Abstract
Gene expression diversity is crucial for crop breeding, yet population genomics has focused primarily on sequence polymorphisms. A single reference genome for RNA-seq cannot handle introgression bias. Here, we conduct RNA-seq for 328 wheat lines, including landraces and elite cultivars from China and the United States, to investigate the expression variation underlying agronomic traits. Leveraging pan-genome resources, we identify 23,296 more transcripts than using the Chinese Spring reference. We construct a pan-gene regulatory atlas through eQTL analysis, revealing the tight regulation of introgressed genes. We identify 299 high-confidence candidate genes for 34 agronomic traits and resistance to 8 Blumeria graminis f. sp. tritici isolates, more than one-fifth of which were absent from the Chinese Spring. Utilizing the Kenong 9204 mutant library, 73.7% of the candidates show significant phenotypic effects. Our work mitigates the reference bias and highlights the impact of breeding-driven directional expression changes on wheat adaptation and improvement.
Similar content being viewed by others
Introduction
Wheat is the most important food crop worldwide, accounting for 19% of the daily caloric intake and 21% of the protein needs of the global population1. In modern wheat breeding, scientists have successfully developed high-yield, disease-resistant, and stress-tolerant wheat varieties. This genetic improvement is particularly essential when facing the dual challenges of climate change and a growing global population2. Wheat is vulnerable to various diseases, including Fusarium head blight, rust, and powdery mildew, in different growing environments. Genetic diversity studies enable breeders to identify and utilize disease resistance genes, leading to the development of wheat varieties with increased resistance. Moreover, when genetic diversity is high, the broad gene pool for wheat to adapt to harsh climate conditions, such as drought and saline soils3. Maintaining and expanding the genetic diversity of wheat is key to preventing genetic bottlenecks and provides abundant genetic resources for future breeding4.
SNP diversity has been extensively characterized in recent years. One of the conclusions is that the introgression of landraces and wild relatives has shaped the genetic diversity of wheat. Landraces, which are a rich genetic resource, have been less influenced by historical and geographical effects, preserving a significant number of genes that have not been widely utilized in modern breeding. These genes can be used to improve the diversity of cultivars, particularly in terms of complex quantitative traits and stress-resistance traits. Cheng et al. conducted whole-genome resequencing on 827 A. E. Watkins landraces and 208 modern varieties to investigate the genetic and phenotypic diversity present in the historical Watkins germplasm collection. This research highlights the unique allelic and haplotypic variation in landraces, thereby providing resources for future breeding efforts5. Niu et al. collected 180 landraces and 175 cultivars to investigate the genetic variation in modern Chinese and American breeding programs through whole-genome resequencing. This study highlights the necessity of conserving and utilizing the genetic diversity of landraces during breeding6. Additionally, the introgression of wild relatives serves as a potential resource for increasing genetic diversity. He et al. performed exome sequencing approximately 1000 hexaploid and tetraploid wheat lines, identifying gene introgression from wild relatives, and highlighting the important contribution of historical gene flow from wild relatives to the adaptive landscape of modern bread wheat7. Most wild relatives from the Triticeae tribe can hybridize with wheat, and through backcrossing or chromosome engineering, chromosome segments carrying specific alleles can be introgressed into the wheat genome. To date, numerous genes, particularly disease resistance genes, have been transferred into wheat from rye, various Triticum species, and the Aegilops, Thinopyrum and Dasypyrum genera8,9,10,11,12,13. However, only a small portion of the existing genetic diversity has been utilized, with many genes and alleles yet to be leveraged for broader trait improvements14.
Despite functional studies of individual genes demonstrating the importance of gene expression regulation in wheat improvement, genome-wide investigations into how breeding selection has shaped the expression landscape remain limited. For example, the regulatory expression of Vernalization 1 (VRN1), VRN2, and VRN3/Flowering locus T1 (FT1) during the vernalization process is essential for determining flowering time and environmental adaptability in wheat15. The photoperiod-insensitive Ppd-1 allele (e.g, Ppd-D1a) has been widely adopted in breeding to reduce sensitivity to day length. Ppd-1 modulates the expression of the flowering activator VRN3/FT1, thereby promoting early flowering under short photoperiods and enhancing grain development and adaptation16,17,18,19. The upregulation of the NAM-ATAF-CUC transcription factor TaNAC100 facilitates the expression of the starch synthesis-related genes TaGBSS1 and TaSUS2, increasing the starch content in seeds. Overexpression of TaNAC100 also affects the total seed protein content, suggesting a role in maintaining the balance between starch and stored protein20. The nitrate-responsive NAC transcription factor TaNAC2-5A positively regulates TaNRT2.5 and TaNRT2.1. The overexpression of TaNAC2-5A significantly increases nitrate absorption, grain nitrogen concentration, and yield, indicating its potential for simultaneously improving productivity and protein content21. Under salt stress, elevated expression of TaSOS1 improves root development and water potential, conferring increased salt tolerance and underscoring its breeding potential22,23. These gene-level insights illustrate the significance of transcriptional regulation for key agronomic traits; however, a comprehensive, genome-wide understanding of how selection has reshaped gene expression remains a critical unmet need for advancing wheat improvement.
A special case of dysregulation in homoeologs was correlated with the agronomic traits of wheat, which was later proven to be due to reference bias24,25. Reference bias refers to the underestimation of transcripts from non-reference alleles during quantification, which potentially compromising the accuracy of subsequent conclusions. This bias is particularly pronounced in complex polyploid genomes, such as those of hexaploid wheat, which harbor highly heterologous gene blocks resulting from the introgression of wild relatives. Consequently, a pan-transcriptome reference was proposed by integrating gene models from Chinese Spring (CS) and nine additional assemblies from the 10+ Wheat Genomes Project. They incorporated only the transcripts of genes with a 1-to-1 orthologous relationship to the genes of Chinese Spring and subsequently merged the quantitative results on the basis of the homology relationships of Chinese Spring25. Their method primarily addressed errors caused by allelic variation, however, they still face limitations regarding presence-absence variations. Despite the abundant genome assemblies of wheat cultivars and related species, RNA sequencing (RNA-seq) of cultivated wheat still uses gene models mostly from one single reference genome, i.e., Chinese Spring.
Here, we utilize 44 genome assemblies to construct a non-redundant pan-gene atlas along with the genes of Chinese Spring. A total of 328 common wheat accessions, selected from previous resequencing study6, are subjected to RNA sequencing. RNA-seq reads are aligned to the pan-gene atlas, and high-density-resequenced SNP data are integrated to construct a genetic regulatory map of gene expression. Introgressed genes are found to be trans-regulated, especially the resistance genes. Through the analysis of 34 field agronomic traits and the seedling resistance to 8 Blumeria graminis f. sp. tritici (Bgt) isolates, we identify 299 high-confidence candidate genes, including 74 non-CS genes. Of these, 86 agronomic trait-associated candidates are validated using the Kenong 9204 mutant library. Studies of genes that are differentially expressed between cultivars and landraces reveal divergent breeding trajectories across countries, with regulatory regions exhibiting stronger selection pressure. Modern breeding significantly alters regulatory networks in cultivars compared with those in landraces. Together, these findings highlight the genetic regulation of the wheat transcriptome and its contribution to breeding outcomes, providing a valuable resource for future wheat improvement.
Results
Single-reference bias underestimates the expression of introgressed genes according to RNA-seq
To investigate the genetic regulation of gene expression in modern wheat breeding improvement, we performed RNA sequencing on 2-week-old seedlings from a panel of 328 common wheat accessions (Supplementary Data 1), comprising 172 representative diverse landraces (LRs), 92 modern Chinese cultivars (MCCs) and 64 modern United States cultivars (USMCs), all of which had been previously whole-genome sequenced6. An average of 77.0 million paired-end Illumina reads (2 × 150 bp) were obtained per sample, followed by quality control and alignment to the Chinese Spring reference genome (IWGSC RefSeq v1.1)26.
The presence of introgressions can lead to reference bias in wheat RNA-seq analysis25. Therefore, we first tested the effect of reference bias for the most widely deployed 1 RS.1BL introgression, where the short arm of Secale cereale L. (rye) chromosome 1R translocated with the short arm of wheat chromosome 1B. Fifty accessions in our panel carried this introgression (Supplementary Data 2). The number of expressed genes (transcripts per million, TPM > 0.5)24,27 across the sliding window along the short arm of Chinese Spring chromosome 1B was significantly lower in the introgression lines than in the non-introgressed lines (Fig. 1a). On average, each 10 Mb window contained 12 expressed genes in the introgressed lines and 20 expressed genes in the non-introgressed lines (two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16) (Fig. 1a). Similarly, the average expression level of the 1RS.1BL translocation was markedly lower than that of the native 1BS arm in non-introgression lines (two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16) (Fig. 1b and Supplementary Fig. 1). Given that rye diverged from diploid wheat approximately 9.6 million years ago28, quantification of gene expression from the rye introgression based on the Chinese Spring reference may result in significant underestimation due to sequence divergence.
a The left panel shows the number of expressed genes in introgression lines (n = 50) and non-introgression lines (n = 278) along chromosome 1BS (0–300 Mb) on the basis of Chinese Spring RefSeq v1.1 as the reference genome, with a sliding window of 10 Mb and a step size of 2.5 Mb. The right panel presents boxplots comparing the number of expressed genes per window between introgression lines (n = 50) and non-introgression lines (n = 278) within the chromosome 1BS (0–240 Mb) (two-sided Wilcoxon rank-sum test, p < 2.2 × 10–16). b Average gene expression of genes shared between introgression and non-introgression lines in the chromosome 1BS (0–240 Mb) region, with Chinese Spring RefSeq v1.1 as the reference genome. c A heatmap displays missing genotype sites, and a boxplot illustrates the genotype loss rate with the Chinese Spring RefSeq v1.1 as the reference genome. In the heatmap, n represents the number of accessions, orange represents modern Chinese cultivars (MCCs), cornflower blue represents modern United States cultivars (USMCs), and green represents landraces (LRs). The upper red color bar indicates introgressed fragments, and four genotype categories are depicted: blue–green for homozygous reference alleles, gold for homozygous variant alleles, light gold for heterozygous variant alleles, and light gray for missing genotypes. The lower boxplot represents the missing genotype rates calculated with a 2 Mb sliding window and a 1 Mb step size, and the sample size for each boxplot matches the number ‘n’ shown in the heatmap above. Comparisons of all introgression regions vs. background regions for chromosomes 1 A, 1B, 2D, 3D, and 5B: two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16. In (a) and (c), the box shows the median and interquartile range (IQR). The end of the top line is the maximum or the third quartile (Q) + 1.5 × IQR. The end of the bottom line denotes either the minimum or the first Q − 1.5 × IQR. The dots are more or less than Q ± 1.5 × IQR. d The number of expressed genes was calculated using a merged reference genome of Chinese Spring (CS) and introgression donors. The method for quantifying expressed genes is consistent with that in Fig. 1a; non-introgression lines include only the 100 samples with the most consistent gene expression (gray lines). The sample sizes for the introgression lines are as follows: chromosomes 1 A (n = 8), 1B (n = 50), 2D (n = 49), 3D (n = 4) and 5B (n = 74) (red lines). The upper line chart shows the number of expressed genes from Chinese Spring in the merged genome, while the lower line chart displays the number of expressed genes from the introgressed reference genome in the merged assembly. ****p < 0.0001. Source data are provided as a Source Data file.
To understand the scale of reference bias due to large segment introgression, we first determined the genomic locations of these introgressions. We utilized the genotype loss rates to identify multiple deletion regions across the Chinese Spring genome. Those regions with continuously high missing rates were considered as potential introgressions29. We focused on five introgressions larger than 20 Mb from known donor species, including 1RS.1AL (chr1A: 0–214.4 Mb) and 1RS.1BL (chr1B: 0–239.3 Mb)30, as well as chr2D (570.1–619.2 Mb) from Aegilops markgrafii31,32, chr3D (500–615.5 Mb), and the terminal ~ 60 Mb of chr3D in LongReach Lancer, which has been confirmed as an introgression from Thinopyrum ponticum31. Additionally, chr5B (497.2–537 Mb) originated from Triticum timopheevii (Fig. 1c and Supplementary Data 2)29. Those regions are more likely to represent introgressions rather than deletions on the basis of two lines of evidence: (1) the missing rate in these regions was significantly higher than that in other genomic regions (all introgression vs. background region comparisons for chromosomes 1 A, 1B, 2D, 3D and 5B: two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16) (Fig. 1c); and (2) the missing rate in protein coding sequences was significantly lower than that in non-coding regions (two-sided Wilcoxon rank-sum test, p value < 0.0001) (Supplementary Fig. 2), indicating the presence of homologous sequences in the collinear regions between Chinese Spring and non-Chinese Spring genomes. Gene expression within these introgressed regions was also systematically underestimated (two-sided Wilcoxon rank-sum test, p value < 0.0001) (Supplementary Figs. 1 and 3).
