Abstract
The integration of organic synthesis with enzymatic catalysis offers a promising route toward efficient and sustainable construction of complex molecules. While organic synthesis enables diverse transformations, enzymatic catalysis enhances stereoselectivity under mild conditions, improving cost-effectiveness and environmental impact. However, current enzymatic synthesis planning algorithms face challenges in formulating robust hybrid organic–enzymatic strategies. Key issues include the difficulty in devising hybrid planning approaches and the reliance on template-based enzyme recommendations, which limits their adaptability across diverse scenarios. Here we show ChemEnzyRetroPlanner, an open-source hybrid synthesis planning platform that combines organic and enzymatic strategies with AI-driven decision-making. The platform features advanced computational modules, including hybrid retrosynthesis planning, reaction condition prediction, plausibility evaluation, enzymatic reaction identification, enzyme recommendation, and in silico validation of enzyme active sites. A central innovation is the RetroRollout* search algorithm, which outperforms existing tools in planning synthesis routes for organic compounds and natural products across multiple datasets. ChemEnzyRetroPlanner provides an intuitive graphical interface and programmatic APIs for scalability, while leveraging the chain-of-thought strategy and the Llama3.1 model to autonomously activate hybrid synthesis strategies for diverse scenarios. The results indicate that this fully automated, open-source system holds potential value for improving the efficiency and sustainability of molecular synthesis.
Similar content being viewed by others
Introduction
Organic synthesis and enzymatic synthesis techniques offer complementary advantages, making their integration a powerful approach for planning the synthesis of complex compounds. Conventional organic synthesis provides a broad spectrum of chemical reactions with wide applicability, while enzymes, as highly efficient biocatalysts, excel in regulating stereochemistry during critical steps, thereby reducing the need for protection and deprotection strategies. Furthermore, enzymatic reactions typically operate under mild conditions, often in water or benign organic solvents, making them not only environmentally sustainable but also more cost-effective compared to organic catalytic reactions that often rely on precious metals. Recent advancements in directed evolution have significantly expanded the chemical space of enzymatic catalysis, enabling enzymes to act on unnatural substrates and meet the demands of increasingly complex synthesis processes. This progress has opened new opportunities for leveraging enzymes in hybrid synthesis strategies, bridging the gap between traditional organic synthesis and biocatalysis to address modern challenges in the design and production of complex molecules.
Most computer-aided synthesis planning algorithms emulate the chemist’s retrosynthetic analysis process, beginning with the target molecule and recursively identifying the most promising reaction precursors until reaching suitable simple molecular building blocks1,2,3,4,5. Data-driven organic synthesis planning systems typically consist of several key components: a single-step prediction model to generate search actions, a reaction plausibility evaluator, multi-step search strategies, a customizable library of molecular building blocks, a reaction condition recommender6,7,8, and a pathway ranking function9,10,11,12,13. While traditional single-step retrosynthesis models are often trained on datasets like USPTO14 or Reaxys15, which may include a limited number of enzymatic reactions, these models struggle to make accurate enzyme reaction predictions. This limitation underscores the need for specialized approaches to integrate enzymatic synthesis planning into existing frameworks effectively.
In recent years, researchers have developed fully enzymatic synthesis planning algorithms16,17, with single-step prediction models trained on specialized enzymatic reaction databases such as KEGG18, MetaNetX19, and Rhea20. While these models are capable of predicting enzymatic reactions, their coverage is limited to a relatively narrow synthesizable chemical space and lacks the breadth of synthetic strategies offered by models trained on conventional organic chemistry datasets. To overcome the limitations of existing algorithms in hybrid synthesis route planning, numerous researchers have begun exploring strategies that integrate organic and enzymatic reactions. For instance, Anand et al.21 employed a mixed-integer linear programming approach to identify optimal synthesis routes from starting materials to target products within a hybrid reaction network constructed from literature-reported organic and enzymatic reactions. While this method is well-suited for the controllable optimization of known partial routes, it lacks the capacity to generate novel synthetic strategies. Levin et al.22 proposed a collaborative prediction framework that jointly utilizes two template-based single-step retrosynthesis models—one trained on organic reactions and the other on enzymatic reactions—to co-design hybrid synthetic pathways. Similarly, Li et al.23 employed a pair of retrosynthesis predictors trained separately on organic and enzymatic data. However, unlike Levin’s integrated strategy, their approach does not invoke both models concurrently during the search process. Instead, a scoring model is first applied to evaluate the efficiency of a complete synthesis route generated by purely organic planning. The step with the lowest efficiency is then identified and replaced using an enzymatic synthesis planning tool. Kreutter et al.24 trained two distinct Triple Transformer Loop architectures based on the Transformer model. These independently handle single-step retrosynthesis, reaction condition recommendation, and forward reaction validation for organic reactions, as well as single-step enzymatic retrosynthesis, enzyme suggestion, and forward verification for enzymatic reactions. During synthesis route planning, the model combines a route penalty score25 with a heuristic best-first tree search to enable effective hybrid organic-enzymatic synthesis route generation. In a different approach, Sankaranarayanan et al.26 integrated a template-based enzymatic reaction identification module into an existing multi-step retrosynthesis framework, enabling the identification of enzyme-catalyzed steps after completing synthesis planning.
Following similar technical approaches, several computational platforms have recently been developed specifically for the planning of hybrid organic-enzymatic synthesis. For instance, RetroBioCat27 offers both interactive and fully automated planning strategies, utilizing template-based predictive algorithms to design synthetic pathways and employing expert-defined enzymatic reaction templates to identify potential enzymatic steps within these pathways. Unlike the Monte Carlo tree search (MCTS) method used in RetroBioCat, BIONAVI28 adopts a search algorithm based on Retro*29, integrating a Transformer-based hybrid organic-enzymatic single-step retrosynthesis prediction model to enhance the success rate of pathway planning. Although research has made advances in the field of hybrid organic-enzymatic synthesis planning, current strategies for identifying enzymatic reactions and recommending enzymes primarily rely on template matching and similarity-based ranking of reactions. This approach of strict pattern matching may severely limit the exploration of other potential hybrid organic-enzymatic synthesis strategies. Additionally, further mechanisms for reaction-enzyme matching validation, such as the identification of enzymatic activity and critical active sites, remain insufficiently addressed with the current methods. This limitation leaves significant room for error and highlights the need for more precise and reliable validation strategies.
Moreover, the level of automation in existing synthesis planning platforms is generally inadequate, requiring experts to invest significant effort at various stages of the planning process, including defining computational tasks and selecting appropriate tools. The automation level of hybrid organic-enzymatic synthesis planning platforms is particularly limited compared to established commercial synthesis planning platforms. Recently, with advancements in general large language models (LLMs) such as ChatGPT30 and Llama331, the development of LLM-driven specialized Agents has emerged. Previous studies have shown that LLMs can autonomously perform a wide range of research-related tasks through prompt engineering, significantly lowering the barrier of domain expertise and improving workflow efficiency32,33,34,35,36,37. In the domain of synthetic chemistry, LLM-driven domain-specific Agents have begun to emerge. For example, ChemCrow34 was the first to demonstrate how LLMs, combined with chain-of-thought reasoning strategies, can effectively orchestrate a variety of synthesis tools to address diverse and complex chemical challenges. Subsequent developments such as Coscientist35 and LLM-RDF36 have further expanded the applicability of LLMs by integrating web browsers, literature analysis modules, code interpreters, and automated synthesis platforms—enabling end-to-end autonomous chemical experiment design, execution, and optimization. In parallel, Ma et al.37 integrated LLMs with knowledge graph technologies to propose, for the first time, an automated framework for route planning and optimization in the field of polymer synthesis, thereby broadening the potential applications of LLMs in materials design. Although the capabilities of such chemistry tool-augmented LLMs in solving synthesis planning problems are limited by the tools they invoke, they still demonstrate the potential to autonomously configure different tools to maximize their utility. Specifically, In the realm of hybrid enzymatic-organic synthesis planning, the scarcity of enzymatic reaction data hinders the ability of single-step retrosynthesis models for enzymatic reactions to support search algorithms in exploring a wide chemical space effectively. At the same time, conventional organic single-step retrosynthesis models are ineffective at predicting enzymatic steps. Therefore, it is essential to explore various hybrid planning strategies, such as integrating single-step retrosynthesis prediction models or strategies for recommending reaction conditions and enzymes. The complex configuration and decision-making processes involved in these tasks can be entrusted to LLMs to improve planning efficiency and prediction accuracy. Developing agents to enhance the automated decision-making capabilities of hybrid enzymatic-organic synthesis planning platforms undoubtedly represents a highly promising direction for future research.
