Abstract
The receptor-binding domain (RBD) of the SARS-CoV-2 spike (S) protein continues to evolve, facilitating antibody evasion. It remains unclear whether any conserved RBD epitopes persist across SARS-CoV-2 variants and whether vaccination and/or breakthrough infection (BTI) can elicit antibodies capable of targeting these conserved regions to counter future variants. Here, using a heterogeneous double-bait single B-cell sorting strategy, we identify a subset of antibodies with broad-spectrum RBD binding, including recognition of SARS-CoV-1 and emerging variants such as EG.5.1, BA.2.86, JN.1, and KP.2/3. These broadly binding antibodies (bbAbs) exhibit elevated levels of somatic hypermutation but are infrequently derived from clonally expanded B lymphocytes. Passive transfer of representative bbAbs reduces viral infection in a male hamster model. Structural analyses reveals that these bbAbs primarily target three distinct, highly conserved RBD epitopes, suggesting potential regions of future mutational pressure and highlighting the presence of conserved and immunogenic RBD conformations that may serve as a foundation for the development of broadly protective vaccines.
Similar content being viewed by others
Introduction
Since the first infection reported in late 2019, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has already caused nearly 800 million confirmed cases and more than 7 million deaths globally as of late 2024. During the past years, the SARS-CoV-2 virus, being remarkably dynamic, rapidly evolved from the original wildtype (WT) strain to a series of variants, including five major variants of concern (VOCs), such as Alpha, Beta, Gamma, Delta, and Omicron1,2, and multiple major Omicron subvariants, such as BA, BQ, XBB, and KP3,4.
Since the Alpha, Beta, Gamma, and Delta variants bear only few mutations within the viral Spike (S) protein, a very low level of antibody evasion was observed, with only few monoclonal antibodies (mAbs) losing neutralizing activity against these VOCs5,6,7,8. However, the later emerged Omicron and its subvariants BA.1, BA.2.12.1, BA.3, BA.4/5, BQ, and XBB are heavily mutated, with more than 30 amino acid substitutions on their S proteins. Nearly half of these mutations are located in the receptor binding domain (RBD), leading to a remarkable reduction of neutralization of many mAbs9,10,11,12. Especially, the BQ and XBB subvariants completely evaded almost all clinically available mAbs, exhibiting striking antibody evasion characteristics9,10,13. Most recently, the newly identified Omicron subvariants, KP.2 and KP.3, with over 60 amino acid substitutions in their S proteins and in total 28 RBD mutations compared to the WT strain, showed robust antibody evasion, even against those JN.1-induced antibodies14,15.
Due to the immune imprinting, the antibody repertoire diversity after repeated SARS-CoV-2 WT vaccination is limited during an Omicron breakthrough infection (BTI)16,17. This immune imprinting could be largely mitigated by repeated Omicron exposure, with a series of Omicron-specific antibodies induced18. Therefore, facing the endlessly emerging variants with remarkable dynamic evolution and striking antibody evasion capacity, a question to be answered is whether our humoral immune system, after rounds of vaccinations and infections, could still generate mAbs broadly binding to RBDs of different SARS-CoV-2 sublineages and exerting antiviral activity against future variants.
Distinct combinations of amino acid mutations on RBD contribute to antibody evasion9,15. Conversely, these RBD mutations might weaken the binding affinity with the host receptor of SARS-CoV-2, angiotensin-converting enzyme 2 (ACE2), which is largely compensated by certain epistatic mutations, such as N501Y19, to maintain the overall ACE2 affinity for the S proteins of different SARS-CoV-2 variants20. Consequently, although the receptor-binding motif (RBM) region within RBD is highly mutated in the Omicron variants, facilitating their immune evasion against many RBM-targeting antibodies, the binding of RBM with ACE2 have been maintained for all reported SARS-CoV-2 variants so far. Thus, in terms of ACE2 binding, the highly mutated RBM region is structurally conserved. On the other hand, though many RBD amino acid residues are highly conserved throughout SARS-CoV-2 lineages, the capacity of certain residues to elicit human antibody response might be limited. Therefore, to evaluate and achieve a broad COVID-19 vaccine, there remains a need to further understand our human antibody immune response against the RBD, and to reveal structurally conserved RBD epitopes with high immunogenicity in human.
Here, we adopted a single-B cell sorting strategy using heterogeneous double baits (SARS-CoV-1 RBD and XBB RBD), and identified a minority of non-clonally expanded mAbs from vaccinated or BTI individuals. These antibodies showed a broad-spectrum of RBD binding activities, recognizing mainly three non-overlapping RBD epitopes, including two highly sequence-conserved epitopes and one partially mutated but structurally conserved epitope (a part of RBM). Our findings suggest that broadly RBD-binding mAbs could be induced post-vaccination and infection (existing as minority antibodies), and provide at least low-level protection against the future SARS-CoV-2 variants and even sarbecoviruses.
Results
Antibody cloning using heterogenous double-bait sorting strategy
Our previous finding showed a negative correlation of neutralizing activities against SARS-CoV-1 and SARS-CoV-2 Omicron for broad neutralizing antibodies (bnAbs) against RBD (Supplementary Fig. 1a)21. Therefore, to isolate a minority of broadly binding antibodies (bbAbs) against RBDs, we proposed to utilize the cross-reactivity against SARS-CoV-1 and one of Omicron variants, XBB (Fig. 1a). To do so, we adopted a heterogenous double-bait single-B-cell sorting strategy by using phycoerythrin (PE)-conjugated SARS-CoV-1 RBD and Allophycocyanin (APC)-conjugated XBB RBD proteins as the two heterogenous bait proteins (Fig. 1b and Supplementary Fig. 1b). Since none of these donors have been exposed to SARS-CoV-1 or XBB at the time of blood donation, the two heterogenous bait proteins could be considered as the unexposed or the future variants of SARS-CoV-2.
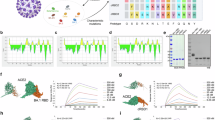
a Phylogenetic tree of the RBD proteins. The RBD proteins of SARS-CoV-1 and XBB, both highlighted, were used in this study as bait proteins for single B cell sorting. b Diagram of the heterogeneous double-bait sorting strategy used in this study. Two heterogenous bait proteins were used to sort single B cells recognizing phycoerythrin (PE)-conjugated SARS-CoV-1 RBD and allophycocyanin (APC)-conjugated XBB RBD proteins. c Immunization history of four donors. The four recruited convalescent donors experienced vaccination of inactivated vaccines, BA.5 breakthrough infection and blood donation. d Frequency of human B cells recognizing heterogenous bait proteins, PE-conjugated SARS-CoV-1 RBD and APC-conjugated XBB RBD, in donor M, N, O, and P. e Initial screening for mAbs with binding activity against both SARS-CoV-1 RBD and XBB RBD. Each pie chart represents a donor with the number indicating the isolated mAbs with naturally paired heavy and light chains. ELISA categorized these mAbs into four types: red, binders against both SARS-CoV-1 RBD and XBB RBD; orange, antibodies binding only XBB RBD; green, antibodies binding only SARS-CoV-1 RBD; and white, mAbs with weak/no binding activity against both RBDs or mAbs with low expression level. f Broad ELISA binding activity. Representative antibodies (each line) isolated from the four BA.5 BTI donors have their ELISA activities against 18 types of RBD proteins and 14 distinct S proteins calculated and shown as half maximal effective concentration (EC50) values. EC50 value is illustrated by different colors: red, <0.05 μg/ml; orange, 0.05–0.5 μg/ml; yellow, 0.5–5 μg/ml; light green, 5–10 μg/ml; and green, >10 μg/ml. Source data are provided as a Source Data file.
The first four volunteers (donor M, N, O, and P) we recruited have been fully vaccinated by inactivated SARS-CoV-2 WT vaccine and experienced BTI during a wave of BA.5 infections from December 2022 to January 2023 in Shanghai after adjustment of the zero-COVID policy (Fig. 1c and Supplementary Fig. 1c)22. Among these donors, donor P received a fourth booster dose of RBD dimer-based protein vaccine23 prior to the BTI (Fig. 1c and Supplementary Fig. 1c). All four individuals donated their blood within around one month after their BTIs. Their sera exhibited potent S- and RBD-binding capacity and in vitro neutralizing activity (Supplementary Fig. 2 and Supplementary Table 1).
The percentages of double-positive cells in these four donors, as shown by flow cytometry, were around 0.014%, 0.028%, 0.087%, and 0.046%, respectively (Fig. 1d). These sorted single cells were subjected to the single-B cell-based antibody cloning, and in total 122 mAbs (22, 18, 55, and 27 mAbs from each donor, respectively) with naturally paired heavy and light chains were finally amplified. Together, by using heterogenous double-bait proteins, SARS-CoV-1 RBD and XBB RBD, to both of which the donors have never been exposed, we sorted a unique population of B cells and cloned 122 natural human mAbs.
Broad binding capacity against RBDs
To characterize these cloned mAbs, designated as XGi antibodies (Supplementary Table 2), we expressed them and performed ELISA against the two RBD bait proteins. Among the 122 mAbs, 54 (44.3%) mAbs (11/22, 7/18, 18/55, and 18/27 from each donor, respectively) showed binding activity against both RBD bait proteins, SARS-CoV-1 RBD and XBB RBD (the red slices in Fig. 1e). Meanwhile, the remaining isolated mAbs showed low expression level or weak binding activity against one or both of the tested RBD proteins (Fig. 1e).
We further determined the binding breadth of these RBD-binding antibodies against S or RBD proteins of 27 major SARS-CoV-2 variants, including WT, Alpha, Beta, Delta, Kappa, Lambda, Omicron, BA.1, BA.2.12.1, BA.2.38.1, BA.4/5, BF.7, BA.2.75, BQ.1.1, CH.1.1, XBB, XBB.1.5, XBB.1.16, XBB.2.3, EG.5, EG.5.1, HK.3, BA.2.86, BA.2.87.1, JN.1, KP.2 and KP.39,24,25,26. Among the 54 SARS-CoV-1/XBB-binding mAbs, most exhibited broad binding activity (Fig. 1f). Moreover, mAbs ELISA against RBD proteins of SARS-CoV-1, WIV1, SHC014, RaTG13, and Pangolin-GD further demonstrated their broad binding activity against sarbecoviruses (Fig. 1f). Importantly, anti-RBD bbAbs were identified in all of these four donors, such as XGi-048 and XGi-201 from donor M, XGi-091 and XGi-100 from donor N, XGi-106 and XGi-113 from donor O, and XGi-074 and XGi-082 from donor P. These bbAbs exhibited half maximal effective concentration (EC50) values lower than 0.1 μg/ml against most tested antigen proteins (Fig. 1f).
Taken together, the XGi antibodies isolated by a heterogeneous double-bait sorting strategy from BTI donors showed broad RBD-binding capacity against SARS-CoV-2, SARS-CoV-2 sublineages, SARS-CoV-1, and other sarbecoviruses.
