Abstract
Myelodysplastic syndrome (MDS) is characterized by bone marrow failure, clonal evolution and leukemic progression, but the pathophysiologic processes driving these events remain incompletely understood. Here, by establishing a comprehensive single-cell transcriptional taxonomy of human MDS, we reveal that inflammatory remodeling of bone marrow stromal niches is a common early feature, irrespective of the genetic driver landscape. We identify an activated CD8-T-cell subset as a source of stromal inflammation via TNF-receptor signaling, which prompts the inflammatory rewiring and loss of repopulating ability of residual normal hematopoietic stem/progenitor cells (HSPC). Mutant HSPCs display relative resistance to this inflammatory stress and reside predominantly in a transcriptional ‘high output’ state, providing a biological framework to their competitive advantage in an inflammatory microenvironment. Consistent with this, stromal inflammation associates with leukemic progression and reduced survival. Our data thus support a model of immune-stromal inflammatory signaling driving tissue failure and clonal evolution in the hematopoietic system. Mechanisms of clonal evolution in myeloid neoplasms remain incompletely understood. Darwinian theory predicts that the (micro)environment of clone-propagating stem cells may contribute to clonal selection. Here, we provide data fitting this model, establishing a relationship between stromal niche inflammation, inflammatory stress in HSPCs, clonal resistance and leukemic evolution in human MDS.
Similar content being viewed by others
Introduction
Myelodysplastic syndrome (MDS) is a clonal disease characterized by inefficient hematopoiesis and a strong propensity to develop acute myeloid leukemia (AML)1,2. It is initiated by (epi)genetic events in hematopoietic stem/progenitor cells (HSPC), including chromosomal aberrations and somatic driver mutations in genes encoding splicing factors or epigenetic regulators3. HSPC clones carrying such genetic lesions can persist in the bone marrow for a long time, eventually outcompeting normal HSPCs, accumulating additional mutations and ultimately transforming into AML4,5. Although it has been clear that the driving genetic events play central roles in the positive selection and expansion of mutant clones, hematopoietic cell intrinsic events may not always be sufficient to drive a competitive advantage. For example, mutations in SF3B1, among the most frequently mutated genes in lower-risk(LR)-MDS6,7, do not confer an obvious competitive advantage to hematopoietic cells in model systems8, suggesting that environmental selective pressure may be required for clonal expansion.
One potential cellular component within the bone marrow ecosystem that forms the direct environment of HSPCs, and may be involved in driving MDS pathogenesis, is bone marrow stromal cells (BMSC)9. BMSCs represent a population of non-hematopoietic cells comprising up to 20% of cellular volume of normal bone marrow (NBM)10. These cells pervade the tissue in extensive networks and physically interact with the vast majority of hematopoietic cells. As a heterogeneous population, their biological functions are closely associated with their surface marker expression and anatomic localization11. A BMSC subset with high expression of leptin receptor (LEPR) is considered as a central regulator of normal hematopoiesis12,13,14. These LEPR+ stromal cells, mostly residing around sinusoidal blood vessels11, serve as the major source of key HSPC regulatory factors such as C-X-C motif chemokine ligand 12 (CXCL12), stem cell factor (SCF, encoded by KITLG) and interleukin-7 (IL7), which are crucial for retention, survival, self-renewal and lineage differentiation of HSPCs. In addition, BMSCs also play a vital role in regulating development and activation of immune cells like natural killer (NK), B, myeloid and dendritic cells (DC)14,15,16,17,18.
BMSCs have been implicated in MDS disease pathogenesis in experimental models. Genetic abnormalities in BMSCs have been demonstrated to be sufficient or required for disease initiation, evolution and propagation in mouse models19,20,21,22,23,24,25, but the cellular heterogeneity of BMSCs in MDS, their predicted interactions with mutant vs. non-mutant HSPCs and adaptive/innate immune cells in human MDS have remained largely elusive.
We recently established a comprehensive taxonomy of the predicted cellular interactions between LEPR+ stromal niches, HSPCs, and adaptive and innate immune cells in the human NBM and described the disruption of these signaling pathways in AML using adapted single-cell RNA sequencing (scRNA-seq)26.
Building on this methodology, we here report a comprehensive cellular taxonomy of the bone marrow ecosystem in human lower risk (LR)-MDS. The data reveal that inflammatory remodeling of stromal stem cell niches contributes to clonal evolution in this disorder by relative resistance of mutated HSPCs to repressive inflammatory signaling, which is associated with leukemic progression and clinical outcome. Our findings are consistent with a model in which differing interactions of mutant and non-mutant cells with their tumor-remodeled inflammatory micro-environment drive clonal selection and disease evolution.
Results
A comprehensive cellular taxonomy of human lower-risk (LR)-MDS
To assess the heterogeneity of HSPCs, their stromal niches and all other cells in the bone marrow, and predict potential cellular interactions, we established a comprehensive cellular taxonomy of human LR-MDS (representing early phases in disease evolution) by performing adapted scRNA-seq on bone marrow aspirates, as recently described26. Bone marrow aspirates of newly diagnosed LR-MDS patients (n = 5) carrying mutations in SF3B1 and/or TET2 (Supplementary Data 1), which are among the most common mutated genes in MDS27, were investigated. To ensure a robust representation of all bone marrow cell populations, including the rare (HSPC) subpopulations and their stromal niches, we used flow cytometry to sort the bone marrow cells into HSPCs (CD45+CD34+), myeloid-lineage cells (CD45+CD34–CD33+/CD117+), non-myeloid hematopoietic cells (CD45+CD34–CD33–CD117-), endothelial cells (EC) (CD45–CD71–CD31+) and BMSCs (CD45–CD71–CD31–) (Fig. 1a). Sorted cells were pooled into two fractions (HSPCs with myeloid cells and non-hematopoietic cells with non-myeloid cells) and were subjected to scRNA-seq in two separate runs, followed by integration into a single dataset. This dataset was subsequently integrated with a NBM scRNA-seq dataset of aspirates from healthy bone marrow donors (n = 4)26. This comprehensive cellular taxonomy comprised 47,950 NBM cells and 57,478 LR-MDS cells (Fig. 1b).
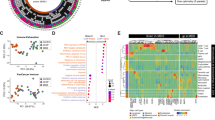
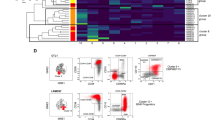
a FACS gating strategy for the sorting of hematopoietic and non-hematopoietic cell populations from bone marrow aspirates of LR-MDS patients at diagnosis for single cell RNA sequencing (scRNA-seq) (adapted from26). Created in BioRender. Bian, Y. (2025) https://BioRender.com/0sa14pd. In order to obtain robust representation of all bone marrow cell subtypes in the scRNA-seq data, bone marrow cells were sorted into HSPCs (CD45+CD34+), myeloid cells (CD45+CD34–CD33+/CD117+), non-myeloid lineage hematopoietic cells (CD45+CD34–CD33–/CD117–), endothelial cells (CD45–CD235a–CD71–CD31+) and non-endothelial bone marrow stromal cell (BMSCs) (CD45–CD235a–CD71–CD31–) fractions. The sorted cells were pooled into two populations and subjected to scRNA-seq, followed by integration of the data into a single dataset. b Uniform Manifold Approximation and Projection (UMAP) plot of mononuclear cells from bone marrow aspirates, representing 47,950 cells from healthy donors (n = 4) and 57,478 cells from LR-MDS patients (n = 5). c Expression of cell-type identifying genes in NBM and LR-MDS across all bone marrow cell subtypes. Cell-type identifying genes are defined as the top 50 most upregulated genes in each cell type in NBM. d Expression of BMSC markers, HSPC markers and lymphocyte markers in NBM and LR-MDS.
Using fully-annotated public datasets as refs. 12,26,28,29, the sequenced cells were classified into 18 subpopulations, including HSPCs (comprising hematopoietic stem cells/multipotent progenitors (HSC/MPP), lymphoid-myeloid primed progenitors (LMPP), megakaryocyte and erythroid progenitors (MEP), common lymphoid progenitors (CLP) and common myeloid progenitors (GMP)), myeloid cells (monoblasts, DCs, CD14+ monocytes and CD16+ monocytes), non-myeloid/lymphoid cells (NKs, CD4+ T, CD8+ T, B and plasma cells), megakaryocytes (Mk), erythroid progenitors and non-hematopoietic cells (EC and BMSC) (Fig. 1b).
The cellular taxonomy of the bone marrow was relatively preserved in LR-MDS, with all cell subtypes found in NBM present in LR-MDS, and exhibiting similar patterns of marker gene expression (top 50 most upregulated genes in each cell subtype versus other cells in NBM) (Fig. 1c). Annotation accuracy was validated by detecting the expression of marker genes “identifying” cell types (Fig. 1d and Supplementary Fig. 1a). Monocytes, for instance, specifically expressed genes encoding the myeloid markers ITGAM (CD11b) and CD14. T cell subsets specifically expressed the T lymphocyte markers CD3D, CD7, IL7R, CD4 and/or CD8A. HSPCs commonly expressed CD34, KIT and FLT3, with genes encoding key stemness regulators such as HLF, MEIS1 and CRHBP were preferentially expressed in the HSC/MPP subset. Genes associated with myeloid lineage development such as MPO, ELANE, AZU1 and LYZ were differentially expressed in GMPs, while erythroid developmental genes such as GATA1, HBD and CA1 were differentially expressed in MEPs and downstream erythroid progenitors.
BMSCs in our dataset highly expressed genes encoding the mesenchymal stromal markers PRRX1, NGFR (CD271), PDGFRA, pre-adipocyte markers LEPR, LPL, and the HSPC regulatory factors CXCL12, KITLG (SCF) and IL7 (Fig. 1d and Supplementary Fig. 1a), but not osteolineage markers (BGLAP, SPP1), chondrocyte differentiation markers (SOX9, ACAN, COL2A1), fibroblast markers (S100A4, SEMA3C) or pericyte markers (NES, CSPG4 and ACTA2)12 (data not shown), indicating that BMSCs from LR-MDS and NBM aspirates are the likely equivalents of LEPR+ sinusoid perivascular BMSCs with high HSPC supporting capacity, as defined in mice11,12,13,14,30.
Inflammatory remodeling of stromal niches in LR-MDS
Establishing the cellular taxonomy of LR-MDS bone marrow enables the transcriptional characterization of disease-specific BMSCs. Differential gene expression analysis between BMSCs from LR-MDS (n = 4/5 MDS samples, no BMSCs could be retrieved from sample MDS04) and NBM (n = 4) showed upregulation of 632 genes and downregulation of 164 genes in the LR-MDS BMSCs (Supplementary Data 2). The upregulated genes included genes related to inflammatory activation, such as ANXA1, TNFAIP8, NFKBIA, GADD45B, INHBA, CXCL8 and CD44 (padj < 0.05) (Fig. 2a). Genes encoding the key HSPC regulatory factors KITLG, ANGPT1 and IL7, on the other hand, were significantly downregulated (Fig. 2a) in LR-MDS BMSCs. Transcriptional signatures indicative of inflammatory activation in LR-MDS were confirmed by gene-set enrichment analysis (GSEA), showing enrichment of NFĸB-associated inflammation (TNFα signaling via NFĸB, Inflammatory responses), as well as transcriptional programs associated with cellular stress and apoptosis (Apoptosis and P53 pathway) (False discovery rate (FDR) < 0.05), while the significant downregulation of signatures associated with oxidative phosphorylation may point at metabolic alterations in MDS BMSCs (Fig. 2b).
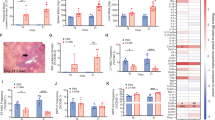
a Volcano plot of differentially expressed genes in BMSCs of LR-MDS versus NBM. Inflammation-associated genes and genes encoding HSPC regulatory factors are highlighted. Differential expression gene analysis is performed at sample level using the pseudoDE R-package. Statistical analysis was performed by a two-sided Wald test and adjusted for FDR. b Enrichment of Hallmark gene signatures in BMSCs of LR-MDS versus NBM. Positive NES (Normalized Enrichment Score) reflects programs enriched in LR-MDS, while negative scores indicate enrichment in NBM. Statistical analysis was performed by a Kolmogorov–Smirnov (K-S) test and adjusted for FDR. c, d Distribution and frequencies of BMSC subsets in NBM (n = 4) and LR-MDS (n = 4). Statistical analysis was performed by a two-sided unpaired Welch’s t-test (d). Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers (d). e Expression of gene signatures reflecting NFĸB-associated inflammatory activation in BMSCs. BMSC cluster-2 (BMSC-2) is characterized by increased expression of transcriptional programs indicative of (NFĸB associated) inflammatory activation. Statistical analysis was performed by a one-sided K-S test (via the GSEA algorithm) and the p-value was corrected for FDR. f Expression of Annexin A1 at the protein level, as assessed by flow cytometry on stromal cells in BM aspirates. NBM (n = 5) and LR-MDS (n = 4). Investigations were performed in patients not included in the scRNA-seq analyses. Statistical analysis was performed by a two-sided unpaired Welch’s t-test. Error bar represents mean ± SEM. g Expression of HSPC niche factors across the BMSC subsets in NBM and LR-MDS. NBM (n = 4) and LR-MDS (n = 4). Statistical analysis was performed by a two-sided Wald test and adjusted for FDR using the PseudoDE R-package at sample level. Source data are provided as a Source Data file.
