Abstract
Western equine encephalitis virus (WEEV), a group of encephalitic alphaviruses that cause severe diseases in humans and equids, historically used the very-low-density lipoprotein receptor (VLDLR) as a receptor during infection. However, current epidemic strains no longer use VLDLR as a receptor. In this study, we identify that LA1, LA2, LA3, and LA5 of VLDLR can directly interact with WEEV. Using cryo-electron microscopy, we investigate the structures of complexes formed between WEEV and VLDLR-LBD or other VLDLR fragments. Our findings show that LA1 and LA2 insert into a cleft formed by two adjacent E2-E1 heterodimers within a single trimeric spike, while LA3 and LA5 interact with the DIII region of WEEV E1. Among VLDLR concatemers, the LA1-5 exhibits the strongest binding affinity for WEEV. Additionally, we find that a single polymorphism in the E2 glycoprotein determines WEEV’s receptor tropism. Mutations E2E181K or E2E81K in the nonpathogenic strain Imperial-181 enhanced its ability to enter via VLDLR. These results enhance our understanding of alphavirus receptor recognition and receptor usage shifts, providing insights for the development of antiviral therapies.
Similar content being viewed by others
Introduction
Alphaviruses are a group of positive-sense single-stranded RNA viruses transmitted by arthropods, which can cause musculoskeletal disease or encephalitis in humans1,2. Encephalitic alphaviruses, such as Eastern Equine Encephalitis Virus (EEEV), Venezuelan Equine Encephalitis Virus (VEEV), and Western Equine Encephalitis Virus (WEEV), cause severe neurological syndromes leading to long-term sequelae or death in both humans and equines. In contrast, arthritic alphaviruses, including Chikungunya Virus (CHIKV), Ross River Virus (RRV), Mayaro virus (MAYV), and O’nyong’nyong virus (ONNV), cause acute or chronic arthritis3,4. WEEV is an important causative agent of viral encephalitis in humans and equids, causing symptoms ranging from mild flu-like illnesses to encephalitis and death5,6. Notably, human WEEV infections can occur directly via aerosol transmission, leading to more severe encephalitis than infection via the subcutaneous route7,8. Encephalitis can result in permanent neurological sequelae, including seizures, paralysis, and intellectual disability3. Despite the significant health burden, the U.S. Food and Drug Administration has not approved any vaccines or antiviral drugs to combat WEEV9.
The alphavirus genome (~12 kb) encodes four non-structural proteins (nsP1–nsP4) for viral replication and five structural proteins (capsid, E3, E2, 6K/TF, and E1) for virus assembly10. The 70 nm viral surface comprises 80 trimeric E2-E1 heterodimer spikes in a T = 4 icosahedral symmetric arrangement11,12. The 240 E1/E2 heterodimers are arranged as 80 trimer spikes on icosahedral 3-fold (i3) and quasi-3-fold (q3) axes of symmetry. Each asymmetric unit (ASU) comprises a complete q3 trimer and an i3 E1/E2 heterodimer11,13,14,15,16,17,18. The E1 protein adopts a fusion loop responsible for cell fusion13, while the E2 protein participates in receptor binding19,20,21,22.
A key step in the alphavirus infection cycle is the engagement with a host receptor that facilitates cell entry23. For example, matrix remodeling-associated protein 8 (MXRA8), a molecule containing an immunoglobulin (Ig)-like domain, is a receptor for several members of the SFV complex (e.g., CHIKV, RRV, MAYV, and ONNV)24. Further studies revealed that low-density lipoprotein receptor class A domain-containing protein 3 (LDLRAD3), very-low-density lipoprotein receptor (VLDLR), apolipoprotein E receptor 2 (ApoER2), and low-density lipoprotein receptor (LDLR) are receptors for multiple alphaviruses25,26,27. For WEEV strains, the mammalian MXRA8 was identified as not a receptor for WEEV28. In contrast, WEEV can use the avian MXRA8 as a receptor in avian reservoirs28. A recent study has shown that protocadherin-10 (PCDH10) is a general mammalian receptor engaged by multiple WEEV strains, and WEEV group A strains can use mammalian VLDLR and ApoER2 as receptors29,30,31. Receptors from the LDL receptor family have similar small LDLR class A (LA) repeat sequences in their extracellular structural domains. The ligand-binding domain (LBD), consisting of 8 LDLR type A (LA) repeats, is necessary and sufficient to mediate infection31. Previous works have also shown how receptors VLDLR interact with E2-E1 glycoproteins of SFV and EEEV32,33,34. In two recent studies35,36, the interaction sites of WEEV McMillan and WEEV Fleming with VLDLR-LA1-2 were reported. However, LA3 and LA5 can also mediate WEEV infection, and the mechanisms of WEEV recognition of LA3 and LA5 remain unclear. Furthermore, the precise mechanisms by which WEEV recognizes VLDLR and its multiple binding patterns are not fully understood.
Here, we conducted detailed interaction analyses and cryo-electron microscopy reconstructions of VLDLR-LBD, VLDLR-LA1-3, VLDLR-LA3, and VLDLR-LA5 in complex with WEEV strain California VLPs and mapped all LA repeats interacting with WEEV California VLPs and revealed their binding sites on WEEV. Additionally, we find that a single polymorphism in the E2 glycoprotein determines WEEV’s receptor tropism. These findings enhance our understanding of the mechanisms of WEEV recognition of VLDLR and its multivalent binding patterns, providing a theoretical foundation for alphavirus receptor recognition and viral therapy.
Results
Cryo-EM structure of VLDLR in complex with WEEV
To elucidate the structural basis of WEEV recognition of VLDLR, we produced WEEV strain California VLPs complex with human VLDLR LBD (LA1-8) fused with a C-terminal human antibody constant region tag (VLDLRLBD-Fc). A total of 4869 cryo-electron micrographs for WEEV in complex with VLDLRLBD-Fc were used for further reconstruction (Supplementary Table 1). Single-particle reconstructions with icosahedral symmetry yielded the nominal resolutions of 5.09 Å using 19,830 particles for WEEV VLPs in complex with VLDLRLBD-Fc. To enhance the densities, we applied symmetry expansion, block-based classification, and local refinement focusing on the q3 spikes, achieving a final resolution of 3.27 Å for the complex (Fig. 1a, b, e; Supplementary Figs. 1, 2, and 3).
