Abstract
Developing selective kinase inhibitors is challenging due to the conserved kinase structures and costly kinome profiling experiments, highlighting the need for accurate prediction of kinase-inhibitor affinity and specificity. Here we present MMCLKin, an attention consistency-guided contrastive learning framework that integrates geometric graph and sequence networks with multi-head attention and multimodal, multiscale contrastive learning to accurately and interpretably predict kinase-inhibitor activity and selectivity. MMCLKin outperforms existing methods across two 3D kinase-drug datasets and demonstrates strong generalizability on ten diverse protein-drug and one mutation-aware datasets, and effectively screens on both known and unknown kinase structures. In-depth analysis of attention coefficients reveals that MMCLKin can identify key residues and molecular functional groups critical for kinase-inhibitor binding. Additionally, ADP-Glo assays confirm that five out of 20 MMCLKin-identified compounds inhibit the pathogenic LRRK2 G2019S mutant, with four exhibiting nanomolar-level potency. Collectively, MMCLKin represents a useful tool for discovering potent and selective kinase inhibitors.
Similar content being viewed by others
Introduction
Kinases play essential roles in various biological processes, and their dysregulation is implicated in numerous progressive diseases, including autoimmune disorders, cancer, and neurological conditions. Therefore, protein kinases have emerged as one of the most prominent drug targets of the 21st century1. However, developing highly efficient and selective kinase inhibitors presents considerable challenges due to the high evolutionary conservation of kinase structures, particularly at the ATP binding site2. Many compounds, initially demonstrating promising activity, often fail during preclinical or clinical trials due to off-target effects stemming from low selectivity3. While wet-lab kinome profiling methods can provide multidimensional structure-activity insights across the human kinome, these experiments are highly costly and labor-intensive, limiting their application to evaluating few compounds4. Consequently, developing precise predictive methods for kinome-wide bioactivity profiling is critical for discovering kinase inhibitors with both high selectivity and strong affinity.
Currently, deep learning methodologies have gained increasing prominence in predicting kinase-inhibitor affinity5, with representations primarily divided into sequence-based6 and graph-based approaches7. Sequence-based methods primarily represent drugs and kinases using SMILES notations and kinase sequences, which are readily accessible and abundant. However, since deep learning algorithms inherently rely on pattern matching, sequence-based approaches often suffer from overfitting to incorrect training modes caused by unlimited degrees of freedom8. By contrast, graph-based methods represent biomolecules as 2D or 3D graphs. While 2D graphs encode atomic features, chemical bonds, and adjacency relationships of molecules9, 3D graphs provide a more nuanced representation of molecular reality by incorporating both topological features and spatial conformation characteristics10. Despite these advantages, acquiring 3D kinase structures is highly resource-intensive, resulting in limited sample availability that hampers model performance. Concurrently, the inherent sparsity of graphs may impede the models from fully capturing intricate biomolecular relationships9. In addition, kinase-drug affinity prediction approaches can also be categorized by interaction granularity, typically into global interaction-based10 and local interaction-based methods11. Global interaction-based models focus on drug features, comprehensive protein kinase information, and the broad interactions between drugs and entire kinases. Incorporating 3D graph representations further enhances their ability to capture the intricate spatial relationships governing drug-kinase interactions. However, encompassing extensive kinase information may increase the risk of introducing substantial noise, potentially interfering with model training and diminishing the focus on the determinants of kinase-drug recognition. Conversely, local interaction-based methods emphasize extracting the biochemical features of the key binding site, drug, along with their interaction information. Nevertheless, by concentrating narrowly on local interactions, these methods may overlook critical global characteristics, such as protein folding states, which are intrinsically correlated with kinase functionality.
Building upon the complementary strengths and limitations outlined above, we propose a node-level Multimodal and Multiscale Contrastive Learning with Attention Consistency (MMCLAC) method to effectively integrate the heterogeneous information while fully accounting for the distinct attributes of sequence and graph representations, as well as the hierarchical nature of local interactions within global ones. This approach is grounded in the premise that each compound-kinase system represents an objectively existing entity, inherently encoding its own interaction information alongside a structured and learnable distribution of attention. Specifically, MMCLAC implements a hierarchical contrastive learning paradigm that operates on attention coefficients extracted from dual molecular representations (sequence-based and 3D graph-based modalities) and multi-scale interaction patterns (local atomic-level and global structural-level features) of kinase-inhibitor complexes. This contrastive learning of the attention coefficients between sequence and 3D graph modalities facilitates the concurrent extraction of sequence information, contextual features, and spatial structural characteristics of both kinases and drugs, while alleviating the adverse effects of high degrees of freedom in kinase sequences and graph sparsity on model performance. In parallel, contrasting local and global interaction-based attention encourages the model to emphasize crucial drug-pocket interactions while preserving an overarching awareness of entire kinase information. Notably, MMCLAC enforces attention consistency through node-level contrastive learning. This strategy extends beyond merely aligning attention distributions across different modalities and scales of kinase-drug interactions within the same structural domain. It also empowers the model to discern subtle variations across various systems, ultimately enhancing its capacity to capture the specificity and selectivity of kinase-drug interactions.
In this work, we develop MMCLKin, a comprehensive framework engineered to enhance the prediction of kinase-inhibitor selectivity and binding affinity by effectively integrating multimodal and multiscale interaction information based on the proposed MMCLAC method. Firstly, two high-quality 3D kinase-drug datasets are constructed to minimize noise and accurately represent structural features. Subsequently, MMCLKin integrates geometric graph networks with sequence networks powered by large language models to capture the spatial structure and evolutionary information of protein kinases, the 3D conformational and chemical characteristics of kinase inhibitors, along with their local and global interaction features, and then quantifies the attention of each component for kinase-drug binding using a multi-head attention mechanism. Grounded in the principle of attention consistency, MMCLKin employs the MMCLAC method to further distill pivotal features across diverse modalities and scales within the same system while discerning subtle variances among different systems. We evaluate MMCLKin on two constructed datasets with three splitting strategies, along with ten structurally diverse protein-drug datasets and one mutation-aware dataset. Results indicate that MMCLKin consistently demonstrates high predictive accuracy for kinase inhibitor selectivity and binding affinity, with strong generalizability across broader protein-drug interaction scenarios. Additional assessments are conducted across three scenarios: (1) structurally resolved kinases, (2) structurally uncharacterized kinases, and (3) the specific mutated kinases. MMCLKin consistently exhibits strong virtual screening performance and predictive accuracy across structurally known, unknown and mutated kinases, highlighting its reliability, generalizability and versatility. This screening capability is further supported by ADP-Glo assays on the pathogenic LRRK2 G2019S mutant, where five of 20 MMCLKin-identified compounds exhibit potent inhibitory activity, with \({{{{\rm{IC}}}}}_{50}\) values of 468 \({{{\rm{nM}}}}\) (LY2025-01), 2.081 \({{{\rm{nM}}}}\) (LY2025-02), 1384 \({{{\rm{nM}}}}\) (LY2025-03), 8.694 \({{{\rm{nM}}}}\) (LY2025-04), 130.3 \({{{\rm{nM}}}}\) (LY2025-05), respectively. Moreover, visualization analyses reveal MMCLKin can effectively identify key interaction features, such as the hinge-region residues of kinases closely associated with specific binding, as well as polar atoms or functional groups of kinase inhibitors that facilitate polar interactions with pocket residues. Overall, MMCLKin exhibits robust predictive accuracy and holds promising potential for extension to other conserved protein families and mutated kinases, particularly in scenarios with limited protein experimental structure data, owing to its independence from crystal structures.
Results
Overview of MMCLKin
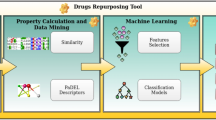
MMCLKin is a deep learning framework designed to accurately predict kinase-inhibitor selectivity and activity by extracting and integrating their critical interaction information across diverse modalities and scales, as illustrated in Fig. 1. To minimize noise, two high-quality 3D kinase-drug datasets, 3DKDavis and 3DKKIBA, are constructed by extracting high-confidence kinase domains and key binding sites from 3D structures predicted by AlphaFold212, and generating minimum-energy conformations for small molecules using LigPrep module (Fig.1a). Subsequently, a geometric graph network module, emphasizing both local and global completeness, is utilized to comprehensively capture the spatial structural features of kinases together with the 3D conformational characteristics of small molecules. In parallel, a sequence network module, incorporating a protein language model and a chemical language model, is utilized to extract evolutionary information from kinase sequences and detailed chemical features of small molecules. Next, MMCLKin employs a multi-head attention mechanism to autonomously learn kinase-inhibitor dependencies at different ranges and quantify the contribution of each element within the complex system to the prediction task. Simultaneously, our proposed MMCLAC method is engineered to ensure thorough and effective integration of spatial structure-based, sequence-based, local and global kinase-drug interaction features by aligning their attention distributions at the node level. This methodology facilitates the model to comprehensively capture the significant interaction features within complexes while allowing the model to effectively differentiate binding patterns among distinct kinase-inhibitor systems, thereby bolstering its interpretability and generalizability (Fig. 1b). Finally, the prediction module integrates interaction information across modalities and scales to generate predictive results. MMCLKin consistently achieves competitive performance in multiple application scenarios (Fig. 1c), including kinase-inhibitor and other protein-drug affinity prediction, kinase inhibitor selectivity profiling, virtual screening on structurally known, unknown and mutated kinases, and interpretability analysis. ADP-Glo assay results also further confirm that five out of 20 MMCLKin-identified compounds effectively inhibit the LRRK2 G2019S mutant, with four demonstrating inhibitory activity at nanomolar concentrations. These findings underscore the predictive accuracy of MMCLKin, particularly highlighting its promise in addressing structurally uncharacterized kinases and clinically significant mutations.
a Construction of two 3D kinase-inhibitor datasets by extracting high-confidence kinase domains and key binding pockets from the AlphaFold2-predicted 3D structures, coupled with the generation of minimum-energy molecular conformations using the LigPrep module within the OPLS4 force field. b MMCLKin employs geometric graph and sequence network modules to extract both local and global interaction features from kinase-drug 3D structures and 1D sequences. A multi-head attention mechanism is subsequently utilized to more comprehensively capture the intricate kinase-drug interaction patterns. Finally, multimodal and multiscale contrastive learning with attention consistency (MMCLAC) approach is integrated to effectively fuse and learn these interaction features, thereby enabling accurate predictions. c Applications of MMCLKin across four different scenarios and interpretability analysis.
MMCLKin achieves robust performance in predicting kinase-drug binding affinity
Firstly, we evaluated the utility and quality of two constructed datasets, 3DKDavis and 3DKKIBA, against the original datasets, Davis13 and KIBA14, which consisted solely of kinase sequences and drug SMILES, using the sequence-based ConPLex model15. ConPLex harnesses the pre-trained protein language model ProtBert16 to encode protein sequence features and molecular fingerprints for drugs, then employs a protein-anchor contrastive co-embedding strategy to co-locate proteins and drugs into a shared latent space to force separation between true interacting partners and decoys. Figure 2a and b present the distribution of five independent runs of the ConPLex model on four datasets conducted with the kinase-drug cold-start split. The results indicate that ConPLex exhibited improved performance on 3DKKIBA and 3DKDavis across five metrics compared to the original datasets, with particularly notable improvements in CI, PCC, and Spearman's coefficients, emphasizing the capacity of the constructed datasets to facilitate the model in capturing critical kinase-inhibitor interaction information.
a Performance comparison of ConPLex on the constructed 3DKDavis dataset versus the original Davis dataset. Five independent replications of each method were performed (n = 5). Box plots show the median as the center lines, upper and lower quartiles as box limits, whiskers as maximum and minimum values, and circles represent individual data points. b Performance comparison of ConPLex on the constructed 3DKKIBA dataset versus the original KIBA dataset. Five independent replications of each method were performed (n = 5). Box plots show the median as the center lines, upper and lower quartiles as box limits, whiskers as maximum and minimum values, and circles represent individual data points. c Comparison of the kinase-drug affinity prediction performance of MMCLKin against other models across three splitting strategies on the 3DKDavis dataset. Three independent replications of each method were performed (n = 3). Data are expressed as mean ± SD. d Comparison of the kinase-drug affinity prediction performance of MMCLKin against other models across three splitting strategies on the low drug similarity LSKIBA dataset. Three independent replications of each method were performed (n = 3). Data are expressed as mean ± SD. All models were rigorously evaluated using a comprehensive set of performance metrics, including the Concordance Index (CI), Mean Absolute Error (MAE), Pearson Correlation Coefficient (PCC), Mean Squared Error (MSE), and Spearman’s rank correlation coefficient (Spearman). Source data are provided as a Source Data file.
