Abstract
YTHDC2, a unique YTH-domain-containing protein that recognizes N6-methyladenosine (m6A) on RNA, plays critical roles in diverse pathological processes and represents a promising therapeutic target. Despite its potential, no potent small-molecule inhibitors have been reported to date. To bridge this gap, we develop EPMolGen, a deep learning-based molecular generative model that explicitly incorporates the electrostatic features of receptor proteins. The model achieves state-of-the-art performance in dry-lab validations. Using EPMolGen, we identify H3, a YTHDC2 inhibitor with an IC50 of 16.84 μM. Subsequent structural optimization of H3 yields DC2-C1, a highly potent compound with an IC50 of 0.168 μM against YTHDC2 and selectivity over other YTH-domain proteins. In cellular assays, DC2-C1 effectively targets YTHDC2. Notably, DC2-C1 treatment substantially reduces the expression levels of multiple target mRNAs of YTHDC2, leading to phenotypic suppression of related cells. Overall, this study highlights the great potential of deep learning in drug discovery and provides a promising lead compound for drug development targeting YTHDC2.
Similar content being viewed by others
Introduction
N6-methyladenosine (m6A) is a common RNA modification in eukaryotic organisms. It dynamically regulates the RNA fate and functions through the coordinated efforts of methyltransferases (writers), demethylases (erasers), and m6A-recognition proteins (readers)1,2. This modulation influences a wide array of biological processes, including cellular differentiation, development, and disease progression3,4. Among these components, m6A readers function as effector proteins and are involved in numerous physiological and pathological processes5,6,7. Currently, proteins containing the YT521-B homology (YTH) domain (YTHDC1-2 and YTHDF1-3) have been identified as the primary readers of m6A-modified mRNAs8,9,10,11.
YTHDC2 is a unique YTH-domain containing m6A reader, which is located in both the nucleus and cytoplasm12,13. It contains multiple functional domains, with its YTH domain being the most notable for its ability to recognize and bind to m6A-modified RNA molecules14. YTHDC2 has been demonstrated to play a crucial role in various biological processes, including RNA metabolism, cellular function and development, and has also been linked to the pathology of numerous diseases15. For instance, multiple studies have highlighted a close association between YTHDC2 and the onset and progression of rheumatoid arthritis (RA)16,17,18. In RA, RA fibroblast-like synoviocytes (RA-FLS) serve as key constituents of the hyperplastic and invasive synovial tissue. The abnormal proliferation and aggressive phenotypes of RA-FLS exacerbate joint destruction. Notably, impairing the m6A binding function of YTHDC2 in RA-FLS reduces the stability of AMIGO2 mRNA, a target of YTHDC2, leading to decreased AMIGO2 protein expression. This downregulation of AMIGO2 inhibits the proliferation, migration, and invasion of RA-FLS, thereby alleviating RA symptoms18. Furthermore, YTHDC2 exhibits a complex role in cancer, acting as either a promoter or suppressor of tumor growth through various mechanisms. For example, numerous studies19,20,21,22,23,24,25,26 have shown that YTHDC2 serves as a tumor suppressor in hepatocellular carcinoma, and papillary thyroid carcinoma, while an oncogene role in gastric cancer, pancreatic ductal adenocarcinoma (PDAC), prostate cancer, nasopharyngeal carcinoma, and cutaneous squamous cell carcinoma. In addition, YTHDC2 has also been implicated in metabolic regulation, such as hepatic lipogenesis27, therefore, contributing to the pathogenesis of some metabolic disorders.
However, current experimental approaches to studying the biological functions of YTHDC2 predominantly rely on the knockdown, knockout, or overexpression of the entire YTHDC2 gene. These methods do not allow for the determination of whether blocking YTHDC2-m6A recognition is responsible for the observed biological effects. Small molecule inhibitors of the YTH domain could serve as ideal tools to address this limitation. To our knowledge, only one pan-YTH protein inhibitor has been reported thus far, which showed weak activity against YTHDC2 with an IC50 of 30 μM28. In this investigation, we aim to discover potent inhibitors specifically targeting YTHDC2.
Traditionally, identifying active compounds against a new target protein has relied on high-throughput screening29, a method that is often costly and time-intensive. Recently, deep learning-based generative models have achieved remarkable success across various domains, including speech processing30,31, computer vision32,33 and natural language processing34,35. In the realm of drug discovery, this technological revolution has led to the development of a number of specialized models tailored for molecular generation36,37,38,39 Among these, receptor-based molecular generative models, such as LiGAN40, Pocket2Mol41, GraphBP42, TargetDiff43, and PocketFlow44, stand out as representative and cutting-edge approaches.
LiGAN converts protein-ligand complexes into three-dimensional (3D) atomic density grids and utilizes a 3D convolutional neural network (CNN) architecture for molecular generation. Pocket2Mol employs an E(3)-equivariant generative framework, outperforming non-equivariant models in capturing molecular graph topology and geometric constraints. GraphBP introduces a local spherical coordinate system to parameterize spatial molecular generation. TargetDiff represents a full-atom 3D equivariant diffusion model for target-aware drug design. PocketFlow applies the idea of autoregressive flow and incorporates chemical knowledge into its framework, with the bioactivities of the molecules it generates having been experimentally validated in wet-lab settings.
Despite their individual strengths and impressive benchmark performances, these models are not without their drawbacks. For instance, LiGAN falls short in achieving the desirable equivariance property, as performing 3D CNNs on atomic density grids lacks inherent equivariance. Additionally, its reliance on discretizing continuous 3D space to construct grids further limits its practical applicability. Models like LiGAN, GraphBP, and TargetDiff depend on Open Babel45 for chemical bond generation, a post-hoc approach which may inadvertently produce undesirable substructures, thereby diminishing drug-likeness of the generated molecules and complicating chemical synthesis. Furthermore, with the exception of PocketFlow, these models have primarily been evaluated in dry-lab settings, lacking validation through wet-lab experiments—a critical step in confirming their relevance to real-world drug discovery. Lastly, none of these models explicitly consider electrostatic features (including electric field intensity and electric potential) of target proteins, which are vital for ligand-protein interactions. To our knowledge, not only the models discussed here but also no existing molecular generative models currently explicitly integrate these electrostatic features into their molecular generation processes.
In this study, we develop EPMolGen, an autoregressive flow-based deep generative model which explicitly incorporates electrostatic features of target proteins into its architecture. The model’s effectiveness is systematically validated through a series of dry-lab experiments. Following this, we employ EPMolGen to generate small molecules targeting the YTH domain of YTHDC2. Subsequent wet-lab experiments lead to the identification of an active compound, H3. Further structural optimization towards H3 results in the discovery of a highly potent compound, which also exhibits cellular activity.
Results
Establishment of the generative model EPMolGen
EPMolGen is an autoregressive flow-based deep learning model designed for pocket-aware molecular generation. Unlike conventional molecular generative approaches, EPMolGen explicitly incorporates the electrostatic features of receptor proteins, which are critical determinants of ligand-receptor interactions. By embedding these features, the model is expected to enhance its capacity to generate biologically active molecules. In this study, we adopted a point-charge model, with the atomic charges derived from the OPLS4 force field46. The electric field intensity and electric potential at any specified location within the binding pocket were calculated by summing the contributions from all individual point charges. This approach ensures a physically accurate depiction of the protein’s electrostatic landscape.
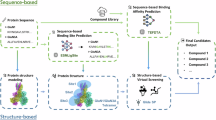
Figure 1 schematically illustrates the workflow of EPMolGen, which consists of six distinct modules. The first one is the encoder module (Fig. 1a), which encodes the context of either a protein pocket or protein-ligand complex using 3D equivariant neural networks that incorporate message passing and graph attention mechanisms. This module captures essential 3D spatial and environmental features of the binding site. The second one is the pivotal atom selection module (Fig. 1b), which selects a pivotal atom based on the encoded context. For an empty protein pocket, the pivotal atom is chosen from among the protein atoms, whereas for protein-ligand complexes, it is selected from existing ligand atoms. The third one is the atom position prediction module (Fig. 1c), which employs a Mixture Density Network (MDN)47 to predict the position of a new atom based on the encoded context from the encoder module and the feature representations from the pivotal atom selection module. This module only considers positions within 2 Å of the selected pivotal atom.
a Encoder module. Here, f denotes the equivariant graph neural networks. Yellow and light green circles represent protein atom nodes and ligand atom nodes, respectively, in the KNN (k-nearest neighbor) graph. Variations in the depth of red color indicate different attention weights. Proteins and molecules are drawn using PyMOL(Version: 3.1). b Pivotal atom selection module. The purple curve represents the protein pocket. c Atom position prediction module. Here, \(k\) represents the different Gaussian mixture components, and \({w}_{k}^{{{{\rm{p}}}}}\) denotes the coefficients of these components. d Electrostatic feature modeling module. \({{{{\bf{E}}}}}_{{{{\rm{sum}}}}}\) and \({\varphi }_{{{{\rm{sum}}}}}\) represent the total electric field intensity and electric potential, respectively, contributed by all atoms in the protein pocket at the predicted atom position. \({{{\bf{v}}}}\) and \(s\) are the vector and scalar context feature of the newly predicted atom position, respectively, derived from the output of the encoder module. e Atom type sampling module samples the atom type of the next generated atom. It is based on autoregressive flow, with only 9 types of atoms (C, N, O, F, P, S, Cl, Br and I) could be possibly sampled. f Bond type sampling module samples a chemical bond type. It is also based on autoregressive flow, with four types of chemical bonds (Single, Double, Triple, and Empty) could be possibly sampled. Here, Empty means no bond between two atoms.
The fourth one is the electrostatic features modeling module (Fig. 1d), which extracts representations of electric field intensity and electric potential, and integrates them with the encoded environmental features. The remaining two modules handle atom type sampling (Fig. 1e) and chemical bond sampling (Fig. 1f), respectively. These two components generate new atoms and chemical bonds guided by features from the previous modules. Notably, the bond type sampling module incorporates geometric constraints and chemical rules to ensure the generation of valid molecular structures. To consider geometric constraints, we introduce a triangular self-attention mechanism inspired by AlphaFold248, for modeling bond type distributions. This mechanism captures three-edge constraint relationships within the molecular graphs. For instance, if there is a triple bond between atoms i and j, it is unlikely that another atom k can simultaneously form a double bond with both atoms. Additionally, our approach limits potential bonding candidates to previously generated ligand atoms located within a 4 Å radius of the newly generated atom. To ensure chemical validity, the bond type sampling module discards proposed bonds that violate atomic valence rules or form unfavorable structural motifs (e.g., O-O, O–N, C = C = C, and three-membered rings). The model then resamples until a valid bond type is selected. This ensures that all generated molecules comply with fundamental chemical principles. Ultimately, the newly generated atom’s position, type, and chemical bonds are incorporated into the environment, serving as inputs for the next generation step.
