regulations and was performed in accordance with the National Research Council’s Guide for the Care and Use of Laboratory Animals and was approved by the University of Colorado Anschutz Medical Campus Institutional Animal Care and Use Committee.
scMultiome sample preparation
Patient samples
Human MB samples were collected and viably banked at the time of surgery at Children’s Colorado with informed written consent from the parents or legally authorized individuals (COM-IRB 95-500) over a 10-year period. Samples were processed to obtain single nuclei for single-cell Multiome sequencing according to the 10X Genomics (Pleasanton, CA) protocol. Briefly, a rice grain size of cyropreserved tumor tissue was homogenized in chilled 0.1X Lysis Buffer with a pellet pestle. Lysis buffer(10 mM Tris-HCl (pH 7.4), 10 mM NaCl, 3 mM MgCl2, 0.1%Tween-20, 0.1% NP-40, 0.01% Digitonin, 1% BSA, 1 mM DTT, 1U/μL RNase Inhibitor) and Lysis Dilution buffer(10 mM Tris-HCl(pH7.4), 10 mM NaCl, 3 mM MgCl2, 1%BSA, 1 mM DTT, 1U/μL RNAse inhibitor). Samples were incubated on ice for 5 min, mixed, then iced again 10 min. Samples are washed with 500μL of chilled wash buffer (10 mM Tris-HCl(pH7.4), 10 mM NaCl, 3 mM MgCl2, 1%BSA, 0.1% Tween-20, 1 mM DTT, 1U/μL RNase inhibitor) and passed through a 40μM strainer twice. The samples were centrifuged at 500 rcf for 5 min at 4 °C. The nuclei pellet was then washed twice with 1 mL of chilled wash buffer. Nuclei were counted on the Countess II FL automated cell counter, and nuclei were diluted in nuclei buffer and submitted for sequencing. The Chromium Next GEM Chip J Single Cell Kit (10xGenomics), Dual Index Kit TT set A, and Chromium Next GEM Single Cell Multiome ATAC + Gene Expression Reagent Bundle were used for sample preparation and library sequencing on the Chromium X.
PDX model
MED411 cell isolation from mouse xenograft was conducted with NeuroCult™ Enzymatic Dissociation Kit (STEMCELL Technologies). Mouse cells were depleted from the single cell suspension with the Mouse Cell Depletion Kit on LS Columns (Miltenyi Biotec). Isolated human cells were then processed according to 10X Genomics nuclei isolation protocol for single-cell Multiome sequencing as described.
scMultiome data processing
The feature-barcode matrix was generated using cellranger-arc (10x Genomics, v2.0.2), aligning the sequenced reads to the human reference genome (10x Cell Ranger reference GRCh38 2020-A-2.0.0) followed by standard procedure using Seurat R package (v4.4.0). Cells with fewer than 200 genes detected were filtered from the scMultiome dataset. Low-quality cells with mapped reads in the mitochondrial genes >10% were removed. Doublets were identified using scDblFinder (v1.16.0). To address the issue of technical differences across samples, the cell count matrix was individually normalized by the total read count and log-transformed, and the top 2000 variable genes were selected for each sample using option “vst” in Seurat’s FindVariableFeatures function. Principal components analysis (PCA) was conducted on the scaled expression matrix. The top 15 principal components (PCs) were used to compute SharedNearestNeighbors (SNNs) in the Seurat R package, and for UMAP embedding. Nucleosome signal was computed with Signac’s Nucleosome Signal function. Low-quality cells were removed from the scMultiome dataset based on the following criteria: <2000 or > 100,000 fragments mapped to peak regions, nucleosome signal > 2, and TSS enrichment <1. MACS2 was employed to recall peaks for each sample. LSI was performed by Signac (v1.10.0). A joint neighbor graph that represents both gene expression and DNA accessibility measurements was obtained using the FindMultiModalNeighbors function in Signac. UMAP was generated PCA dimensional reduction for gene expression and LSI indexing for DNA accessibility. A supplement providing individual clinical and gene expression signature information is provided as Supplementary Data 1, and scATAC peak association as Supplementary Data 2. Enriched functional annotation and Pathway analyses were performed using top 100 differentially expressed genes (DEGs) using clusterProfiler (v4.8.2), referencing GO and KEGG databases. Cis-regulatory regions determined by ATAC-seq was conducted with GREAT57,58. Enrichr59,60,61 and Metascape62.
Cell type annotation
Cells were clustered using FindClusters in Seurat with resolution of 0.5. Clusters were annotated based on known marker genes and integrated datasets24,25,26,27,28,29,30,31.
Trajectory analysis
Cell trajectories were analyzed using Monocle2 (v2.28.0). Selected cells were first down-sampled to a total of 5000 to remove trends driven by different cell abundance in different cell types. The dispersion table function in Moncle2 was applied to calculate gene variances. Top genes with mean expression > 0.1 were selected for further analysis. UMAP creation and pseudotime estimation were performed by the reduceDimension and orderCells functions in Monocle2, respectively. Branched expression analysis modeling (BEAM) was utilized to identify DEGs along pseudotime in different lineages.
