Abstract
Detecting cell-cell communications (CCCs) in single-cell transcriptomics studies is fundamental for understanding the function of multicellular organisms. Here, we introduce FastCCC, a permutation-free framework that enables scalable, robust, and reference-based analysis for identifying critical CCCs and uncovering biological insights. FastCCC relies on fast Fourier transformation-based convolution to compute p-values analytically without permutations, introduces a modular algebraic operation framework to capture a broad spectrum of CCC patterns, and can leverage atlas-scale single cell references to enhance CCC analysis on user-collected datasets. To support routine reference-based CCC analysis, we constructed the first human CCC reference panel, encompassing 19 distinct tissue types, over 450 unique cell types, and approximately 16 million cells. We demonstrate the advantages of FastCCC across multiple datasets, most of which exceed the analytical capabilities of existing CCC methods. In real datasets, FastCCC reliably captures biologically meaningful CCCs, even in highly complex tissue environments, including differential interactions between endothelial and immune cells linked to COVID-19 severity, dynamic communications in thymic tissue during T-cell development, as well as distinct interactions in reference-based CCC analysis.
Similar content being viewed by others
Introduction
Multicellular organisms rely on cell–cell communications (CCCs) to coordinate development, regulate biological functions, maintain homeostasis, and respond to external stimuli1,2,3. CCCs often occur in the form of ligand–receptor interactions (LRIs), where a ligand released from one cell binds to a receptor on another, triggering downstream signaling events that alter transcription factor activity and gene expression in the receiving cells3,4. These interactions enable cells to coordinate their behavior and facilitate the orchestrated activities of different cell types, driving critical biological processes. Dysregulation of CCCs is frequently associated with pathological conditions, leading to compromised tissue repair, immune dysfunction, cancer progression, and neurodegenerative diseases2,5. Consequently, understanding CCCs is fundamental for unraveling the complex biological processes that drive development and disease.
The widespread availability of single cell transcriptomics data has greatly facilitated the study of CCCs, leading to the development of many computational methods for detecting them3,6,7,8,9,10,11,12,13,14,15. These include core tools, such as CellPhoneDB (CPDB)(V16), CellChat9, ICELLNET10, and SingleCellSignalR11; task-specific tools, such as CellCall12 and NicheNet13 that incorporate intracellular signaling and CPDB (V47 and V58) that expands ligand types; as well as other tools that account for broader conditions, such as CellChat(V1.6 and later), iTALK14, and Connectome15, which integrate differential expression analysis to infer CCCs. Despite their differing emphases, these methods all share a common analytic procedure. This procedure begins with a candidate list of LRIs, which can be either user-defined or obtained from existing databases. It proceeds by examining one pair of cell types at a time, calculating a quantity called the communication score (CS) for each LRI. The value of CS is used to determine whether there is coordinated expression of the ligand on cells of one cell type (i.e., senders) and the receptor on cells of the other cell type (i.e., receivers), at levels exceeding what would be expected under the null. Such coordinated expression is suggestive of potential cell-cell communication and interaction.
While the CCC analysis framework is conceptually straightforward, accurately inferring CCCs presents three important challenges. First, determining an appropriate CS threshold is technically challenging. Some tools rely on manually setting a default CS threshold, which is inherently arbitrary, while others employ statistical methods to compute p-values, which are essential for rigorously assessing whether the observed coordinated expression exceeds expectations under the null hypothesis. However, calculating p-values is difficult since CS, derived from ligand-receptor expression levels, has a relatively complicated functional form that complicates direct evaluation of its null distribution. Consequently, most statistical methods rely on permutation tests to estimate p-values. However, permutation approaches are not only time-consuming and computationally demanding but also highly sensitive to the number of permutations performed, affecting the usability, accuracy, and reliability of CCC analysis. Second, almost all existing methods rely on a single CS to evaluate CCC. For example, CPDB calculates the average expression level of the ligand and receptor, whereas CellChat computes their product. While the average or product can capture specific aspects of CCC, relying on a single score reduces flexibility and limits adaptability to diverse datasets, interaction patterns, and biological contexts. Third, large-scale single-cell studies, involving millions of cells, are being conducted, offering unprecedented insights into cellular diversity and function. These studies contain valuable information that could substantially enhance the analysis of CCC. Unfortunately, few methods are capable of scaling to analyze millions of cells efficiently. Moreover, none of the existing methods can leverage these large-scale single-cell datasets to enhance the analysis of user-collected datasets, which are often smaller in size, thereby uncovering additional biological insights. These limitations restrict the discovery of critical intercellular signaling pathways and impede the biological understanding of CCCs.
To address these challenges, we present FastCCC, a highly scalable, permutation-free statistical toolkit tailored to identify critical CCCs in the form of LRIs and uncover novel biological insights in single-cell transcriptomics studies. FastCCC adopts a prior-guided, reference-based statistical design. It leverages curated LRI lists and cell-type annotations to perform interpretable, hypothesis-driven inference grounded in established biological knowledge. It presents a novel analytic solution for computing p-values in CCC analysis, enabling scalable analysis without the need for computationally intensive permutations. It introduces a modular computational framework for CS computation that calculates various CSs through a range of algebraic operations between ligand and receptor expression levels, capturing a broad spectrum of CCC patterns and ensuring robust analysis. Additionally, FastCCC not only enables the analysis of large-scale datasets containing millions of cells, but also introduces, for the first time, reference-based CCC analysis, where large-scale datasets are treated as reference panels to substantially improve CCC analysis on user-collected datasets. To support routine reference-based CCC analysis, we have constructed the human CCC reference panel, which includes 19 distinct tissue types containing approximately 16 million cells across over 450 cell types, enhancing the interpretability and consistency of CCC analysis across diverse biological contexts. We demonstrate the advantages of FastCCC across multiple datasets, most of which exceed the analytical capabilities of existing CCC methods. In real datasets, FastCCC reliably captures biologically meaningful CCCs, even in highly complex tissue environments. Examples include differential interactions between endothelial and immune cells via the C3 ligand and its receptors that are linked to COVID-19 disease severity, dynamic LRIs in thymic tissue during distinct T-cell developmental stages, as well as distinct CCC patterns in reference-based CCC analysis.
Results
FastCCC Overview
FastCCC is described in the Methods, with its technical details provided in the Supplementary information and its core method schematic shown in Fig. 1. FastCCC provides three major advances. First, FastCCC presents a novel, alternative strategy for computing p-values in CCC analysis. By leveraging convolution techniques through Fast Fourier Transform (FFT), FastCCC derives the analytic solution for the null distribution of CS, enabling scalable analytic computation of p-values without requiring computationally intensive permutations. This approach makes FastCCC orders of magnitude faster than existing methods, while enhancing accuracy and robustness. The computational gain of FastCCC further increases with larger datasets, making it particularly suited for large-scale, state-of-the-art single-cell studies that existing methods cannot handle. Second, FastCCC introduces a modular CS computation framework that captures interaction strength through algebraic operations between ligand and receptor expression levels. This framework enables the development of new CS scores beyond common ones by utilizing a wide range of expression summary statistics to capture a broad spectrum of CCC patterns. It also accommodates multi-subunit protein complexes for ligands and receptors, accounting for distinct interaction strengths characterized by different subunits. As such, FastCCC substantially enhances the power and robustness of CCC detection across diverse datasets and biological contexts. Finally, FastCCC not only offers a scalable and effective solution for CCC analysis, enabling the analysis of large-scale datasets containing millions of cells, but also allows these large-scale datasets to serve as reference panels, facilitating more comprehensive and context-aware CCC analysis on user-collected datasets, which may be much smaller in size. To support routine reference-based CCC analysis with FastCCC, we constructed the human CCC reference panel with 19 distinct tissue types and approximately sixteen million cells (Methods, Supplementary Table S2). This reference panel enhances the interpretability and consistency of CCC analysis across diverse biological contexts.
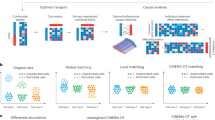
a Core computational workflow. Expression levels of ligand and receptor genes are modeled as random variables X and Y with probability distributions PX and PY. Convolution of distributions and Gaussian approximation via the Central Limit Theorem form the basis for permutation-free null distribution computation of communication scores (CS). Created in BioRender. Zhou, X. (2025) https://BioRender.com/ys4o8uu. b Modular scoring has three layers: (1) gene-level expression summarization using statistics (mean, median, k-th order); (2) aggregation of subunit summaries for multi-gene ligands/receptors into Sligand and Sreceptor; (3) CS calculation using arithmetic or geometric means. c A human CCC reference panel from 19 tissues (~16 million cells, > 450 cell types) stores expression summaries and distributions. d FastCCC derives null distributions and key metrics from this reference for inference. Created in BioRender. Zhou, X. (2025) https://BioRender.com/ys4o8uu. e Reference-based analysis applies the panel to new data to detect significant cell-cell communication, improving robustness and reducing dataset-specific bias. Created in BioRender. Zhou, X. (2025) https://BioRender.com/t50kcmo.
FastCCC ensures accurate and scalable p-value computation
The first important feature of FastCCC is its ability to compute p-values analytically, rather than relying on computationally intensive permutations. To validate the analytic solutions provided by FastCCC, we first focus on the same CS statistic used in CPDB, and compare the p-values from FastCCC, calculated using this single CS statistic, with those from CPDB, which uses permutations. We began by testing the tutorial data used in CPDB (referred to as CPDBTD), which comprises 3312 cells across 40 cell types, nearly half of which are rare, with fewer than 30 cells per cell type, including five extremely rare cell types with fewer than ten cells (Fig. 2a). The cell type composition in the data captures a range of the strategies employed by FastCCC to compute p-values (see Methods; Table S3). As expected, the p-values from FastCCC are highly concordant with those obtained from CPDB (Pearson correlation > 0.995), with the consistency increasing as the number of permutations increases (Fig. S12). For example, when evaluating the set of significant LRIs (p-value < 0.05), the intersection over union (IoU, also known as the Jaccard index) between FastCCC and CPDB, using the default CPDB setting of 1000 permutations, is 96.7%, with precision exceeding 99.3%. These metrics continue to improve as the number of CPDB permutations increases (Fig. 2b, S13, S14): with one million permutations, Pearson correlation increased to 0.988 (Fig. 2c), while IoU and precision reach 97.3% and 99.7%, respectively. Additionally, the correlation for valid LRIs, defined as those where both the non-zero expressed ligand in the sender and the receptor in the receiver exceed a specified percentile, was 0.996 (Fig. S15, Methods). Treating the results of CPDB with one million permutations as ground truth, we found that using 1000–50,000 permutations in CPDB resulted in a notable drop in precision and IoU metrics, particularly under stringent p-value thresholds (Fig. S16). In contrast, the analytical solution provided by FastCCC maintained high accuracy. Notably, CPDB requires at least 10,000 permutations—ten times its default setting—to match FastCCC’s p-value accuracy. Importantly, we also modified CPDB’s CS formulation and validated FastCCC’s p-value results across a range of CS using operations, including order statistics and geometric mean (Fig. S17, Table S3, and Methods).
a UMAP plot of CPDBTD (left) and a line plot shows the corresponding cell counts for each cell type (right). The pink dashed line indicates rare cell types with fewer than 30 cells while the blue dashed line indicates common cell types. b IoU results on the significant LRIs detected between FastCCC and CPDB across varying numbers of permutation tests used in CPDB. Each circle represents an individual experiment (20 replications), with squares and error bars indicating the mean ± SD. c Comparison of FastCCC results with CPDB (1 million permutation tests). Each point represents a significant LRI identified by CPDB. The x-axis shows CPDB p-values on -log10 scale, and the y-axis shows FastCCC results. Pearson correlation coefficient and corresponding p-value are annotated. d UMAP plot of ten additional scRNA-seq datasets, with different colors representing different cell types. The p-value for valid LRIs between FastCCC and CPDB is compared, with Pearson correlation coefficient and corresponding p-value annotated. e Scatter plot displays the number of cells and the number of interaction types in each dataset. f Precision and IoU results for different numbers of permutation tests across all test datasets. g Running time of CPDB and FastCCC, along with data loading time, across all test datasets. h Memory consumption of CPDB and FastCCC, along with the memory used for data loading, across all test datasets. Source data are provided as a Source Data file.
Next, we assessed the consistency of FastCCC with CPDB across ten additional scRNA-seq datasets, encompassing a wide range of cell type numbers, dataset sizes, and ligand-receptor pairs (Fig. 2e, Methods). A high level of consistency was again observed between the p-values from the two methods, with Pearson’s correlation ranging from 0.996 to 0.999, confirming the reliability of FastCCC (Fig. 2d). Moreover, the consistency between the two methods improves with an increasing number of permutations used in CPDB, with precision surpassing 99% and IoU surpassing 97% across all datasets when the number of permutations reached 10,000 (Fig. 2f).