We subsequently assessed the kinase-inhibitor affinity prediction performance of the proposed MMCLKin on these two constructed datasets with three splitting settings (as defined in “Construction of two high-quality 3D kinase-drug datasets”). Comparisons were conducted against six representative baselines spanning three input modalities: sequence (TransformerCPI17, FusionDTA18, PSICHIC19), 2D molecular graphs (GraphDTA20, DrugBAN21), and 3D structural representations (KDBNet22). Specifically, GraphDTA is a widely adopted baseline for drug-target affinity prediction. TransformerCPI is recognized for its resilience to data biases and strong interpretability. FusionDTA incorporates global sequence features and achieves high predictive accuracy on both Davis and KIBA datasets. KDBNet leverages geometric graph networks to model the intricate local spatial and topological structures of kinase-drug interactions. DrugBAN employs conditional domain adversarial learning to align the learned interaction representations across heterogeneous data distributions, enabling strong generalization to novel drug-target pairs. PSICHIC integrates structural constraints to capture the underlying physicochemical mechanisms of protein-drug binding. The details for all baseline methods are provided in Supplementary Note 1.1. The hyperparameters of MMCLKin and all comparative models were meticulously adjusted multiple times to ensure better model fit to the datasets. The mean, standard deviation, and distribution of prediction performance from three independent runs were reported for comparative analysis.
On the 3DKDavis dataset, MMCLKin demonstrated lower predictive standard deviations and consistently outperformed all comparison models across four metrics under both kinase and kinase-drug cold-start splitting strategies (Fig. 2c). Notably, in kinase cold-start setting, MMCLKin substantially outperformed the leading sequence-based method (PSICHIC), achieving a 17.74% reduction in MSE (0.269 vs 0.327) and a 6.14% reduction in MAE (0.260 vs 0.277). In the more challenging kinase-drug cold-start scenario, MMCLKin also reduced MAE by 16.26% relative to the 2D graph-based DrugBAN (0.381 vs 0.455). Additionally, under drug cold-start splitting, MMCLKin also delivered best-in-class performance across MAE, MSE and CI, with MSE and MAE reduced by 0.137 and 0.077, respectively, compared to DrugBAN. To rigorously evaluate generalization capability, we constructed the LSKIBA benchmark - a low-similarity subset of 3DKKIBA containing only compound pairs with a Tanimoto similarity below 0.4 (calculated from SMILES representations) to ensure structural dissimilarity between test and training compounds. The similarity distributions of 3DKDavis and LSKIBA are shown in Supplementary Fig. S1. As shown in Fig. 2d, MMCLKin consistently outperformed other methods in both MAE and MSE across all three splitting strategies. Notably, it achieved an MSE of 0.310 and an MAE of 0.376 under the kinase cold-start setting, and exhibited 8.42% lower MSE relative to 3D geometric model KDBNet22 under the drug cold-start split. Additionally, MMCLKin attained the highest PCC and CI scores under the kinase cold-start scenario, while delivering comparable results under the remaining two splits.
Five-fold cross-validation23 was further performed to objectively evaluate the robustness of our model (denoted as \({{MMCLKi}n}_{5{CV}}\)). On the 3DKDavis dataset under the kinase-drug cold-start split, \({{MMCLKi}n}_{5{CV}}\) achieved a PCC improvement of 0.072 over MMCLKin. On the LSKIBA dataset with the drug cold-start split, \({{MMCLKi}n}_{5{CV}}\) yielded reductions of 0.057 in MSE and 0.024 in MAE compared to MMCLKin. While \({{MMCLKin}}_{5{CV}}\) did not surpass MMCLKin on some evaluation metrics, it consistently outperformed all baseline models. For instance, \({{MMCLKin}}_{5{CV}}\) achieved lower MAE than all compared models across kinase and kinase-drug cold-start splits on LSKIBA and outperformed all comparison models in MSE and PCC under the kinase cold-start split on 3DKDavis. These results highlight that MMCLKin with five-fold cross-validation remains a highly competitive model for kinase-drug affinity prediction.
In addition, we quantified the predictive uncertainty of MMCLKin on the 3DKDavis dataset under three different splitting strategies and examined its Spearman correlation with MAE score, as well as the calibration performance of MMCLKin. A higher Spearman’s coefficient indicates a stronger alignment between uncertainty and MAE, while a smaller deviation from the ideal diagonal on the calibration curve reflects more reliable confidence estimation. As shown in Supplementary Fig. S2, MMCLKin achieved strong positive Spearman correlations, especially under the kinase cold-start (\({\rho }_{{Spearman}}=0.786\)) and kinase-drug cold-start (\({\rho }_{{Spearman}}=0.675\)) settings. These correlations were particularly pronounced in the low-uncertainty regime, where lower predicted uncertainty was associated with higher predictive accuracy. This is further supported by calibration curves exhibiting relatively low miscalibration in these regions. Under the drug cold-start setting, the Spearman correlation between MAE and predicted uncertainty was substantially lower, and MMCLKin consistently underestimated the true error. This miscalibration may stem from the limited number of compounds in 3DKDavis dataset (68 in total), which could impair the ability of MMCLKin to learn well-calibrated uncertainty estimates due to data sparsity.
In conclusion, the constructed 3DKDavis and 3DKKIBA datasets facilitate more effective learning of essential kinase-drug interaction features. On these datasets, MMCLKin consistently delivers accurate and stable kinase-drug affinity predictions across all three data-splitting strategies, even under five-fold cross-validation, underscoring its advanced predictive performance and strong generalization. These results also emphasize the advantage of integrating both sequence and 3D graph representations, as opposed to single-modality inputs, for modeling kinase-drug interactions. Simultaneously, compared with models that focus solely on either local (e.g., KDBNet) or global (e.g., DrugBAN and FusionDTA) interaction patterns, the strong performance of MMCLKin suggests that jointly modeling both local and global features may yield a more comprehensive and informative representation of kinase-drug interactions. In addition, the model also produces well-calibrated uncertainty estimates specifically under the kinase and kinase-drug cold-start settings, as evidenced by strong positive correlations between predicted uncertainty and actual error, along with reliable calibration in low-uncertainty regimes. This enhances the reliability of MMCLKin in real-world drug discovery scenarios.
Contributions of MMCLKin’s components to enhanced predictive performance
We systematically evaluated the contributions of individual components to MMCLKin’s performance on the 3DKDavis dataset under kinase-drug cold-start setting (Supplementary Fig. S3). To ensure unbiased comparison, we assessed MMCLKin variants incorporating individual features (sequence, 3D structure, local and global interactions) while maintaining fixed architecture and hyperparameters. For clarity, the corresponding sub-models were designated as \({{MMCLK}}_{{Seque}},\) \({{MMCLK}}_{3{DGraph}},{{MMCLK}}_{{Local}}\) and \({{MMCLK}}_{{Global}}\), respectively. Results indicated that the performance of all sub-models consistently fell short of MMCLKin across all metrics. Notably, compared to MMCLKin, \({{MMCLK}}_{3{DGraph}}\) and \({{MMCLK}}_{{Local}}\) exhibited substantially higher prediction errors, with observed increases of 29.76% and 22.18% in MAE, and 6.05% and 17.09% elevation in MSE, respectively. In terms of PCC and CI, \({{MMCLK}}_{3{DGraph}}\) showed reductions of 0.120 and 0.061, while \({{MMCLK}}_{{Global}}\) exhibited decreases of 0.026 and 0.047, relative to MMCLKin. In addition, we also assessed the impact of ESM-derived features by constructing an ablated variant, \({{MMCLK}}_{{NOESM}}\), in which the ESM-based embeddings were substituted with a basic index-based encoding of amino acids. The results indicated that \({{MMCLK}}_{{NOESM}}\) exhibited notable performance degradation across four evaluation metrics compared to MMCLKin, underscoring the importance of ESM-derived embeddings to effectively capture informative protein sequence features.
As a key methodological innovation in this study, the underlying significance of MMCLAC approach was also investigated by a comparative analysis of MMCLKin with and without MMCLAC on the 3DKDavis dataset under kinase-drug cold-start splitting (Supplementary Fig. S3b). The results revealed that MMCLAC module contributed substantially to MMCLKin’s capability. Specifically, MMCLKin with MMCLAC exhibited 15.35% and 12.76% reduction in MAE and MSE, respectively, and 5.26% and 17.92% increases in CI and PCC, respectively. Additionally, we examined the impact of MMCLAC on attention-based contrastive losses. The analysis indicated that MMCLKin without MMCLAC exhibited large fluctuations across the four attention-based losses, whereas the inclusion of MMCLAC resulted in the substantial convergence. This stability highlights the effectiveness of MMCLAC in constraining attention weights of the interacting systems across distinct modalities and scales.
In summary, the ESM-based embeddings strengthen MMCLKin to extract informative protein sequence features. The integration of four diverse characterizations of kinase-drug systems, together with the MMCLAC approach, enables effective learning and fusion of the interaction features across different modalities and scales, empowering MMCLKin to achieve strong accuracy, stability and generalizability in predicting kinase-drug binding affinity. Concurrently, the distinct contribution of each representation may also provide valuable insights for future endeavors in multimodal and multiscale feature integration. Collectively, these findings reaffirm the efficacy of MMCLKin and align with our initial hypothesis.
MMCLKin performs well in predicting the selectivity of kinase inhibitors across the human kinome
The development of highly selective kinase inhibitors demonstrates a strong correlation with minimized off-target interactions, establishing kinase selectivity as a critical determinant in kinase-directed drug discovery. Accordingly, we further evaluated the ability of MMCLKin to predict kinase inhibitor selectivity using the 3DKDavis and low-similarity LSKIBA datasets. To achieve broader coverage of human kinome, the drug cold-start splitting method was employed. Simultaneously, a panel of established selectivity metrics, standard score, Gini coefficient, selectivity entropy, and partition index24 (see Supplementary Note 1.2 for definitions), was adopted to holistically assess MMCLKin’s selectivity prediction performance. The standard score quantifies how many kinases a compound binds with affinity exceeding a specified threshold. The Gini coefficient measures the inequality in a compound’s binding affinity distribution across kinases, with higher values indicating more selective binding to a narrow subset. Selectivity entropy assesses the dispersion of binding affinities, where lower entropy values correspond to more selective compounds. The partition index, derived from the association constants, evaluates the preferential binding of a compound to a reference kinase relative to others. While these four metrics provide a comprehensive assessment of compound selectivity across the human kinome, they do not directly evaluate the accuracy of predicted selectivity. To address this, we analyzed the correlation between predicted and experimentally observed selectivity distributions for each metric. This correlation analysis offers a more direct evaluation of model performance, with higher correlation coefficients indicating greater agreement between predicted and experimental selectivity metrics.
