Abstract
Conjugative plasmids can drive the global spread of antimicrobial resistance (AMR) in Enterobacterales. Hypervirulent Klebsiella pneumoniae (hvKp) increasingly acquire AMR plasmids, raising concern about convergent hypervirulent drug-resistant clones. Yet little is known about plasmid transmission dynamics in hvKp. Using an antibiotic-perturbed murine gut model with hvKp and human commensal E. coli, we discover that broad-host range IncP plasmids belonging to different phylogenetic branches (clade I (PTU-P1) and clade II (PTU-P2)) transfer differentially in the gut, mirroring the higher prevalence of PTU-P2 plasmids in human-associated samples. Statistical modelling and experimental results show that secondary transfer by transconjugants sustains gut transmission without continuous donor input. Furthermore, the hvKp capsule exerts a modest effect on transfer in vivo compared to in vitro aerobic conditions. Under anaerobic conditions, hvKp capsule mucoviscosity is markedly reduced, and PTU-P2 plasmids conjugate more efficiently than PTU-P1 counterparts. Our findings reveal that the hypermucoviscous capsule may not substantially impede gene exchange in the gut where microenvironments shape plasmid transfer dynamics, highlighting the pitfalls of extrapolating in vitro data to relevant ecological niches. Our work also emphasizes the high-risk nature of gut-adapted PTU-P2 plasmids and the ease with which hvKp can acquire them, underscoring the need for continued surveillance.
Similar content being viewed by others
Introduction
Plasmids are pivotal in bacterial evolution, imparting advantageous traits such as antimicrobial resistance genes (ARGs), heavy metal resistance, and metabolic capabilities1. Early studies on plasmid biology focused on well-characterized examples like the F-plasmid2 and the IncP plasmid RP43, revealing key mechanisms of transfer, replication, and maintenance. Recently, plasmids have also been explored as vehicles for targeted payload delivery4. At the same time, a growing body of research on clinical plasmids that disseminate ARGs underscores their public health impact. However, the mechanisms driving the success of specific plasmid–host pairings remain poorly understood.
Plasmid-mediated antimicrobial resistance (AMR) is a major concern across Enterobacterales, with Klebsiella pneumoniae and Escherichia coli frequently implicated. K. pneumoniae, often dubbed the “ARG trafficker,” plays a central role in resistance spread5. In carbapenem-resistant K. pneumoniae, specific plasmids are linked to dominant clones, such as pOXA-48-like plasmids with sequence type (ST) 116 and pKpQIL plasmids with the globally prevalent ST258 clone7. The co-occurrence of multiple blaKPC variants on pKpQIL plasmids and the ability of these plasmids to recombine allows K. pneumoniae ST307 to develop simultaneous resistance to both carbapenems and ceftazidime-avibactam while generating heterogeneous resistance profiles that facilitate survival under diverse antibiotic pressures8. In E. coli ST131, IncF plasmids encoding CTX-M extended-spectrum β-lactamases have similarly facilitated the global dissemination of AMR9,10. Multiple recent reports also document the increasing acquisition of multidrug-resistance (MDR) plasmids in hvKp that is of grave concern.
Genomic and experimental data indicate that horizontal plasmid transfer pathways are often specific to particular bacterium-plasmid combinations and are influenced by strain-specific factors11,12. Large-scale studies of bloodstream infections reveal that while plasmid groups are phylogenetically structured13, there is significant intra- and inter-species transmission. Notably, ARG-bearing plasmids often cluster with plasmids lacking ARGs. This suggests that “high-risk” plasmid backbones, regardless of their ARG content, play a central role in horizontal transfer of resistance. Machine learning and mathematical modeling approaches have been utilized to predict plasmid backbones without ARGs that might become high-risk AMR vectors13. While bioinformatics has greatly advanced our understanding of plasmid-host interactions, in vivo studies are indispensable to elucidate the complexities of these relationships. For instance, an investigation of gut isolates from hospital patients showed a trade-off between bacterial fitness and resistance in pOXA-48-mediated carbapenem resistance14. Benz et al. reported that the in vitro conjugation properties of clinically relevant ESBL plasmids, varied by donor-recipient pairings and plasmid types, were generally predictive of in vivo transfer efficiency15. Conversely, Neil et al. found that most plasmids exhibited poor conjugation in vivo despite high in vitro transfer frequencies except for the IncI2 plasmid TP114, which successfully conjugated between E. coli Nissle strains16. Furthermore, very little is known about the transfer dynamics of IncP plasmids except for the canonical IncP1 plasmid RP4, that transfers very poorly in the gut17. Therefore, understanding plasmid transmission dynamics in relevant reservoirs is vital to distill the governing principles of plasmid-host relationships.
We previously documented the prevalence of a non-canonical IncP plasmid pKPC2 in carbapenem-resistant Enterobacterales within hospital settings in Singapore18,19, including multiple hypervirulent K. pneumoniae (hvKp) strains20. pKPC2 demonstrates superior in vitro conjugation dynamics compared to pNDM1, the second most prevalent clinical carbapenemase plasmid in the same collection18. We postulated that pKPC2’s efficient in vitro transfer might explain its clinical dominance, raising concerns that plasmids with similar backbones could pose a significant threat. However, in vivo studies are essential to assess the relevance of in vitro findings and identify emergent properties. Furthermore, the in vivo plasmid transmission properties and dynamics in hvKp are unknown. Here, we examine the transfer dynamics of several clinical carbapenemase plasmids within the intestinal tract, a key clinically relevant niche for plasmid exchange. Focusing on plasmid transfer between a human probiotic E. coli Nissle 1917 (EcN) and hvKp strain SGH10 capable of stable gut colonization to simulate human gut isolates, we evaluated the transmission of pKPC2 and other plasmids in the murine gut. Our findings reveal rapid and stable plasmid transmission of clade II IncP plasmids within hvKp that is not impeded by the hypermucoid capsule nor the presence or absence of donor strains, where anaerobicity is a driver for these behaviors. These behaviors highlight how context and microenvironments greatly impact plasmid conjugation dynamics that likely reflects differential plasmid evolutionary trajectory and adaptation.
Results
Differential dynamics of plasmid conjugation in vitro and in the murine gut between E. coli and hvKp
Plasmid conjugation studies, particularly in murine gut models, have predominantly utilized E. coli or Salmonella as model organisms16,21, with limited exploration into K. pneumoniae. Here, we developed an interspecies transmission model using EcN (a human-derived commensal strain22) and hvKp SGH10 (a hypervirulent gut pathogen) as donor and recipient strains to study plasmid transfer. This approach is particularly relevant given recent clinical reports of hvKp acquiring drug-resistance plasmids and evolving into highly drug-resistant, difficult-to-treat strains.
We first compared the in vitro transfer frequency and stability of our clinical carbapenemase plasmid pKPC2 against two other clinically important plasmids belonging to different incompatibility groups pNDM1 (IncN) and pKpQIL (IncFII), which are major drivers of carbapenem resistance in Enterobacterales23,24. To benchmark their behavior, we also included three prototypical laboratory plasmids RP4-11 (IncP), pOX38 (IncFI), and TP114 (IncI2) commonly used in experimental models. EcN effectively colonizes both murine and human guts and serves as a representative of endogenous gut E. coli strains that could participate in plasmid transfer. SGH10, from a major hvKp lineage (K1 serotype, clonal group 23)5, can colonize an antibiotic-perturbed gut at high levels25 and is a clinically relevant model for gut pathogens. Since capsule production has been reported to hinder plasmid transfer in vitro18,26, we also included a capsule-deficient mutant of SGH10 (SGH10ΔwcaJ) as a recipient in our assays.
In vitro filter-mating assays revealed a range of transfer efficiencies among the plasmids (Fig. 1a). With EcN as donor and wildtype SGH10 as recipient, plasmids pKPC2, RP4-11, and pOX38 achieved relatively high conjugation frequencies, whereas pNDM1, pKpQIL, and TP114 transferred at lower frequencies. Using the non-capsulated SGH10ΔwcaJ as recipient significantly increased the transfer frequencies of pKPC2, RP4-11, pNDM1, and TP114 (Fig. 1a). In contrast, the IncF plasmids (pKpQIL and pOX38) showed no such increase, consistent with prior observations26 that capsule primarily impedes plasmids requiring short pili for conjugation, but not IncF plasmids that use long conjugative pili27.
a Box plots showing the log10-transformed conjugation efficiency of six plasmids (pKPC2, pKpQIL, pNDM1, RP4-11, pOX38, and TP114) measured after 4 h of in vitro filter mating, with equal donor and recipient proportions between E. coli Nissle (EcN) to K. pneumoniae SGH10 (red circles), EcN to non-capsulated mutant SGH10ΔwcaJ (green triangles), and SGH10 to EcN (blue squares). The center line denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents one biological replicate (n = 9 from three independent experiments). Statistical differences were analyzed using one-way ANOVA (Dunnett’s test), comparing EcN–SGH10ΔwcaJ or SGH10–EcN to the reference EcN–SGH10, and are indicated by asterisks on the graph. The exact p values are (EcN–SGH10ΔwcaJ vs. EcN-SGH10, SGH10–EcN vs. EcN-SGH10): pKPC2 (<0.0001, <0.0001); RP4 (<0.0001, <0.0001); pNDM1 (<0.0001; 0.0013); TP114 (<0.0001, <0.0001); pOX38 (0.4328, 0.6078) and pKpQIL (0.7809, <0.0001). b Plasmid retention rates in EcN, SGH10, and SGH10ΔwcaJ over 90 generations, with each dot representing the mean from three independent experiments with 9 biological replicates and error bars indicating standard deviation. c Gut colonization levels of donor (red), recipient (green) and transconjugant (blue) over 3 days. Each dot represents the data from one of the five mice (n = 5 mice per group) and the dotted line indicates the limit of detection. d Boxplot of the log10-transformed transconjugant-to-recipient ratios (T/R) of the same six plasmids from EcN to SGH10 or SGH10ΔwcaJ, and from SGH10 to EcN, with transconjugant frequencies detected in mouse stool at days 1, 2, and 3 post-inoculation, labeled as D1, D2 and D3 on the x-axis. The center line denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents the data from a mouse with a total of five mice per group (dots at the bottom indicate undetectable levels). e Confidence interval plot showing posterior mean estimates as the measure of center, illustrating the statistical model of changes in transconjugant populations over time as a function of the donor, plasmid-free recipient, and transconjugant population for various plasmids transferred between EcN and SGH10. Significant changes are indicated with credible intervals that do not cross zero, annotated as (l-95% CI, u-95% CI). The model for the pKpQIL plasmid transfer from SGH10 to EcN is omitted and marked with hashes due to a lack of transfer events. The data analyzed in the model was from five mice per group.