Sub-clustering of BMSCs from NBM samples confirmed the presence of a BMSC subset (BMSC-0) with high expression of LEPR and HSPC regulatory factors (KITLG, CXCL12) (Fig. 2c and Supplementary Fig. 1b), as recently reported by us26. Expression of the critical HSPC regulatory factor KITLG13 gradually decreased from BMSC subset 0 to 3, with significant lower expression in BMSC 2-3 in comparison to BMSC 0-1 (Supplementary Fig. 1b). In LR-MDS, the relative size of BMSC-2 was significantly increased in comparison to NBM (from 12.6 ± 1.36% in NBM to 38.4 ± 8.2% in LR-MDS), with a concomitant decrease in the relative size of BMSC-1 (from 73.2 ± 6.05% in NBM to 47.6 ± 10.7%, p = 0.011) (Fig. 2c, d).
The relative abundancy of BMSC-2 in LR-MDS prompted us to further characterize this stromal subset. GSEA revealed that cells in BMSC-2 exhibited a significantly increased expression of gene signatures reflecting NFĸB-associated inflammatory activation (padj < 0.05) (Fig. 2e and Supplementary Fig. 1c). Consistent with this finding, the pro-inflammatory cytokines CXCL3, CXCL5, CXCL8, IL6 and CCL2, inflammatory modulator ANXA131, and genes indicative of inflammatory activation (CD44 and TNFAIP8) were among the most significantly upregulated genes in BMSC-2 compared to other clusters in LR-MDS (padj < 0.05) (Supplementary Fig. 1d, e and Supplementary Data 2). Stromal inflammatory features were confirmed in situ at the protein level by showing increased expression of Annexin A1 (encoded by ANXA1, among the most upregulated genes in BMSCs of LR-MDS versus NBM (Fig. 2a and Supplementary Fig. 1d)) in CD45-CD235a-CD271+ BMSCs by flowcytometric assessment of aspirates from LR-MDS patients (Fig. 2f and Supplementary Fig. 1f).
The relative expansion of the inflammatory BMSC-2 subset was associated with a reduction in the expression of KITLG, IL7 and ANGPT1 in BMSC-0, −1 and −2 compared to their counterparts in NBM (Fig. 2g), suggesting the existence of a broader compromise of stromal HSPC supportive capacity in LR-MDS.
Inflammatory remodeling of stromal niches is common in LR-MDS, irrespective of the genetic driver landscape
LR-MDS is a genetically and phenotypically heterogeneous disease triggered by diverse genetic events in HSPCs, of which some, such as mutations in genes encoding epigenetic or splicing regulators (TET2, SF3B1 and ASXL1), have been associated with exaggerated inflammatory signaling32,33,34. We therefore questioned whether the inflammatory remodeling of BMSCs in LR-MDS is associated with certain driver genetic variants or represents a more common feature of LR-MDS, independent of the driver landscape.
To this end, we employed a BMSC bulk RNA-seq dataset from highly purified CD45–CD235a–CD31–CD271+ BMSCs isolated from diagnostic bone marrow aspirates of 45 LR-MDS patients included in the HOVON89 clinical trial (LR-MDS patients (International Prognostic Scoring System (IPSS) low and intermediate-1) (https://www.hovon.nl/en/trials/ho89; NTR1825 (former ID); NL1715 (recent ID); EudraCT 2008-002195-10))35, in comparison to BMSCs of normal controls (n = 10, age-matched healthy donors)20. The extent of BMSC inflammatory remodeling was assessed by constructing an enrichment score (iBMSC score), using single sample Gene Set Variant Analysis (GSVA)36, from genes significantly upregulated in the inflammatory BMSC-2 cluster (Fig. 3a). As expected, the median iBMSC score was significantly elevated in BMSCs from LR-MDS (n = 45) in comparison to NBM (n = 10) (mean iBMSC score 1.53 ± 0.24 vs. 1.21 ± 0.17, p = 0.0002, respectively, although a wide variability in iBMSC-scores was noted among LR-MDS samples (range 0.9–1.89) (Fig. 3b). Consistent with a higher iBMSC-score in LR-MDS, expression of inflammatory associated transcriptional programs (TNFα signaling via NFĸB, Inflammatory Response, and IL6-JAK-STAT3 signaling) and genes (NFKBIA, CD44, ANXA1 and IL8) were significantly upregulated in BMSCs from LR-MDS (Fig. 3c, d). To further demonstrate whether the upregulation of iBMSC score and the pro-inflammatory genes in LR-MDS BMSCs were associated with expansion of the inflammatory BMSC-2 cluster, the CIBERSORTx algorithm was applied to deconvolute the bulk-RNA-seq dataset, showing that the relative size of the BMSC-2 subpopulation was significantly increased in LR-MDS patients, confirming data from the scRNA-seq analyzes (Fig. 2c, d and Supplementary Fig. 1g).
a, b Assessment of BMSC inflammatory remodeling in a larger cohort of LR-MDS patients (LR-MDS (n = 45) and NBM (n = 10)) (data derived from: EGAS00001001926). Genes differentially upregulated in BMSC-2 (in comparison to other BMSC clusters combined) (a) were used to generate a BMSC inflammatory score (iBMSC score). This iBMSC score was calculated from transcriptional data of highly purified stromal cells from the larger cohort of LR-MDS patients, confirming significantly increased, albeit variable, inflammation scores in BMSCs from LR-MDS versus NBM (b). Statistical analysis was performed by a two-sided Wilcoxon rank-sum test. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers (b). c Enrichment of Hallmark gene signatures in BMSCs from LR-MDS versus NBM in bulk RNA-seq. FDR adjusted p-value < 0.05. Positive NES indicates programs enriched in LR-MDS BMSC. Statistical analysis was performed by a K-S test and adjusted for FDR. d Differential expression of genes encoding inflammation-associated proteins in BMSCs from LR-MDS (n = 45) in comparison to NBM (n = 10). Statistical analysis was performed by a two-sided Wald test and adjusted for FDR. TPM = transcript per million. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers. e The level of inflammatory activation of BMSCs, as reflected by the iBMSC score, does not correlate with the mutational landscape or presence of cytogenetic abnormalities in LR-MDS. f iBMSC score in BMSCs of LR-MDS patients aggregated by ‘generic’ subgroups defined by either a mutation in a splice-factor encoding gene and/or an epigenetic regulator (left) or the presence of cytogenetic abnormalities (right). LR-MDS (n = 45). Statistical analysis was performed by a two-sided Wilcoxon rank-sum test. ns, not significant. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers.
The extent of BMSC inflammatory activation was not associated with the driver landscape (as assessed by the TruSight myeloid NGS panel) in this dataset of LR-MDS, with mutations in TET2, SF3B1 and ASXL1 being present across the iBMSC score range (Fig. 3e). In line with this notion, presence of a mutation in genes encoding splice factors and/or epigenetic regulators were not associated with higher iBMSC scores (Fig. 3f, left panel). Similarly, the presence of chromosomal abnormalities (as detected by conventional karyotyping) (Fig. 3f, right panel) or the total number of mutations in the LR-MDS marrow (Fig. 3e) was not associated with the level of stromal inflammatory activation.
Collectively, the data indicate that inflammatory remodeling of stromal niches is a common finding in LR-MDS, albeit at variable levels, with no clear association with the driver landscape.
Network analysis identifies a pro-inflammatory subset of CD8+ T-cells as a putative source of the inflammatory remodeling of BMSCs in LR-MDS
We next interrogated putative mechanisms and cellular sources of BMSC inflammation in LR-MDS. Given that the transcriptional programs TNFα Signaling via NFĸB was among the most upregulated inflammatory gene signatures in BMSCs in LR-MDS (Fig. 2b), we hypothesized that TNFα could be a trigger of the inflammatory activation of stromal niches. Analysis of the cellular taxonomy demonstrated that CD8+ T cells have the highest expression of genes encoding TNFα (TNF) and IFNγ (IFNG) in LR-MDS, followed by CD4+ T-cells and NK cells (Fig. 4a). Interestingly, expression of the canonical receptors for TNFα (TNFRSF1A) and IFNγ (IFNGR1 and IFNGR2) was the highest in BMSCs in comparison to all other bone marrow resident cells (Fig. 4a). Consistent with these findings, ligand-receptor interaction analysis predicted the existence of a strong CD8+ T cell-BMSC IFNγ signaling axis (Supplementary Fig. 2a).
a Expression of genes encoding TNFα, IFNγ (left panel) and their canonical receptors (right panel) in bone marrow cell subtypes in NBM and LR-MDS indicating the highest expression in CD8+ T-cells in LR-MDS and differential expression of their canonical receptors in BMSCs. Size and color of the dot plot represent percentage of expressing cells and level of expression, respectively. b Representative image showing immunofluorescence staining of CD271 and CD8 in the SF3B1-mutated LR-MDS bone marrow (n = 3), demonstrating direct contact between a substantial subset (21–27%, lower panel) of CD8+ T-cells and CD271+ BMSCs. White arrows indicate examples of direct contact between CD8+ T-cells (green) in CD271+ (purple) BMSCs. The nuclei were counterstained with DAPI (blue). Error bar represents mean ± SEM. c, d Comparative CD8+ T-cell gene expression in LR-MDS vs. NBM. CD8+ T cells in LR-MDS have increased expression of transcriptional programs indicative of immune activation, as demonstrated by differential gene expression analysis (c) and GO term analysis (d). Positive NES indicates programs enriched in LR-MDS, while negative scores indicate enrichment in NBM. Statistical analysis was performed by a two-sided Wald test and adjusted for FDR (c). Statistical analysis was performed by a K-S test and adjusted for FDR (d). e Expression of naïve markers, immuno-modulating genes, activation markers and pro-inflammatory cytokines in CD8+ T cells across all samples in NBM (n = 4) and LR-MDS (n = 5). Statistical analysis was performed by a two-sided Wald test with FDR correction. f Distribution and frequency of CD8+ T-cell subsets in NBM (n = 4) and LR-MDS (n = 5) indicating relative expansion of EM CD8+ T cells in LR-MDS. Statistical analysis was performed by a two-sided unpaired Welch’s t-test. ns, not significant. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers. g Expression of naïve markers, immune modulating genes, cytotoxic factors, activation markers and pro-inflammatory cytokines in CD8+ T cell subsets in LR-MDS and NBM indicating differential expression of pro-inflammatory cytokines, including TNF, IFNG and CCL3 in IFNG+ EM CD8+ T cells in LR-MDS. h Frequency of EM cells within the CD8+ T-cell subsets as demonstrated by flow-cytometry. Representative plots in left panel. NBM (n = 5) and LR-MDS (n = 6), from 3 independent experiments. Statistical analysis was performed by a two-sided unpaired Welch’s t-test. Error bar represents mean ± SEM. i Activation of the CD8 T-cell EM subset in LR-MDS, as indicated by increased expression of CD69 by flowcytometric assessment. NBM (n = 5) and LR-MDS (n = 6), from 3 independent experiments. Statistical analysis was performed by a two-sided unpaired Welch’s t-test. Error bar represents mean ± SEM. j Induction of NFKBIA and reduction in KITLG gene expression, recapitulating aspects of stromal inflammation in LR-MDS (n = 4, from 4 independent experiments). Statistical analysis was performed by a one-way ANOVA and Tukey’s multiple comparisons test. ns, not significant. Error bar represents mean ± SEM. k Partial rescue of the inflammatory activation by knockdown of the canonical TNFα-receptor encoded by TNFRSF1A in BMSCs (n = 3, from 3 independent experiments). Statistical analysis was performed by a two-sided unpaired Welch’s t-test. ns, not significant. Error bar represents mean ± SEM. Source data are provided as a Source Data file.
In situ immunofluorescence of the bone marrow of SF3B1-mutated LR-MDS (n = 3) revealed intimate anatomical relationships (defined as direct contact) between a substantial subset of CD8+ T-cells (range 21–27%) and CD271+ BMSCs (Fig. 4b), further suggesting the possibility of active crosstalk between these cell types.
Subclustering of T and NK cell subpopulations resulted in 10 immune subsets based on expression of genes encoding immune cell canonical markers (Supplementary Fig. 2b). Subsets included CCR7+ naïve CD4+ T cells, CXCR3+/CCR6+ Th1/Th17 CD4+ T cells, CCR4+ Th2 CD4+ T cells, FOXP3+CTLA4+ regulatory T cells, CCR7+ naïve CD8+ T cells, Granzyme K (GZMK)+ effector memory (EM) CD8+ T cells, granzyme B (GZMB)+ GLYN+ terminally differentiated effector memory (TEMRA) CD8+ T cells, CD56high NK cells and CD56dim NK cells37,38 (Supplementary Fig. 2b). The relative distribution of these immune subsets appeared to be relatively preserved in LR-MDS (Supplementary Fig. 2c) while TNF and IFNG were predominantly expressed in CD8+ T cell subsets (Fig. 4a and Supplementary Fig. 2d).
