Abstract
Voltage-gated sodium channels initiate action potentials and control electrical signaling throughout the animal kingdom. Fast inactivation is an essential auto-inhibitory mechanism and requisite component of sodium channel physiology. Recent structural and electrophysiological results are inconsistent with the canonical ball and chain model of fast inactivation thus necessitating an updated theoretical framework. Here, we use encoded fluorescence spectroscopy and high-resolution electrophysiology to capture key steps in the fast inactivation mechanism, from voltage-sensor activation to pore occlusion, an ultra-fast process which occurs in less than 2 milliseconds. Upon depolarization, activation of the domain IV voltage sensor initiates cytoplasmic DIII_DIV linker movement and quickly repositions the IFM motif into a hydrophobic pocket adjacent to the pore. This triggers a structural rearrangement of the pocket. The phenylalanine of the IFM motif contacts the pore-forming helices via a hydrophobic interaction with S6 of DIV and an aromatic/hydrophobic interaction with S6 of DIII. These two interactions occur only after both S6 segments rotate, thus exposing the hydrophobic gate into the pore producing the fast inactivation. Based on the current results, we propose an alternative lock and key model to explain the molecular mechanism of fast inactivation.
Similar content being viewed by others
Introduction
The action potential is the elementary unit of bio-excitability. At the molecular level, voltage-gated sodium (Nav) channels are responsible for the rapid upstroke of the action potential. These channels are gated to an open, sodium-conducting conformation by positive changes in transmembrane potential1,2,3. During the rising phase of the action potential, Nav channels operate in a classical positive-feedback manner: opened by membrane depolarization, the Nav channels allow further Na+ entry with consequent membrane depolarization, thus resulting in a cascading, all-or-nothing action potential. In this positive feedback loop, fast inactivation is an essential built-in auto-inhibitory mechanism to terminate the cascade and serves as a molecular kill switch. Milliseconds after the channels are open during activation, fast inactivation drives the channels into the nonconductive, fast inactivated state, facilitating the repolarization by voltage-gated potassium channels. Acquired or inherited abnormalities in fast inactivation can result in severe pathological states4. Additionally, due to the prevalence and diversity of Nav channels, these pathological states can manifest in numerous tissue types with a wide range of symptoms5,6,7,8,9. Hence, a molecular understanding of the fast inactivation mechanism is of fundamental importance.
Originally proposed almost 50 years ago, the ball and chain model predicted that an intracellular located inactivation particle would bind to the open channel at the pore and physically block the ionic conduction10. Later, the identification of the amino acid triad, Isoleucine-Phenylalanine-Methionine (the IFM motif) in the cytoplasmic domain III and domain IV (DIII_DIV) linker as the inactivation particle materialized the abstract concept and provided a molecular framework for fast inactivation11. Numerous studies have since provided strong evidence for this model, and consequently, the ball and chain model has been largely accepted as the textbook explanation for fast inactivation in Nav channels12,13,14,15,16.
However, upon the advent of the cryogenic electron microscopy-mediated resolution revolution in structural biology, discrepancies emerged between the old models and new experimental results17. As of today, essentially all of the available eukaryotic Nav channel structures, likely in the inactivated state due to the lack of external electric field, consistently demonstrate that the IFM motif, the inactivation particle, does not block the permeation pathway as expected18,19,20,21. Instead, the motif docks in an adjacent hydrophobic pocket between the pore and S4-S5 linkers from DIII and DIV, a stark contradiction to the canonical pore-blocking ball and chain model. Our recent work began to address this inconsistency and identified the inactivation gate, a pore-lining double-layered hydrophobic ring at the intracellular mouth of the pore that blocks Na+ ion permeation in the inactivated state, instead of the IFM motif22. However, this is only the final step, and the series of movements involved in linking voltage sensor movement to ultimate channel inactivation remains to be elucidated. The inactivating particle (IFM motif) is necessary for inactivation, but the mechanistic linkage between voltage-sensor activation, the docked IFM motif, and the newly described gate remains poorly resolved. Thus, the lack of molecular description and theoretical framework of the fast inactivation process once again represents a major conceptual gap in Nav channel biophysics and physiology.
Here in this work, we present the sequential conformational changes during the fast inactivation process in mammalian Nav channels. The dynamic nature of the molecular rearrangements between the IFM binding and the gate closing requires techniques that can detect these changes while observing the actual function of the channel. To this end, we utilize site-directed fluorescence in voltage-clamped conditions. With our improved incorporation protocol and optical recording setup, we introduce an environmentally sensitive fluorescent unnatural amino acid, ANAP, into various regions of the Nav channel and monitor the local molecular motions during fast inactivation by rapid fluorescence measurements. ANAP fluorescence recordings reported two types of movements at the IFM location during fast inactivation. A fast movement, triggered by the activation of voltage sensing domain (VSD) in domain IV (DIV), reflects the movement of the DIII_DIV linker region and positions the IFM motif into the appropriate location, conceptually like putting a key into a keyhole, that was exposed by the movement of VSD in DIV. Coupled by hydrogen bonds between DIII_DIV linker and DIV S4_S5 linker, this fast component follows faithfully the movements of DIV VSD, via simultaneously measured gating currents, and proceeds prior to the closure of the fast inactivation gate. The other slower conformational change at the IFM motif triggers the fast inactivation. The aromatic side chain of the phenylalanine residue in the IFM motif forms interactions with two residues in DIII S6 and DIV S6, through T-shaped π-π and hydrophobic interactions. In tandem, these interactions propagate the conformational change from the linker to the pore and lead to the closure of the double-layered fast inactivation gate, similar to the movement of the bolt of the lock. Contrary to our expectations, modification to the interaction between the IFM motif and the hydrophobic pocket does not impede the binding of the motif itself, rather, it appears to influence the effectiveness of the IFM motif at triggering fast inactivation, like creating mismatches between the key and the lock. With these current observations, we propose a lock and key model to describe the sequential conformational changes during fast inactivation in mammalian sodium channels.