Figure 3 presents the Pearson correlations between predicted and ground-truth selectivity metrics across MMCLKin, FusionDTA, TransformerCPI, and KDBNet on two datasets. On the 3DKDavis dataset, MMCLKin substantially outperformed all baselines with respect to the standard score, Gini coefficient, and selectivity entropy, yielding Pearson correlation coefficients of 0.898, 0.537, and 0.651, respectively, indicating its closer concordance between predicted and true selectivity profiles. On the low-similarity LSKIBA dataset, MMCLKin consistently achieved either the highest or equivalent Pearson correlations across all metrics, demonstrating its robustness and strong generalization capacity. Interestingly, all models exhibited near-perfect correlations on the partition index across both datasets, likely because it only emphasizes relative affinity rankings, which are inherently more predictable than precise absolute values. A similar pattern was observed for selectivity entropy on LSKIBA dataset, where Pearson correlations for all models approached 1.0. By contrast, on the 3DKDavis dataset with only 68 unique compounds, selectivity entropy appeared more sensitive to local prediction errors, resulting in greater variability across models.
a Comparison of selectivity prediction performance of kinase inhibitors between MMCLKin and several models on the 3DKDavis dataset (n = 39). b Comparison of selectivity prediction performance of kinase inhibitors between MMCLKin and several models on the low drug similarity LSKIBA dataset (n = 156). Pearson represents the linear correlation between the predicted and experimentally observed selectivity distributions for each metric, while RMSD evaluates their overall deviation. The shaded areas indicate 95% confidence intervals. Source data are provided as a Source Data file.
In summary, MMCLKin more precisely predicts kinase inhibitor selectivity across the human kinome. This facilitates the identification of highly selective kinase inhibitors, while indirectly attesting to its ability to discern subtle differences in binding interactions between inhibitors and diverse kinase targets, thereby providing valuable guidance for the rational design of highly selective kinase inhibitors. Moreover, the divergent behavior observed across selectivity metrics offers insights for future metric selection. For example, the standard score and Gini coefficient appear more sensitive to differences in model performance, potentially providing a more effective basis for evaluating selectivity modeling capabilities.
MMCLKin showcases good generalization capacity on diverse protein structures
In addition to evaluating the predictive performance of MMCLKin on conserved kinase structures, we also examined its generalization ability using ten structurally diverse datasets, including PDBbind dataset (PDBbind v2020 and CASF-201625 datasets), seven target superfamilies26, and two kinase datasets from the IDG-DREAM Drug-Kinase Binding Prediction Challenge27. For PDBbind dataset, CASF-2016 benchmark was employed as the test set, and all overlapping complexes were excluded from both the general and refined sets of PDBbind v2020 to eliminate data leakage. Subsequently, 500 samples were randomly selected from the refined set to serve as the validation set, while the remaining samples were combined with the general set to form the training set. Notably, although MMCLKin does not depend on experimental complex structures, we compared it against both experimental complex-based models and complex-free models to ensure a more comprehensive evaluation. Experimental results for comparison models were derived from previously published studies28,29. For the remaining nine datasets, 3D protein structures and small-molecule conformations were generated following the same protocol employed for 3DKDavis and 3DKKIBA datasets.
Table 1 presents the performance comparison between MMCLKin and previously reported models on the CASF-2016 test set. MMCLKin consistently outperformed all complex-free models across three metrics. Especially, it achieved an MAE of 0.997, which is 0.029 lower than that of the geometry-aware GAABind29. Additionally, MMCLKin recorded an RMSE of 1.291 and a PCC value of 0.807, surpassing other models utilizing sequence (MolTrans, TransformerCPI), graph (GAABind, GraphDTA) or 3D point cloud (KIDA) representations as input. When compared to complex-based models, MMCLKin also exhibited competitive performance, with RMSE and MAE values closely matching those of the optimal model, IGN30.
Table 2 summarizes the comparison between MMCLKin and MMAtt-DTA across seven target superfamilies. MMCLKin outperformed MMAtt-DTA on six of the seven datasets, and consistently achieved the best performance across RMSE, CI, and Spearman correlation on Enzyme, GPCR, Ion channel, Kinase, and Transporter datasets. For instance, MMCLKin reduced RMSE by 7.68%, 11.49%, 5.89% and 7.45% on the Kinase, Transporter, Enzyme and Ion channel datasets, respectively, compared to MMAtt-DTA. In terms of Spearman correlation, improvements of 0.014 and 0.018 were observed on the Kinase and Transporter datasets. Additionally, MMCLKin also attained the highest CI and Spearman correlation on the Epigenetic regulator dataset, with the latter showing a relative increase of 6.38%.
We further conducted a systematic comparison between MMCLKin and models submitted to the IDG-DREAM Drug-Kinase Binding Prediction Challenge across both Round 1 and Round 2 datasets. Notably, samples in the 3DKDavis dataset with affinity values of 5 were excluded to construct a more refined training set, as such values primarily serve as default indicators of insufficient binding evidence rather than true binding affinities13. As shown in Supplementary Fig. S4, MMCLKin achieved competitive performance on both rounds. On the Round 1 dataset, it attained a Spearman correlation of 0.430 and an RMSE of 1.147, ranking second only to the top-performing model. In Round 2, MMCLKin maintained strong predictive accuracy, with a Spearman correlation of 0.482 and an RMSE of 1.068, closely matching the best-performing entries.
These findings emphasize the enhanced predictive accuracy and exceptional generalization capability of MMCLKin on datasets featuring structurally diverse proteins. Simultaneously, its stable and strong performance on protein families such as GPCRs and Transporters further underscores its potential for broader applicability across conserved non-kinase protein families. More importantly, its ability to function independently of experimental structures may present a significant advantage, supporting innovative drug discovery targeting proteins with unresolved crystal structures.
MMCLKin shows strong virtual screening performance and interpretability for two kinase targets with known experimental 3D-structures
To validate the real-world applicability of MMCLKin in drug discovery, we further comprehensively evaluated its virtual screening performance on two kinase targets with experimental structures (LRRK231, HPK132; their detailed information is provided in Supplementary Note 1.3). For the virtual screening, we selected the MMCLKin model with performance closest to the average under kinase cold-start split, and benchmarked it against Schrödinger’s Glide Standard Precision (SP, 2023)33, a state-of-the-art docking tool widely used in drug discovery.
High-resolution wild-type PDB experimental structures (shown in Fig. 4a, 8FO734 for LRRK2 and 7R9N35 for HPK1) were selected as receptors. Inhibitors with experimentally determined dissociation constants (\({K}_{d}\)) below 100 \({{{\rm{nM}}}}\), sourced from BindingDB36, were identified as active molecules (see Supplementary Tables S1–S3). Decoy sets were generated using DUD-E37 based on these active molecules, and subsequently, the resulting decoys were combined with the actives to construct the screening set. Receptor Grid Generator and LigPrep modules were utilized for binding site preparation and to generate the lowest-energy molecular conformations. Virtual screening capability was evaluated using the recall rate of active kinase inhibitors (see Supplementary Note 1.4 for calculation details of recall rate).
a Virtual screening workflow implemented by MMCLKin and Glide SP for LRRK2 and HPK1. b Comparison of recall rates for active molecules targeting LRRK2 (PDB ID: 8FO7) between MMCLKin and Glide SP. c Comparison of recall rates for active molecules targeting HPK1 (PDB ID: 7R9N) between MMCLKin and Glide SP. d Identification of critical residues and functional groups within the 8FO7-BDBM50308060 complex system by MMCLKin. e Identification of critical residues and functional groups within the 7R9N-BDBM4814 complex system by MMCLKin. Two complexes are generated using Glide SP to enhance clarity and intuitive structural insights. Source data are provided as a Source Data file.
The results unequivocally demonstrate a substantial advantage of MMCLKin over Glide SP docking method in identifying active inhibitors for both LRRK2 and HPK1. For LRRK2 (Fig. 4b), Glide SP achieved a recall rate of 36.36% within the top 1% of ranked compounds, whereas MMCLKin reached 45.45%. At the top 5% and 10% thresholds, MMCLKin attained recall rates of 72.73% and 81.82%, markedly outperforming 45.45% of Glide at both thresholds. MMCLKin exhibited equally compelling performance on HPK1 (Fig. 4c), yielding recall rates of 42.86%, 57.14% and 64.29% within the top 1%, 2% and 10% of ranked molecules, respectively, substantially exceeding 28.57%, 35.71% and 42.86% of Glide.
We further mapped the attention coefficients to the residues of these kinase targets and their representative inhibitors to investigate the interpretability of MMCLKin. To facilitate more intuitive visualization, the top 15 residues with the highest attention weights were focused on. For LRRK2 (Fig. 4d), residues Val1946, Glu1948, Ala1950, Lys1952, Ser1954 were situated within the hinge region of kinase, which is essential for the stable binding of ATP and high specificity of many marketed kinase inhibitors. In particular, the identified residues Glu1948 and Ala1950 are pivotal in LRRK2 for forming hydrogen bonds with most kinase inhibitors38. Additional residues, including Glu1902, Ala1904 and Val1905, were found within the β-sheet region of the N-terminal lobe, while residues such as Leu2001, Leu2002, Phe2003, and Ile2015 were located within the β-sheet region of the C-terminal lobe. As for HPK1 (Fig. 4e), residue Glu92 was identified in the hinge region of conserved kinase domain, residues Arg22, Leu23, Gly25, Val31, Val43, Ala44, Leu45, Ile89, Cys90 were located in the β-sheet region of the small lobe, and Leu144, Asn142, Arg152, Leu153 were situated in the β-sheet region of the large lobe. These regions have been extensively documented in prior studies as critical for facilitating kinase-inhibitor specificity and binding stability39. This alignment with experimentally validated critical binding regions underscores the precision of MMCLKin in identifying key residues involved in inhibitor binding.
MMCLKin also exhibits pronounced attention to polar functional groups of inhibitors, such as hydroxyl, amino, imino, tertiary amino, pyrrole and amide groups. These functional groups are widely acknowledged to mediate strong polar interactions with residues in the kinase binding pocket, thereby enhancing kinase-drug binding affinity and selectivity.
Taken together, these findings not only validate the virtual screening ability of MMCLKin, demonstrating its superiority over a leading industry-standard tool in accurately identifying kinase inhibitors, but also underscore its advanced interpretability. Furthermore, the capacity of MMCLKin to effectively capture and exploit key atoms, functional groups and residues within kinase-drug systems also demonstrates its deep learning-driven proficiency in uncovering critical features of protein-ligand interactions, thereby reinforcing its utility for potential kinase inhibitor discovery.
MMCLKin maintains good virtual screening performance and interpretability on two kinase targets lacking experimental 3D-structures
Given the independence of MMCLKin from experimentally resolved structures, we further assessed its robustness on two kinases, NUAK240 and CRK1241, lacking experimental 3D-structures. For these targets (Fig. 5a), AlphaFold2-predicted kinase domain structures were utilized as receptors, with the binding pockets defined by residues within 14 Å of the active site center, as predicted by P2Rank42. Owing to the limited availability of known inhibitors for CRK12, compounds reported by Smith et al.43 were designated as active molecules. Screening sets for both kinases (see Supplementary Tables S1, S4 and S5 for the respective active molecules) were constructed and processed following the same standardized procedures as established for LRRK2 and HPK1.
a Virtual screening workflow implemented by MMCLKin and Glide SP for NUAK2 and CRK12. b Comparison of recall rates of active molecules targeting NUAK2 between MMCLKin and Glide SP. c Comparison of recall rates of active molecules targeting CRK12 between MMCLKin and Glide SP. d Identification of critical residues and functional groups within the NUAK2 complex system by MMCLKin. e Identification of critical residues and functional groups within the CRK12 complex system by MMCLKin. Two complexes were constructed using Glide SP to bolster clarity and yield intuitive structural insights. Source data are provided as a Source Data file.
Impressively, even in the absence of resolved 3D structures for kinases, MMCLKin maintained strong and even more competitive screening capabilities. For NUAK2 (Fig. 5b), MMCLKin achieved markedly higher recall rates of 41.67%, 50% and 66.67% within the top 2%, top 5% and 10% of ranked compounds, compared to 25%, 33.33% and 41.67% of Glide SP. The advantage was even more pronounced for CRK12 (Fig. 5c), where MMCLKin reached recall rates of 53.85% and 84.62% at both the top 5% and 10% thresholds, far exceeding 15.38% and 23.08% of Glide SP. These results not only affirm the strong virtual screening capabilities of MMCLKin across diverse kinase targets but also highlight its consistent efficacy and substantial potential to advance drug discovery targeting structurally unresolved kinases.