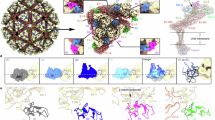
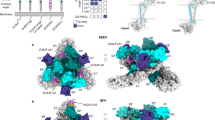
The colored surface representations of cryo-EM maps of WEEV VLPs in complex with VLDLR-LBD (a) or WEEV VLPs alone (c). The densities of WEEV E1 and E2 proteins are colored pale cyan and light blue, whereas the densities of VLDLR-LAs are colored red. The 5-fold, 3-fold, and 2-fold icosahedral axes are indicated by a pentagon, triangles, and an oval, respectively. The i3 spike and q3 spike are indicated by the labels “i3” and “q3” respectively. Cryo-EM maps achieved by the final refinement focusing on the q3 spike in WEEV VLPs in complex with VLDLR-LBD (b) or WEEV VLPs alone (d). e An atomic model of a single E1/E2 heterodimer with non-descript LA domains docked into the experimental electron density map.
Compared to WEEV California VLP alone (Fig. 1c, d), additional densities comparable in size to LA repeats are evident in the q3 spike structure of the WEEV-VLDLR complex (Fig. 1a, b). We fitted the structures of two consecutive LAs into the cryo-EM density (Fig. 1e). The fitted density indicates that the two consecutive LAs are positioned within the cleft formed between the E1/E2 heterodimers and on the tilted scaffold of the E2 A domain.
Mapping of WEEV recognition of VLDLR LAs
To identify which LA repeats interact with the WEEV strain of California, we generated human VLDLR LBD (VLDLRLBD-Fc) and each individual LA repeat (LA-Fc). We then used bio-layer interferometry (BLI) and enzyme-linked immunosorbent assay (ELISA) to measure the direct interactions between each single LA-Fc and WEEV VLPs. The BLI results showed that LA1-Fc, LA2-Fc, LA3-Fc, and LA5-Fc exhibited strong binding affinities for the WEEV strain California, with KD values of 0.11, 1.59, 6.09, and 0.37 nM, respectively (Fig. 2a, b and Supplementary Table 2). The ELISA results and cell-based assays were consistent with the BLI findings (Supplementary Figs. 4 and 5a, b). In contrast, other LA repeats (LA4-Fc, LA6-Fc, LA7-Fc, and LA8-Fc) showed no detectable binding to the WEEV strain California (Fig. 2a, b and Supplementary Figs. 4 and 5a, b).
a Biolayer interferometry binding analysis of individual LAs to WEEV California VLPs. The displayed results were obtained from a single technical replicate and are representative of three biologically independent experiments. The equilibrium dissociation constant (KD) was calculated as the mean ± standard deviation (SD; n = 3). b A schematic diagram showing the ectodomains of VLDLR. The LA repeats are colored according to their binding affinity to WEEV California VLPs.
Structure of VLDLR LA1-3 binding with WEEV
To further investigate the detailed interactions between VLDLR and WEEV VLPs, we selected the LA1-3-Fc construct, which has high binding affinity for WEEV VLPs, for structural analysis. A total of 6073 cryo-electron micrographs of WEEV VLPs in complex with LA1-3-Fc were used for further reconstruction (Supplementary Fig. 1 and Supplementary Table 1). We determined the cryo-EM structure of LA1-3-Fc bound to WEEV q3 spikes at a resolution of 3.03 Å (Fig. 3a, b and Supplementary Figs. 1, 2, and 3). By using the cryo-EM densities as a guide, we built residues 32–110 of LA1-2 in the complex. The models of WEEV E1 (residues 1–439) and E2 (residues 5–419) were also built in the densities.
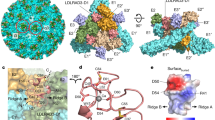
a The colored surface representations of cryo-EM maps of WEEV VLPs in complex with VLDLR-LA1-3. b Cryo-EM maps achieved by the final refinement focusing on the q3 spike in WEEV VLPs in complex with VLDLR-LA1-3. The densities are shown as a colored surface in two perpendicular views. c Ribbon and surface representations depict structural details of the binding interface between LA1-2 (shown in red) and WEEV VLPs. Electrostatic surface potential highlights complementary charge complementarity at the binding interface. Details of interactions of the WEEV VLPs with VLDLR LA1 (d) and LA2 (e). All polypeptides are shown as cartoon diagrams. Interacting residues are displayed as colored sticks with labels. The subscripts of the labels indicate the proteins to which the residues belong. Dashed lines denote the hydrogen bonds or salt bridges. For note, some interactions are not involved in these panels due to clear views and representations. Cryo-EM maps achieved by the final refinement focusing on the q3 spike in WEEV VLPs in complex with VLDLR-LA3 (f) and VLDLR-LA5 (g). The densities are shown as a colored surface in two perpendicular views.
In the structure, the two LA repeats are positioned within the cleft formed between the E2–E1 heterodimers within a single trimeric spike. The EF-like motifs of LA1 and LA2 are calcium-binding motifs, and the calcium ions are chelated by residues in these motifs. The density for the calcium ions is clearly visible in the reconstructed maps (Supplementary Fig. 6a). To validate whether calcium ions are required for the interaction between WEEV VLPs and LA1-3-Fc, we conducted BLI assays under calcium depletion conditions using EGTA, a calcium-specific chelator. The inclusion of 10 mM EGTA in the binding buffer eliminated the formation of the WEEV VLP: LA1-3-Fc complex, confirming that calcium ions play a critical role in stabilizing this interaction (Supplementary Fig. 6b).