Persistence of broadly RBD-binding antibodies after convalescence
Since the first recruited four donors had their blood draw around one month after their BTIs, we then would like to ask whether the memory B cells encoding bbAbs against RBD are persistent in vivo. To do this, we recruited an individual (donor Q) who experienced a BTI during the wave of BA.2 outbreak in spring 2022 in Shanghai27,28 and collected the donor’s PBMCs 10 months after recovery (Fig. 2a and Supplementary Fig. 1c). By using the same heterogenous double-bait sorting strategy, we also identified a small population (0.045%) of double-positive memory B cells recognizing both SARS-CoV-1 RBD and XBB RBD (Fig. 2b). Among the 41 mAbs cloned from these single sorted B cells (Supplementary Table 2), 14 (34.1%) mAbs exhibited binding activity against both SARS-CoV-1 RBD and XBB RBD (the red slice in Fig. 2c). ELISA results showed their broad RBD-binding activity, such as XGi-168, XGi-171 and XGi-185 mAbs (Fig. 2d). Together, these results suggested that the memory B cells encoding bbAbs against SARS-CoV-2 RBD could last for 10 months after convalescence.
a Immunization history for donor Q and R. Donor Q donated her blood ten months after her breakthrough infection of BA.2. Donor R received two doses of inactivated CoronaVac vaccines and experienced no breakthrough infection before blood donation. b Frequency of human B cells recognizing heterogenous bait proteins, PE-conjugated SARS-CoV-1 RBD and APC-conjugated XBB RBD, in donor Q and R. c Initial screening for mAbs (donor Q and R) with binding activity against both SARS-CoV-1 RBD and XBB RBD. Each pie chart represents a donor and the number indicates the isolated mAbs with naturally paired heavy and light chains from that donor. Four categorized mAbs based on ELISAs: red, binders against both SARS-CoV-1 RBD and XBB RBD; orange, antibodies binding only XBB RBD; green, antibodies binding only SARS-CoV-1 RBD; and white, mAbs with weak/no binding activity against both RBDs or mAbs with low expression level. d Broad ELISA binding activity. EC50 values of representative mAbs (each line) isolated from donor Q and R were calculated based on ELISAs against 18 types of RBD proteins and 14 distinct S proteins. The gradient color represents the EC50 values: red, <0.05 μg/ml; orange, 0.05-0.5 μg/ml; yellow, 0.5–5 μg/ml; light green, 5–10 μg/ml; and green, >10 μg/ml. Source data are provided as a Source Data file.
Vaccination-induced broadly RBD-binding antibodies
Anti-RBD bbAbs could be isolated from BTI donors, leading us to investigate whether vaccination only is sufficient to induce such bbAb against RBDs. To answer this question, we investigated a donor (donor R)29 who received only two doses of inactivated vaccines (Fig. 2a and Supplementary Fig. 1c). Flow cytometry using the same B cell sorting strategy showed that the fraction of the double-positive B cells recognizing both SARS-CoV-1 RBD and XBB RBD was around 0.013% in donor R (Fig. 2b), lower than the comparable fractions (0.014-0.087% for donor M-Q) in BTI individuals. Among the 40 mAbs cloned mAbs (Supplementary Table 2), 20 (50%) could bind with the two bait RBD proteins (the red slice in Fig. 2c). Further ELISA showed that these mAbs exhibited broad binding activities against all the tested RBD and S proteins, though their binding potency was lower than those of BTI mAbs (Fig. 2d). Therefore, we concluded that the primary vaccination only was sufficient to induce a variety of broad-spectrum RBD-binding antibodies.
Anti-RBD bbAbs are non-clonally expanded
We then characterized the genetic features of our cloned XGi mAbs by analyzing their immunoglobulin heavy chain V gene (IGHV) and light chain V gene (IGLV) repertoires. Among their IGHV, a preferred IGHV usage was observed, with 34% dominated by IGHV3-7, IGHV3-30, IGHV3-53, IGHV1-18, and IGHV3-48 (Supplementary Fig. 3a). The general IGHV usage frequency for XGi mAbs was similar to that of our previously cloned mAbs from inactivated vaccinated or convalescent individuals by using the dual-color one-bait sorting approach (Supplementary Fig. 3b)29,30. Moreover, a preferential usage of IGHV was also shared among donors (Supplementary Fig. 3c). For example, IGHV3-53 was overrepresented in donors O and P; while IGHV3-7, IGHV1-18, and IGHV3-30 dominated in donors M and N.
We then identified expanded clones of memory B cells encoding closely related XGi antibodies, which shared the same IGHV/IGLV and highly similar CDR3 sequences. In each donor, only a small proportion of XGi mAbs, 0/22 (0%), 3/18 (16.7%), 4/55 (7.3%), 3/27 (11.1%), 0/41 (0%), and 2/40 (5%), respectively, were clonally expanded, while the rest were singlets (Fig. 3a). We then performed a series of comparisons with previously isolated mAbs, including XG2v/XGv mAbs from donors receiving two/three doses of inactivated vaccines29,31, mAbs from donors receiving their two/three doses of mRNA vaccines32,33, and mAbs from convalescent donors 1.3 or 12 months after their recovery34,35. The percentages of clonally expanded XGi mAbs, ranging from 0%-16.7%, were statistically significantly lower than those of previously cloned mAbs (Fig. 3b). For example, 46/84 (54.8%), 78/129 (60.5%), 25/56 (44.6%), and 72/153 (47.1%) XGv antibodies were clonally expanded in each of the four three-dose vaccinees29,36.
a Antibody pie charts for six donors. Each pie chart represents a donor, with the number inside indicating the number of cloned XGi mAbs for this donor. Antibodies with the same IGHV/IGLV usage and closely related CDR3 sequences (see more details in Supplementary Table 2) are grouped together and represented as light purple slice with their percentages shown; while the white slice represents antibody singlets. b Percentage comparisons of antibody clonality. The percentages of antibody clonalities (y-axis) were compared among donors with distinct immunization history. Each dot represents an individual donor. The number above each column indicates the average percentage of antibody clonality. c, d Numbers of somatic nucleotide mutations (y-axis) in the IGHV (c) and IGLV (d) for antibodies from donors with distinct immunization history. Each dot represents a mAb, and the number above each violin column indicates the average number of somatic nucleotide mutations. e, f Amino acid lengths (y-axis) of CDRH3 (e) and CDRL3 (f). Each dot represents a mAb, and the number above each violin column indicates the average amino acid length. Antibodies analyzed includes: XGi mAbs isolated from six donors in this study using heterogeneous double-bait sorting strategy; antibodies isolated from vaccinees after receiving two or three doses of inactivated vaccines29,31; antibodies isolated from mRNA vaccine recipients (two doses of mRNA vaccine, 1.3 months after two doses of mRNA vaccine, 5 months after two doses of mRNA vaccine, and three doses boost vaccination with mRNA vaccine)32,33; antibodies isolated from COVID-19 convalescent donors at 1.3 or 12 months after infection34,35. Data are represented as mean ± SEM. Statistical significance was determined using one-way ANOVA with Dunnett’s multiple comparisons test. Source data are provided as a Source Data file.
Moreover, our selected XGi mAbs (in total 88 among 203 mAbs) with broad RBD-binding activity against both SARS-CoV-1 RBD and XBB RBD exhibited significantly higher levels of IGHV/IGLV nucleotide mutations (Fig. 3c, d) and slightly shorter IGHV CDR3 sequences (Fig. 3e, f).
Taken together, compared with those mAbs previously isolated by using only one type of bait protein, our anti-RBD XGi mAbs identified by using heterogenous double-RBD-bait strategy constituted only a small population among all anti-RBD antibodies in these donors, exhibited broad-spectrum of RBD binding activity, harbored more IGHV/IGLV mutations, and importantly, were barely clonally expanded.
In vitro neutralizing activity
Many previously reported neutralizing antibodies have partially, if not fully, lost their neutralizing capacity against the newly emerged SARS-CoV-2 variants, such as XBB, XBB.1.5, EG.5, BA.2.86, JN.1, KP.2, and KP.39,10,13,24,37,38. To determine the neutralizing activity of our cloned XGi bbAbs, we performed a screening using the pseudovirus-based in vitro neutralization assays against four pseudoviruses, SARS-CoV-1, SARS-CoV-2 WT, BA.2, and XBB (Fig. 4a and Supplementary Fig. 4a). Many of the tested XGi bbAbs exhibited neutralizing activity (IC50 < 10 μg/ml) against each pseudovirus, respectively, but only a few (4 mAbs for SARS-CoV-1, 15 mAbs for SARS-CoV-2 WT, 8 mAbs for BA.2, and 2 mAbs for XBB) showed potent neutralizing activity (IC50 < 0.1 μg/ml) (Supplementary Fig. 4a). In each of the six recruited donors, there are anti-RBD bbAbs broadly neutralizing all four tested pseudoviruses (light purple slices in Fig. 4a).
a In vitro neutralizing activity against pseudotyped SARS-CoV-1, SARS-CoV-2 wildtype (WT), BA.2, and XBB. Each column in the heatmap represents a XGi mAb with broad RBD-binding activity. The gradient color in the heatmap represents IC50 values. The six pie charts below the heatmap correspond to the six donors, and the number of tested antibodies from each donor is indicated inside each pie chart. Antibodies with distinct neutralization breadth are groups into slices of different colors. b In vitro neutralization breadth of the 12 selected XGi antibodies. IC50 values were calculated based on the in vitro neutralization assays against pseudotyped viruses of 13 SARS-CoV-2 variants. c In vitro neutralizing activity against SARS-CoV-1 and SARS-CoV-1/2-related variants, including Pangolin-GD, RaTG13, WIV1 and SHC014. Source data are provided as a Source Data file.
We then selected 12 XGi mAbs and further performed more pseudovirus-based in vitro neutralization assays (Supplementary Fig. 4b, c). Among them, most maintained potent neutralizing activity against Delta and several early Omicron subvariants, such as BA.1, BA.2, BA.4/5 and BF.7 (Fig. 4b). As for the three recent SARS-CoV-2 variants (HK.3, BA.2.86, and JN.1) with remarkable antibody evasion capacity, some XGi mAbs showed drastic reduction of neutralizing activity ( > 10 times compared with the SARS-CoV-2 WT), while others, such as XGi-082, XGi-198, and XGi-199, still neutralized potently (Fig. 4b).
Several XGi bnAbs were identified (Fig. 4b and Supplementary Fig. 4d). XGi-198 and XGi-199 from donor Q exhibited broad neutralizing activity against all 13 tested pseudoviruses of SARS-CoV-2 subvariants, with IC50 values ranging from 0.001-0.009 μg/ml and 0.003-0.268 μg/ml, respectively (Fig. 4b), even though they showed minimal neutralization against SARS-CoV-1 and its related variant WIV1 (IC50 > 10 μg/ml) (Fig. 4c). Meanwhile, XGi-082 (donor P) and XGi-091 (donor N) were highly potent and broad neutralizers, with their IC50 values lower than 1 μg/ml against most of the tested pseudoviruses, including SARS-CoV-1 and several closely-related sarbecoviruses (Fig. 4b, c).
In conclusion, most of our cloned XGi anti-RBD bbAbs were not potent neutralizers, but a few of them exhibited excellent and broad neutralizing activity in vitro.
Three conserved non-overlapping RBD epitopes
The broad RBD-binding activity of our cloned XGi bbAbs suggested that their RBD epitopes were conserved across SARS-CoV-2 variants and even sarbecoviruses, such as SARS-CoV-1. To determine their conserved RBD epitopes, we performed competition ELISAs using 20 representative XGi mAbs. As expected, all of the tested 1st non-biotinylated XGi antibodies efficiently blocked the binding of the 2nd biotinylated version of themselves (yellow rectangles, Fig. 5a). Three groups of non-overlapping epitopes on RBD were identified (red rectangles, Fig. 5a), suggesting three non-competing super-conserved RBD epitopes, SCORE-A, -B and -C.