Evaluation of the performance of EPMolGen
We assessed the performance of EPMolGen by selecting a test set of 10 diverse target proteins which have previously been used to evaluate other generative models40,44. These proteins are represented by their PDB IDs: 1bvr, 1u0f, 1zyu, 2ah9, 2ati, 2hw1, 4bnw, 4i91, 5g3n, and 5lvq. For each binding pocket of these target proteins, we generated 10,000 molecules using EPMolGen, followed by calculating the average values of several key properties of these molecules, including validity of chemical structure, quantitative estimate of drug-likeness (QED)49, octanol-water partition coefficient (logP), and synthetic accessibility (SA)50. All these properties were calculated by utilizing the RDKit tool. To benchmark EPMolGen’s performance, we also evaluated other representative generative models, including LiGAN40, Pocket2Mol41, GraphBP42, TargetDiff43, and PocketFlow44. Additionally, the corresponding property values of molecules from CrossDocked202051 were calculated to serve as reference baselines.
The average values of key properties for molecules generated by different generative models and those from CrossDocked2020 are presented in Table 1. To calculate these averages, we first calculated the results for each property across all molecules generated by each model, and then determined the mean and standard deviation (SD). Remarkably, all molecules generated by EPMolGen and PocketFlow were chemically valid, whereas TargetDiff, GraphBP, and LiGAN did not achieve complete validity. In terms of logP values, all the generative models generated molecules with average logP values within the desirable range of 0-5, thereby meeting the drug-likeness criteria. However, for specific targets, some baseline models, including TargetDiff, Pocket2Mol, PocketFlow, and LiGAN, failed to ensure that average logP values remain within this range (see Supplementary Tables S1-S4). Notably, EPMolGen showed distinct advantages in two critical metrics: QED and SA. QED provides an intuitive, transparent, and straightforward measure of a compound’s drug likeness, where a higher QED value indicates better drug-likeness. Molecules generated by EPMolGen exhibited significantly higher average QED values compared to those generated by other models. SA indicates how easily a molecule can be synthesized, where a lower SA value signifies easier synthesis. Molecules generated by EPMolGen displayed lower average SA values than those produced by other models, indicating their easier synthesis. Furthermore, as evidenced by the distribution analysis in Fig. 2, EPMolGen demonstrated tighter clustering (smaller standard deviations) in both QED and SA. This indicated that EPMolGen could maintain consistently high performance in both metrics compared to other baseline models. Detailed metrics for molecules generated by these generative models for each target are provided in Supplementary Tables S1-S7.
a QED density distributions of molecules generated by EPMolGen, Pocket2Mol, PocketFlow, GraphBP, LiGAN and TargetDiff. b SA density distributions of molecules generated by EPMolGen, Pocket2Mol, PocketFlow, GraphBP, LiGAN and TargetDiff. Data are shown as mean ± SD.
We next assessed the binding affinities of the generated molecules using ChemScore52,53. Table 1 presents the calculated average ChemScore values for all the molecules generated by these models against the 10 target proteins (average ChemScore values for each target see Supplementary Table S8). Four models—EPMolGen, PocketFlow, TargetDiff, and Pocket2Mol—demonstrated high average binding affinities. It is worth noting that GraphBP exhibited abnormal binding affinity values, likely due to many of its generated molecules clashing with protein atoms or locating outside the protein pocket. One might argue that binding affinity is heavily influenced by molecular weight, with larger molecules potentially exhibiting higher binding affinities. To address this, we evaluated ligand efficiency (LE), which measures binding affinity per heavy atom of a molecule and is often used to guide the selection and optimization of fragment hits. The calculated average LE values for all the molecules generated by these models are also presented in Table 1 (details for each target see Supplementary Table S9). Among all generative models, EPMolGen exhibited the highest average LE value, indicating superior performance.
Finally, we analyzed the proportions of molecules containing uncommon substructures within the entire set of generated molecules. These uncommon substructures encompass three-membered rings, eight-membered rings, unexpected rings (depicted in Supplementary Fig. S1a), and condensed rings (comprising ≥4 fused rings, as illustrated in Supplementary Fig. S1b). The presence of these substructures, while occuring in some drug molecules, is often associated with challenges such as high toxicity or synthesis difficulty. As presented in Table 2, EPMolGen generated the lowest proportion of molecules containing three-membered rings, eight-membered rings and unexpected rings among different generative models. Also, the proportion of uncommon structures generated by EPMolGen is nearly comparable to that of CrossDocked2020. Notably, several other baseline models produced a substantial number of molecules with uncommon substructures. For instance, 52.6% of molecules generated by Pocket2Mol contained condensed rings, suggesting a tendency towards generating highly rigid structures. Similarly, GraphBP generated a set of molecules in which 84.9% contained three-membered rings and 64.2% contained unexpected rings. LiGAN generated molecules with three-membered rings and unexpected rings in 50.9% and 32.2% of cases, respectively. Lastly, 7.7% of molecules generated by TargetDiff contain eight-membered rings.
Ablation analyses
In comparison to other reported generative models, a key distinction of our model lies in its explicit consideration of electrostatic features. To assess the influence of these features on model performance, we conducted ablation analyses. To this end, we developed a model, termed the ablation-model, which did not incorporate electrostatic features. We then calculated various properties of molecules generated by the ablation-model, using the same methodology as before. As presented in Tables 1–2, the ablation-model is capable of generating chemically valid molecules with reasonable logP values, and a very small proportion of these molecules contain uncommon substructures. However, it exhibits inferior performance compared to EPMolGen in terms of average QED, SA, LE, and ChemScore values. Notably, the LE and ChemScore values of the ablation-model are significantly lower than those of EPMolGen. The density distributions of QED, SA, LE, and ChemScore values (Supplementary Fig. S2) further show that the ablation-model performs substantially less effectively than EPMolGen. Collectively, the results of the ablation analyses underscore the benefits of explicitly considering electrostatic characteristics in enhancing model performance, particularly with regard to the binding affinities.
Application of EPMolGen provided a hit compound targeting YTHDC2
We subsequently utilized EPMolGen to discover small molecule inhibitors targeting the YTH domain of the m6A reader protein YTHDC2. The 3D structure of the receptor was derived from the crystal structure of the YTH domain of YTHDC2 (PDB ID: 6K6U), with the binding site for the m6A-mRNA substrate defined as the pocket for molecular generation. We generated 5,000 molecules using EPMolGen. From this pool, we selected molecules with QED values exceeding 0.8 and sorted them based on their LE values. Out of the top fifteen molecules, we chose five (H1-H5) that could likely be synthesized easily and quickly. These five molecules were subsequently synthesized and tested for their bioactivity. The synthetic routes for these molecules are detailed in Supplementary Fig. S3–S7.
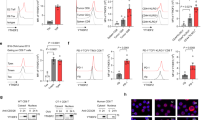
To assess the bioactivity of compounds against YTHDC2, we expressed and purified the recombinant human YTHDC2 YTH-domain protein (Supplementary Fig. S8a–c). Leveraging the YTH-domain’s specific recognition of m6A, we established a fluorescence polarization (FP)-based assay using the fluorescein amidite (FAM)-labeled m6A-mRNA. (Fig. 3a)54,55. For the FP assays, we opted to use the GST-tagged YTHDC2-YTH protein instead of the tag-free YTHDC2-YTH protein, as the latter’s relatively small molecular weight (< 16 kDa) resulted in inadequate signal intensity (Supplementary Fig. S9a). Furthermore, our findings indicated that the GST-tagged YTH domain protein provided enhanced FP signals compared to the tag-free protein, without interfering with the specific recognition of m6A by the YTH protein (Fig. 3b, and Supplementary Fig. S9a–h). Out of the five compounds generated by EPMolGen, only H3 demonstrated inhibitory activity, with an IC50 value of 16.84 ± 2.04 μM (Fig. 3c–g). Subsequently, we conducted structural optimization and structure-activity relationship (SAR) analyses on H3.
a Schematic diagram of the screening method (FP) for small molecule inhibitors of YTHDC2. b Results of FP assay using FAM labeled mRNA/m6A-mRNA, incubated with GST-YTHDC2. Data are shown as the mean ± SD from three technical replicates in three independent experiments. c-g Dose-activity curves of compounds (H1-H5) against YTHDC2 measured by the FP assay. Data are shown as the mean ± SD from three technical replicates in three independent experiments. h Regions selected for the structural optimization of H3. Source data are provided as a Source Data file.
Structural optimization and structure-activity relationship analyses of H3
The structural optimization of H3 was focused on two regions: the benzyl group (R1) and the 4-chlorophenol moiety (R2) (Fig. 3h). In the first step, we optimized R1 while keeping R2 unchanged. We synthesized a total of 13 new compounds (14a-m) with various substituents at R1 (Supplementary Fig. S10). The bioactivities of these compounds are summarized in Fig. 4. Fluorine substitutions at different positions on the benzene ring did not improve the bioactivity (14a-c); but the para-fluorine substitution is relatively more tolerable. Then chloride and bromine were used to replace the para-fluorine, nevertheless the bioactivity was not improved (14d-e). Further substitutions with tert-butyl and phenyl at the para-position led to a complete loss of bioactivity (14f-g). Notably, ester and carboxyl substitutions (14h-i) either enhanced or maintained bioactivity. Next, we introduced double substituents (including an ester) on the benzene ring. The resulting compound 14j exhibited nearly a 6-fold increase in bioactivity compared to the original hit compound (H3). The cyclization product 14k, with para- and meta-substituents, also showed improved bioactivity compared to H3. However, compound 14l, containing a naphthalene group, did not display activity. Surprisingly, when R1 was replaced with a dibenzo[b,e]oxepin-like moiety (14m), the activity increased further, with an IC50 value of 1.56 µM, suggesting that the dibenzo[b,e]oxepin-like moiety is an optimal substituent for R1.
Data are shown as mean ± SD of three independent experiments.
In the second step, we fixed the dibenzo[b,e]oxepin-like moiety at R1 position, and replaced the phenyl group of R2 with a variety of substituents. A total of 14 new compounds (16a-n) were synthesized (Supplementary Fig. S11). Bioactivities of compounds 16a-n are displayed in Fig. 4. Initially, we investigated the impact of the phenol hydroxyl group and observed that the activity was lost when the phenol hydroxyl group was removed (16a), indicating a critical role of the phenol hydroxyl group to the bioactivity. Subsequently, the position of chlorine was changed or removed (16b-e), and we found that the ortho-substitution of chlorine (16d) slightly increased the potency, while other changes resulted in loss of activity. Then, replacement of the chlorine group by fluorine (16f), bromine (16g), methyl (16h), and trifluoromethyl (16i) decreased the potency. Surprisingly, the introduction of chlorine at positions 4 and 6 of the benzene ring (16j) enhanced the activity by approximately 3-fold. We thus introduced different substitutions at these positions of the benzene ring and synthesized four additional compounds (16k-n). Among them, compound 16k (hereafter called DC2-C1) with a 4-bromo-6-chlorophenol at R2 position, displayed the highest potency with an IC50 value of 0.168 µM.
Bioactivity characterization of compound DC2-C1 against YTHDC2
Through structural optimization and SAR studies conducted on the initial hit compound H3, we successfully developed compound DC2-C1 (Fig. 5a). This compound exhibited remarkable inhibitory potency in the FP assay, with an IC50 value of 0.168 ± 0.02 µM (Fig. 5b), marking a 100-fold enhancement in potency relative to the original hit compound H3. In the FP assay for DC2-C1, the fluorescence intensities remained consistently stable, indicating the absence of any fluorescent interference from the compound (Supplementary Fig. S12a–c).
a Chemical structure of DC2-C1. b Dose-activity curves of DC2-C1 measured by the FP assay. Data are shown as the mean ± SD from three technical replicates in three independent experiments. c ITC binding curves for YTHDC2 with DC2-C1. Data are shown as the mean ± SD from three independent experiments. d, e The effect of DC2-C1 on the thermal stability of YTHDC2 was detected using DSF. The concentrations of DC2-C1 from high to low are 6.25 µM, 3.13 µM, 1.56 µM, 0.78 µM, and 0 µM. Data are shown as the mean ± SD from three technical replicates in three independent experiments. f Dose-activity curves of DC2-C1 against m6A “reader” proteins measured by the FP assay. Data are shown as the mean ± SD from three technical replicates in three independent experiments. Source data are provided as a Source Data file.
