Abstract
Breast cancer remains the most prevalent malignancy in women worldwide. Mammography-based early detection plays a pivotal role in improving patient survival outcomes. While large vision-language models offer transformative potential for mammogram visual question answering, the absence of standardized evaluation benchmarks currently makes it hard to fairly compare different large vision-language models’ performance in mammogram interpretation. In this study, we address this critical gap through three key contributions: (1) We introduce MammoVQA, a mammogram visual question-answering dataset that unifies 15 public datasets, comprising 131,847 images (421K question-answering pairs) for image-level cases and 72,518 exams (476K images, 144K question-answering pairs) for exam-level cases. (2) Systematic evaluation of 12 recent high-performance large vision-language models (6 general, 6 medical) reveals diagnostic performance statistically equivalent to random guessing, highlighting their unreliability for mammogram interpretation. (3) Our domain-optimized LLaVA-Mammo achieves average +19.66% weighted accuracy gains over the best recent high-performance model in internal validation, with average +21.21% weighted accuracy improvements in external validation.
Similar content being viewed by others
Introduction
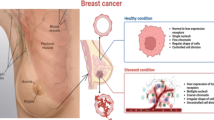
Breast cancer is one of the most significant health challenges globally, with millions of new cases diagnosed each year1,2,3. Early detection is crucial to improving patient outcomes4, and mammography serves as a cornerstone tool for the detection and diagnosis of breast cancer5,6. As a specialized radiographic technique, mammography provides critical visual evidence to detect early signs of cancer, such as masses or calcifications7. However, interpreting mammograms is inherently complex and highly dependent on the expertise of qualified radiologists. Even among experts, diagnoses can involve subjective judgments and inconsistencies. Therefore, developing high-quality mammogram datasets tailored to support model training and evaluation has become an essential requirement to advance intelligent applications in this domain.
In recent years, the rapid development of Large Vision-Language Models (LVLMs) has demonstrated substantial potential in multimodal learning. General-domain LVLMs, such as Flamingo8, BLIP-29, LLaVA10, and MiniGPT-411, have shown remarkable performance in tasks like image-text generation, visual question answering (VQA), and image captioning. These models leverage large-scale pretraining datasets and cross-modal alignment techniques to enhance generalization capabilities in both vision and language tasks. In the medical domain, specifically tailored medical LVLMs have further expanded the application scope of LVLMs. For example, RadFM12 optimizes PMC-LLaMA13 with 16 million radiology image-text pairs, while Med-Flamingo14 enhances OpenFlamingo-9B15 using billions of biomedical image-text pairs to meet the demands of medical image analysis and report generation. LLaVA-Med16 extends LLaVA10 by incorporating a biomedical figure caption dataset extracted from PubMed Central17, significantly improving biomedical image understanding and open-domain dialog capabilities. MedVInT18 strengthens medical imaging analysis and open-ended VQA through visual instruction fine-tuning on 227,000 vision-question-answer pairs from the PMC-VQA18 dataset. Furthermore, the latest GMAI-VL19, developed using the proposed GMAI-VL-5.5M dataset, advances research in multimodal medical representation. Despite these advancements, the application of LVLMs in specific medical fields, i.e., mammograms, remains limited.
To validate the VQA performance of existing LVLMs on mammograms, we investigate a variety of datasets that have supported advancements in mammographic image analysis. Early datasets like MIAS20 and INbreast21 laid the foundation, with MIAS providing 322 images with basic annotations and INbreast offering 410 high-resolution images with detailed lesion labels. Despite their contributions, these datasets are limited in size and diversity. CBIS-DDSM22, built on the previous DDSM dataset, introduced standardized mammograms with detailed annotations such as lesion segmentation and Breast Imaging-Reporting and Data System (BI-RADS23) descriptors, significantly improving the quality of mammogram data. More recent datasets, such as Vindr-Mammo24, CSAW-M25, KAU-BCMD26, and BMCD27, provide thousands of mammograms with comprehensive annotations, addressing the need for larger and more diverse data. Specialized datasets like CDD-CESM28, which focus on contrast-enhanced spectral mammography (CESM) with over 1000 annotated images, are designed for diagnostics in dense breast tissue. In contrast, DMID29 emphasizes multimodal breast imaging, combining digital mammography with other imaging techniques and detailed annotations to support research on integrating complementary imaging modalities for better detection of breast cancer. Large-scale datasets, such as EMBED30, which includes a total of 3.4 million images (with around 480,000 currently available), and the Radiological Society of North America (RSNA) Breast Cancer Detection Challenge dataset31, which contains around 54,700 images, offer substantial resources for AI development. These datasets with diverse scales and imaging modalities have laid a solid foundation for advancing mammogram analysis. However, their lack of design specificity for the VQA task limits their applicability in vision-language research.
In addition, we explore existing medical VQA datasets, such as VQA-RAD32 and SLAKE33, which have laid the foundation for this advancement. VQA-RAD focuses on radiological images but covers a limited range of anatomical regions, while SLAKE expands its coverage to include areas such as the pelvis and neck, though its data diversity remains constrained. Additionally, benchmark datasets like MIMIC-CXR34 and CheXpert35 in the chest radiographic imaging domain have supported numerous multimodal studies. PMC-VQA18 has further contributed to the field by incorporating various imaging modalities and tasks. OmniMedVQA36 and the recent GMAI-VL-5.5M19 dataset attempt to bridge this gap by covering 12 and 13 imaging modalities, respectively, along with multiple anatomical regions and medical professional tasks, making them the largest and most diverse medical VQA datasets to date. Despite these advancements, these datasets contain limited or no mammogram data, which is insufficient for evaluating or enabling model performance in interpreting mammograms.