Supplementary data
Combined trajectory was performed using Monocle3 (v1.4.0)63 and Seurat (v4.9.9)64. Differential expression analysis was performed using the ‘graph_test’ function. Lineage-specific metadata and DEGs were saved for further analysis.
Transcriptional factor analysis
pySCENIC (v0.12.1) was used to infer transcription factors. Genes co-expressed with transcription factors were identified using GENIE3. AUCell was then used to score the activity of each regulon in each cell. Regulons with the highest activity in each group were identified by the Wilcox rank-sum test followed by Bonferroni correction with an adjusted P-value < 0.01.
scRNA sequencing
D458 xenograft tumors were harvested from the cerebellum of mice. Tumors were chopped with razor blades and dissociated in RPMI media. Live cell suspensions were flow sorted based on green fluorescent protein (GFP) expression on a MoFlo XDP-Flex (Beckman Coulter) and 2000/μL GFP+ cells were submitted to the University of Colorado Genomics core for scRNA sequencing with the Chromium X. n = 6.
scRNA-sequencing data processing
Cell Ranger (v6.0.1)65 was used to process the fastq files to cell and gene count tables using unique molecule identifiers (UMIs) aligning to GRCh38 (compiled by 10X Genomics as 2020-A). Downstream quality control and analysis utilized the Seurat (v4.3.0)64, pipeline. Cell Ranger-filtered data was further processed by removing genes identified in fewer than 10 cells. Cell barcodes were removed if they had greater than 75,000 UMIs, fewer than 5000 UMIs, greater than 20% of UMIs from mitochondria or fewer than 1500 genes. The gene counts were normalized using Seurat where counts are divided by each cell’s total counts and multiplied by 10,000 followed by natural-log transformation. Samples were integrated using the RPCA Seurat method followed by clustering. The top 200 enriched genes by cluster were used as input for over-representation analysis66 with gene sets from the MSigDb collection67.
Supplementary data
Analysis performed using Pluto (https://pluto.bio). Data was processed using nf-core/scrnaseq v2.4.0 (https://doi.org/10.5281/zenodo.3568187) of the nf-core collection of workflows68, utilizing reproducible software environments from the Bioconda69 and Biocontainers70 projects. Module score analysis: Gene module activity for progenitor-photoreceptor was generated from the Seurat1 object data slot of the RNA assay, which contains log normalized values, using the AddModuleScore()64. This function calculates a score for each cell that reflects the activity of progenitor-photoreceptor by averaging the expression of 10 genes within the module and normalizing these scores to ensure comparability across all cells. Cell scatter plot: Cell scatter plot showing the activity scores for progenitor-photoreceptor across individual cells, using t-SNE-defined coordinates. Points are colored by the module activity score that was computed from 10 genes. Gene set enrichment analysis (GSEA)67 was calculated using the fgsea R package and the fgseaMultilevel()71. The log2 fold change from the Progenitor - RT vs Progenitor- control differential expression comparison was used to rank genes. C5: Gene Ontology gene sets - biological process gene set collection from the Molecular Signatures Database (MSigDB)72,73 was curated using the msigdbr R package4. Prior to running GSEA, the list of gene sets was filtered to include only gene sets with between 5 and 1000 genes. GSEA Table: GSEA results for each tested gene set are shown in the table. The Adj p-value column contains the false discovery rate (FDR)-adjusted p-value. The NES column contains the normalized enrichment score (NES) computed by GSEA, which represents the magnitude of enrichment as well as the direction. A positive NES indicates more enrichment in the first group, while a negative NES indicates more enrichment in the second group. Genes ranked by log2 fold change along the x-axis, with the vertical ticks representing the location of the genes in this gene set. The heatmap displays the expression of the genes, red being more expressed in the first group and blue more expressed in the second group. The green line shows the enrichment score.
Analysis of single-cell/single-nuclei RNA-Seq data from MED411 PDX model
Data integration, clustering analysis, and plotting were performed in Seurat (5.0.0), running under R version 4.3.1. Single-sample gene set enrichment analysis (ssGSEA, version 10.1.0) was performed using GenePattern (Broad Institute).
Integration and analysis of scRNA-seq data
The gene expression matrix file for scRNA-Seq performed on MED411 patient samples was obtained from Gene Expression Omnibus archive GSE11992615. These data were merged with the GEX single nuclei gene expression data from mouse PDX samples 914 – 918, which were derived from MED411. Data were log-normalized, and SCTransform (v2) was performed in Seurat using percent mitochondrial gene expression as the regression variable. Harmony integration74 was applied but deemed unnecessary, presumably because all samples originated from a single patient. Dataset integration was validated by comparing Uniform Manifold and Projection (UMAP) plots of the patient sample and the untreated PDX control. To generate the clustering analysis, a principal component analysis (PCA) with 30 dimensions was used to generate the UMAP projection. FindClusters was run in Seurat with a resolution of 0.6. Differential expression analysis was performed on the SCT assay using FindAllMarkers with min.pct and logfc threshold set to 0.25. Only positive differential expression (enrichment) was identified. The progenitor_photoreceptor gene signature was compared between the RT and control using ModuleScore and the following genes: (“SOX4”, “TEAD1”, “HSPH1”, “KDM5B”, “KDM4B”, “NFE2L1”, “EVX1”, “SOX11”, “RCOR1”, “BHLHE40”, “CRX”).
ssGSEA analysis
Aggregate expression was computed by gene for each sample. The aggregate data were normalized using the calcNormFactors function of EdgeR with the trimmed mean of M-values “TMM” method. Data were output without log transformation. ssGSEA was performed using GenePattern with no sample normalization and a weighting exponent of 0.75. Output gct files from ssGSEA were filtered, and heatmaps were plotted using pheatmap (1.0.12) in R.
