Abstract
Osteoporosis therapies remain limited by non-specific osteoclast inhibition, which disrupts physiological bone remodeling and increases fracture risk. Here we show that selectively targeting mature osteoclasts via enzyme-responsive nanoparticles could eliminate pathological bone resorption while generating apoptotic bodies (ABs) to promote regeneration. We design OsteoSAVE, a peptide-based nanoparticle integrating a cathepsin K (CTSK)-cleavable linker, a bone-targeting motif, and a self-assembling biphenyl core. Upon CTSK-mediated hydrolysis, OsteoSAVE transforms into nanofibers within osteoclast lysosomes, inducing apoptosis and releasing ABs enriched with PDGF-BB. Mechanistic studies confirm CTSK-selective cleavage and charge-driven cellular specificity. In ovariectomized mice, OsteoSAVE restores bone density by coupling osteoclast elimination with ABs-mediated osteogenesis via PI3K/AKT activation in mesenchymal stem cells. This work establishes enzyme-triggered supramolecular self-assembly as a paradigm for dual-action therapies that recalibrate bone homeostasis without disrupting physiological processes.
Similar content being viewed by others
Introduction
The aging population faces significant health challenges, with osteoporosis posing a critical burden, especially among postmenopausal women1. Characterized by deterioration of bone microstructure and accelerated bone loss, osteoporosis arises from an imbalance between bone resorption and formation, primarily driven by excessive osteoclast (OC) activity2. This imbalance increases fracture risk, contributing to elevated morbidity and mortality3. Current first-line treatments, such as bisphosphonates, indiscriminately inhibit OCs, leading to apoptosis of all bone-resorbing cells4. While effective in slowing disease progression, these therapies disrupt pre-osteoclasts (pre-OCs)-mediated bone remodeling processes5, often resulting in serious complications6,7. Consequently, there is an urgent need for targeted therapeutic strategies that selectively limit excessive osteoclastic activity while maximally preserving pre-OCs activity to maintain physiological bone remodeling.
OCs are macrophage-derived multinucleated cells formed by the fusion of bone marrow-derived monocytes (BMMs), tightly controlled by two key cytokines: receptor activator of nuclear factor κB ligand (RANKL) and monocyte colony-stimulating factor (M-CSF)8. The formation of OCs induced by RANKL involves the differentiation of BMMs into pre-OCs, followed by their fusion into mature OCs9. While pre-OC membranes have been explored as a therapeutic strategy to inhibit the fusion of pre-OCs to form OCs, mature OCs (already formed) may also fuse with pre-OC membranes, potentially reducing the therapeutic efficacy10. Notably, mature OCs exhibit high expression of cathepsin K (CTSK)11, a lysosomal enzyme critical for bone resorption, whereas pre-OCs show minimal CTSK activity9. Despite its pivotal role, there are currently no CTSK-based precision therapies for osteoporosis. Additionally, treating osteoporosis requires bone targeting in vivo to minimize the risk of side effects at non-bone sites12. A previous study demonstrated that a sequence of six aspartic acid residues (D-Asp6) can effectively deliver drugs to bone tissue13. Therefore, we developed a osteoporosis treatment strategy that combines bone targeting capability with CTSK responsiveness.
OCs have a short lifespan and undergo apoptosis after completing their resorptive function14. This process involves nuclear and cytoplasmic condensation, membrane blebbing, and the formation of apoptotic bodies (ABs)15. The contents of ABs are highly variable and may include nucleic acids, proteins, and lipids. Recent studies suggest that pharmacologically induced OCs-derived ABs, generated using agents like Alendronate (ALD) and Staurosporine, can promote bone formation and modulate bone remodeling16,17. However, these approaches lack selectivity, inducing apoptosis in both OCs and pre-OCs, and are unsuitable for in vivo application. Leveraging the unique enzymatic profile of OCs in osteoporosis, we propose a targeted strategy to selectively induce OC apoptosis in vivo and generate therapeutic ABs that restore bone homeostasis.
Here, we introduce OsteoSAVE, a bone-targeted nanoparticle system designed to induce OC apoptosis through CTSK-assisted self-assembly. OsteoSAVE nanoparticles are designed to target bone surface and have passive cell targeting properties due to their negative charge, which allows them to be engulfed in large quantities only by macrophages, such as OCs and pre-OCs. Within in OC lysosomes, CTSK triggers their transformation into nanofibers, induce apoptosis and generating ABs enriched with PDGF-BB. These ABs regulate osteogenic differentiation of bone marrow mesenchymal stem cells (BMSCs), restoring bone homeostasis in osteoporosis. By coupling selective OC elimination with in vivo generation of therapeutic ABs, OsteoSAVE offers a promising strategy for treating osteoporosis and advancing therapies for age-related bone disorders.
Results
Molecular design and characterization of OsteoSAVE-BGD nanoparticles
We developed OsteoSAVE nanoparticles, termed BGD, by integrating a self-assembling sequence with a CTSK-cleavable peptide linker and a bone-targeting sequence (Fig. 1b). The modular structure of BGD includes: i) Bip-FF: This component features a biphenyl group (Bip) and two phenylalanine (FF), enhancing hydrophobicity at the N-terminus18,19,20,21,22. It facilitates the formation of stable self-assembled structures through π-π interactions23,24. Previous study has demonstrated that Bip-FF not only facilitates the self-assembly but also increases membrane disruption when assemblies accumulate locally25. ii) WKYK: A hydrophobic/positively charged sequence to enable lysosomal membrane disruption post-assembly. iii) GHPG↓GPQ: a CTSK-cleavable peptide linker for targeted cleavage26. iv) DDDDDD: A hexa-aspartic acid bone-targeting motif13 to direct nanoparticles to bone surfaces and confer a negative surface charge. BGD nanoparticles respond to CTSK overexpressed by mature OCs, undergoing enzymatic cleavage to form nanofibers that induce apoptosis. The resulting OCs-derived ABs promote osteogenic differentiation of bone marrow mesenchymal stem cells (BMSCs), restoring bone homeostasis (Fig. 1a). The peptides are synthesized by standard Fmoc-based solid-phase peptide synthesis (SPPS) method (Supplementary Fig. 1). We also designed several control peptides to examine the importance of each part (Table S3). The structure, purity, and molecular weight of the peptides are confirmed by liquid chromatography-mass spectrometry (LC-MS) and matrix-assisted laser desorption ionization with time-of-flight mass spectrometry (MALDI-TOF, Supplementary Fig. 2).
a By intravenous injection, the bone-targeted OsteoSAVE-BGD nanoparticles are recruited to the bone surface and passively target OCs. When the nanoparticles are phagocytosed by OCs, CTSK in lysosomes triggers their conversion into nanofibers, inducing OCs apoptosis and producing ABs rich in PDGF-BB. These ABs can activate the PI3K/AKT signaling pathway in BMSCs to promote their osteogenic differentiation, ultimately achieving the goal of restoring bone homeostasis in vivo. b The structure of the nanoparticle with bone targeting and CTSK responsiveness.
Critical micelle concentration (CMC) measurements by Spectrofluorometer (Edinburgh, F55) revealed that BGD (7.8 μM), BAD (11.8 μM), and BGG (14.37 μM) exhibited strong self-assembly capabilities (Fig. 2a and Supplementary Fig. 3). However, WGD (222.83 μM) exhibited weak self-assembly ability (Supplementary Fig. 3). Using TEM to observe the morphological changes of peptides in PBS buffer (pH = 7.4). Transmission electron microscopy (TEM) indicated that both BGD and BAD formed dispersed, nearly spherical nanoparticles in an aqueous solution (pH 7.4), while BGG self-assembled into uniform nanofibers with a diameter about 10 nm (Fig. 2b and Supplementary Fig. 4). Dynamic light scattering (DLS) analysis further confirmed that BGD and BAD formed nanoparticles average sizes of 32.5 and 36.6 nm (Fig. 2c and Supplementary Fig. 5), respectively. Zeta potential measurements indicated that BGD nanoparticles (−32.8 mV) and BAD nanoparticles (−20.8 mV) possessed high surface negative charges, suggesting excellent colloidal stability27 (Fig. 2d and Supplementary Fig. 5).
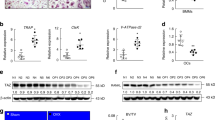
a The CMC measure of BGD. b Transmission electron microscopy (TEM) images of nanoparticles formed by the self-assembly of BGD at pH 7.4 (n = 3 biologically independent samples, scale bars = 50 nm). c The diameter of nanoparticles formed by the self-assembly of BGD. d The zeta potential of nanoparticles formed by the self-assembly of BGD (n = 3 biologically independent samples). e, f TEM images of the reaction mixture of BGD incubated with the CTSK enzyme for 8 h at pH 5.5 (n = 3 biologically independent samples, scale bars = 100 nm). g Reaction progress of BGD in the presence of the CTSK enzyme monitored by LC-MS. The black arrow indicates the peak of the original molecule, and the blue arrow represents the peak of the product after enzymatic cleavage. h The remaining BGD after the enzymatic reaction. The half-life of enzymatic cleavage is 2.29 h. i, j Representative images of TRAP-positive cells in each group in the presence of M-CSF and RANKL. The black dotted circle represents damaged OCs. Scale bar = 400 μm. k Representative confocal images of F-actin in OCs of each group and representative scanning electron microscopy (SEM) images of the resorbed areas in bone slices of each group. Scale bars: 30 and 100 μm. j, l, m Quantification of the number of TRAP-positive OCs (> 3 nuclei) in each group (n = 5 biologically independent samples). k, n, o Quantification analyses of the mean number of nuclei in each OC and the resorbed area (n = 5 biologically independent samples). Data are expressed as mean ± SD. Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test. Source data are provided as a Source Data file.