While the p-values from FastCCC are nearly identical to those from CPDB, FastCCC offers significant computational advantages in terms of both computation time and memory usage. These efficiency gains brought by FastCCC increase as the number of cells in the dataset grows. Specifically, FastCCC is at least 40 times faster than CPDB when the number of cells exceeds 100,000 and nearly 500 times faster on a dataset with two million cells, completing all calculations under 20 minutes in the latter case (Fig. 2g). In fact, the computation of FastCCC is so efficient that a substantial proportion of its required computing resources, approximately 60% of memory and 25% of computing time across all test datasets, are spent on data reading rather than the actual computation (Fig. 2g, h). Additionally, across the eleven smaller datasets, the average memory consumption of FastCCC was 69% of CPDB; while FastCCC is the only method that is scalable to the two large datasets (AIDA and HLCA), each containing over one million cells with thousands of different LRIs.
We extended the computation comparison to the default version of FastCCC, which uses 16 distinct CS values (more below), along with seven existing CCC analysis methods, including CPDB V5 and V4, CellChat, SingleCellSignalR, ICELLNET, NicheNet, and CellCall. These methods were selected based on their superior performance in previous benchmarking studies16,17 and their widespread usage in the field. Compared to these approaches, FastCCC stands out as the only method that provides statistical significance without the need for permutations (Fig. 3a). For the eight smaller datasets, FastCCC was at least ten times faster than nearly all other methods, while also consuming the least memory across datasets (Fig. 3b, c). For the two large datasets, FastCCC was again the only applicable method. In particular, CellChat and CellCall encountered memory errors when processing datasets with 100–200 k cells using default parameters, while SingleCellSignalR timed out when processing datasets over 100 k cells (see Methods). Both NicheNet and CellCall exhibited significant delays and timeouts on datasets larger than 500 k cells. CPDB series showed excessive memory consumption on datasets reaching 1 million cells. In contrast, FastCCC efficiently processed large-scale datasets with high speed and low memory consumption, demonstrating its scalability and practical application in real-world scenarios involving massive scRNA-seq datasets.
The results are shown for the default version of FastCCC with 16 scoring methods a A comparative table outlining the key features, functionalities, and applicability of different CCC analysis methods across test datasets. A ✓indicates the presence of a feature or compatibility, while × indicates the absence or incompatibility. A half tick for CellPhoneDB denotes partial support, such as requiring user-provided DEG data. b Runtime comparison between FastCCC and other methods across 10 datasets. Datasets are sorted by increasing number of cells along the x-axis; runtime (in seconds) is plotted on the y-axis. Points represent mean values and the whiskers extend from the minimum to the maximum values. c Memory usage comparison across the same datasets. Peak memory consumption is shown on the y-axis (in MB). d Comparison of false positive rates (FPR) across eight benchmark datasets. Each point represents one simulation replicate; error bars indicate mean ± S.E.M. Lower values indicate better performance. e Comparison of F1-scores across the same eight datasets. Each point corresponds to an individual simulation replicate, with mean ± S.E.M. indicated. Statistical annotations above the FastCCC results indicate the significance of difference between FastCCC and the best-performing alternative method on the same dataset, using a two-sided Mann-Whitney U tests. These tests assess whether FastCCC achieves significantly better F1-score performance compared to the strongest competing method on each dataset. Source data are provided as a Source Data file.
FastCCC is robust and powerful
The second important feature of FastCCC is its ability to employ a variety of algebraic operations to compute additional CS values beyond those previously used, such as the one used in CPDB. The default version of FastCCC includes a total of 16 CS values, allowing it to capture a wide range of interaction patterns between ligands and receptors across different biological contexts, thus ensuring its power and robustness. Here, we evaluated the accuracy of FastCCC and compared it with existing methods on eight relatively small single cell datasets (excluding AIDA and HLCA), where most other methods were able to produce results. To achieve this, we employed a semi-simulated approach: shuffling cell labels to create null scenarios where none of the LRIs interact with each other, and a small proportion of LRIs were then randomly selected and overexpressed to serve as the truly interacting LRIs, thereby establishing ground truth (see Methods).
We first examined false positive rates (FPR) and recall of various methods across datasets. Nearly all methods achieved FPR below 10% across all datasets (Fig. 3d), with the only exception of SingleCellSignalR, which exhibited an average of 17.4% and 16.0% FPR in the Fibroblast and ECGE datasets, respectively. FastCCC achieved the lowest FPR among all methods, maintaining FPR below 1% across all datasets, except Fibroblast, where the FPR was 1.5%. CellCall also achieved a relatively low FPR, but its recall was nearly zero across all datasets (Fig. S18), suggesting that it failed to detect any signals. Similarly, NicheNet also yielded low recall. In contrast, FastCCC achieved the highest recall while maintaining the lowest FPR, reflecting its superior accuracy in capturing the true interactions.
Beyond recall and FPR, FastCCC consistently outperformed the other methods across all datasets on other evaluation metrics, such as precision, accuracy, specificity, balanced accuracy, and the Jaccard index (Fig. S18). Additional metrics, such as F1 score and Matthews correlation coefficient (MCC), also yielded similar conclusions (Fig. 3e, Fig. S18), supporting FastCCC’s superior performance. For instance, in the Ileum dataset, FastCCC achieved the highest median and mean F1 scores, both at 0.863, outperforming CellChat (median: 0.699, mean: 0.701), CPDB V5/V4 (median: 0.391, mean: 0.413), SingleCellSignalR (median: 0.425, mean: 0.431), ICELLNET (median: 0.574, mean: 0.565), NicheNet (median: 0.027, mean: 0.025), and CellCall (both 0.0 due to no significant output). Notably, among the other methods, methods employing p-values, such as CellChat and CPDB, generally outperformed others, which aligns with previous benchmarking studies16,17, highlighting the value of rigorous statistical approaches in CCC analysis.
To further assess the accuracy and biological relevance of FastCCC, we performed a series of independent evaluations using diverse external data sources. Specifically, we evaluated whether top-ranked LRIs inferred by FastCCC are enriched between spatially adjacent cell types in multiple spatial transcriptomics datasets, aligned with surface protein expression in CITE-seq data, and consistent with in vivo cell-contact measurements from the uLIPSTIC system. In addition, we assessed cross-method concordance across several benchmark datasets to determine whether FastCCC captures widely supported communication signals. These complementary evaluation strategies and results, described in detail in the Supplementary Materials (Section 1, Fig. S1 to S5), demonstrate that FastCCC not only achieves strong statistical performance, but also recovers biologically meaningful cell-cell communications across multiple modalities.
Application of FastCCC to a large-scale COVID-19 dataset
The scalability and effectiveness of FastCCC allows us to conduct CCC analysis on large-scale single-cell datasets within a reasonable time frame and at a fraction of computational cost of the other methods. To further illustrate its utility, we applied FastCCC to two large-scale datasets: a large COVID-1918 dataset, and a thymus dataset19. The second data contains pseudo-time information that requires CCC analysis to be conducted across time points.
We first applied FastCCC to a COVID-19 scRNA-seq dataset comprising 1.46 million cells18 (Fig. 4a), a scale beyond the feasible resource limits of other comparison methods. FastCCC successfully completed all 16 distinct score calculation methods in approximately 40 min, using 120 GB of memory (with the dataset itself occupying 60 GB). This dataset includes samples from 40 patients and five healthy controls, categorized into different stages—disease progression and recovered convalescence (Fig. 4b). The dataset also comprises epithelial (Epi) cells and various immune cells sampled from patients with moderate and severe disease. We explored significant CCCs across different groups to understand changes in cellular communication during COVID-19 progression.
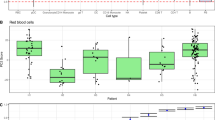
a UMAP plot of the COVID-19 dataset, with colors representing different cell types. b A percent stacked bar chart showing sample groups, disease sampling time points, and COVID-19 severity levels. c Venn diagram of significant LRIs across different groups. The number and proportion of unique LRIs in each group are annotated, with additional labels for intersections comprising more than 10%. d Chord plot of significant LRIs uniquely expressed in the progression severe group compared to the mild group. The color and angle of the arcs represent cell types and the number of significant LRIs involved, respectively. The color of the chords indicates the sender cell type, with the width reflecting the number of LRIs. Arrows point to the receiver cell type, and the main cell types involved in CCC are annotated. e LRIs uniquely expressed in the mild group compared to the severe group. f Pseudo-bulk RNA-seq expression of signaling molecules involved in CCCs during disease progression compared to healthy controls. Data are presented as mean ± SD; n = 17 (group severe(saliva)), five (group mild(saliva)), nine (group severe(lung)) and three (group mild(lung)) biological replicates. P-values were calculated using two-sided Mann-Whitney U tests. g Violin plot showing the expression levels of ligand C3 and its receptor C3AR1 across different groups. h Circle plot of differentially expressed LRIs. The size and color of the circles together represent the significance of LRIs in the mild and severe groups. i Pathways associated with the ligands and receptors differentially expressed in the severe group compared to the mild group are displayed. Source data are provided as a Source Data file.
We first conducted CCC analysis across five sample groups, including healthy controls, patients with mild and severe cases during disease progression, and patients in recovery, categorized as mild and severe convalescence. FastCCC detected a total of 1874 CCCs by integrates 16 CS scores, with 702, 1273, 1026, 996, and 992 CCCs identified in the five groups, respectively (Fig. S23). A significant proportion of CCCs (358 CCCs, 19.1%) were shared across all groups, with 7–170 intersections (mean = 45.4; median = 24) shared between pairs of groups (Fig. S23). Importantly, we observed a progressive divergence from the healthy control group, moving through mild recovery, severe recovery, mild disease, to severe disease (Fig S24), as reflected by the increasing number of unique CCCs detected in these groups. For instance, the healthy control group exhibited only 30 unique CCCs, while the convalescence groups showed approximately 60 unique CCCs. In contrast, the disease progression groups displayed over 200 unique CCCs (Fig. 4c). Additionally, 9.1% (170) of CCCs were shared exclusively between the mild and severe disease progression groups.
We narrow down our focus to examine the CCCs identified by FastCCC in severe and mild COVID-19 case to interrogate the possible molecular pathways underlying disease progression. We found that the CCCs uniquely occurring in severe cases primarily involved interactions among epithelial cells and myeloid cells, such as macrophages (Macro), neutrophils (Neu), monocytes (Mono), and dendritic cells (DCs)(Fig. 4d), In contrast, CCCs occurring in mild cases primarily involved interactions between Macro, DCs, Mono (Fig. 4e), suggesting a different pattern of immune response involvement during COVID-19 progression (Fig. S24). The top signaling molecules involved in CCCs with disease progression compared with healthy controls include ACE2, TMPRSS2, KRT5, ITGB6, and VIM, which emerged as significant ligands or receptors in patients. ACE2 and TMPRSS2 are key components of the COVID-19 pathway20,21,22, with ACE2 serving as the functional receptor for the spike glycoprotein of SARS-CoV-2, while TMPRSS2 serves to cleave the spike protein, facilitating viral entry into host cells. To further investigate, we obtained pseudobulk RNA expression data for each sample and focused on epithelial cells from two primary sources: saliva and lung, capturing nasal epithelial cells and upper airway epithelial cells, respectively. We examined the differential CCC information to explore possible communication mechanisms involving these epithelial cells. We found that (Fig. 4f), while both ACE2 and TMPRSS2 are significantly elevated in COVID-19 patients compared to healthy controls, their expression patterns differ between severe and mild cases: ACE2 is marginally more expressed in severe cases, both in nasal and upper airway epithelial cells, whereas TMPRSS2 expression does not differ significantly between the two groups. Additionally, markers of epithelial-mesenchymal transition (EMT), such as KRT5, ITGB6, and VIM, were associated with COVID-19 severity. Specifically, KRT5 and ITGB6 were more highly expressed in lung tissues of severe patients compared to mild cases, while VIM was highly expressed in the upper airway of severe patients but showed no significant difference in lung tissues. Therefore, epithelial cells in severe cases appear to secrete ligands that enrich EMT pathways compared to mild cases, dovetailing recent studies and suggesting significant differences in disease progression mechanisms between severe and mild cases23,24.
