Abstract
Model bacteria are fundamental for research, but knowledge about their ecology and evolution is often limited. Here, we establish an evolutionary and ecological context for the model organism Caulobacter crescentus—an alphaproteobacterium intensively studied for its dimorphic lifecycle. By analyzing the phylogenetic relatedness and genetic potential of hundreds of Caulobacterales species, we reveal substantial diversity regarding their environmental distribution, morphology, cell development, and metabolism. Our work provides insights into the evolutionary history of morphological features such as the cell curvature determinant crescentin and uncovers a striking case of convergent loss of traits for cellular dimorphism among close relatives of C. crescentus. Moreover, we find that genes for phototrophy are widespread across Caulobacterales and that the new genus Acaudatibacter, described here, includes the first reported Caulobacterales lineage with photoautotrophic potential. Our study advances our understanding of an environmentally widespread bacterial order and sheds light on the evolution of fundamental prokaryotic features.
Similar content being viewed by others
Introduction
Most bacteriological research relies on a limited number of model bacteria, which often represent large taxonomic groups1,2. The bias towards a few species, primarily human pathogens, leaves the vast majority of bacteria understudied, greatly narrowing the scope of known prokaryotic biology. A current challenge in microbiology is to integrate existing knowledge about model bacteria with the immense and unexplored bacterial diversity revealed by the recent advancement of cultivation-independent techniques, such as metagenomics3.
One example of a well-studied model bacterium is Caulobacter crescentus (also known as C. vibrioides4). Since its isolation in the 1960s, it has been studied intensively for its complex lifecycle and today serves as one of the forefront models for bacterial cell and developmental biology5,6. The C. crescentus lifecycle is dimorphic, comprising two morphologically distinct stages: a brief motile dispersive ‘swarmer’ stage and a mature sessile reproducing ‘stalked’ stage7. The stalked cell is named after its thin cell envelope extension (prostheca), which it uses to attach itself to a surface with its holdfast adhesin. Stalked cells always divide asymmetrically to generate swarmer cells from their stalk-distal cell pole. Newborn swarmer cells are capable of exploring their environment using their flagellum, chemotaxis machinery, and adhesive type IV pili (T4P), but cannot reproduce until they have irreversibly differentiated into sessile stalked cells7. This cell differentiation event, which is an integral part of the Caulobacter cell cycle, along with the sophisticated spatiotemporal mechanisms underlying its asymmetric cell division, has attracted extensive research during the past 50 years, which has fundamentally shaped our views on bacterial cell organization and development5,6.
Despite decades of molecular research, the precise ecological relevance and evolutionary history of C. crescentus and its dimorphic lifecycle remain unknown. Research on C. crescentus has relied on two lab-adapted strains (CB15/NA1000) derived from the same pond water isolate8,9. Accordingly, Caulobacter was long assumed to be primarily found in oligotrophic aquatic environments. However, recent reports have shown that the genus Caulobacter is also prevalent in terrestrial environments, including soil and in association with plant material10. Caulobacter belongs to the alphaproteobacterial family Caulobacteraceae, a lineage of aerobic heterotrophs that includes biodegraders, plant commensals, and opportunistic human pathogens11,12,13,14,15. However, the diversity within the Caulobacteraceae and other families of the order Caulobacterales remains poorly studied, despite their ubiquity in nature. Thus, it is unclear to what extent the characteristic features of C. crescentus are shared with its relatives.
Recent advances in cultivation-independent methods, such as genome-resolved metagenomics, have enabled the reconstruction of genomes from a greatly expanded diversity of environmental bacteria, including many uncharacterized Caulobacterales clades. Here, we use publicly available metagenome-assembled genomes (MAGs) and isolate genomes, to investigate the diversity and evolutionary history of the Caulobacterales order, focusing on Caulobacteraceae, its largest family. Our work reveals several unexpected features of this ubiquitous group of bacteria. Notably, our results indicate the convergent loss of flagellar motility and dimorphism among independent Caulobacteraceae lineages, helping to define core modules for cellular dimorphism. Moreover, we find that phototrophy is widespread across the order Caulobacterales (~10% species), and we identify the first reported Caulobacterales lineage with photoautotrophic potential in the new genus Acaudatibacter, revealing a previously unrecognized potential role of Caulobacterales members as primary producers. Together, our work illuminates the diversity and evolution of an environmentally widespread bacterial order and provides new perspectives on the evolution of bacterial lifecycle complexity.
Results
An updated species phylogeny of the Caulobacterales
To explore the genomic and environmental diversity within the family Caulobacteraceae and its position within the order Caulobacterales, we reconstructed an updated species phylogeny using all available high-quality Caulobacterales genomes (see “Methods”). To achieve this, we grouped genomes into species clusters (≥95% average nucleotide identity; ANI)16,17 and kept one representative genome per species (Supplementary Data 1a), resulting in 347 Caulobacterales species, of which 107 are currently described. We then used the representative genomes to infer a maximum-likelihood (ML) phylogeny using 72 previously curated genes conserved across Alphaproteobacteria18, resulting in a robust species phylogeny with high non-parametric bootstrap support (npBS) for most clades (Fig. 1a and Supplementary Fig. S1). In further support of the inferred phylogenetic relationships, the tree was highly concordant with an alternative, less refined ML phylogeny of 117 alphaproteobacterial genes and with an ML phylogeny inferred for concatenated 16S and 23S rRNA genes (Supplementary Figs. S2 and S3).
a ML concatenated phylogenetic tree of 72 conserved single-copy alphaproteobacterial marker genes18 inferred using IQ-TREE109 with the LG + C60 + F + R model of evolution and 100 non-parametric bootstraps (alignment length of 25448 amino acids). The tree was inferred for genome representatives of 347 species clusters (ANI ≥ 95%) from the order Caulobacterales (following GTDB taxonomy), the model strain Caulobacter crescentus CB15, and five outgroup non-Caulobacterales Alphaproteobacteria for rooting. Poindextera gen. nov., Caulobacteraceae genera with ≥ 3 species clusters, and other Caulobacterales families are indicated with colored boxes in the phylogeny. npBS ≥ 70 are indicated with shaded circles, as outlined in the top-left corner, and the thin scale bar indicates the number of substitutions per site in the phylogeny. Genomes of validly described species type strains are marked with orange hexagons (LPSN125, accessed March 21, 2025). Schematic drawings of cell silhouettes of selected species highlight morphological diversity and are drawn roughly to scale (thick scale bar) from microscopy images (Supplementary Fig. S4) and published material (references are listed in Supplementary Table S1). Red cell outlines highlight species misclassified as Caulobacteraceae. The tree is presented alongside all species names and genome accessions in Supplementary Fig. S1. b Color-coded genome metadata. Column 1 distinguishes MAGs in light gray from isolate genomes in dark gray. Column 2 indicates the overarching environmental source for each genome, and column 3 provides more details on the sampling/isolation source. The keyword “algae” also includes Cyanobacteria. See Supplementary Data 2 for keyword definitions and original metadata. c Genome size (color) and estimated genome size (gray) calculated using estimated completeness scores from CheckM83 (‘lineage_wf’). Source data are provided as a Source Data file. d Species clusters with genetic potential for phototrophy (cyan markers); see Fig. 5b.
In the species tree, the first-described Caulobacteraceae genera Asticcacaulis, Brevundimonas, Caulobacter, and Phenylobacterium, formed a distinct clade alongside several uncharacterized genus-level groups (Fig. 1a and Supplementary Fig. S1). Notably, the clade did not include the genera Aquidulcibacter, Pseudaquidulcibacter, and Terricaulis, which were recently assigned to Caulobacteraceae19,20,21. Instead, these genera formed a clade (100% npBS) that clustered sister to the family Hyphomonadaceae (98% npBS), together with Vitreimonas flagellata22 (Hyphomonadaceae), indicating that they do not belong to the family Caulobacteraceae. We instead propose the name Aquidulcibacteraceae fam. nov. for this clade (description found in Supplementary Note 1). Based on our robust and comprehensive species phylogenies (Supplementary Figs. S1 and S2), we propose additional revisions to Caulobacterales taxonomy, including the transfer of the recently described species Phenylobacterium montanum23, which consistently clustered outside of Phenylobacterium and Caulobacter, into the new Caulobacteraceae genus Poindextera gen. nov. (Poindextera montana comb. nov.) (Supplementary Notes 1 and 2). Henceforth, we will refer to the clade composed of Asticcacaulis, Brevundimonas, Caulobacter, Phenylobacterium, and Poindextera gen. nov. as Caulobacteraceae, excluding members of Aquidulcibacteraceae fam. nov.
The refined Caulobacteraceae family comprised 206 species, of which 152 are undescribed, and was notably more species-rich than the other four Caulobacterales families (140 species in total) (Fig. 1a). It was split into two major clades: one including the genera Asticcacaulis, Brevundimonas, and the genus-level clade “17J80-11” (100% npBS), and the other including Caulobacter, Phenylobacterium, Poindextera gen. nov., as well as the genus-level clade “Palsa-881” and six other uncharacterized genera (99% npBS) (Fig. 1a and Supplementary Fig. S1). Due to its distinct features described below, we renamed “Palsa-881” to Acaudatibacter gen. nov. according to the SeqCode24 nomenclature for uncultivated microorganisms (available under the accession ID 9aocwnme; Supplementary Notes 3 and 4).
Environmental distribution of Caulobacterales
To investigate Caulobacterales environmental distribution, we manually compiled and curated metadata (see “Methods”) for all 347 species-cluster genome representatives (Fig. 1b and Supplementary Data 2). We found that Caulobacteraceae differs from other Caulobacterales families in several regards. First, while the other families were predominantly aquatic, Caulobacteraceae included a high proportion (32.4%) of terrestrial lineages (Fig. 1b and Supplementary Fig. S5a). Second, only 5.8% of Caulobacteraceae species representatives came from marine/saline environments, while the other Caulobacterales families (except Aquidulcibacteraceae fam. nov.) were almost exclusively associated with such habitats (Fig. 1b and Supplementary Fig. S5b). Third, unlike other families of the order Caulobacterales, the Caulobacteraceae family contained a considerable fraction of species associated with plant and animal hosts. Using the IMNGS 16S rRNA gene amplicon database25, we similarly found that Caulobacteraceae most frequently had high relative abundance (≥1%) in environments associated with plants and their roots, as well as freshwater, soil, and wastewater (Supplementary Fig. S5c and Supplementary Data 3). Caulobacteraceae sequences were also prevalent in studies of marine, gut, and fecal environments, but less frequently at high relative abundance.
Within the genus Caulobacter, nearly all species came from aquatic freshwater environments or from terrestrial environments closely associated with plants (Fig. 1b, Supplementary Fig. S6a, and Supplementary Data 2). Terrestrial plant-associated Caulobacter species had considerably larger genomes than aquatic species (Fig. 1c and Supplementary Fig. S6b). This is consistent with a previous report for Caulobacter10 and observations that plant-inhabiting Pseudomonadota (Proteobacteria) in general have larger genomes26,27,28, and suggests that distinct Caulobacter lineages are specialized to either plant-associated or aquatic lifestyles. Indeed, terrestrial Caulobacter genomes encoded several virulence and host colonization factors rare among aquatic species, including type I (hlyBD), type III, and type VI secretion systems, and the sulfonate transport system ssuABC (Supplementary Fig. S6a), all suggested mediators of plant–bacterial interactions29,30,31,32,33,34,35.