Interactions of WEEV and VLDLR LA1-3
In the structure of the complex, VLDLR-LA1-2 contacts E1 and E2 in one E2–E1 heterodimer (named as E2–E1) and E2’ in an adjacent heterodimer (named as E2’–E1’). The contact surface area of VLDLR-LA1-2 is calculated to be 791.9 Ų, which is comparable to the contact area of PCDH10-EC1 (1080 Ų)37. The contacting areas on VLDLR-LA1-2 accounting for the interactions with E1, E2, and E2’ are calculated as 171.8 Ų, 480.3 Ų, and 139.8 Ų, respectively.
The interacting residues of VLDLR-LA1-2 include D53, D55, E56, and D57 in LA1, which are involved in calcium ion chelation, and R88, W89, P95, as well as D92, D94, and D96 in LA2, which form coordination bonds with another calcium ion (Supplementary Fig. 7).
At the contact interface in LA1, an exposed lysine residue (K227) of E1 is positioned between the residues D53, D55, and D57. The clustering of these acidic residues around K227 creates a region of strong electrostatic complementarity (Fig. 3c). K227 establishes favorable interactions with LA1 through hydrogen bonds and salt bridges formed with these aspartic acid residues. In addition, the side chain of residue E56 in LA1 forms a hydrogen bond with the main chain of residue R228 in E2. This interaction further stabilizes the binding interface between LA1 and the viral glycoproteins (Fig. 3d and Supplementary Fig. 7).
Within the LA2 binding interface, the side chains of R88 and W89 interact with the main chain of S168 in E2, and the side chain of R88 forms a hydrogen bond with the main chain of K190 in E2, further stabilizing the interaction between LA2 and E2. Additionally, the main chain of P95 forms a hydrogen bond with E2’s K181. Furthermore, the three acidic aspartic acid residues D92, D94, and D96, which are involved in calcium ion coordination, contact the three basic residues K181, K190, and R214 in E2. These interactions form multiple hydrogen bonds and salt bridges that stabilize the LA repeat (Fig. 3e and Supplementary Fig. 7).
To evaluate how interacting residues influence virus‒receptor engagement, we mutated specific residues from VLDLR LA1 and LA2. K562 cells stably expressing these mutant VLDLR proteins were infected with VSV-WEEV (California) at an MOI of 0.2 for 24 h, and infection levels were monitored via flow cytometry. The results indicated that single-point mutations D53A, D55A, and D57A of LA1 completely abolished the ability of VLDLR-LA1 to promote VSV-WEEV (California) infection. In the case of mutation E56A, there was a reduction in infection efficiency compared to the wild-type. Similarly, mutations D92A, D94A, and D96A in of VLDLR-LA2 also prevented promotion of VSV-ΔG-WEEV (California) infection, while R88A showed a slight decrease in infection efficiency relative to the wild-type (Supplementary Fig. 5b). Overall, these experiments highlight the critical roles of residues D53, D55, and D57 in LA1, and D92, D94, and D96 in LA2 for effective recognition of WEEV.
Mapping of the LA3 and LA5 domains and WEEV binding sites
To further confirm the binding sites of LA3 and LA5 with the WEEV strain California, we generated human VLDLR LA3-Fc and LA5-Fc constructs. After incubation with WEEV VLPs, we conducted two independent data collections and reconstructions. Through block-based reconstruction and 3D classification, we obtained density maps of the complexes formed between LA3-Fc and LA5-Fc with WEEV VLPs. The maps revealed additional density at the DIII region of WEEV E1, indicating potential binding sites for LA3 and LA5 (Fig. 3f, g). However, the overall binding proportion was relatively low. We masked one position in q3 and performed 3D classification to observe the extra binding density. This may explain why no additional binding density was observed in the cryo-EM maps of the VLDLRLBD-Fc complex with WEEV VLPs. Regrettably, the densities were insufficient for models building, which might be due to the relatively low binding affinity of LA3 (10.91 μM) and LA5 (7.77 μM) with WEEV California VLPs, compared to VLDLR-LA1-3 (0.47 μM) (Fig. 4 and Supplementary Fig. 4). To further validate the binding sites of LA3 and LA5 with the WEEV strain California, we generated human VLDLR LA3-His, LA5-His, and WEEV California E1-DIII-Fc constructs. BLI measurements demonstrated interactions between VLDLR LA3-His, LA5-His, and WEEV strain California E1-DIII-Fc, with affinity constants of 829.6 nM and 808 nM, respectively (Supplementary Fig. 8). These results indicate that VLDLR LA3 and LA5 can interact with WEEV strain California’s E1-DIII to facilitate viral infection.
Binding of different VLDLR concatemers to WEEV VLPs measured by enzyme-linked immunosorbent assay (ELISA). The equilibrium dissociation constant (KD) was calculated as mean of two biologically independent experiments.
The multivalent binding of VLDLR to WEEV
To better understand how WEEV recognizes the VLDLR receptor and its multiple binding patterns and significance, we produced different VLDLR concatemers, including LA1-2, LA1-3, LA1-4, LA1-5, LA1-8, and LA2-3, as well as LA3-5. ELISA measurements of the binding of these VLDLR concatemers to WEEV VLPs showed that LA1-5 exhibited the highest affinity (0.41 μM). In contrast, the affinity of VLDLR-LA1-2 (1.63 μM) was lower than that of LA1-3 (0.47 μM) and LA1-5 (0.41 μM) (Fig. 4). These results suggest a synergistic enhancement of affinity as more VLDLR segments capable of promoting WEEV infection are included. The results also imply that LA3 and LA5 interact with distinct epitopes on WEEV, separate from those recognized by LA1 and LA2, which is consistent with our structural findings. However, the affinity of VLDLR-LBD (LA1-8) (1.21 μM) was lower than that of LA1-5 (0.41 μM) (Fig. 4). This may be due to non-interactive LA repeats (LA6, LA7, LA8) with WEEV California VLPs, which could cause steric hindrance and prevent binding to the epitopes recognized by WEEV California VLPs, thereby reducing affinity. Furthermore, the exceptional binding affinity of LA1-5 underscores its potential utility in the design of fusion proteins for protective trials aimed at blocking viral infection.