Differential gene expression analysis and gene ontology (GO) analysis showed a significant enrichment of gene signatures reflecting immune activation in CD8+ T cells of LR-MDS (Fig. 4c–e) including upregulation of the expression of canonical immune activation markers such as HLA-DRA, CD69, and TNFRSF9 (encoding 4-1BB), with a concomitant downregulation of genes encoding naïve T-cell markers TCF7, SELL (encoding CD62L), CCR7, and IL7R (Fig. 4e). In line with immune activation, expression of genes encoding the pro-inflammatory cytokines TNFα and/or IFNγ, TGFβ, and CCL3 was significantly elevated in LR-MDS (Fig. 4e), suggesting the CD8+ T cells in LR-MDS harbor a strong pro-inflammatory capacity.
Analysis of the heterogeneity of CD8+ T cells in LR-MDS revealed a relative expansion of the EM CD8+ T cell subpopulation (Fig. 4f and Supplementary Fig. 2c), which included an IFNG+ EM CD8+ subset displaying the highest expression of the immune activation markers TNFRSF9, CD69, HLA-DRA and pro-inflammatory cytokines TNF, IFNG, CCL3 and CCL4, in comparison to other CD8+ T cell subtypes in LR-MDS (Fig. 4g). This is reminiscent of the tissue-resident GZMK+CD8+ cells that drive tissue inflammation and disease pathogenesis in inflammatory disorders like rheumatoid arthritis39. Of further interest, expansion of TEMRA CD8+ T cells producing high levels of cytotoxic factors like GZMB and Perforin-1 to trigger CD8+ T-cell–targeted cell death40,41,42 was not observed in LR-MDS (Fig. 4f and Supplementary Fig. 2e).
Multi-parameter flow-cytometry on primary SF3B1-mutated LR-MDS samples vs. NBM to validate the transcriptional data at the protein level, confirmed relative expansion of the CD8+ T EM (CCR7-CD45RA–) subset with high expression of CD69 and GZMK in LR-MDS (Fig. 4h, i and Supplementary Fig. 2f, g), without relative expansion of the CD8+ TEMRA subset (CCR7-CD45RA+) and without an increase of expression in GZMB and Perforin-1 in comparison to the NBM (Supplementary Fig. 2f), pointing at a potential disbalance between inflammation-driving (EM) and cytotoxic (TEMRA) CD8+-T cell subsets in LR-MDS, as recently described in human inflamed tissues39.
To experimentally address whether the secretome of activated non-clonal T cells can indeed serve as a source of the inflammatory remodeling of stromal niches, normal T-cells from peripheral blood mononuclear cell (PBMC) fractions were activated using anti-CD3/CD8 beads (Supplementary Fig. 3a, b) and supernatant was used to stimulate human BMSCs, either the HS-5 cell line (Fig. 4j) or primary normal human BMSCs (Supplementary Fig. 3c), in vitro. This resulted in a significantly increased expression of NFKBIA, a canonical transcriptional indicator of NFĸB inflammatory activation, and decreased expression of KITLG (Fig. 4j, Supplementary Fig. 3c), recapitulating key transcriptional aspects of the inflammatory remodeling of BMSCs in LR-MDS.
To minimize the potential contribution of secreted factors from other cells than T-cells in the PBMC fraction to this effect, we also performed co-culture experiments with the secretome of purified and anti-CD3/CD8 bead-activated CD3 T-cells, which recapitulated the inflammatory activation of BMSCs (Supplementary Fig. 3d–f).
This purified T-cell secretome-induced inflammatory activation of HS-5 BMSCs was partially rescued by targeted knockdown of TNFRSF1A (Fig. 4k and Supplementary Fig. 3g), the canonical TNFα receptor highly expressed by LEPR+ stromal niches (Fig. 4a), confirming the notion that activated T-cells may drive inflammatory remodeling of stromal niches via TNFα signaling, although other (T-cell secreted) inflammatory factors likely play a role and ancillary contributions of other cell types cannot be excluded.
Inflammatory remodeling of stromal niches is associated with transcriptional signatures indicative of inflammatory stress in HSPCs in LR-MDS
To interrogate the potential impact of the inflammatory BMSC activation on hematopoiesis in LR-MDS, predictive intercellular communication analysis between BMSCs and HSPC subsets was performed, based on the expression of ligand and receptor complexes43 (Supplementary Fig. 4). This revealed a reduction in ligand-receptor signaling pathways governing HSPC retention, maintenance and lineage differentiation, including CXCL12-CXCR4 (for all HSPC subtypes), KITLG-KIT (for HSC/MPPs, LMPPs, GMPs and MEPs), FLT3LG-FLT3 (for LMPPs and GMPs) and IL7-IL7R (for CLPs) signaling (Supplementary Fig. 4). In contrast, JAG1-NOTCH1/NOTCH2 (for HSC/MPPs, LMPPs, GMPs and MEPs) and HGF-CD44 (for HSC/MPPs and LMPPs) signaling, which have been suggested to promote the initiation and progression of MDS21,44,45, were predicted to be enhanced in LR-MDS.
These predictions suggest that inflammatory BMSC activation in LR-MDS may contribute to the failure of normal hematopoiesis and leukemic progression. To interrogate this further, we re-clustered HSPC subsets (Fig. 5a) revealing a relative decrease in size of the HSC/MPP compartment within the HSPC cluster (Supplementary Fig. 5a). HSPC subsets (HSC/MPPs, LMPPs, GMPs, MEPs and CLPs) in LR-MDS demonstrated transcriptional activation of inflammatory stress signaling, as assessed by GSEA (TNFα signaling via NFĸB, Apoptosis and P53 pathway) (Fig. 5b) in both LR-MDS HSC/MPP subset and committed progenitors, including GMPs, MEPs and CLPs in comparison to NBM (Supplementary Fig. 5b). These signatures included ‘leading edge’ genes such as NFKBIA, RIPK2 (a potential activator of NFĸB and apoptosis), RELB (a NFκB subunit), GADD45A and B (responders to environmental stress), TXNIP (regulator of cellular redox signaling) and ATF3 (a stress responsive gene) (Fig. 5c and Supplementary Fig. 5c, Supplementary Data 2). The transcriptional activation linked to inflammatory stress signaling was accompanied by downregulation of gene signatures related to cell proliferation (G2M checkpoint and E2F targets) in LR-MDS HSPCs (Fig. 5b).
a Distribution of HSPC subpopulations in NBM and LR-MDS. b Enrichment of gene signatures indicative of NFĸB associated inflammatory activation and apoptotic stress and depletion of gene signatures indicative of cell-cycle progression in HSPCs (HSC/MPPs, LMPPs, MEPs and GMP clusters combined) in LR-MDS, as demonstrated by Hallmark gene set enrichment analysis. Statistical analysis was performed by a K-S test and adjusted for FDR. c Expression of “leading-edge” genes indicative of inflammatory stress in the overall HSPC population. NBM (n = 4) and LR-MDS (n = 5). Statistical analysis was performed by a two-sided Wald test and adjusted for FDR. PseudoDE R-package was used for differential expression gene analysis at sample levels. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers. d, e HSC/MPP transcriptional heterogeneity demonstrating relative expansion of the ‘high-output’ HSC-2 fraction in LR-MDS. NBM (n = 4) and LR-MDS (n = 5). Statistical analysis was performed by a two-sided unpaired Welch’s t-test. Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers (e). f, g Confirmation of HSPC scRNA-seq transcriptional data in larger LR-MDS dataset. CD45+CD34+ cells were sorted from LR-MDS (n = 38) or NBM (n = 7) and subjected to bulk RNA-seq. Transcriptional rewiring indicative of inflammatory stress/apoptosis and reduced cell cycle progression by differential gene expression analysis (f) and Hallmark geneset enrichment analysis (g). Statistical analysis was performed by a two-sided Wald test and corrected for FDR (f). Statistical analysis was performed by a K-S test, and corrected for FDR (g). h, i HSPC inflammatory stress signatures are linearly correlated with stromal niche inflammation in LR-MDS. Hallmark HSPC signatures are separated by level of stromal inflammation (iBMSC score low vs. high with median value as cutoff). Positive NES indicates programs enriched in iBMSC scorehigh patients, while negative scores indicate enrichment in iBMSC scorelow patients. Statistical analysis was performed by a K-S test and adjusted for FDR (h). Significant linear correlation between the iBMSC score and inflammatory-stress associated genes and gene programs in HSPCs of LR-MDS patients was assessed by a two-sided Spearman’s rank-order correlation analysis. To illustrate these correlations, a least-squares linear regression line with 95% confidence interval shading was fitted to the data (i).
We recently described the use of transcriptional signatures to discern “low-output” from “high-output” HSCs in the human bone marrow26 (Supplementary Fig. 5d, e), in analogy to such transcriptional subsets in mice46 that discern a quiescent HSC population from an HSC subset that actively contributes to hematopoiesis. The HSC compartment in LR-MDS, in comparison to the NBM (Fig. 5d), displayed a relative increase of the “high-output” HSC-2 fraction, with a concomitant relative decline of the HSC-1 subset (Fig. 5e).
To interrogate whether inflammatory activation of BMSCs in LR-MDS is associated with inflammatory signaling in HSPCs, we next performed RNA-seq on CD45+CD34+CD235a- HSPCs isolated from available LR-MDS samples (n = 38) from the HOVON89 trial, in which we performed bulk RNA-seq on sorted BMSCs (Supplementary Fig. 5f). Consistent with findings in the scRNA-seq dataset, LR-MDS HSPCs exhibited enrichment of genes and signatures indicative of (NFĸB-associated) inflammatory activation and cellular stress, with concomitant downregulation of gene signatures related to cell proliferation (G2M checkpoint; E2F targets) (Fig. 5f, g and Supplementary Data 3).
Of interest, comparison of HSPC gene expression in patients with a high iBMSC score (cutoff at median value) to those with a low iBMSC score revealed the enrichment of inflammatory and stress pathways in the iBMSChigh group, while HSPC gene expression from the iBMSClow group displayed relative preservation of metabolic (oxidative phosphorylation) and proliferation (E2F targets; G2M checkpoint) signatures (Fig. 5h). In line with this, spearman correlation analysis revealed a significant correlation between the iBMSC score and NFκB gene signature (TNFα signaling via NFĸB) and the HSPC expression of key genes contributing to this pathway (“leading edge”) inflammatory genes from the same patients (Fig. 5i), demonstrating that the level of BMSC inflammation is linearly correlated with the extent of HSPC inflammatory stress in the LR-MDS marrow.
Stromal inflammation induces inflammatory transcriptional rewiring of normal hematopoietic stem/progenitor cells and attenuates their functional capacity
The association between the inflammatory activation of BMSCs and HSPCs opens the possibility that stromal niche inflammation can actually induce inflammatory stress in normal HSPCs and impact their function. To elucidate this experimentally, NFκB-mediated inflammatory signaling was induced in BMSCs (murine OP9 cell line) by lentiviral overexpression of the constitutively active form of IKK2 (FLAG-IKK2SE) (Fig. 6a), a kinase upstream regulator of NFκB, recapitulating inflammatory alterations in BMSCs in LR-MDS, including overexpression of Nfkbia and Il-6 (Supplementary Fig. 6a), as previously described by us47.
a Experimental outline of the delineation of the effects of stromal inflammatory activation on the transcriptome and function of normal HSPCs in a murine system. Created in BioRender. Bian, Y. (2025) https://BioRender.com/x52bgzc. b Effect of stromal inflammation on the transcriptional wiring of normal HSPCs (Hallmark analysis), recapitulating findings in human LR-MDS. Positive NES indicates programs enriched in murine HSPCs cocultured with IKK2SE-transduced (inflamed) BMSCs, while negative scores indicate enrichment in HSPCs cocultured with EV-transduced (non-inflamed) BMSCs. Statistical analysis was performed by a K-S test and adjusted for FDR. c Definition of the Stromal Inflammation-induced Gene alterations in the HSPC Transcriptome (SIGHT) in LR-MDS comprising the set of genes that was differentially expressed in CD34+ HSPCs in LR-MDS in comparison to their counterparts in NBM and significantly dysregulated in HSPCs exposed to inflamed stromal niches ex vivo. d Implication of SIGHT in inflammatory signaling (MAPK, NFĸB and TNFα) and apoptosis by KEGG overrepresentation analysis. Statistical analysis was performed by a hypergeometric test and adjusted for FDR. e–h Attenuation of HSPC number and function by stromal inflammation. e Reduced numbers of HSPCs after co-culture on inflamed BMSCs (replicates from n = 2 mice performed in triplicates). Reduced repopulating ability of HSPCs, as demonstrated by reduced CFU-C forming ability (scale bar, 5 mm) (f) and in vivo competitive repopulating ability in PB (g) and in BM (h) (16 weeks after competitive transplantation; n = 5 mice/group). Statistical analysis was performed by a two-sided unpaired Welch’s t-test in (e, f, h); statistical analysis was performed by a two-way ANOVA with Sídák’s multiple testing correction in (g). Data are represented as mean ± SEM. Source data are provided as a Source Data file.
