Abstract
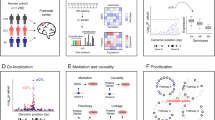
Genetic variants influencing gene expression have been extensively studied at the transcriptional level. How these variants affect downstream processes remains unclear. We quantitated ribosome occupancy in prefrontal cortex samples from the BrainGVEX cohort and integrated these data with transcriptomic and proteomic profiles from the same individuals. Through cis-QTL mapping, we identified genetic variants associated with transcript level (eQTLs), ribosome occupancy (rQTLs), and protein level (pQTLs). Notably, only 34% of eQTLs have their effects propagated to the protein levels, suggesting widespread post-transcriptional attenuation. Using both a gene-based approach and a variant-based approach we identified omics-specific QTLs that associated with brain disorder GWAS signals and found the majority of them to be driven predominantly by transcriptional regulation. Consistently, using a TWAS approach, we identified 74 SCZ risk genes across the three omics layers, 52 were discovered using transcriptome with 68% showing limited impact on protein expression. Our findings indicated that many disease-associated variants act through regulatory mechanisms that do not lead to an observable impact on the protein level.
Similar content being viewed by others
Introduction
Complex diseases such as neuropsychiatric disorders are multi-factorial with genetic components1,2. Large-scale Genome-Wide Association Studies (GWAS) have uncovered thousands of disease-associated loci, signaling a promising era ahead for causal variant identification3. However, efforts in fine mapping these disease risk loci often narrowed down the underlying causal signals to non-coding regions of the genome4,5,6. Regulatory variants in such non-coding regions are therefore the prime candidates for driving the genetic risk of disease etiology. Consequently, integrating gene expression information to pinpoint causal variants or to identify risk genes has become a staple of genetic studies of complex diseases7, with multiple consortia efforts facilitating large-scale gene expression profiling and regulatory element mapping8,9,10. Many powerful methods, such as coloc, PrediXcan, SMR/HEIDI, to name a few, have also been developed to leverage gene expression information for fine mapping GWAS signals or for identifying underlying risk genes11,12,13.
Brain-associated complex disorders, such as neuropsychiatric disorders, are a group of complex diseases that can be highly heritable14. For example, schizophrenia (SCZ), a psychiatric disorder affecting ~1% of the world-wide population15, has a heritability estimate ranged between 60% and 80%, indicating a strong genetic component16,17. Accordingly, recent SCZ GWAS study reported by Trubetskoy et al. identified 287 significant risk loci and prioritized 120 risk genes using functional genomics18. The use of RNA-Seq data from the brain was instrumental for risk gene prioritization by Trubetskoy et al., however, information from downstream gene regulation processes, such as translation rate and protein abundance, was either not utilized or unavailable.
Measuring transcriptional changes as a proxy for gene activity has a long history in molecular biology19. In the context of human genetics, buffering of downstream effects of genetic variants impacting gene expression (i.e., an eQTL) has been shown to be prevalent20. In addition, QTLs specific to protein level have also been reported20,21,22. Together, these observations indicated the importance and potential benefits of including downstream omics types, such as proteomics data, as information sources for fine mapping disease regulatory variants. Indeed, recent studies using multi-omics approaches have demonstrated increased power for risk gene identification among other benefits23,24. Of note, our recent work on genetic variants associated with protein expression level in the prefrontal cortex of the human brain indicated the contribution from non-synonymous coding variants to the changes in protein expression level, and the utility of these protein QTL variants in prioritizing GWAS risk genes for psychiatric disorders22.
Another potential benefit of taking a multi-omics approach for identifying disease risk genes rests in the potential to dissect the fine details of regulatory mechanisms driving the disease-genotype association. Having relevant datasets to illuminate the origin and propagation of genetic impact could potentially arrive at a driver regulatory process from a risk variant to a risk gene. Ribo-seq is a technology that fills in the gap between transcript and protein expression. By adapting RNA-Seq to a ribosome footprinting method, ribo-seq provides transcriptome-wide quantification of ribosome occupancy25,26, which can serve as a proxy for the amount of active translation synthesizing proteins from each mRNA transcript. When analyzed in conjunction with RNA-Seq and quantitative proteomics data, ribo-seq enables identification of translational and post-translational regulatory events20,27, both major steps defining the Central Dogma of molecular biology.
As a part of the consortium efforts to improve our understanding of the genetic basis of neuropsychiatric disorders28, we generated multiple data modalities that included SNP genotyping, RNA-Seq, ribo-seq, and proteomics of postmortem cortical tissue samples of the BrainGVEX cohort, which altogether covered the major steps of the Central Dogma. In conjunction with the quantitative proteomics and transcriptome profiling results that we previously published22,29, here we integrated ribo-seq data as our operational definition for protein synthesis, based on the ribosome occupancy level across the transcriptome, to advance our multi-omics investigation. Our results reveal regulatory properties of common variants and their utility in identifying the regulatory processes in the human brain driving disease risk for brain-associated complex disorders. It offers an opportunity to dissect and appreciate the regulatory information flow in the biological processes from a population perspective. Additional “rules” including an effect attenuation process, is recognized.
Results
Measuring transcriptome-wide ribosome occupancy level in prefrontal cortex of adult human brain to quantify the level of protein synthesis
To investigate regulatory impact of genetic variants on protein synthesis in the prefrontal cortex of the human brain, we performed ribosome profiling on 211 prefrontal cortex samples from the BrainGVEX collection. In total, we collected ~62 billion ribosome profiling reads. Consistent with the expected ribosome footprint size, we found the average insert size of our ribo-seq libraries to range between 27.4 and 29.5 nucleotides. Similar to prior published studies30, we found on average 74 % of unwanted reads from ribosomal RNA, tRNA, and snoRNA, which contributed no information to the translation of protein-coding genes. After removing these unwanted reads, we obtained an average of 30.3 million uniquely mapped informative reads per sample (inter-quartile range: 20.5 ~ 37.6 million reads) (Fig. S1A). Focusing on the informative reads, we found the majority (i.e., 84%) of our ribo-seq reads mapped to coding sequence (CDS) exons (Fig. S1B). Moreover, when visualized in aggregate across annotated coding genes, we found our ribo-seq data to show strong sub-codon periodicity at the expected positions (Fig. S1C). A high proportion of CDS exon reads and strong sub-codon periodicity indicates the enrichment of footprints from ribosomes actively engaged in translating mature mRNA and reflects the quality of the dataset.
Multi-omics cis-QTL mapping identified candidate regulatory variants and revealed translational and post-translational attenuation of eQTL effects
To identify variants associated with inter-individual expression differences, we perform cis-QTL mapping for each omics type independently. From the full dataset we filtered out low coverage genes and removed potentially low quality samples (see details in Data processing section in Methods) to reach a mapping dataset of 416 RNA-Seq samples, 195 ribo-seq samples, 268 proteomics samples, and the corresponding genotype data. Using the mapping dataset, we identified 12,411 eQTLs (out of 16,540 genes we deem sufficiently quantitated), 2849 rQTLs (out of 14,572 genes), and 1036 pQTLs (out of 8013 genes) at FDR < 0.1. Overall, we observed similar genomic feature enrichment patterns of QTL variants across the three omics types (Fig. S2). On the other hand, when focusing on coding variants, we found pQTL to have the largest proportion of coding variants (pQTL: 11.5% vs. eQTL: 3.4% vs. rQTL: 5.1%). Among the coding variants, we found 82.4% were missense variants for pQTL comparing to 56.2% for rQTL and 46.4% for eQTL (Fig. S2). Of note, we found three start loss variants (i.e., nonsynonymous substitutions in start codons) in rQTL, which were not found in eQTL.
Intriguingly, we found drastic differences between omics types in the number of QTLs mapped, with far fewer rQTLs and pQTLs identified, suggesting a reduction of genetic effect along the path of Central Dogma. However, the differences in the number of genes tested between omics types and the differences in sample size complicate the interpretation. To better compare the effects of genetic regulation across data modalities, we identified 185 samples with 7458 genes that were sufficiently quantitated across all three omics types (Fig. S3). Using this subset of data, we found 3253 eQTLs, 1344 rQTLs, and 657 pQTLs at FDR < 0.1 (Fig. 1A, Fig. S4, Data S1, S2, S3). Consistently fewer significant QTLs were identified as we moved downstream the Central Dogma of molecular biology.
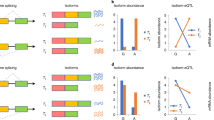
A P-value quantile-quantile plot between the observed (Y-axis) and the expected based on null distribution (X-axis). QTL mapping p-values shown here are permutation-based p-values calculated using QTLtools with beta approximation. The black line indicates the expected distribution of p-values when there are no real QTL signals. The number of cis-QTLs (i.e., the most significantly associated SNP for each gene) identified at 10% FDR is labeled in the top left inset. B Replication rate between QTL types. Proportions of QTLs replicated in the other two omics types are listed in the 3 × 3 matrix. Each row is a discovery omics type and each element of the row correspond to the proportion QTL signals replicated in the omics type specified by the column label. For example, only 34% of the eQTL signals were replicated in the protein data. Ribo: ribosome occupancy. C Effect size of CMC eQTL SNPs in BrainGVEX data. Mean and 95% confidence interval of absolute per allele effect across 5915 CMC eQTL SNPs that were also analyzed in the BrainGVEX dataset is shown.