To evaluate the CTSK-responsive properties of BGD, we examined its morphological changes in MES buffer (pH = 5.5) with and without CTSK. In the absence of CTSK, BGD formed amorphous aggregates when the pH reached to pH 5.5. However, upon addition of CTSK (800 ng/ml), BGD underwent enzymatic cleavage and self-assembled into nanofibers (Fig. 2e, f). Liquid chromatography-mass spectrometry (LC-MS) analysis confirmed the progressive degradation of intact BGD, with the emergence of a new peak corresponding to the cleavage product Bip-FFWKYKGHPG (Fig. 2g and Supplementary Fig. 6). The enzymatic cleavage exhibited a half-life of 2.29 h (Fig. 2h). Notably, the zeta potential of the BGD reaction mixture increased from −22.3 mV to +19.6 mV after CTSK-mediated cleavage, a shift that facilitates lysosomal membrane disruption28 (Supplementary Fig. 7). Moreover, in the presence and absence of CTSK enzyme, the zeta potential of the control peptide BAD is −9.5 and −14.3 mV, respectively (Supplementary Fig. 7).
In vitro inhibition of osteoclastogenesis and bone resorption
We first evaluated the cytotoxicity of BGD, BAD and WGD toward bone marrow-derived monocytes (BMMs) and RAW264.7. BGD showed no significant cytotoxicity against either BMMs and RAW264.7, while BAD demonstrated consistent cytocompatibility across all tested concentration (Supplementary Fig. 8). Since OCs differentiate in the bone marrow cavity, we tested the toxicity of BGD on other myeloid cells, including pre-OCs, BMSCs, and ATDC5 chondrocytes. The results showed that BGD did not show obvious toxicity to other cells, confirming its broad cytocompatibility (Supplementary Fig. 8). In addition, WGD showed no obvious cytotoxicity towards BMMs (Supplementary Fig. 9).
To assess the effect and selectivity of BGD on OCs formed by BMMs and RAW264.7, peptides were added on day 5 of RANKL induction to intervene in OCs (Fig. 2i). TRAP staining and cytotoxicity assay revealed that BGD significantly reduced the number of OCs in a concentration-dependent manners compared to the RANKL control group (Fig. 2j, l and Supplementary Fig. 8). In contrast, BAD showed no significant effect on OCs number (Fig. 2j, m and Supplementary Fig. 8), suggesting the key role of CTSK-responsive motif for OCs death selectively. At the same time, WGD also had no significant effect on OCs number, indicating the importance of the N-terminal self-assembly sequence Bip-FF (Supplementary Fig. 9). Moreover, BGD also had a corresponding effect on RAW264.7-derived OCs (Supplementary Fig. 10). Pretreatment with the CTSK inhibitor Balicatib29 greatly diminished BGD’s inhibitory effect on OCs (Fig. 2j, m and Supplementary Fig. 10), confirming that CTSK-mediated cleavage is essential for BGD’s activity. Cytoskeletal fluorescence staining demonstrated a dramatic reduction in F-actin ring size in the BGD-treated group compared to the RANKL and Balicatib-pretreated groups (Fig. 2k, n), indicating impaired OCs functionality. To further evaluate the impact of BGD on OCs-mediated bone resorption, we conducted a bovine bone resorption assay30. Scanning electron microscopy (SEM) revealed that the bone resorption area in the BGD group was significantly smaller than in the RANKL group (Fig. 2k, o). In order to study whether BGD affects the differentiation process of OCs, we intervened with BGD on days 1 and 3, and performed TRAP staining on day 6. Compared with the RANKL group, the TRAP staining results showed no difference in the number of OCs formed when BGD intervened on days 1 and 3 (Supplementary Fig. 11). In addition, we applied Balicatib to intervene in OCs on day 5 of RANKL induction. The results showed that the application of Balicatib at this stage did not inhibit OCs formation (Supplementary Fig. 12). These findings suggest that BGD can effectively induce OCs apoptosis but does not inhibit RANKL-induced OCs differentiation, with CTSK playing a central role in its mechanism of action.
BGD induces OCs apoptosis via lysosomal self-assembly
To elucidate the mechanism by which BGD induces OCs apoptosis, we performed a series of experiments. Immunofluorescence (IF) images revealed that BGD-Cy5.5 treatment for 24 h disrupted the podosome belt structure of OCs on both cell plates and bone slices (Fig. 3a). Red fluorescence in the extracellular space indicated peptide escape from lysosomes, accompanied by cytoskeletal damage and gradual cell disintegration. To explore the mechanism of nanoparticle endocytosis by OCs, we next used various endocytic inhibitors in co-incubation experiments. After the addition of Filipin III, the mean fluorescence intensity in OCs decreased from 1.27 to 0.045 (Fig. 3b, c), indicating that caveolae-mediated endocytosis contributes to the cellular uptake of BGD. In addition, ethylisopropyl amiloride (EIPA) and chlorpromazine (CPZ), inhibitors of macropinocytosis/clathrin-mediated endocytosis, caused the mean fluorescence intensity in OCs to decrease to 0.24 and 0.1, respectively (Fig. 3b, c). These results suggest that the uptake of BGD by OCs occurs through multiple endocytic pathways. The above effects of BGD were reversed by pretreatment with the CTSK inhibitor Balicatib, confirming the CTSK’s role in BGD’s activity. To further evaluate the effect of BGD on lysosomes, we used the lysosomal membrane marker Lamp1 and the lysosomal enzyme CTSK for IF experiments. In the RANKL group, Lamp1 and CTSK were fused, but after 24 h of intervention with BGD, the co-staining of Lamp1 and CTSK decreased, and OCs showed signs of disintegration and fission (Fig. 3d). Additionally, we found that 2 and 6 h after pretreatment of Balicatib, red fluorescence fused with green fluorescence, indicating that BGD was trapped in the lysosomes (Supplementary Fig. 13), while the cytoskeleton remained relatively intact.
a The representative confocal images show the impaired podosome belt structure of OCs after treatment with BGD. Blue indicates nuclei; green indicates Lamp1; cyan indicates F-actin; red indicates Cy5.5-labeled BGD. White arrows indicate the impaired podosome belt structure, and yellow arrows represent BGD escaping from lysosomes (n = 3 biologically independent samples, Scale bar = 100 μm). b Representative confocal images of BGD uptake by OCs after intervention with different endocytosis inhibitors. Red indicates Cy5.5-labeled BGD. White dotted circles indicate OCs. Scale bar = 100 μm. c Quantification of the mean fluorescence intensity (MFI) of each OC (n = 3 biologically independent samples). d Representative confocal images of OCs treated with BGD. Blue indicates nuclei; green indicates Lamp1; red indicates CTSK. The blue dotted circle represents OCs (n = 3 biologically independent samples, Scale bar = 50 μm). e Representative Bio-EM images of OCs following treatment with BGD for different times. Scale bars: 5 μm, 2 μm, 1 μm, and 500 nm. White arrows indicate the endocytosis process, and yellow arrows highlight the distribution of peptide within endosome. Red arrows denote the disruption of lysosomal membrane, and blue arrows represent self-assembled nanofibers (n = 3 biologically independent samples). f Representative confocal images of apoptosis and necrosis staining of OCs with BGD treatment for different times. Blue indicates healthy cells; green indicates necrotic cells; red indicates apoptotic cells (n = 3 biologically independent samples, Scale bar = 50 μm). g Representative confocal images of Tunel staining of OCs treated with BGD. Blue indicates healthy cells; green indicates apoptotic cells; red indicates F-actin. Scale bars = 100 μm. h Protein expression of cleaved caspase-3 in OCs after BGD intervention. i, j Quantification of the percentage of Tunel-positive nuclei (n = 5 biologically independent samples) and the expression level of cleaved caspase-3 after treatment with BGD (n = 3 biologically independent samples). Data are expressed as mean ± SD. Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test. Source data are provided as a Source Data file.