To address this gap, we introduce MammoVQA, a VQA dataset specifically focused on mammograms. MammoVQA is a multimodal medical imaging dataset designed to meet the unique requirements of the mammogram, comprising 15 mammogram datasets and 565,092 question-answering pairs designed to cover clinically relevant tasks such as BI-RADS classification, breast density assessment, and abnormality detection.
Beyond simply providing a benchmark, the central contribution of this work lies in rigorously evaluating the capabilities of existing LVLMs in interpreting mammography images. While the general-domain models have shown outstanding performance in general VQA tasks, their performance on mammograms remains underexplored. Similarly, the medical-domain models, although tailored for medical applications, have not been extensively tested in mammography-specific contexts. This gap motivates our study: Do LVLMs truly possess sufficient interpretation ability when faced with clinically relevant mammography tasks?
To validate the efficacy of MammoVQA, we systematically evaluated the performance of 12 state-of-the-art LVLMs on this dataset, including 6 general-domain models and 6 medical-domain models. LVLMs face significant challenges when addressing mammography-related questions, with almost all models on various question topics showing performance comparable to random guessing. This indicates that they cannot interpret mammograms effectively. These issues likely derive from the models’ lack of sufficient mammogram data during the training phase. To investigate the impact of domain-specific adaptation on performance, we fine-tune LLaVA-NeXT37 on the MammoVQA training set, aiming to further enhance its performance in mammography-related tasks.
Furthermore, we train and evaluate vision-only models (ResNet-5038 and DINOv239) with linear probing and a multimodal framework ViLT40 that does not utilize large language models on MammoVQA. Unlike LVLMs, the outputs of vision-only models and ViLT-based models are closed-set, meaning their answer spaces are constrained to predefined options. Although vision-only models and ViLT-based models exhibit a certain level of competitiveness in MammoVQA, their overall performance remains significantly inferior to that of the domain-optimized LVLMs. The external validation particularly highlights LLaVA-Mammo’s robustness. These findings indicate that LVLMs have substantial potential for the VQA task on mammograms, as they can handle more open-ended and complex question-answering scenarios. However, they also highlight the limitations of existing LVLMs, particularly in tasks requiring highly domain-specific knowledge, where further optimization and adaptation are still demanded.
In this work, we make the following contributions:
-
We introduce a VQA benchmark designed specifically for mammogram interpretation, MammoVQA, which is created by integrating 15 public mammogram datasets. This comprehensive benchmark combines 131,847 images with 420,923 QA pairs for image-level cases and 72,518 examinations (475,971 images) with 144,169 QA pairs for exam-level cases. By providing both an evaluation framework for assessing LVLMs’ mammogram interpretation capabilities and structured data for domain adaptation, MammoVQA establishes a critical infrastructure for advancing research in AI-assisted mammogram interpretation.
-
We conduct a systematic evaluation of 6 general-domain and 6 medical-domain LVLMs on MammoVQA, revealing that their mammogram interpretation performance was statistically indistinguishable from random guessing. This striking inadequacy highlights the challenges of mammogram interpretation, where subtle anatomical patterns and complex contextual relationships across multiple views differ fundamentally from other domain images.
-
We propose a domain-optimized model, LLaVA-Mammo, which significantly outperforms both general and medical LVLMs in mammogram VQA tasks. Specifically, for internal validation, LLaVA-Mammo achieves performance gains of 38.29% average absolute accuracy (dataset level) and 19.66% average weighted accuracy (question topic level) over the best recent high-performance model in internal validation. External validation across 4 independent external datasets demonstrates that LLaVA-Mammo outperforms the best recent high-performance model by 19.87% average absolute accuracy (dataset level) and 21.21% average weighted accuracy (question topic level). This advancement demonstrates the critical importance of domain-specific design for medical AI and provides a foundation for future development of vision-language systems in mammogram interpretation.
Results
Construct MammoVQA from public mammogram datasets
Recognizing the scarcity of image-text data in mammograms, we aggregate a large number of classification datasets from this domain and convert them into the VQA format to form our MammoVQA dataset. MammoVQA comprises 15 mammogram datasets released by various authoritative medical institutions (shown in Fig. 1a), resulting in a diverse set of images that enable models to learn more generalized representations and validate their performance across a wide range of heterogeneous data. Importantly, all images are sourced from real medical settings, ensuring that MammoVQA is closely aligned with real-world applications. Through manual verification, we re-examined all mammograms to ensure the absence of image corruption, unreadable files, or visual abnormalities. Furthermore, we verified that all bounding box coordinates precisely delineated the target mass regions, and confirmed that all classification labels were accurately mapped to our predefined unified label space. The dataset covers 9 question topics (shown in Fig. 1b and detailed in Supplementary Table 8), including but not limited to BI-RADS classification, density assessment, and abnormality detection, fully reflecting the diversity and complexity of the mammogram. MammoVQA includes 131,847 images and 420,923 QA pairs for image-level cases, as well as 72,518 examinations (475,971 images) and 144,169 QA pairs for exam-level cases, establishing a large-scale dataset.
a Dataset composition statistics, which describe the number of images contained in each sub-dataset of MammoVQA (* indicates the number of examinations) and the distribution of corresponding question topics. b Hierarchical taxonomy of 9 clinically validated question topics organized by diagnostic workflow stages. c The sub-datasets of MammoVQA are categorized into three types according to the format of the provided labels. d An example of question-answer pair generation: for each question topic, four question formats are generated using GPT-4o, and one is randomly selected as the {Question} of the question-answer pair. The {Options} are then randomly ordered from the candidate options and filled into the template to form the final question-answer pair. e An example of a question and its corresponding answer for each of the 9 question topics in MammoVQA.