Co-culture of HSPCs (murine lineage- cells) on IKK2SE-transduced OP9 cells (in comparison to EV-transduced cells) resulted in upregulation of Sca-1 protein expression (Supplementary Fig. 6b), a well-documented consequence of inflammatory activation of HSPCs48, as well as distinct alterations in their transcriptional landscape (Supplementary Data 4), indicating activation of inflammatory programs and downregulation of cell-cycle related programs (Fig. 6b), recapitulating findings in CD34+ HSPCs in human LR-MDS (Fig. 5b,g).
Of all the differentially expressed transcripts in CD34+ HSPCs in human LR-MDS in comparison to CD34+ cells from the NBM, 14.6% (175/1200) was experimentally defined to be induced by stromal inflammation in the co-culture system (Fig. 6c and Supplementary Data 5), suggesting that stromal inflammation may be a significant driver of the transcriptional rewiring of HSPCs in LR-MDS. Consistent with this, a substantial fraction (41%) of these HSPC genes was significantly associated with the degree of stromal inflammation, as indicated by the iBMSC-score (Supplementary Data 6). We dubbed this 175-gene signature Stromal Inflammation-induced Gene alterations in the HSPC Transcriptome) (or SIGHT) and KEGG pathway overrepresentation analysis confirmed its implication in the MAPK, NFĸB, TNFα and apoptotic signaling pathways (Fig. 6d).
Next, we tested the effect of stromal inflammation on the functional capacity of HSPCs by assessing HSPC numbers, in vitro colony formation ability and in vivo competitive repopulating ability (Fig. 6a). Exposure of normal HSPCs to inflammatory BMSCs resulted in a significant reduction of their numbers after a 5-day co-culture period (Fig. 6e), as well as impairment of their repopulating ability, as indicated by a reduced in vitro colony (CFU-C) formation ability (Fig. 6f) and in vivo competitive repopulation ability upon transplantation in mice (Fig. 6g, h).
Collectively, the data demonstrate that inflammatory activation of stromal niches induces inflammatory transcriptional remodeling of HSPCs in human LR-MDS and results in the impairment of their number and repopulation ability.
Clonal hematopoietic stem cells display resistance against stromal niche induced inflammatory stress and reside predominantly in a transcriptional ‘high-output’ state
MDS pathogenesis is characterized by clonal selection and/or expansion, but the underlying mechanisms remain to be fully elucidated, since mutated cells, e.g., SF3B1-mutated HSPCs, do not always have a cell-intrinsic competitive advantage over non-mutated HSPCs. Resistance to inflammatory stress has been dubbed as one potential mechanism of clonal selection34,49 in model systems, but its relevance for human LR-MDS has remained incompletely understood.
To test whether the stroma-induced inflammatory stress has differential consequences for the transcriptional wiring and predicted output of SF3B1 mutant versus residual normal HSPCs, we next compared the transcriptomes of SF3B1 mutated HSPCs to their non-mutant, residual normal counterparts. To this end, mutant cells in the single-cell LR-MDS hierarchy were identified by three different, complementary approaches: i. an in-house developed mutation calling algorithm based on 10X RNA-seq data26, ii. a novel biocomputational method to detect transcriptional consequences of aberrant mRNA splicing (SpliceUp) and iii. Rapid Capture Hybrid sequencing (RaCHseq), a targeted next-generation sequencing method that use long, biotinylated oligonucleotide baits to capture full-length cDNA molecules for long read (nanopore) DNA sequencing50.
These three methods yielded highly consistent results regarding the frequency of mutant cells in distinct cellular subsets in the LR-MDS marrow (Fig. 7a and Supplementary Fig. 7a, b), with mutant cells being (nearly) absent in CD4/CD8 T-cells, plasma cells and non-hematopoietic stromal and endothelial fractions, but robustly represented in all HSPC subsets and more differentiated CD14+/CD16+ myeloid cells (Fig. 7a and Supplementary Fig. 7b), in line with the notion that bone marrow niche cells are of non-mutant origin in LR-MDS. Notably, small subsets of mutant cells were found in the NK and pre-B-cell population, in line with previous reports51,52.
a Identification and distribution of SF3B1-mutated cells within the LR-MDS hierarchy. UMAP distribution (left panel) and population-specific frequencies (right panel) of mutant cells. Mutant cells are identified by combining 10X scRNA-seq, RaCHseq and SpliceUp computational analyses. b Distribution of mutant cells within HSPC subsets (left panel) and differentially expressed gene (Hallmark) programs in mutant cells (vs. ‘non-assignable’ cells) (right panel), revealing mitigation of inflammatory stress pathways and enrichment of gene programs implicated in cell cycle progression in mutant cells. Positive NES reflects programs enriched in mutant HSPCs, while negative scores indicate enrichment in non-assignable HSPCs. All cells in HSC/MPPs, LMPPs, MEPs, and GMP clusters were used in the analysis. Statistical analysis was performed by a K-S test and adjusted for FDR. c, d Distribution of mutant cells within HSC subsets. c UMAP and d distribution per sample indicating relative abundance of Mutant cells in the ‘high-output’ (HSC-2) subset in comparison to the residual normal (‘non-assignable’) subset. LR-MDS (n = 4). Data are presented as boxplots: the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range. Data points beyond this range are shown individually as outliers (d). e HSPC numbers upon co-culture on inflamed (IKK2SE) stromal cells (relative to EV-transduced BMSCs), demonstrating a relative preservation of SF3B1-mutant immunophenotypic HSPCs (CD34+ and CD34+CD38–CD45RA–CD90+). Data represents replicates from NBM (n = 2) and SF3B1-mutated LR-MDS (n = 3), experiments performed in dupli/triplicates for each bone marrow sample. Statistical analysis was performed by a two-sided unpaired Welch’s t-test. Data are presented as mean ± SEM. Source data are provided as a Source Data file.
The three methods resulted in the composite identification of mutant cells in up to 20–25% of hematopoietic cells (in patients with a median VAF of SF3B1 of 40%, range 22–57%) (Fig. 7a and Supplementary Data 1), indicating that a substantial fraction of mutant cells is captured by this approach. Nevertheless, and of note, the “non-assignable” population is expected to still harbor mutated cells, not detected by any of these approaches (due to insufficient coverage at the locus of mutation), thus reducing the sensitivity to detect transcriptional differences between mutant and “non-mutant” cells in this analysis, and likely resulting in the underestimation of the real differences between mutant and non-mutant cells in the LR-MDS marrow.
Mapping of mutant cells within the HSPC cluster (Fig. 7b) revealed that residual normal (“non-assignable”) cells differentially expressed transcripts associated with inflammatory activation (Supplementary Data 7) which was reflected in significant enrichment of hallmark transcriptional signatures indicative of inflammatory activation (TNFα signaling via NFĸB; Interferon gamma response, IL2-STAT5 signaling; IL6-JAK signaling; Inflammatory response). In contrast, HSPCs carrying the SF3B1 mutation differentially expressed many genes associated with cellular activation, proliferation and cell cycle progression, reflected in signatures indicative of cell proliferation (G2M Check Point, E2F Target and mitotic spindle) and oxidative phosphorylation (Fig. 7b). These differences were found to be highly consistent between the three methods to detect mutant cells and found both in the HSC/MPP/LMPP subset and their downstream MEP progeny (Supplementary Fig. 7c, d).
Mutant cells could be detected in the HSC low-output fraction (Fig. 7c), in line with the notion that dormant HSCs are the disease-initiating cells in SF3B1-mutated MDS52, but the majority of mutant cells within the HSC fraction was present in the high-output HSC-2 fraction (Fig. 7c, d), suggesting active contribution to downstream progeny, consistent with their transcriptional wiring indicative of active cycling. In contrast, residual normal HSCs resided predominantly in the low-output HSC subset (Fig. 7d), consistent with the notion of the relative competitive advantage of mutant cells in the LR-MDS hierarchy.
To further address this notion experimentally, we performed co-culture experiments as described in Fig. 6, using SF3B1-mutated HSPCs (CD34+ cells from SF3B1-mutated LR-MDS bone marrow samples with a high VAF (range 39–42%; n = 3), indicating that the vast majority of HSPCs carried the mutation, in comparison to CD34+ cells from NBM (n = 2). Exposure to the IKK2SE-inflamed stromal cells reduced the absolute number of both normal (consistent with murine normal HSPCs; Fig. 6e) and mutant HSPCs, but to a significantly lesser degree in the mutant CD34+CD38-CD45RA-CD90+ immunophenotypic HSC-enriched fraction (Fig. 7e and Supplementary Fig. 7e), supporting the notion of relative resistance against stromal-inflammation-induced suppressive stress.
Collectively, the data support a model in which inflammatory activation of stromal HSPC niches acts as a driver of clonal selection and expansion through the selective repression of residual normal HSPCs, with relative resistance to this repressive signaling of mutated clones providing a relative competitive advantage of mutant cells in the system.
Inflammatory remodeling of stromal niches is associated with leukemic progression and reduced survival in MDS
The proposed relationship between stromal inflammation, bone marrow failure and clonal selection would predict that the level of stromal inflammation relates to outcome, and in particular clonal evolution and leukemic progression, in LR-MDS. To investigate this, we correlated the iBMSC score to overall survival (OS) in our LR-MDS cohort (n = 45) of uniformly treated patients within the HOVON89 clinical trial. Comparison of clinical outcome between iBMSChigh and iBMSClow patients (cutoff at median value) shows a significant correlation between BMSC inflammation and OS (Fig. 8a). Patients with a higher iBMSC score had a significantly shorter survival (median OS 40.7 months vs. 54.6 months, p = 0.037).
a Reduced overall survival (OS) in patients with high levels of stromal inflammation in the HOVON89 “training” cohort as reflected in the iBMSC-score (cutoff median value). Statistical significance was determined by the log-rank test. b–d Generation of a HSPC-SIGHT prognostic score (SIGHT-P) (by using penalized cross-validation with SIGHT as input) to identify a HSPC gene-score comprising 9 genes that linearly correlated with the level of stromal inflammation (c) and outcome (OS) (d) in the ‘training’ set. A linear regression line (with 95%-confidence interval shading) was fitted to the data using least squares to illustrate the correlation between SIGHT-P with stromal inflammation. Two-sided Spearman’s rank-order correlation was used for correlation analysis (c). OS analysis was performed by the log-rank test, patients stratified based on the median SIGHT-P value as cutoff (d). Validation of the prognostic value of the SIGHT-P score in an independent cohort of MDS patients (GSE58831) demonstrating significantly reduced OS (e) and increased leukemic evolution (f) in patients with a high SIGHT-P score (reflecting stromal inflammation). OS analysis was performed by the log-rank test (e). Leukemic evolution was evaluated with a competing risk analysis (competing risk: non-leukemia mortality) and statistical significance determined by a two-sided Gray’s test (f).
Our stromal transcriptional data from a well-annotated cohort of MDS patients is unique and therefore precludes the direct validation of this finding in an independent MDS patient cohort. To study the impact of stromal inflammation on outcome in an independent MDS patient cohort, we therefore exploited our identification of the SIGHT genes in LR-MDS. Using a machine learning method (Lasso regression) with the SIGHT genes as input, a 9-gene gene signature was constructed (named SIGHT-P (prognostic)) using cross-validated penalized regression. SIGHT-P accurately reflected the extent of BMSC inflammation as indicated by Spearman correlation with iBMSC scores (rho=0.72, padj < 0.0001) (Fig. 8b, c). Interestingly, leading genes within this signature (RNF217, JUND and DNAJC2) comprise genes implicated in apoptotic regulation53,54,55,56,57, inflammation58 and cell cycle progression59,60. As expected, given the correlation with the iBMSC-score, patients with a higher SIGHT-P score (cut-off at median value) had significantly reduced survival probability in our training cohort (median OS 25.7 months in SIGHT-Phigh vs. 62.3 months in SIGHT-Plow, p = 0.036 (Fig. 8d)).
The identification of an HSPC signature reflective of stromal inflammation allowed us to validate the prognostic significance of stromal inflammation in a larger cohort of MDS patients of which gene expression of CD34+ HSPCs was available, with broad representation of risk categories and higher rates of leukemic evolution (accession: GSE58831)61. The SIGHT-P score in this cohort was associated with a significantly reduced OS (median OS 36.8 months vs. 84.3 months in the SIGHT-Phigh vs. SIGHT-Plow groups, respectively, p = 0.023) (Fig. 8e), confirming findings in our LR-MDS cohort. Strikingly, patients with a high SIGHT-P score were much more likely to develop AML in comparison to patients with a low SIGHT-P score (5-year AML incidence 33.17% vs. 6.5%, p = 0.0074 (Fig. 8f). A significant correlation was found between the SIGHT-P score and the IPSS score (Supplementary Fig. 8a), but the SIGHT-P score remained of independent prognostic value in LR-MDS patients as indicated by multivariable competing risk analysis (Supplementary Fig. 8b, c).