MMCLKin also exhibited robust and powerful interpretability when applied to predicted kinase structures. Especially, hinge region residues Glu130, Tyr131, Ala132, Arg134, and Asp136 of top 15 critical residues for NUAK2 (Fig. 5d), and residues Pro431, Tyr432, Ala433 of CRK12 were prioritized by MMCLKin (Fig. 5e). Other critical residues identified by MMCLKin were primarily distributed across β-sheet structures of the N- and C-lobes, further demonstrating the adaptability of MMCLKin in generalizing to predicted 3D structures. Furthermore, an in-depth analysis of kinase inhibitors revealed that MMCLKin consistently emphasizes the characteristics of polar functional groups such as amino, ether, keto carbonyl, pyrrole and amide groups. These polar functional groups are predisposed to form key hydrogen bonds or electrostatic interactions with binding pocket, thereby enhancing the specificity and strength of kinase-drug interactions.
In conclusion, these findings underscore the consistent and competitive virtual screening performance of MMCLKin across both experimentally determined and predicted kinase structures. Simultaneously, its ability to autonomously capture key atoms, functional groups and residues from raw kinase-drug data emphasizes its self-learning capability and interpretability, deepening understanding of model predictions. Furthermore, the distinct recognition patterns by MMCLKin across different kinase systems substantiate its capacity to differentiate various protein structures, providing a solid foundation for elucidating the specific kinase-inhibitor interactions. Collectively, these strengths position MMCLKin as a highly promising tool for drug discovery targeting both experimentally resolved and structurally unknown proteins.
MMCLKin-driven discovery of LRRK2 G2019S inhibitors and biological activity evaluation
Residue mutations of kinases are frequently implicated in a wide range of diseases and the occurrence of drug resistance44. Targeted screening against mutant kinases facilitates the identification of lead compounds with mutant-specific activity. Accordingly, beyond investigating the capability of MMCLKin on wild-type (WT) kinases, we systematically assessed its predictive performance on mutant kinase targets from three perspectives: (1) prediction accuracy of inhibitor activities against both LRRK2 WT and its G2019S mutant, a variant strongly implicated in Parkinson’s disease; (2) comprehensive evaluation on a dataset comprising 3082 WT-mutant kinase pairs; and (3) virtual screening and experimental validation of potential inhibitors targeting the LRRK2 G2019S mutant.
We first evaluated the ability of MMCLKin to discriminate the subtle differences between LRRK2 WT and G2019S mutant. Specifically, high-resolution experimental structures (PDB IDs: 8FO7 for WT and 8TZC for G2019S) were selected as receptors. A balanced kinase-inhibitor dataset targeting LRRK2 WT and G2019S mutant, curated from BindingDB, was utilized to fine-tune the MMCLKin model trained on 3DKDavis dataset. Finally, the finetuned MMCLKin model was used to predict the pIC50 values of four active compounds previously identified by our group45. Supplementary Table S6 indicates that the predicted outcomes of MMCLKin for both LRRK2 WT and G2019S mutant align closely with experimental values. For instance, the predicted pIC50 of LY2023-24 against the LRRK2 G2019S mutation is 6.747, and the experimental value is 6.661. The predicted pIC50 value for LRRK2-IN-1 targeting LRRK2 WT is 8.315, closely matching its experimental value of 8.509. Further horizontal analysis revealed that, whether for WT kinases or their mutants, inhibitors with higher experimental pIC50 values consistently exhibited higher predicted values. These findings illustrate that MMCLKin accurately predicts the inhibitory activity of drugs against both LRRK2 WT and G2019S mutant, showcasing its competence to discern the subtle mutational differences and providing a reliable basis for identifying kinase inhibitors with high selectivity and binding affinity towards kinase mutants.
To systematically evaluate the predictive capability of MMCLKin on both WT and mutant kinases, we further curated a mutation-aware dataset, 3DKinMW, comprising 3082 WT-mutant kinase pairs spanning five kinase targets: E2BBR (4 pairs), JAK2 (1 pair), RET (2343 pairs), LRRK2 (591 pairs), and MET (143 pairs). Their 3D structures were obtained following protocols consistent with those used in the 3DKDavis and 3DKKIBA datasets. Model training and evaluation were carried out under drug cold-start split, enabling the model to learn from cases where the same compound interacts with both WT and mutant of a given kinase in the training set, and to predict \(p{{IC}}_{50}\) values of previously unseen compounds against both forms in the test set. The average results of three independent experiments are presented in Supplementary Fig. S5. MMCLKin demonstrated strong predictive performance, achieving a CI of 0.766, MSE of 0.350, PCC of 0.728, and MAE of 0.459.
The practical applicability of MMCLKin in mutated kinase-targeted drug discovery was further investigated by integrating MMCLKin-based virtual screening with biological experimental validation using the ADP-Glo assay. Specifically, the MMCLKin model trained without any LRRK2-related data was employed to screen approximately 180,000 compounds from the ChemDiv library against the LRRK2 G2019S mutant (PDB ID: 8TZC). The top 5,000 candidate compounds (predicted score > 5.9) were subjected to molecular docking using the Glide extra precision mode. Compounds with docking scores below -8.0 were retained and clustered using k-means to maximize structural diversity. Twenty representative compounds were selected based on careful visual inspection of their predicted binding poses, with particular emphasis on key interactions involving Glu1948 and Ala1950 in the hinge region45,46, which are critical for ligand-kinase binding. Subsequently, these compounds were initially evaluated using the ADP-Glo kinase activity assay at 10 μM, a concentration adopted in previous reports47,48, with LRRK2-IN-1 included as a positive control. Of the 20 compounds, five compounds (designated LY2025-01 to LY2025-05) exhibited > 50% inhibition. Their specific IC₅₀ values were determined using a 10-point, three-fold serial dilution. As shown in Fig. 6a, all five candidate compounds display substantial topological diversity, characterized by distinct core scaffolds and substituent patterns. Notably, LY2025-04 (FN-1501) exhibited 100% inhibition at 10 μM, slightly surpassing the positive control LRRK2-IN-1 (99.72%) (Fig.6b). And further biochemical validation also revealed that LY2025-04 (IC₅₀ =8.694 nM) exhibited an inhibitory potency nearly equivalent to LRRK2-IN-1 (IC₅₀ =7.001 nM) (Fig.6c). In addition, LY2025-01, LY2025-02, and LY2025-05 also achieved over 90% inhibition at 10 μM. Especially, LY2025-01, previously unreported as a kinase inhibitor, exhibited an IC₅₀ of 468 nM, suggesting its favorable inhibitory activity and potential as a functional modulator of LRRK2 G2019S. Although previously reported to inhibit ULK1/249, LY2025-02 (sbp-7455) demonstrated markedly greater potency against LRRK2 G2019S (IC₅₀ = 2.081nM), surpassing the reference inhibitor LRRK2-IN-1. This finding supports its enhanced target selectivity and structural compatibility for LRRK2 G2019S. LY2025-05 (Befotertinib), an approved drug for non-small cell lung cancer50, also exhibited substantial inhibitory activity against LRRK2 G2019S (IC₅₀=130.3nM), underscoring its potential therapeutic repurposing in Parkinson’s disease and other LRRK2 G2019S-associated pathologies. Additionally, LY2025-03, showing an IC₅₀ of 1384 nM, may also provide a viable starting point for future optimization.
a Chemical structures of five candidate compounds exhibiting greater than 50% inhibition at 10 \({{{\rm{\mu }}}}{{{\rm{M}}}}\) and the positive control. b Inhibitory ratio of five candidate compounds and the positive control at 10 \({{{\rm{\mu }}}}{{{\rm{M}}}}\) concentration. Three independent replications of each method were performed (n = 3). Data are expressed as mean ± SD. c IC₅₀ values of five candidate compounds and the positive control against LRRK2 G2019S mutant. Three independent replications of each method were performed (n = 3). Data are expressed as mean ± SD. Source data are provided as a Source Data file.
In summary, MMCLKin exhibits strong predictive accuracy and reliable efficacy in identifying potential inhibitors targeting mutant kinases. These findings reinforce its robustness in handling challenging kinase profiling scenarios and position it as a promising tool for mutation-aware drug discovery.
Discussion
Discovering efficient and selective kinase inhibitors remains a critical yet formidable challenge in contemporary biomedical research due to the conserved structure of kinases. The substantial costs of experimental profiling across the human kinome further underscore the imperative need for developing high-precision predictive approaches for kinase-inhibitor binding affinity and selectivity. In this study, we developed a framework, MMCLKin, to predict the activity and selectivity of kinase inhibitors on diverse kinases. This framework leverages geometric graph networks to capture spatial structural features, employs large language model-based sequence networks to extract evolutionary and chemical information, and incorporates a multi-head attention mechanism to model complicated kinase-drug interactions while quantifying the contribution of each element to prediction task. Simultaneously, we further proposed a multimodal and multiscale contrastive learning with attention consistency to effectively integrate these diverse interaction characteristics. Comprehensive evaluations confirm the competitive predictive capabilities of MMCLKin, outperforming other methods in predicting activity and selectivity of kinase inhibitors on two constructed high-quality 3D kinase-drug datasets. The strong prediction performance across ten datasets featuring diverse protein structures and a mutation-aware dataset further showcases the generalizability and adaptability of MMCLKin. Furthermore, MMCLKin exhibits good virtual screening capability for structurally known, unknown as well as challenging mutated kinase targets, and attention coefficient analysis further reveals that MMCLKin can capture key residues and molecular functional groups from raw data, proving its interpretability and autonomous learning ability. Finally, biochemical profiling using ADP-Glo assays substantiated that five out of 20 MMCLKin-identified compounds potently inhibited the LRRK2 G2019S mutant, with four exhibiting nanomolar-level potency, underscoring its practical utility in identifying highly potent mutant kinase inhibitors.
In conclusion, MMCLKin represents a robust and versatile framework for advancing the discovery of highly selective and high-affinity kinase inhibitors. Its strong performance on structurally diverse datasets also suggests promising applicability to other non-kinase protein families. While the integration of multi-scale and multi-modal features improves model representational capacity, this comes with increased computational demands during both training and inference phases compared to sequence-based methods. Moving forward, a key challenge lies in efficiently extracting essential information from these heterogeneous representations and developing more streamlined fusion strategies to improve computational efficiency without compromising predictive performance.
Methods
Construction of two high-quality 3D kinase-drug datasets
KIBA and Davis are two widely recognized kinase-drug affinity datasets that encompass the binding affinities of a single molecule across various kinases, thereby facilitating the elucidation of the binding specificity of a given kinase inhibitor toward multiple kinase targets. However, both datasets are limited to sequence-based representation of drugs and protein kinases, omitting the comprehensive three-dimensional structural information. This limitation may impede the ability to model the intricate conformational landscapes and physiologically relevant interaction patterns.
To address this limitation while minimizing reliance on experimental crystal structures, we implemented a comprehensive workflow to construct two high-quality 3D kinase-drug datasets, 3DKKIBA and 3DKDavis (Fig. 1a and Supplementary Fig. S6). Specifically, duplicate sequences were first removed to prevent data leakage during model evaluation. Subsequently, AlphaFold2-predicted protein kinase structures were used, followed by extraction of kinase domains to minimize the interference from non-kinase domains. An added benefit of this strategy is that kinase domains predicted by AlphaFold2 typically exhibit high confidence scores, greatly reducing errors propagated to downstream modeling. Next, binding pockets were predicted using P2Rank, with optimal site centers identified through scoring and manual verification. To fully exploit the binding information, residues within 20 Å radius of the site center were defined as the binding pocket. For kinase inhibitors, 3D conformations were generated using LigPrep module of Schrödinger, with the minimum-energy conformation selected as the dominant state. Supplementary Figs. S6a, b illustrate the distinct distributions of kinases and small molecules of two datasets. The 3DKDavis dataset quantifies affinity using the \({{pK}}_{d}\) constant \(({{pK}}_{d}=-{\log }_{10}({K}_{d}/{10}^{9}))\), whereas the 3DKKIBA dataset retains the original KIBA score, an integrated metric derived from \({{IC}}_{50}\), \({K}_{i}\) and \({K}_{d}\) values. Notably, for 3DKKIBA, only complexes involving small molecules with Tanimoto similarity scores below 40% were selected to construct the LSKIBA subset for performance evaluation (The similarity distributions of 3DKDavis and LSKIBA were shown in Supplementary Fig. S1).
