Abstract
Freezing-induced enrichment of organic solutes within ice has long been hypothesized to play a pivotal role in prebiotic chemistry and the origins of life, yet direct evidence for the in-ice aggregation of dilute, water-soluble organics has remained elusive. Here, we employ a conformation-sensitive organic phosphor, 2-phenylbenzothiazole iodide (SNI), to uncover the formation of amino-acid nanoaggregates in water ice. Unusual, amino-acid-specific phosphorescence signatures from SNI-guided investigations prompt us to examine the frozen samples with cryo-transmission electron microscopy (cryo-TEM), which directly reveals uniform nanoaggregates. These nanoaggregates create distinct local microenvironments that influence the photophysical properties of SNI, inducing distinct ground-state conformations that lead to conformation-dependent phosphorescence. Complementary theoretical calculations, molecular dynamics simulations, and temperature-variable Raman spectra suggest that amino acids, such as alanine, undergo a temperature-dependent proton-transfer process from their ammonium to carboxylate groups, enhancing their hydrophobicity and triggering aggregation upon freezing. This enrichment of amino acids in ice may represent a prerequisite condition for polymerization, offering new insights into how primitive icy environments could have influenced peptide-based prebiotic chemistry.
Similar content being viewed by others
Introduction
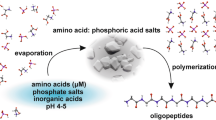
Studies on small organic molecules such as amino acids in ice are of central importance for understanding pre-biotic chemistry1. Water ice matrices not only protect amino acids from cosmic radiation but also facilitate the formation of larger molecules such as peptides and proteins in environments like interstellar clouds, exoplanets, meteorites, and Earth2,3,4,5. For instance, Herrero et al. demonstrated the crucial role of water ice as a medium for the survival of primitive amino acids during the formation of stars and planets6. Calculations made by Kimura and Kitadai on the Gibbs free energy for the polymerization of organic monomers, including amino acids, revealed the thermodynamic favorability of cold-water ice (80–120 K) on surfaces of celestial bodies for peptide formation7. Recently, Rousseau et al. studied the polypeptide formation in clusters of β-alanine amino acids by α-particle impact4. Although these studies confirmed the final products, it remained unclear how amino acids could cluster or aggregate closely enough to react, given that organic substances are many orders of magnitude less abundant than H₂8. One plausible explanation is that freezing water ice excludes organic molecules into the interstices of crystalline ice regions, leading to their enrichment9,10. This process has been observed for various solutes, including certain biomolecules11. While proteins are known to undergo freezing-induced aggregation due to significant entropic changes, it has been uncertain whether amino acids, as smaller molecular entities, exhibit similar behavior. Previous research has provided indirect evidence, based on studies of polypeptides and proteins, that certain amino acids might be prone to phase separation when the temperature is lowered12. However, no direct experimental results ever supported that amino acids phase-separate in water ice; on the contrary, a recent study did not find aggregates in an alanine-water ice mixture13. Here in this study, we present direct observation of microphase separation, i.e., nanoaggregates formation for various amino acids, including alanine. Specifically, we first observed unusual spectroscopic14,15 profiles in SNI phosphorescence16,17,18,19,20,21,22,23 in the solutions, frozen solutions of various naturally occurring amino acids; these solutions were then examined by cryo-TEM and SEM, and were found to contain nanoaggregates of various sizes and shapes; molecular dynamics simulations were employed to give a consistent explanation of the spectral shift accounted for the local chemical environment formed by nanoaggregates. To understand the mechanism of the microphase separation behavior of these hydrophilic molecules, temperature-dependent Raman spectroscopy reveals a sudden disappearance of spectral intensity at 180 K, suggesting a charged-to-neutral amino acid structural transition. These findings support a sequential pathway for prebiotic polypeptide formation in ice24,25,26,27,28,29: amino acid aggregation via nanophase separation as a prerequisite condition, followed by freeze-concentration via ice sublimation, and ultimately polymerization under external energy inputs30,31 (Processes 1–3 in Fig. 1a). Our work highlights this prerequisite first step by providing direct experimental evidence that amino acids indeed form nanoaggregates in ice.
a Proposed Mechanism for Prebiotic Polypeptide Formation in Ice Matrices. The schematic illustrates a potential pathway for the formation of prebiotic polypeptides from amino acids in ice (left). (1) Amino acids in dilute aqueous solutions undergo microphase separation in the crystalline ice matrix. (2) Following aggregation, sublimation of ice leads to the enrichment of amino acid clusters. (3) These aggregates facilitate the polymerization of amino acids into more complex polypeptides, possibly under cosmic radiation or other prebiotic conditions. The inset on the right shows a molecular dynamics snapshot of glutamate molecules (blue: nitrogen atom, red: oxygen atom) and SNI (yellow: sulfur atom) interacting with the surface of ice, where SNI (Iodine atoms have been hidden) experiences changes in ground-state conformations. b The bottom panel highlights the observed phosphorescence (Phos.) shifts in SNI from yellow to green upon conformational changes induced by the rotational motion (left) of SNI interacting with the ice surface. This interaction provides spectroscopic evidence for amino acid nanoaggregation of various microphologies (right), a critical step in the proposed mechanism for peptide polymerization in early Earth conditions. (vibronic energy level: v, singlet state: S, triplet state: T).
In a recent study, we utilized the phosphorescent probe acridinium iodide (ADI) to investigate the microstructural states of water ice and explored its ability to distinguish crystalline and amorphous (glassy) phases from the spectroscopic signatures, as ADI’s phosphorescence properties were highly sensitive to changes in its hydration state (hydrated vs. non-hydrated ADI)10. However, its binary spectral response features pose limitations for more detailed microstructure sensing32,33,34,35,36,37,38,39,40,41,42.