Systematic evaluation of existing LVLMs
To assess the capabilities of existing LVLMs in interpreting mammograms, we select 12 models pre-trained on large-scale datasets. These models, with similar scales and distinct characteristics, have gained widespread recognition for their performance. We conduct zero-shot experiments on the MammoVQA benchmark to evaluate their ability to interpret mammograms. The selected models include 6 general-domain models, namely MiniGPT-4-7B11, BLIP-2-11B9, InstructBLIP-7B41, LLaVA-NeXT-Interleave-7B42, InternVL3-8B43, and Qwen2.5-VL-7B44, along with 6 medical-domain models, including LLaVA-Med-7B16, RadFM-14B12, Med-Flamingo-7B14, MedVInT-TD-7B18, MedDr-40B45, and MedGemma-4B46.
By selecting these diverse models, we aim to comprehensively evaluate their performance in mammogram interpretation. Notably, RadFM and MedDr’s pre-training datasets included a small number of mammograms.
To establish performance benchmarks, we calculate the accuracy of a random guess based on the number of answer categories for each question topic. To thoroughly evaluate model performance and avoid assessment bias due to single-category predictions, we employ both absolute accuracy and weighted accuracy metrics. We also evaluate the macro-F1 score, which exhibits an overall trend consistent with that of weighted accuracy. To maintain conciseness in the main text, the full F1 results are provided exclusively in the supplementary materials. Through detailed analysis of weighted accuracy, we observe a notable phenomenon: the majority of LVLMs perform close to random guess levels across various question topics.
InternVL3 and MedGemma show significant advantages in some tasks. Specifically, in the pathology (breast) task, these 2 models outperform random guessing by 5.82% and 9.80%, respectively. In the pathology (finding) task, the advantages further expand to 10.42% and 13.50%. Moreover, in the abnormality (breast) task, they exceed random guessing by 2.57% and 3.55%, respectively. In the abnormality (finding) task, they also outperform other models (except BLIP-2) by approximately 3–4%.
The experimental results show that most LVLMs perform at near-random levels in mammogram interpretation. Even better-performing models (e.g., InternVL3 and MedGemma) still exhibit insufficient accuracy in most tasks, indicating a lack of domain-specific interpretation of breast lesion patterns. This limitation highlights the importance of improving model performance in mammogram interpretation, which is crucial to achieving reliable AI-assisted early detection of breast cancer.
How does MammoVQA boost the LVLMs’ interpretation ability on mammograms?
To further enhance the model performance in MammoVQA, we fine-tune the LLaVA-NeXT model in the MammoVQA training set, resulting in LLaVA-Mammo (Fig. 2a). Through fine-tuning on the MammoVQA dataset, the model can better learn the associations between the features of mammograms and category words, while adapting to the unique semantics and question types in the MammoVQA task. Moreover, the fine-tuned LLaVA-Mammo can also provide a powerful baseline model for subsequent research, promoting the further development of the mammogram visual question-answering task.
a Architecture of LLaVA-Mammo. b Architecture of ViLT-Mammo. c Architecture of ViLT-Mammo-Expert. d Architecture of R50-Mammo and DINOv2-Mammo. e A VQA sample on MammoVQA. The robot icon and doctor icon are from Flaticon.com, created by Freepik. f The distribution of answer predictions by LLaVA-Mammo on MammoVQA's internal benchmark set. #Source data are provided with this paper.
LLaVA-Mammo demonstrates absolute superiority over existing LVLMs on the MammoVQA internal benchmark set (the output example can be viewed in Fig. 2e, and the overall performance can be viewed in Fig. 3a, b). Specifically, in the background tissue task, the weighted accuracy of LLaVA-Mammo reaches 54.46%, which is 13.01% higher than the best-performing InternVL3 among other LVLMs (21.13% higher than a random guess). In the view task, the weighted accuracies of LLaVA-Mammo reach 98.45%, which is 44.43% (48.45% higher than a random guess) higher than the best-performing MedDr among other LVLMs. In the abnormality (finding) task, the weighted accuracy of LLaVA-Mammo reaches 13.61%, which is 4.89% higher than the best-performing model. In the subtlety, pathology (breast), pathology (finding), masking potential, BI-RADS (breast), Density (breast), laterality, and abnormality (breast) tasks, LLaVA-Mammo achieves weighted accuracies of 37.23%, 46.34%, 70.60%, 27.02%, 32.31%, 66.85%, 99.97%, and 6.72%, respectively. Except for the abnormality (breast) task where it is relatively lower, in other tasks, it is 12.93% (20.56% higher than random guess), 3.21% (13.01% higher than random guess), 23.77% (37.27% higher than random guess), 9.46% (14.52% higher than random guess), 13.65% (15.64% higher than random guess), 35.04% (41.85% higher than random guess), and 45.95% (49.97% higher than random guess) higher, respectively. Overall, in the image case, the average absolute accuracy and weighted accuracy of LLaVA-Mammo reach 73.89% and 50.32%, which is 38.07% and 19.66% (25.66% higher than a random guess) higher than the best-performing MedGemma among LVLMs, showing a significant improvement.
a Weighted accuracy across question topics. b Absolute accuracy across sub-datasets. c Category distribution of question topics in MammoVQA. d–f Comparison of two top existing and three domain-optimized models on MammoVQA benchmarks, showing weighted accuracy and absolute accuracy with 95% confidence intervals (calculated via 9999 bootstrap iterations), where the sample size n represents the total number of test cases. Error bars represent the range (mean ± CI). Dashed (---) and solid (-) lines indicate mean absolute and weighted accuracies, excluding exam-level question topics and EMBED sub-dataset. #Source data are provided with this paper. Detailed performance metrics are available in Supplementary Tables 1-7.