Most cells preferentially phagocytose positively charged particles, but macrophages, including OCs, exhibit strong uptake of negatively charged particles due to their high phagocytic capacity31. To evaluate this, we tested the uptake of BGD by three myeloid cells (BMMs, BMSCs, and ATDC5). Fluorescence imaging confirmed that negatively charged BGD was rarely phagocytosed by these cells (Supplementary Fig. 14), highlighting its selective targeting of OCs. To further investigate the role of surface charge, we replaced the C-terminal hexa-aspartic acid (BGD) with hexa-lysine (BGK) or hexa-asparagine (BGN). BGK formed positively charged assemblies (zeta potential: +11.2 mV), while BGN maintained a negative charge (−26.4 mV) (Supplementary Fig. 15). Cytotoxicity assays revealed that BGN was biocompatible with BMMs, whereas BGK exhibited cytotoxicity at 32 μM (Supplementary Fig. 15). These results suggest that negatively charged peptide assemblies, like BGD, are better suited for targeting multinucleated macrophages such as OCs.
Bio-EM provided direct insights into the intracellular behavior of BGD (Fig. 3e). Within the first 10 min, BGD was observed entering the OCs, evidenced by the accumulation of peptide assemblies on cell membrane and their sparse distribution within developing vesicles. After 2 h, the assemblies were more widely distributed in the endosomes. By 4 h, a pronounced concentration of assemblies within the endosomes was noted, indicated by the darkening of the endosomal structures, alongside early signs of lysosomal integrity damage. After 24 h, extensive lysosomal disruption and nanofiber formation were observed both inside and outside lysosomes.
Lysosomal enzyme leakage into the cytoplasm triggers cell death32. Using an Apoptosis/Necrosis Assay Kit, we observed that BGD induced OC apoptosis (red fluorescence) with minimal necrosis (green fluorescence) (Fig. 3f). Apoptotic features, including membrane blebbing and cell disintegration, were confirmed by white light microscopy (Supplementary Fig. 16). Tunel staining and WB further validated apoptosis, showing increased cleaved Caspase-3 expression after 6 h, which peaked at 24 h (Fig. 3g–j). Compared with the Tunel staining positive control group (ALD), almost only OCs underwent apoptosis in the BGD group, while a large number of OCs and pre-OCs underwent apoptosis in the ALD group (Supplementary Fig. 17). Balicatib pretreatment significantly reduced apoptosis, underscoring the role of CTSK in BGD’s mechanism. To further verify the importance of caspase 3 in BGD-induced OCs apoptosis, we subsequently performed siRNA experiments. OCs were transfected with 10 nM siRNA (caspase 3) for 6 h, and the knockout efficiency was verified by WB, followed by TRAP and Tunel staining. WB showed that 10 nM siRNA could significantly inhibit the expression of caspase 3 protein. In addition, TRAP and Tunel staining showed that the number of BGD-induced apoptosis was significantly reduced after OCs were pretreated with siRNA (Supplementary Fig. 18). This series of experiments confirmed that caspase 3 can play an important role in the process of BGD-induced OCs apoptosis.
BGD-derived ABs promote osteogenic differentiation via PDGF-BB-mediated PI3K/AKT signaling pathway
A large number of ABs are produced during cell apoptosis, and ABs of OCs have the function of stimulating the osteogenic differentiation of BMSCs16. Therefore, we collected ABs of OCs induced by BGD for subsequent experiments. To investigate the biological functions of OC-derived ABs, we collected ABs induced by BGD and the positive control Alendronate (ALD)17, and examine their effects on the differentiation of BMSCs into osteoblasts (Fig. 4a). ABs were identified using Annexin-V/FITC staining and TEM. Confocal microscopy revealed that both BGD-ABs and ALD-ABs were spherical, single-membrane vesicles positive for Annexin-V (Fig. 4b and Supplementary Fig. 19). TEM confirmed the spherical structure of ABs, and DLS showed that the diameters of ABs mostly ranged from 700 nm to 5 µm (Fig. 4c and Supplementary Fig. 20). Flow cytometry further assessed ABs purity, showing distinct size differences between ABs and cells (Fig. 4d). Annexin-V/FITC staining indicated significant phosphatidylserine (PS) exposure in ABs compared to apoptotic cells, while live cells showed minimal PS exposure. Propidium iodide (PI) staining revealed a small proportion of PI-positive ABs, suggesting the presence of nucleic acids in a minor fraction of ABs.
a Schematic diagram of the induction and isolation process of ABs. b Representative confocal image of BGD-ABs stained with Annexin-V/FITC (n = 3 biologically independent samples, Scale bar = 3 μm). c TEM image of BGD-ABs (n = 3 biologically independent samples, Scale bar = 500 nm). d The FSC/SSC analysis, along with Annexin-V/PI staining, was performed to assess live cells, apoptotic cells, and BGD-ABs. e Volcano plots showing the DEPs in the ABs of ALD and BGD groups. The red dots represent DUPs (FC > 2.0), and the blue dots represent DDPs (FC < 0.5). Differential expression was analyzed using a two-sided t-test with Benjamini-Hochberg correction. Among the proteins, PDGF-BB expression levels were significantly upregulated in BGD-ABs compared with ALD-ABs. f KEGG pathway enrichment analysis for BGD-ABs versus ALD-ABs group. Enrichment analysis of differentially expressed genes in KEGG pathways was performed through the hypergeometric test with one-sided test and Benjamini-Hochberg adjustment. g Heat map of DEPs involved in the PI3K-Akt signaling pathway, in which PDGF-BB plays an important role. h BMSCs labeled with Cell Tracker Red were cultured with BGD-ABs labeled with Annexin V/FITC (n = 3 biologically independent samples, Scale bar = 50 μm). i BMSCs were stained with ALP and ARS after being treated with media containing BGD-ABs or ALD-ABs. j, k Quantification of the ALP and ARS activity after treatment with BGD-ABs and ALD-ABs (n = 3 biologically independent samples). l, m WB was used to analyze the expression of ALP, Runx2, p-PI3K, and p-AKT in BMSCs after treatment with BGD-ABs and ALD-ABs. Data are expressed as mean ± SD. Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test. Source data are provided as a Source Data file.
To explore the proteomic composition of BGD-ABs and ALD-ABs, we performed liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis. The resulting mass spectra were processed using Proteome Discoverer 2.5, with database searches against the Mus musculus proteome UniProtKB (UP000000589). A total of 41,135 peptides and 5470 proteins were identified. Hierarchical clustering analysis of the proteomic profiles was visualized in heatmaps (Supplementary Fig. 21). Differentially expressed proteins (DEPs) with statistical significance (P < 0.05) were categorized into up-regulated (FC > 2.0) and down-regulated (FC < 0.5) groups, and visualized and identified in volcano plots. Notably, PDGF-BB was identified as a key component of BGD-ABs, with minimal expression in ALD-ABs (Fig. 4e). Functional annotation of differentially expressed genes (DEGs) using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database revealed enrichment in pathways such as metabolic pathways, phagosome, carbon metabolism, PI3K-Akt signaling, and oxidative phosphorylation (Fig. 4f). Screening for DEPs associated with the PI3K/AKT pathway highlighted PDGF-BB as a distinctly altered protein (Fig. 4g). WB and enzyme-linked immunosorbent assay (ELISA) experiments further confirmed that ABs in the BGD and ALD groups co-expressed PDGF-BB protein, and the PDGF-BB level in the BGD group was significantly higher than that in the ALD group (Supplementary Fig. 20). Previous studies have shown that PDGF-BB activates the PI3K/AKT/mTOR pathway, promoting osteogenic differentiation and bone repair33,34,35. Thus, we hypothesized that PDGF-BB-enriched BGD-ABs activate the PI3K/AKT signaling pathway to enhance BMSCs osteogenesis.
To confirm the role of ABs in osteogenic differentiation, BGD-ABs labeled with Annexin-V/FITC were co-cultured with BMSCs stained with Cell Tracker Red. Confocal microscopy revealed time-dependent internalization of ABs by BMSCs (Fig. 4h). Alizarin Red S (ARS) and alkaline phosphatase (ALP) staining demonstrated that both BGD-ABs and ALD-ABs induced osteogenic differentiation, with BGD-ABs exhibiting significantly higher osteogenic potential (Fig. 4i–k). This difference may be attributed to the higher PDGF-BB content in BGD-ABs. WB analysis confirmed elevated expression of osteogenic markers, including Runx2 and ALP, in BMSCs treated with BGD-ABs compared to ALD-ABs (Fig. 4l). Additionally, BGD-ABs significantly activated the PI3K/AKT pathway, as evidenced by increased phosphorylation of AKT (Fig. 4m). Collectively, these finding demonstrate that BGD-ABs promote osteogenic differentiation of BMSCs through PDGF-BB-mediated activation of the PI3K/AKT signaling pathway, highlighting their therapeutic potential for bone regeneration.
Bone targeting and anti-osteoporosis effects of BGD in vivo
After validating the selectivity of BGD in vitro, we assessed its bone-targeting ability and therapeutic efficacy in vivo. BGG, a control peptide replacing the hexa-aspartic acid bone-targeting motif with hexa-glycine, was used to evaluate the role of targeting sequence in bone homing. First, we evaluated the biosafety of BGD and BGG. Similar with the in vitro experiment of BGD, the in vitro experiments demonstrated the cytocompatibility of BGG (Supplementary Fig. 22). In vivo, short-term (24 h) intravenous administration of BGD and BGG caused no hemolysis or systemic inflammation (Supplementary Figs. 23, 24). Hemolysis tests revealed that both peptides induced a hemolytic rate of less than 5% at 800 μM, confirming their safety for systemic use.
