Abstract
The highly pathogenic avian influenza H5N1 virus clade 2.3.4.4b has been spreading globally since 2022, causing mortality and morbidity in domestic and wild birds, as well as in mammals, which underscores its potential to cause a pandemic. Here, we generate a panel of anti-hemagglutinin (HA) human monoclonal antibodies (mAbs) against the H5 protein of clade 2.3.4.4b. To develop human chimeric antibodies, H2L2 Harbor Mice®, which express human immunoglobulin germline genes, were immunized with H5 and N1 recombinant proteins from A/mallard/New York/22-008760-007- original/2022 H5N1 virus. Through hybridoma technology, sixteen fully human mAbs are generated, most of which show cross-reactivity against H5 proteins from different clade 2.3.4.4 virus variants. Fourteen out of the sixteen mAbs neutralize the virus in vitro. The mAbs with the strongest hemagglutination inhibition activity also demonstrate greater neutralizing capacity and show increased protective effects in vivo when administered prophylactically or therapeutically in a murine H5N1 challenge model. Using cryo-electron microscopy, we identify a cross-clonotype conserved motif that bound a hydrophobic groove on the head domain of H5 HA. Akin to mAbs against severe acute respiratory syndrome coronavirus 2 during the coronavirus 2019 pandemic, these mAbs could serve as treatments in case of a widespread H5N1 epidemic or pandemic.
Similar content being viewed by others
Introduction
Highly pathogenic H5N1 avian influenza virus emerged as a human disease in 19971. After variable H5N1 virus activity in the years before the coronavirus disease 2019 (COVID-19) pandemic, a subclade of H5N1, clade 2.3.4.4b, started to spread globally in 2022. This spread has resulted in significant losses in the poultry industry and endangered millions of wild birds2,3. This virus clade has also spilled over into mammals, causing severe disease and high fatality rates4. In March 2024, the virus was detected in the United States (U.S.) in dairy cattle, spreading among herds with high titers of virus present in the cows’ milk, and in October 2024, the virus was detected in a pig on a backyard farm in Oregon5. Even more alarming is the reported transmission of the virus to humans, posing a risk to workers who are in contact with infected animals and raising concerns about the potential for a pandemic. So far, more than 70 clade 2.3.4.4b human infections have been reported, with the vast majority in the U.S. While antivirals licensed for seasonal influenza are likely effective for H5N1, there are currently no long-acting treatments available that could e.g., be used prophylactically for individuals with a dysfunctional immune system who do not respond optimally to vaccination6,7. However, monoclonal antibodies (mAbs) can be used as long-acting treatment and have been successfully deployed during COVID-19 but also for respiratory syncytial virus (RSV) as well as ebolavirus8,9,10,11. Monoclonal antibodies targeting older clades of H5N1 have been developed, but are likely outpaced by virus evolution12. To address this issue, we have developed H5N1 clade 2.3.4.4b specific human mAbs generated in immunoglobulin germline humanized mice. These mAbs can effectively neutralize clade 2.3.4.4b H5N1 and provide protection in a murine H5N1 challenge model. MAbs have been used extensively and successfully as therapeutics and prophylactics during the COVID-19 pandemic13,14. The mAbs developed here against H5N1 may similarly serve as therapeutics or prophylactics in potential future H5N1 epidemics or pandemics.
Results
Generation and in vitro characterization of mAbs
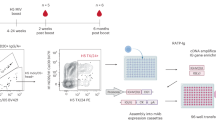
H2L2 Harbor Mice®, which express human-rat chimeric antibody genes15, were immunized with recombinant hemagglutinin H5 and neuraminidase N1 proteins from a clade 2.3.4.4b virus (Supplementary Fig. 1a) and mAbs were generated via hybridoma technology. Initial screening was done using enzyme-linked immunosorbent assays (ELISA) and hemagglutination inhibition (HI) assays, identifying 1977 positive clones. From these, 401 clones were down-selected for secondary screening, of which 30 clones showed strong binding to the H5 protein and had HI activity, leading to their selection for isotyping and sequencing. After removal of IgMs and identical clones, down-selected clones were chosen for variable segment DNA synthesis and cloned into human constant chain containing-expression vectors (human G1/ kappa) for fully human recombinant antibody production. Sixteen recombinant clones were selected as sequence unique lead candidates based on broad H5 activity and recombinant expression. We grouped these sixteen antibodies into clonotypes based on exact amino acid length with >90% amino acid identity in their junctions/complementarity-determining region 3 (CDR3), along with shared V(D)J/VJ gene usage for both their heavy and light chains, resulting in a total of 5 closely related clonotypes (Supplementary Fig. 1d). The identified clonotypes and their corresponding clones were as follows: clonotype 1 (1A1, 12C11, 12G9, 13B9, 13E8), clonotype 2 (1H2, 17E3, 20D10), clonotype 3 (6G1, 7G4, 7G11, 12G8, 17C12), clonotype 4 (7H10, 14B8) and clonotype 5 (12G1). While clonotypes 3, 4 and 5 each utilized the longest heavy chain CDR3s at 19 amino acids (IMGT numbering) and had shared heavy V(D)J usage (VH4-59, DH3-10, and JH6), the sequence variation in their CDR3 was greater than 40%, which was significantly higher than the predicted somatic hypermutation rates (1.7-3.6%) when comparing their VH4-59 variable genes alone to germline (data not shown). This argues for separate clonal ancestries and, thus, their categorization as unique clonotypes. Light chain CDR3 lengths were less varied given the lack of a D gene cluster and averaged 9 amino acids for each clonotype, typical for human kappa chains, even though clonotype 4’s light chain was one residue shorter at 8 amino acids, adding another unique characteristic for this clonotype.