A challenge in comparing between the numbers of QTLs identified from each omics type rests in the fact that not all true effects were identified. While the majority of the QTLs identified here were replicated in independent datasets (Data S4, S5, Fig. S5), which provides experimental support for our discoveries, the level of false negative rates is less clear. Tests replicating QTLs identified from one omics type in the other omics types can better capture the proportion of genetic effects shared between QTL types. We performed replication tests using π1 estimates from the qvalue method31. Overall, we found a high proportion of QTLs replicated in other omics types (Fig. 1B). However, when considering the replication rates with the direction of genetic information flow, we found asymmetric replication rates, with the downstream omics types to replicate less than the upstream omics types. More specifically, we found 84.7% of the rQTLs were replicated at the transcript level, but only 60.2% of the eQTLs were replicated at the ribosome occupancy level. Moreover, while 75.9% of the pQTLs were replicated at the transcript level, only 34.0% of the eQTLs were replicated at the protein level (Fig. 1B). A similar asymmetry in proportion replicated between upstream and downstream omics types was observed when using a direction-aware cutoff-based approach across a wide range of significance cutoffs (Fig. S6). The lower percentages of eQTLs and rQTLs replicated at the protein level indicate potential effect attenuation (i.e., either the inter-individual variation in gene expression becomes smaller and therefore harder to detect or a lack of such effect in the downstream omics types). Such effect attenuation at the protein level has been previously reported in lymphoblastoid cell lines (LCLs)20. Our brain data revealed an additional layer of complexity, where effect attenuation was also observed at the level of mRNA translation.
While our replication tests revealed a trend of effect attenuation for eQTL variants in the downstream phenotypes (Fig. 1B), these same observations could alternatively be explained by differences in statistical power between technologies. An independent piece of evidence that is not sensitive to measurement precision is needed to reach a solid conclusion. Fold change estimates from less precise measurements are noisier but not biased. In other words, by directly comparing QTL impact on expression level change across omics types, we could gain a direct view of effect size attenuation. One issue that remains is that using eQTLs identified from our dataset could bias the results by artificially inflating the variant pool with large eQTL effect sizes, which might arise simply from applying the QTL mapping procedure to the same dataset (i.e., a technical ascertainment bias). Using eQTLs independently identified from prefrontal cortex samples by the CommonMind Consortium (CMC)32, we avoid the ascertainment bias and can therefore compare the effect size of eQTL variants between the three omics types using our dataset. A similar approach was successfully implemented to address this power issue in previous work in LCLs20. Using 5915 CMC eQTLs that were also quantitated in our dataset, we found the eQTL effects on the transcript expression level to be significantly larger than their effects on ribosome occupancy level (per allele log2 fold differences: mRNA 0.2433 [95% CI = 0.2381 ~ 0.2486] vs. ribosome occupancy 0.1836 [95% CI = 0.1794 ~ 0.1878]), which were in turn significantly larger than their effects on protein level (per allele log2 fold differences: 0.1486 [95% CI = 0.1451 ~ 0.1522]) (Fig. 1C, one-sided t-test P < 2.2e−16 for all pairwise comparisons). Moreover, for these CMC eQTLs, we found translational regulation to account for more effect size reduction than post-translational regulation (Fig. 1C). By focusing on the effect sizes of independently identified eQTLs, our results strongly support the presence of downstream mechanisms attenuating eQTL effects both at the ribosome occupancy level (translationally) and at the protein level (post-translationally). A complementary analysis looking at the effect size of pQTL variants identified from an independent study33 further supported our conclusion. We found, in our dataset, a larger effect size at the transcript level when comparing to the fold change at the protein level for these independently identified pQTL SNPs (Fig. S7). We noticed that for these independently identified pQTL SNPs, the eQTL effect attenuation appears to be predominantly translationally regulated (Fig. S7B; mRNA vs. ribosome occupancy one-sided t-test P = 6.6e−15 and ribosome occupancy vs. protein t-test P = 0.4). Similar results were observed across a wide range of significance levels used to define pQTL (Fig. S7).
In contrast to prior studies in LCL20, we found clear translational attenuation of eQTL effects in prefrontal cortex. To further investigate this apparent discrepancy, we cross-referenced the genes associated with eQTL identified in prefrontal cortex samples to those identified in LCL. We found rather limited overlaps: of the 3253 eQTL-associated genes identified in our brain data only 598 (i.e., ~18%) were found associated with a significant eQTL in LCL (Fig. S8A), indicating the possibility that the between-study differences in eQTL discoveries could play a role in the observed differences in translational attenuation of eQTL effects. Indeed, when inspecting the downstream effect of LCL-specific eQTL versus a set of effect size-matched brain-specific eQTL in our brain dataset we found no attenuation of eQTL effect at the ribosome occupancy level among LCL-specific eQTL, while translational attenuation of eQTL effect remains significant among brain-specific eQTLs (Fig. S8B). On the other hand, the shared eQTL found between the two studies showed the expected translational attenuation of eQTL effects in the brain dataset (Fig. S8C), again supporting the notion that the distinct groups of eQTL analyzed between the two studies drive the observed discrepancy.
Identifying omics specific QTLs and their signal colocalization with brain disorder GWAS
The prevalent effect size reduction of eQTLs raised the question of the relevance of these genetic regulations at the organismal level. Because most cellular tasks are executed by proteins, the genetic regulatory effects not reaching the protein level are, presumably, less likely to have an impact on organismal traits. More specifically, we seek the biological relevance of expression-specific and ribosome occupancy level-specific QTLs (i.e., esQTL and rsQTL). To answer this question, we first set out to investigate the relevance of different QTL types in brain-associated complex traits such as neuropsychiatric and neurodevelopmental disorders. By applying mediated expression score regression (MESC)34 to summary statistics from GWAS of four different complex brain disorders and our three types of molecular QTL summary statistics, we estimated for each disorder the proportion of heritability mediated by each of our molecular QTL types (Data S6 and Fig. S9). We noticed that for schizophrenia (SCZ), bipolar disorder (BD), and autism spectrum disorder (ASD), eQTL mediated the most heritability, followed by rQTL, which is then followed by pQTL. For example, using the Trubetskoy et al. SCZ GWAS18, we found our eQTLs to mediate 7.09%, rQTLs to mediate 4.06%, and pQTLs to mediate 2.17% of SCZ heritability (Data S6). On the other hand, for major depressive disorder (MDD), all three QTL types mediated similar proportion of heritability ( ~ 2.5%).
After establishing the relevance for each of the three QTL types in brain-associated disorders, we next sought to identify omics-specific QTLs, in order to further evaluate their relevance in driving disease risk. For example, we aim to identify expression-specific QTLs (i.e., genetic variants that impact transcript level of the linked genes but not the downstream ribosome occupancy level nor protein level) that colocalize with GWAS signals of brain associated disorders. To identify omics-specific QTLs we took a multiple regression approach by first removing effects of the two other omics-types and then test for genotype association with the residual. To further reduce the number of false positives, for each potential omics-specific QTL identified at 10% FDR we further test genotype association with the other two omics-types and remove any variant that has nominally significant association. We identified 1553 loci/genes with expression-specific QTL (esQTL), 155 genes with ribosome occupancy-specific QTL (rsQTL), and 161 genes with protein-specific QTL (psQTL) (Fig. S10). Similar to standard QTLs we found enrichment of esQTL and psQTL in genomic features such as the exons and the 3’UTRs (Fig. S2). In addition, we found even higher proportion of coding variants and missense variants among psQTLs comparing to the enrichment of these categories reported above for pQTL (Fig. 2A). Of note, we found a clear downward shift in 5’UTR enrichment from pQTL to psQTL (odds ratio 2.51 vs. 1.52, P < 5e−6). Similar decrease in 5’UTR variant enrichment was also observed among the other two pairs of standard and omics-specific QTL (i.e., eQTL vs. esQTL and rQTL vs. rsQTL), indicating the possibility that QTL variants in 5’UTR are more likely to be shared across omics types, presumably because they impact protein expression level via transcript level and/or ribosome occupancy level.
A Categorizing QTL SNPs according to coding feature annotations. Bar plots showing the percentage of QTL SNPs in each category out of the total number of QTL SNPs considered. Note the high percentage (28%) of coding variants in psQTL SNPs and the dominating percentage (97.8%) of missense variants among coding psQTL SNPs. B An example SCZ risk gene, GATAD2A, with predominantly transcript-specific QTL effects. The red line indicates the top signal colocalization SNP between eQTL and SCZ GWAS on a Manhattan plot. Boxplots to the right summarize the normalized gene expression level stratified by eQTL SNP genotypes across 185 samples. Ribo: ribosome occupancy. C–E Example genes containing multiple QTL signals of varying omics specificity. Individual credible set SNPs containing 95% of the QTL signal are color-coded on a Manhattan plot with a vertical line indicating the top SNP that contributes the most signal. Boxplots to the right summarize the normalized gene expression level stratified by genotypes of the top SNP across 185 samples. Note that the example gene in (E) is a SCZ risk gene with two credible sets. Only one of the two credible sets’ QTL signal colocalizes with SCZ GWAS signal. The bar plots underneath the expression level boxplots show the log odds ratio +/− standard errors for the association between the credible set top SNP genotype and SCZ, which is calculated based on GWAS summary statistics from Trubetskoy et al. QTL p-values shown in Manhattan plots are nominal p-values obtained from two-sided t-tests of linear regression coefficient using QTLtools for each SNP–gene pair. Boxplots summarizing expression level show the interquartile range (IQR), spanning from the 25th percentile (lower bound) to the 75th percentile (upper bound), with the median (center line) shown inside the box. Whiskers extend to 1.5 × IQR.
