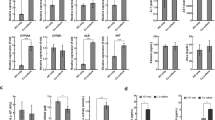
Abstract
Drug-Induced Liver Injury (DILI) remains one of the most critical challenges in drug development, causing patient safety concerns, clinical trial failures and drug withdrawals. We introduce ToxPredictor, a toxicogenomics framework combining RNA-seq data from primary human hepatocytes with pharmacokinetic data to predict dose-resolved DILI risks and safety margins. At its core is DILImap, an RNA-seq library tailored for DILI research, comprising 300 compounds at multiple concentrations. ToxPredictor achieves 88% sensitivity at 100% specificity in blind validation, outperforming state-of-the-art methods. It flagged recent phase III clinical failures, including Evobrutinib, TAK-875, and BMS-986142, overlooked by animal studies. Beyond prediction, ToxPredictor provides mechanistic insights into hepatotoxic pathways, enabling early de-risking and actionable safety decisions. Unlike single-endpoint readouts—even from 3D models—transcriptomics offers a multi-dimensional system-level view of hepatocyte responses, capable of detecting diverse DILI mechanisms not captured by conventional assays. Scalable, actionable, and integrated into a broader AI/ML drug discovery platform, this work establishes toxicogenomics as a promising tool for developing safer therapeutics and addressing one of the most pressing challenges in toxicology.
Similar content being viewed by others
Introduction
Drug-Induced Liver Injury (DILI) presents a poorly understood late-stage challenge in drug development, costing an estimated $350 million annually per pharmaceutical company1. Its rarity and unpredictability in clinical populations, often less than 1 in 10,000 persons, hinder detection in clinical studies, masking its severity until post-market exposure. Animal models fail to identify about half of the pharmaceuticals that exhibit clinical DILI2. This makes DILI a leading cause of drug candidate failure and market withdrawal, impeding the development of new therapies3.
DILI arises from complex, multifactorial mechanisms, involving dose-dependent intrinsic mechanisms and with current methods unpredictable idiosyncratic reactions4,5 influenced by genetic predisposition, environment, and individual health status6. DILI involves various cellular disruptions including mitochondrial dysfunction, oxidative stress, bile acid imbalance, inhibition of specific enzymes or transporters, and reactive metabolites formation7,8. However, the precise mechanisms and contributing factors are not fully delineated4 and the lack of biomarkers hampers early detection9. Pre-clinical methods, such as quantitative structure-activity relationship (QSAR) models, offer low specificity and binary predictions lacking mechanistic insights10,11. In vitro models use diverse cell sources (e.g., HepG2, THLE, HepaRG cells, primary human hepatocytes) with endpoints ranging from cytotoxicity markers (e.g., LDH, ATP) to mechanistic assessments using high-content imaging (HCI) and multi-parametric strategies. 3D liver models aim to better mimic human tissue characteristics and in vivo responses12. However, despite improved physiological relevance, these models remain constrained by low-dimensional readouts—typically a limited panel of markers such as ATP levels, LDH release, or imaging-based features—which fail to capture the full spectrum of molecular responses and often miss the mechanistic cause. This gap continues to result in late-stage drug withdrawals and clinical failures, highlighting the urgent need for more comprehensive and predictive DILI models9.
Inspired by the idea of viewing cells as complex systems, we adopt a machine learning-driven toxicogenomics approach to analyze how DILI-associated compounds alter gene expression, identifying early gene signatures indicative of liver injury. Encompassing the interplay of pathways in response to toxic compounds, we aim to decipher the molecular mechanisms underlying DILI. Utilizing resources like DILIrank13 and LiverTox14 for drug categorization into DILI positives and negatives, we employed the TG-GATES15 microarray database for an initial proof-of-concept, enabling us to accurately predict DILI with 62% sensitivity and 92% specificity in blind validation. Building on this initial success, we created DILImap, a purpose-built and significantly expanded transcriptomic library designed to capture a broader spectrum of DILI mechanisms. DILImap features full-transcriptome RNA-seq data from 300 compounds profiled at multiple concentrations in primary human hepatocytes (PHHs), making it, to the best of our knowledge, the largest toxicogenomics dataset available for DILI modeling.
ToxPredictor, our random forest-based machine learning model trained on DILImap, achieves 88% sensitivity (29/33 DILI positives) at 100% specificity (14/14 DILI negatives) in blind validation. It outperforms 20+ pre-clinical models in a head-to-head comparisons, including mechanistic assays16,17,18,19,20,21, cytotoxicity markers22,23,24,25,26,27, physicochemical properties28, bioactivation29, BSEP30 approaches, and the latest in-silico models31,32,33, effectively identifying DILI risks in drugs previously overlooked by traditional models. To the best of our knowledge, it is the first pre-clinical model to flag DILI risks in high-profile clinical failures, such as Evobrutinib, TAK-875 and BMS-986142, all recently withdrawn in phase III trials due to liver injury despite clean preclinical profiles. The model provides dose-resolved predictions and mechanistic insights, demonstrating its utility for prioritizing safer drug candidates.
Beyond generalization to unseen compounds, the model has a distinct edge in its mechanistic breadth. The model leverages the full transcriptomic landscape to detect DILI-related mechanisms such as mitochondrial dysfunction, oxidative stress, immune activation, and metabolic perturbation—often well before cell death. These advantages are particularly evident when compared to high-content 3D liver assays26,34,35,36, which, while physiologically relevant, are typically constrained to low-dimensional viability or imaging endpoints. In head-to-head comparisons, our model uniquely identifies non-cytotoxic risks missed by 3D assays. This systems-level resolution enables more comprehensive and unbiased detection of toxic liabilities across diverse compound classes.
This work, integral to a broader AI/ML drug discovery platform, aims at enhancing predictive power and operational efficiency in drug development. It showcases that the shift from a single-target to a systems-level perspective holds great promise and positions machine learning in toxicogenomics as significant enhancement to existing methods. To advance the field and foster collaborative innovation, we have made our open-source model and validation data publicly available at dilimap.org, providing a powerful tool for de-risking drug candidates and setting the stage for a paradigm shift in safety evaluations.
Results
DILImap – a human toxicogenomics database for DILI modeling
We have created DILImap, a comprehensive RNA-seq library tailored for drug-induced liver injury (DILI) modeling, encompassing 300 compounds tested at four concentrations. As the most extensive toxicogenomics resource to date, DILImap includes a curated selection of DILI-positive and DILI-negative compounds that span a wide range of known DILI mechanisms, including well-documented liver-injuring drugs and idiosyncratic compounds with no characteristic signature (Fig. 1A).
A We created DILImap, the largest toxicogenomics dataset to date, encompassing RNA-seq profiles from 300 compounds tested at multiple concentrations in primary human hepatocytes. Compounds span a range of DILI mechanisms, idiosyncratic effects, and non-DILI controls, ensuring broad mechanistic coverage. This resource forms the foundation for training our DILI model. B ToxPredictor, our machine learning model trained on the DILImap, learns pathway signatures indicative of DILI risk. C ToxPredictor de-risks new drug candidates by estimating DILI risk and safety margins. In blind validation, it achieved 88% sensitivity (29/33 DILIs detected) and 100% specificity (all 14 non-DILI compounds accurately classified). This framework sets the stage for a paradigm shift in safety evaluations.
All compounds were screened in sandwich-cultured primary human hepatocytes (PHHs), the gold standard and most physiologically relevant in vitro model for liver toxicity, which preserve key hepatic functions such as metabolic activity and bile canaliculi formation37,38. Each compound was tested in triplicate across six concentrations using lactate dehydrogenase (LDH) and Adenosine Triphosphate (ATP) cell viability assays. RNA-seq profiling was performed at four selected doses, spanning the pharmacologically relevant range from therapeutic plasma Cmax to the highest tolerated non-cytotoxic dose just below the IC₁₀ threshold (Supplementary Fig. S1).
We selected a 24-hour post-exposure time point, based on the trade-off between signal strength and cellular viability: earlier time points (e.g., 2 h or 8 h) yield weaker transcriptional responses39, while longer incubations risk hepatocyte de-differentiation and RNA degradation40. This strategy allowed us to capture early transcriptional responses without compromising RNA integrity. Marker analysis confirmed retention of hepatocyte identity at 24 hours. We further ensured data quality by including only wells with sufficient total RNA counts and low mitochondrial RNA content, indicating viable, transcriptionally active cells. This streamlined workflow—including automated solubility testing, viability screening, and IC₁₀-based dose selection—enabled us to profile 300 compounds in four months, including 110 drugs tested preclinically for the first time as part of a systematic benchmark (Supplementary Fig. S2; Methods). To support comprehensive benchmarking, we provide detailed annotations for each compound, including clinical DILI labels13, DILI mechanisms14, molecular information41, consensus plasma Cmax from various studies10,19,20,23,24,25,28,42,43, and DILI classification results from over 20 pre-clinical studies (Supplementary Data S1–S4).
Compounds were categorized based on DILIrank13 and LiverTox14 as follows (Supplementary Fig. S3):
-
Withdrawn DILI: withdrawals or clinical trial failures due to DILI (Most-DILI-Concern; withdrawn).
-
Known DILI: compounds with well-established clinical DILI risk (Most-DILI-Concern or LiverTox score A).
-
Likely DILI: drugs with documented liver injury cases (Most-DILI-Concern or LiverTox score A/B).
-
Idiosyncratic DILI: rare cases without clear dose–response (LiverTox score C/D, <12 case reports).
-
Unlikely DILI: discordant or weak evidence across databases (Less-DILI-concern, but LiverTox score E).
-
No DILI: compounds with no documented hepatotoxicity (No-DILI-Concern; LiverTox score E).
Withdrawn, Known, and Likely DILI serve as positive controls; No DILI as negative controls; while Idiosyncratic and Unlikely DILI, due to their label ambiguity, are excluded from training and reserved for downstream testing.
