Abstract
Complex systems often involve higher-order interactions that go beyond pairwise networks. Triadic interactions, where one node regulates the interaction between two others, are a fundamental form of higher-order dynamics found in many biological systems, from neuron-glia communication to gene regulation and ecosystems. However, triadic interactions have so far been mostly neglected. In this article, we propose the Triadic Perceptron Model (TPM) which shows that triadic interactions can modulate the mutual information between the dynamical states of two connected nodes. Leveraging this result, we formulate the Triadic Interaction Mining (TRIM) algorithm to extract triadic interactions from node metadata, and we apply this framework to gene expression data, finding new candidates for triadic interactions relevant for Acute Myeloid Leukemia. Our findings highlight crucial aspects of triadic interactions that are often ignored, offering a framework that can deepen our understanding of complex systems across biology, ecology, and climate science.
Similar content being viewed by others
Introduction
Higher-order networks1,2,3,4,5 are key to capturing many-body interactions present in complex systems. Inferring higher-order interactions6,7,8,9,10,11 from real, pairwise network datasets is recognized as one of the central challenges in the study of higher-order networks2,12, with wide applicability across different scientific domains, from biology and brain research13,14,15 to finance16,17. Mining higher-order interactions from the exclusive knowledge of the pairwise networks typically involves generative models and Bayesian approaches based on network structural properties6,7,9,11,18. Note, however, that when the inference is performed on the basis of the knowledge of the nodes’ dynamical states8,10, inferring higher-order interactions also requires dynamical considerations.
Triadic interactions19 are a fundamental type of signed higher-order interaction that are gaining increasing attention from the statistical mechanics community19,20,21,22,23,24, since they are not reducible to hyperedges or simplices. A triadic interaction occurs when one or more nodes regulate the interaction between two other nodes. The regulator nodes may either enhance or inhibit the interaction between the other two nodes. Triadic interactions are known to be important in various systems, including: ecosystems25,26,27 where one species can regulate the interaction between two other species; neuronal networks28, where glial cells regulate synaptic transmission between neurons, thereby controlling brain information processing; and gene regulatory networks29,30, where a modulator can promote or inhibit the interaction between a transcription factor and its target gene. There is mounting evidence that triadic interactions can induce collective phenomena and/or modulate dynamical states that reveal important aspects of complex system behavior19,20,21,22,23,24,31,32. An important advance in this line of research is triadic percolation19,20,21, a theoretical framework that captures the non-trivial dynamics of the giant component. Moreover, recent results demonstrate that triadic interactions can have significant effects on stochastic dynamics24 and learning22,23. However, despite the increasing attention that higher-order interactions are receiving, the detection of triadic interactions from network data and node time series, is an important scientific challenge that has not been thoroughly explored29,33,34.
In this article, we formulate the triadic perceptron model (TPM) in which continuous node variables are affected by triadic interactions. Based on the insights gained by investigating this model, we propose an information theoretic approach, leading to the triadic interaction mining (TRIM) algorithm, for mining triadic interactions. The TPM provides evidence of the mechanisms by which a triadic interaction can induce a significant variability of the mutual information between two nodes at the end-points of an edge. The TRIM algorithm leverages this finding and mines triadic interactions using knowledge of the network structure and the dynamical variables associated with the nodes. The significance of each putative triadic interaction is then validated by comparison with two distinct null models.
In this way, the TRIM algorithm can go beyond monotonicity assumptions regarding the functional form of the regulation of the two linked nodes by the third node (which is at the foundation of previously proposed methods29) allowing for broader applications. Significant node triples are also associated with an normalized entropic score function S ∈ [0, 1] that quantifies the spread of the conditional joint distribution functions of the variables at the ends of the regulated edge. We test the TRIM algorithm on the benchmark TPM, demonstrating its efficiency in detecting true triadic interactions. We also use the TRIM model to mine triadic interactions from gene-expression to identify “trigenic” processes35. We demonstrate that the TRIM algorithm is able to detect known interactions as well as propose a set of new candidate interactions that can then be validated experimentally.
Results
Triadic interactions
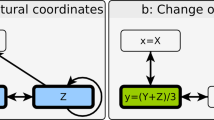
A triadic interaction occurs when one or more nodes modulate (or regulate) the interaction between two other nodes, either positively or negatively. A triadic interaction network is a heterogeneous network composed of a structural network and a regulatory network encoding triadic interactions (Fig. 1). The structural network GS = (V, ES) is formed by a set V of N nodes and a set ES of L edges. The regulatory network GR = (V, ES, ER) is a signed bipartite network with one set of nodes given by V (the nodes of the structural network), and another set of nodes given by ES (the edges of the structural network) connected by the regulatory interactions ER of cardinality \(| {E}_R|=\hat{L}\). The signed regulatory network can be encoded as an L × N matrix K where Kℓi = 1 if node i activates the structural edge ℓ, Kℓi = −1 if node i inhibits the structural edge ℓ and Kℓi = 0 otherwise. If Kℓi = 1, then the node i is called a positive regulator of the edge ℓ, and if Kℓi = −1, then the node i is called a negative regulator of the edge ℓ. It is worth noting that node i ∈ V cannot serve as both a positive and negative regulator for the same edge ℓ at the same time. However, node i can act as a positive regulator for edge ℓ while simultaneously functioning as a negative regulator for a different edge \({\ell }^{{\prime} } \,\ne \; \ell\).
a A triadic interaction occurs when a node Z, called a regulator node, regulates (either positively or negatively) the interaction between two other nodes X and Y. The regulated edge can be conceptualized as a factor node (shown here as a cyan diamond). b A network with triadic interactions can be seen as a network of networks formed by a simple structural network and by a bipartite regulatory network between regulator nodes and regulated edges (factor nodes).