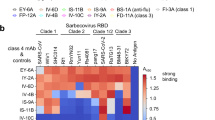
a Competition ELISA among XGi mAbs with broad RBD-binding activity. The antibodies in the x-axis are the 1st non-biotinylated antibody for epitope blocking, while the antibodies in the y-axis are the 2nd biotinylated antibody for signal detection by using streptavidin-HRP. The reference for each 2nd detection antibody used for normalization (100%) is in the right-most lane by using phosphate-buffered saline (PBS) to substitute the 1st blocking antibody. The percentage of 2nd antibody binding: 0-50% in black; 51-75% in gray; and >75% in white. The efficient blocking for all tested XGi mAbs by their own biotinylated versions is marked by yellow rectangles, and three identified non-overlapping XGi antibody groups are marked by red rectangles. b Competition ELISA by using biotinylated S309, SA58, SA55, ADG20, CR3022 or ACE2 as the 2nd antibody for streptavidin-HRP-dependent detection. Three non-competing super-conserved RBD epitopes, SCORE-A, -B, and -C, are labeled.
The non-competing binding capacity between antibodies recognizing distinct SCOREs were further verified by using biolayer interferometry (BLI) competition assays (Supplementary Fig. 5). For example, low levels of competition were observed between XGi-091 (SCORE-A) and XGi-198 (SCORE-B) (Supplementary Fig. 5a), XGi-183 (SCORE-A) and XGi-203 (SCORE-B) (Supplementary Fig. 5b), XGi-203 (SCORE-B) and XGi-171 (SCORE-C) (Supplementary Fig. 5c), and XGi-183 (SCORE-A) and XGi-171 (SCORE-C) (Supplementary Fig. 5d).
Moreover, based on further competition ELISAs, these three identified non-overlapping SCOREs correlated with the epitopes of several well-characterized mAbs, S30939 and SA5816,40 for SCORE-A, SA5516,40 and ADG2041 for SCORE-B, and CR302242 for SCORE-C, respectively (Fig. 5b). Importantly, only the SCORE-B-binding, but not SCORE-A/C-binding, mAbs efficiently blocked the ACE2 binding, highlighting that the RBD/ACE2 interface overlapped with the SCORE-B (Fig. 5b).
Among SCORE-A-binding mAbs, some, such as XGi-091, exhibited potent in vitro neutralizing activity (Fig. 4b, c); while others, such as XGi-183, barely neutralized in our assay (IC50 > 10 μg/ml). Similarly, some SCORE-B-recognizing mAbs, such as XGi-082, XGi-198 and XGi-199, were potent neutralizers (Fig. 4b, c), while others like XGi-203 were not. For SCORE-C-binding mAbs, such as XGi-171 and XGi-185, almost no neutralizing activity was detected in our assay (IC50 > 10 μg/ml).
Taken together, our competition ELISA results suggested three non-competing super-conserved RBD epitopes, SCORE-A, -B and -C, for broad anti-RBD antibody binding.
High-resolution structural characterization of broadly RBD-binding mAbs
To reveal the structural details of the three non-overlapping SCOREs within RBD, we performed structural analysis and first investigated three broadly RBD-binding but not neutralizing mAbs, XGi-183 (SCORE-A), XGi-203 (SCORE-B), and XGi-171 (SCORE-C). We utilized cryo-electron microscopy (Cryo-EM) to determine the high-resolution structures of mAbs in complex with the ectodomain of prefusion-stabilized EG.5.1 S-trimer. Initial negative staining EM revealed that incubation XGi-183 with S-trimer destroyed the S-trimer, whereas XGi-203 and XGi-171 did not. The purified S/mAb complexes for XGi-183 (Supplementary Figs. 6 and 7), XGi-203 (Supplementary Figs. 8 and 9), and XGi-171 (Supplementary Figs. 10 and 11) were used for Cryo-EM structural study.
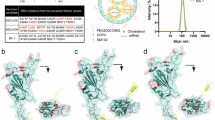
During the cryo-EM data processing of S/XGi-183 (SCORE-A) complex, very few S trimers were observed (Supplementary Fig. 6). The final density map only contains the N-terminal domain (NTD) and RBD of the S protein and the Fab region of XGi-183 (Fig. 6a). The RBD and Fab regions were locally refined, and the resolution was finally improved to be 3.4 Å (Supplementary Fig. 6a). Superimposition of the RBD/XGi-183 complex with the S-trimer in the prefusion state showed that XGi-183 clashed with the NTD of the adjacent protomer, which explains how XGi-183 binding disrupts the integrity of the S-trimer (Supplementary Fig. 7d). To accommodate XGi-183 binding, the RBD undergoes a slight rotation, which likely perturbs the integrity of the prefusion S-trimer. This conformational adjustment may lead to partial opening of the S1 subunit while the S2 region remains intact. Additionally, dimeric complexes were observed in 2D classification (Supplementary Fig. 7e). The XGi-183 binds to the lateral side of the RBD, with a buried surface area (BSA) of 1232 Å2 involving 22 amino acid residues, almost all of which are highly conserved among sarbecoviruses (Fig. 6b and Supplementary Fig. 7c). The paratope of XGi-183 was primarily located on the light chain, creating an extensive network of hydrogen bonds with the RBD. In the framework of 20 hydrogen bonds, T345 demonstrates a robust interaction with the N27 residue on the CDR1 of XGi-183 light chain (Fig. 6b).
a–h Cryo-EM structures of EG.5.1 S trimer in complex with four XGi bbAbs, and their hydrogen bond interactions or salt bridges. Tested mAbs include XGi-183 (a, b), XGi-203 (c, d), XGi-171 (e, f), and XGi-198 (g, h). The hydrogen bond interactions or salt bridges are illustrated as dotted lines. i Epitopes of XGi-183, XGi-203 and XGi-171 on the EG.5.1 RBD.
XGi-183, as a SCORE-A mAb, exhibited a weak neutralizing but broad RBD-binding activity against SARS-CoV-2 variants (Supplementary Fig. 4a). Sequence alignment revealed that the epitope residues targeted by XGi-183 (orange circle in Supplementary Fig. 12) were highly conserved across SARS-CoV-2 variants except in SARS-CoV-1 and certain sarbecoviruses from bat, particularly at A348, N354, R357, K462, T470, and E471 residues (Supplementary Fig. 12). Moreover, XGi-183 had a diminished binding affinity towards BA.2.86, JN.1, KP.2 and KP.3 variants (EC50 > 10 μg/ml in Fig. 2d), likely due to the K356T escape mutation in the S protein (Supplementary Fig. 12), which disrupts the salt bridge between K356 and D57 of XGi-183. Notably, another SCORE-A mAb S309 shares a similar epitope with XGi-183, yet exhibits significantly stronger neutralizing activity. Structural analyses reveal that S309 could stabilize spike in the inactive 1-up-2-down or 3-down conformation via binding to the conserved N343 glycan39,43, probably explaining S309’s superior neutralizing capacity than XGi-183. However, distinct from XGi-183, several XGi mAbs with a similar RBD epitope landscape (SCORE-A), such as XGi-091, XGi-100, XGi-106, and XGi-176, maintained their efficient RBD binding activity against BA.2.86 and its descendant variants (Fig. 1f).
In the S/XGi-203 (SCORE-B) complex, the S-trimer was characterized by two up-RBDs and one down-RBD, each interacting with an XGi-203 Fab (Fig. 6c and Supplementary Fig. 8). The S-trimer/XGi-203-Fab was determined at a resolution of 2.95 Å and the down-RBD/XGi-203 Fab was locally refined to 3.28 Å (Supplementary Fig. 8a). The Fc region was not visible in the final structure due to its intrinsic flexibility. The BSA for RBD/XGi-203 is 781 Å2, encompassing 12 residues (Fig. 6d and Supplementary Fig. 9c). The epitope of XGi-203 was located at the top of the RBD with significant overlap with the RBM region (Supplementary Fig. 9d), and half of the RBD residues for XGi-203 binding (green triangle in Supplementary Fig. 12) were involved in ACE2 interaction. These findings were consistent with our competition ELISA (Fig. 5b) that SCORE-B overlapped with the ACE2-binding epitope on RBD. Interestingly, a di-trimer-like structure was observed in cryo-EM data 2D classification, but not in 3D classification, suggesting weak binding between the trimers (Supplementary Fig. 9e).
Sequence conservation analysis on the XGi-203-binding RBD residues (green triangle in Supplementary Fig. 12) revealed that the Omicron variants harbored unique mutations, including S375F, N440K, and Q498R, while the V445P mutation is predominantly found in XBB and its descendants (Supplementary Fig. 12). However, these mutations failed to decrease the RBD-binding affinity of XGi-203 across different Omicron variants (Fig. 2d). Conversely, XGi-203 lost its efficient binding against the S proteins of SARS-CoV-1 and WIV1 (EC50 > 10 μg/ml in Fig. 2d), possibly due to the presence of mutation residues within its RBD epitope, A372 and R498 in EG.5.1 SARS-CoV-2 (T359 and Y484 in SARS-CoV-1) (Supplementary Fig. 12). Since N439/K440 formed strong hydrogen bonds with the Y66 residue of IGHV CDR2 of XGi-203, and R498, along with P445, contributed to a hydrophobic pocket involving V112/N114 of IGHV CDR3 as well as N109/L112 of IGLV CDR3, the corresponding RBD substitutions in SARS-CoV-1 and WIV1 might potentially disrupt both hydrogen bonding and hydrophobic interaction essential for XGi-203 binding.
The structure of the S-trimer/XGi-171 (SCORE-C) complex was resolved to a resolution of 3.18 Å (Supplementary Fig. 10), with each one of up RBDs interacting with one XGi-171 Fab (Fig. 6e). A locally refined structure of the up-RBD in complex with the XGi-171 Fab was determined to a resolution of 3.65 Å (Supplementary Fig. 10a). The epitope for XGi-171 (blue square in Supplementary Fig. 12) resided within a cryptic region of RBD, permitting its exclusive binding to the up-RBD conformation. The interaction between XGi-171 and the RBD buried a surface area of 781 Å2 and involves 15 RBD residues (Fig. 6f and Supplementary Fig. 11c), all of which were highly conserved across sarbecoviruses except for P384 and T385 (Supplementary Fig. 12). P384 and T385, alongside residues Y369 and S383, were involved in hydrophobic interactions with Y66 of IGHV CDR2, W113 of IGHV CDR3, and M113 of IGLV CDR3. Therefore, due to the high level of sequence conservation of XGi-171-binding epitope across various sarbecoviruses, XGi-171 showed ultra-broad RBD binding activity for all the tested RBD or S proteins of SARS-CoV-2 variants as well as sarbecoviruses (Fig. 2d).
Moreover, the binding of XGi-171 induces an outward rotation of the up-RBD by approximately 16° compared to the apo state of the S-trimer (PDB: 7WVP) (Supplementary Fig. 11e), which might lead to the instability and disassembly of the S-trimer. Consistent with our speculation, we indeed observed a small portion of depolymerized S trimers in the 2D classification (Supplementary Fig. 11f) and a weak neutralizing activity against SARS-CoV-2 WT in vitro (Supplementary Fig. 4a). These observations suggested that, given sufficient time, XGi-171 might promote the dissociation of S-trimer, implying its slow-acting neutralizing activity. Notably, the di-trimer-like structure was observed in negative staining EM and cryo-EM 2D classification (Supplementary Fig. 11a, b and f), suggesting disruption of the di-trimer conformation during the freezing process. Interestingly, another SCORE-C mAb, CR3022, exhibits weak neutralizing activity as XGi-171. Similar to XGi-171, CR3022 binds a cryptic epitope adjacent to the RBD core, inducing conformational flexibility that accelerates Fab-antigen disengagement (half-life <5 min)44. Similarly, CR3022 binding is made possible by the dissociation of the S-trimer into monomers45. This mechanistic parallel supports the model whereby cryptic-epitope antibodies exert weak neutralization primarily through allosteric destabilization rather than direct RBD blockade.