To further substantiate the bioactivity of DC2-C1 in vitro, we employed a battery of biochemical and biophysical techniques. An Isothermal Titration Calorimetry (ITC) assay was utilized to evaluate the binding affinity of DC2-C1 to YTHDC2, revealing a robust binding affinity with an equilibrium dissociation constant (KD) of 0.46 ± 0.10 μM (Fig. 5c). Additionally, a Differential Scanning Fluorimetry (DSF) assay demonstrated that DC2-C1 dose-dependently stabilized the YTHDC2 protein, inducing a thermal shift (∆Tm) of 2.18 ± 0.21 °C at a concentration of 6.25 μM (Fig. 5d, e). Furthermore, based on the FP-based screening assay, DC2-C1 exhibited a more pronounced inhibitory effect on YTHDC2 compared to other YTH-domain-containing m6A readers, underscoring its selectivity for YTHDC2 (Fig. 5f).
Binding mode of DC2-C1 with YTHDC2
To explore the possible binding mode between DC2-C1 and YTHDC2, we performed docking analysis to identify a putative binding mode after unsuccessful co-crystallization attempts. Given that the hit compound was generated in the m6A binding pocket, we hypothesized that the DC2-C1’s binding site is likely to be shared with the m6A-binding site. In order to obtain a more precise binding mode of DC2-C1 and YTHDC2, we conducted induced fit docking using Glide56. Our docking result suggests that DC2-C1 resides perfectly in the m6A binding pocket of YTHDC2, thus could block YTHDC2 from binding to m6A-mRNA (Fig. 6a). In comparison with the generated pose of YTHDC2-H3, it was found that the binding mode of DC2-C1 with YTHDC2 is similar to that of H3 (Supplementary Fig. S13). DC2-C1 forms multiple interactions with the m6A-binding site of YTHDC2 (Fig. 6a). Specifically, the hydroxyl group on 2-amino-4-bromo-6-chlorophenol of DC2-C1 forms a hydrogen bond with residue N1300. The 2-amino-4-bromo-6-chlorophenol engages in hydrophobic interactions with residues I1303, I1309, and L1365. The nitrogen atom on the amide bond of DC2-C1 establishes a hydrogen bond with residue S1311. The oxygen atom on the amide bond forms hydrogen bonds with residues N1300 and S1295. Furthermore, each of the two benzene rings in the dibenzo[b,e]oxepin-like moiety of DC2-C1 form a pi-anion interaction with residue D1402, respectively.
a Predicted binding mode of DC2-C1 with YTHDC2 (the 3D structure of YTHDC2 was taken from PDB ID: 6K6U), proteins and molecules are drawn using PyMOL(Version: 3.1). b DSF analysis of DC2-C1 (6.25 µM) binding to the recombinant mutated YTHDC2 proteins. Data are shown as the mean ± SD from three technical replicates in three independent experiments. c, d UVPD fragmentation propensity comparisons of YTHDC2 7+ versus and YTHDC2-DC2-C1 7+ based on the summed abundances of a/x ions, and the results were mapped to the crystal structure. Blue indicates suppression, and red indicates enhancement of UVPD FYs, the green area represents the possible binding pockets. Data are shown as the mean ± SD from three independent experiments. Source data are provided as a Supplementary Data file. e Schematic summary of the chemical modification towards DC2-C1, providing the negative control compound 16o. f The CETSA assay was carried out with Flag-YTHDC2-(YTH domain) transfected HEK293T cells grown in the presence or absence of DC2-C1/16o. Blots shown are representative of three biological replicates. Full blots can be found in Supplementary Fig. S27. g The DARTS assay was carried out with Flag-YTHDC2-(YTH domain) transfected HEK293T cells grown in the presence or absence of DC2-C1/16o. Blots shown are representative of three biological replicates. Full blots can be found in Supplementary Fig. S28. Source data are provided as a Source Data file. (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; by two-tailed unpaired Student’s t-test).
To verify the predicted binding mode, we conducted mutation experiments. According to the molecular docking results, DC2-C1 establishes crucial interactions with residues D1402, I1303, I1309, L1365, N1300, S1311, and S1295 of YTHDC2. Consequently, we individually mutated these seven residues to alanine and successfully generated six YTHDC2 mutants (I1303A, I1309A, L1365A, N1300A, S1311A, and S1295A); we failed to obtain the D1402A mutant due to compromised protein stability. To assess the bioactivity of DC2-C1, we employed DSF assays using these mutated YTHDC2 proteins. Our results showed that all the six mutations significantly reduced the bioactivity of DC2-C1 (Fig. 6b), thereby validating the binding mode of DC2-C1 with YTHDC2.
To further validate the interaction site of YTHDC2 with DC2-C1, we adopted Nuclear Magnetic Resonance (NMR), a commonly used technology to determine the binding site of a ligand with a receptor protein57. We attempted NMR experiments but encountered challenges due to DC2-C1’s poor water solubility, which precluded meaningful data acquisition under standard NMR conditions. To overcome this limitation, we chemically modified DC2-C1 to generate a more water-soluble analog, 16p (IC50 = 1.88 ± 0.07 µM), and used it for NMR analysis (Supplementary Fig. S14 and S15). The NMR data revealed that amino acid residues N1297, N1300, L1301, K1307, T1312, T1313, K1319, A1323, and I1352—which exhibited chemical shift perturbations (CSPs) upon 16p binding—are located within or adjacent to the m6A recognition pocket (Supplementary Fig. S16a–g and S17a–b). These findings suggest that YTHDC2 interacts with 16p specifically at the m6A recognition pocket.
Although we have established via NMR that the analog 16p of DC2-C1 binds to the m6A recognition pocket of YTHDC2, we cannot definitively confirm whether DC2-C1 itself also targets the same site, given their distinct chemical structures. To address this uncertainty, we employed native top-down mass spectrometry (MS) combined with 193-nm ultraviolet photodissociation (UVPD), a powerful tool for elucidating dynamic structural changes in proteins, which has been applied to the identification of ligand binding sites. A notable advantage of UVPD is its ability to operate without requiring high concentrations of small molecules, making it particularly well-suited for our current investigation58,59,60,61. In this assay, the YTHDC2 protein in solution was directly introduced into the MS system via electrospray ionization (ESI) under native-like conditions. Subsequently, target ions were selectively isolated and exposed to UV radiation to induce fragmentation, thereby providing structural insights into binding sites, conformations, and interaction patterns from the resulting fragment ions. During native MS characterization, YTHDC2 was successfully transferred to the gas phase under native ESI conditions, generating noncovalent complexes containing either DC2-C1 or 16p. Both the YTHDC2 (apo) and its complex ions (holo) with a charge state of 7+ were isolated and subjected to UVPD analysis, achieving over 87% sequence coverage for both apo and holo ions (Supplementary Fig. S18 and S19).
Protein regions engaged in non-covalent interactions following ligand binding typically exhibit reduced fragmentation efficiencies during UVPD. Therefore, the fragmentation efficiency of each backbone linkage in YTHDC2 was quantified based on the intensities of the fragment ions generated by cleavage of that linkage. The results revealed a decrease in fragmentation efficiencies for most YTHDC2 backbones upon binding to DC2-C1 or 16p (Fig. 6c and Supplementary Fig. S20). Notably, the region (highlighted in green) with reduced fragmentation efficiency corresponds to the reported m6A-binding pocket for both compounds (Fig. 6d and Supplementary Fig. S20). These UVPD findings further substantiate that both DC2-C1 and 16p bind to the m6A recognition pocket of YTHDC2.
DC2-C1 binding to YTHDC2 in living cells
Next, we proceeded to examine the cellular activity of DC2-C1. Prior to these investigations, we synthesized a negative control compound, 16o (Supplementary Fig. S11), which is an analog of DC2-C1 (Fig. 6e). Compound 16o was synthesized by removing the halogen atoms from DC2-C1 and introducing a methoxy group. In the FP assay, compound 16o exhibited no obvious activity against YTHDC2 (IC50 > 50 μM) (Supplementary Fig. S21a) and showed no fluorescent interference (Supplementary Fig. S21b–d). We proposed that this inactivity is due to a steric clash between the methoxy substituent and residue I1309 of YTHDC2, based on the predicted binding mode (Fig. 6a and Supplementary Fig. S21e).
The cellular thermal shift assay (CETSA) and drug affinity responsive target stability (DARTS) assays were adopted to examine the direct interaction between DC2-C1 and YTHDC2 in living cells. It is important to note that the YTH domain constitutes only a minor segment of YTHDC2. Consequently, we posited that relying on full-length YTHDC2 Western blot to assess DC2-C1’s binding capacity to the YTH domain in CETSA and DARTS might not provide a specific evaluation. We thus, in these assays, used HEK293T cells transfected with a pCDH-GFP-3xFlag-YTHDC2 (YTH domain) plasmid. DC2-C1 treatment led to substantial shifts of the thermal stability of YTHDC2 protein in CETSA (Fig. 6f and Supplementary Fig. S21f). Moreover, DC2-C1 could block pronase-induced proteolysis of YTHDC2 (Fig. 6g and Supplementary Fig. S21g), further confirming their direct interactions. In contrast, 16o did not exhibit any effect in CETSA and DARTS assays (Fig. 6f, g and Supplementary Fig. S21f, g).
It is important to note that the use of cells overexpressing the YTH domain of YTHDC2 in CETSA and DARTS assays presents certain limitations, as this approach may not accurately capture the genuine binding interaction between DC2-C1 and YTHDC2 in wild-type cells. Consequently, we plan to validate the regulatory impact of DC2-C1 on YTHDC2 specifically in wild-type cells in subsequent experiments.
In vitro cellular activity of DC2-C1 in RA-FLS
As mentioned above, YTHDC2 contributes to RA progression by promoting joint destruction through the hyperproliferative and invasive behavior of RA-FLS. Disruption of YTHDC2’s m6A-binding capacity in RA-FLS destabilizes its target, AMIGO2 mRNA, leading to reduced AMIGO2 protein expression. This downregulation attenuates RA-FLS proliferation, migration, and invasion, thereby alleviating RA symptoms18. Consistent with these findings, CRISPR-mediated knockout of YTHDC2 in MH7A cells (a human RA-FLS line) significantly diminished AMIGO2 protein levels (Supplementary Fig. S22).