For the exam case, only LLaVA-Mammo, LLaVA-NeXT-interleave, InternVL3, Qwen2.5-VL, RadFM, Med-Flamingo, and MedGemma can process the multi-image input. In the BI-RADS (exam) task, LLaVA-Mammo achieves a weighted accuracy that is 1.07% lower than that of Qwen2.5-VL and is 0.85% higher than a random guess. In the density (exam) task, the weighted accuracy is 65.91%, which is 33.70% and 40.91% higher than the best-performing MedGemma among LVLMs and random guess, respectively.
From the perspective of sub-datasets, LLaVA-Mammo achieves an absolute accuracy of 74.33% and 68.49% on average in the breast dataset and the exam dataset, which is 38.29% and 44.48% higher than the best-performing MedGemma among LVLMs.
It is widely recognized that specialized models adopting closed-set outputs outperform large language models (LLMs) with open-ended outputs in classification tasks47. To verify whether this holds for our MammoVQA dataset constructed from classification datasets, we train 2 vision-only models and 2 ViLT40-based models with DINOv239 as the vision backbone on the MammoVQA training set.
For vision-only models, we perform linear probing on ResNet-5038 and DINOv239 using a multi-classification-head-per-model setup (Fig. 2d). For architectures based on ViLT, we first train ViLT-Mammo (Fig. 2b) with the same multi-head approach to handle multiple question types concurrently. Subsequently, we train ViLT-Mammo-Expert (Fig. 2c) using a single-classification-head-per-model strategy, where a dedicated classification head is optimized for each question type to improve specialization. Overall, DINOv2 achieves the best performance, followed by the two ViLT-based models with comparable results, while ResNet-50 performs the worst.
Under the same image-text input setting, a comparison between ViLT-based models and LLaVA-Mammo reveals that, aside from the simplest view and laterality tasks where performance is comparable, LLaVA-Mammo outperforms in other tasks by approximately 6–20% in terms of weighted accuracy. In summary, the weighted accuracy of LLaVA-Mammo on question topics is on average 6.72% and 7.82% higher than that of ViLT-Mammo and ViLT-Mammo-Expert, respectively, and its absolute accuracy is on average 7.09% and 7.01% higher. From the perspective of sub-datasets, the absolute accuracy is on average 9.72% and 7.13% higher.
The results show that ResNet-50 performed poorly, with its predictions consistently biased toward a single class upon statistical analysis. This suggests that the features extracted by ResNet-50 may not be directly suitable for mammography classification tasks. In contrast, DINOv2 achieved performance second only to LLaVA-Mammo. Since MammoVQA is essentially a pure visual classification task, it is intuitive that DINOv2 outperforms ViLT-based models, which require joint representation learning for both image and text, as the introduction of a textual modality may introduce irrelevant noise, and the language model could interfere with visual features. The superior performance of LLaVA-Mammo can be attributed to its powerful generalization capability of the large language model.
External validation
To evaluate the generalizability and reliability of LLaVA-Mammo, we conduct external validation using four independent datasets (DBT48, LAMIS49, MM50, and NLBS51). The external benchmark sets encompass six question topics: BI-RADS (breast), breast density, view, laterality, pathology (breast), and pathology (finding). External validation across 4 external datasets (shown in Fig. 3f) demonstrates that LLaVA-Mammo outperforms the best-performing model by 19.87% in absolute accuracy at the dataset level, and by 26.39% in absolute accuracy and 21.21% in weighted accuracy at the question-topic level. These findings demonstrate that LLaVA-Mammo maintains strong generalization capabilities, consistently outperforming existing models across all tasks and showing superior performance compared to domain-specialized models, thus indicating robust cross-dataset adaptability and reliability for diverse mammogram interpretation tasks.
Discussion
Importance of MammoVQA
MammoVQA is a large-scale mammogram VQA dataset, comprising 131,847 images with 420,923 QA pairs for image-level cases and 72,518 examinations (475,971 images) with 144,169 QA pairs for exam-level cases. MammoVQA addresses pivotal challenges in medical multimodal AI by: (1) establishing a dedicated evaluation framework for assessing LVLMs’ diagnostic capabilities in mammogram interpretation, (2) providing curated training data to adapt general-purpose LVLMs to the specialized domain of breast imaging through structured QA pairs, (3) emulating clinical reading paradigms by including both single-view cases and multi-view cases that reflect radiologists’ reliance on composite information for accurate diagnosis, and (4) creating the foundational infrastructure for developing human-AI collaborative systems where models can assist in preliminary screening while maintaining physician oversight for final decisions.