Strain-specific residues on WEEV: VLDLR interface
Since the first isolation of WEEV, the scale and frequency of its outbreaks, as well as its mammalian virulence, have declined. This is thought to be linked to changes in receptor usage during the virus’s submergence5,6,29,38,39. Highly virulent ancestral WEEV group A strains California, Fleming, McMillan, Y62-33, and CU71-CPA, can engage both human VLDLR/ApoER2 and PCDH1029. The WEEV group B1 strains BFS2005 and BFS932, group B2 strains Montana-64 and 71V1658, group B3 strains 85-452NM and PV012357A, and group C strain CBA87, which cannot utilize VLDLR and ApoER2 as receptors, were found to engage PCDH10 as the receptor29. In contrast, the entry of the WEEV group B3 strains R02V003422B and Imperial-181 cannot be supported by either PCDH10, VLDLR, or ApoER2.
To investigate why group A WEEV strains can use VLDLR/ApoER2 as receptors while other groups cannot, we performed sequence alignments of WEEV strains (Fig. 5a; Supplementary Figs. 9 and 10). And our structural analysis revealed that the lysine residue at position 181 in the E2 glycoprotein play a crucial role in the interaction with LA2. Sequence alignment showed that most group A strains have a lysine (K) at E2 position 181, while strains that cannot use VLDLR/ApoER2 as receptors typically have a glutamic acid (E) at E2 position 181 (Fig. 5a). To validate the importance of this residue for VLDLR binding, we conducted site-directed mutagenesis. We introduced E2K181E mutations into the California strain and found that these mutations abolished the binding of California VLPs to VLDLR-LBD, as measured by BLI (Fig. 5b). Conversely, we performed a reverse mutation (E2E181K) in the Imperial 181 strain and demonstrated that this mutant can bind to VLDLR-LBD (Fig. 5b). These findings highlight the critical role of the E2 glycoprotein’s 181th residue in determining WEEV’s receptor usage and virulence. In two recent studies35,36, it was similarly found that the lysine (K) at E2 position 181 is crucial for WEEV’s recognition of VLDLR. The ability of group A strains to bind VLDLR/ApoER2 appears to be dependent on this lysine residue, which facilitates essential interactions between the virus and these receptors. Our study provides molecular insights into the evolutionary changes in WEEV receptor usage and virulence, which could have implications for understanding the virus’s potential for re-emergence and for developing targeted antiviral strategies.
a Comparative analysis of strain-specific residues of the E2 protein. Sequence alignment of positions 181 and 81 of E2 proteins encoded by WEEV strains as labeled in the figure. White, light pink, light blue, yellow, and light purple backgrounds indicate Group C, A, B1, B2, and B3. Red words denote the strictly strain-specific residues, whereas black words denote the conserved residues. b BLI analysis of the binding of human VLDLR to WEEV VLPs and their mutants. The left panel shows the BLI analysis of the binding between wild-type WEEV California VLPs and their mutants E2K181E with VLDLR LBD. The middle panel shows BLI analysis of the binding of VLDLR LBD to WEEV Imperial 181 VLPs and their mutant E2E181K. The right panel shows BLI analysis of the binding of VLDLR LBD to WEEV Imperial 181 VLPs and their mutantE2 E81K. c The impacts of WEEV E2 protein polymorphisms on recognition by PCDH10 and VLDLR. K562 cells stably expressing VLDLR or PCDH10 were infected with WT or mutant VSV-ΔG-WEEV (California) (MOI = 1) or VSV-ΔG-WEEV (Imperial) (MOI = 0.1) for 24 h. Infection was monitored by flow cytometry. Data are normalized to that of empty vector-transduced cells. Data are mean ± SD from 3 biological repeats (n = 3) and p values are from unpaired two-tailed Student’s t-test. At least three independent experiments were performed with similar results and representative images are shown.
Notably, the Fleming strain, classified as Group A, has a glutamic acid (E) at position 181 of the E2 glycoprotein (Fig. 5b). According to our structural and mutational analyses, an E at position 181 disrupts the interaction between WEEV and VLDLR. Despite this, Fleming can still use VLDLR as a receptor29, suggesting it may have different VLDLR recognition epitopes. Sequence analysis revealed that Fleming has a lysine (K) at position 81 of the E2 glycoprotein, whereas other strains have glutamic acid (E) or aspartic acid (D). We hypothesize that this lysine residue plays a crucial role in VLDLR recognition. To test this hypothesis, we performed a site-directed mutation of position 81 in the E2 glycoprotein of the Imperial 181 strain (E81K). BLI assays demonstrated that this mutation can bind to VLDLR-LBD (Fig. 5b). These findings indicate that the K81 residue in Fleming facilitates its interaction with VLDLR, providing insights into the diverse receptor recognition mechanisms among WEEV strains. Recently reported structures of the WEEV Fleming strain with VLDLR-LA1-2 support our hypothesis (Supplementary Fig. 11)35.
To further evaluate the impact of mutations on viral infectivity, we conducted experiments using K562 cells stably expressing VLDLR or PCDH10. These cells were infected with WT or mutant VSV-ΔG-WEEV (California) at an MOI of 1 or VSV-ΔG-WEEV (Imperial) at an MOI of 0.1 for 24 h. Infection levels were monitored via flow cytometry. The results demonstrated that the E2K181E mutation in the California strain completely abolished its ability to bind to VLDLR. Conversely, the E2E181K and E2E81K mutations in the Imperial 181 strain enhanced its ability to enter via VLDLR (Fig. 5c). These findings highlight how a single amino acid mutation in the E2 protein can alter WEEV’s receptor binding preference. This deepens our understanding of the WEEV submergence phenomenon and the molecular mechanisms governing viral evolution and receptor recognition.