The triadic perceptron model (TPM)
Here, we formulate a model for node dynamics in a network with triadic interactions that we call TPM. The TPM acts as a benchmark to validate the TRIM algorithm proposed here. We assume that each node i of the network is associated with a dynamical variable \({X}_{i}\in {\mathbb{R}}\), and that the dynamical state of the entire network is encoded in the state vector \(X={({X}_{1},{X}_{2}\ldots,{X}_{N})}^{T}\). The topology of the structural network is encoded in the graph Laplacian matrix L with elements
where a is the adjacency matrix of the network of elements aij, and J > 0 is a coupling constant. In the absence of triadic interactions, we assume that the dynamics of the network is associated with a Gaussian process implemented as the Langevin equation
with the Hamiltonian
where Γ > 0, α > 0, and where η(t) indicates uncorrelated Gaussian noise with
for all t and \({t}^{{\prime} }\). The resulting Langevin dynamics are given by
We remark that the Hamiltonian \({{{\mathcal{H}}}}\) has a minimum for X = 0, and its depth increases as the value of α increases. In a deterministic version of the model (Γ = 0), the effect of the structural interactions will not be revealed at stationarity. However, the Langevin dynamics with Γ > 0 encode the topology on the network. Indeed, at equilibrium, the correlation matrix \({C}_{ij}={\mathbb{E}}(({X}_{i}-{\mathbb{E}}({X}_{i}))({X}_{j}-{\mathbb{E}}({X}_{j})))\) is given by
see Supplementary Information (SI) for details. In other words, from the correlation matrix, it is possible to infer the Laplacian, and hence the connectivity of the network.
We now introduce triadic interactions in the TPM. As explained earlier, a triadic interaction occurs when one or more nodes modulate the interaction between another two nodes. To incorporate triadic interactions into the network dynamics, we modify the definition of the Laplacian operator present in the Langevin equation. Namely, we consider the Langevin dynamics
obtained from Eq. (2) by substituting the graph Laplacian L with the triadic Laplacian L(T) whose elements are given by
Moreover, we assume that the coupling constants Jij(X) are determined by a perceptron-like model that considers all the regulatory nodes of the link ℓ = [i, j] and the sign of the regulatory interactions. Specifically, if \({\sum }_{k=1}^{N}{K}_{\ell k}{X}_{k}\ge \hat{T}\) then we set Jij = w+; if instead \({\sum }_{k=1}^{N}{K}_{\ell k}{X}_{k} < \hat{T}\) then we set Jij = w_, with \({w}_{+},{w}_{-}\in {{\mathbb{R}}}_{+}\) and w+ > w−. Thus,
where θ( ⋅ ) is the Heaviside function (θ(x) = 1 if x ≥ 0 and θ(x) = 0 if x < 0). Note that, in the presence of triadic interactions, the stochastic differential equation (7) is not associated with any Hamiltonian, and a stationary state of the dynamics is not guaranteed, making this dynamical process significantly more complex than the original Langevin dynamics given in Eq. (2). The TPM is related to a recently proposed model that captures information propagation in multilayer networks24, but the TPM does not make use of a multilayer representation of the data. Moreover, the TPM is significantly different from models of higher-order interactions previously proposed in the context of consensus dynamics36,37 or contagion dynamics38,39. Indeed, in our framework, triadic interactions between continuous variables are not reducible to standard higher-order interactions because they involve the modulation of the interaction between a pair of nodes. Moreover, this modulation of the interaction is not dependent on the properties of the interacting nodes and their immediate neighbors, as is the case in36,37 or in the machine learning attention mechanism40. On the contrary, the modulation of the interaction is determined by a third regulatory node (or a larger set of regulatory nodes) encoded in the regulatory network.
The TPM for continuous node dynamics in the presence of triadic interactions is very general and comprehensively expresses the modulation of structural interactions by other nodes in the network. Therefore, the dynamics of TPM cannot be reduced to dynamics exclusively determined by pairwise interactions. An important problem that then arises is whether such interactions can be mined from observational data. To address this issue, we will develop a new algorithm—that we call the TRIM algorithm—to identify triadic interactions from data, and we will test its performance on the data generated from the TPM model described above.
Mining triadic interactions with the TRIM algorithm
We propose the TRIM algorithm (see Fig. 2) to mine triadic interactions among triples of nodes. To simplify the notation, we will use the letters X, Y, Z to indicate both nodes as well as their corresponding dynamical variables.