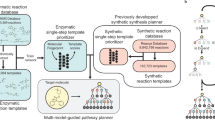
In this study, we developed a hybrid organic-enzymatic synthesis planning platform, ChemEnzyRetroPlanner. An overview of the platform is summarized in Fig. 1. The platform offers seven key computational modules: multi-step synthesis planner, single-step retrosynthesis predictor, reaction condition predictor, reaction rate predictor, enzymatic reaction identifier, enzyme recommender, and enzyme active site annotator (EasIFA38). A schematic illustration of the coordinated logic among modules during synthesis planning is provided in Supplementary Information Fig. S10. Each module can be independently invoked via programmatic API to accommodate automated synthesis planning needs under customized scenarios. Building on this, we utilized Llama3.1:70b in the chain-of-thought approach to leverage multiple tools, constructing an agent capable of executing a set of hybrid enzymatic-organic synthesis planning tasks. Additionally, we developed a user-friendly graphical interface based on Web services. Our contributions are summarized as follows:
-
1.
We developed a search framework, RetroRollout*, guided by an And-Or Tree and a pathway depth scoring function. RetroRollout* incorporates online simulation steps and a sibling node jump search strategy based on Retro*, improving synthesis planning speed and target molecule pathway resolution rate.
-
2.
We integrated various single-step strategies from diverse data sources, enabling the selection of appropriate hybrid organic-enzymatic synthesis strategies tailored to different planning scenarios.
-
3.
We developed a tool chain related to biocatalytic synthesis for pathway enzymatic reaction identification, enzyme recommendation, and in silico validation of enzyme active sites.
-
4.
Through the developed programmatic API, we constructed a fully open-source hybrid organic-enzymatic synthesis planning agent based on Llama3.1:70b. In addition to inheriting ChemCrow’s cheminformatics tools, safety tools, and general tools, our agent offers purely open-source alternatives for reaction tools and supports enzymatic reaction tools.
a The technical framework of ChemEnzyRetroPlanner and its three supported interaction modes: graphical interface, programmatic API, and agent. b The main algorithmic modules included in the platform. API application programming interface.
Results
Benchmark for multi-step planning
We evaluated the multi-step planning capabilities of the platform using test sets that cover three distinct multi-step synthesis scenarios, including two test sets derived from organic synthesis patents and one test set based on natural products. Specifically, these test sets include: the USPTO-multistep-190 test set, initially curated by Chen et al.29 during the evaluation of the Retro* algorithm and widely used in subsequent studies; random subsets of the standard evaluation datasets PaRoutes-Set-n1 and Set-n5 compiled by Genheden et al12. for assessing multi-step retrosynthesis algorithms (containing multi-step synthesis pathway data, single-step reaction training data used during evaluation, and building block molecular datasets); and the natural product test set used by Zeng et al.28 to evaluate the BioNavi platform, along with its accompanying building block molecular dataset.
Benchmark in USPTO-multistep-190 Test Set. In this section, we compared the impact of search frameworks, jump search strategy, types of guiding functions, the quality of buyable building block molecule datasets, and the performance of single-step retrosynthesis models on the search performance for synthetic pathways of target molecules. The chosen baseline algorithm is the original Retro*, and the test molecules were 190 representative complex compound molecules extracted from the USPTO by Chen et al.29. The evaluation results are shown in Table 1.
We denote the search framework without using a guiding function as “Search Frame Name-0”. When limiting the search iterations to 100, using GraphFP as the single-step retrosynthesis policy, and employing the modified Zinc buyable building block molecule dataset, our proposed RetroRollout* algorithm framework successfully resolved a synthesis plan for more molecules than Retro*, regardless of the use of guiding functions. The GraphFP model is a template-based, single-step retrosynthesis prediction model trained on the USPTO-all-remapped dataset (see Supplementary Information S1.1 for details). Comprehensive details about the model architecture, training procedure, and the buyable building block datasets are available in Supplementary Information Section S1.5. Compared to Retro-0, RetroRollout*−0 solved an additional 11 molecules, increasing the resolution rate by 5.79%; RetroRollout solved 12 more molecules than Retro, improving the resolution rate by 6.32%. Comparing the effectiveness of algorithms with and without depth-based guiding functions, it is evident that the guiding functions effectively increased the number of solved target molecules. Specifically, Retro* solved 3 more molecules than Retro*−0, and RetroRollout* solved 4 more molecules than RetroRollout*−0. After increasing the maximum search rounds to 500, the number of molecules solved by RetroRollout* and RetroRollout*−0 further increased, solving 7 and 5 more molecules, respectively, compared to their corresponding plannings at 100 search rounds. More specifically, the 500-round search RetroRollout* resolving 88.42% of the target molecules’ synthetic pathways. We also compared the original Retro* model (using Neuralsym as the single-step policy network, the original configuration of the guiding function model, and the buyable building block molecule dataset) and found that, at the same number of rounds, RetroRollout* solved more molecules than the original Retro* proposed by Chen et al.29, and relied less on guiding functions. RetroRollout*−0, even without using guiding functions, solved 0.53% more molecules than Retro* using a reaction cost guiding function.
In addition, we conducted an ablation study on the jump search strategy to evaluate its specific impact on search performance. In this experiment, we introduced two variants without jump search, named RetroRollout*−0 (W/O JS) and RetroRollout* (W/O JS), corresponding to search frameworks without a guiding function and with a depth-based guiding function, respectively. The results show that, compared to the original RetroRollout*−0 and RetroRollout* with jump search, these variants solved 4.7% and 4.2% fewer target molecules, respectively, demonstrating the effectiveness of the jump search strategy in improving both search efficiency and success rate. We also compared the Retro*−0 and Retro* methods. Under the same one-step prediction model and guiding function settings, our proposed search framework, even without the jump search strategy, still solved 1.1% and 2.1% more target molecules than the original methods, further indicating the intrinsic strength of our search framework. Moreover, we adopted a continuous search strategy to compare the path performance of Retro* and RetroRollout* on the same dataset (shown in Table S14). With an equal number of search iterations (100 rounds), RetroRollout* not only solved more target molecules but also provided an average of 57.8 additional successful pathways per target molecule. Among these, 58% were shorter than the reference (ground-truth) pathways, compared to 51% for Retro*. Notably, even under a limited budget of 100 search iterations, RetroRollout* achieves a higher target molecule resolution rate than Retro* with 350 iterations, outperforming it by 3.16%, while also reducing the average search time by 7.54 s. Although RetroRollout* generates fewer successful routes per target molecule (170.2) compared to Retro* (409.5) under this setting, it still effectively identifies valid synthetic solutions for structurally complex and hard-to-solve targets, thereby maintaining robust overall planning capability. This suggests that the search behavior of RetroRollout* tends to favor deeper exploration, which facilitates earlier discovery of complex pathways without compromising computational efficiency. Collectively, these results highlight the superior balance of RetroRollout* in terms of route quality, search efficiency, and its ability to address challenging synthetic targets, relative to Retro*.