Importantly, all questions in MammoVQA are closed-ended and formulated as fundamental classification tasks. We adopt this simple but critical format, as poor performance on these basic tasks would indicate even greater challenges for more complex, open-ended VQA scenarios. The systematic evaluation of LVLMs on such fundamental tasks is not just a technical exercise, but has tangible clinical implications: As LVLMs demonstrate broad potential in medical image analysis, their ability to reliably interpret medical images directly impacts the safety, generalizability, and trustworthiness of future clinical decision support systems. LVLMs not only have the potential to enable more natural human-computer interactions, such as generating imaging descriptions and preliminary assessments through conversational interfaces, but could also significantly improve the efficiency and interpretability of medical report generation. For these reasons, before deploying such models in real-world applications, it is essential to rigorously and systematically evaluate their foundational capabilities, particularly in high-stakes domains such as healthcare. Performance in basic classification tasks serves as a critical indicator of visual-language alignment and generalization ability in medical concepts.
Performance analysis
Through detailed analysis of the answer distributions of the MedDr and LLaVA-NeXT-interleave, we find that their prediction patterns are unique. When dealing with pathology questions, when the true answer is ‘normal’, the models tend to make correct predictions, while for other categories, the predictions appear random. In the abnormality task, they also show high accuracy for ‘normal’ samples, but for abnormal samples, they tend to predict the most common ‘mass’ category. We believe that the excellent performance of the MedDr model may be attributed to the inclusion of mammogram data in its training set, while the outstanding performance of the LLaVA-NeXT-interleave model lacks a clear explanation.
In the BI-RADS (exam) task, LLaVA-Mammo’s weighted accuracy is 1.07% lower than that of Qwen2.5-VL. This indicates that LLaVA-Mammo does not gain the ability to diagnose the BI-RADS score of the corresponding patient from multiple images through fine-tuning. Based on the good results in the density (exam) task, we can confirm that the model can extract key information from multiple images. However, since judging the BI-RADS score requires very detailed information, and different images in the same exam may vary greatly in key detailed information, we believe that one of the problems is that LLaVA-Mammo cannot determine which image-provided information to use. Secondly, we find that LLaVA-Mammo performs very well in tasks such as view and laterality that only require macro-information for prediction, but performs poorly in problems such as identifying abnormalities in breast images that require detailed information. This shows that LLaVA-Mammo has a poor ability to capture detailed information, which we also consider as one of the problems.
ViLT-Mammo-Expert performs at the level of random guess in the background tissue and masking potential tasks. We attribute this to the imbalanced distribution of the training data. In contrast, ViLT-Mammo achieves good weighted accuracy in these 2 tasks, outperforming a random guess by 19.37% and 9.2%, respectively. We believe that this is because the background tissue, masking potential, and density (breast) tasks share commonalities as they are all related to the density of breast glands. The multi-classification-head design of ViLT-Mammo allows for knowledge sharing, which mitigates the impact of data imbalance.
Reliability analysis of LLaVA-Mammo
We select 2 top-performing models (InternVL3 and MedGemma) from 12 chosen LVLMs and compare their performance on question topics with 3 image-text domain-optimized models. In Fig. 3c, the outer bars represent the results of absolute accuracy, while the inner bars represent the results of weighted accuracy. The proximity of the two bars reflects the reliability of the model’s predictions: the closer the bars, the more stable the model’s predictions. From the figure, it can be observed that for the view and laterality tasks, the two bars for all 5 models are very close, indicating that these tasks are relatively simple and the models can handle them well. However, for the abnormality (breast) and abnormality (finding) tasks, the gaps between the two bars for all 5 models are significant, reflecting the complexity of abnormality tasks, particularly in identifying abnormality in mammograms, where the models’ reliability still has considerable room for improvement.
Further analysis of Fig. 3d reveals that the balance of the distribution of training data has a significant impact on the performance of the trained model. The figure shows that the more balanced the training data distribution, the smaller the gap between the two bars for the 3 trained models, indicating that the balanced training data improves the reliability of model prediction. Furthermore, Fig. 3e shows that LLaVA-Mammo consistently outperforms ViLT-based models on the finding dataset, and the overall performance of the finding dataset is significantly higher than that of the breast dataset. This result reflects LLaVA-Mammo’s limitations in extracting detailed information from breast images.
From Fig. 2f, we can observe that, apart from the view and laterality tasks, the labels of other tasks exhibit a gradual progression from mild to severe. Although the prediction distributions for background tissue and BI-RADS (exam) are less ideal due to uneven training data distribution and relatively high task difficulty, the prediction distributions for other tasks show clear trend characteristics. This indicates that LLaVA-Mammo indeed possesses the ability to distinguish different features in mammograms, especially when handling tasks with varying degrees of severity, where the model can capture hierarchical information in the data.
Limitations and future works
Our study has four primary limitations. First, computational constraints limit our experiments to Vicuna-7B, preventing verification of the scaling law hypothesis with larger LVLMs. Second, performance bias emerges from the imbalance of training data in the question topics. Third, MammoVQA’s classification-based design limits answers to closed categories, restricting LVLMs’ open-ended reasoning potential. Fourth, no systematic investigation was conducted to identify optimal model architectures or training strategies that would maximize MammoVQA’s effectiveness.
These limitations motivate four research priorities: (1) scaling to larger architectures, (2) developing robust data balancing methods, (3) constructing open-ended mammogram VQA datasets to properly assess LVLMs’ mammography interpretation abilities, and (4) using the MammoVQA dataset to develop lightweight LVLMs via knowledge distillation techniques, optimizing for real-time clinical mammogram QA applications.