Results
IFM motif undergoes two distinct movements during fast inactivation
The IFM motif is evolutionarily conserved in Nav channels across the animal kingdom (Fig. 1A and Supplementary Fig. 7). Its importance for fast inactivation has been demonstrated repeatedly in the literature14,23. However, recent structural results have cast uncertainty as to the functional role IFM motif plays in the mechanism of fast inactivation. To describe the conformational changes at the IFM position and to understand its role in fast inactivation, we replaced the phenylalanine in the IFM motif with a fluorescent, environmentally sensitive unnatural amino acid, ANAP (F1304ANAP, numbering based on rat Nav 1.4, all experiments were performed on rNav1.4)24,25. ANAP (3-(6-acetylnaphthalen-2-ylamino)−2-aminopropanoic acid) has a comparable size as tryptophan which allows minimum disturbance upon incorporation (Fig. 1B). More importantly, the emission spectrum of ANAP is modulated by the hydrophobicity of its immediate environment, making ANAP an ideal reporter of molecular movement25 (Fig. 1B). Robust currents were seen from F1304ANAP, and no discernable current was seen in cells where no ANAP was supplied, thus demonstrating high fidelity of encoding (Fig. 1C). F1304ANAP channels showed slowed fast inactivation kinetics with slightly increased steady-state current (Supplementary Table 2), similar to previously reported tryptophan replacement14, and had minimal impact on activation (Fig. 1D–F and Supplementary Table 1). Utilizing site-directed fluorimetry with cut-open voltage clamp technique (VCF), ionic current and fluorescence signal were resolved simultaneously26,27. Fluorescence signals increased upon depolarization, indicating an increase in the environmental hydrophobicity seen by the side chain of ANAP (Fig. 1G). This is consistent with the notion that during fast inactivation the IFM motif transitions from a hydrophilic to more hydrophobic environment. The overlap of the steady-state fluorescence (FV) curve and fast inactivation, tracked via 1-(h-infinity) curve (Fig. 1H), indicates that the movement of the IFM motif directly reports the fast inactivation state of the Nav channels. Additionally, the off-fluorescence signal closely tracks the recovery from inactivation (Supplementary Fig. 1). Interestingly, the fluorescence signals that originate from F1304ANAP channels exhibit two kinetic components, suggesting two distinct conformational changes in the vicinity of the IFM motif occurred during fast inactivation (Fig. 1G, inset). Comparison between the fast inactivation kinetics and the fluorescence signal kinetics revealed that the fast time constant of fluorescence signal was significantly faster than the fast inactivation kinetics. The slow component, on the other hand, closely followed the development of fast inactivation (Fig. 1I).
A Side view of a Nav channel structure (PDB: 7XVF). IFM motif (in pink) resides in DIII_DIV linker (yellow) and docks into a pocket away from the pore. B Size of ANAP is comparable to tryptophan. With the filter set used in the study (details in Methods), the fluorescence is brighter in hydrophobic environment. C Example ionic current traces of wild type (WT), F1304ANAP, and F1304UAG when no ANAP was supplied. Incorporation of ANAP into IFM motif slowed fast inactivation kinetics. Inset shows the voltage protocol with 5 mV voltage steps. D GV curve of WT and F1304ANAP. No significant difference was observed. N = 4 for WT, N = 5 for F1304ANAP. E h-infinity curve for WT and F1304ANAP, with the latter showing a right-shifted Vhalf (∆Vhalf = ~8.5 mV). Inset shows the voltage protocol with 10 mV voltage steps and a 5 s cycle period. N = 4 for WT, N = 5 for F1304ANAP. F Recovery from fast inactivation at −90mV, measured by comparing the peak current during control and test pulse. F1304ANAP recovered faster than WT, suggesting a less stable inactivated state. N = 4 for WT, N = 5 for F1304ANAP. Inset shows the voltage protocol. G Example family of traces of simultaneously recorded ionic current and fluorescence signal from F1304ANAP. An increase of fluorescence intensity indicates a transition from a relative hydrophilic environment to a more hydrophobic one. Two kinetic components can be distinguished at IFM motif. Left inset shows the voltage protocol with 10 mV voltage steps. Right inset shows the fluorescence signal at 60 mV fitted with only one fast component. H Comparison of steady state fluorescence (FV) curve and h-infinity curve. H-infinity curve is shown as 1-(h-infinity) curve, which closely tracks with the FV curve. N = 5. I Comparison of fluorescence signal and fast inactivation kinetics. The fast fluorescence signal is faster than the fast inactivation. The slow component follows more closely the fast inactivation kinetics. N = 5 for fluorescence signal, N = 4 for ionic current. Data is shown as Mean ± SEM.
Fast Movement at IFM Motif Reflects DIII_DIV Linker Movement Following DIV VSD Activation
To further investigate the nature of conformational dynamics at the IFM motif during fast inactivation, we monitored the movement of the IFM-bearing DIII_DIV linker by incorporating ANAP at L1319 position (L1319ANAP, Fig. 2A). ANAP incorporation at position 1319 was well tolerated (Supplementary Fig. 2 and Tables 1 and 2) and opposed to the F1304 position, L1319ANAP channels showed a marked decrease in the fluorescence intensity upon depolarization, suggesting a transition to a more hydrophilic environment (Fig. 2B). This is consistent with the existing structural results since L1319 is depicted as being completely exposed in aqueous environment in the fast inactivated structure (Fig. 2A). More interestingly, there is only one kinetic component in the fluorescence signal at L1319 position. This component showed similar time constants as the fast movement seen in F1304ANAP (Fig. 2C, D). Like the fast fluorescence signal in F1304ANAP channels, the fluorescence signal recorded in L1319ANAP is faster than the development of fast inactivation itself (Fig. 2E). To better understand the origin of this signal, we simultaneously recorded fluorescent signals and gating currents in L1319ANAP channels (Fig. 2F). Upon depolarization, gating charges in voltage sensors rapidly traverse the electric field, creating a fast, transient current, the gating current28,29. Given that gating currents track the movement of VSDs, this can be correlated to the observed fluorescence signals that originate from movements at the cytoplasmic III-IV linker. Gating current recordings from L1319ANAP channels revealed that the voltage-dependent gating charge movement, shown by the QV curve, overlaps closely with the steady-state FV curve (Fig. 2G). Despite the relative distance between the VSD and III-IV linker, the movement seen at L1319 linker region is more closely related to the voltage sensor movements than to fast inactivation. More specifically, the DIV VSD movement appears to directly trigger the DIII_DIV linker movement. When we compared the kinetics of the gating current and the fluorescence recordings, we discovered that the fluorescence signal at position 1319 explicitly followed the slow component of the gating current (Fig. 2H). Due to the asymmetry in Nav channels, voltage sensors from different domains translocate their charges at different rates30,31. Particularly, the activation of the DIV VSD is significantly slower than the other VSDs, giving rise to the slow component of the gating current in Nav channels30,32. The shared kinetics between fluorescence signal at position 1319 and the DIV VSD movement strongly suggest the activation of voltage sensor in DIV initiates the DIII_DIV linker movement during fast inactivation.
