Abstract
The transformation of a two-dimensional epithelial sheet into various three-dimensional structures is a critical process in generating the diversity of animal forms. Previous studies of epithelial folding have revealed diverse mechanisms driven by epithelium-intrinsic or -extrinsic forces. Yet little is known about the biomechanical basis of epithelial splitting, which involves extreme folding and eventually a topological transition breaking the epithelial tube. Here, we leverage tracheal-esophageal separation (TES), a critical and highly conserved morphogenetic event during tetrapod embryogenesis, as a model system for interrogating epithelial tube splitting. We identify an evolutionarily conserved, compressive force exerted by the mesenchyme surrounding the epithelium, as being necessary to drive epithelial constriction and splitting. The compressive force is mediated by localized convergent flow of mesenchymal cells towards the epithelium. Sonic hedgehog (SHH) secreted by the epithelium functions as an attractive cue for mesenchymal cells. Removal of the mesenchyme, inhibition of cell migration, or loss of SHH signaling all abrogate TES, which can be rescued by externally applied pressure. These results unveil the biomechanical basis of epithelial splitting and suggest plausible mesenchymal origins of tracheal-esophageal birth defects.
Similar content being viewed by others
Introduction
The epithelium, a fundamental tissue type across all metazoans, can adopt morphologies ranging from simple, two-dimensional sheets to highly complex, three-dimensional scaffolds, giving rise to the characteristic forms and thus functions of different organs. How the epithelium folds during the morphogenesis of these organs has been a central question in mechanobiology1,2,3. We now know that epithelial folding can be driven by both intrinsic and extrinsic mechanisms. Epithelium-intrinsic forces, such as those generated by apical/basal constriction or directional cell migration, are crucial for many epithelial morphogenetic events, including gastrulation and neural tube closure1,3,4. Conversely, non-epithelial tissue can also generate force to shape the adjacent epithelium, for example, through differential proliferation or cell clustering2,5,6,7.
Despite these growing insights into epithelial folding, very little is known about the extreme case in which epithelial tube deformation reaches a state where closely apposed folded segments form a new junction. A transient junction may relax without changing the topology of the epithelium, whereas a sustained junction can potentially lead to reorganization of epithelial cell polarity, resulting in a topological transition that breaks the continuity of the epithelial tube, which we here define as epithelial splitting. This is seen in tracheal-esophageal separation/septation (TES) in tetrapods, as well as cloacal septation in mammals, in both cases involving the splitting of an epithelial tube into two tubes giving rise to different organs8,9,10. Although the morphological and genetic aspects of TES have been extensively studied, very few studies have examined the biomechanical basis of epithelial splitting in this intriguing system, and the biophysical forces driving the formation and resolution of the epithelial junction remain unclear11.
TES starts with the dorsal-ventral patterning of the foregut epithelium into esophageal and tracheal progenitors at Embryonic Day 9.5 (E9.5) in mouse embryos, mainly attributed to ventral-to-dorsal gradients of BMP and Wnt signaling and ephrin-mediated cell sorting8,9,12,13. Over the next two days in development, the foregut epithelium constricts bilaterally and forms a medial junction (septum) where cells transiently lose and re-establish their apical-basal polarity, resolving the septum to generate the esophagus dorsally and the trachea ventrally (Fig. 1a)14,15. TES is highly conserved in tetrapods, and defects in TES cause a spectrum of congenital tracheal-esophageal malformations impacting 1 in 2500 humans14,15,16. Using mouse and frog embryos as models, previous studies have identified a number of genetic factors essential for the epithelial patterning, sorting, and septal resolution11,13,14,15,16,17,18,19,20,21. So far, there are more than 50 genes associated with tracheal-esophageal birth defects in humans and ~15 genes whose loss-of-function mutations lead to defects in mouse TES14. Among these genes, some are expressed in the epithelium, whereas others are expressed in the surrounding mesenchyme, and some genes do not have an obvious connection to the morphogen gradients patterning the epithelium. It remains an open question whether TES is driven by epithelium-intrinsic forces or requires additional force from the mesenchyme to achieve its extremely folded geometry and to enable splitting. Addressing this question will not only help explain the etiology of TES-related birth defects but also deepen our understanding of the biophysical mechanisms driving epithelial splitting.
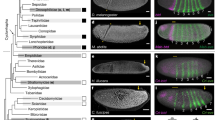
a Schematic of TES. The airway (purple) and the esophagus (green) form through foregut splitting. The inset shows the cross-section of the foregut at the level of TES along the dashed line. The inset was created in BioRender. Yan, R. (2025) https://BioRender.com/cdwttt9. b Whole mount immunofluorescence of E-cadherin (CDH1) of chick and mouse foreguts during TES. Images show lateral views from the right. The red and blue line segments indicate the lengths of the undivided foregut and the trachea. c Quantification of the lengths of the undivided foregut (red) and the trachea (blue) during TES from whole mount images. N = 4 biological replicates for each stage. Data are presented as mean ± standard deviation (SD). d Immunofluorescence of SOX2 and NKX2-1 in transverse sections of the chick and mouse foreguts during TES. e Quantification of SOX2 intensity along the dorsal-ventral axis (N = 6 biological replicates). Data are presented as mean ± SD. f, g Immunofluorescence of cleaved Caspase 3 (CC3) (f) and phospho-Histone H3 (pHH3) (g) in transverse sections of chick and mouse TES. Arrows point to the tracheal-esophageal septum. h Immunofluorescence of phospho-MLC2 (pMLC) in serial transverse sections of chick and mouse foreguts, from the anterior undivided foregut through the posterior separated foregut. i Quantification of the apical-basal ratio of pMLC intensity in the dorsal, medial, and ventral foregut epithelium (N = 7 biological replicates for each species). Data are presented as mean ± SD. Scale bars: 100 µm (b), 50 µm (d, f–h). Double arrows indicate the dorsal-ventral axes for tilted samples.
To this end, we comparatively studied TES in the chick and the mouse to identify evolutionarily conserved biophysical mechanisms of epithelial splitting. Far less prior work has been done on chick TES8. We found it to be morphologically similar to mouse TES, making it feasible to extract evolutionarily conserved mechanisms from the two systems (Fig. 1b). To enable access to biophysical force measurements, we focused on analyzing live tissues and established an ex vivo slice culture system in which TES can be visualized in real-time, pharmacologically treated, and surgically manipulated. We found that although chick and mouse TES differ in many aspects, such as epithelial morphology and cytoskeletal structure, both species rely on forces generated by the mesenchyme to facilitate epithelial constriction, which itself is also sufficient to induce septation. We further discovered that the mesenchymal cells flow convergently towards the epithelium, causing the compressive stresses in both species. Sonic hedgehog (SHH) secreted by the epithelium endogenously attracts surrounding mesenchymal cells and modulates the epithelium’s shape. These results demonstrate the essential role of mesenchymal force in epithelial splitting and suggest that defects in mesenchymal cell migration in response to SHH signaling are a plausible mechanism of tracheal-esophageal malformations.
Results
Morphological and molecular comparison of chick and mouse TES
We first thoroughly characterized chick TES in vivo, such that it can be spatiotemporally aligned to the better-studied mouse system. Chick lung buds emerge at ~E3.0, or Hamburger Hamilton (HH) stage 17–18, and then elongate posteriorly. TES initiates shortly after at E3.5 (HH19), from the lung budding site moving anteriorly, until the TES septum reaches the larynx at E4.5 (HH23–24) (Fig. 1a, b, Supplementary Movie 1). The undivided anterior foregut shortens by ~150 µm in 1.5 days, a length equal to the nascent trachea (Fig. 1c). A similar rate of foregut shortening is observed in mouse TES, but the mouse trachea exhibits additional elongation (Fig. 1b, c, Supplementary Movie 2). The foregut epithelia in both species are pseudostratified (Supplementary Fig. 1a). The foregut morphology is dorsoventrally asymmetrical because of the gradients of BMP and Wnt signaling. Molecularly, the ventral foregut (prospective trachea) of the chick embryo is marked by NKX2-1, and the dorsal foregut (prospective esophagus) is marked by SOX2 (Fig. 1d), markers previously described in the context of mouse TES17,22. In mice, SOX2 is a ubiquitous marker of the foregut epithelium before E9.5, which is then transiently downregulated in the ventral foregut by NKX2.1 between E9.5 and E11.5, coinciding with TES18. Of note, the dorsal-ventral gradient of chick SOX2 is less pronounced than the mouse (Fig. 1e), implying that NKX2-1 may not strongly inhibit SOX2 in the chick as they have been reported to be in the mouse17,19,23. At the septum, many epithelial cells undergo apoptosis, which is rarely observed outside this region (Fig. 1f). Cell proliferation is also halted at the septum in both species, whereas it appears homogenous across the mesenchyme (Fig. 1g), consistent with what has been reported11,24.
We further probed the molecular patterns pertinent to tissue biomechanics, including localization of phosphorylated myosin light chain (p-MLC), F-actin, and extracellular matrix components. In the chick, p-MLC and F-actin are highly enriched on the apical side of the epithelium outside the septum, in which cells lose their polarity (Fig. 1h, Supplementary Fig. 1b). This pattern suggests strong apical constriction in the chick epithelium. By contrast, p-MLC is not highly enriched apically in the mouse foregut (Fig. 1h), despite an apical concentration of actin (Supplementary Fig. 1b). The ratio of apical to basal p-MLC, an indirect measure of apical constriction, indicates a weaker epithelium-intrinsic constrictive force in the mouse than in the chick (Fig. 1i). Electroporation of tdTomato-tagged MLC2 into the chick foregut epithelium revealed a less elongated epithelial morphology in cells undergoing tube splitting compared to the cells outside this region (Supplementary Fig. 1c, d). Hyaluronic acid, which is known for generating fluid pressure within the tissue25, shows a dorsal-ventral gradient in the chick but a homogeneous distribution in the mouse (Supplementary Fig. 1e). Fibronectin, a stiffer component of the extracellular matrix, is ubiquitously expressed in the mesenchyme in both species (Supplementary Fig. 1f), These results suggest that although chick and mouse TES are morphologically similar, they exhibit significant differences related to tissue biomechanics at the molecular and cellular scales. It thus became imperative to examine the tissue-level biomechanics to understand this conserved morphogenetic program.
Evolutionarily conserved, constrictive mesenchymal force deforms the epithelium during TES
We cut freshly dissected foreguts transversely into ~100 µm-thick slices to obtain a cross-sectional view of epithelial splitting in live tissue (Fig. 2a). To probe the force distribution at the epithelium, we surgically removed the surrounding mesenchyme and analyzed the epithelial deformation (Fig. 2a). Intriguingly, the epithelium expanded immediately after mesenchymal removal, indicating that the epithelium was being compressed by the mesenchyme (Fig. 2a, Supplementary Fig. 2a, b). The expansion was more pronounced mediolaterally than dorsoventrally. The results were consistent in the chick and the mouse, and a similar pattern of epithelial deformation was previously reported in Xenopus foregut11, implying that such constrictive mesenchymal force is evolutionarily conserved.
a Stereoscope imaging of the transverse slice of an E3.75 GFP chick foregut before (magenta) and after (green) surgical removal of the mesenchyme. b Two-photon laser ablation of an E3.75 GFP chick foregut slice in the medial or dorsal sub-epithelial mesenchyme (magenta). c Kymographs along the dotted lines in (b). The magenta arrow indicates the initial displacement rate. The green line and double arrows indicate the total epithelial displacement at the interface. The light blue dashed double arrows mark the epithelial thickness. d Quantifications of dorsal and medial ablation measurements as in (c). N = 4 biological replicates. Data are presented as mean ± SD. P-values: Two-tailed Welch’s t-test. Scale bars: 100 µm (a), 50 µm (b).