To investigate bone targeting and the biodistribution of BGD nanoparticles, Cy5.5-labeled BGD and BGG were administered to ovariectomized (OVX) mice 28 days post-surgery (n = 3) (Fig. 5a). Mice euthanized at 6-, 24-, and 36-h post-injection, and tissues (spines, hindlimbs, hearts, livers, and kidneys) were harvested for fluorescence analysis (Fig. 5b). BGD exhibited significant accumulation in the hindlimbs and spine at 6 h, with sustained retention at 36 h, demonstrating robust bone-targeting ability. In contrast, BGG showed minimal bone accumulation due to the mutation of targeting residues. High fluorescence signal in the liver and kidneys suggested hepatic and renal clearance of the peptides. Quantitative analysis of fluorescence signals further confirmed that BGD had a highly efficient bone targeting ability compared with BGG (Fig. 5b). IF analysis of femur sections further confirmed BGD’s bone-targeting specificity. Co-staining with an anti-CTSK antibody to label OCs revealed greater overlap between green (CTSK) and red (Cy5.5) signals in the BGD group compared to BGG, particularly in the bone marrow and on trabecular surfaces (Fig. 5c). These findings demonstrated the bone targeting ability of our design.
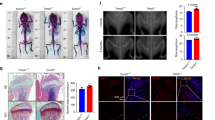
a Schematic diagram of in vivo targeted experiment process. b Representative images depict the biodistribution of BGD and BGG in the heart, liver, kidneys, hindlimbs, and spine at 6, 24, and 36 h after intravenous injection. ROI, region of interest (n = 3 biologically independent samples). c The biodistribution of BGD and BGG in the femur at 6 h after intravenous injection (n = 3 biologically independent samples, Scale bars: 100 and 50 μm). TB, trabecular bone. Red, Cy5.5 labeled BGD and BGG; green, CTSK-positive OCs; blue, DAPI. d Schematic diagram of animal treatment experiment process. e Micro-CT evaluation of the bone masses of the femur in each group. f–j Quantitative measurements of bone microstructure-related parameters: BV/TV, BMD, Tb.N, Tb.Sp, and Tb.Th, in each group (n = 4 biologically independent samples). k H&E and TRAP staining images of the femur in each group. Scale bars, 500 and 100 μm. l, m Quantification of the BV/TV, and N.oc/BS values in each group (n = 5 biologically independent samples). Data are expressed as mean ± SD. Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test. Source data are provided as a Source Data file.
To evaluate the therapeutic potential of BGD, OVX mice were treated with BGD, BGG, or ALD starting at week 5 post-surgery. BGD and BGG were administered intravenously every three days for four weeks, while ALD was injected intraperitoneally (Fig. 5d). Eight weeks post-OVX, mice were euthanized, and femurs, livers, spleens, and kidneys were harvested for analysis. Micro-CT imaging revealed that BGD and ALD significantly improved bone mineral density (BMD), bone volume/total volume (BV/TV), trabecular thickness (Tb.Th), and trabecular number (Tb.N), while reducing trabecular separation (Tb.Sp) compared to the OVX and BGG groups (Fig. 5e–j). Notably, BGD outperformed ALD in all parameters, indicating its superior efficacy in delaying bone loss. BGG showed minimal anti-osteoporotic effects, consistent with its poor bone-targeting ability. Histological analysis of femur sections stained with hematoxylin and eosin (H&E) and tartrate-resistant acid phosphatase (TRAP) confirmed these findings. The OVX group exhibited significant bone loss and increased OC numbers, while BGD treatment effectively reversed these trends (Fig. 5k). Quantitative analysis showed that BV/TV reached 26.74% in the BGD group, compared to 20.92% (ALD), 14.16% (BGG), and 11.08% (OVX) (Fig. 5l). Similarly, the number of OCs per bone surface (N.oc/BS) was significantly lower in the BGD group (3.74%) than in the OVX (7.34%) group (Fig. 5m). H&E staining of the liver, spleen, and kidneys confirmed the biosafety of BGD and BGG after long-term (4-week) intravenous administration (Supplementary Fig. 25). Additionally, no systemic inflammation or abnormalities in liver and kidney function were observed (Supplementary Figs. 26, 27). These results demonstrate that BGD exhibits efficient bone-targeting ability and robust anti-osteoporotic effects in vivo, outperforming ALD in restoring bone mass and structure. Its biosafety profile further supports its potential as a therapeutic agent for osteoporosis.
BGD regulates bone homeostasis in vivo
To investigate the regulatory effects of BGD on bone homeostasis in vivo, OVX mice were treated with BGD, BGG, and ALD at week 5 post-surgery10. To assess the rate of new bone deposition, intraperitoneal injections of calcitonin and alizarin red were administered to label the bone formation lines. As illustrated in Fig. 6a, the BGD group exhibited the greatest distance between the two calcium deposition lines, indicating enhanced bone formation. Quantitative analysis of the mineral apposition rate (MAR) confirmed that BGD promoted a significantly faster rate of bone deposition compared to the other groups (Fig. 6d). Von Kossa staining revealed a marked increase in the number of mineralized bone trabeculae and thicker trabeculae in the BGD group compared to the OVX, BGG, and ALD groups (Fig. 6b). Quantitative analysis of the mineralized area demonstrated that calcium deposition was significantly higher in the BGD group (Fig. 6e). Masson’s trichrome staining further supported these findings, showing a substantial presence of newly formed woven bone in the BGD group (Fig. 6b). Quantitative analysis indicated that collagen volume reached 9.19% in the BGD group, compared to 6.75% (ALD), 4.88% (BGG), and 3.23% (OVX) (Fig. 6f). Immunohistochemistry (IHC) staining showed that the positive expression of p-AKT, Runx2, ALP, and VEGF was the strongest in the BGD group, indicating that both the osteogenic and vascular coupling systems were activated (Fig. 6c, g–i and Supplementary Fig. 28). In contrast, the BGD group exhibited the lowest expression of CTSK, a marker of osteoclast activity, suggesting significant inhibition of bone resorption (Supplementary Fig. 29).
a Alizarin red (red) S and calcein blue (green) fluorescent labeling indicating the width of bone mineral attachment (distance between yellow arrows). TB: trabecular bone. Scale bars = 20 μm. b Von Kossa and Masson’s trichrome staining images of the femur in each group. Scale bars, 500 and 100 μm. c IHC staining for Runx2, p-AKT, and ALP of the femur in each group. Scale bars = 100 μm. d–f The MAR (n = 3 biologically independent samples), the formation of mineralized bone nodules (n = 3 biologically independent samples), and collagen volume were quantified (n = 5 biologically independent samples). g–i Quantitative analysis of the intensity of immunostaining for Runx2, p-AKT, and ALP (n = 5 biologically independent samples). Data are expressed as mean ± SD. Statistical analysis was performed using one-way ANOVA with Tukey’s post hoc test. Source data are provided as a Source Data file.
Discussion
This study introduces OsteoSAVE, a bone-targeted nanoparticle system that leverages enzyme-responsive self-assembly to selectively eliminate mature OCs while in vivo generating pro-osteogenic ABs, thereby restoring bone homeostasis in osteoporosis. Unlike conventional antiresorptives (e.g., bisphosphonates), which indiscriminately inhibit OC lineage cells and disrupt physiological bone remodeling5,7, OsteoSAVE combines precision targeting of pathological bone resorption with ABs-mediated bone formation-a dual mechanism that addresses a critical unmet need in osteoporosis therapy.
OsteoSAVE exploits the overexpression of CTSK, a lysosomal protease central to OCs-mediated bone resorption, to achieve cellular specificity. The nanoparticles’ CTSK-cleavable backbone triggers their transformation into membrane-disruptive nanofibers within mature OC lysosomes, inducing apoptosis while sparing pre-OCs. This specificity is further enhanced by OsteoSAVE’s negative surface charge, which minimizes uptake by non-target cells (e.g., BMSCs, ATDC5) but enables selective phagocytosis by multinucleated OCs. Such spatial control over therapeutic activity preserves pre-OC communication with osteoblasts, maintaining physiological remodeling-a key limitation of current therapies.
