Abstract
Tumor heterogeneity drives drug resistance and relapse, influencing immune evasion and tumor progression. While intratumor heterogeneity has been extensively studied at the genomic level, its functional outcomes and interactions with the tumor microenvironment remain underexplored. In contrast, the functional outcome of heterogeneity and the interplay with the tumor microenvironment have not been addressed. In this study, we integrate multi-region spatial MS-based proteomics of 280 tumor regions, exome sequencing, and imaging to investigate spatial proteomic heterogeneity in breast cancer. Our findings reveal increased proteomic heterogeneity with tumor progression, independent of genomic heterogeneity but closely associated with microenvironmental differences. Integration with immune and stromal imaging highlighted a dynamic interplay where low-grade tumors exhibit constrained immune infiltration, and upon progression to higher grades, macrophages and T cells infiltrate. However, anti-inflammatory pathways involving kynurenine and prostaglandins are more highly expressed in infiltrated regions, suggesting that anti-tumorigenic activities are inhibited. Integration with the global protein network provides potential targetable mediators of immune evasion in breast cancer that can serve as the basis for future development of personalized breast cancer therapies.
Similar content being viewed by others
Introduction
Breast cancer inter-patient diversity has been extensively explored across multiple dimensions, including genomics, transcriptomics, and proteomics1,2,3,4,5,6. Integrated genomic and transcriptomic analyses initially delineated five intrinsic breast cancer subtypes: luminal A, B, HER2-amplified, basal-like, and normal-like4. These subtypes align with the conventional clinical classification based on hormone receptor expression (estrogen and progesterone) and the HER2 receptor tyrosine kinase7. Consequently, luminal tumors are characterized by hormone receptor positivity (HR+), HER2 tumors exhibit amplification of the HER2 oncogene, and triple-negative breast cancer (TNBC) tumors lack expression of these receptors. Over time, these subtypes have been further refined to associate them with their characteristic driver mutations, incorporate additional triple-negative subtypes, or subdivide them into additional distinct clusters5,7. At the proteomic level, multiple studies showed a low correlation between mRNA and protein8,9, which ultimately impacts tumor classification1,2,10,11.
Despite the extensive knowledge of inter-patient variability in breast cancer, the clinical landscape faces challenges, indicating that a straightforward classification approach falls short. Firstly, there is frequent discordance between clinical classification, which relies on immunohistochemistry, and RNA-based classification, resulting in classification ambiguity12,13. Furthermore, pathological examinations frequently reveal subtype transitions between primary tumors and metastases14,15. These phenomena occur due to internal tumor heterogeneity and metastatic dissemination of cells, which may have been overlooked during the initial diagnosis. Alternatively, these phenomena could be linked to cancer cell plasticity and their ability to adapt to new microenvironments.
Prior investigation of intra-tumor heterogeneity (ITH) primarily focused on genomic landscapes, unraveling clonal heterogeneity within single tumors, and elucidating evolutionary processes during cancer progression16,17. Moreover, previous studies showed how heterogeneity can impact the tumor microenvironment. For example, high genomic ITH in melanoma and breast cancer was shown to be associated with reduced anti-tumor immune response, presumably due to inhibitory signals in some cancer clones18,19. While the genetic events form the primary underlying factor for tumor evolution, the tumor phenotype reflects the impact of these events, combined with the impact of other factors such as epigenetic changes and interactions with the tumor microenvironment (TME). Non-genetic heterogeneity reflects the cancer cell plasticity and its adaptation to the tumor metabolic, immune, and stromal microenvironments. Therefore, analysis of non-genetic heterogeneity is critical to better understanding immune evasion mechanisms and therapy resistance, providing a basis for improved patient-tailored therapy.
In this work, we aimed to tackle the question of existing ITH in breast cancer primary tumors, focusing on the causes and consequences of proteomic ITH. We performed microscopy-guided laser microdissection followed by MS-based proteomics of multiple regions per tumor. We identified proteomic heterogeneity, which did not correlate with the genomic heterogeneity of these tumors. Integration with multiplexed imaging showed the marked impact of the TME on the tumor proteome.
Results
To comprehensively profile the heterogeneous proteomic landscape in breast cancer, we developed an analytical framework to examine the proteomic compositions of distinct regions exhibiting variable histopathological characteristics within individual tumors (Fig. 1A). We analyzed 280 microdissected tumor regions from a cohort of 33 breast tumors (Supplementary Data 1). Sections from each tumor block were stained for three key breast cancer receptors: estrogen receptor (ER), progesterone receptor (PR), HER2 (ERBB2), and pan-cytokeratin (panCK). AI-based cell quantification and pathological mapping defined discrete regions within tumors based on their receptor expression profiles. Subsequently, we laser-capture microdissected and analyzed each of these regions as distinct samples. These samples comprised 19 normal-adjacent tissue samples, 12 samples of ductal carcinoma in-situ (DCIS), 10 Grade 1, 139 Grade 2, and 100 Grade 3 tumor regions (Supplementary Data 1). We identified 8263 proteins with relatively uniform receptor expression patterns within each region and overall high cancer cell purity (median = 0.8), as assessed by the ESTIMATE algorithm20 (Supplementary Fig. 1A, Supplementary Data 1).
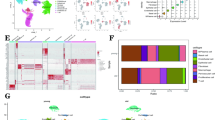
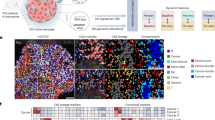
A Schematic representation of the multi-regional proteomic workflow. Proteomic analysis was performed on FFPE tumor sections. H&E staining, Immunohistochemistry of ER, PR, HER2, pan-cytokeratin, and immunofluorescence (IF) on consecutive sections, followed by laser-capture microdissection of histopathologically defined regions from each tumor. Extracted proteins were trypsin-digested, labeled with TMT10plex, mixed with an internal standard, and separated using high pH reverse phase fractionation. Peptide samples were analyzed on the Q Exactive HF MS. Created in BioRender. Neta, H. (2025) https://BioRender.com/by0z6ya. B Volcano plot shows significantly changing proteins between Grade 2 and Grade 3 across both HR+ and TNBC regions. For each protein, a Linear Mixed Effects Model (LMM) was fitted using the “lme4” package in R, with Grade as a fixed effect and random intercepts for patient. Fold changes (Grade 3 vs Grade 2) were extracted from the model estimates. Statistical significance was assessed using Wald t-tests, and raw p-values were adjusted for multiple testing via False Discovery Rate (FDR) method using “qvalue” (v2.38.0) package. Proteins with FDR-adjusted p < 0.05 were considered significant. Proteins upregulated in Grade 3 are colored purple, and those upregulated in Grade 2 are colored green. Selected protein labels are displayed; the full list is available in Supplementary Data 2. Bar plots showing enriched pathways identified by 1D enrichment analysis (FDR q-value < 0.02, two-sided Mann–Whitney test) in (C) triple-negative breast cancer (TNBC) tumor regions and (D) hormone receptor-positive (HR+) tumor regions. The complete enrichment lists are provided in Supplementary Data 2. E Volcano plot shows significantly changing proteins between TNBC and HR+, analyzed using the same methods and statistical tests described in (B). Proteins upregulated in TNBC are shown in orange, and those upregulated in HR+ are shown in light blue. Selected protein labels are displayed; the full list is available in Supplementary Data 2. F Bar plot shows enrichment analysis of HR+ and TNBC in Grade 3 (1D Enrichment analysis with FDR q < 0.02, two-sided Mann–Whitney test). The complete enrichment list is provided in Supplementary Data 2. G Bar plot shows enrichment analysis of HR+ and TNBC in Grade 2 (1D Enrichment analysis with FDR q < 0.02). The complete enrichment list is provided in Supplementary Data 2. Asterisks next to enrichment pathway names were edited for visualization purposes.
Spatial proteomics recapitulates known differences between tumor subtypes and Grades
As an initial assessment of the data, we performed a series of statistical tests to compare tumor grades and subtypes. However, rather than comparing bulk tumors, as previously done, these were performed on tumor regions based on their histopathological characteristics. Over 2400 proteins significantly differed between cancer and normal-adjacent tissues (Supplementary Fig. 1B, Supplementary Data 2). As expected, cancer regions exhibited elevated levels of proliferation markers such as KI67, PCNA, and DNA polymerase, as well as metabolic enzymes (e.g., SCD, PYCR2, GLUL), potentially supporting increased biosynthetic demands. Enrichment analysis showed high expression of nucleotide biosynthesis pathways, DNA repair, chromosome organization and mitochondrial translation in the cancer samples and lipoprotein metabolism, cell adhesion, and humoral immune response in the normal controls (Supplementary Fig. 1C, Supplementary Data 2). In agreement with the known higher proliferation rate of Grade 3 tumors, comparing Grade 3 and Grade 2 proteomes showed high expression of PCNA, MCM proteins, and nucleotide metabolism proteins in both HR+ and TNBC regions in Grade 3 (Fig. 1B, Supplementary Data 2) and lower level of the cell cycle regulator WEE1. Subtype-separated enrichment analysis showed that Grade 3 TNBC also highly expressed MCM complex proteins, DNA replication, pyrimidine metabolism, and proteins related to antigen processing and presentation (Fig. 1C, Supplementary Data 2). HR+ Grade 3 tumors also highly expressed proteins related to antigen processing and presentation, proteasomal complex proteins and proteins involved in DNA biosynthesis (Fig. 1D, Supplementary Data 2).