In this work, we have developed a spectroscopic strategy using 2-(2-hydroxyphenyl)−3-methylbenzo[d]thiazol-3-ium iodide (SNI, Fig. 1b), where SNI exhibits enhanced sensitivity due to its phenol functional group and increased intramolecular motion, allowing it to adopt different ground-state configurations upon interacting with different microenvironments, and thus variable phosphorescence behavior reflecting the specific interactions with the external matrix. Surprisingly, we find dramatically different spectroscopic features for water ice samples containing leucine and isoleucine at a concentration of 22.2 mM, respectively. This tiny variation in molecular structure is unlikely to impose such a spectroscopic shift43,44,45,46, if the phosphorescence emission merely reflects the morphological difference of the ice matrix47. Hence, a more plausible mechanism, which involves the formation of amino-acid aggregates is hypothesized: upon freezing, the amino acids form aggregates, which then provide hydrophobic surfaces for the SNI probe. Consequently, the SNI probe can have a whole spectrum of electronic configurations in the ground/excited states. As Fig. 1b indicates, the SNI probe consists of a phenol electron donor and a benzothiazolium electron acceptor. When the two rings are co-planar, the pKa of the phenol is expected to be reduced, so that the SNI exists as a deprotonated anion under neutral pH when it binds to the smoother surface of leucine aggregates. However, upon binding to the more disordered surface of isoleucine aggregates, forced rotation destroys the co-planar structures and increases the electronic density on the phenol group, which should prefer the protonated form. Accordingly, the phosphorescence emission energy is also expected to change from red-shifted to blue-shifted state. Here in this study, we show that our hypothesis is indeed consistent and may shed light on how polypeptides could have been formed during the early period of the universe.
Results
The SNI salt was synthesized by a one-step SN2 reaction, and its purity was confirmed using nuclear magnetic resonance (NMR) spectroscopy, high-resolution mass spectrometry (HRMS), and single-crystal X-ray diffraction (XRD) (Fig. S1–S4). Optically dilute solutions of SNI in deionized water (7 × 10-⁵ M) exhibited a broad absorption band between 260 and 500 nm, with resolved peaks at 290, 328, and 420 nm, characteristic of the π-π* transition of the conjugated SN+ cation (Fig. S5). At room temperature, the SN+ cation in aqueous solution showed no detectable photoluminescence (Fig. S6), presumably due to a high degree of rotational freedoms between the donor (phenyl ring) and acceptor (benzothiazole ring) planes. However, when the SNI solution (7 × 10-⁵ M in H₂O) was cooled at an average rate of approximately 1 °C/s using liquid nitrogen, the ice exhibited green photoluminescence at thermal equilibrium (77 K) with a broad spectroscopic range spanning the entire visible range (Figure 2a, λem = 546 nm, τ = 2.56 ns). When recorded at a delay time of 3 ms, a phosphorescence emission band (λP = 597 nm) with a lifetime of 25.17 ms was noted. The behavior was very similar to the ADI probe in ice, which has been attributed to dehydration-induced charge-transfer phosphorescence between the halide anion and the organic cation phosphor10.
We applied this probe to the water-amino acid system and selected representative water-soluble amino acids – leucine (Leu), phenylalanine (Phe), histidine (His), glutamic acid (Glu), and glutamine (Gln) – each at a concentration of 55.6 mM (Fig. S7). Our initial observations reveal that the addition of trace leucine caused a reduction in the fluorescence band around 546 nm (τ = 3.61 ns, Figs. S8 and S9), accompanied by a more intense phosphorescence band with well-resolved vibronic progressions at 515 and 596 nm (τ = 55.67 ms), making the photoluminescence appear green to the naked eye (Fig. 2a). When testing amino acids containing aromatic group, the fluorescence band intensity at 540 nm (τ = 3.89 ns, Fig. S9) similarly decreased, while the phosphorescence intensity at 585 nm (τ = 65.31 ms) increased for Phe. For His, the fluorescence band emerged at a blue-shifted maximum of 531 nm (τ = 4.34 ns, Fig. 2b), accompanied by a more intense phosphorescence band with well-resolved vibronic progressions at 563 nm (τ = 69.31 ms), which is also blue-shifted. The photoluminescence of SNI is further blue-shifted in the presence of amino acids with higher acidity, e.g., Specifically, Glu exhibited a fluorescence vibrational peak at 511 nm and induced distinct green phosphorescence at 513 nm (τ = 73.93 ms). Similarly, Gln displayed a fluorescence vibrational peak at 515 nm, accompanied by prominent green phosphorescence at 518 nm (τ = 75.54 ms). We also investigated the phosphorescence lifetime of each sample at a wavelength of 550 nm and found that the addition of amino acids to the SNI aqueous solution increased the lifetime. Specifically, the lifetimes for Leu, Phe, His, Glu, and Gln were 56.28 ms, 81.79 ms, 86.01 ms, 71.01 ms, and 74.11 ms, respectively (Fig. S10). Based on the spectroscopic trend, it appears that as the pH values of the aqueous solutions decrease, i.e., Leu/H2O (6.4), Phe/H2O (6.2), His/H2O (7.2), Glu/H2O (3.1), and Gln/H2O (3.8), there is a progressive blue shift in the photoluminescence emission. Since the SNI has a protonatable phenol group, we suspect that the zwitterionic form of SNI gives rise to the yellow phosphorescence while the cationic, protonated SNI is blue-shifted to the green emission.
a Steady-state (dashed line) and delayed emission (solid line) spectra of SNI in water and after the addition of various amino acids at 77 K. From top to bottom: SNI in H₂O, SNI in Leu/H₂O, and SNI in Phe/H₂O. [SNI] = 7 × 10-⁵ M, [amino acids] = 55.6 mM, λex = 365 nm, delay time = 3 ms; insets: images showing fluorescence (left) and phosphorescence (right) at 77 K excited by a 365 nm UV lamp. b The same as a. From top to bottom: SNI in His/H₂O, SNI in Glu/H₂O, and SNI in Gln/H₂O. c Phosphorescence lifetime curves of SNI aqueous solution in the presence of various amino acids at 77 K. d Corresponding fitted lifetime values from the phosphorescence decay curves in (c) (1: H2O, 2: Leu/H2O, 3: Phe/H2O, 4: His/H2O, 5: Glu/H2O, 6: Gln/H2O). e Time-lapsed phosphorescence images after the addition of Leu and Glu to SNI aqueous solutions at 77 K, with UV on/off cycles showing the emission decay over time.