Convergent losses of dimorphism-related genes in Caulobacteraceae
Having established a robust species phylogeny and the environmental distribution of Caulobacteraceae, we used these insights to assess the extent to which key features of the model organism C. crescentus, such as its well-characterized lifecycle, are representative of the family. During cell development, an obligate motile-to-sessile transition precedes reproduction, and during each cell division, the sessile stalked cell generates motile flagellated offspring (Fig. 2a). In line with this canonical dimorphic lifecycle, most Caulobacteraceae genomes had large sets of genes known to mediate flagellar motility and chemotaxis (Fig. 2b and Supplementary Data 5b, c). However, strikingly, four genomes completely lacked such genes: (i) the immotile soil isolate Phenylobacterium immobile ET 36 and (ii) three freshwater MAGs clustering together in the uncharacterized genus-level clade “Palsa-881” here renamed Acaudatibacter gen. nov. (‘tailless bacterium’) (Fig. 2b) (SeqCode taxonomic descriptions in Supplementary Notes 3 and 4). When also including Acaudatibacter genomes subset from the dataset of Rodríguez-Gijón et al.37, which we refer to as the “extended dataset” (see “Methods”; Supplementary Data 1b), the number of Acaudatibacter species clusters lacking these genes increased to ten, comprising 32 MAGs from diverse sequencing projects (Supplementary Figs. S7a, b and S8). Given the large number of independently sampled and assembled Acaudatibacter genomes specifically lacking these genes, and that motility genes are normally spread out over the chromosome, the observed gene absence patterns are unlikely to stem from assembly artifacts. Moreover, these ten species formed a monophyletic group within Acaudatibacter (100% ultrafast bootstrap support [ufBS]), while the remaining two Acaudatibacter species, which have motility genes, Ac. sp. SZAS_AMP-5 and Ac. sp. CP_BM_RX_R9_33, were deep-branching (Supplementary Fig. S9). Thus, P. immobile and most members (10/12) of the new genus Acaudatibacter lack flagellar genes and are both robustly nested within clades where deeper branching lineages have flagellar genes. Together, this strongly suggests that flagellar motility has been lost in at least two separate Caulobacteraceae lineages. It also shows that the isolate P. immobile ET is immotile due to a complete lack of flagellar genes, in contrast to P. hankyongense and P. soli, which despite being described as immotile38,39 have flagellar genes (Fig. 2b and Supplementary Data 5b, c).
a Lifecycle of Caulobacter crescentus, showcasing the obligate motile-to-sessile (swarmer-to-stalked) cell differentiation that precedes reproduction, as well as the obligate generation of motile offspring through asymmetric cell division. Schematic representation of cellular levels of the intracellular signaling molecule cyclic diguanylate (c-di-GMP) is indicated in purple in the lower bar, highlighting the surge in c-di-GMP levels that drives cell differentiation126. b Presence and absence of genes involved in chemotaxis (mcp and genes with che, cle, and mot names), flagellar motility (genes with fla, flb, flg, flh, fli, and flm names), type IV tight-adherence pilus (T4P; pilA, tadD, and genes with cpa names), holdfast (genes with hfa and hfs names), and cell developmental regulation, as annotated by KEGG KOs (dark gray; from eggNOG-mapper95) and using the reciprocal best blast hit (RBH) algorithm against the C. crescentus CB15 proteome (blue). The total numbers of genes investigated for each functional category are indicated (n). For genera and families, the average presence and absence is shown with linear color gradients. Five outgroup Alphaproteobacteria are included to highlight the broader conservation of genes. The labels of the cell developmental genes have been colored as follows. Purple: master cell cycle regulators; red: absent from both lineages lacking flagellar genes; yellow: absent from both lineages lacking flagellar genes and their close relatives; and gray: remaining regulators. See Supplementary Data 4 and 5a–f for the full dataset, including gene names and functional annotations. c Simplified schematic summary of C. crescentus cell development regulation, showcasing positive and negative regulatory interactions. Developmental regulators are color-coded as in (b). Direct binders of cyclic diguanylate (c-di-GMP) are highlighted with purple text. CckA is displayed in its phosphatase state (*). For simplicity, temporal aspects are disregarded, and most of the specific genes controlled by CtrA, CcrM, DnaA, and GcrA are not indicated. Based on refs. 7,40,41,42,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144.
C. crescentus also produces polarly localized type IV pili (T4P), holdfast adhesin, and the prostheca (stalk) in a cell cycle-dependent manner (Fig. 2a). We found that P. immobile lacked genes for both holdfast and T4P, with the latter otherwise being highly conserved across Caulobacteraceae (Fig. 2b and Supplementary Data 5e, f). Similarly, the Acaudatibacter species lacking flagellar genes also lacked holdfast genes, in contrast to other members of the genus (Fig. 2b, Supplementary Fig. S8, and Supplementary Data 5e, f). Notably, since the genetic basis for the prostheca remains poorly defined, we could not robustly predict the presence of prosthecae among the species in our dataset (Supplementary Data 5i).
To elucidate how the cell developmental program has evolved to accommodate the absence of flagella and polar adhesion pathways, we searched for orthologs of C. crescentus developmental regulators. Remarkably, the genomes lacking flagellar and holdfast genes also lacked orthologs of the same 26 regulators of dimorphic development otherwise largely conserved across Caulobacteraceae, despite representing separate lineages (Fig. 2b, c, Supplementary Figs. S8 and S9, and Supplementary Data 5d). Moreover, except for two proteins (SpbR40 and CpaM41), these regulators have all been associated with the signaling molecule cyclic diguanylate (c-di-GMP), a conserved bacterial regulator of motile–sessile transitions that in C. crescentus drives cell differentiation and orchestrates the asymmetric generation of motile offspring (Fig. 2a)42. This included producers and degraders of c-di-GMP (dgcAB, pleD, and pdeA, respectively), c-di-GMP-binding effectors (popA, shkA, dgrAB, tipF), as well as regulators closely upstream (cckN, fssAB, hmrABCX, mopJ), or downstream (cpdR, rcdA, rpoN, shpA, shkA, spmY, staR) of c-di-GMP (Fig. 2c). Despite the absence of this large set of c-di-GMP-associated genes, the genomes lacking flagellar and holdfast genes still encoded multiple enzymes with canonical c-di-GMP biosynthesis and/or degradation domains (Supplementary Fig. S10). Together, this suggests that these species still maintain c-di-GMP metabolism, but that they might use it for regulating other functions than their core developmental programs. Notably, in addition to the roughly 80–100 characterized cell morphology and development genes, we found 25 uncharacterized genes specifically absent from both non-flagellated Caulobacteraceae lineages (Supplementary Fig. S11 and Supplementary Data 6). These genes encode putative signaling proteins, transcriptional regulators, enzymes, and hypothetical proteins, and are promising candidate genes for potential flagellar motility and/or cellular dimorphism factors.
Given the absence of genes involved in morphological asymmetry among non-flagellated Caulobacteraceae species, we wondered if they also lack reproductive asymmetry, i.e., the production of daughter cells with different reproductive capabilities—a hallmark of the dimorphic lifecycle43. Indeed, when tracking two subsequent cell division cycles of P. immobile by time-lapse microscopy, we found that most daughter cell siblings descending from the first division cycle divided synchronously during the second division cycle (Fig. 3a–c and Supplementary Movie 1), similarly to the monomorphic bacterium Escherichia coli (Fig. 3a–c and Supplementary Movie 2). In contrast, in C. crescentus, one daughter cell consistently divided later than the other daughter cell, as expected (Fig. 3a–c and Supplementary Movie 3). Moreover, while C. crescentus exhibited a bimodal distribution of generation times among individual cells, consistent with the generation time difference between swarmer and stalked cells, monomodal distributions were instead observed for P. immobile and E. coli (Fig. 3d). Lastly, we also found that the cell division site was symmetrically positioned mid-cell in P. immobile, mirroring E. coli, but contrasting C. crescentus, in which it is asymmetrically localized closer to the swarmer cell pole44 (Fig. 3e). These results demonstrate that P. immobile produces symmetric daughter cells, both regarding size and cell division timing.
a Micrographs from timelapses at the time of the first and second cell division events for Caulobacter crescentus CB15 (PYE, 30 °C), P. immobile ET (R2A, 30 °C), and Escherichia coli MG1655 (LB, 37 °C). Cell outline schematics highlight daughter cell lineages. Full timelapses are shown in Supplementary Movies 1–3. Scale bars: 1 µm. b Schematics of the quantification of degrees of replicative asymmetry from time-lapse microscopy. The daughter cells of an initial binary cell division event divide again after the times t1 and t2, respectively. For bacteria with replicative asymmetry among daughter cells, one daughter cell undergoes subsequent cell division at a later time point (t2), than its sibling (t1), i.e., ∆t = t2 − t1 > 0. To compare species with different growth rates, we defined the degree of asymmetry as ∆t / t2, i.e., how much extra time the late-dividing daughter cells spends to divide, as a proportion of their total generation time. c Histograms of degrees of asymmetry calculated for daughter cell pairs as described in (b), from time-lapse experiments conducted as in (a) for C. crescentus, P. immobile, and E. coli in at least three biologically independent replicates (in total n = 433, n = 473, and n = 207 daughter cell pairs, respectively). The right-most bins represent degrees of asymmetry above 50%. Source data are provided as a Source Data file. d Histograms of generation times of individual cells, from daughter cell pairs analyzed in (c). For C. crescentus CB15, the approximate time difference between the bimodal peaks is highlighted. The right-most bins represent generation times above 150 min for C. crescentus and E. coli, or above 20 h for P. immobile. Source data are provided as a Source Data file. e Histograms of relative cell constriction site positions in C. crescentus NA1000 (n = 1900), P. immobile ET (n = 1900), and E. coli MG1655 (n = 1200), divided into 100 bins, quantified from snapshot images of cells grown in batch cultures to mid-exponential phase under growth conditions corresponding to those used for timelapses presented in (a). Vertical lines show mid-cell position ±5%. Schematic cell illustrations highlight that relative constriction site positions are not sorted by cell polarity. Source data are provided as a Source Data file.
Taken together, we show that genes associated with the canonical dimorphic lifecycle of C. crescentus are widespread across Caulobacteraceae, suggesting similar lifecycles among most members. However, our results indicate that P. immobile and a large subclade of Acaudatibacter have lost flagella and polar adhesins, along with specific cell developmental genes, suggesting convergent losses of dimorphic lifecycles. Moreover, we verified that P. immobile—the only cultured representative of these bacteria—reproduces symmetrically, in support of it having a monomorphic lifecycle.
Most Caulobacterales members lack genetic potential for S-layers and crescentin-mediated cell curvature
Other well-studied5 features of C. crescentus are its S-layer45, a cell envelope layer assembled from the protein RsaA, and crescentin46 (CreS), an intermediate filament-like cytoskeletal protein that generates cell curvature (Fig. 4a). We found that these two features were not particularly common among Caulobacteraceae (7.8% and 23.8% of species, respectively) and that they were absent from other Caulobacterales families (Fig. 4b and Supplementary Data 5g, h). Specifically, the S-layer genes rsaADEF were limited to the environmentally widespread (see Fig. 1a, b) C. crescentus–segnis clade and disparate Phenylobacterium species, while creS was present among most Caulobacter and a minority of Brevundimonas species, with only singular hits found in a genome each of Acaudatibacter, “CAIMFV01”, and Phenylobacterium (Fig. 4b).
a Schematic illustration of the cell curvature-generating cytoskeletal protein crescentin (CreS) and S-layer cell envelope protein RsaA in C. crescentus. OM outer membrane, PG peptidoglycan, IM inner membrane. b Presence (triangles) of crescentin (creS; K18642; blue), S-layer (rsaA; K12544; orange), and S-layer secretion machinery (rsaDEF; K12533–K12535; black) across Caulobacterales genomes. The phylogeny shows the tree from Fig. 1a. Phen., Phenylobacterium; Acau., Acaudatibacter; Caul., Caulobacter; CAIM., “CAIMFV01”; Brev., Brevundimonas. See Supplementary Data 4 and 5g, h for the full dataset. c ML tree of crescentin (CreS) homologs inferred using IQ-TREE109 with the Q.pfam + C60 + R9 model of evolution and 1000 ultrafast bootstraps (356 sequences with 902 amino acid alignment positions), and rooted using the major Beta-/Gammaproteobacteria clade. ufBS is indicated with circles and specified with numbers for key branches. Greek letters indicate Pseudomonadota (Proteobacteria) classes, and uppercase letters indicate alphaproteobacterial orders (see Supplementary Fig. S12a). The numbers of sequences per collapsed clade are shown in parentheses. Scale bars here and in (d) indicate the number of substitutions per site in the phylogenies. The full tree is presented in Supplementary Fig. S12a. d ML tree inferred separately for the subset of putative crescentin homologs that form a supported clade (100% ufBS) labeled “Crescentin-like proteins” in the tree shown in (c), using IQ-TREE109 with the JTT + C60 + R5 model of evolution and 1000 ultrafast bootstraps (135 sequences with 407 amino acid alignment positions). ufBS is indicated with numbers for key branches. Schematic cell silhouettes are drawn roughly in scale (thick scale bar: ~1 µm) based on microscopy images presented in Supplementary Fig. S4 for Brevundimonas goettingensis, C. crescentus, and Chelatococcus reniformis, and from published material for remaining species (Methylocystis parva, Methylocystis sp. Rockwell, and Roseiarcus fermentans; references are listed in Supplementary Table S1). gen. genus, fam. family, ord. order. The full tree is presented in Supplementary Fig. S13. e Representative micrographs (from two independent experiments each) of C. crescentus NA1000 and Ch. reniformis B2974T from cultures in mid-exponential phase, grown in PYE at 30 °C and R2A at 28 °C, respectively. Scale bars: 5 µm. f Complementation of C. crescentus NA1000 ΔcreS with native creS or the crescentin homolog from Ch. reniformis B2974T (creSCh.reniformis) expressed from the xylose-inducible Pxyl promoter on the pBXMCS-4 vector. Representative micrographs (from one experiment) are shown for mid-exponential cultures before (0 h) and after the addition of 0.3% xylose. Scale bars: 5 µm. EV empty vector. See Supplementary Fig. S14 for corresponding experiments done in the wild-type NA1000 background.