Comparing between subtypes showed that TNBC regions exhibited higher expression of proteins of the one carbon metabolism pathway (PHGDH and PSAT), along with increased levels of PODXL, an anti-adhesive cell surface sialomucin that promotes tumor cell detachment and enhances migratory potential. (Fig. 1E, Supplementary Data 2). HR+ regions displayed elevated levels of AGR2 and AGR3, both known estrogen receptor (ER)–associated proteins, along with KRT18 and KRT19, which are typical markers of HR+ breast cancer. Grade-separated enrichment analysis showed more substantial differences in Grade 3 tumors, with enrichment of ribosomes and chromatin proteins in TNBC and higher PI3K and typeI interferon in HR+ regions (Fig. 1F, G, Supplementary Data 2). Altogether, these initial comparisons provide a proof-of-concept for the region-based spatial approach as we recapitulate known changes in cancer development and progression and between cancer subtypes and provide a resource of proteome-wide changes that can be further investigated.
Receptor-ITH is reduced, while proteomic-ITH is elevated in high-grade tumors
To uncover the drivers of cancer heterogeneity, we performed unsupervised clustering of tumor regions based on their global proteome profiles. The analysis revealed a distinct hierarchical separation based on tumor grade, receptor expression patterns, and patient of origin. Tumor grade emerged as the most prominent separator, primarily distinguishing Grade 2 from Grade 3 tumors (Fig. 2A). At the secondary level, receptor expression patterns effectively stratified HR+ , TNBC, and HER2+ regions (Fig. 2B). The tertiary level of stratification correlated with the patient of origin, where regions from the same patient generally clustered together (Fig. 2C). Inter-patient variance, however, may have been partially confounded by the TMT set and experimental batch (Supplementary Fig. 1D). We therefore focused our analyses on intra-tumor comparisons, comparing samples within the same TMT set and experimental batch (Supplementary Fig. 1A). Intriguingly, we found multiple TNBC regions clustered with HR+ tumors, indicative of heterogeneous receptor expression within single tumors. While some tumors, particularly Grade 2 tumors, displayed receptor diversity, Grade 3 tumors demonstrated higher receptor homogeneity (Fig. 2A–C).
A Unsupervised clustering of tumor regions, using a tree-and-leaf representation based on a modified minimum variance method, in which each node represents a tumor region. The tree-and-leaf branches are constructed based on Euclidean distances, and the spatial localization of the branches presents the cluster separation. Presented dendrograms are color-coded based on tumor grade. Visualized using “TreeAndLeaf” (v1.6.1), “igraph” (v1.4.2), and “RedeR” (v2.1.4) R packages. B Same as panel A, with coloring based on receptor expression. C Same as panel A, with coloring based on patient identity. D Schematic representation of the receptor-intratumor heterogeneity score (Receptor-ITH) based on immunohistochemistry (IHC) staining and calculated per tumor. E Boxplot showing the distribution of Receptor-ITH scores in Grade 2 (n = 20) and Grade 3 (n = 10) tumors. Boxplots show the median and interquartile range, with individual patient data points overlaid as jittered dots. Statistical comparison between groups was performed using a two-sided Student’s t-test, t(28) = −1.62, p = 0.12, 95% CI [−0.471, 0.0556], Cohen’s d = 0.63. F Boxplots show the distribution of Shannon’s Diversity Index in Grade 2 (n = 141) and Grade 3 (n = 221) tumors from the METABRIC cohort21. Individual patient data points overlaid as jittered dots. Statistical significance was assessed using a two-sided Student’s t-test, t(350) = −2.96, p = 0.003, 95% CI [−0.1158, −0.0235], Cohen’s d = 0.32. G Schematic representation of the proteomic intratumor heterogeneity score (P-ITH) calculated per tumor per TMT set. H Boxplots showing the distribution of P-ITH scores in Grade 2 (n = 16) and Grade 3 (n = 12) tumors. Individual patient data points overlaid as jittered dots. Statistical comparison was performed using a two-sided Student’s t-test, t(26) = 2.25, p = 0.03, 95% CI [0.0037, 0.0813], Cohen’s d = −0.86. I Correlation heatmap of tumor features: grade, cellularity, tumor size, age, Receptor-ITH, P-ITH, and disease-free survival (DFS). Colors indicate Pearson’s correlation coefficients, and statistically significant correlations are marked with asterisks: *p < 0.05, **p < 0.01, ***p < 0.001. The “corrplot” (v0.95) R package was used for visualization.
To determine the statistical significance of the grade-related receptor heterogeneity, we defined a receptor intratumor heterogeneity score named Receptor-ITH. This score accounts for the number of different receptor expression patterns co-existing in a single tumor and the fraction of the most dominant pattern (Fig. 2D). In this way, higher Receptor-ITH occurs when more patterns co-exist and when these are more evenly distributed. A comparison of Receptor-ITH between Grades 2 and 3 showed a trend toward reduced ITH with cancer progression (p = 0.12; Fig. 2E, Supplementary Data 3). Since the number of tumors in our cohort limited our statistical power, we validated our results using imaging mass cytometry data from an independent cohort of 385 breast tumors21. Examining HR+ and TNBC cells, we calculated Shannon’s Diversity Index, which quantifies diversity based on the number of distinct populations and their relative distributions. In agreement with our results, the examination of 385 tumors showed significantly reduced receptor diversity in high-grade compared to lower-grade tumors (p = 0.003; Fig. 2F, Supplementary Data 3). These results suggest potential selective pressures that change tumor subtypes, even in the primary site, before any treatment.
Our clustering results suggested discordance between the receptor expression patterns and the overall proteomic patterns, as often protein-based clusters included regions with distinct receptor expression (Fig. 2C). To quantify these differences, we defined a proteomic intratumor heterogeneity score, or P-ITH score, based on the pair-wise protein Pearson’s correlations (Fig. 2G, Supplementary Data 3). Surprisingly, compared to the reduction observed in Receptor-ITH with increasing grade, P-ITH exhibited a significant increase in Grade 3 tumors compared to Grade 2 tumors (p = 0.03; Fig. 2H). Global Pearson’s correlation analyses showed a significant positive correlation between Receptor-ITH and disease-free survival (DFS) (r = 0.72, p = 0.009), alongside a negative correlation with grade (r = −0.29, p = 0.009). Conversely, P-ITH demonstrated a positive correlation with grade (r = 0.6, p = 0.046) and average tumor cellularity (r = 0.72, p = 0.003). In line with these observations, P-ITH negatively correlated with DFS and was elevated in patients who experienced metastatic disease or relapse (Fig. 2I, Supplementary Fig. 2). In summary, these results underscore that tumor functionalities, as reflected by global proteome profiles, diverge from the receptor expression patterns, offering valuable insights into the complex interplay of these factors in tumorigenesis.
P-ITH is impacted by interactions with the tumor microenvironment
Aiming to decipher the underlying causes of P-ITH, we first examined the potential impact of genomic ITH on the downstream proteomic profiles. Using consecutive FFPE tumor slides, we laser-microdissected the same regions analyzed proteomically and performed exome sequencing. Employing a focused exome library, we profiled 5227 genes across 47 tumor regions extracted from four HR+ tumors. Variant calling indicated comparable results to other breast cancer genomic datasets (e.g., TCGA4) (Supplementary Fig. 3A-C, Supplementary Data 4). In agreement with previous studies, inter-patient differences were more dominant than those between regions within single patients22,23. The vast majority of mutated genes were patient-specific (Supplementary Fig. 3D). Nevertheless, genes that were either shared across all regions of a single patient or common between patients included known breast cancer driver mutations cataloged in the COSMIC database24, such as PIK3CA and TP53 (Supplementary Fig. 3). To further examine whether the proteomic ITH corresponds to the genomic ITH, we plotted the pair-wise ranked cosine similarity between regions within each tumor (Fig. 3A–D, Supplementary Data 4). All tumors exhibited consistently low and non-significant correlations, with correlation coefficients |r| < 0.18 and p > 0.26, implying that the genomic ITH does not impact P-ITH.
A–D Pair-wise correlations between genomic and proteomic ranked cosine similarities across regions within each tumor in four ER+ breast cancer patients. For each patient, the solid line indicates the best-fit linear regression, with the regression equation, Pearson’s r, and p-value shown. The grey shaded area represents the 95% confidence interval of the fit. Pathway enrichment analysis was performed using a two-sided Mann-Whitney test in most constant (E) and most variable (F) proteins, defined by the 10% lowest and highest median absolute deviation (MAD) scores, respectively. Circle size reflects the number of proteins per enriched pathway, and color scale indicates FDR q-values. Pathway names marked with an asterisk (*) were edited for visualization. The complete enrichment results are provided in Supplementary Data 5.
Given that genomic heterogeneity did not directly influence proteomic heterogeneity (P-ITH), we explored alternative drivers of P-ITH by leveraging the proteomic data to identify proteins that distinguish tumor regions within the same patient across multiple tumors. For each protein, we calculated the median absolute deviation (MAD) score in each patient to pinpoint the most and least variable proteins, corresponding to the highest and lowest MAD scores, respectively. Enrichment analysis of the most constant proteins showed processes related to translation, protein folding, turnover, and trafficking (Fig. 3E; Supplementary Fig. 4A, Supplementary Data 5). We defined these as the core cancer proteome not subjected to spatial plasticity; on the opposite side, enrichment analysis of the most variable proteins identified multiple immune-related processes, primarily involved in antigen presentation (e.g., HLA molecules), histone proteins, proteins that may reflect vicinity to blood vessels (e.g., HBB), and metabolic processes (Fig. 3F; Supplementary Fig. 4B, Supplementary Data 5). These results suggest that the constant cancer proteome primarily comprises intrinsic factors that reflect the cancer cells’ basic proliferative and biosynthetic requirements. In contrast, the heterogeneous proteome reflects cellular plasticity shaped by the interactions with the TME.