To accurately quantify gene expression in introgressed regions, we merged the donor genome with the Chinese Spring reference genome (see “Methods”) and reanalyzed gene expression for all the wheat samples. We found that when only the Chinese Spring reference genome was used, the average numbers of expressed genes detected in the introgression lines within the corresponding translocated chromosome segments (chr1A: 0–214.4 Mb; chr1B: 0–239.3 Mb; chr2D: 570.1–619.2 Mb; chr3D: 500–615.5 Mb; and chr5B: 497.2–537 Mb) were 14, 12, 39, 26 and 27 per 10 Mb window, respectively. In contrast, when the merged genome was used as a reference, the average number of expressed genes mapped to these Chinese Spring segments decreased to 4, 4, 22, 12 and 9, while the corresponding donor-derived regions rye (chr1R: 0–280 Mb), Renan (chr2D: 570–635 Mb), Th. elongatum (chr3E: 500–676.9 Mb), and T. timopheevii (chr5G: 435–485 Mb) presented higher average numbers of expressed genes—15, 26, 19 and 29 per 10 Mb window. These results demonstrate that using the merged genome enables a substantial number of mis-mapped transcripts to be correctly assigned to their donor genome origins, effectively recovering actively expressed genes in the introgressed regions across each 10 Mb window (Fig. 1d, Supplementary Fig. 2 and Supplementary Note 1). Thus, the reference bias can be severe when expression in introgression is measured using a single reference genome. The incorporation of gene models from donor species is essential for accurate estimation of expression levels in introgressed regions.
Utilizing pan-genome resources for accurate quantification of gene expression
Recent studies have shown that wild relatives, such as wild emmer, can contribute up to 15% of the wheat genome, and most introgressed fragments are less than 1 Mb in size7,33. In addition to the large introgressed segments from rye, Ae. markgrafii, Th. ponticum and T. timopheevii, the majority of the introgressions in our panel were likely smaller chromosomal fragments. To address the limitations of using a single reference genome and to improve the detection of alien gene expression, we constructed a pan-gene atlas from 44 publicly available Triticeae genomes (Supplementary Data 3). Simply combining multiple genomes as a reference for RNA-seq read mapping is insufficient, as conserved gene sequences across species are often misaligned because of the limitations of short read alignment algorithms. Thus, we developed a workflow for constructing a non-redundant pan-gene atlas.
First, all 107,422 genes from the Chinese Spring reference genome were retained (Fig. 2c). Second, to study the expression patterns of large introgressed chromosomal fragments, 5492 genes from four large introgressions in our wheat panel were included (Fig. 2c). Third, to comprehensively capture small introgressed genes absent from Chinese Spring, non-redundant genes from 39 Triticeae genomes with different ploidy levels were selected through the following two steps: (1) Genes homologous to 107,422 Chinese Spring genes and 5492 large-introgression genes were removed. We used OrthoFinder to classify 190,752 orthologous groups from the 44 genomes on the basis of homology (Fig. 2a, b). In total, 70,930 orthologous groups containing at least one gene from Chinese Spring or the four large introgressions were excluded. The remaining 119,822 orthologous groups, containing genes from 1 to 39 other genomes, were retained (Supplementary Fig. 4a). (2) For each of the 119,822 retained orthologous groups, the longest transcript was selected to represent the group. Among these genes, 53,852 genes were classified as assigned genes, as their orthogroups contained genes from at least two genomes, while 65,970 genes were classified as unassigned genes, whose orthogroups contained genes from only a single genome (Fig. 2c). The three gene sets were integrated to create a non-redundant pan-gene atlas consisting of 232,736 genes (Fig. 2c). This resource enables accurate quantification of gene expression and facilitates functional characterization of alien genomic segments, thereby advancing our understanding of how germplasm diversity shapes expression variation.
a Number of high-confidence genes in the 44 genomes of different wheat varieties and related species at different ploidy levels. b Density distribution of the number of homologous genes in each orthologous group. c Construction of the pan-gene atlas from three parts: (1) all high-confidence genes of Chinese Spring, (2) genes from large introgressed fragments identified in the analysis, and (3) assigned genes and unassigned genes from the 39 remaining genomes. d Number of expressed genes in each genome of the pan-gene atlas for each wheat line (n = 328) of our RNA-seq panel. Each dot represents a wheat line. The asterisk denotes the number of genes with a TPM > 0.5 in more than 5% of the samples at the population level. The box depicts the median and interquartile range (IQR). The whiskers extend to the most extreme data points within 1.5 × IQR from the quartiles, and points outside this range are shown as outliers. e Density distribution of the mapping rates for RNA-seq data from 327 wheat accessions aligned separately to Chinese Spring and the pan-gene atlas as reference gene models. f The number of expressed genes for 327 wheat accessions, separately quantified using Chinese Spring and the pan-gene atlas as a reference. g Heatmap showing the genome proportion of expressed genes for each wheat accession. The bottom color bar indicates the type of wheat accessions, the middle heatmap shows the proportion of expressed genes from different genomes (genome colors are consistent with Fig. 2c), and the top heatmap represents the introgression of foreign fragments in different wheat varieties. Source data are provided as a Source Data file.
To evaluate the alignment accuracy and gene detection improvement, RNA-seq data from all the wheat samples were mapped to the pan-gene atlas. The average alignment rate of the pan-gene atlas was 87.6%, representing a 3.5% improvement over that of the single Chinese Spring reference genome (Fig. 2e and Supplementary Data 4 and 5). A total of 80,128 genes were detected as expressed at population level (Fig. 2d), including 54,237 (67.7%) Chinese Spring genes, 14,137 (17.6%) assigned genes, 10,357 (12.9%) unassigned genes, and 1397 (1.8%) alien genes, with 23,296 more genes detected than the single Chinese Spring reference (Fig. 2f). On a per-sample basis, expressed Chinese Spring genes accounted for 71.66–77.72% of the total, assigned genes for 11.82–15.96%, unassigned genes for 9.34–11.18%, and alien genes for 0.49–1.80%. A total of 53 samples contained more than one introgressed segment (Fig. 2g and Supplementary Data 5 and 6). Interestingly, compared with landraces, the MCCs and USMCs presented significantly greater numbers of expressed genes, particularly for non-Chinese Spring genes (two-sided Wilcoxon rank-sum test, p value < 0.01) (Fig. 2d and Supplementary Figs. 4b and 5), suggesting that modern breeding has increased gene expression diversity.
Gene expression patterns of introgressed segments from rye, Ae. markgrafii, Th. ponticum and T. timopheevii
Introgressed segments from wild relatives can introduce additional genes or regulatory elements into the wheat genome, potentially reshaping gene expression and agronomic traits. Recent studies have shown that species-specific genes within introgressed segments are rarely expressed, whereas genes replacing wheat homologs tend to be downregulated25. To investigate expression changes associated with wild-type relative introgressions, we mapped RNA-seq data from 6 rye samples, 2 Renan samples (excluded from the main text because of the limited sample size, Supplementary Fig. 6d and f), 7 Th. ponticum samples, and 3 T. timopheevii samples to a pan-gene atlas combined with the corresponding donor genome (Supplementary Data 7). We analyzed the expression patterns of 5492 alien genes in both the donor species and their corresponding wheat introgression lines. The proportion of expressed genes within introgressed segments was significantly lower in the wheat lines than in their wild donors (two-tailed Student’s t test, p value < 0.001) (Fig. 3a, f; Supplementary Fig. 6e). Overall, the proportion of expressed species-specific genes was significantly lower than that of conserved genes (two-tailed Student’s t test, p value < 0.0001) (Fig. 3b, g; Supplementary Fig. 6a, g). Moreover, gene conservation was positively associated with the likelihood of expression (Fig. 3c, h; Supplementary Fig. 6h and i), which is consistent with observations in model organisms34,35,36.
a Percentage of expressed genes within the chromosome 1RS (0–280 Mb) region of the rye translocation lines 1RS.1AL, 1RS.1BL and rye lines. b Percentage of expressed specific and conserved genes within the chromosome 1RS (0–280 Mb) region in the rye translocation lines 1RS.1AL, 1RS.1BL, and rye lines. In (a) and (b), each dot represents one sample, and n indicates the number of samples. Two-tailed Student’s t test. c Relationships between the conservation of rye and wheat genes and the number of expressed genes in the 1RS.1BL introgression lines; n indicates the number of samples. d Functional enrichment analysis of differentially expressed introgressed genes between the 1RS.1BL translocation lines and rye lines. Two-sided Fisher’s exact test, Benjamini–Hochberg for multiple comparisons, p-adjust < 0.05. Bubble size corresponds to the number of genes annotated to a given term, and the color scale represents fold enrichment. e RT‒qPCR-based validation of differentially expressed genes between the three 1RS.1BL translocation lines and three rye lines using RT‒qPCR, with Tatublin as the internal control, and the results were calculated using the comparative CT method. RT‒qPCR was performed three times for each sample, and each time as a technical replicate. The error bars represent the mean ± SDs (n = 3 biological replicates). Two-tailed Student’s t test. f–i Analytical images of Th. ponticum introgression lines using Th. elongatum as the reference, corresponding to the analyses in panels (a–d). In (f–h), each dot represents one sample, and n indicates the number of samples. In (f–g) Two-tailed Student’s t test. The box depicts the median and interquartile range (IQR). The whiskers extend to the most extreme data points within 1.5 × IQR from the quartiles. In (i), the top terms, fold enrichment, and bubble size were determined as in (d). Source data are provided as a Source Data file.
To better understand how introgressed segments modulate gene regulation, we analyzed the differences in expression between wheat lines harboring known alien introgressions and their respective donor species. In the 1RS.1BL translocation lines, 136 genes were upregulated and 128 were downregulated, and in the Th. ponticum introgression lines, 234 genes were upregulated and 302 were downregulated. The upregulated genes in both cases were enriched in functions related to disease resistance and environmental adaptation, whereas the downregulated genes were linked to fundamental biological functions (Fig. 3d, i). These findings are consistent with the well-characterized roles of the 1RS.1BL and Th. ponticum introgression segments in increasing disease resistance. Resistance genes such as Lr10 (ortholog: SECCE1Rv1G0000670) for leaf rust resistance and Pm8 (ortholog: SECCE1Rv1G0001880) for powdery mildew were upregulated in the 1RS.1BL lines37,38. These directional changes in gene activity are likely driven by breeding selection, since wild relatives have been continuously utilized to improve the disease resistance of wheat during the past century. In addition, T. timopheevii has also provided valuable resistance genes for wheat breeding39. Gene Ontology enrichment analysis of the differentially expressed genes revealed significant overrepresentation of biological processes related to endogenous stimulus response, including auxin-mediated signaling and hormone regulation, suggesting that T. timopheevii introgressions may influence hormone-associated developmental pathways in the wheat background (Supplementary Fig. 6j).
To verify the enriched differentially expressed genes in the introgression lines, we selected three upregulated resistance genes and three downregulated core metabolic genes from 1RS.1BL for genomic DNA amplification to check for gene presence–absence variation. The three resistance genes were absent in some rye accessions but were consistently present in the introgression lines (Supplementary Fig. 6b). Reverse transcription quantitative PCR (RT‒qPCR) confirmed that SECCE1Rv1G0002890 and SECCE1Rv1G0003770 were significantly more highly expressed in introgression lines than in rye (two-tailed Student’s t test, p value = 1.9 × 10–5 and p value = 1.4 × 10–5) (Fig. 3e; Supplementary Data 8 and 9), whereas SECCE1Rv1G0008680 expression was comparable between the lines in which the gene was present (two-tailed Student’s t test, p value = 0.001) (Supplementary Fig. 6c; Supplementary Data 8 and 9). These results suggest that resistance genes may be transcriptionally activated in some wheat introgression lines. A more plausible explanation is that resistance genes that were originally present only in a subset of donor accessions are enriched in selective breeding, leading to their frequent presence in the introgression lines. As a result, the observed upregulation of resistance genes in the differential expression analysis is likely because most introgression lines carry these genes, whereas only a few donor accessions do. In contrast, the three core metabolic genes were present in both the donor and introgression genomes but were consistently downregulated in the introgression lines, possibly because of functional redundancy with native wheat genes (two-tailed Student’s t test, SECCE1Rv1G0021710 p value = 0.0002 and SECCE1Rv1G0017850 p value = 0.0008 and SECCE1Rv1G0000360 p value = 0.0223) (Fig. 3e; Supplementary Fig. 6b and c; Supplementary Data 8 and 9). These regulatory patterns are consistent among rye and Th. ponticum and T. timopheevii introgressions, suggesting that they might be caused by continuous breeding selection. Although introgression from wild relatives is a common practice in wheat breeding, our results revealed that not every introgressed gene is equally activated. Genes associated with disease resistance and stress adaptation are more likely to be retained and expressed40,41, whereas genes involved in core cellular functions tend to be repressed, likely because of redundancy. We next investigated the how these introgressed genes are regulated by the wheat genome using our population RNA-seq data.
eQTL map of the pan-gene atlas
To identify genetic loci associated with gene expression, we conducted association analysis between the expression levels of each gene and genome-wide SNPs on the basis of the Chinese Spring reference genome. This analysis revealed that expression quantitative trait loci (eQTLs) were significantly associated with transcript abundance. To characterize regulatory architectures, we classified eQTLs into three categories: intergenic-eQTLs, located in intergenic regions; inactive-eQTGs, residing in genes not expressed in the population; and active-eQTGs, located in genes with detectable expression in the population. Genes regulated by eQTLs are called eGenes (Fig. 4a). A total of 45,901 eGenes were identified, including 30,676 (66.8%) from Chinese Spring, 5791 (12.6%) unassigned eGenes, 8184 (17.8%) assigned eGenes, and 1250 (2.7%) alien eGenes (Fig. 4b). At the population scale, 34,227 (42.7%) non-eGenes lacked associated regulatory variants (Supplementary Fig. 7a, b), suggesting that their expression variation might be predominantly influenced by environmental factors. Among all the eGenes, 33,835 (73.7%) eGenes were associated with all three classes of regulatory loci, including intergenic-eQTLs, inactive-eQTGs, and active-eQTGs, and 130 (0.3%) eGenes were regulated exclusively by active-eQTGs and 272 (0.6%) solely by inactive-eQTGs. A total of 7001 (15.2%) eGenes were only regulated by intergenic-eQTLs, emphasizing the critical role of intergenic regions in gene expression regulation (Fig. 4c).