The training dataset includes 249 compounds (111 DILI + , 52 DILI-, 17 unlikely DILI, 69 idiosyncratic DILI) for cross-validation. For blind validation, a separate experiment was conducted using an independent set of 51 compounds (33 DILI + , 14 DILI-, and 4 with unknown labels, including real-world clinical failures). This carefully curated dataset provides a robust foundation for predictive modeling and mechanistic insights into DILI.
ToxPredictor – pathway-level toxicogenomics predicts DILI risk and therapeutic safety margins
ToxPredictor, a machine learning model trained on our DILImap library, predicts DILI risk from pathway-level transcriptional signatures. These signatures are derived through enrichment analysis (WikiPathways44, FDR-adjusted p-values) of genes differentially expressed between compound- vs. DMSO-treated samples using DESeq245, computed for each dose of every compound in DILImap (Fig. 1B; see Methods).
For model training, we used only compounds with unambiguous DILI labels, resulting in a high-confidence training set of 111 DILI+ (Withdrawn, Known, Likely) and 52 DILI− (No DILI), while the remaining training data were held out to assess model robustness. To ensure high-confidence DILI labels, we further restricted training to drug concentrations tested at more than 20x of their clinical Cmax to reduce the risk of false-negative labeling for DILI+ compounds that may appear safe at lower doses. For 5-fold cross-validation, we applied stratified, compound-level splitting to ensure that all doses and replicates of a given compound were held out together in each fold, mimicking real-world generalization to unseen compounds. From 193 tested configurations across eight model classes, we selected a Random Forest classifier for its strong validation AUC, minimal overfitting, and highest consistency across folds. These properties, combined with its interpretability, motivated its choice over more complex boosting and deep learning models (Supplementary Fig. S4).
The final model is an ensemble of 30 Random Forest models (ensemble members) trained on different cross-validation splits, which together enhance generalization and prediction stability. The ensemble size was chosen based on empirical benchmarking that showed stable test AUC and consistency between models (Supplementary Fig. S5). By estimating DILI probabilities across dose levels, ToxPredictor enables calculation of drug safety margins, defined as the ratio between the first predicted DILI dose (i.e., the lowest dose with predicted probability >0.7) and the maximum plasma concentration (Cmax) at therapeutic levels. This provides a transcriptomics-based surrogate of the clinical therapeutic window. A safety margin threshold of 80 provides an actionable classification into high- and low-risk compounds. The probability threshold of 0.7 and margin of safety (MOS) cutoff of 80 were both optimized on held-out training data to reach performance plateaus while minimizing false positives. Our selected MOS threshold of 80, while on the higher end of literature-reported ranges (10–100)17,23,28,34,36, reflects the greater sensitivity of transcriptomic assays compared to cytotoxicity or mechanistic readouts. Since transcriptional changes often occur at lower doses—before overt toxicity—a higher cutoff is needed to avoid false positives and maintain high specificity in a transcriptome-based model (Supplementary Fig. S5). Among available exposure measures, we used total Cmax instead of free Cmax due to its broader availability across compounds. Both measures showed comparable predictive performance, with total Cmax performing slightly better, possibly due to more robust consensus values derived from a greater number of studies (Supplementary Fig. S6).
All model selection, hyperparameter tuning, and threshold optimization were performed exclusively on the training data. For final evaluation, we used a fully independent blind-validation set of 51 compounds (33 DILI + , 14 DILI − , and 4 unknowns), profiled in a separate experiment using separate plates and sequencing runs. This set was withheld from all stages of model development. Compound selection for the validation study was finalized prior to training and intentionally enriched for withdrawals and recent clinical failures. The four unknowns represent compounds currently in clinical use or trials without confirmed DILI liability (Supplementary Table S1).
In blind validation, the model achieved 88% sensitivity, correctly identifying 29 of the 33 DILI+ compounds, and 100% specificity, with all 14 DILI− compounds correctly classified as safe (Fig. 1C).
Identifying withdrawn and idiosyncratic DILIs previously overlooked in animal and clinical studies
These results represent a substantial improvement over our initial proof-of-concept model trained on TG-GATES microarray data, which achieved 62% sensitivity and 92% specificity. Leveraging our DILImap library, ToxPredictor substantially improved both sensitivity of 88% and specificity of 100% on the same validation set (Fig. 2A). In cross-validation of the entire library, the model identified 110 out of 144 DILIs—surpassing the previous 62 out of 144 with TG-GATES—and misclassified only 8 out of 66 non-DILIs (Fig. 2B, Suppl. Figure S7). This enhancement is attributed to DILImap’s larger dataset with broader mechanistic coverage and the higher resolution of RNA-seq over microarrays, enabling better gene detection and a wider quantitative range for expression level changes compared to microarrays46. Notably, post-market withdrawals, missed in both pre-clinical models and clinical trials, were most confidently flagged by our model as high DILI risk.
A Compounds are grouped into four categories based on DILI potential: No DILI (n = 14), likely DILI (n = 15), known DILI (n = 4), and withdrawn due to DILI (n = 14). Each data point represents the safety margin of an individual compound. Boxplots show the median (center line), interquartile range (box), and data within 1.5 × interquartile range (whiskers). The safety margin—calculated as the ratio of the first toxic dose to the therapeutic maximum plasma concentration (Cmax)—effectively separates DILI-positive from non-DILI compounds in blind validation using a safety margin threshold of 80 (dashed line). The model demonstrated 88% sensitivity and 100% specificity in identifying DILI-positive and non-DILI compounds, respectively. B Cross-validation of the entire DILImap library confirms the model’s robust performance across the entire dataset (No DILI: n = 52; likely DILI: n = 73; known DILI: n = 25; withdrawn: n = 13). These results highlight the ability of transcriptomics-based modeling to improve DILI detection beyond traditional methods, offering enhanced sensitivity for withdrawn and known high-risk compounds while maintaining high specificity for non-DILI controls.
Our DILI safety margin and classification is derived from three parameters: Cmax (baseline concentration), cell viability assays (indicating cell death), and transcriptomics (based on differential pathways). To assess each parameter’s contribution, we assessed their ability to classify DILI cases independently. Out of 144 DILIs, 29 were detected solely based on plasma Cmax (>25 μM) and 42 through the LDH cytotoxicity assay (safety margin <80), both at ≥90% specificity. Combining our transcriptomics-based model with Cmax and LDH data was most effective, identifying 110 out of 142 DILIs (safety margin <80), underscoring the added value of toxicogenomics in DILI detection beyond mere cell death (Supplementary Fig. S8). ToxPredictor achieved a ROCAUC of 0.82 in cross-validation, compared to 0.66 for viability alone. In blind validation, it achieved a ROCAUC of 0.96, compared to 0.65 for using viability alone (Supplementary Fig. S9).
Mechanistic dissection of NSAIDS with shared targets but different DILI profiles
Our model highlights distinct DILI profiles among closely related COX-2 inhibitor non-steroidal anti-inflammatory drugs (NSAIDs) and imparts unique mechanistic insights linking predictions to mechanisms such as hepatocellular injury, oxidative stress, and mitochondrial dysfunction. For instance, Valdecoxib, used for cancer pain, shows no DILI risk (Fig. 3A), while Sulindac, an arthritis treatment with rare but established idiosyncratic DILI cases, and Lumiracoxib, withdrawn due to severe liver failures, are flagged as DILI risks with safety margins below the classification threshold of 80 (Fig. 3B).
A Valdecoxib, deemed safe, is predicted no DILI risk across all dose levels, with a margin of safety (MOS) greater than 300. B Sulindac, associated with idiosyncratic DILI, and Lumiracoxib, withdrawn from the market due to severe liver failure, are both flagged as high-risk by the model with MOS values of 31 and 1, respectively. C The predictions are driven by pathways implicated in DILI. Differential pathway signatures highlight key mechanisms underlying DILI risk, include oxidative stress, drug metabolism, receptor-mediated signaling, fat and cholesterol synthesis, and bile acid regulation. Pathway enrichment was performed using Enrichr (gseapy), based on a Fisher’s exact test with Benjamini–Hochberg correction. Color intensity reflects –log10 adjusted p-values. D Dose-resolved DILI risk probabilities illustrate how increasing dosages impact the likelihood of liver injury. Dots indicate mean predictions and blue shaded confidence intervals represent the standard deviations across ensemble models. These predictions align closely with clinical outcomes, demonstrating its utility in DILI risk stratification, pathway-based mechanistic understanding, and guiding safer drug development.
Crucially, it highlights the pathways implicated in DILI, encompassing direct contributors like oxidative stress leading to cell injury, as well as indirect factors such as disturbances in fatty acid metabolism, which can be particularly relevant to explain idiosyncratic effects (Suppl. Table S2). Sulindac, for example, is linked to disruptions in fatty acid synthesis and cholesterol biosynthesis, aligning with recent studies connecting it to hepatic steatosis47. By pinpointing these pathways, the model provides mechanistic insights into idiosyncratic DILI, offering an understanding previously thought unpredictable (Fig. 3C).
The model assesses DILI risks in a dose-resolved manner, revealing how dosage impacts liver injury likelihood. A key demonstration is its accurate prediction of Sulindac’s DILI risk, despite it being believed to be unpredictable in a dose-resolved manner48. These results highlight the model’s ability to deliver actionable predictions and enable targeted optimization of drug safety profiles by focusing on critical pathways (Fig. 3D).