Xenium spatial transcriptomics
Frozen samples were OCT embedded and 5 μm sections were mounted on xenium slides according to 10x Genomics protocol. A custom-designed probe panel14 was hybridized overnight. Slides are washed and probes are amplified via rolling circle amplification. Stained slides are scanned on the Xenium Analyzer. Enrichment of subpopulations determined as in ref. 75.
Cell culture
D425 (RRID:CVCL_1275; male) and D458 (RRID:CVCL_1161; male) cell lines were kindly provided by Dr. Darell D. Bigner (Duke University Medical Center, NC). Cells were cultured in Dulbecco’s modified eagle’s medium (DMEM) supplemented with 10% FBS (Sigma-Aldrich), 1 mM sodium pyruvate (Gibco), 1× penicillin/streptomycin solution (Cellgro), and with 1× L-glutamine (Cellgro). NHA cells were provided by Cynthia Hawkins (University of Toronto Hospital for Sick Children). MED411FH was originally harvested from a large cell anaplastic MB that was molecularly characterized as a Group 3 tumor. All cell lines were cultured at 37 °C with 95% air and 5% CO2. Cell lines were authenticated with STR fingerprinting using the Globalfiler® System (Thermo Fisher Scientific) and processed on ABI 3500Xl Genetic Analyzer, and mycoplasma testing was conducted with the Venor™ GeM Mycoplasma Detection Kit (Cat# MP0025-1kt, Sigma-Aldrich), both of which were done as recently as 2/23/2024. All cell lines were maintained for a maximum of 20 passages for the duration of the experiments.
Animal model
Female athymic Nude-Foxn1nu mice aged 4 to 8 weeks and female NOD-scid gamma (NSGTM) aged 4 to 8 weeks were utilized for orthotopic xenograft studies. For this initial proof-of-principle efficacy screen, we used female mice exclusively to minimize non-tumor biological variance and husbandry-related confounders. Female cohorts allow stable co-housing with markedly reduced aggression, lowering stress-induced corticosterone and inflammation that can alter tumor growth and drug response, and improving retention of cranial implants critical for longitudinal imaging. Using a single sex increases statistical power for go/no-go decisions while reducing compound usage for expensive biologics. Sex/gender differences were not analyzed in this study. Mice were housed in ventilated cages with appropriate nesting enrichments, bedding, regular chow diet, and water according to the University of Colorado regulations. For the D458 resistance and D458IR shNull/shIDH1 survival study, cells were collected and resuspended as a single cell suspension of 20,000 cells/3 µL in serum-free media. MED411 cells were injected at 200,000/ 3 µL in serum-free media. Intracranial injection of cells into outbred athymic Nude-Foxn1nu mice (Jackson Labs strain 07850) or NSG (Jackson Labs strain 005557) was performed at 1.5 mm lateral and 2 mm posterior of lambda at 400 nanoliters/min. Mice were monitored for tumor growth daily and euthanized by asphyxiation and cervical dislocation when 15% weight loss was reached or signs of tumor (hunched back, paralysis, or tilted head) were evident. Littermates were randomly assigned to experimental groups.
Flank study
A total of 250,000 D458IR cells in a 1:1 ratio of Matrigel to serum-free media were injected subcutaneously into the flank of Nude-Foxn1nu mice. Treatment with IDH1-13 at 10 mg/kg daily by i.p. for 4 weeks. Tumor volume was determined by caliper measurements, and tumor weight was measured at the endpoint. The University of Colorado Anschutz Medical Campus mandates that euthanasia for mice is required when the tumor burden is greater than 10% of the animal’s body weight or when a single tumor exceeds 20 mm (2 cm) in any dimension. The maximal tumor burden was not exceeded. All animal procedures were performed in accordance with the National Research Council’s Guide for the Care and Use of Laboratory Animals and were approved by the University of Colorado Anschutz Medical Campus Institutional Animal Care and Use Committee.
Irradiation of animals
Animal radiotherapy was performed using the X-Rad SmART small animal irradiator (Precision X-Ray, North Branford CT). Under isoflurane anesthesia, mice received 2.5 Gy to the cerebellum, in five fractions of 0.5 Gy, delivered on consecutive days with pairs of 10mm-diameter lateral beams.