To test the hypothesis, the SNI emission spectra in the presence of two isomeric amino acids (Leu and Ile) with almost identical pKa values (pKa of α-carboxyl group of ~2.4 and pKa of α-amino group of ~9.6) were measured. Unexpectedly, we found dramatically different spectroscopic features for water ice samples containing the two isomers, respectively, at a concentration of 22.2 mM (Fig. S11). This tiny variation in molecular structure is unlikely to impose any noticeable spectroscopic shift, if the photoluminescence of SNI emission merely reflects the pH values of the ice matrix. Therefore, the pH-induced emission shift hypothesis cannot be correct or complete, at least. Since large π-conjugated rings are, in general, hydrophobic, we reasoned that SNI is hydrophobic enough to interact with microphases composed of amino acids. Despite the structural similarity between Leu and Ile, their packing in the aggregates could vary dramatically, as has been frequently observed for organic crystals48. Therefore, such morphological differences could induce the water-ice microenvironments that leading to conformational differences for the SNI probe molecules. Specifically, the SNI probe contains a phenol donor and a benzothiazolium acceptor, with significant electron density differences depending on its conformational state. This dual nature allows it to interact differently with the amino acid aggregates depending on their surface characteristics. As such, a more planar ground-state conformation allows better electron flow from the donor to the acceptor moiety, which leads to donor deprotonation and red-shifted photoluminescence. Vice versa, a more twisted ground-state conformation inhibits excited-state deprotonation and thus gives rise to blue-shifted emission (Fig. 1b).
To explore the possibility of an SNI-aggregate direct contact, molecular dynamics simulations were first performed within a water-ice matrix (Fig. 3a–f, Figs. S12-S15). The simulations were conducted within a crystalline water-ice framework, represented by gray grids that depict the organized hydrogen-bond network characteristic of hexagonal ice. Our simulations reveal the nucleation and crystallization of an aqueous solution containing amino acids at 240 K (with and without the presence of SNI), as shown in Movies S1–S6. The MD simulations were carried out at 240 K, which is below the melting temperature of ice Ih in the TIP4P-2005 water model (~252 K at 1 atm). This temperature was chosen because it allows the simulation of ice nucleation and growth, which is essential to study ice-induced amino acid aggregation. Additionally, iodide ions dissolve into the solution at 250 K and subsequently enter the ice phase during crystallization. Within this ice matrix, Leu, Ile, and Glu form distinct aggregates at a concentration of 166.8 mM, regardless of the presence of SNI (Figs. S12–S15). Leucine aggregates (Fig. 3a) appeared as compact, hydrophobic clusters, displaying relatively smooth surface morphologies. In contrast, isoleucine aggregates (Fig. 3b), while structurally similar in their molecular composition, exhibited a more disordered surface, indicative of less efficient packing. Meanwhile, glutamic acid aggregates (Fig. 3c) formed more extended, hydrophilic structures, consistent with their charged side chains. The electrostatic potential maps of these aggregates (Figs. 3d–3f) highlight significant differences in surface chemistry. Leucine aggregates displayed predominantly neutral, hydrophobic surfaces (Fig. 3d). Isoleucine aggregates, although hydrophobic, exhibited more varied topography due to structural disorder (Fig. 3e). In contrast, glutamic acid aggregates showed mixed electrostatic features, with prominent negatively charged regions corresponding to carboxylate groups and positively charged areas corresponding to amino groups (Fig. 3f). To provide direct experimental evidence of amino-acid aggregation, the three frozen samples were examined with transmission electron microscopy under cryogenic conditions (cryo-TEM). For pure water at 77 K, the cryo-TEM image appeared very smooth, indicating a lack of observable nanoscale structures (Fig. S16). Similarly, the aqueous solution of SNI (7 × 10-⁵ M) also exhibited a smooth structure, suggesting that SNI does not disrupt the microstructure of water or form aggregates at the nanoscale (Fig. S17). However, the Cryo-TEM images reveal dispersed, elongated leucine nanoaggregates within the ice matrix. A total of 39 aggregates are observed, exhibiting a relatively uniform size distribution with diameters of 3-5 nm and lengths ranging from 5 to 20 nm (Fig. 3g and Fig. S18). This aggregation behavior aligns with leucine’s hydrophobic nature and its tendency to form tightly packed clusters in ice. In contrast, the isoleucine system shows fewer aggregates (20 in total) with a more irregular morphology and slightly smaller sizes, with lengths between 5 and 10 nm (Fig. 3h and Fig. S19). These findings are consistent with molecular dynamics simulations (Fig. 3b), which suggest a looser packing arrangement for isoleucine, resulting in fewer aggregates. For the Glu-containing ice sample, Cryo-TEM images (Fig. 3i and Fig. S20) reveal relatively small aggregates (5-15 nm in length) but in a greater number (38 aggregates), likely due to the charged side chains of Glu enhancing kinetic stability. It is remarkable to observe such nanoaggregates of amino acids at very low molar concentration (Figs. S18–S20), which are also consistent with the results of molecular dynamics simulations. We also performed Cryo-TEM on amino acid solutions without SNI, revealing the presence of small aggregates in all three samples. Leu, Ile, and Glu formed aggregates ranging from 2 to 8 nm in length, with approximately 100-150 aggregates observed (Figs. S21-22). Comparing these results with those obtained after adding SNI (Fig. 3g–i), we found that the aggregate morphology became more distinct upon SNI addition. Grayscale analysis of the images further confirmed that SNI enhanced image contrast (Fig. S23). This is likely due to the electron-rich iodine in SNI, which enhances contrast in Cryo-TEM images. Similar nanoaggregates were also observed in cryo-TEM images of aqueous solutions containing other amino acids, including phenylalanine, histidine, glutamine, valine, and alanine, further supporting the generality of our findings (Figs. S24–28).
Molecular simulation snapshots of SNI in water and after addition of Leu (a), Ile (b), and Glu (c) at 240 K. A locally enlarged surface potential model after adding Leu (d), Ile (e), and Glu (f) to SNI aqueous solution at 240 K, describing the interaction distance between amino acids and probes (Water molecules have been hidden). Cryo-TEM images of SNI in water and after addition of Leu (g), Ile (h), and Glu (i) at 77 K. [SNI] = 7 × 10-⁵ M, [amino acids] = 166.8 mM. The 2D excitation-emission-intensity phosphorescence spectra acquired at 77 K after adding Leu (j), Ile (k), and Glu (l) to SNI aqueous solution. [SNI] = 7 × 10⁻⁵ M, [amino acids] = 55.6 mM, delay time = 3 ms.