Benchmark in PaRoutes Random Selected Test Set. In this section, we employ a standardized multi-step retrosynthesis evaluation method to thoroughly assess the performance of multi-step retrosynthesis algorithm frameworks. We utilized two subsets of reaction paths extracted from the USPTO reaction dataset, PaRoutes Set-n1 and PaRoutes Set-n512 to train the single-step retrosynthesis prediction model GraphFP, respectively. Details regarding the training of this model on PaRoutes Set-n1 and Set-n5 are provided in Supplementary Information S1.1. We used the leaf node molecules from the paths in these two datasets as the buyable building block molecule dataset and trained depth-based guidance functions separately for Set-n1 and Set-n5. The distribution of Set-n1 is closer to the overall distribution of the USPTO reaction dataset, while Set-n5 contains more samples with longer synthesis paths. The evaluation criteria include not only the number of solved target molecules but also the average time for path search, the top-k accuracy of complete synthesis route hits, and two metrics reflecting the quality of the route: the proportion of shorter synthesized paths planned and the average overlap of leaf node molecules. The evaluation results are presented in Table 2. The scoring and ranking of the routes are conducted according to the method reported by Badowski et al.
When the search iterations were restricted to 250, we observed that in Set-n1, RetroRollout* solved 483 out of 500 target molecules, while Retro* only solved 473. The two algorithm frameworks were comparable in the top-1 accuracy, but RetroRollout* achieved higher top-5 and top-10 accuracies, with its planned paths being shorter on average and having a higher overlap with the standard paths at the leaf nodes. In Set-n5, RetroRollout* solved the paths of 490 molecules, whereas Retro* solved 477, with Retro* slightly surpassing our proposed RetroRollout* algorithm in the top-1 accuracy but RetroRollout* leading in top-10 accuracy. Although the two algorithms had their strengths in top-k accuracy for the Set-n5 dataset, the routes quality planned by RetroRollout* was higher, with shorter routes and greater overlap with the standard route leaf nodes. To further objectively evaluate their performance, we increased the search iterations for Retro* to 750, at which point Retro*‘s time spent on searching for synthesis routes for each molecule was significantly more than RetroRollout*‘s 250-iteration search (averaging about 40 to 50 s more per target molecule), yet the number of molecules solved remained lower than RetroRollout*. While the routes’ top-k accuracy and quality slightly surpassed those of RetroRollout*, the differences were not significant. Overall, under the two PaRoutes standardized evaluation methods, RetroRollout* was able to solve more molecular synthesis path problems in a shorter amount of time while maintaining comparable or slightly higher path planning quality than Retro*.
Benchmark in Nature Products Dataset. In this section, we evaluated the ChemEnzyRetroPlanner platform’s ability to resolve molecular pathways using the natural product dataset employed by Zeng et al. and compared its performance with other reported platforms. The experimental setup was identical to that used by Zeng et al.39, utilizing the same buyable building block dataset. ChemEnzyRetroPlanner tested the effectiveness of various single-step strategy models in pathway searches. These single-step models are labeled in the study by the training datasets name they were derived from and include four template-based single-step retrosynthesis prediction algorithms from ASCKOS: Pistachio ringbreaker, BKMS metabolic, Reaxys, and Reaxys biocatalysis. Among them, BKMS metabolic and Reaxys biocatalysis are single-step retrosynthesis prediction models trained on enzymatic reaction datasets. Additionally, we tested the benchmark model trained on the USPTO-all-remapped dataset (GraphFP, same as the benchmark in USPTO-multistep-190 test set) and a Transformer-based single-step retrosynthesis prediction model trained on a mixed dataset of enzymatic and organic reactions, as reported by Zheng et al.39. The test results for this section are presented in Table 3.
The results demonstrate that the ChemEnzyRetroPlanner platform, when combined with RetroRollout* and other mainstream open-source single-step retrosynthesis prediction models, significantly enhances pathway resolution success rates. Specifically, strategies incorporating two different template-based single-step retrosynthesis prediction models (trained on the Reaxys and BKMS metabolic datasets, respectively) outperformed benchmark models such as RetroPathRL40, RetroBioCat27, ASKCOS41, and original Retro*, achieving a pathway resolution success rate of 64.4%, which is notably higher than the best-performing template-based benchmark method, RetroPathRL (59.8%). Furthermore, the RetroRollout* search strategy, when integrated with a template-free method identical to that used in BioNavi-NP, also achieved a higher pathway resolution success rate. The synthesis pathway resolution success rate for the target molecules reached 98.4%, slightly exceeding that of the BioNavi method trained on broader datasets.
Benchmark for enzymatic reaction identification model and enzyme recommender
In this section, we systematically evaluated the performance of the enzymatic reaction identification model and the enzyme recommender embedded within the ChemEnzyRetroPlanner platform and compared them against current state-of-the-art baseline methods.
For the enzymatic reaction identification model, we selected a widely adopted baseline approach based on reaction template matching. This method leverages 135 enzymatic reaction templates released by the RetroBioCat platform and determines whether a given reaction step is enzymatic by checking for a match with any of the predefined templates. We evaluated the performance of the built-in model and this classical method on a test set composed of a mixed dataset: USPTO-1000-TPL (as negative samples) and ECReact (as positive samples). Details of the dataset partitioning are provided in Supplementary Information S1.3, and the evaluation results are summarized in Table 4. The results show that the built-in RXNFP-based enzymatic reaction identification model performs effectively well on the binary classification task of distinguishing enzymatic from non-enzymatic steps. All evaluation metrics exceeded 0.9900, with the exception of recall (0.9895), and the model achieved an inference speed approximately five times faster than the template-matching baseline. In contrast, the classification performance of the template-match-based method was considerably lower, primarily due to its heavy reliance on the discriminatory power of hand-curated reaction rules. As indicated in Table 4, its predictive capability does not meet the high-accuracy demands of the platform. By framing enzymatic reaction identification as a binary classification problem and leveraging learnable reaction fingerprints with attention mechanisms, our model achieves robust and scalable performance, laying a solid foundation for large-scale enzymatic reaction screening within synthesis planning.
For the enzyme recommender, the core task is the classification of enzymatic reactions into their corresponding EC numbers. We compared our method against two representative mainstream approaches: CLAIRE and Selenzyme 2023. CLAIRE is a contrastive learning-based method that encodes enzymatic reactions using a combination of RXNFP and DRFP fingerprints. It employs the triplet margin loss to minimize the embedding distance between samples sharing the same EC number and maximize the distance between samples with different EC numbers. The original implementation of CLAIRE supports prediction up to the third level of EC numbers (EC-L3). In light of the requirement for fine-grained EC number recommendation in enzyme prediction tasks, we modified the CLAIRE algorithm to enable prediction of the complete four-level EC numbers (EC-L4), while preserving its capability to evaluate performance at the EC-L3 level. To ensure a fair comparison, we retrained CLAIRE on the same dataset split used for the enzyme recommender. Selenzyme 2023 is a similarity-based enzyme recommendation tool that identifies the most similar enzymatic reactions from an internal database and returns the associated enzymes as recommendations. We deployed Selenzyme 2023 locally to assess its real-world inference performance, including both predictive accuracy and computational efficiency. Its performance was evaluated at both EC-L3 and EC-L4 levels. The comparative results of all three methods are summarized in Table 5. On the EC-L3 prediction task, the enzyme recommender embedded in ChemEnzyRetroPlanner performed comparably to CLAIRE, achieving Top-1 accuracies of 83.55% and 82.57%, and Top-3 accuracies of 88.81% and 91.95%, respectively. However, in the more challenging EC-L4 prediction, our enzyme recommender achieved a Top-1 accuracy of 65.31%, significantly outperforming CLAIRE’s 52.96%. This performance gap is mainly attributed to the difficulty CLAIRE faces in contrastive learning when insufficient positive samples exist at finer EC number granularity. Selenzyme 2023 achieved a Top-1 accuracy of 72.43% at the EC-L3 level, but its performance dropped substantially at EC-L4, with a Top-1 accuracy of only 26.49%. In terms of inference speed, the enzyme recommender in ChemEnzyRetroPlanner is approximately 10 times faster than CLAIRE and about 4000 times faster than Selenzyme 2023, demonstrating high computational efficiency. This makes it particularly well-suited for integration into synthesis planning workflows, where rapid and frequent predictions over large numbers of reaction nodes are required.