Collectively, the data reveal a significant correlation between the inflammatory activation of stromal niches and the resulting transcriptional rewiring of HSPCs with leukemic evolution and survival in MDS, in support of the view that stromal inflammation is a driver of clonal evolution and leukemic progression in this disease.
Discussion
Tumor-promoting inflammation is considered a cancer-enabling characteristic facilitating initiation, proliferation, and immune evasion of neoplastic cells62. In the hematopoietic system, inflammation has been associated with aging, clonal hematopoiesis and cytopenia63,64. However, the molecular and cellular mechanisms by which inflammation promotes clonal evolution and tumorigenesis, and the relevance of the concept for human disease, remain incompletely understood.
Here, by establishing a comprehensive single-cell transcriptional taxonomy, we relate stromal inflammation to inflammatory stress in HSPCs, leukemic progression and clinical outcome in human LR-MDS, consistent with a model in which inflammatory activation of stem/progenitor cell niches contributes to clonal expansion and leukemic evolution by repressing normal hematopoiesis with relative resistance to inflammatory signaling of mutant cells.
The data indicate that stromal inflammation may contribute to LR-MDS pathogenesis via distinct mechanisms. Targeted depletion of HSPC-regulatory factors from LEPR+ BMSCs causes HSPC repression and bone marrow failure in mouse models14, suggesting that the significant downregulation of expression of HSPC-supportive factors in the inflamed LEPR+ BMSCs in human LR-MDS may contribute to bone marrow failure. Additionally, actors from the inflammatory stromal population (BMSC-2), such as IL-6 and JAG-1, may directly promote progression and leukemic transformation of mutant cells21,65, while repressing residual normal HSPCs. Pro-inflammatory chemokines such as CXCL3, CXCL5, CXCL8 and CCL2 from inflamed BMSCs may further enable recruitment of other pro-inflammatory immune cells to HSPC niches66,67, potentially exaggerating inflammatory stress in the HSPCs.
These mechanisms are likely to differentially affect mutant vs. residual normal HSPC subsets, which is supported by our finding that transcriptional signatures indicative of inflammatory stress and apoptosis were significantly more enriched in the HSPC subset containing non-mutant cells, in comparison to their mutant counterparts and co-culture data showing relative resistance of SF3B1-mutated HSPCs against stromal-inflammation induced repression. This finding may be consistent with, and provide human-disease relevance to, the emerging notion from experimental models that mutant cells may display relative resistance to inflammatory stress as a mechanism of competitive advantage and clonal expansion34,49,68,69,70
It is important to re-emphasize here that the LR-MDS cellular taxonomy does not include all niche cells that constitute the human bone marrow microenvironment. Specifically, endothelial cells and stromal niche subsets with close anatomic relationship to the (trabecular) bone are likely absent or underrepresented in human BM aspirates26 and predicted contributions of these cells to the disruption of hematopoiesis in MDS cannot be inferred based on this data.
The current work, in which we were able to examine the transcriptional wiring of both BMSCs and HSPCs from the same samples, reveals an intricate network of inflammatory signaling that comprises both HSPCs and their stromal niches. The data provide novel insights into the “chicken or egg” question, asking whether stromal inflammation is cause or consequence of inflammation-associated bone marrow failure and clonal evolution, demonstrating that stromal inflammation may be caused by hematopoietic elements (in particular T-cell subsets), but is a direct cause of the transcriptional inflammatory activation and functional decline of residual normal HSPCs.
Of interest, the stromal inflammation was not unique to LR-MDS but was also observed in clonal hematopoiesis, implying stromal inflammation may serve as a driving force of the disease at a very early stage71.
Our finding that the inflammatory imprint of stromal inflammation on hematopoietic elements increases with progression of the disease (reflected by the IPSS score) may imply that not necessarily the nature/source of inflammatory signaling, but its degree, discerns low from high-risk MDS, although we did not examine (stromal) inflammatory signaling specifically in the latter category.
Immune stimulation and activation of T-cells have previously been associated with stromal remodeling, neutrophil extracellular trap formation and progression of NPM1-driven myeloproliferation72. The network analyzes performed in our study suggest a potential direct contribution of activated immune cells, in particular pro-inflammatory (IFNG+ and GZMK+) EM CD8+ T cells, to stromal inflammation. This subset of EM CD8+ T cells in LR-MDS had the highest expression of the pro-inflammatory cytokines IFNγ and TNFα and is predicted to induce inflammatory activation of LEPR+ stromal niches given the differential expression of the canonical receptors for these cytokines in BMSCs. Interestingly, pro-inflammatory GZMK+CD8+ T cells have recently been identified as major, tissue-resident, cytokine producers with low cytotoxic potential that drive inflammation in patients with inflammatory diseases such as rheumatoid arthritis39. It may be hypothesized, based on our data, that immune activation in LR-MDS does not effectively result in expansion of effector CD8+ T cells and upregulation of cytolytic factors (e.g Granzyme B and Perforin-1 (Supplementary Fig. 2e)), resulting in insufficient constraint of clonal progression via elimination of target cells, yet does result in activation of EM CD8+ T cells, contributing to bone marrow failure. This concept, however, awaits experimental definition, including its consideration as a potential mode of action of immune-suppressive drugs like anti-thymocyte globulin (ATG) and cyclosporine73,74 in LR-MDS.
Although they expressed the highest levels of TNFα and IFNγ transcripts in the LR-MDS taxonomy, activated CD8 T-cells are unlikely to be the only cellular source of stromal inflammation in this disease. Other innate/adaptive immune cells, including NK cells that express TNFα and IFNγ, may contribute to stromal inflammatory remodeling. It has further been established that clonal abnormalities in hematopoietic cells can confer exaggerated inflammatory responses to downstream progenitors and myeloid cells32,33,34,75 and that these hematopoietic cells can confer inflammatory stimuli to stromal cells76, consistent with a model in which mutant hematopoietic cells confer inflammatory signaling to their stromal microenvironment and stromal inflammation, in turn, may contribute to bone marrow failure and leukemic evolution.
Stromal niches are dominant and central regulators of HSPC maintenance and proliferation, through close anatomical relationship with HSPCs and well-established molecular interactions. This makes inflammatory remodeling of these cells, including the associated downregulation of key HSPC regulatory factors such as KITLG, of distinct relevance to HSPC behavior. It cannot be excluded, however, that other cells in the LR-MDS taxonomy, including T and NK cell subsets, contribute to the inflammatory activation of HSPCs in LR-MDS in a direct fashion.
Taken together, the comprehensive single cell atlas of human LR-MDS, relating stromal inflammation to hematopoietic stress, leukemic progression and clinical outcome is anticipated to instruct novel approaches for disease prognostication and therapeutic targeting and constitute a valuable resource to provide human disease relevance to findings in diverse experimental systems.
Methods
Human samples
LR-MDS BM aspirates used for RNA-seq were obtained from newly diagnosed patients (age 68–79, median 72) with LR-MDS (IPSS low and intermediate-1) included in the HOVON89 clinical trial (www.hovon.nl; www.trialregister.nl as NTR1825; EudraCT No. 2008-002195-10) assessing the efficacy of lenalidomide with or without erythropoietin and granulocyte-colony stimulating factor in patients. Age-matched normal BM samples were obtained by from healthy donors for allogeneic transplantation. All specimens were collected with informed consent. The study was approved by the ethics committees of the participating institutions (Central ethics committee (MREC, Medical Research Ethics Committee): METC Amsterdam UMC (central review in the Netherlands)) and was conducted in accordance with the Declaration of Helsinki.
SF3B1-mutated LR-MDS BM aspirates with a high VAF (range 39–42%), aged matched normal BM samples used for stroma coculture experiments, and LR-MDS BM biopsies used Immunofluorescence staining were approved by the ethics research committee at Karolinska Institute (2010/427-31/3, 2017/1090-31/4), collected with written informed consent for research use, in accordance with the Declaration of Helsinki.
Peripheral blood (PB) samples were obtained from healthy donors, all specimens were collected with informed consent for research use.
Patient cell isolation for RNA-seq
Viably frozen BM aspirates (mononuclear cell fractions) were thawed in a water bath at 37 °C and washed with warm Dulbecco’s Modified Eagle Medium (DMEM, Gibco) supplemented with 10% Fetal calf serum (FCS). BM cell fractions were isolated as reported before26. For isolation of the hematopoietic fraction, 10% of the thawed BM cells (>20 × 106) from each individual were stained for sorting in PBS supplemented with 0.5% FCS at 4 °C with the following antibodies: CD45-APC (1:20, clone: 2D1, eBioscience), CD34-AF700 (1:50, clone: 581, BioLegend), CD117-PE-CF594 (1:50, clone: YB5.B8, BD Biosciences), CD33-PE (1:50, clone: P67.6, BD Biosciences), CD3-PE-Cy7 (1:50, clone: SK7, BioLegend), CD19-APC-Cy7 (1:50, clone: HIB19, BioLegend), and CD38-FITC (1:50, clone: HIT2, Life Technologies). For the exclusion of dead cells, 7AAD (1:100, Beckman Coulter) was used. The 7AAD−CD45+CD34+ HSPC fraction, 7AAD−CD45+CD34−CD117+ and 7AAD−CD45+CD34−CD33+ myeloid fraction, and 7AAD−CD45+CD34−CD117−CD33− non-myeloid (lymphoid) fraction were sorted in DMEM supplemented with 10% FCS using a FACSAria III (BD Biosciences) and BD FACSDiva version 5.0.
For isolation of the non-hematopoietic fraction, 90% of the thawed cells (>200 × 106) were stained with biotinylated antibodies against CD45 (1:50, clone: HI30, BioLegend) and CD235a (1:50, clone: HIR2, BioLegend) followed by depletion using magnetic anti-biotin beads (20 μL per 107 cells; Miltenyi Biotec) in PBS supplemented with 2% FCS and iMag (BD Biosciences). After depletion, the remaining cells were stained for sorting in PBS containing 0.5% FCS at 4 °C with the following antibodies: streptavidin-AF488 (1:100, Invitrogen), CD45-BV510 (1:50, clone HI30, BioLegend), CD235a-PE-Cy7 (1:50, clone: HI264, BioLegend), CD71-AF700 (1:20, clone: MEM-75, Exbio), CD271-PE (1:50, clone: ME20.4, BioLegend), CD105-APC (1:50, clone: SN6, eBioscience), CD31-APC-Cy7 (1:20, clone: WM59, BioLegend), CD34-eFluor610 (1:50, clone: 4H11, eBioscience), and CD144-V450 (1:50, clone: 55-7H1, BD Biosciences). 7AAD (1:100, Beckman Coulter) was used for dead cell exclusion. FACSAria III (BD Biosciences) and BD FACSDiva version 5.0 were applied for sorting.
For bulk RNA-seq, 7AAD−CD45+Lin–CD235a−CD34+ cells were sorted in TRIzol (Life Technologies) using antibodies described above.
scRNA-seq
Prior to scRNA-seq, BM cell fractions sorted separately from the same donors were pooled into two fractions (non-hematopoietic fraction with lymphoid fraction, and myeloid fraction with HSPC fraction), followed by encapsulation for barcoding and cDNA synthesis using the Chromium Next GEM Single-cell 3′ Reagent kit v3.1 (10X Genomics). 3′ gene-expression library was constructed according to the manufacturer’s recommendations. The quality and quantity of libraries were determined using an Agilent 2100 Bio-analyzer with 2100 Expert version B.02.11.SI811 software and a High-Sensitivity DNA kit. Libraries were sequenced on a NovaSeq 6000 platform (Illumina), paired-end mode, at a sequencing depth of around 45,000 reads per cell, followed by computational alignment using CellRanger (version 3.0.2, 10X Genomics).
Seurat (R package, version 4.0.0)77 was used for data preprocessing and downstream analysis. For data preprocessing, datasets were first subjected to quality control steps that included removing doublets (a high ratio of RNA counts vs. feature numbers; >6) and filtering out apoptotic cells determined by the high transcriptional output of mitochondrial genes (>5% of total). Subsequently, single-cell data from separate runs of the same donor were merged to generate a complete picture that includes all cell types. In order to generate a single cell atlas robustly representing the BM taxonomy in healthy individuals and LR-MDS, the merged datasets of each LR-MDS patient were first integrated using the canonical correlation analysis (CCA)-based integration function in Seurat, and further integrated with a NBM dataset simultaneously generated using the same method26. This was followed by a linear dimensional reduction procedure, including scaling of gene expression across all cells, principal component analysis on the most variable genes (k = 2000) and unsupervised clustering using a shared nearest-neighbor modularity optimization-based clustering algorithm (resolution: 0.3–1). To mitigate the effects of cell-cycle heterogeneity, cell-cycle phase scores were calculated based on canonical markers and were regressed out as described in the Seurat protocol. The data were visualized using uniform manifold approximation and projection for dimension reduction (UMAP)78. Cell types were identified by the Clustifyr R package79 and by measuring expression of canonical markers of particular cell types. Aiming at revealing BMSC heterogeneity at a high resolution, BMSCs were separated using the CellSelector function in Seurat and independently preprocessed/analyzed.