Another top signal identified during CCC analysis is the ligand C3, which directs the interaction between Epi cells and various immune cells, such as Macro, Mono, and DCs. While C3 is typically produced in the liver and also secreted by adipocytes and macrophages as a central component of the immune system25, it displayed an unusually highly expression in ciliated, secretory, and AT2+ epithelial cells in the severe group(Fig. 4g). In mild cases, C3 was only highly expressed in secretory epithelial cells, which constitute less than 1% of the total epithelial cells. The results suggest that C3 activation drives maladaptive immune responses, promoting the release of pro-inflammatory cytokines and recruitment of neutrophils and monocytes into lung tissue26,27,28,29. These immune cells can damage the air-blood barrier, thereby exacerbating ARDS severity in SARS-CoV infections26.
Other signals differentially detected between severe and mild cases include chemotaxis, such as when Epi cells release PPIA as a ligand to bind with BSG (CD147) receptor, directing Macro, Mono, NK cells and others to the site of infection or inflammation. Additionally, severe cases showed elevated expression of CCCs related to chemokines like CCL and CXCL (Fig. 4h). These findings imply the occurrence of a cytokine storm in severe patients, contributing to their worsened clinical phenotypes. In contrast, mild patients exhibited higher expression of leukocyte adhesion-related CCCs, primarily between innate immune cells.
Finally, pathway enrichment analysis of genes involved in these differential CCCs revealed that severe patients activate multiple immune pathways (e.g., TNFα via NF-κB, IL-6/JAK/STAT3), which are likely contribute to cytokine and inflammatory storm formation(Fig. 4i, Figs. S25, S26).
Applying our method to this large-scale dataset demonstrated its practical utility, with the COVID-19 results largely corroborating existing research while uncovering numerous unique signals that may offer new avenues for further study.
Application of FastCCC to a thymus dataset with developmental time series
We applied FastCCC to a large-scale thymus scRNA-seq dataset with pseudo-time information, dividing it into thousands of sliding time windows for detailed characterization. This required CCC analysis to be carried out within each time window, resulting in thousands of analyses—a burdensome task that again is not feasible by other methods—to comprehensively investigate the dynamic cellular interactions during T cell development. This data consisted a total of 255,901 cells19, including thymic epithelial cells (TECs), DCs, various stromal cells, as well as differentiating T cells (Fig. 5a). The T cell population consists of 76,903 thymocytes at different developmental stages, including double negative (DN), double positive (DP), CD4+ single positive (SP), CD8+ SP, and T regulatory (Treg) cells, among others.
a UMAP plot of the thymus dataset, with colors representing different cell types. Developmental T cell lineage and other stromal cells or immune cells are separated by dividing lines. b Illustration of the T lymphocyte developmental process. Created in BioRender. Zhou, X. (2025) https://BioRender.com/mh49697. c UMAP plot of the T cell lineage pseudotime, showing progression from the earliest DN stage to CD4+ or CD8+ lineages. d GO pathway enrichment analysis of signaling molecules involved in key CCCs at each stage. e Violin plot showing expression levels of ligand DLK1, DLL4, and JAG2 across different stromal cell type groups. The corresponding receptor expression levels (NOTCH1/3) and CS p-value over time are depicted, where each blue dot represents the mean receptor expression within a time window, and dashed lines indicate the smoothed -log(p-value) levels for a specific LRI pair. f. Violin and scatter plots showing expression levels of ligands CCL17/19/22, along with their corresponding receptors CCR4/7, as well as the changes in p-value.
Thymic development is a highly regulated process in which T lymphocytes originate from multipotent hematopoietic stem cells located in the bone marrow30. Initially, progenitor cells migrate from the bone marrow to the thymus via the blood, entering through the cortico-medullary junction before moving to the outer cortex. Within the thymus, these progenitors undergo a series of maturation steps19,30. As shown in the Fig. 5b, the thymus is composed of multiple lobules, each containing a distinct outer cortical region and an inner medulla. Developing T-cell precursors reside within a complex epithelial network known as the thymic stroma, a specialized microenvironment crucial for T-cell development. This stroma is rich in signaling pathways that regulate the commitment to the T-cell lineage.
Upon entering the thymus, progenitor cells lack the surface molecules characteristic of mature T cells, and their receptor genes remain unrearranged. These progenitor cells, often referred to as “double-negative” thymocytes due to their lack of CD4 and CD8 expression, eventually differentiate into the major population of αβ T cells and the minor population of γδ T cells. Further differentiation of αβ T cells results in the formation of two distinct functional subsets: CD4+ T cells and CD8+ T cells. Our study primarily focuses on the major αβ T-cell lineage. By analyzing thymocytes as a whole, we examined the interactions between these cells with other stromal cells and immune cells within the thymic microenvironment.
We first examined the relationship between pseudotime and T cells types (Fig. 5c) and found that the inferred pseudotime trajectory captures the expected T cell development from early thymic progenitors (DN (early)), to DP cells, and eventually to either CD8+ or CD4+ SP T cells, which further specialized into subsets including T helper cells (Th) and regulatory T cells (Tregs). Next, we conducted differential expression analysis using FastCCC’s probability toolkit for each T cell subpopulation and performed pathway enrichment analysis on significant ligands and receptors identified at various developmental stages (Fig. 5d). These confirmed ligands and receptors were involved in CCCs that revealed the critical molecular pathways underlying T cell development. Specifically, during the DN stage, T-cell communication predominantly involved essential metabolic processes related to cell survival, proliferation, and development. However, in the DP (quiescent) stage, the number of significant interactions decreased sharply, with the remaining interactions strongly associated with T-cell differentiation, activation, and selection, indicating the onset of T-cell lineage commitment. In the αβ T-cell stage, additional cytokine and kinase regulation pathways emerged, along with signals related to cell adhesion and migration, suggesting that T cells were beginning to mature and migrate out of the cortex. Once T cells differentiated into CD4+ and CD8+ subsets, the signaling landscape became more complex, with an increase in interactions related to immune cell functions and T-cell proliferation. Signals associated with cell adhesion and migration also reappeared, indicating that these mature T cells were preparing to exit the thymus and circulate in the periphery.
FastCCC identified multiple key signaling pathways during T cell development, including the NOTCH and chemokine pathways. First, FastCCC accurately depicted the intense interactions driven by NOTCH1 and NOTCH3 receptors and their ligands, such as DLK1, DLL4, and JAG2, during the DN(early) and DN stage (Fig. 5e). NOTCH receptor signaling is key for T-cell precursors' commitment to the T-cell lineage30. Second, FastCCC identified multiple chemokine receptors, such as CCR4 and CCR7, along with their ligands (e.g., CCL17, CCL22, CCL19), to serve as key players during the migration of post-selection thymocytes into the medulla (Fig. 5f). This result implies the role of chemokine signaling in guiding thymocyte migration to the medulla and ensuring the proper maturation of SP thymocytes. In particular, we observed that CCL19 and CCL21 released from mTECs (medullary TECs) interacted with the CCR7 receptor on naïve lymphocytes to potentially control the migration of developing thymocytes. Such interaction between CCR7 and its ligands started in the αβ T-cell (entry) stage, and the interaction strength peaked during the SP stage. Among the CCR7 ligands, CCL19 is specifically expressed in mTECs compared to cTECs (cortical TECs), and such an expression pattern between mTECs and cTECs likely created a chemotactic gradient that guides the migration of CCR7-bearing naïve T cells31,32,33,34. Interestingly, we found that a subtype of dendritic cells (aDCs) also expressed ligands, such as CCL17, CCL19, and CCL22 (Fig. 5f, Fig. S27), suggesting that this specific DC subtype, but not other DC subtypes, such as pDCs, DC1, and DC2, may play a key role in modulating the thymic niche.
FastCCC identified multiple additional CCCs that also play crucial roles during T cell development (Fig. S27). For instance, FastCCC revealed that P-Selectin Glycoprotein Ligand (SELPLG or PSGL-1), a glycoprotein characterized by mucin-type O-glycan modifications that are expressed in DN cells, interacts with E- and P-selectins (SELP, SELE) on TECs. This interaction mediates cell rolling and tethering, promoting the capture of cells from the bloodstream, providing support that endothelial cells assist T-cell progenitors in entering the thymus during the colonization of lymphoid progenitor cells31. Additionally, FastCCC also detected CD47-SIRPA signaling, which promotes the opening of tight connections between endothelial cells, to mediate the communication between DN and endothelial cells31,35. Regarding CCL21 and CCR7 signaling, similar to the CCL19-CCR7 signal, we discovered that CCL21 is expressed in both type I and type II mTECs, while CCL19 is expressed across all mTEC subtypes. Our method also identified that the CXCR4 receptor and its ligand CXCL12, as well as the CCR9 receptor and its ligand CCL25, are vital signals directing thymocyte migration.
These findings illustrate that thymocytes are not merely passive participants within the thymus; they actively influence the organization of the TECs on which they depend for survival. As developing thymocytes progress through distinct stages, marked by changes in T-cell receptor expression, these surface changes reflect their functional maturation. Specific combinations of cell surface proteins serve as markers for T cells at different stages of differentiation. Collectively, our results suggest that FastCCC effectively captures significant signaling interactions across various developmental stages, providing invaluable insights into the temporal dynamics of these interactions.
Application of FastCCC for reference-based CCC analysis and construction of a human CCC reference panel
The third important feature of FastCCC is its ability to conduct reference-based CCC analysis. With the growing availability of large-scale single-cell RNA sequencing datasets across diverse tissues and samples36, FastCCC can utilize these datasets as reference panels. By leveraging the information contained in the reference panels, FastCCC enhances CCC analysis on user-collected data, which are often smaller in size, yielding more comprehensive and insightful results.
To support reference-based CCC analysis, we first constructed a human CCC reference panel using the CELLxGENE37 platform. Specifically, we curated and organized 16 million singe-cell data from 19 human tissues, containing 452 cell types from 1025 healthy samples and ten distinct techniques (Fig. 6a, b, and Table S2). This effort resulted in the creation of the first human CCC reference panel, which can be used by FastCCC to directly identify significant CCCs by comparing user-collected query data with the reference (see Methods).
a Stacked bar plot showing the proportion of scRNA-seq data from different technical platforms in each tissue CCC reference panel. b Scatter plot showing the number of cells and cell types in each reference panel. c Comparison of results obtained using the reference-based method and conventional CCC analysis in the benchmarking dataset which serves as the ground truth. Data are presented as mean ± S.E.M. d Pairwise comparison between the results of the reference-based method and conventional CCC analysis using the biased datasets. Each point represents a subset containing a group of similar cell types as the query test. Results from conventional CCC analysis on the entire dataset were used as ground truth. P-values are calculated using one-sided paired Student’s t-tests (n = 6). e Examples of communications between cancer epithelial cells and NK cells are demonstrated. Gaussian distributions represent the null expression summaries of ligands and receptors from the reference, with their statistical values mapped onto the query dataset. Blue bands indicate the mapped CS results, with dark blue representing the 95% confidence interval. Correspondingly colored dots show the levels of the respective statistical values in the query data. f Sankey plot showing downregulated CCCs in the query data with cancer epithelial cells as the sender cell type. The first row represents the ligands involved in the downregulated CCCs, while the second row indicates the cell types these ligands act upon via their receptors. Source data are provided as a Source Data file.
Reference-based CCC analysis using the human CCC reference panel with FastCCC is straightforward, requiring users to provide only the raw gene expression count matrix of their query data. The workflow for reference-based CCC inference implemented in FastCCC involves multiple steps. First, FastCCC preprocesses the query data by converting gene expression to expression ranks in each cell (see Methods), a strategy used in expression quantitative trait mapping studies38 and single-cell foundation models39, to minimize batch effects between the query and reference datasets. Second, FastCCC relies on housekeeping genes40,41 to adjust for the remaining differences between the two datasets through mapping the reference-based expression summaries onto the query domain. Finally, based on the number of cell types in the query dataset, FastCCC determines the threshold for declaring signficiant and further maps the threshold to the query data into a probabilistic distribution to determine the mapped significant threshold for CS, thereby identifying significant LRIs in the query data (more details provided in the Methods).
We first validated the use of reference-based CCC analysis using lung data containing approximately 1.7 million cells from our constructed human CCC reference panel in two distinct case studies. In the first case study, we constructed three distinct query datasets by extracting three anatomical regions of the lung, including lung parenchyma, the lower lobe of the left lung, the bronchus, and the upper lobe of the left lung, while retaining the remaining lung data as the reference. In the second case study, we obtained four normal lung datasets not included in our CCC reference, along with three lung-related disease datasets (one of which encompassed fifteen distinct diseases, each analyzed separately; see Methods). Each of these 21 normal or disease datasets was treated as a query dataset. For the reference data, we used either the complete lung data from our human CCC reference as the reference panel or the lung reference data from the first case study. As shown in Fig. 6c, the results from reference-based CCC analysis are highly accurate compared to standard CCC analysis using the query data alone. Specifically, across all normal and disease query datasets, the average precision reached 80%, with an IoU of 80% and a recall of 95%. In addition, in the second case study, for the four external normal datasets and seventeen disease datasets, the results were nearly identical between the two references(Figs. S29, S30), with a slight increase in precision and IoU for the larger reference, demonstrating the robustness of reference-based CCC analysis using the constructed human CCC reference.