Known and novel genes and pathways associated with DILI risk
DILI arises from disruptions in diverse pathways. Our model highlights pathways with high predictive value (AUC ≈ 0.8) strongly associated with DILI risk, including amino acid metabolism (toxic metabolite buildup causing oxidative stress and liver injury), fatty acid biosynthesis (disruptions leading to lipid accumulation and hepatocyte damage), tryptophan metabolism (toxic intermediates driving oxidative stress and inflammation) and ferroptosis (iron-dependent oxidative stress leading to lipid peroxide accumulation)49,50,51,52,53. Additionally, pathway activations highly correlated with predicted DILI risk include nuclear receptor signaling (e.g., PXR/CAR/FXR), one-carbon metabolism, and bile acid regulation—highlighting transcriptional reprogramming and metabolic stress as key contributors to hepatotoxicity54,55,56 (Fig. 4A). When compounds are ranked by predicted DILI probabilities, a clear gradient of pathway activation emerges, revealing distinct enrichment patterns for these biological processes. This correlation reinforces the direct mechanistic relevance and interpretability of the model’s predictions and highlights these pathways as potential drivers of DILI (Fig. 4B).
A Pathways are ranked by their ability to distinguish DILI vs. non-DILI compounds, measured by AUC. To quantify the strength of association between pathway dysregulation and predicted DILI risk, we show the Spearman correlation between pathway activation scores and DILI probability across compounds. This highlights transcriptional and metabolic stress pathways—such as nuclear receptor signaling, one-carbon metabolism, and bile acid regulation—as key contributors to DILI risk. B Compounds are ordered by their DILI risk probability, showing a correlation between DILI risk and the activation of pathways associated with liver injury. Pathway enrichment was performed using Enrichr (gseapy), based on a Fisher’s exact test with Benjamini–Hochberg correction. The most affected pathways include those critical to drug metabolism, oxidative stress and lipid homeostasis. Key pathways include Cytochrome P450 Oxidation, NRF2-ARE regulation for oxidative stress response, and Fatty Acid Beta-Oxidation, highlighting the interplay between detoxification, mitochondrial function, and energy metabolism. Additionally, nuclear receptor pathways such as Pregnane X Receptor (PXR) and Sterol Regulatory Element-Binding Protein (SREBP) signaling reveal disruptions in lipid and bile acid regulation. C Differential gene signatures were computed with DESeq2 using the Wald test, with Benjamini–Hochberg correction. Values are shown as –log10 adjusted p-values multiplied by the sign of the log-fold change. Top 100 genes most frequently upregulated in DILI cases include those related to drug metabolism (e.g., CYP1A2, CYP51A1, UGT1A8, AKR1C1, AKR1C2, AKR1C3), drug transport (e.g., SLC3A2), stress response (e.g., TXNRD1, SRXN1, GLRX, GLRX3, GCLM, HSP90AA1, HSP90AB1, HSPE1), and lipid metabolism (e.g., PLIN2, INSIG1, SREBF1, SCD, LPIN2, FADS2). Less commonly studied but significant genes include those involved in inflammation (e.g., GDF15, TNFRSF12A, S100A10, LITAF), autophagosome formation (e.g., MAP1LC3B, SQSTM1), and mitochondrial function (e.g., VDAC2, RAN, CHCHD10). D Top 100 genes most frequently downregulated in DILI cases include those involved in drug metabolism and detoxification (e.g., CYP4A11, CYP2E1, AKR1C4, GSTA1, GSTA2, UGT2B10, UGT2B15), drug transport (e.g., SLC10A1, SLC22A1, SLC22A7, SLC38A3, SLC27A5), and protein processing (e.g., ALB, AHSG, TTR, APOA1, APOE). Genes associated with amino acid metabolism and mitochondrial function (e.g., MAT1A, ARG1, CPS1, HMGCS2, ACAA1, ACAA2, ECHS1) also show significant downregulation. Key oxidative stress regulators and redox enzyme (e.g., CAT, ABAT, GNMT, and DHTKD1) are also reduced. Pathways related to lipid metabolism (e.g., CIDEB, ACSM2A, ACSM2B, ACSM5) and coagulation or inflammatory responses (e.g., SERPINF2, SERPINC1, SERPINA6, SERPINA10, FGB, FGG, FGA) are prominently affected. Notable genes with less established links to DILI but showing significant downregulation include ITIH4, PGLYRP2, BHMT, and GUCA2B, which could represent novel mechanisms or pathways contributing to the progression of liver injury. The changes in gene expression reflect early cellular alterations that may lead to DILI and highlight the complexity of DILI mechanisms, which encompass multiple aspects of liver function.
To identify genes most significant for DILI, we determined the frequency at which each gene was differentially up- or downregulated across DILI drugs in our library, using an adjusted p-value threshold of 0.05. This analysis focused on the concentrations at which toxic effects were first predicted, aiming to uncover early upstream regulators potentially driving DILI. Most frequently up-regulated genes were associated with drug metabolism, transport, stress response, and lipid metabolism. Novel genes linked to inflammation, autophagy, and mitochondrial dysfunction were also implicated (Fig. 4C). Frequently down-regulated genes include those critical for liver functions such as drug metabolism, transport, lipid metabolism, amino acid metabolism, mitochondrial function, coagulation and inflammatory responses. Altered expression of these genes may serve as early indicators of liver injury and reflect DILI’s multifaceted mechanisms (Fig. 4D).
Establishing that the DILI pathways and genes identified by our model are specific to liver toxicity rather than general toxicity is inherently challenging. However, the model’s precision is evident in its accurate classification of non-DILI compounds with known toxicities in other systems, such as Valdecoxib (cardiovascular toxicity), Bupropion (neurologic and cardiovascular toxicity), and Warfarin (hematologic toxicity)57,58,59, indicating the model’s ability to distinguish liver-specific toxicity from other forms of organ damage.
ToxPredictor accurately flags DILI risks in recent clinical failures and provides dose recommendations
Bruton tyrosine kinase (BTK) inhibitors, despite their promise in oncology and autoimmune diseases, have faced clinical holds due to liver injury. Recent examples include Evobrutinib, BMS-986142, and Orelabrutinib, all of which were withdrawn or put on hold in phase III in 2023 due to DILI cases.
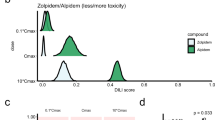
We validated ToxPredictor on four clinical failures: Evobrutinib, BMS-986142, Orelabrutinib, and TAK-875 (type 2 diabetes drug), along with two investigational BTK inhibitors (Rilzabrutinib, Remibrutinib) and two FDA-approved JAK inhibitors (Tofacitinib, Upadacitinib) as negative controls. DILI risk probabilities were assessed at four concentrations and margins of safety (MOS) estimated to classify compounds as high (MOS ≤ 2.5), mid-high (MOS ≤ 12.5), medium (MOS ≤ 80), or low risk (MOS > 80). All clinical failures were flagged as high or medium-high risk with low MOS values, particularly TAK-875, Evobrutinib, and BMS-986142, consistent with their phase III withdrawals. The investigational drugs were classified as medium risk (MOS = 14) and low risk (MOS = 101), which have not yet been linked to DILI in clinical studies yet, while the DILI-negative JAK inhibitors were classified as low risk. These results align closely with their clinical safety profiles (Fig. 5A).
A. ToxPredictor provides outputs including the first DILI concentration, safety margin, and corresponding risk classification. Clinical validation confirms that drugs labeled as high risk by the model, such as TAK-875, Evobrutinib, and BMS-986142, were recently withdrawn and halted in Phase III trials due to DILI. Conversely, low-risk drugs such as Tofacitinib and Upadacitinib, as classified by the model, exhibit no DILI association. B Model-derived DILI risk likelihood curves for representative compounds are plotted against hypothetical Cmax values. These curves delineate dose regimes associated with low, medium, mid-high and high DILI risk, enabling dose recommendations within the low-risk range. Risk classification is determined based on the actual efficacious Cmax of each compound: Evobrutinib is classified as high risk, Rilzabrutinib as medium risk (recommended dose <100 mg q.d.), and Remibrutinib as low risk (recommended dose <155 mg q.d.). This demonstrates the model’s utility in guiding dose selection to minimize DILI risk.
ToxPredictor provides dose-dependent DILI risk curves derived from empirical DILI likelihoods across various hypothetical Cmax values, enabling safe dosing recommendations (see Methods). For instance, Rilzabrutinib is categorized as low risk at doses below 100 mg q.d., which is lower than its efficacious dose of 400 mg. In contrast, Remibrutinib’s efficacious dose of 100 mg falls within the recommended low-risk range of <155 mg q.d. These findings highlight ToxPredictor’s value in informing safe dosing strategies, making it a valuable tool for de-risking new drug candidates (Fig. 5B).
Mapping toxicogenomics in the competitive landscape of existing pre-clinical DILI models
We benchmark our model along two key axes: predictive performance and scalability. Predictive performance, measured by balanced accuracy, reflects the model’s ability to distinguish DILI-positive from DILI-negative compounds. Scalability captures both technical throughput and biological breadth—the capacity to generalize across diverse chemistries and mechanisms, including previously uncharacterized ones (Fig. 6A).
A Balanced accuracy versus scale (number of DILI + /− compounds) across published preclinical models. ToxPredictor achieves the highest performance among both in vitro and in silico methods, outperforming high-content imaging, cytotoxicity, bioactivation-based and structure-based models, demonstrating robust performance even when scaled to hundreds of compounds. B Head-to-head comparison on overlapping compounds shows consistent performance gains over other studies. Models with >80% specificity (marked with *) support credible preclinical de-risking. Results highlight transcriptomics-based toxicogenomics as a leading strategy for mechanistically informed, scalable DILI prediction.