To further investigate the behavior of nanoaggregates, we systematically varied the amino acid molar concentrations and analyzed the resulting structures using cryo-TEM. When the Glu concentration was doubled to 333.6 mM (Fig. S29), a significant increase in aggregate formation was observed. Conversely, reducing this concentration six-fold to 55.6 mM resulted in a sparser distribution of aggregates (Fig. S30). A further 20-fold reduction to 16.7 mM left only a few remaining nanostructures (Fig. S31). Notably, these concentration variations did not significantly affect particle size, which remained consistently within the range of 1–7 nm under all conditions. We also examined the influence of Leu concentration in both the Leu/H₂O and SNI/Leu/H₂O systems. In the Leu/H₂O system, decreasing the concentration to 55.6 mM and 5.6 mM led to a reduction in the number of aggregates, with lengths ranging from 1–4 nm (Fig. S32). Similarly, in the SNI/Leu/H₂O system, lowering the concentration resulted in smaller aggregates, typically 1–5 nm in length (Fig. S33). For comparison, we analyzed peptides and proteins under identical conditions using cryo-TEM (Figs. S34, S35). Interestingly, soybean peptides exhibit sizes comparable to those of the amino acid aggregates observed in our study (Fig. S35).
We also performed additional molecular dynamics simulations on amino acids/water/SNI systems. Based on three independent MD runs, we found that all systems consistently form relatively large aggregates containing more than ten amino acids within ~400 ns (Figs. S37–39). The amino acid aggregates observed in Fig. 3g–i were obtained in vitrified ice. By varying the cooling rate, we also prepared crystalline ice samples (Fig. S40), in which similar aggregates were detected. These results indicate that nanocluster formation occurs in both crystalline and vitrified ice. Therefore, the emergence of nanoclusters during freezing is driven by the combined effects of heterogeneous ice microenvironments and non-equilibrium phase separation, rather than solely by ice crystal exclusion.
To characterize the probe’s molecular conformation, we measured the 2D-phosphorescence spectra of an aqueous SNI solution at 77 K after adding Leu, Ile, and Glu (Figs. 3j–3l), respectively. In these spectra, different ground-state conformations are manifested as various intensity distributions. For example, the SNI solution with Leu (Fig. 3j) shows the strongest intensity at [427 nm, 604 nm]; while in Fig. 3k, there are two prominent intensity distributions at [427 nm, 604 nm] and [292 nm, 590 nm] with the latter being more intense. For Glu (Fig. 3l), however, the strongest intensity is at [292, 510 nm] without the longer wavelength phosphorescence component observed at all. Based on the spectroscopic data, we assign the [427 nm, 604 nm] peak as a contribution from the coplanar structure of the SNI probe and the [292 nm, 510 nm] peak from the twisted ground-state geometry. It can be clearly seen that by interacting with a more ordered and hydrophobic surface such as Leu aggregates, the SNI probe extends a full π-conjugation between the donor and acceptor to generate a phosphorescence maximum at 604 nm (Fig. 3j), as the lowest triplet state energy is usually dependent on the length of electronic conjugation: the longer the conjugation, the lower the triplet-state energy. When the SNI probe interacts with the Ile aggregates, the newly appeared main excitation at 292 nm indicates a significantly reduced π conjugation length (Fig. 3k), consistent with a twisted molecular geometry. The fact that the phosphorescence emission maximum spans a wide range from 500-600 nm also suggests that many different conformations co-exist. For the Glu sample, the most intense spot at [292 nm, 510 nm] is also characteristic of a twisted geometry (Fig. 3l). However, an additional, comparable hot spot at [355, 570 nm] that is relatively weak in the Leu and Ile systems (Fig. 3j and k) could be indicative of a Glu-hydrogen-bonded SNI between the carboxylic acid and the phenol.
To further elucidate how protonation states and molecular conformations influence the photoluminescence behavior of SNI, we employed theoretical calculations based on X-ray diffraction (XRD)-derived coordinates (Figs. 4a and 4b, S41 and S42). By systematically exploring various protonation scenarios and conformational configurations, these calculations provide deeper insight into the interplay of electronic structure, conjugation length, and intermolecular interactions that govern the observed phosphorescence features. The orbital distributions of the lowest singlet (S1) and triplet (T1) excited states of the deprotonated SN+ zwitterion (Fig. 4c) show typical π-π* transitions, with S1 and T1 state energies of 2.69 eV and 1.94 eV, respectively. The large singlet-triplet energy splitting (ΔEST = 0.75 eV) corresponds to an almost pure3π-π* phosphorescence state with a ms-lifetime (τ = 25.17 ms). Calculations of the spin-orbit coupling (SOC) energy indicate that intersystem crossing (ISC) primarily occurs via the S1-T2 channel (SOC energy = 3.0 cm-1). Upon protonation, the S1 state energy increases to 3.18 eV, and that of T1 rises to 2.30 eV (Fig. 4d). These results agree with experimental observations: with a measured pKa value of 6.0, the SNI probe exhibits a blue-shifted phosphorescence emission maximum of 514 nm below pH = 3 and a red-shifted phosphorescence maximum of 630 nm above pH = 7 (Fig. 4e) at 77 K (Fig. S43 and Table S2). However, as has been discussed, the pH effect cannot fully account for the observed photoluminescence difference for the Leu and Ile water-ice samples. A plausible explanation is that the benzene and benzothiazole groups are freely rotatable, which could result in diverse conformations for SNI upon interaction with the nanoaggregates of amino acids. To explore the relationship between SNI luminescence and rotational conformation, we calculated the phosphorescence emission energy as a function of the dihedral angle between the benzothiazole and phenyl rings (Fig. 4f and S44). As the dihedral angle increases, the planes of the two groups become more orthogonal. Specifically, from 0° to 90° or 180° to 270°, the emission energy rises to ~3.0 eV, leading to a blue shift in phosphorescence. Conversely, as the dihedral angle transitions from 90° to 180° or 270° to 360°, the phenyl and benzothiazole groups become increasingly coplanar, leading to a decrease in emission energy to ~2.2 eV. A coupled consequence is that the rotation disrupts the coplanar structure of the SNI and concomitantly increases the electron density on the phenol group, which should be more inclined to the protonation form. As a result, the combined pH and conformation effects together can then be used to explain the experimental phosphorescence findings shown in Fig. 3j–I.