In contrast to the aforementioned mAbs with weak neutralizing capacity, XGi-198, a SCORE-B mAb, exhibited a very broad and potent neutralizing activity (IC50 < 0.01 μg/ml) against all of the tested SARS-CoV-2 variants, including the recent BA.2.86 and JN.1 variants (Fig. 4b). The S/XGi-198 (SCORE-B) complex structure showed that XGi-198 cross-linked two S-trimers into a di-trimer (Fig. 6g and Supplementary Fig. 13). Using masks, two S-trimers were individually refined to the resolutions of 2.98 Å and 3.43 Å, respectively, while the up-RBD associated with the XGi-198 Fab was locally refined to a resolution of 3.24 Å (Supplementary Fig. 13a). Both S-trimers exhibited a conformation characterized by two up-RBDs and one down-RBD, with each RBD engaging an XGi-198 Fab (Fig. 6g).
The RBD epitope of XGi-198 was SCORE-B, similar to that of XGi-203, with both being primarily localized within the RBM region and competing for ACE2 binding (Fig. 6h and Supplementary Fig. 14c). The BSA for RBD/XGi-198 was 681 Å2, involving 13 residues (pink triangle in Supplementary Fig. 12), ten of which were also recognized by XGi-203 (green triangle in Supplementary Fig. 12). Moreover, three residues (P373, H505, and Q506) on the XGi-198 epitope were not recognized by XGi-203, while two residues (A372 and F375) within the XGi-203 epitope were not recognized by XGi-198. Among these residues, Q506 was highly conserved across all analyzed sarbecoviruses, and played a crucial role in stabilizing RBD/XGi-198 interaction by facilitating hydrogen bonding and hydrophobic interactions. This difference on Q506 binding might explain why XGi-198, but not XGi-203, could rapidly crosslink two S trimers to form one di-trimer, providing an explanation for the potent in vitro neutralizing activity of XGi-198.
Cryo-EM and negative staining EM analyses showed that XGi-198, but not XGi-203, efficiently induced di-trimer formation (Supplementary Fig. 19a). Mechanistically, three XGi-198 Fabs binding on one S-trimer establish a stronger cooperative network with three additional interfaces: one of up-RBD-bound XGi-198 Fab forms a 317 Å2 interface with a neighboring down-RBD; while the down-RBD-bound XGi-198 Fab bridges two neighboring up-RBDs with interfaces of 580 Å2 and 348 Å2. Collectively, these additional interactions stabilize the spike in a rigid 2-up-1-down (2u1d) conformation (Supplementary Fig. 19b). In contrast, similar but weaker interactions were observed in XGi-203, explaining its reduced cross-linking capacity. Moreover, functional studies of Fab fragments supported this cryo-EM observation. Both Fab fragments exhibited significant reductions in binding affinity (Supplementary Fig. 19c) and neutralizing activity (Supplementary Fig. 19d) compared to their full-length IgG counterparts. However, XGi-198 exhibited more dramatic reductions of neutralization (70-100-fold for WT, BA.5 and XBB) than XGi-203 (2-54-fold) (Supplementary Fig. 19e), suggesting that XGi-198’s potent neutralization critically depends on efficient cross-linking of S-trimer.
Neither XGi-198 nor XGi-203 exhibited detectable binding to SARS-CoV-1 or WIV-1. To explain this, we compared them with SA55, a mAb with a similar epitope yet exhibiting potent SARS-CoV-1 neutralization (Supplementary Fig. 12). Detailed structural analysis showed that SA55 engaged a cluster of residues highly conserved among SARS-CoV-2, SARS-CoV-1 and WIV-1, including F374, S375, T376, D405, V407, R408, N437, G502, G504, and Y508. In contrast, XGi-198 and XGi-203 interacted with fewer conserved residues (N437, N440, T500, G502, Y505, and Q506 for XGi-198; and S375, N437, N440, T500, and G502 for XGi-203), probably accounting for their weaker binding against SARS-CoV-1 and WIV-1.
In addition to the Cryo-EM structural analysis of four representative mAbs, we also used AlphaFold 3 and performed structure prediction of 23 more XGi mAbs in complex with EG.5 RBD (Supplementary Fig. 15). As shown, the epitopes of seven XGi antibodies (XGi-061, XGi-091, XGi-092, XGi-100, XGi-106, XGi-112 and XGi-176) were mainly overlapped with SCORE-A; fourteen antibodies (XGi-072, XGi-073, XGi-074, XGi-075, XGi-076, XGi-079, XGi-082, XGi-127, XGi-129, XGi-182, XGi-199, XGi-200, XGi-201 and XGi-202) had their epitopes overlapping with SCORE-B; and the rest two of them (XGi-168 and XGi-185) possessed epitopes resembling SCORE-C. These prediction results were consistent with our epitope mapping data determined by competition ELISAs (Fig. 5a and b).
Collectively, XGi-183 (SCORE-A) targeted an epitope on the lateral side of the RBD, disrupting the integrity of the S trimer. XGi-203 (SCORE-B) bound to an epitope overlapping with the RBM, effectively competing for the ACE2 interaction. XGi-171 (SCORE-C) interacted with a cryptic region of the RBD, binding exclusively to the up-RBD conformation. Additionally, XGi-198 (SCORE-B) exhibited potent and broad neutralizing activity, potentially due to its unique ability to robustly cross-link two S-trimers into a di-trimer. Taken together, by studying the cryo-EM structures of four representative bbAbs and predicting 23 antigen-antibody structures by AlphaFold 3, we preliminarily characterized the three non-overlapping highly conserved RBD epitopes, SCORE-A, -B, and -C.
Unbalanced distribution of the three super-conserved RBD epitopes (SCOREs)
To further investigate the identified three non-overlapping SCORE epitopes and deeply explain their conserved molecular basis for broad antibody binding across variants, we first analyzed the binding modes and specificities of several more SCORE-A/B/C-binding mAbs and compared their binding footprints with the level of sequence conservation of their RBD epitopes (Fig. 7a–e).
a Footprints of representative antibodies targeting SCORE-A (RBD-5B). For comparison, the antibody footprints are superimposed onto the same RBD (PDB 6W41) and shown with the same view and angle. On the RBD surface, the SCORE-A is outlined by green lines, and the RBD residues are highlighted according to their degree of sequence conservation among 30 SARS-CoV-2 variants and sarbecoviruses (Supplementary Fig. 12) on a scale from red (highly conserved) to white (weakly conserved). b Footprints of representative antibodies targeting SCORE-B (RBD-3). c Footprint of viral receptor ACE2 on an RBD surface plot. d Footprints of representative antibodies targeting SCORE-C (RBD-7). e Representative mAbs targeting the three super-conserved RBD epitopes, SCORE-A, -B, and -C. XGi-183 (SCORE-A), XGi-203 (SCORE-B), and XGi-171 (SCORE-C) exhibit three different and non-overlapping binding orientations. f–i Footprints of representative antibodies targeting RBD-1, RBD-2 (f), RBD-4, RBD-5A (g), RBD-6 (h), and RBD-8 (i). Each RBD epitope is illustrated by green lines on the RBD. Sequence conservation level of each RBD residue is highlighted based on 30 SARS-CoV-2 variants and sarbecoviruses (Supplementary Fig. 12) on a scale from red (highly conserved) to white (weakly conserved). j Graph-based unsupervised clustering of a series of previously reported mAbs and XGi bbAbs from this study in a variant-dependent manner. Each dot represents an antibody, with distinct RBD epitopes marked by different colors. As new mutations emerge from WT to Omicron subvariants, BA.4/5, XBB, and JN.1, antibodies were gradually evaded, and only three remaining RBD epitopes are still conserved and highly immunogenic. k Graphic summary of RBD epitope landscape. A two-dimensional projection of RBD surface (produced by RIVEM) showed nine RBD epitopes. Based on their immunogenicity (evaluated based on the prevalence of elicited mAbs) and sequence conservation, these epitopes could be into four types: 1) highly conserved and highly immunogenic (RBD-5B and RBD-7); 2) partially mutated but structurally conserved for mAb binding, and highly immunogenic (RBD-3, overlapping with ACE2 binding motif); 3) highly mutated epitopes (RBD-1, RBD-2, RBD-4 and RBD-5A); and 4) highly conserved but lowly immunogenic (RBD-6 and RBD-8).
Consistent with our competition ELISA results (Fig. 5a, b), two previously well-characterized mAbs, S30939 and SA5840, had their epitope footprints overlapped with SCORE-A (Fig. 7a). Their RBD epitopes were characterized as RBD-5B10. Comparing the RBD residues for XGi-183 (orange circle in Supplementary Fig. 12), SA58 (purple circle in Supplementary Fig. 12) and S309 (deep blue circle in Supplementary Fig. 12) revealed two antigenic residues, E340 and T345, were shared and highly conserved across SARS-CoV-2 variants. Combined with the AlphaFold 3-predicted RBD epitopes (Supplementary Fig. 15a) and ELISA results (Fig. 1f and Fig. 2d) for other SCORE-A-binding mAbs, such as XGi-091, XGi-100, XGi-106, and XGi-176, we concluded that the SCORE-A was a relatively conserved region on RBD (Fig. 7a), and that SCORE-A mAbs were commonly induced after SARS-CoV-2 infection or vaccination and exhibited broad binding activity against RBDs of numerous SARS-CoV-2 variants.
Our cloned SCORE-B-binding mAbs, such as XGi-198 and XGi-203, shared a similar RBD epitope and approach angle with a previously isolated mAbs, SA5540, which recognized the RBD-3 epitope (Fig. 7b). In contrast to XGi-198 and XGi-203, monoclonals SA55 and several other SCORE-B mAbs, such as XGi-082, XGi-127, and XGi-201 maintained their potent binding affinity against SARS-CoV-1 RBD. Comparison among XGi-198 (pink triangle in Supplementary Fig. 12), XGi-203 (green triangle in Supplementary Fig. 12) and SA55 (red triangle in Supplementary Fig. 12) revealed shared binding residues on RBD, including N437, N439, P499, N501, G502, and V503, most of which were highly conserved across SARS-CoV-2 variants, and also participated in ACE2 binding (Supplementary Fig. 12). However, according to the epitope footprints of SCORE-B-binding mAbs (Fig. 7b and Supplementary Fig. 15b), the amino acid residues within SCORE-B were only partially conserved (Fig. 7b). Some residues such as N440, V445, Q498, and N501 were highly mutated across SARS-CoV-2 variants and sarbecoviruses (Supplementary Fig. 12). This seemed to resemble the situation of ACE2 binding, during which the SCORE-B residues were highly mutated but perfectly maintained their binding affinity with ACE2 throughout all SARS-CoV-2 variants and all sarbecoviruses (Fig. 7c)46. Therefore, we concluded that, despite being highly mutated, the SCORE-B maintained its structural conservation for ACE2 binding and also for broad antibody recognition.
Besides our cloned XGi-171, XGi-168, and XGi-185, a well-characterized neutralizing antibody targeting RBDs of both SARS-CoV-1 and -2, CR302247, also showed ELISA competition and had its epitope footprint overlapped with SCORE-C (Figs. 5b and 7d). The epitope of CR3022 (yellow square in Supplementary Fig. 12) was highly overlapped with that of XGi-171 (blue square in Supplementary Fig. 12), and was also highly conserved, such as residues Y369, F374, F377, C379, Y380, G381, V382, S383, K386, D427, D428, and F429. Structural comparisons showed the SCORE-C was on the inner RBD surface with no overlap with the ACE2 binding site (Supplementary Fig. 11d). Protein sequence conservation analysis suggested that the amino acid residues within SCORE-C were highly conserved (Fig. 7d), which perfectly explained why the SCORE-C-binding mAbs showed ultra-broad binding affinity against SARS-CoV-2 variants and even sarbecoviruses (Figs. 1f and 2d).