a Schematic model of eQTLs regulating eGenes. Active-eQTG indicates that the regulatory locus is located within a gene and expressed in the population, whereas inactive-eQTG indicates that the locus is within a gene but not expressed in the population. Intergenic-eQTLs refer to regulatory loci located in intergenic regions. b The distribution of eGene types and the number of eGenes. The middle color bar represents the genome classification, with colors consistent with those in Fig. 2c. c Venn diagram showing the distribution of eGenes regulated by three types of eQTLs. d The number of eQTLs for eGenes in the pan-gene atlas. e The outer figure shows the distribution of eQTLs corresponding to each eGene. The inner left panel shows the physical distance distribution between the most significant eQTLs and their eGenes (positive values indicate downstream regions of eGene, and negative values indicate upstream regions of eGene). The inner right panel illustrates the Spearman’s rank correlation coefficient (SCC) distribution between active-eQTGs and their eGenes. Correlation values are shown on the x-axis, and significance (–log10(p)) is shown on the y-axis. f The number of eQTLs for alien introgressed genes calculated for all samples and introgression lines, with the number of eQTLs on each chromosome displayed. We calculated eQTLs on the basis of two groups of lines, namely, ‘All lines’, in which all the wheat lines were used for calculating eQTLs and ‘Only introgression lines’, in which only lines with this introgression were used. g eQTL map of introgressed genes on Renan chr2D. The y-axis represents the position of eSNPs on wheat chromosomes, the dot plot x-axis represents the position of introgressed eGenes on Renan chr2D, and the bar plot x-axis represents the number of eQTLs. h In the line chart above, the x-axis represents the position of eGenes on Renan chr2D, and the y-axis represents the number of active-eQTGs on wheat chromosome 6B. Different colored points correspond to different eGenes in the regulatory network diagram below. The regulatory network diagram below shows the relationships between the eGenes (colored points in the line chart above) and their corresponding active-eQTGs on chromosome 6B. Source data are provided as a Source Data file.
A total of 4,140,408 eQTLs were found to regulate eGenes (Fig. 4d). Most eGenes were associated with a single eQTL (Fig. 4e). The physical proximity between eQTLs and their target eGenes appeared to influence regulatory strength42, with the strongest associations enriched near gene regions, particularly around transcription start sites (Fig. 4e). In this study, eQTLs located on the same chromosome as their target eGene in Chinese Spring were classified as cis-eQTLs, whereas those on different chromosomes were defined as trans-eQTLs. Compared with trans-eQTLs, cis-eQTLs exhibited significantly stronger association signals for the same gene (two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16 for all comparisons) (Supplementary Fig. 7c, d). Analysis of the strongest eQTL signals for Chinese Spring genes revealed that 65.19% were cis-eQTLs (Supplementary Fig. 8a, b).
Furthermore, we identified 2,104,958 pairs of active-eQTGs and eGenes with absolute Spearman correlation coefficients greater than 0.3 (|SCC| > 0.3, p value < 0.05)43, of which 1,779,804 (84.6%) pairs were positively correlated and 325,154 (15.4%) pairs were negatively correlated (Fig. 4e). A total of 84% of intergenic-eQTLs were located within transposable element (TE) regions, suggesting that TEs contribute to regulatory variation and potentially impact agronomical traits. The composition of TE types among intergenic eQTLs was consistent with previous report in wheat26, indicating the widespread distribution of regulatory elements across TE-rich regions (Supplementary Fig. 7e).
Trans-regulation of introgressed genes from rye, Ae. markgrafii, Th. ponticum and T. timopheevii
To investigate how introgressed genes are regulated by the wheat genome, we calculated eQTLs separately for only the introgression lines of 1RS.1BL, Ae. markgrafii, and T. timopheevii (Supplementary Note 2). For comparison, we also performed eQTL analysis for introgressed genes across all the wheat samples at the population scale. In total, 499 eGenes were located on 1RS across the entire population, whereas only 243 eGenes were detected in the 1RS.1BL introgression lines (Supplementary Fig. 9a). The number of eQTLs identified from the introgression lines was markedly lower than that obtained from the full population (Supplementary Fig. 9b). eQTLs from the full dataset were strongly enriched on chromosomes 1 A, 1B, 2D, 3D, and 5B (Fig. 4f, g; Supplementary Fig. 9c, e, and f), a pattern likely influenced by the population structure. These signals largely reflected the presence–absence variation of introgressed segments rather than true regulatory interactions. In contrast, eQTLs calculated from only introgression lines were more evenly distributed across the genome (Figs. 4f, g; Supplementary Fig. 9c, e), providing a more accurate view of how the wheat genome regulates introgressed genes.
Since the three alien introgression segments (rye: (chr1R: 0–280 Mb), Renan: (chr2D: 570–635 Mb), and T. timopheevii: (chr5G: 435–485 Mb)) underwent translocations with the wheat genome, all eQTLs located on the wheat chromosomes were treated as trans-eQTLs. However, because these introgressed segments share high sequence homology with specific wheat chromosomes, we performed synteny analysis to clarify their relationships. The results revealed strong collinearity between the introgressed segments and their homologous wheat chromosomes, whereas no collinearity was observed with non-homologous chromosomes (Supplementary Fig. 10). Given the known translocations, eQTLs located on Chinese Spring chromosomes 1B, 2D, and 5B were approximated as cis-eQTLs under our study’s definitions, whereas those on other chromosomes were considered as trans-eQTLs.
We then compared the regulatory signals from cis-eQTLs, trans-eQTLs, and eQTLs on homoeologous chromosomes. The analysis revealed that the regulatory signals of the cis-eQTLs were significantly stronger than those of the trans-eQTLs (two-sided Wilcoxon rank-sum test, p value < 0.0001). Compared with those from homoeologous chromosomes, the signals from cis-eQTLs were not always stronger (Supplementary Fig. 11a, b, and c), suggesting that both eQTLs on translocated chromosomes and those on homoeologous chromosomes play important roles in regulating introgressed genes.
Genes introgressed from Ae. markgrafii into chromosome 2D included a high number of eQTLs originating from chromosome 6B in the introgression lines (Fig. 4g). A total of 4 genes were associated with 700 eQTLs, including 255 active-eQTGs (Fig. 4g). The regulatory network between active-eQTGs and eGenes was enriched for metabolic processes (Fig. 4h). Additionally, the active-eQTG TraesCS5A02G120000 emerged as a regulatory hotspot, controlling 94 1RS eGenes across all 1RS.1BL lines (Supplementary Fig. 9c). This active-eQTG is located in the centromeric region of chromosome 5A, which displays genotypic divergence relative to the reference genome in both the 1RS.1AL and 1RS.1BL lines. These genotype differences are specific to introgression lines (Supplementary Fig. 9d), suggesting that the centromere of chromosome 5 A may influence the ability to integrate external chromosomes. Overall, the regulation of introgressed gene involves multiple chromosomes, reflecting a complex trans-regulatory effects in wheat. These findings offer mechanistic insights into the adaptive regulation of introgressed genes and provide a framework for improving stress resistance through targeted breeding.
Integrative modeling identified candidate genes from the pan-gene atlas for agronomic traits
Recent studies have shown that both transcriptome-wide association studies (TWASs) and summary data-based Mendelian randomization (SMR) integrate GWASs and eQTL data to identify genes associated with complex traits: in TWASs, gene expression prediction and transcriptome-wide association analysis are used to uncover associations between cis-regulated gene expression and traits, whereas in SMR, Mendelian randomization is used to explore potential causal relationships between gene expression and traits44,45. Here, we integrated GWASs, TWASs, SMR, and Spearman correlation coefficients (SCCs) between gene expression levels and phenotypic values using an eQTL map from the pan-gene atlas to identify candidate genes, including both Chinese Spring and non-Chinese Spring genes, that contribute to phenotypic variation (Fig. 5a). We identified 42 phenotypes, including 34 field agronomic traits and resistance phenotypes for 8 Bgt isolates (Supplementary Data 10). Genes identified by at least two methods were considered high-confidence candidates. A total of 260 high-confidence candidate genes were obtained for 34 field agronomic traits, including 71 non-Chinese Spring genes (Fig. 5b, c; Supplementary Figs. 12 and 13; Supplementary Data 11, 12, 13, 14, and 15). Given that most cloned powdery mildew resistance genes encode nucleotide-binding site leucine-rich repeat (NLR) proteins or kinases46, we designated 39 of such genes as high-confidence resistance candidates, including 3 from non-Chinese Spring segments (Fig. 5b, c; Supplementary Fig. 14; Supplementary Data 16, 17, 18, 19, and 20).
a Schematic diagram for the prediction of candidate gene. The GWAS QTL candidate regions include signals from eQTG and intergenic-eQTL. Candidate genes are predicted using cis-eQTLs through the TWAS, and both cis- and trans-eQTLs through SMR. b Venn diagram of the distribution of candidate genes predicted by the GWAS, the TWAS, SMR and the Spearman correlation coefficient (SCC) between expression levels and phenotypic values. c The number of candidate genes classified by the pan-gene atlas. d Three eQTL regulatory patterns assist in the identification of candidate genes. R1 represents candidate genes regulated by both gene and intergenic regions, R2 represents candidate genes regulated only by the gene region, and R3 represents candidate genes regulated only by the intergenic region. e Types and numbers of regulatory patterns for the candidate genes. f Verification of candidate genes using the Kenong 9204 mutant library. The phenotypes of the mutants and wild-type plants for the predicted candidate genes were analyzed using a two-tailed Student’s t test, with the red line representing the ttest distribution for the candidate genes. The blue–green line represents the t test p value distribution for the randomly selected genes. The same number of random genes were randomly selected as the candidate genes. This process was repeated 10 times. g Proportion of candidate genes validated by the Kenong 9204 mutant library compared with the average proportion of validated genes from ten random sets. h Representative candidate genes verified using the Kenong 9204 mutant library. The candidate genes were predicted to be associated with multiple agronomic traits and were validated for relevant traits (TKW, GW, GN, SPN, and GA) in the mutant library. The x-axis shows WT (wild-type) lines and the corresponding mutant lines of candidate genes, with n indicating the sample size. WT lines were used as the control, and the mutant lines of each candidate gene were compared to the WT line using a two-tailed Student’s t test. Red points represent premature termination (high effect) and yellow points represent non-synonymous mutations (moderate effect) in the candidate gene. The blue–green points represent wild type. The box depicts the median and interquartile range (IQR). The whiskers extend to the most extreme data points within 1.5 × IQR from the quartiles. Source data are provided as a Source Data file.
GWASs can identify only significant SNPs, whereas TWASs and SMR can directly identify candidate genes45,47. Among the candidate genes with eQTLs, regulatory relationships were classified into three models. In total, 97.3% of the candidates associated with 34 field agronomic traits and 8 Bgt isolates were identified as eGenes, with 95.3% regulated by both genetic and intergenic regions. A total of 2.0% of the genes were regulated exclusively by intergenic regions, highlighting the critical role of non-genic regions (Fig. 5d, e). Ppd-D1, which is regulated under the R1 model, was identified as a candidate gene for heading date according to both TWASs and SMR analyses. Ppd-D1, previously reported to be involved in the photoperiod response17, harbored a frameshift mutation (CCGACG → C) that significantly altered its expression level (two-sided Wilcoxon rank-sum test, p value = 6.3 × 10–13), ultimately affecting heading date (two-sided Wilcoxon rank-sum test, p value = 1.8 × 10–14) (Supplementary Fig. 15a). Using the SMR approach, we also identified TaGL3-5B, a gene associated with both grain length and width. TaGL3-5B is known to regulate grain size48 (Supplementary Fig. 15b). Furthermore, we further detected previously powdery mildew resistance genes, including the Pm4 allele49 (TraesSYM2A03G00828360 from SY Mattis and TraesAK58CH2A01G622200 from Aikang 58; Supplementary Fig. 16) and Pm3 (TraesCS1A02G008100)38 using SMR, as well as Pm5 alleles (e.g., TraesCS7B02G441700)50 using GWASs. In addition, we identified a candidate gene, TraesCS3D02G201900, for biomass per plant at the jointing stage that lacked SNP variation across the panel but was regulated by a cis-eQTL. TWASs and SMR linked this gene to phenotypic variation, suggesting that eQTLs may influence traits by modulating gene expression. Importantly, the identification of cloned genes absent from the Chinese Spring reference genome underscores the utility of the pan-gene atlas for transcriptome read mapping.