Benchmark for reaction plausibility evaluator
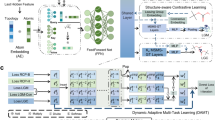
We first constructed a benchmark dataset (referred to as the template-shuffling dataset) for evaluating the baseline performance of reaction plausibility evaluators, using reaction data from the USPTO. Negative samples were generated using a template-shuffling strategy (A schematic illustration of the data generation process is shown in Fig. S5). The resulting dataset contains a total of 1,842,509 reaction samples, with positive samples entirely derived from USPTO data (accounting for 70%) and negative samples comprising the remaining 30%. The dataset was randomly split into training, validation, and test sets at a ratio of 8:1:1, and was used to identify the optimal reaction representation and model architecture. We evaluated three types of reaction representations: Morgan fingerprints, RXNFP, and DRFP. Among them, the model based on Morgan fingerprints and a dual-branch multilayer perceptron architecture (see Fig. S6 for model structure, referred to as RXN Filter-benchmark) achieved the best overall performance. On the test set of the benchmark dataset, this model achieved an AUC of 0.9583, precision of 0.9155, recall of 0.9160, and f1 score of 0.9145, slightly outperforming the models based on RXNFP and DRFP.
After determining the optimal model structure and reaction representation, we proceeded to train the model on a larger-scale reaction plausibility prediction dataset (referred to as the Faiss-Template sampling dataset), which was constructed using a combination of Faiss-based similarity sampling and a reaction template selection model. This dataset contains 2,745,104 samples and was randomly split into training, validation, and test sets at a ratio of 9:0.5:0.5, with a balanced 1:1 ratio of positive to negative samples. On the internal test set of this dataset, the model RXN Filter-deployed achieved an AUC of 0.9093, precision of 0.9331, recall of 0.8330, and f1 score of 0.8329, demonstrating strong and balanced classification performance.
To further compare the models’ ability to distinguish between different types of reaction samples, we extracted 44,280 samples from the test set of the Faiss-Template sampling dataset to construct a common test set with no overlapping with the benchmark dataset. This test set contains three types of samples: true reactions, plausible positive reactions generated with high confidence by the template selection neural network, and implausible negative reactions that follow a reaction template pattern but are chemically unrealistic (low confidence by the template selection neural network). The sample ratio was 4:1:5. We compared the performance of RXN Filter-benchmark, RXN Filter-deployed, and the built-in fast filter in the ASKCOSv2 system. The results are summarized in Table S13. The RXN Filter-deployed model demonstrated the most balanced performance on this test set, with an AUC of 0.9000, accuracy of 0.8209, precision of 0.8211, recall of 0.8212, and f1 score of 0.8210. In contrast, the RXN Filter-benchmark model, trained only on the template-shuffling data, exhibited a much higher false-positive rate: although it achieved a recall of 0.9441, its accuracy dropped to 0.6420 and precision to 0.5885. Similarly, the ASKCOSv2 fast filter displayed comparable characteristics, with an accuracy of 0.6487, precision of 0.6008, and recall of 0.8865, indicating limited effectiveness in identifying implausible reactions. In summary, RXN Filter-deployed provides more accurate and robust classification, making it a more suitable filtering module for detecting chemically implausible reactions in practical synthesis planning workflows.
Web server interface
For input and options, the platform supports directly pasting SMILES, as well as clicking the “Draw” button to sketch a molecule in the JSME container. In this study, the molecule CCCC( = O)O is used as an input example, as shown in Fig. 2a. Users can click the “Options” button to select several parameters for the hybrid synthesis planning platform to run. These include “Keep search after solved one route”, “Use reaction plausibility evaluator”, “Use guiding function”, “Predict reaction conditions”, “Identify enzymatic reactions”, and “Recommend enzymes”. Users can select from four predefined purchasable building block datasets provided by the platform. Multiple datasets can be chosen simultaneously. Additionally, the platform offers seven optional single-step retrosynthetic prediction models, which include three trained on conventional organic reaction datasets—GraphFP (USPTO-all-remapped), Pistachio Ringbreaker, and Reaxys—as well as three models trained on enzymatic reaction datasets—Transformer (USPTO-NPL + BioChem), BKMS Metabolic, and Reaxys Biocatalysis (these models are named based on the datasets used for training, employing MLP and reaction template-based single-step algorithms). Details of the models are provided in Supplementary Information S1.1. In the single-step retrosynthetic model selection box, multiple models can be selected simultaneously for collaborative search. Combining enzymatic reaction single-step retrosynthesis models (e.g., BKMS Metabolic) with organic reaction single-step retrosynthesis models (e.g., Reaxys) facilitates the planning of hybrid synthetic pathways. Additionally, users can set the number of iterations for the search algorithm as needed. After configuring all parameters, clicking the “Submit” button initiates the synthetic route search for the input target molecule. All algorithm outputs are displayed in the “Log” box, and the platform returns a key for viewing the results. Once the search is complete, the “View Results” button becomes active. Clicking this button redirects users to the results interface, where they can view the computational results by uploading the corresponding key.
a Parameter selection interface, and log output interface; b The synthesis pathway and enzyme active site display interface of the platform.
For results display, after the user provides the corresponding key file, the interface will jump to the results page, as shown in the upper part of Fig. 2b. The results page contains three main sections: a dropdown menu to switch between multiple synthesis routes, an interactive synthesis route display diagram, and a single-step reaction display box. For each displayed synthesis route, users can click on reaction nodes (enzymatic reactions are shown in green, conventional organic reactions in yellow) to view detailed information about that step. This information is displayed in the single-step reaction display box, which sequentially shows the single-step reaction equation, the platform-recommended reaction conditions, the identification results for enzymatic reactions, and the enzyme recommendation results. Once a single-step reaction is identified as enzymatic, the “Enzyme Recommend” button becomes active. Clicking this button, the platform will call the UniProt database API to find enzymes under the recommended EC number and use the EasIFA algorithm to predict active sites. At this point, the enzyme structure display column will pop up, allowing interactive viewing of the enzyme’s structure. Detailed active site information is also displayed in a table below the structure.
Synthesis planning case study
To further evaluate the utility of ChemEnzyRetroPlanner, we applied it to six pharmaceutically relevant or bioactive compounds for retrosynthetic route planning. The algorithm employs a template-based, interpretable single-step retrosynthesis predictor integrating Reaxys and BKMS metabolic reaction knowledge bases. The selected starting material library consists of the building blocks used in Repaired Zinc and BioNav28, with an additional constraint limiting the maximum number of carbon atoms in each compound to eight. The number of search iterations was set to 10. All proposed synthetic routes successfully matched literature-reported syntheses. Among them, the predicted routes for Esmolol, Desmethylxanthohumol, and 3-(Benzenesulfonyl)−6-methyl-4-(4-methylpiperidin-1-yl)quinoline reflected hybrid enzymatic-organic synthetic strategies (Supplementary Information Fig. S12), whereas those for Celiprolol, S-(-)-Warfarin, and AZD7545 corresponded to fully organic pathways (Supplementary Information Fig. S13).
In the case of Esmolol, the synthesis commenced from commercially available starting materials including p-iodophenol, methyl acrylate, allyl bromide, and isopropylamine (highlighted by dashed boxes in Fig. S12). The strategy began with esterification and nucleophilic substitution to construct a key intermediate featuring an aromatic core and an allyl side chain. This was followed by regioselective epoxidation of the terminal double bond, catalyzed by a monooxygenase (UniProt ID: A0A1D6GQ67; EC 1.14.14.159), to generate epoxide intermediate. A final ring-opening reaction with isopropylamine afforded the β-blocker scaffold of Esmolol. For Desmethylxanthohumol, three commercially available building blocks were utilized: trihydroxyaryl ketone, phenylpropene ether, and dimethylallyl diphosphate (DMAPP) (dashed boxes in the figure). A literature-supported aldol condensation assembled the chromanone core intermediate. Subsequently, a nucleophilic substitution between the chromanone and DMAPP yielded the prenylated natural product derivative Desmethylxanthohumol. This step was plausibly catalyzed by a prenyltransferase (UniProt ID: A0A0B5A051; EC 2.5.1.136), as proposed by ChemEnzyRetroPlanner, consistent with known enzymatic specificity. In the third example, targeting 3-(Benzenesulfonyl)−6-methyl-4-(4-methylpiperidin-1-yl)quinoline, the synthetic route initiated from commercially available compounds, including benzenesulfonitrile, methanol, triethyl orthoformate, p-toluidine, and p-methylpiperidine (dashed boxes). Benzenesulfonitrile was first hydrolyzed by nitrilase (UniProt ID: Q42966; EC 3.5.5.1) to yield the corresponding acid, followed by esterification with methanol to generate methyl benzenesulfonylacetate. In parallel, p-toluidine underwent condensation and cyclization with an electrophilic ketone intermediate to form the quinolinone core. Subsequent dehydrative chlorination using phosphoryl chloride (POCl₃) yielded a chloroquinolinone intermediate, which was finally subjected to nucleophilic substitution with p-methylpiperidine to introduce the key piperidine side chain, completing the synthesis.