RBD could induce highly potent neutralizing antibodies48,49. On the surface of RBD, many distinct non-overlapping epitopes have been reported50,51. To understand why our cloned XGi mAbs with broad RBD-binding activity mainly targeted these three RBD epitopes (SCORE-A, -B, and -C), we characterized the binding footprints of a series of representative mAbs targeting the other six RBD epitopes (Supplementary Fig. 16) and compared their RBD-binding breadth and epitope sequence conversation level.
A series of previously isolated mAbs targeting RBD-1 (CB6, BD-704, S2K146 and CT-P59 in Fig. 7f) and RBD-2 (XGv347, XGv051 and REGN10933 in Fig. 7f) drastically lost their binding activity against almost all Omicron subvariants (Supplementary Fig. 16). This could be explained by the fact that both RBD-1 and RBD-2 epitopes overlapped with RBM and showed high mutation frequencies and low levels of sequence conservation (Fig. 7f). For example, monoclonal CB6 recognizing RBD-1 showed no binding affinity with BA.2 S protein due to its K417N and Q493R mutations.
Similarly, RBD-4-targeting mAbs (P2B-2F6 and XGv-282 in Fig. 7g) and RBD-5A-targeting mAbs (LY-CoV1404, XG005, REGN10987, XGv264, XGv265, COV2-2130 in Fig. 7g) were evaded by BQ.1.1 and XBB, consistent with the fact that both RBD-4 and RBD-5A epitopes covered amino acid residues highly mutated in Omicron subvariants (Fig. 7g). For example, R346T and V445P substitutions on BQ.1.1 and XBB partially evaded the antibody binding of XGv282 and LY-CoV1404.
In contrast, RBD-6-binding mAbs (COVA1-16, G32Q4, and DH1047 in Fig. 7h) and RBD-8-binding mAbs (BIOLS56, IMCAS74, and S2H97 in Fig. 7i) retained their binding to the RBDs of Omicron subvariants. Adjacent to the SCORE-C (RBD-7 epitope), both RBD-6 and RBD-8 epitopes were also on the inner RBD surface and the amino acid residues within these epitopes were highly conserved (Fig. 7h, i). However, in our study, we did not isolate any XGi mAbs targeting RBD-6 and RBD-8, and even the number of previously isolated RBD-6- and RBD-8-targeting mAbs was very few10. Moreover, we found some amino acid residues on the surface of RBD, such as R355, K378, C379, Y396, R466, S514, F515, and L518, which were highly conserved across SARS-CoV-2 variants and even sarbecoviruses but did not participate in any antibody recognition (Supplementary Fig. 17). These observations implied a low level of immunogenicity of some highly conserved RBD residues, such as RBD-6 and RBD-8, resulting in a less efficient selection of corresponding B cells from repertoire during antibody maturation.
In summary, as new variants emerged, the RBD-binding antibodies were gradually evaded (Fig. 7j), especially for mAbs targeting RBD-1, RBD-2, RBD-4, and RBD-5A epitopes due to the low sequence conservation of these epitopes (Fig. 7k). However, although new RBD epitopes appeared and induced Omicron subvariant-specific RBD-binding antibodies, there were still three remaining super-conserved RBD epitopes to induce a broad antibody response from the original WT to the most recent KP.2/3 variants. These RBD epitopes included two highly sequence-conserved epitopes, RBD-5A and RBD-7, and one partially mutated but structurally conserved RBD epitope (overlapped with the ACE2-binding motif), RBD-3 (Fig. 7j and k). Meanwhile, RBD-6, RBD-8 and some other RBD residues were highly conserved but exhibited low immunogenicity (Fig. 7k).
In vivo protection by broadly RBD-binding mAbs
To determine whether our isolated anti-RBD bbAbs could prevent infection in vivo, we utilized the Syrian golden hamster model and injected them with a single XGi antibody or only phosphate-buffered saline (PBS) as a negative control before infection with SARS-CoV-2 Omicron EG.5.1 authentic virus (Fig. 8a, b). Four days after virus inoculation, the hamster lung tissue and nasal turbinate were harvested to measure the viral subgenomic E (sgE) transcript levels and infectious virus titers by using quantitative reverse transcription polymerase chain reaction (qRT-PCR) and 50% tissue culture infectious dose (TCID50) assay, respectively (Fig. 8c, d).
a Diagram of prophylaxis protocols for male golden Syrian hamsters intranasally (i.n.) challenged with Omicron EG.5.1 eight h after antibody injection (20 mg/kg for each mAb) through intraperitoneal injection (i.p.). Nasal turbinate (NT) and lung tissues were collected at day 4 post-viral challenge to measure the viral load and viral antigen expression. b List of five broadly RBD-binding XGi antibodies tested against Omicron EG.5.1 in golden Syrian hamsters. c, d SARS-CoV-2 viral sgE gene copies and infectious virus titers (n = 5). Quantitative reverse transcription polymerase chain reaction (qRT-PCR) was used to quantify Omicron EG.5.1 viral mRNA load (c), while the infectious virus titer was determined by TCID50 assay (d). e Immunofluorescence staining for viral nucleocapsid (N) protein (n = 3). SARS-CoV-2 N protein in hamster lungs was stained with the specific antibody and visualized under fluorescence microscopy. Cell nuclei are stained with DAPI. Scale bar, 200 μm. Bar graph represents fluorescent intensity of viral N protein expression quantified using ImageJ. f Histopathological staining of hamster lungs after mono-prophylaxis (n = 5). Representative manifestation of lung damages including multi-focal inflammation, alveolar wall congestion, proteinaceous exudation, epithelial damage, and alveolar wall collapse were denoted by different symbols as indicated. Scale bar, 200 μm. Bar graph represents lung pathological changes quantified by a semi-quantitative histology scoring system as previously reported64,65. Each data point represents one biological repeat. Data represents mean ± SEM from the indicated number of biological repeats. Statistical significance was determined using one-way ANOVA with Dunnett’s multiple comparisons test. Source data are provided as a Source Data file.
We first tested two bnAbs, XGi-082 and XGi-198, both recognizing SCORE-B and exhibiting potent and broad neutralizing activity, with their IC50 values ranging from 0.007-0.712 μg/ml and 0.001-0.009 μg/ml, respectively (Fig. 4b). Compared with the PBS control, pre-exposure prophylaxis with either XGi-082 or XGi-198 significantly decreased the viral sgE expression and infectious virus titer in hamster lungs and nasal turbinates (Fig. 8c, d). Especially, XGi-198 robustly suppressed the viral replication in vivo, with two logs lower viral sgE expression in lung and one log lower viral sgE expression in nasal turbinate compared with those of XGi-082 (Fig. 8c).
To determine whether the bbAbs but not bnAbs could also provide protection against infections in vivo, we chose three XGi antibodies, XGi-183 (SCORE-A), XGi-203 (SCORE-B), and XGi-185 (SCORE-C) and evaluated their four-day outcome for mono-prophylaxis. The five hamsters that received XGi-183 showed reduced viral sgE expression and infectious virus titers in lung of hamsters (Fig. 8c, d). Similar results were observed in hamsters receiving XGi-203 (Fig. 8c, d). However, animals that received XGi-185 mAb had comparable levels of viral sgE and infectious virus as negative control of PBS only (Fig. 8c, d). Together, these results suggested that broadly binding mAbs, XGi-183 and XGi-203, but not XGi-185, could provide antiviral effect against Omicron EG.5.1 in vivo.
We further performed immunofluorescence staining of the infected hamster lungs against the viral nucleocapsid (N) protein and showed that all tested XGi antibodies, except XGi-203 and XGi-185, significantly reduced the level of the viral N protein in hamster lungs compared with the PBS control (Fig. 8e). The expression of viral N protein was marginally detected in the XGi-198-treated lung samples (Fig. 8e), further confirming the robust viral suppression capacity of XGi-198.
In addition, Hematoxylin and Eosin (H&E) staining was conducted to reveal the histopathological changes in hamster lungs. In the PBS controls, typical virus-induced lung damages were observed, including multifocal inflammation, collapse or congestion of the alveolar wall, proteinaceous exudation in the alveolar cavity, and epithelial damage in small bronchioles (Fig. 8f). On the contrary, in mono-prophylaxis groups (except XGi-185), these histopathological changes in hamster lungs were significantly attenuated (Fig. 8f).
Combined with the results that these mAbs showed low polyreactivity and autoreactivity (Supplementary Fig. 18), we concluded that our cloned XGi mAbs with broad RBD-binding activity could confer antiviral protection in vivo and reduce virus-induced lung damages, regardless of their neutralizing capacities.
Discussion
Although the World Health Organization (WHO) has declared the end of COVID-19 pandemic, SARS-CoV-2 will be likely to persist for a long time and continue to evolve with new escape mutations in the foreseeable future, and the infections and diseases caused by the newly emerging SARS-CoV-2 variants will still be a global threat. A realistic question to be answered is that whether the population who have been exposed to either SARS-CoV-2 vaccination or infection or both have already developed mAbs targeting the future variants. For example, could the individuals experienced BA.5 BTI develop anti-XBB, or even anti-JN.1, antibodies? In this study, we used SARS-CoV-1 RBD and XBB RBD as bait proteins, and isolated a minority of antibodies which exhibited broad RBD binding activity against SARS-CoV-2 variants, including the newly emerged variants, such as EG.5.1, BA.2.86, JN.1, and KP.2/3. Since the donors had never been exposed to these newly emerged Omicron variants at the time of the blood donation, our in vitro and in vivo neutralization results emphasized that our adaptive immune system could develop rare mAbs that bind, neutralize, and protect against future variants.
Another question to be answered is more fundamental. For the most recently emerged Omicron subvariants, such as KP.2 and KP.3, over 60 mutations have accumulated in their S proteins, and around half are located on RBD. Are there any RBD epitopes remaining conserved across SARS-CoV-2 variants? In our study, the characterization of our cloned XGi anti-RBD bbAbs revealed three non-overlapping RBD epitopes, two (SCORE-A and SCORE-C) with highly conserved amino acid residues, and one (SCORE-B) with partially mutated RBD residues but conserved epitope structure for antibody binding. Moreover, other RBD epitopes were either highly mutated for broad antibody recognition or less immunogenic to induce an antibody response. Therefore, these findings help us to annotate various RBD epitopes for their conservation across SARS-CoV-2 variants and even sarbecoviruses, and for their immunogenicity to induce antibody recognition.
Our identified XGi mAbs with broad RBD binding activity were rare and, importantly, non-clonally expanded (not sharing the same IGHV/IGLV and highly similar CDR3 sequences). Moreover, these XGi mAbs had higher levels of somatic hypermutation on IGHV and IGLV, implying the crucial role of some key IGHV/IGLV mutations for their breadth of RBD-binding activity. Therefore, we speculate that the mAbs encoded by the sister B cells (cells originating from the same progenitor) of our isolated XGi-encoding B cells showed no broad RBD binding activity.
As many clinically developed anti-SARS-CoV-2 mAbs have been evaded, new bnAbs are urgently needed for prevention and treatment. To examine the neutralizing activity against all major SARS-CoV-2 one mAb by one mAb is obviously time consuming and resource-wasting. Therefore, how to efficiently isolate and screen for a bnAb neutralizing the existing major SARS-CoV-2 variants and the potential future variants? In a previous study, we proposed to use the cross-neutralization against SARS-CoV-1 and SARS-CoV-2 Omicron as criteria for the initial screening for bNAbs21. Following this proposed strategy, instead of using only one type of bait S protein (two distinct fluorescent dyes were usually coupled to eliminate the B cells binding fluorescent dyes and to increase the specificity) as our previous studies have performed29,30,33,35, we adopted a heterogenous double-bait sorting strategy in this study, using PE-conjugated SARS-CoV-1 RBD and APC-conjugated XBB RBD. Finally, a series of bnAbs, such as XGi-082, XGi-091, XGi-198, and XGi-199, were successfully identified, proving this strategy works.