We demonstrate that OsteoSAVE-generated ABs, enriched with PDGF-BB, activate the PI3K/AKT pathway in BMSCs, driving osteogenic differentiation and bone formation. Compared to ABs induced by ALD, OsteoSAVE-derived ABs exhibit superior osteogenic potential due to their higher PDGF-BB content, a finding supported by proteomic analyses, molecular biological studies, and functional assays. The abundance of PDGF-BB in OsteoSAVE-derived ABs may be attributed to the fact that intracellular PDGF-BB is primarily degraded in lysosomes. Since BGD can damage lysosomes, this leads to reduced degradation of PDGF-BB, which is then released along with the ABs. Compared to ABs induced by ALD, OsteoSAVE-derived ABs exhibit superior osteogenic potential due to their higher PDGF-BB content, a finding supported by proteomic analyses, molecular biological studies, and functional assays. The reason why OsteoSAVE-derived ABs was rich in PDGF-BB may be that intracellular PDGF-BB is mainly degraded in lysosomes36, while BGD can damage lysosomes, resulting in reduced PDGF-BB degradation, which is then released with ABs. This contrasts with prior strategies using non-selective pharmacological agents (e.g., staurosporine) to generate ABs in vitro, which lack translational applicability and disrupt OC lineage cells indiscriminately. Thus, unlike prior strategies relying on exogenous AB administration or non-selective in vitro induction, OsteoSAVE nanoparticles exploit the pathological bone microenvironment to generate ABs directly at the site of imbalance. This approach harnesses the body’s natural apoptotic machinery to deliver osteogenic signals (e.g., PDGF-BB), creating a self-reinforcing cycle of resorption inhibition and bone formation. In OVX mice-a model mimicking postmenopausal osteoporosis-intravenous OsteoSAVE administration achieved robust bone targeting, significantly improving bone mineral density, trabecular thickness, and bone volume while reducing OC numbers. Crucially, OsteoSAVE showed no systemic toxicity, with no hemolysis, organ damage, or inflammatory responses observed after short- or long-term use. Its bone-specific accumulation and clearance via hepatic/renal pathways further underscore its clinical translatability. Recent articles reported that in osteoarthritis, OCs-ABs promote subchondral bone reconstruction through the RANKL reverse signaling pathway, aggravating osteoarthritis37. In bone tumor diseases, OCs-ABs exhibit an immunosuppressive function and promote secondary metastasis of breast cancer38. The above studies suggest that OC-derived ABs play a wide range of roles in bone-related diseases, with mechanisms of action that vary significantly, warranting further investigation in the future.
In summary, OsteoSAVE exemplifies the potential of enzyme-responsive biomaterials to recalibrate bone homeostasis through cell-selective targeting and engineered extracellular vesicle signaling. By coupling precision antiresorption with pro-osteogenic cues, this platform offers a regenerative alternative to traditional osteoporosis therapies, with broad implications for treating age-related skeletal disorders.
Methods
Materials
RAW264.7 cell was kindly provided by Stem Cell Bank, Chinese Academy of Sciences (TCM13). ATDC5 chondrogenic cell was obtained from Sigma-Aldrich (99072806-1VL). The following materials were used: Fetal bovine serum (FBS) (Gibco, USA), α-minimal essential medium (α-MEM) (Gibco, USA), recombinant receptor activator of nuclear factor κB ligand (RANKL) (R&D Systems, USA), macrophage colony-stimulating factor (M-CSF) (R&D Systems, USA), Calcein (Sigma, USA), Alizarin Red S (Sigma, USA), Sterile bone slices (IDS, UK), β-glycerophosphate (Aladdin, China), L-ascorbic acid (Aladdin, China). ActinRed™ 555 ReadyProbes™ Reagent and Cell Tracker Red were obtained from Thermo (San Jose, USA). Recombinant human CTSK (219461) were purchased from Merck (China). CTSK inhibitor Balicatib (HY-15100) and Alendronate (ALD) (HY-11101) were purchased from MCE (Shanghai, China). Staining antibodies and kits were summarized in Supplementary Tables 1, 2.
Peptide synthesis
Peptides (Table. S3) were synthesized via Fmoc-based solid-phase peptide synthesis (SPPS) employing a microwave peptide synthesizer (Liberty Blue 2.0, CEM Corporation) as previously described39. During SPPS, the N-terminus was conjugated with biphenyl using 4-biphenylacetic acid (catalog no. 196487, Sigma-Aldrich). Crude peptides were cleaved from 2-chlorotrityl chloride resin for 1 h using a cleavage reagent (trifluoroacetic acid (TFA): triisopropylsilane (TIS): H2O = 95%: 2.5%: 2.5) and then concentrated with ice-cold ether. Purification was carried out using high-performance liquid chromatography (HPLC) on an Agilent 1260 Infinity II Manual Preparative system, utilizing a mobile phase consisting of CH3CN and water, each containing 0.1% TFA. The purified peptide was subjected to lyophilization and subsequently analyzed using mass spectrometry (MS) on an Agilent InfinityLab LC/MSD system, with the MSD detector configured to positive ion mode. MALDI TOF of each peptide was obtained on Bruker’s FLEX series MALDI-TOF systems.
Material characterization
The critical micelle concentration (CMC) of the peptides was evaluated with a spectrofluorometer (Edinburgh Instruments, F55). Initially, BGD, BAD, BGG, and WGD peptides were dissolved in phosphate-buffered saline (PBS) solution at a concentration of 2.56 mM and then diluted eight times two-fold. After adjusting the pH of the solutions to 7, 100 μL of the peptide solution was combined with 2 μL of rhodamine 6 G and incubated for 24 h to determine the CMC. The CMC was calculated by measuring the wavelength that showed the maximum optical density (OD) value within the 515 to 560 nm range. Moreover, the average size and ζ-potential of the peptides at the specified concentrations were determined using a Zetasizer (Malvern Instruments, U.K.).
To study the morphology of the peptide assemblies, a Talos L120C G2 transmission electron microscope (TEM) from Thermo Fisher Scientific was used. For sample preparation, BGD, BAD, and BGG peptides were dissolved in PBS solution at a concentration of 64 μM and incubated for 8 h. Following incubation, 10 μL of the peptide suspension was deposited onto 200-mesh copper grids and subsequently stained with uranyl acetate for 2 min. After allowing the grids to dry, they were examined under the TEM for morphological analysis.
In vitro study of enzymolysis
Based on previous studies, in vitro enzyme cleavage assays were conducted40,41. BGD (64 μM) was incubated with CTSK (800 ng/ml) in MES buffer (50 mM, pH 5.5), supplemented with 0.1% DMSO, 2 mM ethylene diamine tetraacetic acid (EDTA), 4 mM dithiothreitol (DTT) at 37 °C. The ζ-potential of BGD and BAD in the reaction mixture was determined using a Zetasizer (Malvern Instruments, U.K.). At specified time intervals, 100 μL of the reaction mixture was collected. To stop the enzymatic reaction, 100 μL of methanol was added to each sample. The samples were then analyzed using Liquid Chromatography-Mass Spectrometry (LC-MS). During the LC-MS analysis, peaks corresponding to both the original molecule (the compound before conversion) and the product (the compound of the conversion reaction) were identified. The ratio of remaining peptide was calculated by integrating the areas under the respective peaks for the original molecule and product at each time point.
Cells culture
All animal experiments received approval from the Animal Experimental Ethics Committee at Zhejiang Chinese Medicine University (IACUC-20230731-01). BMMs were isolated from the tibiae and femora of 6-week-old mice, then incubated with α-MEM complete medium supplemented with M-CSF (20 ng/ml). After 48 h, the medium was changed, leaving adherent BMMs. Similarly, BMSCs were harvested from the tibia and femur, expanded in α-MEM complete medium for subsequent experiments.
Cytotoxicity Assay
To test the toxicity of each peptide on cells within the bone marrow cavity, including BMMs, pre-OCs, OCs, BMSCs, and ATDC5 chondrocytes, CCK-8 assays were performed according to the instructions provided by the manufacturer. Cells (104 cells per well) were seeded into 96-well plates and incubated overnight. Afterwards, the medium was replaced with the medium containing the indicated peptides concentrations and incubation for 24 h. Following this, 10 μL of CCK-8 reagent was introduced to each well, followed by a 4-h incubation at 37 °C, after which absorbance was recorded at 450 nm using a multimode plate reader.
OCs differentiation assay
BMMs, at a density of 104 cells per well, were incubated overnight in a 96-well plate using α-MEM complete medium with 20 ng/ml of M-CSF (Day 0). On day 1, the cells were stimulated with RANKL (50 ng/ml), and OCs typically began to form on day 5 after RANKL stimulation (the medium was replaced every 2 days). In order to study whether BGD affects the differentiation process of OCs, we intervened with BGD on days 1 and 3, and performed TRAP staining on day 6. Additionally, we investigated whether Balicatib itself could affect the differentiation process of OCs by intervening with 20 nM Balicatib on day 5 and performing TRAP staining on day 6. To evaluate the ability of the peptides to induce OCs apoptosis, new culture medium containing 64 μM BGD, BAD, and WGD was added on day 5, and performed TRAP staining on day 6. In addition, OCs in the CTSK inhibitor pretreatment group were first treated with Balicatib (20 nM) for 12 h, followed by peptide intervention experiments. Following incubation, the cells were fixed with 4% paraformaldehyde for 20 min, after which OCs were identified using TRAP staining. TRAP-positive multinucleated cells (with more than three nuclei) were counted and considered to be mature OCs. For immunofluorescence (IF) staining of F-actin, the ActinRed™ 555 ReadyProbes™ Reagent was applied. A confocal fluorescence microscope (Leica, DMi8, Germany) was used to capture images.