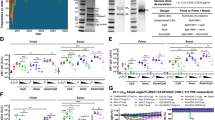
These mAbs were produced, and their binding activity was assessed against a panel of H5 proteins from virus of different clades (Fig. 1a). The mAbs showed robust binding to H5 HAs from recent clade 2.3.4.4b isolates including to HAs from viruses recently detected in birds in New York City, New York, U.S.16. Of note, robust binding to the HA of a U.S. bovine H5N1 isolate was detected as well. Most mAbs also showed strong binding to HAs from older clade 2.3.4.4 isolates. This included avian strains from 2014 from the Netherlands and the U.S. and an H5N6 human isolate from a 2016 case in China. However, no binding was observed with HAs from older clade 1 and 2 strains, HA from the 2.3.2.1c clade responsible for recent human cases in Cambodia, or H1 HA from a seasonal H1N1 strain (used as a negative control). The phylogenetic relationship of these HA proteins is shown in Supplementary Fig. 2. Most clonotypes, except for the one containing mAbs 7H10 and 14B8, showed strong binding to H5 of clade 2.3.4.4b H5N1 viruses. Clones belonging to the same clonotype typically exhibited similar binding patterns.
a Binding breadth of anti-H5 mAbs to H5 protein from avian isolates (purple), a cattle isolate (orange), and human isolates (blue), was evaluated by ELISA in duplicates. As positive control, an anti-HA antibody, CR911432 was used. Data shown is minimal binding concentration (ug/ml), defined as the lowest concentration with a signal greater than 3 standard deviations (SD) above blanks. b MAbs ability to block the virus-host receptor interaction was assessed by hemagglutination inhibition (HI) assay in duplicates. As positive control, serum (1:10) from a mouse infected with A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant) was used. Data shows minimal HI concentration (ug/ml), defined as the lowest concentration that shows HI activity. c Efficacy of the anti-H5 mAbs to inhibit viral replication was evaluated using an in vitro neutralization assay against A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant), in duplicates, with controls identical to part A. Data shows minimal neutralizing titer (ug/ml), defined as the lowest concentration where neutralization is observed. d MAbs capacity to inhibit neuraminidase (NA) activity of clade 2.3.4.4b virus was tested in duplicates, with an anti-NA antibody, 1G0117, as positive control. Data shown is the 50% inhibitory concentration (IC50) (ug/ml). Negative control for all the assays was an anti-SARS-CoV-2 spike antibody18. Source data are provided as a Source Data file.
Further characterization of the mAbs was performed in vitro using HI and microneutralization assays. An HI assay was run to assess whether the mAbs inhibit the interaction of the virus with the host receptor (Fig. 1b). This assay was conducted with several virus strains. Antibodies showed HI activity against A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant), here referred to as A/bald eagle/FL/W22-134-OP/2022 and viruses isolated from dairy cattle (clade 2.3.4.4b also) in a similar manner. As expected, in accordance with the binding data, no HI activity was observed against the A/Vietnam/1203/2004 virus belonging to clade 1. A/Victoria/4897/2022, a human seasonal H1N1, was used as an irrelevant virus control. No HI activity was detected against this strain demonstrating specificity to clade 2.3.4.4 H5N1 viruses. Of note, the clonotype containing clones 7H10 and 14B8, not only had low binding, but also showed no HI activity. Additionally, an in vitro microneutralization assay was performed to assess the ability of the mAbs to disrupt the viral life cycle, potentially through various mechanisms (Fig. 1c). We tested whether the antibodies could prevent the virus from entering the cell or block the virus post initial infection, or both. The same fourteen mAbs with the highest HI activity also exhibited greater neutralizing capacity and inhibited viral replication in cell culture at different stages of the virus life cycle. MAb 1A1 from clonotype 1, as well as all the clones belonging to clonotype 2, 3, and 5 with the strongest HI activity, demonstrated greater neutralizing capacity, with a minimal neutralizing titer of up to 0.012 ng/ml being observed. As the final assay, we assessed whether the antibodies inhibited the neuraminidase (NA) enzymatic activity through steric hindrance. Clone 1H2 from clonotype 2 and 12G1 from clonotype 5 showed strong inhibitory activity, similar to the previously characterized pan-NA antibody, 1G0117, here used as a positive control. 7H10 and 14B8 from clonotype 4 exhibited low or no NA inhibitory activity. The mAbs did not inhibit when tested using a small molecule substrate, suggesting that the inhibition observed was through steric hindrance rather than direct interaction with the active site of the NA (Supplementary Fig. 3).
Human anti-H5 mAbs provide protection in a murine H5N1 challenge model when given prophylactically or therapeutically
To further investigate the antiviral activity of the mAbs in vivo, we tested selected mAbs from different clonotypes in both prophylactic and therapeutic settings in an H5N1 challenge model in 6-week-old BALB/c mice. The mAbs with the strongest binding and highest neutralization activity from each clonotype, 1A1 (clonotype 1), 20D10 (clonotype 2), 6G1 (clonotype 3) and 12G1 (clonotype 5), were selected for in vivo studies. Mab 7H10 (clonotype 4) was added as a representative of a binding but non-neutralizing mAb. In the prophylactic study, the mAbs were administered at four different doses via the intraperitoneal route, 4 hours prior to intranasal challenge with 5 × 50% lethal doses (LD50) of A/bald eagle/FL/W22-134-OP/2022. Mice were monitored for weight loss and survival for 14 days to assess protection (Fig. 2a–c). The antibodies that were effective neutralizers in vitro, 1A1, 20D10, 6G1 and 12G1, also demonstrated the ability to protect mice in a dose-dependent manner. Mice administered with 5 mg/kg, 1 mg/kg, and 0.3 mg/kg of the indicated antibodies showed no morbidity and complete survival. Of note, mice injected with antibodies belonging to clonotype 1 (1A1) and clonotype 3 (6G1) even demonstrated protection at a very low dose of 0.1 mg/kg. This level of protection was not observed in mice administered with the non-neutralizing mAb 7H10 or with an isotype control antibody (hG1/hk, anti-severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike mAb18), indicating that protection in this case is associated with the administration of neutralizing antibodies.