On the S protein, many epitopes have been identified and characterized. Many antibodies have been evaded because their recognized epitopes are highly mutated. However, on the other hand, many mAbs targeting highly conserved regions and showing broad S-binding activity are not potent neutralizers. For example, many anti-S bbAbs target highly conserved SCORE-A, or SCORE-C, or “cold spots” outside of RBD (such as subdomain-1, fusion peptide, and stem helix) exhibited broad S-binding activity but weak neutralizing capacity52,53,54. Therefore, in this study, identifying and characterizing the RBD epitope, such as SCORE-B, that are highly conserved and also capable of inducing potent neutralizing mAbs improved our understanding about SARS-CoV-2 evolution and shed light on better evaluation and further development of broad vaccines.
Methods
Volunteer recruitment and blood collection
Volunteer recruitment and blood draws were performed at the Huashan Hospital of Fudan University, and all experiments related to human samples were performed at Fudan University and approved by the institutional Research Ethics Committee (approval number: 2022-C005). Six donors in total were recruited in this study, and all participants voluntarily consented to take part in a blood draw and a subsequent research study. Donor M, N, and O had received three doses of inactivated COVID-19 vaccine, Sinopharm (Vero Cell), manufactured by Wuhan Institute of Biological Products, and experienced BA.5 breakthrough infection in Shanghai after China eased a series of stringent restrictions at the end of 202222,55. Besides the three doses of inactivated COVID-19 vaccine, Donor P received the fourth dose of RBD dimer-based protein vaccine produced by Anhui Zhifei Longcom Biopharmaceutical23. Donor Q had been vaccinated by three doses of the inactivated COVID-19 vaccine, CoronaVac (Vero Cell), manufactured by Sinovac Biotech, prior to the breakthrough infection during the Omicron BA.2 outbreak in spring 2022 in Shanghai27,28. Donor R received only two doses the inactivated COVID-19 vaccine, CoronaVac (Vero Cell) and donated blood within one month after vaccination29. No serum screening for cross-reactivity or cross-neutralization was performed for the selection of these donors. After blood draw, human peripheral blood mononuclear cells (PBMCs) were isolated by using a cell separation tube with frit barrier as previously reported56. The isolated PBMCs were resuspended in 90% heat-inactivated fetal bovine serum (FBS) supplemented with 10% dimethyl sulfoxide (DMSO) and then cryopreserved in liquid nitrogen.
Protein expression and purification
The expression plasmids were constructed for the expression of S ectodomain (residues 14-1208) and RBD (residues 319-541) proteins of SARS-CoV-1, SARS-CoV-2, and their related variants. To enhance the conformation stability of S trimer, the proline substitutions at 817, 892, 899, 942, 986, and 987, ‘GSAS’ substitutions at the S1/S2 furin cleavage site (residues 682-685), and a C-terminal T4 fibritin trimerization domain (foldon) were introduced in the S constructs. For protein expression, the corresponding plasmids were transfected into human embryonic kidney 293 F (HEK293F) cells, which were cultured with serum-free OPM-293-CD05 medium (OPM Biosciences) at 37 °C in 5% CO2 with shaking at 100 rpm. The cell supernatants were collected five days post-transfection, and further purified by affinity chromatography29. The purified proteins were used in ELISAs, single B cell sorting experiments, and cryo-EM studies.
Bait protein preparation and single B cell sorting
Avi-tagged SARS-CoV-1 and XBB RBD expressed in HEK293F cells were biotinylated using BirA Biotin-Protein Ligase kit (Avidity). The excess of unbound biotin was removed by using a Zeba Spin Desalting column (Thermo Fisher Scientific) according to the manufacturer’s instructions. To prepare the fluorophore-conjugated bait proteins for single B cell sorting, the bait proteins of SARS-CoV-1 RBD and XBB RBD were prepared by incubating biotinylated RBD proteins with streptavidin-phycoerythrin (PE) (eBioscience) or streptavidin-allophycocyanin (APC) (BD Biosciences), respectively.
Human B cell purification and the following single B cell sorting by bait proteins conjugated with two distinct fluorescent dyes were performed as previously described30,56. Briefly, B lymphocytes were selected from PBMCs with CD19 MicroBeads (Miltenyi Biotec). Sequential incubation at 4 °C with human Fc block (BD Biosciences), bait protein-PE/APC, and anti-CD20-PECy7 (BD Biosciences) was performed, followed by the single-cell sorting of CD20-PECy7+ SARS-RBD-PE+ XBB-RBD-APC+ B cells into 96-well plates using an Aria II (BD Biosciences). The sorted single B cells were stored at -80 °C.
Antibody cloning, sequencing and production
Antibody cloning from the sorted single cells and the production of mAbs were performed as previously reported30,56. The amplified PCR products from each single B cell were loaded onto 2% agarose gel for electrophoresis and purified for Sanger sequencing. All the sequencing results of heavy and kappa/lambda light chains were analyzed by IMGT/V-QUEST57 and IgBlast58, and the V(D)J gene segment and CDR3 sequences of each antibody were determined. All cloned antibodies were subjected to vector construction and antibody expression as previously described59. Besides our cloned antibodies, a broad and potent neutralizing antibody to HIV-1 as a negative control, 3BNC11760, and several previously reported anti-SARS-CoV-2 RBD mAbs, including SA55, SA5816,40, BD57-012916, CR302242, S30939, ADG2041, XGv289, XGv347, XGv405, and XGv42129,36 were also prepared by transient transfection of HEK293F cells. The HEK293F cells were cultured with HEK293 serum-free OPM-293-CD05 medium (OPM Biosciences) at 37 °C in 5% CO2 with shaking at 100 rpm. After seven days, the cultured medium was harvested, and the secreted antibodies were purified by using Protein G Sepharose 4 Fast Flow resin (Cytiva) and further verified by using Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE).
ELISA and competition ELISA
For ELISA, the binding activities of the recombinant IgG antibodies against RBD or S proteins of SARS-CoV-1, SARS-CoV-2 and their related variants were measured as previously reported30. Briefly, plates were coated with antigen proteins (10 μg/ml) in phosphate-buffered saline (PBS) overnight at 4 °C, and then blocked with 3% bovine serum albumin (BSA) in PBS. The three-fold serially diluted antibody with a maximum concentration of 10 μg/ml was added for incubation for one h at room temperature. Visualization was carried out by using horseradish peroxidase (HRP)-conjugated goat anti-human IgG (Thermo Fisher Scientific). The EC50 and area under the curve (AUC) values were calculated for each antibody by using PRISM software to evaluate the antigen-binding capacity.
For competition ELISA, plates were coated with 2 μg/ml SARS-CoV-2 RBD and then blocked with 3% BSA in PBS. Plates were incubated with 15 μg/ml 1st blocking antibody (60 μg/ml for CR3022) for two h, and afterwards 0.25 μg/ml biotinylated 2nd antibodies (15 μg/ml for CR3022) were directly added for half an h incubation at room temperature. Detection was performed using streptavidin-HRP (BD Biosciences). PBS buffer replacing the 1st blocking antibody was used as a reference for normalization, while the anti-HIV antibody 3BNC11760 with no blocking capacity served as a negative control.
Preparation of SARS-CoV-2 and SARS-CoV-1 pseudotyped viruses
The pseudotyped viruses were produced as previously reported61. Briefly, plasmids encoding the S proteins of SARS-CoV-2, SARS-CoV-1, and their related variants were synthesized. Expi293F cells were cultured to a density of 3 \(\times\) 106/mL before transfection with the specified S gene using polyethylenimine. Cells were then incubated overnight at 37 °C with 8% CO2. VSV-G pseudo-typed ΔG-luciferase (G*ΔG-luciferase, Kerafast) was utilized to infect the cells in DMEM at a multiplicity of infection of 5 for 4 h, followed by three washes with PBS/2% FBS. After 24 h, the transfection supernatant was collected and clarified by centrifugation. Each viral stock was subsequently incubated with 20% I1 hybridoma (anti-VSV-G; CRL-2700, ATCC) supernatant for one h at 37 °C to neutralize the contaminating VSV-G pseudotyped ΔG-luciferase viruses before titration and aliquoting for storage at -80 °C.
In vitro neutralization assay using pseudotyped viruses
Neutralization assays were performed by incubating pseudoviruses with serial dilutions of mAbs and assessing the reduction in luciferase gene expression, as previously described62. Briefly, Vero-E6 cells were seeded in a 96-well plate at a concentration of 2 \(\times\) 104 cells per well. After 24 h, pseudoviruses were incubated with serial dilutions of the specified mAbs in triplicate for 30 minutes at 37 °C. The antibody-pseudovirus mixture was then added to cultured cells and incubated for an additional 24 h. Subsequently, the luminescence was measured using the Luciferase Assay System (RG062M, Beyotime). IC50 value was defined as the dilution at which the relative light units (after subtraction of the background in the virus-free cell-only wells) were reduced by 50% compared to the virus-only control wells. IC50 values were calculated using nonlinear regression in GraphPad Prism.
Authentic SARS-CoV-2 virus and biosafety
SARS-CoV-2 Omicron EG.5.1 (GISAID: EPI_ISL_18461518) strain was isolated from the respiratory tract specimens of laboratory-confirmed COVID-19 patients in Hong Kong. Omicron EG.5.1 was cultured and titrated by performing plaque assay. All experiments, including authentic virus, were performed according to the approved standard operating procedures of the Biosafety Level 3 facility at the Department of Microbiology, the University of Hong Kong63.
Animal prophylaxis experiment
The use of all animals in this study was approved by the Committee on the Use of Live Animals in Teaching and Research (CULATR) of the University of Hong Kong under CULATR 22-139. The six-week-old male Syrian golden hamsters obtained from the Center for Comparative Medicine Research (CCMR) at the University of Hong Kong were housed in cages equipped with individual ventilation systems. Moreover, the hamsters were kept at 65% humidity and at ambient temperatures of 21-23 °C with the 12-h day-night cycle for proper care and management. The group size of hamsters was determined based on statistical power analysis and our prior experience64,65.
For mono-prophylaxis experiments, hamsters were intraperitoneally injected with one dose of PBS or the corresponding antibodies (20 mg/kg for each antibody) eight h before virus inoculation. For virus inoculation, intranasal virus infection was performed after intraperitoneal ketamine (100 mg/kg) and xylazine (10 mg/kg) anesthesia. Virus-infected hamsters were euthanized at four days post-inoculation to collect the nasal turbinate (NT) and lung tissues. The collected samples were used to determine the viral gene expression level and the infectious virus titer, and were also used for immunofluorescence staining and hematoxylin and eosin (H&E) staining as described previously66.