The siRNA specific to Caspase 3, and the negative control siRNA were custom-made by TsingkeBio, China (Table. S4). For siRNA transfection in OCs, BMMs (106 cells per well) were cultured in a 6-well plate with 20 ng/ml M-CSF (Day 0). On day 1, the cells were stimulated with RANKL (50 ng/ml), and OCs typically began to form on day 5 after RANKL stimulation (the medium was replaced every 2 days). Briefly, cells were washed with PBS after OCs formation and then transfected with siRNA/Lipofectamine complex. Specifically, siRNA and Lipofectamine (Thermo Fisher Scientific, MA, USA) were first diluted in optiMEM (Gibco, USA) to a final concentration of 10 nM and allowed to stand at room temperature for 20 min to form complexes. Cells were then treated with 10 nM siRNA for 6 h. After 6 h, cells were treated with fresh medium containing FBS, M-CSF, and RANKL for 24 h. After 24 h, transfection efficiency was detected by WB. In addition, we investigated whether siRNA pre-treatment would affect the ability of BGD to induce OCs apoptosis. Briefly, we first pre-treated cells with siRNA on day 5 of RANKL-induced OCs, treated cells with BGD on day 6, and finally performed TRAP staining.
In addition, RAW264.7 (5 × 103 cells/well) were incubated overnight in a 96-well plate using α-MEM complete medium (Day 0). On day 1, the cells were stimulated with RANKL (50 ng/ml), and OCs typically began to form on day 5 after RANKL stimulation (the medium was replaced every 2 days). To evaluate the ability of BGD induces OCs apoptosis, new culture medium containing BGD was added on day 5, and TRAP staining was performed and counted on day 6.
Bone slice resorption assay
Sterile bone slices (DT-1BON1000-96) were employed in the pit formation assay. First, these bone slices were carefully placed on the bottom of a 96-well plate, and then BMMs (104 cells per well) were seeded on top of them and incubated with 20 ng/ml M-CSF overnight. To induce the formation of OCs, RANKL at a concentration of 50 ng/ml was added, and the medium was refreshed every two days. On the fifth day, OCs were exposed to 64 μM of either BGD or BAD, along with RANKL and Balicatib, for an additional 48 h. On day 7, the bone slices were first digested and then brushed to remove the remaining cells. Scanning electron microscopy (SEM) was utilized to observe the bone resorption pits on bone slices. Subsequently, ImageJ was employed to quantify the bone resorption pits in each slice. At least 3 slices were quantified per group.
Cellular uptake
To analyze the cellular distribution of BGD within OCs, BMMs (104 cells per well) were placed in a 96-well plate and subjected to M-CSF and RANKL stimulation for 5 days. On day 5, OCs were treated with Cy5.5-labeled BGD (16 μM) in the presence of absence of Balicatib for different times. Then, cells were next fixed with 4% PFA and stained with a primary LAMP1 antibody at a 1:200 dilution, as well as F-actin. Following this, a goat anti-rabbit Alexa Fluor 488 secondary antibody, diluted 1:1000, was used for one h at room temperature in the dark. After being stained with 4′,6-Diamidino-2-phenylindole (DAPI), the cells were visualized using a confocal fluorescence microscope (Leica, DMi8, Germany). Moreover, we used Cy5.5-labeled BGD to intervene in BMMs, BMSCs and ATDC5 and observed the uptake of various cells.
The method to evaluate the endocytosis mode was previously described39. BMMs, at a density of 104 cells per well, were incubated overnight in a 96-well plate using α-MEM medium with 20 ng/ml of M-CSF (Day 0). On day 1, the cells were stimulated with RANKL (50 ng/mL), and OCs typically began to form on day 5 after RANKL stimulation (the medium was replaced every 2 days). Briefly, various endocytosis inhibitors such as MβCD, Amiloride, EIPA (ethyl-isopropyl-amiloride), CPZ (chlorpromazine), and Filipin III were pre-incubated for 30 min. Subsequently, fresh medium containing Cy5.5-labeled BGD was added to each well to a final concentration of 16 μM, and fluorescence images were taken after 30 min of incubation. The mean fluorescence intensity (MFI) of each OC was calculated.
The Bio-EM (Talos L120C G2 transmission electron microscope) was employed to observe the endocytosis and cytoplasmic release of the peptide. OCs were induced from BMMs (106 cells per well) in 6-well plates using RANKL and M-CSF stimulation, followed by incubation with BGD (64 μM). At the indicated time points, cells were collected with 900 μL PBS and 100 μL FBS. The resulting cell suspension was centrifuged at 400 g for 10 min. The obtained pellet was fixed in a fixation buffer, followed by embedding, staining, and imaging procedures.
OCs apoptosis detection
On day 5 of RANKL stimulation of BMMs, the medium was changed to fresh medium containing 64 μM BGD in the presence and absence of Balicatib or siRNA pre-treatment and culture continued for the indicated time points. In the apoptosis experiment, ALD (500 μM)-induced OCs apoptosis was used as the positive control group. To observe the intracellular distribution of CTSK, cells were fixed and washed and stained with LAMP1 (dilution 1:200), CTSK (dilution 1:200), and DAPI. Next, secondary antibodies goat anti-rabbit Alexa Fluor 488 and goat anti-rat Alexa Fluor 555, both diluted at 1:1000, were applied for one h at 37 °C in the dark. Finally, CLSM imaging was performed. To observe the phenotype of cellular death, cells was stained for apoptosis and necrosis with the Apoptosis/Necrosis Detection Kit. To observe the apoptosis, cells were stained for apoptosis with the Tunel apoptosis assay kit before CLSM imaging (Leica, DMi8, German).
To form OCs, BMMs (106 cells per well) were cultured in a 6-well plate and treated with RANKL and M-CSF for a duration of 5 days. OCs were exposed to 64 μM BGD for specified durations. The cells were subsequently lysed with radioimmunoprecipitation (RIPA) lysis buffer to extract the proteins, followed by WB experiment to detect the expression of cleaved-caspase 3.
Extraction and identification of ABs
To trigger apoptosis in OCs, they were treated with 64 μM BGD for 24 h at 37 °C, using ALD (500 μM) as a positive control. Following the incubation, the supernatants from the cells were collected and centrifuged at 300 g for 15 min to eliminate cell debris. The supernatant obtained was then centrifuged again at 3000 g for 20 min, and the ABs were collected for further analysis. Using the DLS (Malvern, UK) to measure the size distribution of BGD-induced ABs. ABs were characterized through staining with Annexin-V/FITC and examined using confocal microscopy. For flow cytometry, the ABs were processed using the Annexin V-FITC/PI kit following the manufacturer’s instructions. Briefly, live cells were the negative control and ALD induced apoptotic cells were the positive control. The amount of ABs was then analyzed and quantified using a flow cytometer analyzer (CytoFLEX LX, Beckman Colter). After quantification, ABs-media was generated by dissolving 5 × 106 ABs into 50 mL complete medium for further in vitro study16.
LC-MS/MS analysis of ABs
The protein concentrations in ABs generated from BGD or ALD were determined using a bicinchoninic acid (BCA) protein assay (Catalog No. 23225, Thermo Scientific). A quantity of 10 µg of total protein was reduced with 10 mM dithiothreitol at 60 °C for 30 min, followed by alkylation using 20 mM iodoacetamide at room temperature in the dark for 30 min. Purification was performed employing SP3 technology as described42. The purified proteins were digested with trypsin (Promega, Catalog No. V5280) using a 1:50 enzyme-to-protein ratio (w/w) at room temperature overnight. The resulting peptides were then lyophilized in preparation for LC-MS/MS analysis.
For LC-MS/MS analysis, peptide separation was performed using a 72 min gradient elution at a flow rate of 0.300 µL/min on a Thermo Vanquish Neo integrated nano-HPLC system. This system was directly interfaced with the Thermo Exploris 480 mass spectrometer. Separation was achieved using a custom-designed fused silica capillary column with an inner diameter of 75 µm and a length of 150 mm, packed with C-18 resin (300 Å, 3 µm, Varian, Lexington, MA). In mobile phase A, there was 0.1% formic acid dissolved in water, while mobile phase B included 80% acetonitrile along with 0.1% formic acid. Using Xcalibur 4.1 software, the mass spectrometer functioned in data-dependent acquisition mode with FAIMS set at CV = −45. A comprehensive mass spectrum was obtained using the Orbitrap (350–1800 m/z, 60,000 resolution), followed by MS/MS scans at 15,000 resolution, with a cycle time of 2 s and 30% normalized collision energy.