To validate the reliability of the candidate genes identified from the pan-gene atlas, we leveraged the indexed EMS mutant library of the wheat cultivar Kenong 920451. Among the 260 agronomic candidate genes, 138 had 164 homologs in the Kenong 9204 genome, and corresponding mutants were identified from approximately 2000 EMS lines. Among these genes, 133 homologous genes were associated with at least five mutant lines carrying either knockout or non-synonymous mutations and were subsequently used for validation. We evaluated seven traits, namely, grain area, grain length, grain perimeter, grain width, spikelet number, thousand-kernel weight and grain number, in both the mutants and the wild-type controls. Significant phenotypic differences were observed for 98 (73.7%) of the 133 homologous genes, corresponding to 86 candidates, including 8 non-Chinese Spring genes. By comparison, only 36.4% of the randomly selected genes demonstrated this level of phenotypic change, highlighting the high reliability of our candidates (Fig. 5f, g; Supplementary Fig. 17a and b; Supplementary Data 21, 22, and 23; Supplementary Note 3). Among the validated genes, 44 Kenong 9204 homologs carried premature termination mutations that resulted in significant phenotypic variation (Supplementary Data 22). For instance, the TraesKN5A01HG39180 mutant exhibited a pronounced reduction in both spikelet number and spike grain number, with the strongest observed in lines with premature termination. Mutants of TraesKN6A01HG11540 and TraesKN6A01HG11620 presented significant decreases in thousand-kernel weight and grain width, respectively (two-tailed Student’s t test, TraesKN6A01HG11540: (TKW, p value = 1.5 × 10–5; GW, p value = 0.0003) and TraesKN6A01HG11620: (TKW, p value = 0.0003; GW, p value = 0.0027)) (Fig. 5h and Supplementary Data 21, 22, and 23). In summary, we provided a robust method for identifying high-confidence candidate genes, including those from exogenous sources, and validated them using a functional mutant library, offering valuable genetic resources for improving wheat.
Differentially expressed genes and modules between different sub-populations
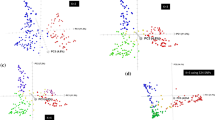
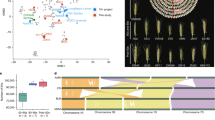
To investigate the impact of breeding on transcriptomic divergence, we first examined genes whose expression was significantly up- or downregulated across sub-populations. We are also wondering whether differentially expressed genes (DEGs) are involved in trait improvement. First, we observed dramatic expression changes between sub-populations, which is consistent with genetic divergence. A total of 6514 DEGs were identified between MCCs and LRs, of which 2893 (44.4%) were non-Chinese Spring genes; 5631 DEGs were found between USMCs and LRs, including 2354 (41.8%) non-Chinese Spring DEGs; and 4018 DEGs were detected between USMCs and MCCs, including 2246 (55.9%) non-Chinese Spring DEGs (Fig. 6a). These results suggest that the divergence between breeding programs (MCCs vs. USMCs) is smaller than that between cultivars and landraces. The high percentage of non-Chinese Spring DEGs (41.8%–55.9%) highlights the limitations of relying on a single reference genome in RNA-seq analyses.
a Number of DEGs for each pair of sub-populations. b Sharing of DEGs between sub-populations. c UMAP plot of the dimensionality reduction analysis of the DEG expression matrix. d Heatmap of the 8 modes of DEGs. The numbers on the left represent the count of DEGs. The dashed box in the middle highlights representative cloned genes of known function among the DEGs, and the right side shows functional enrichment for each module (M3–M8) (two-sided Fisher’s exact test, Benjamini–Hochberg for multiple comparisons, p-adjust < 0.05. The bubble size corresponds to the number of genes annotated to a given term, and the color scale represents fold enrichment). e FST values of genic regions for DEGs and non-DEGs: for MCCs vs. LRs, DEGs (n = 3564) and non-DEGs (n = 49,972); for USMCs vs. LRs, DEGs (n = 3222) and non-DEGs (n = 50,309); and for USMCs vs. MCCs, DEGs (n = 1721) and non-DEGs (n = 51,413). Two-sided Wilcoxon rank-sum test, p < 2.2 × 10–16 for all comparisons (DEGs vs. non-DEGs). Dots within the boxes indicate the FST values. f FST values of eQTLs for DEGs and non-DEGs. For MCCs vs. LRs, DEGs (n = 4715) and non-DEGs (n = 35,893); for USMCs vs. LRs, DEGs (n = 4162) and non-DEGs (n = 36,288); and for USMCs vs. MCCs, DEGs (n = 3304) and non-DEGs (n = 36,715). Two-sided Wilcoxon rank-sum test, p < 2.2 × 10–16 for all comparisons (DEGs vs. non-DEGs). Dots within the boxes indicate the FST values. g Relationships between the FST values of DEGs and the degree of differential expression. The data are presented as the mean values ± SEM. h Relationships between the FST values of eQTLs for DEGs and non-DEGs and the strength of eQTL signals. The data are presented as the mean values ± SEM. i XP-CLR scores of DEGs and non-DEGs: for MCCs vs. LRs, DEGs (n = 1788) and non-DEGs (n = 17,974); for USMCs vs. LRs, DEGs (n = 1596) and non-DEGs (n = 19,813). Two-sided Wilcoxon rank-sum test: p = 1.35 × 10–6 for MCCs vs. LRs and p = 2.89 × 10–4 for USMCs vs. LRs (DEGs vs. non-DEGs). Dots within the boxes indicate XP-CLR scores. j XP-CLR scores of eQTLs for DEGs and non-DEGs: for MCCs vs. LRs, DEGs (n = 4196) and non-DEGs (n = 31,363); for USMC vs. LR, DEGs (n = 3641) and non-DEGs (n = 31,887). Two-sided Wilcoxon rank-sum test: p = 5.02 × 10–8 for MCC vs. LR and 7.54 × 10–4 for USMC vs. LR (DEGs vs. non-DEGs). Dots within the boxes indicate XP-CLR scores. k Relationships between the XP-CLR scores of DEGs and the degree of differential expression. The data are presented as the mean values ± SEM. l Relationships between the XP-CLR scores of eQTLs for DEGs and non-DEGs. The data are presented as the mean values ± SEM. All boxes depict the median and interquartile range (IQR). The whiskers extend to the most extreme data points within 1.5 × IQR from the quartiles, and points outside this range are shown as outliers. Source data are provided as a Source Data file.
Second, we investigated whether the DEGs were genetically regulated by eQTLs. More than 81% of the DEGs between the cultivars and landraces were associated with at least one eQTL, which is significantly greater than the percentage associated with the genome-wide background (57%; two-proportion z-test, p value < 2.2 × 10–16) (Fig. 6a and Supplementary Fig. 7a). In addition, 94% of the DEGs between MCCs and USMCs were linked to eQTLs, suggesting that expression divergence between these breeding programs is driven largely genetically. The strong enrichment of eQTL-regulated DEGs across comparisons highlights the role of breeding selection in shaping transcriptional landscapes. Consistently, uniform manifold approximation and projection (UMAP)-based dimensionality reduction using these DEGs clearly separated the sub-populations, indicating that transcriptomic variation mirrors the underlying genetic structure (Fig. 6b, c).
To further dissect the transcriptional trajectories shaped by breeding in China and the United States, the DEGs were classified into eight co-expression modules (Fig. 6d). Modules M1 represent 31 genes whose expression is up-regulated exclusively in Chinese cultivars but down-regulated in American cultivars; and M2 represent 32 genes whose expression is up-regulated exclusively in American cultivars but down-regulated in Chinese cultivars, suggesting that the breeding directions of wheat in China and the United States do not significantly differ. Modules M3 and M4 contained genes whose expression was consistently upregulated or downregulated, respectively, in both MCCs and USMCs compared with that in landraces (LRs). The genes in M3 were enriched for abiotic stress and water response pathways, suggesting a shared emphasis on increasing stress resilience at the seedling stage. Conversely, M4 genes were enriched for secondary metabolism and cell wall organization, reflecting potential growth-defense trade-offs selected during modern breeding. Cloned genes within these modules reinforced these functional distinctions. M3 includes Lr34, TaAGL22, VRN2, SVP1, and WRKY45, which regulate environmental responses and flowering time52,53,54,55,56,57,58. M4 contained TaSnRK2.3-1 A, FUL2, VRN1, VRN3, and TaSuSy4, which are associated with flowering induction and secondary metabolism (Fig. 6d)6,59,60. VRN2 was upregulated in cultivars, whereas VRN1 and VRN3 were downregulated, which is consistent with a shift from spring wheat, predominantly in landraces, to winter wheat in modern cultivars61 (Supplementary Figs. 18 and 19). These findings were also validated using 1,034 accessions from previously published datasets5 (Supplementary Fig. 20). This trend reflects a breeding-driven transition toward a winter growth habit, which allows vernalization-dependent flowering and improved nutrient accumulation, thereby contributing to increased yield potential62,63.
Modules M5 and M7 captured divergent breeding directions between MCCs and USMCs. M5 genes were specifically upregulated in MCCs and enriched in resistance-related pathways, indicating that Chinese breeders have placed greater emphasis on disease resistance. M7 genes were upregulated in USMCs and enriched in post-embryonic and seed development functions, suggesting a focus on improving developmental traits and yield potential in U.S. breeding programs. For example, M5 included rye-derived 1RS genes, which are more prevalent in MCCs but also present in USMCs, highlighting the widespread introgression of alien resistance loci. M7 contained TaGRP-2, TaAGL18-A1, and Lr6752,64, which are associated with flowering time and resistance. In addition, M6 contained key yield- and quality-related genes such as TaISA2, GW5, Ppd-D1, TaAP2, TaTAR2.1-3 A, and TaGASR7-A117,65,66,67,68,69,70, whose expression was downregulated at the seedling stage in MCCs compared with that in LRs, possibly reflecting delayed expression until later developmental stages (Fig. 6d). In summary, these expression patterns illustrate that while both MCCs and USMCs have acquired resistance genes during modern breeding, resistance improvement has been emphasized more for MCCs, whereas developmental regulation has been prioritized for USMCs.
Population divergence and selection of differentially expressed genes
To investigate the genetic differentiation of DEGs under modern breeding, we performed an FST analysis between DEGs and non-DEGs. The FST values of the genomic regions containing DEGs were significantly greater than the FST values of the regions harboring non-DEGs (two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16 for MCCs vs. LRs, USMCs vs. LRs, and USMCs vs. MCCs) (Fig. 6e). Similarly, the FST values of the eQTL regulating DEGs were also significantly greater than those of the eQTLs associated with non-DEGs (two-sided Wilcoxon rank-sum test, p value < 2.2 × 10–16 for MCCs vs. LRs, USMCs vs. LRs, and USMCs vs. MCCs), indicating that not only were the DEGs more genetically differentiated but also that their regulatory loci experienced stronger divergence (Fig. 6f). Furthermore, as the FST values increased, the signals of the DEGs or their eQTLs increased, whereas the signals of the eQTLs regulating non-DEGs were significantly decreased (Fig. 6g, h).
To further whether DEGs are under selection, we conducted maximum likelihood ratio XP-CLR analysis to compare cultivars with landraces. The XP-CLR scores of the genomic regions containing DEGs were significantly greater than those of the non-DEGs in both MCCs vs. LRs (two-sided Wilcoxon rank-sum test, p value = 1.35 × 10–6 for MCCs vs. LRs, and p value = 2.89 × 10–4 for USMCs vs. LRs) (Fig. 6i). Similarly, compared with non-DEGs, eQTLs regulating DEGs had significantly higher XP-CLR scores (two-sided Wilcoxon rank-sum test, p value = 5.02 × 10–8 for MCCs vs. LRs, and p value = 7.54 × 10–4 for USMCs vs. LRs), indicating that DEGs and their regulatory loci were selected during the breeding process (Fig. 6j). Moreover, as the XP-CLR scores of the genomic regions containing DEGs increased, the signals of the DEGs increased; although there was only a slight upward trend in the signals of eQTLs regulating DEGs with increasing XP-CLR scores, the signals of eQTLs for non-DEGs showed a slight downward trend (Fig. 6k, l). Collectively, these results indicate that both DEGs and their associated eQTLs have undergone significant genetic differentiation and experienced stronger selection pressure during the breeding process.
Modern breeding reshaped the gene regulatory network of wheat
To investigate changes in gene co-expression and regulatory networks during modern breeding, we analyzed changes in co-expression patterns across sub-populations. Specifically, we calculated the number of correlations between active-eQTGs and eGenes with a |SCC| > 0.3 and a p value < 0.05 across different sub-populations (see Methods). A total of 1148 DEGs between LRs and MCCs, and 741 DEGs between LRs and USMCs, were not only regulated by active-eQTGs but were also identified as candidate genes for agronomic traits by at least one method (Fig. 7a). The 1148 candidate genes were regulated by 52,713 active-eQTGs, with 25,405 pairs (48.2%) and 43,156 pairs (81.8%) of active-eQTGs and eGenes showing co-expression (|SCC| > 0.3 and pvalue < 0.05) in LRs and MCCs, respectively (Supplementary Data 23 and 24). The proportion of co-expressed pairs in the MCCs was significantly greater than that in the LRs (two-proportion z-test, p value < 2.2 × 10–16) (Fig. 7b). Similarly, the 741 candidate genes were regulated by 32,636 active-eQTGs, with 16,054 pairs (49.2%) and 27,018 pairs (82.8%) of active-eQTGs and eGenes showing significant co-expression in LRs and USMCs, respectively (Supplementary Data 25). The proportion of co-expressed genes in USMCs was significantly greater than that in LRs (two-proportion z-test, p value < 2.2 × 10–16) (Fig. 7b). The co-expression regulatory networks of known genes such as TaISA2, involved in starch biosynthesis67, and VRN1-5A, associated with vernalization57, underwent substantial rewiring from LRs to MCCs or USMCs (Fig. 7c), further indicating that regulatory networks have been modified during modern breeding.
a Venn diagram of candidate genes, eGenes regulated by active-eQTGs, and DEGs. b Proportion of candidate genes in the orange blocks of the panel (Fig. 7a) where the Spearman’s rank correlation coefficient between active-eQTGs and eGene has a |SCC| > 0.3 and p value < 0.05. Differences in proportions were assessed using a one-sided two-proportion z-test, with p < 2.2 × 10–16. c Gene network from the orange blocks in Fig. 7a, showing only genes that satisfy p-GWAS < 1 × 10–5, p-TWAS < 1 × 10–4, p-SMR < 1 × 10–5, and |SCC| > 0.6. Nodes represent eGenes or active-eQTGs. Orange edges indicate positive expression correlations (SCC > 0.3 and p value < 0.05) between eGenes and active-eQTGs, whereas the blue edges represent the negative correlation (SCC < -0.3 and p value < 0.05). No edges were drawn between eGenes and active-eQTGs that did not meet these thresholds. To reduce complexity, only eGenes associated with agronomic traits that met all of the following criteria were visualized: GWAS p value < 1 × 10–5, TWAS p value < 1 × 10–4, SMR p value < 1 × 10–5, |SCC| > 0.6, and classification as DEGs. The small regulatory network on the right shows co-expression between the cloned genes (TaISA2 and VRN1-5A) and active-eQTGs. MCCs stands for Modern Chinese Cultivars. USMCs stands for United States Modern Cultivars. LRs stands for landraces. Source data are provided as a Source Data file.