One important advantage of FastCCC’s reference-based approach is its ability to capture a diverse range of cell types, facilitating the generation of more accurate background distributions and, consequently, more precise CCC analysis. Compared to analyzing the query data alone, incorporating a reference helps mitigate inherent biases in CCC analysis that may arise from the relatively small size or specific experimental conditions of the query data. For example, certain query datasets may involve selective sorting of single cells, where precise isolation of specific cells from a heterogeneous population is conducted, thus artificially inflating the mean CS for all LRI pairs associated with these cell types in the null distribution. Such inflation of null statistics can result in a loss of power and failure to detect potentially up-regulated interactions. By leveraging our comprehensive human CCC reference, which encompasses a diverse range of cell types, such biases can be effectively mitigated.
To illustrate this benefit, we used the dataset from Teichmann et al., which contains a total of ~318 k cells across 38 cell subtypes that belong to six major cell type groups: endothelial, epithelial, fibroblast, lymphoid, myeloid, and muscle. We conducted conventional CCC analysis in the full dataset and treated the results as the ground truth. In addition, we split this dataset into six subsets based on the major cell type groups and conducted independent CCC analyses for each subset, which exhibited pronounced imbalances in cell type proportions compared to the full dataset. This imbalance significantly affected the baseline levels of CS statistics for certain LRI sets associated with specific cell types, potentially introducing biases when applying CCC analysis directly to the subdivided datasets rather than the full dataset. Ideally, an accurate inference method should yield consistent CCC results regardless of the cell types present in the dataset, leading to similar results as inferred from the full dataset. However, in some extreme cases, such as myeloid and muscle major group, the IoU and recall from CCC analysis using the query data alone dropped to below 60% when compared to the results of the full dataset used as ground truth (Fig. 6d). In contrast, inference using the human CCC reference panel consistently outperformed inference using query data alone across all analyses. Specifically, using the lung CCC reference panel achieved an average consistency exceeding 80% compared to the ground truth obtained from the full dataset. For all subdivided datasets, recall consistently surpassed 90%, while precision remained at the same level. Even in extreme scenarios – where the query dataset contained only a single cell type and conventional CCC methods failed to produce any autocrine interaction results, reference-based CCC analysis using the human CCC reference panel maintained an average IoU exceeding 80% for single-cell-type query datasets (Fig. S31).
To further demonstrate the benefits of FastCCC’s reference-based approach, we utilized the breast reference from our human CCC reference panel, consisting of over 3 million cells and 51 cell types from 323 healthy samples, to analyze an independent breast tumor dataset42 obtained externally. Despite the differences in the resolution and ontology of cell type annotations between the reference and query datasets, FastCCC’s versatile reference-based approach enables flexible merging of subtypes or mapping of cell type labels to align the meta-information of the reference and query datasets. In the reference-based analysis, we found that significant LRIs in breast cancer were predominantly observed between specific immune and stromal cell types and cancer epithelial cells. For instance, cancer epithelial cells, as senders, influence immune and stromal cells through receptors, such as TIGIT, ITGB1, PIK3CB, and a range of chemokines and cytokines (Fig. S32)—findings consistent with previous research43,44,45,46,47,48. These receptors are involved in pathways extensively linked to cancer progression, including cell adhesion molecules, whose disruption is a hallmark of cancer44,45; to PI3K-Akt signaling, a key regulator of survival, metastasis, and metabolism46,47; and to ECM-receptor interaction, crucial in breast cancer development48 (Other top enriched KEGG pathways see Fig. S33). Compared to conventional CCC analysis using large-scale normal breast tissue samples, reference-based CCC analysis enables precise identification of contributions to significant CS statistics. For example, in LRIs between cancer epithelial and NK cells, such as ICAM3-ITGB2 and ICAM3-ITGAL, both ligands and receptors are expressed at significantly higher levels in the query dataset than in normal conditions(Fig. 6e). In these cases, CS in the query dataset are over ten times the standard deviation beyond the CS threshold from the reference, indicating a strong communication between these two cell clusters.
The second important advantage of FastCCC’s reference-based analysis lies in its ability to detect reduced CCC levels in the query data compared to the reference, which is composed of normal tissues. Specifically, conventional methods focus only on identifying significantly enhanced CCCs exceeding a certain threshold. In contrast, FastCCC leverages a CCC reference panel constructed from normal tissues to perform additional analysis, enabling the identification of CCCs with significantly reduced interactions relative to the reference, in addition to significantly enhanced CCCs. For instance, in ANXA1-CXCR3, THBS1-ITGB1, and ITGB1-CD46 interactions between cancer epithelial and NK cells(Fig. 6e), ligand expression in cancer cells is significantly lower in the query dataset compared to the reference, regardless of receptor expression levels. The overall CS values fall more than two standard deviations below the normal reference threshold, suggesting markedly reduced communication levels.
Notably, across the query dataset, 12 cancer epithelial ligands, including ANXA1, THBS1, ITGB1, and ADAM17, were consistently associated with all other immune cells through LRIs that fell below 1.96 times the standard deviation of the normal reference threshold (Fig. 6f). Importantly, literature extensively links these ligands’ overexpression with increased tumor growth, enhanced tumor cell migration, aggressiveness, and poor prognosis49,50,51,52,53,54,55,56. To further validate the clinical relevance of FastCCC results, we separately analyzed single-cell data from triple-negative breast cancer (TNBC) patients—a more aggressive breast cancer subtype with a worse prognosis—accounting for less than 30% of the total dataset. Reference-based CCC analysis in TNBC samples revealed notable differences: LRIs associated with most of the aforementioned ligands, such as ANXA1, THBS1, and NAMPT, were no longer significantly downregulated (Fig. S35). Additionally, ITGB1 and ADAM17-related LRIs showed a dramatic reduction in the number of significantly decreased interactions, involving only 2-4 CCCs. For these remaining downregulated LRIs, the reduction primarily resulted from decreased receptor expression.
These findings demonstrate that reference-based comparisons effectively provide background distribution information, closely approximating the results of direct CCC inference methods applied to datasets with large, diverse cell populations, align with clinical phenotypes and offer mechanistic insights into tumor behavior and microenvironmental changes. This reference-based approach effectively mitigates biases associated with small datasets, providing a more comprehensive characterization of CCCs. This enables researchers to obtain a more accurate representation of CCCs and facilitates a deeper understanding of intercellular communication.
Discussion
We have presented FastCCC, which represents a significant advancement in computational frameworks for CCC inference from single-cell transcriptomic data. By circumventing the need for permutation-based statistical testing, FastCCC offers a highly efficient and analytically rigorous alternative that substantially enhances scalability, flexibility, and robustness. These methodological improvements are not merely technical refinements—they directly address long-standing limitations in CCC analysis and open the door to new biological discovery in the era of atlas-scale single-cell datasets.
A central innovation of FastCCC lies in its analytic computation of p-values for CS, replacing the computationally intensive and often unstable permutation strategies commonly used in tools like CPDB. This not only accelerates computation by several orders of magnitude but also ensures more consistent and reproducible inference. As datasets grow to millions of cells across hundreds of cell types, this computational scalability becomes essential for timely and routine CCC analysis.
Another key strength of FastCCC is its flexible, modular design. Rather than relying on a single definition of CS, FastCCC incorporates a wide range of algebraic operations between ligand and receptor expression summaries—such as mean, quantile statistics, and geometric combinations—allowing the framework to model diverse interaction patterns. This flexibility enables FastCCC to capture biologically relevant signals across a wide spectrum of scenarios, including those driven by rare subpopulations or constrained by multi-subunit bottlenecks, which are often overlooked by traditional approaches. The aggregation of multiple CS metrics using the Cauchy combination test (CCT) further enhances statistical power while maintaining type I error control under correlation.
FastCCC also pioneers reference-based CCC analysis, allowing large-scale, biologically diverse single-cell datasets to serve as reference panels for smaller user-collected datasets. This paradigm shift not only improves the sensitivity and specificity of CCC inference but also facilitates comparative studies across tissues, disease states, or perturbations. Our construction of the first human CCC reference panel, encompassing 19 tissue types, over 450 cell types, and 16 million cells, demonstrates the feasibility and power of this approach. Through empirical evaluation in both real and simulated settings, we demonstrate that reference-based analysis mitigates biases arising from small sample sizes and unbalanced cell-type compositions, enabling more accurate and context-aware discovery of intercellular signaling.
Nonetheless, FastCCC’s design reflects several key assumptions and limitations, many of which are shared by other existing CCC methods. First, the framework currently infers communication based solely on transcriptomic co-expression of ligands and receptors. While this enables broad applicability, it does not guarantee physical interaction or functional signaling. For instance, the presence of a transcript does not confirm surface protein abundance, subcellular localization, or proximity-based activation. FastCCC does not yet incorporate orthogonal modalities, such as spatial transcriptomics, proximity ligation assays, or surface proteomics (e.g., CITE-seq), which are critical for refining the interpretation of CCC predictions.
Second, FastCCC assumes independence between ligand and receptor expression under the null hypothesis, including the independence of subunits in multi-gene complexes. Although empirical data suggest that most LRIs exhibit weak or no correlation, and this assumption has minimal impact on accuracy in practice, it may oversimplify cases with tightly co-regulated expression modules. Our analysis reveals that the independence assumption is conservative in the presence of positive correlation, thereby narrowing the null and increasing statistical power. Nevertheless, future extensions could incorporate co-regulation-aware null models when joint distributions are estimable.
Third, while FastCCC demonstrates strong concordance with permutation-based methods and high evaluation performance in orthogonal data types (e.g., spatial adjacency, CITE-seq protein data, in vivo contact assays), it is important to emphasize that no universally accepted ground truth exists for CCC inference1,3,17,57,58. As such, we encourage cautious interpretation of results and the use of complementary validation strategies whenever possible.
Looking forward, FastCCC offers a flexible foundation for future methodological expansion. Incorporating spatial coordinates and neighborhood structure from spatial transcriptomics will allow FastCCC to model spatially resolved communication landscapes and define spatial CCC reference panels. Furthermore, integrating additional molecular modalities—such as surface proteomics, epigenetic states, or spatially-resolved chromatin contacts—will enable a more holistic reconstruction of intercellular signaling. Finally, expanding FastCCC to non-human species and developmental atlases will extend its utility to comparative and evolutionary studies.
In conclusion, FastCCC is a fast, robust, and versatile toolkit for CCC analysis in large-scale single-cell transcriptomics. By addressing long-standing computational and methodological bottlenecks, it enables researchers to more fully harness the growing wealth of single-cell RNA-seq data and uncover new insights into the complex signaling networks that govern multicellular biology.
Methods
A general CCC testing framework
FastCCC is a biologically informed, prior-gufFFT ided statistical framework for cell-cell communication. It operates by integrating three types of inputs: (1) a curated list of known LRI pairs, (2) annotated cell types or clusters, and (3) user-provided scRNA-seq profiles. These inputs provide the biological context necessary to compute CSs and assess statistical significance using analytical null distributions. We begin with a candidate list of ligand-receptor pairs, which can be either user-defined or obtained from existing databases. For each ligand l and receptor r, we analyze one pair of cell types at a time. For each such quadruplet, we compute a CS to quantify the coordinated expression of ligand l in cell type ca and receptor r in cell type cb. The CS score is represented in general form as follows:
where \(s({x}_{{c}_{a},l})\) is a scalar and represents the gene expression summary of ligand l across cells of cell type ca; \(s({x}_{{c}_{b},r})\) is also a scalar and represents the gene expression summary of receptor r across cells of cell type cb; and h( ⋅ , ⋅ ) is a function that measures the coordinated expression between \(s({x}_{{c}_{a},l})\) and \(s({x}_{{c}_{b},r})\).