Our model outperforms a wide range of pre-clinical DILI models, including mechanistic assays16,17,18,19,20,21, cytotoxicity markers22,23,24,25,26,27, physicochemical properties28, bioactivation29 and BSEP30 approaches. In a head-to-head comparison across matched compound sets, it identified 46 out of 66 DILI cases versus the 27 out of 66 identified by Xu et al.17 HCI assay (49/66 vs 27/66); it shows superior performance over Garside et al.20 HCI assay (37/46 vs 29/46 DILIs), Vorrink et al.26 cytotoxicity assay using CD spheroids (37/43 vs 30/43 DILIs), Sakatis et al.29 bioactivation endpoint GSH adduct (47/65 vs 25/65) as well as their combined assay integrating covalent binding and dose (47/65 vs 32/65). When compared to Kohonen et al.’s transcriptomics-based cytotoxicity model60, our approach showed improved sensitivity (26/36 vs. 16/36 DILIs). These comparisons, all at 100% specificity evaluated on the same compounds, underscore the added value of our systems-level, mechanism-agnostic readout (Fig. 6B; Supplementary Table S3).
Structure-based in silico models such as TxGemma31, DILIGeNN32 and DILIPredictor33 underperform in vitro-based approaches in our benchmark. To assess real-world generalizability, we evaluated them on 314 independent compounds (45 DILI + , 269 DILI − ) primarily annotated via LiverTox (scores A/B as DILI + , E as DILI − ); TxGemma was also tested on an expanded set (143 DILI + , 536 DILI − ). All showed limited specificity: DILIGeNN (84% sensitivity, 28% specificity), DILIPredictor (80% sensitivity, 29% specificity), and TxGemma-27B (57% sensitivity, 37% specificity). These findings are slightly below the balanced accuracy of 0.59 reported by Seal et al. (2024)33 for DILIPredictor. On a benchmark subset of unseen compounds overlapping with DILImap (n = 97), TxGemma reached 63% sensitivity (39/62) and 57% specificity (20/35), while our model achieved 76% sensitivity (47/62) and 86% specificity (30/35). Similarly, DILIGeNN showed perfect sensitivity (5/5) at moderate specificity (2/3), while our model reached 100% on both (5/5 and 3/3). DILIPredictor reached complete sensitivity (23/23) but at the expense of poor specificity (1/7), while ToxPredictor maintained high sensitivity (20/23) at markedly higher specificity (5/7). Low specificity is a key limitation of structure-based models, which lack biological context and tend to over-call toxicity. This results in false positives for commonly prescribed drugs with no risk of hepatotoxicity, such as biotin (flagged DILI+ by DILIPredictor), vitamin D (flagged DILI+ by DILIGeNN), and pemetrexed (flagged DILI+ by all three models). Moreover, they provide only binary outputs, without dose or mechanistic insight. In contrast, transcriptomics enables dose-resolved predictions, mechanistic interpretability, and safety margin estimation—critical for evaluating toxic liabilities and guiding follow-up experiments (Supplementary Fig. S10; Supplementary Data S4).
3D liver systems offer important physiological context. High-content imaging in 3D models, such as those by Walker et al.34 and Ewart et al.35, achieves similar performance on small, curated panels (Walker: 23/27 vs. 23/27; Ewart: 11/14 vs. 13/14). However, their limited scalability constrains their utility to a broader range of DILI mechanisms. They may perform well on narrow, curated panels, but struggle with unknown mechanisms or mechanisms not captured by the low-dimensional endpoint, as shown in the following comparison. To explore the unique capabilities of 2D transcriptomics vs 3D cytotoxicity assays, we conducted direct compound-level comparisons with larger 3D screening studies: Vorrink et al.26 and Fäs et al.36 In Vorrink et al. 3D cytotoxicity uniquely detected 3 compounds (Fialuridine, Methotrexate, Trazodone), all linked to cytotoxic effects that result in acute cell death. Conversely, our model uniquely flagged 10 compounds—including fluconazole, phenytoin, and zileuton—associated with immune activation, metabolic stress, or enzyme modulation, which are not readily captured by viability endpoints. A similar pattern emerged in the Fäs et al. study: 3D cytotoxicity exclusively identified 4 compounds (e.g., Haloperidol, Fialuridine) whose toxicities depend on structural or metabolic context. Our model uniquely identified 5 compounds (e.g., Cimetidine, Fluconazole, Ximelagatran) marked by subtle transcriptional responses rather than overt cell death. These comparisons highlight a key limitation of fixed single-endpoint models: while effective in narrow contexts, they struggle with broader chemical and mechanistic diversity. Our transcriptomic approach, by contrast, offers systems-level resolution that generalizes across DILI pathways—not only detecting known cytotoxic responses but also uncovering less immediate, non-lethal mechanisms often missed by traditional assays (Supplementary Data S4).
As a result of its unbiased modeling, our approach shows improved detection of idiosyncratic compounds—a class of toxicities that often escape detection in targeted or phenotypically narrow assays. These compounds, many of which are associated with extremely rare clinical incidence (<12 case reports), present a significant challenge for preclinical screening. Our model identified 29 out of 65 of those cases (44%) the highest detection rate among all evaluated models, while maintaining a specificity of 88% (Supplementary Fig. S11A).
Next, we analyzed how combining toxicogenomics with orthogonal assays further enhances detection. The three most effective combinations include pairing our model with Walker et al.’s 3D-based HCI assay34 to improve DILI detection from 23/27 to 26/27 cases, pairing with Persson et al.’s 2D-based HCI assay19 to improve DILI detection from 28/37 to 30/37 cases, and pairing with Sakatis et al. GSH depletion assay29 to increase detection from 47/65 to 53/65. Such strategic combinations could raise balanced accuracy to as high as 98% (Supplementary Fig. S11B).
Based on these insights, we propose a tiered de-risking funnel strategy that begins with straightforward endpoints, such as PK data (e.g., Cmax <25 μM) and cytotoxicity assays, to flag overt hepatotoxicity. For candidates showing no early toxicity signals, toxicogenomics provides the most comprehensive and unbiased assessment of DILI risk—capturing both known and novel mechanisms. For a select few advanced candidates, with sufficient resources, applying toxicogenomics in advanced 3D liver models may offer the most accurate prediction of in vivo responses. This strategy ensures a resource-efficient and mechanistically broad DILI risk assessment in drug development.
Discussion
Our toxicogenomics approach offers a comprehensive and unbiased perspective on cellular responses, providing rich information for a nuanced understanding of liver toxicity, including idiosyncratic reactions with unknown mechanisms. By shifting from single-target analyses to a systems-level viewpoint, we demonstrate that applying machine learning to toxicogenomics holds great promise as a significant enhancement over existing toxicology methods. It enables dose-specific predictions and safety margins, moving beyond binary DILI/No-DILI assessments. Its extensive mechanistic coverage presents a substantial advantage over current pre-clinical models. By creating a comprehensive toxicogenomics library specifically designed for DILI research, we achieved notable improvements in predicting DILI risks compared to state-of-the-art methods. The high sensitivity (88%) and specificity (100%) obtained in blind validation, along with the identification of DILI in withdrawn drugs and clinical failures overlooked in animal and clinical studies, underscore its practical utility in early drug development phases. We consider this an important step forward in predictive toxicology, positioning our approach as a significant advancement in the field. The scalability and adaptability of our method, a central component of a larger AI/ML platform, are designed to enhance the predictive power and efficiency of drug development pipelines.
While our approach represents a meaningful step forward, it is important to acknowledge the inherent limitations of our method. First, the multifactorial nature of DILI, involving genetic, environmental, and lifestyle factors, means even the most advanced models cannot capture the full spectrum of potential mechanisms. Our approach, though comprehensive, does not account for all interindividual variability or rare genetic predispositions contributing to DILI risk. Second, the predictive power of our model is constrained by the completeness of the DILImap library. Gaps in the database, particularly in relation to poorly documented idiosyncratic reactions, can limit the model’s accuracy. Third, our reliance on a 2D hepatocyte culture system may fail to replicate the complex interactions between hepatocytes and other cell types or tissues that can drive certain DILI mechanisms. Fourth, the exposure timepoint of 24 hours limit the detection of delayed or immune-mediated toxicities. Fifth, as with any preclinical model, the ultimate test of its utility lies in its ability to predict clinical outcomes, a domain where uncertainties and unpredicted variables can significantly impact performance.
Looking ahead, toxicogenomics holds tremendous promise for advancing our understanding and prediction of DILI. Integrating RNA-seq with advanced 3D culture systems—such as spheroids and liver-on-chip platforms—may enable longer drug exposures and the inclusion of non-parenchymal cells like Kupffer cells and hepatic stellate cells. These co-culture models are essential for capturing immune activation, inflammation, and fibrosis—hallmarks of idiosyncratic and chronic DILI that hepatocyte monocultures cannot recapitulate. As long 3D models rely on high-content imaging or cytotoxicity endpoints that capture specific phenotypic responses they remain limited in mechanistic scope and scalability. Their performance is often demonstrated on small, curated compound sets with known mechanisms, which may not translate to broader chemical space. In contrast, transcriptomics provides a scalable, unbiased readout capable of detecting diverse DILI mechanisms, including those not captured by existing assays. As RNA-seq becomes more feasible in physiologically relevant 3D systems, we anticipate a powerful synergy—combining the mechanistic breadth of transcriptomics with the physiological relevance of 3D models. Our work lays the foundation for such integration, demonstrating that transcriptomics alone can robustly capture DILI risk across a wide range of mechanisms.
Combining RNA-seq data with structural information of molecules could enable a deeper understanding of the interactions between drugs and cellular components, facilitating more accurate predictions of toxicity. Using early estimates as surrogate for plasma Cmax, such as target activity, could help derive safety margins earlier in the drug discovery process. The development of multivariate models that include DILI regulatory networks represents another exciting frontier. Such models can incorporate the complex interplay of genes, proteins, and metabolic pathways involved in DILI. Furthermore, exploring additional data types, such as chromatin accessibility, proteomics, or metabolomics, could yield further insights into DILI mechanisms.