To identify genes with omics-specific QTL effects driving disease risk, we perform, for each gene containing omics-specific QTL signal, coloc analyses with default prior11 between the nominal p-value from cis-QTL mapping and each brain-associated disorder GWAS. We performed separate analysis using either the nominal p-value from the omics-specific cis-QTL mapping or the standard cis-QTL mapping. At a colocalization posterior probability cutoff of 70% we identified risk genes for each omics-type by brain disorder combination for both approaches (Data S7). We found the most risk genes from esQTL loci, with 14 genes showing signal colocalization between SCZ GWAS signals and either standard or omics-specific eQTL signals, and 14 genes for BD, while the remaining combinations resulted in single-digit colocalizations (Data S7). For the majority of the risk genes, we observed consistent results between the two approaches (Data S7). On the other hand, we found a few risk genes that have GWAS colocalization with only standard QTL signals, indicating the possibility of GWAS signals colocalizing with QTL signals that were shared with, and therefore regressed out by, other omics types at these loci. In other words, some of these omics-specific QTL were identified from loci containing shared QTL signals that colocalized with GWAS signal.
Because regulation of a gene is often modulated by multiple genetic variants, to evaluate the consequence of overall cis-QTL impact on gene expression, we use PrediXcan to estimate aggregate genetic regulatory effects for each gene. To identify genes that have omics-specific QTL effects, for each omics type, we built PrediXcan models using normalized quantifications with and without regressing out signals from the other omics types. A significant PrediXcan model fit on residuals indicates the potential presence of omics-specific QTL signals (i.e., QTL signals remained after regression). Using a high PrediXcan model fit R2 cutoff of 0.1, we identified 1098 candidate genes containing omics-specific QTL signals. To further identify genes with predominantly omics-specific QTL effects, we then remove, from the candidate gene list, genes that have either shared QTL effects or additional QTL effects from other omics types by filtering out genes with significant PrediXcan model built from any of the two other omics types. Using this approach we identified 382 esQTL genes, 36 rsQTL genes, and 51 psQTL genes that have predominantly omics-specific QTL effect. Consistently, these predominantly omics-specific QTL effect genes identified using the PrediXcan gene-based approach are enriched of omics-specific QTL variants identified from the SNP-based multiple regression approach (Fig. S11). Colocalization with brain-associated complex disorder GWAS identified 11 risk genes with predominantly omics-specific QTL effects (Data S8). We found most signal colocalization with SCZ, which resulted in 7 SCZ risk genes with predominantly esQTL effects (Fig. 2B).
For the remaining 629 candidate genes that potentially contain both omics-specific and shared QTL signals or contain independent omics-specific signals from multiple omics types (i.e., the candidate omics-specific QTL effect genes filtered out from the above step), we use SuSiE to identify independent QTL signals from each omics for each gene (i.e., SuSiE 95% credible sets). By separating out independent signals from the same gene using SuSiE we can then operate on the credible set level to identify additional omics-specific QTL signals that colocalize with complex disorder GWAS. We identified a total of 829 credible sets from 594 candidate genes (Fig. S12). For each credible set, we perform QTL signal colocalization between omics types using coloc. We found 404 credible sets to contain QTL signals that are shared with at least one other omics type (i.e., PPH4 > 0.7), while 96 contain QTL signals that are shared across all three omics types (Data S9). Of note, of the 594 candidate genes, we found 170 genes to have at least two credible sets. Among these genes with multiple credible sets, we found examples of separate omics-specific QTL signals from different omics types (Fig. 2C) and example genes that contain both omics-specific and shared QTL signals (Fig. 2D).
To further investigate the relevance of these QTL credible sets in brain-associated disorders, we search for colocalization with brain-associated disorder GWAS signals. Of the 829 credible sets we found 25 (from 25 genes) to have signal colocalization with at least one of the four brain-associated complex disorders tested (Data S10). Of these 25 GWAS colocalization signals, 13 have QTL signals shared between at least two omics types. Upon visually inspecting the nominal p-values from QTL mapping and brain-associated disorder GWAS signals at these risk loci we found a few genes with apparently differential colocalization results between credible sets. For example, we found RCBTB1 to have two eQTL credible sets while only one of the two credible sets contains eQTL signals colocalizing with SCZ GWAS (PPH4 = 0.8). Interestingly, the GWAS colocalizing eQTL for the RCBTB1 locus appears to be omics-specific while the other credible set appears to contain a QTL signal that is shared between transcript expression and ribosome occupancy (PPH4 = 0.7) (Fig. 2E).
Functional genomics identification of Schizophrenia risk genes
To further investigate the relevance of attenuated eQTLs in brain disease risk, we next took a complementary approach by first identifying risk genes from each omics type separately and then investigating the relevant regulatory processes driving disease risks. Given that we observed the highest number of risk genes for SCZ from our omics-specific QTL analyses, here we focused on SCZ. We applied S-PrediXcan35 on the Trubetskoy et al. SCZ GWAS data and our BrainGVEX multi-omics data. At 5% family-wise error rate, we found 52, 29, and 16 SCZ risk genes, respectively from RNA-Seq, ribo-seq, and proteomics data (Fig. 3A, Fig. S13). Consistently, when comparing the corresponding GWAS-QTL coloc results between genes that passed and genes that failed the S-PrediXcan tests, we found S-PrediXcan risk genes to have higher colocalization posterior probability (Fig. S14). Among S-PrediXcan risk genes, only four genes, NEK4, KHK, CNNM2, and DARS2 were consistently identified as SCZ risk genes from all three omics types (Fig. 3B). The majority (74.3%) of the risk genes were identified only from one of the three omics types. This limited sharing in risk gene identification between omics types is consistently observed across multiple significance levels and is in clear contrast to the amount of signal sharing found between QTL types (Fig. 1B; Fig. S15; Data S11, S12, S13).
A Manhattan plots showing significance level (i.e., -log10 FWER from S-PrediXcan) of gene expression-schizophrenia association across the genome for genes that pass the 5% FWER significance cutoff for each omics type. P-values from S-PrediXcan were calculated based on two-sided Z tests. The black horizontal dotted line indicated the significance cutoff (5% Bonferroni-adjusted p-value). B Venn diagram illustrating the number of overlapping risk genes identified between omics types. C Boxplots summarizing normalized gene expression level stratified by eQTL SNP genotypes across 185 samples for KHK. Boxes represent the interquartile range (IQR), spanning from the 25th percentile (lower bound) to the 75th percentile (upper bound), with the median (center line) shown inside the box. Whiskers extend to 1.5 × IQR. D Manhattan plots showing nominal p-value distribution for each QTL type and schizophrenia GWAS for the 1 Mb QTL mapping window flanking KHK. The red line indicates the position of the lead colocalization SNP between eQTL and schizophrenia GWAS.
Among the 74 risk genes we identified using S-PrediXcan (i.e., the union of the risk genes identified from each of the three omics types), 44 have previously been reported in GWAS studies as either the mapped genes or as one of the nearby genes under the GWAS peak (Data S14)18,23,29,36,37,38,39. Of these previously reported genes, 27 matched the mapped genes, while the remaining 17 nominated an alternative candidate gene for each GWAS locus (Note that for some of these 17 loci, the original GWAS signals were mapped to an intergenic region). Comparing our results to other published multi-omics SCZ risk gene identification studies, we found 15 matched to the risk genes identified by Giambartolomei et al. which used RNA-Seq and DNA methylation data from the prefrontal cortex of the human brain23, and 12 matched to the 120 prioritized SCZ risk genes from Trubetskoy et al., which used colocalization with eQTL and Hi-C data18. When comparing our results to other published TWAS SCZ risk gene identification studies, we found 19 matched to the risk genes identified by Gandal et al.29, and 21 matched to the risk genes identified by Gusev et al39. (See Data S14 for an extended list of risk gene comparisons to prior TWAS studies). On the other hand, for 22 of our 74 risk genes (30%), we found no match to the known risk gene list (Data S14, bottom rows), which we compiled based on previous GWAS, TWAS, and other functional genomics risk gene identification studies18,23,29,36,37,38,39. These “no match” risk genes have relatively weak SCZ GWAS signals and are therefore challenging to identify without the additional information provided by our multi-omics QTL dataset. For example, we found strong colocalization between a modest SCZ GWAS signal at 2p23.3 and all three types of molecular QTLs of the gene KHK (Fig. 3D). Intriguingly, the KHK pQTL is in the opposite direction of the KHK eQTL and rQTL (Fig. 3C), suggesting a strong post-translational process regulating its protein level in the opposite direction to the transcriptional regulation. KHK, Ketohexokinase, plays a pivotal role in fructose metabolism and has been hypothesized to contribute to neuronal glycolysis and the loss of neuronal functions40. In addition to KHK, other SCZ risk genes have also been reported to participate in neuronal/ cognitive functions: For example, in neuroglial cell lines, silencing of SNX32 leads to defects in neuronal differentiation41. As another example, NSF, N-Ethylmaleimide Sensitive Factor, which encodes a vesicle fusing ATPase, was reported as a causal factor in intelligence traits42.