Electron density distribution and energy levels of the frontier molecular orbitals of the deprotonated SNI zwitterion (a) and protonated SNI (b). Electron and hole distributions for excited states and the spin-orbit coupling (SOC) matrix of the deprotonated SNI zwitterion (c) and protonated SNI (d). e Comparison of phosphorescence emission wavelengths of SNI in water at different pH levels at 77 K. [SNI] = 7 × 10−⁵ M, λex = 365 nm, delay time = 3 ms. f Relationship between phosphorescence emission energy and the dihedral angle between benzothiazole and phenyl rings.
Finally, we show that the nanoaggregation, or microphase separation, of amino acids in water ice is most likely caused by an intramolecular proton transfer49 process, i.e., amino acids in an aqueous solution exist in the form of zwitterion at room temperature, but as the temperature decreases, protons from the positively charged amino group transfer to the negatively charged carboxylate group (The model inserted in Fig. 5a), increasing their hydrophobicity and thus the tendency to aggregate. One of the main driving forces for this proton transfer process is the sharply decreased hydration ability of water molecules at a reduced temperature, so that the zwitterion form is no longer stabilized. In a previous work, we have observed that the hydration of acridinium iodide is largely suppressed at 110 K10. For zwitterionic amino acids, however, we expect the temperature to be higher than 110 K since the charges are not permanent. To validate this hypothesis, we conducted temperature-dependent Raman spectroscopy of an aqueous solution of alanine due to its excellent solubility (Fig. 5a). At a concentration of 0.50 M, we observed strong peaks at 864, 1318, 1370, 1427, and 1477 cm-¹, which gradually weakened and vanished abruptly at 180 K. In addition, a blue shift for peaks at 864 and 935 cm-1 (assigned as the stretching motion of C-C-C)50 to 874 and 945 cm−1 was also noted51. Such changes are consistent with a transition from the zwitterionic to the neutral form of alanine for two main reasons. First, in its zwitterionic state, alanine is more polarizable due to the presence of the carboxylate anion. Upon transitioning to the neutral form, protonation of the carboxylate group reduces polarizability. Secondly, a higher frequency in the motion of the carbons connecting to carbonyl group clearly indicates reduced electronic density, consistent with protonation. Temperature-dependent Raman spectroscopy at different concentrations supports proton transfer as a mode of intramolecular transfer (Fig. S45). Therefore, these spectral changes provide strong evidence for low-temperature-induced proton transfer. We note, however, that the observed disappearance of Raman signals could also be attributed to physical effects such as enhanced light scattering upon freezing and changes in crystal orientation. This phenomenon is therefore likely the result of a combination of these factors.
a Temperature-dependent Raman spectra of alanine in water, [Ala] = 0.50 M, λex = 532 nm. b Steady-state (dashed line) and delayed emission (solid line) spectra of SNI in solid-state amino acids at 77 K. From top to bottom: SNI@Leu, SNI@Ile, and SNI@Glu, SNI/amino acids (mass ratio) = 1/500, λex = 365 nm, delay time = 3 ms. Inserts: stick-and-ball chemical structures of related amino acids. SEM images of freeze-dried aqueous solutions of leucine (c), isoleucine (d), and glutamic acid (e), illustrating variations in morphologies and surface structures across different amino acids, [amino acids] = 166.8 mM.
As this point, we have experimentally verified that, for one, amino acids form nanoaggregates at low temperature due to an intramolecular proton transfer process to increase hydrophobicity, and that, for another, the SNI probe exhibits both pH- and conformation-dependent photoluminescence, particular phosphorescence. We now show that the SNI probe exhibits almost identical phosphorescence features when the frozen solutions of Leu, Ile, and Glu were sublimed to yield solid-state amino acids. As shown in Fig. 5b, SNI@Leu exhibits phosphorescence at 599 nm (τ = 1.56 ms), while SNI@Ile and SNI@Glu display phosphorescence emission maxima at 573 nm (τ = 1.31 ms) and 525 nm (τ = 1.16 ms), demonstrating notable differences in emission wavelengths (Figs. S46-S47). These results align with the phosphorescence colors of SNI in amino acid aqueous solutions (Fig. S48): SNI emits yellow phosphorescence (596 nm) in Leu/H₂O, greenish yellow (570 nm) in Ile/H2O and green phosphorescence (515 nm) in Glu/H₂O (Fig. 5b). This consistency indicates that, even in the solid state, amino acids can influence the ground-state electronic configuration of the SNI probe through short-range interactions. For a closer examination of the large-scale aggregation behavior of amino acids in the ice matrix, we used scanning electron microscopy (SEM) to examine solid samples obtained after the lyophilization of amino acid aqueous solutions. Figures 5c–5e show the microstructures of the sublimed samples, which reveal significant differences in morphologies. The SEM images of leucine (Figs. 5c, S49) reveal plate-like structures ranging from 1 to 10 um in length. At higher magnification (center and right), these plates appear irregular in shape with smooth surfaces.
In contrast, isoleucine aggregates (Figs. 5d, S50) exhibit similar block-like structures but are comparatively smaller, ranging from 1 to 5 um. Glutamic acid forms significantly smaller and more dispersed aggregates (Figs. 5e, S51), with lengths between 100 and 500 nm. The observed morphological variations among leucine, isoleucine, and glutamic acid indicate that subtle structural differences between amino acids may influence packing behavior.
Discussion
In summary, this study demonstrates that freezing induces the formation of amino acid nanoaggregates within water ice, a process critically influenced by the conformation-dependent behavior of the organic phosphor SNI. By integrating phosphorescence spectroscopy, cryo-transmission electron microscopy (Cryo-TEM), and molecular simulations, we have provided direct evidence that amino acids such as leucine, isoleucine, and glutamic acid aggregate into uniform nano-sized structures upon freezing. These nanoaggregates interact specifically with the SNI probe, which adopts distinct conformations based on the aggregate’s morphology, resulting in characteristic phosphorescence signatures. Our findings indicate that under freezing conditions, amino acids undergo a proton-transfer from their ammonium to carboxylate groups, increasing their hydrophobicity and facilitating aggregation. This mechanism not only enhances the local concentration and stabilization of amino acids in ice but also creates conducive environments for prebiotic polymerization processes like peptide formation. The conformation-dependent phosphorescence of SNI offers a spectroscopic tool for probing molecular interactions within ice matrices. The current study signifies ice as a medium for prebiotic molecular evolution and provides foundational insights into the physical and chemical processes that may have contributed to the origin of life in low-temperature environments.
