Abstract
Succinate: menaquinone oxidoreductase (SQR) couples the oxidation of succinate with the reduction of menaquinone (MK) as part of the TCA cycle and the aerobic respiratory chain in MK-containing bacteria and archaea. Despite its significance, questions persist regarding the electron and proton transfer mechanisms that drive the endergonic MK reduction by succinate. In this study, we determine cryo-EM structures of succinate dehydrogenase (SDH) from Chloroflexus aurantiacus (CaSDH), a facultative filamentous anoxygenic phototroph (FAP) that forms one of the earliest branches of photosynthetic bacteria. The structures of trimeric CaSDH, resolved in both apo- and MK-bound forms, reveal a single membrane-anchoring subunit containing two b-type hemes, a canonical QP site, and a QD site with atypical location, configuration and specificity, each bound to MK molecules. Using structural analysis, EPR, and enzymatic assays, we uncover electron transfer pathways connecting succinate oxidation to MK reduction at the QP and QD sites. These findings provide structural insights into the electron and proton transfer mechanisms of MK-dependent diheme SQRs and establish a foundation for structure-based inhibitor design and antibacterial drug development targeting these enzymes.
Similar content being viewed by others
Introduction
Succinate: quinone oxidoreductase (SQR, EC 1.3.5.1) and quinol: fumarate reductase (QFR) are closely related enzymes that couple the oxidation of succinate to fumarate with the reduction of quinone to hydroquinone (quinol) as well as the reverse reaction1,2. These enzymes are found across all kingdoms of life and play key roles in aerobic and anaerobic metabolic pathways3. SQR, also known as mitochondrial Complex II, participates in aerobic metabolism as part of the tricarboxylic acid (TCA) cycle and respiratory chain, spanning bacteria to mammals4. In contrast, QFR supports bacterial anaerobic respiration by using fumarate as the terminal electron acceptor to oxidize various donor substrates (e.g., NADH, H2, or formate)5. Defects in SQRs and QFRs are associated with impaired electron transfer, metabolic disorders, and overproduction of reactive oxygen species (ROS) in mitochondria, causing neurodegenerative diseases and tumor development in human6,7,8,9,10. These enzymes also serve as potential therapeutic targets in human pathogens like Mycobacterium smegmatis, Helicobacter pylori, and Campylobacter jejuni11,12,13.
While both SQRs and QFRs share a conserved architecture of a soluble domain and a membrane domain, their roles diverge significantly based on cellular context and metabolic demands14,15,16,17,18,19,20,21,22. The soluble domain includes two hydrophilic subunits: a flavoprotein harboring a covalently attached flavin adenine dinucleotide (FAD) cofactor at the succinate-fumarate interconversion site, and an iron-sulfur protein coordinating three iron-sulfur clusters ([2Fe-2S], [4Fe-4S], and [3Fe-4S])14,15,16,17,18,19,20,21,22. Depending on the reaction direction, this soluble domain functions either as succinate dehydrogenase (SDH) or fumarate reductase (FRD). The membrane domain, exhibiting greater variability among species, contains one or two membrane-anchoring subunits with b-type hemes and at least two binding sites for quinones (a “proximal” QP site and a “distal” QD site)14,18,19.
SQRs and QFRs are categorized into three functional classes based on quinone composition and in vivo roles1,23. Class 1 includes SQRs from mammalian mitochondria and many prokaryotes, which catalyze an electrogenic reaction of coupling succinate oxidation to high-potential quinones like ubiquinone (UQ)15,16,17,20,21. Class 2 comprises QFRs, which favorably oxidize low-potential menaquinol (MKH2) or rhodoquinol by fumarate14,18. Class 3 enzymes couple succinate oxidation to the reduction of low-potential MK in an endergonic reaction driven by proton potential. These include the diheme SQRs from Gram-positive bacteria (e.g. Bacillus subtilis24, Bacillus licheniformis25, and Mycobacterium smegmatis19) and sulfur-reducing δ-proteobacteria (e.g. Desulfovibrio vulgaris26 and Geobacter sulfurreducens27) that contain MK as the respiratory quinone. While structural studies of M. smegmatis Sdh2 have revealed features of a three-subunit membrane-anchoring domain, such as its two b-type hemes, two QD sites, and a QP site that is blocked by SdhF19, simpler diheme SQRs with a single membrane-anchoring subunit remain poorly understood.
The bacterium Chloroflexus aurantiacus represents an excellent model for exploring these mechanisms. As a member of the green filamentous anoxygenic phototrophs (FAPs), this organism contains only MK (predominantly MK8 and MK10) and is metabolically versatile in adapting to aerobic and anaerobic conditions28,29,30. During aerobic respiration, C. aurantiacus uses the TCA cycle to produce NADH for the respiratory electron transfer, while under anaerobic light conditions, it employs the 3-hydroxypropionate (3-HP) cycle for autotrophic carbon fixation31,32. A shared step between these pathways involves the conversion of succinyl-CoA to malate, where succinate is oxidized by a diheme SQR named CaSDH, which contains a single membrane-anchoring subunit with two b-type hemes (Fig. 1a). Proteomic time-course analyses suggest that CaSDH is constitutively expressed during transitions between chemoheterotrophic respiratory growth and photoheterotrophic growth, indicating its dual function under aerobic and anaerobic conditions31. Like the diheme SQRs from B. subtilis, B. licheniformis, and G. sulfurreducens, CaSDH catalyzes both succinate oxidation and fumarate reduction in vitro33. However, the architecture and coupling mechanism of this enzyme are unknown.
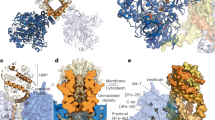
a Diagram illustrating the CaSDH activity in coupling succinate oxidation and menaquinone (MK) reduction in the respiratory chain and tricarboxylic acid (TCA) cycle under aerobic conditions, as well as the autotrophic 3-hydroxypropionate (3-HP) cycle under anaerobic conditions. Electron transfer and proton translocation between respiratory complexes are omitted for simplicity. b EPR characterization of CaSDH in the air-oxidized (black trace) and dithionite-reduced (red trace) states, and in the presence of succinate alone (blue trace) or succinate with MK (magenta trace). The peak at g = 4.29 likely attributed to mononuclear high-spin Fe3+ ion in sites of low symmetry, which is commonly found in a variety of non-heme iron metalloproteins. To highlight signals in the low-field region, the signals between 900 and 2300 Gauss were expanded, and the signal intensities of three traces were multiplied by a factor of 2 for comparison. EPR conditions: temperature, 15 K; microwave power, 10 mW; modulation amplitude, 10 G. c Cryo-EM map of CaSDH shown from the front (left) and bottom (right) views, illustrating the dimensions of the complex. SdhA, SdhB, and SdhC are colored wheat, sea green, and slate blue, respectively. d, e Cartoon representations of the apo-form (d) and the MK-bound form (e) of CaSDH. Cofactors and lipids are shown as sticks and color-coded as follows: flavin adenine dinucleotide (FAD), blue; succinate, orange; iron-sulfur clusters, orange-brown; b-type hemes, red; phosphatidylglycerol (PG), brown; phosphatidylethanolamine (PE), cyan; MK in the QP site, violet; MK in the QD site, green.
In this study, we determined cryo-EM structures of CaSDH in its apo- and MK-bound forms with both QP and QD sites bound to MK molecules. Our findings reveal a canonical QP site and a QD site with unusual location, configuration, and specificity, and electron transfer mechanisms of MK reduction at both sites, providing a structural basis for understanding the coupling mechanisms of MK-dependent diheme SQRs.
Results
Purification and characterization of CaSDH as succinate menaquinone oxidoreductase
CaSDH was extracted and purified from phototrophically grown C. aurantiacus J-10-fl cells using a modified protocol based on previously described methods33. Blue native polyacrylamide gel electrophoresis (BN-PAGE) of the purified complex revealed a single band with an estimated molecular weight of ~440 kDa, consistent with a CaSDH trimer (Supplementary Fig. 1a). Analysis by tricine–sodium dodecyl sulfate (SDS)-PAGE indicated three distinct bands corresponding to the subunits SdhA, SdhB, and SdhC (Supplementary Fig. 1b). These subunits were further confirmed by peptide mass fingerprinting (PMF) of protein bands excised from the tricine–SDS-PAGE gel (Supplementary Table 1).
Spectral analysis revealed an absorption peak at 415 nm, corresponding to air-oxidized hemes bound to CaSDH. Reduction with sodium dithionite shifted this peak to 425 nm and produced an additional peak at 555 nm, indicative of heme reduction (Supplementary Fig. 1c). Enzymatic activity was confirmed using the succinate-2,6-dichlorophenolindophenol (DCPIP) assay, with catalytic constant (kcat) and Km values of 1.87 ± 0.12 s-1 and 0.12 ± 0.01 μM, respectively for succinate (Supplementary Fig. 1d). Similar spectral shifts were observed upon the addition of reduced menaquinone-4 (MKH2), indicating that CaSDH is capable of oxidizing MKH2 (Supplementary Fig. 1e). Fumarate reductase activity was also detected, demonstrated by the oxidation of dithionite-reduced benzyl viologen (BV) by fumarate (Supplementary Fig. 1f). The Km value for fumarate is 0.981 ± 0.340 mM, which is higher than the bifunctional QFRs from C. jejuni (with a reported Km value of 0.1 mM)34 and E. coli (with a reported Km value of 20 ± 2 μM)35.