Intercellular communications were predicted based on expression of ligand and receptor isoforms using Cellchat and CellphoneDB43,80. For differential expression gene analysis between samples, a DEseq281-based pseuDE package was developed for the aggregation of single-cell level gene counts and application of differential expression (DE) analysis (software is available at https://github.com/weversMJW/pseude). Differentially expressed genes were considered present when the FDR-corrected p-value was ≤0.05. For GSEA82 between groups, variable genes were identified using Seurat’s Findmarker function and ranked using the formula −sign(log2foldchange) × log10(P-value). The R package fgsea was used for the analysis with a permutation of 10,000 using predefined gene sets from the Molecular Signatures Database (MSigDB 6.2) as input. Gene enrichment scores for individual cells were calculated using Seurat’s AddModuleScore function, which calculates the score by counting the average expression levels of provided genes, subtracted by the aggregated expression of randomly selected control genes.
Identification of SF3B1 mutant cells by 10x scRNA-seq
Identification of cells carrying mutations in SF3B1 was performed using an in-house–developed tool. In short, alignment results produced by the Cell Ranger pipeline in the form of a BAM file and a list of accepted cellular barcodes (CB) were used as an input. Aligned reads, representing sequenced cDNA molecules, were extracted at the mutation position and screened for the known SF3B1 mutations taking into account the cellular barcode and unique molecular identifier (UMI), irrespective of whether the cell is of hematopoietic origin. Following UMI-based consensus sequence building, mutant-carrying reads were assigned to the cell of origin based on the CB.
For each read overlapping the position of the mutation of interest, a reference sequence and a modified mutant sequence were constructed. Both sequences are constructed taking the local splicing structure into account. Local optimal alignment of the read of interest was performed on both the reference sequence and on the modified mutant sequence containing the known mutant. The read is classified as wild-type or mutant depending on the highest alignment score.
Cells with at least one mutant read detected were classified as mutants and those with no mutation detected were classified as “non-assignable (NA)”. Software is available at https://github.com/RemcoHoogenboezem/annotate_bam_statistics_sc.
Identification of SF3B1 mutant cells by RaCH-sequencing
RaCH-seq was performed as previously described in detail83. In short, all known isoforms for the genes of interest were selected from the GENCODE v40 transcript database based on GRCh38. Genes of interest included in the custom panel are: ASXL1, BCOR, BCORL1, CEBPA, DNMT3A, FLT3, IDH1, IDH2, JAK2, KIT, KRAS, NRAS, PTPN11, RUNX1, SF3B1, SRSF2, STAG2, TET2, TP53, U2AF1, WT1, ZRSR2. Next, we used a modified version of the RaCH-seq R script for designing panels (https://github.com/HongkePn/RaCHseq_Probe_Design) to extract transcript sequences of the genes of interest. In brief, capture probes are 120 nucleotides (nts) in length. For mid-sized exons, >120 and <1000 nts, we included the complete exonic sequence for capture. For large-sized exons, >1000 nts, we aimed to have at least a single capture probe per kilobase (kb), therefore large-sized exons were dissected into 1 kb or smaller portions and used as distinct sequences for capture. For small-sized exons, ≤120 nts, depending on the isoform, we concatenated the up- or downstream exon to the current exon to generate a capture sequence larger than 120 nts. All sequences extracted for capture were submitted to the IDT xGen Hyb Panel Design Tool as separate entries for probe design. The subsequent results were downloaded from the IDT website and further filtered. Since numerous exons are shared between distinct isoforms of a gene, multiple probes selected by the IDT xGen Hyb Panel Design Tool are the exact same. These were deduplicated, leaving only a single probe. Furthermore, probes with a GC-content closer to average GC-content over all probes were preferred. While this procedure removes probes with the exact same sequence and those with either too low or too high GC-content, it still results in the retention of probes being merely a single-based shift of the other. We next aligned the retained probes against GRCh38 and the GENCODE v40 transcript database using STAR 2.7.584 with default settings, with the exception of –outSAMmultNmax 1. Probes aligning outside the genes of interest or an alignment score ≤5 were discarded using the SAMtools API85. Finally, we developed an in-house algorithm that discards the lower-scoring probe (according to IDT) when: (I) it overlaps with another probe for ≥70%, (II) both probes measure the same transcript isoforms. A probe was selected at random in case the IDT scores are the same and all the above criteria hold. Ultimately, the final probe selection was submitted to be designed as a custom IDT xGen MRD Hybridization panel.
10 μL of full-length 10x Genomics cDNA library was used as input for full-length amplification using the primers 10× Partial R1: CTACACGACGCTCTTCCGATCT and T5′ PCR Primer IIA: AAGCAGTGGTATCAACGCAGAG. AMPure XP bead-based reagent (Beckman Coulter) to each 1–2 μg of amplified cDNA was added in a 1.8× bead/sample ratio for cleanup. The bead pellet was washed twice with 500 μL EtOH and air dried. To the dried sample, 8.5 μl 2× Hybridization Buffer, 2.7 μl Hybridization Buffer Enhancer (xGen Hybridization and Wash Kit, IDT, cat#1080577), 1 μl of each Blocking Oligo (1000 μM, xGen Integrated DNA Technologies [IDT]): PolyA: TTTTTTTTTTTTTTTTTTTTTTTTTTTTTT/3InvdT/, PR1: CTACACGCGCTCTTCCGATCT and SO:AAGCAGTGGTATCAACGCAGAGTAC were added. After resuspending, this elution mix was transferred to a magnet. Hereafter, 13 μL eluted sample was transferred to a new clean PCR strip and heated at 95 °C for 10 min to denature the cDNA. Custom Lockdown Probe Panel (4 μl; xGen IDT) was added to the sample and hybridized to the cDNA at 65 °C for 16 h with the heated lid set to 75 °C.
The probe pulldown with streptavidin beads was performed using xGen Lockdown Hybridization and Wash Kit (IDT). The captured sample and beads were resuspended in 50 μl EB Buffer (Qiagen) and amplified with the LA Taq DNA Polymerase Hot-Start (Takara) using the primers:
FPSfilA: 5′-ACTAAAGGCCATTACGGCCTACACGACGCTCTTCCGATCT,
RPSfilBr: 5′-TTACAGGCCGTAATGGCCAAGCAGTGGTATCAACGCAGAGTA.
Full-length cDNA libraries were barcoded with the PCR-cDNA barcoding kit 24 (Nanopore Technologies, cat #SQK-PCB111.24). Pools of 5 barcoded captured samples were sequenced on PromethION flow cells R9.4.1 (Nanopore Technologies, cat#FLO-PRO002). Samples were sequenced for 42–72 h on the PromethION P24, and basecalling was performed in parallel using Dorado 0.8.2 with a minimum basecalling set at 9.
The raw FASTQ files were used as input for an adapted version of the FLAMES R pipeline86. Due to the higher sequencing and basecalling error rate of Nanopore sequencing, we made the following adjustments to the FLAMES pipeline. The number of allowed errors present in the UMI-sequence before it was recognized as separate molecules is set to 2. As a consequence, more molecules were therefore recognized as potential PCR duplicates resulting in the filtering of more reads. We additionally tweaked the FLAMES pipeline to achieve higher throughput by increasing the number of threads used for sorting BAM files with the SAMtools API. After adjustment, we aligned the Nanopore raw FASTQ files against GRCh38, using the GENCODE v40 transcript database GTF file, list of 10× filtered cellular barcodes as input. Ultimately, in the first pass, this resulted in a UMI-deduplicated FASTQ file. This filtered FASTQ file was used as input in the second pass of the FLAMES pipeline resulting in an aligned BAM file containing UMI-deduplicated reads. These reads were annotated by an in-house algorithm to add the cellular barcode and UMI as SAMtools tags to reads, resulting in an annotated BAM file. This annotated BAM file was used as input into AnnotateBamStatistics for single cells, to extract mutation details per cell: https://github.com/RemcoHoogenboezem/annotate_bam_statistics_sc.
Using the extract cellular barcodes, mutation information was superimposed on top of the single cells analyzed using the standard 10× genomics workflow to learn which cells carried the mutation of interest.
Single-cell prediction of SF3B1 mutant status via mis-splicing event quantification (SpliceUp)
In parallel with the direct identification of mutant SF3B1 cells, SF3B1 mutation status was predicted through single-cell quantification of cryptic 3′ alternative splice site (A3SS) usage via the SpliceUp biocomputational tool87 to amplify the detection of mutant cells and further increase differential expression analysis power. Percent-splice-in (PSI) values were quantified on a per-cell basis for each of 36 signature SF3B1 mutant A3SS events88 and used as predictor variables in a multiple logistic regression model with SF3B1 mutant sample status as the response variable. Single-cell SF3B1 mutant status was predicted at a significance threshold of 0.01.
Bulk RNA-sequencing
RNA was extracted using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions, in combination of isolation with GenElute LPA (Sigma-Aldrich). cDNA was prepared using the SMARTer procedure through the SMART-Seq v4 Ultra Low Input RNA Kit for Illumina Sequencing (Takara Bio). Quantity and quality of cDNA production were assessed using the Agilent 2100 Bio-analyzer and the High Sensitivity DNA kit. cDNA libraries were generated using the TruSeq Sample Preparation v2 guide (Illumina) and paired-end sequencing on a NovaSeq 6000 (Illumina). Adaptor sequences and polyT tails were trimmed from unprocessed reads using fqtrim version 0.9.7. (http://ccb.jhu.edu/software/fqtrim/). Read counts and transcripts per million (TPM) were determined with Salmon version 1.2.189. Gene count estimates were determined from Salmon output with the tximport R package version 1.1690. These gene count estimates were in turn normalized, prefiltered according to standard practices, and used for determining the differential gene expression between groups of interest through the DESeq2 package (version 1.28)81 using default parameters. GSEA was performed with GSEA software (version 3.0, Broad Institute) using predefined gene sets from the Molecular Signatures Database (MSigDB 6.2). Gene lists were ranked on the basis of the log2FC made available through the DESeq2 package. Classic enrichment statistics with 1000 permutations were used to determine significant enrichment within gene sets. GSVA program (R package) was applied to calculate the iBMSC score in single samples using genes significantly upregulated (padj < 0.05) in BMSC-2 vs. other BMSC clusters in LR-MDS. Deconvolution of the bulk-RNAseq dataset was performed using CibersortX algorithms in S mode, with scRNA-seq data on BMSCs serving as inputs91.
Flow cytometry analysis for mouse and human HSPCs and T-cells
For flow cytometry analysis of mouse Lin– co-cultured with or without IKK2SE inflamed BMSC (OP9 (ATCC, CRL-2749)): Lin– cells were stained with following antibodies: lineage cocktail containing biotinylated antibodies against-Biotin-BV510 (B220 Biotin, 1:25, clone: RA3-6B2, BD Biosciences; CD3e Biotin, 1:25, clone: 145-2C11, BD Biosciences; CD4 Biotin, 1:25, clone: GK1.5, BD Biosciences; CD8a Biotin, 1:25, clone: 53-6.7, BD Biosciences; Ly-6G Ly-6C/Gr1 Biotin, 1:25, clone: RB6-8C5, BD Biosciences; CD11b/Mac1 Biotin, 1:25, clone: M1/70, BD Biosciences; TER-119 Biotin, 1:25, clone: TER-119, BD Biosciences), Sca-1-BV421 (1:100, clone: D7, BioLegend), c-Kit-PE-TR(1:100, clone: 2B8, BD Biosciences), CD48-AF700 (1:100, clone: HM48-1, BioLegend), CD150-PE (1:100, clone: TC15-12F12.2, BioLegend), CD45.1-APC (1:100, clone: A20, Sony), CD45.2-PE-Cy7 (1:50, clone: 104, BioLegend); Streptavidin Pacific Orange (1:200, Life Technologies), 7AAD-Percpcy5.5 (1:100, Beckman Coulter). Samples were measured using the FACS Symphony A5 Cell Analyzer and analyzed with a FlowJo_v10.6.1 program.
For flow cytometry analysis of mouse PBMCs: mouse PBMCs were stained with following antibodies: CD45.1-APC-Cy7 (1:100, clone: A20, BioLegend), Gr-1-APC-R700 (1:100, clone: RB6-8C5, BioLegend), CD11b-PE-Cy7 (1:100, clone: M1/70, BioLegend), CD115-APC (1:100, clone: AFS98, BioLegend), B220-BV421 (1:100, clone: RA3-6B2, BioLegend), CD3e-BV510 (1:25, clone: 145-2C1, BD Biosciences), CD45.2-FITC (1:100, clone:104, BioLegend), 7AAD-Percp-Cy5.5 (1:100, Beckman Coulter). Samples were measured using FACS Symphony A5 Cell Analyzer and analyzed with a FlowJo_v10.6.1 program.