In the retrosynthetic planning of the antihypertensive drug Celiprolol, ChemEnzyRetroPlanner identified a feasible route starting from commercially available 1-(2-hydroxy-5-nitrophenyl)ethanone, tert-butylamine, and diethylcarbamoyl chloride (highlighted in dashed boxes in Fig. S13). The synthesis begins with epoxidation of the phenolic hydroxyl group in 1-(2-hydroxy-5-nitrophenyl)ethanone, as reported by Sayyed et al.42, introducing an epoxide moiety. The resulting intermediate then undergoes a nucleophilic ring-opening reaction with tert-butylamine to form a tertiary butyl amide-containing intermediate. Subsequent reduction of the nitro group yields the corresponding aniline derivative, which reacts with N-ethyl-N-chloroacetamide in an amidation step to install the urea-containing side chain, thus completing the synthesis of Celiprolol. In the fifth case study, ChemEnzyRetroPlanner was applied to the synthesis of the anticoagulant drug S-(-)-warfarin, using commercially available o-hydroxyacetophenone, diethyl carbonate, benzaldehyde, and acetone as starting materials. The route initiates with a cyclization-condensation reaction between o-hydroxyacetophenone and diethyl carbonate to form 4-hydroxy[1]benzopyran-2-one. In parallel, benzaldehyde and acetone undergo an aldol condensation to afford 1-phenylbut-1-en-3-one. These two intermediates are subsequently coupled through a Michael addition, effectively yielding the final product, S-(-)-warfarin. In the sixth case, ChemEnzyRetroPlanner successfully reproduced the full synthetic route of AZD7545 as reported by Patel et al.43. The synthesis begins with three commercially available building blocks—2-chloroaniline, dimethylamine, and (R)−3,3,3-trifluoro-2-hydroxy-2-methylpropionic acid—and proceeds through a four-step sequence to afford AZD7545, fully consistent with the literature-reported synthesis.
Collectively, these six representative examples not only demonstrate the broad applicability of ChemEnzyRetroPlanner in retrosynthetic planning for complex bioactive compounds but also highlight its dual capability in designing both hybrid enzymatic–organic routes and conventional organic syntheses with high accuracy and practicality.
Agent case study
ChemEnzyRetroPlanner includes an agent APP based on the programmatic API, which, in addition to performing the synthesis planning tasks supported by ChemCrow with fully offline deployment, can also handle enzymatic step planning and validation tasks within reaction steps. Figure 3a illustrates the workflow in which the agent autonomously evaluates the feasibility of a single-step reaction, determines whether it is an enzymatic reaction, and recommends enzymes. The agent first uses the ReactionRater tool to assess whether the synthesis reaction of 4-(hydroxymethyl)−2-furancarboxaldehyde phosphate (4-HFC-P) is feasible. Once the reaction is confirmed to be feasible, the EnzymaticRXNIdentifier is used to determine whether this reaction step could be catalyzed by an enzyme (i.e., whether it is an enzymatic reaction). The tool identifies with a confidence of 0.9975 that this single-step reaction is an enzymatic reaction. The agent then invokes EnzymeRecommender, which recommends five enzyme categories most likely to catalyze this reaction, with the most likely enzyme having an EC number of 4.2.3.153. All the reasoning results are accurate. Figure 3b further demonstrates the application of the EasIFA module in recommending the most likely enzyme entity and the use of active site prediction to check the presence of active sites in the recommended enzyme for the given enzymatic reaction. The results show that the EasIFA module recommends the enzyme with UniProt ID Q58499, (5-formylfuran-3-yl)methyl phosphate synthase, which harbors two active sites, LYS27 and LYS85, corresponding to the reaction site information recorded in UniProt.
a Inference cases for reaction feasibility judgment, enzymatic reaction identification, and enzyme recommendation. The reaction case presented is the synthesis of 4-(hydroxymethyl)−2-furancarboxaldehyde phosphate (4-HFC-P). b Individual enzyme recommendations for the synthesis of 4-HFC-P, following the predicted most probable EC Number: 4.2.3.153 from the case in panel a. The recommendation includes the verification of the active site.
Additionally, we also present the reasoning results of the ChemEnzyRetroPlanner agent for multi-step synthesis planning tasks in Supplementary Information Fig. S14. Similar to ChemCrow, the Agent can autonomously query the SMILES information of compounds, perform safety checks, and finally use the programmatic API of the ChemEnzyRetroPlanner synthesis planning tool to generate synthetic routes and summarize the relevant synthetic steps. ChemEnzyRetroPlanner and its agent platform support fully offline deployment and have the potential to enable a fully autonomous hybrid enzymatic-organic synthesis planning automation platform.
Moreover, in more complex synthesis planning tasks, the ChemEnzyRetroPlanner Agent demonstrates strong automated decision-making capabilities. This system can autonomously determine and optimize hybrid configuration strategies for single-step retrosynthesis prediction models, enabling more efficient planning of synthesis pathways for molecules residing in various chemical spaces. For instance, Fig. 4 illustrates the Agent’s analysis of the synthesis pathway for the natural product phenoxy radical VII (an intermediate in the biosynthesis of hispidol from isoliquiritigenin in soybean). The Agent first evaluates whether the compound is a controlled chemical. After confirming its low similarity to controlled chemicals, it invokes the RetroPlannerRetrosynthesis tool for pathway planning. Initially, the Agent employed a purely organic synthesis planning strategy but failed to generate a pathway for this natural product. Subsequently, the Agent autonomously determined to switch to a hybrid synthesis planning strategy, combining organic synthesis and enzymatic synthesis approaches. This successfully generated 131 candidate synthesis pathways, among which the optimal pathway precisely matched the actual biosynthetic pathway of the natural product phenoxy radical VII. Additionally, the Agent comprehensively summarized the fundamental reaction conditions required for each step of this optimal pathway and appropriately recommended the types of enzymes to prioritize if an enzymatic synthesis strategy is adopted. In the Supplementary Information Fig. S15, we further present the Agent’s analysis of the synthesis pathway for another natural product, along with the synthesis pathway diagrams for the two natural products shown in Fig. S16. These cases not only highlight the potential advantages of hybrid synthesis planning strategies over purely organic synthesis strategies in resolving synthesis pathway challenges but also demonstrate the Agent’s robust automation capabilities in making complex configuration decisions.
The Agent first invokes the CotrolChemCheck tool to determine whether the input molecule is a regulated chemical. It then invokes the RetroPlannerRetrosynthesis tool for retrosynthetic analysis. When the pure organic synthesis planning configuration fails to yield results, the Agent automatically adopts a mixed synthesis strategy, ultimately successfully obtaining a synthesis planning outcome, which corresponds to an actual synthetic case.
Discussion
In this study, we developed a hybrid enzymatic-organic synthesis planning platform, ChemEnzyRetroPlanner, which integrates a comprehensive set of organic synthesis planning tools along with computational and validation tools for enzymatic synthesis planning. It enables joint search functionality between the organic single-step retrosynthetic prediction model and the enzymatic reaction single-step retrosynthetic prediction model. ChemEnzyRetroPlanner demonstrates significant advantages over comparable algorithmic platforms across multiple test sets related to organic synthesis and natural products. Additionally, we developed an automated configuration process and built a user-friendly graphical interface based on a web server to enhance the user experience. Furthermore, we have developed the programmatic API for platform tools and a fully open-source agent application, offering strong potential for autonomous laboratory deployment and scalability. In summary, ChemEnzyRetroPlanner can plan hybrid synthetic routes that combine organic reactions and enzymatic reactions, supporting reaction condition prediction, enzyme category recommendations, and enzymatic active site annotation for synthesis pathway planning. This open-source platform is designed to assist pharmaceutical researchers in identifying more rational, cost-effective, and environmentally sustainable synthetic routes, while also providing essential active site validation information for the application of enzymatic reactions, thereby offering synthesis solutions with a sound foundation.