The limited distribution of crescentin prompted us to investigate its evolutionary origin. For this, we inferred a phylogeny of putative crescentin homologs (Fig. 4c and Supplementary Figs. S12a and S13). The resulting tree included phage tail tape-measure proteins (TMPs) from Alpha-, Beta-, and Gammaproteobacteria, and a fully-supported clade (100% ufBS) of sequences from Caulobacteraceae and the alphaproteobacterial order Hyphomicrobiales (synonym Rhizobiales), referred to here as ‘crescentin-like’, which included all proteins with the crescentin Pfam47 domain (PF19220) (Supplementary Fig. S12b). TMPs are structural proteins that form extended coiled coils inside phage tails to regulate their length48, reminiscent of the extended coiled coils of crescentin49. Crescentin-like proteins clustered with alphaproteobacterial TMPs, in lieu of most beta-/gammaproteobacterial TMPs, but with low support (81% ufBS) (Fig. 4c). While the simple repetitive primary sequences of coiled-coil proteins make them challenging to analyze phylogenetically46, these results could hint at a potential phage ancestry of crescentin.
Apart from Caulobacteraceae sequences, which formed a distinct clade (100% ufBS), remaining crescentin-like proteins came from 13 bacterial families of the alphaproteobacterial order Hyphomicrobiales (Fig. 4d, Supplementary Fig. S13). Strikingly, the Hyphomicrobiales species included isolates previously shown to have curved morphologies (Fig. 4d, e), such as Methylocystis parva12, Methylocystis sp. Rockwell50, Roseiarcus fermentans51, and Chelatococcus reniformis52, whose crescentin-like protein is possibly deep-branching (90% ufBS) (Fig. 4c). To experimentally test if Hyphomicrobiales crescentin homologs can induce cell curvature, we complemented a non-curved ΔcreS mutant of C. crescentus with the creS homolog of Ch. reniformis. Indeed, creSCh.reniformis expression was sufficient to restore cell curvature in the ΔcreS mutant (Fig. 4f and Supplementary Fig. S14). Expression progressively generated left-handed cell spiraling upon longer incubation—a phenotype characteristic of creS overexpression53—confirming the presence of curvature-inducing crescentin outside of Caulobacteraceae. Together, our results reveal a restricted phylogenetic distribution of crescentin-like proteins across bacteria, being limited to Hyphomicrobiales and Caulobacteraceae (Fig. 4c).
Phototrophy is widespread among Caulobacterales
In contrast to the unpigmented colonies of C. crescentus, some Caulobacteraceae strains produce carotenoid pigments ranging from yellow to reddish orange (Supplementary Data 7 and 8b). When analyzing the distribution of carotenoid biosynthesis genes, we found that many species possessed crtCDF (Supplementary Data 8c)—genes used by photosynthetic bacteria to make carotenoids aiding light capture and electron transfer, and acting as photodamage protectants54,55. This observation, together with recent reports of phototrophic potential among individual Caulobacterales species56,57,58,59, prompted us to systematically investigate the distribution of phototrophy genes across Caulobacterales.
Unexpectedly, we found type II anoxygenic photosynthesis genes in 10% (35/347) of Caulobacterales species representative genomes. This included reaction center (RC) subunits L, M, and H (pufLMH), light-harvesting complex I (LH1) α/β subunits (pufAB), chlorophyll transporters (pucC), and numerous bacteriochlorophyll biosynthesis enzymes (acsF and bch genes) (Fig. 5a,b and Supplementary Data 8d–g). Phototrophy genes always co-occurred with a combination of the aforementioned carotenoid genes crtCDF (Fig. 5b), which were otherwise absent among Caulobacterales (Supplementary Data 8c–g). Furthermore, photosynthesis genes were generally organized in a gene cluster (Fig. 5c and Supplementary Fig. S15), as is typical for Pseudomonadota60.
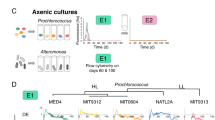
a Schematic illustration of a type II anoxygenic photosystem. LH2 light-harvesting complex II, RC–LH1 reaction center-light harvesting complex I, BChl bacteriochlorophyll, Q ubiquinone/ubiquinol electron carrier, Q red. quinone reductase, NDH NADH dehydrogenase, Cyt. bc1 cytochrome bc1, ∆p proton gradient. Roman numerals indicate orthologs of mitochondrial respiratory chain complexes. b Presence (color) and absence (light gray) of type II anoxygenic photosynthesis genes (KEGG orthologs) among putative phototrophs (top panel) and all remaining genomes averaged over each Caulobacterales family (bottom panel). Species phylogeny shows the tree from Fig. 1a pruned to include only relevant species. See Supplementary Data 4 and 8 for the full dataset. c Synteny of photosynthesis gene clusters (PGCs) from Acaudatibacter gen. nov. phototrophs compared to B. subvibrioides CB81T, showing that LH2 and carbon fixation genes are encoded inside the Acaudatibacter PGCs. Gray boxes link orthologs. Blue boxes highlight sequences aligned using progressiveMauve105 to map the contigs of the incompletely assembled PGC of Ac. boreus against the fully assembled PGC of Ac. lapponiensis. Contigs are marked with “c” followed by the contig number. Color coding corresponds to panel (b). d Colony spots and absorbance scans of colony material from putative phototrophs (bold), as well as non-phototrophic pigmented (CB-RT) and unpigmented (Pickett K-248T and CB15) control strains after 6 days on PYE agar at 23 °C in ambient light. The insert highlights the peaks at 871 nm (major) and 803 nm (minor), characteristic of bacteriochlorophyll a145, for B. variabilis CB17T. Source data are provided as a Source Data file. e The Caulobacterales-containing clade of an ML tree of photosynthetic reaction center subunit PufM. The phylogeny was inferred using IQ-TREE109 with the WAG + C60 + R9 model of evolution and 1000 ultrafast bootstraps (278 sequences with 295 amino acid positions). ufBS is indicated with shaded circles and specified with numbers for key branches. The clade comprising Caulobacterales phototrophs (ufBS = 46%) was pruned from the tree, and unsupported branches (ufBS < 80%) were deleted. The number of sequences per collapsed subclade is shown in parentheses. Uppercase letters represent Caulobacterales families; see panel (b). The asterisk indicates a clade in which a second copy of PufM from Hyphomonas sp. DS1.007 clusters with three Sphingomonadales sequences. The scale bar indicates the number of substitutions per site. The full tree is presented in Supplementary Fig. S17.
We found phototrophic potential in members of all Caulobacterales families, totaling at least 16 genera (Figs. 1d and 5b and Supplementary Data 8). This mostly included uncultured species of previously uncharacterized lineages, but also characterized Aquidulcibacter, Brevundimonas, and Oceanicaulis (Alkalicaulis) species, such as the dark-pigmented Brevundimonas isolates B. bacteroides CB7T, B. subvibrioides CB81T, and B. variabilis CB17T. Absorbance scans of cell suspensions of these strains, when grown aerobically in ambient light, confirmed the production of a-type bacteriochlorophyll in B. variabilis CB17T (Fig. 5d). However, B. bacteroides CB7T and B. subvibrioides CB81T lacked detectable bacteriochlorophyll, suggesting that phototrophy was not induced for these species under the chosen growth conditions.
Notably, we also found phototrophy genes among the three putatively monomorphic Acaudatibacter species of our core dataset (Fig. 5b and Supplementary Fig. S16; see Fig. 2b). In addition to RC–LH1 supercomplex genes, the three Acaudatibacter species also encoded the α and β subunits (pucAB) of the auxiliary light-harvesting complex II (LH2) (Fig. 5a–c and Supplementary Data 8f, g). Unexpectedly, two of the species, Ac. aquilonius and Ac. lapponiensis, had genes for RuBisCO (cbbLS) and the complete Calvin-Benson-Bassham (CBB) cycle for carbon fixation (Fig. 5b and Supplementary Data 8h and 9a, b). This suggests that they might not only capture light energy primarily for ATP synthesis, as is most common for alphaproteobacterial phototrophs (photoheterotrophy)61, but that they can also use light to fix carbon (Fig. 5a). In support of such photoautotrophic metabolism, Acaudatibacter CBB cycle genes were organized inside the photosynthesis gene cluster (PGC) (Fig. 5c), which also included accessory genes for red-type RuBisCO activase (cbbX) and XuBP phosphatase (cbbY). The genome of Ac. boreus, the third Acaudatibacter species, probably also encodes a complete CBB cycle, since we could identify orthologs for all enzymatic steps except for cbbLS (Supplementary Data 9a, b), which was likely incompletely assembled, since part of the cbbL sequence was detected on a contig edge (Fig. 5c). Notably, complete and near-complete CBB cycles were also found among two Vitreimonas phototrophs (Fig. 5b, Supplementary Data 8h and 9a, b). Acaudatibacter PGCs were highly syntenic to other Caulobacteraceae PGCs, with the Acaudatibacter-specific LH2 and carbon fixation genes being organized outside of the syntenic region (Fig. 5c and Supplementary Fig. S15), suggesting related origins of the PGCs despite photosystem complexity differences.
To investigate the evolutionary history of Caulobacterales phototrophy genes, we inferred phylogenies for the reaction center protein PufM and bacteriochlorophyll enzyme BchY. Aside from Maricaulaceae (M), Caulobacterales PufM orthologs clustered together (96% ufBS), with individual genera forming subclades, suggesting close relatedness of PufM across most Caulobacterales (Fig. 5e and Supplementary Fig. S17). The Caulobacterales clade was in turn affiliated (46% ufBS) with a clade comprising sequences from Maricaulaceae (M) and Sphingomonadales (an alphaproteobacterial order) (Fig. 5e and Supplementary Fig. S17). The affiliation with Sphingomonadales was consistent and well-supported (97% ufBS) in another PufM tree (Supplementary Figs. S18a and S19), and matches previous phylogenetic analyses that included B. subvibrioides62. However, in our alternative PufM tree, Hyphomonas (H) and “Parvularculaceae” (P) sequences clustered outside the main Caulobacterales clade (Supplementary Figs. S18a and S19). Similar to PufM, in BchY phylogenies, most Caulobacterales sequences clustered with each other and with Sphingomonadales, although with larger numbers of Caulobacterales sequences branching outside with unsupported positions (Supplementary Figs. S18b, c and S20–S21). While we were unable to further resolve the topology of Caulobacterales PufM/BchY, the patchy but widespread representation of phylogenetically affiliated photosynthesis genes across all major Caulobacterales clades (Fig. 5b, e and Supplementary Data 8a–h), organized in largely syntenic gene clusters (Fig. 5c and Supplementary Fig. S15), could suggest that the ancestor of Caulobacterales was potentially a phototroph. It is also possible that horizontal transfer of the PGC has occurred between different Caulobacterales lineages, followed by vertical inheritance within individual clades, as horizontal transfers are more frequent between closely related lineages63.