a Six-week-old female BALB/c mice (n = 5/group) were injected intraperitoneally with different amounts of anti-H5 mAb. As a negative control, an anti-SARS-CoV-2 spike mAb was used and the data of the shared negative control group is plotted alongside the experimental groups. After 4 h, mice were infected with 5 x 50% lethal doses (LD50) of A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant) virus. Body weight percentage b and survival c were plotted over 14 days post-infection. Data are presented as the mean (n = 5), with error bars showing the SD of the mean. The dotted line in b indicates the maximum body weight loss (25%) allowed in the experiment, defined as the humane endpoint. d Six-week-old female BALB/c mice (n = 3/group) received 5 mg/kg of anti-H5 mAbs 4 h before infection, and anti-SARS-CoV-2 mAb was used as a control, as in Part a. Lungs were harvested on day 3 and day 5 post-infection. e Viral titers in lungs from individual mice were determined as log10 plaque forming units (PFU)/ml and samples were analyzed in duplicate. Dots represent values from individual mice while bars show geometric means (n = 3) with geometric SD. The dashed line at the y-axis indicates the limit of detection (LOD), defined by half of the lowest dilution at which a positive assay response is observed. For e, statistical analyses were performed using a two-way ANOVA test ((*) p < 0.05, (ns) no significant). a, d illustration created with BioRender.com. Source data are provided as a Source Data file.
We then wanted to study if these mAbs could reduce viral replication in the lungs of animals. Experiments were performed similarly to the one described above with mAbs dosed 4 hours prior to challenge with 5 LD50 of A/bald eagle/FL/W22-134-OP/2022 virus. Lungs were harvested on days 3 and 5 post-infection, and viral titers were determined using plaque assay (Fig. 2d, e). Viral titers were observed to be 4 logs lower in lung homogenates of mice treated with mAbs 20D10, 6G1, or 12G1 3 days post-infection, compared to the isotype control group. At day 3, we also observed a significant (3 log) reduction in viral titers in the lung homogenates of mice that received mAb 1A1. By day 5, no virus was detectable in the lungs of 20D10, 6G1, and 12G1 treated animals. No reduction was observed for mAb 7H10 group compared to the control one on day 5. This finding suggests that the antibodies inhibit replication of H5N1 in the lungs, likely associated to their high neutralizing efficiency in cell culture.
To further assess the potential therapeutic benefits of the mAbs, mice were intranasally challenged with 5 LD50 of A/bald eagle/FL/W22-134-OP/2022 virus. MAbs were subsequently administered on days 2, 3, or 4 post-infection, and the mice were monitored for weight loss and survival for 14 days to evaluate the protective effect (Fig. 3). All mice that received 6G1 mAb on day 2 post-infection survived, and 80% of mice that received 20D10 and 12G1 at the same time point survived as well. Only 40% of mice that received mAb 1A1 at day 2 post-infection survived. Mice treated with 20D10, 6G1, and 12G1 antibodies on day 3 and day 4 post-infection, when significant weight loss had already occurred, showed varying levels of survival, ranging from 20% to 60%. All the mice treated with 7H10 or anti-SARS-CoV-2 antibodies reached the ethical endpoint on days 5 and day 6 post-infection, consistent with the results from the prophylactic treatment study.
a Six-week-old female BALB/c mice (n = 5/group) were injected intraperitoneally with 5 mg/kg of anti-H5 mAb and as a negative control anti-SARS-CoV-2 spike mAb was used. The data of the shared negative control group is plotted alongside the experimental groups. The antibodies were administered on day 2, 3, and 4 after infecting the mice with 5 LD50 of A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant) virus (created with BioRender.com). The percentage of initial body weight b and survival c were plotted over 14 days post-infection. Data are presented as the mean (n = 5), with error bars showing the SD of the mean. The dotted line in b indicates the maximum body weight loss (25%) allowed in the experiment, defined as the humane endpoint. Arrows indicate the time points at which mAbs were administered. Source data are provided as a Source Data file.
MAbs converge at the HA head
Negative stain electron microscopy revealed that 13 representative mAbs from clonotypes 1-3 and 5 had binding footprints overlapping (clonotype 1) or proximal to the receptor binding site (RBS) (clonotypes 2, 3, and 5) when complexed with recombinantly expressed HA from A/Jiangsu/NJ210/2023 H5N1 (Jiangsu H5), a clade 2.3.4.4b clinical isolate from an infected human patient in Jiangsu, China, which is consistent with their competition pattern (Supplementary Figs. 4–6). Representative mAbs from clonotypes 1–3 and 5 were also complexed with recombinant HA from A/red-tailed hawk/New York/NYCVH 22-8477/2022, another clade 2.3.4.4b virus HA, indicating that the binding footprint remained consistent across strains (Supplementary Figs. 5, 7). To understand the molecular basis of antigen engagement, we solved three high resolution cryo-EM structures from complexes with mAbs 20D10 (clonotype 2) at 3.4 Å resolution, 6G1 (clonotype 3) at 3.2 Å resolution, and 12G1 (clonotype 5) at 3.06 Å resolution (Fig. 4, Supplementary Fig. 8, Supplementary Table 1). All non-van der Waals interactions between the heavy and light chains with the H5 head are illustrated in Fig. 4a–f. Antigen engagement by all three mAbs was mediated primarily by heavy chain complementarity-determining region 3 (CDRH3) and light chain complementarity-determining region 2 (CDRL2) with additional interactions involving CDRH1 and CDRL1. Generally, CDRH1 and CDRH3 were found to insert into a hydrophobic groove composed of residues 123-129 and 164-171 on Jiangsu H5 while CDRL1 and CDRL2 engaged the cleft between adjoining protomers. While there is minor variation in the specific residues that define epitope-paratope contacts for each clonotype, all heavy chains were predicted to interact with H5 residues S125B, S167, and N169; light chain contacts were more variable across clonotypes, but all CDRL2s engaged T163. Indeed, all three mAbs use the same CDRH1 Y32 residue to engage S125B and the same CDRL2 Y49 residue to engage T163. Interestingly, S67 in light chain framework region 3 (FR3) of 6G1 is predicted to form a sidechain-to-sidechain hydrogen bond with N187 with the adjoining protomer (Fig. 4d). In total, each clonotype is predicted to form eleven non-van der Waals interactions with nearly identical regions in the H5 head with clonotype 2 forming the greatest number of heavy chain mediated contacts and clonotype 5 forming the most light chain mediated contacts.