Methods
Materials
2-(2-Hydroxyphenyl)benzothiazole, iodomethane were obtained from Aladdin Reagent (Shanghai) Co., Ltd. All amino acids were acquired from Sigma-Aldrich LLC. Bovine serum albumin and peptides were acquired from Shanghai Macklin Biochemical Technology. All other reagents and solvents were acquired from Sinopharm Chemical Reagent Co., Ltd and used as received. Water was deionized with a Milli-Q SP reagent water system (Millipore) to a specific resistivity of 18.2 MΩ.cm. All amino acid aqueous solution samples in this article were dissolved by ultrasound and heating, and then filtered through an aqueous phase filter membrane with a pore size of 0.1 μm.
Instrumentation
NMR spectra were recorded on a Bruker AV400 NMR spectrometer operated in the Fourier transform mode. NMR spectra were recorded at 400 MHz for 1H and 101 MHz for 13C. 1H NMR spectra are represented as follows: chemical shift, multiplicity (s = singlet, d = doublet, t = triplet, q = quartet, m = multiplet, br = broad), integration, and coupling constant (J) in Hertz (Hz). 1H NMR spectra were referenced to the signal for residual protio DMSO at 2.50 ppm. Electrospray ionization (ESI) mass spectra were recorded on an Acquity UPLC-Xevo G2 QT mass spectrometer (Waters). UV/Vis absorption spectra were recorded on a PerkinElmer Lambda 465 UV-Vis spectrometer. Steady-state emission spectra were recorded on a Horiba FluoroMax-4 spectrofluorometer (Horiba Scientific). The light source (xenon lamp) that supplies UV excitation is focused onto the entrance slit of the excitation monochromator with an elliptical mirror. According to the manufacturer, the light source is a vertically mounted 150-W ozone-free cw xenon arc lamp. The distance between excitation and the sample is ~120 cm, and the integration time is 0.1 s. Phosphorescence lifetime data were acquired with a 1 MHz LED laser with the excitation peak at 374 nm (SpectraLED-370). Lifetime data were analyzed with Data Station v6.6 (Horiba Scientific). The delayed emission spectra were recorded on a Horiba FluoroMax-4 spectrofluorometer as well, using a 10-W xenon flash lamp as the excitation source with an integration time of 0.1 s, delay time = 3 ms. Single crystal samples were obtained by slow evaporative crystallization of compounds in methanol, and single crystal data was collected on a Bruker Smart APEXII CCD diffractometer using graphite monochromated Cu- Kα radiation (λ = 1.54178 Å). Raman spectrum is measured by a self-built instrument. In brief, a CW laser (Coherent, 532 nm) with a power of 5 W was employed as the excitation light. The Raman scattering light was dispersed by a triple-monochromator system (Acton Research, TriplePro) and recorded in the region of 800–1700 cm−1 by a liquid nitrogen-cooled CCD detector (Princeton Instruments, Spec-10:100B) at a spectral resolution of ~1 cm−1. The Raman shift was carefully calibrated using the standard spectral line of a mercury lamp. SEM images used for the morphology study were obtained via ZEISS GeminiSEM 500 Schottky field emission scanning electron microscope.
Cryo-TEM sample preparation and data collection
3 μL of sample (SNI aqueous solution and after addition of amino acids) was applied to freshly glow-discharged QUANTIFOIL (R1.2/1.3, 300 mesh, holey carbon films) copper grids. The grids were blotted with filter paper for 3.5 s and zero blotting force. Then, the grids were plunged into liquid ethane cooled by liquid nitrogen using a Vitrobot Mark IV (FEI) under 100% humidity at 8 °C. Micrographs were collected with EPU 2 software on a Glacios 200 kV transmission electron microscope (Thermo Fisher) equipped with a Falcon3 camera.
Theoretical calculation methods
Molecular dynamics (MD) simulations were conducted using the OPLS-AA force field and the TIP4P-2005 water model with the GROMACS 2018 package. The study focused on the impact of ice growth on the behavior of Leu, Ile, and Glu, as well as the influence of C14H12NOSI. To enhance ice nucleation rate, the crystal/liquid coexistence state was employed. An ice Ih slab was generated using Genice v.2 software, and Leu, Ile, and Glu molecules were individually solvated in the liquid water slab alongside C14H12NOSI molecules. Each system underwent equilibration for 1 ns at 250 K. Subsequently, the ice/water system was equilibrated for 400 ns at 240 K under ambient pressure. All simulations were carried out in the isothermal-isobaric (NPT) ensemble using the V-rescale thermostat and Parrinello-Rahman barostat. Periodic boundary conditions (PBC) were applied in the x, y, and z directions. A 1.0 nm cutoff was applied for Lennard-Jones interactions, and long-range electrostatics were treated with the Particle-Mesh-Ewald (PME) method.
Conformational searches for the molecules in aqueous solution were conducted using the GFN-xTB method with the molclus software, and the lowest-energy conformer was selected for subsequent quantum chemical calculations.
Ground-state (S0) geometry optimizations were performed using Gaussian 16 (revision C.03) at the B3LYP/def2-SVP level of theory with empirical dispersion correction (GD3BJ), incorporating solvent effects via the SMD model for water. Vibrational frequency analyses were conducted to confirm that the optimized ground-state geometries exhibited no imaginary frequencies, ensuring stable minima. The distribution and energies of the molecular frontier orbitals (HOMO and LUMO) were calculated based on the optimized ground-state geometry using the same B3LYP/def2-SVP level of theory with GD3BJ and SMD solvation.
The geometry optimizations for the first excited singlet (S1) and triplet (T1) states were performed using Gaussian 16 (revision C.03) at the ωB97XD/def2-TZVP level of theory, incorporating solvent effects via the SMD model for water. Time-dependent density functional theory (TD-DFT) within the Tamm-Dancoff approximation (TDA) was employed, considering 10 singlet or triplet states with the lowest-energy root selected, and the IOp(9/40 = 4) option was used to facilitate hole-charge analysis, as described in [Carbon 2020, 10.1016/j.carbon.2020.05.023], with subsequent processing using Multiwfn software. Visualization of the results was performed using VMD software.