We then assessed the effect of DC2-C1 on the binding between YTHDC2 and AMIGO2 mRNA using the RIP-qPCR (RNA immunoprecipitation-qPCR) assay. DC2-C1 treatment markedly reduced YTHDC2-AMIGO2 mRNA binding in MH7A cells (Fig. 7a). RNA stability assays further revealed that DC2-C1 substantially accelerated AMIGO2 mRNA degradation (Fig. 7b), accompanied by dose-dependent reductions in AMIGO2 mRNA (Fig. 7c) and protein levels, while GAPDH remained unaffected (Fig. 7d, e). In contrast, the negative control compound 16o showed no obvious effects in these assays (Fig. 7a-d). These results demonstrate that DC2-C1 intracellularly inhibits YTHDC2 binding to AMIGO2 mRNA, disrupting its stabilizing role and reducing AMIGO2 expression. It is necessary to mention that we cannot completely rule out the possibility that DC2-C1 may affect the expression of AMIGO2 through other regulatory mechanisms.
a Interaction analyses between YTHDC2 and AMIGO2 mRNA in RA-FLS (treated with DMSO, DC2-C1 or 16o) determined by the RIP-qPCR assay. Data are shown as the mean ± SD from three technical replicates in three independent experiments. b The RNA decay was detected in DC2-C1- or 16o-treated RA-FLS after incubation with actinomycin D for the indicated times (normalized to 0h). Data are shown as the mean ± SD from four technical replicates in four independent experiments. c AMIGO2 mRNA expression was validated by RT-qPCR in DC2-C1- or 16o-treated RA-FLS. Data are shown as the mean ± SD from four technical replicates in four independent experiments. d Western blot analysis of AMIGO2 expression in DC2-C1- or 16o-treated RA-FLS. Blots shown are representative of three biological replicates. Full blots can be found in Supplementary Fig. S29. e Quantitative analysis of Western blot for MH7A cells. Data are shown as the mean ± SD from three independent experiments. f Colony formation assay of MH7A cells treated with DC2-C1 or 16o. g Quantitative analysis of colony formation for MH7A cells. Data are shown as the mean ± SD from three independent experiments. h Abilities of migration and invasion of MH7A cells treated with DC2-C1 or 16o. Scale bar, 50 μm. i Quantitative analysis of migration and invasion for MH7A cells. Data are shown as the mean ± SD from three independent experiments. Source data are provided as a Source Data file. (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; by two-tailed unpaired Student’s t-test).
We finally evaluated DC2-C1’s impact on MH7A cell phenotype using functional assays. In the colony formation assay, DC2-C1 substantially inhibited MH7A cell clonogenic potential (Fig. 7f, g). Transwell assays revealed that DC2-C1 significantly impaired both cellular migration and invasive capacity (Fig. 7h, i). In all the assays, the negative control compound 16o had no discernible effects (Fig. 7f–i).
In vitro cellular activity of DC2-C1 in PDAC cells and hepatocytes
We next investigated the effects of DC2-C1 on PDAC cells, building on prior evidence that YTHDC2 knockout suppresses malignant phenotypes in this context. Mechanistically, YTHDC2 promotes histone H3 lysine 4 trimethylation (H3K4me3) in an m6A-dependent manner, facilitating local chromatin accessibility and oncogene expression in PDAC cells62. Consistent with these findings, CRISPR-mediated YTHDC2 knockout in PDAC cells significantly reduced H3K4me3 levels (Supplementary Fig. S23a, b). Treatment of PANC-1 and MiaPACA-2 cells with DC2-C1 induced a dose-dependent decrease in H3K4me3 expression (Fig. 8a, b). Functionally, DC2-C1 markedly suppressed the clonogenic potential and invasion of both cell lines, whereas the negative control compound 16o exhibited no noticeable inhibitory effects (Fig. 8c, d and Supplementary Fig. S23c, d).
a Western blot analyses of H3K4me3 expression in DC2-C1-treated PANC-1 or MiaPACA-2 cells. Blots shown are representative of three biological replicates. Full blots can be found in Supplementary Fig. S30. b Quantitative analysis of Western blot for PANC-1 or MiaPACA-2 cells. Data are shown as the mean ± SD from three technical replicates in three independent experiments. c Colony formation assay of PANC-1 or MiaPACA-2 cells treated with DC2-C1 or 16o. d Abilities of invasion of PANC-1 or MiaPACA-2 cells treated with DC2-C1 or 16o. Scale bar, 50 μm. e-g ACC1, FASN and SREBP-1C mRNA expression were determined by RT-qPCR in DC2-C1-treated HepG2 cells. Data are shown as the mean ± SD from three technical replicates in three independent experiments. h Western blot analyses of ID2 expression in DC2-C1-treated PANC-1 cells; Western blot analyses of FZD7 expression in DC2-C1-treated HGC27 cells. Blots shown are representative of three biological replicates. Full blots can be found in Supplementary Fig. S31. i FZD7 mRNA expression was determined by RT-qPCR in DC2-C1-treated HGC27 cells. Data are shown as the mean ± SD from four technical replicates in four independent experiments. j ID2 mRNA expression was validated by RT-qPCR in DC2-C1-treated PANC-1 cells. Data are shown as the mean ± SD from four technical replicates in four independent experiments. Source data are provided as a Source Data file. (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; by two-tailed unpaired Student's t-test).
Previous studies established that YTHDC2 plays a critical role in regulating hepatic lipogenesis and triglyceride (TG) homeostasis. Specifically, YTHDC2 knockdown in HepG2 cells was shown to upregulate mRNA levels of key lipogenic genes—ACC1, FASN, and SREBP-1C27. We confirmed these results via CRISPR-mediated YTHDC2 knockout in HepG2 cells (Supplementary Fig. S24a-d). Consistent with these, DC2-C1 treatment induced a dose-dependent increase in ACC1, FASN, and SREBP-1C mRNA levels (Fig. 8e-g).
Finally, we examined the selectivity of DC2-C1 in living cells. To this end, we selected two mRNAs as examples, which are targets of other YTH proteins but not targets of YTHDC2. These RNAs are FZD7 and ID2, targets of YTHDF1 and YTHDF2, respectively63,64. Our results showed that DC2-C1 treatment did not alter mRNA/protein expression of FZD7 in HGC27 cells and that of ID2 in PANC-1 cells (Fig. 8h-j and Supplementary Fig. S25a-d), indicating good selectivity in living cells.
Discussion
The m6A-modification of RNA plays a pivotal role in various physiological and pathological processes. Dysregulation of the proteins governing m6A modifications has been implicated in a myriad of diseases, encompassing immune disorders, cancer, metabolic syndromes, and neurological conditions. The development of inhibitors targeting m6A regulatory proteins is of great significance for the treatment of major diseases caused by m6A dysregulation and for in-depth research in this field. Currently, multiple potent inhibitors have been reported for the methyltransferase METTL365, demethylases FTO66,67, and ALKBH568,69, as well as other YTH family proteins (YTHDF1-357,70,71, and YTHDC172. These inhibitors have shown promising therapeutic potential in various diseases. However, no potent inhibitors targeting YTHDC2 have been reported to date.
Identifying active compounds against a novel target such as YTHDC2 presents a formidable challenge. To swiftly obtain hit compounds while circumventing the high costs associated with high-throughput screening, we leveraged an AI-based generative model to design new ligands in this study. To achieve this, we developed a cutting-edge deep learning generative model named EPMolGen, which sets itself apart from other established models by explicitly incorporating electrostatic features. Ablation experiments underscored the advantages of considering electrostatic characteristics in enhancing model performance. Utilizing this model, we generated small-molecule inhibitors targeting YTHDC2. From the generated molecules, five compounds were selected for chemical synthesis, with one compound exhibiting activity with an IC50 of 16.84 μM. Further structural optimization and SAR studies led to the discovery of a potent YTHDC2 inhibitor, DC2-C1, boasting an IC50 of 0.168 μM. This compound demonstrated good selectivity against other YTH proteins and exhibited low cytotoxicity (Supplementary Fig. S26a). Notably, DC2-C1 significantly reduced the expression levels of multiple targets of YTHDC2, thereby inhibiting the phenotypes of related cells. However, DC2-C1 serves only as a lead compound and warrants further investigation; in silico evaluation of its pharmacokinetic properties and toxicity revealed that while it possesses good drug-like potential, it also exhibits some less favorable properties, such as poor water solubility, and the inhibition of certain metabolic enzymes (see Supplementary Fig. S26b-c and Supplementary Table S10). Additionally, in the current version of EPMolGen, we incorporated only electrostatic contributions from receptor proteins due to the characteristics of the autoregressive flow-based model, which demonstrated marked performance improvements over alternative models. We hypothesize that considering electrostatic interactions from both receptor proteins and small-molecule ligands could further enhance performance, a feature we plan to incorporate in future updates. In conclusion, our proposed molecular generative model facilitated the discovery of a potent and cell-active lead compound against YTHDC2, underscoring the successful application of deep learning in the early stages of drug discovery.
Methods
Data sets and preparation
We pretrained and fine-tuned the model using data selected from the ZINC(downloaded in December, 2022)73 and CrossDocked2020 database, respectively. The molecules selected from the ZINC database adhere to the following criteria: (1) containing no more than the nine types of atoms: C, N, O, F, P, S, Cl, Br, and I; (2) having molecular weights ranging from 300 to 600 Daltons; (3) exhibiting a QED score above 0.6; (4) possessing a maximum ring size of less than 8; and (5) having a smallest set of smallest rings (SSSR) fewer than 6. Among them, criteria (1)-(3) are related to the common drug-like properties. Criterion (4) was applied because molecules containing rings with 8 or more members are often more challenging to synthesize and, in some cases, exhibit reduced chemical stability compared to smaller-ringed analogs. Criterion (5) was used because molecules with an SSSR of 6 or more are typically difficult to synthesize and may have a higher propensity for cytotoxicity relative to those with smaller SSSR (such as many natural product molecules).
Regarding the CrossDocked2020 dataset, the selected complexes should meet the following requirements: the RMSD (root mean square deviation) of binding poses should be less than 1\({\rm A}\); no more than 9 types of atoms, namely C, N, O, F, P, S, Cl, Br, and I, can be included in small molecules.
The statistical summary for pretraining and fine-tuning datasets is shown in Supplementary Table S11.
The protein-ligand complex was modeled using heterogeneous K-nearest neighbor (KNN) graphs, wherein we explicitly accounted for the covalent bonds between atoms, rather than treating them solely as a point cloud model. In a KNN graph, nodes and edges are key components: protein and ligand atoms were represented as nodes, while the connections between these nodes were depicted as edges. Further details regarding the attributes and descriptions of these heterogeneous KNN graphs are provided in Supplementary Table S12. Finally, protein preparation was performed using the Glide OPLS4 force field46 to determine the partial charge of each atom within the protein pocket.
Encoder and pivotal atom selection
The encoder module of EPMolGen is constructed by 6 blocks which could extract geometric features of either proteins or protein-ligand complexes. Each block is mainly built up by an equivariant graph neural network with graph attention and message passing mechanisms. Message passing mechanism combines the messages between atoms to extract the geometric features of neighbor atoms. Graph attention mechanism is used for further capturing the local and global geometric features. The details of graph attention and message passing mechanism are displayed in section 4 in the Supplementary Information.
The pivotal atom selection module is mainly constructed using the same type of equivariant neural network as encoder. It takes the vector and scalar features from the encoder module as its input. During the initial autoregressive step, when no ligand atoms are present in the protein pocket, the module selects a pivotal atom from protein pocket atoms. In all subsequent steps, the selection is made from existing ligand atoms.