Viral transduction
IDH1 stable knockdown cell lines were established using gene-specific mission shRNAs (Cat# TRCN0000027249, CCTTTGTATCTGAGCACCAAA) (Cat# TRCN0000027253, GCTTTGGAAGAAGTCTCTATT) or a non-targeting control (Sigma-Aldrich). HEK293FT cells were plated at 5 × 106 cells/10 cm dish 1 day before transfection in complete media (DMEM, 10% FBS, 5% penicillin and streptomycin). Lipofectamine 3000 (Thermo Fisher Scientific, Cat#L3000008) was diluted in Opti-MEM at a ratio of 1:16.6. P3000 was diluted at 1:25 in Opti-MEM along with 1μg of pMD2.G, 2 μg of psPAX2, and 1.2 μg of pLenti-shIDH1 expression vector. Both mixes were combined and incubated for 15 mins at 25 °C. The mixture was added dropwise to HEK293FT cells in fresh media, 24 h later, the media was replaced with DMEM/2.5% FBS. Viral particles were collected at 48 h in a 15 mL conical and centrifuged for 5 min at 500 x g. Viruses were filtered through a 0.45 μm filter and stored in 1 mL aliquots at -80 °C. To generate the stable knockdown cell lines, cells were plated at 5 × 105 in a 6-well dish in growth medium containing 8 µg/mL polybrene. Viral particles were added with 8 µg/mL polybrene and incubated for 7 h. Transduced cells were disaggregated, cultured, and selected with 2 µg/mL puromycin for 14 days. Transformants were confirmed by qRT-PCR and western blot.
Western blot
Cells were lysed in RIPA buffer (Pierce, Thermo Fisher Scientific) containing an EDTA-free protease inhibitor (Roche), and protein concentrations were determined with the BCA Protein Assay Kit (Pierce, Thermo Fisher). 30 µg of total protein was separated on a 4-20% gradient SDS-PAGE (BioRad). Primary antibodies α-IDH1 1:1000 (#8137), α-H3K27me3 1:1000 (#9733), α-H3K27ac 1:1000 (#8173), and α-H3 1:1000 (#9715) (Cell Signaling Technology) were exposed overnight at 4 °C. Secondary antibody, α-rabbit-HRP 1:2000 Cat#7074 or α-β-Actin-HRP 1:2000 #12262 (Cell Signaling Technology), was exposed for one hour at RT. Blots were developed with Luminata Forte Western HRP (Millipore) and imaged using Syngene GBox Chemi-SL1.4 gel doc. Western blots quantified using ImageJ76. Proteins were normalized to loading controls, then compared to shNull or DMSO.
Quantitative reverse transcription PCR
Total RNA was isolated from 1 × 106-1 × 107 cells with the RNeasy Mini kit (Qiagen). 2 μg of isolated RNA was used to make cDNA using the High Capacity Reverse Transcription Kit (Thermo Fisher Scientific) as per manufacturer’s instructions. cDNA was then diluted to 20 ng/μL for qRT-PCR reactions. TaqMan Universal Master Mix II, no UNG (Thermo Fisher Scientific) was used with gene-specific TaqMan primers for all qRT-PCR reactions. 5μL of the diluted cDNA was used per 20μL PCR reaction. Reactions were conducted in triplicate and performed on a QuantStudio 5 Real Time PCR machine (Thermo Fisher Scientific). Analysis of cDNA transcripts was done using the comparative Ct method.
Compounds
IDH305 (HY-104036), IDH1 inhibitor 2 (compound 13) (HY-128661), GSK321 (HY-18888) were all obtained from MedChemExpress. Chemical compounds were diluted in dimethylsulfoxide DMSO at 10 mM stock concentration. Working concentrations were then diluted in serum-free media. Exogenous α-Ketoglutaric Acid (Sigma Aldrich) stock concentration of 1 M in PBS, was diluted to 2 mM in media.
Survival enhancement ratio and methylcellulose assay
D458, D458IR, D425, and D425IR cells were exposed to irradiation with the Precision MultiRad350. Dosages included 0, 2, 4, 6, and 8 Gy. 48 hours after exposure 1000 up to 10,000 cells/3 mL were plated in a 1:1 mixture of 2.6% methylcellulose and complete growth medium. Cells were allowed to grow for 10 days; colonies were stained with nitrotetrazolium blue chloride (Sigma-Aldrich) at 1.5 mg/mL in PBS for 24 h at 37 °C and counted.
Extreme limiting dilution assay
Cells were seeded in a 96-well ultra-low-attachment round-bottom tissue culture plate in serum-free media at 1 cell/well, 10 cells/well, and 100 cells/well. Cells were allowed to grow for 14 days, then the number of neurosphere in the wells were counted. ELDA software (http://bioinf.wehi.edu.au/software/elda/) was used to calculate the comparative self-renewal potential of cells77.
Neurosphere growth assay
D458IR shNull, D458IR shIDH1, D425IR shNull, D425IR shIDH1 D458 or D425 shNull and shCDK7 cells were serially diluted at 100,10, and single-cell suspensions in neurosphere growth media (neurobasal medium, B-27+vitamin A, L-glutamine, pen/strep, epidermal growth factor (EGF), and fibroblast growth factor (FGF). Cells were grown for 14 days with media replacement every 3 days at 37 °C. At 14 days, spheres were collected, disassociated, and replated at 100, 10, and single-cell suspensions; thereafter, spheres were allowed to grow for an additional 14 days. Neurosphere proliferation and real-time monitoring were performed using the Incucyte® S3 live cell imaging system (Essen Bioscience).