The analysis of mass spectra data was conducted with Proteome Discoverer 2.5, which allowed for searches in the Mus musculus (mouse) proteome database from UniProtKB (UP000000589). For the Sequest search, the parameters were adjusted to a precursor mass tolerance of 10 ppm, a fragment ion tolerance of 0.02 Da, and up to 2 internal cleavage sites were allowed. Cysteine alkylation was used as a constant modification, whereas methionine oxidation was considered a variable modification. A filtering threshold of 1% false discovery rate (FDR) was applied to peptide identifications. Additional details can be requested from the corresponding author.
ABs induce osteogenic differentiation of BMSCs in vitro
To assess the effect of BGD induced OCs-derived ABs on the osteogenesis of BMSCs, 5 × 103 BMSCs were incubated in 96-well plates. To study the phagocytosis of ABs by BMSCs, the BMSCs were marked with Cell Tracker Red for 30 min at 21 °C. Annexin V/FITC-labeled ABs were then co-incubated with these labeled BMSCs for 24 h. Afterward, the cells were washed and examined with confocal microscopy to confirm ABs uptake.
Secondly, osteogenic medium containing ABs was used to induce BMSCs osteogenic differentiation. Osteogenic medium was based on complete medium containing 10 mM β-glycerophosphate and 50 μg/mL L-ascorbic acid. When the cell confluence reaches more than 80%, basal medium was changed to ABs-containing osteogenic medium and induced osteogenic differentiation for 7 days and 14 days (medium was replaced every 2 days). Then, ALP activity was assessed using an ALP staining kit and calcium deposit was stained with Alizarin Red S (ARS) after 7 and 14 days of induction, respectively. Image J was used to measure the intensities of ALP and ARS staining. To detect the protein levels, 106 BMSCs were incubated in 6-well plates, then treated with ABs-containing osteogenic medium for 7 days. Then, the total proteins were harvested for WB detection.
WB and Elisa assay
The cells or ABs were washed with PBS, and then RIPA lysis buffer was added to extract proteins for subsequent WB and ELISA experiments. Protein levels of active + pro Caspase 3, pro Caspase 3, PDGF-BB, ALP, Runx2, PI3K, p-PI3K, AKT, p-AKT, β-actin, and Histone H3 were assessed using WB. The proteins used for WB experiments were first separated by SDS-PAGE electrophoresis, and then the proteins were transferred to a polyvinylidene fluoride membrane and blocked with skim milk. The membranes underwent overnight incubation with the primary antibodies mentioned above (1:1000) at 4 °C. Subsequently, they were washed and incubated with HRP-conjugated secondary antibodies (1:10000) for 2 h at room temperature. Super ECL Plus (Beyotime, China) was used to detect the chemiluminescence signals of the antibodies. Images were taken with an ImageQuant LAS 4000 (GE Healthcare, Silverwater, Australia) and analyzed with ImageJ software.
In addition, the PDGF-BB concentration in ABs was also determined by ELISA. Serum level of PDGF-BB were determined by ELISA using a mouse-specific ELISA kit according to the manufacturer’s instructions.
Hemolysis assay
To achieve a final concentration of 800 μM, BGD and BGG were dissolved in PBS and then subjected to three 2-fold dilutions. A tenfold dilution was carried out on 2 ml of freshly collected blood from a New Zealand rabbit using PBS. Subsequently, the RBC pellet was obtained through centrifugation and rinsed eight times with PBS prior to being incubated with varying concentrations of peptides for 3 h at 37 °C. For controls, water was set as positive and PBS as negative. The mixture was centrifuged at 1000 g for 10 min, and the supernatant’s absorbance at 570 nm was measured to assess the hemolysis rate.
Bone targeting fluorescence imaging experiment in vivo
Using an in vivo IVIS spectrum imaging system, the specific bone binding of BGD in vivo was confirmed. BGD-Cy5.5 and BGG-Cy5.5 were administered intravenously to the mice at a dose of 0.77 mg/kg via the tail vein. The spine, hindlimbs, heart, liver, and kidneys were collected and examined in vitro to study the distribution of BGD and BGG at 6, 24, and 36 h post-injection. Living Image software was used to quantify the fluorescence intensity. Additionally, bone tissue sections were created to assess the in vivo distribution of BGD 6 h post-injection. Further staining of femur sections with anti-CTSK (1:200) and DAPI was performed to visualize BGD colocalization with OCs, followed by CLSM imaging.
OVX mice model establishment
Animals were housed in the Laboratory Animal Center of Zhejiang Chinese Medical University. Food and water were available ad libitum. Mice were housed in a specific pathogen-free (SPF) environment with a 12-h light/dark cycle, a temperature controlled at 24 ± 2 °C, and a humidity controlled at 50–60%. Twelve-week-old female mice were anesthetized with an intraperitoneal injection of zoletil at a dose of 50 mg/kg, then underwent OVX. The sham group mice had only the skin and muscles cut without removing the ovaries. The mice were randomized and assigned to five groups (n = 5 per group): the sham group, the OVX group, the OVX with BGD group, the OVX with BGG group, and the OVX with ALD group. Four weeks after OVX surgery, mice in the BGD and BGG groups were administered a tail vein injection of peptides at 11.5 mg/kg every 3 days for 4 weeks, while mice in the ALD group were given intraperitoneal injections of ALD (10 mg/kg) as a positive control43. Additionally, 0.9% NaCl (NS) was injected into the tail veins of mice in the sham and OVX groups as a control. Following four weeks of medication, mice in each group were executed for major organs (liver, spleen, and kidney), peripheral blood, serum and bilateral femurs collection. Subsequently, blood counts and the functions of liver and kidney were analyzed. According to the manufacturer’s instructions, an ELISA was utilized to evaluate the serum levels of IL-1β, IL-6, TNF-α, and IL-10 using a mouse-specific ELISA kit.
Micro-CT scanning and analysis
The Skyscan 1176 micro-CT instrument (Bruker, Kontich, Belgium) was used to scan the separated femurs of each group with a resolution of 4000 × 2672 pixels and an isotropic voxel size of 10 mm. 3D images were reconstructed through NRecon and analyzed using CTAn software. The areas of 0.5 mm below the distal femoral growth plate and 1 mm in height were selected for quantitative analysis. Morphometric analysis was undertaken using bone mineral density (BMD), trabecular thickness (Tb. Th), trabecular number (Tb. N), trabecular separation (Tb. Sp), and trabecular bone volume/tissue volume (BV/TV).
Histology, immunohistochemistry, and fluorescence labeling
Fluorescent dual-labeling of the newly formed mineralization was performed in this study44. Briefly, Calcein (30 mg/kg) and Alizarin Red S (30 mg/kg) were administrated to the mice by intraperitoneal injection at 4- and 8-week post-surgery. The subsequent procedures included fixing, dehydrating, and sectioning the bone samples, using fluorescence microscopy to visualize new mineralization through fluorescent labeling, and then applying Von Kossa staining.
Then, other femurs were fixed and decalcified, dehydrated, and embedded. Subsequently, femur tissue sections with a thickness of 4 μm were stained using Masson, TRAP, hematoxylin and eosin (H&E), IF, and immunohistochemistry (IHC). For IHC staining, paraffin sections underwent incubation in 0.01 M citrate buffer for 4 h at 60 °C, and then inactivated in 3% H2O2 for 20 min. Next, sections were incubated with primary antibodies against Runx2 (1:200), ALP (1:200), p-AKT (1:200), CTSK (1:200), and VEGF (1:200) at 4 °C overnight. The corresponding secondary antibody was utilized for a period of 2 h at room temperature. Finally, hematoxylin was used as a counterstain after applying DAB staining. Two Veteran physicians were recruited to measure the number of OCs per bone surface area (N. oc/BS), the trabecular region, the collagen region, and the mean density of Runx2, ALP, p-AKT, CTSK, and VEGF. Quantification of positive staining was calculated using ImageJ software.
Acute and chronic toxicity assay in vivo
For chronic toxicity assay, major organs (liver, spleen, and kidney) were collected from mice treated with 4 weeks of BGD and BGG interventions in the animal experiments described above. Subsequently, the tissues were fixed and subjected to H&E staining. Serum concentrations of IL-1β, IL-6, TNF-α, and IL-10 were measured by ELISA. For acute toxicity assay, 12-week-old healthy mice were given a single injection of the peptides (11.5 mg/kg). 24 h later, the serum was executed and collected for ELISA testing.
Statistical analysis
Data were expressed as mean ± standard deviation (SD). Our results were analyzed in Graph pad Prism 7 using independent samples t-test or one-way ANOVA and Tukey’s post hoc test to make multiple comparisons. The experiments were repeated independently at least three times with similar results unless otherwise mentioned, and the micrographs were representative data from individual samples (the n was provided in figure legends). Significance levels were denoted as P < 0.05, P < 0.01, P < 0.001, and P < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data is available for Figures and Supplementary Figs. in the associated source data file. Source data are provided with this paper. The mass spectrometry proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD069800.
References
Reid, I. R. A broader strategy for osteoporosis interventions. Nat. Rev. Endocrinol. 16, 333–339 (2020).