Atom position prediction
To predict the new atom position, we first estimate the relative position of the new atom to the pivotal atom. Here EPMolGen models the probability distribution of three-dimensional relative atom coordinates based on Gaussian Mixture Model:
Here, \(n\) is the number of Gaussian components, \(\Delta {{{{\bf{r}}}}}^{{{{\rm{p}}}}}\) is the relative three-dimensional coordinates from the selected pivotal atom \({{{\rm{p}}}}\). \(k\) denotes different Gaussian mixture components, \({w}_{k}^{{{{\rm{p}}}}}\) represents the coefficient of the component, \({{{{\boldsymbol{\mu }}}}}_{k}^{{{{\rm{p}}}}}\) is the mean vector of the component, \({{{{\boldsymbol{\Sigma }}}}}_{k}^{{{{\rm{p}}}}}\) stands for the covariance matrix of the component. In EPMolGen, n is set to 3, indicating that 3 components are considered to model the probability distribution of three-dimensional relative atom coordinates.
We applied a Mixture Density Network (MDN) to realize the Gaussian Mixed Model. The MDN directly predicts \({w}_{k}^{{{{\rm{p}}}}}\), \({{{{\boldsymbol{\mu }}}}}_{k}^{{{{\rm{p}}}}}\) and \({{{{\boldsymbol{\Sigma }}}}}_{k}^{{{{\rm{p}}}}}\) for each component \(k\). In order to keep the whole molecular generating process equivariant, we directly select the \({{{{\boldsymbol{\mu }}}}}_{k}^{{{{\rm{p}}}}}\) with the maximum \({w}_{k}^{{{{\rm{p}}}}}\), as the predicted relative coordinates of the new atoms. The details of MDN are shown in the Supplementary Methods.
Electrostatic features modeling
As shown in Fig. 1d, electrostatic features modeling module is mainly constructed by 4 blocks. Two of them (\({\vec{f}}_{1}\), \({\vec{f}}_{2}\)) are for vectors, which are VN-MLPs74, while the other two (\({f}_{1}\), \({f}_{2}\)) are for scalars, which are MLPs. A VN-MLP is constructed by a vector neuron for down-sampling vector features, an activation function for vector features and a vector neuron for up-sampling vector features. An MLP is constructed by a linear layer for down-sampling scalar features, an activation function for scalar features and a linear layer for up-sampling scalar features:
Where \({f}_{{{{\rm{VN}}}}1}^{\downarrow }\) and \({f}_{{{{\rm{VN}}}}2}^{\downarrow }\) are vector neurons which down-sample vector features; \({f}_{{{{\rm{VN}}}}1}^{\uparrow }\) and \({f}_{{{{\rm{VN}}}}2}^{\uparrow }\) are vector neurons which up-sample vector features. \({f}_{{{{\rm{lin}}}}1}^{\downarrow }\) and \({f}_{{{{\rm{lin}}}}2}^{\downarrow }\) are linear layers which down-sample scalar features; \({f}_{{{{\rm{lin}}}}1}^{\uparrow }\) and \({f}_{{{{\rm{lin}}}}2}^{\uparrow }\) are linear layers which up-sample scalar features. \({\sigma }_{{{{\rm{v}}}}}\) is an activation function using VN-LeakyReLU for vector features, while \({\sigma }_{{{{\rm{s}}}}}\) is an activation function using LeakyReLU for scalar features.
\({{{{\bf{E}}}}}_{{{{\rm{sum}}}}}\) and \({\varphi }_{{{{\rm{sum}}}}}\) denote the electric field intensity and electric potential, respectively, which are calculated by summing up the corresponding values generated by various point charges:
Where \({Q}_{i}\) is the partial charge of an atom in the protein pocket. \({C}_{i}\in {{\mathbb{R}}}^{3}\) is the coordinates of pocket atom i. \({C}_{{{{\rm{pred}}}}}\in {{\mathbb{R}}}^{3}\) is the predicted new atom coordinates. \({d}_{i}\) is the distance between \({C}_{i}\) and \({C}_{{{{\rm{pred}}}}}\). \(\varepsilon\) is the permittivity. Here the permittivity of water was used.
The \({{{\bf{v}}}}\) and \({{{\rm{s}}}}\) are the vector and scalar features of context of \({C}_{{{{\rm{pred}}}}}\):
where \({h}_{i}\) and \({{{{\bf{h}}}}}_{i}\) denote the scalar and vector features of the i-th neighbor of \({C}_{{{{\rm{pred}}}}}\) in the KNN graph, which are computed using \({C}_{{{{\rm{pred}}}}}\) and \({C}_{{{{\rm{ctx}}}}}\). \({C}_{{{{\rm{ctx}}}}}\) are the positions of existed nodes.
Atom type sampling and bond type sampling
EPMolGen applied a special type of flow-based model, autoregressive flow, to model the atom type and bond type distributions as well as to sample new atom and bond types. The objective of flow-based model is to build an invertible transformation between the real-world distribution \({{{\bf{x}}}} \sim {p}_{{{{\bf{x}}}}}({{{\bf{x}}}})\) and a base distribution \({{{\boldsymbol{z}}}} \sim {p}_{{{{\bf{z}}}}}({{{\bf{z}}}})\). A normalizing flow is a type of flow-based model, which aims to transform a base distribution (normal distribution for example) into a complex distribution by utilizing a sequence of invertible transformation functions.
In EPMolGen, both atom type sampling and bond type sampling are implemented based on normalizing flows. In each case, the entire sequence of flow transformations is structured as an autoregressive process, which is an autoregressive flow. For the \(i\)-th generation step in autoregressive flow, the result \({{{{\bf{x}}}}}_{i}\) is based on the results of the previous \(i-1\) steps: \({{{{\bf{x}}}}}_{1:i-1}\). As shown in Formula:
where \(\otimes\) is element-wise product. Therefore,
During these processes, we only considered scalar features in the invertible transformations, while vector features provide additional information in the generation of new atom and bond types.
Training method
To address the challenge posed by limited protein-ligand complex data, we adopted a transfer learning approach for model training, which comprises two stages: pretraining and fine-tuning.
During both pretraining and fine-tuning, the datasets were randomly divided into training and valid set. The loss of valid set was used to monitor the training status of model, such as adjusting learning rate, preventing overfitting or drastic changes of gradient. After applying the filtering criteria to the ZINC dataset, a total of 10,019,815 compounds were randomly selected as the pretraining dataset. From this pool, 10,000 samples were randomly divided as the valid set, while the remaining 10,009,815 compounds constituted the training set. In the fine-tuning stage, a total of 157,960 protein-ligand complexes remained after filtering from CrossDocked2020 dataset, we randomly selected 100 protein-ligand complexes as the valid set, while the remaining samples were used for training (see Supplementary Table S11).
During the pretraining stage, we set the batch size to 128. It’s worth noting that we employed gradient accumulation during fine-tuning. This was necessary because, unlike the pretraining dataset, the fine-tuning dataset contains protein-ligand complexes, necessitating small batch sizes to avoid GPU memory overflow. However, small batch size may compromise the model’s stability and training convergence speed. Gradient accumulation mitigates this issue by dividing the original batch into several mini-batches, calculating the gradients for each mini-batch in sequence, and then accumulating them for backpropagation. Specifically, we set the mini-batch size to 1, with gradient accumulation performing over 8 mini-batches.
During the model training phase, EPMolGen takes protein pockets and ligand trajectories as input, both of which are represented using heterogeneous KNN graphs (k = 16). It is worth noting that in the pretraining stage, we designate the protein pockets as blank, as the pretraining data samples do not include protein pockets. Self-supervised learning is employed throughout the model training process. At each step, an atom and its covalent bonds are masked, and EPMolGen is trained to recover the masked elements based on the remaining ones.
Training of EPMolGen requires optimizing four loss functions simultaneously. The overall loss function \({L}_{{{{\rm{total}}}}}\) is the summation of four loss functions:
Here, \({L}_{{{{\rm{piv}}}}}\) denotes the binary cross entropy loss of the predicted pivotal atom. \({L}_{{{{\rm{pos}}}}}\) represents the loss of the new atom position, which is the negative log likelihood of the masked atom positions. \({L}_{{{{\rm{atom}}}}}\) is the negative log likelihood of masked atom, which indicates the loss of new atom type. The loss of new bond type, denoted as \({L}_{{{{\rm{bond}}}}}\), is the negative log likelihood of masked chemical bond.
We mainly applied PyTorch and PyTorch Geometric to construct EPMolGen. An NVIDIA A800 80 G GPU was adopted to train the model. The Adam optimizer was used for gradient descent optimization. Specifically, the initial learning rate was set as \(2\times {10}^{-4}\), the minimum learning rate was set as \(1\times {10}^{-5}\), the decay rate was set as 0.6 and the patience was set as 10. Gradient clipping was applied to the model training process to avoid gradient explosion and ensure training stability.
Protein expression and purification
Human YTHDC2 (residues 1287-1423) was subcloned into the pGEX-6P-1 vector. Proteins were over-expressed in Escherichia coli BL21 (DE3) cells grown in Luria Broth media. The media was treated with Ampicillin. Protein expression was induced with 0.2 mM isopropyl-1-thio-D-galactopyranoside for 15 h at 16 °C. GST-tag could be removed by PreScission Protease. All recombinant proteins were further purified by SuperdexTM 200 Increase 10/300 GL (GE Healthcare). The purified proteins were stored at -80 °C in the buffer containing 20 mM Tris-HCl pH 7.5 and 150 mM NaCl. The instrument used for protein purification was the ÄKTA pure (Cytiva), and the software used for data collection was UNICORN 7.0. The mutated YTHDC2 proteins were expressed and purified using the same method. The expression and purification conditions for Human YTHDF1 (residues 366-558), Human YTHDF2 (residues 383-553), Human YTHDF3 (residues 393-585), and Human YTHDC1 (residues 345-509) were consistent with those for YTHDC2. The plasmids were purchased from Tsingke Biotechnology Co., Ltd. UV absorption spectra of proteins were collected using NanoDrop One (Thermo Scientific).
Fluorescence polarization (FP) assay
Fluorescence Polarization (FP) experiments were performed in the buffer containing 20 mM HEPES pH 7.5, 180 mM NaCl. GST-tagged YTHDC2 proteins (650 nM) were incubated with 7 nM probe (FAM-m6A-mRNA). FAM-m6A-mRNA sequence: FAM-UUCUUCUGUGG-(m6A)-CUGUG. The FAM tag is a fluorescent dye whose degree of polarization changes with binding status, thereby indicating molecular interactions. Experiments were performed in 384-well black plates and FP reading was performed in the Plate Reader (CLARIOstar PLUS). For the compound screening assay, compounds and protein were incubated for 30 min, and then the probe was added to the assay system for further incubation for 2 h. To minimize the risk of RNase contamination, RNase-free water was used for preparing mRNA solutions, and RNase-free consumables were used in FP assays. The data were fitted using GraphPad Prism software.
In the FP assays, to ensure that all YTH family proteins could specifically recognize m6A to the same extent, the experimental conditions for different proteins were not entirely consistent (the concentration of NaCl affects the specific recognition of m6A by YTH family proteins). For YTHDF1-3 and YTHDC1 in the FP assays, the protein concentration was 300 nM, and the substrate concentration was 7 nM. The buffer conditions for other proteins in the FP assays were as follows: YTHDF1 (20 mM HEPES pH 7.5, 380 mM NaCl), YTHDF2 (20 mM HEPES pH 7.5, 200 mM NaCl), YTHDF3 (20 mM HEPES pH 7.5, 250 mM NaCl), and YTHDC1 (20 mM HEPES pH 7.5, 180 mM NaCl).