As these advanced models and datasets become integrated into the early stages of drug development, we anticipate a decrease in liver-related adverse events, improved efficiency in drug development, significant cost savings, and, most importantly, enhanced patient safety. The ongoing evolution of toxicogenomics approaches, bolstered by advancements in computational machine learning methods and multi-omics technologies, marks an important step toward more predictive drug safety evaluation—one that has the potential to support the development of safer therapeutics and improved patient outcomes.
Methods
Experimental workflow
We employed a systematic, high-throughput approach using cryopreserved primary human hepatocytes (PHHs) to study transcriptional changes underlying drug-induced liver injury (DILI).
Human tissue sourcing and ethical compliance
Cryopreserved PHHs were obtained from LifeNet Health (Virginia Beach, VA, USA) under standard provider agreements. LifeNet Health procured tissues under informed donor consent and Institutional Review Board (IRB) approval in accordance with U.S. regulations. Cellarity did not create any new cell lines for this publication.
Cell culture overview
Cryopreserved PHHs were selected for high viability (>95%), long-term plateability (10–15 days), compatibility with 96-well formats, and Grade A quality. Donor 1917277-01, a 37-year-old Caucasian female (BMI 26), was used. Hepatocytes were cultured in collagen I-coated 96-well plates and maintained in a sandwich configuration (collagen base with a 100 μg/mL Matrigel overlay) to preserve hepatocyte function. Cells were matured for three days with daily media changes before compound treatment. After 24 h of treatment, cells were either assayed for viability (LDH/ATP) or lysed for RNA extraction and sequencing (Supplementary Fig. S1A).
Cell culture protocol
Human hepatocyte thawing media, human hepatocyte culture media, HHCM supplement, human hepatocyte plating media, HHPM supplement (LifeNet Health) and Pen/Strep (Gibco) were thawed and filtered before plating. PHHs were counted using a Luna-FL Cell Counter and Acridine Orange/Propidium Iodide Stain (Logo Bio). Cells were plated at a cell density of 0.5 million cells / mL in collagen I-coated 96-well plates (Gibco). Cells were left to attach in the incubator for 6 hours and then replaced with maintenance medium (LifeNet Health). The next day, cells were overlayed with a thin coat of Matrigel and left to incubate for an additional day with a daily media change to allow for full maturation of cells. On Day 3, cells were treated with compounds at multiple concentrations for 24 h and then were either taken down for either viability testing (LDH and ATP readouts) or lysed for RNA sequencing.
Experimental setup
The objective of this experiment was to screen 300 compounds to create a hepatotoxicity intervention library for building a predictive DILI model. PHHs were first screened for maximum tolerated dose (MTD) in six-point log curves (0.01 μM to 1 mM, Figure. S1B). MTD was defined as the highest concentration before observing >10% cell death in the LDH assay. RNA sequencing was conducted on cells treated with compound concentrations ranging from Cmax to MTD to evaluate the safety margin between therapeutic and toxic doses.
Compound dissolution and plate preparation
All compounds were purchased from MedChemExpress (MCE). Compound preparation was performed in-house. Compound dissolution was prepared manually fresh on the day of compound treatment to avoid freeze/thaw cycles. In a previous DMSO tolerance test, 0.5% of DMSO in compound was dictated as the ideal concentration. Compound dissolution with DMSO started at a highest concentration of 1000 µM. If compounds were not solubilized at the 200x stock, dissolution was attempted at 100x and 50x concentrations. The compounds were added to a barcoded plate and transferred to the Hamilton MicroLab Star liquid handler for titration (Fig. S1B). The plate layout for titration varied between toxicity screens and RNA sequencing runs (Fig. S1C). Automation was used to prepare compound dilution and compound treatment.
Lactate dehydrogenase (LDH) viability assay
As primary viability assay, we employed the non-radioactive cytotoxicity assay from Promega, which utilizes lactate dehydrogenase (LDH) as a marker for cell death. For reagent preparation, we thawed LDH buffer at 4 °C overnight and used it to reconstitute the LDH substrate bottles by adding 12 mL, which were then stored at –20 °C. We utilized designated untreated cells to establish the 100% lysate positive control, against which we normalized all subsequently treated cells for % viability calculations. To these designated cell wells, we added 10x Lysis buffer in a 1:10 volume ratio. We then mixed 50 μL of the collected sample with 50 μL of LDH substrate in a flat-bottom tissue culture plate. Plates were covered by and incubated at room temperature for 30 min. Following the incubation, we added 50 μL of thawed Stop solution and used a SpectraMax i3x plate reader with absorbance settings at 490 nm to read the plate.
CellTiter-Glo (CTG) viability assay
Another orthogonal viability assay that was utilized for toxicity screening was CellTiter-Glo (CTG) Luminescent Cell Viability Assay (Promega) which determined the ATP content within the wells. The CTG viability assay is a terminal endpoint due to the lysis of cells. To prepare the reagent, the CTG buffer and substrate were thawed at room temperature and 10 mL of the buffer was resuspended in the substrate and stored at –20 °C freezer. CTG aliquots were thawed the day of the takedown, and DPBS (Gibco) was added to CTG substrate at a 1:1 ratio. Following the removal of the contents of the culture plate, 100 uL of CTG and PBS mix was added across all the wells of the plate. Following the lysis, plates were covered with aluminum foil and set on an orbital shaker for 2 min at 400 rpm. Following the 2 min, the plates remained covered and were left at room temperature for 10 minutes. The plates were then read on the SpectraMax i3x using the luminescence and Standard Opaque settings.
Cell lysis for RNA extraction
RLT buffer (Qiagen) and 2-mercaptoethanol (Thermofisher Scientific) were prepared to create a RLT + 1% BME reagent. 140 uL of lysis buffer was added across all wells of the plate using a multichannel pipette. After ensuring the cells were lysed under a microscope, the 140 uL of lysate was transferred to an Eppendorf twin.tec 96-well PCR plate (Fisher Scientific) and placed into a −80 °C freezer.
Library preparation and sequencing (SMART-Seq)
Total RNA was prepared from cell lysates in 96-well plates using a QIAcube HT robotic workstation (Qiagen) in conjunction with RNeasy 96 QIAcube HT kits (Qiagen) according to the manufacturer’s recommended protocol. SMART-Seq DE3 libraries were prepared from total RNA according to the manufacturer’s protocol. Briefly, for each row of the plate, polyadenylated mRNA was selected using uniquely barcoded oligo(dT) primers. First strand cDNA was generated via reverse transcription; double-stranded cDNA was created via template switching with limited cycles of PCR amplification. Each row of samples was then pooled and subjected to transposon-based fragmentation using the Nextera XT DNA Library Preparation Kit (Illumina). Libraries were then PCR amplified using unique combinations of Illumina P5 and P7 barcodes and mixed in equimolar pools prior to sequencing. Sequencing was performed on an Illumina NovaSeq6000 using custom read lengths of 89 bp (Read1) and 26 bp (Read2).
Computational workflow
Cmax annotations
Cmax values were compiled from multiple resources, and the median of these values was used to derive a consensus total Cmax. The following resources contributed to computing the consensus Cmax: Drug information from the National Center for Advancing Translational Sciences (NCATS), Porceddu et al. (2012), Khetani et al. (2013), Persson et al. (2013), Aleo et al. (2014), Garside et al. (2014), Gustafsson et al. (2014), Chen et al. (2014), Shah et al. (2015), Camenisch et al. (2019), Dixit et al. (2019), Aleo et al. (2020), Williams et al. (2020), and Smit et al. (2020). For compounds where no studies provided reliable Cmax estimates or where estimates varied significantly across studies, additional manual annotations were performed by searching PubChem and relevant clinical studies.
DILI categorization
DILIrank and LiverTox serve as key resources for categorizing drugs as DILI positive or negative. DILIrank, developed by the FDA, classifies over 1000 drugs into four levels of DILI concern — most, less, no concern, and undetermined — based on historical liver injury data. LiverTox, created by the NIDDK, is an exhaustive online database with detailed information on these drugs, including their clinical manifestations, mechanisms of action and likelihood of causing DILI. It also offers a system to assess the likelihood of DILI, from well-known cases to idiosyncratic drugs without a characteristic signature to drugs considered safe. These categorizations serve as DILI endpoints for the development and refinement of our model. Compounds were systematically categorized based on DILIrank and LiverTox into the following categories:
Withdrawn DILI: Market withdrawals and clinical failures due to DILI (Most-DILI-Concern; e.g., Troglitazone).
-
Known DILI: Compounds with well-established DILI risk (Most-DILI-Concern in DILIrank or LiverTox score A; e.g., Isoniazid at therapeutic doses, Acetaminophen at overdose levels).
-
Likely DILI: Compounds with documented cases of DILI in specific contexts (Most-DILI-Concern or LiverTox score A/B; e.g., Progesterone).
-
Idiosyncratic DILI: Rare, unpredictable DILI without clear dose-response (LiverTox score C/D; <12 case reports).
-
Unlikely DILI: Discordantly labeled safe in LiverTox (score E) but Less-DILI-concern in DILIrank.
-
No DILI: Compounds with no clinical or preclinical documented evidence of DILI (No-DILI-Concern in DILIrank; LiverTox score E).
The categories Withdrawn DILI, Known DILI, and Likely DILI serve as positive controls and No DILI as negative control. Our models were trained to differentiate between positive and negative control compounds. The categories Unlikely DILI and Idiosyncratic DILI, due to their label ambiguity, were excluded from model training and reserved for downstream testing.