a–b Map and model for clonotype 2 mAb 20D10 complexed with A/Jiangsu/NJ210/2023 H5 protein. The primary protomer is depicted in light gray with adjoining protomers depicted in gray. c–d Map, model, and antigen contacts identified in the H5 complex formed with clonotype 3 mAb 6G1. e–f Map, model, and antigen contacts identified in the H5 complex formed with clonotype 5 mAb 12G1. a–f All non-van der Waals 20D10 heavy and light chain contacts with antigen are depicted as dashed lines. Hydrogen bonds depicted in gray, hydrophobic interactions depicted in turquoise. All antibody residues are presented in Kabat numbering and all H5 residues in H3 numbering.
While we were unable to build an atomic model for mAb 1A1 (clonotype 1) our cryoEM map allowed us to identify the epitope footprint and compare it to clonotypes 2, 3, and 5 (Fig. 5a, Supplementary Fig. 6). Our data revealed that clonotypes 2, 3, and 5 had nearly identical binding footprints proximal to the protomer interface while clonotype 1 had a distinct footprint that encompasses the RBS (Fig. 5a). Alignment of the 20D10, 6G1, and 12G1 complexes highlights how epitopes, angles of approach, and binding modes are structurally conserved across all three clonotypes (Fig. 5b). This is further illustrated through comparison of the heavy and light chain CDRs involved in antigen engagement, which contain a series of tyrosine residues and hydrophobic side chains across all three models including an LxYYY motif in CDRH3 (Fig. 5c). Although clonotype 4 was not structurally characterized, it also contains the LxYYY motif, providing further evidence of the importance of CDRH3 in mediating antigen binding. The relative hydrophobicity of the antigen surface engaged by CDRH3 and CDRL2 is visualized through a surface rendering, highlighting the prominence of the hydrophobic groove (Fig. 5d). Through sequence conservation analyses, we found that both the hydrophobic groove targeted by the heavy chain CDRs and the residues targeted by the light chain CDRs were highly conserved across the clade 2.3.4.4b strains included in this study (Fig. 5e). In accordance with our functional data, these regions were not conserved in A/Wisconsin/67/2022 H1 or H5 clades 1, 2.3.2.1c and 2.1.3.2 and residue I166 within the hydrophobic groove is particularly variable (Supplementary Fig. 10).
a Footprints for clonotype 1 (salmon) and clonotypes 2, 3, and 5 (fuchsia) on a surface rendering of A/Jiangsu/NJ210/2023 H5. b Shared architecture of antigen engagement by the variable fragments of mAbs 20D10 (pink), 6G1 (green), and 12G1 (blue) colored by clonotype. Heavy chains are presented in a darker shade than light chains. c Heavy and light chain complementarity determining regions making direct contact with antigen. Residue labels in 20D10 numbering. d Hydrophobic surface renderings of the binding pocket and CDRH3 residues involved in direct engagement. Residue labels are provided using 20D10 numbering. e Sequence conservation of clade 2.3.4.4b H5 sequences from strains used for functional characterization as determined by sequence alignment using ClustalW mapped onto a surface rendering of A/Jiangsu/NJ210/2023 H5. Residue variability increases from white (AL2CO score 0.306) to dark blue (AL2CO score -4.895).
Discussion
Here, we developed a panel of sixteen human antibodies against clade 2.3.4.4b highly pathogenic avian influenza (HPAI) H5N1 virus using an antibody germline humanized mouse system. A spleen from an H2L2 Harbor Mouse® was used for hybridoma fusion and full human antibody production, with the ultimate goal for prophylactic and/or therapeutic applications. Most of the generated antibodies showed broad binding within clade 2.3.4.4b HAs and many even bound to HAs from historic clade 2.3.4.4 strains. No cross-binding to older H5 clades, clade 1, or to clade 2.3.2.1c was observed. Of note, clade 2.3.2.1c is responsible for a handful of human infections in Southeast Asia-in contrast to more than 70 human infections with clade 2.3.4.4b. Most mAbs showed strong in vitro antiviral activity in HI, neutralization, and neuraminidase inhibition assays. Importantly, the HI active antibodies also had strong HI activity against two recent dairy cattle isolates. In vivo, all tested mAbs with in vitro neutralizing activity showed prophylactic activity against a lethal H5N1 challenge and provided a survival benefit when given as late as 4 days post challenge, except for mAb 1A1 (clonotype 1), which only provided protection prophylactically. This limitation could perhaps be attributed to the half-life of the different antibodies affecting their effectiveness. MAbs from clonotype 4 showed limited binding to clade 2.2.4.4b or older clade 2.3.4.4 H5 proteins; however, they did not have neutralization activity in vitro, resulting in a failure to provide protection in vivo. Structural analysis revealed that mAbs from clonotypes 2, 3 and 5 used a conserved motif to bind a hydrophobic groove on the HA head adjacent to the promoter interface while the clonotype 1 mAb bound directly to the RBS. So far in the U.S. more than 70 humans have been infected with clade 2.3.4.4b H5N1 virus with several severe infections19,20,21 including two mortalities22,23. While no human-to-human spread has been detected so far, mammalian adaptations have been found in clade 2.3.4.4b viruses that could increase the pandemic potential of the virus. In addition, a reassortment event between seasonal H1N1 or H3N2 with H5N1 could lead to the emergence of a virus that is both pathogenic and transmissible in humans (of note, the last three influenza pandemics were caused by such reassortants)24. The antibodies described here, especially mAbs 20D10, 6G1, and 12G1, could be used as therapeutic treatment for severe zoonotic human H5N1 cases on their own or in addition to other antivirals like neuraminidase or endonuclease inhibitors25,26. Since 20D10, 6G1, and 12G1 target the same footprint, it is unlikely that using them together in a cocktail would be beneficial. But, especially for prophylactic treatment, one of them could be combined with mAb 1A1, which targets a different footprint and has shown potent protection when given prophylactically. These mAbs could also be combined with other anti-influenza virus mAbs with different targets e.g., in the stalk domain of HA27 or on the viral NA28. In case of an H5N1 pandemic, it is likely that vaccines would be rolled out fast and that short acting antivirals like neuraminidase or cap-snatching inhibitors would be available. However, similar to COVID-19, it is likely that a proportion of the population with underlying diseases or those that are immunocompromised will not be able to mount a sufficient response to vaccination and can not be kept on antivirals for long periods of time. A long-acting mAb prophylaxis, as used for COVID-19 or respiratory syncytial virus based on mAbs like 20D10, 6G1, or 12G1 could be used to protect vulnerable individuals during an H5N1 pandemic.