It is crucial to emphasize that MammoVQA is fundamentally a technical benchmark designed to evaluate AI model performance, not a study comparing diagnostic accuracy against experienced radiologists. Therefore, any reported ‘superior accuracy’ should be strictly interpreted as superior technical performance on this specific benchmark task and must not be misconstrued as evidence of superior clinical diagnostic utility. Superior performance on the MammoVQA benchmark is a necessary initial step. Nevertheless, it is critically important to recognize that this represents technical proficiency rather than proven clinical utility. The definitive evidence for any model’s diagnostic value must ultimately be established through prospective, patient-centered clinical studies.
Methods
This project has been reviewed and approved by the Human and Artefacts Research Ethics Committee (HAREC). The protocol number is HREP-2025-0025.
MammoVQA construction
As shown in Fig. 1c, we use classification labels of each dataset to categorize all datasets based on the label types. Specifically, datasets where each label corresponds to an individual image are categorized as breast datasets. In contrast, datasets where the labels correspond to an entire examination are classified as exam datasets, provided that each examination contains no more than 15 images. VinDr-Mammo and RSNA datasets include labels for both examinations and individual images. To ensure data balance, these datasets are treated as breast datasets. For the breast datasets, if bounding boxes and corresponding labels for the findings are provided, the finding regions are cropped and used to construct the finding datasets. Additionally, since the finding dataset only includes confirmed abnormal cases and lacks normal cases, random crops of the same size as the findings are taken from the original images to create normal cases. This structured approach ensures that the MammoVQA is well organized and covers a wide range of mammography-related tasks, facilitating effective model evaluation. All datasets are presented in Tables 1 and 2.
MammoVQA splits
In constructing MammoVQA, the internal data is divided into training, validation, and internal benchmark sets in a 7:1:2 ratio. This division specifically applies to the 11 internal datasets, ensuring that each sub-dataset is proportionally represented across all three sets. Additionally, MammoVQA includes 4 external datasets (as shown in Fig. 1a) that are reserved exclusively for external validation, providing a comprehensive evaluation framework that assesses model generalization across diverse data distributions. Furthermore, this division guarantees comprehensive coverage of all question topics, such as BI-RADS classification and density assessments, within each internal split. This is particularly important because not every mammogram contains labels for all question topics. By ensuring a balanced distribution of question topics across the splits, the design maximizes the representation of all labels, enabling the model to learn and be evaluated on multiple tasks effectively. This approach minimizes the risk of certain sub-datasets or question topics disproportionately influencing model performance and allows for a thorough evaluation of the model’s generalization capabilities across varying data distributions and tasks. Maintaining a proportional representation of sub-datasets and balanced coverage of all question topics also helps mitigate potential data biases, providing a more realistic reflection of the model’s ability to handle the diversity and complexity inherent in mammogram analysis.
Question-Answer pair generation
To construct the question-answer (QA) pairs based on the identified question topics, we leverage the category information of each question topic to guide the process, as shown in Fig. 1d. For each question topic, we identify whether it originates from a breast dataset, a finding dataset, or an exam dataset, and accordingly use GPT-4o52 to generate four corresponding question templates. For question topics from the breast dataset and finding dataset, the templates use terms such as ‘image’ or ‘mammogram’, whereas, for exam datasets, these terms are replaced with ‘exam’ to maintain consistency with the dataset context. To better evaluate the performance of LVLMs, we construct our QA pairs in a multiple-choice format. Specifically, for each dataset entry in MammoVQA, we construct prompts based on whether the question topic corresponds to a single-choice or multiple-choice question. For single-choice questions, the prompt is designed to ensure concise and evaluable answers, adopting the following structure: ‘This is a mammography-related medical question with several options, only one of which is correct. Select the correct answer and respond with just the chosen option, without any further explanation. ### Question: {Question} ### Options: {Options}. ### Answer:’. For multiple-choice questions, a similar structure is used with slight modifications to the phrasing of the instructions. Here, {Question} represents the question generated by GPT-4o, and {Options} is a randomized list of all possible options for the corresponding question topic. For example, for the question topic ‘Laterality’, {Options} could take the form of ‘A: Left, B: Right’ or ‘A: Right, B: Left’, depending on the random shuffle. Randomizing the order of the options is crucial to avoid biases where LVLMs might consistently predict the same choice (e.g., ‘A’) due to a tendency toward fixed option orders, thus ensuring more realistic performance evaluation metrics. By carefully designing the prompts and randomizing the orders of the options, our objective is to minimize biases and maximize the evaluability of the responses of the LVLMs, thus improving the reliability of our benchmark results. The examples of QA pairs of each question topic can be viewed in Fig. 1e.
Evaluation metrics
We employ three evaluation metrics in our study:
where N is the total number of samples, scorei ∈ {0, 1} indicates prediction correctness, and \(| {{{{\mathcal{C}}}}}_{i}|\) is the number of samples in the i-th sample’s category. Additionally, we report Macro F1-score results in the supplementary tables for comprehensive performance evaluation.
Evaluation method
Since the outputs of LVLMs are in the form of open-ended text, to obtain reliable and accurate model performance, we adopt a two-step evaluation approach for single-choice questions. First, we use difflib.SequenceMatcher to calculate the similarity (the ratio of the longest common subsequence to the total length of both texts) between the predicted text and each option, and then sort the options in descending order of similarity. If there is only one option with the highest similarity, we select this option as the final output. If there are multiple options with the highest similarity, we use the fuzzywuzzy library to calculate the similarity (Levenshtein Distance) again. If the options with the highest similarity are still not unique, the output will be ‘make no choice’ (judged as incorrect). For multiple-choice questions, we use the keyword-matching method. A prediction is considered correct if and only if all the correct answers appear in the model’s output.