Discussion
Receptor recognition is a pivotal step in viral entry mechanisms. Most WEEV strains are capable of engaging human PCDH10 as a receptor, while highly virulent ancestral WEEV strains can interact with both human PCDH10 and VLDLR/ApoER229.In this study, our cryo-EM analysis has elucidated the structural basis of WEEV (California strain) recognition of VLDLR, uncovering the binding sites for VLDLR-LA1, LA2, LA3, and LA5 on WEEV. Furthermore, our comparative analysis of WEEV’s recognition of PCDH10 and VLDLR-LA1-2 reveals that they share a common binding epitope on WEEV glycoproteins (Supplementary Fig. 11a)37. This epitope is situated in the cleft formed by two adjacent E2–E1 heterodimers within a single trimeric spike and is highly structurally conserved among different WEEV strains, making it an ideal drug target.
VLDLR/ApoER2 serves as a receptor for multiple alphaviruses, including SFV, EEEV, and WEEV. Previous studies have detailed how VLDLR interacts with the E2–E1 glycoproteins of SFV and EEEV32,33,34. Our comparative analysis reveals that WEEV California strain and EEEV bind to VLDLR-LA1-2 differently. Specifically, WEEV interacts with LA2 via the E2 protein on the same side as E1, while EEEV binds to LA2 through the E2’ protein on the opposite side of E140 (Supplementary Fig. 11b). However, the recently reported binding mode of the WEEV Fleming strain to VLDLR-LA1-2 is different with WEEV California strain and EEEV (Supplementary Fig. 11c)35. In the WEEV Fleming strain, the K81 residue on the E2 protein forms a salt bridge with the aspartic acid residues in the calcium-binding region of LA2, enabling VLDLR recognition (Supplementary Fig. 11d). Our mutation of E81K in the Imperial 181 strain confirmed this interaction, as it enabled the strain to utilize VLDLR (Fig. 5c). These variations in binding modes highlight the diverse evolutionary strategies alphaviruses have adopted to optimize their interaction with VLDLR, enhancing our understanding of viral adaptation and receptor recognition mechanisms. Notably, the binding of WEEV to LA3 and LA5 shares similarities with SFV’s binding to LA3, as both target the DIII region of the WEEV E1 protein32 (Supplementary Fig. 11e, f). Despite the insufficient densities of WEEV in complex with LA3 and LA5 for models building (Fig. 3f, g and Supplementary Figs. 11c, d), we used BLI to measure the interactions between VLDLR LA3-His, LA5-His, and WEEV strain California E1-DIII-Fc. The affinity constants were determined to be 829.6 nM and 808 nM, respectively (Supplementary Fig. 8). These results indicate that VLDLR LA3 and LA5 can interact with the E1-DIII region of the WEEV strain California to facilitate viral infection.
Understanding these receptor recognition mechanisms and their structural bases not only deepens our knowledge of alphavirus entry but also offers valuable targets for the development of antiviral therapies and vaccines. Future studies could explore the dynamic interplay between viral evolution and receptor usage, as well as the potential for developing inhibitors that target these conserved binding epitopes.
Methods
Preparation of WEEV VLPs
The gene encoding the WEEV capsid and envelope proteins (strain California, GenBank KJ554965.1) and Imperial 181 (GenBank: GQ287641.1) was codon-optimized and synthesized (GenScript, USA). The synthesized WEEV gene was cloned into the pCAGGS vector. HEK293F cells were cultured in SMM 293 T-II medium (Sino Biological Inc., China) at 37 °C under 5% CO2. The cells (500 ml culture) were seeded at 2 × 106 cells per ml and were subsequently transfected with a mixture of 1 mg of WEEV VLP plasmid and 3 mg polyethyleneimine (PEI; Polysciences, USA). After 96 h of transfection, the supernatant was collected by centrifugation at 1000 g for 15 min. The supernatant was centrifuged at 3000 g for 15 min and sterile-filtered using a 0.45 μM filter to remove the cell debris. The VLPs were subsequently pelleted through a 30% (w/v) sucrose cushion at 140,000 g for 3 hours at 4 °C using a SW32 Ti rotor (Beckman Coulter, USA). The resultant VLPs pellets were resuspended in the buffer (20 mM HEPES, 150 mM NaCl, pH 7.5), and were further purified by ultracentrifugation on a 20–60% (w/v) sucrose density gradient using a SW41 Ti rotor (Beckman Coulter, USA) at 168,000 g for 12 h at 4 °C. The band of VLPs was carefully extracted using a syringe, buffer-exchanged into the buffer containing 20 mM HEPES, 150 mM NaCl, pH 7.5, and concentrated with an Amicon Ultra-4 300-kDa cut-off centrifugal concentrator (Millipore, USA). The particle integrity was checked using negative stain electron microscopy, and VLPs were always used within 7 days of purification.
Preparation of the recombinant proteins
The gene encoding human VLDLR (GenBank NP_003374.3) LA1-8 (residues 1–355), LA1 (residues 1–69), LA1-2 (residues 1–113), LA1-3 (residues 1–152), LA1-4 (residues 1–190), LA1-5 (residues1–236), LA1-6 (residues1–275), LA2 (residues 70–110), LA3 (residues 111–151), LA4 (residues152–190), LA5 (residues 191–236), LA6 (residues 237–275), LA7 (residues 276–315), LA8 (residues 316–355), LA2-3 (residues 70–151), and LA3-5 (residues 111–236) was codon-optimized and synthesized (SupraGene, China). The genes were cloned into the pCDNA3.4 vector with the native signal peptide sequence, followed by a human IgG1 Fc region. To increase VLDLR-Fc fusion protein purification yields, constructs were co-transfected with human LRPAP1 (RAP) chaperone gene (residues 1–353) (GenBank NP_002328) at a (4:1ratio). HEK293F cells (500 ml culture) were transfected with a mixture containing 1 mg of plasmids and 3 mg PEI at a cell density of 2.5 × 106 cells/ml. The supernatant was collected 72 h after transfection, centrifuged at 3000 g for 20 min, and sterile-filtered using a 0.45 μM filter to remove the cell debris and purified using protein A Sepharose 4B (GenScript, USA). The column was washed with 25 column volumes of 1× TBS (20 mM Tris pH 8.0, 150 mM NaCl) and 2 mM CaCl2, 50 column volumes of high-salt buffer (20 mM Tris pH 8.0, 500 mM NaCl, 10 mM EDTA) to strip LRPAP1, followed by 25 column volumes of 1× TBS and 10 mM CaCl2. After elution, the target protein was concentrated and purified by Superdex-200 size exclusion chromatography column (GE Healthcare, USA) in the buffer containing 20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH 7.5. Protein purity was assessed by SDS-PAGE analysis.