In the FP results corresponding to Fig. 3b and Supplementary Fig. S9a-h, the concentration of FAM-m6A-mRNA or FAM-mRNA was constant (7 nM), and different protein concentrations were used for the FP assays.
Differential scanning fluorimetry (DSF) assay
DSF experiments were performed on the RT-PCR detection system. The fluorescent dye selected was SYPRO orange (Sigma, 5000×, S5692). DSF experiments were performed in the buffer containing 20 mM HEPES pH7.5, 30 mM NaCl. The excitation and emission wavelengths are 492 nm and 610 nm, respectively. The reaction mixture, which consisted of 2 μM YTHDC2 proteins (wild-type and mutant), 5× SYPRO orange, and the DC2-C1, was heated from 25 to 95 °C. The heating rate was 1 °C per 15 seconds. The Boltzmann equation was employed to calculate the inflection point of the transition curve (Tm). Data analysis was carried out using GraphPad Prism software.
Isothermal titration calorimetry (ITC) assay
ITC experiments were performed on a MicroCal PEAQ-ITC instrument. The buffer conditions were 20 mM Tris-HCl (pH 7.5) and 150 mM NaCl. The titration was performed by injecting the YTHDC2 protein (580 μM) into a reaction cell containing the DC2-C1 (5-20 μM). The MicroCal Origin software was utilized to analyze the calculated equilibrium dissociation constant (KD).
Induced fit docking
The binding mode of DC2-C1 with YTHDC2 (PDB ID: 6K6U) was predicted through induced fit docking, performed using Schrödinger software. The molecular structure of DC2-C1 was prepared using the LigPrep module with its default settings. Specifically, the OPLS4 force field46 was used, possible ionization states were generated for a target pH range of 7.0 ± 2.0, the molecule underwent desalting, and all potential tautomeric forms were considered. For protein preparation, water molecules were initially removed. Subsequently, the protein was processed using the Protein Preparation Workflow with its default parameters, again utilizing the OPLS4 force field. The simulation pH was 7.4, and the maximum number of states to be automatically processed was 1.
In the induced fit docking procedure, all parameters were maintained at their default values. The energy window designated for sampling ring conformations was 2.5 kcal/mol. The van der Waals scaling factors were 0.50 for both the receptor and the ligand.
Native UVPD-MS analysis
The protein samples were buffer-exchanged into 200 mM NH4Ac (pH 7.0) using Bio-spin columns and were then diluted to 10 µM. Subsequently, the protein samples were incubated with 500 µM compounds at room temperature for 120 min. All the MS data were collected using a Thermo Fusion Lumos mass spectrometer equipped with UVPD as reported previously60,61,75. The incubated samples were directly infused into MS by nano spray. The full mass spectra and fragment mass spectra were both collected by the orbitrap analyzer with a resolution of 500000. The temperature of the transfer capillary was set to 200°C. The protein ions with the charge state of 8+ were isolated with an isolation width of 4 m/z (± 1.5 m/z) and subjected to single pulse (1.2 mJ) 193 nm laser shoot. The fragment mass spectra were obtained by averaging 500 transients across a scan range of 200-4000 m/z.
Native UVPD-MS data analysis
Raw MS data were converted to mzML format using MSConvert (ProteoWizard). Deconvolution was performed with TopFD (TopPIC Suite v1.6.2) to extract monoisotopic masses of fragment ions. The deconvoluted spectra were calibrated and searched against an in silico-generated library of expected fragment ions (including a, a + 1, a + 2, b, c, x, x + 1, y, y-1, y-2, z, z + 1) with a 2 ppm mass tolerance. Fragment ion identifications were validated using the Pearson correlation coefficient (PCC ≥ 0.7) between experimental and theoretical isotope distributions. For replicate consistency, ions detected in < 16 of 20 scans were discarded. Intensity normalization was applied using the median method. The UVPD fragment yield (FY) for each residue was derived from the summed intensities of a, a + 1, b, c, x, x + 1, y, y-1, z, z + 1 ions. Differential fragmentation between YTHDC2 and complex was assessed via Welch’s t-test (p < 0.01). All analyses were implemented in custom R scripts.
15N-Heteronuclear single-quantum coherence spectroscopy
15N-heteronuclear single-quantum coherence (HSQC) NMR spectra were recorded at 298 K on a 700 MHz Avance III Bruker spectrometer equipped with a 5 mm QCI-F CryoProbe (Ettlingen, Germany). 1H-15N HSQC spectra were acquired using the Bruker pulse program hsqcfpf3gpphwg with D1 = 1 s, DS = 16, NS = 32, TD = 2048 and 256, and SW = 16.2300 and 34.0000 ppm for the direct 1H and indirect 15N dimensions, respectively. The total acquisition time for each 1H-15N HSQC experiment was 2 h 34 min. The 1H-15N HSQC spectra were processed using Bruker TopSpin 4.5.0 software (Bruker biospin, Rheinstetten, Germany), including automatic zero filling, Fourier transformation, manual phase-correction, and chemical shift calibration. 15N-labeled YTHDC2 protein for NMR studies was prepared and purified using the same protocol as for unlabeled protein with the exception that the bacteria were grown on M9 minimal media supported with 1 g/L of 15NH4Cl. Protein samples were prepared in a 20 mM Tris-HCl, 100 mM NaCl, and 1 mM DTT, 1% DMSO-d6, and 0.02% NaN3 solution at pH 7 in 5% D2O. The final concentration of the protein was 0.4 mM. Then, HSQC NMR spectra of the protein at its free state were recorded. After that, the tested compound dissolved in DMSO-d6 final (concentration: 0.8 mM) was added into the above solution, which was incubated at 4 °C for 1 h and then centrifuged. HSQC NMR spectra of the protein-compound complex were recorded. The HSQC spectrum of free YTHDC2, which is publicly available from the Biological Magnetic Resonance Data Bank (http://www.bmrb.wisc.edu/; access number: 36310), was used as the reference for the assignment of chemical shifts. The ∆δ-value computed as the following equation was used to quantify the chemical shift of residues on the 2D HSQC spectra.
Cell culture
hTERT-HPNE, PANC-1, DU145, MiaPACA-2, BxPC-3, HepG2, and HEK293T were obtained from the American Type Culture Collection (ATCC); LX-2 was acquired from Sigma-Aldrich; HGC27 was sourced from the European Collection of Authenticated Cell Cultures (ECACC); MKN−45 was obtained from the German Collection of Microorganisms and Cell Cultures (DSMZ); and MH7A was originally established by the RIKEN BioResource Research Center. Unless otherwise specified, LX-2, PANC-1, MH7A, MiaPACA-2, HepG2, and HEK293T were cultured in high-glucose DMEM (Gibco) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. MKN-45, hTERT-HPNE, HGC27, and BxPC-3 were maintained in RPMI-1640 (Gibco) with 10% FBS and 1% penicillin/streptomycin, while DU145 was cultured in MEM (Gibco) containing the same supplements. All cells were maintained in a humidified incubator at 37 °C with 5% CO₂. All cells were negative for mycoplasma, and these cell lines are not among those commonly misidentified by International Cell Line Authentication Committee (ICLAC).
Cell viability assays
Cell viability was assessed using the MTT assay to evaluate cell proliferation. Cells were seeded in 96-well plates at a density of 2×103 to 5×103 cells per well. After overnight incubation at 37 °C to allow cell attachment, test compounds were added, and cells were treated for 72 h. Subsequently, 20 μL of MTT solution (5 mg/mL in saline, Sigma) was added to each well, followed by incubation for 2-4 h at 37 °C. To solubilize the formazan crystals, 50 μL of SDS solution (10% w/v in water with 0.1% HCl) was added per well, and plates were incubated overnight at 37 °C. Finally, absorbance was measured at 570 nm using a multiscan spectrum reader (BMG Labtech), and cell viability was calculated as a percentage relative to the control group.
Cell migration and invasion assays
For invasion assays, matrigel (LABLEAD, MG2237) was thawed at 4 °C for 12 h, diluted 1:8 with serum-free DMEM, and added to transwell chambers (8-μm pore) (LABSELECT, 14341). MH7A, PANC-1, and MiaPACA-2 cells were serum-starved overnight. Cells (MH7A: 2.5×104 cells/mL, PANC-1: 5×104 cells/mL, MiaPACA-2: 2.5×104 cells/mL) in 200 μL serum-free medium were seeded into uncoated (migration) or matrigel-coated (invasion) transwell inserts, placed in lower chambers with 600 μL DMEM with 10% FBS, and treated with DMSO, DC2-C1 (25 μM), or 16o (25 μM). After 48 h at 37 °C, cells penetrating the filter of the inserts were fixed, stained with crystal violet (Beyotime, C0121), imaged via inverted microscope (Evident, Olympus IX73), and quantified.
Cell colony formation assay
MH7A cells were seeded at a density of 1000 cells per well in 12-well plates. PANC-1 and MiaPACA-2 cells were seeded at a density of 400 cells per well in 12-well plates. Following overnight incubation at 37 °C to allow cell attachment, cells were treated with DMSO, DC2-C1 (25 μM), or 16o (25 μM). Cultures were maintained for 10-14 days with medium changed every 2 days to ensure nutrient supply. Once visible colonies formed, cells were fixed and stained, and the images were captured.
Virus infection
The sgRNA sequences for transfection were: sgDC2-1 (tgagtaccattctgtaagtg), sgDC2-2 (atgtgaattgccttgaacca), sgDC2-3 (tatgacttactggatgatgg). The recombinant plasmids (containing the above sgRNA sequences in lentiCRISPR v2-CMV-ZsGreen-Puro vector) were purchased from Tsingke Biotechnology Co., Ltd. HEK293T cells were co-transfected with psPAX2, pMD2.G and the lentiviral plasmids using Hieff Trans® Liposomal 2000 Transfection Reagent (YEASEN, 40802ES03) in Opti-MEM for virus packaging. Viral supernatants were collected and filtered through 0.45 μm membranes to remove cell debris. Target cells were seeded in 6-well plates to achieve 30−50% confluency, then infected with 200 μL of viral supernatant per well in the presence of 6-8 μg/mL polybrene (YEASEN, 40804ES76) to enhance infection efficiency. After 24 h, the medium was replaced, and cells were cultured for an additional 1-2 days before selection with 2 μg/mL puromycin (YEASEN, 60209ES10) for 3-5 days to establish stable knockout cell lines.
Transient transfection
The coding sequence for the YTH domain of YTHDC2 (amino acid residues 1287-1423) was inserted into the PCDH-GFP-Puro-3xFlag vector to generate overexpression plasmids. HEK293T cells were transfected for 24 h with 4 µg overexpression plasmid and Hieff Trans® Liposomal 2000 Transfection Reagent. The cells then were used for CETSA and DARTS experiments as described below.