To thoroughly assess the predictive performance of MMCLKin on kinase targets, in accordance with Luo et al.22, 3DKDavis and LSKIBA were divided into training and test sets at a 4:1 ratio, adopting three distinct splitting strategies (Supplementary Fig. S6c). Drug cold-start indicates that the test set excludes any identical drugs present in the training set. Kinase cold-start signifies that the test set omits any protein kinases from the training set. Kinase-drug cold-start denotes that the test set contains neither kinases nor drugs overlapping with the training set. Additionally, to further assess the generalization capability of MMCLKin, we trained it on the PDBbind v2020 subset and tested it on the structurally diverse CASF-2016 benchmark. For seven target superfamilies, we followed the same protocol as MMAtt-DTA, randomly splitting each dataset into training and test sets at a 4:1 ratio.
3D graph and sequence representations of protein kinases and binding pockets
Each protein kinase or binding pocket was represented as a 3D graph and a sequence to comprehensively encode its structural and biochemical properties. A 3D graph is defined as \(G=\left(V,E,P\right),\) where \(V={[{v}_{1},{v}_{2},\cdots,{v}_{n}]}^{T}\in {{\mathbb{R}}}^{n\times 6}\) is the node feature matrix, with each node corresponding to an amino acid residue. The edge feature matrix, \(E={[{e}_{1},{e}_{2},\cdots,{e}_{m}]}^{T}\in {{\mathbb{R}}}^{m\times 32}\), defines edges e based on spatial proximity, where an edge exists if one node is among the 30 nearest neighbors of another. The position matrix \(P={[{p}_{1},{p}_{2},\cdots,{p}_{n}]}^{T}\in {{\mathbb{R}}}^{n\times 3}\) denotes the spatial coordinates of all residues. To fully capture the conformational characteristics of protein kinases or binding pockets, both local completeness and global completeness were leveraged to represent their overall spatial structures, as proposed by Wang et al.51. This method has been proven to effectively distinguish naturally occurring conformers. Specifically, local completeness is characterized by spherical coordinate \(({d}_{{ij}},{\theta }_{{ij}},{\varPhi }_{{ij}})\), which is derived from node features, edge indices, and positional coordinates, to describe the relative position of node i and its 1-hop neighborhood. Global completeness is achieved by further incorporating the edge rotation angle \({\tau }_{{ij}}\), providing a comprehensive representation of spatial orientations. These variables are defined as follows:
Where \({P}_{i}\) and \({P}_{j}\) are the position coordinates of nodes i and j, respectively. \({f}_{i}\) and \({s}_{i}\) are the first and second nearest neighbors of node i, \({f}_{i/j}\) denotes the nearest neighbor of node i excluding j, and \({f}_{j/i}\) denotes the nearest neighbor of node j except i, \({{\mbox{plane}}}_{{f}_{i},i,{s}_{i}}\) refers to the plane formed by \({f}_{i},i,{s}_{i}\), with similar definitions for other planes. This approach can effectively capture complete geometric structure information while significantly reducing computational complexity.
For sequence representation, ESM model52, a cutting-edge protein language model pretrained on 250 million data, was employed to extract rich biological evolutionary features and contextual information from kinases and binding pockets. The resulting embeddings can effectively encode structural, functional, and evolutionary properties, which have proven beneficial for tasks such as functional prediction and structural modeling22. The specific features of protein kinases and binding pockets are summarized in Supplementary Table S7.
3D graph and sequence representations of kinase inhibitors
Kinase inhibitors are similarly characterized using both 3D graphs and SMILES notations. 3D graph for an inhibitor is also represented as \(G=\left(V,E,P\right)\), where \(V={[{v}_{1},{v}_{2},\cdots,{v}_{n}]}^{T}\in {{\mathbb{R}}}^{n\times 75}\) denotes the feature matrix of n molecular atoms, with each node having 75 dimensional features. \(E={[{e}_{1},{e}_{2},\cdots,{e}_{m}]}^{T}\in {{\mathbb{R}}}^{m\times 8}\) represents the edge feature set between nodes, where each edge possesses 8 characteristic dimensions and m is the number of chemical bonds in the molecule. The position matrix P is constructed analogously to that used for proteins. We also incorporated local spherical coordinates \(({d}_{{ij}},{\theta }_{{ij}},{\varPhi }_{{ij}})\) and edge rotation angle \({\tau }_{{ij}}\) to characterize the local and global completeness of molecules. For SMILES, the ChemBERTa-2 model53, pretrained on 10 million compounds from PubChem, was employed to extract 384 dimensional chemical information. ChemBERTa-2, built upon the RoBERTa transformer54, leverages semi-supervised pre-training of language models to learn molecular fingerprints. This model has been extensively applied in drug screening, property prediction, and other chemical-related tasks, demonstrating strong scalability and efficiency55. The specific meaning of each molecular feature dimension is detailed in Supplementary Table S7.
MMCLKin architecture
MMCLKin is designed for accurate prediction of kinase-drug activity and selectivity via the effective extraction and integration of interaction features across diverse modalities and scales. It consists of five primary components:
Geometric graph network module
In this module, we developed EComENet (Fig. 7a), a geometric graph neural network built upon the ComENet framework51. ComENet is a graph neural network that leverages quantum-inspired basis functions to comprehensively represent 3D molecular conformation by achieving both local and global completeness. However, it focuses solely on node features, neglecting edge features that are essential for determining molecular properties, such as bond types, which influence electron distribution and chemical reactivity. EComENet addresses this limitation by integrating edge features into the geometric graph representation, enabling the extraction of more complete conformational features of chemical entities.
a Geometric graph network module models the local and global spatial interactions of kinases and drugs using 3D kinases, 3D binding pockets, and 3D molecules. b Sequence network module leverages large language models and BiLSTMs to extract evolutionary information from kinase and pocket sequences, alongside chemical features from SMILES. c A multi-head attention mechanism is applied to further identify dependency relationships across varying ranges within kinase-drug interaction systems operating at diverse modalities and scales, while quantifying the contribution of each component to the prediction task. d Prediction module is used to generate the predictive results based on the concatenated interaction features from various modalities and scales. e Multimodal and multiscale contrastive learning with attention consistency (MMCLAC) method aligns attention coefficients across different modalities and scales for elements within the same domain, ensuring the model effectively captures kinase-drug interaction features from diverse perspectives while distinguishing binding differences among diverse interaction systems. \({{\mathbb{R}}}_{13{kd}},{{\mathbb{R}}}_{13{pd}},{{\mathbb{R}}}_{1{kpd}}\) and \({{\mathbb{R}}}_{3{kpd}}\) denote the shared domains of four paired interactions (1D and 3D kinase-drug interactions, 1D and 3D pocket-drug interactions, 1D kinase-drug and 1D pocket-drug interactions, 3D kinase-drug and 3D pocket-drug interactions) used for contrastive learning, respectively, and \({{\mathbb{P}}}_{{kd}-1d}^{13{kd}},{{\mathbb{P}}}_{{kd}-3d}^{13{kd}},{{\mathbb{P}}}_{{pd}-1d}^{13{pd}},{{\mathbb{P}}}_{{kd}-3d}^{13{kd}},{{{\mathbb{P}}}_{{kd}-1d}^{1{kpd}},{{\mathbb{P}}}_{{pd}-1d}^{1{kpd}}{\mathbb{,}}{\mathbb{P}}}_{{kd}-3d}^{3{kpd}},{{\mathbb{P}}}_{{pd}-3d}^{3{kpd}}\) represent the corresponding relative attention probability sets within the defined shared domains.
Specifically, EComENet begins by incorporating node features, edge features, and edge indices as inputs, and utilizes message passing neural networks to aggregate the features of target node, their neighboring nodes, and the associated edges, yielding a new node feature \({v}_{i,j,{e}_{{ij}}}\) that incorporates bond information. Subsequently, the aggregated features are passed through a ReLU activation function to introduce nonlinearity, followed by a linear layer with bias to transform the graph information into a new feature space. The formulas are:
Where θ and β are the learnable parameter matrix, \({v}_{i}\) is the feature of target node i, \({v}_{j}\) is the feature of neighbor node j, \(N\left(i\right)\) is all adjacent nodes of node i, \({h}_{\theta }\) is the neural network, \({e}_{i,j}\) is the edge feature connecting node i and node j, b is the bias vector.
Given the pivotal role of distance as a geometric feature, two associated tuples \(({d}_{{ij}},{\theta }_{{ij}},{\Phi }_{{ij}})\) and \({d}_{{ij}},{\pi }_{{ij}}\) are utilized as inputs to capture local and global structural features of biomolecular conformations. Then TBF and SBF convert these raw geometric data into physically meaningful vectors.
Where TBF and \({{{\rm{S}}}}{{{\rm{BF}}}}\) denote the basis functions for the tuples \(({d}_{{ij}},{\theta }_{{ij}},{\Phi }_{{ij}})\) and \({d}_{{ij}},{\pi }_{{ij}}\), respectively. \({{{{\rm{j}}}}}_{\vartheta }\left(\cdot \right)\) represents the spherical Bessel function of order \(\vartheta\), \(c\) is the cutoff value, and \({\rho }_{\vartheta n}\) is the \(n-{th}\) root of the Bessel function of order \(\vartheta,{{{{\rm{Y}}}}}_{\vartheta }^{m}\) is a spherical harmonic function of degree \(m\) and order \(\vartheta\).
These vectors, along with \({v}_{i,j,{e}_{{ij}}}^{{\prime} }\) and the edge indices, are then fed into the interaction blocks. Within each block, \({v}_{i,j,{e}_{{ij}}}^{{\prime} }\), \({F}_{i,j,{local}}\), \({F}_{i,j,{global}}\) are further updated via the linear layer. Then the local and global graph convolution layers take the updated node matrix \({v}_{i,j,{e}_{{ij}}}^{{\prime\prime} }\) as input and employ the vector derived from the basis function as edge weights of the convolution layer to extract the local and global conformational features. Following that, the obtained features are linearly transformed and subjected to nonlinear activation through Swish activation function. Next, the local and global features are concatenated, and residual connections are applied to sum the input features \({v}_{i,j,{e}_{{ij}}}^{{\prime\prime} }\) with the concatenated features, enhancing the robustness and feature-learning capability of our model. Finally, several linear layers and GraphNorm56 are applied to down-project and regularize the features. The specific formulas are as follows:
Local completeness:
Global completeness:
Concatenate and down-project:
Where \(\beta,{\theta }_{1},{\theta }_{2},\alpha\) are the learnable parameter matrices, b is the bias vector, \({||}\) represents the concatenation operation, \(\zeta (\cdot )\) denotes a sequence of four MLP layers, \(E[v]\) represents the mean of the input features, \(\odot\) signifies the element-wise multiplication, \({Var}\)[⋅] measures the dispersion of samples around the mean, \(\epsilon\) is a numerical stability constant ensuring stable computations, \(\gamma\) can scale the normalized features to adjust the importance of each feature. μ serves as the shifting parameter, acting as a bias term after normalization.