Methods
Implementation of RetroRollout* multi-step search framework
We optimized the search framework based on the AND-OR tree and the score updating strategy of Retro*, aiming to alleviate Retro*‘s heavy reliance on value function estimation to identify the most promising molecular nodes. The algorithmic framework is illustrated in Fig. 5a. We introduced a MCTS-like Rollout step to enhance the algorithm’s adaptability for the synthesis route planning. The modified search process now consists of four steps: Selection, Expansion, Rollout, and Update. The “Selection” and “Update” steps remain consistent with the Retro* algorithm, while a jump search strategy targeting the sibling nodes of successful nodes is introduced during the Rollout step to enhance search efficiency. Specifically, when one of a pair of sibling nodes is confirmed to be successful (i.e., purchasable or synthesizable), that node is prioritized for expansion by the algorithm. Additionally, the guiding function in these two phases is replaced with a synthesis route depth-based guiding function, which estimates the number of steps from the molecular node to the purchasable dataset. This guiding function is implemented using a multilayer perceptron classification model based on molecular fingerprints, with details of the model architecture and training dataset provided in the Supplementary Information S1.2. The pseudocode of the RetroRollout* algorithm has been provided in Supplementary Information S1.8.
a Schematic illustration of algorithmic improvements in RetroRollout*. Key steps include Selection, Expansion, Rollout, and Update, with a schematic illustration of the jump search process during Rollout; the algorithm pseudocode is provided in Supplementary Information S1.8. Here, \(m\) denotes a molecular node, \({V}_{{all}}\) represents the global cost list, and \({V}_{{sub}}\) represents the cumulative subtree cost of all child nodes corresponding to the molecular node \(m\). b Toolsets implemented in the ChemEnzyRetroPlanner agent. These toolsets comprise molecular tools, reaction tools, safety assessment tools, general tools, and enzymatic reaction tools, collectively enabling flexible and extensible agent behavior. RXN: reaction.
Implementation of the hybrid single-step retrosynthesis algorithm
The platform integrates single-step retrosynthesis prediction models trained on six distinct dataset distributions, including five template-based prediction algorithms and one transformer-based single-step retrosynthesis prediction algorithm. Among the five template-based algorithms, two are trained on datasets specific to enzymatic reactions, while the Transformer-based algorithm is trained on a mixed dataset of organic and enzymatic reactions. For details regarding the algorithms and their names as displayed on the platform, please refer to the Supplementary Information S1.1. The platform supports the simultaneous use of multiple single-step algorithms in multi-step search strategies, with the prediction results unified and ranked based on normalization of Softmax probability scores.
Implementation of the reaction condition recommendation module
We embedded two reaction condition recommendation models in ChemEnzyRetroPlanner, namely the reaction condition recommender (RCR)7 trained on Reaxys and Parrot6 trained on USPTO-Condition. We performed adaptations to embed the models within the synthesis planning framework.
For the embedding of the two reaction condition recommendation models, we adopted a post-planning offline inference method. Specifically, for the results obtained from the synthesis planning steps, we traverse the entire reaction tree, collecting each single-step reaction and converting them into reaction SMILES44. Then, we call the API of the reaction condition recommendation model to predict each single-step reaction, and attach the prediction results as reaction attributes back into the original reaction tree data structure.
Implementation of the enzymatic reaction identification model and enzyme recommender
We developed an enzymatic reaction identification model and an enzyme recommender to identify steps in the reaction tree that can be catalyzed by enzymes and to recommend the EC Number of enzymes for enzymatic reactions, respectively. We utilized the BERT-based learnable fingerprint RXNFP45 to represent reactions, and then employed multilayer perceptron to accomplish the identification of enzyme reactions (a binary classification task to predict whether it is an enzymatic reaction) and the recommendation of enzyme types (a multi-class classification task to predict the enzyme’s EC Number). For the enzymatic reaction identification model, we used a combined dataset consisting of USPTO-1000-TPL45 and ECReact17, where reactions from USPTO-1000-TPL serve as negative samples and reactions from ECReact are treated as positive samples. For the enzyme recommender, we used ECReact as an EC Number multi-class classification dataset. The details of the model architecture, datasets used, and implementation of the enzymatic reaction identification model and the enzyme recommender can be found in Supplementary Information S1.3.
For the enzymatic reaction identification model, we conducted a comparative evaluation using a reaction template-matching-based method under the same dataset split. This baseline method, similar to the approach proposed by Sankaranarayanan et al.26, utilizes the 135 enzymatic reaction templates provided by the RetroBioCat27 platform as identification rules. By matching reaction templates, the method determines whether a target reaction step qualifies as an enzymatic reaction. If the step conforms to one of the reaction templates, it is classified as an enzymatic step; otherwise, it is considered a conventional organic reaction.
For the enzyme recommender, we selected two mainstream methods as baselines for comparison: CLAIRE46 and Selenzyme 202347,48. CLAIRE46 is a state-of-the-art enzymatic reaction classification model based on contrastive learning and deep learning strategies. While the original version supports prediction of the first three levels of EC numbers, we modified it to enable prediction of the complete four-level EC numbers. Selenzyme47, on the other hand, is a widely used similarity-based enzyme recommendation model. Its upgraded version, Selenzyme 202348,49, has been integrated into the hybrid chemoenzymatic synthesis planning platform BioNavi28 as default enzyme recommendation module. We evaluated the enzyme recommendation performance of all three methods on the same test set, analyzing their accuracy in predicting both the first three levels and the full four-level EC numbers to comprehensively assess their recommendation capabilities. For implementation details of the baseline methods discussed in this section, please refer to Supplementary Information S1.3.
Integration of enzyme activity site annotation model
To facilitate the selection of suitable enzymes for catalyzing specific reactions and in designing directed evolution, we have integrated the enzyme active site annotation tool EasIFA38 into the ChemEnzyRetroPlanner platform. When the enzymatic reaction identification model classifies a single-step reaction as enzymatic, the platform subsequently activates the enzyme recommendation model to predict the most relevant EC numbers for the reaction. Among these recommended enzyme categories, the enzymatic reaction similarities associated with enzyme entries under the corresponding EC numbers in the UniProt database are calculated and compared. The enzymatic reaction most similar to the input reaction is identified, and its corresponding enzyme is retrieved for recommendation. Furthermore, using the EasIFA model, the active sites of these recommended enzymes most relevant to the reaction are identified as preliminary verification information for enzymatic activity. EasIFA is provided with the SMILES sequence of the single-step enzymatic reaction and the PDB structure data of the most relevant enzyme inferred through AlphaFold, subsequently annotating the potential active sites of the recommended enzymes. This active site information serves as an important reference for the final selection of enzymes and their directed evolution design. In the implementation, we used UniProt’s programmatic access API to query the enzyme sequence and structure information under a specified EC Number and employed PDB structure data stored in the AlphaFold Database50 to reduce the computational load of ab initio inference calculation for annotating active sites of enzymes without PDB structures. We utilized Py3Dmol for visualizing the enzyme structure and its active sites.
Implementation of reaction plausibility evaluator
To suppress the implausible predictions that may occasionally arise from the single-step retrosynthesis model, we implemented a reaction plausibility evaluator. We first constructed a benchmark dataset for evaluating the reaction plausibility evaluator based on the US patents dataset (1976–2016). The dataset was cleaned and augmented with atom mappings and reaction template annotations. In this benchmark dataset, negative samples account for 30% of the total. The strategy for constructing positive and negative samples is detailed in Supplementary Information Section S1.4. The dataset was split into training, validation, and test sets at a ratio of 8:1:1.