An analytical pipeline of multiplexed imaging to define the TME landscape
Identifying highly variable proteins has underscored the potential impact of interactions with the TME on the proteome. To examine the link between the proteome of tumor regions and the features of the TME, we performed multiplexed imaging of TME components: T cells (CD3+), macrophages (CD68+), fibroblasts (αSMA+), endothelial cells (CD31+) and cancer cells (panCK) (Fig. 4A). We developed a computational pipeline to transfer the proteomic regions of interest (P-ROIs) from the images of H&E stained and immunohistochemistry (IHC)-stained tissue sections to the consecutive immuno-fluorescent images. We then developed two deep-learning-based analytical pipelines for quantifying cell components and blood vessels (see Methods). These analyses were conducted at three levels: the entire tumor slide, the P-ROI, and the region’s neighborhood surrounding the P-ROI (Fig. 4B, Supplementary Data 6).
A Schematic representation of the multiplexed imaging workflow. Consecutive slides were stained with two antibody panels and DAPI. Quantification was composed of two workflows. The first workflow was used for cell quantification and involved cell segmentation, followed by cell classification and quantification. The second workflow, for blood vessel quantification, was pixel-based on and quantified the area of blood vessels in the tumors. B Schematic representation of the analytical pipeline for multilayer proteomics and imaging integration. Scale = 200 µm. C–F Dot plots show the quantification of TME components in whole-slide images: CD3+ T cells percent (C), CD68+ macrophages percent (D), αSMA+ fibroblasts percent (E), and blood vessels (BV) area percent. Each dot represents an individual slide from a patient. Patient identities are color-coded, and the same slide order is maintained across all graphs. Data represent n = 11 patients with a total of 47 slides. G Correlation heatmap of percent CD3+ cells, CD68+ cells, BV, αSMA+ cells, and pan-cytokeratin (panCK+) cancer cells in the whole tissue analysis. Values represent average percentages across multiple FFPE tissue blocks per patient. Colors indicate Spearman’s correlation coefficients, and statistically significant correlations are marked with asterisks: *p < 0.05, **p < 0.01, ***p < 0.001. The “corrplot” (v0.95) R package was used for visualization. H Boxplots depict the distribution of percent CD3 + T cells and CD68+ macrophages in BV-Low (light pink, n = 61) and BV-High (coral, n = 60) regions. Each box shows the median, interquartile range, and whiskers indicate data variability, with individual data points overlaid as jittered dots. Data represent cell type percentages in P-ROI regions. Statistical comparisons were performed using a two-sided Student’s t-test for CD3 (t(119) = 2.23, p = 0.028, 95% CI [0.43, 7.18], Cohen’s d = 0.41), and a two-sided Welch’s t-test for CD68 due to unequal variance (Welch’s t(108.89) = 3.53, p = 6 × 10-4, 95% CI [1.17, 4.15], Glass’s Δ = 0.78). Linear mixed-effects models (LMMs) accounting for patient as a random effect were also applied to obtain adjusted p-values. I Violin plots combined with boxplots show the distribution of percent CD3+ , CD68+ , and αSMA+ cells in Grade 2 vs. Grade 3 tumors at the whole slide level. Boxplots indicate the median (center line), interquartile range (IQR; box limits), and whiskers extending to 1.5 × IQR; points beyond whiskers represent outliers. Two-sided Student’s t-tests were performed to compare T cells (t(44) = −3.31, p = 0.002, Cohen’s d = −0.98; Grade 2, n = 20; Grade 3, n = 26), macrophages (t(44) = −2.41, p = 0.02, Cohen’s d = −0.72; Grade 2, n = 20; Grade 3, n = 26), and fibroblasts (t(45) = 6.03, p = 2.8 ×10-7,Cohen’s d = 1.77; Grade 2, n = 21; Grade 3, n = 26). Linear mixed model (LMMs) accounting for the patient as a random effect were applied to obtain adjusted p-values. J Violin plots combined with boxplots show the distribution of averaged T-cell, fibroblast, and macrophage percentages per METABRIC patient in the imaging mass cytometry dataset21. A minimum threshold of 500 single cells per patient was applied to filter out insignificant observations. Patients with zero values and outliers (defined using the IQR) were excluded. Two-sided Student’s t-tests were used to compare CD3+ T cells (t(246) = −1.78, p = 0.077, Cohen’s d = −0.23), and αSMA+ fibroblasts (t(294) = 2.32, p = 0.021, Cohen’s d = 0.28) between Grade 2 (n = 141) and Grade 3 (n = 211) patients. For CD68+ macrophages, a two-sided Welch’s t-test was applied due to unequal variances (t(279.74) = −3.472, p = 6×10-4, Glass’s Δ = −0.345).
In total, we spatially profiled 47 tumor slices, encompassing 121 proteomic regions and their respective neighborhoods, all originating from 11 patients within our cohort. Initially, we quantified cell numbers and blood vessels, irrespective of the P-ROI, to assess the involvement of all TME components. Quantitation of T cells, macrophages, blood vessels, and fibroblasts at the slide level unveiled a broad spectrum of T-cell infiltration levels, ranging from “cold” tumors with no T cells to tumors with >40% CD3-positive cells (Fig. 4C, Supplementary Data 6). The numbers of macrophages (CD68+) exhibited a variation from 0.5% to 18% in the entire slide, while the count of fibroblasts (αSMA+) demonstrated the widest range, spanning from 2.4% to 64.7% (Fig. 4D, E, Supplementary Data 6). Our calculation of the blood vessel area indicated an average of 1-2% of the total tissue area, with only two tissue sections displaying higher percentages (Fig. 4F, Supplementary Data 6). Examining the correlations among these TME components revealed a general trend: immune components, such as CD68+ macrophages and CD3+ T cells, were positively correlated (Spearman’s ρ = 0.38, p = 0.25), although this association was not statistically significant. However, panCK+ epithelial cells were significantly negatively correlated with both macrophages (Spearman’s ρ = −0.64, p = 0.034) and T cells (Spearman’s ρ = −0.91, p < 0.001). In contrast, αSMA+ fibroblasts were negatively correlated with CD3+ T cells (Spearman’s ρ = −0.66, p = 0.028; Fig. 4G). Additionally, a trend of positive correlation was observed between blood vessels (BV) and fibroblasts (Spearman’s ρ = 0.35). These analyses reaffirm the previously published positive impact of cancer-associated fibroblasts (CAFs) on vascular growth25 and the negative correlation between tissue fibrosis and immune infiltration26. We found insignificant correlations between immune infiltrates and blood vessels, potentially due to the skewed distribution of percent blood vessel values. However, when tumors were separated into blood-vessel-high (BV-high) and blood-vessel-low (BV-low) groups, BV-low regions showed significantly higher numbers of T cells (p = 0.028) and macrophages (p < 0.001) compared with BV-high regions (Fig. 4H). This observation suggests potential recruitment of T cells and macrophages into regions with fewer blood vessels that might be more hypoxic. Interestingly, analysis of the immune infiltration separated by cancer grade showed higher T-cell and macrophage levels in Grade 3 compared to Grade 2 tumors (T cells: p = 0.076; Macrophages: p = 0.02) alongside fewer fibroblasts (p = 2.774 × 10−7) (Fig. 4I). We further validated these results using imaging mass cytometry data of 358 patients21, which showed higher immune infiltration (T cells: p = 0.077; Macrophages: p = 0.0006) and decreased fibroblasts (p = 0.02) in higher grades (Fig. 4J). These intriguing findings suggest that cancer-associated fibroblasts decrease with cancer progression and immune infiltration increases with progression and tumor diversification.
Examination of the imaging results within the P-ROIs showed that some P-ROIs include T cells, macrophages and fibroblasts. We investigated the extent of contribution of these non-cancer cells to the measured proteome and the proteomic heterogeneity. We used two algorithms to determine the contribution of immune and stromal components to the proteomic profiles and compared these scores to the imaging-based “ground truth”. The ESTIMATE algorithm provides an immune score, stromal score, and tumor purity, and the ConsensusTME algorithm provides additional cell-specific scores. A comparison of immune scores from the two algorithms showed a strong positive correlation (Pearson’s r = 0.87). We used the ConsensusTME algorithm to predict the extent of T cells, macrophages and fibroblasts in the tissues, based on the proteomic data. We found a positive correlation between the predicted T-cell score and the imaging data. However, macrophage and fibroblast scores showed no correlation with the imaging-based observations (Supplementary Fig. 5A–D). Given that the imaging and proteomics are not performed on the same tissue section, some discrepancies are expected, but it also suggests that this algorithm cannot be used to predict the percentages of macrophages and fibroblasts in the entire cohort. The positive correlation between T-cell imaging and the CD3-score allowed extrapolation of the T-cell percentages across all proteomic samples. Overall, we found a median T-cell count of 6.5%, suggesting a low impact on the overall proteomic signal. Despite the relatively low signal of non-cancer cells in the P-ROI, we corrected the proteomic signal to account for tumor purity (see Methods). Recalculating the P-ITH showed a slight reduction in overall P-ITH; however, the differences between tumor grades persisted (Supplementary Fig. 5E). No significant P-ITH differences were observed between subtypes (as defined clinically, Supplementary Fig. 5F). Therefore, we cannot exclude some contribution of non-cancer cells to the overall proteomic signals, but our results show that this contribution is minor.