We further confirmed this finding with localized laser ablation of the sub-epithelial mesenchyme. When a fraction of the medial mesenchyme was ablated, the outer edge of the epithelium moved towards the ablated site, reminiscent of the bulk tissue removal assay (Fig. 2b, c, Supplementary Fig. 2b, c and Supplementary Movies 3–5). By comparison, we observed no significant epithelial deformation when the dorsal mesenchyme was ablated (Fig. 2b–d, Supplementary Fig. 2b–d). Quantitative analyses of the initial displacement rate and total displacement suggest that a compressive mesenchymal force existed near the medial region of the epithelium, whose loss led to an outward movement of the interface mainly contributed by the thickening of the epithelium (Fig. 2d, Supplementary Fig. 2d). Altogether, we conclude that the mediolateral constriction by the mesenchyme is an evolutionarily conserved feature of TES.
The mesenchymal force is essential for TES ex vivo
Having demonstrated the presence of a constrictive mesenchymal force, we next asked whether this force is functionally important to TES morphogenesis. This requires a system allowing real-time monitoring of TES as well as experimental perturbations. To that end, we developed an ex vivo culture of foregut slices in Matrigel, which, for at least 24 h, preserved the dorsoventral patterning of SOX2 and NKX2-1 and the spatial distribution of cell proliferation (Fig. 3a, Supplementary Fig. 3a, b). There was detectable, but insubstantial, apoptosis in the ex vivo culture (Supplementary Fig. 3c). Importantly, the slice culture recapitulates TES morphogenesis in vivo for both chick and mouse, which can be visualized by two-photon live imaging (Fig. 3b, Supplementary Movies 6, 7). The decay of image intensity over long periods of imaging appears to be due to photobleaching. The observed morphological changes were not due to the focusing plane shift during acquisition, as we could track individual 200-nm fluorescent beads embedded with the slice for more than 12 h (Supplementary Fig. 3d, e, Supplementary Movie 8), and consistent morphological changes were observed across different imaging planes (Supplementary Fig. 3f). To quantitatively characterize the morphological parameters of the epithelium, we used a machine learning-based automated segmentation pipeline to extract epithelial morphology from the time-lapse imaging data (Supplementary Movie 6, see also “Methods”). The epithelium narrows to create negative curvatures in the medial region. The distance between the bilateral curvature minima, defined as the neck width, continuously decreases during TES, whereas the area of the epithelium exhibits minimal changes (Fig. 3a, c, d). The dorsoventral length of the epithelium thus increases, and we use the dimensionless ratio of the epithelial perimeter to the square root of the area (perimeter-area ratio) to indicate the elongation of the epithelium (Fig. 3e). As the epithelium narrows during TES, the neck curvature also decreases monotonically (Fig. 3f). Although TES at different time points have distinct epithelial morphologies (Fig. 3b, Supplementary Fig. 4), the shape evolution in the neck width and the perimeter-area ratio are highly similar, especially after the formation of the septum (Fig. 3c, e).
a Schematics of foregut slice culture. The white contour depicts the perimeter of the epithelium, the black shade indicates the area enclosed by the epithelium, and the distance between the two orange dots defines the neck width. b Live imaging of GFP chick and nTnG mouse foregut slices in culture. The moment of formation of the epithelial septum is defined as time 0. The red ellipse highlights a small population of elongated mesenchymal cells. c–f Quantification of the neck width (c), the area enclosed by the epithelium (d), the ratio between the epithelial perimeter and the square root of its area (e, referred to as the perimeter-area ratio below), and the normalized neck curvature of the epithelium (f) from chick slice culture videos. Data are shown as median ± median absolute deviation (MAD). N = 10 biological replicates for E3.5, N = 14 for E3.75, N = 9 for E4.0. The dotted lines indicate time 0 when the epithelial septum forms, which is used to align different samples temporally. g Live imaging of E3.75 GFP chick slices with surgical removal of the mesenchyme, in the absence or the presence of bilateral pressure by a collagen gel trough. h, i Quantification of the neck width (h) and the perimeter-area ratio (i) of mesenchyme-removed slices without (magenta) or with external bilateral force (blue). Each curve shows an individual sample (N = 5 for slices with only mesenchyme removal, N = 4 for slices with mesenchyme removal and external pressure). Scale bars: 50 µm.
Leveraging the slice culture system, we then asked how the absence of the mesenchyme would affect TES. Strikingly, mesenchyme-reduced foregut slices failed to complete TES (Fig. 3g). Instead, their initial constriction in the epithelium often reversed over time, even for those with an existing septum (Fig. 3g, Supplementary Movies 9, 10). This suggests the constrictive mesenchymal force may be essential for TES ex vivo. However, an alternative explanation could be that the reduction of the mesenchyme also diminished the mesenchymal signaling molecules, which are potentially important to TES. To distinguish between these possibilities, we squeezed mesenchyme-removed slices into a narrow trough made of a stiff collagen gel, which provides external forces in the mediolateral direction as the remaining mesenchyme proliferates in such a bilaterally confined space. Such external compression rescued the morphological progression of TES (Fig. 3g–i, Supplementary Fig. 5 and Supplementary Movie 11), indicating an essential biomechanical role of the foregut mesenchyme in epithelial splitting during TES.
Directed cell migration underlies the essential compressive mesenchymal force
We next investigated the cellular basis of the mesenchymal force. It is well understood that local compressive force can be generated by differential proliferation within the tissue5,26,27. We thus performed pulsed 5-ethynyl-2'-deoxyuridine (EdU) labeling to test this possibility. We found that the medial mesenchymal cells did not proliferate at a faster rate than other parts of the mesenchyme (Fig. 4a, b), in line with previous findings in the mouse using proliferation markers11,24, and thus cannot explain the mediolateral enrichment and the anisotropy of mesenchymal force that specifically narrows the prospective septum. Moreover, when we inhibited cell proliferation with aphidicolin in the slice culture, the number of mitotic cells decreased (Fig. 4c), but the morphological features of TES were not affected (Fig. 4d–f, Supplementary Movie 12). Another possibility that mesenchymal cells undergo active elongation to generate pressure was not supported at the population level, even though some cells appeared elongated as they invaded the basement membrane (circle in Fig. 3b, Supplementary Fig. 6).
a Fluorescence imaging of transverse sections of chick and mouse slices ex vivo fixed after 10 µM EdU labeling for 2 h. The dashed lines mark manually defined dorsal (D), medial (M), and ventral (V) regions for quantification. b Quantification of EdU-positive cells in the mesenchyme along the dorsoventral axis. Data are presented as mean ± SD. N = 4 biological replicates (2 measurements per sample). c Immunostaining of phospho-Histone H3 in E3.75 chick slices cultured with or without 3 µM aphidicolin for 24 h. d Live imaging of an E3.5 GFP chick slice treated with 3 µM aphidicolin. e, f Quantification of the neck width (e) and the perimeter-area ratio (f) of aphidicolin-treated E3.75 chick slices overlaid with E3.75 controls (as in Fig. 3c, e). Control data are shown as median ± MAD. Individual samples (N = 5) are shown for aphidicolin. Scale bars: 50 µm.
An alternative source of localized force could be directional cell flow28. We hypothesized that if the mesenchymal cells converged towards the epithelium, they could generate a mediolateral pressure potentially driving TES. Harnessing the single-cell resolution in our live imaging of the slice culture, we performed particle image velocimetry (PIV) analysis to infer the temporally resolved velocity field of mesenchymal cells (Fig. 5a, Supplementary Movie 13). This analysis revealed that the mesenchymal cells indeed converged towards the epithelium, and the tissue-level strain inferred from the velocity field was initiated in the mesenchyme, which then propagates to the epithelium (Fig. 5a, Supplementary Fig. 7a, b). As the chick mesenchymal cells are more densely packed than the mouse, the source of tissue strain in the chick appeared less pronounced, but the velocity field showed the same trend (Supplementary Fig. 7c). Focusing on the mouse, we independently validated the convergent mesenchymal motion by single-particle tracking of nuclear-labeled cells (Fig. 5b, Supplementary Movie 14). Analysis of the tracked cells’ trajectories showed that the cells converged towards the epithelium with varied average velocities yet strongly biased angle distribution (Fig. 5c–e). Mean squared displacement (MSD) analysis further revealed an apparent super-diffusion of cells with a scaling exponent of ~1.2, suggesting active cell migration (Fig. 5f). To rule out the potential artifacts by analyzing the three-dimensional (3D) tissue in 2D, we also performed cell segmentation and tracking in 3D, which yielded migration patterns consistent with 2D analysis (Supplementary Fig. 8, Supplementary Movie 15). However, as many datasets were not suitable for 3D analysis due to the deterioration of image contrast in deeper tissue, we performed most analyses in 2D.
a Particle image velocimetry (PIV) analysis of an E10.5 nTnG mouse foregut slice culture and mapping of the strain rate along the mediolateral axis (εxx). Cyan indicates compressive strain and red indicates expansive strain. Arrows indicate the shift of the compressive strain from the mesenchyme to the epithelium. The color wheel indicates the directions of PIV vectors. b Live imaging of an E10.5 nTnG mouse slice overlaid with cellular trajectories colored by their average speeds. c Individual cell trajectories in the left and right boxes in (b). Trajectories are color-coded by relative time, with each trajectory’s origin designated as 0. d Average velocity vectors of individual cell trajectories in (c). Vectors are scaled with the speed and color-coded by their directions according to the color wheel. e Angular histograms the velocity vectors in (d). f Mean squared displacement (MSD) plots of cell trajectories in the left and right boxes in (b). The black lines and shades show the population averages and standard deviations. The red dashed lines indicate the power-law fitting results. Frame time: 7.5 min. g Live imaging of E3.75 GFP chick slices with 2.5 µM FAK inhibitor (PF-573228), in the absence or the presence of bilateral pressure by a collagen gel trough. h, i Quantification of the neck width (h) and the perimeter-area ratio (i) of FAK inhibitor-treated slices without or with external bilateral force compared with E3.75 controls (as in Fig. 3c, e). Averaged curves are shown for FAK inhibitor-only samples as median ± MAD (N = 7), and individual curves are shown for slices with FAK inhibitor and external pressure (N = 4). j Immunofluorescence of GM130 (GOLGA2) in a chick foregut section. k Zoomed-in view of the red box in (j) overlaid with mesenchymal cell orientation vectors indicated by arrows colored in grayscale. 0° marks direction towards the epithelium. l, m Distributions of cell orientation in control (l, N = 11) and FAK inhibitor-treated (m, N = 8) slice culture. Each colored line represents one biological replicate, and the black line shows the mean ± SD. The dashed purple line marks the expected uniform distribution of angles for random cell orientation (null hypothesis). P-values are calculated by the Chi-square test against a uniform distribution (df = 10 for control, df = 7 for FAK inhibitor). Scale bars: 50 µm.
The above results indicate that the directional flow of mesenchymal cells towards the epithelium may underlie TES. To test the functional relevance of the mesenchymal flow, we inhibited cell migration with the focal adhesion kinase (FAK) inhibitor PF-573228 in the slice culture. FAK is an enzyme essential for cell migration whose activity is particularly high in the mesenchyme (Supplementary Fig. 7d). In both chick and mouse, inhibition of cell migration significantly reduced the mesenchymal flow, in terms of both speed and directionality (Supplementary Fig. 7e, f). In all treated samples, TES was stalled at the epithelial narrowing stage (Fig. 5g–i, Supplementary Fig. 7e, f and Supplementary Movie 16). As with the surgical removal of the mesenchyme, the effect of FAK inhibition can be rescued by external pressure (Fig. 5g–i, Supplementary Fig. 5e and Supplementary Movie 17).