Electron paramagnetic resonance (EPR) analysis of air-oxidized CaSDH exhibited a prominent peak of an oxidized [3Fe-4S]1+ cluster with a g value of 2.01, and two peaks attributed to the gz components of the low-spin heme (g = 3.23) and high-spin heme (g = 5.96) (Fig. 1b). These three EPR signals disappeared upon addition of either sodium dithionite or succinate, indicating the reduction of hemes and [3Fe-4S]1+ cluster after accepting electrons transferred from dithionite or succinate. Concomitantly, EPR signals corresponding to reduced [2Fe-2S]1+ and reduced [4Fe-4S]1+ clusters were detected for dithionite-reduced samples at low temperatures (Fig. 1b and Supplementary Fig. 2). The incubation of air-oxidized CaSDH with succinate, however, failed to reduce the [2Fe-2S]1+ and [4Fe-4S]1+ clusters (Fig. 1b). Subsequent addition of MK restored part of the signals of [3Fe-4S]1+ and high-spin heme (g = 5.96), indicating re-oxidation of the iron-sulfur clusters and hemes once electrons are transferred to MK (Fig. 1b). These results indicate that the b-type hemes and iron-sulfur clusters are redox-active and they likely participate in the electron transfer processes upon succinate oxidation and MK reduction. This is consistent with previously characterized MK-dependent diheme SQRs from M. smegmatis19, B. licheniformis25, and B. subtilis36.
CaSDH composition and overall structure
To investigate the structural basis of CaSDH in catalyzing the succinate oxidation by MK, the purified CaSDH was analyzed with single particle cryo-EM (Supplementary Fig. 3). We first obtained a 2.62 Å cryo-EM map of the “apo-form CaSDH”, which contains an endogenously bound succinate substrate but an empty QP site (Supplementary Fig. 3a). To resolve the MK-binding sites, CaSDH was incubated with MK4 or menaquinone-7 (MK7), yielding cryo-EM structures of the “MK4-bound CaSDH” at 2.90 Å and the “MK7-bound CaSDH” at 2.74 Å, respectively (Supplementary Fig. 4). We also reconstructed a 2.69 Å cryo-EM map of the “lipid-bound CaSDH” by changing the detergent in anion exchange chromatography (Supplementary Fig. 3b). This structure contains a second succinate molecule, and the QP and QD sites were resolved with a dodecyl-β-D-maltopyranoside (DDM) and a phosphatidylethanolamine (PE) molecule, respectively. These final reconstructed cryo-EM maps enabled us to build accurate models of all protein sidechains, cofactors and lipid molecules bound in CaSDH (Supplementary Figs. 5 and 6a).
The overall structure of CaSDH reveals a spike-like homotrimer with dimensions of 125 × 120 × 120 Å (Fig. 1c). The trimeric assembly is similar to those observed in Escherichia coli SQR15 and M. smegmatis Sdh219. Each monomer comprises three subunits: an FAD-binding protein (SdhA), an iron-sulfur protein (SdhB), and a membrane-anchored protein (SdhC) (Fig. 1c, d). On the cytoplasmic side, SdhA binds FAD and a succinate molecule in the apo-form structure, while SdhB accommodates three iron-sulfur clusters: [2Fe-2S], [4Fe-4S], and [3Fe-4S] (Fig. 1d). Within the membrane region, SdhC contains five transmembrane (TM) helices with two b-type hemes, designated as proximal heme bP and distal heme bD (Fig. 1d).
In the apo-form and lipid-bound structures, each SdhC harbors one phosphatidylglycerol (PG) and two PE molecules at the trimer interface. The PG is located at the interface of SdhB and SdhC on the cytoplasmic side, while the two PE molecules are sandwiched between the TM1 and TM2 helices of adjacent SdhC subunits on the periplasmic side (Fig. 1d; Supplementary Fig. 6b, c). In the MK4- and MK7-bound structures, two MK molecules were resolved per subunit: one at the QP site near the [3Fe-4S] cluster and the other at an atypical QD site near the heme bD (Fig. 1d, e). The spatial arrangement of redox centers forms an electron transfer pathway linking the succinate-binding site in SdhA to the MK-binding site in SdhC (Fig. 1d, e).
SdhA structure and the succinate binding pocket conformational changes
SdhA catalyzes the conversion of succinate to fumarate and the reduction of FAD. Similar to previously reported SQRs from Homo sapiens21, Sus scrofa16, E. coli15, and M. smegmatis19, SdhA exhibits a Rossmann-fold architecture comprising four domains: an FAD-binding domain, a capping domain, a helical domain, and a C-terminal domain (Fig. 2a).
a Overall structure of SdhA, which includes an N-terminal extension of two α-helices (N-helix), a FAD-binding domain (beige), a capping domain (golden yellow), a helical domain (light pink), and a C-terminal domain (orange). FAD is shown in blue stick form. b Succinate and FAD binding pocket. Key residues coordinating succinate (orange) and FAD (blue) are labeled and shown in stick form, with hydrogen bonding interactions indicated by dashed lines. c Conformational changes in the succinate-binding site between the apo-form (white), lipid-bound (gray), and MK-bound (wheat) forms. Loop regions (F167GGA170 and S304E305S306) are shown in cartoon form, with coordinating residues Glu-A305 and Ser-A306 shown in stick form. d Superposition of SdhA (wheat) with those from E. coli (PDB code 1NEK, light pink), Homo sapiens (PDB 8GS8, white), Sus scrofa (PDB 1ZOY, gray) SQRs, M. smegmatis Sdh2 (PDB 6LUM, light blue), and D. gigas (PDB 5XMJ, light green) QFR. The superimposed SdhA structures are displayed as surface. N-helix regions are highlighted in cartoon form. e Spatial organization of the SdhA N-helix in the CaSDH trimer (left) and its interaction with adjacent monomers (right). Zoom in view shows interacting residues (stick) and hydrogen bonding interactions (dashed). f Overall structure of SdhB (left) and coordination of iron-sulfur clusters (right). Iron-sulfur clusters are shown as orange-brown spheres, and coordinating cysteine residues are shown in stick form. Edge-to-edge distances are indicated with orange dashed lines.
In the apo-form structure, the succinate substrate is coordinated by hydrogen bonds with the hydroxyl group of Glu-A305 and the O4 atom of the FAD isoalloxazine ring, which is further stabilized by a hydrogen bond with the main chain nitrogen of Gly-A98 (Fig. 2b). The FAD cofactor is immobilized through hydrogen bonding interactions with Ala-A460, Glu-A445, Met-A211, Ser-A61, Gln-A85, and Ser-A93 from the FAD-binding domain. The flavin C8M methyl group is covalently attached to His-A92, a residue strictly conserved in the A subunits of previously reported SQRs (Fig. 2b, Supplementary Fig. 7a). A second succinate molecule is resolved near the S304E305S306 loop in the lipid-bound structure, in which the Glu-A305 hydroxyl group is flipped ~2.9 Å relative to that in the apo-form structure, while the conformation of the FAD isoalloxazine ring is not changed (Fig. 2c). Importantly, the succinate substrate is absent in both the MK4- and MK7-bound structures (Fig. 1e). Compared to the apo-form and MK-bound structure, the MK-bound SdhA exhibited striking conformational changes in the loop regions F167GGA170 and S304E305S306. Notably, the Glu-A305 residue shifted approximately 3.2 Å away from the succinate-binding site in the apo-form structure (Fig. 2c). These changes opened the substrate-binding pocket, which likely accounts for the absence of succinate or the release of fumarate upon MK binding.
Compared to bacterial SQRs, SdhA contains an N-terminal extension of ~50 amino acid residues (Met-A1 to Lys-A53) and an insertion sequence (Lys-A317 to Glu-A330) between the β13 and β14 strands of the capping domain (Supplementary Fig. 7a). The N-terminal extension forms two short α-helices (Arg-A35 to Glu-A42, and Arg-A51 to Lys-A53) linked by loops. This is distinct from E. coli and M. smegmatis SdhA, which both lack an N-terminal extension but instead contain a β-strand at the N-terminus (Fig. 2d). We, therefore, name the N-terminal extension in CaSDH SdhA as “N-helix”. Each N-helix stretches along and tightly embraces the C-terminal domain of SdhA (Fig. 2d). At the trimer interface, each N-helix forms hydrogen bonds with α5 and its connecting loop from the C-terminal domain of an adjacent CaSDH protomer (Fig. 2e).
In contrast, although Homo sapiens and Sus scrofa SQRs also contain an N-terminal extension, these enzymes are monomeric, and their N-terminal extensions (Ser52-Asp63 in Homo sapiens21 and Ile51-Asp63 in Sus scrofa16) remain structurally unresolved (Supplementary Fig. 7a, b). The lack of an N-terminal extension in the trimeric M. smegmatis Sdh219 suggests that the N-helix is a feature specific for CaSDH, contributing to its trimer formation. Additionally, the CaSDH-specific insertion folds into a coiled-coil structure exposed on the exterior surface of each SdhA, lacking functional contacts with adjacent protomers in the CaSDH trimer (Supplementary Fig. 7c).