Switching the mating direction, with SGH10 carrying the various plasmids as donor to EcN, generally yielded slightly higher transfer frequencies for most plasmids relative to the original pairing where EcN as the donor (Fig. 1a). A notable exception was pKpQIL, which transferred at a much lower frequency from SGH10 to EcN, whereas TP114 transferred at a much higher frequency in this orientation. We next examined plasmid stability without selection over ~90 generations of growth in LB broth (serial passaging) in EcN, SGH10, and SGH10ΔwcaJ (Fig. 1b). All plasmids except pOX38 remained stably maintained in all hosts. pOX38 was gradually lost in both SGH10 and SGH10ΔwcaJ, and it imposed the greatest growth fitness cost on all host strains (Supplementary Fig. 1). It is worth noting that both SGH10 and EcN harbor endogenous plasmids. SGH10 carries a large 231 kb non–self-transmissible virulence plasmid with IncH1B and IncFIB(K) replicons28, which underpins its hypervirulent phenotype. EcN carries two small cryptic plasmids, pMUT1 (3.2 kb) and pMUT2 (5.5 kb), of unknown function, bearing ColE1- and ColE2-like replicons29. These endogenous plasmids were not cured from the strains for the purpose of our experiments, as they are intrinsic to the identity of each strain, particularly the large virulence plasmid of SGH10. Importantly, the replicons encoded by these endogenous plasmids are distinct from those tested in this study, with the exception of pOX38, whose IncFI replicon proved unstable in SGH10 (Fig. 1b), likely due to incompatibility with the resident virulence plasmid.
Having established the in vitro behavior, we next compared plasmid transfer in vivo using a mouse gut colonization model. We inoculated mice with SGH10 (or SGH10ΔwcaJ) and EcN via oral gavage, staggered by 6 h to confine conjugation to the gut. Stool samples on days 1, 2, and 3 post-inoculation were plated to enumerate donors, recipients, and transconjugants. Both donor and recipient populations reached ~10¹⁰ colony-forming units (CFU) per gram of stool by day 1 and remained at this high level through day 3 (Fig. 1c), indicating successful co-colonization. Importantly, SGH10 and SGH10ΔwcaJ colonized at similar levels, suggesting the capsule does not impact gut colonization efficiency30. Notably, when SGH10 was the donor, the EcN population on day 1 was about ten-fold lower than SGH10, but it caught up by day 2. Transconjugants mostly emerged by day 1 and remained at similar levels thereafter.
In the mouse gut, the clinical plasmids pKPC2, pKpQIL, and pNDM1 exhibited higher mean transconjugant-to-recipient ratios (T/R) than the laboratory plasmids (RP4-11, pOX38, TP114), with most comparisons reaching statistical significance on at least one of the three measured days (Fig. 1d and Supplementary Data 1). The non-capsulated SGH10ΔwcaJ accepted pKPC2 ~ 10-fold more efficiently than wildtype SGH10, although this difference was only statistically significant on day 3 (Supplementary Data 2). Thus, the hvKp capsule may have influenced plasmid transfer in the gut, although the effect appears significantly less pronounced than that observed under in vitro conditions. When SGH10 was the donor and EcN the recipient, pKPC2 showed the highest T/R over the 3-day period, whereas the T/R of most other plasmids were significantly much lower (Fig. 1d and Supplementary Data 1). pKpQIL transfer from SGH10 to EcN was virtually undetectable, likely because EcN lacks the outer membrane receptor (OmpK36) required for mating pair stabilization by the pKpQIL-encoded type IV secretion system31. In contrast, TP114 transferred more readily to EcN, possibly due to a favorable interaction between its type IV pilus adhesin and a complementary EcN surface receptor32. These in vivo findings confirm that plasmid transfer can depend strongly on recipient-specific factors, as also observed in vitro.
To dissect the contributions of each bacterial subpopulation to overall conjugation outcomes, we performed a Bayesian regression analysis to model changes in transconjugant counts over time as a function of donor, plasmid-free recipient, and transconjugant densities (Fig. 1e). In the EcN→SGH10 mating, the size of the initial transconjugant population emerged as the strongest predictor of the next-day increase in transconjugants for pKPC2, pKpQIL, and RP4-11. In the SGH10→EcN mating, transconjugant population size was the primary predictor for pOX38, while donor population size had a marginal but statistically significant effect for pKPC2. Other factors had minimal or no significant influence. We note that the small sample size limits the precision of some estimates. Nonetheless, whenever the transconjugant population had a significant effect, its effect size surpassed that of donor or recipient populations, especially for pKPC2 and RP4. These results imply that once a transconjugant subpopulation is established, further spread via transconjugants (either secondary transfer or their clonal expansion) may be more critical for plasmid dissemination in the gut than continuous input from the original donor.
Secondary transfer among recipients drives pKPC2 spread in the gut
Conjugative transfer in a mixed population can occur via primary transfer (directly from an original donor) and secondary transfer (from an initial transconjugant to other plasmid-free recipients). A study using Salmonella in a murine gut model demonstrated that rare donors can achieve dramatic transfer efficiency when recipient densities are high, with up to 99% transconjugants observed within 2–3 days where donor re-seeding is the rate-limiting step33. It is possible that this involves initial transfer from rare donors to available recipients, followed by rapid spread through secondary transfer among the recipient population to achieve such high transconjugant frequencies. This phenomenon, where secondary transfer may play an important role in plasmid spread, could occur in our model with comparable size of the donor and recipient populations. This was predicted by our mathematical modeling (Fig. 1e). To directly test secondary transfer’s role, we focused on pKPC2, the plasmid with the highest gut transfer frequency and the largest predicted secondary contribution. We constructed a transfer-deficient variant of pKPC2 by deleting the key mating gene traF (plasmid pKPC2ΔtraF) and introduced it into EcN. As expected, pKPC2ΔtraF exhibited no detectable conjugation in vitro (Supplementary Fig. 2a). Providing traF in trans on a non-mobilizable plasmid (pACYC_traF) fully restored transfer of pKPC2ΔtraF from EcN into SGH10 or SGH10ΔwcaJ to wildtype pKPC2 levels, confirming that the traF deletion specifically abolishes mating ability. When we used SGH10 or SGH10ΔwcaJ as donors carrying pKPC2ΔtraF, no transfer occurred unless traF was complemented in trans, which again rescued conjugation. Notably, complementation of pKPC2ΔtraF in SGH10ΔwcaJ restored transfer to EcN to approximately an order of magnitude lower than wildtype pKPC2 levels, highlighting that secondary transfer of EcN-to-EcN is prominent for this pairing. In contrast, transfer from SGH10 to EcN remained low despite traF complementation in SGH10 (reflecting the capsule’s effect on primary transfer). The traF deletion did not affect plasmid stability (Supplementary Fig. 2b) or bacterial growth (Supplementary Fig. 2c) in either EcN or SGH10.
We next compared plasmid transfer in vivo with and without secondary transfer. Mice were colonized with donors carrying either wildtype pKPC2 (allowing both primary and secondary transfer) or pKPC2ΔtraF + pACYC_traF (allowing only primary transfer). We tested three mating pair scenarios: EcN to SGH10, EcN to SGH10ΔwcaJ, and SGH10 to EcN, measuring donor, recipient, and transconjugant CFUs on days 1, 2, and 3 (Fig. 2a). The pACYC_traF complement was progressively lost in donors in vivo, but a substantial fraction of donors retained it throughout the 3-day experiment especially on day 1 (Supplementary Fig. 3). Across all pairings, the T/R with pKPC2ΔtraF (primary transfer only) was dramatically lower than with intact pKPC2 every day (Fig. 2b). We further validated these results using strains with chromosomally integrated traF to overcome progressive pACYC_traF loss over the days. We used EcN::traF and SGH10::traF (EcN and SGH10 with chromosomally integrated traF respectively) to restore conjugative ability to pKPC2ΔtraF without relying on the unstable pACYC_traF complementation plasmid. We first confirmed that under in vitro conditions (Supplementary Fig. 4), pKPC2ΔtraF in strains with integrated traF conjugated at frequencies similar to those complemented with pACYC_traF plasmids. We then compared in vivo transfer rates across five donor-recipient pairs: (1) EcN pKPC2 to SGH10 [wild-type control], (2) EcN pACYC_traF pKPC2ΔtraF to SGH10 [unstable donor, secondary transfer deficient], (3) EcN pACYC_traF pKPC2ΔtraF to SGH10::traF [unstable donor, secondary transfer proficient], (4) EcN::traF pKPC2ΔtraF to SGH10 [stable donor, secondary transfer deficient], and (5) EcN::traF pKPC2ΔtraF to SGH10::traF [stable donor, secondary transfer proficient] (Fig. 2c). This experimental design allowed us to dissect the contributions of donor stability versus recipient secondary transfer capability. Pair (1) served as the baseline comparison for pKPC2 transfer. Pairs (2) and (3) modeled progressive donor loss caused by pACYC_traF instability, enabling assessment of its effect on T/R in the absence (pair 2) or presence (pair 3) of a recipient capable of secondary transfer. Pairs (4) and (5) represented scenarios with stable donors that maintain traF expression over time.
a Donor (yellow), recipient (red), and transconjugant (green) counts from stools days 1, 2 and 3 for pKPC2ΔtraF transfer from EcN-SGH10, EcN-SGH10ΔwcaJ, and SGH10-EcN. Each group included five mice and dotted line indicates detection limit. b Log10-transformed transconjugant-to-recipient ratios (T/R) showing primary transfer (donor with pACYC_traF pKPC2ΔtraF) versus primary + secondary transfer (donor with intact plasmid) across days 1, 2 and 3. Each group included five biological replicates. Each point represents an individual biological replicate, and the center denotes the mean of log10-transformed values with error bars showing ± SD on the log scale. The symbols at the bottom of the graph represent undetectable levels. Statistical differences between primary and primary + secondary transfer frequencies were analyzed using an unpaired t-test with Welch’s correction (two-sided) for each day. The exact p values (D1, D2, D3): EcN-SGH10 (0.0033, 0.0011, 0.0006); EcN-SGH10ΔwcaJ (0.0405, 0.0169, 0.0002); SGH10-EcN (0.0157, 0.0002, 0.0002). c Donor, recipient, and transconjugant counts from five mice per group for (1) EcN pKPC2 to SGH10; (2) EcN pACYC_traF pKPC2ΔtraF to SGH10 or (3) to SGH10::traF; and (4) EcN::traF pKPC2ΔtraF to SGH10 or (5) to SGH10::traF measured on days 1, 2 and 3 in mouse stool. The subset of EcN pACYC_traF donors harboring both pKPC2ΔtraF and pACYC_traF is denoted as Donor* (pink dotted line). d Corresponding log10-transformed T/R for the five groups on days 1, 2 and 3. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each group included five biological replicates. Statistical analysis was conducted using a one-way ANOVA with Tukey’s multiple-comparison test. Although multiple pairwise comparisons were performed, only the results comparing each group with group (1) on the same day are shown in the graph. Additional pairwise comparisons (group 2 vs. 3, group 4 vs. 5, group 2 vs. 4, and group 3 vs. 5) are presented in Supplementary Data 3. The exact p values (D1, D2, D3): group 2 vs. 1 (<0.0001, <0.0001, <0.0001); group 3 vs. 1 (0.3221, 0.9888, 0.9875); group 4 vs. 1 (<0.0001, <0.0001, <0.0001); group 5 vs. 1 (0.0211, 0.8771, 0.7888). e Donor (yellow), recipient (blue), and transconjugant (green) counts from gastrointestinal sites (duodenum, jejunum, ileum, cecum, proximal/distal colon) and stool of three mice on day 1. f Log10-transformed T/R of pKPC2-mScarlet3 from EcN to SGH10::sfGFP in gastrointestinal sites and stool of mice on day 1. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Statistical differences were analyzed using one-way ANOVA (Dunnett’s test), with stool samples as the reference group and p values are shown as asterisks (n = 3 mice for gastrointestinal sites and n = 8 for stools). The exact p values (vs. stool): duodenum (0.0003); jejunum (0.0117); ileum (0.2468); cecum (0.9953); proximal colon (0.8709); distal colon (0.9760). g Imaging of cecum and colon sections using maximum intensity projection showing SGH10::sfGFP recipients (green), EcN pKPC2-mScarlet3 donors (red), transconjugants (yellow, white arrowheads in insets), nuclei (blue), and actin (magenta); scale bars = 100 µm (main) and 5 µm (inset). Inset images show a single Z-slice. The distribution of the donor and recipient populations were examined from five mice with consistent results. b, d, f indicate significance (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns not significant).