Relative quantification of viral gene expression
Expression of sgE transcript level was quantified as previously described67. Briefly, probe-based qRT-PCR targeting the subgenomic transcript of the SARS-CoV-2 envelope (sgE) gene was performed, normalizing against the hamster reference gene, β-actin. Primers and probes: sgE-forward, 5’-CGA TCT CTT GTA GAT CTG TTC TC-3’; sgE-reverse, 5’- ATA TTG CAG CAG TAC GCA CAC A-3’; sgE-probe, FAM-ACA CTA GCC ATC CTT ACT GCG CTT CG-BBQ; β-actin-forward, 5’-ATG GCC AGG TCA TCA CCA TTG-3’; β-actin-reverse, 5’-CAG GAA GGA AGG CTG GAA AAG-3’. Total RNA was extracted from homogenized animal tissues using RNeasy Mini Kit (Qiagen), and qRT-PCR was performed using QuantiNova Probe RT-PCR Kit (Qiagen) or QuantiNova SYBR Green RT-PCR Kit (Qiagen) with the LightCycler 480 Real-Time PCR System (Roche). Cycle threshold (Ct) values were obtained, and relative expression was calculated using the 2−ΔCt method: ΔCt=CtsgE − Ctβ-actin.
Immunofluorescence staining and histology
Infected hamster lungs were fixed by 4% paraformaldehyde (PFA) for 24 h. The fixed samples were then washed and embedded in paraffin with a TP1020 Leica semi-enclosed benchtop tissue processor. Embedded samples were sectioned with a microtome (Thermo Fisher Scientific) and placed on microscope slides for drying at 37 °C overnight. Sample slides were dewaxed by washing in serially diluted xylene, ethanol, and double-distilled water. For antigens unmasking, diluted antigen unmasking solution H-3300 (Vector Laboratories) was heated to 85 °C, and then the slides were immersed into the solution and boiled for 90 sec. Afterwards, the unmasked slides were blocked with Sudan Black B and 5% FBS, respectively for 30 min. The rabbit serum against SARS-CoV-2 nucleocapsid (N) protein was applied onto the slides and incubated overnight at 4 °C. Fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit IgG (Thermo Fisher Scientific) was used as a secondary antibody. To stain the cell nucleus, ProLong Diamond Antifade Mountant with DAPI (Thermo Fisher Scientific) was used during slide mounting. Subsequently, fluorescent images were taken with the Olympus BX53 fluorescence microscope. Fluorescent signal intensity of N protein expression was quantified with ImageJ.
For hematoxylin and eosin (H&E) staining, slides were stained with Gill’s hematoxylin and eosin. Images were taken using the Olympus BX53 light microscope. Five hamsters were sampled in each group, and four to six slides from each animal were used for histology analysis as we previously described68. Histopathological scores were quantified using a semi-quantitative method as previously reported64,65.
Determination of infectious virus titer
VeroE6-TMPRSS2 was obtained from the Japanese Collection of Research Bioresources (JCRB) Cell Bank, and was cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS, 100 units penicillin, 100 μg/ml streptomycin, and 2% G418. To measure the infectious virus titer, hamster nasal turbinate and lung tissues were homogenized and centrifuged to collect the supernatants. Ten-fold serially diluted supernatants were then inoculated into monolayered VeroE6-TMPRSS2 cells in 96-well plates. Cytopathic effect (CPE) was observed at four days post-infection, and the median tissue culture infectious dose (TCID 50) was determined as previously reported69.
Polyreactivity and autoreactivity
Polyreactivity and autoreactivity assays were performed as previously described70. For polyreactivity assays, binding affinity of mAbs against six antigens, including single-stranded DNA (ssDNA), double-stranded DNA (dsDNA), insulin, keyhole limpet hemocyanin (KLH), lipopolysaccharides (LPS), and cardiolipin, was measured using ELISA assays. For autoreactivity assays, selected antibodies were tested by using a FACS-based assay. Briefly, harvested HEp-2 cells were sequentially incubated with the tested antibodies (10 μg/ml) and PE-conjugated goat anti-human IgG Fc detection antibodies (eBioscience), and subsequently analyzed by flow cytometry analysis using BD Fortessa Analyzer. 3H9 and mGO53 antibodies were used as positive and negative controls, respectively.
We also performed an immunofluorescence staining assay for autoreactivity on HEp-2 cells. Slides were firstly incubated with antibodies at a concentration of 10 μg/ml and then detected with Alexa647-conjugated F(ab’)2 goat anti-human IgG-Fc antibody (Jackson ImmunoResearch). Cells were then imaged using the BZ-X series fluorescence microscope (Keyence).
S protein purification and cryo-EM sample preparation
We expressed and purified the prefusion-stabilized trimeric ectodomains of the EG.5.1 S trimer, which incorporate GSAS and 6 P mutations, alongside the T4 fibritin trimerization domain to enhance structural stability. The purified EG.5.1 S trimer was subsequently mixed with the XGi-183, XGi-203, XGi-171, and XGi-198, respectively, at a molar ratio of 1:1.2, followed by incubation at 4 °C for a duration of 0.5 to 1.5 h. Subsequently, the S trimer-antibody complexes were isolated using a Superose 6 Increase 10/300 GL column and diluted to a concentration of 0.6-0.7 mg/mL in 20 mM Tris buffer at pH 8.0 with 200 mM NaCl. \(\beta\)-OG was added to S-XGi-203 complex to a final concentration of 0.05%. A 3 μl sample of the complex was placed onto a holey amorphous nickel-titanium alloy film supported by 400-mesh gold grids that had been glow-discharged. The sample was then vitrified by plunging into liquid ethane using a Vitrobot IV (FEI, Thermo Fisher Scientific) with a blot force of -3 and 2.0 s blot time.
Cryo-EM data collection and processing
Data were collected using a TITAN Krios G4 Transmission electron microscopy (Thermo Fisher Scientific) operated at 300 kV, equipped with a Falcon 4i and a Selectris X Imaging filter (Thermo Fisher Scientific), setting to a slit width of 20 eV. EPU software at 300 kV in AFIS mode was used for automated data collection. EER movie stack was collected in super-resolution mode, at a nominal magnification of 130,000×, corresponding to a physical pixel size of 0.932 Å, dose fractioned to 1080 frames. Defocus ranges from -1.0 μm to -3.0 μm, with a total dose of about 50 e–/Å2. The movie stacks were binned 2 × 2, dose-weighted, and motion-corrected using MotionCorr271 within Relion72 (v3.1). Data processing was carried out using cryoSPARC73 (v4.2.1). After 2D classification, particles with good qualities were selected for global 3D reconstruction and then subjected to non-uniform refinement. To improve the density surrounding the RBD-Fab region, UCSF Chimera74 (v1.16) and Relion (v3.1) were used to generate the masks, and local refinement was then performed using cryoSPARC (v4.2.1). The reported resolutions above were based on the gold-standard Fourier shell correlation (FSC) 0.143 criterion. All the visualizations of the map are performed in UCSF Chimera (v1.16). Sharpened maps are generated using DeepEMhancer75. All the data processing procedures were summarized in Supplementary Fig. 6 for XGi-183, Supplementary Fig. 8 for XGi-203, Supplementary Fig. 10 for XGi-171, and Supplementary Fig. 13 for XGi-198.
Model building and refinement
The initial model was generated utilizing SWISS-MODEL76 in conjunction with the structure of the Omicron S protein (PDB: 7WVN). The initial models were integrated into the corresponding maps employing UCSF Chimera, followed by manual adjustments in COOT77. Subsequent to this, multiple rounds of real-space refinement were carried out using the PHENIX78 software. Model validation was also conducted within the PHENIX. Figures were created using both UCSF Chimera and UCSF ChimeraX79. Detailed statistics regarding data collection and model refinement are provided in Supplementary Table 4. The residues of the antibody were numbered according to the IMGT® (International ImMunoGeneTics information system®) unique numbering.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The coordinates of RBD-XGi-183 Fab complex, S-XGi-203 trimer complex, S-XGi-203 down-RAb1 local refinement, S-XGi-171 trimer complex, S-XGi-171 RAb1 local refinement, S-XGi-198 dimer of trimers complex, S-XGi-198 up-RAb1 local refinement have been deposited in the Protein Data Bank (PDB) with the accession numbers: 9KZE, 9L15, 9L07, 9KZZ, 9KZD, 9L2L, and 9L05, respectively; their corresponding maps have been deposited in the Electron Microscopy Data Bank (EMDB) with the accession numbers: EMD-62661, EMD-62734, EMD-62729, EMD-62731, EMD-62733, EMD-62691, EMD-62680, EMD-62660, EMD-62777, EMD-62744, EMD-62745, EMD-62746, EMD-62687, Source data are provided with this paper.
References
Carabelli, A. M. et al. SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat. Rev. Microbiol 21, 162–177 (2023).
Tao, K. et al. The biological and clinical significance of emerging SARS-CoV-2 variants. Nat. Rev. Genet 22, 757–773 (2021).
Roemer, C. et al. SARS-CoV-2 evolution in the Omicron era. Nat. Microbiol 8, 1952–1959 (2023).
Xia, S., Wang, L., Zhu, Y., Lu, L. & Jiang, S. Origin, virological features, immune evasion and intervention of SARS-CoV-2 Omicron sublineages. Signal Transduct. Target Ther. 7, 241 (2022).
Mlcochova, P. et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 599, 114–119 (2021).
Dejnirattisai, W. et al. Antibody evasion by the P.1 strain of SARS-CoV-2. Cell 184, 2939–2954.e2939 (2021).
Wang, P. et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 593, 130–135 (2021).
Chen, R. E. et al. In vivo monoclonal antibody efficacy against SARS-CoV-2 variant strains. Nature 596, 103–108 (2021).
Wang, Q. et al. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 186, 279–286.e278 (2023).
He, Q. et al. An updated atlas of antibody evasion by SARS-CoV-2 Omicron sub-variants including BQ.1.1 and XBB. Cell Rep. Med. 4, 100991 (2023).
Planas, D. et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 602, 671–675 (2022).
Cao, Y. et al. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. Nature 608, 593–602 (2022).
Wang, Q. et al. Antibody neutralisation of emerging SARS-CoV-2 subvariants: EG.5.1 and XBC.1.6. Lancet Infect. Dis. 23, e397–e398 (2023).
Qu, P. et al. Immune evasion, infectivity, and fusogenicity of SARS-CoV-2 BA.2.86 and FLip variants. Cell, https://doi.org/10.1016/j.cell.2023.12.026 (2024).
Jian, F. et al. Evolving antibody response to SARS-CoV-2 antigenic shift from XBB to JN.1. Nature 637, 921–929 (2024).
Cao, Y. et al. Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution. Nature 614, 521–529 (2023).
Gao, B. et al. Repeated vaccination of inactivated SARS-CoV-2 vaccine dampens neutralizing antibodies against Omicron variants in breakthrough infection. Cell Res. 33, 258–261 (2023).
Yisimayi, A. et al. Repeated Omicron exposures override ancestral SARS-CoV-2 immune imprinting. Nature 625, 148–156 (2024).
Starr, T. N. et al. Shifting mutational constraints in the SARS-CoV-2 receptor-binding domain during viral evolution. Science 377, 420–424 (2022).
Xue, S., Han, Y., Wu, F. & Wang, Q. Mutations in the SARS-CoV-2 spike receptor binding domain and their delicate balance between ACE2 affinity and antibody evasion. Protein Cell 15, 403–418 (2024).
Li, S. et al. Characterization of cross-reactive monoclonal antibodies against SARS-CoV-1 and SARS-CoV-2: Implication for rational design and development of pan-sarbecovirus vaccines and neutralizing antibodies. J. Med Virol. 95, e28440 (2023).
Lu, G. et al. Primary assessment of the diversity of Omicron sublineages and the epidemiologic features of autumn/winter 2022 COVID-19 wave in Chinese mainland. Front Med. 17, 758–767 (2023).
Dai, L. et al. Efficacy and Safety of the RBD-Dimer-Based Covid-19 Vaccine ZF2001 in Adults. N. Engl. J. Med. 386, 2097–2111 (2022).