The TRIM algorithm identifies triples of nodes X, Y, and Z involved in a putative triadic interaction, starting from the knowledge of the structural network and the dynamical variables associated with its nodes. For each putative triple of nodes involved in a triadic interaction (a), which belongs to a network whose structure and dynamics are known (b), we study the functional behavior of the conditional mutual information MIz (c), and assess the significance of the observed modulations of MIz with respect to a null model (d). Given a predetermined confidence level, we can use these statistics to identify significant triadic interactions (e). This procedure can be extended to different triples of the network, thereby identifying the triadic interactions present in it (f).
Given a structural edge between nodes X and Y, our goal is to determine a confidence level for the existence of a triadic interaction involving an edge between node X and node Y with respect to a potential regulator node Z. Specifically, we aim to determine whether the node Z regulates the edge between node X and node Y, given the dynamical variables X, Y and Z associated with these nodes. To do so, given a time series associated with node Z, we first sort the Z-values, and define P bins in terms of the quantiles of z, chosen in such a way that each bin mz comprises the same number of data points (ranging in our analyses from 30 to 100). We indicate with zm the quantile of Z corresponding to the percentile m/P. Therefore, each bin mz indicates data in which Z ranges in the interval [zm, zm+1). We indicate with μ(x∣zm), μ(y∣zm) and μ(x, y∣zm) the probability density of the variables X, Y and the joint probability density of the variables X and Y in each mz bin.
A triadic interaction is taken to occur when the node Z affects the strength of the interaction between the other two nodes X and Y. Consequently, our starting point is to consider the mutual information between the dynamical variables X and Y conditional to the specific value of the dynamical variable Z. We thus consider the quantity MIz(m) = MI(X, Y∣Z = zm) defined as
In order to estimate this quantity, we rely on non-parametric methods based on entropy estimation from k-nearest neighbors41,42,43 (see SI for details). For each triple of nodes, we visualize the mutual information MIz computed as a function of the m/P-th quantiles zm and fit this function with a decision tree comprising r splits.
In the absence of triadic interactions, we expect MIz to be approximately constant as a function of the m/P-th quantiles zm, while, in the presence of triadic interactions, we expect this quantity to vary significantly as a function of zm. The discretised conditional mutual information CMI between X and Y conditioned on Z can be written as
where p(zm) = 1/P indicates the probability that the Z value falls in the mz bin. This quantity indicates important information about the interaction between the nodes X and Y when combined with the information coming from the mutual information MI, given by
where μ(x), μ(y) are the probability density functions for X and Y, and μ(x, y) is the joint probability density function of the variables X and Y. The conditional mutual information, however, is not sensitive to variations in MIz and does not therefore provide the information needed to detect triadic interactions. In order to overcome this limitation, we define the following two quantities that measure how much the mutual information between X and Y conditioned on Z ∈ [zm, zm+1) changes as zm varies. Specifically, we consider:
-
(1)
the standard deviation Σ of MIz, defined as
$$\begin{array}{r}\Sigma=\sqrt{{\sum }_{m=0}^{P-1}p({z}_{m}){\left[MIz(m)-\left\langle MIz\right\rangle \right]}^{2}};\end{array}$$(11) -
(2)
the difference T between the maximum and average value of MIz, given by
$$\begin{array}{r}T={\max }_{m=0,\ldots,P-1}\left\vert MIz(m)-\left\langle MIz\right\rangle \right\vert .\end{array}$$(12)
The quantities Σ and T collectively measure the strength of the triadic interaction under question and can thus be used to mine triadic interactions in synthetic as well as in real data. In order to assess the significance of the putative triadic interaction, we compare the observed values of these variables to the results obtained with given null models. In order to determine if the observed values are significant with respect to a given null model, we compute the scores ΘΣ, ΘT, given by
and the p-values
Note that if we consider \({{{\mathcal{N}}}}\) realizations of the null model, we cannot estimate probabilities smaller than \(1/{{{\mathcal{N}}}}\). Therefore, if in our null model we observe no value of Σran larger than the true data Σ, we set the conservative estimate \({p}_{\Sigma }=1/{{{\mathcal{N}}}}\). A similar procedure is applied also to pT.
To assess this significance, we consider two types of null models. The first is the randomization null model obtained by shuffling the Z values, to give \({{{\mathcal{N}}}}\) randomization of the data, i.e., we use surrogate data for testing44,45. The second is the maximum likelihood Gaussian null model between the three nodes involved in the triple X,Y,Z. Specifically, the Gaussian null model uses the mean and covariance of the timeseries of X, Y and Z to define a multivariate normal distribution from which samples are randomly drawn, thereby providing surrogate timeseries values for the considered triple. We note that the use of these two null models also allows us to identify non-monotonic relationships between MIz and z, thereby going beyond underlying monotonic assumptions made elsewhere46. The first null model disrupts the temporal correlations between the timeseries of the node Z and the timeseries the two nodes X and Y at the endpoints of the considered edges. Therefore, this null model is robust with respect to the presence of possible outliers in the dataset. However, this first null model may overlook confounding network effects that affect correlations between the dynamical variables. The second null model more efficiently captures correlations between the dynamical state of the three considered nodes due to network effects, but is more sensitive to the presence of outliers in the data. To increase confidence, we therefore combined the insights coming from both these null models (see SI for details).