Our results revealed that secondary transfer capability dramatically influences plasmid spread. When secondary transfer was absent (pairs 2 and 4), the T/R were significantly lower than the wild-type control (Fig. 2d and Supplementary Data 3). It is worth noting that in pairs 2 and 4, SGH10 recipients that acquire pKPC2ΔtraF become conjugatively deficient and cannot participate in subsequent horizontal transfer events, but they retain the ability to propagate the plasmid vertically to their progeny through cell division. The consistently low T/R observed in these pairs, despite unimpeded vertical transmission, demonstrate that clonal expansion through vertical inheritance plays a negligible role in sustaining plasmid populations under these experimental conditions. Conversely, when secondary transfer was present (pairs 3 and 5), the T/R were comparable to the wild-type pair (1) (Fig. 2d and Supplementary Data 3). Notably, T/R were similar between pairs (3) and (5), despite pair (3) having donors that progressively lose pACYC_traF over time (Fig. 2c). This suggests that the number of competent original donors is less critical in the long run than the recipients’ ability to undergo secondary transfer for sustaining transconjugant populations in the gut environment.
To visualize the spatial context of conjugation in vivo, we fluorescently tagged the donor plasmid and recipient bacteria. We constructed pKPC2-mScarlet3 (red fluorescent protein marker) and introduced it into EcN, and we tagged SGH10 with a stable chromosomal sfGFP (green fluorescent protein). Control experiments verified that these modifications did not alter conjugation efficiency (Supplementary Fig. 5a) and produced distinct, strong fluorescent signals (Supplementary Fig. 5b). We then co-colonized mice with the red EcN donor and green SGH10 recipient and examined different gut segments on day 1. Plating results showed that donor and recipient densities were low in the duodenum and jejunum, higher in the ileum, and highest in the cecum and colon. Transconjugants were recovered primarily in the cecum and colon, in numbers proportional to donor and recipient densities (Fig. 2e). Notably, stool samples closely mirrored the T/R observed in the cecum and colon (Fig. 2f), confirming that stool can be used as a proxy for gut-level transfer. Direct imaging of gut tissues corroborated these findings: in cecum and colon sections, donors and recipients were intermingled (with SGH10 predominating and associated with fecal matter) and transconjugant bacteria (appearing as yellow due to co-localization of red and green) were visible within the lumen (Fig. 2g). These microscopy observations provide direct evidence of in situ plasmid transfer in the gut environment.
pKPC2 diverges from canonical IncP plasmids in phylogeny and global distribution
IncP plasmids have recently been classified into two major clades (I and II) based on phylogenetic differences and distinct features34. Prototypical IncP plasmids like RP4 and R751 define clade I, whereas non-canonical plasmids like pKPC2 belong to clade II. Interestingly, pKPC2 has become prevalent in clinical isolates18, whereas typical clade I IncP plasmids are now seldom encountered in clinical settings (even though historically RP4 and R751 were first isolated from hospital patients35,36). Instead, clade I plasmids are more commonly found in environmental contexts such as wastewater treatment plants37, the rhizosphere38, and agricultural environments39.
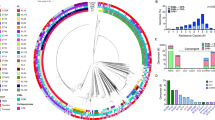
In our experiments, pKPC2 and RP4 exhibited similar conjugation efficiency in vitro (Fig. 1a), yet pKPC2 exhibited higher T/R than RP4 in the mouse gut (Fig. 1d). This suggests that IncP clade II plasmids like pKPC2 may have evolved traits for more efficient transmission in mammalian guts compared to clade I plasmids. To explore this idea, we compared the prevalence of clade II vs clade I IncP plasmids in large genomic datasets. We searched ~1.93 million assembled bacterial genomes from the European Nucleotide Archive (ENA)40 for sequences matching the conserved core of IncP plasmids. The query consisted of concatenated sequences of 22 core type IV secretion system (T4SS) genes (traMLKJIGFEDC; trbNLJIHGFEDCBA)34 plus the replication initiation gene trfA, taken from representative clade II (pKPC2) and clade I (RP4) plasmids. Using ≥95% nucleotide identity and ≥95% coverage as cutoffs, we identified 2809 pKPC2-like sequences versus only 121 RP4-like sequences in the ENA dataset (Fig. 3a; Supplementary Fig. 6a–c). The majority of pKPC2-like hits came from human-associated samples (1298), with fewer from animals (113) and environmental sources (50). In contrast, most RP4-like hits were from laboratory isolates (73), with only 15 from human samples. Thus, within this collection, pKPC2-related plasmids are ubiquitous across clinical and non-clinical contexts, whereas RP4-related plasmids are relatively rare outside of lab strains.
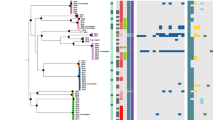
a Sankey diagram showing the total number of IncP clade I RP4-like sequences (n = 121 sequences) and pKPC2-like sequences (n = 2809 sequences) identified in the European Nucleotide Archive (up to May 2023) as well as the source of the isolates. b ARG distribution in pKPC2-like sequences. c Phylogenetic tree of IncP clade I and clade II plasmids based on 22 conserved T4SS genes plus trfA. Scale bar denotes nucleotides substitution per site. d Venn diagram illustrating the number of sequences that meet the threshold criteria of 95% query coverage and nucleotide identity for each respective plasmid group. The non-overlapping sections represent the unique sequences for each plasmid group that meet the thresholds, while the overlapping regions indicate shared sequences that satisfy the criteria across two or more plasmid groups.
We next examined the taxonomic distribution of these hits (Fig. 3a). As expected, pKPC2-like sequences were predominantly found in Enterobacterales genera: Escherichia (1034), Klebsiella (865), and Salmonella (660). However, a few occurred in more distantly related bacteria such as Vibrio cholerae, indicating that the host range can extend beyond the Enterobacterales. RP4-like sequences were mainly confined to Escherichia (82 total, mostly laboratory strains) and were sporadically present in other genera, reflecting RP4’s known broad host range41. Notably, consistent with prior reports34 that clade II plasmids do not readily transfer to Pseudomonas, we found no pKPC2-like sequences in Pseudomonas genomes. The detection of pKPC2-related sequences in Vibrio, however, hints that clade II IncP plasmids may occasionally move into other non-Enterobacterales hosts. Geographic metadata (Supplementary Fig. 6d) indicated that the majority of pKPC2-like and RP4-like sequences in ENA originated from Singapore, the USA, and the UK (likely reflecting sampling biases), but collectively they spanned all major continents, underscoring a global distribution. We also observed that pKPC2-like sequences from human-associated isolates carried a richer complement of ARGs compared to those from animal or environmental sources (Fig. 3b; Supplementary Fig. 6e).
As plasmids carrying the same replicon and T4SS may not always be similar at the overall plasmid genomic level, we sought to determine whether pKPC2-like and RP4-like sequences identified earlier exhibit whole-genome similarity. For this analysis, the sequences were categorized into four groups based on their similarity to reference T4SS and replicon sequences: pKPC2_95 (≥95% identity and coverage to pKPC2 components), pKPC2_sub95 (≥80% identity and coverage to pKPC2 components but <95% for identity or coverage), RP4_95 (≥95% identity and coverage to RP4 components), and RP4_sub95 (≥80 identity and coverage to RP4 components but <95% for identity or coverage). We also included pKPC2 and RP4 as reference sequences for comparison. Pairwise average nucleotide identity (ANI) calculations were performed across all sequences using FastANI, followed by network clustering using Louvain community detection applied to similarity relationships meeting thresholds of ≥70% ANI and ≥50% coverage. Network analysis revealed pKPC2-like and RP4-like sequences forming distinct clusters with complete absence of inter-cluster connections, confirming evolutionary divergence at the whole-genome level between pKPC2 and RP4 (Supplementary Fig. 7). The pKPC2-like sequences segregated into two separate clusters: Cluster 0 predominantly contained pKPC2_95 sequences including the reference pKPC2 sequence, while Cluster 1 was dominated by pKPC2_sub95 sequences. Both pKPC2 clusters maintained internal connectivity through ≥70% ANI and ≥50% coverage relationships, indicating they represent evolutionarily related but distinct plasmid variants with higher intra-cluster than inter-cluster genomic similarity. In contrast, RP4-like sequences formed a single cohesive cluster (Cluster 2) comprising both RP4_95 and RP4_sub95 sequences including the reference RP4 sequence, suggesting less genomic diversification within the RP4-like sequences compared to pKPC2-like sequences. This clustering pattern demonstrates that whole-genome architecture, rather than solely replicon and T4SS similarity, governs the evolutionary relationships among these plasmids.
To further validate the prevalence difference between clade II and clade I plasmids, we scanned the IMG/PR plasmid database42 (a non-redundant set of ~700,000 plasmid sequences from isolate and metagenome data) using a slightly relaxed BLAST threshold (≥95% query coverage and ≥60% identity to the core genes and replicon). Consistent with the ENA survey, we found 85 pKPC2-like plasmids versus only three RP4-like plasmids in IMG/PR (Supplementary Fig. 8). To see if this dominance of clade II holds for other members of each clade, we included two additional clade I plasmids (pMNCN064 and pYKBP039, isolated from environmental samples in Japan34) and two clade II plasmids (pVS0309 and pVS1035, that we have isolated from Salmonella causing diarrhea in Vietnam43) in the ENA search. A phylogenetic tree of these plasmids alongside RP4 and pKPC2 illustrates their relationships (Fig. 3c). We found that sequences related to the clade II examples were consistently more numerous than those related to the clade I examples in the global data (Fig. 3d).