Additionally, among the 665 candidate genes for powdery mildew resistance, 438 were regulated by 32,860 active-eQTGs (|SCC| > 0.3, p value < 0.05). In the landrace group, MCC group and USMC group, 22,194 pairs (67.5%), 26,144 pairs (79.6%) and 24,249 pairs (73.8%) of active-eQTGs and eGenes were co-expressed (|SCC|> 0.3, p value < 0.05), respectively (Supplementary Data 26). Compared with the LR group, the MCC and USMC groups had significantly greater proportions of co-expression relationships (two-proportion z-test, p value < 2.2 × 10–16 for both the MCCs vs. LRs and the USMCs vs. LRs) (Supplementary Fig. 21a and b). The regulatory networks of the cloned resistance genes Pm4 and Pm5 also differed between landraces and cultivars49,50. However, their co-expression networks appeared relatively simple, which may be attributed to the fact that powdery mildew resistance being a qualitative trait controlled by major-effect genes (Supplementary Fig. 21c). Taken together, these findings suggest that modern breeding has substantially altered co-expression and regulatory networks.
Discussion
Although several high-quality wheat reference genomes have been released, a pan-genome is not yet available for use in aligning short reads from next-generation sequencing (NGS)25. Frequent hybridization among wheat germplasms, including introgressions from wild relatives and domesticated progenitors, complicates accurate gene expression quantification using a single reference genome25. Previous population transcriptome studies predominantly rely on the Chinese Spring reference model24,71 and inevitably fail to capture the substantial expression diversity present in non-Chinese Spring samples. This loss may exclude critical insights pertaining to modern breeding improvements, such as segments derived from rye 1RS, Ae. markgrafii, Th. ponticum, T. timopheevii, and other sources.
To address this, we merged the gene models of Chinese Spring with non-redundant gene models from 43 Triticeae genomes. Unlike conventional pan-genome studies that require the annotation of all genomes using a unified pipeline72,73, our approach directly incorporates published gene models to construct a pan-gene catalog. This strategy not only effectively quantifies a larger number of non-Chinese Spring genes but also significantly reduces the computational burden and time required by traditional pan-genome construction. However, since these genomes were generated from independent studies with diverse annotation methodologies, some genes classified as ‘unassigned’ in this study may not be truly unique to a single genome. The ‘unassigned’ designation is solely a result of OrthoFinder analysis and does not fully capture the genomic differences among the 44 genomes. A comprehensive comparison of genomic differences would require a classical pan-genome analysis pipeline. Nevertheless, for the purpose of this study, identifying additional transcripts and candidate genes and leveraging the integrated pan-gene atlas provides a simpler and more efficient solution.
Moreover, the classification results of OrthoFinder for the pan-gene atlas have a minimal effect on our main conclusions. The major advances enabled by the pan-gene atlas, including the detection of 23,296 additional expressed genes and the identification of 74 non-Chinese Spring candidate genes, primarily result from the inclusion of diverse gene models. Each candidate gene is a member of an orthogroup, representing a group of homologous genes. The classification of genes into assigned and unassigned categories was therefore designed to elucidate homologous relationships within these orthogroups. This is the key reason why the pan-gene atlas substantially improves the efficiency of candidate gene discovery. Additionally, analyses of eQTLs, breeding selection, and regulatory networks focused on population-level patterns. Thus, the misclassification of unassigned genes does not affect our major conclusions. The pan-gene atlas offers a lightweight, traceable and user-friendly resource that greatly reduces the limitations of single-reference approaches while conserving both time and computational resources.
Orthogroups frequently harbor highly similar transcripts, leading to ambiguous multi-mapping of short RNA-seq reads and making it difficult to assign reads to specific transcripts. Recent studies on exploring expression diversity in wheat and barley conducted gene annotation for each individual line and therefore provided more accurate results72,73. However, in population-scale transcriptomic studies involving hundreds of wheat landraces and cultivars, conducting gene annotation for each line is nearly impossible at the current stage. For these reasons, selecting the longest or most highly expressed transcript as the representative sequence is widely adopted, as it reduces redundancy, improves computational efficiency, and facilitates downstream eQTL and association analyses.
Recently, Cheng et al.5 demonstrated that modern wheat cultivars are derived primarily from two of the seven ancestral groups, AG2 and AG5. The ancestral composition of our wheat panel was previously characterized by Cheng et al.5, who classified the accessions using genotype data from Niu et al.6. In our panel, MCC varieties align with AG2, USMC varieties align with AG5, and landraces align mainly with AG1, AG3, and AG4, with a minor fraction distributed across other groups. Thus, the 328 accessions used in this study collectively represent five of the seven ancestral groups (AG1–AG5), capturing broad ancestral diversity. To improve alignment accuracy and representation of ancestral variation, we incorporated 44 reference genomes into our pan-reference, several of which correspond to key ancestral groups in our panel. For instance, Chinese Spring and Norin61 represent AG1, representing landrace-associated haplotypes; Mace, Lancer, Jagger, CDC Landmark, and CDC Stanley represent AG5, facilitating analysis of USMC lines; and ArinaLrFor, SY Mattis, and Julius align with AG2, representing MCC diversity. The inclusion of these representative genomes increases mapping fidelity and enables more comprehensive characterization of haplotype diversity across the panel. In addition, the integration of diploid and tetraploid progenitors involved in wheat evolution further strengthens the ability of our pan-gene atlas to capture genetic variation across wheat germplasms, including lineage-specific and introgressed alleles that are often overlooked in modern breeding.
Introgression in the wheat genome has been intensively studied in recent years because of the decreasing cost of next-generation sequencing technologies6,7,33,74, but the genome-wide pattern of gene expression for introgression has rarely been studied24,25. Our population-level gene expression analysis revealed a lack of consistency between the expression of introgressed genes and the goal of breeding improvement. Genes related to environmental stress or disease resistance are often favored by breeders. The corresponding chromosomal segments are introduced into common wheat through hybridization and tend to be transcriptionally activated, thereby increasing stress tolerance and disease resistance. In contrast, genes involved in basic cellular processes are frequently downregulated or silenced in introgression lines, possibly because of functional redundancy with the native wheat genome. These changes might be achieved through trans-eQTLs located on other wheat chromosomes. We propose that the optimal expression of introgressed genes depends on the presence of favorable alleles at these trans-eQTL loci. In other words, our calculations suggested that the gene expression activity of introgressed alleles depends on the genetic background, which is a phenomenon frequently observed in distant hybrid breeding75,76. Our results provide clues for designing future breeding plans by selecting hybrid parents with favorable alleles at those trans-eQTLs. eQTLs for introgressed genes were analyzed both at the population level and only among lines with that introgression. Because introgressed segments are present only in a subset of samples, population-level eQTLs likely exhibit pronounced population structure. In a comparison, eQTLs computed using only introgression lines are distributed more uniformly across the genome. Therefore, eQTLs for introgressed segments calculated using all samples should be interpreted with caution.
The pan-gene atlas revealed that 22% of the transcripts expressed in seedling tissues are not encoded by the Chinese Spring reference. In total, more than twenty thousand non-Chinese Spring transcripts were detected in our panel of 327 wheat lines. Although numerous RNA-seq and GWASs have been conducted in wheat, non-Chinese Spring genes have always been ignored in previous studies6,24,71. Because GWASs rely solely on SNP information present in the Chinese Spring genome to associate with phenotypes, it can identify only candidate genes located within the Chinese Spring genome. In contrast, by using TWASs, SMR, and correlation analyses between gene expression and phenotypes, we were able to identify candidate genes that are not present in the Chinese Spring genome. The use of transcriptomic data enabled us to identify a greater number of non-Chinese Spring candidate genes. However, as the non-Chinese Spring genes could not be assigned to the same orthogroups as the Chinese Spring genes were, we did not perform synteny analysis for them with Chinese Spring. Instead, we approximated their positions using the most significant eQTL signals, with the accuracy of this approach potentially reaching up to 65.19% (Supplementary Fig. 8a, b). Nevertheless, this strategy only partially addresses this limitation, and further methodological improvements are needed in future studies to identify additional non-Chinese Spring candidate genes. Given that the RNA-seq data used in this study were collected at the seedling stage, the detection of expression changes associated with developmental traits was limited. However, introgressed genes often exhibit presence–absence variation in expression, which was effectively captured in this study. On the basis of an integrative prediction pipeline similar to that used in our previous work24, we obtained candidate genes for agronomic traits and disease resistance traits, which included 71 non-Chinese Spring genes for a set of 34 agronomic traits, and 3 non-Chinese Spring genes for a set of 8 Bgt isolates. Functional validation using an indexed EMS mutant library revealed that 98 out of 133 homologous candidate genes with at least five mutant lines exhibited significant trait differences between the wild type and mutant, supporting the predictive value of our candidate gene set.
Compared with our previous eQTL study24, in this work, whole-genome resequencing-derived SNP data was employed, which enabled a more comprehensive detection of regulatory elements, including those located in distal intergenic regions. While intergenic SNPs are often excluded in genomic studies74, we found that approximately 2.0% of candidate genes were regulated exclusively by intergenic eQTLs, and that their trait associations were mediated through such loci. These results highlight the importance of non-coding regions in gene regulation and their potential application in marker-assisted selection. Although the Chinese Spring v1.1 reference genome, which was used for variant calling, is based on short-read sequencing and may contain assembly errors, particularly in intergenic regions, it remains the most widely adopted reference in wheat genomics and ensures compatibility with major genotyping platforms such as the 1000 Wheat Exomes Project7 and the Watkins panel5. More than 99% of the resequencing reads were successfully mapped to the Chinese Spring reference (Supplementary Fig. 22), indicating minimal reference bias. Moreover, most identified eQTLs are located in well-assembled genic regions; therefore, the potential assembly errors in Chinese Spring v1.1 do not affect the main conclusions of our study.
Modern breeding involves not only the selection of target genes but also changes in transcriptome profiles at the population level. Approximately 10% of the genes exhibited differential expression between different sub-populations. Differences exist in the direction of changes between the cultivars from China and those from the United States, which is likely attributed to varying breeding objectives and environmental adaptability. While common regulatory changes were detected in genes involved in stress responses and photoperiod regulation (e.g., VRN1, VRN2, and WRKY45)57,58, U.S. cultivars tended to favor genes related to developmental regulation (e.g., TaGRP-2)52, whereas breeders in China favor genes related to disease resistance and yield (e.g., Lr34, TaAGL12)55,77. These trends highlight how breeding objectives have shaped the direction of transcriptomic changes. Moreover, compared with non-DEGs, DEGs exhibited greater genetic differentiation and stronger signatures of selection, a pattern that extended to their regulatory regions. These findings suggest that breeding has exerted a widespread influence on gene regulatory networks. In addition to individual expression changes, breeding altered gene co-expression and regulatory network architecture. The number of co-expression pairs involving active-eQTGs and eGenes was significantly lower in landraces than in cultivars, suggesting that modern breeding has increased network complexity. Future breeding strategies may benefit from considering not only gene-specific selection but also the optimization of broader regulatory networks.
Methods
Wheat germplasm and phenotypes
We selected a panel of 328 Triticum aestivum accessions from previously published whole-genome resequencing dataset comprising 355 accessions6, including 92 modern Chinese cultivars (MCCs), 64 modern United States cultivars (USMCs), and 172 landraces (LRs) from 13 countries worldwide, representing a wide range of genetic diversity (Supplementary Data 1).
A total of 42 phenotypic traits were assessed, encompassing 34 field agronomic traits and 8 seedling-stage resistance traits to different Bgt isolates. The planting and phenotypic measurements of the field agronomic traits were conducted concurrently with those reported in previous studies6, and the phenotypes for 20 of these traits have already been published. These include four grain-related traits: grain length (GL), grain roundness (GRO), grain width (GW), and grain number (GN); seven yield-related traits: thousand-kernel weight (TKW), harvest index (HI), yield per plant (YPP), spikelet number (SPN), awn length (AL), sterile spikelet number (SSN), and biomass per plant (BPP); and nine growth and development traits: anthesis days (AD), heading days (HD), flag leaf length (FLL), flag leaf width (FLW), plant height (PH), peduncle length (PL), stem diameter (SD), tiller number at the jointing stage (TNJS), and tiller number at the seedling stage (TNSS)6.