We complemented the live-imaging results with fixed-tissue analysis, using the vector formed by the Golgi apparatus and the nucleus as an indicator of mesenchymal cell polarity29 (Fig. 5j, k). We found that the mesenchymal cells collectively oriented towards the epithelium, producing an angular bias in their polarity vectors (Fig. 5l), which was absent in FAK inhibitor-treated samples (Fig. 5m).
In conclusion, through a comprehensive series of biophysical and pharmacological perturbations in the ex vivo slice culture system, we have demonstrated that epithelial splitting in TES is initially driven by the convergent mesenchymal pressure due to directional cell flow, which is critical to narrow the epithelium mediolaterally to form the septum and enable tube splitting.
SHH signaling is essential for generating the convergent mesenchymal force
A clue to the molecular regulation of the directional cell movements underlying TES came from consideration of congenital birth defects in which the foregut fails to separate and develops as a fused tube. Laryngo-tracheo-esophageal cleft (LTEC) is a relatively rare human anomaly where an abnormal connection between the airway and esophagus forms due to a failure of TES14,15. Notably, the LTEC phenotype is closely modeled in mice deficient in SHH signaling, implicating this pathway in achieving normal TES11,23,30,31,32. Although esophageal atresia and tracheal-esophageal fistula (EA/TEF), involving an incomplete TES, is much more commonly observed in human birth defects14,15, the complete loss of TES in LTEC offers a simpler model to identify the most critical components of TES. We first confirmed that SHH signaling, indicated by the expression of the Ptch1/PTCH1 gene, was widespread in the foregut mesenchyme in both mouse and chick; and that it was lost upon genetic knockout of Shh in the mouse or in ovo pharmacological inhibition of SHH signaling in the chick (Fig. 6a, Supplementary Fig. 9a). Notably, the loss of SHH signaling also dramatically reduced the number of mesenchymal cells, with less significant impact on foregut patterning (Fig. 6b, Supplementary Fig. 9b). The mesenchymal hypoplasia was mainly due to a lack of proliferation, rather than excessive cell death (Supplementary Fig. 9c–f). Importantly, the mesenchyme removal assay showed that the mesenchymal compressive force was lost upon SHH inhibition (Fig. 6c–e). Collective cell polarity towards the mesenchyme was also lost with SHH inhibition (Fig. 6f). Live imaging of cyclopamine-treated chick and Shh-knockout mouse foregut slices further revealed that SHH signaling is essential for the narrowing of the epithelium, as its loss led to stalling or even relaxation of the epithelial morphology (Fig. 6g–i, Supplementary Fig. 9g and Supplementary Movie 18). In these cases, both the isotropic, proliferative force and the directional, migrative force were diminished, yet the epithelium still exhibited slight medial constriction, suggesting epithelium-intrinsic mechanical heterogeneities. Application of external pressure can partly rescue the LTEC phenotype, allowing the formation of the epithelial septum between the forming esophagus and airway, although the resolution of the septum took significantly longer than in wildtype, implying a role for active flow in septum resolution that was not fully recapitulated by bilateral, proliferative pressure (Fig. 6g, Supplementary Movie 19). PIV analysis further showed that SHH signaling is necessary for the convergent flow pattern of the mesenchyme (Supplementary Fig. 9h).
a HCR-FISH of PTCH1 in transverse sections of E4.0 chick foreguts with or without in ovo SHH inhibitor (cyclopamine) treatment. b Immunofluorescence of SOX2 and NKX2-1 in transverse sections of E3.75 chick slices with or without in ovo SHH inhibitor treatment. c Stereoscope imaging of a control and a SHH inhibitor-treated GFP chick foregut slice before (magenta) and after (green) surgical removal of the mesenchyme. d Quantification of epithelial neck widths (defined in Fig. 3a) before and after mesenchyme removal (N = 5 biological replicates for control, N = 4 for SHH inhibition). P-values are calculated by a multiple paired t-test. e Comparison of epithelial neck width changes calculated from (d). Data are presented as mean ± SD. P-value is calculated by the Mann-Whitney test. f Distributions of mesenchymal cell orientation in SHH inhibitor-treated slice culture as in Fig. 5l (N = 7 biological replicates). Data are presented as mean ± SD. P-value is calculated by the Chi-square test against uniform distribution (df = 6). g Live imaging of E3.75 GFP chick slices treated with SHH inhibitor in ovo, in the absence (top) or the presence (bottom) of bilateral pressure by a collagen gel trough. h, i Quantification of the neck width (h) and the perimeter-area ratio (i) of SHH inhibitor-treated slices without or with external bilateral force compared with E3.75 controls (as in Fig. 3c, e). Each curve shows an individual sample (N = 5 for SHH inhibitor-only samples, and N = 3 for SHH inhibitor with bilateral pressure). Control data are presented as median ± MAD. Scale bars: 50 µm.
The dorsal sub-epithelial mesenchyme is sensitive to SHH signaling and contributes to the mesenchymal force
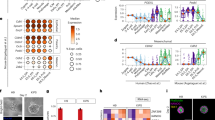
Recent studies uncovered multiple cell types in the developing foregut mesenchyme, which could be potentially responsible for driving the SHH-dependent convergent flow23,33. To understand which cell type responds to SHH signaling to enact the directed movement, we performed single-cell RNA sequencing (scRNA-seq) of foregut slices from normal and cyclopamine-treated chick embryos and used graph-based clustering to establish cell groupings putatively representing distinct cell types (Fig. 7a). We then spatially mapped the identified cell populations in the foregut by staining for marker genes representative of each cluster (Fig. 7b, Supplementary Figs. 10a, b, 11a). We found that the foregut mesenchyme is composed of sub-epithelial and peripheral populations along the radial axis from the epithelium, in addition to the dorsoventral axis corresponding to the esophageal and tracheal cell types. Upon SHH inhibition, the cell population whose size is most affected was the dorsal sub-epithelial mesenchyme, whereas the fractions of other major cell populations remained balanced (Fig. 7c, d, Supplementary Fig. 10c). The dorsal sub-epithelial mesenchyme, marked by NKX6-1, also express SHH response genes including PTCH1/2 and FOXF1/2, which were dramatically downregulated with cyclopamine treatment (Fig. 7e). Although smooth muscle differentiation has not yet begun during TES, we found early expression of smooth muscle actin (ACTA2) in a subset of dorsal sub-epithelial mesenchyme, which plummeted upon SHH inhibition (Supplementary Fig. 10d). This may underlie the smooth muscle defects observed in SHH mutants34,35. Gene ontology analysis of differentially expressed genes between control and cyclopamine-treated dorsal sub-epithelial mesenchyme revealed that pathways related to cell migration, including extracellular matrix (ECM)-receptor interaction, axon guidance, and focal adhesion, were significantly downregulated with SHH inhibition, suggesting a role of SHH in directing cell migration (Fig. 7f). As SHH is secreted by the epithelium and sensed by the mesenchyme, we further analyzed intercellular communication pathways between dorsal epithelium and dorsal sub-epithelial mesenchyme/peripheral mesenchyme (Supplementary Fig. 11b). Among the predominant signaling pathways between these cell types, including SHH, semaphorin, and specific ECM-receptor interactions, only SHH signaling was dramatically weakened by cyclopamine treatment (Supplementary Fig. 11b). Immunostaining confirmed that the dorsal sub-epithelial mesenchyme was almost absent with in ovo cyclopamine treatment, with the remained mesenchyme being the peripheral mesenchyme marked by TBX1 (Fig. 7g). In such cases, the remaining mesenchyme is insufficient to generate the compressive force critical to TES.
a UMAP of single-cell transcriptomes from E3.5 and E4.0 chick foregut slices. b Schematics of the spatial distribution of major epithelial and mesenchymal cell types in the foregut based on HCR-FISH and immunofluorescence of select marker genes. c Fractions of the dorsal sub-epithelial mesenchyme in individual embryos recovered from scRNA-seq data. N = 6 embryos for Control-E3.5, N = 4 for Control-E4.0, N = 9 for SHH inhibition-E4.0. Data are presented as mean ± SD. P values are calculated by one-way ANOVA with Dunnett’s test. d Cell composition analysis of E3.5 control, E4.0 control, and E4.0 cyclopamine-treated samples. e Volcano plot highlighting differentially expressed genes (DEG) between control and cyclopamine-treated samples. Genes with log2-fold change > 1 and Padj < 10-4 are marked red. f Gene ontology analysis of downregulated DEGs by SHH inhibition. g Immunofluorescence of NKX6-1 and TBX1 in transverse sections of E4.0 chick foreguts with or without in ovo cyclopamine treatment. h Live imaging of an E3.75 GFP chick slice with surgical removal of the dorsal mesenchyme in the absence or presence of bilateral pressure. i, j Quantification of the neck width (i) and the perimeter-area ratio (j) of dorsal mesenchyme-removed slices without or with external bilateral force. N = 8 samples for dorsal mesenchyme removal and N = 6 for dorsal mesenchyme removal with compression. Data are presented as median ± MAD. Scale bars: 50 µm.
These results suggest that SHH is essential for the specification and proliferation of the dorsal sub-epithelial mesenchyme, the loss of which leads to defective mesenchymal force, and hence a failure to enable TES. To directly test the functional importance of this population, which spans the dorsal and medial domains of the foregut (Fig. 7g), we surgically removed the dorsal and medial mesenchyme, while keeping the ventral part intact (Fig. 7h, Supplementary Movie 20). Strikingly, this treatment is sufficient to halt TES, verifying that the major source of the compressive mesenchymal force comes from the dorsal/medial mesenchyme (Fig. 7h). Quantitative analysis shows that although the ventral mesenchyme is still present, the initial epithelial neck width increases and its reduction slows (Fig. 7i, Fig. 3c), both of which are rescued by applying additional bilateral pressure (Fig. 7i, j, Supplementary Movie 21). These results suggest a significant and essential contribution of mesenchymal force from the dorsal sub-epithelial mesenchyme. The presence of medial constriction rather than dorsal compression can be partly explained by the substantially greater number of cells surrounding the epithelium bilaterally than dorsally (Fig. 7g, Supplementary Fig. 10b).