Cellular thermal shift assay (CETSA)
HEK293T cells transiently transfected with the YTH domain of YTHDC2 for 24 h were treated with DC2-C1 (30 μM), 16o (30 μM), or DMSO for an additional 24 h at 37 °C. Cells were harvested by centrifugation, washed twice with ice-cold PBS, and resuspended in PBS supplemented with 1× protease inhibitor cocktail (Sigma-Aldrich, P8340) and 1 mM PMSF (TargetMol, T0789). Cell suspensions were aliquoted into PCR tubes and subjected to thermal challenge at varied temperatures for 3 min. Samples were then lysed by three freeze-thaw cycles (liquid nitrogen/37 °C) and centrifuged at 4 °C. Supernatants containing soluble proteins were mixed with SDS-PAGE loading buffer, heated at 95 °C for 8 min, and analyzed by Western blot.
Drug affinity responsive target stability (DARTS)
HEK293T cells transiently transfected with the YTH domain of YTHDC2 for 24 h were treated with DC2-C1(30 μM), 16o(30 μM), or DMSO for an additional 24 h at 37 °C. The cells were collected by centrifugation, washed twice with PBS, and cells were lysed in 200 μL NP-40 lysis buffer (Beyotime, P0013F) containing 1× protease inhibitor cocktail, and 1 mM PMSF on ice for 20 min. And each soluble protein was collected and normalized to 3 mg/mL using a BCA assay (Beyotime, P0012). The protein was divided into 50 μL aliquots per tube, and pronase (Roche, PRON-RO) was added at final concentrations of 0-10% (w/w) relative to total protein. Samples were incubated at 37 °C for 30 min, and reactions were terminated by adding 20× protease inhibitor cocktail, followed by ice incubation for 10 min. Finally, the proteins were mixed with SDS-PAGE loading buffer and heated at 95 °C for 8 min. The results were analyzed by western blotting assay as described in Western blot analysis.
Western blot (WB) assay
MH7A, HGC27, PANC-1, and MiaPACA-2 cells were treated with compounds at different concentrations in complete medium for 48 h. Whole cell lysates were extracted using RIPA lysis buffer (Beyotime, P0013C) supplemented with 1× protease inhibitor cocktail and 1 mM PMSF. Protein concentrations were quantified by BCA assay (Beyotime, P0012), and equal amounts of protein were separated by SDS-PAGE on 10–12% polyacrylamide Tris-glycine gels before transfer to PVDF membranes (Millipore, IPVH00010). Membranes were then blocked with TBS-T containing 5% nonfat dry milk for 1 h. Then, the membranes were incubated with primary antibodies for 12 h at 4 °C. After being washed three times with TBS-T, the membranes were incubated with the corresponding secondary antibodies for 1 h at 37 °C, and washed three times with TBS-T subsequently. Finally, the immune-bands were visualized with the SuperLumia ECL plus HRP Substrate Kit (Abbkine, K22030) and imaged with a chemiluminescence detector (FUSION-FX6.EDGE V.070). The knockout cells were directly lysed with RIPA buffer, and the above procedure was repeated. Antibodies used: YTHDC2 antibody (ZenBio, R27443), AMIGO2 antibody (ImmunoWay, YN2372), ID2 antibody (HUABIO, M1301-2), H3K4me3 antibody (Abcam, ab8580), FZD7 antibody (Proteintech, 16974-1-AP), DYKDDDDK Tag (Flag-tag) antibody (Cell Signaling Technology, 14793), GAPDH antibody (Proteintech, 60004-1-Ig), HRP-conjugated Affinipure Goat Anti-Rabbit IgG(H + L) (Proteintech, SA00001-2), HRP-conjugated Affinipure Goat Anti-Mouse IgG(H + L) (Proteintech, SA00001-1).
Real-time quantitative polymerase chain reaction (RT-qPCR)
The cells at logarithmic growth stage were inoculated into 6-well plates at a density of 1× 105 cells per well. Cells were treated with test compounds for 24 h. Total RNA was extracted from cells according to the manufacturer’s instruction of the Cell Total RNA Isolation Kit (Foregene, RE-03113). Then, total RNA was converted into cDNA using the Hifair® III 1st Strand cDNA Synthesis SuperMix for qPCR (gDNA digester plus) (YEASEN, 11141ES60). Subsequently, all qRT-PCR experiments were conducted strictly according to the instructions of the Hieff® qPCR SYBR Green Master Mix (No Rox) (YEASEN, 11201ES08). Gene expression was calculated using the comparative ΔΔCt method with GAPDH for normalization. All primers of the selected genes were listed as follows: for GAPDH (h), forward, 5′-GCCCAATACGACCAAATCC-3′; reverse, 5′-AGCCACATCGCTCAGACAC-3′; for AMIGO2 (h), forward, 5′-GTGTCTGGACCCCATCCTTA-3′; reverse, 5′- CCCATCACTTCGTGCTTTTT-3′;for FZD7 (h), forward, 5′-TGGTGCTCTACTTCTTCGG-3′; reverse, 5′-AGGATAGTGATGGTCTTGACG-3′; for ID2 (h), forward, 5′-CCGTGAGGTCCGTTAGGAAA-3′; reverse, 5′-TGAGCTTGGAGTAGCAGTCG-3′; for FASN (h), forward, 5′-TGCCCTGAGCTGGACTACTT-3′; reverse, 5′-AAAGCTGCTCAGGACCATGT-3′; for SREBP-1C(h), forward, 5′-GACAGCCCAGTCTTTGAGGA-3′; reverse, 5′-CAGGACAGGCAGAGGAAGAC-3′; for ACC1(h), forward, 5′-TCACACCTGAAGACCTTAAAGCC-3′; reverse, 5′-AGCCCACACTGCTTGTACTG-3′.
RNA immunoprecipitation (RIP-qPCR) assay
RIP-qPCR was performed using the PureBinding® RNA Immunoprecipitation Kit (GENESEED, P0102) according to the manufacturer’s protocol. MH7A cells and PANC-1 cells (1×107 cells/sample), which were treated for 12 h with DMSO, DC2-C1 (30 µM) or 16o (30 µM), were incubated with YTHDC2 antibody or IgG antibody (Beyotime, A7016) at 4 °C for 2 h according to the instructions, and the corresponding RNA was extracted and then qRT-PCR was performed. The formula for calculating the RIP-qPCR result is as follows:
RNA stability assay
MH7A and HepG2 cells were seeded in 12-well plates and pretreated with DMSO, DC2-C1 (30 μM), or 16o (30 μM) for 24 h. Transcription was inhibited by adding actinomycin D (MedChemExpress, HY-17559) at a final concentration of 5 μg/mL. Cells were harvested at 0, 2, 4, 6, and 8 h post-treatment, and total RNA was extracted and used for qRT-PCR. Relative mRNA levels were normalized to the starting point of treatment.
Determination of solubility
The solubility of compounds was determined using the shake-flask method with LC-MS/MS quantification. Briefly, supersaturated aqueous solutions of the compounds were prepared, and were agitated at 37 °C in an orbital shaker (200 rpm) for 48 h. Following incubation, samples were prepared for LC-MS/MS analysis. Calibration curves were generated using serial dilutions of standard compounds to quantify the soluble concentration, with each measurement performed in triplicate to ensure reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The pretraining and fine-tuning datasets are available on Zenodo(https://doi.org/10.5281/zenodo.15921759)76. The crystal structures of YTHDC2 used in this study are available in the Protein Data Bank (PDB) under accession code 6K6U. The ZINC database is available at https://zinc20.docking.org. The CossDocked2020 database is available at https://bits.csb.pitt.edu/files/crossdock2020. Source data are provided with this paper as a Source Data file. Supplementary Data files are also available under Supplementary Data 1 - Supplementary Data 9.
Code availability
The codes are available on Github (https://github.com/atzhenyuyang/EPMolGen). A citable version with a DOI is available on Zenodo (https://doi.org/10.5281/zenodo.17176305)77 with MIT License.
References
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Zhu, Z., Huo, F. & Pei, D. Function and evolution of RNA N6-methyladenosine modification. Int. J. Biol. Sci. 16, 1929–1940 (2020).
Wang, J. et al. M6A mRNA methylation regulates the development of gestational diabetes mellitus in Han Chinese women. Genomics 113, 1048–1056 (2021).
Jiang, X. et al. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target. Ther. 6, 74 (2021).
Deng, L. et al. m6A modifcation: recent advances, anticancer targeted drug discovery and beyond. Mol. Cancer 21, 52 (2022).
Sikorski, V., Selberg, S., Lalowski, M., Karelson, M. & Kankuri, E. The structure and function of YTHDF epitranscriptomic m6A readers. Trends Pharmacol. Sci. 44, 335–353 (2023).
Fang, Z. et al. Role of m6A writers, erasers and readers in cancer. Exp. Hematol. Oncol. 11, 45 (2022).
Liu, S. et al. The roles and mechanisms of YTH domain-containing proteins in cancer development and progression. Am. J. Cancer Res. 10, 1068–1084 (2020).
Luo, Y. et al. A novel MYCN-YTHDF1 cascade contributes to retinoblastoma tumor growth by eliciting m6A -dependent activation of multiple oncogenes. Sci. China Life Sci. 66, 2138–2151 (2023).
Liao, S., Sun, H. & Xu, C. YTH Domain: A Family of N6-methyladenosine (m6A) Readers. Genomics Proteom. Bioinforma. 16, 99–107 (2018).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Kretschmer, J. et al. The m(6)A reader protein YTHDC2 interacts with the small ribosomal subunit and the 5′-3′ exoribonuclease XRN1. RNA 24, 1339–1350 (2018).
Hsu, P. J. et al. Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res 27, 1115–1127 (2017).
Ma, C., Liao, S. & Zhu, Z. Crystal structure of human YTHDC2 YTH domain. Biochem. Biophys. Res. Commun. 518, 678–684 (2019).
Wu, X. et al. The biological function of the N6-Methyladenosine reader YTHDC2 and its role in diseases. J. Transl. Med. 22, 490 (2024).
Geng, Q. et al. Diagnostic gene signatures and aberrant pathway activation based on m6A methylation regulators in rheumatoid arthritis. Front. Immunol. 13, 1041284 (2022).
Xiao, J., Cai, X., Wang, R., Zhou, W. & Ye, Z. ALKBH5-YTHDF2 m6A modification axis inhibits rheumatoid arthritis progression by suppressing NLRP3. Biochem. Biophys. Res. Commun. 668, 70–76 (2023).
Miao, T. et al. METTL3 knockdown suppresses RA-FLS activation through m6A-YTHDC2-mediated regulation of AMIGO2. Biochim. Biophys. Acta Mol. Basis. Dis. 1870, 167112 (2024).
Li, Y. et al. Autophagy activation is required for N6-methyladenosine modifcation to regulate ferroptosis in hepatocellular carcinoma. Redox Biol. 69, 102971 (2024).
Zhou, G. & Wang, S. YTHDC2 retards cell proliferation and triggers apoptosis in papillary thyroid cancer by regulating CYLD-mediated inactivation of Akt signaling. Appl. Biochem. Biotechnol. 196, 588–603 (2024).
Zhou, X. et al. The m6A methyltransferase METTL3 drives thyroid cancer progression and lymph node metastasis by targeting LINC00894. Cancer Cell Int 24, 47 (2024).
Li, Q. et al. METTL16 inhibits papillary thyroid cancer tumorigenicity through m(6)A/YTHDC2/SCD1-regulated lipid metabolism. Cell Mol. Life Sci. 81, 81 (2024).