α-Ketoglutarate assay
D458, D458IR, D425, and D425IR cells were treated for 48 hours with IDH305 or GSK321 at IC50 or DMSO. α-KG standards were prepared at 0, 2, 4, 6, 8, and 10 nmole/well in 50μL of α-KG assay buffer. Cells were collected and resuspended in 100μL of ice-cold α-KG buffer. 50μL of sample was mixed with 50 μL of reaction mix and incubated at 37 °C in the dark for 30 mins as per the manufacturer’s instructions (Sigma Aldrich, α-Ketoglutarate Quantitation Kit, Cat#MAK541). Absorbance was measured at 570 nm.
α-Ketoglutarate rescue
2.5×105 D458IR cells were exposed to 1.5 μM IDH1-13, 2 mM α-KG, or combined for 48 hours then irradiated at 4 Gy. Cells are collected after 24 h, washed with PBS, and then resuspended 1 × 106 cells/mL. Cells were stained with propidium iodide. Live cells are captured on the Cytoflex flow cytometer. D458IR neurosphere proliferation with 1.5 μM IDH1-13, 2 mM α-KG, or combined was monitored over 6 days on the Incucyte® S3 live cell imaging system (Essen Bioscience).
ALDEFLUOR™ assay
D458, D458IR, D425, and D425IR cells were treated for 48 hours with IDH305 at IC50 or DMSO. Cells were collected and then diluted at 1 × 106/mL in ALDEFLUOUR assay buffer. Cells were exposed to 5μL of ALDEFLUOR reagent/mL for 60 mins at 37 °C as per the manufacturer’s instructions (STEMCELL technologies, ALDEFLUOR™ Kit (Cat#01700)). Samples were counterstained with Draq5™ (eBioscience™). Following incubation, samples were assayed on the CytoFLEX Flow Cytometer. 10000 events were collected. Gating and analysis were performed using FlowJo Software.
IC50 determination
Cells were plated at 1000 cells/well in a 96-well dish and exposed to DMSO, IDH305, GSK321, or IDH1-13 for 48 h at 0, 0.1, 0.25, 0.5, 1, 2, 5, 7, 10, 20, and 30 µM. Cell viability was assessed with the CellTiter 96® Aqueous One Solution Cell Proliferation Assay #G3580 (Promega) as per the manufacturer’s instructions.
Mass spectrometry and 13C5, 15N2 L-glutamine trace
D458 (Sensitive) and D458IR (IR-resistant) cells were plated at 1 × 106 in NSA media with DMSO or 7μM IDH305. At 24 h NSA media was replaced with glutamine-deficient NSA media + 13C5, 15N2 L-glutamine (0.5 mM) + 7μM IDH305 or DMSO. Cell pellets were collected after 4 h, and samples were snap-frozen. For high-throughput polar metabolite profiling, D458, D458IR, D425, and D425IR cell pellets were collected and snap-frozen. Samples were submitted to the University of Colorado School of Medicine Metabolomics Core for analysis. Metabolites were extracted using 5:3:2 methanol:acetonitrile: water and profiled using a Thermo Vanquish UHPLC coupled to a Thermo Orbitrap Exploris 120 mass spectrometer as previously described78. Statistical and enrichment analyses on metabolite profiles were conducted using Metaboanalyst79. Flux analysis was performed by the University of Colorado School of Medicine Metabolomics core. Metabolite profiling mass spectrometry and glutamine trace flux analysis are provided as Supplementary Data 3 and 4.
ATAC-sequencing
D458 and D458IR cells were exposed to IDH305 or GSK321 at IC50 for 48 h. 50,000 cells were collected and washed in cold 1x PBS. Cell pellets were resuspended in 450μL of cold hypotonic buffer (10 mM Tris-HCL, pH7.4, 10 mM NaCl, 3 mM MgCl2). Next, 50μL of 1% IGEPAL CA-630 was added and inverted to mix. Samples were iced for 10 min, centrifuged for 10 mins at 4 °C. Cell pellets were resuspended in the transposition reaction mix and incubated at 37 °C for 60 min. Following transposition, the DNA was isolated, purified, and prepped for sequencing. The sequencing library is purified with AMpure XP beads (Millipore). ATAC assay kit (Cell Biologics™ ATAC-Seq Kit (Cat# CB6936)).