A To monitor the movement of DIIII_DIV linker region during fast inactivation, ANAP was incorporated at L1319 position, in the short alpha helix. B Example traces of L1319ANAP ionic current and fluorescence signal. Inset shows the voltage protocol. C Comparison of fluorescence signal from F1304ANAP and L1319ANAP. Inset shows a scaled comparison of the fluorescence signals at 40 mV. D The fast component of the fluorescence signal from F1304ANAP is not significantly different from the signal at position 1319 at 60 mV. Two-tailed Welch’s t-test was used, p = 0.6419. E Comparison of the fluorescence signal and the fast inactivation kinetics of L1319ANAP. The fluorescence signal was always faster than the fast inactivation development. N = 7 for fluorescence signal and N = 5 for ionic current. F Example traces of gating current measured from L1319ANAP. Inset shows the voltage protocol with 10 mV voltage steps. Framed inset shows a scaled comparison of fluorescence signal and gating current at 40 mV where gating charge moved in a fast single exponential manner. G Comparison of voltage-dependent gating charge movement (QV curve), FV curve of L1319ANAP, and 1-h-infinity curve. Clearly, the fluorescence signal is following more closely the gating charge movement. N = 9 for QV, N = 10 for FV, and N = 6 for h-infinity curve. H Comparison of fluorescence kinetics of L1319ANAP and gating charge movement. The fluorescence kinetics are similar to the slow component of gating charge movement which is contributed mostly by DIV VSD movement. N = 9. Data is shown as Mean ± SEM.
Backbone hydrogen bonding couples the DIII_DIV linker and DIV VSD movement
Since the DIII_DIV linker movement closely correlates with the activation of DIV VSD, there must exist interactions that couple these two structural components. A close structural analysis of this region revealed the existence of two hydrogen bonds between the DIII_DIV linker and the DIV S4_S5 linker (Fig. 3A). Specifically, residue N1477 at the elbow region of the DIV S4_S5 linker forms a hydrogen bond directly with the F1304 in the bound IFM motif and reciprocally, Q1309 at the DIII_DIV linker forms the other hydrogen bond with P1473 in DIV (Fig. 3A). Interestingly, both hydrogen bonds are formed between the nitrogen in the primary amide in the side chain and the carbonyl oxygen in the backbone. Removal of the primary amide in the side chain in both positions result in impaired fast inactivation (Fig. 3B). In the case of Q1309L and N1477D, the channels exhibit an IQM type phenotype with most of the fast inactivation absent11,31 (Fig. 3C, D and F, Supplementary Fig. 4 and Supplementary Table 2) without drastic change in GV curves (Fig. 3C and Supplementary Table 1). The mutation N1477D essentially exchanges this primary amide with a carboxylic group. At physiological pH used in our study, this group would be >99% deprotonated to the COO- carboxylate. Thus, there is no hydrogen atom in the N1477D sidechain to act as a hydrogen donor. As a result, N1477D sodium channels showed very little fast inactivation. We do however acknowledge that the N1477D substitution does not serve as a perfect mimic of a deprotonated asparagine sidechain as it also introduces the COO- in place of the amide group. An analysis of the recovery from fast inactivation for Q1309A channels further demonstrates that the identified hydrogen bond is essential for the stabilization of the fast inactivated state (Fig. 3E). In the absence of this interaction, the channels recover from fast inactivation exceedingly fast. Evidently, these hydrogen bonds are important for fast inactivation and likely serve as the functional bridges between the DIV VSD and the intracellular linker. To further demonstrate this, we disrupted one of the hydrogen bonds and investigated its impact on DIII_DIV linker movement. In L1319ANAP_Q1309A, fast inactivation becomes slower and incomplete (Fig. 3F). The fluorescence signal became much smaller (Fig. 3F) and, more importantly, progressed slower and recovered faster compared to L1319ANAP (Fig. 3F, G), demonstrating the identified interactions are indeed involved in coupling between the voltage sensor in DIV (via the DIV S4-S5 linker) and DIII_DIV linker.
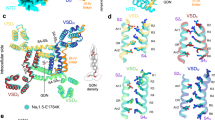
A Structural analysis reveals a network of hydrogen bonds between DIII_DIV linker (yellow) and DIV S4_S5 linker (cyan). Q1309 from the DIII_DIV linker forms a potential hydrogen bond with the backbone carbonyl group of P1473 DIV S4_S5 linker while N1477 in the same region forms directly a putative hydrogen bond with the backbone of the phenylalanine in IFM motif (pink). B Example ionic current from Q1309A, Q1309L, and N1477D respectively. Inset shows the voltage protocol. C GV curves for Q1309, Q1309L and N1477D. Activation of the mutant channels is not significantly altered compared to WT. N = 5 for Q1309A, N = 4 for Q1309L and N = 5 for N1477D. D h-infinity curves for the indicated channels. N = 4 for Q1309A, N = 4 for Q1309L and N = 6 for N1477D. E Q1309A recoveries from fast inactivation significantly faster than WT, suggesting a destabilized inactivated state. N = 3 for Q1309A. F Example traces of ionic current and fluorescence signal from L1319ANAP_Q1309A. Inset shows the voltage protocol. G Comparison of fluorescence kinetics of L1319ANAP and L1319ANAP_Q1309A. Mildly disrupting the hydrogen bond, L1319ANAP_Q1309A showed slowed fluorescence kinetics than L1319ANAP, suggesting the identified hydrogen bonds are important for the coupling of the DIV VSD and DIII_DIV linker. N = 6. Data is shown as Mean ± SEM.
IFM motif relays its conformational changes to the pore through hydrophobic and T-shaped π-π interactions
With the molecular underpinning of the fast component elucidated, we set out to investigate the slower conformational change at the IFM motif that triggers fast inactivation. Our previous work demonstrated the specialized role of the pore-lining hydrophobic residues in DIII and DIV S6 in fast inactivation22 (Fig. 4A). Therefore, residues in DIII and DIV S6 that could form potential interactions with the IFM motif might be involved in the coupling between the IFM motif and the fasti inactivation. We performed mutagenesis screenings on the DIII and DIV residues adjacent to the IFM. The screening revealed two residues that were particularly important for fast inactivation.
A Interactions of IFM motif (pink) with F1291 in DIII S6 (green) and I1589 in DIV S6 (olive). Previously identified fast inactivate is highlighted in red circle. Relative positions between IFM and F1291 suggest a potential T-shaped π-π interaction. B–E Example of ionic current from F1291 and I1589 mutants. In F1291A, F1291W and I1589A, significant fast inactivation removal was observed. Intriguingly, replacement of a single hydrogen atom to fluorine at the para position of F1304 also led to incomplete inactivation, suggesting the existence of the suspected aromatic interaction. Inset shows voltage protocol. F Double mutant of F1291A_I1589A led to complete removal of fast inactivation. No significant current decay was observed after 30 ms of depolarization. G GV curves of the all the mutants shown previously. N = 4 for WT and F1291A_I1589A, N = 5 for I1589A, F1291A and F1304ParaF_Phe, N = 7 for I1291W. H h-infinity curves for the mutants shown previously. N = 4 for WT and F1291A_I1589A, N = 5 for I1589A, F1291A and F1304ParaF_Phe, N = 7 for I1291W. I Example of ionic current from F1304ANAP_F1291W. Fast inactivation was largely absent. J Fluorescence signal from F1304ANAP_F1291W fitted with one exponential component. Different from F1304ANAP, fluorescence signals from F1304_F1291W can be adequately described by a mono exponential. By disrupting the interactions of IFM motif with the F1291, the second component of the fluorescence signal seen in F1304ANAP is eliminated. Data is shown as Mean ± SEM.