In the training data, DILI positives include the highest concentrations and those above 20x Cmax, while DILI negatives include the lowest concentrations and those below 80x Cmax. This approach ensures unambiguous labeling while minimizing bias from overdose signatures in negatives and inactive signatures in positives. Using these categorizations, our model was trained to distinguish between 177 positive and 93 negative control data points. An additional set of 70 compounds known for their idiosyncratic effects, each linked to fewer than 12 case reports, were exclusively used for testing. Idiosyncratic hepatotoxicity, characterized by its unpredictability and lack of dose-response or temporal patterns, often leads to drug withdrawals despite thorough clinical testing. An independent in-house experiment was conducted for blind validation, comprising 46 compounds: 32 DILI positives and 14 negatives. This included well-established DILI positives/negatives and a real-world set with 4 recent clinical failures.
Toxicogenomics resources and initial efforts
Human toxicogenomics benefits from resources such as TG-GATES, CMap and L1000. CMap and L1000 catalogue genomic responses in in vitro cell lines, primarily cancer lines, and can be used to study cytotoxicity, however, their utility for DILI is limited as our initial efforts showed poor predictive performance likely due to the lack of physiologically relevant cell types and relevant range of concentrations. In contrast, TG-GATES provides a rich microarray database tailored for DILI research, including data from primary human hepatocytes (PHH) exposed to 158 compounds. Our initial proof-of-concept, utilizing TG-GATES data, yielded meaningful predictions. In a rigorously designed in-house pilot experiment involving 46 compounds for blind validation, our model demonstrated a 62% sensitivity in accurately identifying DILI positives and an 92% specificity for negative control compounds, demonstrating the potential of toxicogenomics in identifying DILI risks during drug development.
Quality control
We have created DILImap, our RNA-seq library encompassing 300 compounds, to improve the predictive power and mechanistic coverage of our ML model. We applied stringent quality control metrics and filtering criteria to retain only high-quality RNA-seq samples:
-
1.
Total RNA counts > mean −2.5 standard deviations (~700,000 counts).
-
2.
Mitochondrial RNA fraction <9%.
-
3.
Correlation between replicates >0.99.
In addition to these technical filters, we performed a hepatocyte fidelity check: each sample was assessed for expression of liver-specific marker genes and the preservation of a hepatocyte-like transcriptional profile.
Differential signatures
Samples passing all QC filters were used to compute compound-specific differential expression signatures. We used DESeq2, a standard RNA-seq analysis tool, which models read counts using a negative binomial distribution. DESeq2 adjusts for library size differences and estimates gene-specific dispersion, enabling accurate detection of differentially expressed genes (DEGs).
For each compound-dose combination, we compared treated samples against matched DMSO controls from the same plate. This plate-specific normalization controls for potential batch effects and ensures that the derived signatures reflect treatment-specific transcriptional responses.
Pathway signatures
Pathway enrichment analysis is a statistical approach used to identify whether specific biological pathways are significantly enriched with differentially expressed genes (DEGs) in RNA-seq data. The significance of this enrichment is assessed using p-values, which indicate the likelihood that the observed enrichment occurred by chance. In our analysis, we utilized the widely adopted hypergeometric test to compute p-values. This test calculates the probability of observing the number of DEGs in a given pathway, considering the total number of genes in the pathway and the overall number of DEGs in the dataset. To derive pathway signatures from p-values, we applied a -log10 transformation of the adjusted p-value, or False Discovery Rate (FDR). The FDR accounts for multiple testing, controlling the proportion of false positives in the analysis. The resulting -log10(FDR) scores serve as the input to our model, providing a robust representation of pathway enrichment.
ToxPredictor classifier for cross-validation
To develop ToxPredictor, we evaluated a range of machine learning models, including Logistic Regression, Support Vector Classifier (SVC), Random Forest, Gradient Boosting Classifier, Hist Gradient Boosting Classifier, XGBoost Classifier, LGBM Classifier and Multi-Layer Perceptron (MLP). Each model was evaluated using a 5-fold stratified compound-level cross-validation strategy, ensuring that all samples from a single compound were held out together in each fold to prevent data leakage. This compound-level splitting reflects a realistic generalization scenario for novel compound prediction.
We optimized hyperparameters using grid search, assessing performance across the following metrics:
-
Area Under the ROC Curve (AUC): Measures the overall ability to distinguish DILI from No-DILI compounds.
-
Precision: Fraction of predicted DILI compounds that are truly DILI-positive, relevant for minimizing false positives.
-
Recall: Fraction of true DILI compounds that are correctly identified, reflecting the model’s ability to avoid false negatives.
-
Inter-fold correlation: Quantifies agreement in predicted probabilities between models trained on different splits.
-
Monotonic dose response: Assesses whether increasing doses of a compound correspond to increasing predicted DILI risk.
Model selection strategy
-
Generalizability: prioritize high validation AUC with low train–validation gap to avoid overfitting → Best models had AUC ≥ 0.74 (RF, HistGB, LGBM, XGB); RF showed the lowest gap (~0.09 vs. 0.16–0.20 for GBMs).
-
Stability: require high inter-fold correlation for consistent predictions across folds → RF achieved 0.98–1.0, higher than GBMs (0.8–0.9).
-
Clinical utility: balance precision (reduce false positives) and recall (reduce false negatives) → RF performance was comparable to other top models.
-
Interpretability: prefer simpler, easier-to-interpret models at similar performance → RF is more interpretable than boosting methods.
Final choice: Random Forest was selected as the most balanced and robust model, combining strong validation AUC, minimal overfitting, high stability, and interpretability. Although XGBoost and LightGBM reached similar AUCs, they showed greater overfitting (train-validation AUC gap) and lower stability (inter-fold correlation). These characteristics along with its ability to handle non-linear relationships, its robustness to noise, and its suitability for datasets of moderate size, make Random Forest the most robust choice for generalization to unseen compounds.
The optimal hyperparameters were:
n_estimators = 100, max_depth = 2, min_samples_split = 2, min_samples_leaf = 1
This combination of performance and robustness established Random Forest as the base model for ToxPredictor.
ToxPredictor ensemble modeling for blind validation
To ensure predictive robustness, we employed a 30-model Random Forest ensemble using bootstrap aggregation (bagging). Each model was trained on a unique bootstrap sample from the training data, ensuring diversity in training instances and mitigating overfitting. We selected 30 folds to strike a balance between accuracy and ensemble stability. In our benchmarks, ensembles with 30 independent learners consistently achieved smooth, reproducible predictions and stable dose-responses. All fold models used the hyperparameters optimized during cross-validation. At prediction time on unseen compounds, model outputs are averaged with equal weighting, producing a consensus decision that mitigated individual model variance, ensuring robust performance by combining the strengths of each base learner, leading to stable and reliable classification of DILI vs. No-DILI. In blind validation, where predictions were made on compounds not seen during training or model selection, the ensemble demonstrated high AUC, dose consistency, and biological plausibility, confirming its utility for real-world DILI risk assessment.
DILI risk probabilities
ToxPredictor outputs a predicted DILI probability, defined as the average prediction across the 30 Random Forest models in the ensemble. This probability reflects the model’s confidence that a compound induces liver injury, based on its pathway perturbation profile. To enable binary classification for downstream evaluation and decision-making, we defined a DILI risk threshold of 0.7. Compound-dose pairs with predicted probabilities ≥0.7 are considered DILI-positive, while those <0.7 are considered DILI-negative. This threshold was empirically selected to maximize the trade-off between sensitivity and specificity in our benchmark datasets, ensuring robust identification of true DILI compounds while minimizing false positives.
DILI safety margins
ToxPredictor computes a Margin of Safety (MOS) for each compound, defined as the ratio between the first DILI dose and its maximum plasma concentration (Cmax) at therapeutic levels. The first DILI dose is the lowest dose at which a compound crosses the 0.7 DILI probability threshold, indicating transcriptional evidence of hepatotoxicity. We use total Cmax values rather than free (unbound) Cmax, as both approaches yield comparable predictive performance, but total Cmax offers broader availability across clinical data sources and aligns with common reporting in literature.
For validation and benchmarking, to classify a compound as DILI, we selected a Margin of Safety (MOS) threshold of 80, determined empirically from the training set. This value reflects an optimized trade-off between sensitivity and specificity—minimizing false positives while reaching a performance plateau that captures the majority of true DILI liabilities. Importantly, this threshold is consistent with values commonly used in the literature, where MOS cutoffs typically range from 10 to 100 depending on context, placing our choice within accepted toxicological standards.
Prioritization of predictive features/pathways
To identify and prioritize features and pathways relevant to DILI risk prediction, feature importance within the Random Forest ensemble was assessed using statistical significance, impurity reduction, permutation importance, and direct discriminative power, ensuring a comprehensive and biologically meaningful selection of key predictors of DILI risk:
-
Statistical Significance: Features were prioritized if they demonstrated at least one significant differential occurrence across compounds, with a p-value threshold of 0.001.
-
Mean Decrease in Impurity (MDI): A standard metric for tree-based models that quantifies the reduction in impurity (e.g., Gini impurity) achieved by splits on specific features. The MDI values were averaged across all trees in the ensemble, providing a robust measure of a feature’s overall predictive contribution. Features with higher MDI scores were deemed more important for the model’s decision-making.
-
Permutation Importance: The values of each feature were shuffled individually, and the resulting change in model performance, specifically the Area Under the Curve (AUC), was measured. A greater reduction in AUC indicated a higher importance of the feature, offering additional insight into the features that most strongly influence the model’s predictions.
-
Discriminative Power (AUC): Each feature’s ability to distinguish between DILI and no-DILI categories was evaluated directly by calculating its individual AUC score, ensuring that features with strong discriminative capabilities were prioritized.
-
Spearman correlation: Correlation of pathway signatures with predicted DILI risk.