Using this benchmark dataset, we evaluated three types of reaction representations and their corresponding deep learning architectures: models based on Morgan fingerprints, RXNFP45, and DRFP51. The model architectures are illustrated in Figs. S6 and S7. Among them, the model based on Morgan fingerprints (hereafter referred to as RXN Filter-benchmark) employs a dual-branch design that processes product fingerprints and reaction difference fingerprints separately. This model outperformed the others across multiple evaluation metrics. Detailed benchmark results are provided in Table S10.
After identifying the optimal reaction representation and model architecture, we constructed a larger-scale dataset based on USPTO reaction data to better reflect real-world scenarios encountered in synthesis planning. This dataset was generated by combining similarity-based sampling with a deep learning–based reaction template prediction strategy, aiming to enrich the model’s ability to distinguish samples near the decision boundary. Specifically, we used the Faiss algorithm52 to efficiently perform similarity searches on reactants, generating new reactant combinations as candidate reaction pairs. A reaction template selection neural network was then used to suggest the most probable reaction templates. Based on the predicted probability scores, a threshold was applied: reactions above the threshold were considered additional positive samples near the decision boundary, while those below were treated as negative samples. The overall data construction workflow is shown in Fig. S8, and the composition of the final training, validation, and test sets is summarized in Table S11. These were split at a ratio of 9:0.5:0.5, with a balanced 1:1 ratio between positive and negative samples.
The model trained on this extended dataset is referred to as RXN Filter-deployed. In addition, we conducted a comparative evaluation on an external test set, comparing RXN Filter-benchmark, RXN Filter-deployed, and the built-in fast reaction filtering model from the ASKCOSv2 platform (ASKCOSv2 fast filter)53. Implementation details are provided in Supplementary Information Section S1.4.
Implementation details of platform
The platform was developed on the Ubuntu 22.04 OS using Python version 3.8.18. A graphical user interface was built utilizing the Flask framework, CSS, and HTML5, while the programmatic API functionality was implemented through Flask. All deep learning models were constructed using PyTorch version 2.1.2, with GPU acceleration achieved via CUDA version 12.1. The core of the platform is deployed through Docker containers. Additionally, some template-based single-step retrosynthesis prediction models, derived from the template_relevance subproject of ASKCOS v2, were deployed using Torch Server. All template-related computational tasks were carried out using RDChiral54 and RDKit55.
Implementation of agent
The Agent component is based on the ChemCrow framework, with its core model replaced from ChatGPT to the open-source Llama3.1:70b (deployed via Ollama). Additionally, by encapsulating the Request functionality, the Agent enables the invocation of ChemEnzyRetroPlanner’s seven module tools, serving as a replacement for the paid IBM Chemistry synthesis planning tools used in ChemCrow. Furthermore, the Agent introduces tools for enzymatic reaction identification, enzyme recommendation, and enzyme active site detection and annotation. Complete set of tool support is presented in Fig. 5b. We also developed a standalone Agent dialogue interface based on Streamlit, where users can interact with the chatbot to perform various tasks related to hybrid enzymatic-organic synthesis planning. The chatbot interface is shown in Fig. S9. We also present the system prompt used in the implementation of the Agent in Supplementary Information S1.9.
Operational requirements
The ChemEnzyRetroPlanner system is modular and can be deployed in two independent components. The first is the platform web server, which hosts the web interface and all predictive tools described herein. It primarily handles user interaction and the persistent invocation of models. The second is the Agent client, which is responsible for dialogue-based reasoning and the execution of chain-of-thought logic.
The platform web server can be reliably operated on a standard consumer-grade computer. Empirical tests confirmed that it can be stably deployed on a Dell OptiPlex-7090 workstation equipped with a single NVIDIA RTX 3060 GPU (12 GB VRAM), an 8-core Intel® Core™ i7-11700 CPU @ 2.50 GHz, and 32 GB of RAM. In contrast, the Agent client runs LLMs with up to 70 billion parameters, requiring significantly more computational resources. Recommended deployment settings include either a single NVIDIA GPU with at least 48 GB of VRAM (e.g., RTX A6000-48GB, A100-80GB, or H100-80GB), or an inference server equipped with at least three NVIDIA GPUs, each with 24 GB of VRAM (e.g., RTX 3090-24GB or RTX 4090-24GB), to ensure efficient inference performance.
All predictive models integrated into the system were trained on a server equipped with NVIDIA Tesla A100-SXM4-80GB GPUs and 64 core AMD EPYC 7742 CPUs @ 2.25 GHz. Inference benchmarking was performed on the aforementioned Dell OptiPlex-7090 desktop.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ChemEnzyRetroPlanner web platform is available at http://cadd.zju.edu.cn/retroplanner, with an alternate access link at http://cadd.iddd.group/retroplanner. The detailed platform documentation can be found at http://cadd.iddd.group/retroplanner/help. The precompiled environment and model weights required to run ChemEnzyRetroPlanner are available in the Hugging Face repository at https://doi.org/10.57967/hf/666756. Source data for Figs. 3, 4, S14 and S15 are provided with this paper. Source data are provided with this paper.
Code availability
The source code of the ChemEnzyRetroPlanner platform is available at https://github.com/wangxr0526/ChemEnzyRetroPlanner and archived on https://zenodo.org/records/1733174757 under the MIT license. The implementation of the agent core and user interface is accessible at https://github.com/wangxr0526/ChemEnzyRetroPlanner_agent and archived on https://doi.org/10.5281/zenodo.1741645658, also under the MIT license.
References
Corey, E. J., Long, A. K. & Rubenstein, S. D. Computer-assisted analysis in organic synthesis. Science 228, 408–418 (1985).
Corey, E. J. & Todd Wipke, W. Computer-assisted design of complex organic syntheses. Science 166, 178–192 (1969).
Corey, E. J. The logic of chemical synthesis: multistep synthesis of complex carbogenic molecules (Nobel Lecture). Angew. Chem. Int. Ed. Engl. 30, 455–465 (1991).
Mikulak-Klucznik, B. et al. Computational planning of the synthesis of complex natural products. Nature 588, 83–88 (2020).
Corey, E., Wipke, W. T., Cramer, R. D. III & Howe, W. J. Computer-assisted synthetic analysis. Facile man-machine communication of chemical structure by interactive computer graphics. J. Am. Chem. Soc. 94, 421–430 (1972).
Wang, X. et al. Generic interpretable reaction condition predictions with open reaction condition datasets and unsupervised learning of reaction center. Research 6, 0231 (2023).
Gao, H. et al. Using machine learning to predict suitable conditions for organic reactions. ACS Cent. Sci. 4, 1465–1476 (2018).
Pensak, D. A. & Corey, E. J. LHASA—Logic and heuristics applied to synthetic analysis. In Computer-Assisted Organic Synthesis. (eds Wipke, W. T. & Howe, W. J.) (American Chemical Society, 1977).
Saigiridharan, L. et al. AiZynthFinder 4.0: developments based on learnings from 3 years of industrial application. J. Cheminform. 16, 57 (2024).
Genheden, S. et al. AiZynthFinder: a fast, robust and flexible open-source software for retrosynthetic planning. J. Cheminform. 12, 70 (2020).
Coley, C. W., Green, W. H. & Jensen, K. F. Machine learning in computer-aided synthesis planning. Acc. Chem. Res. 51, 1281–1289 (2018).
Genheden, S. & Bjerrum, E. PaRoutes: towards a framework for benchmarking retrosynthesis route predictions. Digital Discov. 1, 527–539 (2022).
Yin, X. et al. Enhancing generic reaction yield prediction through reaction condition-based contrastive learning. Research 7, 0292 (2024).
Lowe, D. Chemical reactions from US patents (1976-Sep 2016). https://figshare.com/articles/Chemical_ reactions_from_US_patents_1976-Sep2016_/5104873 (2017).
Goodman, J. Computer software review: Reaxys. J. Chem. Inf. Model. 49, 2897–2898 (2009).