Cut and run-sequencing
D458 and D458IR were exposed to 10μM 1DH305, or 1μM GSK321 for 48 h; 5 × 105 cells were collected for analysis by CUT&RUN (EpiCypher, CUTANA™, CUT&RUN Kit Version 4 #14-1048) per the manufacturer’s instructions. Antibodies against α-H3K4me3 and IgG controls were validated antibodies from EpiCypher. α-H3K27me3 (Cell Signaling technologies, #9733) and α-H3K27ac (Cell signaling technologies, #8173). CUT&RUN library prep was performed with the NEBNext®Ultra™II DNA Library Prep Kit for Illumina® (NEB #E7645L) and NEBNext Multiplex Oligos for Illumina (NEB#E7350). Sequencing was performed on a Novaseq 6000. CUT&RUN-seq reads were aligned to the reference human genome hg38 using BOWTIE (v.2.3.4.1). Aligned reads were stripped of duplicate reads using Sambamba (v.0.6.8). Peaks were called using the program MACS (v2.1.2), with the narrow peak mode using matched input controls and a q-value of 0.00001. Peaks in the blacklisted genomic regions identified by the ENCODE consortium were excluded using bedtools. For downstream analysis and visualization, bamCoverage was used to generate bigwig files, and density maps were produced using IGV tools. Group 3 MB enhancers were defined based on H3K27ac signals. Regions within 1 kb of RefSeq TSS locations and peaks with strong H3K4me3 signals typical of active promoters were subtracted from these signals. Annotation and visualization of the peaks were conducted using ChIPseeker (v3.18). Differentially marked genes were calculated using DiffBind and DESeq2, based on the threshold of FDR < 0.05 and FC ≥ 2.
Statistics and reproducibility
All experiments were performed with at least three independent replications, except where indicated. All data were collected in Excel, and Graphpad Prism 10.3.1 statistical software was used for analysis. Data are presented as the mean ± standard deviation (SD). P-values < 0.05 were considered significant. Unpaired, two-tailed t-tests were used for two-group comparisons, and ANOVA analysis and Dunnett’s test were performed for multiple group comparisons. Patient statistical analyses were performed using the Wilcox rank-sum test. Multiple comparison corrections were applied using the Bonferroni method.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All raw snMultiome, bulk RNA, ATAC, and Cut and Run sequencing files and processed data generated in this study are available through GEO under the submission super series (GSE283621) https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE283621. The publicly available Hovestadt bulk RNA sequencing dataset was obtained from Gene Expression Omnibus (GSE119926) https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE119926. Okonechnikov35 and Cavalli3 RNA_sequencing datasets were accessed through R2: Genomics Analysis and Visualization Platform (http://r2.amc.nl)’. The remaining data are available within the Supplementary Data or Source Data file. Source data are provided with this paper.
References
Gajjar, A. et al. Outcomes by clinical and molecular features in children with medulloblastoma treated with risk-adapted therapy: results of an international phase III trial (SJMB03). J. Clin. Oncol. 39, JCO2001372 (2021).
Northcott, P. A. et al. The whole-genome landscape of medulloblastoma subtypes. Nature 547, 311–317 (2017).
Cavalli, F. M. G. et al. Intertumoral Heterogeneity within Medulloblastoma Subgroups. Cancer Cell 31, 737–754.e6 (2017).
Cho, Y. J. et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J. Clin. Oncol. 29, 1424–1430 (2011).
Schwalbe, E. C. et al. Molecular and clinical heterogeneity within MYC-family amplified medulloblastoma is associated with survival outcomes: A multicenter cohort study. Neuro-Oncol. 27, 222–236 (2024).
Gajjar, A. J. & Robinson, G. W. Medulloblastoma-translating discoveries from the bench to the bedside. Nat. Rev. Clin. Oncol. 11, 714–722 (2014).
Dhall, G. et al. Outcome of children less than three years old at diagnosis with non-metastatic medulloblastoma treated with chemotherapy on the “Head Start” I and II protocols. Pediatr. Blood Cancer 50, 1169–1175 (2008).
Hill, R. M. et al. Time, pattern, and outcome of medulloblastoma relapse and their association with tumour biology at diagnosis and therapy: a multicentre cohort study. Lancet Child Adolesc. Heal. 4, 865–874 (2020).
Huybrechts, S. et al. Prognostic clinical and biologic features for overall survival after relapse in childhood medulloblastoma. Cancers 13, 53 (2020).
Johnston, D. L. et al. Survival Following Tumor Recurrence in Children With Medulloblastoma. J. Pediatr. Hematol.solOncol. 40, e159–e163 (2018).
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Morrissy, A. S. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016).
Borgenvik, A. et al. Dormant SOX9-positive cells facilitate MYC-driven recurrence of medulloblastoma. Cancer Res 82, 4586–4603 (2022).
Riemondy, K. A. et al. Neoplastic and immune single-cell transcriptomics define subgroup-specific intra-tumoral heterogeneity of childhood medulloblastoma. Neuro-Oncol. 24, 273–286 (2021).
Hovestadt, V. et al. Resolving medulloblastoma cellular architecture by single-cell genomics. Nature 572, 74–79 (2019).
Williamson, D. et al. Medulloblastoma group 3 and 4 tumors comprise a clinically and biologically significant expression continuum reflecting human cerebellar development. Cell Rep. 40, 111162 (2022).
Richardson, S. et al. Emergence and maintenance of actionable genetic drivers at medulloblastoma relapse. Neuro-Oncol. 24, 153–165 (2021).
Bakhshinyan, D. et al. Temporal profiling of therapy resistance in human medulloblastoma identifies novel targetable drivers of recurrence. Sci. Adv. 7, eabi5568 (2021).