Yang, S. et al. Fast evolution of SARS-CoV-2 BA.2·86 to JN.1 under heavy immune pressure. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(23)00744-2 (2023).
Yamasoba, D. et al. Virological characteristics of the SARS-CoV-2 omicron XBB.1.16 variant. Lancet Infect. Dis. 23, 655–656 (2023).
Wang, Q. et al. Antigenicity and receptor affinity of SARS-CoV-2 BA.2.86 spike. Nature 624, 639–644 (2023).
Ling, Y. et al. The Omicron BA.2.2.1 subvariant drove the wave of SARS-CoV-2 outbreak in Shanghai during spring 2022. Cell Discov. 8, 97 (2022).
Zhang, X., Zhang, W. & Chen, S. Shanghai’s life-saving efforts against the current omicron wave of the COVID-19 pandemic. Lancet 399, 2011–2012 (2022).
Wang, K. et al. Memory B cell repertoire from triple vaccinees against diverse SARS-CoV-2 variants. Nature 603, 919–925 (2022).
Zhou, Y. et al. Enhancement versus neutralization by SARS-CoV-2 antibodies from a convalescent donor associates with distinct epitopes on the RBD. Cell Rep. 34, 108699 (2021).
Wang, K. et al. A third dose of inactivated vaccine augments the potency, breadth, and duration of anamnestic responses against SARS-CoV-2. medRxiv, 2021.2009.2002.21261735 (2021).
Muecksch, F. et al. Increased memory B cell potency and breadth after a SARS-CoV-2 mRNA boost. Nature 607, 128–134 (2022).
Wang, Z. et al. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 592, 616–622 (2021).
Robbiani, D. F. et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature 584, 437–442 (2020).
Wang, Z. et al. Naturally enhanced neutralizing breadth against SARS-CoV-2 one year after infection. Nature 595, 426–431 (2021).
Jia, Z. et al. A third dose of inactivated vaccine augments the potency, breadth, and duration of anamnestic responses against SARS-CoV-2. Protein Cell, https://doi.org/10.1093/procel/pwae033 (2024).
Yang, S. et al. Antigenicity and infectivity characterisation of SARS-CoV-2 BA.2.86. Lancet Infect. Dis. 23, e457–e459 (2023).
Yang, J. et al. Low levels of neutralizing antibodies against XBB Omicron subvariants after BA.5 infection. Signal Transduct. Target Ther. 8, 252 (2023).
Pinto, D. et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 583, 290–295 (2020).
Cao, Y. et al. Rational identification of potent and broad sarbecovirus-neutralizing antibody cocktails from SARS convalescents. Cell Rep. 41, 111845 (2022).
Yuan, M. et al. A broad and potent neutralization epitope in SARS-related coronaviruses. Proc. Natl Acad. Sci. USA 119, e2205784119 (2022).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Zhao, Z. et al. Omicron SARS-CoV-2 mutations stabilize spike up-RBD conformation and lead to a non-RBM-binding monoclonal antibody escape. Nat. Commun. 13, 4958 (2022).
Huo, J. et al. Neutralization of SARS-CoV-2 by Destruction of the Prefusion Spike. Cell Host Microbe. 28, 497 (2020).
Wrobel, A. G. et al. Antibody-mediated disruption of the SARS-CoV-2 spike glycoprotein. Nat. Commun. 11, 5337 (2020).
Starr, T. N. et al. Deep mutational scanning of sars-cov-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell 182, 1295–1310.e1220 (2020).
Tian, X. et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 9, 382–385 (2020).
He, Y., Lu, H., Siddiqui, P., Zhou, Y. & Jiang, S. Receptor-binding domain of severe acute respiratory syndrome coronavirus spike protein contains multiple conformation-dependent epitopes that induce highly potent neutralizing antibodies. J. Immunol. 174, 4908–4915 (2005).
Liu, Z. et al. RBD-Fc-based COVID-19 vaccine candidate induces highly potent SARS-CoV-2 neutralizing antibody response. Signal Transduct. Target Ther. 5, 282 (2020).
Hastie, K. M. et al. Defining variant-resistant epitopes targeted by SARS-CoV-2 antibodies: A global consortium study. Science 374, 472–478 (2021).
Barnes, C. O. et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 588, 682–687 (2020).
Ju, B. et al. Infection with wild-type SARS-CoV-2 elicits broadly neutralizing and protective antibodies against omicron subvariants. Nat. Immunol. 24, 690–699 (2023).
Bianchini, F. et al. Human neutralizing antibodies to cold linear epitopes and subdomain 1 of the SARS-CoV-2 spike glycoprotein. Sci. Immunol. 8, eade0958 (2023).
Zhou, P. et al. Broadly neutralizing anti-S2 antibodies protect against all three human betacoronaviruses that cause deadly disease. Immunity 56, 669–686.e667 (2023).
Cai, J. et al. Risk of reinfection and severity with the predominant BA.5 Omicron subvariant China, from December 2022 to January 2023. Emerg. Microbes Infect. 13, 2292071 (2024).
Wang, Q. et al. A Combination of Human Broadly Neutralizing Antibodies against Hepatitis B Virus HBsAg with Distinct Epitopes Suppresses Escape Mutations. Cell Host Microbe. 28, 335–349.e336 (2020).
Brochet, X., Lefranc, M. P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–508 (2008).
Ye, J., Ma, N., Madden, T. L. & Ostell, J. M. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–40 (2013).
Zhou, Y. et al. Single-Cell Sorting of HBsAg-Binding Memory B Cells from human peripheral blood mononuclear cells and antibody cloning. STAR Protoc. 1, 100129 (2020).
Scheid, J. F. et al. Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333, 1633–1637 (2011).
Zhao, X. et al. Potent and broadly neutralizing antibodies against sarbecoviruses induced by sequential COVID-19 vaccination. Cell Discov. 10, 14 (2024).
Wang, X. et al. Neutralization of distinct Omicron sublineages by longitudinal vaccination sera. J. Med. Virol. 94, 5090–5092 (2022).
Chu, H. et al. Coronaviruses exploit a host cysteine-aspartic protease for replication. Nature 609, 785–792 (2022).
Shuai, H. et al. The viral fitness and intrinsic pathogenicity of dominant SARS-CoV-2 Omicron sublineages BA.1, BA.2, and BA.5. EBioMedicine 95, 104753 (2023).
Hu, B. et al. Divergent trajectory of replication and intrinsic pathogenicity of SARS-CoV-2 Omicron post-BA.2/5 subvariants in the upper and lower respiratory tract. EBioMedicine 99, 104916 (2024).
Chan, J. F. et al. Virological features and pathogenicity of SARS-CoV-2 Omicron BA.2. Cell Rep. Med. 3, 100743 (2022).
Liu, Y. et al. Lineage-specific pathogenicity, immune evasion, and virological features of SARS-CoV-2 BA.2.86/JN.1 and EG.5.1/HK.3. Nat. Commun. 15, 8728 (2024).
Chan, J. F. et al. Altered host protease determinants for SARS-CoV-2 Omicron. Sci. Adv. 9, eadd3867 (2023).
Shuai, H. et al. Attenuated replication and pathogenicity of SARS-CoV-2 B.1.1.529 Omicron. Nature 603, 693–699 (2022).
Molinos-Albert, L. M. et al. Anti-V1/V3-glycan broadly HIV-1 neutralizing antibodies in a post-treatment controller. Cell Host Microbe 31, 1275–1287.e1278 (2023).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, https://doi.org/10.7554/eLife.42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D. Biol. Crystallogr 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
We thank the Center of Cryo-Electron Microscopy, core facility of Shanghai Medical College, Fudan University, for the support on cryo-EM data collection. This work was supported by National Key Research and Development Program (2021YFA1301404 and 2023YFC0872600 to Q.W., 2023YFC2605400 and 2023YFC3404000 to X.Z., 2021YFC0866100 and 2023YFC3041600 to H.C., and 2021YFC2302500 to L.S.), National Natural Science Foundation of China (32070947 and 32370943 to Q.W., 32300121 to X.Z., 32270142 to P.W., 32122001 to H.C., and 92469108 to L.S.), Shanghai Municipal Science and Technology Major Project (ZD2021CY001 to F.W.), Shanghai Rising-Star Program (22QA1408800 to P.W.), Shanghai Municipal Health Commission (202240081 to Y.X.), the Program of Science and Technology Cooperation with Hong Kong, Macao and Taiwan (23410760500 to P.W.), and R&D Program of Guangzhou Laboratory (SRPG22-003 to L.S.). This work was also supported by the Non-profit Central Research Institute Fund of the Chinese Academy of Medical Sciences (2023-PT310-02 to Q.W.). H.C. acknowledges funding from the General Research Fund (17118621, 17119122), Collaborative Research Fund (C7103-22G), and Theme-Based Research Scheme (T11-709/21-N), the Research Grants Council of the Hong Kong Special Administrative Region. H.C. also acknowledges funding from Health@InnoHK, Innovation and Technology Commission, the Government of the Hong Kong Special Administrative Region. The funding agencies had no role in study design, data collection, data analysis, data interpretation, or manuscript writing, and the content is solely the responsibility of the authors. We acknowledge the use of NIH BIOART assets: “NIAID Visual & Medical Arts. (10/7/2024). B Cell. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/50”, “NIAID Visual & Medical Arts. (10/7/2024). BCR. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/51”, “NIAID Visual & Medical Arts. (10/7/2024). Syringe. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/505”, “NIAID Visual & Medical Arts. (10/7/2024). SARS-CoV-2. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/464”, “NIAID Visual & Medical Arts. (10/7/2024). BDBlood Vial. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/52”, “NIAID Visual & Medical Arts. (10/7/2024). Lab Hamster. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/278”, “NIAID Visual & Medical Arts. (10/7/2024). Plasma B Cell. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/409”, “NIAID Visual & Medical Arts. (10/7/2024). RNA. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/452”, and “NIAID Visual & Medical Arts. (10/7/2024). 96 Well Plate. NIAID NIH BIOART Source. bioart.niaid.nih.gov/bioart/7”.
Author information
Authors and Affiliations
Contributions
Q.W., H.C., P.W., and L.S. conceived and designed the study. M.X., Y.Q., X.Z., J.S., and Q.Z. performed most of the experiments and analyzed data together with T.Y.O., Z.W., Y.Y., W.Z, C.T.M., Y.X., S.J. and L.L. R.X., M.X., S.S., Y.Z., Y.Z., and A.X. recruited volunteers and organized blood collection. M.X., Q.Z., J.H., and L.L. performed single B-cell sorting, antibody cloning, antibody expression, and purification. M.X., Q.Z., Y.Z., and L.L. performed the ELISA and competition ELISA. X.Z. and J.L. constructed the SARS-CoV-2 pseudovirus panel and performed pseudovirus neutralization assays. Y.Q. and Q.M. performed the antibody structural studies. J.S. and Y.L. executed the animal protection studies and participated in data analysis. Q.W., M.X., Y.Q., X.Z., J.S., Y.L., and Q.Z. had full access to data in the study, generated figures and tables, and took responsibility for the integrity and accuracy of the data presentation. Q.W., H.C., P.W., L.S., F.W., and M.X. participated in the discussion of the results and wrote the manuscript. All authors read and approved this version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Q.W., M.X., F.W., L.S., H.C., and P.W. have a provisional patent application. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xavier Saelens and Renhong Yan for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xie, M., Qiu, Y., Zhao, X. et al. Orphan broadly RBD-binding antibodies annotate three remaining conserved RBD epitopes along SARS-CoV-2 evolution. Nat Commun 16, 10566 (2025). https://doi.org/10.1038/s41467-025-65596-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65596-0