SHH induces directional migration of the foregut mesenchymal cells to deform the epithelium
Our results showed that SHH is essential for the specification and proliferation of the dorsal sub-epithelial mesenchyme, which in itself can potentially generate static crowding pressure on the epithelium. To test whether, in addition, SHH plays a role in inducing directional mesenchymal cell migration essential to TES, we implanted beads loaded with SHH ligand into the foregut mesenchyme (Fig. 8a). Subsequently, 24 h post-implantation, the mesenchyme near the SHH bead showed significantly higher density than the contralateral side, whereas implantation of an empty bead exhibited no such effect (Fig. 8b). The increased cell density was not due to enhanced cell proliferation by SHH because the fraction of dividing cells did not increase near the beads (Supplementary Fig. 12a, b), potentially because the endogenous SHH level was sufficient to sustain a relatively high proliferation rate. Live imaging further revealed that mesenchymal cells near the bead moved towards the SHH source, instead of moving towards the epithelium on the contralateral side (Fig. 8c–e, Supplementary Movie 22). Such effect was not reproduced with an empty bead (Supplementary Fig. 12c–f, Supplementary Movie 23). The cell polarity vectors show a significant bias towards the SHH bead at an ~60° angle (Fig. 8f–h), potentially due to cell packing density, which prevents unlimited persistent migration towards the bead. These data strongly suggest that foregut mesenchymal cells are attracted to the source of SHH activity.
a HCR-FISH of PTCH1 in an E3.5 chick foregut slice 24 h after implantation of an Affi-Gel bead loaded with recombinant mouse SHH protein. b Comparison of mesenchymal cell density near the implanted bead (with or without SHH) and the contralateral side. N = 7 biological replicates for control, N = 5 for SHH. P-values are calculated by a paired t-test. c PIV analysis of an E3.75 GFP chick slice with an implanted SHH-bead. The color wheel indicates the directions of PIV vectors. d Live imaging of an SHH-bead-implanted GFP chick slice overlaid with cellular trajectories colored by their average speeds. e Average velocity vectors of individual cell trajectories in the blue and red boxes of (d). Vectors are scaled with the speed and color-coded by their directions according to the color wheel. The dashed circle marks the bead. f Immunofluorescence of GM130 (GOLGA2) in a chick foregut section with a SHH bead. g, h Distributions of mesenchymal cell orientation in slices implanted with control beads (g, N = 4 biological replicates) and SHH beads (h, N = 6 biological replicates). Data are presented as mean ± SD. P-values are calculated by Chi-square test against uniform distribution (df = 3 for Control, df = 4 for SHH bead). i, j In ovo electroporation of plasmids encoding mCherry (i), SHH (i), or HHIP (j) into the right epithelium of the chick foregut. Frontal views of the foreguts by stereo microscopy show the effective electroporation (i). Immunofluorescence of SHH (i) and HCR-FISH of HHIP and PTCH1 (j) confirmed the efficient expression of the constructs. Arrows indicate the asymmetrical deformation of the epithelium induced by SHH overexpression. The dashed lines bisect the foregut lumen as a reference for measuring epithelial distances to the midline. k Comparisons of right (electroporated) and left (non-electroporated) epithelium-to-midline distances with control (mCherry, N = 5 biological replicates), SHH (N = 7 biological replicates), and HHIP (N = 6 biological replicates) electroporation. Data are presented as mean ± SD. P-values are calculated by One-way ANOVA with Dunnett’s test. l Working model of TES initiation. SHH secreted from the foregut epithelium induces the dorsal sub-epithelial mesenchyme’s migration towards the epithelium, generating bilateral compressive force essential to TES. Created in BioRender. Yan, R. (2025) https://BioRender.com/cdwttt9. Scale bars: 50 µm, except those marked on the images.
Next, we attempted to perturb the symmetry of SHH expression to test whether the strength of SHH signaling correlates with epithelial deformation. When we overexpressed SHH, but not mCherry as negative control, in the right foregut epithelium by electroporation, the right foregut epithelium was more deformed than the left side, implying the induction of a stronger mesenchymal force with the increase of SHH signaling (Fig. 8i). This result shows that the active flow is sufficient to enhance local deformation of the epithelium independent of the proliferative force. Conversely, when the SHH antagonist Hedgehog Interacting Protein (HHIP) was overexpressed, SHH signaling near the electroporated site was attenuated, resulting in a slightly less deformed epithelium compared to the unaffected side, potentially due to the inability of HHIP to fully inhibit SHH secretion (Fig. 8j, k). Therefore, induction of directional cell migration by physiological SHH signaling is sufficient to initiate epithelial deformation for tube splitting. It is plausible that the mesenchymal cells are chemotactic to SHH secreted by the epithelium, and as the epithelium constricts, a negative curvature then concentrates SHH to generate a positive mechanochemical feedback, which facilitates epithelial splitting. To test this hypothesis, we developed a theoretical model to quantify how tissue curvature affects the concentration of secreted SHH (Supplementary Note 1). Analytical predictions and numerical calculations both indicate that within physiological curvature ranges, negative curvatures from −0.01 µm−1 to −0.04 µm−1 can effectively increase local SHH concentration by 10% to 40% compared with a flat interface (Supplementary Fig. 13). This positive feedback between mesenchymal force and epithelial constriction may ensure the robustness of TES, which is essential for the individual’s viability after birth.
Discussion
In search of the biomechanical basis of epithelial tube splitting, we focused on TES, a critical epithelial splitting event in foregut development. We established an ex vivo slice culture system to observe and perturb TES with single-cell resolution. This enabled us to uncover an unappreciated role of mesenchymal force due to convergent cell migration. Gaining insights from mouse models with defective TES, we identified SHH signaling as both the driver of the convergent cell flow and the maintainer of mesenchymal proliferation. Our results provide a biomechanical mechanism bridging the existing genetic understanding of TES and its morphogenetic outcome, demonstrating how SHH signaling from the epithelium to the mesenchyme transforms into convergent mesenchymal force driving TES (Fig. 8l).
Epithelial tube splitting morphogenesis requires mesenchymal force
The mechanism of epithelial folding has attracted the attention of biophysicists and developmental biologists for decades. Early work highlighted that the asymmetric distribution of actomyosin within the epithelium can drive folding, such as in neural tube closure and intestinal crypt morphogenesis36,37. Anisotropic proliferation of the epithelial cells is also known to fold the epithelium through buckling27,38. Besides the epithelium-intrinsic mechanisms, we have just begun to appreciate the biomechanical role of the adjacent mesenchyme, whose signaling role in epithelial-mesenchymal interaction is much better understood5,6,39,40,41. For example, spontaneous clustering of mesenchymal cells underlies the initial intestinal villus formation in the mouse7,42,43. Although either epithelium-intrinsic or -extrinsic forces can fold the epithelium in various systems, it remains unknown whether either, or both forces, are sufficient to drive epithelial tube splitting, which starts with high-curvature folding and ends up with a topological transition in the epithelium. The poor understanding of epithelial splitting is at least in part due to the lack of a tractable experimental system. TES, where the dorsoventrally patterned foregut epithelium splits into the dorsal esophagus and the ventral trachea, is among the few biological examples of epithelial splitting in normal development8,10. The esophageal and tracheal progenitors sort into distinct domains via actomyosin contractility downstream of heterotypic ephrin signaling, yet in vitro reconstituted cell domains remain connected, suggesting additional force is required for initiating epithelial splitting13,44.
Our ex vivo slice culture system recapitulates the tissue dynamics of TES characterized in vivo, and importantly, allows cross-sectional imaging with single-cell resolution. By surgical and pharmacological manipulations of the foregut mesenchyme in slice culture, we demonstrate that the cell flow-dependent mesenchymal force is crucial to epithelial narrowing in TES to form the epithelial septum. The septal epithelial cells then undergo apical-basal remodeling and apoptosis to resolve into two tubes11,24. It is noteworthy that the critical role of mesenchymal force does not imply that epithelium-intrinsic processes are unimportant. Indeed, previous genetic studies have identified essential epithelial mechanisms in TES, especially the proper patterning of the SOX2 and NKX2.1 domains13,20,45. Therefore, TES requires both mesenchymal and epithelial forces, which may also hold true in other epithelial splitting systems, such as cloacal septation of the embryonic hindgut10.
SHH signaling integrates molecular patterning of the foregut and TES morphogenesis
Beyond elucidating the biomechanical basis of TES, we also sought to connect the biophysical mechanism to the existing knowledge about foregut patterning through genetic studies. It was demonstrated almost two decades ago that the gradients of BMP and Wnt signaling in the mesenchyme instruct the dorsoventral patterning of the foregut epithelium into SOX2+ esophageal and NKX2-1+ ventral domains, which secretes SHH to sustain the mesenchyme19,22,23,46,47. Deficient SHH signaling has been observed in human patients with TES defects as well as in genetic mouse and frog models, where the foregut mesenchyme exhibits various degrees of hypoplasia depending on the severity of SHH reduction11,48. Although these studies have implicated a role for the mesenchyme in TES morphogenesis, the mechanisms have not been fully investigated. In our mouse and chick models of SHH deficiency, the foregut epithelium fails to narrow bilaterally to initiate TES, in a similar fashion to the mesenchyme-reduced wild-type sample in slice culture. Combining scRNA-seq, bead implantation, and targeted electroporation, we show that the dorsal sub-epithelial mesenchyme is the main effector of SHH in TES morphogenesis. SHH signaling can directly induce mesenchymal cell migration and is crucial for dorsal mesenchymal cell proliferation49,50,51. Both roles are essential to TES. Inhibition of the migratory force by the FAK inhibitor fully blocks TES (Fig. 5g–i), whereas a reduction in mesenchymal cell number by surgical dissection can also delay TES (Fig. 7h–j). By contrast, blocking cell proliferation at later stages does not prevent TES (Fig. 4), suggesting a critical threshold of mesenchymal cell number in TES. We propose that the SHH-dependent mesenchymal proliferation generates an isotropic pressure that constrains the epithelium, whereas the SHH-dependent convergent flow constricts the epithelium more mediolaterally in TES. The exact positioning of the septum at the dorsoventral boundary of the epithelium is likely initiated by ephrin-mediated epithelial surface tension differences13,44, which can be further reinforced by the curvature-sensitive, mechanochemical feedback (Supplementary Fig. 13). Our results thus bridge the gap between molecular patterning and tissue morphogenesis in TES, demonstrating how epithelial-mesenchymal interactions function synergistically to enable robust epithelial splitting.
Mechanobiology of morphogenesis sheds light on the etiology of structural birth defects
In the study of structural birth defects, much of the focus has been on evaluating individual disease-causing genes and their roles in normal morphogenesis, taking advantage of the rapidly expanding base of clinical genetics data as a starting point52,53. While this paradigm has provided valuable insights into the molecular underpinnings of many conditions, critical aspects of the underlying mechanisms might be missed when pertinent genes have pleiotropic effects essential for embryonic development or early postnatal life. For example, although genetic disruptions of SHH signaling (Shh, Gli2), WNT signaling (Wnt2/2b), and ephrin signaling (Efnb2) all cause a fused foregut tube in mouse embryos with nearly 100% penetrance, these pathways are underrepresented or absent in the clinical data of TES-related birth defects, likely because these pathways are critical for other aspects of embryonic development14,15. Consequently, studying normal tissue morphogenesis is crucial for understanding the full spectrum of factors contributing to proper organ formation. Mechanobiological processes, in particular, play an essential role in shaping tissues and organs during development, but these mechanisms remain underappreciated in the development of many organ systems. By focusing on the mechanobiology of physiological and pathological TES morphogenesis, our work unveils the vital role of mesenchymal force in shaping the foregut epithelium, deepening our understanding of the etiology of foregut birth defects. The ex vivo system that enables biomechanical and biochemical perturbations of TES lays the foundation for future work to elucidate the biomechanical and cellular mechanisms underlying common foregut malformations, such as EA/TEF.
Methods
All animal studies were performed in compliance with the protocols approved by the Institutional Animal Care and Use Committee at Harvard Medical School (IS00000122-9).
Chick embryos
Fertilized Specific Pathogen Free (SPF) white Leghorn chicken eggs (Charles River/AVS Bio) and transgenic Roslin Green (expressing cytoplasmic GFP) chicken eggs (Susan Chapman, Clemson University) were used. Eggs were incubated in a 38 °C humidified chamber before collection.
Mouse embryos
All mouse lines used were obtained from the Jackson Laboratory: C57BL/6 J (#000664), ROSAnT-nG (nTnG, #023537), Shh-GFP-Cre (#005622), and Shh-CreERT2 (#005623). We crossed Shh-CreERT2 with nTnG to generate the strain ShhCreER/+;nTnG/nTnG, which was then crossed with Shh-GFP-Cre to generate the Shh-knockout embryos and littermate controls. All mice were housed in a specific pathogen-free (SPF) facility at Harvard Medical School under a 12-h light/12-h dark cycle, at an ambient temperature of 20–24 °C with 40–60% relative humidity. Animals had ad libitum access to food and water. Timed pregnancy was set up for all experiments using females of 3–6 months old. The morning when a vaginal plug was observed was designated as E0.5. The gender of the experimented embryos was not determined, as it is not a known factor relevant to TES in mouse development.