SdhB is highly conserved with three iron-sulfur clusters
The structure of SdhB, including its three iron-sulfur clusters, aligns with previously resolved SQR and QFR structures14,15,16,19,21,22 (Supplementary Fig. 8a), indicating structural and functional conservation of the iron-sulfur proteins. The N-terminal domain of SdhB is composed of a three-stranded β-sheet and an α-helix, with a [2Fe-2S] cluster covalently coordinated by residues Cys-B58, Cys-B63, Cys-B66, and Cys-B84 (Fig. 2f). The C-terminal α-helical domain contains seven α-helices linked by loops, where the [4Fe-4S] cluster is stabilized by conserved residues Cys-B159, Cys-B162, Cys-B165, and Cys-B224, and the [3Fe-4S] cluster is ligated to residues Cys-B169, Cys-B214, and Cys-B220 (Fig. 2f and Supplementary Fig. 8b). The [2Fe-2S] cluster is positioned ~9.8 Å from the FAD cofactor (Fig. 2f) and is presumed to facilitate electron transfer from the FAD to the [4Fe-4S] cluster. The three iron-sulfur clusters are arranged linearly with edge-to-edge distances of less than 14.0 Å (Fig. 2f), enabling efficient electron orbital coupling and energy resonance37,38.
SdhC possesses a single membrane-anchoring subunit with two b-type hemes
As a member of the type B SQRs, CaSDH contains a single membrane-anchoring subunit, SdhC (Fig. 3a). Each SdhC comprises five TM helices, designated TM1 to TM5, which host two b-type hemes (bP and bD) within the first four-helix bundle (Fig. 3b). The two b-type hemes are arranged perpendicularly, with an edge-to-edge distance of 3.3 Å (Fig. 3b). This configuration is similar to that observed in M. smegmatis Sdh219 and QFRs from W. succinogenes14 and D. gigas18.
a Overview of subunit composition in previously reported SQR and QFR structures. Conserved hydrophilic subunits A and B are shown in gray, and transmembrane (TM) subunits are color-coded. b Cartoon representations of the SdhC structure (left) and the coordination of b-type hemes (right). Proximal heme bP and distal heme bD are shown in red stick form. Edge-to-edge distances and hydrogen bonding interactions are indicated with dashed lines. c Topology diagram of the five transmembrane helices (TM1-TM5) of SdhC. Axial ligands to heme irons are shown with black lines, and residues coordinating propionate groups are shown with dashed lines. d Superposition of SdhC subunit (slate blue) with those from W. succinogenes (PDB 1QLB, orange) and D. gigas (PDB 5XMJ, gold) QFRs. The SdhC subunits (cartoon) and b-type hemes (stick) are shown. e Top view of superimposed TM helices in SdhC with the membrane regions of previously reported SQR structures. Alignment differences in TM3 and M. smegmatis SdhF are noted. f Superposition of SdhC (slate blue) with subunits from M. smegmatis Sdh2 (PDB 6LUM, light green), Sus scrofa SQR (PDB 1ZOY, light blue), and E. coli SQR (PDB 1NEK, light pink). TM3 and other structural features are highlighted. The unaligned TM3 and the parallel helix that blocks the heme bD binding site in M. smegmatis and Sus scrofa SQRs are shown in a cartoon. The b-type hemes and the cardiolipin in E. coli SQR are shown in stick form.
The iron center of heme bP is axially coordinated by His-C81 and His-C188 on TM2 and TM4, while the axial ligands for heme bD are His-C32 and His-C135 on TM1 and TM3, respectively (Fig. 3b, c). The propionate groups of hemes bP and bD are stabilized through hydrogen bonding interactions with Thr-C199/Lys-C15 and Arg-C54/Arg-C71, respectively (Fig. 3b, c). Sequence alignment reveals conservation of the axial ligands for hemes bP and bD, except His-C81, with those in W. succinogenes and D. gigas QFRs, as well as MK-dependent SQRs from B. subtilis, B. licheniformis, D. vulgaris, and G. sulfurreducens (Supplementary Figs. 9 and 10). Structural superpositions of SdhC with those of W. succinogenes14 and D. gigas18 QFRs yield root mean square deviations of 1.356 Å and 1.243 Å, respectively. Striking conformational differences are observed in the N-terminal α-helix (α1) that interacts with SdhB, a horizontal helix on the cytoplasmic side, and the TM2 and TM3 helices on the periplasmic side (Fig. 3d).
SQRs from Homo sapiens21, Sus scrofa16, Gallus gallus17, and E. coli15, as well as the E. coli QFR22, contain two membrane-anchoring subunits (SdhC and SdhD), each with three TM helices. In contrast, M. smegmatis Sdh2 includes an additional subunit, SdhF, which blocks the canonical QP site19 (Fig. 3a). Structural alignment of the transmembrane regions of these SQRs reveals a mismatch in the SdhC TM3 and the single TM helix of SdhF from M. smegmatis Sdh2 (Fig. 3e and Supplementary Fig. 11a). CaSDH SdhC also lacks the N-terminal α-helix (α1) that interacts with SdhB on the cytoplasmic side (Supplementary Fig. 11a). Additionally, a single heme bP has been resolved in UQ-dependent SQRs, in which the heme bD binding site is either occupied by a parallel helix from the SdhD TM3 or replaced by two cardiolipin acyl groups in E. coli SQR15 (Fig. 3f, Supplementary Fig. 11b). Conversely, the heme bD binding site is occupied by a MK7 molecule in E. coli QFR22 (Supplementary Fig. 11c). These observations indicate that CaSDH possesses a single membrane-anchoring subunit, with its subunit composition, heme arrangement, and spatial organization closely resemble those of W. succinogenes14 and D. gigas18 QFRs.
Menaquinone (MK) binding sites
Previous studies of SQRs and QFRs have identified at least two quinone binding sites within the membrane-anchoring subunit16,19,22. A proximal quinone site (QP), located near the [3Fe-4S] cluster, has been identified with bound quinone molecules in UQ-dependent SQR structures15,16,17,21. A second distal quinone binding site (QD) has been identified by the binding of an inhibitor TTFA2 in the crystal structure of Sus scrofa SQR and by a bound MK7 molecule in E. coli QFR22. In M. smegmatis Sdh2, two QD sites have been resolved with bound MK9 molecules: QD1 is positioned ~15 Å from heme bD on the periplasmic side of SdhD, and QD2 is located within the trimer interface19. Although two MK-binding sites have been identified in E. coli QFR, both sites are different from the UQ binding sites in the SQRs22.
Comparison of CaSDH monomer with superimposed SQR structures around the [3Fe-4S] cluster readily revealed the location of the QP site at the SdhB and SdhC interface (Fig. 4a, b). This site was empty in the apo-form and was occupied by a DDM molecule in the lipid-bound structure, but it was resolved with an MK molecule with clear EM densities in both the MK4- and MK7-bound structures (Fig. 4c, Supplementary Fig. 12a). Upon MK binding, no significant conformational changes occurred at the QP site, apart from minor shifts in a solvent-exposed area spanning residues Leu-C96 to Tyr-C119 (Fig. 4d). The MK head group is buried within a pocket composed of residues Pro-B170 and Asn-B171 (SdhB), Leu-C96, Val-C104, Arg-C112, Asp-C118, and Tyr-C119 (SdhC) (Fig. 4c–e, Supplementary Fig. 9). While the alkyl carbon tail of MK aligns along a slot between the TM2 and TM3 helices (Fig. 4c). Key interactions include hydrogen bonding between the O4 carbonyl group and Asp-C118 (2.7 Å), and proximity of the O1 carbonyl group to the Tyr-C119 hydroxyl group (4.7 Å). The amino acid residues Asp-C118 and Tyr-C119 (Asp-D82 and Tyr-D83 in E. coli numbering) are critical for proton transfer and UQ reduction, as demonstrated by site-directed mutagenesis studies39. The spatial organization of the QP site relative to [3Fe-4S] and heme bP resembles that of SQRs from Homo sapiens, Sus scrofa, G. gallus, and E. coli (Fig. 4a, b, e), suggesting a similar mechanism of electron transfer from succinate to the QP site.
a Structural comparison of the CaSDH monomer with previously reported SQRs around the [3Fe-4S] cluster, readily revealing the location of the QP binding site. The monomeric structures of CaSDH (blue) and E. coli (PDB 1NEK, light pink) SQR are shown in cartoon form, while Homo sapiens (PDB 8GS8, white), Sus scrofa (PDB 1ZOY, light blue), and Gallus gallus (PDB 1YQ3, steel blue) SQRs are shown as surface. b Superposition of redox centers and the resolved quinone molecules in SQRs from panel a, representing the conserved spatial organization of iron-sulfur clusters, heme bP, and the QP site. c Enlarged view of the QP site with a bound MK fitted in clear EM densities. The MK molecule (violet) is shown in stick form, and the QP site (blue) is shown as surface with the surrounding amino acid residues labeled. d Comparison of the QP site in apo-form (white) and the MK-bound (blue) structures. Amino acid residues essential for MK binding are shown in stick form and hydrogen bonding interactions are indicated with dashed lines. e Diagram of the MK coordination, showing edge-to-edge distances between the QP site, [3Fe-4S], and heme bP. f Structural superposition of CaSDH with W. succinogenes (PDB 1QLB, orange) and D. gigas (PDB 5XMJ, gold) QFRs. W. succinogenes and D. gigas QFRs both contain an N-terminal α-helix (α1) that interacts with SdhB, and an additional horizontal helix that blocks the equivalent QP site on the cytoplasmic side. On the periplasmic side, two closely positioned TM helices (TM2 and TM3) narrow the equivalent QD site in CaSDH.