Analyses of multi-omics dataset reveal regulatory mechanisms of schizophrenia risk genes
While PrediXcan is a powerful tool for GWAS risk gene identification, it does not control for potential horizontal pleiotropy43. To this end, we performed two-sample Mendelian Randomization (MR) with Egger regression to replicate the risk genes we identified from using S-PrediXcan. Egger regression includes an intercept term, which can be used to evaluate the level of horizontal pleiotropy44. For each risk gene we first used LD-clumping45 to identify the top QTL SNP from each clump (i.e., a group of linked variants defined by the LD cutoff) to serve as strong genetic instruments46. We then tested for the causal relationship between gene regulation (i.e., QTL signal) and SCZ (i.e., GWAS). We used an operational definition of a causal effect based on the MR test results (see Methods). At 10% FDR, of the 97 gene-by-omics combinations (i.e., a total of 74 risk genes, including some discovered from more than one omics type), 67.0% (65/97) passed the MR test. Of those 33.0% that failed the MR test, 37.5% (12 out of 32) failed because of horizontal pleiotropy identified by the Egger intercept test (Data S15). A total of 53 risk genes were replicated in at least one of the three omics types. Note that in order to increase our power, we used a relaxed R2 cutoff of 0.5 for LD-clumping and adjusted for the linkage between instrument variables in our MR tests using generalized IVW and Egger47. Similar causal effects were observed when using either a stringent R2 cutoff of 0.01 or using SuSiE (the Sum of Single Effects)48 as an alternative approach to select instrument SNPs (Fig. S16). However, these more stringent criteria on instrument variable independence led to lower power to detect and fewer genes available to test (Data S16). For example, when using SuSiE for selecting instrument variables, only 40 genes were tested because some of the risk genes had no fine-mapped QTL SNPs according to SuSiE.
A key strength of using a multi-omics QTL approach to identify GWAS risk genes rests in the possibility of further narrowing down the potential regulatory mechanisms. To this end, we further examined the likely causal mechanisms for the 53 replicated SCZ risk genes using one-sample Mendelian Randomization to infer causality between QTL types (Data S17). We focused our analysis on independently testing for causal effects between neighboring QTL types following the direction of information flow of the Central Dogma (i.e., mRNA -> ribosome occupancy, and ribosome occupancy -> protein). Here, we used fine-mapped QTL SNPs identified from the exposure omics types (i.e., the upstream omics types) as instrument variables for one-sample MR analysis. Among the 53 MR-Egger replicated SCZ risk genes, we found 21 genes with significant causal effects both from eQTL to rQTL (i.e. the upstream pathway) and from rQTL to pQTL (i.e. the downstream pathway) (both-passed risk genes; Fig. 4A). Significant causal effects detected from both pathways suggest transcriptionally regulated protein level differences as the potential mechanisms driving these risk genes in SCZ etiology (see an example in Fig. 4B). On the other hand, 25 and 7 replicated risk genes have either significant causal effects only in one of the two pathways (single-passed risk genes; Fig. 4A) or have no significant causal effects (none-passed risk genes; Fig. 4A), respectively.
A Summaries of MR test results. Left panel: Two-sample MR replication of S-PrediXcan SCZ risk gene discoveries. Each circle represents a S-PrediXcan discovery. A solid-colored circle indicates successful replication in a MR test. Right panel: One-sample MR test results for causality between omics types. Shown are MR effect size +/− the corresponding standard errors estimated using 185 samples. Red data points indicate significant test results. Ribo: ribosome occupancy. B, C Example SCZ risk genes driven by either transcriptional regulation of protein expression (B) or transcript-specific effects (C). Manhattan plots on the left show nominal p-values for all variants in the QTL mapping window for each omics and SCZ GWAS, with the red vertical line indicating the top signal colocalization SNP between QTL and GWAS. Boxplots on the right summarize the normalized gene expression level stratified by QTL SNP genotype across 185 samples. Boxes represent the interquartile range (IQR), spanning from the 25th percentile (lower bound) to the 75th percentile (upper bound), with the median (center line) shown inside the box. Whiskers extend to 1.5 × IQR. The effect size estimates from the corresponding one-sample MR causality tests between each pair of omics and two-sample MR tests replicating the risk gene discoveries are shown in the plot beneath. MR effect +/− the corresponding standard errors were estimated using 185 samples.
Of note, for 23 of the 25 single-passed risk genes, we found significant causal effects in the upstream pathway (mRNA -> ribosome occupancy). This asymmetry is reminiscent of the eQTL effect attenuation described in the prior sections. A failed test could indicate either a true lack of effect or a lack of statistical power. To take a closer look, we directly assessed the effect size, the noise level, and instrument strength of the one-sample MR test results. When comparing to the 21 both-passed risk genes, we found clearly smaller effect sizes for the failed tests of both the 7 none-passed risk genes (average 0.577 vs 0.058, t-test P < 10−3, Fig. S17A) and the 23 single-passed risk genes (average 0.524 vs 0.116, t-test P < 10−3, Fig. S18A). Note that in both cases the inter-quartile range of the estimated causal effects for the failed tests covered zero (Figs. S17A, S18A). For these same comparison groups, we found no significant differences in instrument strength (Figs. S17C, S18C) and found slightly higher noise level in the failed MR tests (Fig. S17B, S18B). When considered together, these observations (i.e., clearly smaller effect size and slightly elevated noise level for the failed tests) indicate that many of these test results reflect a true lack of causal effect, while in some cases weak effects were obscured by noise. In other words, many of these risk genes are likely to be driven by either omics-specific QTLs or attenuated eQTLs. Case in point, we found SF3B1 to have strong signal colocalization between eQTLs and SCZ GWAS without clear QTL signals in either ribo-seq or proteomics data (Fig. 4C).
Discussion
Using a panel of postmortem prefrontal cortex samples, we found clear evidence of post-transcriptional attenuation of eQTL effects in the human brain. Many of the differences found in transcriptomes were not present in proteomes. This observation echoes earlier work in HapMap lymphoblastoid cell lines20 and extends the prior conclusion from in vitro cell lines to complex human tissues. Importantly, distinct from the earlier work in lymphoblastoid cell lines, which found rQTL to mostly track with eQTL, we found clear attenuation of eQTL effect in ribo-seq data, indicating that translational regulatory processes are involved in eQTL effect attenuation in the human brain. Our further investigation on this observation indicated that differences in the set of eQTL (eGENE) identified between the two studies explains the discrepancy. However, it remains to be determined whether the discrepancy observed between the two studies reflects a true difference between the tissue/cell types used in these studies, or effects rooted in differences in sample complexity, or effects from other unknown unaccounted for variations, or simply incidental. Prevalent translational attenuation of variant impact on transcriptional gene expression level has previously been reported in budding yeast49,50. However, results from follow-up studies51 appear to present an inconsistent picture. Here, using replication tests for individual eQTLs and testing for aggregate effect size of eQTLs independently identified from CMC, we provided strong evidence supporting prevalent translational attenuation of eQTL effects in the human brain. Importantly, by focusing on comparing QTL effects (i.e., between genotype fold change) in aggregate, our analyses are robust to power issues resulted from measurement imprecision (random noise). On the other hand, our conclusions could still be sensitive to potential biases in the dataset, which could result from technical issues that cannot be randomized. For this reason, we paid close attention to the impact of sequencing coverage (for sequencing data) and ratio compression (for quantitative mass spectrometry data) on our major conclusion. Of note, our quantitative mass spectrometry study employed an analytical procedure that adjusts for contaminating background signals in order to correct for potential ratio compression in quantification52. Consistently, among low-abundance proteins, which are more susceptible to the influence of ratio compression, we did not find a disproportional decrease in pQTL effect size (Fig. S19C). Similarly, while genes with low expression level are more sensitive to issues related to low sequencing coverage, we found little to no correlation between eQTL/rQTL effect size and expression level (Figs. S19A, S19B). Although ample precautions and multiple distinct approaches were taken to confirm the observed effect size attenuation, it remains possible that unknown, unaccounted for omics-specific features could potentially confound the results. In other words, although operating at the molecular level, our study remained observational. Future work replicating this observation and elucidating the molecular mechanisms of post-transcriptional attenuation of eQTL effects is needed to provide a clear understanding of the phenomenon.