The features from four iterative interaction blocks are fed into the self-atom layer, which comprises four MLP layers paired with a Swish activation function. Each layer updates node features and projects them into a new dimensional space, and the resulting features serve as the input for the two-layer GraphSAGE57 network. The \({{{\mathcal{l}}}}\)-th layer can be expressed as:
The GraphSAGE network concatenates the target node features with the aggregated features of its neighbors, and fuses them through a fully connected layer, endowing the model with strong expressive capacity. Additionally, since the network is built on an inductive framework, it can efficiently generate node embeddings for previously unseen data, enhancing the generalization ability of our model. The formula for the \({{{\mathcal{l}}}}\)-th GraphSAGE layer is:
Where \({W}_{1}^{{{{\mathscr{(}}}}{{{\mathcal{l}}}}{{{\mathscr{)}}}}}\) and \({W}_{2}^{{{{\mathscr{(}}}}{{{\mathcal{l}}}}{{{\mathscr{)}}}}}\) are the learnable parameter matrix of the \({{{\mathcal{l}}}}\)-th layer, \({{{\mathcal{N}}}}\left(i\right)\) denotes the set of neighboring nodes of node \(i\), \({v}_{j,{ecom}}^{({{{\mathcal{l}}}})}\) represents the feature vector of neighboring nodes \(j\), \({v}_{i,{ecom}}^{({{{\mathcal{l}}}}{-}1)}\) is the embedding of node \(i\) from the layer \({{{\mathcal{l}}}}{{{\mathscr{-}}}}1\).
Finally, the features of protein kinases, pockets and drugs processed by EComENet and GraphSAGE networks are concatenated to obtain both kinase-drug and pocket-drug feature matrices. Layer normalization58 and bidirectional long short-term memory network (BiLSTM)59 are then utilized to normalize and extract their temporal features and contextual information, thereby learning the global and local interactions between protein kinases and drugs. The BiLSTM, composed of two independent LSTMs operating in forward and backward directions, ensures that each time-step output is affected by current, previous, and subsequent states. This bidirectional processing enhances the ability of the model to capture and integrate long-range and short-range dependencies, facilitating a deeper understanding of complicated kinase-drug interactions.
Where \({{{\rm{||}}}}\) represents the concatenation operation, \({V}_{{eg}}^{k}\) denotes the kinase feature processed by EComENet and GraphSAGE networks, \({V}_{{eg}}^{p}\) is the pocket feature, \({V}_{{eg}}^{d}\) represents the drug feature.
Sequence network module based on large language models
In the sequence network module (Fig.7b), the pretrained chemical language model ChemBERTa-2 is harnessed to extract enriched chemical representations from small molecules. The resulting embedding vectors \({h}_{l}\) are subsequently transformed through a linear layer, followed by a LeakyReLU activation function to introduce nonlinearity. To capture intramolecular dependencies encoded within SMILES, a BiLSTM layer is employed.
For protein kinases and binding pocket sequences, the pretrained protein language model ESM is leveraged to derive evolutionary information. This is followed by two BiLSTM layers designed to capture contextual dependencies within the sequences, with a dropout function incorporated to mitigate overfitting.
Where \({h}_{k}\) and \({h}_{p}\) are the evolutionary features of kinase and binding pocket, respectively.
Similar to the geometric graph network module, features from binding pockets, protein kinases, and small molecules are concatenated to generate the pocket-drug and kinase-drug characteristics, allowing the subsequent multi-head attention mechanism module to effectively learn and model global and local interactions.
Multi-head attention mechanism module
To thoroughly investigate kinase-drug interactions and elucidate the contribution of each component within the complex system to its binding affinity, a multi-head attention mechanism60 was implemented (Fig. 7c). Specifically, this mechanism first partitions the input space into \(h\) independent subspaces, each processed by Scaled Dot-Product Attention60. Within each subspace, the input matrix is projected into query, key, and value matrices \({Q}_{i},{K}_{i},{V}_{i}\) via trainable projection matrices \({W}_{i}^{q},{W}_{i}^{k},{W}_{i}^{v}.\) Subsequently, the key matrix is transposed, and its dot product with the query matrix yields a similarity matrix whose elements indicate the alignment between corresponding query and key vectors. To mitigate potential gradient explosion or vanishing issues, these similarity matrices are scaled to prevent excessively large or small values from destabilizing the training process. The scaled weights are then normalized using the softmax function to produce a probability distribution, i.e., the attention weights, that assigns higher weights to residues or molecular atoms most pertinent to the prediction task and lower weights to less important elements. Next, the value vector \({V}_{i}\) is aggregated with the attention weights through a weighted summation, resulting in an updated value vector that incorporates the contribution of each element to the prediction task. Finally, the updated value vectors from \(h\) heads are concatenated, and the resulting vector undergoes a linear transformation and is fed into the downstream prediction module. The formulas for each attention head are as follows:
The formula for multi-head attention is as follows:
Where \({Q}_{i},{K}_{i},{V}_{i}\) represent the query, key and value matrices of the i-th head, \({d}_{k}\) is the dimensionality of the column vectors in matrices \({Q}_{i}\) and \({K}_{i}\). \(h\) denotes the number of attention heads. \({{Head}}_{i}\) signifies the output of the i-th head, and \({W}^{o}\) is the output transformation matrix used to integrate the outputs of all heads.
This methodology enables each attention head to concentrate on distinct subspace features, allowing the model to recognize diverse interaction patterns between protein kinases and drugs. Additionally, the attention weights pinpoint critical kinase residues and molecular atoms, offering valuable insights into the mechanisms underlying kinase-drug interactions.
Multimodal and multiscale contrastive learning with node-level attention consistency (MMCLAC)
We believe that, as a biologically meaningful entity, a kinase-drug complex possesses the intrinsic interaction information alongside a structured and learnable distribution of attention. Consequently, regardless of whether a complex is represented as a sequence or a 3D graph, the attention distribution within the same structural domain should remain consistent. Building on this hypothesis, we implemented the contrastive learning of node-level attention weights for 3D and 1D kinase-drug as well as 3D and 1D pocket-drug interactions (Fig. 7e). It aims to integrate local and global interaction patterns across sequence and graph modalities, while eliminating the negative impacts associated with the inherent limitations of each modality by imposing the attention consistency constraint. Furthermore, since local interactions are inherently a subset of global interactions in a complex, the distribution of attention weights for pocket-drug, extracted from the kinase-drug interaction, is expected to align with that derived from its independently modeled pocket-drug interaction. Consequently, we further implemented the contrastive learning between 3D kinase-drug and 3D pocket-drug interactions, as well as between 1D kinase-drug and 1D pocket-drug interactions. The linkages between local and global interactions were designed to emphasize critical interactions between binding sites and drugs while ensuring that the model remains attentive to the overall biochemical context of the protein kinase. Simultaneously, incorporating node-level contrastive learning enables the model to capture subtle differences between diverse systems, enhancing its interpretability and predictive performance for kinase-drug specificity and selectivity.
Specifically, to expand the differentiation of attention weights among various elements, the unscaled attention weights were selected to perform node-level contrastive learning, and the coefficients across different dimensions for each node were summed to derive its final attention weight. Subsequently, the elements from two contrastive learning items were aligned to determine their maximal intersection, defining a shared structural domain. The attention coefficients of the elements within this domain were then extracted according to their indices. Recognizing that the model may exhibit variations in capturing the interactions of different modalities and scales, the attention weights of each element within the same domain were normalized. This normalization ensures that, during comparisons, our approach focuses solely on the relative contribution of each element to the prediction task, rather than on their absolute values. The corresponding formulas are provided below:
Where \(M\) represents the weight dimensions for each node, \({H}_{i,m}\) is the unscaled attention matrix of node \(i\), \({{Att}}_{i}^{{domain}}\) denotes the summed attention weight of node \(i\), the symbol \(\odot\) indicates element-wise multiplication, \({Var}\)[⋅] measures the dispersion of the samples. The constant \(\epsilon\) ensures numerical stability, \(\gamma\) scales the normalized features, \(\partial\) serves as the shifting parameter. \({P}_{i}^{{domain}}\) indicates the relative attention probability of node \(i\) within the domain.
Accordingly, for each interaction pair used for contrastive learning, relative attention probability sets can be derived based on their shared domains. Taking the 1D and 3D kinase-drug interactions as an example, let their shared domain be denoted as \({{\mathbb{R}}}_{13{kd}}\), with their corresponding index sets within this domain defined as \({{{{\rm Z}}}}_{1}\subseteq \{{x}_{1,}{x}_{2,}\ldots,{x}_{n}\}\) and \({{{{\rm Z}}}}_{2}\subseteq \{{y}_{1,}{y}_{2,}\ldots,{y}_{n}\}\), respectively. The relative attention probability sets corresponding to both index sets can be expressed as:
Similarly, the shared domain between 1D and 3D pocket-drug interactions can be denoted as \({{\mathbb{R}}}_{13{pd}}\), with the corresponding index sets given by \({{{{\rm Z}}}}_{3}\subseteq \{{x}_{1,}{x}_{2,}\ldots,{x}_{m}\}\) and \({{{{\rm Z}}}}_{4}\subseteq \{{y}_{1,}{y}_{2,}\ldots,{y}_{m}\}\), respectively. Their associated relative attention probability sets can be defined as:
For 1D kinase-drug and 1D pocket-drug interactions, the shared domain can be denoted as \({{\mathbb{R}}}_{1{kpd}}\), with the corresponding index sets \({{{{\rm Z}}}}_{5}\subseteq \{{x}_{1,}{x}_{2,}\ldots,{x}_{q}\}\) and \({{{{\rm Z}}}}_{6}\subseteq \{{y}_{1,}{y}_{2,}\ldots,{y}_{q}\}\), and their relative attention probability sets are:
Likewise, for 3D kinase-drug and 3D pocket-drug interactions, the shared domain is denoted as \({{\mathbb{R}}}_{3{kpd}}\), with the corresponding index sets \({{{{\rm Z}}}}_{7}\subseteq \{{x}_{1,}{x}_{2,}\ldots,{x}_{\omega }\}\) and \({{{{\rm Z}}}}_{8}\subseteq \{{y}_{1,}{y}_{2,}\ldots,{y}_{\omega }\}\). Their relative attention probability sets can refer to:
Here, \(n,m,q,\omega\) denote the number of elements within the shared domains of four respective contrastive interaction pairs.
Ultimately, four pairs of node-level attention weight sets were constructed and employed to perform the contrastive learning. By aligning these relative attention probabilities across different modalities and scales, this strategy promotes more effective representation learning and improves the generalizability of our model.
Prediction module
The prediction module integrates both structure-based and sequence-based kinase-drug interaction features at local and global levels (Fig. 7d), to predict binding affinity of kinase-drug pairs and the selectivity of kinase inhibitors across human kinome. Specifically, it concatenates these four distinct interaction features and applies layer normalization to the fused high-dimensional features. This normalization, which enforces a zero mean and unit variance, enhances model stability and accelerates convergence during training.
Where \({{{{\mathcal{H}}}}}_{3d}^{{kl}},{{{{\mathcal{H}}}}}_{3d}^{{pl}},{{{{\mathcal{H}}}}}_{1d}^{{kl}},{{{{\mathcal{H}}}}}_{1d}^{{pl}}\) represent the interaction features of four levels processed by multi-head attention mechanism, \(\odot\) is the element-wise multiplication, \({Var}\)[⋅] measures the dispersion of the samples, \(\epsilon\) denotes the numerical stability constant, \(\gamma\) and \(\mu\) are the learnable scaling and shifting parameters.
Following that, an Adaptive Max Pooling operation is utilized to extract key information \({{{{\mathcal{H}}}}}_{{KL}}^{P}\) from the fused features \({{{{\mathcal{H}}}}}_{{KL}}^{N}\) while effectively reducing their dimensionality. Finally, the prediction layer, composed of three-layer fully connected neural network with ELU activation function, performs a nonlinear transformation on the pooled features to predict the binding affinity between kinases and drugs. The formulas for the prediction layer are:
Where \(\beta\) is the parameter matrix to be learned, \(b\) is the bias vector.