A second MK molecule is resolved with clear EM densities in a cleft formed by the TM2 and TM3 helices and their connecting loop (Pro-C154 to Tyr-C159) on the periplasmic side of SdhC (Fig. 5a, b; Supplementary Fig. 12b). The MK head group interacts sterically with Pro-C154, His-C155, and Tyr-C159, while the alkyl tail is extending along a hydrophobic slot formed by residues Ile-C132, Ile-C136, Thr-C140, Phe-C141 (TM3), Val-C78, Leu-C75, Leu-C74, and Trp-C68 (TM2) (Fig. 5a). In the lipid-bound structure, this hydrophobic slot is occupied by one of the fatty acid tails of a PE molecule (Fig. 5a, Supplementary Fig. 12c). MK binding induced a minor shift (~0.7 Å) in the Glu-C64 hydroxyl group, enabling a weak hydrogen bond with the O1 carbonyl group (~3.6 Å) (Fig. 5a, b, Supplementary Fig. 12b). This residue, while critical for MK binding in CaSDH, is not conserved with Glu-C86 of M. smegmatis Sdh2, which mediates proton transfer at the QD1 site19.
a Enlarged view of the QD site with a bound menaquinone-7 (MK7) fitted in clear EM densities. The hemes (red), the MK7 molecule (green), and surrounding residues are shown in stick form, and hydrogen bonding interactions with Glu-C64 are indicated with dashed lines. The QD site (blue) is shown in cartoon form but as surface in the inset. b Structural superposition of the QD site in the apo-form (white) and MK4-bound (blue) structures. The PE molecule (cyan) in the apo-form, the MK4 (dark green), and surrounding amino acid residues are shown in stick form. The QD site is shown in cartoon form but shown as surface in the inset. c Superposition of CaSDH SdhC (slate blue) with E. coli QFR (PDB 1L0V, yellow), revealing overlap of heme bD with MK7-binding site. CaSDH SdhC is shown as surface, the bound PEs (cyan), QP, and QD sites are shown in stick form. The SdhC of E. coli QFR is shown in cartoon form. d, e Superposition of CaSDH SdhC (slate blue) with the membrane anchor of M. smegmatis Sdh2 (PDB 6LUM, light green). d Horizontal helix between the TM1 and TM2 of CaSDH SdhC (cartoon form) blocks the equivalent QD1 site in Sdh2 (surface form). e A PE molecule in CaSDH SdhC (surface form) occupies the equivalent QD2 site of Sdh2 (cartoon form). The bound menaquinone-9 (MK9, yellow) in Sdh2, and PEs (cyan) in CaSDH SdhC are shown in stick form.
Although the QP site is highly conserved with UQ-dependent SQRs, the QD site exhibits distinct localization and specificity compared to previously reported structures15,16,17,20,21. This QD site is located on the opposite side of the E. coli QFR QD site, and it also does not align with the QD1 or QD2 site of M. smegmatis Sdh2 (Fig. 5c, d; Supplementary Fig. 11c, d). Compared to M. smegmatis SdhC and SdhD, CaSDH SdhC contains a horizontal helix connecting the TM1 and TM2 helices on the periplasmic side, it blocks the entry of the equivalent QD1 site in Sdh2 (Fig. 5d, Supplementary Fig. 11d). The equivalent QD2 site is occupied by one fatty acid tail of the PE molecule in both the apo-form and lipid-bound CaSDH (Fig. 5e, Supplementary Fig. 11d). The SdhC architecture and heme composition of CaSDH are identical to those of W. succinogenes14 and D. gigas18 QFRs. In these QFRs, the equivalent QP site is blocked by a horizontal helix connecting the TM2 and TM3 helices on the cytoplasmic side, and the equivalent QD site is narrowed down by the closely positioned TM2 and TM3 helices on the periplasmic side (Figs. 3d and 4f). These findings suggest that CaSDH has evolved a canonical QP site and an unusual QD site, distinct from previously reported SQR and QFR structures.
Electron transfer pathways
The identification of MK binding sites demonstrates potential electron transfer pathways within each CaSDH monomer, which include redox centers such as FAD, [2Fe-2S], [4Fe-4S], [3Fe-4S], two MK molecules at the QP and QD sites, and two b-type hemes (Fig. 6a, Supplementary Fig. 13). Enzymatic and EPR analyses have revealed that the b-type hemes and iron-sulfur clusters in CaSDH are redox-active and likely participating in the electron transfer processes upon succinate oxidation and MK reduction (Fig. 1b). The shortest edge-to-edge distance between redox centers in adjacent CaSDH monomers within the trimer is ~38.0 Å (Supplementary Fig. 13), indicating that electron transfer likely occurs within individual monomers.
a Overview of redox centers in CaSDH trimer. The redox centers are shown in stick form and edge-to-edge distances are indicated with dashed lines. b, c Putative electron transfer pathways within CaSDH. The intra-protein electron tunneling pathways from succinate oxidation to MK reduction at the QD site (b) and QP site (c) were predicted to be present. Through-bond processes and through-space jumps are represented by cyan solid and dashed lines, respectively. The redox centers and the amino acid residues necessary for mediating electron transfer are shown in stick form and labeled with abbreviated subunit names in parentheses. d The total free energies (kJ/mol) of CaSDH binding to MK4 and MK8 at the QP site, at the QD site, and at both the QP and QD sites. Binding free energies were averaged over 20 frames extracted every 0.5 ns between 40–50 ns of a 50 ns trajectory after equilibration. e, f Proposed electron transfer mechanisms of CaSDH in catalyzing the succinate oxidation and MK reduction at the QD (e) and QP (f) sites.
Electrons generated from succinate oxidation are first transferred to FAD for its reduction. The edge-to-edge distances between FAD, [2Fe-2S], [4Fe-4S], and [3Fe-4S] clusters are all less than ~10.6 Å (Fig. 6a), enabling efficient physiological electron transfer37,38. At the [3Fe-4S] cluster, the electron transfer pathway branches, allowing electrons to either reduce MK at the QP site, or transfer via heme bP and bD to reduce MK at the QD site (Fig. 6b, c). To explore the electron transfer pathways within CaSDH, the Pathways plugin in visual molecular dynamics (VMD) was used40,41. Calculations identified a plausible tunneling pathway originating from FAD and proceeding through covalently bound Cys-B58 and Cys-B63 to [2Fe-2S] (Fig. 6b, c). This is consistent with the spatial proximity of the [2Fe-2S] cluster to the FAD cofactor (~9.8 Å). Electrons accepted by [2Fe-2S] are subsequently transferred through Cys-B162, Cys-B165, and Cys-B169 to the [3Fe-4S] cluster, and ultimately to MK in the QP site for reduction (Fig. 6c). For the electron transfer route from [3Fe-4S] to heme bP, the electrons are transferred via Asn-B213 to heme bP and along heme bD to Glu-64, which accepts electrons for reduction of MK at the QD site (Fig. 6b).
In the structure, the edge-to-edge distance from the [3Fe-4S] cluster to the MK O4 carbonyl group at QP site is 9.8 Å, shorter than the 12.6 Å distance to heme bP (Fig. 6a). This arrangement mirrors that of Homo sapiens21, Sus scrofa16, Gallus gallus17, and E. coli15 SQRs (Fig. 4a, b), where electron transfer from [3Fe-4S] (+65 mV in E. coli SQR) to ubiquinone (UQ, +100 mV) is preferred over transfer to the low potential heme bP (+35 mV). While the redox potential of MK ( − 80 mV) is much lower than that of UQ, the redox potentials of hemes bP and bD in CaSDH (−40 mV and −190 mV, respectively)33 are even lower. Although EPR analysis has revealed the participation of the b-type hemes in the electron transfer of succinate oxidation by MK (Fig. 1b), the electron transfer routes and the priority of the two MK-binding sites are unclear.
To distinguish the QP and QD sites in mediating MK binding and reduction, molecular dynamics (MD) simulations were performed for the CaSDH monomer bound to substrates MK4 or/and C. aurantiacus predominant quinone MK8, and cofactors ([3Fe-4S], [2Fe-2S], [4Fe-4S], FAD, and Heme). Due to the differences in the MD starting structures and the bound substrates, as well as the substrate-induced conformational changes, CaSDH in the binding complexes of MK4/8 at the QP site, MK4/8 at the QD site, and MK4/8 at both the QP and QD sites showed different RMSD profiles (Supplementary Fig. 14). But CaSDH together with the bound cofactors showed converged RMSD profiles in the second half of all MD simulations, indicating the simulation process reached the equilibration stage (Supplementary Fig. 14). An MM/PBSA calculation42 revealed that the binding free energies of MK8 or/and MK4 to the QD site are both lower than to the QP site (Fig. 6d, Supplementary Table 3). The binding free energy of MK8 to the QP and QD sites was further decomposed to identify per-residue contributions (Supplementary Fig. 15a, b). Compared to the QP site, the lower binding free energy of MK8 binding to the QD site is contributed by more hydrophobic residues (Supplementary Fig. 15a, b).