Model training detail and hyperparameter setting
To obtain the LLaVA-Mammo model, we adopt the Low-Rank Adaptation (LoRa)53 to fine-tune the LLM component of LLaVA-NeXT while freezing the parameters of the other parts of the model. Specifically, we set the LoRa hyperparameters as follows: both lora_alpha and lora_r were set to 8, and lora_dropout was set to 0.05. During training, the total number of epochs was set to 1, the batch size was 16, the learning rate was 2 × 10−5, and the maximum text length was 32768. The entire fine-tuning process took approximately 10 days and used four NVIDIA L20 (48GB). For the two ViLT-based models and two vision-only models, we froze the parameters of the vision backbone for training. Tables 1 and 2 The number of training epochs and the batch size were the same as those for LLaVA-Mammo, which were 1 and 16, respectively, and the learning rate was set to 0.001. All training processes were implemented using Python 3.9, PyTorch 2.5.1, and CUDA 12.2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sub-datasets of MammoVQA are publicly available, and you can access them through the links in Table 1 and Table 2. The complete MammoVQA can be accessed at https://github.com/PiggyJerry/MammoVQA. Source data are provided with this paper.
Code availability
The trained model and source code can be accessed at https://github.com/PiggyJerry/MammoVQAwith the https://doi.org/10.5281/zenodo.1738474054.
References
Broeders, M. et al. The impact of mammographic screening on breast cancer mortality in europe: a review of observational studies. J. Med. Screen. 19, 14–25 (2012).
Morra, L. et al. Breast cancer: computer-aided detection with digital breast tomosynthesis. Radiology 277, 56–63 (2015).
Chon, J.-W. et al. Effect of silk fibroin hydrolysate on the apoptosis of mcf-7 human breast cancer cells. Int. J. Ind. Entomol. 27, 228–236 (2013).
Ginsburg, O. et al. Breast cancer early detection: a phased approach to implementation. Cancer 126, 2379–2393 (2020).
Jalalian, A. et al. Computer-aided detection/diagnosis of breast cancer in mammography and ultrasound: a review. Clin. imaging 37, 420–426 (2013).
Contrast-Enhanced Mammography. Diagnosis and staging of breast cancer: When and how to use mammography. Diseases of the Chest, Breast, Heart and Vessels 2019-2022: Diagnostic and Interventional Imaging, page 155 (2019).
Alghaib, H. A., Scott, M. & Adhami, R. R. An overview of mammogram analysis. IEEE Potentials 35, 21–28 (2016).
Alayrac, J.-B. et al. Flamingo: a visual language model for few-shot learning. Adv. Neural Inf. Process. Syst. 35, 23716–23736 (2022).
Li, J., Li, D., Savarese, S. & Hoi, S. Blip-2: Bootstrapping language-image pre-training with frozen image encoders and large language models. In International conference on machine learning, pages 19730–19742. PMLR, (2023).
Liu, H., Li, C., Wu, Q. & Lee, Y. J. Visual instruction tuning. Advances in Neural Information Processing Systems 36 (2024).
Zhu, D., Chen, J., Shen, X., Li, X. & Elhoseiny, M. Minigpt-4: Enhancing vision-language understanding with advanced large language models. In The Twelfth International Conference on Learning Representations, https://openreview.net/forum?id=1tZbq88f27 (2024).
Wu, C., Zhang, C., Zhang, Y., Wang, Y. & Xie, W. Towards generalist foundation model for radiology by leveraging web-scale 2d\&3d medical data. Nat. Commun. 16, 7866 (2025).
Wu, C. et al. Pmc-llama: toward building open-source language models for medicine. J. Am. Med. Inform. Assoc. 31, 1833–1843 (2024).
Moor, M. et al. Med-flamingo: a multimodal medical few-shot learner. In Machine Learning for Health (ML4H), pages 353–367. PMLR, (2023).
Awadalla, A. et al. Openflamingo: An open-source framework for training large autoregressive vision-language models. arXiv preprint arXiv:2308.01390, (2023).
Li, C. et al. Llava-med: Training a large language-and-vision assistant for biomedicine in one day. Advances in Neural Information Processing Systems, 36 (2024).
Roberts, R. J. Pubmed central: The genbank of the published literature, (2001).
Zhang, X. et al. Development of a large-scale medical visual question-answering dataset. Commun. Med. 4, 277 (2024).
Li, T. et al. Gmai-vl & gmai-vl-5.5 m: A large vision-language model and a comprehensive multimodal dataset towards general medical ai. arXiv preprint arXiv:2411.14522, (2024).
Suckling, J. The mammographic images analysis society digital mammogram database. In Exerpta Medica. International Congress Series, 1994, volume 1069, pages 375–378 (1994).
Moreira, I. C. et al. Inbreast: toward a full-field digital mammographic database. Academic Radiol. 19, 236–248 (2012).
Lee, R. S. et al. A curated mammography data set for use in computer-aided detection and diagnosis research. Sci. data 4, 1–9 (2017).
Liberman, L. & Menell, J. H. Breast imaging reporting and data system (bi-rads). Radiologic Clin. 40, 409–430 (2002).
Nguyen, H. T. et al. Vindr-mammo: A large-scale benchmark dataset for computer-aided diagnosis in full-field digital mammography. Sci. Data 10, 277 (2023).