Plasmids
pCAGGS-mCherry was a gift from Phil Sharp (Addgene plasmid #41583). pCAGGS-SHH and pCAGGS-HHIP was made by replacing mCherry with chick SHH (cloned from Addgene plasmid #13991, Primer 1: TTCTGCAGTCGACGGTACCGCGGGCCCGGGATCCAATGGACGAAATGCTGCTGTTGAC; Primer 2: CACCTTCTGATAGGCAGCCTGCACCTGAGGAGTGCTCAGCTGGCCGGTGC) or mouse HHIP (Origene, MC203592, Primer 1: GCAGCTAGCCGCCACCATGCTGAAGATGCTCTCGTTTAAG; Primer 2: GACGGATCCCTATACAATGTAACTTGTGAGATCCAGC) using Gibson assembly or restriction/ligation cloning. pCAGGS-MLC2-tdTomato was made by cloning mouse MLC2-tdTomato (from Addgene plasmid #58108, Primer 1: GCAGAATTCGCCACCATGTCGAGCAAGAGAGCC; Primer 2: CGCTTACTTGTACAGCTCGTC) to pCAGGS. The plasmids will be deposited to Addgene. QIAGEN Plasmid Maxi Kit was used to purify plasmids from in-house prepared DH5-alpha cells.
Foregut slice culture
Fresh chick or mouse embryos were collected and promptly dissected in chilled Dissection Medium (4% fetal bovine serum [Peak Serum, PS-FB2] and 100X-diluted penicillin-streptomycin [Gibco, 15140-122] in DMEM with HEPES [Gibco, 21063029]) to isolate the foregut, from the pharyngeal arches to the upper stomach. The foregut was then manually sliced with a surgical blade (Aspen Surgical, 371111) transversally to obtain slices of 100–150 µm thickness. The slice containing the septum and the anterior undivided foregut was embedded in 50 µL 50% Matrigel (Corning, 356231) diluted with the Dissection Medium on a 35-mm glass bottom dish (MatTek, P35G-1.5-14-C), with the undivided foregut side facing up. Once the slice was settled in Matrigel at room temperature, the dish was incubated at 37 °C in a humidified chamber for 30 min to solidify the Matrigel. 2-mL of culture medium for chick (5% chick embryo extract [US Biological, C3999], 3% fetal bovine serum, 0.25 µM LDN-193189 [Cayman Chemical, 11802], and 100X-diluted penicillin-streptomycin in FluoroBrite DMEM [Gibco, A1896701]) or mouse (100 ng/mL EGF [R&D Systems, 236-EG], 5% fetal bovine serum, 0.25 µM LDN-193189, and 100X-diluted penicillin-streptomycin in FluoroBrite DMEM) was carefully added to the dish, and the sample was incubated at 37 °C with 5% CO2 prior to imaging.
Surgical removal of foregut mesenchyme
Foregut slices were trimmed with fine forceps (Fine Science Tools, 11252-00) to reduce the mesenchyme without impacting the epithelium. Stereo images before or immediately after mesenchyme removal were taken on a Leica stereoscope (M165 FC) with an LED light source (Lumencor, SOLA), and the images were aligned using a custom MATLAB code. The removal of chick foregut mesenchyme can be nearly complete, as the tissue is relatively large and the epithelium is robust, whereas for the mouse foregut, we usually left some mesenchyme to avoid tearing and breaking the fragile epithelium.
Laser ablation
Laser ablation was performed with the Leica Stellaris 8 multiphoton microscope with the Insight X3 dual beam laser and the 25X/1.0NA water immersion objective. GFP chick slices were excited at 960-nm with 7% power. A video before ablation was first recorded as a control. Laser ablation was then performed with 800-nm excitation at 70%–80% power depending on the depth of the imaging plane, in a defined ROI marking the sub-epithelial mesenchyme. The ROI was ablated for ~0.1 s per 4-s frame for 3–10 frames, with the transmission light image being recorded during ablation to monitor the damage to cells. Once the cells were ablated, the system was switched to the imaging mode as before ablation, with a recording rate of 4–12 s per frame.
The post-ablation video was drift-corrected with the Fiji StackReg plugin (https://bigwww.epfl.ch/thevenaz/stackreg)54. Kymographs were generated with the Multi Kymograph function in Fiji (https://biii.eu/multi-kymograph) along lines normal to the contour of the epithelium crossing the ablated region.
We quantified three characteristics of recoil after ablation. First, the initial displacement rate was determined by the slope of the tangent line at the kymograph’s base, which reflects the force imbalance at the tissue interface and the epithelial viscosity. Second, we measured the total epithelial displacement by tracking the movement of the basal side of the epithelium, which is a consequence of force imbalance and epithelial elasticity55. However, quantitative fitting of the tissue displacement by a simple Kevin-Voigt model is complicated by the simultaneous expansion of the epithelial width in both directions due to the lack of resisting force on the apical side of the epithelium.
Time-lapse imaging of the foregut slice culture
GFP chick slices were excited at 960-nm with 7% power, and nTnG mouse slices were excited at 1040-nm with 7% power on the above-mentioned two-photon microscope. A Z stack of 30-60 µm with 10–15 µm step size was recorded for each slice, starting from ~30 µm below the tissue surface to avoid potential artifacts from cutting. Samples were imaged every 10–30 min for 12-16 h. The XY resolution ranged from 250 nm to 500 nm per pixel. The time-lapse videos were used for quantitative analysis of the morphological evolution of the foregut (see “Morphometric analysis of the foregut epithelium”).
Bilateral compression of foregut slices
We made a narrow trough in a collagen gel pad to compress the foregut slice. The collagen gel was made with a solution containing 2 mg/mL rat tail collagen (Gibco, A1048301), 1X PBS (Invitrogen, AM9625), and 17 µM sodium hydroxide (Supelco, SX0607N) in water. 1 mL collagen solution was spread on a 35-mm glass-bottom dish and incubated at 37 °C for 30 min. The solidified gel pad was buffered at room temperature with 2 mL Dissection Medium, then 1 mL culture medium. A 1-mm long, 200–300-µm wide trough was made in the gel pad with a fine tungsten needle (Fine Science Tools, 10130-05), and the foregut slice was gently squeezed bilaterally with forceps to fit into the trough with the dorsoventral axis parallel to the long axis of the trough. The sample was incubated at 37 °C for several hours before live imaging as described above.
We also used the collagen gel to calibrate the stability of the imaging setup. We embedded 200-nm red fluorescent beads (Invitrogen, F8810) at 1:10,000 dilution in collagen solution prior to gelation. During live imaging, the beads were imaged with the 1040-nm laser and tracked with TrackMate to generate the trajectories.
Pharmacological treatment of foregut slices
We used drugs at concentrations that effectively inhibited the biological target with minimal adverse effects on the tissue integrity and viability. 3 µM aphidicolin (Tocris, 5736) or 2.5 µM PF-573228 (MedChemExpress, HY-10461) was added to the slice culture medium 2–3 h prior to live imaging for 12–24 h.
In ovo cyclopamine treatment
To inhibit SHH signaling in vivo, we developed an in ovo cyclopamine treatment protocol for chick embryos. 3 mL of albumen was drawn from the bottom of the egg at E2 to prevent the embryo from sticking to the eggshell. The egg was sealed with tape and incubated to E3 when it was windowed. 50 µL 10 mM cyclopamine (ApexBio, A8340) dissolved in ethanol was mixed with 250 µL PBS by gentle pipetting, and the mixture, usually in a form of cloudy suspension, was carefully added on top of the chick embryo. The treated egg was immediately sealed and returned to the incubator. We found that this protocol effectively inhibited SHH signaling, yielding phenotypes highly similar to the Shh-knockout mouse embryos. The timing and dose of cyclopamine addition could be tuned to inhibit SHH signaling in a controlled manner.
Bead implantation of foregut slices
A 20-µL aliquot of 100–200 mesh Affi-Gel blue beads (Bio-Rad, 1537302) was rinsed with 1.5 mL PBS overnight at 4 °C. 10 µL of 2 mg/mL recombinant mouse SHH N-terminus (R&D Systems, 464-SH) was dropped on the lid of a plastic dish placed on ice. Due to the small size of the foregut tissue, we picked the smallest beads to transfer into the SHH droplet, incubating on ice for 1–2 h. The loaded beads were implanted to the dorsal mesenchyme of foregut slices using fine forceps, making sure the implanted bead was in the middle of the slice to prevent dislodging of the bead. Implanted slices were then embedded and cultured in Matrigel as describe above. As control, the PBS-rinsed beads were transferred to a droplet of PBS on ice prior to implantation.
Slices were fixed 24 h post-implantation for HCR of chick PTCH1. To quantify mesenchymal cell density, we counted the number of DAPI-positive nuclei within a ring 10 µm to 50 µm away from the bead surface, and then divided the total cell number by the area of the ring. On the contralateral side without the bead, a square of equivalent area was drawn to count the cells.
In ovo electroporation of the foregut epithelium
The plasmid mixture for electroporation was made with 4 µg/µL plasmid, 0.5% Fast Green FCF (Sigma-Aldrich, F7252), and 3% sucrose (Sigma-Aldrich, S8501) in TE buffer. E2 chick embryos (HH10-12) were lowered and windowed. The embryo was counterstained with an injection of ~30 µL 20X diluted ink (Pelikan, 211862) in PBS into the yolk below the embryo. A mouth pipette with a thin tip, made with a glass capillary (FHC Inc, 30-30-0) pulled by a micropipette puller (Sutter Instrument, P-97), was loaded with the plasmid mixture, which was injected into the foregut lumen dorsally until the dye flowed out from the anterior intestinal portal. A parallel needle electrode (Bulldog Bio, CUY560-5-0.5) was inserted into the yolk parallel to the anterior-posterior axis of the embryo, with the positive electrode on the right side of the embryo. Electroporation was performed with three 50-V, 8-ms, 100-ms apart poring pulses, followed by five 20-V, 8-ms, 100-ms apart transfer pulses, all towards the positive electrode (Nepa Gene, NEPA21). A thin layer of fresh albumen was added to the top of the embryo to prevent drying, and the egg was taped and incubated to the desired stage.
EdU labeling and detection
10 µM EdU (5-ethynyl-2'-deoxyuridine, Invitrogen, A10044) was added to the culture medium for foregut slices. After two hours of incubation at 37 °C, the sample was fixed, embedded, and sectioned as described in “Immunofluorescence of tissue sections”. EdU labeling was performed with the Click-iT EdU Cell Proliferation Kit for Imaging with Alexa Fluor 488 (Invitrogen, C10337) according to the manufacturer’s protocol.
To quantify the fraction of EdU-positive cells, we manually counted the total number of cells labeled by DAPI and the EdU-positive cells in 30 × 30 µm2 boxes within the dorsal, medial, or ventral mesenchyme. At least three boxes were counted for each region in each slice, and the results were averaged as one biological replicate.