To determine whether clade I and clade II IncP plasmids fall under different plasmid taxonomic units (PTUs), a classification system proposed to group plasmids into consistent evolutionarily meaningful categories, we analyzed these sequences using the COPLA pipeline44,45. However, COPLA could not assign PTUs to several plasmids due to insufficient cluster sizes. Specifically, pKPC2 was assigned PTU-?, denoting a novel unnamed PTU, whereas RP4 remained unassigned because its cluster contained only 3 plasmids (sHSBM size of 3), falling below COPLA’s minimum threshold of 4 members required for PTU designation. To complement this analysis, we performed network clustering using Louvain community detection with thresholds of ≥70% ANI and ≥50% coverage, incorporating the respective COPLA PTU assignments for visualization. Network analysis revealed that clade I and clade II plasmids predominantly segregated into two distinct clusters, with one notable exception: pQKH54, a clade I plasmid that failed to cluster with other clade I sequences, suggesting it is genetically more divergent despite having related T4SS and replicon sequences (Supplementary Fig. 9). This clustering pattern demonstrates that clade I and clade II plasmids exhibit high overall genomic similarity within their respective groups but maintain distinct evolutionary trajectories between clades. Among the 12 clade I plasmids that formed a cohesive cluster, 5 were assigned PTU-P1 by COPLA, while the remaining 7 were left undefined due to insufficient sHSBM cluster sizes. Similarly, among the 10 clade II plasmids clustering together, 8 received PTU-? assignments from COPLA, indicating they belong to a novel PTU, while 2 remained undefined. We propose the designation “PTU-P2” for the clade II plasmids to reflect their evolutionary relatedness to the IncP family and to acknowledge that they constitute a distinct taxonomic unit from clade I, of which most members belong to PTU-P1. In this study, we subsequently refer to IncP clade I plasmids that cluster with PTU-P1 as PTU-P1, and to clade II plasmids that cluster with pKPC2 as PTU-P2.
We also compared representative PTU-P1 and PTU-P2 plasmid sequences using a whole-genome alignment visualization tool (AliTV)46. As expected, homology between PTU-P1 and PTU-P2 plasmids was largely confined to the conserved backbone (transfer, replication, maintenance, and regulatory regions). Even within these shared regions, the sequence identity was modest (Supplementary Fig. 10), indicating substantial evolutionary divergence. The accessory gene content differed greatly between the PTUs, although each PTU harbors some specific hypothetical genes of unknown function that are commonly shared among plasmids within that PTU.
PTU-P2 plasmids transfer more efficiently in the gut than PTU-P1 plasmids
The prevalence and dominance of PTU-P2 over PTU-P1 plasmids may be linked to differences in transmission efficiency. We hypothesized that the greater success of PTU-P2 plasmids might stem from higher transfer efficiency, particularly under gut conditions. To investigate this, we directly compared three PTU-P1 (RP4-11, pMNCN064, pYKBP039) and three PTU-P2 plasmids (pKPC2, pVS0309, pVS1035) in our experimental model. In vitro, all six plasmids transferred at relatively high frequencies after 4 h of mating on filters (Fig. 4a), with some variability but no systematic difference under these aerobic conditions. We also assessed plasmid stability and fitness costs associated with each plasmid. In EcN grown in rich medium, two plasmids (PTU-P1 pMNCN064 and PTU-P2 pVS1035) showed gradual instability over ~90 generations, whereas the others remained fully stable. Notably, all plasmids were stably maintained in SGH10 (Fig. 4b). The same two plasmids (pMNCN064 and pVS1035) imposed the greatest fitness burden on EcN (slower growth relative to the no-plasmid control), and even RP4-11 and pYKBP039 caused a measurable growth defect in EcN. By contrast, none of the plasmids appreciably affected the growth of SGH10 (Supplementary Fig. 11).
a Log10-transformed conjugation efficiency (transconjugant per recipient) of plasmid transfer from E. coli Nissle (EcN) to K. pneumoniae SGH10. Conjugation assays for PTU-P1 (RP4-11, pMNCN064, pYKBP039) and PTU-P2 (pKPC2, pVS0309, pVS1035) plasmids were performed on nitrocellulose membranes atop LB agar, incubated at 37 °C for 4 h. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents one biological replicate (n = 9) from three independent experiments. Statistical differences were analyzed using one-way ANOVA with Tukey’s correction, with significant p values indicated by asterisks. The exact p values: pKPC2 vs. pMNCN064 (<0.0001); pKPC2 vs. pVS0309 (<0.0001); pMNCN064 vs. pVS1035 (0.0452); pMNCN064 vs. RP4-11 (<0.0001); pVS0309 vs. pVS1035 (0.0385); pVS0309 vs. RP4-11 (<0.0001). b Plasmid retention in EcN and SGH10 over 90 generations in LB, based on three independent experiments. Each dot represents the mean of three biological replicates, with error bars showing standard deviation. c Log10-transformed transconjugant-to-recipient ratios (T/R) of PTU-P1 and PTU-P2 plasmids from EcN to SGH10 measured in mouse stool (n = 5 per group) on days 1, 2, and 3. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Symbols at the bottom indicate the absence of transconjugants. PTUs (n = 5 mice per plasmid) were compared using an unpaired t-test with Welch’s correction (two-sided) with significant p values shown as asterisks. The exact p values (D1, D2, D3): PTU-P1 vs. PTU-P2 (<0.0001, <0.0001, <0.0001).
In the mouse gut colonization experiments, however, a clear difference emerged. The PTU-P2 plasmids consistently achieved higher T/R than the PTU-P1 plasmids (Fig. 4c). This disparity occurred even though their in vitro transfer efficiencies were comparable. These results suggest that PTU-P2 plasmids possess adaptations enabling more efficient transfer in the gut environment than PTU-P1 plasmids.
Anaerobicity approximates plasmid transfer dynamics in the mammalian gut
A major environmental difference between our standard lab matings and the gut is the presence or absence of oxygen. We hypothesized that anaerobic conditions might underlie the superior gut transfer ability of PTU-P2 plasmids. Indeed, when we performed in vitro conjugation assays under strictly anaerobic conditions (Fig. 5a), PTU-P1 plasmids (RP4-11, pMNCN064, pYKBP039) transferred at significantly lower frequencies than the PTU-P2 plasmids (pKPC2, pVS0309, pVS1035), mirroring the pattern observed in vivo (Fig. 4c). Previous studies have similarly noted that certain PTU-P1 plasmids (e.g., pB10 and pB136) suffer reduced transfer efficiency in anaerobiosis47,48. Thus, oxygen availability appears to be a crucial factor. PTU-P1 plasmids, which perform well in oxic environments, are at a disadvantage in the anoxic gut, whereas PTU-P2 plasmids maintain efficient transfer under anaerobic conditions consistent with their success in clinical (gut-associated) settings versus PTU-P1’s prominence in environmental niches.
a Log10-transformed conjugation efficiency of PTU-P1 (RP4-11, pMNCN064 and pYKBP039) and PTU-P2 (pKPC2, pVS0309 and pVS1035) plasmids from EcN to SGH10 measured under anaerobic conditions after 4 h of filter mating. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents one biological replicate from a total of nine biological replicates for pKPC2 and RP4-11 or a total of twelve biological replicates for pMNCN064, pYKBP039, pVS0309 and pVS1035 from three independent experiments. Significant differences between PTU-P1 (n = 9 for RP4-11, n = 12 for pMNCN064 and pYKBP039) and PTU-P2 (n = 9 for pKPC2, n = 12 for pVS0309 and pVS1035) plasmids were determined using an unpaired t-test with Welch’s correction (two-sided) and are indicated by asterisks. The exact p value (PTU-P1 vs. PTU-P2): <0.0001. b Log10-transformed conjugation efficiency of pKPC2, RP4-11, pNDM1, and TP114 from EcN into SGH10 and SGH10ΔwcaJ measured under anaerobic conditions after 4 h of filter mating. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents one biological replicate (n = 9) from three independent experiments. Significant differences between the anaerobic transfer efficiencies were determined using an unpaired t-test with Welch’s correction (two-sided) and are marked with asterisks. The exact p values (SGH10 vs. SGH10 ΔwcaJ): pKPC2 (0.9019); pNDM1 (0.9061); RP4-11 (0.0004); TP114 (0.0140). For comparison, aerobic data from Fig. 1a are shown in light overlay. Insets display the fold-change in conjugation efficiency into SGH10ΔwcaJ relative to SGH10 under respective aerobic and anaerobic conditions. c Mucoviscosity tests (supernatant-to-culture OD600 ratio) and d uronic acid quantification (µg/mL per 109 CFU) for SGH10 grown aerobically and anaerobically, with SGH10ΔwcaJ serving as the negative control. The center line of the boxplot in c denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Error bars in d represent mean ± SD. Each dot represents a biological replicate (n = 7 each for SGH10 aerobic and anaerobic; n = 5 SGH10ΔwcaJ) from three independent experiments. For both (c, d), one-way ANOVA with Tukey’s correction was used to assess statistical significance, with significant p values shown as asterisks. The exact p values (c, d): aerobic vs. anaerobic (<0.0001, 0.1408); aerobic vs. control (<0.0001, <0.0001); anaerobic vs. control (0.0673, <0.0001). e Log10-transformed conjugation efficiency of pKPC2, pNDM1, RP4-11 and TP114 from EcN into SGH10 (blue circles) and its mutants (SGH10ΔrmpA (orange triangles), SGH10ΔrmpC (pink squares), SGH10ΔrmpD (green plus signs), and SGH10ΔwcaJ (square with cross)) measured under aerobic conditions after 4 h of filter mating. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents a biological replicate (n = 9) from three independent experiments. Statistical significance was assessed using one-way ANOVA (Dunnett’s test), with SGH10 as the reference group. Asterisks indicate statistical significance. The exact p values (SGH10 vs. SGH10ΔrmpA, SGH10 vs. SGH10ΔrmpC, SGH10 vs. SGH10ΔrmpD, SGH10 vs. SGH10ΔwcaJ): pKPC2 (<0.0001, 0.0282, 0.0044, <0.0001); pNDM1 (0.0107, 0.4998, 0.1765, <0.0001); RP4-11 (0.0007, 0.9986, 0.1992, <0.0001); TP114 (<0.0001, 0.0227, 0.6183, <0.0001). f Mucoviscosity tests (supernatant-to-culture OD600 ratio) and g uronic acid quantification (µg/mL per 109 CFU) for aerobically grown SGH10, SGH10ΔrmpA, SGH10ΔrmpC, SGH10ΔrmpD, and SGH10ΔwcaJ. The center line of the boxplot denotes the median, the box bounds indicate the 25th and 75th percentiles, and the whiskers extend to 1.5X the IQR. Each dot represents one biological replicate (n = 4) and the error bars represent mean ± SD. Statistical significance for (f, g) was assessed using one-way ANOVA (Dunnett’s test), with SGH10 as the reference group. Asterisks indicate statistical significance (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001), while “ns” denotes no significant difference. The exact p values for (f): SGH10 vs. SGH10ΔrmpA (<0.0001); SGH10 vs. SGH10ΔrmpC (0.9759); SGH10 vs. SGH10ΔrmpD (<0.0001); SGH10 vs. SGH10ΔwcaJ (<0.0001). The exact p values for (g): SGH10 vs. SGH10ΔrmpA (0.0165); SGH10 vs. SGH10ΔrmpC (0.0123); SGH10 vs. SGH10ΔrmpD (0.8007); SGH10 vs. SGH10ΔwcaJ (<0.0001).