The sequence encoding the WEEV California E1-DIII (E1 residues 290-393, GenBank accession number KJ554965.1) was codon-optimized and synthesized (SupraGene, China). The genes were cloned into the pCDNA3.4 vector with the human IL-2 signal peptide sequence, followed by a human IgG1 Fc region. Soluble protein was purified from the culture supernatant using protein A Sepharose 4B (GenScript, USA).
The gene encoding human VLDLR (GenBank NP_003374.3) LA3 (residues 111–151) and LA5 (residues 191–236) was codon-optimized and synthesized (SupraGene, China). The genes were cloned into the pCAGGS vector with the native signal peptide sequence, followed by a 6× His tag. Constructs were co-transfected with human LRPAP1 (RAP) chaperone gene (residues 1–353) (GenBank NP_002328) at a 4:1 ratio and proteins were purified from the culture supernatant using Ni Sepharose resin (17371202, Cytiva).
Bio-layer interferometry (BLI) analysis
BLI analyses were performed by the ForteBio Octet Red system (ForteBio Inc., USA). The purified Fc-tagged VLDLR-LAs or WEEV California E1-DIII proteins (100 nM) were immobilized onto the AHC biosensors (Sartorius 18-5060) in 200 μl loading buffer (20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, 0.1% BSA, pH 7.5) for 100 s. Coated sensor tips were dipped into the kinetic buffer for a baseline measurement of 100 s, then dipped into wells containing the various concentrations of WEEV VLPs or VLDLR-LAs-His for 480 s. Subsequently, a buffer wash of 300 s was applied to allow the dissociation of molecules from the sensor. Data analysis was performed with the software Octet (v.6.4, ForteBio) using a standard 1:1 binding model. Two independent experiments were conducted for each sample.
Titration ELISA
The binding affinity of VLDLR-LAs was determined by titration ELISA. WEEV California VLPs were diluted to 2 μg/ml using an ELISA Coating Buffer (Solarbio, C1050), and the VLPs were immobilized on an ELISA plate (100 μl per well) and placed overnight at 4 °C. PBS containing 5% FBS was used to block the coated plate at 37 °C for 1 h. Serial diluents of different fragments of recombinant VLDLR were prepared with PBS containing 5% FBS. The gradient diluent was added to the plate and incubated at 37 °C for 1 h. Then it was washed again with PBS for 5 times, and horseradish-peroxidase-conjugated anti-human IgG (Promega, W4031) diluted with 5% FBS in PBS at a ratio of 1:4000 (100 μl per well), and incubated at 37 °C for 1 h. After washing with PBS for 5 times, 100 μl Supersensitive TMB Chromogen Solution (Beyotime, P0208) was added for incubation at room temperature, and then 100 μl ELISA Stop Solution for TMB Substrate (Beyotime, P0215) was added to stop the reaction and the absorbance was read at 450 nm optical density.
Preparation of WEEV pseudotype virus
BHK-21 cells were transfected with plasmids encoding E3-E2-6K-E1 of WEEV for 24 h followed by infection with VSV- ΔG-eGFP viruses. One hour post infection, cells were washed with PBS for 5 times to remove uninfected viral particles and then cultured with DMEM containing 2% FBS. Supernatants containing pseudotyped VSV-ΔG-eGFP viruses were harvested 24–36 h later. Virus titers were quantified on Vero cells based on the number of eGFP-positive cells30.
Flow cytometry
For detection of VSV-ΔG-WEEV infection, K562 cells stably expressing human PCDH10, VLDLR or various VLDLR truncations were infected with VSV-ΔG-WEEV for 24 h. Cells were fixed with 4% paraformaldehyde for 15 min, washed with PBS and processed on BD LSRFortessa, and analysed using FlowJo software.
Western blot
Equal amounts of cell lysates were mixed with 2× SDS loading buffer, heated at 95 °C for 30 min, and loaded onto 10% SDS-PAGE gels for running at 135 V. Proteins were transferred to nitrocellulose membrane (Millipore #IPVH00010) via wet electroblotting (300 mA, 1 h, 4 °C). Membranes were blocked with 5% non-fat dry milk in TBST for 0.5 h at room temperature (RT), followed by incubation with primary antibodies: Mouse anti-Flag mAb (Sigma-Aldrich #F3165, 1:2000 dilution from 0.5 μg/μL stock) and rabbit anti-β-actin mAb (ABclonal #AC026, 1:5000 dilution from 1 μg/μL stock) at 4 °C for 1–2 h. After washing three times with TBST (10 min each), membranes were incubated with HRP-conjugated secondary antibodies (Pierce #31430/#31460, 1:2000) for 1 h at RT. Final washes were performed 4 times with TBST (10 min each). Protein bands were detected using ECL substrate (Advansta # K-12043-D20).