Although our study was not designed to answer questions regarding the underlying mechanisms of the observed translational and post-translational attenuation of eQTL effect, the high prevalence of the observed eQTL effect attenuation when considered in conjunction with the low prevalence of coding mutations in the human population provides some hints. Since a compensatory variant driving translational or post-translational attenuation is expected to be either a coding mutation that impact translation elongation rates or protein turnover rates or a UTR variant that impacts translation initiation rate, the apparent mismatch in incident prevalence indicated that compensatory variants are unlikely to explain the majority of the effect size attenuation and therefore implies a broad-spectrum mechanism that reduce variation at the protein level across genes. Consistently, we observed significant translational and post-translational attenuation of eQTL effect across all PhastCons score strata on top of a clear trend of decreased QTL effect size as the evolution conservation scores increase (Fig. S20). Our current work focuses on the other important follow-up question of whether the attenuated eQTLs are functionally (biologically) relevant. We attempted to address this question by exploring the relevance of attenuated eQTLs in brain-associated disorders, such as SCZ, a neuropsychiatric disorder that is highly heritable. We took multiple approaches to identify risk genes that have either omics-specific QTL signals or attenuated eQTLs. In one approach, we used regression-based methods to identify omics-specific QTL performing separate analyses either at the SNP level or using PrediXcan to aggregate variant impact at the gene level. In both cases, we found the most number of colocalization between esQTL and complex disorders (i.e., comparing to rsQTL and psQTL). From the gene level analysis we further identified interesting example genes containing multiple QTL signals with varying level of omics-specificity, including an example gene, RCBTB1. We found two strong eQTL signals at RCBTB1, while one potentially shared and one potentially omics-specific. Only the apparently omics-specific eQTL was found to colocalize with SCZ GWAS. These examples illustrate the complexity of variant impact on gene regulation and disease risks. A regression-based approach, however, could produce false positive omics-specific QTL simply from noisy predictors. Although we implemented checks and filters to remove these likely false positives, this caveat is important to keep in mind when interpreting these results.
In another approach, we conducted our search by identifying risk genes from all quantitated genes using a TWAS approach, S-PrediXcan, separately for each omics. We then tested for causal regulatory mechanisms driving the risk gene using QTL SNPs as instrumental variable in a one-sample Mendelian Randomization test. Before testing for causal regulatory mechanisms, we replicated the TWAS risk genes using two-sample Mendelian Randomization with Egger intercept tests. About 30% of the risk genes that we identified here have not previously been reported as SCZ risk genes. The use of our functional genomics dataset is key to the discovery of these SCZ risk genes because of their relatively weak GWAS signals (see Fig. 3D for an example). As a reminder, we would like to emphasize the fact that individual risk gene identification is dependent on the choice of significance cutoffs, which is to some extent arbitrary. We used a rather stringent family-wised error rate for risk gene identification in order to reduce the number of false positive results that could mislead researchers interested in molecular biology follow-up studies.
To identify causal regulatory mechanisms for each replicated risk gene, we further tested causality between QTL types. We found 21 risk genes that are likely contributing to SCZ risk through transcriptionally regulated protein levels. On the other hand, we also identified 7 genes that show no significant results from MR tests (candidate omics-specific risk genes) and 23 genes that have significant MR test results only from transcription to translation (candidate post-translationally-attenuated risk genes). In essence, attempts to identify omics-specific risk genes (or omics-specific QTLs) are dealing with the challenge of separating true negatives from false negatives. As such, the interpretation of the failed MR tests is challenging. Our subsequent analyses looking at comparing instrument strength, noise level, and effect size between passed and failed MR tests indicated comparable instrument strength, slightly elevated noise level in the failed tests, and clearly smaller effect size in the failed tests. In other words, it remains possible that some of the failed tests could be reflecting a small effect size obscured by the elevated noise level. In addition to the issues with false negative results, some of the discoveries could be false positives to begin with. Although we have good confidence with our FDR and FWER estimates, given that our test statistics for QTL mapping and risk gene identification are well-calibrated (Figs. S4, S13), pleiotropy could introduce positive results from a different underlying cause (i.e., true effects on SCZ risk but false positive risk genes).
Power issues, however, do not explain the whole story. As was consistently observed throughout our study, when viewed in aggregate, we see clear effect size differences between omics types, both for QTLs and for causal effects from MR tests. These effect size estimates are not influenced by significance cutoffs and are not biased by power differences. Such general trends are also unlikely to simply be a result of spurious associations or made up entirely of false positives. Our results therefore, bring forth an interesting mechanistic question: how do attenuated eQTL variants impact SCZ without changing protein levels? One interesting possibility is that the biologically relevant traits here are protein synthesis rates and protein turnover rates. An association between genotype and protein synthesis rates could manifest in the form of an attenuated eQTL, if the differences in per transcript protein synthesis rate (e.g., differences in ribosome elongation rate or translation initiation rate) appears to offset the differences introduced at the transcript level, which in turn could resulted in a lack of differences in ribosome occupancy level. Similarly, a protein turnover QTL could also manifest in the form of a rQTL attenuated at the protein level. In other words, colocalization between an attenuated rQTL and a GWAS signal could actually be reflecting a colocalization of the GWAS signal with a protein turnover QTL. Testing this hypothesis requires a direct measurement of protein turnover rate. We hope by presenting the current results, our findings can inspire future studies on this topic to better understand the regulatory processes from DNA to brain-associated disorders.
Methods
Data sources
Our study complies with all relevant ethical regulations. The SNP genotypes22, RNA-Seq29, and quantitative mass spectrometry22 data generated from prefrontal cortex tissue samples of the PsychENCODE BrainGVEX cohort were downloaded from the National Institute of Mental Health (NIMH) Data Archive (see Data availability). According to the NIMH Data Archive policy, all data distributed by NIMH Data Archive are de-identified and were collected under Independent Review Boards (IRB)-approved protocols with consent for data sharing. See Data S18 for a summary of the number of samples in each dataset and their respective overlap with the samples in the genotype data. The BrainGVEX cohort includes 420 Caucasians, 2 Hispanics, 1 African American, 3 Asian Americans, and 14 unknown-origin individuals. We also used GWAS summary statistics data of schizophrenia18, bipolar disorder53, autism spectrum disorder54, and major depression disorder55 obtained from the Psychiatric Genomics Consortium.
Ribosome profiling
The biological samples used in this study were from the BrainGVEX cohort, which was obtained from the Stanley Medical Research Institute (SMRI) and the Banner Sun Health Research Institute (BSHRI). Participant consent and compensation were managed by the original providers. Ribosome Profiling experiments were performed using Illumina’s TrueSeq Ribo Profile (Mammalian) Kit. TrueSeq Ribo Profile (Mammalian) Kit was developed for cell lines. We adapted the protocol to frozen tissue samples with MP Biomedicals™ FastPrep-24™ Classic Bead Beating Grinder and Lysis System. Specifically, 60–80 mg of frozen brain tissue was homogenized in Lysing Matrix D tubes containing 800 µl polysome lysis buffer. The Lysing tubes were placed on the FastPrep-24™ homogenizer set at 4.0 m/s with 20 s increments until no visible chunks of tissue remained. Tissue lysate was incubated on ice for 10 min followed by centrifugation at 20,000 g at 4 °C for 10 min to pellet cell debris. Clarified lysate was subsequently used for ribo-seq library preparation following TrueSeq Ribo Profile (Mammalian) Kit instructions. Indexed libraries were pooled and then sequenced on an Illumina HiSeq 4000. Note that the experimental protocol for TrueSeq Ribo Profile (Mammalian) Kit that we followed is a modified version of the previous ribo-seq protocol published by Ingolia and colleagues26, and it has the following key modifications: Monosome isolation was performed using Sephacryl S400 spin columns (GE27–5140-01) on a tabletop centrifuge instead of ultra-high speed centrifugation in sucrose cushion. Ribosomal RNA depletion was carried out by using Ribo-Zero Magnetic Kits and this step is moved up to right after the monosomal RNA isolation step and before the Ribosome Protected Fragment gel isolation step.
Data processing, gene expression quantification, and normalization
For Genotype data, we obtained Genotype VCF files from PsychENCODE NIMH Data Archive (see Data availability). The VCF file includes data from 420 individuals genotyped using a combination of 3 different platforms. Details on genotyping experiments and data processing can be found in our companion manuscripts22,29. Briefly, data from whole-genome sequencing (WGS), Human PsychChip platform (a custom version of the Illumina Infinium CoreExome-24 v1.1 BeadChip), and Affymetrix GeneChip Mapping 5.0 K Array were separately imputed using Minimac3 with Haplotype Reference Consortium (HRC) reference panel and then merged using PLINK with filters for variant call quality, imputation quality, minor allele frequency, and relatedness. Only variants with genotype (or imputed) R2 > 0.3, Hardy-Weinberg Equilibrium > 10−6 and minor allele frequency (MAF) > 0.01 are kept. The combined and filtered VCF file contains genotype data of 8,108,028 SNPs for 420 individuals. We further evaluated potential effects of additional confounding factors on genotype data (Fig. S21).