For flow cytometry analysis of human T cells from PBMCs, human PBMCs were stained with the following antibodies: CD3-PE-Cy7 (1:50, clone: SK7, BioLegend), CD4-Perp-Cy5.5 (1:50, clone: OKT4, eBioscience), CD8-PE/BYG584 (1:50, clone: RPA-T8, eBioscience), CD69-FITC (1:50, clone: FN50, BioLegend), DAPI (1:100000, BioLegend). Samples were measured using the FACS Symphony A5 Cell Analyzer and analyzed with the FlowJo_v10.6.1 program.
For flow cytometry analysis of human T cells from BM, BM cells were washed in PBS for Zombie Aqua-BV510 (1:500, BioLegend) staining for 20 min on ice, then were washed and resuspended in PBS supplemented with 0.5% FCS at 4 °C with the following antibodies: Surface marker staining: CD3-PE-Cy5 (1:50, clone: UCHT1, BioLegend), CD8-PE-Cy7 (1:50, clone: SK1, BioLegend), CD69-BYG584 (1:50, clone: FN50, BioLegend), CCR7-APC (1:50, clone: G043H7, BioLegend), CD45RA-APC-H7 (1:50, clone: HI100, BD Bioscience), CD4-BUV395 (1:50, clone: SK3, BD Bioscience); Intracellular staining: the above samples were resuspended in 100 μl Fixation/Permeabilisation solution, incubated on ice for 20 min and then were washed with 1 ml 1X PermWash buffer, spin down and resuspended in 100 μl 1X PermWash buffer, and stained with the following antibodies on ice for 20 min: Granzyme B-APC-R700 (1:50, clone: QA16A02, BioLegend); Granzyme K-BB515 (1:20, clone: GM26E7, BioLegend), Perforin-BV421(1:50, clone: B-D48, Biolegend). Samples were measured using FACS Symphony A5 Cell Analyzer and analyzed with a FlowJo_v10.6.1 program.
For flow cytometry analysis of human purified CD34+ cells from BM co-cultured with (IKK2SE-transduced or EV) BMSC (OP-9), the cells were resuspended in PBS supplemented with 0.5% FCS at 4 °C and stained with the following antibodies: CD45-PE-Cy7 (1:100, clone: HI31, BD pharmingen), CD34-APC-R700 (1:100, clone: 581; BioLegend), CD38-BB700 (1:100, clone: HIT2, BioLegend), CD90-BYG584 (1:100, clone: eBio5E10, eBioscience), CD45RA-APC-Cy7 (1:100, clone: HI100, BD Bioscience). For the exclusion of dead cells, DAPI (1:100,000, BioLegend) was used. The immunophenotype and absolute cell counts were determined by flow cytometry using flow-count fluorosphere beads (Beckman 755 Coulter). The OP9 stromal cells were excluded based on GFP expression. Samples were analyzed by FACS Aria III (BD 530 Biosciences) analyzed with the FlowJo_v10.6.1 program.
T cell activation and isolation of supernatant
PBMCs were isolated from healthy donors and activated by Dynabeads™ Human T-Activator CD3/CD28 (Thermo Fisher Scientific, 11161D). 48 h post activation, the beads were removed and supernatant was collected and frozen in −80 °C. Supernatant from the immune culture was used to stimulate either healthy primary BMSCs or the human stromal cell line HS-5 (ATCC, CRL-3611) in vitro.
Lentiviral transduction of OP9 BMSCs and coculture with murine HSPCs
The plasmid containing the mutant form of IKK-2 (Flag-IKK-2 S177E S181E; referred hereafter as IKK2SE), which has previously been demonstrated to result in constitutive NFκB activation92,93 was obtained from Addgene. The IKK2SE fragment was recloned into the pCIG3 (pCMV-IRES-GFP version 3) plasmid (a gift from Felicia Goodrum (Addgene plasmid # 78264)), resulting in pCIG-IKK2SE. pCIG-IKK2SE and empty pCIG3 (pCIG-EV) were used to transfect HEK-293T cells with psPAX2 and pMD2G to produced lentivirus. OP9 cells were transduced with lentivirus containing either pCIG-IKK2SE or pCIG3, resulting in OP-9-IKK2SE and OP9-EV. Transduction efficiency was determined by measuring GFP expression with FACS. 2 days after transduction, 100,000 OP9 were seeded overnight and then cocultured with 200,000 wild-type C57BL/6J (CD45.2) murine bone marrow Lin– cells (using a mouse Lineage Cell Depletion Kit (Miltenyi Biotec)). After 5 days of coculture, the immunophenotype and absolute cell counts were determined by flow cytometry using flow-count fluorosphere beads (Beckman Coulter).
CFU-c assay
After 5 days of coculture, 800 mouse Lin– cells from EV or IKKSE coculture conditions were sorted and resuspended in IMDM (Thermo Fisher Scientific). This cell suspension was mixed with MethoCult™ GF M3434 (STEMCELL Technologies), which allowed the growth of colonies from all three lineages, and triplicate dishes were plated. The MethoCult plates were kept at 37 °C in a 5% CO2 incubator for 2 weeks until colony counting under a light microscope (Zeiss).
Mice and transplantation experiments
Three-month-old male C57BL/6 (CD45.2; Stock No. 000664) and female B6.SJL-Ptprca Pepcb/BoyJ (CD45.1; Stock No. 002014) mice were purchased from Charles River and maintained in specific pathogen-free conditions in the Experimental Animal Center of the Erasmus MC (EDC). Mice were housed in groups of maximum five animals (at least 100 cm2 per animal) under a standard 12 h light/dark cycle, with access to food and water ad libitum and an ambient temperature of 21–23 °C with air humidity maintained between 40 and 70%. All mice were euthanized by cervical dislocation.
Mouse hematopoietic cells (CD45.2) from EV or IKK2SE coculture conditions were sorted, 10,000 CD45.2+ cells were mixed with 100,000 CD45.1+ BM mononuclear cells and intravenously injected into lethally irradiated (8.5 Gy) age-matched CD45.1 mice. Chimerism of CD45.2+ cells was determined by monthly peripheral blood assessments. All work was performed in accordance with Dutch legislation and following institutional ethical approval by Animal Welfare/Ethics Committee of the EDC (IvD and DEC Rotterdam).
Human BM CD34+ HSPCs coculture with stroma cell line
Viably cryopreserved BM aspirates (mononuclear cell fractions) were thawed in a water bath at 37 °C and washed with warm DMEM supplemented with 10% FCS. CD34+ fractions were enriched by the CD34 MicroBead Kit (Miltenyi) and LS MACS separation columns (Miltenyi Biotec) to achieve optimal purity. CD34+ HSPCs were seeded on the lentiviral transduced OP-9 cell line (OP-9-IKK2SE and OP9-EV). The CD34+ cells (seeding density ~15,000 cells/well) were cultured in StemSpan™ SFEM (STEMCELLTM TECHNOLOGIES) medium supplemented with (Human SCF, 50 ng/ml; Human thrombopoietin, 50 ng/ml; Human Flt3 Ligand, 50 ng/ml). Cultures were kept at 37 °C in a 5% CO2 incubator for 2.5 days until further analysis.
Immunofluorescence staining of human bone marrow biopsies
MDS bone marrow biopsy samples derived from excess diagnostic material at the Department of Hematology were fixed in formaldehyde and processed for paraffin embedding at the Department of Clinical Pathology and Cancer Diagnostics (Karolinska University Hospital, Huddinge). Sections were cut with a thickness of 6 µm and mounted on glass slides. Prior to staining, these were deparaffinized in xylene and rehydrated through a graded ethanol series to distilled water. Antigen retrieval was performed using Tris-EDTA buffer (pH 9.0) in a water bath. Pre-blocking, sections were treated with TrueBlack (Biotium, 23007) to reduce autofluorescence according to manufacturer instruction. Blocking took place with 10% FBS in 0.5% BSA for 60 min at room temperature. Directly conjugated antibodies were diluted in blocking buffer and applied overnight at 4 °C in a humidified chamber. After 3 rounds of washing steps with PBS, nuclear staining was performed with DAPI. Slides were mounted using antifade mounting medium (abcam, AB104135) and stored at 4 °C until imaging. Sections treated in parallel without antibody staining were used to assess autofluorescence. Images were acquired on a single point scanning confocal Nikon Ti2 microscope with 2× GaAsP detectors (green and red). Fiji (ImageJ) was used to run the image analysis pipeline.
Quantitative PCR
Total RNA was isolated using Trizol reagent (Ambion), followed by cDNA synthesis using SuperScript II Reverse Transcriptase II kit (Invitrogen). Q-PCR was performed using Fast SYBR Green master mix (Applied Biosystems) on a 7500 Fast Real-Time PCR system (Applied Biosystems) or QIAquant 96 5plex system with the following primer sets:
Primers for human genes:
KITLG: Fw: 5′-AGTCCTGAGAAAGGGAAGGC-3′, Rv: 5′-GGCTCCAAAAGCAAAGCCAA-3′; NFKBIA: Fw: 5′-CCCTACACCTTGCCTGTGAG-3′, Rv: 5′-TAGACACGTGTGGCCATTGT-3′; IKK2: Fw: 5′-TCCGATGGCACAATCAGGAAA-3′, Rv: 5′-ATCTGGATCTCCAGGCACCA-3′; GAPDH: Fw: 5′-GTCGGAGTCAACGGATT-3′, Rv: 5′-AAGCTTCCCGTTCTCAG-3′; TNFRSF1A: Fw: 5′-ACGAGTGTGTCTCCTGTAGT-3’, Rv: 5′-CACTGTGGTGCCTGAGTCC-3′.
Primers for murine genes:
Il6: Fw: 5′-TCGGAGGCTTAATTACACA-3′, Rv: 5′-CTGGCTTTGTCTTTCTTGTT-3′; Nfkbia: Fw: 5′-TGAAGGACGAGGAGTACGAGC-3′, Rv: 5′-TTCGTGGATGATTGCCAAGTG-3′; Gapdh: Fw: 5′-AAGGTCATCCCAGAGCTGAA-3′, Rv: 5′-CTGCTTCACCACCTTCTTGA-3′.
Clinical outcome analyses
The SIGHT-P score was generated using a penalized Lasso regression algorithm with fourfold cross-validation (penalized R package) with SIGHT genes and iBMSC scores as inputs. The median score value was used to dichotomize the LR-MDS cohort into low- and high-score groups. These groups were labeled as SIGHT-Plow and SIGHT-Phigh, respectively. The log-rank test was used to assess statistical differences between the survival distributions, a p-value ≤ 0.05 was considered statistically significant. For clinical outcome analysis using publicly available datasets, GSE58831 dataset was acquired using R package GEOquery. Cumulative incidence of leukemic transformation was analyzed using competing risk analysis (competing with non-leukemia death) using tidycmprsk R package. Gray’s test was used to determine statistical significance.
Statistical analysis
Statistical analysis was performed using Prism 10 (GraphPad Software) and/or R. Unless otherwise specified, a two-sided Student’s t-test (parametric test) or Wilcoxon rank-sum test (nonparametric test) was used for two-group comparisons. For analysis involving multiple groups, one-way or two-way ANOVA (multiple comparisons) was used with FDR correction. For differential gene expression analysis, a two-sided Wald test with FDR correction was applied. The Kolmogorov–Smirnov (K-S) test with FDR correction was used for GSEA analysis. Two-sided Spearman’s rank-order correlation analysis with least-squares linear regression was conducted for correlation analysis. Log-rank test and two-sided Gray’s test for competing risks were used for survival analysis, as appropriate. For data displayed in boxplots, the central line marks the median, the box spans the 25th to 75th percentiles (interquartile range), and whiskers extend to the smallest and largest values within 1.5× the interquartile range; data points beyond this range are shown individually as outliers. All results in bar graphs are presented as mean ± SEM.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study were deposited and publicly available in European Genome-Phenome Archive (EGA) at accession number EGAS00001008181 and EGAS00001008182. The data analyzed in this study were obtained from the database of Gene-Expression Omnibus at GSE58831 and from EGA at EGAS00001001926. All other data supporting the findings of this study are cited in Methods, in supplementary documents, or are available upon request from the author. Source data are provided with this paper.
Code availability
Customized code used for analysis of the bioinformatic data are available at https://github.com/LanpengChen/LRMDS-scRNAseq-analysis.
References
Sperling, A. S., Gibson, C. J. & Ebert, B. L. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat. Rev. Cancer 17, 5–19 (2017).
Nimer, S. D. Myelodysplastic syndromes. Blood 111, 4841–4851 (2008).
Woll, P. S. et al. Myelodysplastic syndromes are propagated by rare and distinct human cancer stem cells in vivo. Cancer Cell 25, 794–808 (2014).
Walter, M. J. et al. Clonal architecture of secondary acute myeloid leukemia. N. Engl. J. Med. 366, 1090–1098 (2012).
Menssen, A. J. & Walter, M. J. Genetics of progression from MDS to secondary leukemia. Blood 136, 50–60 (2020).
Papaemmanuil, E. et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 122, 3616–3627 (2013).
Haferlach, T. et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia 28, 241–247 (2014).
Obeng, E. A. et al. Physiologic expression of Sf3b1(K700E) causes impaired erythropoiesis, aberrant splicing, and sensitivity to therapeutic spliceosome modulation. Cancer Cell 30, 404–417 (2016).