Spin-orbit coupling (SOC) matrix elements were computed using ORCA 5.0.4 based on the optimized S1 geometry, employing the ωB97X-D3 functional with the DKH2 scalar relativistic Hamiltonian for all-electron relativistic corrections, the DKH-def2-TZVP basis set (supplemented with SARC-DKH-TZVP for iodine-containing systems), SARC/J auxiliary basis, and RIJCOSX approximation. The CPCM solvation model with SMD parameters for water was applied, and TD-DFT calculations included 30 roots with SOC enabled under TDA. For molecules containing iodine, the SOC computation utilized the DKH2 Hamiltonian to account for all-electron scalar relativistic effects. In the calculation under solvent environment, the source of molecular coordinates can be said to be based on single crystal coordinates, and conformational search was conducted.
Data availability
All relevant data generated in this study are provided in the supplementary information and are also available from the authors upon request. Source data are provided as a Source Data file. CCDC 2495887 contains the supplementary crystallographic data for this paper. These data are provided free of charge by The Cambridge Crystallographic Data Centre. Source data are provided with this paper.
References
Ioppolo, S. et al. A non-energetic mechanism for glycine formation in the interstellar medium. Nat. Astron. 5, 197–205 (2021).
Krishnamurthy, R. & Hud, N. V. Introduction: Chemical evolution and the origins of life. Chem. Rev. 120, 4613–4615 (2020).
Kaiser, R. I., Stockton, A. M., Kim, Y. S., Jensen, E. C. & Mathies, R. A. On the formation of dipeptides in interstellar model ices. Astrophys. J. 765, 111 (2013).
Rousseau, P. et al. Polypeptide formation in clusters of β-alanine amino acids by single ion impact. Nat. Commun. 11, 3818 (2020).
Frenkel-Pinter, M., Samanta, M., Ashkenasy, G. & Leman, L. J. Prebiotic Peptides: Molecular Hubs in the Origin of Life. Chem. Rev. 120, 4707–4765 (2020).
Maté, B., Tanarro, I., Escribano, R., Moreno, M. A. & Herrero, V. J. Stability of extraterrestrial glycine under energetic particle radiation estimated from 2 kev electron bombardment experiments. Astrophys. J. 806, 151 (2015).
Kimura, J. & Kitadai, N. Polymerization of building blocks of life on Europa and other icy moons. Astrobiology 15, 430–441 (2015).
Tian, F., Toon, O. B., Pavlov, A. A. & De Sterck, H. A hydrogen-rich early earth atmosphere. Science 308, 1014–1017 (2005).
Kim, K. et al. Nitrite-Induced Activation of Iodate into Molecular Iodine in Frozen Solution. Environ. Sci. Technol. 53, 4892–4900 (2019).
Liu, H. et al. Water-Ice Microstructures and Hydration States of Acridinium Iodide Studied by Phosphorescence Spectroscopy. Angew. Chem. lnt. Ed. 63, e202405314 (2024).
Thorat, A. A., Munjal, B., Geders, T. W. & Suryanarayanan, R. Freezing-induced protein aggregation - Role of pH shift and potential mitigation strategies. J. Control. Release 323, 591–599 (2020).
Hong, Y. et al. Hydrophobicity of arginine leads to reentrant liquid-liquid phase separation behaviors of arginine-rich proteins. Nat. Commun. 13, 7326 (2022).
Hao, J., Giovenco, E., Pedreira-Segade, U., Montagnac, G. & Daniel, I. Compatibility of Amino Acids in Ice Ih: Implications for the Origin of Life. Astrobiology 18, 381–392 (2018).
Baryshnikov, G., Minaev, B. & Ågren, H. Theory and Calculation of the Phosphorescence Phenomenon. Chem. Rev. 117, 6500–6537 (2017).
Zhao, W., He, Z. & Tang, B. Z. Room-temperature phosphorescence from organic aggregates. Nat. Rev. Mater. 5, 869–885 (2020).
Zhang, Y. et al. Ultraviolet irradiation-responsive dynamic ultralong organic phosphorescence in polymeric systems. Nat. Commun. 12, 2297 (2021).
Zhang, Y. et al. Cross-linked polyphosphazene nanospheres boosting long-lived organic room-temperature phosphorescence. J. Am. Chem. Soc. 144, 6107–6117 (2022).
Zhang, X. et al. Ultralong UV/mechano-excited room temperature phosphorescence from purely organic cluster excitons. Nat. Commun. 10, 5161 (2019).
Zhang, K. Y. et al. Long-lived emissive probes for time-resolved photoluminescence bioimaging and biosensing. Chem. Rev. 118, 1770–1839 (2018).
Zhang, G., Palmer, G. M., Dewhirst, M. W. & Fraser, C. L. A dual-emissive-materials design concept enables tumour hypoxia imaging. Nat. Mater. 8, 747–751 (2009).
Yu, J. et al. Efficient visible-light-activated ultra-long room-temperature phosphorescence triggered by multi-esterification. Angew. Chem. lnt. Ed. n/a, e202316647 (2023).
Zhou, B. & Yan, D. Hydrogen-bonded two-component ionic crystals showing enhanced long-lived room-temperature phosphorescence via TADF-assisted Förster resonance energy transfer. Adv. Funct. Mater. 29, 1807599 (2019).
Zhou, B., Xiao, G. & Yan, D. Boosting wide-range tunable long-afterglow in 1D metal–organic halide micro/nanocrystals for space/time-resolved information photonics. Adv. Mater. 33, 2007571 (2021).
Bencs, F. et al. Chemical evolution of early macromolecules: from prebiotic oligopeptides to self-organizing biosystems via amyloid formation. Chem. Eur. J. 31, e202404669 (2025).
Sato, A. et al. First-principles study of the formation of glycine-producing radicals from common interstellar species. Mol. Astrophys. 10, 11–19 (2018).
Wołos, A. et al. Synthetic connectivity, emergence, and self-regeneration in the network of prebiotic chemistry. Science 369, eaaw1955 (2020).
Sebaaly, A. P., van Rijn, F., Hanna, K. & Boily, J.-F. Ice as a kinetic and mechanistic driver of oxalate-promoted iron oxyhydroxide dissolution. Proc. Natl. Acad. Sci. USA 122, e2507588122 (2025).