For RNA-Seq data, we obtained raw reads in FASTQ format from the PsychENCODE NIMH Data Archive (see Data availability). We then process the raw reads by removing the adapter sequence using cutadapt (v1.12) code “cutadapt –minimum-length=5 –quality-base=33 -q 30 -a AGATCGGAAGAGCACACGTCTGAACTCCAGTCA -A AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGT”. The cutadapt processed reads were then mapped onto GENCODE Release 19 (GRCh37.p13) genome with transcript annotations from the corresponding version of GTF files using STAR (v2.4.2a). We used RSEM software (v1.2.29) to quantify read counts for each gene56. The cpm function in the R package “limma” was used to calculate the log-transformed counts per million (CPM). We filtered out genes with CPM < 1 in more than 75% of the samples and samples with network connectivity57 z score < −5 (Fig. S22A), which resulted in a total of 17,207 genes from 426 individuals in the quantification table. We then used normalize.quantiles function in the R package “preprocessCore”58 to normalize expression level for each sample. DRAMS software was used to detect and correct mixed-up samples59, which resulted in a final count of 423 individuals.
For ribo-seq data, we used cutadapt (v1.12) to remove adapter sequence from raw reads with code “cutadapt -a AGATCGGAAGAGCACACGTCT –quality-base=33 –minimum-length=25 –discard-untrimmed”. The processed reads were then mapped against a FASTA file of rRNA, tRNA, and snoRNA sequences downloaded from NCBI using bowtie260 to filter out uninformative reads. The filtered reads were mapped to GENCODE Release 19 (GRCh37.p13) genome with corresponding transcript annotation GTF file using STAR (v2.4.2a). The uniquely mapped reads, as defined by the “NH:i:1” flag of the alignment files, were kept for subsequent analysis. We used the featureCounts function in the R package “subreads”61 to calculate gene level read counts for ribosome occupancy. The cpm function in the R package “limma” was used to calculate log-transformed CPM value. We filtered out genes with CPM < 1 in more than 75% of the samples and samples with network connectivity57 z score < −3.5 (Fig. S22B), which resulted in a total of 15,171 genes quantitated from 209 individuals in the quantification table. We then used the normalize.quantiles function in the R package “preprocessCore”58 to normalize ribosome occupancy level for each sample. DRAMS software was used to detect and correct mixed-up samples59, which resulted in a final count of 199 individuals.
For quantitative mass spectrometry data, we obtained a protein quantification table from the PsychENCODE NIMH Data Archive (see Data availability). This table includes abundance quantification for 11,572 proteins from 268 individuals. The MS/MS raw data were searched against a composite target/decoy human proteome database. The target database contains 83,955 entries, including both canonical proteins and isoforms downloaded from the UniProt database. Since we employed shotgun proteomics, we identified peptides rather than full-length proteins. In cases where a peptide could be mapped to multiple homologous proteins, we followed the rule of parsimony: the peptide was first assigned to the canonical protein. If no canonical form was defined, the peptide was assigned to the protein isoform with the highest uniquely mapped PSM (Peptide-Spectrum Match). For quantification, we summarized all peptides that are mapped to the representative protein. These data processing steps for producing the mass spectrometry quantification table are detailed in Luo et al.22. We further log-transformed protein abundance for each sample. We filtered out genes with protein abundance <1 in more than 75% of the samples and samples with network connectivity57 z score < −6 (Fig. S22C, note that all genes and samples passed the filtering cutoff). We then used the normalize.quantiles function in the R package “preprocessCore”58 to normalize protein level for each sample. DRAMS software was used to detect mixed-up samples59 and found none. We matched protein IDs to gene IDs according to the UCSC database (hg19 version), which resulted in protein-level quantifications for 8330 genes. When matching pQTL signals to genes for comparison with eQTLs/rQTLs, we used the pQTL signals associated with the protein isoform that has the highest median abundance.
QTL mapping
Estimating and adjusting for unwanted factors
We used the R package “PEER”62 to estimate hidden factors for RNA-Seq, ribo-seq, and proteomics data separately. The principle of selecting unwanted hidden factors was to maximize the variance explained with the least number of factors. We identified 30, 29, and 19 hidden factors to remove from RNA-Seq, ribo-seq, and mass spectrometry data, respectively (Fig. S23). For each gene from each omics type, we adjusted the expression level by fitting the selected hidden factors as predictors in a linear model and taking the residuals as the adjusted expression level. The adjusted expression levels were then further centered by mean and scaled by standard deviation.
Genotype association tests
We identified cis-region expression QTLs (eQTLs), ribosome occupancy QTLs (rQTLs), and protein QTLs (pQTLs) separately using QTLTools (1.3.1)63. Because each gene can encode several protein isoforms, we selected the protein isoform with the highest median abundance as the representative protein. For each gene, we defined cis-region as the region ranging from 1 Mb upstream of the 5’ end of the gene to 1 Mb downstream of the 3’ end of the gene (i.e., the length of the gene body plus 2 Mb in size). We tested all common SNPs (MAF > 0.05) within the cis-region for each gene. We performed 10,000 permutations to create null distributions to obtain permutation-based p-values, and further adjusted permutation-based p-values used the beta-approximation approach implemented in QTLTools to calculate empirical p-values. For each gene, we selected the most significant SNP to represent the QTL signal (i.e., the variant with the smallest empirical p-value) and then calculated a genome-wide FDR using the qvalue function of the R package “qvalue”.
Annotation and enrichment analyses of QTL variants
We used Ensembl Variant Effect Predictor (VEP) version 105 to annotate QTL SNPs. Each variant was assigned multiple annotations, as each variant could map to multiple transcript isoforms of the same gene. We perform three separate analyses with different counting schemes, counting all annotations, counting only the most frequent annotations, counting only the annotation with the most severe consequence, to ensure the percentage estimates and conclusions of enrichment are robust.
For genomic feature enrichment analysis, we extracted 5’ UTRs, exons, introns, 3’ UTRs, and intergenic regions from the GENCODE v19 GTF file. The exon regions were extracted directly from entries in the third column of the GTF file (column label: “exon”). Similarly, gene regions were extracted directly from the GTF file. Intron regions were calculated by removing exon regions from gene regions, while intergenic regions were obtained by removing gene regions from the genome. The 5’ UTR and 3’ UTR coordinates were generated using the fiveUTRsByTranscript() and threeUTRsByTranscript() functions from the R package “GenomicFeatures”, respectively. We used bedtools intersect -wa to identify QTL SNPs overlapping each genomic feature. We used Fisher’s exact test to evaluate the statistical significance of QTL SNP enrichment in each genomic feature. For genic features (e.g., 5’ UTRs, exons, introns, 3’ UTRs) total genic counts were used as the background; for intergenic features, total cis-region (i.e., mapping window) counts were used as the background.
Estimating mediated heritability of each omics
We used MESC to estimate the proportion of heritability mediated by different omics separately for four different brain-associated disorders33. In the first step, we calculated the overall expression score using the unwanted-factor-adjusted-expression data (see QTL mapping section) as the individual-level gene expression data and the corresponding BrainGVEX genotype data. We used the 1000 genome phase 3 genotype data to calculate LD scores in order to match BrainGVEX genotype data to GWAS genotype data. In the second step, we used the overall expression score calculated in the previous step and GWAS summary statistics to estimate heritability mediated by each omics.
Identifying omics-specific QTL
SNP-based approach
We used QTLTools to identify omics-specific QTL variants following the same procedure as standard QTL mapping, which is described in the above section, except that the gene expression quantification for the omics type of interest was adjusted by regressing out signals from the other two omics types. The regression was achieved by including peer-adjusted normalized quantification of the other two omics types as covariates in QTLTools. For each gene, we selected the SNP with the most significant genotype-expression association as the candidate omics-specific QTL SNP and used Benjamini-Hochberg procedure to control for false discovery rates. To further remove false positives resulted from ineffective removal of signals from the other two omics types, likely due to poor linear model fit, we removed candidate omics-specific QTLs with nominal significant association (p-values < 0.01) with any of the other two omics types.
Gene-based PrediXcan approach
Building prediction models: We used the PrediXcan software13 to separately build gene expression prediction models based on RNA-Seq, ribo-seq, and quantitative mass spectrometry data. We built these models for each gene for each omics type, with and without regressing out signals from the other two omics types (i.e., for each gene using data of each omics type, we build both a standard model and a residual model). For each model, an Elastic Net algorithm was used for feature selection from SNPs located within the cis-region defined for each gene (i.e., gene body +/− 1 Mb flanking regions) based on results from a ten-fold cross-validation. Weights were produced for every selected SNP of each gene, which were used in the prediction models. For each model, we calculated the Pearson correlation between the predicted and the observed gene expression (Cross-validation R, Rcv), which was considered a metric for model prediction accuracy. Unless otherwise specified, we consider models with Rcv2 above the PrediXcan default 0.01 threshold as models describing genes with significant genetic contribution to expression regulation13. We produced prediction models for the unified set of 7458 genes from 185 samples.