Acaudatibacter species with photoautotrophic potential inhabit anoxic layers of stratified boreal lakes in high relative abundance
Given the unusual genetic potential for photoautotrophic metabolism and the lack of dimorphic lifecycle genes in Ac. aquilonius, Ac. boreus, and Ac. lapponiensis, we wondered if this Acaudatibacter lineage inhabits an ecological niche atypical for Caulobacteraceae. To this end, we analyzed a dataset of 24,050 prokaryotic species-cluster genome representatives that have been competitively mapped against 636 freshwater metagenomes (Supplementary Figs. S22a, b)37. In the dataset, Ac. aquilonius, Ac. boreus, Ac. lapponiensis, as well as a fourth species cluster (Ac. sp. 23796) not part of our initial core dataset (Supplementary Fig. S7), were among the most prevalent and relatively abundant Caulobacteraceae freshwater species (Supplementary Fig. S23).
The MAGs of these four Acaudatibacter species derive from short-read metagenomes collected together with water chemistry measurements along the water columns of stratified freshwater bodies64. Genomes of these species recruited reads from metagenomes from twelve boreal freshwater bodies from Northern Sweden, Finland, and Canada (Fig. 6 and Supplementary Figs. S22c and S24). In most cases, the Acaudatibacter species were most abundant right below the oxycline (Fig. 6c). This pattern contrasts with the predominantly aerobic lifestyles of Caulobacteraceae11, but is typical for anoxygenic phototrophs that can be found in the illuminated upper parts of the hypolimnion, where they use reduced compounds found under anoxic conditions as electron donors65. Given these patterns, we suspect that Ac. sp. 23796 could also encode a PGC that was undetected due to genome incompleteness (51.0–92.5% completeness of species cluster members) (Supplementary Fig. S7a). In some sample series, Ac. aquilonius and Ac. sp. 23796 were most abundant in the oxic upper layers (Fig. 6c), suggesting that these species are not obligate anaerobes. Indeed, consistent with a facultatively aerobic lifestyle, all four Acaudatibacter species representatives have the genetic potential for aerobic respiration (Fig. 6a, Supplementary Fig. S16, and Supplementary Data 8i, j). Moreover, the non-homogeneous water column distribution of the Acaudatibacter species suggests that they actively position themselves, despite lacking flagellar genes, perhaps by regulating buoyancy through pilus-mediated cellular aggregation, as previously reported for other anoxygenic phototrophs65.
a Overview of the genetic potential of Acaudatibacter species present in stratified freshwater bodies (based on Figs. 2b and 5b, Supplementary Figs. S8 and S16, and Supplementary Data 9). The genetic potential of C. crescentus is simply included as a point of reference. Question marks emphasize that, given their genome incompleteness (Supplementary Fig. S7a), the genome assemblies of Ac. sp. 23796 could lack photosynthetic potential due to the photosynthesis gene cluster not being assembled. T4P type IV pilus, cyt. cytochrome. b Geographical regions with stratified freshwater bodies containing putatively photoautotrophic Acaudatibacter species in metagenomes from Buck et al.64 Numbers within square brackets denote the number of Acaudatibacter-containing freshwater bodies in the region (see Supplementary Fig. S22c). c Relative abundance of Acaudatibacter species among metagenomes sampled across the water column of stratified freshwater bodies. Dashed lines show O2 concentrations measured during sampling. For each sampling series, the name of the freshwater body, geographical region (parentheses), lake code (italics), and sampling date are indicated. Colored circles represent geographical regions shown in (b). Additional Canadian freshwater bodies with Acaudatibacter presence, but lacking depth profiles, are presented in Supplementary Fig. S24. Source data are provided as a Source Data file.
Discussion
Knowledge about the ecology and evolutionary context of many well-characterized bacterial model organisms is surprisingly limited, restricting our ability to apply insights from experimental studies to the diversity of environmental bacteria. In this study, we provide a comprehensive portrait of the evolution and environmental distribution of the model organism C. crescentus and members of its alphaproteobacterial family Caulobacteraceae and order Caulobacterales. Our work sheds light on the biology of hundreds of previously uncharacterized species and uncovers unexpected diversity within this bacterial order.
C. crescentus has long served as a model for bacterial cell biology6 and has significantly advanced our understanding of bacterial development and morphogenesis. We found that some of the characteristic morphological features of C. crescentus, including its S-layer and crescentin-mediated curved shape, are not common characteristics across Caulobacteraceae and are even absent from other Caulobacterales families (Fig. 4b). However, we identified and experimentally confirmed the presence of crescentin-like proteins in the sister order Hyphomicrobiales (Fig. 4d–f). Crescentin-mediated cell curvature is thought to improve flagellar motility5,46 and pilus-mediated surface colonization66 in C. crescentus, but interestingly, two of the curved crescentin-encoding Hyphomicrobiales species (Ch. reniformis and R. fermentans) lack genes for flagella and pili (Supplementary Fig. S25). Thus, crescentin-mediated curvature might provide yet another selective advantage in these species. Additionally, our phylogenetic analyses could indicate that crescentin has evolved from a prophage protein, either initially in Hyphomicrobiales, later being horizontally transferred to Caulobacteraceae, or in their common ancestor, with extensive loss in the Caulobacterales lineages. While this finding requires further investigation, it would explain the taxonomically limited distribution of crescentin among bacteria (Fig. 4c).
While our data revealed a limited distribution of crescentin and the S-layer, most of the numerous genes important for the motile–sessile dimorphic lifecycle of C. crescentus are present across both Caulobacteraceae and Caulobacterales (Fig. 2b and Supplementary Data 5a–f). However, we identified eleven species from two evolutionarily independent lineages that appear to have lost essentially all genes known to mediate flagellar motility, chemotaxis, and polar holdfast adhesion, alongside over 20 developmental regulators. This includes ten species clusters of the herein described genus Acaudatibacter gen. nov., and P. immobile (genus Phenylobacterium)—the only cultured representative—which we experimentally verified to lack dimorphic reproductive asymmetry (Fig. 3). Thus, our findings uncover the first clear example of a lineage (P. immobile) in which an obligate dimorphic lifecycle has reverted to a simpler monomorphic lifecycle. Moreover, the remarkably similar large absence of genes among the Acaudatibacter species compared to P. immobile strongly suggests convergent loss of dimorphism and reveals core modules specifically used for establishing bacterial dimorphism. The nearly 100 morphogenesis and developmental genes absent from these lineages are distributed across the chromosome, rather than being part of one or a few gene clusters, indicating that their combined absence is the result of numerous mutations driven by considerable evolutionary pressure. Dimorphism is common to the alphaproteobacterial Caulobacterales–Hyphomicrobiales–Rhodobacterales–Sphingomonadales superclade, estimated to have diverged ~1.5 billion years ago67,68, and analogous dimorphic lifecycles are found in diverse bacteria69,70,71,72,73, yet it remains unclear precisely what has driven the evolution of these complex bacterial lifecycles. Understanding the evolutionary processes that revert dimorphic lineages into monomorphs will help clarify the selective advantages and disadvantages of bacterial dimorphism.
It is noteworthy that virtually all regulatory developmental modules that were found to be lost in P. immobile and Acaudatibacter species through convergent evolution are intimately linked to the signaling molecule c-di-GMP (Fig. 2c), a key driver of motile–sessile transitions in bacteria, including the dimorphic lifecycle of C. crescentus42. Yet, we found that these lineages still have multiple genes encoding canonical c-di-GMP enzymes (Supplementary Fig. S10), indicating extensive restructuring of the c-di-GMP signaling network to orchestrate other functions than the dimorphic lifecycle. Precisely how this large rewiring of cellular signaling pathways has been accomplished is an exciting open question that will be worthwhile to address by exploiting the experimental tractability of P. immobile and its close phylogenetic relationship with the well-established model organism C. crescentus.
The new genus Acaudatibacter not only stands out regarding its inferred lifecycle, but also its metabolic potential. In three of the putatively monomorphic Acaudatibacter species, we have identified photosynthesis gene clusters containing CBB cycle genes supporting carbon fixation (Fig. 5c). Together with our finding that these species inhabit both oxic and anoxic layers of stratified freshwater bodies in high relative abundance (Fig. 6 and Supplementary Fig. S23), this suggests possible roles for these bacteria as primary producers—a previously unrecognized role for Caulobacterales. While the Acaudatibacter species stood out with their potential for full photosynthesis, including carbon fixation (Supplementary Fig. S15 and Supplementary Data 8 and 9), we found that phototrophic potential is surprisingly widespread in Caulobacterales (~10% of species across all major clades), most of which remain uncultured (Fig. 1b, d). This discovery is in line with recent reports uncovering phototrophic potential in other groups of environmental bacteria through cultivation-independent comparative genomics74. It is noteworthy that the putative Caulobacterales phototrophs include species with various morphologies and lifecycles, comprising both dimorphic and putatively monomorphic species, with or without prosthecae, even including members of prosthecate budding lineages (Figs. 1a, d and 5b). This raises questions as to how photosystem assembly is coordinated with the dimorphic developmental program. For example, the prosthecae (stalks), which are extensions of the cell envelope that increase cellular membrane surface area, might participate in light capture, similar to the intracellular membrane invaginations produced by classical phototrophic bacteria61. Moreover, our finding that Caulobacterales photosynthesis genes mostly form one supported clade (Fig. 5e and Supplementary Fig. S18) suggests that phototrophy could represent an ancestral Caulobacterales trait maintained in a minority of extant members. Anoxygenic type II photosynthesis has an ancient and complex history in Alphaproteobacteria, which remains unresolved62. Incorporating the rich diversity of Caulobacterales phototrophs into phylogenetic investigations is expected to shed new light on the evolution of bacterial photosynthesis.
Methods
Bacterial strains and growth conditions
Bacterial strains listed in Supplementary Table S2 were routinely grown using peptone yeast extract (PYE; 0.2% (w/v) Bacto peptone, 0.1% (w/v) Bacto yeast extract, 1 mM MgSO4, 0.5 mM CaCl2) or in R2A75 (at pH 7.2, or adjusted to pH 6.0 using 37% HCl before autoclaving), or LB (Carl Roth), as specified. When necessary, media were solidified with 1.5% (w/v) Difco agar (BD). For plasmid selection and maintenance, media were supplemented with gentamicin (15 µg/ml for LB broth, 20 µg/ml for LB agar, 0.625 µg/ml for PYE broth, 5 µg/ml for PYE agar). Liquid cultures were incubated in flat-bottomed E-flasks, with 200 rpm orbital shaking. Unless specified, cultures were incubated at 30 °C. Strain stocks were preserved in 10% DMSO at −80 °C.
Strain construction
Oligonucleotides (synthesized at Eurofins Genomics) are listed in Supplementary Table S3. The creS gene from C. crescentus NA1000 (KJ1) and the creS homolog (WP_188608507.1; IEW34_RS07480) from Ch. reniformis B2974T (KJ1171) were amplified from genomic DNA using primer pairs OJH27/OJH28 and OJH29/OJH30, respectively. The creS fragments were then inserted by Gibson assembly76 into the pBXMCS-4 vector77 linearized by PCR using primers OJH25 and OJH26, creating plasmids pBXMCS-4-Pxyl-creS and pBXMCS-4-Pxyl-creSCh.reniformis. The resulting plasmids and the original pBXMCS-4 empty vector (EV) were transformed into C. crescentus NA1000 (KJ1) and C. crescentus NA1000 ΔcreS (KJ1179) by electroporation.