Bada, J. L., Bigham, C. & Miller, S. L. Impact melting of frozen oceans on the early Earth: implications for the origin of life. Proc. Natl. Acad. Sci. USA 91, 1248–1250 (1994).
Gao, B., Wyttenbach, T. & Bowers, M. T. Protonated Arginine and protonated lysine: hydration and its effect on the stability of salt-bridge structures. J. Phys. Chem. B 113, 9995–10000 (2009).
Ciesla, F. J. & Sandford, S. A. Organic synthesis via irradiation and warming of ice grains in the solar Nebula. Science 336, 452–454 (2012).
Singh, A., Kaur, S., Kaur, J. & Singh, P. Transformation of gas-phase amino acid clusters to dipeptides: a nice approach to demonstrate the formation of prebiotic peptides. Rapid Commun. Mass Sp. 28, 2019–2023 (2014).
Gu, L. et al. Colour-tunable ultra-long organic phosphorescence of a single-component molecular crystal. Nat. Photonics 13, 406–411 (2019).
Yang, J. et al. The influence of the molecular packing on the room temperature phosphorescence of purely organic luminogens. Nat. Commun. 9, 840 (2018).
Xu, Z. et al. Supercooled liquids with dynamic room temperature phosphorescence using terminal hydroxyl engineering. Angew. Chem. Int. Ed. 62, e202301564 (2023).
Wang, J., Huang, Z., Ma, X. & Tian, H. Visible-light-excited room-temperature phosphorescence in water by Cucurbit[8]uril-mediated supramolecular assembly. Angew. Chem. Int. Ed. 59, 9928–9933 (2020).
Su, Y. et al. Ultralong room temperature phosphorescence from amorphous organic materials toward confidential information encryption and decryption. Sci. Adv. 4, eaas9732 (2018).
Nie, X. et al. Kinetic and thermodynamic control of tetraphenylethene aggregation-induced emission behaviors. Aggregate 3, e165 (2022).
Ma, L. et al. A universal strategy for tunable persistent luminescent materials via radiative energy transfer. Angew. Chem. Int. Ed. 61, e202115748 (2022).
Liu, D. et al. Molecular core–shell structure design: Facilitating delayed fluorescence in aggregates toward highly efficient solution-processed OLEDs. Aggregate 3, e164 (2022).
Li, D. et al. Completely aqueous processable stimulus responsive organic room temperature phosphorescence materials with tunable afterglow color. Nat. Commun. 13, 347 (2022).
Li, D. et al. Amorphous metal-free room-temperature phosphorescent small molecules with multicolor photoluminescence via a host–guest and dual-emission strategy. J. Am. Chem. Soc. 140, 1916–1923 (2018).
Kabe, R. & Adachi, C. Organic long persistent luminescence. Nature 550, 384–387 (2017).
Cai, S., Yao, X., Ma, H., Shi, H. & An, Z. Manipulating intermolecular interactions for ultralong organic phosphorescence. Aggregate 4, e320 (2023).
Ding, B., Ma, L., Huang, Z., Ma, X. & Tian, H. Engendering persistent organic room temperature phosphorescence by trace ingredient incorporation. Sci. Adv. 7, eabf9668 (2021).
Dai, W. et al. Controllable modulation of efficient phosphorescence through dynamic metal-ligand coordination for reversible anti-counterfeiting printing of thermal development. Adv. Funct. Mater. 33, 2210102 (2023).
Chen, X. et al. Versatile room-temperature-phosphorescent materials prepared from n-substituted naphthalimides: emission enhancement and chemical conjugation. Angew. Chem. Int. Ed. 55, 9872–9876 (2016).
Su, H. et al. Functional roles of polymers in room-temperature phosphorescent materials: modulation of intersystem crossing, air sensitivity and biological activity. Angew. Chem. Int. Ed. 62, e202218712 (2023).
Gan, N. et al. Manipulating the stacking of triplet chromophores in the crystal form for ultralong organic phosphorescence. Angew. Chem. Int. Ed. 58, 14140–14145 (2019).
Kumar, M., Sinha, A. & Francisco, J. S. Role of double hydrogen atom transfer reactions in atmospheric chemistry. Acc. Chem. Res. 49, 877–883 (2016).
Kumar, S. et al. Infrared, Raman and electronic spectra of alanine: A comparison with ab intio calculation. J. Mol. Struct. 791, 23–29 (2006).
Wen, C.-I., Yu, C.-N., Thirumalaivasan, N. & Hiramatsu, H. 532-nm-excited hyper-Raman spectroscopy of amino acids. J. Raman Spectrosc. 52, 641–654 (2021).
Acknowledgements
We thank the National Natural Science Foundation (21975238 to G. Z. and 22473064 to W. Z.), Innovation Program for Quantum Science and Technology (2021ZD0303301 to G. Z.), and the Fundamental Research Funds for the Central Universities (WK9990000099 to G. Z.) for financial support. This research was also supported by the advanced computing resources provided by the Supercomputing Center of the USTC. The authors are particularly grateful to Senior Engineer Zhongliang Zhu at the School of Life Sciences at USTC for his contribution to the single-crystal X-ray diffraction data analysis. The authors also thank Senior Engineer Shengquan Fu at the School of Life Sciences at USTC for his contribution to the scanning electron microscope data analysis.
Author information
Authors and Affiliations
Contributions
H. L. prepared and characterized the samples, processed the experimental data, and designed the figures. H. S. carried out all theoretical calculations. N. C. assisted in Raman spectra measurements. B. Z. provided helpful discussions. A. C. assisted in photo taking. X. C. assisted in PL measurements. X. P. Z. provided a helpful discussion on the figures. X. G. Z. assisted in Raman experiments and provided helpful discussions on the data of Raman spectroscopy. Y. G. assisted in Cryo-TEM measurements. W. Z. assisted in the molecular dynamics simulation. Y. L. provided important guidance for studying the structure of water ice. W. Z. and G. Z. are responsible for funding acquisitions. H. L.and G. Z. revised the manuscript. G. Z. supervised the whole work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hai-Tao Feng, Tong Zhu, and the other anonymous reviewer(s) for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, H., Su, H., Chen, N. et al. Confirmation-dependent organic phosphor reveals amino acid nanoaggregates in ice with insight for prebiotic chemistry. Nat Commun 16, 10873 (2025). https://doi.org/10.1038/s41467-025-65885-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65885-8