The 14 other field agronomic trait phenotypes are described in this study, namely, seven grain-related traits: grain filling days (GFD), grain area (GA), grain diameter (GD), grain perimeter (GP), and grain color, which included grain red (GR), grain green (GG), and grain blue (GB); five yield-related traits: yield per head per plant (YHPP), biomass per plant at the jointing stage (BPPJS), biomass per tiller at the jointing stage (BPTJS), biomass per plant at the seedling stage (BPPSS), and biomass per tiller at the seedling stage (BPTSS); and two growth and development traits: life cycle (LC) and days from heading to anthesis (HAD). All 328 common wheat accessions were grown for three consecutive years (2013–2016) in Zhao County, Shijiazhuang, Hebei Province, China (38°05′N, 114°52′E). The accessions were randomly arranged in plots with 110 × 25 cm row and column spacing, each with three independent replicates. Agronomic traits were evaluated using five centrally located plants per plot. The best linear unbiased estimates (BLUEs) were derived using a model with fixed genotype effects and random effects for each year.
We isolated and purified eight Bgt isolates from fields across different provinces in China, designated B040A1, B056A1, B080A1, B094A1, B099A1, B114A2, B132A2 and B138A1 (Supplementary Data 10). We planted the 328 wheat accessions in rectangular trays with three replicates, providing 14 of light at 22 °C and 10 h of darkness at 18 °C. At the one-leaf stage, seedlings of all wheat lines were inoculated with one Bgt isolate. Infection types were recorded 10 days post-inoculation using a 0–4 scale, where 0–2 indicates resistance and 3–4 indicates susceptibility. Higher scores represent stronger disease susceptibility, and the mean value of three individual plants per variety was used for subsequent analysis (Supplementary Fig. 14a)50,78.
RNA-seq sequencing and data preprocessing
A total of 328 wheat lines were grown in trays, with each line having three biological replicates, maintained under a 14-h light and 10-h dark cycle at 22 °C and 18 °C, respectively. Leaf tissues were sampled at the two-week seedling stage, and the leaves from the three biological replicates were averaged for RNA extraction using the FastPure Universal Plant Total RNA Isolation Kit. The extracted RNA was subsequently used to construct libraries via the BGI Optimal Series Dual Module mRNA Library Construction Kit (LR00R96) and sequenced on the DNBSEQ-T7 platform, generating 2 × 150 bp reads. One accession (1699B) was excluded from subsequent analyses because of potential contamination, resulting in a final set of 327 high-quality RNA-seq samples.
In parallel, six rye accessions and seven Thinopyrum ponticum accessions were also grown under identical conditions for RNA sequencing. Additionally, we obtained published RNA data from previous studies39,79, which included leaf samples from 2 Renan samples79 and 3 T. timopheevii samples39 (Supplementary Data 7), all of which were obtained at the 2–3-week-old seedling stage. The software Salmon v1.8.080, which uses pseudoalignment techniques on RNA-seq reads to reference gene models was used to quantify the transcript abundance.
Identification of large introgression events using whole-genome resequencing data
The genotypes of 327 wheat accessions were obtained from a previous study6 in which those wheat accessions were whole-genome-sequenced and genotyped on the basis of the Chinese Spring reference genome (IWGSC RefSeq v1.1). After filtering out variants with a missing rate >25% and heterozygosity >30%, variants with an MAF > 0.05 were retained for this study, resulting in a total of 26,788,626 variants, including 24,744,215 SNPs and 2,044,411 InDels.
To identify large introgressed segments that are not present in Chinese Spring, a genome-wide heatmap of genotype calls was constructed using the ComplexHeatmap R package81,82, with a window size of 1 SNP/Mb, revealing >20 Mb of missing segments. To quantify these deletions, we calculated the missing rate across the genome using a 2 Mb sliding window with a 1 Mb step size, compared the missing rates in deleted versus non-deleted regions, and tested for significance using the two-sided Wilcoxon rank-sum test.
Reference genome model of large introgressed segments
In this study, four large introgressed segments previously reported39,79,83,84 in wheat were analyzed using the genomes of corresponding donor species as reference models. We used the genomes of Secale cereale L.83, the French cultivar Renan (instead of Ae. markgrafii, because the terminal end of Renan 2D has been reported to possibly originate from the introgression of Ae. markgrafii without a published genome)79, Thinopyrum elongatum (instead of Th. ponticum, because the genome of Th. ponticum has not been published, and Th. elongatum is closely related to Th. ponticum)84, and Triticum timopheevii39. These genomes were merged with the Chinese Spring reference genome (IWGSC RefSeq v1.1). Afterward, on the basis of the merged gene models, the gene expression levels of 328 wheat samples were calculated using Salmon v1.8.080. The number of expressed genes with expression (transcripts per million, TPM > 0.5) was then counted in 10 Mb windows with a 2.5 Mb step along each chromosome.
Construction of the pan-gene atlas
To complement the identification of large introgressed fragments (>20 Mb), which could be readily detected on the basis of genotype loss rates, we sought to capture smaller introgressed regions that could not be accurately assessed using the Chinese Spring gene model alone. To this end, we integrated gene models from 44 published Triticeae genomes. A key selection criterion for these genomes was the availability of RNA-seq-supported gene annotations.
These included hexaploid wheat genomes such as Chinese Spring (IWGSC RefSeq v1.1, 2n = 6x = 42, AABBDD)26, Kenong 920485, Aikang 5886, and 9 cultivars from The 10+ Wheat Genomes Project (ArinaLrFor, Jagger, Julius, LongReach Lancer, CDC Landmark, Mace, Norin 61, CDC Stanley, SY Mattis)31,72, the South African bread wheat cultivar Kariega87, the Tibetan semi-wild wheat (Triticum aestivum ssp. tibetanum Shao) accession Zang181788. Among 17 representative Chinese cultivars with high-quality genome assemblies, four (XY6, AMN, JM22, and ZM16) were included on the basis of the availability of RNA-seq-supported annotations89. We also incorporated the synthetic hexaploid wheat-derived cultivar Chuanmai 104 and the backbone breeding parent line Zhou8425B90,91.
In addition, we included tetraploid genomes (T. turgidum ssp. dicoccoides and T. turgidum L. ssp. durum, 2n = 4x = 28, AABB)92,93, along with diploid genomes such as T. urartu (2n = 2x = 14, AA)94, 3 T. monococcum (TA10622, TA299, PI 306540, 2n = 2x = 14, AA)95,96, 6 genomes of Aegilops speltoides relatives (TS01, Ae. bicornis, Ae. longissimi, Ae. searsii, Ae. sharonensis, Ae. speltoides (Y2032), 2n = 2x = 14, SS)97,98 and 8 Aegilops tauschii genomes (AL8/78, AY17, AY61, T093, XJ02, TA1675, TA2576, TA10171, 2n = 2x = 14, DD)99,100,101.
Furthermore, we incorporated four gene models from previously characterized large introgressed segments: Secale cereale L. (rye, 2n = 2x = 14, RR, chr1R: 0–280 Mb)83, the French cultivar Renan (2n = 6x = 42, AABBDD, chr2D: 570–635 Mb, used as a proxy for Ae. markgrafii because of the absence of its reference genome)79, Thinopyrum elongatum (2n = 2x = 14, EE, chr3E: 500–676 Mb) (a close relative of Th. ponticum)84, and T. timopheevii (2n = 4x = 28, AtAtGG, chr5G: 435–485 Mb)39.
The pan-gene atlas was constructed in three parts. The first part included all the high-confidence genes from Chinese Spring. The second part included high-confidence genes from the four large introgressed segments listed above. The third part consisted of non-redundant, high-confidence genes from the remaining 19 hexaploid, 2 tetraploid, and 18 diploid genomes.
To remove redundant genes, we used OrthoFinder to cluster all genes from the 44 genomes102,103. For orthogroups containing genes from Chinese Spring or from the four large introgressed regions, only the Chinese Spring or introgressed genes were retained, and these orthogroups were excluded from the non-redundant set. For orthogroups lacking Chinese Spring or introgressed genes, the longest transcript was selected as the representative isoform, following previous transcriptomic studies, to minimize redundancy from repetitive sequences and reduce the impact of short-read multi-mapping104,105,106. Therefore, the longest transcript was selected as the representative gene, and its expression level was significantly greater than that of other genes (Supplementary Note 4, Supplementary Fig. 23). The genes from all three parts were then merged to create the final pan-gene atlas. Using this pan-gene atlas as the reference, we employed Salmon v1.8.0 to assess the transcript abundance of 328 wheat samples through pseudoalignment80, which was used for all subsequent analyses.
Quantification of gene expression in introgressed segments
To investigate gene expression in introgressed segments, we used OrthoFinder to identify homologous high-confidence genes between Secale cereale L. (rye; chr1R: 0–280 Mb) and Chinese Spring102,103. Genes that are homologous between rye and wheat are considered conserved genes, whereas those that are unique to rye are called specific genes. To assess the relationship between gene sequence similarity and expression conservation, we selected 1RS genes whose alignment lengths exceeded 80% and whose sequence identities were greater than 70% relative to those of their wheat homologs.
We performed differential expression analysis of chromosome 1RS genes between the 1RS.1AL and 1RS.1BL introgression lines and the 6 rye accessions. Transcript abundances were first log2 transformed (TPM + 0.1), after which the expression matrix was normalized using the normalize.quantiles.robust method. Differential expression analysis was conducted using the two-sided Wilcoxon rank-sum test107, comparing gene expression between the 1RS.1AL line and rye 1RS and between the 1RS.1BL line and rye 1RS. The log2-fold change (log2FC) was calculated as the difference in average gene expression between the two groups. Genes with adjusted p values (Padj) < 0.05 and absolute log2FC > 1 were defined as differentially expressed genes (DEGs). The same analytical pipeline was applied to the other introgressed gene sets. We annotated genes from rye, T. elongatum and T. timopheevii using InterProScan (v5.66–98.0)108. Gene Ontology (GO) enrichment analysis of the DEGs was performed using TBtools109.
Characterization of rye genes
To assess the presence and expression of rye-derived resistance genes and genes involved in fundamental biological processes in introgression lines and the donor genome, three resistance-related genes (SECCE1Rv1G0002890, SECCE1Rv1G0003770, and SECCE1Rv1G0008680) and three genes related to fundamental biological activities (SECCE1Rv1G00021710, SECCE1Rv1G00017850, and SECCE1Rv1G0000360) were selected for genomic DNA validation and reverse transcription quantitative PCR (RT‒qPCR) analysis. Gene-specific primers were designed the basis of both the genomic and coding sequences of the six target genes (Supplementary Data 8). Genomic DNA amplification was performed using ten rye accessions and ten 1RS.1BL introgression lines. For RT‒qPCR analysis, three rye accessions and three 1RS.1BL introgression lines were used (Supplementary Data 9). The wheat TaTublin gene was used as the endogenous control, and relative expression levels were calculated using the comparative CT method110.
Detection of eQTLs
Gene expression data were log2-transformed and normalized using robust quantile normalization in R. To account for hidden confounding factors, we applied the probabilistic estimation of expression residuals (PEER) method111 and used the resulting residuals to assess the genetic regulation of gene expression. For eQTL mapping, we associated PEER-corrected expression residuals with filtered SNP genotypes using the MatrixEQTL R package (useModel = modelLINEAR)112. The first five principal components (PCs) of the SNP matrix, representing population structure, were included as covariates. A false discovery rate (FDR) threshold of <1 × 10–5 was used to define significant eQTLs in the population-level analysis, whereas a less stringent threshold (FDR < 1 × 10–2) was adopted for the introgression lines because of their smaller sample size.
We classified SNPs associated with gene expression into three categories. First, SNPs were annotated using SnpEff (v.5.0e)113 and classified as intergenic SNPs or genic SNPs. Genic SNPs were further classified the basis of gene activity: if a gene was expressed (TPM > 0.5) in more than 5% of samples, it was considered active, and the corresponding SNPs were labeled as active SNPs; otherwise, genes and their SNPs were classified as inactive. We subsequently merged the significant SNPs of the three types using strict criteria. Intergenic SNPs were merged on the basis of the criteria of continuous SNPs within <100 kb, a minimum of three SNPs, and an LD > 0.2, retaining the most significant SNP as the intergenic-eQTL. For inactive SNPs, the same merging strategy as for intergenic SNPs was applied, and the most significant SNP from each cluster was retained as the inactive-eQTG. Active SNPs were processed in two steps: (1) High-impact SNPs (as defined by SnpEff) were retained as active-eQTGs. (2) For non-high-impact SNPs, we calculated the Spearman correlation coefficient (SCC) between the expression levels of the SNP-harbored gene and its target eGene. SNPs located within genes and significantly associated with eGenes (|SCC| > 0.3, p value < 0.05) were considered as regulatory. The most significant SNP was retained as the active-eQTG. Finally, all intergenic-eQTLs, inactive-eQTGs, and active-eQTGs are collectively referred to as eQTLs. eQTLs located on the same chromosome as their associated eGene were classified as cis-eQTLs, whereas those located on different chromosomes were classified as trans-eQTLs. This classification was applied only to eGenes from the Chinese Spring reference genome.
Synteny analysis of alien introgression segments
Synteny analysis was conducted using JCVI114 to investigate the chromosomal correspondence between alien introgression segments and the Chinese Spring reference genome. Gene sequences from three introgressed segments (rye: (chr1R: 0–280 Mb), Renan: (chr2D: 570–635 Mb), and T. timopheevii: (chr5G: 435–485 Mb)) were subjected to pairwise synteny searches against all annotated genes in the Chinese Spring genome using JCVI114 with default parameters. The resulting paired gene files were subsequently used to generate macrosynteny visualizations, and the default settings were used.