Specifically, the hydrophobic alkyl tail of MK8 at the QP site is buried in the phospholipid bilayer of the membrane, with one side contacted by the amino acid residues Thr-C89, Tyr-C119, Phe-C123, Val-C126, and Phe-C130 (Supplementary Fig. 15c). While in the QD site, the MK8 alkyl tail is stabilized in a slot between TM2 and TM3 helices, forming extensive hydrophobic interactions with Trp-C68, Phe-C72, Leu-C74, Leu-C75, Val-C78, Phe-C79, Phe-C129, Phe-C130, Ile-C132, Ile-C133, Ile-C136, Leu-C137, and Phe-C141 (Supplementary Fig. 15d). These hydrophobic interactions help to stabilize the MK8 conformation and give rise to a much lower binding free energy for the QD site than that for the QP site. Therefore, the MD simulations indicate that the QD site is more favorable for MK binding. This aligns with our EPR analysis that the b-type hemes participate in the electron transfer from succinate oxidation to reduce the MK at the QD site of CaSDH (Fig. 1b).
Discussion
Unlike SQRs that catalyze the energetically favorable electrogenic reaction of succinate oxidation by high-potential quinones, class 3 enzymes catalyze succinate oxidation by low-potential MK, an endergonic reaction that remains incompletely understood. In this study, we determined the cryo-EM structures of an unusual MK-dependent SQR, CaSDH, in both the apo- and MK-bound forms, identifying a canonical QP site and a previously unrecognized QD site within a single membrane-anchoring subunit containing two b-type hemes. Through structural, EPR, and enzymatic analyses, we uncovered an uncommon electron and proton transfer mechanism for MK reduction in diheme SQRs. Electrons released from succinate oxidation are transferred sequentially via FAD, three iron-sulfur clusters, and two b-type hemes to the MK bound at the QD site for its reduction. Distinct from previously characterized diheme SQRs and QFRs, CaSDH possesses a less conserved Glu64 residue in the QD site for MK binding and reduction.
The subunit and heme composition of CaSDH closely resemble those of MK-dependent SQRs from Gram-positive bacteria (B. subtilis, B. licheniformis) and sulfur-reducing δ-proteobacteria (D. vulgaris, G. sulfurreducens), all of which contain a single membrane-anchoring subunit with two b-type hemes but unknown quinone binding sites24,25,26,27. Here, we provide the first detailed identification of the location and spatial organization of two MK binding sites in these enzymes. While the QP site conforms to canonical configurations seen in previously studied SQRs15,16,17,21, the QD site is distinct in location and specificity compared to those in E. coli QFR22 and M. smegmatis Sdh219. Sequence alignment revealed that the residues forming the MK binding sites are not conserved in previously studied SQRs and QFRs, but they are conserved in the SQRs and QFRs from the Chloroflexi clade (Supplementary Figs. 9 and 10, and Supplementary Table 4). These analyses indicate that the QD site configuration is specific to Chloroflexi enzymes, which probably employ a MK reduction mechanism different from previously reported SQRs and QFRs. The phylogenetic position of Chloroflexi SQRs (Supplementary Fig. 16) further suggests this QD site could be an adaptation to the physiological conditions.
Recent studies on B. subtilis SQR have demonstrated that the coupling between electron and proton transfer depends on membrane energization and localization of MK/MKH2 binding43. No proton potential is required when MK reduction occurs at the QP site in a de-energized membrane, whereas MK reduction at the QD site operates only in an energized membrane, using energy stored by other sites of the respiratory chain43. Similar to B. subtilis, C. aurantiacus contains a cytochrome aa3 oxidase in the aerobic respiratory chain (Supplementary Fig. 17), it is capable of translocating 2H+/e– to create a higher PMF, which could facilitate MK reduction at the QD site. Although MK molecules were resolved at both QP and QD sites, EPR analyses demonstrated that electrons released during succinate oxidation are transferred from the [3Fe-4S] cluster via b-type hemes to reduce the MK (Fig. 1b). MD simulations further revealed that the QD site is more favorable for MK binding and reduction. This aligns with previous studies that only one of the two identified quinone sites in SQRs and QFRs is typically functional14,18,19.
The MK reduction at the QD site has also been observed in M. smegmatis Sdh219 and diheme SQRs from B. subtilis24 and B. licheniformis25. To drive the endergonic reaction of succinate oxidation by MK, the transmembrane electron transfer via the two b-type hemes is accompanied by a proton potential across the membrane25,44. Although these enzymes have a Glu residue (corresponding to Glu66 in W. succinogenes QFR14) at the QD site for MK reduction, they all lack the residue homologous to Glu180 in W. succinogenes QFR for the E-pathway-producing proton potential45,46,47. Despite this, Bacilli SQRs can drive MK reduction at the QD site using a proton potential across the membrane43,48. In M. smegmatis Sdh2, Glu-C86 near the QD1 site is proposed to function as the proton entry point. The QD2 site, which is located at the trimer interface, may facilitate proton transfer between the ring C-propionate group of the bD heme and the surrounding proton-accepting residues19. In contrast, CaSDH lacks the residues homologous to Glu66 (Arg54 in CaSDH) and Glu180 (Ala186 in CaSDH). Instead, it possesses a less conserved Glu64 residue in an unusual QD site for MK binding and reduction. The lack of an E-pathway Glu residue suggests MK reduction at the QD site in CaSDH requires an electrochemical proton potential across the membrane (Fig. 6e). This structural arrangement differentiates CaSDH from previously characterized diheme SQRs.
In B. subtilis SQR, MK reduction at the QP site occurs only when the QD site is blocked by the inhibitor 2-n-heptyl-4 hydroxyquinoline N-oxide (HQNO) or when the membrane is deenergized without the formation of proton potential36,48. During MK reduction at the QP site, both protons and electrons originate from the cytoplasmic side of the membrane43,44,48. Considering the edge-to-edge distance from the [3Fe-4S] cluster to the MK at the QP site is shorter than that to the heme bP (Fig. 6a), we cannot exclude the possibility of MK reduction at the QP site of CaSDH. In the QP site, the MK head group is coordinated by Asp-C118 and Tyr-C119, which undergo sidechain conformational change upon MK binding (Fig. 3c), indicating their essential roles in MK binding and reduction. As the QP site is near the cytoplasmic side, these residues are well-positioned to accept protons from the cytoplasm, contributing to drive the endergonic reduction of MK (Fig. 6f).
Considering only one of the two quinone sites is functional, the two MK molecules bound to CaSDH exhibit distinct redox states upon succinate oxidation and MK reduction: the MK at the QP site is fully reduced after accepting electrons from succinate oxidation, whereas the MK at the QD site remains oxidized. These analyses indicate that the two MK binding sites differ in their affinity for oxidized and reduced substrates, allowing them to adapt to specific physiological conditions. For instance, under abiotic stresses such as high light conditions that de-energize the photosynthetic membrane, MK reduction can occur at the QP site without consuming proton potential. Conversely, under aerobic conditions, the QD site is used for MK reduction, driven by a transmembrane electrochemical proton potential established by the respiratory chain (Fig. 6e, f).
This study reveals the structural basis for a noncanonical diheme SQR that catalyzes the endergonic reaction of succinate oxidation by MK. Similar to CaSDH, the QFRs from human pathogens Helicobacter pylori, which causes gastritis and peptic ulcer diseases, and Campylobacter jejuni, which causes gastroenteritis and Guillain–Barré syndrome, both contain two hydrophilic subunits and a single membrane-anchoring subunit with two b-type hemes11,34,49. However, their MK binding and reduction mechanisms remain unknown. Given the structural and functional similarities between CaSDH and diheme SQRs and QFRs from MK-utilizing bacteria and archaea, our findings not only advance our understanding of electron and proton transfer mechanisms in MK-dependent SQRs and QFRs, but also provide a foundation for structure-based inhibitor design and the development of antibacterial and fungicidal agents.
Methods
Extraction and purification of Chloroflexus aurantiacus succinate dehydrogenase (CaSDH)
Chloroflexus aurantiacus J-10-fl cells were cultured anaerobically at 50 °C under 180 μmol m-2 s-1 illumination in a modified PE medium33. The cells were collected by centrifugation at 9874 × g for 10 min, and were resuspended in buffer containing 50 mM Tris-HCl pH 8.0 (buffer A) and then homogenized using a high-pressure homogenizer (Union, China). The cell extracts were centrifuged at 23, 807 × g for 15 min to remove unbroken cells and insoluble lipids. The whole membrane components were harvested by centrifugation at 200,000 × g for 2 h (Ti 70 rotor, 45,000 rpm) at 4 °C.