For each triple, the function MIz(m) is fitted with a decision tree with two splits. In this way, three different intervals of values of Z are identified, each corresponding to a distinct functional behavior of the correlation functions between the variables X and Y. While our method in principle allows for more than two splits of the decision tree, for illustrative purposes we have chosen two splits since this is the minimum number of splits needed to capture non-trivial functional behavior in MIz, such as non-monotonicity. In practice, this choice of two splits will also be the best choice when data is limited, such as the gene expression data we will analyze in the following section.
We also further characterize significant triples by examining their normalized entropic score function S ∈ [0, 1], which is used to characterize their corresponding functional behavior. Specifically, the entropic score S classifies the diversity of each of the joint distribution functions of X and Y conditioned on Z for each interval obtained through the decision tree (see Methods for details).
As we will discuss below, the algorithm performs well on data obtained from the TPM. In this case, we also observe that true triadic triples are characterized by a high entropic score S. On real data, the results obtained with the TRIM algorithm using randomized surrogate data might neglect potentially important network effects, this shortcoming is mitigated by performing an additional validation using the Gaussian null model and the entropic score S (see SI for the full pipeline of TRIM).
Validation of the TRIM algorithm on the triadic perceptron model
In order to discuss the phenomenology of the TPM, we first considered a representative network (see Fig. 3) of N = 10 nodes, L = 12 edges and \(\hat{L}=5\) triadic interactions (each formed by a single node regulating a single edge) on top of which we consider the TPM.
We consider a network with N = 10 nodes, L = 12 edges, and \(\hat{L}=5\) triadic interactions (a). b, c Display the effect of triadic interactions on the Mutual Information profile MIz. b shows MIz for the triple [4, 9, 5] involved in a positive triadic interaction. c Shows MIz for the triple [1, 2, 6] that is not involved in a triadic interaction. In all panels, simulations were run to \({t}_{\max }=4,000\) with a timestep of dt = 10−2. For the analysis we consider 40, 000 time steps. The parameters of the model are: \(\alpha=0.05,\hat{T}=1{0}^{-3},\Gamma=1{0}^{-2}\), w+ = 8, w− = 0.5, number of bins P = 400.
We found that data obtained from the TPM on this network shows a strong dependence of MIz(mz) on mz for the triples of nodes involved in triadic interactions, with greater significance for smaller values of α. Figure 3 shows the difference between the MIz(m) profile of a triple that is involved in a triadic interaction compared to a triple that is not, demonstrating how triadic interactions modulate the MIz(mz) profile.
Moreover, Fig. 4 shows, for a given triadic interaction involving nodes X, Y and Z, the joint distributions μδ(X, Y) of X and Y for each interval δ of values Z determined by the decision tree. The results provide evidence of this interesting dynamical behavior of the triadic model in the case of a positive regulatory interaction. Note that the analysis of the form of the function MIz(mz) also allows us to distinguish between positive and negative regulatory interactions, which are associated with an increase or a decrease in MIz for larger values of mz, respectively.
Results for the triple [4, 9, 5] of the network in Fig. 3 of the main text, which is triadic, are shown. The joint distributions of variables X and Y conditional on the values of Z are shown in a. b Shows the behavior of MIz as a function of the values of zm, which clearly departs from the constant behavior expected in absence of triadic interactions. c Presents the decision tree for fitting the MIz functional behavior and determining the range of values of Z for which the most significant differences among the joint distributions of the variables X and Y conditioned on Z are observed. The parameters used are the same as in Fig. 3.
In the Supplementary Figs. S1 and S2, we display further examples of the function MIz for triadic triples. We observe the increased variability of the MIz functional behavior as the parameter Γ is raised, i.e., the noise increases.
These results confirm the main general principle on which the TRIM algorithm is based, i.e., that the conditional mutual information MIz is modulated by triadic interactions. To make this observation precise, we examined the performance of the TRIM algorithm in mining triadic interactions from synthetic data. We first considered the network shown in Fig. 3, and using the score ΘΣ, we evaluated the receiver operating characteristic (ROC) curve and precision recall (PR) curve for different values of the dynamical parameters (see Fig. 5). Both the ROC curve and the PR curve (which addresses the limitations of the ROC curve for imbalanced datasets) indicate that the TRIM algorithm performs well on data produced by the TPM, with a better performance for higher values of α.
We consider the network in Fig. 3a. The time series obtained by integrating the stochastic dynamics of the proposed dynamical model for triadic interactions (Eq. (7)) are analyzed with the TRIM algorithm. a Displays the receiver operating characteristic curve (ROC curve) obtained by running TRIM with P = 400 bins and \({{{\mathcal{N}}}}=1{0}^{3}\) realizations of the null model on these synthetic time series, using ΘΣ to score for different parameters values indicated in the legend. b Displays the corresponding precision-recall curve (PR curve) obtained by running TRIM with the same parameters. The timeseries are simulated up to a maximum time tmax = 4000 with a dt = 10−2. For the analysis, we consider 40, 000 time steps (see the SI for details). The parameter of the model are: \(\hat{T}=1{0}^{-3}\), w+ = 8, w− = 0.5, and α and Γ as indicated in the figure legend.
For all parameter values, we noticed that false positives are more likely to involve short-range triples, i.e., triples in which the regulator node Z is close (in the structural network) to the end-points X and Y of the target edge.