GWAS
A GWAS was performed using a linear mixed model that addressed both population structure and kinship for all 34 field agronomic traits and 8 seedling-stage Bgt isolate phenotypes, employing the ‘--mlma’ parameter in GCTA (v1.94.1)115. The first five principal components were used to control for population structure. A kinship matrix was generated on the basis of a set of independent SNPs to capture relatedness among individuals. If two consecutive significant SNPs were located less than 2 Mb apart, they were grouped into a single QTL. The QTL interval was defined by SNPs with a p value < 1 × 10–4, and the most significant SNP within the interval was designated as the lead variant. QTLs were retained only if the lead SNP had a p value < 1 × 10–6; otherwise, they were excluded. Genes harboring significant SNPs within each QTL interval were identified as candidate genes.
TWAS
Transcriptome-wide association studies (TWASs) provide a framework for identifying significant cis genetic variant correlations between gene expression and phenotype. For this research, Fusion software was utilized to carry out the TWAS (http://gusevlab.org/projects/fusion/)44. The program requires the computation of gene expression weights, reflecting the pre-modeled relationships between SNPs and gene expression levels, which are then integrated with GWAS data to estimate the associations between genes and phenotypic traits.
For Chinese Spring genes, we computed expression weights using SNPs within a 2 Mb window centered on each gene and their expression levels across the population. For non-Chinese Spring genes, the genomic positions of their most significant eQTL signals in the Chinese Spring reference genome were used as proxies. SNPs and expression levels within 2 Mb of the eQTL peak were used to model expression weights. Heritability calculations were carried out using GCTA (v1.94.1)115, and expression weights were derived from methods including top1, blup, lasso, and enet. We subsequently extracted the SNP data from the GWAS results of 42 traits, including A1 (first allele) and A2 (second allele), and computed the Z-scores with the formula:
β represents the estimated effect size of the allele on the trait, and SE denotes the standard error of the effect size estimate.The FUSION.test.R script from the FUSION program was used to separately analyze each GWAS summary dataset and expression weight file to predict candidate genes (http://gusevlab.org/projects/fusion/). Genes a TWAS p value < 1 × 10–3 were considered candidate genes.
SMR
We performed a summary data-based Mendelian randomization analysis (SMR) to investigate the association between gene expression and trait variation, utilizing summary-level data from our eQTL mapping study and GWAS results for 34 field agronomic traits and 8 seedling-stage Bgt isolate phenotypes, employing GCTA (v1.94.1)115. For the physical location information of non-Chinese Spring genes, we approximated the positions of the strongest eQTL signals from Chinese Spring as the physical locations. The summary-level statistics of these two GWAS datasets were analyzed using the SMR commands ‘-cis-wind 10,000’ for cis-eQTL and ‘--trans-wind 5000’ for trans-eQTL whose SMR p value was <1 × 10–4 were designated as candidate genes.
Validation of candidate genes using the EMS mutant library
To validate candidate genes, we identified homologs in the Kenong 9204 genome with ≥ 98% sequence identity and ≥90% coverage relative to the candidate genes. Homologous genes with at least five non-synonymous mutants in the indexed Kenong 9204 EMS library were selected for further analysis51. For each gene, a two-tailed Student’s t test was conducted between the mutant (n > 5) and wild-type samples (n = 15). To account for background variation, an equal number of control genes without mutations were randomly selected, and the same analysis was applied. We evaluated the effects of candidate gene mutations on seven agronomic traits, namely, spikelet number (SPN), grain number (GN), grain area (GA), grain perimeter (GP), grain length (GL), grain width (GW), and thousand-kernel weight (TKW). Given the high Pearson correlation coefficient among the 34 field agronomic traits (Supplementary Fig. 12), mutations in candidate genes affecting one trait often influence multiple traits. Thus, candidate genes associated with these agronomic traits were tested across all seven selected traits.
Detection of DEGs between cultivars and landraces
We identified differentially expressed genes (DEGs) between landraces (LRs) and two cultivar groups (MCCs and USMCs) using the two-sided Wilcoxon rank-sum test. The expression levels were normalized as described above. DEGs were defined on the basis of an adjusted p value < 0.05 and an absolute log2-fold change >1, where log2-fold change represents the ratio of the mean expression between two populations. DEGs were categorized into four types across the eight models, with the landrace gene used as a reference. The first category (M1 and M2) includes genes whose regulation is inconsistent between MCCs and USMCs relative to landraces; the second category (M3 and M4) includes genes whose regulation is consistent. The third category (M5 and M6) consists of genes that are differentially expressed between MCCs and landraces, with no significant differences for USMCs and landraces. The fourth category (M7 and M8) consists of that are differentially expressed between USMCs and landraces, with no significant differences for MCCs and landraces. Finally, Gene Ontology (GO) enrichment analysis was conducted for the 8 models of DEGs using TBtools109.
Dimensionality reduction of population expression
We used population-specific DEGs identified from the MCC, USMC, and LR comparisons to perform dimensionality reduction analysis. The expression values were normalized as described above. UMAP was applied using the umap function from the R package uwot to visualize expression divergence across populations116.
Population genetics analysis
Given the large LD distance and dense SNP coverage, we filtered SNPs on the basis of the criteria from published work6, retaining 5,749,696 SNPs/InDels for principal component analysis using PLINK v1.9117. The genetic differentiation (FST) between sub-populations (MCCs vs. LRs, USMCs vs. LRs, USMCs vs. MCCs) was calculated using a 20-kb sliding window and a step size of 10 kb with VCFtools (v0.1.16)118. To evaluate the genetic differentiation of DEGs, non-DEGs, and regulatory regions under modern breeding, we compared the FST values for these categories. To analyze the relationships between the signals of DEGs and the most significant eQTL signals with FST value, we paired the signals of DEGs and the most significant eQTLs signals with their corresponding FST value, sorting them by FST value. A custom script119 was used to split the data into ten bins, calculating the mean and standard error of the signals of the DEGs and top eQTL signals were calculated as the FST value changed.
Detection of selective sweeps between landraces and cultivars
We applied the XP-CLR method (https://github.com/hardingnj/xpclr), a Python-based composite likelihood approach120, to identify selective sweeps during modern wheat breeding. For this analysis, landraces were considered the reference group, with MCCs and USMCs acting as the query groups. We scanned for selective sweeps with a step size of 10 kb and a 20 kb sliding window across each chromosome (--size 20,000; --step 10,000). We extracted XP-CLR scores for genomic regions containing DEGs to evaluate whether they were under selection during breeding. Additionally, XP-CLR scores were obtained for regions harboring the most significant eQTLs of DEGs to assess potential selection on regulatory regions. Sites with XP-CLR scores equal to zero were excluded. The remaining scores were ranked and divided into 20 equal bins. For each bin, the mean and standard error of scores for both DEGs and eQTLs were calculated using a custom script119.
Construction of gene co-expression regulatory networks
Co-expression regulatory networks for each sub-population were constructed based on the regulatory pairs of active-eQTGs and their corresponding eGenes identified at the population level in the study. Active-eQTG–eGene pairs were defined as those showing significant expression correlations across all samples (|SCC| > 0.3 and p value < 0.05). For each active-eQTG–eGene pair, expression correlations were recalculated within each sub-populations (MCCs, USMCs and LRs). Pairs with |SCC| > 0.3 and p value < 0.05 within a sub-population were considered co-expressed. To ensure comparability across sub-populations, the same set of active-eQTG–eGene pairs identified at the population level was used for all sub-population analyses.
For constructing the regulatory networks of agronomic trait–related candidate genes, eGenes that were also differentially expressed (LRs vs. MCCs and LRs vs. USMCs) were selected, and their correlations with corresponding active-eQTGs were computed within each sub-population. For the powdery mildew resistance network, due to the limited number of candidate genes, the eGenes regulated by active-eQTGs at the population level were considered, and their correlations were recalculated across sub-populations. The resulting co-expression relationships were visualized in Gephi (https://gephi.org/).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA-seq data generated in this study have been deposited in the Genome Sequence Archive (GSA)121 at the National Genomics Data Center122, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession CRA022107 (328 wheat accessions), CRA025944 (6 rye accessions), and CRA022106 (7 Th. ponticum accessions). The results, including the sequence of pan-gene atlas, orthologous genes and eQTLs, have been uploaded to the Figshare database [https://figshare.com/s/ee89a74381fea08c4c4e]. Source data are provided with this paper.
References
Tadesse, W. et al. Genetic gains in wheat breeding and its role in feeding the world. Crop Breed. Genet. Genom. 1, e190005 (2019).
Tester, M. & Langridge, P. Breeding technologies to increase crop production in a changing world. Science 327, 818–822 (2010).
Lopes, M. S. et al. Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J. Exp. Bot. 66, 3477–3486 (2015).
Reif, J. C. et al. Wheat genetic diversity trends during domestication and breeding. Theor. Appl. Genet. 110, 859–864 (2005).
Cheng, S. et al. Harnessing landrace diversity empowers wheat breeding. Nature 632, 823–831 (2024).
Niu, J. et al. Whole-genome sequencing of diverse wheat accessions uncovers genetic changes during modern breeding in China and the United States. Plant Cell 35, 4199–4216 (2023).
He, F. et al. Exome sequencing highlights the role of wild-relative introgression in shaping the adaptive landscape of the wheat genome. Nat. Genet. 51, 896–904 (2019).
Tiwari, V. K. et al. SNP Discovery for mapping alien introgressions in wheat. BMC Genomics 15, 273 (2014).
Delibes, A. et al. Hessian fly-resistance gene transferred from chromosome 4Mv of Aegilops ventricosa to Triticum aestivum. Theor. Appl. Genet. 94, 858–864 (1997).
Rani, K. et al. A novel leaf rust resistance gene introgressed from Aegilops markgrafii maps on chromosome arm 2AS of wheat. Theor. Appl. Genet. 133, 2685–2694 (2020).
Chen, G. et al. Molecular cytogenetic identification of a novel dwarf wheat line with introgressed Thinopyrum ponticum chromatin. J. Biosci. 37, 149–155 (2012).
Grosso, V., Farina, A., Gennaro, A., Giorgi, D. & Lucretti, S. Flow sorting and molecular cytogenetic identification of individual chromosomes of Dasypyrum villosum L. (H. villosa) by a single DNA probe. PLoS One 7, e50151 (2012).
Rabinovich, S. V. Importance of wheat-rye translocations for breeding modern cultivar of Triticum aestivum L. Euphytica 100, 323–340 (1998).
Molnár-Láng, M., Ceoloni, C. & Doležel, J. Alien Introgression in Wheat. Springer ISBN: 978-3-319-23493-9, Preface (2015).
Chen, F. et al. Molecular characterization of vernalization and response genes in bread wheat from the Yellow and Huai Valley of China. BMC Plant Biol. 13, 199 (2013).
Gauley, A. et al. Photoperiod-1 regulates the wheat inflorescence transcriptome to influence spikelet architecture and flowering time. Curr. Biol. 34, 2330–2343.e2334 (2024).
Boden, S. A. et al. Ppd-1 is a key regulator of inflorescence architecture and paired spikelet development in wheat. Nat. Plants 1, 14016 (2015).
Gauley, A. & Boden, S. A. Stepwise increases in FT1 expression regulate seasonal progression of flowering in wheat (Triticum aestivum). New Phytol. 229, 1163–1176 (2021).
Li, C. & Dubcovsky, J. Wheat FT protein regulates VRN1 transcription through interactions with FDL2. Plant J. 55, 543–554 (2008).
Li, J. et al. TaNAC100 acts as an integrator of seed protein and starch synthesis exerting pleiotropic effects on agronomic traits in wheat. Plant J. 108, 829–840 (2021).
He, X. et al. The nitrate-inducible NAC transcription factor TaNAC2-5A controls nitrate response and increases wheat yield. Plant Physiol. 169, 1991–2005 (2015).
Zhang, Z. et al. Insights into salinity tolerance in wheat. Genes 15, 573 (2024).
Jiang, W. et al. Conservation and divergence of the TaSOS1 gene family in salt stress response in wheat (Triticum aestivum L. Physiol. Mol. Biol. 27, 1245–1260 (2021).
He, F. et al. Genomic variants affecting homoeologous gene expression dosage contribute to agronomic trait variation in allopolyploid wheat. Nat. Commun. 13, 826 (2022).
Coombes, B., Lux, T., Akhunov, E. & Hall, A. Introgressions lead to reference bias in wheat RNA-seq analysis. BMC Biol. 22, 56 (2024).
International Wheat Genome Sequencing Consortium (IWGSC). Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361, eaar7191 (2018).
Ramírez-González, R. H. et al. The transcriptional landscape of polyploid wheat. Science 361, eaar6089 (2018).
Li, G. et al. A high-quality genome assembly highlights rye genomic characteristics and agronomically important genes. Nat. Genet. 53, 574–584 (2021).
Heuberger, M. et al. Analysis of a global wheat panel reveals a highly diverse introgression landscape and provides evidence for inter-homoeologue chromosomal recombination. Theor. Appl. Genet. 137, 236 (2024).
Friebe, B., Zeller, F. J. & Kunzmann, R. Transfer of the 1BL/1RS wheat-rye-translocation from hexaploid bread wheat to tetraploid durum wheat. Theor. Appl. Genet. 74, 423–425 (1987).