The pellets were resuspended in buffer A and solubilized by adding 3% Triton X-100 at room temperature for 1 h with gentle stirring and then were ultra-centrifuged at 200,000 × g for 1 h at 4 °C. The supernatant containing CaSDH was filtered through a 0.22 μm Millipore filter and diluted with the 5-fold volume of buffer A. Then the diluted supernatant was subsequently loaded on a prepacked anion exchange chromatography column (Hitrap Q HP, GE Health) that had been pre-equilibrated with buffer A containing 0.25% Triton X-100 (buffer B). The “lipid-bound” CaSDH was loaded on a prepacked anion exchange chromatography column (Hitrap Q HP, GE Health) that had been pre-equilibrated with buffer A containing 0.1% DDM (buffer C). The “apo-form” CaSDH and “lipid-bound” CaSDH was eluted from the column with 0.2 M NaCl in buffer B and buffer C, respectively. And further purified by gel filtration through a Uniodex 200 PG 16/60 size-exclusion column (Union, China) in buffer containing 25 mM Tris-HCl pH 8.0, 150 mM NaCl, 0.02% n-Dodecyl-β-D-maltopyranoside (DDM, Anatrace). The whole preparation procedure was monitored by measuring the absorption spectrum from 250 to 900 nm. The fractions verified by blue-native PAGE and SDS-PAGE were used for cryo-EM analyses and the other experiments.
Peptide mass fingerprinting (PMF) analysis of CaSDH
The identity of SDS-PAGE and blue-native PAGE bands was confirmed by peptide mass fingerprinting (PMF) using MALDI-TOF mass spectroscopy. Each gel band was analyzed once. The stained bands from the SDS-PAGE and blue-native PAGE were excised, destained and washed with 50% acetonitrile (v/v) in 50 mM aqueous NH4HCO3 until clear. To reduce the proteins, the gel pieces were dehydrated with 100 μL of 100% acetonitrile for 5 min and then were incubated with 10 mM dithiothreitol in 100 mM NH4HCO3 at 56 °C for 60 min. Then cysteine residues in the proteins were further alkylated by 55 mM iodoacetamide in 100 mM NH4HCO3 at room temperature in dark for 45 min. Then the gel pieces were washed with 50 mM NH4HCO3 and dehydrated with 100% acetonitrile, the proteins were digested with 10 ng/μL trypsin in 50 mM NH4HCO3 at 37 °C overnight. The reaction was terminated with 1% trifluoroacetic acid, then the peptides were extracted with 50% acetonitrile/5% formic acid before being dried by 100% acetonitrile and resuspended in 2% acetonitrile/0.1% formic acid.
The tryptic peptides were dissolved in 0.1% formic acid (solvent A) and directly loaded onto a home-made reversed-phase analytical column (15-cm length, 75 μm i.d.). Then the peptides were separated by a gradient comprised of an increase from 6% to 23% solvent B (0.1% formic acid in 98% acetonitrile) over 16 min, 23% to 35% in 8 min and climbing to 80% in 3 min, and then was holding at 80% for the last 3 min were used to separate these peptides, at a constant flow rate of 300 nL/min on an EASY-nLC 1000 UPLC system (Thermo Fisher Scientific, USA). The peptides were then subjected to NSI source followed by tandem mass spectrometry (MS/MS) in HFX (Thermo Fisher Scientific, USA) coupled online to the UPLC. The electrospray voltage applied was 2.0 kV. The m/z scan range was 350 to 1800 for full scan, and intact peptides were detected in the Orbitrap (Thermo Fisher Scientific, USA) at a resolution of 70,000. Peptides were then selected for MS/MS using NCE setting as 28 and the fragments were detected in the Orbitrap at a resolution of 17,500. A data-dependent procedure that alternated between one MS scan followed by 20 MS/MS scans with 15.0 s dynamic exclusion. Automatic gain control (AGC) was set at 5E4. The resulting MS/MS data were processed using MaxQuant Version 2.0.150,51. Carbamidomethyl on Cys were specified as fixed modification and oxidation on Met, deamidation (NQ) and acetyl (Protein N-term) were specified as variable modifications. Min. delta score for modified peptides was set 8. Other parameters were set to the default setting.
Cryo-electron microscopy
3 μL aliquots of CaSDH at 10 mg/mL were applied to freshly glow-discharged GiG R1.2/1.3 300-mesh gold holey nickel titanium grid (Zhenjiang Lehua Technology Co. Ltd., China). Each grid was blotted for 3 s with a blot force of 2 under 100% humidity at 4 °C before being plunge-frozen in liquid ethane with a Mark IV Vitrobot system (Thermo Fisher/FEI). To obtain the MK4-bound and MK7-bound CaSDH, the purified CaSDH was incubated with MK4 or MK7 at a molar ratio of 1:10 before preparing the cryo-EM grids.
Cryo-EM images of the apo-form CaSDH and the lipid-bound CaSDH were recorded on a 300 kV Titan Krios microscope (FEI) equipped with a Falcon 4 detector (Thermo Fisher Scientific, USA) in electron counting mode. Image stacks were automatically collected using EPU software (Thermo Fisher Scientific, USA). A calibrated magnification of 130,000× was used for imaging, corresponding to a pixel size of 0.93 Å. Each movie was dose-fractionated to 40 frames with a total dose rate of 50 e− Å-2 and an exposure time of 6 s in a defocus range from −0.8 to −1.6 μm. In total, 2028 movies of the apo-form and 2624 movies of the lipid-bound datasets were collected. Further details are given in Supplementary Table 2.
Cryo-EM images of the MK-bound CaSDH were recorded on a 300 kV Titan Krios microscope (FEI) equipped with a Falcon 4 detector (Thermo Fisher Scientific, USA) in electron counting mode. Image stacks were automatically collected using EPU software (Thermo Fisher Scientific, USA). A calibrated magnification of 120,000× was used for imaging, corresponding to a pixel size of 0.64 Å. Each movie was dose-fractionated to 40 frames with a total dose rate of 50 e− Å-2 and an exposure time of 4.1 s in a defocus range of −0.8 to −1.6 μm. In total, 6954 movies of the MK4-bound and 2364 movies of the MK7-bound datasets were collected. Further details are given in Supplementary Table 2.
Image processing
Motion correction and contrast transfer function (CTF) estimation were performed using CryoSPARC v3.3.252. For the apo-form and lipid-bound datasets, 867,865 and 242,221 particles were automatically selected from 2028 and 2624 micrographs, respectively, and were extracted and subjected to two-dimensional (2D) classifications. After several iterations of 2D, 77,387 and 82,226 particles were selected to generate the initial model in 3D ab initio reconstruction, respectively. Several iterations of homogeneous refinement and non-uniform refinement53 using 51,634 and 69,415 particles generated the maps with overall resolutions at 2.62 Å and 2.69 Å, respectively, based on the gold standard Fourier shell correlation (FSC) cut-off of 0.14354. Local resolution was estimated with Resmap55.
For the MK-bound datasets, 1,671,875 particles and 555,591 particles were automatically selected from 6954 and 2364 micrographs of the MK4-bound and the MK7-bound CaSDH, respectively, then were extracted and subjected to two-dimensional (2D) classifications. After several iterations of 2D, 78,470 and 32,964 particles were selected to generate initial model in 3D ab initio reconstruction, respectively. Several iterations of homogeneous refinement and non-uniform refinement53 of 44,018 and 25,554 particles generated the maps with overall resolutions at 2.90 Å and 2.74 Å, respectively, based on the gold standard Fourier shell correlation (FSC) cut-off of 0.14354. Local resolution was estimated with Resmap55.
Model building, refinement, and validation
The atomic model of apo-form CaSDH was built by fitting M. smegmatis Sdh2 structure19 (PDB: 6LUM) into the cryo-EM density map in UCSF Chimera56. Based on the density map, the structure model of CaSDH, including the amino acid residues, and cofactors were manually built and adjusted in Coot57. Using the refined model of apo-form CaSDH as a reference, the structures of the MK-bound and lipid-bound CaSDH were manually built in Coot. Then, real-space refinement in PHENIX58,59 was used for model refinement with intra-cofactor and protein-cofactor geometric constraints. The refinement statistics are summarized in Supplementary Table 2. All figures were drawn in PyMOL (The PyMOL Molecular Graphics System, Version 2.5.2, Schrödinger, LLC.), UCSF chimera60 or ChimeraX61.
Enzymatic analyses of CaSDH
The succinate: quinone oxidoreductase activity of CaSDH was determined using a succinate-2,6-dichlorophenolindophenol (DCPIP) assay according to the method previously described35. The reaction was carried out with 0.05 mg/mL CaSDH, 0.025 to 1.6 mM succinic acid, 0.1 mM MK4, and 100 μM DCPIP in buffer C. The reaction was initiated by the addition of DCPIP, and all assays were performed at 37°C. Data and figures were processed with GraphPad Prism 9.