Whole-mount immunofluorescence
Whole embryos (chick embryos before E3.5 and mouse embryos before E10.0) or dissected foreguts (chick embryos after E4.0 and mouse embryos after E10.5) were fixed in 4% paraformaldehyde (PFA, Electron Microscopy Sciences, 15710) diluted in PBS (Gibco, 10010023) overnight at 4 °C. The samples were then washed three times with PBS (10 min each at room temperature), and blocked with Blocking Buffer (10% bovine calf serum [Gibco, 16010159] and 1% Triton X-100 [Sigma-Aldrich, T8787] in PBS) for one hour at room temperature. Mouse anti-E-Cadherin (BD Biosciences, 610182) was diluted 1:100 in the Blocking Buffer and labeled the samples for two days at 4 °C. The samples were then washed three times with the Blocking Buffer (1 h each at room temperature) and stained with donkey anti-mouse-Cy3 (Jackson ImmunoResearch, 715-165-150) diluted 1:400 in the Blocking Buffer for one day at 4 °C. After six times of washing with 5X diluted Blocking Buffer, the samples were serially dehydrated with 50% methanol (Fisher Scientific, A433P-4)/50% PBS, 80% methanol/20% water (Invitrogen, 10977015), and 100% methanol (1 h each step at 4 °C). Optical clearing was performed with the CytoVista Tissue Clearing kit (Invitrogen, V11322) per the manufacturer’s instruction.
Cleared samples were transferred to a 50-mm glass-bottom dish (MatTek, P50G-1.5-30-F) filled with the CytoVista Tissue Clearing Enhancer solution (part of the Tissue Clearing kit) and imaged on a Nikon Ti inverted microscope with a W1 spinning disk scanner (Yokogawa, CSU-W1) using a 20x objective (Nikon, MRD70270) and 561-nm excitation. A Z-stack of ~0.6–1.0 µm step size was acquired for each sample. Three-dimensional rendering of the image was performed in the Arivis Vision4D software (Zeiss), with the Z step size multiplied by 1.5 to correct for the refractive index mismatch. Markers were manually set at the pharyngeal arch-foregut junction, tracheal-esophageal septum, and the tracheal-bronchial junction to measure the undivided foregut length and the trachea length.
Immunofluorescence of tissue sections
Whole embryos, dissected foreguts, or ex vivo cultured foregut slices were fixed in 4% paraformaldehyde diluted in PBS overnight at 4 °C. Fixed tissue was washed three times with PBS (10 min each at room temperature), and cryopreserved with a series of 15% sucrose (Sigma-Aldrich, S8501) in PBS, 30% sucrose in PBS, and 1:1 O.C.T. Compound (Sakura Finetek, 4583):30% sucrose in PBS (1 h each at room temperature). Samples were transferred to and embedded in O.C.T. Compound, frozen in the dry ice-ethanol bath, before stored at −80 °C. Frozen samples were transversally sectioned using a cryostat (Leica, CM3050 S) to 14–20 µm thickness onto Superfrost Plus glass slides (Fisher Scientific, 12-550-15). The sample slides were baked at 50 °C for 20 min to dry, and then rinsed three times with PBS (5 min each at room temperature). For labeling phospho-MLC, TBX1, and SHH, antigen retrieval was performed with 1X citrate buffer (Abcam, 64214) in a boiling steamer (Amazon, B00DPX8UBA) for 10 min. The sample was then cooled down to room temperature and rinsed two times with PBS (5 min each at room temperature). For immunolabeling, the samples were first permeabilized with 0.1% Triton X-100 in PBS for 20 min, and blocked with the Blocking Buffer (5% donkey serum [Jackson ImmunoResearch, 017-000-121] and 0.3% Triton X-100 in PBS) for one hour. The primary antibodies were diluted in the Blocking Buffer and incubated with the sample overnight at 4 °C. The labeled samples were washed with the Washing Buffer (10X-diluted Blocking Buffer in PBS) three times (10 min each at room temperature), and labeled with secondary antibodies (including 10 µg/mL DAPI [Invitrogen, D1306] or 1:100 phalloidin-Alexa Fluor 488 [Invitrogen, A12379]) diluted in the Blocking Buffer and incubated for two hours at room temperature. After three washes with the Washing Buffer, the samples were mounted in ProLong Diamond Antifade Mountant (Invitrogen, P36970) with #1 coverglass (VWR, 48393-106).
Primary antibodies used were: rat anti-SOX2 (1:300, Invitrogen, 14-9811-82), rabbit anti-NKX2-1 (1:300, Abcam, 76013), rabbit anti-cleaved Caspase 3 (1:300, Cell Signaling Technology, 9661), rabbit anti-phospho-Histone H3 (1:300, Cell Signaling Technology, 3377), rabbit anti phospho-MLC2 (1:100, Cell Signaling Technology, 3674), rabbit anti-mCherry (1:500, Abcam, 167453, for labeling tdTomato), biotinylated Hyaluronic Acid Binding Protein (1:200, Sigma-Aldrich, 385911), mouse anti-GM130 (1:150, BD Biosciences, 610822), rabbit anti-phospho-FAK (1:200, Invitrogen, 700255), goat anti-NKX6-1 (10 µg/mL, R&D Systems, AF5857), rabbit anti-TBX1 (1:50, Invitrogen, 34-9800), mouse anti-ISL1 (1:30, DSHB, 40.2D6), mouse anti-ALDH1A2 (1:50, Santa Cruz Biotechnology, sc-393204), goat anti-SHH (1:100, R&D Systems, AF464). Dye-conjugated secondary antibodies were from Jackson ImmunoResearch (with Alexa Fluor 647, Cy3, or Alexa Fluor 488) and used at 1:300 dilution, except the streptavidin-Alexa Fluor 647 for HABP detection (1:300, Invitrogen, S32357).
Tissue sections were imaged with a Nikon Ti inverted microscope with a W1 spinning disk scanner (Yokogawa, CSU-W1) using a 20x or 40x objective (Nikon, MRH01401). Z-stacks with 0.3–0.9 µm step size were acquired, and maximum intensity projection was performed with a custom Python program.
Hybridization chain reaction fluorescent in situ hybridization (HCR-FISH)
Probes for HCR-FISH were designed by the insitu_probe_generator (https://github.com/rwnull/insitu_probe_generator)56 using the coding sequences of chick PTCH1, chick ROR2, chick PRICKLE1, mouse Ptch1, and mouse Hhip from the Ensembl Release 112. Generated probes were blasted against the respective species’ transcriptome to exclude potential off-target probes. The designed probes with HCR 3.0 barcode sequences (Molecular Instruments) were synthesized as 50 pmol oligo pools (oPools) by Integrated DNA Technologies and reconstituted in the TE buffer (Qiagen, 19086) to 1 µM.
Dissected foregut tissues were fixed in a 1.7 mL microcentrifuge tube with 4% PFA for one hour at room temperature, and washed twice with PBS (5 min each). The samples were then dehydrated and permeabilized with 70% ethanol (VWR, V1001) in PBS for one hour at room temperature. The samples were equilibrated with the probe wash buffer (Molecular Instruments) for 10 min at room temperature and then with the probe hybridization buffer (Molecular Instruments) for 30 min at 37 °C (without rocking). Probes were diluted to 40 nM in 100 µL of hybridization buffer, incubating the samples overnight at 37 °C without rocking. Labeled samples were washed twice with the wash buffer, then twice with 5X SSCT (5X Saline-Sodium Citrate buffer [Invitrogen, 15557044] and 0.1% Tween 20 [Sigma-Aldrich, P9416] diluted in PBS), and finally with the HCR amplification buffer (Molecular Instruments), 20 min each at room temperature. The HCR amplifiers with Alexa Fluor 647 or Alexa Fluor 546 (Molecular Instruments) were denatured at 95 °C for 90 s and annealed at room temperature for 30 min. The amplifiers were diluted in the amplification buffer at 37 nM per strand, which was incubated with the sample overnight at room temperature without rocking. Two 5X SSCT washes were performed before embedding and sectioning in O.C.T. A 30-min DAPI labeling in the Blocking Buffer was performed on the sections before mounting.
Single-cell RNA sequencing (scRNA-seq)
Library preparation and sequencing
Control E3.5 (N = 6), E4.0 (N = 4), and cyclopamine-treated E4.0 (N = 9) foreguts were dissected and dissociated to single cells with TrypLE (Gibco, 12604013) at 37 °C with intermittent trituration with a 25 Gauge needle. Dissociated cells were spun down at 800 g for 3 min at 4 °C and washed twice with cold PBS. We performed MULTI-seq lipid barcode labeling on ice to multiplex the samples as described (Sigma-Aldrich, LMO001)57. After rinsing off the unlabeled barcode and anchors, the cell suspension was passed through a 40 µm Falcon cell strainer (VWR, 21008-949), spun at 800 g for 5 min at 4 °C, and resuspended to 20 µL at a density of ~500 cells/µL. All cells were used for Gel Bead-In Emulsions (GEM) with a Chromium Controller (10x Genomics) per manufacturer’s instructions. Library construction was performed with the Chromium Next GEM Single Cell 3’ Kit (v3.1) with dual index (10x Genomics). Quality control of the library was performed by the Biopolymers Facility at Harvard Medical School.
The 10-nM library was sequenced with the NovaSeq 6000 platform at the Biopolymers Facility at Harvard Medical School, with the configuration of 28 bp for cell barcode 1 and UMI, 8 bp for i7 index, 10 bp for i5 index, and 90 bp for the transcript.
Demultiplexing by embryo and sample origin (MULTI-seq)
All libraries were mapped using Cell Ranger (7.1.0) and the associated custom reference that includes EGFP (see Data Availability). Samples of different groups were demultiplexed using the deMULTIplex package in RStudio (https://github.com/chris-mcginnis-ucsf/MULTI-seq) based on the counts of MULTI-seq barcodes. Leveraging the intrinsic genetic variation of chick embryos, individual cells were assigned to different embryos within each sample using the vireo package (https://github.com/single-cell-genetics/vireo)58. Doublets identified from MULTI-seq or vireo analysis were excluded from downstream analysis.
Quality control and clustering
Single-cell data was processed with Seurat V4 in RStudio (https://github.com/satijalab/seurat)59. A set of criteria were used to select high quality cells for downstream analysis: nCount_RNA > = 1000, nFeature_RNA > = 500, log10GenesPerUMI > 0.80, percent.mito <0.18, percent.rbc <0.20. After filtration, we ended up with 8,021 cells for control E3.5, 4915 cells for control E4.0, and 4829 cells for cyclopamine-treated E4.0. Differently treated samples were integrated using SCTransform of Seurat after regression of cell cycle and mitochondrial fractions. We used Uniform Manifold Approximation and Projection (UMAP) to perform dimension reduction of the integrated data. Louvain algorithm was used to cluster cells with 40 principal components based on the elbow plot, with a resolution of 0.6, resulting in 15 clusters. We annotated the clusters by their marker genes identified by FindMarkers, and verified their spatial distribution by HCR or immunostaining. For each cell cluster, we computed their abundance in total cells from the same embryo inferred by demultiplexing.
Differentially expressed gene (DEG) analysis
Given the demultiplexed biological replicates in each group, we performed pseudo-bulk DEG analysis using the glmGamPoi package in RStudio (https://github.com/const-ae/glmGamPoi)60. Cells of the dorsal sub-epithelial mesenchyme cluster were grouped into control (E3.5 and E4.0) and cyclopamine. Genes with adjusted P-values < 0.001 were identified as DEGs. Volcano plot was generated by the EnhancedVolcano package (https://github.com/kevinblighe/EnhancedVolcano). Control group-enriched DEGs were imported into Shiny GO V0.77 (http://bioinformatics.sdstate.edu/go/)61 using the AmiGo 2 gene ontology database (https://amigo.geneontology.org/amigo) with a false discovery rate threshold of 0.1.
For ACTA2 expression analysis, as this gene is expressed at a very low level at E4, prior to smooth muscle differentiation, we performed DEG analysis with FindMarkers in Seurat.