Methods
Cells
Madin-Darby canine kidney (MDCK) cells (ATCC #CCL-34) were used for in vitro microneutralization assays and viral titer determination. Expi293F cells were used for antibody production.
Recombinant glycoproteins
Recombinant hemagglutinin (HA) and neuraminidase (NA) proteins were produced using the baculovirus expression system as described in ref. 29. References of the proteins from the indicated virus isolates, such as accession GISAID numbers and GenBank numbers, are indicated in the supplementary material. Briefly, the protein constructs for HAs were designed with an N-terminal signal peptide, the HA ectodomain, a C-terminal thrombin cleavage site, a T4 trimerization domain and a hexahistidine tag. Notably, for the recombinant H5 proteins, the polybasic cleavage site in the HA ectodomain was replaced by a single alanine. The NA protein constructs were designed with an N-terminal signal peptide, a 6x-His tag, a VASP (vasodilator-stimulated phosphoprotein) tetramerization domain, a thrombin cleavage site, and the NA head domain. The constructs were cloned into shuttle vectors, and recombinant bacmids were generated and transfected into Sf9 cells to produce baculoviruses. These HA-expressing baculoviruses were passaged in Sf9 cells and then used to infect BTI-TN-5B1-4 cells for protein production. Proteins were purified from the cell supernatants via affinity chromatography using Ni2 + -nitrilotriacetic acid (NTA) agarose beads (QIAGEN) three days post-infection, as described in ref. 29.
Viruses
A/bald eagle/FL/W22-134-OP/2022 is a 6:2 reassortant H5N1 virus with A/PR/8/1934 rescued via reverse genetics, with a monobasic cleavage site replacing the polybasic cleavage site30. Similarly, A/Vietnam/1203/2004 virus retains surface H5N1 glycoproteins from A/Vietnam/1203/2004 (with a monobasic cleavage site replacing the polybasic cleavage site in the HA) while using the A/PR/8/34 backbone. Both viruses were used for in vitro neutralization assays, in addition the first one mentioned here was used for in vivo studies. To assess the hemagglutination inhibition activity of our antibodies against full-length viruses, we included authentic bovine viruses; A/bovine/Ohio/B24OSU-467/2024 and A/bovine/Ohio/B24OSU-323/2024.
Immunizations and hybridoma generation
To generate human antibodies, H2L2 Harbor Mice®, human antibody transgenic mice (Harbor BioMed, Cambridge, MA) were used in collaboration between the Icahn School of Medicine at Mount Sinai and Harbor BioMed. The H2L2® is a chimeric transgenic mouse containing a selected human variable gene segment loci of the heavy and kappa antibody chains along with the rat heavy and kappa constant gene segment loci, producing antibodies with similar diversity seen in human antibody immunity15,31. Eight- to twelve-week-old H2L2 Harbor Mice® mice were immunized with HA and NA from A/mallard/New York/22-008760-007-original/2022 (H5N1) adjuvanted with 40ug of poly I:C (InVivoGen, San Diego, CA) by the intraperitoneal route which we have found optimal in stimulating B cells in the spleen. Each mouse received a prime followed by two boosts, and blood was collected two weeks after each boost to monitor serum antibody titers via enzyme-linked immunosorbent assay (ELISA) and hemagglutination inhibition (HI) analysis. One mouse was selected for hybridoma fusion and received two final boosts consisting of 20ug of the H5 protein at -4 and -2 days before being bled, euthanized by Institutional Animal Care and Use Committee (IACUC) approved methods, and the spleen was harvested for splenic fusions. The spleen was processed to a single-cell suspension, and hybridomas were generated following a standard protocol (StemCell Technologies, Vancouver, BC, Canada). Briefly, individual B cell clones were grown on soft agar and selected for screening using a robotic ClonaCell Easy Pick instrument (Hamilton/Stem Cell Technology). Individual clones were expanded, and the supernatant was used to screen for H5 binding (ELISA) and HI activity. All animal studies were approved by the Icahn School of Medicine IACUC.
Sequencing and humanizing of antibodies
Sequencing was done using Switching Mechanism At 5’ End of RNA Template (SMARTer 5’) Rapid Amplification of cDNA Ends (RACE) technology (Takara Bio USA) adapted for antibodies to amplify the variable genes from heavy and kappa chains for each hybridoma. Briefly, RNA was extracted from hybridomas using QIAGEN RNeasy Mini Kit (QIAGEN, Valencia, CA), followed by first strand cDNA synthesis with isotype-specific constant gene 3’ primers, the SMARTer II A Oligonucleotide, and SMARTscribe reverse transcriptase. Amplification of cDNA was performed with SeqAmp DNA Polymerase (Takara), using a nested 3’ primer and a 5’ universal primer. Purified PCR products were then Sanger sequenced using 3’ constant gene primers (Azenta/GeneWiz, South Plainfield, NJ). Sequence results were blasted against the IMGT human databank of germline genes using V-Quest (http://imgt.org) and analyzed for clonality based on CDR3/junction identity and V(D)J usage. Unique clones from each clonal family were selected, and DNA was synthesized and cloned into in-house antibody expression vectors (pTWIST-hG1/hck) containing a human IgG1 constant region and kappa light chain constant region (Twist Biosciences, South San Francisco, CA).