Region-wise, TME analyses link the proteomic landscape with the tumor vascularization
To better understand the cancer-TME interactions, we delved deeper into these associations at the region level, establishing links between the spatial phenotypes and the proteomic landscape of the tumors. Region-wise quantification of blood vessels included the P-ROIs and the surrounding rings (500 µm), since the P-ROIs were largely devoid of blood vessels. To identify proteins associated with BV%, we performed a Pearson’s correlation analysis and identified 1103 proteins significantly correlated with BV% (Supplementary Data 6), of which 95% showed negative correlations. Among these proteins, we found PIK3CA, IL13RA1, LIMK1, SLC16A1, SOX4, YBX2, MKI67, SCD, CDK1, and ENO2, which are known for their association with hypoxia or breast cancer (Fig. 5A). PIK3CA, frequently mutated or amplified gene in breast cancer, displayed a strong positive correlation with BV%. PIK3CA has been implicated in cancer cell survival, metastasis, and angiogenesis27. Its association with BV% in our data could suggest its potential involvement in tumor vascularization. Similarly, IL13RA1 exhibited a strong positive correlation with BV% (Fig. 5A). In agreement, previous studies have shown IL13RA1 is down-regulated under hypoxic conditions28. LIMK1, which has been shown to contribute to tumor angiogenesis in human breast cancer cells28, demonstrated increased expression in BV-rich regions. In contrast, the vast majority of these proteins showed a negative correlation with BV%, suggesting that hypoxic conditions, or reduced blood vessels in the TME, may drive their upregulation as part of an adaptive response to response to oxygen and nutrient deprivation. Among these proteins we found the hallmark proteins of proliferating cells MKI67, the mitotic cell cycle regulator CDK1. SOX4, a transcription factor known as a regulator of epithelial-mesenchymal transition (EMT) was shown to promote the growth and metastasis of breast cancer29,30. From a metabolic perspective, we observed upregulation of the glycolytic enzyme ENO2 and the lactate transporter SLC16A1 (both key components of the Warburg effect) in BV-low regions, consistent with the expected metabolic adaptation to hypoxia. In line with these changes, we also detected increased expression of SCD, a key enzyme in lipid biosynthesis that supports membrane formation and energy storage in rapidly proliferating cells. Thus, vascular and hypoxic niches within tumors harbor distinct proteomic signatures with implications for tumor progression and therapy response.
A Scatter plots show Pearson’s correlation between log2 protein expression values and blood vessel percentages in the P-ROI and the surrounding ring area. Each dot represents a valid data point. The solid line represents the linear regression fit, with the grey shaded area representing the 95% confidence interval. Pearson’s r, two-sided p-value from a t-test on the correlation coefficient, and FDR-adjusted p-values are shown. B Representative immunofluorescent images after cell segmentation and cell type classification show the spatial distribution of CD3+ T cells (cyan), CD68+ macrophages (magenta), panCK+ cancer cells (green), and other cell populations (red). Among 47 tumor slides from 11 patients, patient BC1 is shown as an example. The immune phenotype classification of each region is indicated. Scale = 200 µm. C Dot plot example of immune phenotypes of BC1 based on percentages of CD3+ T cells and CD68+ macrophages within the P-ROIs and surrounding rings. D Heatmap of 414 significantly changing proteins associated with immune phenotypes, as determined by a three-way ANOVA, not confounded by Grade and molecular subtype. Color scale denotes Tukey’s HSD post-hoc score values Colors represent Tukey’s HSD post-hoc scores. Visualization was generated using the “ComplexHeatmap” (v2.22.0) R package. E Heatmap of 658 significantly changing proteins between immune phenotypes in TNBC P-ROIs. Color scale denotes Tukey’s HSD post-hoc score. F Heatmap of 2822 significantly changing proteins between immune phenotypes in HR+ P-ROIs. Color scale denotes Tukey’s HSD post-hoc score values. Protein network representation of the Kynurenine pathway, Aryl Hydrocarbon Receptor (AHR), and Prostaglandins in TNBC (G) and HR+ (H) subtypes. Each node is divided into four immune-phenotype quadrants, color-coded by Tukey’s HSD post-hoc scores. Protein-protein interactions were obtained from the STRING database, and visualized in Cytoscape (v3.10.0) using the Omics Visualizer app63.
Proteomics of four cancer-immune phenotypes
Quantifying the numbers of T cells and macrophages in the TME provides the extent of immune infiltration. However, this approach lacks spatial information regarding their distribution and localization in the tissue. Previous studies in breast cancer and other cancer types have described three T-cell-related immune phenotypes: T-cell infiltrated (hot tumors), cold tumors (devoid of T-cells), and T-cell excluded or compartmentalized tumors, where large numbers of T-cells are located outside large cancer cell clusters31,32. Examining immune phenotypes at the region level enabled their association with the proteomic landscape of each phenotype. To this end, we quantified T cells and macrophages within each proteomic region and additionally measured cell abundance in a ring surrounding the P-ROI to discriminate between cold tumors and immune-cell-excluded regions. The same pipeline was applied to 121 regions and their surrounding rings for both T cells and macrophages (Supplementary Fig. 6A, B). Combining the T cell and macrophage phenotypes resulted in four main immune phenotypes: T-cell and macrophage infiltrated (‘Infiltrated’), T cell and macrophage cold (Cold), T cell compartmentalized, and macrophage infiltrated (CD3COMPCD68INF), and T-cell cold and macrophage infiltrated (CD3COLDCD68INF) (Fig. 5B, C, Supplementary Fig. 6C, Supplementary Data 6). In agreement with ref. 33, we found that almost all immune infiltrated regions included macrophages, whereas not all of them included T cells. To identify the proteomic determinants of the immune phenotypes, we first performed a three-way ANOVA test to detect the significantly changing proteins, while correcting for the confounding factors of grade and receptor expression (Fig. 5D, Supplementary Data 7). We found 414 proteins significantly associated with the immune subtype but not confounded by grade and subtype. To examine the potential contribution of proteins originating from the T cells and macrophages, we compared the 2289 proteins significantly associated with immune phenotypes (irrespective of grade and subtype) with publicly available signatures of these cell types. Of the T-cell signature proteins, fewer than 0.8% were present in the immune phenotype-significant list (19 proteins). Similarly, fewer than 6.8% of macrophage signature proteins were included in the immune phenotype-significant list (155 proteins, Supplementary Fig. 7A). Interestingly, only 94 macrophage signature proteins and none of the T-cell signature proteins were elevated in the immune-infiltrated regions compared with the cold regions (Supplementary Fig. 7B, Supplementary Data 8). These results suggest that less than 7.6% of the proteomic signal associated with immune differences originates from the immune cells themselves, while the remaining signal likely reflects the impact of the immune cells on the cancer cells.
When we examined the differences within each subtype, we found 2433 and 658 proteins significantly associated with immune phenotype in HR+ and TNBC tumor regions, respectively. Notably, TNBC appeared to be most affected by the infiltration of T cells, as the proteomes of macrophage-infiltrated tumors resembled the proteomes of cold tumors. In contrast, the proteomic changes in HR+ tumors suggested that these tumors are affected by both macrophage- and T-cell infiltration (Fig. 5E, F, Supplementary Data 7). Enrichment analysis showed that infiltrated regions (both TNBC and HR+) highly express antigen presentation proteins and IFN-gamma signaling proteins (Supplementary Fig. 7C-D, Supplementary Data 7). Interestingly, we found a network of proteins involved in arachidonic acid metabolism, linked to the aryl hydrocarbon receptor (AHR) and kynurenine pathway proteins, which were more highly expressed in infiltrated regions of both TNBC and HR+ P-ROIs (Fig. 5G, H). These pathways elicit immunosuppressive signals, suggesting that these tumors continue to progress despite high infiltration due to their suppression of cytotoxic T-cell activities.
Overall, our MS-based proteomic approach, integrated with imaging data, provides spatial snapshots of tumors in different tumor states. These snapshots enhance our understanding of cancer progression and interactions between cancer cells and the TME. Our data-rich resource highlights potential mechanisms involved in immune evasion at different tumor grades, shedding light on pathways that may contribute to tumor progression and reduced patient survival.
Discussion
Intratumor heterogeneity has been mainly studied at the genomic level. However, recent studies highlight the role of the cancer epigenome, transcriptome, and immune microenvironment in shaping functional ITH. Here, we consider the importance of both genetic and non-genetic ITH and their role in tumor evolution and present the rationale for a broad research focus beyond the cancer genome. Our results are in line with previous studies that address the difference between functional variation, conferring the Darwinian fitness effect that brings about an important change in the tumor phenotype, and ‘non- functional’ variation, which is often presented by the cancer genome34,35. While the genomic variability can initiate clonal divergence, the cancer phenotype is largely dictated by the TME through epigenetic and post-transcriptional regulation. Our study provides a comprehensive view of breast cancer’s spatial intratumor proteomic heterogeneity. It shows the intricate dynamics of the tumor ecosystem across different molecular subtypes and disease states. These findings contribute to the understanding of breast cancer phenotypes at the functional level, revealing an additional dimension of intratumor heterogeneity that may drive treatment resistance and relapse. By integrating an MS-based proteomic approach with imaging-based immune and stromal characterization, we illuminated the influence of the tumor microenvironment on cancer cell plasticity and, consequently, patient outcomes. Using multi-region proteomic analysis of 280 cancer regions from 33 patients, we examined the proteomic phenotypes in the complex tumor microenvironment rather than using simplified controlled systems of animal models or co-cultures that typically examine one TME component at a time. The proteomic data suggests how cancer cells, immune cells, cancer-associated fibroblasts, and blood vessels might affect each other to impact cancer outcomes. Furthermore, we show how the cancer-TME components might evolve with cancer progression from Grade 2 to Grade 3 tumors.
Despite the unique multi-region and multi-layered analyses of many breast cancer samples, our study suffers from several limitations. Primarily, the patient cohort is limited, which reduces the strength of inter-patient comparisons and the analysis of the clinical association of P-ITH. Moreover, the genomic analyses were limited to four HR+ tumors, and therefore, we cannot be confident about the broader relevance to TNBC as well. To address this, whenever available, we validated our findings in a large cohort of 385 tumors using publicly available imaging mass cytometry data21. Additionally, the use of consecutive slides for proteomics, imaging, and genomic analyses poses a limitation, as we do not take the exact same cells for analysis. However, given that the overall P-ROI encompasses several mm2, the impact of using different tissue sections (up to 50 µm apart) has a minor effect. Lastly, the use of TMT labeling and the use of two sample preparation methods induced some batch effect. To mitigate this, most analyses compared samples within a patient and a TMT set. In this way, we minimize the impact of these potential confounders.