Microscopy and image analysis
Cells were spotted onto 1% (w/v) agarose pads prepared with appropriate growth media, and phase-contrast images were acquired using an ECLIPSE Ti inverted research microscope (Nikon) equipped with a Plan Apo λ 100× Oil Ph3 DM (1.45 NA) objective (Nikon) and a Zyla sCMOS camera (Andor). For time-lapse imaging, cells were spotted onto growth medium agarose pads prepared inside Gene Frames (65 µl, 1.5 × 1.6 cm; Thermo Fisher) on glass slides, sealed with a glass coverslip, incubated at the specified temperatures, and illuminated only during imaging (a few seconds). For timelapses of C. crescentus, E. coli, and P. immobile, imaging intervals of 30 s, 20 s, and 5 min were used, respectively. Relative constriction site position was determined using the plugin MicrobeJ (v5.13l) in Fiji (ImageJ; v2.1.0/1.53c). Differences in replication timing for daughter cell pairs were calculated from manually tracked timelapses, as outlined in Fig. 3b, by following daughter cells stemming from the first cell division events after being spotted onto the agarose. Time-lapse images (Fig. 3a) and videos (Supplementary Movies 1–3) were subjected to linear stack alignment with SIFT using default settings in Fiji (ImageJ; v1.54 f).
Pigment analyses
For agar spots, material from PYE agar colonies incubated at 23 °C for 6 days was suspended in PYE and diluted to an OD600 of 0.1, before 10 µl was spotted onto a PYE agar plate. The resulting growth patches were photographed after 6 days at 23 °C in ambient light. For in vivo absorbance scans, PYE plate cultures were grown at 23 °C for 6 days in ambient light before cells were suspended in 10 mM Tris-HCl buffer (pH 7.8). To reduce light scattering, the cell suspension was mixed with glycerol in a 3:7 (suspension:glycerol) ratio, before one-nanometer absorbance spectrum scans (200–1000 nm) were recorded using a Spark multimode microplate reader (TECAN) at 23 °C.
Isolation and sequencing of Caulobacter strains
A pre-enrichment of Lake Erken (Norrtälje, Sweden) microorganisms was established in dissolved organic matter produced by Microcystis aeruginosa (cyanobacterium) in a previous study78 on February 28th, 2019. The pre-enrichment was incubated at 20 °C in 12-h light–dark cycles. The pre-enrichment was then diluted a thousand-fold to 1000 cells/ml on April 29th, 2019, for a second round of enrichment with DOM from M. aeruginosa, and incubated under the same conditions. On October 7th, 2021, the presence of prosthecate dimorphic bacteria in the second batch enrichment culture was confirmed by microscopy, and culture liquid was streaked on PYE and incubated in ambient light at 23 °C for 6 days. Seven colonies were re-streaked and incubated for another 6 days, before a well-isolated single colony from each of the resulting seven strains was inoculated into 10 ml PYE broth and incubated at 23 °C with 200 rpm shaking. After three days, cultures were preserved in 10% DMSO at −80 °C. Out of seven strains, three were yellow-pigmented rosette-forming vibrioid prosthecate dimorphic bacteria (strains ErkDOM-C, ErkDOM-E, and ErkDOM-YI). Frozen stocks of these three strains were streaked onto PYE and incubated for 6 days (23 °C) before inoculating single colonies into 25 ml PYE broth, followed by incubation at 23 °C, 200 rpm. Cells were harvested after three days from 8 ml exponentially growing culture (OD600 ≈ 0.2; cultures had been back-diluted with fresh PYE, after two days, to maintain exponential growth), and genomic DNA was extracted using Genomic tips 20/G (Qiagen) as described by Hallgren et al.79 Long-read genome sequencing (Sequel system, Pacific Biosciences) and genome assembly was done at the National Genomics Infrastructure in Uppsala (Sweden). For this, DNA was sheared using Megaruptor 3 (Diagenode), and fragments below 10 kb were removed using a PippinHT (Sage Science). The sequencing library was prepared using the SMRTbell Express Template Prep Kit 2.0 (Pacific Biosciences, USA) and sequenced on the Sequel single-molecule real-time cell platform (8 M v3, 4 cells; Pacific Biosciences, USA). Sequencing of strains ErkDOM-C, ErkDOM-E, and ErkDOM-YI yielded 99,762 reads totaling 2,973,558,276 bp, 120,349 reads totaling 3,547,634,714 bp, and 99,713 reads totaling 2,975,639,866 bp, respectively. The reads were assembled using Flye80 v2.8.3 (default parameters). For strain ErkDOM-C, this resulted in a 4,064,099 bp genome (31× mean coverage depth), comprising a circular chromosome contig (3,918,693 bp) and a circular plasmid contig (145,406 bp). For strain ErkDOM-E, this resulted in a genome comprising a single circular chromosome contig (4,052,756 bp; 42× mean coverage depth). For strain ErkDOM-YI, this resulted in a 4,064,094 bp genome (32× mean coverage depth), comprising a circular chromosome contig (3,918,688 bp) and a circular plasmid contig (145,406 bp). The genomes were made publicly available on GenBank (BioProject accession: PRJNA1228543; assemblies: GCA_048541825.1, GCA_048541895.1, and GCA_048541805.1).
Selection of genomes
An initial set of 768 Caulobacterales genomes was collected as follows (Supplementary Data 1a). First, Caulobacterales MAGs were retrieved from metagenomes sequenced from Lake Erken (Sweden) microbial model community cultures78 (n = 8). Additionally, genomes from Caulobacter strains ErkDOM-C, ErkDOM-E, and ErkDOM-YI (n = 3) isolated from one of the Lake Erken microbial model community cultures were included (see above). Lastly, genome assemblies identified as Caulobacterales in the Genome Taxonomy Database (GTDB) Release 07-RS20781 (genomes released by July 12th, 2021) were retrieved accordingly from NCBI82 RefSeq and GenBank databases in April 2022, with the exception of 20 genomes that were suppressed (n = 747). Of these genomes, 65 corresponded to MAGs annotated as Caulobacterales in the stratfreshDB64 of stratified freshwater metagenomes. Thus, 58% (65/112) of the stratfreshDB Caulobacterales MAGs passed GTDB quality checks. Additional assemblies annotated as Caulobacteraceae and released after the GTDB release were also retrieved from NCBI (n = 10).
The completeness and contamination of these genomes were then estimated using the CheckM83 v1.1.3 “lineage_wf” method and the “taxonomy_wf” method with the Caulobacterales order-specific dataset (except for alphaproteobacterial outgroup genomes, where the alphaproteobacterial class-specific dataset was used), which relies on Prodigal84 for protein prediction and HMMER85 for identifying marker genes. The 573 genomes with an estimated completeness ≥ 90% and contamination ≤ 5% (“taxonomy_wf”) were selected for further analysis (Supplementary Data 1a). The genomes were then de-replicated to obtain species clusters at 95% average nucleotide identity (ANI) using dRep86 v3.2.2 with the option “-sa 0.95”. This resulted in selected genome representatives of 207 Caulobacteraceae species clusters and 140 outgroup Caulobacterales species clusters (including GCF_019455445.1, which was misclassified as Caulobacteraceae in NCBI) (Supplementary Data 1a). Changes were made to the automatically selected genome representatives whereby they were replaced with the species type strain, if one was available and had not been selected as the representative (applies to B. aurantiaca DSM 4731T, B. bullata, HAMBI 262T, B. denitrificans TAR-002T, B. diminuta ATCC 11568T, B. huaxiensis 090558T, B. vesicularis NBRC 12165 T, C. vibrioides DSM 9893T [CB51T], He. mobilis M65T, He. pelagia LA220T, Hy. atlantica 22II1-22F38T, “Hy. pacifica” T16B2T, Hy. polymorpha PS728T, and M. maris DSM 4734T). An additional five Alphaproteobacteria genomes were included as an outgroup. This set of Caulobacterales genomes is here referred to as the “core dataset”
Freshwater metagenome analyses, assigning Acaudatibacter gen. nov. species clusters, and defining the “extended dataset” of genomes
Data for the presence and absence in freshwater metagenomes, as well as relative abundance estimations, of Caulobacteraceae species clusters and Acaudatibacter gen. nov. species clusters, were subset from the dataset of Rodríguez-Gijón et al.37 (Supplementary Data 1b). In short, the dataset comprises 80,561 prokaryotic genomes of medium-to-high-quality (>50% completeness and <5% contamination) according to CheckM83 v1.1.3 “lineage_wf”. The estimated genome size of all 80,561 genomes was calculated by dividing the assembly size by the estimated completeness provided by CheckM, and the genomic guanosine and cytosine (G + C) content was provided by CheckM. All medium-to-high quality genomes were grouped into species clusters at ANI ≥ 95%, using FastANI87 v1.34 and mOTUpan88 v0.3.2, obtaining 24,050 species clusters in total, each being represented by the highest-quality genome of the species cluster (i.e., the species representative). This dataset was then mapped against a dataset of 636 short-read metagenomes from globally distributed freshwater environments using Strobealign89 v0.11.0, detecting the presence of 9028 species in at least one metagenome. Further details can be found in Rodríguez-Gijón et al.37 The subset of Acaudatibacter gen. nov. genomes available in this dataset is here referred to as the “extended dataset” (Supplementary Fig. S7 and Supplementary Data 1b), and was annotated and analyzed further. With regards to which Acaudatibacter gen. nov. genomes are represented, the “extended dataset” principally differs from the “core dataset” of genomes in three regards: (i) it also includes genomes from Nayfach et al.90, (ii) it also includes newly re-binned stratfreshDB64 MAGs (see Rodríguez-Gijón et al.37), and (iii) it had a less stringent genome quality threshold (>50% completeness and <5% contamination, instead of the >95% completeness and <5% contamination threshold used for the “core dataset”).
Genome annotation
Protein sequences (retrieved April 2022) and genome annotations (GenBank flat files; retrieved January 2024) were downloaded from NCBI82 where available. In cases where they were unavailable for newly generated MAGs and genomes, Prokka v.1.14.6 (with the options “--compliant”, “--kingdom Bacteria”, “--gcode 11”, “--addgenes”) was used to call proteins (using Prodigal91). Resulting protein sequences were assigned to KEGG orthologs92 (KOs) using GhostKOALA93 v2.2 (v3.0 was used for annotations presented in Supplementary Fig. S16) and to eggNOG94 v5.0 non-supervised orthologous groups (NOGs), as well as KOs, using eggNOG-mapper (emapper)95 v2.1.5 (v2.1.12 was used for annotations presented in Supplementary Figs. S8, S16, and S25) (with the options “-m diamond” and “--sensmode more-sensitive”). Overviews of NOG and KO presence across genomes were generated using custom scripts (https://github.com/jennahd/anno-utils), and can be found in Supplementary Data 10–12. Pathway completeness was estimated using KEGG Decoder96 v1.3 from GhostKOALA KO predictions (Supplementary Data 13). Protein domains were predicted using InterProScan97 v5.52.-86.0 (with the options “-appl Pfam, TIGRFAM, PANTHER”, “-iprlookup”, and “-pa”). Proteins were further annotated with top hits from NCBI’s nr database82 (April 2022) using DIAMOND98 blastp v2.0.9 (with the options “--max-target-seqs 1”, and “--more-sensitive”). All protein annotations were then compiled using a custom script (https://github.com/jennahd/anno-utils). Reciprocal best blast hits99,100 (RBH) against the C. crescentus CB15 assembly GCF_000006905.1 were determined using blastp101 v2.15.0+ “-max_target_seqs 1 -evalue 1e-3” for protein sequences from each genome using custom scripts (https://github.com/j-hallgren/make-RBH) and can be found in Supplementary Data 14. To map locus IDs for the two C. crescentus laboratory strains CB15 and NA1000 (“CC” and “CCNA” numbers, respectively), RBH was also run for CB15 against the NA1000 genome assembly GCF_000022005.1. Pairwise ANIs were calculated using FastANI87 v1.33 with default options (Supplementary Data 15). Pairwise average amino acid identities (AAI) were calculated using EzAAI102 v1.2.3 ‘extract’ and ‘calculate’ modules with default options (Supplementary Data 16). Genome statistics were calculated using Quast103 v5.0.2, dRep86 v3.2.2 (as outlined above), and CheckM83 v1.1.3 (with the methods “lineage_wf” and “taxonomy_wf”, as outlined above). Gene synteny maps were plotted using the R package gggenomes104 v1.0.0 and edited in Adobe Illustrator 2025 for clarity. For this, gene coordinates were retrieved from GenBank flat files (.gbff) from NCBI, or from the GenBank files (.gbk) provided by Prokka when necessary, and syntenic regions were identified using blastp101 v2.15.0+ (with the options “-evalue 1”, “-task blastp-fast”, and “-max_target_seqs 1”) and through manual curation. For B. subvibrioides ATCC 15264T, the pufB gene was manually drawn based on the coordinates from GCF_000144605.1 assembly version 07/24/2021. progressiveMauve105 (Mauve snapshot_2015-02-25 build 0) was used to align Acaudatibacter PGC contigs.