Generation of recombinant (r)mAbs in 293 suspension cells
DNA plasmids of the light and heavy chains were combined with Expifectamine1000 (Gibco) in a total of 24 ml of Opti-minimal essential media (MEM) and transfected into 3.5 × 106 cell/ml in a final volume of 200mlof Expi293 Expression Medium (Gibco). The next day, Enhancer 1 and Enhancer 2 were added (Gibco).
ELISA
96-well plates (Immulon 4 HBX; Thermo Scientific) were coated with 50 ul of recombinant protein (2 ug/ml) and incubated at 4 °C overnight. The next day, 100 ul of serially diluted IgG, starting at a concentration of 30 ug/ml, was added to the plates in duplicate. Binding of the antibodies to the protein was evaluated using a horseradish peroxidase (HRP)-labeled anti-human IgG (Sigma) as secondary antibody, diluted 1:3000. The positive control was an anti-HA antibody32, while the negative control was an anti-SARS-CoV-2 spike antibody18.
HI assay
First, an HA assay was performed using turkey RBCs to determine the viral HA titer. Next, 25ul of serially diluted IgG, starting at a concentration of 30 ug/ml or 25 ug/ml were incubated with 8 HA units (HAU) of the virus for 1 hour (h) at room temperature (RT). Then, 50ul of 0.5% turkey RBCs were added, and the plates were placed at 4 °C for 45 minutes. Virus-negative wells were analyzed for pellet formation.
In vitro microneutralization assay
Neutralization of clade 2.3.4.4b (H5N1-PR8) virus with human IgG was measured as previously described33. Serially diluted mAbs at a starting concentration of 25 ug/ml were incubated in 100ul with 100 times 3.82 × 104 TCID50/50 ul of the virus for 1h at RT followed by inoculation of MDCK cells plated on 96-well plates (Corning). After 48 h of incubation at 37 °C, an HA assay was performed to detect the presence of the virus. The positive control was an anti-HA antibody32, while the negative control was an anti-SARS-CoV-2 spike antibody18.
Neuraminidase inhibition (NI) assay
First, a neuraminidase assay of the clade 2.3.4.4b (H5N1-PR8) virus was performed to determine the optimal virus concentration to be used in the NI assay. To measure the inhibitory activity of the antibodies, serially diluted mAbs (starting concentration 30 ug/ml) were incubated in 50ul for 6 h at 37 °C with an equal volume of the selected virus dilution in fetuin-coated plates. The percentage of enzymatic inhibition was calculated relating the number in experimental wells to the mean number of virus only wells and wells with sample diluent only. The half inhibitory concentration (IC50) was calculated based on dose-response curves, using top and bottom constraints of 0 and 100% in GraphPad Prism 10.
In vivo experiments
All experiments were performed according to protocols approved by the Icahn School of Medicine at Mount Sinai IACUC. Six-week old female BALB/c mice were housed for at least one week before experiments (sourced from The Jackson Laboratory), and infections were performed on animals anesthetized with a ketamine-xylazine mixture. In prophylactic studies, mice were divided into four groups of five, receiving 5mg/kg, 1mg/kg, 0.3mg/kg, and 0.1mg/kg of anti-H5 mAbs intraperitoneally. An irrelevant antibody, an anti-SARS-CoV-2 spike mAb, was used to measure any non-specific protection. After 4 h, mice were inoculated intranasally with 5 × 50% lethal doses (LD50) of A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant) virus in a total volume of 50ul of phosphate buffered saline (PBS). Mice were monitored daily for body weight and mortality until day 14 post-infection or until all animals died, with those losing 25% or more of their initial body weight being euthanized. Mice that showed severe clinical signs, such as a ruffled coat, hunched posture, orbital tightening, and lack of reaction to stimuli, were euthanized at 24% weight loss since they were only weighted once every day and likely to cross the endpoint within the next few hours. In therapeutic studies, three experimental groups of five mice were first infected with 5 LD50 of A/bald eagle/FL/W22-134-OP/2022 (H5N1-A/PR/8/34 reassortant) virus and mice were then treated at 2, 3, or 4 days post-infection with 5 mg/kg of anti-H5 mAbs and an irrelevant antibody, an anti-SARS-CoV-2 mAb that served as a control to measure any non-specific protection. Mice were monitored similarly daily for body weight and mortality. To analyze viral clearance in lungs, as for prophylactic studies, groups of three mice received 5 mg/kg of anti-H5 mAbs intraperitoneally, followed by intranasal inoculation with 5 LD50 of virus in 50ul of PBS. Mice were euthanized on day 3 or 5 post-infection for lung titer analysis. Lungs were harvested and homogenized in 0.8 ml of PBS using a BeadBlaster 24 (Benchmark) homogenizer. The homogenates were centrifuged (15 min, 16,100 × g, 4 °C) to remove cellular debris and stored in single-use aliquots at −80 °C. Infectious virus titers were determined by plaque assay (see supplementary material).
Immune complexing and negative stain electron microscopy (EM)
Monoclonal antibody-binding fragments (Fab) and H5 protein (A/Jiangsu/NJ210/2023 or A/Red-tailed hawk/New York/NYCVH 22-8477/2022) were complexed at a 3:1 molar ratio for 1h at RT. For nsEM grids, 3ul of immune complexes were applied to glow-discharged, carbon-coated 400 mesh copper grids at a concentration of ~10-20ug/ml (Electron Microscopy Services). Excess sample was blotted, and the complexes were stained with 2% w/v uranyl formate for 60s twice. Imaging was performed on a Talos 200 C or Tecnai Spirit T12 electron microscope. Micrographs were collected using Leginon. For each complex, 30k to 100k particles were picked and stacked using Appion and processed to generate 2D classes and 3D reconstructions using Relion 3.0 or Relion 4.0b1. Initial reference models were based on PDB: 6E7G and 4K62. Composite 3D reconstructions were made with UCSF ChimeraX.