In DIII S6, when F1291 is mutated to either smaller or bigger residues, significant amount of fast inactivation was removed (Fig. 4B, C, H and Supplementary Table 2), suggesting that the precise size of the side chain is important for this interaction. Additionally, structural analysis based on the published structure33 also pointed out the possibility of a T-shaped π-π interaction between the phenylalanine in the IFM motif and F1291 (Fig. 4A). To precisely target this potential aromatic interaction, we utilized another unnatural amino acid, 4-fluorophenylalanine (ParaF-Phe). Substitution of the hydrogen at the para position on the aromatic ring targets the positive charge on the edge of the aromatic ring and would be predicted to weaken a T-shaped π-π interaction with a minimal impact on the volume of side chain34. In support of this notion, substitution of only one atom in F1304 via encoding of ParaF-Phe led to not only slowed, but also incomplete fast inactivation and faster recovery from inactivation (Supplementary Fig. 3 and Supplementary Table 2), suggesting a functional role for this π-π interaction between F1304 and F1291 (Fig. 4D, H).
Mutagenesis in DIV S6 revealed that changing I1589 led to a similar phenotype observed in DIII S6 mutations. I1589A severely impeded fast inactivation and showed large non-inactivating steady-state current (Fig. 4E, H and Supplementary Table 2). As the inactivation gate is made of large hydrophobic residues in S6 of both domain III and IV, the combination of F1291A in S6-DIII and I1589A in S6-DIV should eliminate fast inactivation. Indeed, combining both mutations in a single construct, F1291A_I1589A, produced channels where fast inactivation was completely removed. No significant reduction in current amplitude was seen after 30 ms of depolarization (Fig. 4F).
Our mutagenesis experiments on F1291 and I1589 confirmed their importance for fast inactivation. However, these results did not directly demonstrate those are residues coupled the IFM motif to the fast inactivation gate at the pore. To provide direct evidence, we investigated the movement of the IFM motif in the absence of the identified interactions. We recorded the ionic current and fluorescence signal in F1304ANAP_F1291W (Fig. 4I, J). Fast inactivation was largely removed in F1304ANAP_F1291W, but still, clear fluorescence signals were resolved from the IFM position. Evidently, disrupting the interaction between F1291 and F1304 did not abolish the binding of the IFM motif. The kinetics of the fluorescence signals were significantly faster than the residual fast inactivation. These observations are consistent with our previous results, suggesting that the fast component of the IFM movement was triggered by the DIV VSD and coupled by the hydrogen bonds, instead of interactions in the hydrophobic pocket. More importantly, now with disrupted interaction with F1291, the IFM motif movement showed only one fast fluorescence component, and the second slow movement seen before in F1304ANAP alone was largely absent (Fig. 4J), providing direct evidence that the identified residues are responsible for coupling the IFM motif and the pore.
Fast inactivation process requires sequential conformational changes
Finally, we monitored the movement of the pore during fast inactivation by encoding ANAP at Q1293, a position at the bottom of the pore-lining DIII S6 segment (Fig. 5A). Incorporation of ANAP at Q1293 led to minimal alteration in fast inactivation (Fig. 5B, D) and activation (Fig. 5C). The fluorescence signal from Q1293ANAP mirrored the development of fast inactivation, suggesting the movement seen at this part of the S6 helix reflects mostly conformational changes associated with fast inactivation instead of activation (Fig. 5E). Interestingly, there was a ~ 1 ms delay to the onset of the fast-inactivation-associated fluorescence signal, which was not observed in other ANAP mutants. This delay is similar to the previous observation in pronase treated giant squid axon35 (Fig. 5B inset). The observed delay also resembles the classical Cole-Moore shift36 and indicates that additional conformational changes are required before channel can enter into the fast inactivated state. It is likely that the two movements seen at the IFM position happen in a sequential manner in that the DIII_DIV linker needs to move to position the IFM at the right location before the second, slower conformation change can happen and triggers the fast inactivation. Supporting this notion, in the case of L1319ANAP_F1291A (Fig. 5F), the fluorescence signal was unaltered by abolishing the second, slower interaction (Fig. 5G), suggesting the DIII_DIV linker movement happened upstream of the second interaction, and the fast inactivation process is best described as sequential conformational changes.
A To track the pore movement during fast inactivation, ANAP was incorporated at Q1293 position at the bottom of DIII S6 (green). B Example of ionic current and fluorescence signal recorded from Q1293ANAP. Inset shows overlapped comparison of ionic current and fluorescence signal at 60 mV. There is a ~ 1 ms delay to the onset of fluorescence signal, suggesting a sequential movement for fast inactivation. C, D GV curve and h-infinity curve for Q1293ANAP. No drastic difference was observed. N = 5 for Q1293ANAP. E Comparison of fluorescence kinetics and fast inactivation kinetics. Fluorescence kinetics follows closely the fast inactivation development, suggesting the conformational change observed at this position reports mostly pore movement associated with fast inactivation. N = 10. F) Example of ionic current and fluorescence signal from L1319ANAP_F1291A. G Comparison of fluorescence kinetics between L1319ANAP and L1319ANAP_F1291A. No significant difference in kinetics is seen in the DIII_DIV linker movement after abolishing the interaction between IFM and F1291, favoring a sequential model for fast inactivation. N = 7 for L1319ANAP_F1291A and N = 8 for L1319ANAP. Data is shown as Mean ± SEM.
Discussion
The IFM motif couples the DIV_VSD to the fast inactivation gate
The interconnectivity between VSD function and fast inactivation has long been established. From the pronase perfused giant squid axon35 to charge neutralization experiments32 on DIV VSD, it is clear that VSD activation is required for fast inactivation. Further, fast inactivation can also influence VSD function, as has been shown in studies on charge immobilization10,37. Our results indicate that the IFM motif couples the voltage-driven movement of the DIV VSD to the inner mouth of the pore. By incorporating ANAP directly into the IFM motif, we find that the resulting fluorescence signal reveals two sequential movements at IFM motif during fast inactivation. For the study of sodium channel inactivation, ANAP enables the tracking of ultra-fast conformational changes on the sub- millisecond time scale, limited by the speed of the voltage clamp. These data show that the fast component of the fluorescence originates from the DIII_DIV linker that is tightly coupled to DIV VSD activation and the slower movement that closely aligns with the closure of the fast inactivation gate in the pore. Evidently, the movement of the IFM motif transduces the activation of the DIV VSD directly to closure of the inner entrance to the pore. These two movements also explain the domain specific charge immobilization during fast inactivation shown previously in MTS modification experiments10,38. It is likely that the movement of the DIII_DIV linker triggers the immobilization of DIV VSD by binding of the IFM with the DIV S4-S5 linker, while the closure of the fast inactivation gate immobilizes DIII VSD. We also note that while VCF combined with ANAP serves as a powerful technique for assessing otherwise electrically silent protein conformational dynamics, we need to point out some potential caveats. The size of ANAP, though similar, is somewhat larger than tryptophan (Fig. 1B) and incorporation of ANAP could impact channel gating, similar to any other site-directed mutation (Supplementary Tables 1 and 2). In our experiments, we have focused on sites of ANAP incorporation where encoding has a relatively minor impact on fast inactivation, such as the F1304ANAP.