In this study, we introduced a measure of tumor heterogeneity, P-ITH (proteomic intratumor heterogeneity), which provides an additional dimension to understanding tumor dynamics. Previous studies focusing on breast cancer genomics have shown that breast cancer genomic ITH increases with tumor progression36,37 and that heterogeneity is associated with lower immune activation19. However, these studies have been limited only to the genomic layer. Here, we provide views on ITH at the proteomic level. Our imaging data, along with independent validation by imaging mass cytometry data, showed that receptor expression heterogeneity is reduced from Grade 2 to Grade 3. At the same time, P-ITH is elevated with grade and higher cellularity, but it does not correlate with the genomic ITH. Heterogeneous receptor expression is known in routine clinical diagnosis and treatment. Tumor classification as ER-positive or HER2-positive is typically based on a small percentage of the tumor, leaving a substantial fraction of the cells uncharacterized. Receptor expression profiles have been demonstrated to vary within the same patients, particularly between primary tumors and their corresponding metastatic lesions38,39. However, very little is known about Receptor-ITH at the primary tumor level, which may contribute to treatment resistance and relapse when targeting the receptors directly (e.g., with tamoxifen). For instance, clinical studies show that often, tumors with favorable prognoses, namely Grade 2 luminal A tumors, can relapse with highly aggressive phenotypes. Our results suggest that receptor expression heterogeneity, already in primary tumors, might be the origin of these aggressive metastases.
The increase in P-ITH with grade might explain cancer phenotypic divergence and raises the need for more personalized tumor characterization. Proteomic characterization of the highly variable proteins suggested that these are primarily associated with interactions with the TME. Multiplexed imaging of T cells, macrophages, fibroblasts, and blood vessels sheds light on the TME contribution at different spatial levels (P-ROI, neighborhood, and whole tissue). According to the imaging data and the external validation on a large cohort, while T-cell and macrophage percentages increased, fibroblasts and BV percentages decreased in Grade 3. However, the proteomic data revealed immunosuppressive signals in these infiltrated tumors, including expression of checkpoint proteins and activation of the prostaglandin and kynurenine pathways. The kynurenine pathway plays a major role in various cancer types, including breast cancer, creating an immunologically vulnerable microenvironment that supports cancer cell growth and promotes T-reg differentiation and infiltration40,41. In agreement with our findings, Wagner et al. showed high PDL1+ tumor-associated macrophages (TAMs) in high-grade breast tumors42. With the integration of the MS-based proteomic data, we add a system-level view and provide this resource as the basis for identifying additional components of cancer-TME interactions and for developing improved therapies for these complex tumors.
Methods
This research complies with all relevant ethical regulations and approved by the Institutional Review Board of Sheba Medical Center, Tel Aviv University, Weizmann Institute of Science, and the Israel Ministry of Health.
Clinical sample collection and tumor mapping
Formalin-fixed paraffin-embedded (FFPE) tissue samples were retrospectively collected from 33 breast cancer patients with primary tumors, with no pretreatment and various pathological characteristics. All samples were collected from the Institute of Pathology at the Sheba Medical Center and the Israel National Biobank for Research (MIDGAM). The use of these samples for research was approved by the Institutional Review Board of Sheba Medical Center (approval SMC-7509-09, head of committee Prof. Dror Haratz), the Tel Aviv University and Weizmann Institute of Science ethics committees per the Declaration of Helsinki ethical guidelines. Tumor sections from five patients with hormone receptor-positive and triple-negative breast cancer diagnoses were obtained from the MIDGAM according to protocol no. 130-2013, following approval by the Ministry of Health. Clinical samples and clinical data were collected upon the patient’s informed consent to collect, store, and distribute samples. As the extent of proteomic heterogeneity was unknown a priori, a sample size calculation was not performed to determine the number of patients included.
To map the histopathological characteristics of each tumor region, stained slides were assessed by trained breast pathologists from the Sheba Medical Center, and receptor staining status was determined based on the Allred scoring system. H&E-stained slides were analyzed using Elston and Ellis semi-quantitative method to assess histological grade. Combining these scores, we created histopathological maps and defined distinct tumor regions according to the following criteria: receptor expression (per region) and tumor grade (per region). These visually represent intratumoral heterogeneity for downstream processing by laser capture microdissection (LCM). Tumor regions defined based on receptor expression included triple-negative (TN) (n = 68), hormone receptor-positive (HR+) (n = 172), and HER2+ regions (n = 18). We also dissected normal-adjacent tissue samples (n = 19) and regions lacking defined receptor status, labeled as “NA” (n = 3). All tumor-containing paraffin blocks were collected from every patient (4-5 blocks per patient). The criteria for selecting regions for further processing was defined as 70% enrichment of cancer cells (within a region) with a minimal size requirement of the target regions to provide sufficient material for proteomics. Detailed clinical information of each patient is included in Supplementary Data 1. Disease-free survival was calculated from the date of primary treatment to the date of relapse or until the patient’s death. For each tissue block, multiple slides were prepared to facilitate various analyses. The total number of slides obtained from each tissue block was as follows: i) Hematoxylin and Eosin (H&E) and Immunohistochemistry (IHC): five slides, each 3.5 μm thick; ii) Proteomics: one slide, 8 μm thick; iii) Immunofluorescence imaging: two slides, each 3.5 μm thick; iv) Exome Sequencing (four patients): five slides, each 8 μm thick. Given this arrangement, the distance between the proteomic and the immunofluorescence sections did not exceed 40–50 μm.
Immunohistochemistry (IHC) staining
Five consecutive 3.5 μm tissue sections were sliced from all cancer-containing paraffin blocks from each patient (average 4–5 blocks per patient), mounted on positively charged glass slides, and dried overnight at 37°C. Slides were mapped by staining with H&E, anti-ER (DAKO), anti-PR (DAKO), anti-HER2 (DAKO), and anti-cytokeratin (Cell Marque). Staining was performed on the BOND-RX automated staining platform using a Bond Polymer Refine Detection automated kit (Leica Biosystems). H&E staining was performed manually using a standard protocol.
IHC Image analysis
DAB-stained slides were scanned using the Leica Aperio VERSA Digital Pathology Scanner (Aperio Technologies Inc.). Staining quantification of all selected regions was performed using the eSlide Manager software via the Aperio image analysis algorithms: Nuclear (ER, PR), Membrane (HER2), and Cytoplasmic (CK). Regions were defined by a trained pathologist and manually annotated within the software. Regions were considered positive for the antibody staining if they achieved a staining cutoff threshold of 10% and were defined as high-intensity staining (+3) based on the machine-learning algorithmic scoring.
Laser capture microdissection (LCM)
PEN-membrane glass slides allowed combined UV and IR laser microdissection. Tissue sections (8 μm thick) were mounted on the slides, dried, and deparaffinized with xylene, followed by a series of graded ethanol washes. Staining was performed with the Paradise Plus system (Thermo Fisher Scientific). Stained tumor samples were microdissected using the ArcturusXT laser capture microdissection system (Thermo Fisher Scientific). This LCM system allowed us to harvest relatively pure target populations of cancer cells using precise ultraviolet (UV) and infrared (IR) lasers under direct microscopic visualization.
Proteomics sample preparation
Microdissected tumor sections were collected into Eppendorf LoBind microcentrifuge tubes, lysed, and digested using two protocols: trifluoroethanol (TFE) or SDS in two sample batches. Protocol I included lysis with 50% 2,2,2- TFE in 50 mM ammonium bicarbonate buffer; the samples were boiled at 95 °C for 1 h, sonicated for 10 cycles in a Bioruptor sonicator (Diagenode), and then centrifuged at 17,000 g for 20 minutes to pellet tissue debris. Lysates were transferred to fresh LoBind tubes, reduced with 5 mM dithiothreitol (DTT), and alkylated with 15 mM iodoacetamide (IAA) for 30 minutes, followed by overnight “in-solution” digestion using LysC-trypsin mix as described43. Protocol II included lysis in 4% Sodium dodecyl sulfate (SDS) in 25 mM HEPES (pH = 8) lysis buffer, incubation for 1.5 h at 95 °C, reduction, alkylation, and digestion following the Single Pot Solid Phase Sample Preparation (SP3) protocol44,45. SeraMag Hydrophilic and Hydrophobic beads (GE Healthcare) mix was used at a concentration of 100 µg/µl for overnight “on-bead” digestion using LysC-trypsin mix (Promega). Tryptic peptides from 280 regions were labeled using 10plex TMT reagent and assembled into 38 TMT-labeled sample sets. Each set was composed of nine channels for the different regions and one channel for an internal standard. The internal standard was always labeled with the 131-C reagent. Assembled TMT-labeled peptides were then fractionated offline into eight fractions using high pH reverse-phase chromatography (PierceTM High pH Reversed-Phase Peptide Fractionation kit), following the manufacturer’s instructions. Fractionation was followed by LC-MS analysis.