CryoEM grid preparation and imaging
Fabs of 12G1 and 20D10 were incubated with Jiangsu H5 and CR9114 Fab at a 3:1:3 molar ratio for 1h at RT prior to cryoEM grid preparation and placed on ice. Fabs of 1A1 and 6G1 were co-incubated with Jiangsu H5 and CR911432 at a molar ratio of 3:3:1:3. A Vitrobot Mark IV (Thermo Fisher Scientific) was used to vitrify all samples at chamber temperature of 4 °C, 100% humidity. To decrease orientation bias, immune complexes (~0.7–0.85 mg/ml) were mixed with n-octyl-beta-D-glucoside (OBG, Anatrace) to a final OBG concentration of 0.1% before freezing. Datasets were collected on a 200 kV Glacios (Thermo Fisher) equipped with a Falcon IV direct electron detector. Data for the 12G1 complex and the 1A1 + 6G1 co-complex were collected using EPU at 190,000× magnification, with a pixel size of 0.725 Å and an exposure dose of 45 e − /Ų. The 20D10 complex was collected similarly with a pixel size of 0.718 Å. Each dataset contained approximately 2000 to 7000 movie micrographs.
Statistics & reproducibility
Bar graphs are displayed as individual values and the average is presented as the geometric mean, while the line graph data are shown as geometric means. Errors are represented by the standard deviation (SD) of the geometric mean. The experiments had sufficient power (n = 5 mice per group) to detect clear differences between groups. No statistical method was used to predetermine the sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to group allocation during experiments and outcome assessment. All statistical analyses were performed using GraphPad Prism 10.
Ethics statement
Our research adheres to all ethical regulations. Specifically, animal experiments were conducted in accordance with protocols approved by the Institutional Animal Care and Use Committee of the Icahn School of Medicine at Mount Sinai (protocol #2014-0255). Additional Methods can be found in the Supplementary Methods Section of the Supplementary Information file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting our work are available in the paper and Supplementary Figs. 1–10. nsEM and cryoEM maps and models are deposited in the Electron Microscopy DataBank under accession IDs EMD-48510-48512, 48679-48690, 48692-48696: EMD-48510, EMD-48511, EMD-48512, EMD-48679, EMD-48680, EMD-48681, EMD-48682, EMD-48683, EMD-48684, EMD-48685, EMD-48686, EMD-48687, EMD-48688, EMD-48689, EMD-48690, EMD-48692, EMD-48693, EMD-48694, EMD-48695 [https://www.ebi.ac.uk/emdb/EMD-48692], and EMD-48696 and in the Protein DataBank under accession IDs 9MQ1, 9MQ2, and 9MQ3: 9MQ1, 9MQ2, and 9MQ3. Source data are provided with this paper.
References
Sims, L. D. et al. Avian influenza in Hong Kong 1997-2002. Avian Dis. 47, 832–838 (2003).
Webby, R. J. & Uyeki, T. M. An update on highly pathogenic avian influenza A(H5N1) virus, clade 2.3.4.4b. J. Infect. Dis. 230, 533–542 (2024).
Confirmations of highly pathogenic avian influenza in commercial and backyard flocks., https://www.aphis.usda.gov/livestock-poultry-disease/avian/avian-influenza/hpai-detections/commercial-backyard-flocks (2024).
Plaza, P. I., Gamarra-Toledo, V., Eugui, J. R. & Lambertucci, S. A. Recent changes in patterns of mammal infection with highly pathogenic avian influenza A(H5N1) virus worldwide. Emerg. Infect. Dis. 30, 444–452 (2024).
Federal and State Veterinary Agencies Share Update on HPAI Detections in Oregon Backyard Farm, Including First H5N1 Detections in Swine, https://www.aphis.usda.gov/news/agency-announcements/federal-state-veterinary-agencies-share-update-hpai-detections-oregon (2024).
CDC A(H5N1) Bird Flu Response Update November 18, 2024, https://www.cdc.gov/bird-flu/spotlights/h5n1-response-11152024.html (2024).
Adisasmito, W. et al. Effectiveness of antiviral treatment in human influenza A(H5N1) infections: analysis of a Global Patient Registry. J. Infect. Dis. 202, 1154–1160 (2010).
Subramaniam, S., Saville, J. W., Feng, F. & Freiburger, L. Therapeutic antibodies for infectious diseases: recent past, present, and future. Biochemistry 64, 3487–3494 (2025).
Casadevall, A. & Focosi, D. Lessons from the use of monoclonal antibodies to SARS-CoV-2 spike protein during the COVID-19 pandemic. Annu Rev. Med 76, 1–12 (2025).
Sevendal, A. T. K., Hurley, S., Bartlett, A. W., Rawlinson, W. & Walker, G. J. Systematic review of the efficacy and safety of RSV-specific monoclonal antibodies and antivirals in development. Rev. Med Virol. 34, e2576 (2024).
Rijal, P. & Donnellan, F. R. A review of broadly protective monoclonal antibodies to treat Ebola virus disease. Curr. Opin. Virol. 61, 101339 (2023).
Ellebedy, A. H. et al. Adjuvanted H5N1 influenza vaccine enhances both cross-reactive memory B cell and strain-specific naive B cell responses in humans. Proc. Natl. Acad. Sci. USA 117, 17957–17964 (2020).
Wang, C. et al. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 11, 2251 (2020).
Focosi, D. et al. Monoclonal antibody therapies against SARS-CoV-2. Lancet Infect. Dis. 22, e311–e326 (2022).
Widjaja, I. et al. Towards a solution to MERS: protective human monoclonal antibodies targeting different domains and functions of the MERS-coronavirus spike glycoprotein. Emerg. Microbes Infect. 8, 516–530 (2019).