We noticed that the hvKp capsule’s inhibitory effect on plasmid transfer, evident in vitro (Fig. 1a), was less prominent in the in vivo gut environment (Fig. 1c). A possible explanation is that anaerobic conditions in the gut alter capsule properties. We therefore repeated conjugation experiments under anaerobiosis using representative plasmids whose transfer was capsule-inhibited in aerobic assays (pKPC2, RP4-11, pNDM1, TP114). Strikingly, under anaerobic conditions the capsule’s barrier effect was greatly diminished. pKPC2 and pNDM1 transfer into capsulated SGH10 vs non-capsulated SGH10ΔwcaJ became equivalent, and the transfer deficits observed for RP4-11 and TP114 with a capsule were substantially reduced (Fig. 5b). Correspondingly, SGH10 cultured anaerobically had a much lower mucoviscosity than when cultured aerobically (Fig. 5c), even though the total amount of capsular polysaccharide (quantified by uronic acid content) was similar between aerobic and anaerobic growth (Fig. 5d).
To determine whether mucoviscosity rather than the capsule amount inhibits plasmid transfer, we examined the transfer of pKPC2, pNDM1, RP4-11, and TP114 into several capsule regulator mutants under aerobic in vitro conditions by making use of a series of mutants that can distinguish between capsule mucoviscosity versus capsule amount. We tested an rmpA-deficient mutant (SGH10ΔrmpA) with reduced mucoviscosity and capsule, an rmpC-deficient mutant (SGH10ΔrmpC) with reduced capsule but retained mucoviscosity, and an rmpD-deficient mutant (SGH10ΔrmpD) with abolished mucoviscosity but intact capsule (Fig. 5e). Transfer into SGH10ΔrmpA increased compared to wildtype SGH10, though still lower than that observed in SGH10ΔwcaJ which lacks both mucoviscosity and capsule. SGH10ΔrmpC exhibited only a slight increase in line with expectations as mucoviscosity is retained in this mutant. However, transfer into SGH10ΔrmpD was slightly reduced compared to wildtype. Measurements of mucoviscosity (Fig. 5f) and uronic acid content (Fig. 5g) confirmed that these mutants behaved consistently with previous reports49.
One possible explanation is that the rmpD mutation, despite affecting mucoviscosity, may not fully recapitulate the changes induced by anaerobic growth on the capsule, and thus does not significantly increase transfer efficiency. As the rmpC mutation did increase conjugation slightly, it could also mean that both the capsule amount and mucoviscosity need to be titrated to just the right amount for increased conjugation to take place, and this is not completely recapitulated by the individual mutations but is more accurately represented by the rmpA mutation. The rmpA mutation did increase conjugation significantly, though still not to the extent observed under anaerobic conditions. This could mean that besides the influence of the rmp locus, other factors contribute to nullify the effect of capsule. Anaerobic growth likely alters capsule physical properties, reducing the barrier to conjugation by making the cell surface less obstructive. This change may facilitate donor-recipient interactions in a manner similar to capsule-null conditions, enhancing plasmid transfer. Together, these results suggest that in anaerobic environments such as the mammalian gut, the K. pneumoniae capsule presents a reduced barrier to plasmid transfer.
Discussion
In our recent surveillance of carbapenemase plasmids in Singapore hospitals, we found pKPC2 to be the most prevalent carbapenem-resistance plasmid across multiple Enterobacterales species18, and we identified it as a non-canonical IncP plasmid. Historically, most attention has focused on plasmids from the IncF, IncA, IncC, IncL, IncM, IncI, IncH and IncN groups as dominant vectors of clinical AMR spread, with hardly any implicating IncP plasmids50,51. In this study, by comparing pKPC2 with other dominant carbapenemase plasmids of various incompatibility types, we found that conjugation efficiencies measured in vitro do not invariably mirror the transfer dynamics within the gastrointestinal tract. However, specific findings like the preferential transfer of TP114 into EcN and pKpQIL into SGH10 align with in vivo results, demonstrating in vitro assays can provide valuable insights into recipient-specific mating factors that are relevant in an in vivo context.
One of our key findings is the discovery that PTU-P2 plasmids transmit very well in the murine gut and this highlights stark differences between closely related PTU-P1 plasmids. The clinically successful PTU-P2 plasmid pKPC2 vastly outperformed the prototypical PTU-P1 plasmid RP4 in the gut, despite both carrying similar transfer machinery and having comparable efficiency in standard lab matings. Combined with the large-scale genomics survey showing pKPC2-like plasmids are common in human-associated bacteria, whereas RP4-like plasmids are rare, it appears that PTU-P2 plasmids have undergone selection for success in host-associated (particularly gut) settings, unlike PTU-P1 plasmids which seem to thrive mostly in environmental contexts. In other words, even within the broad-host-range IncP family, subtle genetic differences translate to major ecological specialization. Indeed, the additional PTU-P2 plasmids we tested (from clinical isolates) all transferred better in vivo than the environmental PTU-P1 plasmids, reinforcing the idea of niche-specific adaptation. Our experiments pinpoint anaerobic conditions as a critical factor underlying these differences. PTU-P2 plasmids maintained high conjugation rates when we removed oxygen, whereas PTU-P1 plasmids did not. This likely explains why PTU-P2 plasmids flourish in the anaerobic mammalian gut and have become prevalent in hospitals, whereas PTU-P1 plasmids are largely confined to oxygen-rich external environments.
Another of our key findings is the importance of secondary transfer in sustaining plasmid spread in the gut. Both our statistical modeling and the traF-deletion experiments showed that after the initial donor-to-recipient conjugation, further dissemination of pKPC2 depended largely on transconjugants acting as new donors. This phenomenon aligns with findings from a recent study using RP4 in a biofilm model, which showed that secondary transfer is particularly pronounced under conditions of continuous nutrient flow52. We propose that this reflects the gut’s spatial and metabolic structuring, where abundant nutrients may enhance secondary plasmid transfer. Despite co-residence of donor and recipient populations, direct contact may be restricted by interspecies competition or contact-dependent inhibition mechanisms, such as bacteriocin production or type VI secretion systems, which inhibit close competitors. Thus, once a transconjugant arises, it may have better access to nearby plasmid-free bacteria of its own species, enabling plasmid propagation within that community. In practical terms, the first inter-species transfer event seeds a new population of plasmid carriers, and thereafter the plasmid can persist and spread even if the original donor strain dwindles. This mode of dissemination, where subsequent intrapopulation transfers among recipients drive the maintenance of a plasmid, emphasizes the need to consider community context (not just donor-recipient pair attributes) when evaluating plasmid transmission potential.
More broadly, our work also revises the paradigm of the hypermucoviscous capsule as a barrier to gene exchange. Its impact is highly context dependent and modulated by environmental cues such as oxygen availability. While in vitro assays under aerobic conditions consistently show reduced conjugation into hvKp due to its capsule18,26,53, epidemiological genomics clearly demonstrate that hvKp lineages do acquire resistance plasmids in real-world settings54,55,56. We found that under anaerobic conditions, the inhibitory effect of the capsule was markedly reduced. Although the capsule remained present, it became far less mucoviscous. This phenotypic shift aligns with other studies under microaerobic growth conditions reporting reduced expression of capsule biosynthesis genes with key regulators downregulated in low-oxygen environments57, potentially through repression by the oxygen-sensitive transcription factor FNR58. In contrast, in oxygen-rich environments such as the lung, modulation of capsule production and mucoviscosity can be critical for virulence to enhance resistance to phagocytosis and complement30,59. However, this may inadvertently limit HGT by hindering conjugation. These findings raise the possibility that hvKp experiences niche-specific trade-offs through environmentally tuned regulation of its capsule. Such context-dependent modulation of capsule traits may be a key evolutionary strategy for balancing colonization, gene acquisition, and virulence across distinct body sites. HvKp is clearly capable of participating in conjugative transfer in vivo, meaning that its observed lower carriage of resistance elements60 might owe more to ecological exposure (limited contact with resistance plasmids) than to any intrinsic barrier. If and when hvKp does acquire a conjugative resistance plasmid, it could potentially use that plasmid as a bridge to mobilize its own large virulence plasmid (which are usually non-conjugative) into other K. pneumoniae strains61. This scenario would foster dangerous hybrids combining hypervirulence and drug resistance causing a public health nightmare.
There are some limitations to our study. First, our use of a broad-spectrum antibiotic to perturb the murine microbiota undoubtedly exaggerates plasmid transfer by abolishing colonization resistance and permitting Enterobacterales overgrowth. Experimental models of gut inflammation and human cohort data show that antibiotic-induced disruption of the microbiota unleashes dramatically elevated rates of HGT62,63,64. In our antibiotic-treated mice, EcN and SGH10 co-occurred at high density with minimal apparent competition, an ecological context that may mirror long-term antibiotic therapy in patients, where diverse Enterobacterales species bloom and freely exchange plasmids65. In contrast, plasmid dissemination is likely far more constrained in an intact, unperturbed microbiota. This study did not address the influence of an unperturbed microbiota on plasmid transfer, nor its potential impact on the transfer into capsulated versus non-capsulated strains. Second, in our model, the donor-to-recipient ratio does not differ substantially. A previous study examining plasmid transfer of Salmonella in the mouse gut demonstrated a positive correlation between donor density and transconjugant yield62. The relative contribution of secondary transfer in the gut may differ under conditions where recipients are rare and donors are abundant. Because plasmid transfer requires physical contact, a high donor density could facilitate primary transfer events, unless donors and recipients occupy distinct ecological niches within the gut, in which case the dynamics may favor secondary transfer. The balance between primary and secondary transfer is therefore likely context-dependent and may also be influenced by microbiome composition. In an intact microbiome, competition for ecological niches could substantially alter transfer dynamics. While our model, with roughly comparable donor and recipient abundances, demonstrated transconjugant-driven spread, other scenarios may produce different outcomes. For example, when the ecological niche available to recipients is much smaller than that available to donors, such as in competitive microbiome environments where donors have colonization advantages, primary donor-to-recipient transfer may outpace transconjugant-to-recipient transfer. This consideration is particularly relevant in natural gut microbiomes, where intense niche competition may constrain the ability of transconjugants to establish themselves. Future studies investigating the conjugation dynamics of hvKp in more complex microbial communities will be essential to determine how these principles operate across diverse ecological contexts. Third, although our anaerobic in vitro conjugation results align with in vivo observations, they remain simplified models that cannot capture the full complexity of the gut environment including its spatial microheterogeneity, mucosal architecture, and dynamic nutrient gradients. While we observed that anaerobicity influences mucoviscosity under in vitro conditions, the interplay of factors in the gut milieu is likely to be far more intricate, and anaerobicity may not even be the primary factor influencing capsule dynamics. It is possible that other capsular biosynthetic machinery and regulatory factors apart from rmpA influence capsule dynamics in the gut, contributing to improved conjugation. Lastly, the rmpD deficient mutant did not achieve the high conjugation frequencies seen under anaerobic conditions hints at unappreciated roles for capsule biophysics such as polysaccharide arrangement, permeability, or regulation that may differ between aerobic and anaerobic conditions. Dissecting how capsule properties differ between aerobic and anaerobic in vitro conditions, and how they manifest in the complex gut environment will be an important goal for future studies. It is also possible that anaerobicity increased the process of conjugation that is independent of capsule but influenced by other bacterial factors. In terms of the differential response between PTU-P1 and PTU-P2 plasmids under anaerobicity, we did not find clear patterns of oxygen-sensitive regulators FNR or ArcA consensus sequences on the plasmids that would indicate differential plasmid gene regulation under anaerobic condition. This indicates a much more complex interaction likely with bacterial chromosomal genes that could influence their conjugation capability and awaits further investigation.