Identifying and categorizing genes containing omics-specific genetic effects: We defined genes with significant residual model as candidate genes containing omics-specific QTL signals using a high Rcv2 cutoff of 0.1 in conjunction with a FDR cutoff of 0.05 for the model fit. Of these candidate genes we further classify them by the model fit results from the other omics types. If no significant standard models were built from any of the two other omics-types, we consider the gene as a predominantly omics-specific QTL gene. For example, for a gene with significant residual model built from RNA-Seq data (eQTL signal), if there are no significant standard model built from either ribo-seq nor proteomics data we consider the gene as a predominantly transcript-specific QTL gene (esQTL). For candidate genes that have significant model built from any of the other two omics types we consider these candidate genes as potentially harboring multiple QTL signals from different omics types (either omics-specific or shared QTL effects). To separate out independent signals from each gene we used SuSiE to identify 95% credible sets representing each individual signals. To categorize these independent signals as shared or omics-specific, we then perform coloc analysis with data from the other omics-types for each credible set using SNPs in modest LD (squared correlation > 0.01) with the credible set of interest to ensure sufficient background SNPs and excluding variants that are in linkage (squared correlation > 0.1) with any other credible sets. For each credible set we also performed coloc analysis with brain-associated disorder GWAS, in order to identify QTL signals driving disease risks among these credible sets. See the next section for details on signal colocalization analysis using coloc.
Colocalization
We used coloc11 to detect signal colocalization between brain-associated disorder GWAS and each QTL type at the cis-region of each QTL gene. For each QTL gene, for all common SNPs (MAF > 0.05) within the cis-region, we use QTL effects and GWAS summary statistics of brain-associated disorders as input for coloc analysis. To estimate QTL effects, we calculated, from a linear model fit using SNP genotype as the predictor for gene quantifications, the predictor regression coefficient and its corresponding square of standard error. We then used the coloc.abf function in the R package “coloc”11 to calculate the posterior probability of each hypothesis using the default prior. We use posterior probability of 70% for the colocalization hypothesis (i.e., PPH4) as the cutoff for reporting our colocalization findings.
TWAS identification of SCZ risk genes
We used S-PrediXcan34 to perform gene-level association tests based on the prediction models built using our brain prefrontal cortex multi-omics data, which is described in the “Gene-based PrediXcan approach” section, and the SCZ GWAS summary statistics data from PGC318. The association tests were performed separately for each omics. For protein data, we performed an omnibus test to incorporate p-values of all protein isoforms together to calculate a single p-value for the corresponding gene. The family-wise error rates for SCZ risk genes were calculated using Bonferroni correction on nominal p-values.
Two-sample Mendelian randomization (MR)
To identify causal relationships between each omics type and SCZ we used MR analysis. Here, we used fine-mapped QTL SNPs as instruments, gene expression quantification at each omics type as exposure, and SCZ GWAS signal as outcome.
More specifically, we took the following steps to test for a causal relationship between gene regulation at each omics level and SCZ:
Step 1: Here we used two methods for selecting instrument SNPs: an LD clumping approach and a SuSiE approach. For the LD clumping approach, for each gene of each QTL type, we used PLINK with “–clump-kb 1000 –clump-r2 0.5” parameters and a QTL nominal p-value cutoff to define instrument SNPs from separate QTL signals. Since our goal is to test as many risk genes as we reasonably can, we used a dynamic (greedy) approach in defining the QTL p-value cutoff for instrument SNP selection. We first set a stringent p-value cutoff at 10−4, which identified the majority of our instrument SNPs. For genes that don’t have at least 3 instrument SNPs identified, we incrementally relaxed the p-value cutoff first to 10−3 and then to 10−2. See Data S15 for details on the p-value cutoffs used for each omics type by risk gene combination. Independent of the above-mentioned, we performed a separate MR analysis using LD clumping with r2 < 0.01 to ensure independence between instrument SNPs and a p-value cutoff of 0.05. For the SuSiE approach, we used susie_rss() function with default parameters in the R package “SuSiE”47.
Step 2: We then used the mr_steiger() function in the R package “TwoSampleMR” to test the causal direction of selected instrument SNPs. For genes failing the Steiger test, we applied the steiger_filtering() function to verify the direction of each instrument SNP and excluded those with incorrect directions, ensuring the explained variance by the instrument SNPs was greater for the exposure than for the outcome.
Step 3: For each gene, we used harmonise_data() function in R package “TwoSampleMR”64 to harmonize QTLs of each omics type and SCZ GWAS SNPs to be in the same direction (i.e., effect relative to the same allele).
Step 4: We then performed two-sample MR for each gene by omics type combination separately. Two-sample MR analysis was done using the R package “MendelianRandomization”65. We used mr_ivw(data, correl = TRUE) function to conduct generalized IVW and mr_egger(data,correl = TRUE) function to conduct generalized Egger (see next step). Generalized IVW accounts for linkage between instrument SNPs, which enables the MR analysis to use linked instrument SNPs.
Step 5: We used the intercept test (i.e., the Egger method) to test for horizontal pleiotropy, and used predictor coefficient and its corresponding p value from IVW to determine the effect size and significance of causal effects for each omics type on SCZ.
Step 6: We used Benjamini-Hochberg control procedure to adjust the nominal p-values from IVW for multiple testing.
Step 7: We define a gene by omics type combination as causal for SCZ if the causal effect test passed the significance cutoff of FDR < 0.1 and the Egger intercept test passed the intercept p-value > 0.05 cutoff.
One-sample Mendelian randomization
For one-sample MR we used Two-stage predictor substitution (TSPS) approach, which is a method to find causal relationships between omics types. We performed the following analysis in two iterations, both following the direction of genetic information flow. In the first iteration, we tested causal relationships between transcript level and ribosome occupancy level (i.e., mRNA -> ribosome occupancy). In the second iteration, we tested causal relationships between ribosome occupancy level and protein level (i.e., ribosome occupancy -> protein).
Step 1: We used LD clumping for instrument SNP selection as described in the previous section. Here we tested two pathways (mRNA -> ribosome occupancy and ribosome occupancy -> protein). For mRNA -> ribosome occupancy, we used the SNPs of eQTL with p-value < 10−4 and R2 < 0.5 from LD. In order to test as many risk genes as we reasonably can, the same dynamic p-value cutoff approach (i.e., the same as the above-described for two-sample MR) was used for instrument SNP selection. See Data S17 for details on the p-value cutoffs used for each risk gene.
Step 2: We then used the mr_steiger() function in the R package “TwoSampleMR” to test the causal direction of selected instrument SNPs. For genes failing the Steiger test, we applied the steiger_filtering() function to verify the direction of each instrument SNP and excluded those with incorrect directions, ensuring the explained variance by the instrument SNPs was greater for the exposure than for the outcome.
Step 3: We then combined genotype and quantification data of the relevant omics types into a dataframe: datmr for mRNA -> ribosome occupancy pathway, datrp for ribosome occupancy -> protein pathway.
Step 4: For each gene, we then set formula in R package “OneSampleMR”66: tsps (“ribosome occupancy ~ mRNA | SNP1 + SNP2 + SNP3 + …+SNPn”, data = datmr) and tsps (“protein ~ ribosome occupancy | SNP1 + SNP2 + SNP3 + …+SNPm”, data = datrp).
Step 5: We used the summary() function to get the slope (i.e., the regression coefficient) and the corresponding p-value of the slope from each tsps object.
Step 6: We used Benjamini-Hochberg control procedure to adjust the p-value of the slope for multiple testing.
Step 7: We defined a relationship for each gene by pathway combination as causal if the test results passed the significance cutoff of FDR < 0.1.
Step 8: We also used R package “ivreg” with the same dataset and formula as the above-described for tsps objects to calculate F-statistics for each gene. The calculated F-statistics were used to check instrument strength.
Statistics and reproducibility
Details of the statistical analyses, including the number of replicates and software used, are provided above. No statistical method was used to predetermine sample size. Genes with low expression levels and outlier samples were excluded from the analyses. The experiments were not randomized, as this study involved observational omics data from human postmortem brain samples. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ribo-seq raw files generated from this study have been deposited in the National Institute of Mental Health Data Archive (NDA) (https://nda.nih.gov/edit_collection.html?id=5032). The SNP genotype, RNA-Seq and MS/MS raw data from PsychENCODE BrainGVEX cohort can also be found in the link above. The raw data are available under restricted access due to participant privacy and data sharing agreements, and can be obtained by submitting a data access request via the NDA portal. For questions regarding the request, please contact NDA at NDAHelp@mail.nih.gov. Data access requests are typically reviewed within 10 business days from receipt of the request, and applicants will be notified of the decision by the NDA. The processed gene expression data are available at https://www.synapse.org/Synapse:syn50698163/files/. The result files generated in this study are provided in the Supplementary Data/Source Data files. Source data are provided with this paper.
Code availability
All code used in the analyses are available at https://github.com/qiumanL/multi-omics-qtl.
References
A. Fialkowski, T. M. Beasley, H. K. Tiwari, Emery and Rimoin’s Principles and Practice of Medical Genetics and Genomics (Seventh Edition) Ch. 11, (Academic Press, 2019).
Brown, A. S. The environment and susceptibility to schizophrenia. Prog. Neurobiol. 93, 23–58 (2011).