Cell-cell communication analysis
LIANA framework (https://github.com/saezlab/liana-py)62,63 was used to analyze potential ligand-receptor communication pathways between the dorsal epithelium and the dorsal mesenchymal cell types. CellPhoneDB64 with 100 permutations was used to identify ligand-receptor pairs, and those with P-values < 0.05 were selected.
Image analysis of immunofluorescence tissue sections
SOX2 gradient
SOX2 intensities were quantified in 12 equal, dorsal-ventrally spaced bins along the foregut epithelium in Fiji and normalized to the most dorsal bin.
pMLC apical-basal ratio
Line profiles normal to the epithelial surface were drawn in Fiji, and the local peak intensity values at the apical and the basal side of the epithelium were taken. The ratio between the apical and basal intensity of the same tissue was calculated, which reflects the extent of apical constriction.
Cell roundness
MLC2-tdTomato-electroporated cells were manually segmented, and cell roundness was calculated in Fiji (https://imagej.net/ij/docs/menus/analyze.html) by:
Cell orientation analysis
Relative positions of the Golgi and the nucleus were used to define cell orientation. Nuclei were segmented with StarDist in TrackMate. The Golgi apparatus was segmented with the LoG detector in TrackMate. The coordinates of the segmented structures were exported to MATLAB, and we wrote a custom code to pair the Golgi and the nucleus with the costMatrix function. Due to the thinness of sections, we often had extra, unpaired nuclei. We thus defined the nucleus-Golgi vectors and computed the angular histograms of their directions. For the epithelium-mesenchyme interface, 0 degree was defined as the vector pointing normal to the epithelium. For bead implantation, 0 degree was defined as the vector pointing towards the center of the bead.
pHH3 and CC3 analysis
As pHH3 and CC3 labels are sparse within the tissue, we define their frequencies by the number of positive cells per unit area of the mesenchyme. Positive cells were counted manually, and the area of mesenchyme was measured in Fiji.
Epithelium right-left distance ratio with one-sided electroporation
We defined the midline of the epithelium by connecting the dorsal and ventral points with the highest curvature. The one-sided distance was measured as the perpendicular line segment from the center of the electroporated cell patch’s basal surface to the midline. The ratio of right (electroporated) to left (non-electroporated) distances was then calculated.
Morphometric analysis of the foregut epithelium
The time-lapse videos of TES in slice culture were first processed in Fiji, and the Segment Anything Model 2 (SAM 2, https://github.com/facebookresearch/sam2)65 was applied to segment the epithelium as a mask. The mask was then processed with the Open Source Computer Vision Library (OpenCV, https://github.com/opencv/opencv) to extract the contour and conduct downstream morphometric analyses.
Video pre-processing
The Bleach Correction function in Fiji66 was used to correct for photobleaching, and StackReg to correct translational and rotational drifts in the rigid body mode, given that the morphological change of the slice was small between two consecutive frames. The corrected video was then cropped to center the epithelium and Gaussian blurred to reduce cellular granularity for higher segmentation accuracy.
Automated segmentation by SAM 2
The pre-trained sam2.1_hiera_large model of SAM 2 was used to segment the foregut epithelium in the time-lapse video. At least one point on the epithelial region was selected as a guide. However, when a single point selection led to misidentification, additional points were used to refine the segmentation. In some cases, bounding boxes were applied in conjunction with points to further constrain the region and isolate the epithelium accurately. This combination of point selection and bounding boxes ensured precise segmentation yielding a > 95% success rate for all chick videos. For the remaining frames, we manually segmented the epithelium in Fiji and replaced the wrong segmentations. The segmentation results were saved as binary masks.
However, segmentation of mouse videos remained challenging because the nTnG mouse slices had nuclear labeling, which made the epithelium more granular than the GFP chick slices. Therefore, in this work, we only quantify the morphological evolution of chick TES.
Morphometric analysis
From the binary segmentation, we found the outline of the shape using the OpenCV wrapper in Python. For each frame of a given sample, we fitted an ellipse using the fitEllipse function in OpenCV. The height and width of the rectangle in which the ellipse is inscribed was used as measures of the shape’s dimensions. We chose to fit an ellipse because we found it yields a more representative measure of the shape than, for example, a rectangle enclosing the 2D point set. Additionally, we computed the arc length and area of the shape using OpenCV.
To calculate the curvature of an outline, we first fitted a B-spline to the 2D point set of each frame in a given video. This was done using the function BSplineFunction in Mathematica. The degree of the B-spline was chosen to be one-quarter of the total number of points in the outline for a given frame. We found this value to be a good compromise between smoothing the noisy data sufficiently for the following operations (namely, computing the curvature) and preserving the high-curvature areas in the neck region of interest. For example, the area and arc length of the fitted curve agree well with the values obtained from the discrete points using OpenCV described above. This thus resulted in a smooth curve γ(s), with arc-length parametrization s. From this, the curvature was computed using the equation:
To define the neck region, we proceeded as follows: For each frame, we first selected the point s = s0 at which the curvature reaches its global minimum. We then selected a second point s1 from the region s = s0 + 0.5 ± 0.2 at which the curvature takes its local minimum in this region. We followed this procedure instead of simply picking the two smallest-curvature points to reduce errors due to noise and segmentation inaccuracies. We found that this method is robust, and the points identified in this way correspond well with those identified manually. Overall, across all videos, the procedure only failed in a few isolated frames when segmentation issues occur. The two identified points then define the neck. We repeated this procedure for all frames of a given video, resulting in a time evolution of the neck curvatures. To mitigate the impact of segmentation errors and noise, we performed a moving median over three frames for the two neck curvatures. The curvature was normalized by multiplication with the initial neck width of the epithelium to account for size variation in different samples.
To average the quantities from several videos of a specific developmental stage, we first interpolated the data obtained from each video. This is necessary since the videos were recorded at different temporal resolutions. Before interpolating, we removed outliers in the data, defined as points that deviate more than 2.5 standard deviations from the average of a given video. This helps reduce the effect of segmentation errors. We then aligned the videos in time using the manually identified frame at which the transition occurs. We computed the median and median deviation (MAD) across all videos for a given point in time and repeated this for all time points to minimize the effect of outliers due to inaccurate segmentation or contour extraction. For experimental groups with N < 6, individual curves were plotted instead of the population average.
Image central moments
We chose to use the central moments for extracting morphological information from the binary epithelial masks67. Central moments were calculated as:
Here \((\bar{x},\bar{y})\) is the mask centroid, \(I\left(x,y\right)\) represents the pixel intensity (1 or 0) at \(\left(x,y\right)\). The second-order central moments were normalized as:
PIV analysis
Drift-corrected, contrast-enhanced time-lapse videos were imported to PIVlab V2.62 in MATLAB68. The image frames were first pre-processed to highlight the individual cells, with CLAHE with a window size of 25 pixels, highpass filtering with a kernel size of 35 pixels, and Wiener2 denoising with a window size of 5 pixels. The PIV settings used were: FFT window deformation algorithm, Pass 1 with 128-pixel integration area and 64-pixel step size, Pass 2 with 64-pixel integration area and 32-pixel step size, and high correlation robustness. The calculated velocity field was filtered by X and Y velocities and a correlation coefficient filter of 0.4 to exclude outliers. The velocity fields were then temporally averaged with a window of one hour. Velocity vectors were recolored by their directions using Cyclic Color Map in MATLAB (by Chad Greene, https://www.mathworks.com/matlabcentral/fileexchange/57020-cyclic-color-map).
The strain rate and the divergence were computed based on the gradients of the averaged velocity fields with an additional 2 × 2 spatial binning using a custom Python code. We also calculated the eigenvectors of the two-dimensional strain rate tensor to indicate the principal direction of strain.
Single-cell tracking and trajectory analysis
Cells were tracked in Fiji TrackMate using the StarDist detector without frame gaps69. The trajectories and average velocity vectors were plotted with a custom MATLAB code. The angular histograms were computed with the velocity vectors in MATLAB. Mean squared displacements (MSD) were plotted for each trajectory (starting time defined as 0) with a population average and SD. The average MSD-t curve was fitted by a power function describing anomalous diffusion70:
Here, \(\alpha=1\) indicates normal diffusion, whereas \(\alpha=2\) indicates ballistic motion. \(1 < \alpha < 2\) suggests super-diffusion, which can be due to active cell migration in a crowded environment.
We also quantified the cell shape in GFP chick samples with good image contrasts. Similarly, cells were tracked with StarDist in TrackMate, and the aspect ratio (defined as the inverse of cell roundness) was calculated and plotted for individual cells.
For tracking cells in 3D, we took live imaging stacks with 2-µm Z step size (8–9 Z planes in total). The image stack was corrected for translational drift using Correct 3D drift71 (https://imagej.net/plugins/correct-3d-drift). Cells were segmented in 3D using a custom Arivis pipeline including background subtraction, denoising, and the Blob Finder segmenter. The segmented cells were then tracked in Arivis using the nearest neighbor algorithm with a maximal inter-frame displacement of 5 µm. The cell tracks were then exported to MATLAB for plotting and angle calculations.
Numerical solution of SHH diffusion
The two-dimensional diffusion equation was solved numerically using the NDSolve function in Wolfram Mathematica 14.2 with the default options.
Statistics and Reproducibility
The number of samples used for each experiment and statistical tests were indicated in the figure legends. All microscopy images are representative of at least four independent experiments. The sample sizes were not pre-determined. GraphPad Prism software was used to plot the data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The chick scRNA-seq data have been deposited to the Gene Expression Omnibus of NCBI under the access code GSE307539. For inquiries about other data or materials used in this study, please reach out to the corresponding authors. Source data are provided with this paper.
Code availability
The custom codes used for image processing and data analysis are available on GitHub (https://github.com/yanrui21/Foregut_septation_N).
References
Heisenberg, C. P. & Bellaïche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Tozluoglu, M. & Mao, Y. On folding morphogenesis, a mechanical problem. Philos. Trans. R. Soc. B Biol. Sci. 375, 20190564 (2020).
Lecuit, T., Lenne, P. F. & Munro, E. Force generation, transmission, and integration during cell and tissue morphogenesis. Annu. Rev. Cell Dev. Biol. 27, 157–184 (2011).
Pearl, E. J., Li, J. & Green, J. B. A. Cellular systems for epithelial invagination. Philos. Trans. R. Soc. B Biol. Sci. 372, 20150526 (2017).
Durel, J. F. & Nerurkar, N. L. Mechanobiology of vertebrate gut morphogenesis. Curr. Opin. Genet. Dev. 63, 45–52 (2020).
Hughes, A. J. et al. Engineered tissue folding by mechanical compaction of the mesenchyme. Dev. Cell 44, 165–178.e6 (2018).
Huycke, T. R. et al. Patterning and folding of intestinal villi by active mesenchymal dewetting. Cell 187, 3072–3089.e20 (2024).
Billmyre, K. K., Hutson, M. & Klingensmith, J. One shall become two: separation of the esophagus and trachea from the common foregut tube. Dev. Dyn. 244, 277–288 (2015).
Que, J. The initial establishment and epithelial morphogenesis of the esophagus: a new model of tracheal-esophageal separation and transition of simple columnar into stratified squamous epithelium in the developing esophagus. Wiley Interdiscip. Rev. Dev. Biol. 4, 419–430 (2015).
Seifert, A. W., Bouldin, C. M., Choi, K. S., Harfe, B. D. & Cohn, M. J. Multiphasic and tissue-specific roles of sonic hedgehog in cloacal septation and external genitalia development. Development 136, 3949–3957 (2009).