Functional identification and validation of the binding site for IFM motif
In this work, we further elucidated a network of hydrogen bonds that forms the structural connection between DIV VSD and cytoplasmic DIII_DIV linker. One intriguing observation is that in both hydrogen bonds, it is the carbonyl groups in the peptide backbone that serve as the H-bond acceptors. In particular, N1477 at DIV S4_S5 linker forms a direct interaction with the backbone carbonyl group of the phenylalanine in IFM motif. One natural conclusion from this observation would be that the exact identity of the residue at the second position of IFM motif shouldn’t influence the formation of the hydrogen bonds and subsequently shouldn’t impede the binding of IFM motif. The binding of IFM motif has been traditionally associated with interactions between the side chain of the phenylalanine residue and its surrounding hydrophobic pocket. However, our results argue that the driving force of the IFM motif binding comes mostly from the hydrogen bond formation with the peptide backbones. In other words, the binding site of the IFM motif is formed not by the hydrophobic pocket but rather Q1309 in DIII_DIV linker and N1477 in DIV S4_S5 linker. It has been observed in molecular dynamic simulations before that mutation N1662D, a disease causing mutant in hNav1.2 (equivalent as N1477 in rNav1.4), led to a significantly decreased dwell time of IFM within the hydrophobic pocket39, a finding consistent with our electrophysiological results. Removing one of the hydrogen bonds in Q1309A led to an destabilized fast inactivated state as manifest in an increased recovery from inactivation. Further, the fluorescence data, which arguably track motions in this area, also support this conclusion. Disruption of the hydrophobic pocket at F1291, while resulting in significant removal of fast inactivation (F1291W), didn’t impede the binding of IFM motif in F1304ANAP_F1291W (Fig. 4J). On the other hand, a mild mutation Q1309A clearly led to impaired coupling between DIV VSD and DIII_DIV linker as was seen in L1319ANAP_Q1309A. Finally, charge immobilization experiments with IQM channels also demonstrated that despite almost complete removal of fast inactivation from ionic current, ~30% (in contrast to 60% in WT) of total gating charge still became immobilized after 24 ms of depolarization (Supplementary Fig. 4). Clearly, even in IQM channels with diminished fast inactivation, some vestigial conformational features along the fast inactivation pathway are in place. In the scenario, the DIII_DIV linker might still bind to the identified residues, leading to some degree of charge immobilization, and likely only in the DIV VSD13,38, but as no further movement triggers the closure of the fast inactivation gate, the immobilization of DIII does not occur. Taken together, the present data strongly favors N1477 and P1473 in DIV S4_S5 linker as the binding sites for IFM motif, instead of a hydrophobic pocket.
Molecular mechanism of fast inactivation
The current and available data are therefore consistent with a lock and key model for fast inactivation in voltage-gated sodium channels. Figure 6 is a schematic representation of the sequence of events that lead to fast inactivation. Upon depolarization, the sensors of domain I, II and III activate and the channel opens (Fig. 6A, E, I). The domain IV sensor next activates30,32 and in doing so exposes the binding site for IFM motif at P1470 and N1477 in DIV S4_S5 linker, similar to a keyhole becoming available (Fig. 6 B, F, J). The IFM-containing DIII_DIV linker, likely mobile in aqueous solution, binds to DIV S4_S5 linker via the H-bonds (Fig. 6 C, G) and positions IFM motif, the key, into the hydrophobic pocket, the keyhole (Fig. 6K). The binding of IFM motif allows the interactions between F1291 and I1589 via additional hydrophobic and aromatic forces, leading to a second conformational change at the hydrophobic pocket. In F1291Q channels, despite some steady state current, the majority of fast inactivation persists, dissimilar to F1304Q (IQM) phenotype (Supplementary Fig. 5). This asymmetric effect suggests that another conformational change must occur to support the interactions between F1291 and I1589. One possibility is that DIII S6 and DIV S6 helices undergo a rotational movement to position F291 and I1589 into the hydrophobic pocket interacting with the IFM motif and thus locking the two S6 helices in position. At the same time, rotations of the large hydrophobic amino acids forming the inactivation gate into the conduction pathway, interrupts the flow of Na+ ions, thus producing fast inactivation22 (Fig. 6D, H, L). Modifications to either IFM motif or the hydrophobic pocket would lead to incompatibility between the key and the lock, leading to impaired inactivation.
Left column is a schematic diagram of the sequence of events. The Middle column shows details of the residues involved. The Right column shows the lock and key analogy. Upon depolarization, VSD from DI to DIII activates and subsequently opens the channel. Given that DIV VSD resides in the resting state (A and E), the hydrogen bond mediated binding pocket for IFM motif is shielded (the lock is covered, I). Maintaining the depolarization, it leads to the activation DIV VSD which reorganizes the DIV S4_S5 linker and displays P1470 and N1477 towards the intracellular aqueous environment (B and F). This process is similar to opening a guard on the lock and exposing the keyhole (J). Now the DIII_DIV linker, possibly mobile in the aqueous environment, binds to the DIV S4_S5 linker positioning the IFM motif, the key, into the hydrophobic pocket (C and G), into the keyhole (K). The binding of IFM motif allows the interactions of F1291 and I1589 with the phenylalanine in IFM motif, creating a rotation of the pore-forming S6 helices in DIII and DIV (D and H), like turning the key in the keyhole (L). This helical rotation in turn locates the hydrophobic gate (yellow residues in D and H and blocking gate in L) in the permeation pathway, blocking the Na+ permeation as a result.
In reverse, the recovery from inactivation requires disengagement of the IFM motif. Results of F1304ANAP demonstrate that the recovery from inactivation and off-fluorescence signal share similar time course, supporting this idea (Supplementary Fig. 1). The two major conformational changes that occurred during fast inactivation are responsible for immobilization of DIV VSD and DIII VSD respectively as suggested by the IQM gating current result13,37 (Supplementary Fig. 4). Under hyperpolarized voltages, a transient unbinding of the IFM motif would allow the deactivation of voltage sensors in DIII and DIV that are previously immobilized. This inward movement of the sensors subsequently abolish the binding site for IFM motif, similar to covering the keyhole on the lock, and allows the channels to recover from the fast inactivated state.