Workflow
In general, we developed a comprehensive paradigm designed to effectively integrate multimodal and multiscale kinase-drug interaction features by incorporating multiple strategies. These strategies encompass minimizing noise in dataset construction from predicted structures, extracting information from multiple modalities, capturing intricate local and global interaction patterns, quantifying the contribution of individual element to the prediction task, while leveraging node-level MMCLAC method to fuse features across modalities and scales and to detect inter-system heterogeneity. Notably, this framework eliminates the dependency on experimental structures, offering significant value for the discovery and screening of therapeutic drugs targeting proteins without structural data. Additionally, we contend that this paradigm holds substantial potential for addressing analogous challenges in other conserved protein families.
Multimodal and multiscale contrastive loss functions
Building on our proposed framework, we developed a contrastive loss function to maximize the consistency between positive pairs that share the same interaction domain, while differentiating them from negative pairs involving different pocket-drug interactions. Specifically, given a batch containing N complexes, we calculated eight attention weights for each system, forming four distinct pairs of attention weights: \(\{{{{{\rm{\delta }}}}}_{{{kd}}_{i}}^{1d},{{{{\rm{\delta }}}}}_{{{kd}}_{i}}^{3d}\}\), \(\{{{{{\rm{\delta }}}}}_{{{pd}}_{i}}^{1d},{{{{\rm{\delta }}}}}_{{{pd}}_{i}}^{3d}\}\), \(\{{{{\delta }}}_{{{kpd}}_{i}}^{1d},{{{{\rm{\delta }}}}}_{{{pd}}_{i}}^{1d}\}\), {\({\delta }_{{{kpd}}_{i}}^{3d},{{\delta }}_{{{pd}}_{i}}^{3d}\)}, where \({{{\delta }}}_{{{kd}}_{i}}^{1d}\) and \({{{\delta }}}_{{{kd}}_{i}}^{3d}\) represent the attention weights of 1D and 3D kinase-drug interactions, \({{{{\rm{\delta }}}}}_{{{pd}}_{i}}^{1d}\) and \({{{{\rm{\delta }}}}}_{{{pd}}_{i}}^{3d}\) denote the attention weights of 1D and 3D pocket-drug interactions, \({{{{\rm{\delta }}}}}_{{{kpd}}_{i}}^{1d}\) refers to the attention weights for elements in the 1D kinase-drug system that aligns with 1D pocket-drug interaction domain, \({{{{\rm{\delta }}}}}_{{{kpd}}_{i}}^{3d}\) represents the attention weights for elements in the 3D kinase-drug system that aligns with the 3D pocket-drug composition.
Taking the attention weights of 1D and 3D kinase-drug interactions as an example, we derived the contrastive loss function to enforce their alignment and consistency, as shown below:
Where 〈·〉 denotes the inner product to measure the similarity, \(N\) represents the number of samples in the batch, \(\tau\) is a scale parameter. Since \({{{{\rm{\delta }}}}}_{{{kd}}_{i}}^{1d}\) and \({{{{\rm{\delta }}}}}_{{{kd}}_{i}}^{3d}\) are two embeddings derived from 1D sequence and 3D spatial structure of the same complex, they are regarded as a positive pair, while all other samples in the batch are treated as negative pairs. The contrastive loss functions of attention weights are similarly defined for the remaining three pairs: 1D and 3D pocket-drug interactions, 3D kinase-drug and 3D pocket-drug interactions and 1D kinase-drug and 1D pocket-drug interactions.
This approach enables the model to effectively capture interaction characteristics within a complex system while discerning binding disparities across distinct systems from multiple perspectives by maximizing the similarity between positive pairs and minimizing the similarity between negative pairs. In addition, a mean squared error loss (MSELoss) was also incorporated to quantify the discrepancy between the predicted and experimental values.
Where \(K\) represents the number of samples, \({x}_{i}\) and \({y}_{i}\) are the predicted and true values. Ultimately, the final loss function was constructed by integrating the affinity loss with multimodal and multiscale attention contrast losses.
Where \(N,M,Q,W\) represent the number of comparison pairs at different scales and modalities, \({{{{\rm{\S }}}}}_{1},{{{{\rm{\S }}}}}_{2},{{{{\rm{\S }}}}}_{3},{{{{\rm{\S }}}}}_{4}\) are coefficients that adjust the contribution of the individual contrastive loss component. This comprehensive loss function not only evaluates the accuracy of our model in predicting kinase-drug affinity and kinase inhibitor selectivity, but also bolsters its ability to capture the binding features intrinsic to a single interaction system while distinguishing among diverse interaction systems.
Hyperparameter tuning, model training and evaluation metrics
The AdamW optimizer61 was utilized for gradient descent. It combines momentum with adaptive learning rate techniques to effectively control model complexity via weight decay adjustments and automatic learning rate tuning, thereby accelerating model convergence. The CosineAnnealingWarmRestarts algorithm was employed to periodically restart the learning rate during training, enabling the model to escape local minima and more effectively explore the global optimum, ultimately improving the model performance. To prevent overfitting, an early stopping strategy was implemented, terminating training once improvements in model performance plateaued.
In both affinity and selectivity prediction analyses, Optuna62 was employed for small-scale hyperparameter optimization of MMCLKin, targeting hyperparameters including batch size, learning rate, learning rate decay, hidden dimensions, and the weight coefficients of four attention contrast loss functions. The hyperparameter configuration obtained from this search was applied to model training and evaluation on both LSKIBA and 3DKDavis datasets. Additionally, extensive hyperparameter tuning was also conducted for all baseline models to ensure their optimal adaptation to the constructed datasets. Ablation studies were conducted to investigate the contribution of individual components on model performance, with all other settings, including architecture and optimization parameters, unchanged. To comprehensively assess model performance, multiple evaluation metrics were employed, including the mean absolute error (MAE), Concordance Index (CI), mean squared error (MSE), root mean square error (RMSE), Pearson Correlation Coefficient (PCC), Spearman’s rank Correlation Coefficient (Spearman), standard score, Gini coefficient, selectivity entropy and partition index. Detailed definitions of these metrics are provided in Supplementary Note 1.2.
ADP-Glo assay
The inhibitory activity of 20 selected compounds against the LRRK2 G2019S mutant was tested using ADP-Glo kinase assay. First, the inhibition rate of each compound was measured at a concentration of 10 \({{{\rm{\mu }}}}{{{\rm{M}}}}\), and LRRK2-IN-1 was used as a positive control. Subsequently, compounds exhibiting 50% inhibition were selected, and their IC₅₀ was further determined at 10 concentration points produced by 3-fold serial dilution. The ADP-Glo assay protocol was as follows: 2× ATP/Substrate and 2× kinase solutions were prepared in kinase reaction buffer. Using the Echo 655 system, 100 \({{{\rm{nL}}}}\) of each compound dilution was transferred to a 384-well assay plate. After centrifugation, 5 \({{{\rm{\mu }}}}{{{\rm{L}}}}\) of 2× kinase solution was added, followed by centrifugation at 1000 × g for 1 min and incubation at 25 °C for 10 min. Then, 5 \({{{\rm{\mu }}}}{{{\rm{L}}}}\) of 2× ATP/substrate solution was added, centrifuged again (1000 × g, 1 min), and incubated at 25 °C for 120 min. Subsequently, 5 \({{{\rm{\mu }}}}{{{\rm{L}}}}\) of ADP-Glo reagent and 10 \({{{\rm{\mu }}}}{{{\rm{L}}}}\) of Kinase Detection Reagent were sequentially added, with each step followed by centrifugation (1000 × g, 1 min) and incubation at 25 °C for 40 min. Luminescence was measured using a BMG microplate reader to assess kinase activity. All experiments were tested three times, and the average values were reported to ensure data accuracy and reproducibility. The percent inhibition (\(\%{{{\rm{Inh}}}}\)) was calculated using the following formula:
Where CW denotes chemiluminescence value of the sample, \(\overline{{{{\rm{HC}}}}}\) refers to the mean conversion rate without inhibitor (reaction mixture containing 1% DMSO, kinase, substrate, and ATP). \(\overline{{{{\rm{LC}}}}}\) is the mean conversion rate without kinase and inhibitor (reaction mixture containing 1% DMSO, substrate, and ATP). Subsequently, IC₅₀ values were determined by fitting the calculated %Inhibition values and log of compound concentrations to nonlinear regression (dose response - variable slope) with GraphPad 8.063.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Datasets generated in this study are deposited via Zenodo at https://doi.org/10.5281/zenodo.17290773, and are also available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets. Two constructed 3D kinase-drug binding affinity datasets, 3DKDavis and 3DKKIBA, are available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/3DKDavis and https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/3DKKIBA. The fine-tuned datasets of inhibitors targeting wild-type LRRK2 and LRRK2 G2019S mutant are available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/LRRK2_mutant. The PDB structures 8FO7, 7R9N, 8TZC, as well as the predicted structures AF-Q9H093-F1 [https://www.uniprot.org/uniprotkb/Q9H093/entry#structure] and AF-Q382V0-F1 [https://www.uniprot.org/uniprotkb/Q382V0/entry#structure], are publicly available. The PDBbind v2020 and CASF-2016 datasets are downloaded from http://www.pdbbind.org.cn/. The two kinase-drug affinity datasets, Davis and KIBA, are available in the Therapeutics Data Commons64 under accession code https://tdcommons.ai/multi_pred_tasks/dti/. The 3D datasets of seven target superfamilies are available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/seven_target_superfamilies. The 3D datasets of Round 1 and Round 2 based on IDG-DREAM Drug-Kinase Binding Prediction Challenge are available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/DREAM_kinases. The 3DKinMW dataset, comprising 3082 wild-type-mutant kinase pairs spanning five kinase proteins, is available at https://github.com/Yanara-Tian/MMCLKin/tree/main/datasets/wt_mutant_3082. Source data are provided with this paper.
Code availability
The source code of MMCLKin is available at https://github.com/Yanara-Tian/MMCLKin, which has also been deposited in the Zenodo under accession code https://zenodo.org/records/17290774 or via https://doi.org/10.5281/zenodo.17290773. P2Rank (https://github.com/rdk/p2rank) is used for predicting the kinase pockets. ESM and gvp (https://github.com/drorlab/gvp) employed to extract kinase features in this study, are publicly available under the MIT license. The code about ComENet used to develop MMCLKin is publicly available and has been deposited in DIG at https://github.com/divelab/DIG under the GPL license. Structures are visualized by Pymol v3.1.0 (https://www.pymol.org/). Data are analyzed using numpy v1.24.1 (https://numpy.org/), pandas v2.0.3 (https://pandas.pydata.org/), seaborn v0.13.2 (https://seaborn.pydata.org/), scikit-learn v1.3.2 (https://scikit-learn.org/) and matplotlib v3.7.5 (https://matplotlib.org/).
References
Cohen, P. Protein kinases—the major drug targets of the twenty-first century. Nat. Rev. Drug Discov. 1, 309–315 (2002).
Gough, N. R. & Kalodimos, C. G. Exploring the conformational landscape of protein kinases. Curr. Opin. Struct. Biol. 88, 102890 (2024).
Zhong, G., Chang, X., Xie, W. & Zhou, X. Targeted protein degradation: advances in drug discovery and clinical practice. Signal Transduct. Target. Ther. 9, 308 (2024).
Ren, Q. et al. KinomeMETA: meta-learning enhanced kinome-wide polypharmacology profiling. Brief. Bioinform. 25, bbad461 (2024).
Liu, S. et al. Versatile framework for drug–target interaction prediction by considering domain-specific features. J. Chem. Inf. Model. 64, 5646–5656 (2024).
Monteiro, N. R., Oliveira, J. L. & Arrais, J. P. TAG-DTA: Binding-region-guided strategy to predict drug-target affinity using transformers. Expert Syst. Appl. 238, 122334 (2024).
Yang, Z., Zhong, W., Zhao, L. & Chen, C. Y.-C. MGraphDTA: deep multiscale graph neural network for explainable drug–target binding affinity prediction. Chem. Sci. 13, 816–833 (2022).