The CS statistic described in equation (1) is quite versatile, accommodating various functional forms for h( ⋅ ) and s( ⋅ ). For s( ⋅ ), several summary statistics can be used to provide a point summary of ligand/receptor gene expression levels across cells in a given cell type. In the default version of FastCCC, we consider four such statistics: mean, median, 3rd quartile, and 90th percentile. These statistics effectively capture diverse cell-cell interaction patterns; for example, the mean expression reflects average expression across cells, capturing interactions that are prevalent in a large fraction of cells, while the 90th percentile is effective in capturing interactions among a small subset of cells with high expression levels. Importantly, these statistics are also applicable when the ligand or receptor is a molecular complex comprising multiple subunits. In such case, FastCCC computes the point summary for each subunit separately and then considers either the minimum or average expression across the subunits as the final s( ⋅ ). The minimum expression is effective in scenarios where all subunits must be expressed for the ligand or receptor complex to function, whereas the average expression among all subunits offers a more lenient assessment that captures the overall activity across the components. Again, while the default version of FastCCC offers minimum and average as two options, its framework is general and can easily incorporate additional alternatives.
For h( ⋅ ), we consider two distinct functional forms. The first form is the arithmetic average of the ligand and receptor expression summaries, represented as
The second form is the geometric average of the ligand and receptor expression summaries, represented as
The arithmetic average is particularly effective in capturing the central tendency of the data and in detecting strong interactions where either the ligand or receptor is highly expressed. In contrast, the geometric average reflects interactions that may scale multiplicatively, providing a better representation of ligand-receptor levels that span several orders of magnitude. This property is particularly important in biological settings, where effective signaling requires the simultaneous high expression of both the ligand and receptor. For example, immune checkpoint interactions, such as PD-L1-PD-1 require both PD-L1 (on tumor or myeloid cells) and PD-1 (on T cells) to be co-expressed at sufficient levels for functional suppression to occur59. Similarly, in Notch-Delta signaling, downstream activation only arises when both neighboring cells express their respective components in tandem60. These cases reflect multiplicative dependencies, for which the geometric mean is better suited than the arithmetic mean. This intuition can also be illustrated through a simple numerical example. Suppose the ligand expression is 0.1 and the receptor expression is 10: The arithmetic mean is (0.1 + 10)/2 = 5.05; The geometric mean is \(\sqrt{0.1\times 10}=1\). Now, if the ligand expression decreases tenfold to 0.01, the arithmetic mean becomes 5.005—a change of less than 1%—while the geometric mean drops to 0.316, a 70% reduction. This example illustrates how the geometric mean penalizes imbalanced expression and more accurately captures the interaction strength that depends on the simultaneous co-expression of ligand and receptor. Such behavior aligns with biological settings where either component being low renders the signaling ineffective (More details in Supplementary Materials, Table S1).
Thanks to the general framework of FastCCC, it includes many previously proposed CS statistics for CCC as special cases. For example, when \(s({x}_{{c}_{a},l})={\bar{x}}_{{c}_{a},l}\) and \(s({x}_{{c}_{b},r})={\bar{x}}_{{c}_{b},r}\) are defined as the average expression measurement of ligand and receptor, respectively, and h( ⋅ ) follows the same format as equation (2), representing the average of these expression measurements, then the CS in equation (1) simplifies to the test statistics used in CPDB. Importantly, the statistical power to detect CCC inevitably depends on how well these chosen functions of h( ⋅ ) and s( ⋅ ) capture the true interaction pattern between ligand and receptor for specific cell type pairs. Unfortunately, the true underlying CCC patterns for any ligand-receptor and cell type pair are unknown and may vary considerably across quadruplets. To ensure robust identification of CCC across various scenarios, we incorporated two distinct h( ⋅ ) functions, four s( ⋅ ) functions for single-unit ligand or receptor, and 2 s( ⋅ ) aggregation functions for ligand or receptor complex. This combination results in a total of 16 scoring methods implemented in FastCCC (detailed below and also Table S4). We fit our model for each combination of h( ⋅ ), s( ⋅ ), and aggregation function, calculate the corresponding p-value, and combine these 16 p-values using the CCT, a robust method for aggregating dependent p-values with analytic type I error control under arbitrary correlation structures61.
Specifically, we compute the Cauchy statistic \({T}_{CCT}={\sum }_{i=1}^{16}{w}_{i}\tan (0.5-{p}_{i})\pi\), where equal weights wi = 1/16 are used by default. The final global p-value is then obtained via the transformation \({p}_{CCT}=1/2-{\tan }^{-1}({T}_{CCT})/\pi\), leveraging the stability of the Cauchy distribution under summation.
The Cauchy rule takes advantage of the fact that the combination of Cauchy random variables also follows a Cauchy distribution regardless of whether these random variables are correlated or not. Therefore, the Cauchy combination rule enables us to combine multiple potentially correlated p-values into a single p-value without compromising type I error control61. This approach enables flexibility in capturing various patterns of interaction based on different expression metrics and interaction models.
Detailed choices of s( ⋅ )
We provide the detailed functional forms for s( ⋅ ) in this section. For a single-unit ligand or receptor, we consider four different choices for s( ⋅ ). The first choice is the mean, represented as
where the subscript g ∈ {l, r} represents a single unit of either ligand or receptor; the subscript c ∈ {ca, cb} represents either cell type ca or cb; nc represents the number of cells belonging to cell type c; xj,g represents the expression level of ligand or receptor for the jth cell in cell type c; and \({\bar{x}}_{c,\cdot }\) represents the average expression level of ligand or receptor for cell type c.
The second to fourth choices for s( ⋅ ) are three order statistics of gene expression, represented as the median, 3rd quartile and 90th percentile from an arbitrary expression distribution. Specifically, for a quantile value q (q = 50, 75, or 90), we compute the kth order statistic in the form of
where \({x}_{c,{g}_{(k)}}\) represents the kth value in the sorted gene expression levels of the ligand or receptor within the corresponding cell type, arranged in ascending order; the value of k is rounded down to determine the rank position to be calculated.
In the setting when either the ligand or receptor is in the form of a multi-subunit complex, the expression summary statistics for the complex is aggregated by taking either the average or the minimum of its subunits. Let ng represent the number of subunits; we have the following two choices to compute the final s( ⋅ ):
As described above, we offer a diverse set of options to provide users with maximum flexibility. However, we emphasize that FastCCC is not intended to require users to select a single formulation. Instead, in its default mode, it computes all 16 combinations of the aforementioned modules. Therefore, for most users—particularly those without prior knowledge of the dominant interaction mechanism—we recommend using the default combined mode. Advanced users may further investigate which scoring scheme is most appropriate for a specific interaction. Additional details are provided in Supplementary Material Section 3 and Table S1.
Convolution-based p-value calculation for C S statistics
After obtaining the above CS statistics, the next step is to test whether the observed CS statistic is as extreme as, or more extreme than, what would be expected under the null hypothesis. Let na denote the number of cells in cell type ca, and nb the number of cells in cell type cb. The null hypothesis is defined based on two cell types with the same sample sizes na and nb, but with ligand and receptor expression randomly drawn from the entire distribution across all cells. The standard approach for computing the corresponding p-value involves randomly sampling na and nb cells from the population, which is achieved practically by permutating cell type labels or shuffling the cell type identities, and then recalculating the CS statistic. This permutation process is repeated at least thousands of times to construct a null distribution of CS statistics. The proportion of CS statistics from the permuted null distribution that exceeds the observed CS gives the p-value. Unfortunately, this permutation-based p-value computation procedure is computationally intensive and can be sensitive to the number of permutations employed.
Here, we take an alternative approach to obtain the distribution of CS statistics under the null using scalable analytic solutions rather than permutation-based numeric solutions. Specifically, we first denote fh( ⋅ ) as the probability density function (PDF) for CS statistics under the null. Our objective is to compute this PDF, which allows us to further compute the p-value corresponding to an observed CS statistic: \(p\,{\mbox{-value}}\,=\int_{t}^{\infty }{f}_{h}(x)dx\) for an observed CS statistic equal to t. To achieve this, we derive our key insights from the fact that fh( ⋅ ) function can be effectively expressed as a convolution of two separate functions. To see this, we first recognize that the first h( ⋅ ) function defined in equation (2) represents the average of the summary levels of ligand and receptor, while the second h( ⋅ ) function defined in equation (3) is similarly the average of the two, but on the log scale. Therefore, for an h( ⋅ ) function defined in equation (2), we have
where \({f}_{{c}_{a},l}(\cdot )\) denotes the PDF describing the distribution of the summary expression level of ligand in cell type ca; \({f}_{{c}_{b},r}(\cdot )\) denotes the PDF describing the distribution of the summary expression level of receptor in cell type cb; and f*g denotes the convolution of two functions, defined as
For an h( ⋅ ) function defined in equation (3), we similarly have
where \({f}_{{c}_{a},l}(\cdot )\) denotes the PDF describing the distribution of the log-scale summary expression level of ligand in cell type ca; and \({f}_{{c}_{b},r}(\cdot )\) denotes the PDF describing the distribution of the log-scale summary expression level of receptor in cell type cb.
Using the above insights, we first obtain the two PDF functions \({f}_{{c}_{a},l}(\cdot )\) and \({f}_{{c}_{b},r}(\cdot )\) separately. These functions depend on the specific choice of s( ⋅ ) and the presence or absence of a multi-subunit complex, as detailed in later sections. Afterwards, we compute the PDF fh( ⋅ ) based on their convolution. In the simplest case where fh( ⋅ ) function defined in equation (2) represents the average of the summary levels of ligand and receptor and where none of the two cell types is rare, this fh( ⋅ ) function can be directly derived based on asymptotic properties and is in the form of a normal distribution. In the more general case, we utilize FFT for computation, accommodating any arbitrary distributional forms and ensuring scalable computation. We note that FastCCC assumes independence between the expression summaries of different components under the null hypothesis. While real biological systems may exhibit some degree of subunit co-regulation, this assumption simplifies derivation and ensures analytical tractability. As demonstrated in the Results (Fig. 2) and Supplementary Material (section 4, Figs. S6–S10), empirical analyses show that most ligand-receptor pairs in real datasets exhibit weak or no correlation, supporting the suitability of this modeling choice.
Probability density functions for single-unit ligand or receptor with different choices of s( ⋅ )
Here, we provide details for obtaining the probability density function fs(x) (s = ca,l or cb,r) for the summary expression level of a single-unit ligand or receptor in a specific cell type. To simplify presentation, we focus on the distribution of the summary expression level of x rather than the log-scale summary expression level of \(\log x\), as the derivations and resulting forms are largely similar.
First, we consider the case where the s( ⋅ ) function represents the expression mean as described in equation (4). For non-rare cell types where the number of cells exceeds 30, we apply the central limit theorem (CLT) to obtain the sampling distribution of the mean as an asymptotical normal distribution. Such normal distribution has mean μg and variance \({\sigma }_{g}^{2}/{n}_{c}\), where μg and \({\sigma }_{g}^{2}\) are the mean and variance of gene expression in all cells regardless of the cell type \(\left(g\in \{l,r\}\right)\), respectively, and nc represents the number of cells in the cell type of focus \(\left(c\in \{{c}_{a},{c}_{b}\}\right)\). For rare cell types where the number of cells does not exceed 30, we cannot apply CLT. Instead, we use convolution to calculate the exact distribution of fs(x). Because observing a mean value of \({\bar{x}}_{c,g}\) is equivalent to observing the corresponding total expression value of \({n}_{c}{\bar{x}}_{c,g}\), we have \({f}_{s}({\bar{x}}_{c,g})={f}_{g}^{{n}_{c}}({n}_{c}{\bar{x}}_{c,g})\), where \({f}_{g}^{{n}_{c}}(\cdot )\) is defined as a convolution across nc cells in the form of
where fg represents the PDF function of expression level for each cell; and ∗ denotes convolution operation defined in equation (9) earlier. We compute such a convolution based on FFT on any arbitrary distribution fg. Specifically, we apply log-transformed normalization to the expression profiles, discretize fg by partitioning the domain into evenly spaced segments with a minimum precision of 0.01, count the frequency of each segment to capture the discrete probability of expression within each segment, and apply FFT to perform convolution and calculate the PDF of fs(x).
Next, we consider the case where the s( ⋅ ) function represents the median, 3rd quartile, or 90th percentile of the expression level. Here, we denote q as the percentile of interest (q=50, 75, or 90) and obtain the corresponding kth order statistic, where k = ⌊q/100 × (nc − 1) + 1⌋. We aim to obtain the PDF for kth order statistic from an arbitrary discrete distribution, regardless of whether the cell type is rare or common. To do so, we denote X1, X2, ⋯ , Xn as n observed expression values for the ligand or receptor in the cell type of focus. We sort them such that X(1) < X(2) < ⋯ < X(n). We denote f(x) as its PDF. The distribution for kth order statistic from the expression level is62:
However, this formulation assumes that the underlying variable is continuous and that all sampled values are distinct. In contrast, single-cell transcriptomic data often exhibit discrete gene expression values with a high proportion of zeros, and repeated observations across cells are common. Therefore, the direct application of Equation (12) is not suitable for such datasets.