Empirical DILI likelihoods for dose recommendations
Empirical cumulative DILI likelihoods are calculated to inform dose recommendations that minimize the risk of DILI. For each compound, safety margins are computed at varying hypothetical Cmax values as the ratio of the model-predicted first DILI dose to the corresponding Cmax. For each safety margin, two cumulative percentages are derived: (1) the percentage of DILI compounds with safety margins above the given value, which increases monotonically from 0 to 1 as Cmax increases, and (2) the percentage of non-DILI compounds with safety margins below the same value, which decreases monotonically from 1 to 0. The empirical cumulative DILI likelihood at each safety margin is calculated as the difference between these two percentages, effectively representing the relative enrichment of DILI compounds compared to non-DILI compounds at or above a given margin. This approach captures the overall relationship between safety margins and DILI likelihood across the dataset, rather than focusing on isolated points, thereby enabling robust differentiation between high-risk and low-risk compounds. The resulting cumulative risk profiles provide a quantitative framework to guide dose selection.
Benchmarking against structure-based in-silico methods
We benchmarked ToxPredictor against three state-of-the-art structure-based models: DILIGeNN, a graph neural network trained on molecular graphs; DILIPredictor, a random forest ensemble model that integrates chemical structure, physicochemical properties, pharmacokinetic parameters and predicted proxy-DILI data; and TxGemma, a generalist large language model (LLM) fine-tuned on biomedical tasks from the Therapeutic Data Commons (TDC). The evaluation aimed to assess the predictive performance of each approach on overlapping and unseen compounds using balanced accuracy, sensitivity, and specificity.
DILIPredictor: Random Forest ensemble integrating proxy-DILI labels and chemical structure
A Random Forest model trained on nine proxy-DILI labels (e.g., mitochondrial toxicity, BSEP inhibition) in conjunction with chemical structural features derived from SMILES strings of 1111 DILI compounds. We reproduced the model from its public repository, converting the Poetry environment to Conda (via poetry2conda) and using scikit-learn v1.2.0 to match the pretrained model version. As compound names were not provided, we standardized SMILES and generated InChIKeys (non-isotopic, non-stereochemical layer) to identify 483 unseen compounds; 6 failed at inference, 6 more were deduplicated, resulting in 471 (98 DILI + , 373 DILI − ). This approach may still introduce limited data leakage if SMILES differ from those used during training. For benchmarking against other in-silico chemistry models, we focused on 314 compounds identified as unseen by both DILIGeNN and DILIPredictor, to provide a more conservative evaluation set that helps reduce the risk of data leakage; particularly important for DILIPredictor, which lacked compound identifiers and used a final model re-trained on the test set. For benchmarking against ToxPredicor, we took the overlap of the 471 compounds with DILImap yielding 30 compounds (23 DILI + , 7 DILI − ).
DILIGeNN: Graph Neural Network on Molecular Graphs
A graph neural network (GNN) trained on molecular graphs derived from SMILES strings of 1167 DILI compounds. We benchmarked the best-performing GraphSAGE models obtained after sequential warm starts, and used the recommended custom molecular graph representations of SMILES for each compound. To identify unseen compounds, we cross-referenced compound names, SMILES and InChIKeys (non-isotopic, non-stereochemical layer), and identified 349 unseen compounds; 18 failed inference, yielding a final evaluation set of 331 compounds (47 DILI + , 284 DILI-). Of these, 8 (5 DILI + , 3 DILI − ) overlapped with DILImap and were used for benchmarking. Final predictions were based on the mean probability and majority vote across four GraphSAGE models. In addition, we benchmarked DILIGeNN against DILIPredictor on 314 shared unseen compounds (45 DILI + , 269 DILI-) to evaluate real-world performance of in silico models across a broader evaluation set.
TxGemma: Generalist Language Models for Biomedical Tasks
A suite of generalist large language models (LLMs), fine-tuned from Gemma-2 (2B, 9B, 27B parameters) on biomedical tasks from the Therapeutic Data Commons (TDC). For DILI classification, models were trained on 475 compounds (325 train, 54 validation, 96 test). To avoid data leakage, we used SMILES and InChIKeys to select 715 unseen compounds (178 DILI + , 537 DILI-); however, note that data leakage could not be fully ruled out. Of these, 97 compounds (69 DILI + , 35 DILI-) overlapped with DILImap and were used for benchmarking. TxGemma models were deployed on Google Cloud Vertex AI using the official cookbook (https://github.com/google-gemini/gemma-cookbook/tree/main/TxGemma) with recommended prompts. The 27B achieved the best performance and was used for comparison with ToxPredictor; AUROC was not computed due to the model only providing DILI label outputs rather than prediction probability outputs. Performance improvements may be possible with additional fine-tuning and prompt engineering.
Data analysis software used in this study
The study used NumPy, pandas, and SciPy for computation, AnnData for single-cell data handling, scikit-learn = 1.4.0 for machine learning, and matplotlib/seaborn for visualization. Dask enabled scalable computing and quilt3 managed dataset access. DESeq2 supported differential expression and gseapy pathway enrichment. XGBoost and LightGBM were used for model comparison to select the optimal base model for ToxPredictor. Benchmarking against in-silico methods employed PyTorch Geometric for graph modeling, Captum for interpretability, and RDKit for cheminformatics.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All datasets and trained models are accessible via an S3 bucket for seamless integration with Jupyter notebooks, with detailed descriptions and access instructions provided at https://www.dilimap.org. In addition, the data have been deposited in GEO under a Creative Commons license (GSE308567). Processed training data (pathway-level signatures used as model input) and both raw and processed validation data are provided to enable full reproducibility of all model training and validation steps. The raw training data (gene expression count matrices) contain proprietary information and are not publicly available. Academic researchers may request access for internal, non-commercial use via DILImap@cellarity.com, with requests reviewed within 4–8 weeks. Approved data are available for 2 weeks and must be deleted within 6 months. Commercial access requires a data-sharing agreement. The data from the Open TG-GATEs database (http://dbarchive.biosciencedbc.jp/en/open-tggates/download.html)15 were used in an initial proof-of-concept to establish a toxicogenomics baseline performance, whereas all results reported in this manuscript are based on the internally generated data described above. The use of all datasets in this study complies with the terms and conditions of their respective repositories and data providers.
Code availability
All code, reproducibility notebooks, and results are available at https://www.dilimap.org, which serves as the central access point for this work. The full Python implementation is hosted at https://www.github.com/Cellarity/DILImap, with reproducibility notebooks and results at https://www.github.com/Cellarity/DILImap_reproducibility. Both repositories are also archived at https://zenodo.org/records/1729052061.
References
A New Standard. http://tools.thermofisher.com/content/sfs/brochures/D01834~.pdf.
Olson, H. et al. Concordance of the toxicity of pharmaceuticals in humans and in animals. Regul. Toxicol. Pharmacol. 32, 56–67 (2000).
Onakpoya, I. J., Heneghan, C. J. & Aronson, J. K. Post-marketing withdrawal of 462 medicinal products because of adverse drug reactions: a systematic review of the world literature. BMC Med. 14, 10 (2016).
Funk, C. & Roth, A. Current limitations and future opportunities for prediction of DILI from in vitro. Arch. Toxicol. 91, 131–142 (2017).
Robles-Díaz, M., Medina-Caliz, I., Stephens, C., Andrade, R. J. & Lucena, M. I. Biomarkers in DILI: One more step forward. Front. Pharmacol. 7, 267 (2016).
Chalasani, N. & Björnsson, E. Risk factors for idiosyncratic drug-induced liver injury. Gastroenterology 138, 2246–2259 (2010).
Mosedale, M. & Watkins, P. B. Drug-induced liver injury: Advances in mechanistic understanding that will inform risk management. Clin. Pharmacol. Ther. 101, 469–480 (2017).
Allison, R. et al. Drug induced liver injury - a 2023 update. J. Toxicol. Environ. Health B Crit. Rev. 26, 442–467 (2023).
Weaver, R. J. et al. Managing the challenge of drug-induced liver injury: a roadmap for the development and deployment of preclinical predictive models. Nat. Rev. Drug Discov. 19, 131–148 (2020).
Chen, M. et al. Quantitative structure-activity relationship models for predicting drug-induced liver injury based on FDA-approved drug labeling annotation and using a large collection of drugs. Toxicol. Sci. 136, 242–249 (2013).
Kim, E. & Nam, H. Prediction models for drug-induced hepatotoxicity by using weighted molecular fingerprints. BMC Bioinforma. 18, 227 (2017).
Yang, S., Ooka, M., Margolis, R. J. & Xia, M. Liver three-dimensional cellular models for high-throughput chemical testing. Cell Rep. Methods 3, 100432 (2023).
Chen, M. et al. DILIrank: the largest reference drug list ranked by the risk for developing drug-induced liver injury in humans. Drug Discov. Today 21, 648–653 (2016).
Serrano, J. LiverTox: An online information resource and a site for case report submission on drug-induced liver injury. Clin. Liver Dis. (Hoboken) 4, 22–25 (2014).
Igarashi, Y. et al. Open TG-GATEs: a large-scale toxicogenomics database. Nucleic Acids Res 43, D921–D927 (2015).
O’Brien, P. J. et al. High concordance of drug-induced human hepatotoxicity with in vitro cytotoxicity measured in a novel cell-based model using high content screening. Arch. Toxicol. 80, 580–604 (2006).
Xu, J. J. et al. Cellular imaging predictions of clinical drug-induced liver injury. Toxicol. Sci. 105, 97–105 (2008).
Tolosa, L. et al. Development of a multiparametric cell-based protocol to screen and classify the hepatotoxicity potential of drugs. Toxicol. Sci. 127, 187–198 (2012).
Persson, M., Løye, A. F., Mow, T. & Hornberg, J. J. A high content screening assay to predict human drug-induced liver injury during drug discovery. J. Pharmacol. Toxicol. Methods 68, 302–313 (2013).
Garside, H. et al. Evaluation of the use of imaging parameters for the detection of compound-induced hepatotoxicity in 384-well cultures of HepG2 cells and cryopreserved primary human hepatocytes. Toxicol. Vitr. 28, 171–181 (2014).