LC-MS-based proteomics
LC-MS/MS analysis was performed using high-performance liquid chromatography (Easy nLC 1000 HPLC system) coupled online to a Q-Exactive HF mass spectrometer (Thermo Fisher Scientific) through the EASY-Spray ionization source. Peptides were separated on 75 μm × 50 cm long EASY-spray PepMap columns (Thermo Fisher Scientific) and loaded with Buffer A (0.1% formic acid). Peptides were eluted with a 140-minute gradient of water-acetonitrile, and each batch of samples, composed of eight fractions, was analyzed for a total of 20 hours. We slightly modified the gradients for each fraction as follows: for fractions 1 and 2, 5-30% buffer B (80% acetonitrile/0.1% formic acid); for fraction 3, 5–31% buffer B; for fraction 4, 5-32% buffer B; for fraction 5, 5–34% buffer B; for fraction 6, 5-36% Buffer B; for fraction 7-8, 5-38% buffer B. All fractions were eluted at a 200 ml/min flow rate at 40 °C. MS data were acquired in a data-dependent mode, and the acquisition method included a full scan event at a resolution of 120,000, followed by top 10 MS/MS scans at a resolution of 60,000. MS acquisition was performed in a blinded manner, namely, patient clinical characteristics were blinded.
Data processing: proteomics raw MS data processing
Raw MS files were analyzed using MaxQuant software (v2.0.3.0) with the Andromeda search engine46. The Uniprot database (human version from 2021 with 95,057 entries) was used for searching MS/MS spectra. Modifications for 10plex TMT labels were defined in MaxQuant and included the label impurities as indicated for each label batch. A decoy database was used to determine a 1% false discovery rate for protein identification and peptide spectrum matches. We included carbamidomethyl cysteine as a fixed modification and N-terminal acetylation and methionine oxidation as variable modifications with a maximum of five modifications per peptide. A maximum of two missed cleavages was allowed. The weighted median normalization method for TMT datasets, using the reference channel, was used as described47. The complete protein table is provided in Supplementary Data 9.
Statistics and reproducibility
All statistical tests were performed using Perseus (v1.6.15.0), Python (v3.12.1), and R (v2024.04.2). We used the proteinGroups MaxQuant output table for the analysis. The initial data processing phase involved the removal of low-quality samples, followed by filtration of proteins originating from the decoy database (reverse proteins), those solely identified by modification sites, and common contaminants. Protein expression values underwent a log2-transformation. The dataset was further refined to retain proteins with a minimum of 50% valid values, resulting in 5475 protein groups. Subsequently, extracellular matrix proteins were removed to prevent the distortion of overall ratio distribution caused by the high and variable expression of extracellular proteins, resulting in 5245 protein groups (Supplementary Data 9).
Additionally, we normalized the expression of protein groups in each region using a two-component Gaussian mixture model2 implemented with the”DMwR” (v0.4.1),”grid” (v4.4.2), and “lattice” (v0.22.6) R packages. Finally, missing values were imputed through k-nearest neighbor (KNN) imputation, employing a k = 17 (the square root of 280 regions). This procedure resulted in a complete dataset, which served as the basis for subsequent downstream analyses.
No statistical method was used to predetermine sample size. The original clinical cohort comprised 363 P-ROIs from 37 patients. Due to technical issues during sample processing, 280 samples passed quality control and were included in the analysis. No data was excluded from the analysis beyond this quality control step, unless otherwise specified for a particular comparison. The experiments were not randomized. During MS acquisition, investigators were blinded to the clinical parameters, whereas all subsequent analyses were not blinded.
Statistical analysis
For all statistical analyses, unless stated otherwise, p-values were corrected for multiple testing using the false discovery rate (FDR) method, with a threshold of 0.05 and S0 set to 0.1. Where applicable, two-sided Student’s T-tests or Welch’s T-tests were applied as appropriate for equal or unequal variances, respectively. Effect sizes were computed using the “effsize” (v0.8.0) R package. Principal component analysis (PCA) was performed on log2-transformed, scaled, and centered intensity values after KNN imputation (as described above).
Unsupervised clustering
Unsupervised clustering was conducted using the modified minimum variance method (Ward.D2). The imputed matrix was utilized, and Euclidean distances between regions were computed. Subsequently, the modified Ward’s method was applied to generate groups to minimize cluster variance. To visually represent our cohort of 280 regions, we employed a “Tree-and-Leaf” dendrogram wherein each node represents a tumor region. The tree branches were constructed based on Euclidean distances and cluster separation, depicting the spatial localization of the branches. For visualization purposes, we utilized the “TreeAndLeaf” (v1.6.1), “igraph” (v1.4.2), and “RedeR” (v2.1.4) R packages.
Supervised analysis
Differences between regions defined as distinct receptor subtypes and between grades were assessed using a Linear Mixed-Effects Model (LMM) implemented with the “lme4” (v1.1.37) and “lmerTest” (v3.1.3) R packages. Standardized effect sizes (standardized β coefficients) were computed using the “parameters” (v0.27.0) package. Model fit and assumption checks were performed using the “performance” (v0.15.0) R package. By including patient as a random effect, the model accounted for the non-independence of regions originating from the same patient. Statistical significance in volcano plots was determined using permutation-based FDR with a q-value < 0.05. One-dimensional annotation enrichment analysis was conducted on fold-change values, applying a Benjamini–Hochberg FDR threshold of 0.02, using Perseus software. Visualizations were generated using the “ggplot2” (v3.5.2) R package.
Constant and variable proteins
We used the median absolute deviation (MAD) score per protein to evaluate which proteins contribute to intratumor heterogeneity. We ranked the MAD scores of all proteins in each tumor. Using the sum of ranked proteins across all patients, we selected the lowest and highest 10% as constant and variable protein groups, respectively. Functional enrichment analysis of these groups was performed using g:Profiler to identify overrepresented biological pathways or GO/KEGG terms relative to all detected proteins in our dataset (one-sided Fisher’s exact test, FDR q < 0.05). The results were visualized in Cytoscape (v3.10.0) using the EnrichmentMap plugin, with a node FDR cutoff of 0.05 and an edge cutoff (Jaccard similarity index) of 0.25. Redundant processes were removed from the resulting networks.
ITH Scores
We derived three intra-tumor heterogeneity (ITH) scores to quantify intratumor heterogeneity at different levels. For breast cancer receptor subtypes (Receptor-ITH), we computed the number of co-existing subtypes per tumor. Subsequently, we obtained the area measure of each subtype using the eSlide Manager software. The score was calculated as follows:
Where F corresponds to the largest fraction of subtypes from the total area of the tumor, and n is the number of subtypes co-existing in the same tumor. The obtained scores were calculated per tumor.
Based on imaging mass cytometry dataset, we employed Shannon’s Diversity Index to evaluate hormone receptor intratumor heterogeneity (ITH). We filtered single cells into two categories, HR+ and HR-, based on their descriptors. Subsequently, we utilized these two classes to compute the Shannon Diversity Index for each patient using the “vegan” (v2.7.1) R package.
Lastly, for the proteomic ITH score (P-ITH) calculation, we computed the correlations between all pairs of tumor regions using the log2-transformed, normalized, and imputed dataset comprising 5475 protein groups. Subsequently, we averaged data from two non-overlapping pairs with the lowest correlation per tumor to alleviate any subjective effect of multi-sampling. The score was calculated per tumor and per TMT set. The score was calculated as follows:
Where mean(LowCor) corresponds to the average of two lowest pairwise proteome correlation within a tumor.
To account for tumor purity in P-ITH calculations, data correction was performed using tumor purity scores derived from the ESTIMATE algorithm. The Pseudo-Replicate Pseudo-Sample (PRPS) method48 was then applied to estimate and remove unwanted variation from the data. Negative control genes, genes that strongly correlate with tumor purity, were defined based on the absolute Pearson’s correlation coefficient greater than 0.6. Two pseudo-samples (average high and average low tumor purity) were created, forming a pseudo-replicate set. P-ITH values were recalculated using the corrected dataset.
Genomic sample preparation and bioinformatic analysis
We selected samples from four ER+ patients that sum up to 53 regions. Three regions of patient BC5 were pathologically defined as ductal carcinoma in situ (DCIS) and thus were not included in downstream analyses. FFPE samples from the four patients were mounted onto PEN-membrane glass slides for downstream LCM using UV and IR lasers under constant microscopic evaluation. Due to low amounts of intact DNA per slide, LCM was performed on five 8 μm-thick consecutive slides from each sample. Samples were deparaffinized using xylene followed by a series of descending ethanol concentrations, and subsequently stained with the Paradise Plus system (Thermo Fisher Scientific) prior to LCM. Microdissected samples were collected and stored at −80 °C.
Focused exome sequencing
DNA was purified from FFPE samples using the Maxwell® 16 FFPE Plus LEV DNA Purification Kit (Promega# AS1135) on a Maxwell MDx robot (Promega). Fifty ng of tumor gDNA was sheared by Covaris S220 system. The SureSelect XT HS Target Enrichment system was used for DNA library preparation and target-enriched library. Next-generation sequencing (NGS) was carried out on the Illumina NextSeq 550 platform using 75 bp pair-end reads following the manufacturer’s instructions.
DNA purification from lymphocyte samples for germline control was performed using the QIAprep spin miniprep kit (Qiagen). One hundred ng gDNA was sheared by Covaris (S220). The SureSelect XT HS Target Enrichment system was used for DNA library preparation and target-enriched library. NGS was carried out using the Illumina Nextseq 550 platform with paired-end reads of 75 bp according to the manufacturer’s instructions. Library preparation and sequencing were performed at the Genomics Technology Center of the Medicine Faculty in Bar-Ilan University, Zefat, Israel.