To address this limitation, we extended the classical formulation to accommodate discrete-valued distributions with ties, we create a reference sample, denoted as Y1, Y2, ⋯ , Yn, which are n samples randomly drawn from a uniform distribution U(0, 1). We also sort them such that Y(1) < Y(2) < ⋯ < Y(n). The distribution for kth order statistic from this reference distribution can be derived as follows
We connect the expression data to the reference distribution through the cumulative distribution functions (CDF) of X and Y. Specifically, we denote FX and GY as the CDF of X and Y, respectively, with range at [0, 1]. We identify the map function between the two as FX(x) = GY(y). It is straightforward to deduce that
Therefore, we have:
With the above equation, we obtain the cumulative distribution for the kth order statistic for a set of uniform random variables, and map x to y through CDF of the gene expression level to further obtain the cumulative distribution of the PDF for X(k).
To facilitate computation, we discretize the interval [0, 1] evenly into m=100,000 segments, enabling us to compute the CDF for the kth order statistic of a set of uniform random variables. This approach allows us to further calculate the CDF and subsequently the PDF for the summary expression level. In practice, for i = 1, 2, ⋯ , m, we compute \({g}_{{Y}_{(k)}}(i/m)\). For a random variable \(\tilde{Y}\) with a discrete uniform distribution over the possible values i = 1, 2, ⋯ , m, the approximate solution for the CDF of the kth order statistic of \(\tilde{Y}\) is given by:
where Z is the normalization constant used to ensure that the resulting distribution integrates to 1, maintaining the properties of a valid probability distribution. Thus, we can calculate the probability mass function \({p}_{{\tilde{X}}_{(k)}}\) for the kth order statistic of any discrete gene expression random variable \(\tilde{X}\), obtained from the data, with all possible values x1, x2, ⋯ , xl (x1 < x2 < ⋯ < xl). Specifically, we have:
Probability density functions for ligands or receptors with multiple subunits
For ligands or receptors with multiple subunits, we first consider the case where the expression summary is the average of s( ⋅ ) for all subunits as defined in equation (6). In the simplest case where the expression summary for each subunit follows a normal distribution, the distribution for the average is also a normal distribution: the mean equals the average of the means of each subunit and the variance is 1/n2 times the sum of variances of each subunit. In the general case where the expression summary of each subunit follows an arbitrary distribution, the distribution for the average is equivalent to the convolution of ng discrete distributions in the form of
where fc,g( ⋅ ) denotes the PDF for each subunit g (g ∈ {1, ⋯ , ng}) in cell type c.
Next, we consider the case where the expression summary is the minimum of s( ⋅ ) across subunits as defined in equation (7). We obtain the CDF for the minimum of ng random variables from the ng subunits as
where, fs(x) represents the PDF of the summary statistic for each subunit, and Fs(x) is the corresponding CDF. Note that \({F}_{s}^{{\prime} }(x)={f}_{s}(x)\) by definition, the derivative of the CDF recovers the PDF.
Valid candidate LRIs
Before running FastCCC, we perform ligand-receptor pair filtering to focus on a set of valid candidate LRIs for interaction analysis. To do so, we follow existing approaches to filter out genes with a large amount of zero expression in certain cell types when considering them as candidate ligands or receptors. In FastCCC, we use CPDB’s default threshold of 10%, meaning that if a gene’s expression is zero in more than 90% of cells within a given cell type, we no longer consider LRIs involved by that gene for that cell type, and its p-value is set to 1. These LRIs that are involved in the actual computation are referred to as “valid LRIs,” as mentioned in the main section. All of our evaluation results are based on valid LRIs. Additionally, FastCCC provides an option to follow the recommendations of difference-assembly-based tools and use our probability distribution toolkit to directly screen for differentially expressed genes (DEGs). Specifically, FastCCC can be used to estimate the null distribution of gene expression within each cell type cluster, enabling direct calculation of p-values for evaluating expression levels without relying on external DEG detection tools. Unlike other CCC methods (e.g., CellChat, ICELLNET) that require prior filtering of significantly upregulated genes, FastCCC internally evaluates the statistical significance of ligand and receptor expression based on a mathematically principled probabilistic framework. This allows FastCCC to efficiently pinpoint DEGs across specific cell types and further incorporate DEGs as an additional filter for candidate LRIs to ensure a rigorous selection process. In particular, such a filter requires both ligands and receptors to be highly expressed in their respective cell clusters, thereby reducing false signals. Users have the option to choose whether to include DEGs as a selection criterion based on their specific analysis needs. This design offers flexibility, allowing researchers to adapt the toolkit to a wide range of biological and computational scenarios by freely combining different operational modules in a modular modeling framework and analytically deriving the null distribution of the scores.
Reference-based CCC inference
Reference-based CCC analysis in FastCCC requires both query data and reference data. The query data, typically a smaller user-collected dataset, is provided as a raw gene expression count matrix. The reference data, significantly larger in scale, is provided and processed through FastCCC. The reference data includes essential information, such as the reference tissue name, LRIs DB used, the types and numbers of cell types included, the PDFs of gene expression, and some precomputed information necessary for the FastCCC inference workflow. No reference expression profile data or meta info is stored or used in the reference panel, ensuring scalability and conciseness in data representation. To support reference-based CCC analysis, we have constructed a human CCC reference panel, with details provided in the next section.
For both query and reference data, FastCCC performs basic quality control to filter out cells with abnormally low gene expression levels (i.e., cells with < 50 expressed genes). Afterwards, FastCCC applies a rank-based alignment strategy to mitigate sensitivity to batch effects and platform-specific biases between query and reference datasets. This transformation preserves gene expression order while discarding magnitude, enabling scale-invariant comparison across datasets. Similar rank-based representations have also been adopted by recent methods, such as scGPT39 and AUCell63. Specifically, in each dataset, it sorts the non-zero expressed genes in ascending order, divides them evenly into nbin predefined intervals, and assigns each gene a rank based on its interval. The smallest non-zero expression value was assigned a rank of 1, and the maximum value was set to nbin. Genes with zero expression remained as 0.
While the rank-based batch correction procedure is effective, the normalized query and reference data inevitably retain some remaining level of noise. To further improve batch correction, we utilize housekeeping genes40 to estimate the remaining noise and directly align the two datasets. To do so, we assume that the mean expression of each housekeeping gene g in the reference (Xr,g) and query (Xq,g) datasets satisfies
where δ follows a normal distribution with a mean of 0 and a standard deviation proportional to Xr,g:
We estimate the parameter k using all the housekeeping gene means, which are recorded during the reference construction step, and the corresponding values in the query data. k is set to a minimum value (default set to 3) if the estimated value falls below it, mitigating excessive uncertainty.
Once the noise parameter k is estimated, all statistics in the reference data can be mapped to the query domain using properties of a Gaussian distribution. Specifically, a point statistic in the reference, Xref, is mapped to a corresponding value in the query data, denoted as Xquery, as follows:
Importantly, the above equation not only gives out the mapped value Xquery but also provides a confidence interval for the mapped value, allowing us to readily assess uncertainty.
With the above setup, we calculate the CS statistics for each LRI in the query dataset using equations (1), (2), (4), and (6), along with the corresponding ligand and receptor expression summary values s( ⋅ ), with a precision of 0.01. In parallel, we use the null distribution of expression summary of ligands and receptors obtained from the reference data to calculate a CS significance threshold corresponding to a p-value of 0.05. This threshold in the reference data is mapped to the query data using equation (24) to yield both the query data threshold and the 95% confidence interval associated with this mapped threshold. Afterwards, we use the confidence interval to compute the lower and upper bounds associated with the mapped query data threshold. Additionally, we also obtained the null expression summaries of ligand and receptor from reference and mapped them using equation (24). Based on the relationship between the CS values of LRIs in the query dataset and the corresponding mapped CS threshold confidence interval, we considered the following five scenarios to determine significance (a schematic workflow is shown in Fig. S28). 1. LRIs with CS values above the upper bound are considered as significantly communicating. 2. Those with CS values below the lower bound are deemed insignificant. For LRIs with CS values within the threshold interval, we conducted further comparison with the reference to determine significance. First, we check whether the corresponding sender-receiver’s LRI is significant in the reference. 3. If both the sender and receiver cell types exist and the LRI is significant in the reference, the ligand and receptor’s expression summaries in the query dataset are compared with the lower bound of the mapped corresponding null summaries from the reference. And if both intensities exceed the lower bound, the LRI is considered significant. 4. On the other hand, if the LRI is not significant in the reference and the ligand and receptor’s expression summaries in the query dataset do not exceed the upper bound, the LRI is considered insignificant. 5. In cases where the conditions above do not apply, such as when CS falls within the mapped threshold interval while the sender or receiver cell type is absent from the reference, or if ligand and receptor expression comparisons with the reference yield conflicting results (e.g., one increases while the other decreases, leading to a CS value near the threshold) – the significance of the LRI is assessed using query data alone by comparing the CS value to its null distribution.
Through the above process, FastCCC enables reference-based CCC analysis by comparing the query and reference data, enabling the identification of CCC signals that conventional CCC analysis might overlook. Reference-based CCC analysis is particularly valuable in scenarios where such CCCs are systematic and global, rendering them challenging to detect when using the query dataset alone as the background.
Construction of human CCC reference
To facilitate reference-based CCC analysis with FastCCC, we constructed a human CCC reference panel using the CellxGene37 platform (version dated 2024-07-01). Specifically, we first obtained 19 single-cell RNA sequencing datasets from the platform, each representing a different normal human tissue, with each dataset containing at least 150,000 cells (an exception is stomach tissue, which contains over 60,000 cells and is included as a reference due to its frequent use in studies). In the process, we selected “Homo_sapiens” as the species, “normal” as the disease condition, set “is_primary_data” to false to use prefiltered data, and used the “tissue_general” option to select datasets for the following tissues: brain, breast, eye, lung, heart, liver, endocrine gland (or thymus), blood, kidney, respiratory system, spleen, bone marrow, skin of body, small intestine, lymph node, adipose tissue, colon, nose, and prostate gland. For brain tissues, we further refined the selection using the “tissue” option to include two representative regions: the midbrain and the cerebral cortex. Detailed information about the reference data for these different tissues is provided in the Supplementary information. All datasets used HGNC symbols as gene names, which were retained during processing. For cell type information, we followed the CellxGene platform guidelines, using the cell type labels provided by the platform according to the Cell Type Ontology standard as input for FastCCC. Additionally, we employed the LRIs database provided by CPDB v58 as the candidate list for constructing the reference. For each dataset in turn, we conducted rank-based batch correction and normalization desribed in the previous section. We then obtain PDFs and summary information from the data in the form of the mean of the ranked gene expression. We only retained the necessary information required by FastCCC for reference construction, as detailed in the previous section, to ensure scalability and conciseness in the reference. For example, raw count matrices and other dataset-specific information were neither saved nor used during reference-based CCC analysis.
We used the lung dataset from the constructed human CCC reference panel to illustrate reference-based CCC analysis in two distinct case studies. In the first case study, we extracted lobe-related and other positions from the dataset to serve as a query, and used the remaining lung data as the reference (which contains over 860,000 cells). In the second case study, we downloaded supplementary lung normal datasets excluded from our CCC reference panel to serve as the query data, mimicking the smaller size typical of user-collected dataset. We ensured no cell overlap between the reference and query data by using “observation_joinid”, a unique identifier assigned to each cell, which is required when uploading data to the CellxGene platform.
Simulations and benchmarking: data sources, designs, and evaluations
Data sources
We first conducted comprehensive simulations and benchmarking using eleven datasets, covering distinct biological contexts and scenarios. These datasets encompass cell numbers ranging from several thousand to two million, with the number of cell types varying from under five to over 50, thereby covering the majority of scenarios encountered in practical applications. The datasets include:
(1) CPDB tutorial dataset, designated as CPDBTD. This dataset is an example dataset provided by CPDB, used to validate the correctness of our algorithm’s theoretical framework and implementation. It contains 3312 cells across 40 cell types. Together with CPDB and a modified version based on the k-th order statistic, this dataset was used to verify equations ((8)-(21)).
(2) Human Ileum dataset. This dataset is part of a scRNA-seq study measuring human intestine64, especially the ileum segmentation. It contains 16,819 genes and 5980 cells across seven cell types.
(3) Human Fibroblast dataset. This dataset is part of an integrated scRNA-seq analysis of the normal gastrointestinal (GI) tract, esophagus and stomach measured by 10X Genomics65. It contains 33,234 genes and 5754 cells across three cell types.