These results indicate that the TRIM algorithm is effective in identifying triadic interactions in a small network generated using the TPM. To examine the scalability of this methodology we also tested the TRIM algorithm on a much larger model network. To this end, we consider a random Erdös-Renyi network of 100 nodes, and average degree c = 4, to which we added 25 random triadic interactions (i.e., between randomly chosen nodes to randomly chosen edges), imposing the condition that each edge is at most regulated by a single node for simplicity. The results of this analysis are shown in Fig. 6a, in which we provide statistics for all possible node triples in the network (the majority of which are not triadic interactions). For each edge, we retained only the 5 most significant triples according to ΘΣ. By conditioning on the value of the third node, for each of these connected nodes we also record the conditional mutual information CMI. In Fig. 6, each considered triple corresponds to a point, color coded according to the value of S. Stars indicate triples that are involved in a triadic interactions (see SI for details). True triadic interactions are found for triples with high ΘΣ, while CMI span between high and intermediate values. This result confirms the very good performance of the TRIM algorithm on the data coming form the TPM.
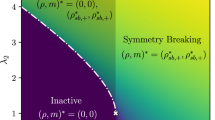
a Each data point represents a given triple of nodes X, Y and Z. The y-axis shows ΘΣ, while the x-axis shows the CMI between X and Y. The color of each point corresponds to the value of S, which characterizes the entropic score of the triple. The synthetic data comes from a structural random Erdös-Renyi network with 100 nodes, and average degree c = 4, to which 25 triadic interactions between random edges and random nodes have been added. For the modeling of the network we used α = 0.06, Γ = 1.4 × 10−2, tmax = 1500, w+ = 18, w− = 0.2 and for the analysis with TRIM we looked at 3000 data points and P = 100 bins. We display top 5 triples for each edge according to ΘΣ that are below our p threshold for the randomization null model. The triples below are represented in the scatter plot, and they all display an entropic score S > 0.5. Stars are the true triadic triples, which are characterized by high ΘΣ. Crosses are the triples that can be excluded by performing TRIM with the Gaussian Null model. b Histogram of the ΘΣ-values for all the triples of the network (in light blue), and for the triples corresponding to the 25 true triadic interactions only (in dark blue). c Histogram of the ΘΣ-values observed in a network of the same topology and with the same dynamical parameters for which all the triadic interactions have been removed (orange).
To test the statistical robustness of the TRIM algorithm, we also conducted the same analysis (i.e., on the same structural network with the same dynamical parameters) in which all triadic interactions were removed. The results of this analysis are shown in Fig. 6c. In this case, and as expected, the TRIM algorithm did not identify any statistically significant triadic interactions. This analysis indicates that the TRIM algorithm is able to identify true triadic interactions with a low false positive discovery rate (compare Fig. 6b, c).
Detecting triadic interactions in gene-expression data
Searching for triadic interactions in gene-expression is a problem of major interest in biology. For instance, understanding the extent to which a modulator promotes or inhibits the interplay between a transcription factor and its target gene is crucial for deciphering gene regulation mechanisms29. In order to address this question with our method, we considered a gene-expression dataset associated with acute myeloid leukemia (AML), extracted from the grand gene regulatory network database47,48.
Exhaustive mining of all potential triadic interactions from every putative triple of nodes in the AML dataset is computationally very demanding (it would require testing of >260M triples) and likely, due to the sheer number of triples being tested, to result in false positives and/or interactions of less biological importance. Moreover, such a brute-force approach would not account for other sources of important biological information, such as putative interactions derived from other experimental sources. To account for such information, we therefore focused our analysis on edges between nodes associated with known biophysical interactions, as identified in the human protein–protein interaction network (PPI)47. Because proteins function through physical and functional interactions, PPI networks provide a biologically meaningful framework for interpreting gene expression changes in terms of coordinated molecular mechanisms49,50. To do this, we considered the connected subgraph of the human PPI network that contains all the genes/proteins included in the AML gene expression data and their associated edges. This network, which contains 622 nodes and 42, 511 edges, formed the structural network for our analysis51. To start, we focused on triples involving genes known to be associated with AML, in which the end-points X and Y of the target are directly connected in the PPI network (see SI for details).
We then selected additional triples according to their positions in the PPI network’s maximum spanning tree (MST), which only includes 621 edges (see Fig. 7). In order to focus on triples for which network effects are likely to be less pronounced, for each edge in the MST, connecting gene/protein X with gene/protein Y we considered all genes Z within a distance of 4 from both the X and Y as candidate regulatory nodes, i.e., the third node in the triple (see SI for details). For each considered triple of genes, we assessed its significance using ΘΣ as the significance score, with P = 5 bins, using \({{{\mathcal{N}}}}=5\times 1{0}^{3}\) realizations of the randomization null model (very similar results were obtained using ΘT as the significance score, see Supplementary Figs. S3-S4 for a comparison).