Solovieff, N., Cotsapas, C., Lee, P. H., Purcell, S. M. & Smoller, J. W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Heckman, M. G. et al. Fine-mapping of the non-coding variation driving the Caucasian LRRK2 GWAS signal in Parkinson’s disease. Parkinsonism Relat. Disord. 83, 22–30 (2021).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Wang, L. et al. Brain Banks Spur New Frontiers in Neuropsychiatric Research and Strategies for Analysis and Validation. Genomics Proteom. Bioinforma. 17, 402–414 (2019).
ENCODE Project Consortium et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Giambartolomei, C. et al. Bayesian test for Colocalisation between pairs of genetic association studies using summary statistics. PloS Genet. 10, e1004383 (2014).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Bray, N. J. & O’Donovan, M. C. The genetics of neuropsychiatric disorders. Brain Neurosci. Adv. 2, 2398212818799271 (2018).
Kahn, R. S. et al. Schizophrenia. Nat. Rev. Dis. Prim. 1, 15067 (2015).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Sullivan, P. F., Kendler, K. S. & Neale, M. C. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch. Gen. Psychiatry 60, 1187–1192 (2003).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Wolf, J. B. W. Principles of transcriptome analysis and gene expression quantification: an RNA-seq tutorial. Mol. Ecol. Resour. 13, 559–572 (2013).
Battle, A. et al. Impact of regulatory variation from RNA to protein. Science 347, 664–667 (2015).
Ferkingstad, E. et al. Large-scale integration of the plasma proteome with genetics and disease. Nat. Genet. 53, 1712–1721 (2021).
Luo, J. et al. Genetic regulation of human brain proteome reveals proteins implicated in psychiatric disorders. Mol Psychiatry. 29, 3330–3343 (2024).
Giambartolomei, C. et al. A Bayesian framework for multiple trait colocalization from summary association statistics. Bioinformatics 34, 2538–2545 (2018).
Foley, C. N. et al. A fast and efficient colocalization algorithm for identifying shared genetic risk factors across multiple traits. Nat. Commun. 12, 764 (2021).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Cenik, C. et al. Integrative analysis of RNA, translation, and protein levels reveals distinct regulatory variation across humans. Genome Res. 25, 1610–1621 (2015).
Akbarian, S. et al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
Gandal, M.J. et al., Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 362, eaat8127 (2018).
Brar, G. A. & Weissman, J. S. Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat. Rev. Mol. Cell Biol. 16, 651–664 (2015).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. Usa. 100, 9440–9445 (2003).
Hoffman, G. E. et al. CommonMind Consortium provides transcriptomic and epigenomic data for Schizophrenia and Bipolar Disorder. Sci. Data. 6, 180 (2019).
Robins, C. et al. Genetic control of the human brain proteome. Am. J. Hum. Genet. 108, 400–410 (2021).
Yao, D. W., O’Connor, J., Price, A. L. & Gusev, A. Quantifying genetic effects on disease mediated by assayed gene expression levels. Nat. Genet. 52, 626–633 (2020).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat. Commun. 9, 1825 (2018).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Lam, M. et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat. Genet. 51, 1670–1678 (2019).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
Johnson, R. J. et al. Cerebral Fructose Metabolism as a Potential Mechanism Driving Alzheimer’s Disease. Front Aging Neurosci. 2, 560865 (2020).
Sugatha, J. et al. Insights into cargo sorting by SNX32 and its role in neurite outgrowth. eLife 12, e84396 (2023).
Korologou-Linden, R., Leyden, G. M., Relton, C. L., Richmond, R. C. & Richardson, T. G. Multi-omics analyses of cognitive traits and psychiatric disorders highlights brain-dependent mechanisms. Hum. Mol. Genet. 32, 885–896 (2021).
Zhou, D. et al. A unified framework for joint-tissue transcriptome-wide association and Mendelian randomization analysis. Nat. Genet. 52, 1239–1246 (2020).
Burgess, S. & Thompson, S. G. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur. J. Epidemiol. 32, 377–389 (2017).
Purcell, S. et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Gkatzionis, A., Burgess, S. & Newcombe, P. J. Statistical methods for cis-Mendelian randomization with two-sample summary-level data. Genet. Epidemiol. 47, 3–25 (2023).
Burgess, S., Dudbridge, F. & Thompson, S. G. Combining information on multiple instrumental variables in Mendelian randomization: comparison of allele score and summarized data methods. Stat. Med. 35, 1880–1906 (2016).
Zou, Y., Carbonetto, P., Wang, G. & Stephens, M. Fine-mapping from summary data with the “Sum of Single Effects” model. PloS Genet 18, e1010299 (2022).
Artieri, C. G. & Fraser, H. B. Evolution at two levels of gene expression in yeast. Genome Res. 24, 411–421 (2014).
McManus, C. J., May, G. E., Spealman, P. & Shteyman, A. Ribosome profiling reveals post-transcriptional buffering of divergent gene expression in yeast. Genome Res. 24, 422–430 (2014).
Albert, F. W., Muzzey, D., Weissman, J. S. & Kruglyak, L. Genetic influences on translation in yeast. PloS Genet. 10, e1004692 (2014).
Niu, M. et al. Extensive peptide fractionation and y 1 ion-based interference detection method for enabling accurate quantification by isobaric labeling and mass spectrometry. Anal. Chem. 89, 2956–2963 (2017).
Mullins, N. et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet 53, 817–829 (2021).
Autism Spectrum Disorders Working Group of The Psychiatric Genomics Consortium. Meta-analysis of GWAS of over 16,000 individuals with autism spectrum disorder highlights a novel locus at 10q24.32 and a significant overlap with schizophrenia. Mol Autism 8, 21 (2017).
Meng, X. et al. Multi-ancestry genome-wide association study of major depression aids locus discovery, fine mapping, gene prioritization and causal inference. Nat. Genet. 56, 222–233 (2024).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Oldham, M. C., Langfelder, P. & Horvath, S. Network methods for describing sample relationships in genomic datasets: application to Huntington’s disease. BMC Syst. Biol. 6, 63 (2012).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Jiang, Y. et al. DRAMS: A tool to detect and re-align mixed-up samples for integrative studies of multi-omics data. PloS Comput. Biol. 16, e1007522 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Delaneau, O. et al. A complete tool set for molecular QTL discovery and analysis. Nat. Commun. 8, 15452 (2017).
Pierce, B. L. & Burgess, S. Efficient Design for Mendelian Randomization Studies: Subsample and 2-Sample Instrumental Variable Estimators. Am. J. Epidemiol. 178, 1177–1184 (2013).
Yavorska, O. & Burgess, S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol. 46, 1734–1739 (2017).
Terza, J. V., Basu, A. & Rathouz, P. J. Two-stage residual inclusion estimation: Addressing endogeneity in health econometric modeling. J. Health Econ. 27, 531–543 (2008).
Acknowledgements
We thank Majd Alsayed, Miguel Brown, Dominic Fitzgerald, Amber Thomas, and Mimi Brown for their assistance with RNA-Seq data production and processing. We thank Richard Kopp at SUNY Upstate Medical University for his help with wordsmithing. We thank all of the donors and their relatives who participated in the brain collections at Stanley Medical Research Institute and Banner Sun Health Research Institute. We thank Drs. Maree J. Webster and Thomas Beach for their support to make the tissue available. We thank the High Performance Computing Center of Central South University for partial support of the calculation. This project was conducted as part of the PsychENCODE Consortium, which is responsible for the maintenance and distribution of the data used in this project. This project has received funding from National Key Research and Development Program of China (2024YFA1108000) (C.C.), National Natural Science Foundation of China (Grants No. 82022024) (C.C.), SUNY Empire Innovation Program (C.L.), National Institutes of Health grants [U01MH122591 (C.L.), U01MH116489 (K.P.W.), R01MH110920 (C.L.), R01MH126459 (C.L.), U01MH103340 (C.L., K.P.W.), R01MH109715 (D.P.), R01MH110555 (D.P.), R21MH129817 (D.P.), R01GM139980 (S.H.W.)].
Author information
Authors and Affiliations
Contributions
C.L., K.P.W., J.P., S.H.W., B.L., X.W., and C.C. were responsible for Conceptualization. A.W.S., L.C., D.Z., R.C., F.W., M.X., M.N., D.P., Y.W., R.V., C.Z., K.G., and G.G. were responsible for sample collection, DNA extraction, mRNA extraction, mRNA binding with ribosome extraction, protein extraction, sequencing, mass spectrum, and raw data processing. S.H.W., Q.L., Y.J., C.C., and A.W.S. were responsible for the investigation of analysis. Q.L. was responsible for visualization of results. S.H.W., C.L., C.C., B.L., and K.P.W. were responsible for supervision. S.H.W., Q.L., Y.J., and C.L. were responsible for original draft writing. S.H.W., C.L., C.C., and Q.L. were responsible for review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xin He, Atsushi Takata and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, Q., Jiang, Y., Shieh, A.W. et al. Impact of common variants on brain gene expression from RNA to protein to schizophrenia risk. Nat Commun 16, 10773 (2025). https://doi.org/10.1038/s41467-025-65818-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65818-5
This article is cited by
-
Functional implications of polygenic risk for schizophrenia in human neurons
Nature Communications (2026)