Schadt, S. et al. Minimizing DILI risk in drug discovery — A screening tool for drug candidates. Toxicol. Vitr. 30, 429–437 (2015).
Proctor, W. R. et al. Utility of spherical human liver microtissues for prediction of clinical drug-induced liver injury. Arch. Toxicol. 91, 2849–2863 (2017).
Khetani, S. R. et al. Use of micropatterned cocultures to detect compounds that cause drug-induced liver injury in humans. Toxicol. Sci. 132, 107–117 (2013).
Porceddu, M. et al. Prediction of liver injury induced by chemicals in human with a multiparametric assay on isolated mouse liver mitochondria. Toxicol. Sci. 129, 332–345 (2012).
Gustafsson, F., Foster, A. J., Sarda, S., Bridgland-Taylor, M. H. & Kenna, J. G. A correlation between the in vitro drug toxicity of drugs to cell lines that express human P450s and their propensity to cause liver injury in humans. Toxicol. Sci. 137, 189–211 (2014).
Vorrink, S. U., Zhou, Y., Ingelman-Sundberg, M. & Lauschke, V. M. Prediction of drug-induced hepatotoxicity using long-term stable primary hepatic 3D spheroid cultures in chemically defined conditions. Toxicol. Sci. 163, 655–665 (2018).
Albrecht, W. et al. Prediction of human drug-induced liver injury (DILI) in relation to oral doses and blood concentrations. Arch. Toxicol. 93, 1609–1637 (2019).
Aleo, M. D. et al. Moving beyond binary predictions of human drug-induced liver injury (DILI) toward contrasting relative risk potential. Chem. Res. Toxicol. 33, 223–238 (2020).
Sakatis, M. Z. et al. Preclinical strategy to reduce clinical hepatotoxicity using in vitro bioactivation data for >200 compounds. Chem. Res. Toxicol. 25, 2067–2082 (2012).
Dawson, S., Stahl, S., Paul, N., Barber, J. & Kenna, J. G. In vitro inhibition of the bile salt export pump correlates with risk of cholestatic drug-induced liver injury in humans. Drug Metab. Dispos. 40, 130–138 (2012).
Wang, E. et al. TxGemma: Efficient and Agentic LLMs for Therapeutics. arXiv [cs.AI] (2025).
Lee, T. & Posma, J. Improving drug-induced liver injury prediction using graph neural networks with augmented graph features from molecular optimisation. ChemRxiv https://doi.org/10.26434/chemrxiv-2024-d12gk-v2 (2025).
Seal, S. et al. Improved detection of drug-induced liver injury by integrating predicted in vivo and in vitro data. Chem. Res. Toxicol. 37, 1290–1305 (2024).
Walker, P. A., Ryder, S., Lavado, A., Dilworth, C. & Riley, R. J. The evolution of strategies to minimise the risk of human drug-induced liver injury (DILI) in drug discovery and development. Arch. Toxicol. 94, 2559–2585 (2020).
Ewart, L. et al. Performance assessment and economic analysis of a human Liver-Chip for predictive toxicology. Commun. Med. (Lond.) 2, 154 (2022).
Fäs, L. et al. Physiological liver microtissue 384-well microplate system for preclinical hepatotoxicity assessment of therapeutic small molecule drugs. Toxicol. Sci. 203, 79–87 (2025).
Wilkening, S., Stahl, F. & Bader, A. Comparison of primary human hepatocytes and hepatoma cell line Hepg2 with regard to their biotransformation properties. Drug Metab. Dispos. 31, 1035–1042 (2003).
Olsavsky, K. M. et al. Gene expression profiling and differentiation assessment in primary human hepatocyte cultures, established hepatoma cell lines, and human liver tissues. Toxicol. Appl. Pharmacol. 222, 42–56 (2007).
Grinberg, M. et al. Toxicogenomics directory of chemically exposed human hepatocytes. Arch. Toxicol. 88, 2261–2287 (2014).
Kiamehr, M. et al. Dedifferentiation of primary hepatocytes is accompanied with reorganization of lipid metabolism indicated by altered molecular lipid and miRNA profiles. Int. J. Mol. Sci. 20, 2910 (2019).
Zdrazil, B. et al. The ChEMBL Database in 2023: a drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res 52, D1180–D1192 (2024).
Shah, F. et al. Setting clinical exposure levels of concern for drug-induced liver injury (DILI) using mechanistic in vitro assays. Toxicol. Sci. 147, 500–514 (2015).
Williams, D. P., Lazic, S. E., Foster, A. J., Semenova, E. & Morgan, P. Predicting drug-induced liver injury with Bayesian machine learning. Chem. Res. Toxicol. 33, 239–248 (2020).
Kelder, T. et al. WikiPathways: building research communities on biological pathways. Nucleic Acids Res. 40, D1301–D1307 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Rao, M. S. et al. Comparison of RNA-Seq and microarray gene expression platforms for the toxicogenomic evaluation of liver from short-term rat toxicity studies. Front. Genet. 9, 636 (2018).
Allard, J. et al. Drug-induced hepatic steatosis in absence of severe mitochondrial dysfunction in HepaRG cells: proof of multiple mechanism-based toxicity. Cell Biol. Toxicol. 37, 151–175 (2021).
Bethesda (MD): National Institute of Diabetes and Digestive and Kidney Diseases. in LiverTox: Clinical and Research Information on Drug-Induced Liver Injury (2012).
Hliwa, A., Ramos-Molina, B., Laski, D., Mika, A. & Sledzinski, T. The role of fatty acids in non-alcoholic fatty liver disease progression: An update. Int. J. Mol. Sci. 22, 6900 (2021).
Osborne, T. F. & Espenshade, P. J. Lipid balance must be just right to prevent development of severe liver damage. J. Clin. Investig. 132, 11 (2022).
Shang, H. et al. Gut microbiota-derived tryptophan metabolites alleviate liver injury via AhR/Nrf2 activation in pyrrolizidine alkaloids-induced sinusoidal obstruction syndrome. Cell Biosci. 13, 1 (2023).
Zhu, L. et al. The emerging role of ferroptosis in various chronic liver diseases: Opportunity or challenge. J. Inflamm. Res. 16, 381–389 (2023).
Chen, S.-S. et al. Serum metabolomic analysis of chronic drug-induced liver injury with or without cirrhosis. Front. Med. (Lausanne) 8, 640799 (2021).
Wagner, M., Zollner, G. & Trauner, M. Nuclear receptors in liver disease. Hepatology 53, 1023–1034 (2011).
da Silva, R. P., Eudy, B. J. & Deminice, R. One-carbon metabolism in fatty liver disease and fibrosis: One-carbon to rule them all. J. Nutr. 150, 994–1003 (2020).
Cai, S.-Y. & Boyer, J. L. The role of bile acids in cholestatic liver injury. Ann. Transl. Med. 9, 737 (2021).
Ray, W. A., Griffin, M. R. & Stein, C. M. Cardiovascular toxicity of valdecoxib. N. Engl. J. Med. 351, 2767 (2004).
Shrier, M., Díaz, J. E. & Tsarouhas, N. Cardiotoxicity associated with bupropion overdose. Ann. Emerg. Med. 35, 100 (2000).
Levine, M., Pizon, A. F., Padilla-Jones, A. & Ruha, A.-M. Warfarin overdose: a 25-year experience. J. Med. Toxicol. 10, 156–164 (2014).
Kohonen, P. et al. A transcriptomics data-driven gene space accurately predicts liver cytopathology and drug-induced liver injury. Nat. Commun. 8, 15932 (2017).
Bergen, V., Srikrishnan, S. & Cellarity Inc. Cellarity/DILImap. (Zenodo, 2025). https://doi.org/10.5281/ZENODO.17290520.
Acknowledgements
We sincerely thank Bill Pennie and Atli Thorarensen for bringing in pivotal ideas that shaped the work. We appreciate Govinda Bhisetti for the thorough review of the manuscript. Our gratitude extends to Robb Nicewonger for his work on the chemistry SOP, Cameron Reilly for his support with RNA extraction and automation, Laura Isacco for coordinating data generation timelines, Thao Tran and Wynter Guess for their assistance with compound screening, Winnie Lee and Brian Yi for their work on RNA extraction, all of which contributed to data generation foundational to this work. Finally, we thank the reviewers for their thoughtful and constructive requests, which helped refine and strengthen the work.
Author information
Authors and Affiliations
Contributions
V.B. and K.K. conceived and co-led the project. V.B. developed the framework and authored the manuscript. K.K. directed the design and execution of the cell culture experiments. S.S. implemented and ran benchmarking against state-of-the-art in silico models. O.B. contributed to the study’s conceptual framework. S.A. contributed to the development of the culture system and conducted viability screens. M.R. established the initial experiments and contributed to RNA-seq method selection. C.F. implemented automation for data generation workflows. H.B. led compound management, screening and contributed to the chemistry SOP. N.R. performed library construction for sequencing. T.F. contributed to compound management and screening. N.L. established the Dotmatics workflows. M.C. supervised the data generation activities. N.P. was involved in the original conception of using transcriptomics to model off-target effects. M.G. provided strategic direction. M.Z. provided oversight for the machine learning aspects and strategic direction.
Corresponding authors
Ethics declarations
Competing interests
All authors were employees of Cellarity Inc. at the time the study was performed and hold equity in the company. The research was funded by Cellarity. V.B., K.K., and O.B. are inventors on patent application WO2025/024525, related to methods for DILI prediction, assigned to Cellarity.
Peer review
Peer review information
Nature Communications thanks Volker Lauschke, James Dear and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bergen, V., Kodella, K., Srikrishnan, S. et al. A large-scale human toxicogenomics resource for drug-induced liver injury prediction. Nat Commun 16, 9860 (2025). https://doi.org/10.1038/s41467-025-65690-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65690-3