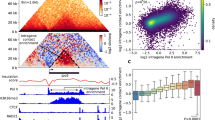
Figure 8 shows the results of the TRIM algorithm for those triples with pΣ < 0.001. Note that for each selected edge, only the top 5 triples ranked according to the ΘΣ score are depicted. Squares indicate triples chosen from biologically relevant genes for AML. The triples deemed insignificant according to the Gaussian null model, are not shown here. The interested reader can see their visualization in Fig. S6 of the SI. Figure 8 provide examples of conditional distributions of two illustrative triples that rank high in TRIM. Both triples show evidence of a triadic interaction: the triple in panel b is a member of the MST, while panel c shows an example of a triple chosen from known biologically relevant genes for AML. Further example triples are shown in the SI. Interestingly, among the significant triples, we also detected triples in which the modulation of the mutual information is non-monotonic (see SI for details).
a Shows the results of TRIM for the significant triples in the AML dataset. The scatter plot shows ΘΣ (y-axis) versus the CMI (x-axis). The colour of each point corresponds to the value of its entropic score S. Here we display only those triples with p-value 0.001 or less in the randomization null model and that have not been excluded by the Gaussian null model (for details about these triples see SI). Circles are triples whose links all appear in the minimum spanning tree, and squares indicate triples involving genes with biological relevance. b, c Display the conditional distributions for two example triples: both are identified by the TRIM algorithm with high significance, suggesting a meaningful biological association. Panel b shows the triple X = GATA1, Y = KLF1, Z = ETV1. According to the randomized surrogate null model, this triadic interaction has pΣ-value 0.001, ΘΣ = 4.75, Σ = 0.44, S = 0.64; panel c shows the results for the triple X = HOXB3, Y = MEIS1, Z = GLIS3 involving two biologically relevant genes. According to the randomized surrogate null model, it has ΘΣ = 3.98, pΣ = 0.001, Σ = 0.38, S = 0.60.
Many of the genes involved in the 50 highest ranking triples have already been linked to AML in the literature (see SI for Table S3 for a list of highly significant triples and Table S4 with links to the literature associating the involved genes with AML). In total, 84% of the top 50 Triples include at least one gene that has a known association with AML.
Discussion
This work provides a comprehensive information theory-based framework to model and mine triadic interactions directly from dynamic observations. The TPM we propose demonstrates that the presence of a triadic interaction leads to systematic variations in the mutual information between the two end nodes of the edge involved (X and Y). Via this model, we have shown that to detect triadic interactions, it is necessary to go beyond standard pairwise measures, such as the mutual information. Importantly, standard higher-order statistical measures, such as the conditional mutual information, which accounts for the average effect of the third regulatory node Z on the mutual information between the target nodes X and Y, are also insufficient to identify triadic interactions. Our proposed approach, implemented in the TRIM algorithm, mines triadic interactions by identifying statistically significant variations in the mutual information between the two linked nodes conditioned on the third regulator node.
To demonstrate the efficacy of this algorithm, we have tested and validated it on a new dynamical model (that we denote the TPM) and shown how it can identify triadic interactions in randomly generated triadic interaction networks. We also used it to mine putative triadic interactions from gene expression data and connect the putative interactions with meaningful biology.
From the network theory point of view, this work opens new perspectives in the active field of modeling and inference of higher-order interactions and can be extended in many different directions, for instance by exploring the effect of triadic interactions on the dynamical state of nodes associated with discrete variables or including time delays in the regulation. From the biological point of view, our results may inspire further information-theoretic approaches to genetic regulatory network inference. Investigating the extent to which triadic interactions are tissue-specific, and if certain regulatory patterns are conserved across different tissues could yield valuable insights. Our proposed approach could also be used to mine triadic interactions in other domains, such as finance or climate, where triadic interactions also have a significant role.
Methods
Entropic score S for significant triples
In order to identify and classify the significant triples [X, Y, Z] involving node X and Y whose interaction is modulated by node Z, we introduce an entropic score function S which characterizes how diverse the conditional joint distributions μδ(X, Y) of X and Y conditioned on Z in each of the obtained intervals δ ∈ {1, 2, 3} are. Dividing the plane X, Y in P2 squares (i, j) (by binning X and Y in P bins each) with \({n}_{ij}^{(\delta )}\) data points, we can calculate the participation ratio \({Y}_{2}^{(\delta )}\)52,53
where \({{{{\mathcal{N}}}}}^{(\delta )}={\sum }_{i=1}^{P}{\sum }_{j=1}^{P}{n}_{ij}^{(\delta )}\). The inverse of the partition function is known to measure the effective number of square bins in which the distribution is localized. We can then introduce the normalized entropic score S as
The entropy S is low if all the conditional distributions μδ(X, Y) are very localized while it acquires large values if all the conditional distributions are delocalized. We adopt a threshold S = 0.5 in order to retain triples with S > 0.5, indicating that in average the conditional distributions associated to these triples have more than \(\sqrt{{P}^{2}}\) significantly populated bins.
Data availability
The data used in this work a gene-expression dataset associated with Acute Myeloid Leukemia (AML), and the human Protein-Protein Interaction network (PPI) publicly available from the Grand Gene Regulatory Network Database47 and included together with our code in https://github.com/anthbapt/TRIM.
Code availability
The Python package TRIM generated in this study have been deposited in the GitHub database under accession code https://github.com/anthbapt/TRIM.