In conclusion, this study highlights three key findings. First, pKPC2 forms a distinct PTU within the IncP group (where most canonical IncP plasmids correspond to PTU-P1), and these PTU-P2 plasmids transfer more efficiently in the gut than PTU-P1 plasmids. This disparity reflects their differential sensitivity to anaerobic conditions, establishing oxygen availability as an important determinant of HGT for such plasmids in the gut environment. Second, pKPC2 and other PTU-P2 plasmids are significantly more prevalent in human-associated samples as observed from the global sequence database, whereas RP4-like PTU-P1 plasmids are underrepresented in these niches. This distribution supports ecological specialization within the IncP group, with pKPC2 representing a group of plasmids adapted to anoxic gut conditions. Third, the hypervirulent K. pneumoniae capsule functions as a more permeable barrier to plasmid transfer than previously recognized, as transfer into capsulated recipients shows less pronounced differences compared with non-capsulated recipients under in vivo gut conditions. Together, these findings demonstrate how microenvironmental factors fundamentally reshape plasmid transmission dynamics, with important implications for predicting and mitigating antimicrobial resistance dissemination in clinical settings.
Methods
Strains and plasmids
K. pneumoniae SGH10 and E. coli Nissle were cultured on LB agar plates at 37 °C. Strains harboring the tested plasmids were grown on LB agar supplemented with 50 μg/mL kanamycin (Km) for selection. The bacterial strains and plasmids used in this study are detailed in Supplementary Table 1.
In vitro conjugation
Donor and recipient strains were streaked on LB agar plates with and without 50 μg/mL kanamycin (Km), respectively, and incubated overnight at 37 °C. For the conjugation assay, 20 μL of the donor-recipient mixture, prepared in 1X PBS at a 1:1 ratio (OD600 = 0.5), was spotted onto a nitrocellulose membrane placed on LB agar and incubated for 4 h at 37 °C. After incubation, the bacterial mixture was plated on appropriate selective agar containing a combination of Km (50 μg/mL), fosfomycin (Fm, 40 μg/mL) or chloramphenicol (Cm, 50 μg/mL) to select for recipient and transconjugant populations. The conjugation frequency was calculated by dividing the number of transconjugants by the total number of recipients. For anaerobic conjugation experiments, bacterial cultures were pre-incubated overnight at 37 °C under anaerobic conditions prior to the conjugation assays. The anaerobic conjugation procedure mirrored the standard aerobic method described previously, with the key difference that the strains were pre-grown in the Coy Vinyl Anaerobic Chamber at a controlled temperature of 37 °C before the conjugation experiment. While the conjugation assays were conducted entirely within the chamber, the antibiotic selection plates were incubated aerobically at 37 °C to maintain the efficacy of antibiotic selection.
Mucoviscosity and uronic acid quantification
The SGH10 strains were grown as described for the conjugation experiment. Both aerobically and anaerobically grown strains were resuspended in 1 × PBS before measurements. For mucoviscosity, we centrifuged samples at 2000 × g for 5 min. The mucoviscosity was calculated as the ratio of the supernatant OD600 after centrifugation to the initial culture OD600, with a lower ratio indicating low mucoviscosity and a higher ratio indicating high mucoviscosity. For capsule quantification, we measured uronic acid content by mixing 500 μL of culture with 100 μL of extraction buffer (100 mM citric acid, pH 2, and 1% Zwittergent 3–14) and incubating the mixture at 56 °C for 20 min. After centrifugation at 17,000 × g for 5 min at 4 °C, 200 μL of the supernatant was mixed with 800 μL of ice-cold ethanol. The mixture was incubated on ice for 20 min and centrifuged again. The pellet was then washed with 1 mL of ice-cold 70% ethanol and dissolved in 100 μL of water. Next, a hydrolysis solution containing 0.0125 M sodium tetraborate decahydrate in concentrated H₂SO₄ was added. The mixture was boiled at 100 °C for 5 min, followed by the addition of 10 μL of 0.15% carbazole in absolute ethanol, and then boiled again at 100 °C for 15 min. After spinning down the sample, 200 μL was transferred for OD measurement at 520 nm using a Tecan Infinite M200 plate reader. The OD values were converted to μg/10⁹ CFU using a standard curve prepared with known glucuronic acid standards and the CFU count of the culture. An aerobically grown non-capsulated mutant was used as a negative control for both assays.
In vitro plasmid stability
Plasmid-bearing bacterial strains were initially grown overnight in LB with 50 μg/mL Km, designated as Generation 0. Each subsequent day, the bacterial culture was subcultured at 1:210 dilution into 5 mL LB, resulting in an increment of 10 generations daily, as previously described66. At generations 0, 30, 60, and 90, the bacterial cultures were serially diluted and plated on LB agar with and without 50 μg/mL Km. Plasmid stability was assessed by calculating the ratio of Km-resistant bacteria to the total bacterial count.
Bacterial growth assays
Bacterial cultures were grown in LB medium supplemented with 50 μg/mL Km for plasmid-bearing strains and in LB alone for the no-plasmid control. The cultures were incubated overnight in a shaking incubator at 37 °C and 150 rpm. We measured the optical density at 600 nm (OD600) of the overnight culture and diluted it to approximately an OD600 of 0.001 in LB medium. We then added 200 μL of the diluted cultures to a 96-well plate and incubated it at 37 °C in a Tecan Infinite M200 plate reader. Absorbance at OD600 was measured hourly for 24 h.
Mouse model for plasmid transfer
The animal research procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) at Comparative Medicine, National University of Singapore (protocol number: R21-1498). Female C57BL/6J mice, aged 7–8 weeks, were obtained from the Jackson Laboratory through InVivos Pte. Ltd. They were housed in groups of up to five per cage under specific pathogen-free conditions with a 12 h dark/light cycle. Room temperature is maintained at 20–24 °C and humidity around 40–60%. They were given access to food and water ad libitum.
After an acclimatization period, the mice were administered 100 μL of 1X PBS containing 2.5 mg of ampicillin sodium salt (Sigma) via oral gavage for three consecutive days, under isoflurane anesthesia. One day after the final ampicillin dose, the mice were orally gavaged with either SGH10 or SGH10∆wcaJ, at a concentration of 104 or 105 CFU, respectively. Six hours later, E. coli Nissle KN02 was gavaged at a density ten times higher than that of the hvKp strains. Stool samples were collected in 2 mL 1.4 mm ceramic OmniTubes (Omni International) on days 1, 2, and 3 after the donor-recipient gavage. Samples were homogenized using a Bead Ruptor (Omni International) for two 30-second cycles at medium setting. The resulting stool slurry was serially diluted and plated on LB agar with appropriate selective antibiotics (a combination of 50 μg/mL Km, Cm or Fm) to quantify donor, recipient, and transconjugant populations.
Plasmid modification
E. coli MG1655::CmR carrying pKPC2 was mixed with E. coli S17–1 λpir harboring pR6KTc, which contained homologous regions to the upstream and downstream sequences of the traF gene on pKPC2 (pR6KTc_traF). The pR6KTc_traF plasmid was constructed using HiFi assembly of EcoRI/PstI-digested pR6KTc vector, along with PCR-amplified upstream and downstream regions of traF, using Phanta Flash Super-Fidelity DNA polymerase (Vazyme). The insert was validated via Sanger sequencing by a commercial company (1st Base). After 2 h incubation at 37 °C, the MG1655 transconjugants were selected on LB agar plates containing 10 μg/mL tetracycline (Tc), 50 μg/mL Cm and Km. To remove the integrated pR6K fragment, the strain was grown in LB with 15% sucrose, followed by colony PCR to confirm deletion of traF. The complementation plasmid pACYC_traF was assembled by combining PCR-amplified fragments of pACYC184, the gentamicin (Gm) resistance gene from pRK2_AraE, and the traF gene from pKPC2. The assembled plasmid was introduced into E. coli HST08 competent cells via heat shock and selected on LB agar with 35 μg/mL Gm. A list of primers used is provided in Supplementary Table 2.
Chromosomal insertion of traF
The plasmid backbone of pUC18R6KT-mini-Tn7T-Km, containing the R6K origin and the Tn7 transposon, was PCR-amplified together with an insert fragment derived from pACYC_traF that included the traF gene and a Gm resistance marker. The two fragments were assembled using HiFi DNA assembly, and the resulting construct was introduced into E. coli S17–1 λpir by electroporation. For chromosomal integration, S17–1 λpir carrying the assembled plasmid was mixed with either SGH10 or EcN KN02, along with a helper strain harboring pJMP1039, which encodes the Tn7 transposase genes and also carries the R6K origin. Triparental mating was performed, and transconjugants of SGH10 and EcN were selected on antibiotic-containing plates. Successful integration of traF into the chromosome was confirmed by PCR and further validated phenotypically using the pKPC2ΔtraF plasmid.
Visualization of in situ conjugation by microscopy
Mice were subjected to the same procedure as previously described and gavaged with SGH10::sfGFP (recipient) and E. coli Nissle KN03 (donor) carrying pKPC2-mScarlet3. One day after the bacterial gavage, stool samples were collected prior to euthanizing the mice using CO2. Gastrointestinal tissues, including the duodenum, jejunum, ileum, cecum, proximal colon and distal colon, were harvested. Portions of these tissues were homogenized using a Bead Ruptor (Omni International) to quantify donor, recipient and transconjugant populations, while the remaining tissues were prepared for microscopy. Each gut segment was dissected and immediately snap-frozen in Tissue-Tek O.C.T. (Sakura) within cryomolds using liquid nitrogen and stored at −80 °C. For the cecum, the distal portion was dissected, and for the colon segments, regions containing a fecal pellet were collected whenever possible. Confocal microscopy was performed using the Leica Thunder Imaging System and the Stellaris Confocal Microscope, with image processing carried out via LAS X Core Software.