It’s worth noting that in this scheme, we didn’t assign specific roles to the C terminus domain (CTD) of the channel. It has been shown that CTD has profound impact on the fast inactivation, especially among different isoform of the Nav channels40,41. In a chimeric channel structure, the CTD was shown to be involved in two important switches that control fast inactivation42. However, in most available mammalian Nav structures, the CTD remains poorly resolved, a feature which has impeded our understanding on the role CTD plays. Clearly additional work is necessary to elucidate the exact function of CTD.
Alternative fast inactivated state in NaV channels
Here, we addressed the molecular pathway from open channels to the fast inactivated state. It is important to point out that closed Nav channels can also inactivate, bypassing the open state altogether, and this is essential for physiological processes10,43,44. Reversal of inactivation by movement through closed states prevents Na+ conduction upon hyperpolarization and determines the absolute refractory period in firing neurons. Structural work suggests a similar pore conformation in the closed inactivated state and open inactivated state19,33. Fluorescent measurements on individual sensor movements have demonstrated that DIV VSD can activate at more hyperpolarized voltages compared to the other VSDs30. Then, as the movement of DIV VSD alone seems to be enough to trigger inactivation, while opening requires all the other VSDs to be activated, it is possible that fast inactivation may occur regardless of an open or a closed pore. When we monitored the pore movement at Q1293 position, two components of fluorescence signals were observed upon hyperpolarization (Fig. 5B). It is possible these two components capture the transitions from open inactivated state to closed inactivated state and from closed inactivated state to closed state, respectively. However, further investigations are required to elucidate the exact structural rearrangement at the pore during these transitions. But most likely, the observed sequential conformational changes also occur during closed state inactivation.
Evolutionary and isoform conservation of the identified fast inactivation apparatus
We compared Nav channels from Bilateria, Cnidaria, and Choanoflagellata (Supplementary Fig. 7) and intriguingly found the herein identified residues are highly conserved, some even more conserved than the IFM motif itself. F1291, N1477 and Q1309 are conserved across all sequences compared, even in the ancestral form of Nav channel in Monosiga brevicollis45. In the case of purple sea urchin, Strongylocentrotus purpuratus, an isoleucine replaces the phenylalanine in the IFM motif but nevertheless, all three residues remain conserved at the equivalent positions. I1589, on the other hand, is less conserved. In the ancestral Nav channel, a valine is found at 1589 equivalent position, and valine is the only alternative amino acid found at this position across different phyla. The high conservation of the herein identified fast inactivation components strongly supports the idea that the molecular mechanism described here in mammalian Nav1.4 channel is conserved across the animal kingdom and appears at the same time as the evolutionary invention of the Nav channel in Choanoflagellata.
Methods
Site-directed mutagenesis and cRNA in vitro synthesis
rNav1.4 in pBSTA vector, flanked by Xenopus laevis β-globin sequences, was used in this study for all the physiological experiments22. Point mutations were generated using QuikChangeTM mutagenesis method (Agilent). The PCR products were first digested by DpnI and were used to transform the XL10-gold ultra-competent cells (Agilent). After ampicillin resistance screening, plasmids were purified from the colonies using standard miniprep protocols (Macherey-Nagel). Purified plasmids were sent to Plasmidsaurus for whole plasmid sequencing to confirm the introduction of the mutations. Verified plasmids were first linearized and later in vitro transcribed to cRNA with mMESSAGE mMACHINE™ T7 Transcription Kit (Thermo Fisher Scientific).
Xenopus laevis oocyte preparation and channel expression
Ovaries of Xenopus laevis were purchased from XENOPUS1 (Dexter, Michigan). The follicular membrane was removed using collagenase type II (Worthington Biochemical Corporation) at 2 mg/mL with bovine serum albumin at 1 mg/mL (BSA). After defolliculation, stage V–VI oocytes were then selected and microinjected with 50-100 ng cRNA. Injected oocytes were incubated at 18 °C for 1–5 days in SOS solution (in mM: 100 NaCl, 5 KCl, 2 CaCl2, 0.1 EDTA, and 10 HEPES at pH 7.4) supplemented with 50 µg/mL gentamicin. Unless otherwise stated, all chemicals were purchased from Sigma-Aldrich.
Cut-open voltage clamp on Xenopus laevis oocytes
Macroscopic ionic and gating current were recorded using the cut-open voltage clamp technique46. Micropipettes filled with 3 M CsCl, with resistance between 0.6 and 1.2 MΩ were used to measure the internal voltage of the oocytes. Current data were first filtered online at 20 kHz with a low-pass 4 pole Bessel filter and sampled by a 16-bit A/D (USB-1604; Measurement Computing, Norton, MA) converter at 1 MHz. A feedbackback Peltier device was used to control the temperature at 13 ± 1 °C for all experiments. For ionic current experiments, unless otherwise stated, were conducted in external solution consisted of in mM: 28 Na methylsulfonate (MES), 92 N-methyl-D-glucamine (NMG) MES, 2 Ca MES, 10 HEPES, 0.1 EDTA, pH = 7.4 and internal solution consisted of in mM: 12 Na MES, 108 NMG MES, 10 HEPES, 2 EGTA, pH = 7.4. The capacitive transient was manually compensated with a dedicated circuit and further removed by an online P/−4 protocol with a holding voltage of either −80 or −90 mV10. For gating current experiments, all experiments were conducted in external solutions consisted of in mM: 120 NMG MES, 2 Ca MES, 10 HEPES, 0.1 EDTA, 750 nM TTX pH = 7.4 and in internal solution consisted of in mM: 120 NMG MES, 10 HEPES, 2 EGTA, pH = 7.4. In this case, an online P/4 at 40 mV was used to subtract the capacitive transients. Voltage clamp speed measured by capacitive transient current gave a time constant around 70 µs.
Unnatural amino acid incorporation
Two methods were used to incorporate unnatural amino acids into Nav channels. For ANAP, two separate cytoplasm injections were performed. The first oocyte injection introduced four different components: folded synthetic tRNA as RNA at 2 ng/nL (Integrated DNA Technologies, Iowa City, IA), ANAP synthetase cRNA at 0.3 ng/nL, mutated Xenopus laevis release factor 1 (E55D) cRNA at 0.3 ng/nL, and the unnatural amino acid ANAP itself in methyl ester form (Cayman Chemical) at 2 mM. 24 hours after the first injection, Nav1.4 channel cRNA bearing the amber stop codon mutation at the desired location was injected at 1 ng/nL. 24 to 72 h after the second injection, the oocytes were used in voltage clamp fluorimetry experiments.