References
Bianconi, G. Elements in the structure and dynamics of complex networks. in Higher-Order Networks (Cambridge University Press, 2021).
Battiston, F. et al. The physics of higher-order interactions in complex systems. Nat. Phys. 17, 1093–1098 (2021).
Battiston, F. et al. Networks beyond pairwise interactions: structure and dynamics. Phys. Rep. 874, 1–92 (2020).
Torres, L., Blevins, A. S., Bassett, D. & Eliassi-Rad, T. The why, how, and when of representations for complex systems. SIAM Rev. 63, 435–485 (2021).
Bick, C., Gross, E., Harrington, H. A. & Schaub, M. T. What are higher-order networks? SIAM Rev. 65, 3 (2023).
Young, J.-G., Petri, G. & Peixoto, T. P. Hypergraph reconstruction from network data. Commun. Phys. 4, 135 (2021).
Contisciani, M., Battiston, F. & De Bacco, C. Inference of hyperedges and overlapping communities in hypergraphs. Nat. Commun. 13, 7229 (2022).
Malizia, F. et al. Reconstructing higher-order interactions in coupled dynamical systems. Nat. Commun. 15, 5184 (2024).
Musciotto, F., Battiston, F. & Mantegna, R. N. Detecting informative higher-order interactions in statistically validated hypergraphs. Commun. Phys. 4, 218 (2021).
Delabays, R., De Pasquale, G., Dörfler, F. & Zhang, Y. Hypergraph reconstruction from dynamics. 16, 2691 (2025).
Lizotte, S., Young, J.-G. & Allard, A. Hypergraph reconstruction from uncertain pairwise observations. Sci. Rep. 13, 21364 (2023).
Rosas, F. E. et al. Disentangling high-order mechanisms and high-order behaviours in complex systems. Nat. Phys. 18, 476–477 (2022).
Rosas, F. E., Mediano, P. A. M., Gastpar, M. & Jensen, H. J. Quantifying high-order interdependencies via multivariate extensions of the mutual information. Phys. Rev. E 100, 032305 (2019).
Stramaglia, S., Scagliarini, T., Daniels, B. C. & Marinazzo, D. Quantifying dynamical high-order interdependencies from the o-information: an application to neural spiking dynamics. Front. Physiol. 11, 595736 (2021).
Olbrich, E., Bertschinger, N. & Rauh, J. Information decomposition and synergy. Entropy 17, 3501–3517 (2015).
Massara, G. P., Di Matteo, T. & Aste, T. Network filtering for big data: triangulated maximally filtered graph. J. Complex Netw. 5, 161–178 (2016).
Tumminello, M., Aste, T., Di Matteo, T. & Mantegna, R. N. A tool for filtering information in complex systems. Proc. Natl. Acad. Sci. USA 102, 10421–10426 (2005).
Wegner, A. E. & Olhede, S. C. Nonparametric inference of higher order interaction patterns in networks. Commun. Phys. 7, 258 (2024).
Sun, H., Radicchi, F., Kurths, J. & Bianconi, G. The dynamic nature of percolation on networks with triadic interactions. Nat. Commun. 14, 1308 (2023).
Millán, A. P., Sun, H., Torres, J. J. & Bianconi, G. Triadic percolation induces dynamical topological patterns in higher-order networks. PNAS Nexus 3, 270 (2024).
Sun, H. & Bianconi, G. Higher-order triadic percolation on random hypergraphs. Phys. Rev. E 110, 064315 (2024).
Herron, L., Sartori, P. & Xue, B. K. Robust retrieval of dynamic sequences through interaction modulation. PRX Life 1, 023012 (2023).
Kozachkov, L., Slotine, J. J. & Krotov, D. Neuron-astrocyte associative memory. Proc. Natl. Acad. Sci. 122, e2417788122 (2025).
Nicoletti, G. & Busiello, D. M. Information propagation in multilayer systems with higher-order interactions across timescales. Phys. Rev. X 14, 021007 (2024).
Bairey, E., Kelsic, E. D. & Kishony, R. High-order species interactions shape ecosystem diversity. Nat. Commun. 7, 12285 (2016).
Grilli, J., Barabás, G., Michalska-Smith, M. J. & Allesina, S. Higher-order interactions stabilize dynamics in competitive network models. Nature 548, 210–213 (2017).
Letten, A. D. & Stouffer, D. B. The mechanistic basis for higher-order interactions and non-additivity in competitive communities. Ecol. Lett. 22, 423–436 (2019).
Cho, W.-H., Barcelon, E. & Lee, S. J. Optogenetic glia manipulation: possibilities and future prospects. Exp. Neurobiol. 25, 197–204 (2016).
Wang, K. et al. Genome-wide identification of post-translational modulators of transcription factor activity in human B cells. Nat. Biotechnol. 27, 829–837 (2009).
Giorgi, F. M. et al. Inferring protein modulation from gene expression data using conditional mutual information. PLoS ONE 9, e109569 (2014).
Gao, J. et al. Triadic percolation in computer virus spreading dynamics. Chin. Phys. B 34, 028701 (2025).
Iskrzyński, M., Puchalska, A., Grzelik, A. & Mutlu, G. Pangraphs as models of higher-order interactions. Preprint at https://doi.org/10.48550/arXiv.2502.10141 (2025).