Determination of pKPC2-like and RP4-like sequences in global databases
The ENA sequences (up to May 2023) of 1,932,812 assemblies40 and their associated metadata were retrieved on 13 April 2024 from https://github.com/AllTheBacteria/ AllTheBacteria to create a local blast database. BLASTN (BLAST + 2.13.0) was performed using the concatenated core T4SS genes (traMLKJIGFEDC; trbNLJIHGFEDCBA) and the trfA replicon sequences from IncP clade I and II plasmids as queries. Matching sequences were filtered using a threshold of 95% for both percent identity and query coverage. Metadata for sequences meeting these criteria were subsequently retrieved for further analysis. ARG carriage was assessed using ResFinder v4.6.067,68. BLAST results, along with data on sources, contig lengths, and geographical distributions, were visualized using ggplot269, Matplotlib70, or Plotly. Additionally, the core T4SS and trfA nucleotide sequences were aligned with ClustalW, and a phylogenetic tree was constructed using the maximum-likelihood method in Mega 1171 with default parameters. Furthermore, 699,973 plasmid sequences from the non-redundant plasmid database IMG/PR42 (https://genome.jgi.doe.gov/portal/IMG_PR), derived from genomes and metagenomes, were retrieved on 16 September 2024 to create another local BLAST database for a similar BLASTN analysis, applying thresholds of 95% query coverage and 60% nucleotide identity.
Plasmid sequence network analysis
Fasta sequences were retrieved from the ENA databases, comprising 2809 pKPC2_95 sequences (≥95% identity and coverage to T4SS and replicon), 2427 pKPC2_sub95 sequences (≥80% identity and coverage, but <95%), 121 RP4_95 sequences, and 10 RP4_sub95 sequences, with pKPC2 and RP4 included as references, totaling 5369 sequences. We performed all-vs-all pairwise genomic similarity using FastANI v1.3472, generating 14.4 million unique undirected comparisons. Results were filtered by ANI ≥ 70% and coverage ≥50%. To handle the large dataset, a stratified sampling approach was applied for network visualization: up to 20,000 intra-cluster edges to maintain connectivity, 10,000 inter-cluster edges to illustrate relationships, 5000 edges involving reference sequences, and 3000 additional edges with ANI ≥ 95%. The network was constructed using NetworkX v3.2, with nodes representing plasmid sequences and edges representing similarity based on ANI and coverage. Clustering was performed using the Louvain community detection algorithm in NetworkX, which iteratively optimizes modularity in two phases: (1) local optimization, where each node is assigned to its own community and moved to neighboring communities to maximize modularity gain, and (2) community aggregation, where nodes in the same community are merged into super-nodes, repeating until no further improvement is possible73. For clade I and II plasmids, sequences were further analyzed using command-line COPLA and counterchecked against the output from Web-COPLA (https://castillo.dicom.unican.es/copla/)44,45. PTU assignments from COPLA were used to label nodes in the network.
Bayesian regression model
Statistical tests on the experimental data were conducted using BRMS v2.22.074, installed in R v4.4.075 on a MacOS machine. We implemented a hierarchical Bayesian regression model76 to examine the relationship between Change_Transconjugants (outcome) and three predictors: Donor_Count, True_Recipient_Count, and Transconjugants_Count.
We modeled per-mouse day-to-day changes in transconjugant counts as the outcome. For each subsequent day (D1 → D2 and D2 → D3), predictors were the donor, recipient and transconjugant counts measured on the preceding day. Because no D0 measurements were available, the model did not include D0 → D1. The model included random intercepts for Plasmid and Donor_Recipient_Pair to account for the hierarchical structure, with variance components estimated at both levels. We specified weakly informative normal priors (mean = 0, SD = 10) for all fixed effects and the intercept. The posterior was sampled using Hamiltonian Monte Carlo with 4 chains of 4000 iterations each (1000 warmup), adapt_delta = 0.99, and max_treedepth = 15. All chains converged successfully (R-hat <1.01, high ESS). Effect sizes are reported as unstandardized regression coefficients with 95% credible intervals, representing the expected change in transconjugant counts per unit increase in each predictor.
Statistical analysis
Statistical differences were analyzed using a two-tailed t-test with Welch’s correction for comparisons between two groups, one-way ANOVA with Tukey’s correction for comparisons involving more than two groups and one-way ANOVA with Dunnett’s correction for multiple groups compared to a reference group. For conjugation frequency, log10-transformed data were used for statistical analysis. In cases where the conjugation frequency was 0 and could not be log10-transformed, a value of −7, representing the lowest frequency, was used in place of 0 for the purposes of analysis. The statistical analyses were performed using the built-in function on GraphPad Prism v.10.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The outputs of the ANI analysis and network clustering generated in this study have been deposited in the Zenodo database available at https://zenodo.org/records/1706031277. The NCBI accession ID of the plasmids used in the analysis of the IncP T4SS and replicon against the global dataset are as follows: pKPC2 is NZ_MN542377.1 [https://www.ncbi.nlm.nih.gov/ nuccore/NZ_MN542377.1] and RP4 is BN000925.1 [https://www.ncbi.nlm.nih.gov/nuccore/ BN000925.1]. Source data are provided with this paper.
Code availability
All R code used for data cleaning, transformations, and modeling for the Bayesian Regression Model is available in the GitHub repository: https://github.com/mlgh-sg/plasmids-amr-modelling78.
References
Carattoli, A. Plasmids and the spread of resistance. Int. J. Med. Microbiol. 303, 298–304 (2013).
Meynell, E. & Datta, N. The relation of resistance transfer factors to the F-factor (sex-factor) of Escherichia coli K12. Genet. Res. 7, 134–140 (1966).
Burkardt, H. J., Riess, G. & Puhler, A. Relationship of group P1 plasmids revealed by heteroduplex experiments: RP1, RP4, R68 and RK2 are identical. J. Gen. Microbiol. 114, 341–348 (1979).
Neil, K. et al. High-efficiency delivery of CRISPR-Cas9 by engineered probiotics enables precise microbiome editing. Mol. Syst. Biol. 17, e10335 (2021).
Lam, M. M. C. et al. Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination. Nat. Commun. 9, 2703 (2018).
Leon-Sampedro, R. et al. Pervasive transmission of a carbapenem resistance plasmid in the gut microbiota of hospitalized patients. Nat. Microbiol. 6, 606–616 (2021).
Chen, L. et al. Comparative genomic analysis of KPC-encoding pKpQIL-like plasmids and their distribution in New Jersey and New York Hospitals. Antimicrob. Agents Chemother. 58, 2871–2877 (2014).
Arcari, G. et al. Multiplicity of bla(KPC) Genes and pKpQIL Plasmid Plasticity in the Development of Ceftazidime-Avibactam and Meropenem Coresistance in Klebsiella pneumoniae Sequence Type 307. Antimicrob. Agents Chemother. 67, e0036823 (2023).
Palkovicova, J. et al. Fitness effects of blaCTX-M-15-harbouring F2:A1:B- plasmids on their native Escherichia coli ST131 H30Rx hosts. J. Antimicrob. Chemother. 77, 2960–2963 (2022).
Pitout, J. D. D. & Finn, T. J. The evolutionary puzzle of Escherichia coli ST131. Infect. Genet. Evol. 81, 104265 (2020).
DelaFuente, J., Diaz-Colunga, J., Sanchez, A. & San Millan, A. Global epistasis in plasmid-mediated antimicrobial resistance. Mol. Syst. Biol. 20, 311–320 (2024).
Benz, F. & Hall, A. R. Host-specific plasmid evolution explains the variable spread of clinical antibiotic-resistance plasmids. Proc. Natl. Acad. Sci. USA 120, e2212147120 (2023).
Lipworth, S. et al. The plasmidome associated with Gram-negative bloodstream infections: a large-scale observational study using complete plasmid assemblies. Nat. Commun. 15, 1612 (2024).
DelaFuente, J. et al. Within-patient evolution of plasmid-mediated antimicrobial resistance. Nat. Ecol. Evol. 6, 1980–1991 (2022).
Benz, F. et al. Plasmid- and strain-specific factors drive variation in ESBL-plasmid spread in vitro and in vivo. ISME J. 15, 862–878 (2021).
Neil, K., Allard, N., Grenier, F., Burrus, V. & Rodrigue, S. Highly efficient gene transfer in the mouse gut microbiota is enabled by the Incl(2) conjugative plasmid TP114. Commun. Biol. 3, 523 (2020).
Neil, K., Allard, N. & Rodrigue, S. Molecular mechanisms influencing bacterial conjugation in the intestinal microbiota. Front. Microbiol. 12, 673260 (2021).
Yong, M. et al. Dominant carbapenemase-encoding plasmids in clinical Enterobacterales isolates and hypervirulent Klebsiella pneumoniae, Singapore. Emerg. Infect. Dis. 28, 1578–1588 (2022).
Octavia, S. et al. Klebsiella pneumoniae and Klebsiella quasipneumoniae define the population structure of blaKPC-2Klebsiella: a 5 year retrospective genomic study in Singapore. J. Antimicrob. Chemother. 74, 3205–3210 (2019).
Chen, Y. et al. Acquisition of plasmid with carbapenem-resistance gene blaKPC2 in hypervirulent Klebsiella pneumoniae, Singapore. Emerg. Infect. Dis. 26, 549–559 (2020).
Gul, E. et al. Differences in carbon metabolic capacity fuel co-existence and plasmid transfer between Salmonella strains in the mouse gut. Cell Host Microbe 31, 1140–1153 e1143 (2023).
Sonnenborn, U. Escherichia coli strain Nissle 1917-from bench to bedside and back: history of a special Escherichia coli strain with probiotic properties. FEMS Microbiol. Lett. 363, https://doi.org/10.1093/femsle/fnw212 (2016).
Netikul, T. et al. Characterization of an IncN2-type blaNDM-(1)-carrying plasmid in Escherichia coli ST131 and Klebsiella pneumoniae ST11 and ST15 isolates in Thailand. J. Antimicrob. Chemother. 69, 3161–3163 (2014).
Leavitt, A., Chmelnitsky, I., Ofek, I., Carmeli, Y. & Navon-Venezia, S. Plasmid pKpQIL encoding KPC-3 and TEM-1 confers carbapenem resistance in an extremely drug-resistant epidemic Klebsiella pneumoniae strain. J. Antimicrob. Chemother. 65, 243–248 (2010).