Nasr, T. et al. Endosome-mediated epithelial remodeling downstream of Hedgehog-Gli is required for tracheoesophageal separation. Dev. Cell 51, 665–674 (2019).
Zorn, A. M. & Wells, J. M. Vertebrate endoderm development and organ formation. Annu. Rev. Cell Dev. Biol. 25, 221–251 (2009).
Lewis, A. E., Kuwahara, A., Franzosi, J. & Bush, J. O. Tracheal separation is driven by NKX2-1-mediated repression of Efnb2 and regulation of endodermal cell sorting. Cell Rep. 38, 110510 (2022).
Durkin, N. & De Coppi, P. Anatomy and embryology of tracheo-esophageal fistula. Semin. Pediatr. Surg. 31, 151231 (2022).
Edwards, N. A. et al. Developmental basis of trachea-esophageal birth defects. Dev. Biol. 477, 85–97 (2021).
van Lennep, M. et al. Oesophageal atresia. Nat. Rev. Dis. Prim. 5, 26 (2019).
Que, J. et al. Multiple dose-dependent roles for Sox2 in the patterning and differentiation of anterior foregut endoderm. Development 134, 2521–2531 (2007).
Jacobs, I. J., Ku, W. Y. & Que, J. Genetic and cellular mechanisms regulating anterior foregut and esophageal development. Dev. Biol. 369, 54–64 (2012).
Domyan, E. T. et al. Signaling through BMP receptors promotes respiratory identity in the foregut via repression of Sox2. Development 138, 971–981 (2011).
Kim, E. et al. Isl1 regulation of Nkx2.1 in the early foregut epithelium is required for trachea-esophageal separation and lung lobation. Dev. Cell 51, 675–683 (2019).
Fausett, S. R., Brunet, L. J. & Klingensmith, J. BMP antagonism by noggin is required in presumptive notochord cells for mammalian foregut morphogenesis. Dev. Biol. 391, 111–124 (2014).
Que, J., Luo, X., Schwartz, R. J. & Hogan, B. L. M. Multiple roles for Sox2 in the developing and adult mouse trachea. Development 136, 1899–1907 (2009).
Kuwahara, A. et al. Delineating the early transcriptional specification of the mammalian trachea and esophagus. Elife 9, e55526 (2020).
Ioannides, A. S. et al. Foregut separation and tracheo-oesophageal malformations: The role of tracheal outgrowth, dorso-ventral patterning and programmed cell death. Dev. Biol. 337, 351–362 (2010).
Munjal, A., Hannezo, E., Tsai, T. Y. C., Mitchison, T. J. & Megason, S. G. Extracellular hyaluronate pressure shaped by cellular tethers drives tissue morphogenesis. Cell 184, 6313–6325.e18 (2021).
Gill, H. K. et al. The developmental mechanics of divergent buckling patterns in the chick gut. Proc. Natl Acad. Sci. Usa. 121, e2310992121 (2024).
Shroff, N. P. et al. Proliferation-driven mechanical compression induces signalling centre formation during mammalian organ development. Nat. Cell Biol. 26, 519–529 (2024).
Villedieu, A., Bosveld, F. & Bellaïche, Y. Mechanical induction and competence in epithelial morphogenesis. Curr. Opin. Genet. Dev. 63, 36–44 (2020).
Kishimoto, K. et al. Synchronized mesenchymal cell polarization and differentiation shape the formation of the murine trachea and esophagus. Nat. Commun. 9, 2816 (2018).
Litingtung, Y., Lei, L., Westphal, H. & Chiang, C. Sonic hedgehog is essential to foregut development. Nat. Genet. 20, 58–61 (1998).
Motoyama, J. et al. Essential function of Gli2 and Gli3 in the formation of the lung, trachea and oesophagus. Nat. Genet. 20, 54–57 (1998).
Ioannides, A. S., Henderson, D. J., Spitz, L. & Copp, A. J. Role of Sonic hedgehog in the development of the trachea and oesophagus. J. Pediatr. Surg. 38, 29–36 (2003).
Han, L. et al. Single-cell transcriptomics identifies a signaling network coordinating endoderm and mesoderm diversification during foregut organogenesis. Nat. Commun. 11, 4158 (2020).
Mao, J., Kim, B. M., Rajurkar, M., Shivdasani, R. A. & McMahon, A. P. Hedgehog signaling controls mesenchymal growth in the developing mammalian digestive tract. Development 137, 1721–1729 (2010).
Han, L. et al. Osr1 functions downstream of the Hedgehog pathway to regulate foregut development. Dev. Biol. 427, 72–83 (2017).
Roellig, D. et al. Force-generating apoptotic cells orchestrate avian neural tube bending. Dev. Cell 57, 707–718.e6 (2022).
Yang, Q. et al. Cell fate coordinates mechano-osmotic forces in intestinal crypt formation. Nat. Cell Biol. 23, 733–744 (2021).
Gill, H. K. et al. Hox gene activity directs physical forces to differentially shape chick small and large intestinal epithelia. Dev. Cell 59, 1–16 (2024).
Villeneuve, C. et al. Mechanical forces across compartments coordinate cell shape and fate transitions to generate tissue architecture. Nat. Cell Biol. 26, 207–218 (2024).
Mammoto, T. et al. Mechanochemical control of mesenchymal condensation and embryonic tooth organ formation. Dev. Cell 21, 758–769 (2011).
Ribatti, D. & Santoiemma, M. Epithelial-mesenchymal interactions: a fundamental developmental biology mechanism. Int. J. Dev. Biol. 58, 303–306 (2014).
Walton, K. D. et al. Hedgehog-responsive mesenchymal clusters direct patterning and emergence of intestinal villi. Proc. Natl Acad. Sci. Usa. 109, 15817–15822 (2012).
Rao-Bhatia, A. et al. Hedgehog-activated Fat4 and PCP pathways mediate mesenchymal cell clustering and villus formation in gut development. Dev. Cell 52, 647–658.e6 (2020).
Kindberg, A. A. et al. Eph/ephrin regulates cellular organization by actomyosin contractility effects on cell contacts. J. Cell Biol. 220, e202005216 (2021).
Edwards, N. A. et al. Disrupted endosomal trafficking of the Vangl-Celsr polarity complex underlies congenital anomalies in Xenopus trachea-esophageal morphogenesis. Dev. Cell 60, 1–16 (2025).
Harris-Johnson, K. S., Domyan, E. T., Vezina, C. M. & Sun, X. β-Catenin promotes respiratory progenitor identity in mouse foregut. Proc. Natl Acad. Sci. USA 106, 16287–16292 (2009).
Li, Y., Litingtung, Y., ten Dijke, P. & Chiang, C. Aberrant Bmp signaling and notochord delamination in the pathogenesis of esophageal atresia. Dev. Dyn. 236, 746–754 (2007).
Mahlapuu, M., Enerbäck, S. & Carlsson, P. Haploinsufficiency of the forkhead gene Foxf1, a target for sonic hedgehog signaling, causes lung and foregut malformations. Development 128, 2397–2406 (2001).
Bermudez, O., Hennen, E., Koch, I., Lindner, M. & Eickelberg, O. Gli1 mediates lung cancer cell proliferation and Sonic Hedgehog-dependent mesenchymal cell activation. PLoS One 8, e63226 (2013).
Jeng, K. S., Chang, C. F. & Lin, S. S. Sonic hedgehog signaling in organogenesis, tumors, and tumor microenvironments. Int. J. Mol. Sci. 21, 758 (2020).
Chen, J. S. et al. Sonic hedgehog signaling pathway induces cell migration and invasion through focal adhesion kinase/AKT signaling-mediated activation of matrix metalloproteinase (MMP)-2 and MMP-9 in liver cancer. Carcinogenesis 34, 10–19 (2013).
Nees, S. N., Jelin, E. & Chung, W. K. Genetics of common birth defects in newborns. in Principles of Neonatology 677–689 (Elsevier Inc., 2023).
Webber, D. M. et al. Developments in our understanding of the genetic basis of birth defects. Birth Defects Res. Part A - Clin. Mol. Teratol. 103, 680–691 (2015).
Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. image Process. 7, 27–41 (1998).
Liang, X., Michael, M. & Gomez, G. A. Measurement of mechanical tension at cell-cell junctions using two-photon laser ablation. Bio-Protoc. 6, e2068 (2016).
Kuehn, E. et al. Segment number threshold determines juvenile onset of germline cluster expansion in Platynereis dumerilii. J. Exp. Zool. Part B Mol. Dev. Evol. 338, 225–240 (2022).
McGinnis, C. S. et al. MULTI-seq: Sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
Huang, Y., McCarthy, D. J. & Stegle, O. Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol. 20, 273 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Ahlmann-Eltze, C. & Huber, W. glmGamPoi: fitting Gamma-Poisson generalized linear models on single cell count data. Bioinformatics 36, 5701–5702 (2020).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Dimitrov, D. et al. LIANA+ provides an all-in-one framework for cell–cell communication inference. Nat. Cell Biol. 26, 1613–1622 (2024).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-Seq data. Nat. Commun. 13, 3224 (2022).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Ravi, N. et al. Sam 2: Segment anything in images and videos. International Conference on Learning Representations (ICLR) (2025).
Miura, K. Bleach correction ImageJ plugin for compensating the photobleaching of time-lapse sequences. F1000Research 9, 1494 (2020).
Flusser, J. Moment invariants in image analysis. Proc. World Acad. Sci. Eng. Technol. 11, 196–201 (2006).
Thielicke, W. & Sonntag, R. Particle image velocimetry for MATLAB: accuracy and enhanced algorithms in PIVlab. J. Open Res. Softw. 9, 12 (2021).
Ershov, D. et al. TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines. Nat. Methods 19, 829–832 (2022).
Codling, E. A., Plank, M. J. & Benhamou, S. Random walk models in biology. J. R. Soc. Interface 5, 813–834 (2008).
Parslow, A., Cardona, A. & Bryson-Richardson, R. J. Sample drift correction following 4D confocal time-lapse imaging. J. Vis. Exp 12, 51086 (2014).
Acknowledgments
We thank the MicRoN Facility of Harvard Medical School for offering equipment and training for microscopy experiments, the Biopolymers Facility of Harvard Medical School for performing next-generation sequencing, and the Center for Comparative Medicine of Harvard Medical School for maintenance of the mouse colonies. We thank members of the Tabin group and the Mahadevan group for their discussion and suggestions. This work was funded by the NIH grant HD087234 (C.J.T.), the Helen Hay Whitney Fellowship (R.Y.), the NWO Rubicon grant (L.A.H.), the Simons Foundation (L.M.), and the Henri Seydoux Fund (L.M.).
Author information
Authors and Affiliations
Contributions
R.Y. conceived the project. R.Y. and C.J.T. designed the experiments and wrote the draft of the manuscript. R.Y. performed all experiments, analyzed the data, and wrote the code for the plot figures. L.A.H. performed analysis and quantification of the foregut morphology in slice culture and gave advice on image processing and PIV analysis. P.O. wrote the code for computing the strain rate tensor from the PIV data. D.L. wrote the code for SAM 2 segmentation. C.L. performed initial processing and analysis of the single-cell RNA-sequencing data. H.K.G. gave advice and technical support on in ovo electroporation and slice culture. A.M., N.L.N., and L.M. gave advice on experimental design and data analysis. L.M. and C.J.T. supervised the study. All authors read the manuscript and gave suggestions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Pierre Recouvreux and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yan, R., Hoffmann, L.A., Oikonomou, P. et al. Convergent flow-mediated mesenchymal force drives embryonic foregut constriction and splitting. Nat Commun 16, 10643 (2025). https://doi.org/10.1038/s41467-025-65644-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65644-9