Meade, P. S. et al. Detection of clade 2.3.4.4b highly pathogenic H5N1 influenza virus in New York City. J. Virol. 98, e0062624 (2024).
Stadlbauer, D. et al. Antibodies targeting the neuraminidase active site inhibit influenza H3N2 viruses with an S245N glycosylation site. Nat. Commun. 13, 7864 (2022).
Clark, J. J. et al. Protective effect and molecular mechanisms of human non-neutralizing cross-reactive spike antibodies elicited by SARS-CoV-2 mRNA vaccination. Cell Rep. 43, 114922 (2024).
Assessment of risk associated with recent influenza A(H5N1) clade 2.3.4.4b viruses https://cdn.who.int/media/docs/default-source/influenza/avian-and-other-zoonotic-influenza/h5-risk-assessment-dec-2022.pdf (2022).
CDC Confirms First Severe Case of H5N1 Bird Flu in the United States, https://www.cdc.gov/media/releases/2024/m1218-h5n1-flu.html (2024).
Jassem, A. N. et al. Critical Illness in an Adolescent with Influenza A(H5N1) Virus Infection. N. Engl. J. Med. https://doi.org/10.1056/NEJMc2415890 (2024).
Castillo, A. et al. The first case of human infection with H5N1 avian Influenza A virus in Chile. J. Travel Med. 30, https://doi.org/10.1093/jtm/taad083 (2023).
Bruno, A. et al. First case of human infection with highly pathogenic H5 avian Influenza A virus in South America: A new zoonotic pandemic threat for 2023? J. Travel Med. 30, https://doi.org/10.1093/jtm/taad032 (2023).
Kilbourne, E. D. Influenza pandemics of the 20th century. Emerg. Infect. Dis. 12, 9–14 (2006).
Andreev, K. et al. Genotypic and phenotypic susceptibility of emerging avian influenza A viruses to neuraminidase and cap-dependent endonuclease inhibitors. Antivir. Res. 229, 105959 (2024).
Gu, C. et al. A human isolate of bovine H5N1 is transmissible and lethal in animal models. Nature 636, 711–718 (2024).
Tan, S. K. et al. A randomized, placebo-controlled trial to evaluate the safety and efficacy of VIR-2482 in healthy adults for prevention of influenza A illness (PENINSULA). Clin. Infect. Dis. 79, 1054–1061 (2024).
Stadlbauer, D. et al. Broadly protective human antibodies that target the active site of influenza virus neuraminidase. Science 366, 499–504 (2019).
Margine, I., Palese, P. & Krammer, F. Expression of functional recombinant hemagglutinin and neuraminidase proteins from the novel H7N9 influenza virus using the baculovirus expression system. J. Vis. Exp. e51112 https://doi.org/10.3791/51112 (2013).
Amanat, F., Meade, P., Strohmeier, S. & Krammer, F. Cross-reactive antibodies binding to H4 hemagglutinin protect against a lethal H4N6 influenza virus challenge in the mouse model. Emerg. Microbes Infect. 8, 155–168 (2019).
Duty, J. A. et al. Discovery and intranasal administration of a SARS-CoV-2 broadly acting neutralizing antibody with activity against multiple Omicron subvariants. Med. 3, 705–721.e11 (2022).
Dreyfus, C. et al. Highly conserved protective epitopes on influenza B viruses. Science 337, 1343–1348 (2012).
Nachbagauer, R. et al. Defining the antibody cross-reactome directed against the influenza virus surface glycoproteins. Nat. Immunol. 18, 464–473 (2017).
Acknowledgements
Work in the Krammer laboratory was funded by institutional funding, NIAID Centers of Excellence for Influenza Research and Response (CEIRR, 75N93021C00014, reagent generation, F.K.) and initial isolation of New York City H5N1 viruses was funded by Flu Lab. Work in the Ward laboratory was funded by NIAID Collaborative Influenza Vaccine Innovation Centers (CIVIC 75N39019C00051, A.B.W.).
Author information
Authors and Affiliations
Contributions
Study design: G.P.A., J.A.D. and F.K. Acquisition of data: G.P.A., A.N.L., T.Y., D.B., M.L., K.B., P.J.M.B., A.J.R., T.J., R.W., C.M., J.H. and J.A.D. Analysis and interpretation of data: G.P.A., A.N.L., T.Y.; D.B., M.L., K.B., P.J.M.B., A.J.R., T.J., R.W., C.M., J.H., A.B.W., J.A.D. and F.K. Drafting of the manuscript: G.P.A., J.A.D. and F.K. Study supervision: J.A.D. and F.K. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Icahn School of Medicine at Mount Sinai has filed patent applications regarding the described H5 mAbs which list G.P.A., J.A.D. and F.K. as inventors. The Icahn School of Medicine at Mount Sinai has filed patent applications relating to SARS-CoV-2 serological assays, NDV-based SARS-CoV-2 vaccines influenza virus vaccines and influenza virus therapeutics which list F.K. as co-inventor and F.K. has received royalty payments from some of these patents. Mount Sinai has spun out a company, Castlevax, to develop SARS-CoV-2 vaccines. F.K. is co-founder and scientific advisory board member of Castlevax. F.K. has consulted for Merck, GSK, Sanofi, Curevac, Gritstone, Seqirus and Pfizer and is currently consulting for 3rd Rock Ventures and Avimex. The Krammer laboratory is also collaborating with Dynavax on influenza vaccine development. ABW has received royalty payments for the licensure of a prefusion coronavirus spike stabilization technology for which he is a co-inventor. A.B.W. is currently consulting for Third Rock Ventures and Merida Biosciences. J.H. and A.B.W. are consultants for Third Rock Ventures. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Alzua, G.P., León, A.N., Yellin, T. et al. Human monoclonal antibodies that target clade 2.3.4.4b H5N1 hemagglutinin. Nat Commun 17, 135 (2026). https://doi.org/10.1038/s41467-025-66829-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66829-y