Walker Marcella, D. & Shane, E. Postmenopausal Osteoporosis. N. Engl. J. Med. 389, 1979–1991 (2023).
Dovjak, P. et al. Prediction of Fragility Fractures and Mortality in a Cohort of Geriatric Patients. J. Cachexia, Sarcopenia Muscle 15, 2803–2814 (2024).
Rogers, M. J., Crockett, J. C., Coxon, F. P. & Mönkkönen, J. Biochemical and molecular mechanisms of action of bisphosphonates. Bone 49, 34–41 (2011).
Xie, H. et al. PDGF-BB secreted by preosteoclasts induces angiogenesis during coupling with osteogenesis. Nat. Med. 20, 1270–1278 (2014).
Black Dennis, M. et al. Atypical Femur Fracture Risk versus Fragility Fracture Prevention with Bisphosphonates. N. Engl. J. Med. 383, 743–753 (2020).
Armamento-Villareal, R., Napoli, N., Panwar, V. & Novack, D. Suppressed Bone Turnover during Alendronate Therapy for High-Turnover Osteoporosis. N. Engl. J. Med. 355, 2048–2050 (2006).
Boyle, W. J., Simonet, W. S. & Lacey, D. L. Osteoclast differentiation and activation. Nature 423, 337–342 (2003).
Tsukasaki, M. et al. Stepwise cell fate decision pathways during osteoclastogenesis at single-cell resolution. Nat. Metab. 2, 1382–1390 (2020).
Wang, Q. et al. Suppression of osteoclast multinucleation via a posttranscriptional regulation–based spatiotemporally selective delivery system. Sci. Adv. 8, eabn3333 (2022).
Teitelbaum, S. L. Bone Resorption by Osteoclasts. Science 289, 1504–1508 (2000).
Seeman, E. & Martin, T. J. Antiresorptive and anabolic agents in the prevention and reversal of bone fragility. Nat. Rev. Rheumatol. 15, 225–236 (2019).
Kasugai, S., Fujisawa, R., Waki, Y., Miyamoto, K. I. & Ohya, K. Selective drug delivery system to bone: small peptide (Asp)6 conjugation*. J. Bone Miner. Res. 15, 936–943 (2000).
Manolagas, S. C. Birth and death of bone cells: basic regulatory mechanisms and implications for the pathogenesis and treatment of osteoporosis*. Endocr. Rev. 21, 115–137 (2000).
Atkin-Smith, G. K. et al. A novel mechanism of generating extracellular vesicles during apoptosis via a beads-on-a-string membrane structure. Nat. Commun. 6, 7439 (2015).
Ma, Q. et al. Osteoclast-derived apoptotic bodies couple bone resorption and formation in bone remodeling. Bone Res. 9, 5 (2021).
Ma, Q. et al. Mature osteoclast–derived apoptotic bodies promote osteogenic differentiation via RANKL-mediated reverse signaling. J. Biol. Chem. 294, 11240–11247 (2019).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Gao, Y., Shi, J., Yuan, D. & Xu, B. Imaging enzyme-triggered self-assembly of small molecules inside live cells. Nat. Commun. 3, 1033 (2012).
Kim, S. et al. Supramolecular senolytics via intracellular oligomerization of peptides in response to elevated reactive oxygen species levels in aging cells. J. Am. Chem. Soc. 145, 21991–22008 (2023).
Feng, Z., Zhang, T., Wang, H. & Xu, B. Supramolecular catalysis and dynamic assemblies for medicine. Chem. Soc. Rev. 46, 6470–6479 (2017).
Yuan, Y. & Liang, G. A biocompatible, highly efficient click reaction and its applications. Org. Biomol. Chem. 12, 865–871 (2014).
Schnaider, L. et al. Self-assembling dipeptide antibacterial nanostructures with membrane disrupting activity. Nat. Commun. 8, 1365 (2017).
Smith, A. M. et al. Fmoc-diphenylalanine self assembles to a hydrogel via a novel architecture based on π–π interlocked β-sheets. Adv. Mater. 20, 37–41 (2008).
Chen, D. et al. Inhalable SPRAY nanoparticles by modular peptide assemblies reverse alveolar inflammation in lethal Gram-negative bacteria infection. Sci. Adv. 10, eado1749 (2024).
Lecaille, F., Weidauer, E., Juliano, M. A., Brömme, D. & Lalmanach, G. Probing cathepsin K activity with a selective substrate spanning its active site. Biochem J. 375, 307–312 (2003).
Zhu, Y. et al. Negative surface shielded polymeric micelles with colloidal stability for intracellular endosomal/lysosomal escape. Mol. Pharm. 15, 5374–5386 (2018).
Kim, D., Kim, S., Park, G., Choi, H. & Ryu, J.-H. Spatiotemporal Self-Assembly of Peptide Amphiphiles by Carbonic Anhydrase IX-Targeting Induces Cancer-Lysosomal Membrane Disruption. JACS Au 2, 2539–2547 (2022).
Rünger, T. M. et al. Morphea-like skin reactions in patients treated with the cathepsin K inhibitor balicatib. J. Am. Acad. Dermatol. 66, e89–e96 (2012).
Zhang, P. et al. Gallic acid inhibits osteoclastogenesis and prevents ovariectomy-induced bone loss. Front Endocrinol. (Lausanne) 13, 963237 (2022).
Thiele, L. et al. Evaluation of particle uptake in human blood monocyte-derived cells in vitro. Does phagocytosis activity of dendritic cells measure up with macrophages? J. Controlled Release 76, 59–71 (2001).
Meyer, N. et al. Autophagy activation, lipotoxicity and lysosomal membrane permeabilization synergize to promote pimozide- and loperamide-induced glioma cell death. Autophagy 17, 3424–3443 (2021).
Zhang, H. et al. Polyhedron-Like Biomaterials for Innervated and vascularized bone regeneration. Adv. Mater. 35, 2302716 (2023).
Wu, X. et al. Enhancing calvarial defects repair with PDGF-BB mimetic peptide hydrogels. J. Controlled Release 370, 277–286 (2024).
Zhao, S.-J. et al. Macrophage MSR1 promotes BMSC osteogenic differentiation and M2-like polarization by activating PI3K/AKT/GSK3β/β-catenin pathway. Theranostics 10, 17–35 (2020).
Motegi, S. -i., Garfield, S., Feng, X., Sárdy, M. & Udey, M. C. Potentiation of Platelet-Derived Growth Factor Receptor-β Signaling Mediated by Integrin-Associated MFG-E8. Arteriosclerosis, Thrombosis, Vasc. Biol. 31, 2653–2664 (2011).
Ai, H. et al. Osteoclast-derived apoptotic bodies accelerate the pathological progression of osteoarthritis via disturbing subchondral bone remodeling. J. Orthop. Transl. 51, 108–118 (2025).
Wu, Y. et al. Osteoclast-derived apoptotic bodies inhibit naive CD8+ T cell activation via Siglec15, promoting breast cancer secondary metastasis. Cell Rep. Med. 4, 101165 (2023).
Wang, H. et al. Integrating enzymatic self-assembly and mitochondria targeting for selectively killing cancer cells without acquired drug resistance. J. Am. Chem. Soc. 138, 16046–16055 (2016).
Koo, S. et al. Real-time live imaging of osteoclast activation via cathepsin K activity in bone diseases. Angew. Chem. Int. Ed. 63, e202318459 (2024).
Song, Z. et al. Cathepsin K-activated probe for fluoro-photoacoustic imaging of early osteolytic metastasis. Adv. Sci. 10, 2300217 (2023).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
Li, J. et al. Epimedin A inhibits the PI3K/AKT/NF-κB signalling axis and osteoclast differentiation by negatively regulating TRAF6 expression. Mol. Med. 30, 125 (2024).
Shao, X. et al. Rescuing SERCA2 pump deficiency improves bone mechano-responsiveness in type 2 diabetes by shaping osteocyte calcium dynamics. Nat. Commun. 15, 890 (2024).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (U24A2076 to H.M., 82272145 to H.M., 82205136 to P.Z.). We thank the Instrumentation and Service Center for Molecular Sciences, Instrumentation and Service Center for Physical Sciences, and Biomedical Research Core Facilities at Westlake University for assistance with measurements.
Author information
Authors and Affiliations
Contributions
H.M.W. conceptualized, supervised, and founded the project. J.J.C. provided intellectual support and revised the original work. P.Z. and D.H.C. designed the experiments and collected the data. Y.F. and S.S.L. synthesized the compounds. Z.A.Z. and H.F.L. conducted some animal experiments. H.M.W., P.Z. and D.H.C. analyzed the date and wrote the manuscript with input from the other authors. All the authors discussed the results and had agreement on the final version of the manuscript. All the authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ce Dou and the other anonymous reviewer(s) for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, P., Chen, D., Fang, Y. et al. Bone homeostasis regulation by in vivo generation of osteoclast-derived apoptotic bodies using targeted self-assembly. Nat Commun 17, 88 (2026). https://doi.org/10.1038/s41467-025-66791-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66791-9