Variant calling
Reads were trimmed using cutadapt and aligned to the H. sapiens genome (b37, downloaded from the GATK bundle v2.8) using bwa mem (v0.7.15, with -a and -M tags). The alignment files were sorted, and duplicates were marked using Sambamba (v0.6.0). In order to reduce the contamination from mouse DNA, reads with soft clips and NM > 6 were discarded from the “INV” samples. Variants were called using lancet (v1.0.7, --primary-alignment-only, --XA-tag-filter --min-alt-count-tumor=6, --max-alt-count-normal=1, --min-vaf-tumor=0.05, --max_vaf_normal = 0.01, --min_coverage_tumor=10, --min-coverage-normal=10, --max-coverage-tumor=2000, --max_coverage_normal=2000, --min_phred_fisher=5). Variants were filtered further based on frequency greater than 0.135 for I1044 and 0.075 for I1218. Variants were annotated using Ensembl’s VEP release 93. All variants with variant allele frequency (VAF) lower than 0.2 were filtered out for downstream analyses. Variant calling was performed by the Mantoux Bioinformatics Institute of the Nancy and Stephen Grand Israel National Center for Personalized Medicine, Weizmann Institute of Science.
Multiplexed immunofluorescence staining and imaging
Tissue sections were deparaffinized in xylene and rehydrated through decreasing ethanol concentrations. Endogenous peroxide blocking was performed with 3% H2O2 and 1% HCL in methanol, followed by heat-induced antigen retrieval in 10 mM citric acid (pH = 6). Blocking was performed with 20% normal horse serum (NHS) and 0.1% Triton X-100, with an additional biotin-blocking step for secondary biotinylated antibodies. Primary antibodies were diluted in 2% NHS and 0.1% Triton X-100 and incubated overnight.
CD15 (Invitrogen, MA5-11789) was used at a 1:50 dilution, followed by HRP-conjugated donkey anti-mouse secondary antibody (Jackson ImmunoResearch, 115-035-146) and OPAL 650 fluorophore (Akoya, FP1496001KT). αSMA expression was detected with αSMA antibody (Sigma-Aldrich, A2547) at a 1:100 dilution, using donkey anti-mouse secondary antibody (Jackson ImmunoResearch, 115-035-146) and OPAL 570 (Akoya, FP1488001KT). CD31 (Cell Signaling Technologies, 3528) was used at 1:1000 dilution, followed by biotinylated donkey anti-mouse (Jackson ImmunoResearch, 715-065-151) and streptavidin-conjugated cy2 (Jackson ImmunoResearch, 016-220-084). CD3 (Atlas Antibodies, HPA043955) was used at a 1:500 dilution, CD68 (Cell Signaling Technologies, 76437) at 1:200 dilution, each followed by HRP-conjugated donkey anti-rabbit secondary (Jackson ImmunoResearch, 711-035-152) and OPAL 650 (Akoya, FP1496001KT) or OPAL 570 (Akoya, FP1488001KT), respectively. Pan-cytokeratin (Cell Marque, CMC3132050) was used at 1:100 dilution, followed by biotinylated donkey anti-mouse secondary (Jackson ImmunoResearch, 715-065-151) and streptavidin-conjugated cy2 (Jackson ImmunoResearch, 016-220-084). Antibody removal was performed with 10 minutes of microwave treatment with citric acid (pH=6), allowing repetition of the staining protocol from the blocking step. Hoechst-stained nuclei and imaging using the Panoramic Scan II (3DHistech).
Fluorescence image analysis and quantification
H-DAB slides (ER, PR, HER2, or panCK) were scanned on a Leica Aperio VERSA Digital Pathology Scanner with a 20× objective (0.2715 µm pixel size, 24-bit color) and loaded onto QuPath (v0.3.2)49. Aperio ROIs were imported from an XML file. Fluorescently stained slides were imaged on a Pannoramic SCAN 150 scanner (3DHistech) with a Plan-Apochromat 20× objective (1× camera adapter) and a pco.edge 4.2 4MP camera at 0.326 µm pixel size, with seven focal planes (5 µm steps), flat-field correction, and intensity compensation. Filter settings were: Set 1 — Dapi-Q (20 ms, 9%), Cy5-Q (85 ms, 100%), FITC (20 ms, 10%), TRITC (20 ms, 45%); Set 2 — Dapi-Q (20 ms, 9%), Cy5-Q (20 ms, 68%), FITC (25 ms, 55%), TRITC (20 ms, 10%). All channels were acquired with 8-bit JPEG (90% quality). Nuclear segmentation on the fluorescent slides was performed using StarDist (v0.8.0)50,51, followed by expansion implemented in the QuPath StarDist extension. The model used ZeroCostDL4Mic52 StarDist notebook. The machine learning-based pixel classifier was trained on CD31 and αSMA-positive blood vessels on a training panel of stitched 500*500μm ROIs. The pixel-classified image areas were converted to objects (minimum object size of 100 µm2, minimum hole size of 20 µm2. The machine learning-based object classifier was trained in QuPath (v0.3.2) on a training panel of stitched 500*500μm ROIs. The stromal panel included classes for αSMA, RBCs, and “others.” The immune panel included classes for panCK (tumor), CD3, CD68, “others”, RBCs, and “ignore”; CD15+ cells were excluded due to rarity. The whole slide image registration to either set of fluorescent panels was performed using Warpy - Whole Slide Image (WSI) Registration With ImageJ/Fiji53, Elastix54BigDataViewer55and QuPath. Transferred ROIs were then expanded by 500 μm². The total number of classified cells and the area covered by blood vessel objects were quantified. For whole-slide marker correlations, the data was averaged per patient, with approximately four FFPE blocks imaged per patient (SD = \(\pm\)1.5). All raw/processed images, ROI metadata, segmentation masks, and full acquisition/analysis details are available in the BioImage Archive (S-BIAD1176) and via the linked GitHub repository.
Proteomic region classifications
Immune phenotype
Using spatial multiplexed imaging, we analyzed the distribution of CD3+ and CD68+ cells in the TME across 121 regions. We identified distinct immune phenotypes: Infiltrated (n = 38), Cold (n = 22), CD3COLDCD68INF (n = 47), CD3COMPCD68INF (n = 11), and CD3INFCD68COLD (n = 3). These phenotypes were defined based on the density of CD3 and CD68 cells per area, with a CD3 cutoff of 600 cells/mm2 and a CD68 cutoff of 250 cells/mm2. Regions with cell numbers below the cutoff were classified as ‘Cold,’ while those above the cutoff were classified as ‘Infiltrated’. The phenotype ‘CD3COLDCD68INF’ is based on the mentioned cutoff having a low number of T cells and high levels of macrophages. Based on the cutoffs, the CD3COLDCD68INF phenotype exhibited low T-cell numbers and high macrophage levels. Due to the limited number of regions (3) characterized as ‘CD3INFCD68COLD’ and the associated statistical power concerns, we excluded these regions from further analysis, resulting in 118 regions. To define the CD3COMPCD68INF phenotype, we introduced an additional criterion: having a ring-to-P-ROI CD3+ ratio greater than 1.2. Thus, ‘CD3 compartmentalized’ regions were defined by low T-cell abundance in the P-ROIs, had a ring-to-P-ROI ratio ≥ 1.2, and elevated macrophage levels. One-way ANOVA, using Immune Phenotype as a factor, was performed to identify proteins significantly associated with immune infiltration in the P-ROIs. To isolate the effects of Immune Phenotype while accounting for confounding factors such as Grade and molecular subtype, a three-way ANOVA was conducted. Proteins were filtered based on significant p-values (p < 0.05) for the Immune Phenotype factor and non-significant p-values (p > 0.05) for the grade and subtype factors. Hierarchical clustering was performed following k-means preprocessing (k = 200), using Euclidean distances and average linkage for both rows and columns.
Stroma phenotype
For BV regional classification, we calculated the BV% within the P-ROI and the adjacent 500 μm expanded ring. Samples with values above the median were classified as “BV-high,” while those below the median were classified as “BV-low”. Similarly, fibrotic classification as “Fibro-high” and “Fibro-low” was based on the \({{{\rm{\alpha }}}}\)SMA% in the P-ROIs, using the same median-based methodology. In the BV classification, the 500 μm ring was included to better capture local vascularization surrounding the P-ROI. Since blood vessels are easily detected in H&E, we avoided cutting through them for proteomics, making the ring essential for a representative BV% measurement.
Immune cell proteomic contributions
To estimate the proteomic contributions of immune cells within the P-ROIs, the ConsensusTME algorithm was applied for tumor microenvironment cell deconvolution56. Prior to applying the algorithm, the proteomic data, normalized to the TMT reference channel, was adjusted by multiplying each expression value by the mean intensity of the reference channel, followed by log2 transformation. The resulting matrix included 8014 proteins. For correlating the ConsensusTME results with CD3% “ground truth” measurements from the imaging cohort, the T-cell normalized enrichment score was calculated as the sum of CD4, CD8, gamma-delta, and regulatory T cells scores. A Random Sample Consensus (RANSAC) regression approach was then used to predict the CD3% in P-ROIs not included in the imaging dataset. The RANSACRegressor function from Python’s “linear_model” module in scikit-learn (v1.3.2) was configured to use a minimum of 50% of the data for model fitting, a residual threshold of 10, and a maximum of 100 iterations. For macrophage enrichment score correlation with CD68%, the score was based on the summed values of M1, M2, monocytes, and macrophages for each region, and the T-cell signature included CD4, CD8, and broad T-cell lineage proteins. Macrophage and T-cell signatures were defined using the Human Protein Atlas database (https://www.proteinatlas.org/), comprising 926 and 408 proteins, respectively (Supplementary Data 8).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All mass spectrometry raw data generated in this study have been deposited in the ProteomeXchange repository under accession code PXD024190 https://www.ebi.ac.uk/pride/archive/projects/PXD024190. Supplementary Data files include full protein lists for each analysis and variant information from the focused exome analysis. Raw genomics data could not be made publicly available due to ethical constraints of the funding agency. Imaging data generated in this study have been deposited in the bioimage-archive repository under accession code S-BIAD1176 https://doi.org/10.6019/S-BIAD1176. Source data are provided with this paper.