(4) Human ECGE dataset. This dataset collects snRNA-seq data measured in human entorhinal cortex (EC) and ganglionic eminence (GE) germinal zones66. It contains 21,563 genes and 7618 cells across ten cell types.
(5) Human Breast dataset, designated as BreastB, is a subset of data from the Human Breast Cell Atlas (HBCA) study67. This dataset includes B cells and comprises 33,234 genes and 12,510 cells across five cell types.
(6) Human HCA Kidney dataset, designated as Kidney. This dataset contains scRNA-seq data from normal regions of kidney tumor-nephrectomy samples collected from 14 individuals68. It contains 37,073 genes and 48,783 cells across 28 cell types.
(7) Human AnG dataset, designated as Angular. This dataset is part of an snRNA-seq study using SMART-seqv4 RNA-sequencing that measures eight cortical areas in the human brain69. It focuses on the angular gyrus (AnG) area and includes 20,981 genes and 110,752 cells across 18 cell types.
(8) Human Fetal Retina dataset, designated as Retina. This dataset comprises single-nucleus multiome data from the developing human retina, obtained from 12 donors using 10X Chromium Single-Cell Multiome ATAC + RNA-seq70. Our study uses only snRNA-seq data, which includes 36,503 genes and 205,619 cells across eight cell types.
(9) Human GBmap dataset. This dataset consists solely of scRNA-seq data obtained via 10X Genomics, covering a large-scale measurement of 240 patients diagnosed with glioblastoma71. The dataset consists of 28,045 genes and 338,564 cells across 17 cell types.
(10) Human AIDA (Asian Immune Diversity Atlas) dataset. This dataset includes scRNA-seq data obtained via 10X Genomics, encompassing 503 healthy donors from seven population groups across five countries72. It contains 36,266 genes and 1,058,909 cells across 33 cell types. A subset version was also used for running CPDB. For this analysis, we focused on 2000 highly variable genes (HVGs), designated as AIDA_hvg.
(11) Human HLCA (integrated Human Lung Cell Atlas) dataset. This dataset integrates 49 studies collected from the human respiratory system into a large-scale atlas data73. It consists of 56,295 genes and 2,282,447 cells across 51 cell types. Similarly, CPDB was applied to 2000 HVGs, designated as HLCA_hvg.
Besides the above 11 datasets, we also used another 11 datasets for benchmarking our human CCC reference panel. The datasets include:
(1) Disease samples from MDCs altas: this dataset integrates 73,872 cells and 11 cell types. It is a partial dataset from the multi-tissue myeloid-derived cells single-cell atlas of tumor and healthy samples developed by Boroni et al.74 We filtered cells from lung tissue with a lung cancer phenotype for testing, and this dataset is therefore referred to as Boroni et al.’s lung cancer dataset.
(2) Disease samples from COVID-19 dataset: this dataset is the same one used in our case study when we applied FastCCC to large-scale COVID-19 data18. We only used data from the lungs of critically ill patients, containing 42,757 cells and 19 cell types. We designated this data as Zhang et al.’s COVID-19.
(3) Disease samples from Human HLCA dataset: this dataset consists of single-cell RNA sequencing data from various lung-related diseases in the HLCA73, including pulmonary fibrosis, lung adenocarcinoma, and 13 other diseases. Each disease is treated as a separate query dataset. For consistency in naming, in the reference benchmarking study, we used the format “Author + Disease Name,” for example, “Luecken et al.’s cystic fibrosis.”
(4) Normal samples from MDCs atlas: this dataset contains 87,370 cells and 11 cell types. It is a partial dataset from the MDCs atlas mentioned earlier. We filtered cells from normal lung tissue for testing, and we referred to this data as Boroni et al.’s dataset.
(5) Normal samples from LungMAP: this dataset is a large-scale snRNA-seq dataset of the human lung from healthy donors of approximately 30 weeks, 3 years, and 30 years of age75. We used the healthy samples as the query for testing, and it contains 46,500 cells and 27 cell types. It is referred to as Wang et al.’s dataset.
(6) Normal lung samples from HCA: this dataset consists of 39,778 cells and nine cell types derived from normal lung tissue76. It is referred to as Eils et al.’s dataset.
(7) Normal samples from Human HLCA dataset: this dataset comprises the normal samples from the HLCA, containing over 333,468 cells. It is designated as Luecken et al.’s dataset.
(8) Normal lung samples from Tabula Sapiens: Tabula Sapiens is a benchmark first-draft human cell atlas comprising over 1.1 million cells from 28 organs of 24 normal human subjects77. For our analysis, we used the normal lung samples from this atlas, containing 65,847 cells and 34 cell types, referred to as Pisco et al.’s dataset.
(9) Normal lung samples from CellHint: this dataset, organized by Teichmann et al., encompasses 12 tissues from 38 datasets, forming a meticulously curated cross-tissue database with approximately 3.7 million cells78. We extracted 318,426 cells from normal lung samples for our analysis. To differentiate it from another dataset curated by the same group, we refer to this dataset as Teichmann et al.’s 1.
(10) Normal samples from human lung immune cells: this dataset profiled human embryonic and fetal lung immune cells using scRNA-seq, smFISH, and immunohistochemistry79. For our analysis, we retained normal samples, which include 670,749 cells. As this dataset is also organized by Teichmann et al., we refer to it as Teichmann et al.’s 2.
(11) Primary breast tumor atlas: this dataset integrated a single-cell RNA sequencing atlas of the primary breast tumor microenvironment containing 236,363 cells from 119 biopsy samples across eight datasets42. We utilized it as a case study to explore the outcomes of reference-based CCC analysis using the human breast CCC reference.
Simulation and benchmarking design
We designed simulations and benchmarking for method comparison on each dataset described in the previous section. All statistical comparisons of performance metrics between different methods were conducted using a two-sided Mann-Whitney U-test to assess the significance of differences. This approach was applied consistently across all subsequent experiments in this study, unless otherwise specified. We considered three distinct sets of comparisons:
(1) Method evaluation. We first validated the analytic solution provided by FastCCC with the permutation-based approach CPDB on the same CS used in CPDB. We used the same log-transformed transcriptomic expression data, the same version of the LRIs database, and the same set of valid LRIs described in the earlier section for both methods. We conducted the evaluation experiments using a standard Linux server running Ubuntu 20.04, equipped with an Intel(R) Xeon(R) CPU E5-2620 v2 and 250GB of memory.
In the comparison, we kept the parameters used in FastCCC consistent with those used in the default setting of CPDB. For example, FastCCC also used the same mean function for ligand and receptor expression summary statistics, the same minimum function for multi-subunit complexes, and the same arithmetic mean as used in CPDB. All other CPDB parameters were set to their defaults, and we varied the number of permutations used in CPDB (default: 1000) to examine their influence on CPDB’s final results. In this benchmarking analysis, CPDB results obtained using one million permutations were treated as the ground truth to validate the theoretical derivation and software implementation of FastCCC, as well as CPDB results generated with varying numbers of permutations. We calculated precision and IoU for the results generated by FastCCC and CPDB and assessed the correlation between the p-values produced by the two methods.
(2)Simulation comparison. We compared the performance of all methods using the test datasets described earlier. For each dataset, we utilized the raw gene expression count matrix as input and applied a semi-simulated approach, where the ground truth is known, following Liu et al.16.
We began by randomly shuffling the cell type labels, ensuring that all LRIs were null and no interaction signals were present. We then randomly selected one cell type pair to exhibit LRIs in a specific direction, with ligands in cell type A communicating to receptors in cell type B. For the selected cell type pair, 30 LRIs were randomly chosen from the candidate LRIs database to serve as significant LRIs, where the expression level of cells from the interacting cell type pair was multiplied by a factor of two. Given the inherent sparsity of the data, we identified cells with zero expression counts for either the ligand or receptor within the interacting cell type pair. From these, 60% were randomly selected, and their zero expression values were replaced. The replacement value was computed by first taking the mean of five randomly sampled non-zero gene expression values and then multiplying it by a random factor drawn from a uniform distribution (0.6, 1.2). This adjustment ensured that the ligand and receptor expression levels for the interacting cell type pair were higher than those for a randomly chosen cell type pair. Following these modifications, log1p-normalization was performed, and the normalized counts were used as input for all compared CCC tools (this normalization step was skipped for methods requiring a count matrix as input). Ten simulation replicates were conducted for each dataset.
All methods were executed on a high-performance computing cluster with Ubuntu 20.04 and Intel(R) Xeon(R) CPU E5-2683 v3. Tasks were submitted via Slurm, with each node allocating 500 GB of memory. Because some methods rely on built-in LRI databases that are difficult to replace, we applied all methods to the intersection of the LRI database used by each method.
(3)Reference-based CCC benchmarking. Here, we evaluated the performance of FastCCC in reference-based CCC analysis using the human CCC reference, comparing it with conventional CCC analysis that relies solely on the user-provided query dataset. Here, we treated the results in the conventional CCC analysis as the ground truth in testing scenarios with biased data subsets.
Evaluation metrics
We evaluated the performance of different methods using multiple evaluation metrics. Specifically, based on the ground truth, we first calculated true positive (TP), true negative (TN), false positive (FP) and false negative (FN). Here, TP represents the number of true LRIs correctly detected; FN represents the number of true LRIs not detected; FP represents the number of false interactions detected; and TN represents the number of false interactions not detected. Afterwards, we calculated nine distinct evaluation metrics including accuracy, precision, recall, specificity, macroF1, balanced accuracy, Jaccard index (or intersection over union), Mathew’s correlation coefficient (MCC), and false positive rate (FPR), detailed below:
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used in this study are publicly available. Most of the datasets used for case analysis and comparative experiments in this study were downloaded from the CellxGene37 platform (https://cellxgene.cziscience.com/). The specific datasets and their sources are as follows: CPDBTD is available at https://github.com/ventolab/CellphoneDB/blob/v5.0.1/notebooks/data_tutorial.zip. Ileum is available from https://cellxgene.cziscience.com/collections/ff668d5d-5b3f-49ee-a007-ff0664bf35ec. Fibroblast dataset is available from https://cellxgene.cziscience.com/collections/a18474f4-ff1e-4864-af69-270b956cee5b. ECGE is available from https://cellxgene.cziscience.com/collections/cae8bad0-39e9-4771-85a7-822b0e06de9f. BreastB is available from https://cellxgene.cziscience.com/collections/4195ab4c-20bd-4cd3-8b3d-65601277e731. Kidney dataset is available at https://explore.data.humancellatlas.org/projects/29ed827b-c539-4f4c-bb6b-ce8f9173dfb7. Angular is available from https://cellxgene.cziscience.com/collections/d17249d2-0e6e-4500-abb8-e6c93fa1ac6f. Retina is available from https://cellxgene.cziscience.com/collections/5900dda8-2dc3-4770-b604-084eac1c2c82. GBmap is available from https://cellxgene.cziscience.com/collections/999f2a15-3d7e-440b-96ae-2c806799c08c. AIDA is available at https://cellxgene.cziscience.com/collections/ced320a1-29f3-47c1-a735-513c7084d508. HLCA is available at https://cellxgene.cziscience.com/collections/6f6d381a-7701-4781-935c-db10d30de293. Disease and normal samples of MDCs atlas can be downloaded through https://cellxgene.cziscience.com/collections/3f7c572c-cd73-4b51-a313-207c7f20f188. LungMAP is available at https://cellxgene.cziscience.com/collections/625f6bf4-2f33-4942-962e-35243d284837. Normal lung samples from HCA is available at https://explore.data.humancellatlas.org/projects/58028aa8-0ed2-49ca-b60f-15e2ed5989d5. Tabula Sapiens is available at https://tabula-sapiens.sf.czbiohub.org/, and we use the normal lung sample. Two normal lung datasetes organized by Teichmann et al. is available at https://cellxgene.cziscience.com/collections/854c0855-23ad-4362-8b77-6b1639e7a9fcand https://cellxgene.cziscience.com/collections/ec329aed-22bc-4d6e-8935-8282dcb1acac. Primary breast tumor atlas is available at https://zenodo.org/records/10672250. For the LRI database, we used version 5.0.0 provided by CellPhoneDB as the primary data source for CCC analysis case studies and reference construction. This database can be downloaded from https://github.com/ventolab/CellphoneDB/blob/master/notebooks/T0_DownloadDB.ipynb. Additionally, version 4.1.0 of the database was also downloaded and used specifically for method comparisons paired with CellPhoneDB v4. In the reference-based breast cancer CCC analysis, we also utilized the LRI database from NicheNet v1.1.1 to maintain alignment with the original study. The candidata LRI pairs is extracted from source code at https://github.com/saeyslab/nichenetr/releases/tag/v1.1.1. Source data are provided with this paper.