Yuan, W. et al. The N6-methyladenosine reader protein YTHDC2 promotes gastric cancer progression via enhancing YAP mRNA translation. Transl. Oncol. 16, 101308 (2022).
Cun, Y., Guo, W., Ma, B., Okuno, Y. & Wang, J. Decoding the specificity of m6A RNA methylation and its implication in cancer therapy. Mol. Ther. 32, 2461–2469 (2024).
He, J. J. et al. m(6)A Reader YTHDC2 promotes radiotherapy resistance of nasopharyngeal carcinoma via activating IGF1R/AKT/S6 signaling axis. Front. Oncol. 10, 1166 (2020).
Yang, Z., Verghese, M., Yang, S., Shah, P. & He, Y. Y. The m(6) A reader YTHDC2 regulates UVB-induced DNA damage repair and histone modifcation. Photochem. Photobiol. 100, 1031–1040 (2024).
Zhou, B. et al. N6 -methyladenosine reader protein YT521-B homology domain-containing 2 suppresses liver steatosis by regulation of mRNA stability of lipogenic genes. Hepatol. (Baltim., Md.) 73, 91–103 (2021).
Wang, C. H. & Zhou, H. Discovery of a new inhibitor for YTH domain-containing m6A RNA readers. RSC Chem. Biol. 5, 914–923 (2024).
Blay, V., Tolani, B., Ho, S. P. & Arkin, M. R. High-throughput screening: today’s biochemical and cell-based approaches. Drug Discov. Today 25, 1807–1821 (2020).
Wali, A. et al. Generative adversarial networks for speech processing: a review. Computer Speech Lang. 72, 101308 (2022). ISSN 0885-2308.
Nortier, B., Sadeghi, M. & Serizel, R. Unsupervised Speech Enhancement with Diffusion-Based Generative Models. IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). 12481–12485 (2024).
Ding, M. et al. Cogview: mastering text-to-image generation via transformers. Adv. neural Inf. Process. Syst. 34, 19822–19835 (2021).
Chen, X. et al. Activating More Pixels In Image Super-resolution Transformer. In Proceedings Of The Ieee/cvf Conference On Computer Vision and Pattern Recognition. 22367–22377 (2023).
Houlsby, N. et al. Parameter-efficient Transfer Learning For NLP. In Proceedings of International Conference on Machine Learning. 2790–2799 (2019).
Joshi, V., Peters, M. & Hopkins, M. 2018. In Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics (1, Long Papers), pages 1190–1199, Melbourne, Australia. Association for Computational Linguistics.
Tong, X. et al. Generative models for De Novo drug design. J. Med Chem. 64, 14011–14027 (2021).
Bian, Y. & Xie, X. Q. Generative chemistry: drug discovery with deep learning generative models. J. Mol. Model 27, 71 (2021).
Cheng, Y., Gong, Y., Liu, Y., Song, B. & Zou Q. Molecular design in drug discovery: a comprehensive review of deep generative models. Brief. Bioinform. 22, bbab344 (2021).
Gangwal, A. & Lavecchia, A. Unleashing the power of generative AI in drug discovery. Drug Discov. Today 29, 103992 (2024).
Ragoza, M., Masuda, T. & Koes, D. R. Generating 3D molecules conditional on receptor binding sites with deep generative models. Chem. Sci. 13, 2701–2713 (2022).
Peng, X. et al. Pocket2mol: efficient molecular sampling based on 3D protein pockets. In Proceedings of the International Conference on Machine Learning 162, 17644–17655 (2022).
Liu, M. et al. Generating 3D molecules for target protein binding. In: Proceedings of the 39th International Conference on Machine Learning 162, 13912–13924 (PMLR, 2022).
Guan, J. et al. 3 d equivariant diffusion for target-aware molecule generation and affinity prediction. The Eleventh International Conference on Learning Representations (2023).
Jiang, Y. et al. PocketFlow is a data-and-knowledge-driven structure-based molecular generative model. Nat. Mach. Intell. 6, 326–337 (2024).
O’Boyle, N. M. et al. Open Babel: an open chemical toolbox. J. Cheminform. 3, 33 (2011).
Lu, C. et al. OPLS4: improving force field accuracy on challenging regimes of chemical space. J. Chem. Theory Comput. 17, 4291–4300 (2021).
Bishop, C. M. et al. Mixture density networks. Technical Report. https://publications.aston.ac.uk/id/eprint/373/ (Aston University, 1994).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Bickerton, G. R. et al. Quantifying the chemical beauty of drugs. Nat. Chem. 4, 90–98 (2012).
Ertl, P. & Schufenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J. Cheminform. 1, 8 (2009).
Francoeur, P. G. et al. Three-dimensional convolutional neural networks and a cross-docked data set for structure-based drug design. J. Chem. Inf. Model. 60, 4200–4215 (2020).
Eldridge, M. D. et al. Empirical scoring functions: I. the development of a fast empirical scoring function to estimate the binding afinity of ligands in receptor complexes. J. Comput. -Aided Mol. Des. 11, 425–445 (1997).
Hartshorn, M. J. et al. Diverse, high-quality test set for the validation of protein-ligand docking performance. J. Med. Chem. 50, 726–741 (2007).
Worpenberg, L. et al. Ythdf is a N6-methyladenosine reader that modulates Fmr1 target mRNA selection and restricts axonal growth in Drosophila. Embo. J. 40, e104975 (2021).
Zhu, T. et al. Crystal structure of the YTH domain of YTHDF2 reveals mechanism for recognition of N6-methyladenosine. Cell Res 24, 1493–1496 (2014).
Repasky, M. P., Shelley, M. & Friesner, R. A. Flexible ligand docking with glide. Curr. Protoc. Bioinforma. 18, 8.12.1–8.12.36 (2007).
Micaelli, M. et al. Small-molecule ebselen binds to YTHDF proteins interfering with the recognition of N6-methyladenosine-modified RNAs. ACS Pharmacol. Transl. Sci. 5, 872–891 (2022).
Chen, X. et al. Motif-dependent immune co-receptor interactome profiling by photoaffinity chemical proteomics. Cell chem. Biol. 29, 1024–1036.e5 (2022).
Cammarata, M. B., Thyer, R., Rosenberg, J., Ellington, A. & Brodbelt, J. S. Structural characterization of dihydrofolate reductase complexes by top-down ultraviolet photodissociation mass spectrometry. J. Am. Chem. Soc. 137, 9128–9135 (2015).
Luo, P. et al. Time-resolved ultraviolet photodissociation mass spectrometry probes the mutation-induced alterations in protein stability and unfolding dynamics. J. Am. Chem. Soc. 146, 8832–8838 (2024).
Xie, Y. et al. Ultraviolet photodissociation mass spectrometry captures the acyl chain length-dependent conformation dynamics of acyl carrier protein. J. Am. Chem. Soc. 147, 16760–16765 (2025).
Li, R. et al. Super-enhancer RNA m6A promotes local chromatin accessibility and oncogene transcription in pancreatic ductal adenocarcinoma. Nat. Genet. 55, 2224–2234 (2023).
Pi, J. et al. YTHDF1 promotes gastric carcinogenesis by controlling translation of FZD7. Cancer Res. 81, 2651–2665 (2021).
Chen, J. et al. METTL3 promotes pancreatic cancer proliferation and stemness by increasing stability of ID2 mRNA in a m6A-dependent manner. Cancer Lett. 565, 216222 (2023).
Yankova, E. et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature 593, 597–601 (2021).
Su, R. et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38, 79–96 (2020).
Huang, Y. et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell 35, 677–691 (2019).
Wu, H. et al. Discovery of covalent and cell-active ALKBH5 inhibitors with potent antileukemia effects in vivo. Angew. Chem. Int. Ed. Engl. 64, e202424928 (2025).
Fang, Z. et al. Discovery of a potent, selective and cell active inhibitor of m6A demethylase ALKBH5. Eur. J. Med. Chem. 238, 114446 (2022).
Wang, L. et al. YTHDF2 inhibition potentiates radiotherapy antitumor efficacy. Cancer Cell 41, 1294–1308 (2023).
Zou, Z. et al. FMRP phosphorylation modulates neuronal translation through YTHDF1. Mol. Cell. 83, 4304–4317 (2023).
Zálešák, F. et al. Structure-based design of a potent and selective YTHDC1 ligand. J. Med. Chem. 67, 9516–9535 (2024).
Irwin, J. J. et al. ZINC20—a free ultralarge-scale chemical database for ligand discovery. J. Chem. Inf. Model. 60, 6065–6073 (2020).
Deng, C. et al. Vector Neurons: A General Framework For So (3)-equivariant Networks. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 12180–12189 (2021).
Yang, S. et al. In-cell mass spectrometry and ultraviolet photodissociation navigates the intracellular protein heterogeneity. J. Am. Chem. Soc. 147, 4714–4719 (2025).
Yang, Z. et al. Deep learning-assisted discovery of a potent and cell-active inhibitor of RNA N6-methyladenosine recognition protein YTHDC2. Pretraining and fine-tuning datasets for EPMolGen, https://doi.org/10.5281/zenodo.15921759 (2025).
Yang, Z. et al. Deep learning-assisted discovery of a potent and cell-active inhibitor of RNA N6-methyladenosine recognition protein YTHDC2. EPMolGen model, https://doi.org/10.5281/zenodo.17176305 (2025).
Acknowledgements
This work was supported by National Key R&D Program of China (2023YFF1204905, Y.J.; 2022YFA1304600, Z.L.), National Natural Science Foundation of China (grant no: 82404514, Y.J.; 82404409, W.S.), the Natural Science Foundation of Sichuan Province (grant no. 24NSFSC6411, Y.J.), the China Postdoctoral Science Foundation (Certificate Number: 2024M752190, Y.J.). The authors thank Qianlun Pu and Fei Fu (Core Facility of West China Hospital Sichuan University) for supports. The authors thank Prof. Yun Tang (East China University of Science and Technology) for his support and help in using Schrödinger. The authors thank the staff members of the Biological Mass Spectrometry System (https://cstr.cn/31127.02.DCLS.ESBMS) at the Dalian Coherent Light Source (https://cstr.cn/31127.02.DCLS), for providing technical support and assistance in data collection and analysis. The authors thank Prof. Zhongliang Zhu (University of Science and Technology of China) for providing the YTHDC2 plasmid.
Author information
Authors and Affiliations
Contributions
Z.Y. developed the algorithm, and implemented EPMolGen. W.S., Y.L., and M.Y. conducted the bioactivity assays. Q.H. and Z.Y. synthesized the chemical compounds. Y.Y. completed the NMR assays. H.Z., Z.L. and F.W. completed the Native UVPD-MS analysis. Z.Y., W.S., Q.H., Y.L., M.Y., Y.Y., X.Z., H.Z., Z.L., F.W. and Y.J. analyzed the data. Z.Y., W.S. and Q.H. wrote the paper. Y.J., Y.Z. and R.C. revised the paper. All authors read and agreed on the content of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Denise Sighel, Alessandro Quattrone and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Z., Sun, W., Huang, Q. et al. Deep learning-assisted discovery of a potent and cell-active inhibitor of RNA N6-methyladenosine recognition protein YTHDC2. Nat Commun 17, 46 (2026). https://doi.org/10.1038/s41467-025-65542-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65542-0