Species phylogenies
An initial phylogenetic tree was inferred using GToTree106 v1.6.20 with the included set of 117 alphaproteobacterial single-copy marker genes. In short, the pipeline predicted genes with Prodigal84 v2.6.3, then target genes were identified with HMMER385 v3.3.2, aligned individually with Muscle107 v5.1.linux64, trimmed with TrimAl108 v1.4.rev15, and then concatenated into a supermatrix alignment. An ML phylogeny was then inferred using IQ-TREE109 v1.6.12 with the LG + C60 + F + G model of evolution, selected using ModelFinder110 from LG profile mixture models (C10 to C60)111, with or without empirical base frequencies (+F), and with 1000 ultrafast bootstraps112 (Supplementary Fig. S2).
A manually refined ML species phylogeny was then inferred using the AlphaCOG dataset of 72 single-copy marker genes compiled by Martijn et al.18 Protein sequences corresponding to each of the 72 COGs were extracted based on eggNOG NOG annotations. Individual ML gene phylogenies were inferred for each of the 72 marker genes by aligning with MAFFT113 L-INS-i v7.407 (poorly aligned sequences were manually removed), trimming with trimAl108 v1.4.1, and tree inference using IQ-TREE109 v2.2.0 with models selected using ModelFinder110 as outlined above for the initial species phylogeny, but including Gamma distributed (+G) or FreeRates (+R) models. Resulting phylogenies were visualized using a custom script (https://github.com/jennahd/tree-utils), and putative paralogs, contamination, long-branching, horizontal transfers, and duplicate sequences were manually removed (Supplementary Data 17). Individually, gene phylogenies were then inferred again as above and manually inspected. Trimmed alignments were then concatenated into a supermatrix alignment using PhyKIT114 v1.11.7 with the “create_concat” option. A concatenated ML species phylogeny was then inferred as outlined for the individual gene phylogenies. An additional phylogeny was inferred using the posterior mean site frequency115 (PMSF) approximation of the selected model (LG + C60 + F + R) with 100 non-parametric bootstraps (Fig. 1a and Supplementary Fig. S1).
An additional species phylogeny was inferred with the “extended dataset” of putative Acaudatibacter gen. nov. (GTDB taxon “g__Palsa-881”) genomes to confirm their phylogenetic placement and to infer their species relationships (Supplementary Figs. S7a, b, Supplementary Data 1b). The following genomes were included: representatives of the twelve Acaudatibacter species clusters from the “extended dataset” (Supplementary Figs. S7a, b and Supplementary Data 1b), all 54 Caulobacteraceae species type strains of the “core dataset” (Supplementary Data 1a), the 16 species representatives from uncharacterized Caulobacteraceae genus-level clades of the “core dataset”, and three outgroup Caulobacterales species type strains for rooting. A set of 117 alphaproteobacterial single-copy marker genes was retrieved, and a concatenated alignment was generated using GToTree106 v1.8.6, as outlined above. An ML species phylogeny was then inferred using IQ-TREE109 v1.6.12 with the LG + C60 + F + G model of evolution and 1000 ultrafast bootstraps112 (Supplementary Fig. S9).
A species phylogeny was also inferred based on concatenated 16S and 23S rRNA genes to provide further insights into the phylogenetic relationships between Acaudatibacter gen. nov., Caulobacter, and Phenylobacterium. rRNA genes were extracted from both the species genome representatives of the “core” and “extended” datasets (Supplementary Data 18) using Barrnap v0.9 (https://github.com/tseemann/barrnap) with a cutoff of 50% of each gene (“--reject 0.5”), and using bacterial rRNA gene models (“--kingdom bac”). Based on long branches in initial phylogenies, the five alphaproteobacterial outgroup genomes and three genomes (GCA_014763715.1, Rhodobacterales bacterium DE15.006; GCA_016791445.1, Hyphomonadaceae bacterium new MAG-150; GCF_014323565.1, Hyphobacterium sp. CCMP332) with divergent rRNA genes were removed from the analysis. The remaining 16S (n = 220) and 23S (n = 203) rRNA gene sequences were individually aligned using MAFFT E-INS-i113 v7.407 and trimmed with trimAl108 v1.4.1 using the “-gappyout” option. 16S and 23S rRNA genes from the same genome were then concatenated using PhyKIT114 v1.11.7. An ML phylogeny was then inferred using IQ-TREE109 v2.2.2.6 with the GTR + I + R7 model of evolution, selected using ModelFinder110 from GTR models116 and with 1000 ultrafast bootstraps112 (Supplementary Fig. S3).
Gene alignments and phylogenies
Protein sequence alignments of EAL (Pfam domain PF00563) and GGDEF (Pfam domain PF00990) domain-containing proteins (Supplementary Fig. S10) were done using Clustal Omega117 and visualized in Jalview118 ver.2.11.4.1 using the “Clustal” color option. For gene phylogenies, protein sequences were retrieved from the Caulobacterales species representative dataset corresponding to CreS (KEGG KO K18642 and/or Pfam domain PF19220), BchY (KEGG KO K11334 and/or PF00148 (BchYZBN)), and PufM (KEGG KO K08928 (PufL), K08929 (PufM), and/or Pfam domain PF00124). Initial gene phylogenies were then inferred together with corresponding Pfam domain sequences from the Pfam47 database to identify which sequences from the species representatives corresponded to which gene homologs. Sequences were aligned using MAFFT113 L-INS-i v7.407, poorly aligned positions trimmed using trimAl108 v1.4.1 with the “-gt 0.2” option, and ML phylogenies inferred with IQ-TREE109 v2.2.0 using the LG + C10 + G + F model of evolution. Sequences corresponding to each gene family of interest (CreS, BchY, or PufM) were extracted and then searched against NCBI’s nr database82 using DIAMOND98 blastp v2.0.9 with the options “--more-sensitive” and “--max-target-seqs 2000”. Unique hits with greater than 25% sequence identity and 30% coverage were collected using EDirect119 v15.1 “efetch”. Sequence redundancy of hits was reduced to 80% using CD-HIT120 v4.8.1. Sequences were combined with the Pfam and species representative sequences and aligned using FAMSA121 v2.2.2, poorly aligned positions were trimmed using trimAl108 1.4.1 with the “-gt 0.2” option, sequences with less than 30% coverage of the resulting alignment were removed using a custom script (https://github.com/jennahd/tree-utils), and ML phylogenies were inferred with IQ-TREE109 v2.2.0 using the LG + C10 + G + F model of evolution and with 1000 ultrafast bootstraps112. Sequences from nr falling in the clades of interest, together with sequences from Caulobacterales species representatives, were selected (the original non-truncated sequences corresponding to the Pfam domain reference sequences were also captured by the nr search and they were removed from further analyses). Sequences with a “partial” keyword in the sequence header were removed, and alignment was performed using MAFFT113 L-INS-i v7.407. Sequences that were poorly aligned, or that appeared truncated, were manually removed, and alignments were rerun. Trimming was then performed using trimAl108 1.4.1 with both the “-gt 0.5” and “-gappyout” options. ML phylogenies were then inferred using IQ-TREE109 v2.2.0 with model selection using ModelFinder110 from LG, WAG, JTT, and Q.pfam profile mixture models (C20, C40, and C60), and up to 10 FreeRate categories, and with 1000 ultrafast bootstraps112. For CreS, an additional phylogeny was inferred as outlined above using only sequences from the clade with members encoding the Crescentin Pfam protein domain (PF19220). The Q.pfam+C60 + R9, JTT + C60 + R5, WAG + C60 + R10, and WAG + C60 + R9 models of evolution were selected for CreS_ALL (all sequences), CreS_SUBSET (the subset described above), BchY, and PufM, respectively.
Metadata collection and environmental distribution
Sampling metadata were manually collected between May and June 2024 from NCBI BioSample (https://www.ncbi.nlm.nih.gov/biosample/) and BioProject (https://www.ncbi.nlm.nih.gov/bioproject/) databases, as well as from listed publications and the JGI GOLD platform (https://gold.jgi.doe.gov/) when necessary, and can be found in Supplementary Data 2. Genomes were manually classified into simple broad environmental categories, based on the available sampling information. A meta-analysis of previous 16S rRNA gene amplicon studies was done using the Integrated Microbial Next-Generation Sequencing Platform (IMNGS) database25, with a Taxonomy job submitted on April 8, 2022, using the query “Bacteria/Proteobacteria/Alphaproteobacteria/Caulobacterales/Caulobacteraceae”, for which the results can be found in Supplementary Data 3.
Statistics and data visualization
Phylogenetic trees, including protein domains (formatted from InterProScan output using a custom script: https://github.com/jennahd/tree-utils), were visualized with iTOL122 and annotated in Adobe Illustrator 2025. Plots were generated using the R123 v4.4.0 packages ggplot2, gggenomes, and maps in Rstudio124 v2024.04.0.735, GraphPad Prism v10.3.0, and Microsoft Excel for Mac v16.92, and edited in Adobe Illustrator 2025 for clarity.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genome data used in this work is publicly available and was obtained from NCBI RefSeq (https://www.ncbi.nlm.nih.gov/refseq/), NCBI Genbank (https://www.ncbi.nlm.nih.gov/genbank/), the JGI Genome Portal (https://genome.jgi.doe.gov/portal/), Lake Erken model community MAGs (https://doi.org/10.17044/scilifelab.19923161.v1), and from sequenced Lake Erken Caulobacter isolates made available at Genbank (BioProject accession: PRJNA1228543; assemblies: GCA_048541825.1, GCA_048541895.1, and GCA_048541805.1). Additional data supporting the findings of this work is provided in the Supplementary Information, and as a tabular data file containing Supplementary Data 1–4, 6–7, and 10–18, as a PDF file containing Supplementary Data 5, 8 and 9, and as three video files containing Supplementary Movies 1–3. Additional raw data files, including genome annotations, sequence alignments, and phylogenetic tree files, are publicly available at Researchdata.se hosted by the Swedish National Data Service (https://doi.org/10.58141/0bz5-dc62). Source data are provided with this paper.
Code availability
Custom scripts used for genome annotation and phylogenetic analyses are available on GitHub (https://github.com/jennahd/anno-utils and https://github.com/jennahd/tree-utils). Additional custom scripts used to run RBH analysis are available on GitHub (https://github.com/j-hallgren/make-RBH).
Change history
30 March 2026
In the version of Supplementary Data initially published alongside this article, Supplementary Data 5, 8, 9 were missing and are now restored in the HTML version of this article.
References
Jensen, P. A. Ten species comprise half of the bacteriology literature, leaving most species unstudied. Preprint at https://doi.org/10.1101/2025.01.04.631297 (2025).
Callaway, E. These are the 20 most-studied bacteria—the majority have been ignored. Nature 637, 770–771 (2025).
Eren, A. M. & Banfield, J. F. Modern microbiology: embracing complexity through integration across scales. Cell 187, 5151–5170 (2024).
Abraham, W.-R. et al. Phylogeny and polyphasic taxonomy of Caulobacter species. Proposal of Maricaulis gen. nov. with Maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera Brevundimonas and Caulobacter. Int. J. Syst. Evol. Microbiol. 49, 1053–1073 (1999).
Govers, S. K. & Jacobs-Wagner, C. Caulobacter crescentus: model system extraordinaire. Curr. Biol. 30, R1151–R1158 (2020).
Barrows, J. M. & Goley, E. D. Synchronized swarmers and sticky stalks: Caulobacter crescentus as a model for bacterial cell biology. J. Bacteriol. 205, e00384–00322 (2023).
Curtis, P. D. & Brun, Y. V. Getting in the loop: regulation of development in Caulobacter crescentus. Microbiol. Mol. Biol. Rev. 74, 13–41 (2010).