Code availability
Image processing code is available via the following Github link: https://github.com/WIS-MICC-CellObservatory/BreastCancerHeterogeneity?tab=readme-ov-file.
References
Yanovich, G. et al. Clinical proteomics of breast cancer reveals a novel layer of breast cancer classification. Cancer Res. 78, 6001–6010 (2018).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Krug, K. et al. Proteogenomic landscape of breast cancer tumorigenesis and targeted therapy. Cell 183, 1436–1456 e1431 (2020).
TCGA Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Curtis, C. et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Prat, A. & Perou, C. M. Deconstructing the molecular portraits of breast cancer. Mol. Oncol. 5, 5–23 (2011).
Arad, G. & Geiger, T. Functional impact of Protein-RNA variation in clinical cancer analyses. Mol. Cell Proteom. 22, 100587 (2023).
Wang, J. et al. Proteome Profiling Outperforms Transcriptome Profiling for Coexpression Based Gene Function Prediction. Mol. Cell Proteom. 16, 121–134 (2017).
Tyanova, S. et al. Proteomic maps of breast cancer subtypes. Nat. Commun. 7, 10259 (2016).
Johansson, H. J. et al. Breast cancer quantitative proteome and proteogenomic landscape. Nat. Commun. 10, 1600 (2019).
Rivenbark, A. G., O’Connor, S. M. & Coleman, W. B. Molecular and cellular heterogeneity in breast cancer: challenges for personalized medicine. Am. J. Pathol. 183, 1113–1124 (2013).
Lopez-Tarruella, S. et al. Correlation between breast cancer subtypes determined by immunohistochemistry and n-COUNTER PAM50 assay: a real-world study. Breast Cancer Res. Treat. 203, 163–172 (2024).
Schrijver, W. A. et al. Receptor conversion in distant breast cancer metastases: a systematic review and meta-analysis. JNCI: J. Natl. Cancer Inst. 110, 568–580 (2018).
Cejalvo, J. M. et al. Intrinsic subtypes and gene expression profiles in primary and metastatic breast cancer. Cancer Res. 77, 2213–2221 (2017).
Pasha, N. & Turner, N. C. Understanding and overcoming tumor heterogeneity in metastatic breast cancer treatment. Nat. Cancer 2, 680–692 (2021).
Swanton, C. Intratumor heterogeneity: evolution through space and time. Cancer Res. 72, 4875–4882 (2012).
Wolf, Y. et al. UVB-induced tumor heterogeneity diminishes immune response in melanoma. Cell 179, 219–235 e221 (2019).
McDonald, K. A. et al. Tumor heterogeneity correlates with less immune response and worse survival in breast cancer patients. Ann. Surg. Oncol. 26, 2191–2199 (2019).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Ali, H. R. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat. Cancer 1, 163–175 (2020).
Karthik, G.-M. et al. Intra-tumor heterogeneity in breast cancer has limited impact on transcriptomic-based molecular profiling. BMC cancer 17, 1–11 (2017).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Tate, J. G. et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47, D941–D947 (2018).
Sewell-Loftin, M. K. et al. Cancer-associated fibroblasts support vascular growth through mechanical force. Sci. Rep. 7, 12574 (2017).
Jiang, H., Hegde, S. & DeNardo, D. G. Tumor-associated fibrosis as a regulator of tumor immunity and response to immunotherapy. Cancer Immunol. Immunother. 66, 1037–1048 (2017).
Mishra R., Patel H., Alanazi S., Kilroy M. K., Garrett J. T. PI3K Inhibitors in cancer: clinical implications and adverse effects. Int. J. Mol. Sci. 22, (2021).
Bagheri-Yarmand, R., Mazumdar, A., Sahin, A. A. & Kumar, R. LIM kinase 1 increases tumor metastasis of human breast cancer cells via regulation of the urokinase-type plasminogen activator system. Int. J. Cancer 118, 2703–2710 (2006).
Zhang, J. et al. SOX4 induces epithelial–mesenchymal transition and contributes to breast cancer progression. Cancer Res. 72, 4597–4608 (2012).
Zhang, J. et al. SOX4 promotes the growth and metastasis of breast cancer. Cancer Cell Int. 20, 1–11 (2020).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387. e1319 (2018).
Li, X. et al. Infiltration of CD8(+) T cells into tumor cell clusters in triple-negative breast cancer. Proc. Natl. Acad. Sci. USA 116, 3678–3687 (2019).
Peranzoni, E. et al. Macrophages impede CD8 T cells from reaching tumor cells and limit the efficacy of anti-PD-1 treatment. Proc. Natl. Acad. Sci. USA 115, E4041–E4050 (2018).
Black, J. R. M. & McGranahan, N. Genetic and non-genetic clonal diversity in cancer evolution. Nat. Rev. Cancer 21, 379–392 (2021).
Karras, P., Black, J. R. M., McGranahan, N. & Marine, J. C. Decoding the interplay between genetic and non-genetic drivers of metastasis. Nature 629, 543–554 (2024).
Dagogo-Jack, I. & Shaw, A. T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15, 81–94 (2018).
Lüönd, F., Tiede, S. & Christofori, G. Breast cancer as an example of tumour heterogeneity and tumour cell plasticity during malignant progression. Br. J. Cancer 125, 164–175 (2021).
Zardavas, D., Irrthum, A., Swanton, C. & Piccart, M. Clinical management of breast cancer heterogeneity. Nat. Rev. Clin. Oncol. 12, 381–394 (2015).
Aurilio, G. et al. A meta-analysis of oestrogen receptor, progesterone receptor and human epidermal growth factor receptor 2 discordance between primary breast cancer and metastases. Eur. J. Cancer 50, 277–289 (2014).
Ala, M. The footprint of kynurenine pathway in every cancer: A new target for chemotherapy. Eur. J. Pharmacol. 896, 173921 (2021).
Girithar H.-N. et al. Involvement of the kynurenine pathway in breast cancer: updates on clinical research and trials. Br. J. Cancer, 1–19 (2023).
Wagner, J. et al. A single-cell atlas of the tumor and immune ecosystem of human breast cancer. Cell 177, 1330–1345. e1318 (2019).
Shenoy, A. et al. Proteomic patterns associated with response to breast cancer neoadjuvant treatment. Mol. Syst. Biol. 16, e9443 (2020).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Jiang, L. et al. A quantitative proteome map of the human body. Cell 183, 269–283.e219 (2020).
Molania, R. et al. Removing unwanted variation from large-scale RNA sequencing data with PRPS. Nat. Biotechnol. 41, 82–95 (2023).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 1–7 (2017).
Schmidt U., Weigert M., Broaddus C., Myers G. Cell detection with star-convex polygons. In: Medical Image Computing and Computer Assisted Intervention–MICCAI 2018: 21st International Conference, Granada, Spain, September 16-20, 2018, Proceedings, Part II 11). Springer (2018).
Weigert M., Schmidt U. Nuclei instance segmentation and classification in histopathology images with Stardist. In: 2022 IEEE International Symposium on Biomedical Imaging Challenges (ISBIC)). IEEE (2022).
von Chamier, L. et al. Democratising deep learning for microscopy with ZeroCostDL4Mic. Nat. Commun. 12, 2276 (2021).
Chiaruttini, N. et al. An open-source whole slide image registration workflow at cellular precision using Fiji, QuPath and Elastix. Front. Comput Sci. 3, 780026 (2022).
Klein, S., Staring, M., Murphy, K., Viergever, M. A. & Pluim, J. P. Elastix: a toolbox for intensity-based medical image registration. IEEE Trans. Med. Imaging 29, 196–205 (2009).
Pietzsch, T., Saalfeld, S., Preibisch, S. & Tomancak, P. BigDataViewer: visualization and processing for large image data sets. Nat. methods 12, 481–483 (2015).
Jimenez-Sanchez, A., Cast, O. & Miller, M. L. Comprehensive benchmarking and integration of tumor microenvironment cell estimation methods. Cancer Res. 79, 6238–6246 (2019).
Acknowledgements
We thank the members of the Geiger lab for fruitful discussions and technical assistance. We thank Dr. Gilgi Friedlander from the Montoux Bioinformatics team at the Nancy and Stephen Grand Israel National Center for Personalized Medicine for genomic data analyses. We thank Dr. Adi Zundelevich for assistance with patient cohort assembly, members of the Pathology Department at the Sheba Medical Center for assistance in tissue selection and processing, and the Israel National Biobank for Research (MIDGAM) for assistance in tissue selection. We thank the Satchi-Fainaro lab at Tel Aviv University for assistance in tissue staining. Figure 1 was created with BioRender.com. We thank the Constantiner Institute for partial support. This work was funded by European Council ERC-starting grant # 639534 and the European Council ERC-consolidator grant # 101044574.
Author information
Authors and Affiliations
Contributions
M.M. performed the experiments and analyzed the proteomics data; S.K. performed the data analysis and wrote the paper; V.M. performed the immunofluorescence staining and image analyses; M.M., S.K., V.M., T.G. conceptualized the research and interpreted the results; G.A. performed the genomic analyses; D.N., K.K., I.M., B.S., I.B. performed the pathological analyses; A.S., D.P., N.I., R.S. performed computational analyses; O.G. developed image analysis pipelines; L.F.A. assisted in multiplexed imaging; YA assisted in image processing; M.M., E.G.Y., T.G. designed the patient cohort; T.G. supervised the research, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks anonymous reviewer(s) for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mardamshina, M., Karagach, S., Mohan, V. et al. Integrated spatial proteomic analysis of breast cancer heterogeneity unravels cancer cell phenotypic plasticity. Nat Commun 16, 10482 (2025). https://doi.org/10.1038/s41467-025-65477-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65477-6