Cryo-EM sample preparation and data collection
The WEEV California VLPs were incubated with VLDLR LA or LAs for 3 min on ice. Subsequently, the samples (4 µL) in the buffer (20 mM HEPES, 150 mM NaCl, pH 7.5) were applied onto a H2/O2 glow-discharged, 300-mesh Quantifoil R1.2/1.3 holey NiTi grids (Quantifoil, Micro Tools GmbH, Germany). The grid was then blotted for 5.0 s with a blot force of 0 at 8 °C and 100% humidity and plunge-frozen in liquid ethane using a Vitrobot (Thermo Fisher Scientific, USA). Cryo-EM data for WEEV VLPs apo and in complex with LA1-3 and LA3 were collected using a 300 kV Titan Krios G4 electron microscope (Thermo Fisher Scientific, USA) equipped with a Gatan K3 camera. The images were recorded at ×81,000 magnification with a super-resolution pixel size of 1.055 Å/pixel. The exposure time was set to 2.7 s, delivering a total accumulated dose of 50 electrons per Ų. Movies were automatically captured using the EPU software, with a defocus range from −2.0 to −1.0 μm. For WEEV VLPs in complex with LBD and LA5, data were collected on the same 300 kV Titan Krios G4 electron microscope and equipped with a Falcon 4i camera and a Quantum LS energy filter (10 eV slit width). The images were recorded at ×105,000 magnification with a super-resolution pixel size of 1.2 Å/pixel. The exposure time was set to 8.72 s with a total accumulated dose of 50 electrons per Å2.
Cryo-EM data processing
All dose-fractioned images were motion-corrected and dose-weighted by patch motion correction, and the contrast transfer function (CTF) of each micrograph was estimated by patch CTF estimation in cryoSPARC41. The micrographs were manually selected for further processing. Particles were picked using a template-based method. Two rounds of 2D classification were performed to refine the particle selection. For heterogeneous refinement, the volume of WEEV-71V1658 VLP (EMD-62791) with low-pass filtering and two junk volumes generated from poor classes were used as references. The good particles were subjected to non-uniform refinement with I1 symmetry to obtain the density of the whole VLPs. To further improve the resolution, block-based reconstruction42 was performed around the 5-fold axes region of the particles. In Relion3.1.243, the particles underwent 3D classification to select high-quality classes, which were then merged particles and subjected to local refinement with C5 symmetry. The particles used for local refinement were symmetry-expanded with C5 symmetry, and all trimeric spikes from the 5-fold axes region were extracted. Local refinement with C1 symmetry was performed to optimize the density for the trimeric spikes. To further enhance the receptor resolution, focused classification without alignment was conducted in Relion3.1.2, and the particles with the clearest receptor density were selected and merged for final local refinement with C1 symmetry. Automatic local sharpening in EMReady44 was performed to improve the densities. The local resolutions were estimated in cryoSPARC. The workflow is shown in Supplementary Fig. 1.
Model building and refinement
To build the structures of WEEV VLP and its complex with VLDLR-LA1-2, the coordinates of E1 and E2 proteins of WEEV strain CBA87 VLP (PDB: 8DEE28), as well as LA1-2 domains of human VLDLR (PDB: 8UFC33), were individually placed and fitted by rigid-body refinement into the cryo-EM maps in UCSF Chimera45. The models were rebuilt using Coot46 with the guidance of the cryo-EM maps and were refined with real-space refinement using Phenix47. The data validation statistics are shown in Supplementary Table 1. Structural figures were generated with Pymol48, UCSF Chimera, and ChimeraX49.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM density maps and the structures were deposited into the Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) with the accession numbers 9UIH and EMD-64194 for WEEV California strain VLPs, 9UIO, EMD-64199 for the complex with VLDLR-LA1-3, 9WSL, EMD-66198 for the complex with VLDLR-LBD, EMD-64263 for the complex with VLDLR-LA3 and EMD-64264 for the complex with VLDLR-LA5. Accession codes of previously published structures used in this study are EMD-62791, 8UFC, and 8DEE. Source data are provided as a Source data file. Source data are provided with this paper.
References
Chen, W. et al. Arthritogenic alphaviruses: new insights into arthritis and bone pathology. Trends Microbiol. 23, 35–43 (2015).
Powers, A. M. et al. Evolutionary relationships and systematics of the alphaviruses. J. Virol. 75, 10118–10131 (2001).
Ronca, S. E., Dineley, K. T. & Paessler, S. Neurological sequelae resulting from encephalitic alphavirus infection. Front. Microbiol. 7, 959 (2016).
Mejía, C. R. & López-Vélez, R. Tropical arthritogenic alphaviruses. Reumatol. Clin. 14, 97–105 (2018).
Bergren, N. A. et al. Submergence” of Western equine encephalitis virus: Evidence of positive selection argues against genetic drift and fitness reductions. PLoS Pathog. 16, e1008102 (2020).
Reisen, W. K. & Wheeler, S. S. Surveys for antibodies against mosquitoborne encephalitis viruses in california birds, 1996-2013. Vector Borne Zoonotic Dis. 16, 264–282 (2016).
Reed, D. S. et al. Aerosol exposure to western equine encephalitis virus causes fever and encephalitis in cynomolgus macaques. J. Infect. Dis. 192, 1173–1182 (2005).
Carey, B. D., Bakovic, A., Callahan, V., Narayanan, A. & Kehn-Hall, K. New World alphavirus protein interactomes from a therapeutic perspective. Antivir. Res. 163, 125–139 (2019).
Zanotto, P. M. A. & Leite, L. C. C. The challenges imposed by dengue, zika, and chikungunya to Brazil. Front. Immunol. 9, 1964 (2018).
Strauss, J. H. & Strauss, E. G. The alphaviruses: gene expression, replication, and evolution. Microbiol. Rev. 58, 491–562 (1994).
Voss, J. E. et al. Glycoprotein organization of chikungunya virus particles revealed by X-ray crystallography. Nature 468, 709–712 (2010).
Basore, K. et al. Cryo-EM structure of chikungunya virus in complex with the Mxra8 receptor. Cell 177, 1725–1737.e1716 (2019).
Lescar, J. et al. The Fusion glycoprotein shell of Semliki Forest virus: an icosahedral assembly primed for fusogenic activation at endosomal pH. Cell 105, 137–148 (2001).
Li, L., Jose, J., Xiang, Y., Kuhn, R. J. & Rossmann, M. G. Structural changes of envelope proteins during alphavirus fusion. Nature 468, 705–708 (2010).