Code availability
The code used to develop the model, perform the analyses and generate results in this study is publicly available and has been deposited in Github at https://github.com/Svvord/FastCCC,under the MIT license. Codes for the version of FastCCC used in this paper are also deposited at Zenodo80(https://doi.org/10.5281/zenodo.17329122).
References
Armingol, E., Officer, A., Harismendy, O. & Lewis, N. E. Deciphering cell-cell interactions and communication from gene expression. Nat. Rev. Genet. 22, 71–88 (2021).
Su, J. et al. Cell-cell communication: new insights and clinical implications. Signal Transduct. Target. Ther. 9, 196 (2024).
Armingol, E., Baghdassarian, H. M. & Lewis, N. E. The diversification of methods for studying cell-cell interactions and communication. Nat. Rev. Genet. 25, 381–400 (2024).
Alberts, B. et al. Essential Cell Biology (Garland Science, 2015).
O’Shea, J. J., Holland, S. M. & Staudt, L. M. Jaks and stats in immunity, immunodeficiency, and cancer. N. Engl. J. Med. 368, 161–170 (2013).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Garcia-Alonso, L. et al. Single-cell roadmap of human gonadal development. Nature 607, 540–547 (2022).
Troulé, K. et al. CellPhoneDB v5: inferring cell–cell communication from single-cell multiomics data. Nat. Protoc. https://doi.org/10.1038/s41596-024-01137-1 (2025).
Jin, S. et al. Inference and analysis of cell-cell communication using cellchat. Nat. Commun. 12, 1088 (2021).
Noël, F. et al. Dissection of intercellular communication using the transcriptome-based framework icellnet. Nat. Commun. 12, 1089 (2021).
Cabello-Aguilar, S. et al. Singlecellsignalr: inference of intercellular networks from single-cell transcriptomics. Nucleic acids Res. 48, e55–e55 (2020).
Zhang, Y. et al. Cellcall: integrating paired ligand–receptor and transcription factor activities for cell–cell communication. Nucleic Acids Res. 49, 8520–8534 (2021).
Browaeys, R., Saelens, W. & Saeys, Y. Nichenet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Wang, Y. et al. Italk: an R package to characterize and illustrate intercellular communication. BioRxiv 507871 (2019).
Raredon, M. S. B. et al. Computation and visualization of cell–cell signaling topologies in single-cell systems data using connectome. Sci. Rep. 12, 4187 (2022).
Liu, Z., Sun, D. & Wang, C. Evaluation of cell-cell interaction methods by integrating single-cell RNA sequencing data with spatial information. Genome Biol. 23, 218 (2022).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-seq data. Nat. Commun. 13, 3224 (2022).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913 (2021).
Park, J.-E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Jackson, C. B., Farzan, M., Chen, B. & Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 23, 3–20 (2022).
Hoffmann, M. et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 181, 271–280 (2020).
Zheng, M. ACE2 and COVID-19 Susceptibility and Severity. Aging Dis. 13, 360 (2022).
Woodall, M. N. J. et al. Age-specific nasal epithelial responses to SARS-CoV-2 infection. Nat. Microbiol. 9, 1293–1311 (2024).
Stewart, C. A. et al. Lung cancer models reveal SARS-CoV-2-induced EMT contributes to COVID-19 pathophysiology (2020).
Hertle, E., van Greevenbroek, M. M. J. & Stehouwer, C. D. A. Complement C3: an emerging risk factor in cardiometabolic disease. Diabetologia 55, 881–884 (2012).
Risitano, A. M. et al. Complement as a target in COVID-19? Nat. Rev. Immunol. 20, 343–344 (2020).
Gralinski, L. E. et al. Complement activation contributes to severe acute respiratory syndrome coronavirus pathogenesis. mBio 9, e01753–18 (2018).
Li, G. et al. Coronavirus infections and immune responses. J. Med. Virol. 92, 424–432 (2020).
Mastellos, D. C., Ricklin, D. & Lambris, J. D. Clinical promise of next-generation complement therapeutics. Nat. Rev. Drug Discov. 18, 707–729 (2019).
Murphy, K. & Weaver, C.Janeway’s immunobiology (Garland Science, 2016).
Tong, Q., Yao, L., Su, M., Yang, Y.-G. & Sun, L. Thymocyte migration and emigration. Immunol. Lett. 267, 106861 (2024).
Uehara, S., Grinberg, A., Farber, J. M. & Love, P. E. A role for ccr9 in t lymphocyte development and migration. J. Immunol. 168, 2811–2819 (2002).
Schwarz, B. A. et al. Selective thymus settling regulated by cytokine and chemokine receptors. J. Immunol. 178, 2008–2017 (2007).
Zlotoff, D. A. et al. CCR7 and CCR9 together recruit hematopoietic progenitors to the adult thymus. Blood J. Am. Soc. Hematol. 115, 1897–1905 (2010).
Takada, T. et al. Roles of the complex formation of SHPS-1 with SHP-2 in insulin-stimulated mitogen-activated protein kinase activation. J. Biol. Chem. 273, 9234–9242 (1998).
Lotfollahi, M., Hao, Y., Theis, F. J. & Satija, R. The future of rapid and automated single-cell data analysis using reference mapping. Cell 187, 2343–2358 (2024).
CZI Single-Cell Biology Program et al. CZ CELL×GENE discover: a single-cell data platform for scalable exploration, analysis and modeling of aggregated data 53, 886–900 (2023).
Shang, L. et al. Genetic architecture of gene expression in European and African Americans: an EQTL mapping study in Genoa. Am. J. Hum. Genet. 106, 496–512 (2020).
Cui, H. et al. SCGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1470–1480 (2024).
Eisenberg, E. & Levanon, E. Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Joshi, C. J., Ke, W., Drangowska-Way, A., O’Rourke, E. J. & Lewis, N. E. What are housekeeping genes? PLoS Comput. Biol. 18, e1010295 (2022).
Xu, L. et al. A comprehensive single-cell breast tumor atlas defines epithelial and immune heterogeneity and interactions predicting anti-PD-1 therapy response. Cell Rep. Med. 5, 101511 (2024).
Liu, Y. et al. Role of nectin-4 protein in cancer. Int. J. Oncol. 59, 1–14 (2021).
Janiszewska, M., Primi, M. C. & Izard, T. Cell adhesion in cancer: beyond the migration of single cells. J. Biol. Chem. 295, 2495–2505 (2020).
Li, D.-M. & Feng, Y.-M. Signaling mechanism of cell adhesion molecules in breast cancer metastasis: potential therapeutic targets. Breast Cancer Res. Treat. 128, 7–21 (2011).
He, Y. et al. Targeting PI3K/AKT signal transduction for cancer therapy. Signal Transduct. Target. Ther. 6, 425 (2021).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Bao, Y. et al. Transcriptome profiling revealed multiple genes and ECM-receptor interaction pathways that may be associated with breast cancer. Cell. Mol. Biol. Lett. 24, 1–20 (2019).
Yom, C. K. et al. Clinical significance of annexin A1 expression in breast cancer. J. Breast Cancer 14, 262 (2011).
Moraes, L. A. et al. Annexin-A1 enhances breast cancer growth and migration by promoting alternative macrophage polarization in the tumour microenvironment. Sci. Rep. 7, 17925 (2017).
Yee, K. O. et al. The effect of thrombospondin-1 on breast cancer metastasis. Breast Cancer Res. Treat. 114, 85–96 (2009).
Su, C. et al. Integrinβ-1 in disorders and cancers: molecular mechanisms and therapeutic targets. Cell Commun. Signal. 22, 71 (2024).
Zhang, X.-X., Luo, J.-H. & Wu, L.-Q. Fn1 overexpression is correlated with unfavorable prognosis and immune infiltrates in breast cancer. Front. Genet. 13, 913659 (2022).
Zheng, X. et al. Adam17 promotes breast cancer cell malignant phenotype through EGFR-PI3K-akt activation. Cancer Biol. Ther. 8, 1045–1054 (2009).
Mattern, J. et al. Adam15 mediates upregulation of claudin-1 expression in breast cancer cells. Sci. Rep. 9, 12540 (2019).
Hong, S. et al. Nampt suppresses glucose deprivation-induced oxidative stress by increasing NADPH levels in breast cancer. Oncogene 35, 3544–3554 (2016).
Almet, A. A., Cang, Z., Jin, S. & Nie, Q. The landscape of cell–cell communication through single-cell transcriptomics. Curr. Opin. Syst. Biol. 26, 12–23 (2021).
Cesaro, G. et al. Advances and challenges in cell–cell communication inference: a comprehensive review of tools, resources, and future directions. Brief. Bioinforma. 26, bbaf280 (2025).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer cell 27, 450–461 (2015).
Bray, S. J. Notch signalling: a simple pathway becomes complex. Nat. Rev. Mol. Cell Biol. 7, 678–689 (2006).
Liu, Y. & Xie, J. Cauchy combination test: a powerful test with analytic p-value calculation under arbitrary dependency structures. J. Am. Stat. Assoc. 115, 393–402 (2020).
David, H. & Nagaraja, H. Order statistics 297 (Wiley, 2003).
Aibar, S. et al. Scenic: single-cell regulatory network inference and clustering. Nat. methods 14, 1083–1086 (2017).
Wang, Y. et al. Single-cell transcriptome analysis reveals differential nutrient absorption functions in human intestine. J. Exp. Med. 217, e20191130 (2019).
Nowicki-Osuch, K. et al. Single-cell RNA sequencing unifies developmental programs of esophageal and gastric intestinal metaplasia. Cancer Discov. 13, 1346–1363 (2023).
Nascimento, M. A. et al. Protracted neuronal recruitment in the temporal lobes of young children. Nature 626, 1056–1065 (2024).
Kumar, T. et al. A spatially resolved single-cell genomic atlas of the adult human breast. Nature 620, 181–191 (2023).
HCA seed network precise tumor-nephrectomy samples. https://explore.data.humancellatlas.org/projects/29ed827b-c539-4f4c-bb6b-ce8f9173dfb7.
Jorstad, N. L. et al. Transcriptomic cytoarchitecture reveals principles of human neocortex organization. Science 382, eadf6812 (2023).
Chen, R. et al. Single cell multiome atlas of the human developing retina. Nat. Commun. 15, 6792 (2023).
Ruiz-Moreno, C. et al. Harmonized single-cell landscape, intercellular crosstalk and tumor architecture of glioblastoma. BioRxiv 2022–08 (2022).
Kock, K. H. et al. Single-cell analysis of human diversity in circulating immune cells. bioRxiv 2024–06 (2024).
Sikkema, L. et al. An integrated cell atlas of the lung in health and disease. Nat. Med. 29, 1563–1577 (2023).
Guimarães, G. R. et al. Single-cell resolution characterization of myeloid-derived cell states with implication in cancer outcome. Nat. Commun. 15, 5694 (2024).
Wang, A. et al. Single-cell multiomic profiling of human lungs reveals cell-type-specific and age-dynamic control of sars-cov2 host genes. Elife 9, e62522 (2020).
Lukassen, S. et al. SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J. 39, e105114 (2020).
Consortium, T. T. S. et al. The tabula sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Xu, C. et al. Automatic cell-type harmonization and integration across human cell atlas datasets. Cell 186, 5876–5891 (2023).
Barnes, J. L. et al. Early human lung immune cell development and its role in epithelial cell fate. Sci. Immunol. 8, eadf9988 (2023).
Hou, S. & Ma, W. FastCCC: a permutation-free framework for scalable, robust, and reference-based cell-cell communication analysis in single cell transcriptomics studies. Zenodo https://doi.org/10.5281/zenodo.17329122 (2025).
Acknowledgements
This study was supported by the National Institutes of Health (NIH) grants R01GM126553, R01HG011883, R01HG009124, and R01GM144960 (all to X.Z.).
Author information
Authors and Affiliations
Contributions
X.Z. and S.H. designed this project. S.H. and X.Z. contributed key ideas. S.H. conceived the algorithm and developed the software. W.M. and S.H. compared the proposed algorithm with existing methods to evaluate its performance. S.H. performed data analysis and interpreted data. W.M. contributed to the literature review and the collection of experimental validation data. S.H. drafted the manuscript with input from all authors and contributed to the creation and formatting of the figures. X.Z. supervised the project, initiated the work, wrote and edited the manuscript. And all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks Jae Kyoung Kim, Vassili Soumelis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hou, S., Ma, W. & Zhou, X. FastCCC: a permutation-free framework for scalable, robust, and reference-based cell-cell communication analysis in single cell transcriptomics studies. Nat Commun 16, 11428 (2025). https://doi.org/10.1038/s41467-025-66272-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66272-z