The encoding of 4-fluoro-L-phenylalanine was appended to synthetic pyl tRNA as previously reported47,48. Briefly, the cyanomethyl ester of N-Boc-4-fluoro-L-Phe was reacted with the dinucleotide pdCpA in dimethylformamide, purified on HPLC, and deprotected with trifluoroacetic acid. A 3 mM stock solution on the amino acid-dinucleotide chimera in dimethylsulfoxide was prepared and added to T4 RNA ligase and pyl tRNA lacking the terminal CA nucleotides to assemble the full-length tRNA with 4-fluoro-L-Phe attached. Pellets (1 nmol) were stored dry at −78 °C until needed. Lyophilized conjugated tRNA was resuspended in 3 mM sodium acetate at pH 5.5 to an approximate concentration of 2.5 µg/µL. Conjugated tRNA and cRNA of the mutated channel were introduced simultaneously into the oocytes, and after 24 to 48 h of expression, oocytes were used in voltage-clamp experiments.
Voltage clamp fluorimetry experiments
An avalanche photodiode or a photomultiplier was used to record the fluorescence signal. A 365 nM LED (Thor Lab) was used as the excitation light source. An AT440DC dichroic mirror purchased from Chromas (Chroma Technology Corp., Bellows Falls, VT, USA) was used to split the beam, and a 470/35 bandpass optical filter was used as the emission filter. No excitation filter was used. In the case of the photomultiplier (H10304-20-NN, Hamamatsu, Japan), the output signal was first processed by a home-built I to V converter, filtered by a low-pass 8 pole Bessel filter at 10 kHz, and sampled by the AD converter every 1 µs. Photobleaching was later subtracted with a Matlab routine and then digitally filtered at 5 kHz or 1 kHz.
Data analysis
Data recorded in the work is analyzed by GraphPad 10 (Prism), Excel (Microsoft), Matlab R2023a (Mathworks), ChimeraX 1.8 (UCSF), Snapgene (V7.0), and in-house software (Analysis 2.0 and GPatchM64_v1.1).
In all experiments, N represents the number of different Xenopus oocytes tested. An N = 5, for instance, represents that 5 different oocytes were measured experimentally and their results analyzed.
i) GV curves calculation: Ionic conductance was calculated by dividing the peak ionic current by the driving force determined experimentally. After normalization, GV curves were fitted with a two-state model described below,
where R is the ideal gas constant, T is the absolute temperature in Kelvin, z is the apparent charge, F is the Faraday constant.
ii) FV curve calculation: Steady state fluorescence signal was normalized to maximum fluorescence and then described with the model below,
where R, T, F and z are as previously defined
iii) Steady state fast inactivation curve (h-infinity curve) was calculated by plotting normalized peak Na+ current during test pulse after 50 ms conditioning pulse voltage and fitted using a two-state model,
where Base is the steady state current, and R, T, F and z are as previously defined.
iv) The time constant of fast inactivation, gating charge movement or fluorescence kinetics was calculated by fitting the signals with either one or two exponential decays using the general equation below,
where A1, τ1, A2, and τ2 represent amplitudes and time constants of the first and second components, respectively. Base represents the steady state current or fluorescence. When one exponential was used, only A1, and τ1 were fitted to the data.
v) Recovery from fast inactivation was calculated by dividing the peak current in the control pulses by the peak current during test pulses. The data was then described by either one or two component association model,
where A1, τ1, A2, and τ2 represent amplitudes and time constants of the first and second components, respectively. When one exponential was used, only A1, and τ1 were fitted to the data.
vi) Statistics: All the data shown in the figures were plotted as Mean ± SEM. The N value represents the number of individual cells tested. Comparison of kinetics from different mutants was done by the Welch’s t-test, which assumes a Gaussian distribution of the two compared samples but doesn’t assume the variance in two populations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Protein structures used in the study were published under the access code 7XVF All data used to generate the main and Supplementary Figs. are submitted as separate Source Data file. The detailed fitting parameters and the goodness of fitting are also provided in the Source data. The raw current/fluorescence data and resources presented in the paper will be made available upon request. Source data are provided with this paper.
References
Bean, B. P. The action potential in mammalian central neurons. Nat. Rev. Neurosci. 8, 451–465 (2007).
Hodgkin, A. L. & Huxley, A. F. Currents carried by sodium and potassium ions through the membrane of the giant axon of Loligo. J. Physiol. 116, 449–472 (1952).
Hodgkin, A. L. & Huxley, A. F. The components of membrane conductance in the giant axon of Loligo. J. Physiol. 116, 473–496 (1952).
De Lera Ruiz, M. & Kraus, R. L. Voltage-gated sodium channels: structure, function, pharmacology, and clinical indications. J. Med. Chem. 58, 7093–7118 (2015).
Fertleman, C. R. et al. SCN9A mutations in paroxysmal extreme pain disorder: allelic variants underlie distinct channel defects and phenotypes. Neuron 52, 767–774 (2006).
Waxman, S. G. Painful Na-channelopathies: an expanding universe. Trends Mol. Med. 19, 406–409 (2013).
Phillips, L. & Trivedi, J. R. Skeletal muscle channelopathies. Neurotherapeutics 15, 954–965 (2018).
George, A. L. Molecular basis of inherited epilepsy. Arch. Neurol. 61, 473–478 (2004).
Kyndt, F. et al. Novel SCN5A mutation leading either to isolated cardiac conduction defect or Brugada syndrome in a large French family. Circulation 104, 3081–3086 (2001).
Armstrong, C. M. & Bezanilla, F. Inactivation of the sodium gating current. J. Gen. Physiol. 4, 865–876 (1977).
West, J. W. et al. A cluster of Hydrophobic amino acid residues required for fast Na+-channel inactivation. Proc. Natl. Acad. Sci. USA 89, 10910–10914 (1992).
Horn, R., Ding, S. & Gruber, H. J. Immobilizing the moving parts of voltage-gated ion channels. J. Gen. Physiol. 116, 461–476 (2000).
Sheets, M. F., Kyle, J. W. & Hanck, D. A. The role of the putative inactivation lid in sodium channel gating current immobilization. J. Gen. Physiol. 115, 609–619 (2000).
Kellenberger, S., West, J. W., Scheuer, T. & Catterall, W. A. Molecular analysis of the putative inactivation particle in the inactivation gate of brain type IIA Na+ channels. J. Gen. Physiol. 109, 589–605 (1997).
Kellenberger, S., Scheuer, T. & Catterall, W. A. Movement of the Na+ channel inactivation gate during inactivation. J. Biol. Chem. 271, 30971–30979 (1996).
Chahine, M., Deschênes, I., Trottier, E., Chen, L. Q. & Kallen, R. G. Restoration of fast inactivation in an inactivation-defective human heart sodium channel by the cysteine modifying reagent Benzyl-MTS: Analysis of IFM-ICM mutation. Biochem. Biophys. Res. Commun. 233, 606–610 (1997).
Kühlbrandt, W. The resolution revolution. Science 343, 1443–1444 (2014).
Pan, X. et al. Structure of the human voltage-gated sodium channel Nav1.4 in complex with β1. Science 362, eaau2486 (2018).