Roussel, A. et al. Structure and interactions at the viral surface of the envelope protein E1 of Semliki Forest virus. Structure 14, 75–86 (2006).
Kostyuchenko, V. A. et al. The structure of barmah forest virus as revealed by cryo-electron microscopy at a 6-angstrom resolution has detailed transmembrane protein architecture and interactions. J. Virol. 85, 9327–9333 (2011).
Pletnev, S. V. et al. Locations of carbohydrate sites on alphavirus glycoproteins show that E1 forms an icosahedral scaffold. Cell 105, 127–136 (2001).
Zhang, R. et al. 4.4 Å cryo-EM structure of an enveloped alphavirus Venezuelan equine encephalitis virus. EMBO J. 30, 3854–3863 (2011).
Smith, T. J. et al. Putative receptor binding sites on alphaviruses as visualized by cryoelectron microscopy. Proc. Natl. Acad. Sci. USA 92, 10648–10652 (1995).
Zhang, W., Heil, M., Kuhn, R. J. & Baker, T. S. Heparin binding sites on Ross River virus revealed by electron cryo-microscopy. Virology 332, 511–518 (2005).
DeTulleo, L. & Kirchhausen, T. The clathrin endocytic pathway in viral infection. EMBO J. 17, 4585–4593 (1998).
Lee, R. C. et al. Mosquito cellular factors and functions in mediating the infectious entry of chikungunya virus. PLoS Negl. Trop. Dis. 7, e2050 (2013).
Zimmerman, O., Holmes, A. C., Kafai, N. M., Adams, L. J. & Diamond, M. S. Entry receptors - the gateway to alphavirus infection. J. Clin. Investig. 133, e16530710 (2023).
Zhang, R. et al. Mxra8 is a receptor for multiple arthritogenic alphaviruses. Nature 557, 570–574 (2018).
Clark, L. E. et al. VLDLR and ApoER2 are receptors for multiple alphaviruses. Nature 602, 475–480 (2022).
Ma, H. et al. LDLRAD3 is a receptor for Venezuelan equine encephalitis virus. Nature 588, 308–314 (2020).
Zhai, X. F. et al. LDLR is used as a cell entry receptor by multiple alphaviruses. Nat. Commun. 15, 622 (2024).
Zimmerman, O. et al. Vertebrate-class-specific binding modes of the alphavirus receptor MXRA8. Cell 186, 4818–4833.e4825 (2023).
Li, W. et al. Shifts in receptors during submergence of an encephalitic arbovirus. Nature, https://doi.org/10.1038/s41586-024-07740-2 (2024).
Yang, Y. et al. PCDH10 is a neuronal receptor for western equine encephalitis virus. Cell Res. 34, 802–805 (2024).
Palakurty, S. et al. The VLDLR entry receptor is required for the pathogenesis of multiple encephalitic alphaviruses. Cell Rep. 43, 114809 (2024).
Cao, D., Ma, B., Cao, Z., Zhang, X. & Xiang, Y. Structure of Semliki Forest virus in complex with its receptor VLDLR. Cell 186, 2208–2218.e2215 (2023).
Adams, L. J. et al. Structural and functional basis of VLDLR usage by eastern equine encephalitis virus. Cell 187, 360–374.e319 (2024).
Yang, P. et al. Structural basis for VLDLR recognition by eastern equine encephalitis virus. Nat. Commun. 15, 6548 (2024).
Raju, S. et al. Structural basis for plasticity in receptor engagement by an encephalitic alphavirus. Cell 188, 2943–2956.e24 (2025).
Fan, X. et al. Molecular basis for shifted receptor recognition by an encephalitic arbovirus. Cell, https://doi.org/10.1016/j.cell.2025.03.029 (2025).
Liang, S. et al. Structural basis for engagement of western equine encephalitis virus with the PCDH10 receptor. Nat. Commun. 16, 6290 (2025).
Nagata, L. P. et al. Infectivity variation and genetic diversity among strains of Western equine encephalitis virus. J. Gen. Virol. 87, 2353–2361 (2006).
Logue, C. H. et al. Virulence variation among isolates of western equine encephalitis virus in an outbred mouse model. J. Gen. Virol. 90, 1848–1858 (2009).
Cao, D. et al. The receptor VLDLR binds eastern equine encephalitis virus through multiple distinct modes. Nat. Commun. 15, 6866 (2024).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zhu, D. et al. Pushing the resolution limit by correcting the Ewald sphere effect in single-particle Cryo-EM reconstructions. Nat. Commun. 9, 1552 (2018).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
He, J., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Schrodinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32188101) to Z.L., the National Key R&D Program of China (2023YFC2306100 and 2024YFA1306500) to Y.W. and Y.Y. respectively, National Natural Science Foundation of China (32273098) and Natural Science Foundation of Jilin Province (20230101368JC) to Y.Z. We thank the Electron Microscopy Facilities of Guangzhou Laboratory for the cryo-EM data collection.
Author information
Authors and Affiliations
Contributions
Y.W., S.L. and Z.L. conceived the project and designed the experiments. S.L., Z.X., J.H., Z.W., D.L., J.Y., J.Z. and J.B. prepared the WEEV VLPs and receptor proteins. X.L., S.L., Y.Y., J.Y., Z.X., Y.Z., C.H. and L.Z. performed the BLI analysis, ELISA analysis, and cell-based assays. S.L. and Z.X. prepared the cryo-EM samples and collected the cryo-EM data. S.L. processed cryo-EM data and built the atomic model. Y.W., S.L. and Z.L. analyzed the data. S.L., Z.L. and Y.W. wrote the manuscript. All authors discussed the experiments, read, and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Laurie A. Silva, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, S., Xu, Z., Liu, X. et al. Structural insights into VLDLR recognition by western equine encephalitis virus. Nat Commun 17, 435 (2026). https://doi.org/10.1038/s41467-025-66330-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66330-6