Marks, M. E. et al. The genetic basis of laboratory adaptation in Caulobacter crescentus. J. Bacteriol. 192, 3678–3688 (2010).
Poindexter, J. S. Biological properties and classification of the Caulobacter group. Bacteriol. Rev. 28, 231–295 (1964).
Wilhelm, R. C. Following the terrestrial tracks of Caulobacter - redefining the ecology of a reputed aquatic oligotroph. ISME J. 12, 3025–3037 (2018).
Abraham, W.-R., Rohde, M., Bennasar, A. The family Caulobacteraceae. In: The Prokaryotes 4th edn (eds Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E. & Thompson, F.) Ch. 7 (Springer, Verlag, 2014).
Garrity, G. M., Bell, J. A. & Lilburn, T. Order V. Caulobacterales. In: Bergey’s Manual of Systematic Bacteriology 2 edn (eds Brenner, D. J., Krieg, N. R. & Staley, J. T.) (Springer New York, New York, 2005).
Poindexter, J. S. The caulobacters: ubiquitous unusual bacteria. Microbiol. Rev. 45, 123–179 (1981).
Luo, D. et al. Plant growth promotion driven by a novel Caulobacter strain. Mol. Plant Microbe Interact. 32, 1162–1174 (2019).
Ryan, M. P. & Pembroke, J. T. Brevundimonas spp: emerging global opportunistic pathogens. Virulence 9, 480–493 (2018).
Goris, J. et al. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91 (2007).
Konstantinidis, K. T. & Tiedje, J. M. Genomic insights that advance the species definition for prokaryotes. Proc. Natl. Acad. Sci. USA 102, 2567–2572 (2005).
Martijn, J., Vosseberg, J., Guy, L., Offre, P. & Ettema, T. J. G. Deep mitochondrial origin outside the sampled alphaproteobacteria. Nature 557, 101–105 (2018).
Cai, H., Shi, Y., Wang, Y., Cui, H. & Jiang, H. Aquidulcibacter paucihalophilus gen. nov., sp. nov., a novel member of family Caulobacteraceae isolated from cyanobacterial aggregates in a eutrophic lake. Antonie van. Leeuwenhoek 110, 1169–1177 (2017).
Liu, Y. et al. Pseudaquidulcibacter saccharophilus gen. nov., sp. nov., a novel member of family Caulobacteraceae, isolated from a water purification facility with supplement of starch as a carbon source. Int. J. Syst. Evol. Microbiol. https://doi.org/10.1099/ijsem.0.005264 (2022).
Vieira, S. et al. Terricaulis silvestris gen. nov., sp. nov., a novel prosthecate, budding member of the family Caulobacteraceae isolated from forest soil. Int. J. Syst. Evolut. Microbiol. 70, 4966–4977 (2020).
Asem, M. D. et al. Vitreimonas flagellata gen. nov., sp. nov., a novel member of the family Hyphomonadaceae isolated from an activated sludge sample. Int. J. Syst. Evolut. Microbiol. 70, 2632–2639 (2020).
Tang, Y. et al. Phenylobacterium montanum sp. nov., an oligotrophic, slightly acidophilic mesophile isolated from sandy soil. Int. J. Syst. Evol. Microbiol. https://doi.org/10.1099/ijsem.0.006463 (2024).
Hedlund, B. P. et al. SeqCode: a nomenclatural code for prokaryotes described from sequence data. Nat. Microbiol. 7, 1702–1708 (2022).
Lagkouvardos, I. et al. IMNGS: a comprehensive open resource of processed 16S rRNA microbial profiles for ecology and diversity studies. Sci. Rep. 6, 33721 (2016).
Levy, A. et al. Genomic features of bacterial adaptation to plants. Nat. Genet. 50, 138–150 (2018).
Rodríguez-Gijón, A. et al. A genomic perspective across earth’s microbiomes reveals that genome size in archaea and bacteria is linked to ecosystem type and trophic strategy. Front. Microbiol. 12, 761869 (2022).
Boussau, B., Karlberg, E. O., Frank, A. C., Legault, B.-A. & Andersson, S. G. E. Computational inference of scenarios for α-proteobacterial genome evolution. Proc. Natl. Acad. Sci. USA 101, 9722–9727 (2004).
Tófoli De Araújo, F. et al. Structural and physiological analyses of the alkanesulphonate-binding protein (SsuA) of the citrus pathogen Xanthomonas citri. PLoS ONE 8, e80083 (2013).
Durham, B. P. et al. Cryptic carbon and sulfur cycling between surface ocean plankton. Proc. Natl. Acad. Sci. USA 112, 453–457 (2015).
Warshan, D. et al. Feathermoss and epiphytic Nostoc cooperate differently: expanding the spectrum of plant–cyanobacteria symbiosis. ISME J. 11, 2821–2833 (2017).
Bernal, P., Llamas, M. A. & Filloux, A. Type VI secretion systems in plant-associated bacteria. Environ. Microbiol. 20, 1–15 (2018).
Deng, W. et al. Assembly, structure, function and regulation of type III secretion systems. Nat. Rev. Microbiol. 15, 323–337 (2017).
Porter, S. S., Dupin, S. E., Denison, R. F., Kiers, E. T. & Sachs, J. L. Host-imposed control mechanisms in legume–rhizobia symbiosis. Nat. Microbiol. 9, 1929–1939 (2024).
Linhartová, I. et al. RTX proteins: a highly diverse family secreted by a common mechanism. FEMS Microbiol. Rev. 34, 1076–1112 (2010).
Lingens, F. et al. Phenylobacterium immobile gen. nov., sp. nov., a gram-negative bacterium that degrades the herbicide chloridazon. Int. J. Syst. Bacteriol. 35, 26–39 (1985).
Rodríguez-Gijón, A. et al. The ecological success of freshwater microorganisms is mediated by streamlining and biotic interactions. Sci. Rep. 15, 34482 (2025).
Choi, G.-M., Lee, S.-Y., Choi, K. D. & Im, W.-T. Phenylobacterium hankyongense sp. nov., isolated from ginseng field soil. Int. J. Syst. Evolut. Microbiol. 68, 125–130 (2018).
Li, X. et al. Phenylobacterium soli sp. nov., isolated from arsenic and cadmium contaminated farmland soil. Int. J. Syst. Evol. Microbiol. 69, 1398–1403 (2019).
Wang, H. & Bowman, G. R. SpbR overproduction reveals the importance of proteolytic degradation for cell pole development and chromosome segregation in Caulobacter crescentus. Mol. Microbiol. 111, 1700–1714 (2019).
Mignolet, J. et al. Functional dichotomy and distinct nanoscale assemblies of a cell cycle-controlled bipolar zinc-finger regulator. eLife 5, e18647 (2016).
Jenal, U., Reinders, A. & Lori, C. Cyclic di-GMP: second messenger extraordinaire. Nat. Rev. Microbiol. 15, 271–284 (2017).
Van Teeseling, M. C. F. & Thanbichler, M. Generating asymmetry in a changing environment: cell cycle regulation in dimorphic alphaproteobacteria. Biol. Chem. 401, 1349–1363 (2020).
Terrana, B. & Newton, A. Pattern of unequal cell division and development in Caulobacter crescentus. Dev. Biol. 44, 380–385 (1975).
Bharat, T. A. M. et al. Structure of the hexagonal surface layer on Caulobacter crescentus cells. Nat. Microbiol. 2, 17059 (2017).
Ausmees, N., Kuhn, J. R. & Jacobs-Wagner, C. The bacterial cytoskeleton: an intermediate filament-like function in. Cell Shape. Cell 115, 705–713 (2003).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Wang, C. et al. Architecture of the bacteriophage lambda tail. Structure 32, 35–46.e33 (2024).
Liu, Y., van den Ent, F. & Löwe, J. Filament structure and subcellular organization of the bacterial intermediate filament–like protein crescentin. Proc. Natl. Acad. Sci. 121, e2309984121 (2024).
Lazic, M. et al. In vivo quantification of polyhydroxybutyrate (PHB) in the alphaproteobacterial methanotroph, Methylocystis sp. Rockwell. Appl. Microbiol. Biotechnol. 106, 811–819 (2022).
Kulichevskaya, I. S., Danilova, O. V., Tereshina, V. M., Kevbrin, V. V. & Dedysh, S. N. Descriptions of Roseiarcus fermentans gen. nov., sp. nov., a bacteriochlorophyll a-containing fermentative bacterium related phylogenetically to alphaproteobacterial methanotrophs, and of the family Roseiarcaceae fam. nov. Int. J. Syst. Evol. Microbiol. 64, 2558–2565 (2014).
Gu, Z. et al. Chelatococcus reniformis sp. nov., isolated from a glacier. Int. J. Syst. Evol. Microbiol. 66, 4525–4529 (2016).
Cabeen, M. T. et al. Bacterial cell curvature through mechanical control of cell growth. EMBO J. 28, 1208–1219 (2009).
Koblízek, M., Zeng, Y., Horák, A. & Oborník, M. Regressive evolution of photosynthesis in the Roseobacter clade. In: Advances in Botanical Research (ed Beatty JT) Ch 13 (Academic Press, 2013).
Liu, L.-N., Bracun, L. & Li, M. Structural diversity and modularity of photosynthetic RC−LH1 complexes. Trends Microbiol. 32, 38–52 (2024).
Nagashima, S. & Nagashima, K. V. P. Comparison of photosynthesis gene clusters retrieved from total genome sequences of purple bacteria. In: Advances in Botanical Research (ed Beatty, J. T.) Ch 5 (Academic Press, 2013).
Tanabe, Y., Yamaguchi, H. & Watanabe, M. M. Draft Genome Sequence of “Candidatus Phycosocius bacilliformis,” an alphaproteobacterial ectosymbiont of the hydrocarbon-producing green alga Botryococcus braunii. Genome Announc. 6, e00396–00318 (2018).
Kuzyk, S. B. et al. Prosthecate aerobic anoxygenic phototrophs Photocaulis sulfatitolerans gen. nov. sp. nov. and Photocaulis rubescens sp. nov. isolated from alpine meromictic lakes in British Columbia, Canada. Arch. Microbiol. 204, 444 (2022).
Tanabe, Y., Yamaguchi, H., Yoshida, M., Kai, A. & Okazaki, Y. Characterization of a bloom-associated alphaproteobacterial lineage, ‘Candidatus Phycosocius’: insights into freshwater algal-bacterial interactions. ISME Commun. 3, 20 (2023).
Brinkmann, H., Göker, M., Koblížek, M., Wagner-Döbler, I. & Petersen, J. Horizontal operon transfer, plasmids, and the evolution of photosynthesis in Rhodobacteraceae. ISME J. 12, 1994–2010 (2018).
Hiraishi, A. Biodiversity and taxonomy. In: Anoxygenic Phototrophic Bacteria (eds Shimada K., Takaichi S.) Ch. 2 (Academic Press, 2024).
Imhoff, J. F., Rahn, T., Künzel, S. & Neulinger, S. C. Phylogeny of anoxygenic photosynthesis based on sequences of photosynthetic reaction center proteins and a key enzyme in bacteriochlorophyll biosynthesis, the chlorophyllide reductase. Microorganisms 7, 576 (2019).
Soucy, S. M., Huang, J. & Gogarten, J. P. Horizontal gene transfer: building the web of life. Nat. Rev. Genet. 16, 472–482 (2015).
Buck, M. et al. Comprehensive dataset of shotgun metagenomes from oxygen stratified freshwater lakes and ponds. Sci. Data 8, 131 (2021).
Van Gemerden, H. & Mas, J. Ecology of phototrophic sulfur bacteria. In: Anoxygenic Photosynthetic Bacteria (eds Blankenship, R. E., Madigan, M. T. & Bauer C. E.) (Springer Netherlands, 1995).
Persat, A., Stone, H. A. & Gitai, Z. The curved shape of Caulobacter crescentus enhances surface colonization in flow. Nat. Commun. 5, 3824 (2014).
Hallgren, J. & Jonas, K. Diversity and evolution of alphaproteobacterial dimorphism. Curr. Opin. Microbiol. 88, 102661 (2025).