Kenett, D. Y., Huang, X., Vodenska, I., Havlin, S. & Stanley, H. E. Partial correlation analysis: applications for financial markets. Quant. Financ. 15, 569–578 (2015).
Zhao, J., Zhou, Y., Zhang, X. & Chen, L. Part mutual information for quantifying direct associations in networks. Proc. Natl. Acad. Sci. USA 113, 5130–5135 (2016).
Kuzmin, E. et al. Systematic analysis of complex genetic interactions. Science 360, eaao1729 (2018).
Neuhäuser, L., Lambiotte, R. & Schaub, M.T. Consensus dynamics and opinion formation on hypergraphs. In Higher-Order Systems 347–376 (Springer, 2022).
Neuhäuser, L., Mellor, A. & Lambiotte, R. Multibody interactions and nonlinear consensus dynamics on networked systems. Phys. Rev. E 101, 032310 (2020).
Iacopini, I., Petri, G., Barrat, A. & Latora, V. Simplicial models of social contagion. Nat. Commun. 10, 2485 (2019).
de Arruda, G. F., Petri, G. & Moreno, Y. Social contagion models on hypergraphs. Phys. Rev. Res. 2, 023032 (2020).
Veličković, P. et al. Graph attention networks. Preprint at https://doi.org/10.48550/arXiv.1710.10903 (2017).
Kozachenko, L. F. & Leonenko, N. N. Sample estimate of the entropy of a random vector. Probl. Peredachi Informatsii 23, 9–16 (1987).
Kraskov, A., Stögbauer, H. & Grassberger, P. Estimating mutual information. Phys. Rev. E 69, 066138 (2004).
Ross, B. C. Mutual information between discrete and continuous data sets. PloS ONE 9, e87357 (2014).
Kurths, J. & Herzel, H. An attractor in a solar time series. Phys. D Nonlinear Phenom. 25, 165–172 (1987).
Theiler, J., Eubank, S., Longtin, André, Galdrikian, B. & Farmer, J. D. Testing for nonlinearity in time series: the method of surrogate data. Phys. D Nonlinear Phenom. 58, 77–94 (1992).
Wang, H., Lin, S.-Y., Hu, F.-F., Guo, A.-Y. & Hu, H. The expression and regulation of hox genes and membrane proteins among different cytogenetic groups of acute myeloid leukemia. Mol. Genet. Genom. Med. 8, e1365 (2020).
Ben Guebila, M. et al. “Grand gene regulatory network database" (2023).
Ben Guebila, M. et al. Grand: a database of gene regulatory network models across human conditions. Nucleic Acids Res. 50, D610–D621 (2022).
Safari-Alighiarloo, N., Taghizadeh, M., Rezaei-Tavirani, M., Goliaei, B. & Peyvandi, A. A. Protein-protein interaction networks (PPI) and complex diseases. Gastroenterol. Hepatol. Bed Bench 7, 17 (2014).
Sevimoglu, T. & Arga, K. Y. The role of protein interaction networks in systems biomedicine. Comput. Struct. Biotechnol. J. 11, 22–27 (2014).
Grigoriev, A. A relationship between gene expression and protein interactions on the proteome scale: analysis of the bacteriophage T7 and the yeast Saccharomyces cerevisiae. Nucleic Acids Res. 29, 3513–3519 (2001).
Barthelemy, M., Gondran, B. & Guichard, E. Spatial structure of the internet traffic. Phys. A Stat. Mech. Appl. 319, 633–642 (2003).
Derrida, B. & Flyvbjerg, H. Statistical properties of randomly broken objects and of multivalley structures in disordered systems. J. Phys. A Math. Gen. 20, 5273 (1987).
Alharbi, R. A., Pettengell, R., Pandha, H. S. & Morgan, R. The role of hox genes in normal hematopoiesis and acute leukemia. Leukemia 27, 1000–1008 (2013).
Guo, H. et al. Pbx3 is essential for leukemia stem cell maintenance in MLL-rearranged leukemia. Int. J. Cancer 141, 324–335 (2017).
Xiang, P. et al. Elucidating the importance and regulation of key enhancers for human meis1 expression. Leukemia 36, 1980–1989 (2022).
Acknowledgements
This work was sponsored by the Turing-Roche Strategic Partnership. M.N. was funded by the UKRI/BBSRC Collaborative Training Partnership in AI for Drug Discovery and Queen Mary University of London. This research utilized Queen Mary’s Apocrita HPC facility, supported by QMUL Research-IT, https://doi.org/10.5281/zenodo.438045.
Author information
Authors and Affiliations
Contributions
G.B. designed the research, M.N., A.B., and J.Y. conducted the numerical analysis, M.N., A.B., and J.Y. wrote the codes, M.N. finalized the figures, M.N., A.B., J.Y., J.K., R.S.G., B.D.M., and G.B. conducted the research, and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Niedostatek, M., Baptista, A., Yamamoto, J. et al. Mining higher-order triadic interactions. Nat Commun 16, 11613 (2025). https://doi.org/10.1038/s41467-025-66577-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66577-z
This article is cited by
-
Collective dynamics on higher-order networks
Nature Reviews Physics (2026)