Pronk, E. & Raaijmakers, M. The mesenchymal niche in MDS. Blood 133, 1031–1038 (2019).
Gomariz, A. et al. Quantitative spatial analysis of haematopoiesis-regulating stromal cells in the bone marrow microenvironment by 3D microscopy. Nat. Commun. 9, 2532 (2018).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Baryawno, N. et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932.e1916 (2019).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Comazzetto, S. et al. Restricted hematopoietic progenitors and erythropoiesis require SCF from leptin receptor+ niche cells in the bone marrow. Cell Stem Cell 24, 477–486.e476 (2019).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Noda, M. et al. CXCL12-CXCR4 chemokine signaling is essential for NK-cell development in adult mice. Blood 117, 451–458 (2011).
Kohara, H. et al. Development of plasmacytoid dendritic cells in bone marrow stromal cell niches requires CXCL12-CXCR4 chemokine signaling. Blood 110, 4153–4160 (2007).
Tokoyoda, K., Hauser, A. E., Nakayama, T. & Radbruch, A. Organization of immunological memory by bone marrow stroma. Nat. Rev. Immunol. 10, 193–200 (2010).
Raaijmakers, M. H. et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 464, 852–857 (2010).
Zambetti, N. A. et al. Mesenchymal inflammation drives genotoxic stress in hematopoietic stem cells and predicts disease evolution in human pre-leukemia. Cell Stem Cell 19, 613–627 (2016).
Kode, A. et al. Leukaemogenesis induced by an activating beta-catenin mutation in osteoblasts. Nature 506, 240–244 (2014).
Dong, L. et al. Leukaemogenic effects of Ptpn11 activating mutations in the stem cell microenvironment. Nature 539, 304–308 (2016).
Medyouf, H. et al. Myelodysplastic cells in patients reprogram mesenchymal stromal cells to establish a transplantable stem cell niche disease unit. Cell Stem Cell 14, 824–837 (2014).
Raaijmakers, M. H. Disease progression in myelodysplastic syndromes: do mesenchymal cells pave the way?. Cell Stem Cell 14, 695–697 (2014).
Kfoury, Y. S. et al. The bone marrow stroma in human myelodysplastic syndrome reveals alterations that regulate disease progression. Blood Adv. 7, 6608–6623 (2023).
Chen, L. et al. A single-cell taxonomy predicts inflammatory niche remodeling to drive tissue failure and outcome in human AML. Blood Cancer Discov. 4, 394–417 (2023).
Ogawa, S. Genetics of MDS. Blood 133, 1049–1059 (2019).
Pellin, D. et al. A comprehensive single cell transcriptional landscape of human hematopoietic progenitors. Nat. Commun. 10, 2395 (2019).
Chen, L. et al. Transcriptional diversity during lineage commitment of human blood progenitors. Science 345, 1251033 (2014).
Tikhonova, A. N. et al. The bone marrow microenvironment at single-cell resolution. Nature 569, 222–228 (2019).
Yang, Y. H. et al. Annexin 1 negatively regulates IL-6 expression via effects on p38 MAPK and MAPK phosphatase-1. J. Immunol. 177, 8148–8153 (2006).
Fuster, J. J. et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 355, 842–847 (2017).
Choudhary, G. S. et al. Activation of targetable inflammatory immune signaling is seen in myelodysplastic syndromes with SF3B1 mutations. eLife 11, e78136 (2022).
Avagyan, S. et al. Resistance to inflammation underlies enhanced fitness in clonal hematopoiesis. Science 374, 768–772 (2021).
van de Loosdrecht, A. A. et al. Determinants of lenalidomide response with or without erythropoiesis-stimulating agents in myelodysplastic syndromes: the HOVON89 trial. Leukemia 38, 840–850 (2024).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Bukhari, S. et al. Single-cell RNA sequencing reveals distinct T cell populations in immune-related adverse events of checkpoint inhibitors. Cell Rep. Med. 4, 100868 (2023).
Andreatta, M. et al. Interpretation of T cell states from single-cell transcriptomics data using reference atlases. Nat. Commun. 12, 2965 (2021).
Jonsson, A. H. et al. Granzyme K(+) CD8 T cells form a core population in inflamed human tissue. Sci. Transl. Med. 14, eabo0686 (2022).
Thiery, J. et al. Perforin pores in the endosomal membrane trigger the release of endocytosed granzyme B into the cytosol of target cells. Nat. Immunol. 12, 770–777 (2011).
Raskov, H., Orhan, A., Christensen, J. P. & Gogenur, I. Cytotoxic CD8(+) T cells in cancer and cancer immunotherapy. Br. J. Cancer 124, 359–367 (2021).
Prokhnevska, N. et al. CD8(+) T cell activation in cancer comprises an initial activation phase in lymph nodes followed by effector differentiation within the tumor. Immunity 56, 107–124 e105 (2023).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Weimar, I. S. et al. Hepatocyte growth factor/scatter factor (HGF/SF) affects proliferation and migration of myeloid leukemic cells. Leukemia 12, 1195–1203 (1998).
Verstovsek, S. et al. Plasma hepatocyte growth factor is a prognostic factor in patients with acute myeloid leukemia but not in patients with myelodysplastic syndrome. Leukemia 15, 1165–1170 (2001).
Rodriguez-Fraticelli, A. E. et al. Single-cell lineage tracing unveils a role for TCF15 in haematopoiesis. Nature 583, 585–589 (2020).
Ping, Z. et al. Activation of NF-kappaB driven inflammatory programs in mesenchymal elements attenuates hematopoiesis in low-risk myelodysplastic syndromes. Leukemia 33, 536–541 (2019).
Kanayama, M. et al. CD86-based analysis enables observation of bona fide hematopoietic responses. Blood 136, 1144–1154 (2020).
Muto, T. et al. Adaptive response to inflammation contributes to sustained myelopoiesis and confers a competitive advantage in myelodysplastic syndrome HSCs. Nat. Immunol. 21, 535–545 (2020).
Thijssen, R. et al. Single-cell multiomics reveal the scale of multilayered adaptations enabling CLL relapse during venetoclax therapy. Blood 140, 2127–2141 (2022).
Boy, M. et al. Myelodysplastic syndrome associated TET2 mutations affect NK cell function and genome methylation. Nat. Commun. 14, 588 (2023).
Mortera-Blanco, T. et al. SF3B1-initiating mutations in MDS-RSs target lymphomyeloid hematopoietic stem cells. Blood 130, 881–890 (2017).
Fontanari Krause, L. M. et al. Identification and characterization of OSTL (RNF217) encoding a RING-IBR-RING protein adjacent to a translocation breakpoint involving ETV6 in childhood ALL. Sci. Rep. 4, 6565 (2014).
Zhou, H. et al. Role of c-Fos/JunD in protecting stress-induced cell death. Cell Prolif. 40, 431–444 (2007).
Weitzman, J. B., Fiette, L., Matsuo, K. & Yaniv, M. JunD protects cells from p53-dependent senescence and apoptosis. Mol. Cell 6, 1109–1119 (2000).
Imamura, T. et al. Overexpression of ZRF1 is related to tumor malignant potential and a poor outcome of gastric carcinoma. Carcinogenesis 39, 263–271 (2018).
Demajo, S. et al. ZRF1 controls the retinoic acid pathway and regulates leukemogenic potential in acute myeloid leukemia. Oncogene 33, 5501–5510 (2014).
Al Moussawi, K. et al. Mutant Ras and inflammation-driven skin tumorigenesis is suppressed via a JNK-iASPP-AP1 axis. Cell Rep. 41, 111503 (2022).
Meixner, A., Karreth, F., Kenner, L., Penninger, J. M. & Wagner, E. F. Jun and JunD-dependent functions in cell proliferation and stress response. Cell Death Differ. 17, 1409–1419 (2010).
Ribeiro, J. D. et al. ZRF1 controls oncogene-induced senescence through the INK4-ARF locus. Oncogene 32, 2161–2168 (2013).
Gerstung, M. et al. Combining gene mutation with gene expression data improves outcome prediction in myelodysplastic syndromes. Nat. Commun. 6, 5901 (2015).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Pietras, E. M. Inflammation: a key regulator of hematopoietic stem cell fate in health and disease. Blood 130, 1693–1698 (2017).
Trowbridge, J. J. & Starczynowski, D. T. Innate immune pathways and inflammation in hematopoietic aging, clonal hematopoiesis, and MDS. J. Exp. Med. 218, e20201544 (2021).
Mei, Y. et al. Bone marrow-confined IL-6 signaling mediates the progression of myelodysplastic syndromes to acute myeloid leukemia. J. Clin. Invest. 132, e152673 (2022).
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774 (2011).
Metzemaekers, M., Gouwy, M. & Proost, P. Neutrophil chemoattractant receptors in health and disease: double-edged swords. Cell Mol. Immunol. 17, 433–450 (2020).
Hormaechea-Agulla, D. et al. Chronic infection drives Dnmt3a-loss-of-function clonal hematopoiesis via IFNgamma signaling. Cell Stem Cell 28, 1428–1442.e1426 (2021).
Caiado, F. et al. Aging drives Tet2+/- clonal hematopoiesis via IL-1 signaling. Blood 141, 886–903 (2023).
Zhang, C. R. et al. Txnip enhances fitness of Dnmt3a-mutant hematopoietic stem cells via p21. Blood Cancer Discov. 3, 220–239 (2022).
Prummel, K. D. et al. Inflammatory stromal and T cells mediate human bone marrow niche remodeling in clonal hematopoiesis and myelodysplasia. Nat. Commun. https://doi.org/10.1038/s41467-025-65803-y (2025).
Tripodo, C. et al. Persistent immune stimulation exacerbates genetically driven myeloproliferative disorders via stromal remodeling. Cancer Res. 77, 3685–3699 (2017).
Molldrem, J. J. et al. Antithymocyte globulin for patients with myelodysplastic syndrome. Br. J. Haematol. 99, 699–705 (1997).
Stahl, M. et al. The use of immunosuppressive therapy in MDS: clinical outcomes and their predictors in a large international patient cohort. Blood Adv. 2, 1765–1772 (2018).
Bellissimo, D. C. et al. Runx1 negatively regulates inflammatory cytokine production by neutrophils in response to Toll-like receptor signaling. Blood Adv. 4, 1145–1158 (2020).
Ribezzo, F. et al. Rps14, Csnk1a1 and miRNA145/miRNA146a deficiency cooperate in the clinical phenotype and activation of the innate immune system in the 5q- syndrome. Leukemia 33, 1759–1772 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Fu, R. et al. clustifyr: an R package for automated single-cell RNA sequencing cluster classification. F1000Res 9, 223 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Peng, H. et al. Single-cell Rapid Capture Hybridization sequencing reliably detects isoform usage and coding mutations in targeted genes. Genome Res 35, 942–955 (2025).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Tian, L. et al. Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing. Genome Biol. 22, 310 (2021).
Kholmatov, M. et al. SpliceUp: predicting SF3B1 mutations in individual cells via aberrant splice site activation from scRNA-seq data. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2025.09.22.677806v1 (2025).
Moura, P. L. et al. Erythroid differentiation enhances RNA Mis-splicing in SF3B1-mutant myelodysplastic syndromes with ring sideroblasts. Cancer Res. 84, 211–225 (2024).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Mercurio, F. et al. IKK-1 and IKK-2: cytokine-activated IkappaB kinases essential for NF-kappaB activation. Science 278, 860–866 (1997).
Bosman, M. C. et al. The TAK1-NF-kappaB axis as therapeutic target for AML. Blood 124, 3130–3140 (2014).
Acknowledgements
The authors wish to thank Michael Vermeulen for technical assistance; the HOVON/SAKK leukemia working group and all its members and participating sites for conducting the HOVON89 trial; the Josephine Nefkens Precision Cancer Treatment Program for infrastructural support and members of the Erasmus MC Department of Hematology for providing scientific discussion. This work was supported by grants from the Dutch Cancer Society (KWF Kankerbestrijding), Amsterdam, the Netherlands (grant EMCR 10488 and 11092 to M.H.G.P.R. and 2008-4333 to A.A.v.d.L.).
Author information
Authors and Affiliations
Contributions
L.C., Y.B., and M.H.G.P.R. conceived the study and designed experiments; L.C., Y.B., E.P., C.v.D., T.V.D.v.T., E.M.B., D.B., and S.F. performed experiments; L.C., Y.B. performed data analysis; L.C., Y.B., R.M.H., and M.A.S. performed the bioinformatical analysis; A.O.d.G., T.M.W., M.K., J.B.Z., P.L.M., E.H.L., A.A.v.d.L., and J.H.J. provided samples and scientific inputs; L.C., Y.B., and M.H.G.P.R. wrote the manuscript, M.H.G.P.R. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yuxin Miao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, L., Bian, Y., Pronk, E. et al. An inflammatory T-cell-stromal axis contributes to hematopoietic stem/progenitor cell failure and clonal evolution in human myelodysplastic syndrome. Nat Commun 16, 10041 (2025). https://doi.org/10.1038/s41467-025-65802-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65802-z