Barouch-Bentov, R. & Sauer, K. Mechanisms of drug resistance in kinases. Expert Opin. Investig. Drugs 20, 153–208 (2011).
Pribluda, A., Cruz, C. C. dela & Jackson, E. L. Intratumoral heterogeneity: from diversity comes resistance. Clin. Cancer Res 21, 2916–2923 (2015).
Garcia, T. B. et al. Increased HDAC activity and c-MYC expression mediate acquired resistance to wee1 inhibition in acute leukemia. Front. Oncol. 10, 296 (2020).
Moncharmont, C. et al. Targeting a cornerstone of radiation resistance: cancer stem cell. Cancer Lett. 322, 139–147 (2012).
Pajonk, F., Vlashi, E. & McBride, W. H. Radiation resistance of cancer stem cells: the 4 R’s of radiobiology revisited. Stem Cells 28, 639–648 (2010).
Trevino, A. E. et al. Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 184, 5053–5069.e23 (2021).
Eze, U. C., Bhaduri, A., Haeussler, M., Nowakowski, T. J. & Kriegstein, A. R. Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat. Neurosci. 24, 584–594 (2021).
Zhang, L. et al. Single-cell transcriptomics in medulloblastoma reveals tumor-initiating progenitors and oncogenic cascades during tumorigenesis and relapse. Cancer Cell 36, 302–318.e7 (2019).
Luo, Z. et al. Human fetal cerebellar cell atlas informs medulloblastoma origin and oncogenesis. Nature 612, 787–794 (2022).
Hendrikse, L. D. et al. Failure of human rhombic lip differentiation underlies medulloblastoma formation. Nature 609, 1021–1028 (2022).
Fan, X. et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res 28, 730–745 (2018).
Smith, K. S. et al. Unified rhombic lip origins of group 3 and group 4 medulloblastoma. Nature 609, 1012–1020 (2022).
Wingate, R. J. T. The rhombic lip and early cerebellar development. Curr. Opin. Neurobiol. 11, 82–88 (2001).
Wang, R. et al. Single-cell dissection of intratumoral heterogeneity and lineage diversity in metastatic gastric adenocarcinoma. Nat. Med. 27, 141–151 (2021).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Shih, D. J. et al. Cytogenetic prognostication within medulloblastoma subgroups. J. Clin. Oncol. 32, 886–896 (2014).
Okonechnikov, K. et al. Comparison of transcriptome profiles between medulloblastoma primary and recurrent tumors uncovers novel variance effects in relapses. Acta Neuropathol. Commun. 11, 7 (2023).
Zuo, W.-F. et al. Heat shock proteins as hallmarks of cancer: insights from molecular mechanisms to therapeutic strategies. J. Hematol. Oncol. 17, 81 (2024).
Kunachowicz, D. et al. Heat shock proteins, a double-edged sword: significance in cancer progression, chemotherapy resistance and novel therapeutic perspectives. Cancers 16, 1500 (2024).
Cho, Y. S. et al. Discovery and evaluation of clinical candidate idh305, a brain penetrant mutant IDH1 inhibitor. ACS Med. Chem. Lett. 8, 1116–1121 (2017).
Jakob, C. G. et al. Novel modes of inhibition of wild-type isocitrate dehydrogenase 1 (idh1): direct covalent modification of His315. J. Med. Chem. 61, 6647–6657 (2018).
Dunican, D. S. et al. Bivalent promoter hypermethylation in cancer is linked to the H327me3/H3K4me3 ratio in embryonic stem cells. BMC Biol. 18, 25 (2020).
Blanco, E., González-Ramírez, M., Alcaine-Colet, A., Aranda, S. & Croce, L. D. The bivalent genome: characterization, structure, and regulation. Trends Genet 36, 118–131 (2020).
Macrae, T. A., Fothergill-Robinson, J. & Ramalho-Santos, M. Regulation, functions and transmission of bivalent chromatin during mammalian development. Nat. Rev. Mol. Cell Biol. 24, 6–26 (2023).
Bunt, J. et al. OTX2 sustains a bivalent-like state of OTX2-bound promoters in medulloblastoma by maintaining their H3K27me3 levels. Acta Neuropathol. 125, 385–394 (2013).
Russo, M. et al. Cancer drug-tolerant persister cells: from biological questions to clinical opportunities. Nat. Rev. Cancer 24, 694–717 (2024).
Pu, Y. et al. Drug-tolerant persister cells in cancer: the cutting edges and future directions. Nat. Rev. Clin. Oncol. 20, 799–813 (2023).
Okonechnikov, K. et al. Oncogene aberrations drive medulloblastoma progression, not initiation. Nature 1–11 https://doi.org/10.1038/s41586-025-08973-5 (2025).
Carey, B. W., Finley, L. W., Cross, J. R., Allis, C. D. & Thompson, C. B. Intracellular alpha-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2015).
Baksh, S. C. & Finley, L. W. S. Metabolic Coordination of Cell Fate by α-Ketoglutarate-Dependent Dioxygenases. Trends Cell Biol. 31, 24–36 (2021).