Wang, S. & Luo, H. Dating Alphaproteobacteria evolution with eukaryotic fossils. Nat. Commun. 12, 3324 (2021).
Volland, J.-M. et al. A centimeter-long bacterium with DNA contained in metabolically active, membrane-bound organelles. Science 376, 1453–1458 (2022).
Bailey, J. V. et al. Dimorphism in methane seep-dwelling ecotypes of the largest known bacteria. ISME J. 5, 1926–1935 (2011).
Wiegand, S., Jogler, M. & Jogler, C. On the maverick planctomycetes. FEMS Microbiol. Rev. 42, 739–760 (2018).
Wiegand, S. et al. Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology. Nat. Microbiol. 5, 126–140 (2019).
Nkamba, I. et al. Intracellular offspring released from SFB filaments are flagellated. Nat. Microbiol. 5, 34–39 (2019).
Li, L. et al. Globally distributed Myxococcota with photosynthesis gene clusters illuminate the origin and evolution of a potentially chimeric lifestyle. Nat. Commun. 14, 6450 (2023).
Reasoner, D. J. & Geldreich, E. E. A new medium for the enumeration and subculture of bacteria from potable water. Appl. Environ. Microbiol. 49, 1–7 (1985).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Thanbichler, M., Iniesta, A. A. & Shapiro, L. A comprehensive set of plasmids for vanillate- and xylose-inducible gene expression in Caulobacter crescentus. Nucleic Acids Res. 35, e137–e137 (2007).
Garcia, S. L. et al. Taxonomic and functional diversity of aquatic heterotrophs is sustained by dissolved organic matter chemodiversity. Preprint at https://doi.org/10.1101/2022.03.21.485019 (2022).
Hallgren, J., Daneby, F. & Jonas, K. Complete genome sequences of the prosthecate dimorphic bacteria Brevundimonas intermedia CB63T and Brevundimonas staleyi FWC43T. Microbiol. Resour. Announc. 13, e0105024 (2024).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2021).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information in 2023. Nucleic Acids Res. 51, D29–D38 (2022).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Hyatt, D., LoCascio, P. F., Hauser, L. J. & Uberbacher, E. C. Gene and translation initiation site prediction in metagenomic sequences. Bioinformatics 28, 2223–2230 (2012).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Buck, M., Mehrshad, M. & Bertilsson, S. mOTUpan: a robust Bayesian approach to leverage metagenome-assembled genomes for core-genome estimation. NAR Genomics Bioinforma. 4, lqac060 (2022).
Sahlin, K. Strobealign: flexible seed size enables ultra-fast and accurate read alignment. Genome Biol. 23, 260 (2022).
Nayfach, S. et al. A genomic catalog of Earth’s microbiomes. Nat. Biotechnol. 39, 499–509 (2021).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119 (2010).
Kanehisa, M. Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Graham, E. D., Heidelberg, J. F. & Tully, B. J. Potential for primary productivity in a globally-distributed bacterial phototroph. ISME J. 12, 1861–1866 (2018).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Rivera, M. C., Jain, R., Moore, J. E. & Lake, J. A. Genomic evidence for two functionally distinct gene classes. Proc. Natl. Acad. Sci. USA 95, 6239–6244 (1998).
Hirsh, A. E. & Fraser, H. B. Protein dispensability and rate of evolution. Nature 411, 1046–1049 (2001).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinforma. 10, 421 (2009).
Kim, D., Park, S. & Chun, J. Introducing EzAAI: a pipeline for high throughput calculations of prokaryotic average amino acid identity. J. Microbiol. 59, 476–480 (2021).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Hackl, T., Ankenbrand, M., van Adrichem, B., Wilkins, D. & Haslinger K. gggenomes: effective and versatile visualizations for comparative genomics. Preprint at http://arxiv.org/abs/2411.13556 (2024).
Darling, A. E., Mau, B. & Perna, N. T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5, e11147 (2010).
Lee, M. D. GToTree: a user-friendly workflow for phylogenomics. Bioinformatics 35, 4162–4164 (2019).
Edgar, R. C. High-Accuracy Alignment Ensembles Enable Unbiased Assessments of Sequence Homology and Phylogeny (Cold Spring Harbor Laboratory, 2021).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2014).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Si Quang, L., Gascuel, O. & Lartillot, N. Empirical profile mixture models for phylogenetic reconstruction. Bioinformatics 24, 2317–2323 (2008).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Steenwyk, J. L. et al. PhyKIT: a broadly applicable UNIX shell toolkit for processing and analyzing phylogenomic data. Bioinformatics 37, 2325–2331 (2021).
Wang, H.-C., Minh, B. Q., Susko, E. & Roger, A. J. Modeling site heterogeneity with posterior mean site frequency profiles accelerates accurate phylogenomic estimation. Syst. Biol. 67, 216–235 (2018).
Tavaré, S. Some probabilistic and statistical problems in the analysis of DNA sequences. Lectures Math. Life Sci. 17, 57–86 (1986).
Madeira, F. et al. The EMBL-EBI Job Dispatcher Sequence analysis tools framework in 2024. Nucleic Acids Res. 52, W521–W525 (2024).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Kans J. Entrez direct: E-utilities on the Unix Command Line. In: Entrez Programming Utilities Help [internet] https://www.ncbi.nlm.nih.gov/books/NBK179288/ (2013).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Deorowicz, S., Debudaj-Grabysz, A. & Gudyś, A. FAMSA: Fast and accurate multiple sequence alignment of huge protein families. Sci. Rep. 6, 33964 (2016).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
R Core Team. R: a Language and Environment for Statistical Computing 4.4.0 (2024-04-24) edn (R Foundation for Statistical Computing, 2024).
Posit Team. RStudio: Integrated Development Environment for R (Posit Software, PBC, Boston, 2024).
Parte, A. C., Sardà Carbasse, J., Meier-Kolthoff, J. P., Reimer, L. C. & Göker, M. List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int. J. Syst. Evolut. Microbiol. 70, 5607–5612 (2020).
Abel, S. et al. Bi-modal distribution of the second messenger c-di-GMP controls cell fate and asymmetry during the Caulobacter cell cycle. PLoS Genet. 9, e1003744 (2013).
Laub, M. T., Chen, S. L., Shapiro, L. & McAdams, H. H. Genes directly controlled by CtrA, a master regulator of the Caulobacter cell cycle. Proc. Natl. Acad. Sci. USA 99, 4632–4637 (2002).
Viollier, P. H. & Shapiro, L. A lytic transglycosylase homologue, PleA, is required for the assembly of pili and the flagellum at the Caulobacter crescentus cell pole. Mol. Microbiol. 49, 331–345 (2003).
Ohta, N. & Newton, A. The core dimerization domains of histidine kinases contain recognition specificity for the cognate response regulator. J. Bacteriol. 185, 4424–4431 (2003).
Biondi, E. G. et al. A phosphorelay system controls stalk biogenesis during cell cycle progression in Caulobacter crescentus. Mol. Microbiol. 59, 386–401 (2006).
Christen, M. et al. DgrA is a member of a new family of cyclic diguanosine monophosphate receptors and controls flagellar motor function in Caulobacter crescentus. Proc. Natl. Acad. Sci. USA 104, 4112–4117 (2007).
Radhakrishnan, S. K., Thanbichler, M. & Viollier, P. H. The dynamic interplay between a cell fate determinant and a lysozyme homolog drives the asymmetric division cycle of Caulobacter crescentus. Genes Dev. 22, 212–225 (2008).
Fiebig, A. et al. A cell cycle and nutritional checkpoint controlling bacterial surface adhesion. PLoS Genet. 10, e1004101 (2014).
Joshi, K. K., Bergé, M., Radhakrishnan, S. K., Viollier, P. H. & Chien, P. An adaptor hierarchy regulates proteolysis during a bacterial cell cycle. Cell 163, 419–431 (2015).
Lasker, K., Mann, T. H. & Shapiro, L. An intracellular compass spatially coordinates cell cycle modules in Caulobacter crescentus. Curr. Opin. Microbiol. 33, 131–139 (2016).
Janakiraman, B., Mignolet, J., Narayanan, S., Viollier, P. H. & Radhakrishnan, S. K. In-phase oscillation of global regulons is orchestrated by a pole-specific organizer. Proc. Natl. Acad. Sci. USA 113, 12550–12555 (2016).
Perez, A. M. et al. A localized complex of two protein oligomers controls the orientation of cell polarity. mBio 8, e02238–02216 (2017).
Coppine, J. et al. Regulation of bacterial cell cycle progression by redundant phosphatases. J. Bacteriol. 202, e00345–20 (2020).
Leicht, O. et al. Integrative and quantitative view of the CtrA regulatory network in a stalked budding bacterium. PLOS Genet. 16, e1008724 (2020).
Kaczmarczyk, A. et al. Precise timing of transcription by c-di-GMP coordinates cell cycle and morphogenesis in Caulobacter. Nat. Commun. 11, 816 (2020).
Hershey, D. M., Fiebig, A. & Crosson, S. Flagellar perturbations activate adhesion through two distinct pathways in Caulobacter crescentus. mBio 12, e03266–20 (2021).
Guzzo, M., Sanderlin, A. G., Castro, L. K. & Laub, M. T. Activation of a signaling pathway by the physical translocation of a chromosome. Dev. Cell 56, 2145–2159.e7 (2021).
Lu, N. et al. Scaffold-scaffold interaction facilitates cell polarity development in Caulobacter crescentus. mBio 14, e03218–e03222 (2023).
Zappa, S. et al. The HmrABCX pathway regulates the transition between motile and sessile lifestyles in Caulobacter crescentus by a mechanism independent of hfiA transcription. mBio 15, e0100224 (2024).
Nagashima K. Light-harvesting systems. In: Anoxygenic Phototrophic Bacteria (eds Shimada, K. & Takaichi, S.) Ch. 7 (Academic Press, 2024).
Acknowledgements
We thank Emilie Falconer for assistance with microscopy, as well as Christine Jacobs-Wagner and Martin Thanbichler for sharing strains and plasmids. Genomic analyses and phylogenetic inferences were enabled through resources provided by the Swedish National Infrastructure for Computing (SNIC) at UPPMAX through the storage project SNIC 2021/6-99 and the computation project SNIC 2022/5-137, partially funded by the Swedish Research Council through grant agreement no. 2018-05973. We acknowledge support of the National Genomics Infrastructure (NGI)/Uppsala Genome Center and UPPMAX for providing assistance in massive parallel sequencing and computational infrastructure. Work performed at NGI/Uppsala Genome Center has been funded by RFI/VR and Science for Life Laboratory, Sweden. The work was financially supported by major funding from the Strategic Research Areas (SFO) program/Science for Life Laboratory distributed through Stockholm University to K.J. and S.L.G., and project grants from Stockholm University–Region Stockholm (FoUI-992787) and the Swedish Research Council (2020-03545; 2024-04942) to K.J. The work was further supported by an international postdoc grant from the Swedish Research Council (VR grant 2022-06250) and a non-stipendiary EMBO Long Term Fellowship (ALTF 740-2022) to J.E.D.
Funding
Open access funding provided by Stockholm University.
Author information
Authors and Affiliations
Contributions
J.H.: conceptualization, methodology, software, formal analysis, investigation (all experimental work, comparative genomics, taxonomic descriptions, and meta-analyses), data curation, writing—original draft, writing—review and editing, and visualization. J.E.D.: conceptualization, methodology, software, formal analysis, investigation (all phylogenetic analyses), data curation, writing—review and editing, and funding acquisition. A.R.-G.: formal analysis, investigation (freshwater metagenome analyses), and writing—review and editing. J.N.: conceptualization and investigation. S.G.: conceptualization, resources, writing—review and editing, supervision, project administration, and funding acquisition. K.J.: conceptualization, resources, writing—original draft, writing—review and editing, supervision, project administration, and funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Roland Wilhelm and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hallgren, J., Dharamshi, J.E., Rodríguez-Gijón, A. et al. Widespread potential for phototrophy and convergent reduction of lifecycle complexity in the dimorphic order Caulobacterales. Nat Commun 16, 11003 (2025). https://doi.org/10.1038/s41467-025-65642-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65642-x