Sorkhei, M. et al. Csaw-m: An ordinal classification dataset for benchmarking mammographic masking of cancer. Adv. Neural. Inf. Process. Syst. (2021).
Alsolami, A. S. et al. King abdulaziz university breast cancer mammogram dataset (kau-bcmd). Data 6, 111 (2021).
Loizidou, K., Skouroumouni, G., Pitris, C. & Nikolaou, C. Digital subtraction of temporally sequential mammograms for improved detection and classification of microcalcifications. Eur. Radiol. Exp. 5, 1–12 (2021).
Khaled, R. et al. Categorized contrast enhanced mammography dataset for diagnostic and artificial intelligence research. Sci. data 9, 122 (2022).
Oza, P. et al. Digital mammography dataset for breast cancer diagnosis research (dmid) with breast mass segmentation analysis. Biomed. Eng. Lett. 14, 317–330 (2024).
Jeong, J. J. et al. The emory breast imaging dataset (embed): A racially diverse, granular dataset of 3.4 million screening and diagnostic mammographic images. Radiology: Artif. Intell. 5, e220047 (2023).
RSNA: Radiological Society of North America. Rsna screening mammography breast cancer detection ai challenge. https://www.rsna.org/rsnai/ai-image-challenge/screening-mammography-breast-cancer-detection-ai-challenge, (2023).
Lau, J. J., Gayen, S., Ben Abacha, A. & Demner-Fushman, D. A dataset of clinically generated visual questions and answers about radiology images. Sci. data 5, 1–10 (2018).
Liu, B. et al. Slake: A semantically-labeled knowledge-enhanced dataset for medical visual question answering. In 2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI), pages 1650–1654 (IEEE, 2021).
Johnson, A. E. W. et al. Mimic-cxr, a de-identified publicly available database of chest radiographs with free-text reports. Sci. data 6, 317 (2019).
Irvin, J. et al. Chexpert: A large chest radiograph dataset with uncertainty labels and expert comparison. In Proceedings of the AAAI conference on artificial intelligence, volume 33, pages 590–597 (2019).
Hu, Y. et al. Omnimedvqa: A new large-scale comprehensive evaluation benchmark for medical lvlm. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pages 22170–22183 (2024).
Liu, H. et al. Llava-next: Improved reasoning, ocr, and world knowledge, (2024).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, pages 770–778 (2016).
Oquab, M. et al. Dinov2: Learning robust visual features without supervision. Trans. Mach. Learn. Res. https://openreview.net/forum?id=a68SUt6zFt (2024).
Kim, W., Son, B. & Kim, I. Vilt: Vision-and-language transformer without convolution or region supervision. In International conference on machine learning, pages 5583–5594. PMLR, (2021).
Dai, W. et al. Instructblip: Towards general-purpose vision-language models with instruction tuning, URL https://arxiv.org/abs/2305.06500 (2023).
Li, F. et al. Llava-interleave: Tackling multi-image, video, and 3d in large multimodal models. The Thirteenth Int. Conf. on Learn. Representations (2024).
Zhu, J. et al. Internvl3: Exploring advanced training and test-time recipes for open-source multimodal models. arXiv preprint arXiv:2504.10479, (2025).
Bai, S. et al. Qwen2. 5-vl technical report. arXiv preprint arXiv:2502.13923, (2025).
He, S. et al. Meddr: Diagnosis-guided bootstrapping for large-scale medical vision-language learning. arXiv e-prints, pages arXiv–2404, (2024).
Sellergren, A. et al. Medgemma technical report. arXiv preprint arXiv:2507.05201, (2025).
Brown, T. et al. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 33, 1877–1901 (2020).
Buda, M. et al. A data set and deep learning algorithm for the detection of masses and architectural distortions in digital breast tomosynthesis images. JAMA network open 4, e2119100 (2021).
Imane, O. et al. Lamis-dmdb: A new full field digital mammography database for breast cancer ai-cad researches. Biomed. Signal Process. Control 90, 105823 (2024).
Aqdar, K. B. et al. Mammogram mastery: a robust dataset for breast cancer detection and medical education. Data Brief. 55, 110633 (2024).
Kendall, E. et al. Full field digital mammography dataset from a population screening program. Scientific Data 12, 1479 (2025).
Achiam, J. et al. Gpt-4 technical report. arXiv preprint arXiv:2303.08774, (2023).
Hu, E. J. et al. Lora: Low-rank adaptation of large language models. ICLR 1, 3 (2022).
Zhu, J. Piggyjerry/mammovqa: Code for: A benchmark for breast cancer screening and diagnosis in mammogram visual question answering, October https://doi.org/10.5281/zenodo.17384740 (2025).
Acknowledgements
This work was supported by Hong Kong Innovation and Technology Commission (Project No. MHP/002/22) and National Key R&D Program of China (Project No. 2023YFE0204000).
Author information
Authors and Affiliations
Contributions
J.Z. and F.H. collected the datasets and prepared the initial manuscript. J.Z. designed and executed the experiments and performed experimental analysis. Q.L. and H.C. supervised the research, provided critical guidance, and reviewed/refined the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Philippe Autier, Pengtao Xie, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, J., Huang, F., Luo, Q. et al. A Benchmark for Breast Cancer Screening and Diagnosis in Mammogram Visual Question Answering. Nat Commun 16, 11683 (2025). https://doi.org/10.1038/s41467-025-66507-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66507-z