Zheng, S. et al. Deep learning driven biosynthetic pathways navigation for natural products with BioNavi-NP. Nat. Commun. 13, 3342–3342 (2022).
Probst, D. et al. Biocatalysed synthesis planning using data-driven learning. Nat. Commun. 13, 1–11 (2022).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Moretti, S. et al. MetaNetX/MNXref: unified namespace for metabolites and biochemical reactions in the context of metabolic models. Nucleic Acids Res. 49, D570–D574 (2021).
Alcántara, R. et al. Rhea—a manually curated resource of biochemical reactions. Nucleic Acids Res. 40, D754–D760 (2012).
Anand, M., Upadhyay, V. & Maranas, C. D. minChemBio: expanding chemical synthesis with chemo-enzymatic pathways using minimal transitions. ACS Synth. Biol. 14, 756–770 (2025).
Levin, I. et al. Merging enzymatic and synthetic chemistry with computational synthesis planning. Nat. Commun. 13, 7747 (2022).
Li, H., Liu, X., Jiang, G. & Zhao, H. Chemoenzymatic synthesis planning guided by reaction type score. J. Chem. Inf. Model. 64, 9240–9248 (2024).
Kreutter, D. & Reymond, J.-L. Chemoenzymatic multistep retrosynthesis with transformer loops. Chem. Sci. 15, 18031–18047 (2024).
Kreutter, D. & Reymond, J.-L. Multistep retrosynthesis combining a disconnection aware triple transformer loop with a route penalty score guided tree search. Chem. Sci. 14, 9959–9969 (2023).
Sankaranarayanan, K. & Jensen, K. F. Computer-assisted multistep chemoenzymatic retrosynthesis using a chemical synthesis planner. Chem. Sci. 14, 6467–6475 (2023).
Finnigan, W., Hepworth, L. J., Flitsch, S. L. & Turner, N. J. RetroBioCat as a computer-aided synthesis planning tool for biocatalytic reactions and cascades. Nat. Catal. 4, 98–104 (2021).
Zeng, T., Jin, Z., Zheng, S., Yu, T. & Wu, R. Developing BioNavi for hybrid retrosynthesis planning. JACS Au 4, 2492–2502 (2024).
Chen, B., Li, C., Dai, H. & Song, L. Retro*: learning retrosynthetic planning with neural guided A* search. In International Conference on Machine Learning. (eds Daumé III, H. & Singh, A.) (PMLR, 2020).
Achiam, J. et al. Gpt-4 technical report. Preprint at https://arxiv.org/abs/2303.08774 (2023).
Grattafiori, A. et al. The Llama 3 Herd of Models. Preprint at arXiv 2407.21783 (2024).
Jin, Q., Yang, Y., Chen, Q. & Lu, Z. Genegpt: augmenting large language models with domain tools for improved access to biomedical information. Bioinformatics 40, btae075 (2024).
Liu, Z., Chai, Y., & Li, J. Toward automated simulation research workflow through LLM prompt engineering design. J. Chem. Inf. Model. 65, 114–124 (2024).
Bran, A. et al. Augmenting large language models with chemistry tools. Nat. Mach. Intell. 6, 1–11 (2024).
Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
Ruan, Y. et al. An automatic end-to-end chemical synthesis development platform powered by large language models. Nat. Commun. 15, 10160 (2024).
Ma, Q., Zhou, Y. & Li, J. Automated retrosynthesis planning of macromolecules using large language models and knowledge graphs. Macromol. Rapid Commun. 2500065 (2025).
Wang, X. et al. Multi-modal deep learning enables efficient and accurate annotation of enzymatic active sites. Nat. Commun. 15, 7348 (2024).
Zheng, S. et al. Deep learning driven biosynthetic pathways navigation for natural products with BioNavi-NP. Nat. Commun. 13, 3342 (2022).
Koch, M., Duigou, T. & Faulon, J.-L. Reinforcement learning for bioretrosynthesis. ACS Synth. Biol. 9, 157–168 (2019).
Levin, I., Liu, M., Voigt, C. A. & Coley, C. W. Merging enzymatic and synthetic chemistry with computational synthesis planning. Nat. Commun. 13, 7747 (2022).
Sayyed, I. A. et al. Asymmetric synthesis of aryloxypropanolamines via OsO4-catalyzed asymmetric dihydroxylation. Tetrahedron 61, 2831–2838 (2005).
Patel, B. et al. Process development and scale-up of AZD7545, a PDK inhibitor. Org. Process Res. Dev. 16, 447–460 (2012).
Weininger, D. SMILES, a chemical language and information system: 1: introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
Schwaller, P. et al. Mapping the space of chemical reactions using attention-based neural networks. Nat. Mach. Intell. 3, 144–152 (2021).
Zeng, Z., Guo, J., Jin, J. & Luo, X. CLAIRE: a contrastive learning-based predictor for EC number of chemical reactions. J. Cheminform. 17, 2 (2025).
Carbonell, P. et al. Selenzyme: enzyme selection tool for pathway design. Bioinformatics 34, 2153–2154 (2018).
Stoney, R. Selenzyme 2023 (accessed 16 June 2025); https://github.com/RuthStoney/selenzyme2023.git (2023).
Stoney, R. A., Hanko, E. K. R., Carbonell, P. & Breitling, R. SelenzymeRF: updated enzyme suggestion software for unbalanced biochemical reactions. Comput. Struct. Biotechnol. J. 21, 5868–5876 (2023).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2021).
Probst, D., Schwaller, P. & Reymond, J.-L. Reaction classification and yield prediction using the differential reaction fingerprint DRFP. Digital Discov. 1, 91–97 (2022).
Johnson, J., Douze, M. & Jégou, H. Billion-scale similarity search with GPUs. IEEE Trans. Big Data 7, 535–547 (2019).
Tu, Z. et al. ASKCOS: open-source, data-driven synthesis planning. Acc. Chem. Res. 58, 1764–1775 (2025).
Coley, C. W., Green, W. H. & Jensen, K. F. RDChiral: an RDKit wrapper for handling stereochemistry in retrosynthetic template extraction and application. J. Chem. Inf. Model. 59, 2529–2537 (2019).
Landrum, G. et al. rdkit/rdkit: 2024_09_4 (Q3 2024) Release_2024_09_4 edn. Zenodo (2024).
Wang, X. et al. ChemEnzyRetroPlanner_metadata (Revision f0ef914). Hugging Face https://doi.org/10.57967/hf/6667 (2025).
Wang, X. et al. ChemEnzyRetroPlanner (accessed 22 October 2025). Zenodo https://doi.org/10.5281/zenodo.17331747 (2025).
Wang, X. et al. ChemEnzyRetroPlanner Agent (accessed 22 October 2025). Zenodo https://doi.org/10.5281/zenodo.17416456 (2025).
Schwaller, P. et al. Predicting retrosynthetic pathways using transformer-based models and a hyper-graph exploration strategy. Chem. Sci. 11, 3316–3325 (2020).
Acknowledgements
This work was financially supported by National Key R&D Program of China (2024YFA1307501 to T.H.), National Natural Science Foundation of China (22220102001 to T.H., 22373085 to C.-Y.H., 22503082 to X.W.), and Postdoctoral Fellowship Program of CPSF (GZC20252369 to X.W.).
Author information
Authors and Affiliations
Contributions
X.W. contributed to the main ideas, algorithm design, coding, and writing of the manuscript. X.Yin developed the enzyme recommendation models, performed benchmarking, co-wrote related sections, and participated in website and API testing. X.Z., H.Z., S.G., Z.W., O.Z., W.Q and Y.H. participated in discussions on method implementation and model evaluation. Y.L., D.J. and M.W. helped with data collection and validation. H.L. provided computational resources. X.Yao supervised the project, provided computational resources, and contributed to manuscript revisions. C.-Y.H. and T.H. conceived and guided the overall research, contributed to the paper revisions, and were responsible for the overall quality and direction of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Yin, X., Zhang, X. et al. A virtual platform for automated hybrid organic-enzymatic synthesis planning. Nat Commun 16, 10929 (2025). https://doi.org/10.1038/s41467-025-65898-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65898-3