Walkowiak, S. et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 588, 277–283 (2020).
Keilwagen, J. et al. Detecting major introgressions in wheat and their putative origins using coverage analysis. Sci. Rep. 12, 1908 (2022).
Zhou, Y. et al. Triticum population sequencing provides insights into wheat adaptation. Nat. Genet. 52, 1412–1422 (2020).
Subramanian, S. & Kumar, S. Gene expression intensity shapes evolutionary rates of the proteins encoded by the vertebrate genome. Genetics 168, 373–381 (2004).
Crow, M., Suresh, H., Lee, J. & Gillis, J. Coexpression reveals conserved gene programs that co-vary with cell type across kingdoms. Nucleic Acids Res. 50, 4302–4314 (2022).
Martin, T. & Fraser, H. B. Comparative expression profiling reveals widespread coordinated evolution of gene expression across eukaryotes. Nat. Commun. 9, 4963 (2018).
Feuillet, C. et al. Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc. Natl. Acad. Sci. 100, 15253–15258 (2003).
Hurni, S. et al. Rye Pm8 and wheat Pm3 are orthologous genes and show evolutionary conservation of resistance function against powdery mildew. Plant J. 76, 957–969 (2013).
Grewal, S. et al. Chromosome-scale genome assembly of bread wheat’s wild relative Triticum timopheevii. Sci. Data 11, 420 (2024).
Hajjar, R. & Hodgkin, T. The use of wild relatives in crop improvement: a survey of developments over the last 20 years. Euphytica 156, 1–13 (2007).
Crespo-Herrera, L. A., Garkava-Gustavsson, L. & Åhman, I. A systematic review of rye (Secale cereale L.) as a source of resistance to pathogens and pests in wheat (Triticum aestivum L.). Hered 154, 14 (2017).
Sun, G. et al. A role for heritable transcriptomic variation in maize adaptation to temperate environments. Genome Biol. 24, 55 (2023).
Mukaka, M. M. Statistics corner: A guide to appropriate use of correlation coefficient in medical research. Malawi Med. J. 24, 69–71 (2012).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Keller, B., Wicker, T. & Krattinger, S. G. Advances in Wheat and Pathogen Genomics: Implications for Disease Control. Annu. Rev. Phytopathol. 56, 67–87 (2018).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
Wang, C. et al. A superior allele of the wheat gene TaGL3.3-5B, selected in the breeding process, contributes to seed size and weight. Theor. Appl. Genet. 135, 1879–1891 (2022).
Sánchez-Martín, J. et al. Wheat Pm4 resistance to powdery mildew is controlled by alternative splice variants encoding chimeric proteins. Nat. Plants 7, 327–341 (2021).
Xie, J. et al. A rare single nucleotide variant in Pm5e confers powdery mildew resistance in common wheat. New Phytol. 228, 1011–1026 (2020).
Wang, D. et al. Boosting wheat functional genomics via an indexed EMS mutant library of KN9204. Plant Commun. 4, 100593 (2023).
Xiao, J. et al. O-GlcNAc-mediated interaction between VER2 and TaGRP2 elicits TaVRN1 mRNA accumulation during vernalization in winter wheat. Nat. Commun. 5, 4572 (2014).
Zhao, T. et al. Characterization and expression of 42 MADS-box genes in wheat (Triticum aestivum L.). Mol. Genet. Genomics 276, 334–350 (2006).
Li, K. et al. Interactions between SQUAMOSA and SHORT VEGETATIVE PHASE MADS-box proteins regulate meristem transitions during wheat spike development. Plant Cell 33, 3621–3644 (2021).
Lagudah, E. S. et al. Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor. Appl. Genet. 119, 889–898 (2009).
Sharma, N. et al. A Flowering Locus C homolog is a vernalization-regulated repressor in brachypodium and is cold regulated in wheat. Plant Physiol. 173, 1301–1315 (2017).
Dubcovsky, J. et al. Effect of photoperiod on the regulation of wheat vernalization genes VRN1 and VRN2. Plant Mol. Biol. 60, 469–480 (2006).
Gupta, S. et al. Deciphering genome-wide WRKY gene family of Triticum aestivum L. and their functional role in response to Abiotic stress. Genes Genom. 41, 79–94 (2019).
Li, C. et al. Wheat VRN1, FUL2 and FUL3 play critical and redundant roles in spikelet development and spike determinacy. Development 146, dev175398 (2019).
Yan, L. et al. The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc. Natl. Acad. Sci. USA 103, 19581–19586 (2006).
Trevaskis, B., Hemming, M. N., Dennis, E. S. & Peacock, W. J. The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci. 12, 352–357 (2007).
Entz, M. H. & Fowler, D. B. Agronomic performance of winter versus spring wheat. Agron. J. 83, 527–532 (1991).
Watson, D. J., Thorne, G. N. & French, S. A. W. Analysis of growth and yield of winter and spring wheats. Ann. Bot. 27, 1–22 (1963).
Herrera-Foessel, S. A. et al. Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor. Appl. Genet. 127, 781–789 (2014).
Shao, A. et al. The Auxin Biosynthetic TRYPTOPHAN AMINOTRANSFERASE RELATED TaTAR2.1-3A Increases Grain Yield of Wheat. Plant Physiol. 174, 2274–2288 (2017).
Mizuno, N. et al. Loss-of-Function Mutations in Three Homoeologous PHYTOCLOCK 1 Genes in Common Wheat Are Associated with the Extra-Early Flowering Phenotype. PLoS One 11, e0165618 (2016).
Liu, G. et al. Virus-induced gene silencing identifies an important role of the tarsr1 transcription factor in starch synthesis in bread wheat. Int. J. Mol. Sci. 17, 1557 (2016).
Liu, J. et al. GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat. Plants 3, 17043 (2017).
Tian, X. et al. Molecular Mapping of Reduced Plant Height Gene Rht24 in Bread Wheat. Front. Plant Sci. 8, 1379 (2017).
Hu, J. et al. A barley stripe mosaic virus-based guide RNA delivery system for targeted mutagenesis in wheat and maize. Mol. Plant Pathol. 20, 1463–1474 (2019).
Zhao, P. et al. Integration of genome-wide association study, linkage analysis, and population transcriptome analysis to reveal the TaFMO1-5B modulating seminal root growth in bread wheat. Plant J. 116, 1385–1400 (2023).
White, B. et al. De novo annotation reveals transcriptomic complexity across the hexaploid wheat pan-genome. Nat. Commun. 16, 8538 (2025).
Guo, W. et al. A barley pan-transcriptome reveals layers of genotype-dependent transcriptional complexity. Nat. Genet. 57, 441–450 (2025).
Hao, C. et al. Resequencing of 145 Landmark Cultivars Reveals Asymmetric Sub-genome Selection and Strong Founder Genotype Effects on Wheat Breeding in China. Mol. Plant 13, 1733–1751 (2020).
Coombes, B. et al. Whole-genome sequencing uncovers the structural and transcriptomic landscape of hexaploid wheat/Ambylopyrum muticum introgression lines. Plant Biotechnol. J. 21, 482–496 (2023).
Simonov, A. V., Pshenichnikova, T. A. & Lapochkina, I. F. Genetic analysis of the traits introgressed from Aegilops speltoides Tausch. to bread wheat and determined by chromosome 5A genes. Russ. J. Genet. 45, 799–804 (2009).
Wang, X. et al. Transcriptome analysis of tomato flower pedicel tissues reveals abscission zone-specific modulation of key meristem activity genes. PLoS One 8, e55238 (2013).
Xie, J. et al. A biGWAS strategy reveals the genetic architecture of the interaction between wheat and Blumeria graminis f. sp. tritici[J]. bioRxiv, 2025.04.09.647224 (2025).
Aury, J. M. et al. Long-read and chromosome-scale assembly of the hexaploid wheat genome achieves high resolution for research and breeding. GigaScience 11, giac034 (2022).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gu, Z. Complex heatmap visualization. iMeta 1, e43 (2022).
Rabanus-Wallace, M. T. et al. Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat. Genet. 53, 564–573 (2021).
Wang, H. et al. Horizontal gene transfer of Fhb7 from fungus underlies Fusarium head blight resistance in wheat. Science 368, eaba5435 (2020).
Shi, X. et al. Comparative genomic and transcriptomic analyses uncover the molecular basis of high nitrogen-use efficiency in the wheat cultivar Kenong 9204. Mol. Plant 15, 1440–1456 (2022).
Jia, J. et al. Genome resources for the elite bread wheat cultivar Aikang 58 and mining of elite homeologous haplotypes for accelerating wheat improvement. Mol. Plant 16, 1893–1910 (2023).
Athiyannan, N. et al. Long-read genome sequencing of bread wheat facilitates disease resistance gene cloning. Nat. Genet. 54, 227–231 (2022).
Guo, W. et al. Origin and adaptation to high altitude of Tibetan semi-wild wheat. Nat. Commun. 11, 5085 (2020).
Jiao, C. et al. Pan-genome bridges wheat structural variations with habitat and breeding. Nature 637, 384–393 (2025).
Liu, Z. et al. Chromosome-level assembly of the synthetic hexaploid wheat-derived cultivar Chuanmai 104. Sci. Data 11, 670 (2024).
Li, G. et al. Genomic analysis of Zhou8425B, a key founder parent, reveals its genetic contributions to elite agronomic traits in wheat breeding. Plant Commun. 6, 101222 (2025).
Avni, R. et al. Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357, 93–97 (2017).
Maccaferri, M. et al. Durum wheat genome highlights past domestication signatures and future improvement targets. Nat. Genet. 51, 885–895 (2019).
Ling, H.-Q. et al. Genome sequence of the progenitor of wheat A subgenome Triticum urartu. Nature 557, 424–428 (2018).
Ahmed, H. I. et al. Einkorn genomics sheds light on history of the oldest domesticated wheat. Nature 620, 830–838 (2023).
Wang, X. et al. A near-complete genome sequence of einkorn wheat provides insight into the evolution of wheat A subgenomes. Plant Commun. 5, 100768 (2024).
Li, L.-F. et al. Genome sequences of five Sitopsis species of Aegilops and the origin of polyploid wheat B subgenome. Mol. Plant 15, 488–503 (2022).
Yang, Y. et al. Genome sequencing of Sitopsis species provides insights into their contribution to the B subgenome of bread wheat. Plant Commun. 4, 100567 (2023).
Luo, M.-C. et al. Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature 551, 498–502 (2017).
Zhou, Y. et al. Introgressing the Aegilops tauschii genome into wheat as a basis for cereal improvement. Nat. Plants 7, 774–786 (2021).
Cavalet-Giorsa, E. et al. Origin and evolution of the bread wheat D genome. Nature 633, 848–855 (2024).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Pozo, F., Rodriguez, J. M., Martínez Gómez, L., Vázquez, J. & Tress, M. L. APPRIS principal isoforms and MANE Select transcripts define reference splice variants. Bioinformatics 38, ii89–ii94 (2022).
Pozo, F., Rodriguez, J. M., Vázquez, J. & Tress, M. L. Clinical variant interpretation and biologically relevant reference transcripts. NPJ Genom. Med. 7, 59 (2022).
Gilbert, D. Longest protein, longest transcript or most expression, for accurate gene reconstruction of transcriptomes? bioRxiv https://doi.org/10.1101/829184 (2019).
Li, Y., Ge, X., Peng, F., Li, W. & Li, J. J. Exaggerated false positives by popular differential expression methods when analyzing human population samples. Genome Biol. 23, 79 (2022).
Mulder N, Apweiler R. InterPro and InterProScan[J]. Comparative Genomics, 59-70 (2007).
Chen, C. et al. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Tang, H. et al. JCVI: A versatile toolkit for comparative genomics analysis. iMeta 3, e211 (2024).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: A tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
He, F. Data binning script. Zenodo. https://doi.org/10.5281/zenodo.17332971 (2025).
Chen, H., Patterson, N. & Reich, D. Population differentiation as a test for selective sweeps. Genome Res. 20, 393–402 (2010).
Chen, T. et al. The genome sequence archive family: Toward explosive data growth and diverse data types. Genomics, Proteom. Bioinforma. 19, 578–583 (2021).
Members & Partners, C.-N. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2023).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2023YFF1000100 to F.H. and 2024YFE0115100 to F.H.), the National Natural Science Foundation of China (32472130 to F.H. and U24A20391 to X.L.), the Yazhouwan National Laboratory project (2310JM01 to S.M.).
Author information
Authors and Affiliations
Contributions
F.H. and H.-Q.L. designed and supervised the research, and edited the manuscript. Z.Z. leads the data analysis and drafted the manuscript with the input from S.M. M.Y. contributed to the overall strategy of data analysis and interpretation of the results. C.Z. provided the phenotypes of 8 Bgt isolates. Y.Y. and X.Z. conducted the RT‒qPCR experiments. J.X. participated in part of the bioinformatics analysis. Y.S., L.W., and X.Z. contributed to the interpretation of gene function. Q.Z. provided materials for Th. ponticum. S.Z. and H.-Q.L. supplied field agronomic traits data for our RNAseq panel. H.W., X.L., and N.J. provided trait for EMS mutant lines. J.W., Y.Z. helped editing the manuscript. All the authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Anthony Hall, Manuel Spannagl, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., Ma, S., Yin, M. et al. Population-scale gene expression analysis reveals the contribution of expression diversity to the modern wheat improvement. Nat Commun 16, 11133 (2025). https://doi.org/10.1038/s41467-025-66100-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66100-4