Huang, G. et al. High-resolution structures of human Nav1.7 reveal gating modulation through α-π helical transition of S6IV. Cell Rep. 39, 110735 (2022).
Jiang, D. et al. Open-state structure and pore gating mechanism of the cardiac sodium channel. Cell 184, 5151–5162.e11 (2021).
Jiang, D. et al. Structure of the cardiac sodium channel. Cell 180, 122–134.e10 (2020).
Liu, Y., Bassetto, C. A. Z., Pinto, B. I. & Bezanilla, F. A mechanistic reinterpretation of fast inactivation in voltage-gated Na+ channels. Nat. Commun. 14, 1–13 (2023).
Patton, D. E., West, J. W., Catterall, W. A. & Goldin, A. L. Amino acid residues required for fast Na+-channel inactivation: charge neutralizations and deletions in the III-IV linker. Proc. Natl. Acad. Sci. USA 89, 10905–10909 (1992).
Chatterjee, A., Guo, J., Lee, H. S. & Schultz, P. G. A genetically encoded fluorescent probe in mammalian cells. J. Am. Chem. Soc. 135, 12540–12543 (2013).
Hyun, S. L., Guo, J., Lemke, E. A., Dimla, R. D. & Schultz, P. G. Genetic incorporation of a small, environmentally sensitive, fluorescent probe into proteins in Saccharomyces cerevisiae. J. Am. Chem. Soc. 131, 12921–12923 (2009).
Cha, A. & Bezanilla, F. Characterizing voltage-dependent conformational changes in the shakerk+ channel with fluorescence. Neuron 19, 1127–1140 (1997).
Kalstrup, T. & Blunck, R. Voltage-clamp fluorometry in Xenopus oocytes using fluorescent unnatural amino acids. J. Vis. Exp. 2017, 1–9 (2017).
Bezanilla, F. Gating currents. J. Gen. Physiol. 150, 911–932 (2018).
Armstrong, C. M. & Bezanilla, F. Currents related to movement of the gating particles of the sodium channels. Nature 242, 459–461 (1973).
Chanda, B. & Bezanilla, F. Tracking voltage-dependent conformational changes in skeletal muscle sodium channel during activation. J. Gen. Physiol. 120, 629–645 (2002).
Liu, Y. & Bezanilla, F. A sodium channel mutant removes fast inactivation with the inactivation particle bound. J. Gen. Physiol. 157, e202413667 (2025).
Capes, D. L., Goldschen-Ohm, M. P., Arcisio-Miranda, M., Bezanilla, F. & Chanda, B. Domain IV voltage-sensor movement is both sufficient and rate limiting for fast inactivation in sodium channels. J. Gen. Physiol. 142, 101–112 (2013).
Huang, G. et al. Unwinding and spiral sliding of S4 and domain rotation of VSD. 1–9 https://doi.org/10.1073/pnas.2209164119/-/DCSupplemental.Published (2022).
Galles, G. D. et al. Tuning phenylalanine fluorination to assess aromatic contributions to protein function and stability in cells. Nat. Commun. 14, 1–12 (2023).
Armstrong, C. M., Bezanilla, F. & Rojas, E. Destruction of sodium conductance inactivation in squid axons perfused with pronase. J. Gen. Physiol. 62, 375–391 (1973).
Cole, K. S. & Moore, J. W. Potassium ion current in the squid giant axon: dynamic characteristic. Biophys. J. 1, 1–14 (1960).
Albert, C., Ruben, P. C., George, A. L., Fujimoto, E. & Bezanilla, F. Voltage sensors in domains III and IV, but not I and II, are immobilized by Na+ channel fast inactivation. Neuron 22, 73–87 (1999).
Sheets, M. F. & Hanck, D. A. Charge immobilization of the voltage sensor in domain IV is independent of sodium current inactivation. J. Physiol. 563, 83–93 (2005).
Berecki, G. et al. Nav1.2 channel mutations preventing fast inactivation lead to SCN2A encephalopathy. Brain 148, 212–226 (2025).
Mantegazza, M., Yu, F. H., Catterall, W. A. & Scheuer, T. Role of the C-terminal domain in inactivation of brain and cardiac sodium channels. Proc. Natl. Acad. Sci. USA 98, 15348–15353 (2001).
Choi, J. S., Tyrrell, L., Waxman, S. G. & Dib-Hajj, S. D. Functional role of the C-terminus of voltage-gated sodium channel Na v1.8. FEBS Lett. 572, 256–260 (2004).
Clairfeuille, T. et al. Structural basis of a-scorpion toxin action on Na v channels. Science 363, eaav8573 (2019).
Armstrong, C. M. Na channel inactivation from open and closed states. Proc. Natl. Acad. Sci. USA 103, 17991–17996 (2006).
Horn, R., Vandenberg, C. A. & Lange, K. Statistical analysis of single sodium channels. Effects of N-bromoacetamide. Biophys. J. 45, 323–336 (1984).
Liebeskind, B. J., Hillis, D. M. & Zakon, H. H. Evolution of sodium channels predates the origin of nervous systems in animals. Proc. Natl. Acad. Sci. USA 108, 9154–9159 (2011).
Stefani, E. & Bezanilla, F. Cut-open oocyte voltage-clamp technique. Methods Enzymol. 293, 300–318 (1998).
Hohsaka, T., Kajihara, D., Ashizuka, Y., Murakami, H. & Sisido, M. Efficient incorporation of nonnatural amino acids with large aromatic groups into streptavidin in in vitro protein synthesizing systems https://doi.org/10.1021/JA9813109.S001 (1998).
Robertson, S. A., Ellman, J. A. & Schultz, P. G. A general and efficient route for chemical aminoacylation of transfer RNAs. J. Am. Chem. Soc. 113, 2722–2729 (1991).
Acknowledgements
We would like to thank Gethiely Gasparini and Hlafira Polishchuk for oocyte preparation, DNA mutation and RNA preparation. Y.L. and F.B. are supported by NIH grant R01-GM030376 and National Science Foundation Award QuBBE QLCI (NSF OMA-2121044). C.A.A. and J.D.G. are supported by R35GM148239.
Author information
Authors and Affiliations
Contributions
Y.L. performed electrophysiological and voltage clamp fluorimetry experiments. J.D.G. performed the tRNA ligation experiments. Y.L. analyzed the data. Y.L. and F.B. conceptualized the project. Y.L., C.A.A., and F.B. interpreted the data. Y.L. wrote the original draft. Y.L., J.D.G., C.A.A., and F.B. edited and reviewed the paper C.A.A. and F.B. supervised the project and oversaw funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflicting interests.
Peer review
Peer review information
Nature Communications thanks Bruce Bean and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Galpin, J.D., Ahern, C.A. et al. Molecular basis of sodium channel inactivation. Nat Commun 16, 10565 (2025). https://doi.org/10.1038/s41467-025-65587-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65587-1