The fumarate reductase activity of CaSDH was measured using benzyl viologen-linked reductase assays as described previously49. The N2 gas was flushed through a quartz cuvette, and the reagents were added with a syringe through the stopper. The reaction mixture contained 75 mM sodium phosphate buffer (pH 6.8), 0.2 mM benzyl viologen, and 0.075 mg/mL CaSDH. Then, the freshly prepared sodium dithionite at 20 mM concentration was injected into the cuvette until the absorbance of reduced benzyl viologen at 585 nm reached 0.9 to 1.0, representing production of the half-reduced benzyl viologen. The reaction was initiated by adding a deoxygenized solution containing 0.25, 0.5, 1, 2.5, 5 mM sodium fumarate, and the absorbance of the reduced benzyl viologen (8.65 cm-1·mM-1) at 585 nm was recorded for 560 s. All the enzymatic data were obtained from triplicate experiments.
Electron paramagnetic resonance (EPR) measurements
Low-temperature electron paramagnetic resonance (EPR) spectrum were acquired using a Bruker X-band (9.4 GHz) EMX plus 10/12 spectrometer equipped with an Oxford Instruments ESR 910 continuous helium-flow cryostat19,62. A cylindrical resonator (ER4119hs TE011) was used for EPR data collection. The purified CaSDH sample was placed into quartz EPR tubes (Wilmad, 707-SQ-250 M, 3 mm i.d., 4 mm o.d.) for measurements. The temperature was set at 15 K for EPR characterization of the metal centers. Other experimental conditions were set as following: microwave power, 10 mW; modulation amplitude, 10 Gauss; modulation frequency, 100 kHz; resonance frequency, ~9.404 GHz. For each sample, multiple scans were accumulated to obtain a good S/N ratio. All spectra were normalized according to their respective scan numbers before comparison.
Molecular dynamics simulation and binding free-energy calculation
Molecular dynamics (MD) models were built from the atomic model of the MK-bound CaSDH monomer with cofactors ([3Fe-4S], [2Fe-2S], [4Fe-4S], FAD, and Heme). Topologies for the cofactors were taken verbatim from the high-level quantum/RESP parameter set reported by Carvalho et al.63, and combined with the CHARMM36m64 force-field for the protein. Two menaquinone analogues—MK4 and MK8—were introduced into the system by aligning their coordinates to the corresponding MK molecule in the reference structure. The parameters of the cofactors were generated using CGenFF65 (version 4.6), and penalty scores were manually curated to ensure accuracy. MD simulations were performed on the binding structures in a membrane system constructed with 70% POPE and 30% POPG using the CHARMM-GUI66,67,68; overlapping lipid and water atoms were deleted and the system was neutralized with 150 mM NaCl. Six simulation boxes were constructed: (i) MK4 bound at QD and QP sites, (ii) MK4 at QD site only, (iii) MK4 at QP site, (iv) MK8 at QD and QP sites, (v) MK8 at QD site, and (vi) MK8 at QP site. Each simulation system was placed in a box and then surrounded by explicit water molecules modeled using the TIP3P water model69.
All MD runs were performed with GROMACS 202370,71 at 323 K (50 °C) and 1 bar using the Nosé–Hoover thermostat (τ = 1 ps) and Parrinello–Rahman barostat (τ = 5 ps, semi-isotropic). After steepest-descent minimization, each system underwent 1 ns NVT and 5 ns NPT restraints, followed by 50 ns of production with 2 fs time-steps and PME electrostatics (1.2 nm cut-off). Trajectories were analyzed for backbone RMSD, per-residue RMSF, hydrogen-bond persistence. Binding free energetics were quantified every 50 ps with the single-trajectory MM/PBSA protocol in g_mmpbsa42 and decomposed into per-residue contributions; the 20 largest |ΔG| residues for each system were visualized with matplotlib bar-plots. The calculated binding free energies are summarized in Supplementary Table 3, and the system setup statistics of MD simulations are summarized in Supplementary Table 5.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM maps and atomic coordinates of Chloroflexus aurantiacus succinate dehydrogenase (CaSDH) in the apo-form, lipid-bound, MK7-bound, MK4-bound forms have been deposited into the Electron Microscopy Data Bank (accession codes, EMD-62932, EMD-62933, EMD-62934, and EMD-62935) and the Protein Data Bank (PDB) (accession codes, 9LAY, 9LAZ, 9LB0, and 9LB1), respectively. The final output coordinate files of MD simulations have been provided as Supplementary Data 1. PDB codes of previously published structures used in this study are 1NEK, 1ZOY, 8GS8, 6LUM, 5XMJ, 1QLB, 1YQ3 and 1L0V. Source data are provided as a Source Data file. Other data are available from the corresponding authors upon request. Source data are provided with this paper.
References
Hägerhäll, C. Succinate: quinone oxidoreductases. Var. a Conserve. theme Biochim Biophys. Acta 1320, 107–141 (1997).
Saraste, M. Oxidative phosphorylation at the fin de siècle. Science 283, 1488–1493 (1999).
van Hellemond, J. J., van der Klei, A., van Weelden, S. W. & Tielens, A. G. Biochemical and evolutionary aspects of anaerobically functioning mitochondria. Philos. Trans. R. Soc. Lond. B Biol. Sci. 358, 205–213 (2003).
Hederstedt, L. Structural biology. Complex II is complex too. Science 299, 671–672 (2003).
Kröger, A. Fumarate as terminal acceptor of phosphorylative electron transport. Biochim Biophys. Acta 505, 129–145 (1978).
Dröse, S. Differential effects of complex II on mitochondrial ROS production and their relation to cardioprotective pre- and postconditioning. Biochim Biophys. Acta 1827, 578–587 (2013).
Siebels, I. & Dröse, S. Q-site inhibitor induced ROS production of mitochondrial complex II is attenuated by TCA cycle dicarboxylates. Biochim Biophys. Acta 1827, 1156–1164 (2013).
Moosavi, B., Zhu, X. L., Yang, W. C. & Yang, G. F. Molecular pathogenesis of tumorigenesis caused by succinate dehydrogenase defect. Eur. J. Cell Biol. 99, 151057 (2020).
Jodeiri Farshbaf, M. & Kiani-Esfahani, A. Succinate dehydrogenase: Prospect for neurodegenerative diseases. Mitochondrion 42, 77–83 (2018).
Esteban-Amo, M. J., Jiménez-Cuadrado, P., Serrano-Lorenzo, P., de la Fuente, M. & Simarro, M. Succinate dehydrogenase and human disease: novel insights into a well-known enzyme. Biomedicines 12, 2050 (2024).
Mendz, G. L., Hazell, S. L. & Srinivasan, S. Fumarate reductase: a target for therapeutic intervention against Helicobacter pylori. Arch. Biochem Biophys. 321, 153–159 (1995).
Cook, G. M., Greening, C., Hards, K. & Berney, M. Energetics of pathogenic bacteria and opportunities for drug development. Adv. Micro. Physiol. 65, 1–62 (2014).
van der Stel, A. X. et al. The Campylobacter jejuni RacRS system regulates fumarate utilization in a low oxygen environment. Environ. Microbiol 17, 1049–1064 (2015).
Lancaster, C. R., Kröger, A., Auer, M. & Michel, H. Structure of fumarate reductase from Wolinella succinogenes at 2.2 A resolution. Nature 402, 377–385 (1999).
Yankovskaya, V. et al. Architecture of succinate dehydrogenase and reactive oxygen species generation. Science 299, 700–704 (2003).
Sun, F. et al. Crystal structure of mitochondrial respiratory membrane protein complex II. Cell 121, 1043–1057 (2005).
Huang, L. S. et al. 3-nitropropionic acid is a suicide inhibitor of mitochondrial respiration that, upon oxidation by complex II, forms a covalent adduct with a catalytic base arginine in the active site of the enzyme. J. Biol. Chem. 281, 5965–5972 (2006).
Guan, H. H. et al. Structural insights into the electron/proton transfer pathways in the quinol:fumarate reductase from Desulfovibrio gigas. Sci. Rep. 8, 14935 (2018).
Gong, H. et al. Cryo-EM structure of trimeric Mycobacterium smegmatis succinate dehydrogenase with a membrane-anchor SdhF. Nat. Commun. 11, 4245 (2020).
Zhou, X. et al. Architecture of the mycobacterial succinate dehydrogenase with a membrane-embedded Rieske FeS cluster. Proc. Natl Acad. Sci. USA 118, e2022308118 (2021).
Du, Z. et al. Structure of the human respiratory complex II. Proc. Natl Acad. Sci. USA 120, e2216713120 (2023).
Iverson, T. M., Luna-Chavez, C., Cecchini, G. & Rees, D. C. Structure of the Escherichia coli fumarate reductase respiratory complex. Science 284, 1961–1966 (1999).
Lancaster, C. R. & Kröger, A. Succinate: quinone oxidoreductases: new insights from X-ray crystal structures. Biochim Biophys. Acta 1459, 422–431 (2000).
Hägerhäll, C., Aasa, R., von Wachenfeldt, C. & Hederstedt, L. Two hemes in Bacillus subtilis succinate:menaquinone oxidoreductase (complex II). Biochemistry 31, 7411–7421 (1992).
Madej, M. G. et al. Experimental evidence for proton motive force-dependent catalysis by the diheme-containing succinate:menaquinone oxidoreductase from the Gram-positive bacterium Bacillus licheniformis. Biochemistry 45, 15049–15055 (2006).