Tan, Y. H., Chen, Y., Chu, W. H. W., Sham, L. T. & Gan, Y. H. Cell envelope defects of different capsule-null mutants in K1 hypervirulent Klebsiella pneumoniae can affect bacterial pathogenesis. Mol. Microbiol. https://doi.org/10.1111/mmi.14447 (2020).
Haudiquet, M. et al. Capsules and their traits shape phage susceptibility and plasmid conjugation efficiency. Nat. Commun. 15, 2032 (2024).
Bradley, D. E. Morphological and serological relationships of conjugative pili. Plasmid 4, 155–169 (1980).
Lam, M. M. C. et al. Tracking key virulence loci encoding aerobactin and salmochelin siderophore synthesis in Klebsiella pneumoniae. Genome Med 10, 77 (2018).
Blum-Oehler, G. et al. Development of strain-specific PCR reactions for the detection of the probiotic Escherichia coli strain Nissle 1917 in fecal samples. Res. Microbiol. 154, 59–66 (2003).
Lim, C. et al. Essentiality of the virulence plasmid-encoded factors in disease pathogenesis of the major lineage of hypervirulent Klebsiella pneumoniae varies in different infection niches. EBioMedicine 115, 105683 (2025).
Low, W. W. et al. Mating pair stabilization mediates bacterial conjugation species specificity. Nat. Microbiol. 7, 1016–1027 (2022).
Allard, N., Neil, K., Grenier, F. & Rodrigue, S. The type IV pilus of plasmid TP114 displays adhesins conferring conjugation specificity and is important for DNA transfer in the mouse gut microbiota. Microbiol. Spectr. 10, e0230321 (2022).
Bakkeren, E. et al. Salmonella persisters promote the spread of antibiotic resistance plasmids in the gut. Nature 573, 276–280 (2019).
Hayakawa, M. et al. Hitherto-unnoticed self-transmissible plasmids widely distributed among different environments in Japan. Appl. Environ. Microbiol. 88, e0111422 (2022).
Datta, N., Hedges, R. W., Shaw, E. J., Sykes, R. B. & Richmond, M. H. Properties of an R factor from Pseudomonas aeruginosa. J. Bacteriol. 108, 1244–1249 (1971).
Jobanputra, R. S. & Datta, N. Trimethoprim R factors in enterobacteria from clinical specimens. J. Med Microbiol 7, 169–177 (1974).
Bahl, M. I., Burmolle, M., Meisner, A., Hansen, L. H. & Sorensen, S. J. All IncP-1 plasmid subgroups, including the novel epsilon subgroup, are prevalent in the influent of a Danish wastewater treatment plant. Plasmid 62, 134–139 (2009).
Shintani, M. et al. Plant species-dependent increased abundance and diversity of IncP-1 plasmids in the rhizosphere: new insights into their role and ecology. Front. Microbiol. 11, 590776 (2020).
Heuer, H. et al. IncP-1epsilon plasmids are important vectors of antibiotic resistance genes in agricultural systems: diversification driven by class 1 integron gene cassettes. Front Microbiol 3, 2 (2012).
Hunt, M., Lima, L., Shen, W., Lees, J. & Iqbal, Z. AllTheBacteria—all bacterial genomes assembled, available and searchable. BioRXiV, https://doi.org/10.1101/2024.03.08.584059 (2024).
Klumper, U. et al. Broad host range plasmids can invade an unexpectedly diverse fraction of a soil bacterial community. ISME J. 9, 934–945 (2015).
Camargo, A. P. et al. IMG/PR: a database of plasmids from genomes and metagenomes with rich annotations and metadata. Nucleic Acids Res 52, D164–D173 (2024).
Duong, V. T. et al. Genomic serotyping, clinical manifestations, and antimicrobial resistance of nontyphoidal Salmonella gastroenteritis in hospitalized children in Ho Chi Minh City, Vietnam. J. Clin. Microbiol. 58, https://doi.org/10.1128/JCM.01465-20 (2020).
Redondo-Salvo, S. et al. COPLA, a taxonomic classifier of plasmids. BMC Bioinform. 22, 390 (2021).
Redondo-Salvo, S. et al. Pathways for horizontal gene transfer in bacteria revealed by a global map of their plasmids. Nat. Commun. 11, 3602 (2020).
Ankenbrand, M. J., Hohlfeld, S., Hackl, T. & Förster, F. AliTV—interactive visualization of whole genome comparisons. PeerJ Comput. Sci. 3, https://doi.org/10.7717/peerj-cs.116 (2017).
Krol, J. E. et al. Increased transfer of a multidrug resistance plasmid in Escherichia coli biofilms at the air-liquid interface. Appl. Environ. Microbiol. 77, 5079–5088 (2011).
Ochi, K. et al. Oxygen concentration affects frequency and range of transconjugants for the incompatibility (Inc) P-1 and P-7 plasmids pBP136 and pCAR1. Biosci. Biotechnol. Biochem. 85, 1005–1015 (2021).
Walker, K. A., Treat, L. P., Sepulveda, V. E. & Miller, V. L. The small protein RmpD drives hypermucoviscosity in Klebsiella pneumoniae. mBio 11, https://doi.org/10.1128/mBio.01750-20 (2020).
Carattoli, A. Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 2227–2238 (2009).
Rozwandowicz, M. et al. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 73, 1121–1137 (2018).
Djermoun, S. et al. Biofilm architecture determines the dissemination of conjugative plasmids. Proc. Natl. Acad. Sci. USA 122, e2417452122 (2025).
Haudiquet, M., Buffet, A., Rendueles, O. & Rocha, E. P. C. Interplay between the cell envelope and mobile genetic elements shapes gene flow in populations of the nosocomial pathogen Klebsiella pneumoniae. PLoS Biol. 19, e3001276 (2021).
Galvez-Silva, M. et al. Carbapenem-resistant hypervirulent ST23 Klebsiella pneumoniae with a highly transmissible dual-carbapenemase plasmid in Chile. Biol. Res 57, 7 (2024).
Wahl, A. et al. Presence of hypervirulence-associated determinants in Klebsiella pneumoniae from hospitalised patients in Germany. Int. J. Med. Microbiol. 314, 151601 (2024).
Du, P., Liu, C., Fan, S., Baker, S. & Guo, J. The role of plasmid and resistance gene acquisition in the emergence of ST23 multi-drug resistant, hypervirulent Klebsiella pneumoniae. Microbiol Spectr. 10, e0192921 (2022).
Sun, W. et al. Microaerobic-mediated suppression of Klebsiella pneumoniae mucoviscosity is restored by rmpD overexpression. J. Appl. Microbiol. 135, https://doi.org/10.1093/jambio/lxae192 (2024).
Lin, T. H. et al. FNR-dependent RmpA and RmpA2 regulation of capsule polysaccharide biosynthesis in Klebsiella pneumoniae. Front. Microbiol. 10, 2436 (2019).
Zierke, L., Mourad, R., Kohler, T. P., Musken, M. & Hammerschmidt, S. Influence of the polysaccharide capsule on virulence and fitness of Klebsiella pneumoniae. Front. Microbiol. 16, 1450984 (2025).
Wyres, K. L. et al. Distinct evolutionary dynamics of horizontal gene transfer in drug resistant and virulent clones of Klebsiella pneumoniae. PLoS Genet. 15, e1008114 (2019).
Xu, Y. et al. Mobilization of the nonconjugative virulence plasmid from hypervirulent Klebsiella pneumoniae. Genome Med. 13, 119 (2021).
Stecher, B. et al. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc. Natl. Acad. Sci. USA 109, 1269–1274 (2012).
Zeng, M. Y., Inohara, N. & Nunez, G. Mechanisms of inflammation-driven bacterial dysbiosis in the gut. Mucosal Immunol. 10, 18–26 (2017).
Groussin, M. et al. Elevated rates of horizontal gene transfer in the industrialized human microbiome. Cell 184, 2053–2067 e2018 (2021).
Kang, J. T. L. et al. Long-term ecological and evolutionary dynamics in the gut microbiomes of carbapenemase-producing Enterobacteriaceae colonized subjects. Nat. Microbiol. 7, 1516–1524 (2022).
De Gelder, L., Ponciano, J. M., Joyce, P. & Top, E. M. Stability of a promiscuous plasmid in different hosts: no guarantee for a long-term relationship. Microbiology 153, 452–463 (2007).
Bortolaia, V. et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 75, 3491–3500 (2020).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag New York, 2016).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Jain, C., Rodriguez, R. L., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008, https://doi.org/10.1088/1742-5468/2008/10/p10008 (2008).
Bürkner, P.-C. brms: an R package for Bayesian multilevel models using Stan. J. Stat. Softw. 80, https://doi.org/10.18637/jss.v080.i01 (2017).
The R Project for Statistical Computing., https://www.R-project.org/.
Gelman, A. et al. Bayesian Data Analysis (3rd edition). https://doi.org/10.1201/b16018 (Chapman and Hall/CRC, 2013).
Yong, M. G. & Y. H. Supplementary Data for Differential Plasmid Transmission between IncP Plasmid Clades in the Mammalian Gut Involving Hypervirulent Klebsiella Pneumoniae Demonstrates Plasmid-specific and Bacterial Capsule Adaptations to Distinct Ecological Niches. https://zenodo.org/records/17060312 (2025).
Mishra, S. mlgh-sg/plasmids-amr-modelling: Pre-release, https://doi.org/10.5281/zenodo.17460639 (2025).
Acknowledgements
This study was supported by National Medical Research Council Grant OFIRG20NOV-0045 awarded to Y.H.G. and the Australian Research Council Laureate Fellowship FL210100258 awarded to D.L. G.O. is supported by NUS President’s Graduate Fellowship for his PhD study. We would like to extend our sincere gratitude to Professor Masaki Shintani for generously providing pMNCN064 and pYKBP039 and to Professor Sebastien Rodrigue for the gift of E. coli Nissle. We thank the Carbapenemase-producing Enterobacteriaceae in Singapore (CaPES) study group for their support.
Author information
Authors and Affiliations
Contributions
Y.H.G. conceptualized the study, co-designed the experiments, secured funding, and co-wrote the initial manuscript draft. M.Y. conducted most of the laboratory work, co-designed the experiments, and co-wrote the first manuscript draft. W.W.L. assisted with the mouse experiments and capsule quantification. G.W., S.M., C.L., and D.L. supported the design and execution of gastrointestinal tissue processing and imaging. G.O., G.C., C.C., Y.C., H.C.T., and T.D.P. contributed to the in vitro work and plasmid isolation. S.M. aided in statistical modeling. All authors participated in subsequent revisions and reviewed the final manuscript prior to submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Raul Fernandez-Lopez and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yong, M., Low, W.W., Mishra, S. et al. Differential gut transmission of IncP plasmid clades involving hypervirulent Klebsiella pneumoniae reveals plasmid-specific ecological adaptation. Nat Commun 16, 11353 (2025). https://doi.org/10.1038/s41467-025-66413-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66413-4