Alzial, G. et al. Wild-type isocitrate dehydrogenase under the spotlight in glioblastoma. Oncogene 41, 613–621 (2022).
Dang, D. et al. Isocitrate dehydrogenase 1 primes group-3 medulloblastomas for cuproptosis. Cancer Cell https://doi.org/10.1016/j.ccell.2025.04.013 (2025).
Calvert, A. E. et al. Cancer-Associated IDH1 Promotes Growth and Resistance to Targeted Therapies in the Absence of Mutation. Cell Rep. 19, 1858–1873 (2017).
Zarei, M. et al. IDH1 inhibition potentiates chemotherapy efficacy in pancreatic cancer. Cancer Res 84, 3072–3085 (2024).
Zhang, C. et al. Wild-type IDH1 maintains NSCLC stemness and chemoresistance through activation of the serine biosynthetic pathway. Sci. Transl. Med. 15, eade4113 (2023).
Vaziri-Gohar, A. et al. Limited nutrient availability in the tumor microenvironment renders pancreatic tumors sensitive to allosteric IDH1 inhibitors. Nat. Cancer 3, 852–865 (2022).
Jones, C. L., Inguva, A. & Jordan, C. T. Targeting energy metabolism in cancer stem cells: progress and challenges in leukemia and solid tumors. Cell Stem Cell 28, 378–393 (2021).
Intlekofer, A. M. & Finley, L. W. S. Metabolic signatures of cancer cells and stem cells. Nat. Metab. 1, 177–188 (2019).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Tanigawa, Y., Dyer, E. S. & Bejerano, G. WhichTF is functionally important in your open chromatin data?. PLoS Comput. Biol. 18, e1010378 (2022).
Xie, Z. et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44, W90–W97 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform 14, 128 (2013).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10, 1523 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov 2, 100141 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Team, T. B. et al. Bioconda: sustainable and comprehensive software distribution for the life sciences. Nat. Methods 15, 475–476 (2018).
Leprevost, F. et al. BioContainers: an open-source and community-driven framework for software standardization. Bioinformatics 33, 2580–2582 (2017).
Korotkevich, G. et al. Fast gene set enrichment analysis. bioRxiv 060012 https://doi.org/10.1101/060012 (2021).
Liberzon, A. et al. The Molecular Signatures Database Hallmark Gene Set Collection. Cell Syst. 1, 417–425 (2015).
Dolgalev, I. msigdbr: MSigDB Gene Sets for Multiple Organisms in a Tidy Data Format. CRAN: Contrib. Packag. https://doi.org/10.32614/cran.package.msigdbr (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Fu, R. et al. Spatial transcriptomic analysis delineates epithelial and mesenchymal subpopulations and transition stages in childhood ependymoma. Neuro-Oncol. 25, 786–798 (2022).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Hu, Y. & Smyth, G. K. ELDA: Extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Nemkov, T., Reisz, J. A., Gehrke, S., Hansen, K. C. & D’Alessandro, A. High-throughput metabolomics: isocratic and gradient mass spectrometry-based methods. Methods Mol. Biol. 1978, 13–26 (2019).
Pang, Z. et al. MetaboAnalystR 4.0: a unified LC-MS workflow for global metabolomics. Nat. Commun. 15, 3675 (2024).
Acknowledgements
The authors wish to thank the University of Colorado School of Medicine Metabolomics Core, and the Bioinformatics and Biostatistics, and the Human Immune Monitoring Shared Resource for their contributions to this manuscript. This study was partly supported by the National Institutes of Health Cancer Center Support Grant P30CA046934, and the following shared resources: Small Animal Image-Guided Radiation Therapy (RRID:SCR_021996), Flow Cytometry (RRID: SCR_022035), Mass Spectrometry Metabolomics Facility (RRID: SCR_021988), Bioinformatics and Biostatistics (RRID: SCR_021983), Genomics Shared Resource (RRID: SCR_021984), and the Human Immune Monitoring Shared Resource (RRID:SCR_021985). We acknowledge ABiosciences for their contribution to the analysis of scMultiome patient samples. This study was supported by the Morgan Adams Foundation (BV, DW, JD, RV), the Cancer League of Colorado (BV), and NIH R01 NS091219 (RV).
Author information
Authors and Affiliations
Contributions
B.V. and R.V. conceived the study and wrote the manuscript. B.V., D.W., J.D., P.B., and A.G. wrote the methods. D.W. performed analysis on CUT&RUN studies. A.G. analyzed scRNAseq. J.D. and B.V. analyzed the scMultiome dataset and performed enrichment analysis Fig. 5G, H. A.P. and B.B. performed in vivo mouse injection studies. B.V. and K.S. performed in vitro studies. A.D. and N.F. provided the spatial transcriptomics data and analysis. P.C.B. and N.D. performed analysis of the scRNA-seq trajectories.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Veo, B., Wang, D., DeSisto, J. et al. Single-cell multi-omics identifies metabolism-linked epigenetic reprogramming as a driver of therapy-resistant medulloblastoma. Nat Commun 16, 10470 (2025). https://doi.org/10.1038/s41467-025-65466-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65466-9