Zaunmüller, T., Kelly, D. J., Glöckner, F. O. & Unden, G. Succinate dehydrogenase functioning by a reverse redox loop mechanism and fumarate reductase in sulphate-reducing bacteria. Microbiol. (Read.) 152, 2443–2453 (2006).
Butler, J. E. et al. Genetic characterization of a single bifunctional enzyme for fumarate reduction and succinate oxidation in Geobacter sulfurreducens and engineering of fumarate reduction in Geobacter metallireducens. J. Bacteriol. 188, 450–455 (2006).
Hale, M. B., Blankenship, R. E. & Fuller, R. C. Menaquinone is the sole quinone in the facultatively aerobic green photosynthetic bacterium Chloroflexus aurantiacus. Biochimica et. Biophysica Acta (BBA) - Bioenerg. 723, 376–382 (1983).
Pierson, B. K. & Castenholz, R. W. A phototrophic gliding filamentous bacterium of hot springs, Chloroflexus aurantiacus, gen. and sp. nov. Arch. Microbiol 100, 5–24 (1974).
Tang, K. H. et al. Complete genome sequence of the filamentous anoxygenic phototrophic bacterium Chloroflexus aurantiacus. BMC Genomics 12, 334 (2011).
Kawai, S., Shimamura, S., Shimane, Y. & Tsukatani, Y. Proteomic time-course analysis of the filamentous anoxygenic phototrophic bacterium, chloroflexus aurantiacus, during the transition from respiration to phototrophy. Microorg. 10 1288, https://doi.org/10.3390/microorganisms10071288 (2022).
Zarzycki, J., Brecht, V., Müller, M. & Fuchs, G. Identifying the missing steps of the autotrophic 3-hydroxypropionate CO2 fixation cycle in Chloroflexus aurantiacus. Proc. Natl Acad. Sci. USA 106, 21317–21322 (2009).
Xin, Y., Lu, Y. K., Fromme, R., Fromme, P. & Blankenship, R. E. Purification, characterization and crystallization of menaquinol:fumarate oxidoreductase from the green filamentous photosynthetic bacterium Chloroflexus aurantiacus. Biochim Biophys. Acta 1787, 86–96 (2009).
Mileni, M. et al. Heterologous production in Wolinella succinogenes and characterization of the quinol:fumarate reductase enzymes from Helicobacter pylori and Campylobacter jejuni. Biochem J. 395, 191–201 (2006).
Maklashina, E. et al. Fumarate reductase and succinate oxidase activity of Escherichia coli complex II homologs are perturbed differently by mutation of the flavin binding domain. J. Biol. Chem. 281, 11357–11365 (2006).
Smirnova, I. A., Hagerhall, C., Konstantinov, A. A. & Hederstedt, L. HOQNO interaction with cytochrome b in succinate:menaquinone oxidoreductase from Bacillus subtilis. FEBS Lett. 359, 23–26 (1995).
Page, C. C., Moser, C. C., Chen, X. & Dutton, P. L. Natural engineering principles of electron tunnelling in biological oxidation-reduction. Nature 402, 47–52 (1999).
Osyczka, A., Moser, C. C., Daldal, F. & Dutton, P. L. Reversible redox energy coupling in electron transfer chains. Nature 427, 607–612 (2004).
Tran, Q. M., Rothery, R. A., Maklashina, E., Cecchini, G. & Weiner, J. H. The quinone binding site in Escherichia coli succinate dehydrogenase is required for electron transfer to the heme b. J. Biol. Chem. 281, 32310–32317 (2006).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 33–38 (1996)
Balabin, I. A., Hu, X. & Beratan, D. N. Exploring biological electron transfer pathway dynamics with the Pathways plugin for VMD. J. Comput Chem. 33, 906–910 (2012).
Kumari, R., Kumar, R., Consortium, O. S. D. D. & Lynn, A. g_mmpbsa A GROMACS tool for high-throughput MM-PBSA calculations. J. Chem. Inf. modeling 54, 1951–1962 (2014).
Azarkina, N. V. Requirement of Bacillus subtilis succinate:menaquinone oxidoreductase activity for membrane energization depends on the direction of catalysis. Biochim Biophys. Acta Bioenerg. 1866, 149522 (2025).
Schirawski, J. & Unden, G. Menaquinone-dependent succinate dehydrogenase of bacteria catalyzes reversed electron transport driven by the proton potential. Eur. J. Biochem. 257, 210–215 (1998).
Lancaster, C. R. et al. Experimental support for the “E pathway hypothesis” of coupled transmembrane e- and H+ transfer in dihemic quinol:fumarate reductase. Proc. Natl Acad. Sci. USA 102, 18860–18865 (2005).
Lancaster, C. R. et al. Essential role of Glu-C66 for menaquinol oxidation indicates transmembrane electrochemical potential generation by Wolinella succinogenes fumarate reductase. Proc. Natl Acad. Sci. USA 97, 13051–13056 (2000).
Madej, M. G., Nasiri, H. R., Hilgendorff, N. S., Schwalbe, H. & Lancaster, C. R. Evidence for transmembrane proton transfer in a dihaem-containing membrane protein complex. EMBO J. 25, 4963–4970 (2006).
Christenson, A., Gustavsson, T., Gorton, L. & Hagerhall, C. Direct and mediated electron transfer between intact succinate:quinone oxidoreductase from Bacillus subtilis and a surface modified gold electrode reveals redox state-dependent conformational changes. Biochim Biophys. Acta 1777, 1203–1210 (2008).
Weingarten, R. A., Taveirne, M. E. & Olson, J. W. The dual-functioning fumarate reductase is the sole succinate:quinone reductase in Campylobacter jejuni and is required for full host colonization. J. Bacteriol. 191, 5293–5300 (2009).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S. et al. Visualization of LC-MS/MS proteomics data in MaxQuant. Proteomics 15, 1453–1456 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
van Heel, M. & Schatz, M. Fourier shell correlation threshold criteria. J. Struct. Biol. 151, 250–262 (2005).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: Phenix.real-space-refine., https://doi.org/10.3390/microorganisms10071288 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl Crystallogr 40, 658–674 (2007).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Gong, H. et al. An electron transfer path connects subunits of a mycobacterial respiratory supercomplex. Science 362, eaat8923 (2018).
Carvalho, A. T. & Swart, M. Electronic structure investigation and parametrization of biologically relevant iron-sulfur clusters. J. Chem. Inf. Model 54, 613–620 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Vanommeslaeghe, K. & MacKerell, A. D. Jr Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model 52, 3144–3154 (2012).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. theory Comput. 12, 405–413 (2015).
Kim, S. et al. CHARMM-GUI ligand reader and modeler for CHARMM force field generation of small molecules. J. Comput Chem. 38, 1879–1886 (2017).
Wu, E. L. et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput Chem. 35, 1997–2004 (2014).
Mark, P. & Nilsson, L. Structure and Dynamics of the TIP3P, SPC, and SPC/E Water Models at 298 K. J. Phys. Chem. A 105, 9954–9960 (2001).
Berendsen, H. J., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys. Commun. 91, 43–56 (1995).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Acknowledgements
This work is supported by grants from the National Natural Science Foundation of China (32471270 and 32171227 to X.L.X., 22437005 to C.L.T.), the Strategic Priority Research Program of Chinese Academy of Sciences (XDB0960201 to L.Y.), Zhejiang Provincial Natural Science Foundation of China (RG26C020001 and LR22C020002 to X.L.X.), Youth Innovation Promotion Association of Chinese Academy of Sciences (2022455 to L.Y.), and Interdisciplinary Research Project of Hangzhou Normal University (2024JCXK04 to X.L.X.). We thank Professor Robert E. Blankenship at Washington University for helpful discussions. We also thank Dr. Shenghai Chang in the Center of Cryo-Electron Microscopy (CCEM), Zhejiang University, for his technical assistance on Cryo-EM data collection. We also thank Shuimu BioSciences for Cryo-EM facility access and technical support during image acquisition. The image processing, model building, structural, and biochemical studies were performed at Hangzhou Normal University.
Author information
Authors and Affiliations
Contributions
X.L.X. initiated the project and supervised all experiments. X.Z. performed the single particle analyses and determined the cryo-EM structures of CaSDH, and assisted in writing the manuscript. J.Y.W. prepared the cryo-EM samples and measured the enzymatic activity of CaSDH. J.M.W. and H.M.H. isolated and purified CaSDH. L.Y. and A.K.L. performed the electron paramagnetic resonance (EPR) and spectroelectrochemical analyses. X.H. and X.J.F. performed and analyzed the MD simulations. Y.Y.Y. and X.K.P. assisted in the sample preparation, spectral, and enzymatic analyses of CaSDH. X.L.X., C.L.T., and Y.Y.X. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stephane Grimaldi, who co-reviewed with Maryam Seif-Eddine; and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Wu, J., Wang, J. et al. Structural basis of menaquinone reduction by succinate dehydrogenase from Chloroflexus aurantiacus. Nat Commun 16, 10782 (2025). https://doi.org/10.1038/s41467-025-65828-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65828-3