Corso G., Stärk H., Jing B., Barzilay R., Jaakkola T. Diffdock: diffusion steps, twists, and turns for molecular docking. Proceedings International Conference on Learning Representations (ICLR, 2023).
Zhang, H., Lu, G., Zhan, M. & Zhang, B. Semi-supervised classification of graph convolutional networks with Laplacian rank constraints. Neural Process. Lett. 54, 2645–2656 (2022).
Theisen, R., Wang, T., Ravikumar, B., Rahman, R. & Cichońska, A. Leveraging multiple data types for improved compound-kinase bioactivity prediction. Nat. Commun. 15, 7596 (2024).
Li, S. et al. PocketAnchor: learning structure-based pocket representations for protein-ligand interaction prediction. Cell Syst. 14, 692–705.e696 (2023).
Yang, Z., Zeng, X., Zhao, Y. & Chen, R. AlphaFold2 and its applications in the fields of biology and medicine. Signal Transduct. Target. Ther. 8, 115 (2023).
Davis, M. I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046–1051 (2011).
Tang, J. et al. Making sense of large-scale kinase inhibitor bioactivity data sets: a comparative and integrative analysis. J. Chem. Inf. Model. 54, 735–743 (2014).
Singh, R., Sledzieski, S., Bryson, B., Cowen, L. & Berger, B. Contrastive learning in protein language space predicts interactions between drugs and protein targets. Proc. Natl. Acad. Sci. USA 120, e2220778120 (2023).
Brandes, N., Ofer, D., Peleg, Y., Rappoport, N. & Linial, M. ProteinBERT: a universal deep-learning model of protein sequence and function. Bioinformatics 38, 2102–2110 (2022).
Chen, L. et al. TransformerCPI: improving compound–protein interaction prediction by sequence-based deep learning with self-attention mechanism and label reversal experiments. Bioinformatics 36, 4406–4414 (2020).
Yuan, W., Chen, G. & Chen, C. Y.-C. FusionDTA: attention-based feature polymerizer and knowledge distillation for drug-target binding affinity prediction. Brief. Bioinform 23, bbab506 (2022).
Koh, H. Y., Nguyen, A. T. N., Pan, S., May, L. T. & Webb, G. I. PSICHIC: physicochemical graph neural network for learning protein-ligand interaction fingerprints from sequence data. Nat. Mach. Intell. 6, 673–687 (2024).
Nguyen, T. et al. GraphDTA: predicting drug–target binding affinity with graph neural networks. Bioinformatics 37, 1140–1147 (2021).
Bai, P., Miljković, F., John, B. & Lu, H. Interpretable bilinear attention network with domain adaptation improves drug–target prediction. Nat. Mach. Intell. 5, 126–136 (2023).
Luo, Y., Liu, Y. & Peng, J. Calibrated geometric deep learning improves kinase–drug binding predictions. Nat. Mach. Intell. 5, 1390–1401 (2023).
Tanoli, Z., Schulman, A. & Aittokallio, T. Validation guidelines for drug-target prediction methods. Expert Opin. Drug Discov. 20, 31–45 (2025).
Wang, T., Pulkkinen, O. I. & Aittokallio, T. Target-specific compound selectivity for multi-target drug discovery and repurposing. Front. Pharmacol. 13, 1003480 (2022).
Su, M. et al. Comparative assessment of scoring functions: the CASF-2016 update. J. Chem. Inf. Model. 59, 895–913 (2018).
Schulman, A., Rousu, J., Aittokallio, T. & Tanoli, Z. Attention-based approach to predict drug-target interactions across seven target superfamilies. Bioinformatics 40, btae496 (2024).
Cichońska, A. et al. Crowdsourced mapping of unexplored target space of kinase inhibitors. Nat. Commun. 12, 3307 (2021).
Lu, R. et al. Improving drug-target affinity prediction via feature fusion and knowledge distillation. Brief. Bioinform. 24, bbad145 (2023).
Tan, H., Wang, Z. & Hu, G. GAABind: a geometry-aware attention-based network for accurate protein–ligand binding pose and binding affinity prediction. Brief. Bioinform. 25, bbad462 (2023).
Jiang, D. et al. Interactiongraphnet: a novel and efficient deep graph representation learning framework for accurate protein-ligand interaction predictions. J. Med. Chem. 64, 18209–18232 (2021).
Tolosa, E., Vila, M., Klein, C. & Rascol, O. LRRK2 in Parkinson disease: challenges of clinical trials. Nat. Rev. Neurol. 16, 97–107 (2020).
Si, J. et al. Hematopoietic progenitor kinase1 (HPK1) mediates T cell dysfunction and is a druggable target for T cell-based immunotherapies. Cancer cell 38, 551–566.e511 (2020).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Zhu, H., Hixson, P., Ma, W. & Sun, J. Pharmacology of LRRK2 with type I and II kinase inhibitors revealed by cryo-EM. Cell Discov. 10, 10 (2024).
Chan, B. K. et al. Discovery of spiro-azaindoline inhibitors of hematopoietic progenitor kinase 1 (HPK1). ACS Med. Chem. Lett. 13, 84–91 (2021).
Liu, T. et al. BindingDB in 2024: a FAIR knowledgebase of protein-small molecule binding data. Nucleic Acids Res. 53, D1633–D1644 (2025).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Tan, S., Gong, X., Liu, H. & Yao, X. Identification of novel LRRK2 inhibitors by structure-based virtual screening and alchemical free energy calculation. Phys. Chem. Chem. Phys. 26, 19775–19786 (2024).
Liao, J. J.-L. Molecular recognition of protein kinase binding pockets for design of potent and selective kinase inhibitors. J. Med. Chem. 50, 409–424 (2007).
Yuan, W. C. et al. NUAK2 is a critical YAP target in liver cancer. Nat. Commun. 9, 4834 (2018).
Wyllie, S. et al. Cyclin-dependent kinase 12 is a drug target for visceral leishmaniasis. Nature 560, 192–197 (2018).
Krivák, R. & Hoksza, D. P2Rank: machine learning based tool for rapid and accurate prediction of ligand binding sites from protein structure. J. Cheminform. 10, 1–12 (2018).
Smith, A. et al. Repositioning of a diaminothiazole series confirmed to target the cyclin-dependent kinase CRK12 for use in the treatment of African animal trypanosomiasis. J. Med. Chem. 65, 5606–5624 (2022).
Lahiry, P., Torkamani, A., Schork, N. J. & Hegele, R. A. Kinase mutations in human disease: interpreting genotype–phenotype relationships. Nat. Rev. Genet. 11, 60–74 (2010).
Gong, X. et al. Discovery of potent LRRK2 inhibitors by ensemble virtual screening strategy and bioactivity evaluation. Eur. J. Med. Chem. 279, 116812 (2024).
Sanz Murillo, M. et al. Inhibition of Parkinson’s disease–related LRRK2 by type I and type II kinase inhibitors: activity and structures. Sci. Adv. 9, eadk6191 (2023).
Quereda, V. et al. Therapeutic targeting of CDK12/CDK13 in triple-negative breast cancer. Cancer cell 36, 545–558.e547 (2019).
Deng, X. et al. Characterization of a selective inhibitor of the Parkinson’s disease kinase LRRK2. Nat. Chem. Biol. 7, 203–205 (2011).
Ren, H. et al. Design, synthesis, and characterization of an orally active dual-specific ULK1/2 autophagy inhibitor that synergizes with the PARP inhibitor olaparib for the treatment of triple-negative breast cancer. J. Med. Chem. 63, 14609–14625 (2020).
Blair, H. A. Befotertinib: first approval. Drugs 83, 1433–1437 (2023).
Wang, L., Liu, Y., Lin, Y., Liu, H. Ji, S. ComENet: towards complete and efficient message passing for 3D molecular graphs. In Proc. 36th International Conference on Neural Information Processing Systems. 47 (NeurIPS, 2022).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl. Acad. Sci. USA 118, e2016239118 (2021).
Ahmad W., Simon E., Chithrananda S., Grand G., Ramsundar B. Chemberta-2: towards chemical foundation models. arXiv preprint arXiv:220901712, (2022).
Liu Y. et al. RoBERTa: A robustly optimized BERT pretraining approach. arXiv preprint arXiv:190711692, (2019).
Rollins, Z. A., Cheng, A. C. & Metwally, E. MolPROP: molecular property prediction with multimodal language and graph fusion. J. Cheminform. 16, 56 (2024).
Cai T. et al. Graphnorm: a principled approach to accelerating graph neural network training. In Proc. 38th International Conference on Machine Learning (ICML), 1204–1215 (PMLR, 2021).
Hamilton, W., Ying, Z. & Leskovec, J. Inductive representation learning on large graphs. Adv. Neural Inf. Process. Syst. 30, 1024–1034 (2017).
Lei Ba J, Kiros JR, Hinton GE. Layer normalization. ArXiv preprints, arXiv: 1607.06450 (2016).
Zhang, S., Zheng, D., Hu, X. & Yang, M. Bidirectional long short-term memory networks for relation classification. Proc. 29th Pac. Asia Conf. Lang. Inf. Comput. 1, 73–78 (2015).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 5998–6008 (2017).
Loshchilov, I. & Hutter, F. Decoupled weight decay regularization. In Int. Conf. Learn. Represent. (ICLR, 2019).
Akiba T., Sano S., Yanase T., Ohta T., Koyama M. Optuna: a next-generation hyperparameter optimization framework. In Proc. 25th ACM SIGKDD Int. Conf. Knowl. Discov. Data Min. (KDD), 2623–2631 (2019).
Swift, M. L. GraphPad Prism, data analysis, and scientific graphing. J. Chem. Inf. Comput. Sci. 37, 411–412 (1997).
Huang K. et al. Therapeutics data commons: machine learning datasets and tasks for drug discovery and development. In NeurIPS Datasets and Benchmarks (NeurIPS, 2021).
Seo, S., Choi, J., Park, S. & Ahn, J. Binding affinity prediction for protein–ligand complex using deep attention mechanism based on intermolecular interactions. BMC Bioinforma. 22, 1–15 (2021).
Stepniewska-Dziubinska, M. M., Zielenkiewicz, P. & Siedlecki, P. Development and evaluation of a deep learning model for protein–ligand binding affinity prediction. Bioinformatics 34, 3666–3674 (2018).
Wang, Z. et al. OnionNet-2: a convolutional neural network model for predicting protein-ligand binding affinity based on residue-atom contacting shells. Front. Chem. 9, 753002 (2021).
Yang, Z., Zhong, W., Lv, Q., Dong, T. & Yu-Chian Chen, C. Geometric interaction graph neural network for predicting protein–ligand binding affinities from 3d structures (gign). J. Phys. Chem. Lett. 14, 2020–2033 (2023).
Huang, K., Xiao, C., Glass, L. M. & Sun, J. MolTrans: molecular interaction transformer for drug–target interaction prediction. Bioinformatics 37, 830–836 (2021).
Wang, P. et al. Structure-aware multimodal deep learning for drug–protein interaction prediction. J. Chem. Inf. Model. 62, 1308–1317 (2022).
Acknowledgements
This work was supported by Macao Science and Technology Development Fund (0043/2023/AFJ), Macao Polytechnic University (No. RP/ FCA-02/2023). This funding was provided to H.L.
Author information
Authors and Affiliations
Contributions
H.L. and Y.T. conceived the research project. Y.T. developed the MMCLKin framework, evaluated it against baseline methods, applied it to various scenarios, and drafted the manuscript. H.L., X.Y., and H.T. were responsible for project administration, funding acquisition, and resource management. X.G. and Y.T. contributed to the screening and biological validation of the potential kinase inhibitors. W.Z., X.J., X.G., R.L., and Q.L. collected some data. Y.Y., X.G., H.L., Y.L., X.W., and J.A. assisted in the completion of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tero Aittokallio and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tian, Y., Lu, R., Gong, X. et al. Enhancing kinase-inhibitor activity and selectivity prediction through contrastive learning. Nat Commun 16, 10860 (2025). https://doi.org/10.1038/s41467-025-65869-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65869-8